\n", - "
\n", - "However, **there are ways** to modify the mutable contents of the tuple without raising the `TypeError`, the solution is the `.extend()` method, or alternatively `.append()` (for lists):" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "tup = ([],)\n", - "print('tup before: ', tup)\n", - "tup[0].extend([1])\n", - "print('tup after: ', tup)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "tup before: ([],)\n", - "tup after: ([1],)\n" - ] - } - ], - "prompt_number": 44 - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "tup = ([],)\n", - "print('tup before: ', tup)\n", - "tup[0].append(1)\n", - "print('tup after: ', tup)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "tup before: ([],)\n", - "tup after: ([1],)\n" - ] - } - ], - "prompt_number": 5 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "### Explanation\n", - "\n", - "**A. Jesse Jiryu Davis** has a nice explanation for this phenomenon (Original source: [http://emptysqua.re/blog/python-increment-is-weird-part-ii/](http://emptysqua.re/blog/python-increment-is-weird-part-ii/))\n", - "\n", - "If we try to extend the list via `+=` *\"then the statement executes `STORE_SUBSCR`, which calls the C function `PyObject_SetItem`, which checks if the object supports item assignment. In our case the object is a tuple, so `PyObject_SetItem` throws the `TypeError`. Mystery solved.\"*" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "#### One more note about the `immutable` status of tuples. Tuples are famous for being immutable. However, how comes that this code works?" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "my_tup = (1,)\n", - "my_tup += (4,)\n", - "my_tup = my_tup + (5,)\n", - "print(my_tup)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "(1, 4, 5)\n" - ] - } - ], - "prompt_number": 6 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "What happens \"behind\" the curtains is that the tuple is not modified, but every time a new object is generated, which will inherit the old \"name tag\":" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "my_tup = (1,)\n", - "print(id(my_tup))\n", - "my_tup += (4,)\n", - "print(id(my_tup))\n", - "my_tup = my_tup + (5,)\n", - "print(id(my_tup))" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "4337381840\n", - "4357415496\n", - "4357289952\n" - ] - } - ], - "prompt_number": 8 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "
\n", - "\n" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "## List comprehensions are fast, but generators are faster!?" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\"List comprehensions are fast, but generators are faster!?\" - No, not really (or significantly, see the benchmarks below). So what's the reason to prefer one over the other?\n", - "- use lists if you want to use the plethora of list methods \n", - "- use generators when you are dealing with huge collections to avoid memory issues" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "import timeit\n", - "\n", - "def plainlist(n=100000):\n", - " my_list = []\n", - " for i in range(n):\n", - " if i % 5 == 0:\n", - " my_list.append(i)\n", - " return my_list\n", - "\n", - "def listcompr(n=100000):\n", - " my_list = [i for i in range(n) if i % 5 == 0]\n", - " return my_list\n", - "\n", - "def generator(n=100000):\n", - " my_gen = (i for i in range(n) if i % 5 == 0)\n", - " return my_gen\n", - "\n", - "def generator_yield(n=100000):\n", - " for i in range(n):\n", - " if i % 5 == 0:\n", - " yield i" - ], - "language": "python", - "metadata": {}, - "outputs": [], - "prompt_number": 11 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "#### To be fair to the list, let us exhaust the generators:" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "def test_plainlist(plain_list):\n", - " for i in plain_list():\n", - " pass\n", - "\n", - "def test_listcompr(listcompr):\n", - " for i in listcompr():\n", - " pass\n", - "\n", - "def test_generator(generator):\n", - " for i in generator():\n", - " pass\n", - "\n", - "def test_generator_yield(generator_yield):\n", - " for i in generator_yield():\n", - " pass\n", - "\n", - "print('plain_list: ', end = '')\n", - "%timeit test_plainlist(plainlist)\n", - "print('\\nlistcompr: ', end = '')\n", - "%timeit test_listcompr(listcompr)\n", - "print('\\ngenerator: ', end = '')\n", - "%timeit test_generator(generator)\n", - "print('\\ngenerator_yield: ', end = '')\n", - "%timeit test_generator_yield(generator_yield)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "plain_list: 10 loops, best of 3: 22.4 ms per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "\n", - "listcompr: 10 loops, best of 3: 20.8 ms per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "\n", - "generator: 10 loops, best of 3: 22 ms per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "\n", - "generator_yield: 10 loops, best of 3: 21.9 ms per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n" - ] - } - ], - "prompt_number": 13 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "
\n", - "" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "## Public vs. private class methods and name mangling\n", - "\n" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Who has not stumbled across this quote \"we are all consenting adults here\" in the Python community, yet? Unlike in other languages like C++ (sorry, there are many more, but that's one I am most familiar with), we can't really protect class methods from being used outside the class (i.e., by the API user). \n", - "All we can do is to indicate methods as private to make clear that they are better not used outside the class, but it is really up to the class user, since \"we are all consenting adults here\"! \n", - "So, when we want to mark a class method as private, we can put a single underscore in front of it. \n", - "If we additionally want to avoid name clashes with other classes that might use the same method names, we can prefix the name with a double-underscore to invoke the name mangling.\n", - "\n", - "This doesn't prevent the class user to access this class member though, but he has to know the trick and also knows that it his own risk...\n", - "\n", - "Let the following example illustrate what I mean:" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "class my_class():\n", - " def public_method(self):\n", - " print('Hello public world!')\n", - " def __private_method(self):\n", - " print('Hello private world!')\n", - " def call_private_method_in_class(self):\n", - " self.__private_method()\n", - " \n", - "my_instance = my_class()\n", - "\n", - "my_instance.public_method()\n", - "my_instance._my_class__private_method()\n", - "my_instance.call_private_method_in_class()" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "Hello public world!\n", - "Hello private world!\n", - "Hello private world!\n" - ] - } - ], - "prompt_number": 11 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "
\n", - "" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "## The consequences of modifying a list when looping through it" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "It can be really dangerous to modify a list when iterating through it - this is a very common pitfall that can cause unintended behavior! \n", - "Look at the following examples, and for a fun exercise: try to figure out what is going on before you skip to the solution!" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "a = [1, 2, 3, 4, 5]\n", - "for i in a:\n", - " if not i % 2:\n", - " a.remove(i)\n", - "print(a)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "[1, 3, 5]\n" - ] - } - ], - "prompt_number": 3 - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "b = [2, 4, 5, 6]\n", - "for i in b:\n", - " if not i % 2:\n", - " b.remove(i)\n", - "print(b)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "[4, 5]\n" - ] - } - ], - "prompt_number": 4 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "
\n", - "**The solution** is that we are iterating through the list index by index, and if we remove one of the items in-between, we inevitably mess around with the indexing, look at the following example, and it will become clear:" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "b = [2, 4, 5, 6]\n", - "for index, item in enumerate(b):\n", - " print(index, item)\n", - " if not item % 2:\n", - " b.remove(item)\n", - "print(b)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "0 2\n", - "1 5\n", - "2 6\n", - "[4, 5]\n" - ] - } - ], - "prompt_number": 7 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "
\n", - "" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "## Dynamic binding and typos in variable names\n" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Be careful, dynamic binding is convenient, but can also quickly become dangerous!" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "print('first list:')\n", - "for i in range(3):\n", - " print(i)\n", - " \n", - "print('\\nsecond list:')\n", - "for j in range(3):\n", - " print(i) # I (intentionally) made typo here!" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "first list:\n", - "0\n", - "1\n", - "2\n", - "\n", - "second list:\n", - "2\n", - "2\n", - "2\n" - ] - } - ], - "prompt_number": 14 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "
\n", - "" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "## List slicing using indexes that are \"out of range\"" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "As we have all encountered it 1 (x10000) time(s) in our live, the infamous `IndexError`:" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "my_list = [1, 2, 3, 4, 5]\n", - "print(my_list[5])" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "ename": "IndexError", - "evalue": "list index out of range", - "output_type": "pyerr", - "traceback": [ - "\u001b[0;31m---------------------------------------------------------------------------\u001b[0m\n\u001b[0;31mIndexError\u001b[0m Traceback (most recent call last)", - "\u001b[0;32m
\n", - "
\n", - "" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "## Reusing global variable names and `UnboundLocalErrors`" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Usually, it is no problem to access global variables in the local scope of a function:" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "def my_func():\n", - " print(var)\n", - "\n", - "var = 'global'\n", - "my_func()" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "global\n" - ] - } - ], - "prompt_number": 37 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "And is also no problem to use the same variable name in the local scope without affecting the local counterpart: " - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "def my_func():\n", - " var = 'locally changed'\n", - "\n", - "var = 'global'\n", - "my_func()\n", - "print(var)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "global\n" - ] - } - ], - "prompt_number": 38 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "But we have to be careful if we use a variable name that occurs in the global scope, and we want to access it in the local function scope if we want to reuse this name:" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "def my_func():\n", - " print(var) # want to access global variable\n", - " var = 'locally changed' # but Python thinks we forgot to define the local variable!\n", - " \n", - "var = 'global'\n", - "my_func()" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "ename": "UnboundLocalError", - "evalue": "local variable 'var' referenced before assignment", - "output_type": "pyerr", - "traceback": [ - "\u001b[0;31m---------------------------------------------------------------------------\u001b[0m\n\u001b[0;31mUnboundLocalError\u001b[0m Traceback (most recent call last)", - "\u001b[0;32m
\n", - "
\n", - "" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "## Creating copies of mutable objects\n", - "\n" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Let's assume a scenario where we want to duplicate sub`list`s of values stored in another list. If we want to create independent sub`list` object, using the arithmetic multiplication operator could lead to rather unexpected (or undesired) results:" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "my_list1 = [[1, 2, 3]] * 2\n", - "\n", - "print('initially ---> ', my_list1)\n", - "\n", - "# modify the 1st element of the 2nd sublist\n", - "my_list1[1][0] = 'a'\n", - "print(\"after my_list1[1][0] = 'a' ---> \", my_list1)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "initially ---> [[1, 2, 3], [1, 2, 3]]\n", - "after my_list1[1][0] = 'a' ---> [['a', 2, 3], ['a', 2, 3]]\n" - ] - } - ], - "prompt_number": 24 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "
\n", - "In this case, we should better create \"new\" objects:" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "my_list2 = [[1, 2, 3] for i in range(2)]\n", - "\n", - "print('initially: ---> ', my_list2)\n", - "\n", - "# modify the 1st element of the 2nd sublist\n", - "my_list2[1][0] = 'a'\n", - "print(\"after my_list2[1][0] = 'a': ---> \", my_list2)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "initially: ---> [[1, 2, 3], [1, 2, 3]]\n", - "after my_list2[1][0] = 'a': ---> [[1, 2, 3], ['a', 2, 3]]\n" - ] - } - ], - "prompt_number": 25 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "And here is the proof:" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "for a,b in zip(my_list1, my_list2):\n", - " print('id my_list1: {}, id my_list2: {}'.format(id(a), id(b)))" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "id my_list1: 4350764680, id my_list2: 4350766472\n", - "id my_list1: 4350764680, id my_list2: 4350766664\n" - ] - } - ], - "prompt_number": 26 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "
\n", - "" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "## Key differences between Python 2 and 3\n", - "\n" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "There are some good articles already that are summarizing the differences between Python 2 and 3, e.g., \n", - "- [https://wiki.python.org/moin/Python2orPython3](https://wiki.python.org/moin/Python2orPython3)\n", - "- [https://docs.python.org/3.0/whatsnew/3.0.html](https://docs.python.org/3.0/whatsnew/3.0.html)\n", - "- [http://python3porting.com/differences.html](http://python3porting.com/differences.html)\n", - "- [https://docs.python.org/3/howto/pyporting.html](https://docs.python.org/3/howto/pyporting.html) \n", - "etc.\n", - "\n", - "But it might be still worthwhile, especially for Python newcomers, to take a look at some of those!\n", - "(Note: the the code was executed in Python 3.4.0 and Python 2.7.5 and copied from interactive shell sessions.)" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "### Unicode...\n", - "####- Python 2: \n", - "We have ASCII `str()` types, separate `unicode()`, but no `byte` type\n", - "####- Python 3: \n", - "Now, we finally have Unicode (utf-8) `str`ings, and 2 byte classes: `byte` and `bytearray`s" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "#############\n", - "# Python 2\n", - "#############\n", - "\n", - ">>> type(unicode('is like a python3 str()'))\n", - "
\n", - "
\n", - "" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "## Function annotations - What are those `->`'s in my Python code?\n" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Have you ever seen any Python code that used colons inside the parantheses of a function definition?" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "def foo1(x: 'insert x here', y: 'insert x^2 here'):\n", - " print('Hello, World')\n", - " return" - ], - "language": "python", - "metadata": {}, - "outputs": [], - "prompt_number": 8 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "And what about the fancy arrow here?" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "def foo2(x, y) -> 'Hi!':\n", - " print('Hello, World')\n", - " return" - ], - "language": "python", - "metadata": {}, - "outputs": [], - "prompt_number": 10 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Q: Is this valid Python syntax? \n", - "A: Yes!\n", - " \n", - " \n", - "Q: So, what happens if I *just call* the function? \n", - "A: Nothing!\n", - " \n", - "Here is the proof!" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "foo1(1,2)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "Hello, World\n" - ] - } - ], - "prompt_number": 9 - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "foo2(1,2) " - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "Hello, World\n" - ] - } - ], - "prompt_number": 11 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "**So, those are function annotations ... ** \n", - "- the colon for the function parameters \n", - "- the arrow for the return value \n", - "\n", - "You probably will never make use of them (or at least very rarely). Usually, we write good function documentations below the function as a docstring - or at least this is how I would do it (okay this case is a little bit extreme, I have to admit):" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "def is_palindrome(a):\n", - " \"\"\"\n", - " Case-and punctuation insensitive check if a string is a palindrom.\n", - " \n", - " Keyword arguments:\n", - " a (str): The string to be checked if it is a palindrome.\n", - " \n", - " Returns `True` if input string is a palindrome, else False.\n", - " \n", - " \"\"\"\n", - " stripped_str = [l for l in my_str.lower() if l.isalpha()]\n", - " return stripped_str == stripped_str[::-1]\n", - " " - ], - "language": "python", - "metadata": {}, - "outputs": [] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "However, function annotations can be useful to indicate that work is still in progress in some cases. But they are optional and I see them very very rarely.\n", - "\n", - "As it is stated in [PEP3107](http://legacy.python.org/dev/peps/pep-3107/#fundamentals-of-function-annotations):\n", - "\n", - "1. Function annotations, both for parameters and return values, are completely optional.\n", - "\n", - "2. Function annotations are nothing more than a way of associating arbitrary Python expressions with various parts of a function at compile-time.\n" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "The nice thing about function annotations is their `__annotations__` attribute, which is dictionary of all the parameters and/or the `return` value you annotated." - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "foo1.__annotations__" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "metadata": {}, - "output_type": "pyout", - "prompt_number": 17, - "text": [ - "{'y': 'insert x^2 here', 'x': 'insert x here'}" - ] - } - ], - "prompt_number": 17 - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "foo2.__annotations__" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "metadata": {}, - "output_type": "pyout", - "prompt_number": 18, - "text": [ - "{'return': 'Hi!'}" - ] - } - ], - "prompt_number": 18 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "**When are they useful?**" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Function annotations can be useful for a couple of things \n", - "- Documentation in general\n", - "- pre-condition testing\n", - "- [type checking](http://legacy.python.org/dev/peps/pep-0362/#annotation-checker)\n", - " \n", - "..." - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "
\n", - "" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "## Abortive statements in `finally` blocks" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Python's `try-except-finally` blocks are very handy for catching and handling errors. The `finally` block is always executed whether an `exception` has been raised or not as illustrated in the following example." - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "def try_finally1():\n", - " try:\n", - " print('in try:')\n", - " print('do some stuff')\n", - " float('abc')\n", - " except ValueError:\n", - " print('an error occurred')\n", - " else:\n", - " print('no error occurred')\n", - " finally:\n", - " print('always execute finally')\n", - " \n", - "try_finally1()" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "in try:\n", - "do some stuff\n", - "an error occurred\n", - "always execute finally\n" - ] - } - ], - "prompt_number": 24 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "
\n", - "But can you also guess what will be printed in the next code cell?" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "def try_finally2():\n", - " try:\n", - " print(\"do some stuff in try block\")\n", - " return \"return from try block\"\n", - " finally:\n", - " print(\"do some stuff in finally block\")\n", - " return \"always execute finally\"\n", - " \n", - "print(try_finally2())" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "do some stuff in try block\n", - "do some stuff in finally block\n", - "always execute finally\n" - ] - } - ], - "prompt_number": 21 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "Here, the abortive `return` statement in the `finally` block simply overrules the `return` in the `try` block, since **`finally` is guaranteed to always be executed.** So, be careful using abortive statements in `finally` blocks!" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "
\n", - "" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "#Assigning types to variables as values" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "I am not yet sure in which context this can be useful, but it is a nice fun fact to know that we can assign types as values to variables." - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "a_var = str\n", - "a_var(123)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "metadata": {}, - "output_type": "pyout", - "prompt_number": 1, - "text": [ - "'123'" - ] - } - ], - "prompt_number": 1 - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "from random import choice\n", - "\n", - "a, b, c = float, int, str\n", - "for i in range(5):\n", - " j = choice([a,b,c])(i)\n", - " print(j, type(j))" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "0
\n", - "
\n", - "
\n", - "
\n" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "# Changelog" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "#### 04/27/2014\n", - "- minor fixes of typos\n", - "- single- vs. double-underscore clarification in the private class section." - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [], - "language": "python", - "metadata": {}, - "outputs": [], - "prompt_number": 1 - }, - { - "cell_type": "code", - "collapsed": false, - "input": [], - "language": "python", - "metadata": {}, - "outputs": [] - } - ], - "metadata": {} - } - ] -} \ No newline at end of file diff --git a/.ipynb_checkpoints/palindrome_timeit-checkpoint.ipynb b/.ipynb_checkpoints/palindrome_timeit-checkpoint.ipynb deleted file mode 100644 index ef0e33f..0000000 --- a/.ipynb_checkpoints/palindrome_timeit-checkpoint.ipynb +++ /dev/null @@ -1,194 +0,0 @@ -{ - "metadata": { - "name": "", - "signature": "sha256:6ea19109869c82ee989c8ea0599ec49401e74246a542ad0b7b05f6ef464bda19" - }, - "nbformat": 3, - "nbformat_minor": 0, - "worksheets": [ - { - "cells": [ - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Sebastian Raschka 04/2014\n", - "\n", - "#Timing different Implementations of palindrome functions" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "import re\n", - "import timeit\n", - "import string\n", - "\n", - "# All functions return True if an input string is a palindrome. Else returns False.\n", - "\n", - "\n", - "\n", - "####\n", - "#### case-insensitive ignoring punctuation characters\n", - "####\n", - "\n", - "def palindrome_short(my_str):\n", - " stripped_str = \"\".join(l.lower() for l in my_str if l.isalpha())\n", - " return stripped_str == stripped_str[::-1]\n", - "\n", - "def palindrome_regex(my_str):\n", - " return re.sub('\\W', '', my_str.lower()) == re.sub('\\W', '', my_str[::-1].lower())\n", - "\n", - "def palindrome_stringlib(my_str):\n", - " LOWERS = set(string.ascii_lowercase)\n", - " letters = [c for c in my_str.lower() if c in LOWERS]\n", - " return letters == letters[::-1]\n", - "\n", - "LOWERS = set(string.ascii_lowercase)\n", - "def palindrome_stringlib2(my_str):\n", - " letters = [c for c in my_str.lower() if c in LOWERS]\n", - " return letters == letters[::-1]\n", - "\n", - "def palindrome_isalpha(my_str):\n", - " stripped_str = [l for l in my_str.lower() if l.isalpha()]\n", - " return stripped_str == stripped_str[::-1]\n", - "\n", - "\n", - "\n", - "####\n", - "#### functions considering all characters (case-sensitive)\n", - "####\n", - "\n", - "def palindrome_reverse1(my_str):\n", - " return my_str == my_str[::-1]\n", - "\n", - "def palindrome_reverse2(my_str):\n", - " return my_str == ''.join(reversed(my_str))\n", - "\n", - "def palindrome_recurs(my_str):\n", - " if len(my_str) < 2:\n", - " return True\n", - " if my_str[0] != my_str[-1]:\n", - " return False\n", - " return palindrome(my_str[1:-1])\n", - "\n", - "\n", - "\n" - ], - "language": "python", - "metadata": {}, - "outputs": [], - "prompt_number": 10 - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "test_str = \"Go hang a salami. I'm a lasagna hog.\"\n", - "\n", - "print('case-insensitive functions ignoring punctuation characters')\n", - "%timeit palindrome_short(test_str)\n", - "%timeit palindrome_regex(test_str)\n", - "%timeit palindrome_stringlib(test_str)\n", - "%timeit palindrome_stringlib2(test_str)\n", - "%timeit palindrome_isalpha(test_str)\n", - "\n", - "print('\\n\\nfunctions considering all characters (case-sensitive)')\n", - "%timeit palindrome_reverse1(test_str)\n", - "%timeit palindrome_reverse2(test_str)\n", - "%timeit palindrome_recurs(test_str)\n" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "case-insensitive functions ignoring punctuation characters\n", - "100000 loops, best of 3: 15.3 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "100000 loops, best of 3: 19.9 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "100000 loops, best of 3: 13.5 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "100000 loops, best of 3: 8.58 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "100000 loops, best of 3: 9.42 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "\n", - "\n", - "functions considering all characters (case-sensitive)\n", - "1000000 loops, best of 3: 508 ns per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "100000 loops, best of 3: 3.08 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "1000000 loops, best of 3: 480 ns per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n" - ] - } - ], - "prompt_number": 11 - }, - { - "cell_type": "code", - "collapsed": false, - "input": [], - "language": "python", - "metadata": {}, - "outputs": [] - } - ], - "metadata": {} - } - ] -} \ No newline at end of file diff --git a/.ipynb_checkpoints/python_true_false-checkpoint.ipynb b/.ipynb_checkpoints/python_true_false-checkpoint.ipynb deleted file mode 100644 index b2786a2..0000000 --- a/.ipynb_checkpoints/python_true_false-checkpoint.ipynb +++ /dev/null @@ -1,578 +0,0 @@ -{ - "metadata": { - "name": "" - }, - "nbformat": 3, - "nbformat_minor": 0, - "worksheets": [ - { - "cells": [ - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Sebastian Raschka, 03/2014 \n", - "Code was executed in Python 3.4.0" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "###`True` and `False` in the `datetime` module\n", - "\n", - "Pointed out in a nice article **\"A false midnight\"** at [http://lwn.net/SubscriberLink/590299/bf73fe823974acea/](http://lwn.net/SubscriberLink/590299/bf73fe823974acea/):\n", - "\n", - "*\"it often comes as a big surprise for programmers to find (sometimes by way of a hard-to-reproduce bug) that, \n", - "unlike any other time value, midnight (i.e. datetime.time(0,0,0)) is False. \n", - "A long discussion on the python-ideas mailing list shows that, while surprising, \n", - "that behavior is desirable\u2014at least in some quarters.\"*" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "import datetime\n", - "\n", - "print('\"datetime.time(0,0,0)\" (Midnight) evaluates to', bool(datetime.time(0,0,0)))\n", - "\n", - "print('\"datetime.time(1,0,0)\" (1 am) evaluates to', bool(datetime.time(1,0,0)))" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\"datetime.time(0,0,0)\" (Midnight) evaluates to False\n", - "\"datetime.time(1,0,0)\" (1 am) evaluates to True\n" - ] - } - ], - "prompt_number": 17 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "## Boolean `True`" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "my_true_val = True\n", - "\n", - "\n", - "print('my_true_val == True:', my_true_val == True)\n", - "print('my_true_val is True:', my_true_val is True)\n", - "\n", - "print('my_true_val == None:', my_true_val == None)\n", - "print('my_true_val is None:', my_true_val is None)\n", - "\n", - "print('my_true_val == False:', my_true_val == False)\n", - "print('my_true_val is False:', my_true_val is False)\n", - "\n", - "print(my_true_val\n", - "if my_true_val:\n", - " print('\"if my_true_val:\" is True')\n", - "else:\n", - " print('\"if my_true_val:\" is False')\n", - " \n", - "if not my_true_val:\n", - " print('\"if not my_true_val:\" is True')\n", - "else:\n", - " print('\"if not my_true_val:\" is False')" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "my_true_val == True: True\n", - "my_true_val is True: True\n", - "my_true_val == None: False\n", - "my_true_val is None: False\n", - "my_true_val == False: False\n", - "my_true_val is False: False\n", - "\"if my_true_val:\" is True\n", - "\"if not my_true_val:\" is False\n" - ] - } - ], - "prompt_number": 83 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "## Boolean `False`" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "my_false_val = False\n", - "\n", - "\n", - "print('my_false_val == True:', my_false_val == True)\n", - "print('my_false_val is True:', my_false_val is True)\n", - "\n", - "print('my_false_val == None:', my_false_val == None)\n", - "print('my_false_val is None:', my_false_val is None)\n", - "\n", - "print('my_false_val == False:', my_false_val == False)\n", - "print('my_false_val is False:', my_false_val is False)\n", - "\n", - "\n", - "if my_false_val:\n", - " print('\"if my_false_val:\" is True')\n", - "else:\n", - " print('\"if my_false_val:\" is False')\n", - " \n", - "if not my_false_val:\n", - " print('\"if not my_false_val:\" is True')\n", - "else:\n", - " print('\"if not my_false_val:\" is False')" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "my_false_val == True: False\n", - "my_false_val is True: False\n", - "my_false_val == None: False\n", - "my_false_val is None: False\n", - "my_false_val == False: True\n", - "my_false_val is False: True\n", - "\"if my_false_val:\" is False\n", - "\"if not my_false_val:\" is True\n" - ] - } - ], - "prompt_number": 76 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "## `None` 'value'" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "my_none_var = None\n", - "\n", - "print('my_none_var == True:', my_none_var == True)\n", - "print('my_none_var is True:', my_none_var is True)\n", - "\n", - "print('my_none_var == None:', my_none_var == None)\n", - "print('my_none_var is None:', my_none_var is None)\n", - "\n", - "print('my_none_var == False:', my_none_var == False)\n", - "print('my_none_var is False:', my_none_var is False)\n", - "\n", - "\n", - "if my_none_var:\n", - " print('\"if my_none_var:\" is True')\n", - "else:\n", - " print('\"if my_none_var:\" is False')\n", - "\n", - "if not my_none_var:\n", - " print('\"if not my_none_var:\" is True')\n", - "else:\n", - " print('\"if not my_none_var:\" is False')" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "my_none_var == True: False\n", - "my_none_var is True: False\n", - "my_none_var == None: True\n", - "my_none_var is None: True\n", - "my_none_var == False: False\n", - "my_none_var is False: False\n", - "\"if my_none_var:\" is False\n", - "\"if not my_none_var:\" is True\n" - ] - } - ], - "prompt_number": 62 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "## Empty String" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "my_empty_string = \"\"\n", - "\n", - "print('my_empty_string == True:', my_empty_string == True)\n", - "print('my_empty_string is True:', my_empty_string is True)\n", - "\n", - "print('my_empty_string == None:', my_empty_string == None)\n", - "print('my_empty_string is None:', my_empty_string is None)\n", - "\n", - "print('my_empty_string == False:', my_empty_string == False)\n", - "print('my_empty_string is False:', my_empty_string is False)\n", - "\n", - "\n", - "if my_empty_string:\n", - " print('\"if my_empty_string:\" is True')\n", - "else:\n", - " print('\"if my_empty_string:\" is False')\n", - " \n", - "if not my_empty_string:\n", - " print('\"if not my_empty_string:\" is True')\n", - "else:\n", - " print('\"if not my_empty_string:\" is False')" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "my_empty_string == True: False\n", - "my_empty_string is True: False\n", - "my_empty_string == None: False\n", - "my_empty_string is None: False\n", - "my_empty_string == False: False\n", - "my_empty_string is False: False\n", - "\"if my_empty_string:\" is False\n", - "\"if my_empty_string:\" is True\n" - ] - } - ], - "prompt_number": 61 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "## Empty List\n", - "It is generally not a good idea to use the `==` to check for empty lists..." - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "my_empty_list = []\n", - "\n", - "\n", - "print('my_empty_list == True:', my_empty_list == True)\n", - "print('my_empty_list is True:', my_empty_list is True)\n", - "\n", - "print('my_empty_list == None:', my_empty_list == None)\n", - "print('my_empty_list is None:', my_empty_list is None)\n", - "\n", - "print('my_empty_list == False:', my_empty_list == False)\n", - "print('my_empty_list is False:', my_empty_list is False)\n", - "\n", - "\n", - "if my_empty_list:\n", - " print('\"if my_empty_list:\" is True')\n", - "else:\n", - " print('\"if my_empty_list:\" is False')\n", - " \n", - "if not my_empty_list:\n", - " print('\"if not my_empty_list:\" is True')\n", - "else:\n", - " print('\"if not my_empty_list:\" is False')\n", - "\n", - "\n", - " \n" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "my_empty_list == True: False\n", - "my_empty_list is True: False\n", - "my_empty_list == None: False\n", - "my_empty_list is None: False\n", - "my_empty_list == False: False\n", - "my_empty_list is False: False\n", - "\"if my_empty_list:\" is False\n", - "\"if not my_empty_list:\" is True\n" - ] - } - ], - "prompt_number": 67 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "## [0]-List" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "my_zero_list = [0]\n", - "\n", - "\n", - "print('my_zero_list == True:', my_zero_list == True)\n", - "print('my_zero_list is True:', my_zero_list is True)\n", - "\n", - "print('my_zero_list == None:', my_zero_list == None)\n", - "print('my_zero_list is None:', my_zero_list is None)\n", - "\n", - "print('my_zero_list == False:', my_zero_list == False)\n", - "print('my_zero_list is False:', my_zero_list is False)\n", - "\n", - "\n", - "if my_zero_list:\n", - " print('\"if my_zero_list:\" is True')\n", - "else:\n", - " print('\"if my_zero_list:\" is False')\n", - " \n", - "if not my_zero_list:\n", - " print('\"if not my_zero_list:\" is True')\n", - "else:\n", - " print('\"if not my_zero_list:\" is False')" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "my_zero_list == True: False\n", - "my_zero_list is True: False\n", - "my_zero_list == None: False\n", - "my_zero_list is None: False\n", - "my_zero_list == False: False\n", - "my_zero_list is False: False\n", - "\"if my_zero_list:\" is True\n", - "\"if not my_zero_list:\" is False\n" - ] - } - ], - "prompt_number": 70 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "## List comparison \n", - "List comparisons are a handy way to show the difference between `==` and `is`. \n", - "While `==` is rather evaluating the equality of the value, `is` is checking if two objects are equal.\n", - "The examples below show that we can assign a pointer to the same list object by using `=`, e.g., `list1 = list2`. \n", - "a) If we want to make a **shallow** copy of the list values, we have to make a little tweak: `list1 = list2[:]`, or \n", - "b) a **deepcopy** via `list1 = copy.deepcopy(list2)`\n", - "\n", - "Possibly the best explanation of shallow vs. deep copies I've read so far:\n", - "\n", - "*** \"Shallow copies duplicate as little as possible. A shallow copy of a collection is a copy of the collection structure, not the elements. With a shallow copy, two collections now share the individual elements.\n", - "Deep copies duplicate everything. A deep copy of a collection is two collections with all of the elements in the original collection duplicated.\"***\n", - "\n", - "(via [S.Lott](http://stackoverflow.com/users/10661/s-lott) on [StackOverflow](http://stackoverflow.com/questions/184710/what-is-the-difference-between-a-deep-copy-and-a-shallow-copy))" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "###a) Shallow vs. deep copies for simple elements \n", - "List modification of the original list doesn't affect \n", - "shallow copies or deep copies if the list contains literals." - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "from copy import deepcopy\n", - "\n", - "my_first_list = [1]\n", - "my_second_list = [1]\n", - "print('my_first_list == my_second_list:', my_first_list == my_second_list)\n", - "print('my_first_list is my_second_list:', my_first_list is my_second_list)\n", - "\n", - "my_third_list = my_first_list\n", - "print('my_first_list == my_third_list:', my_first_list == my_third_list)\n", - "print('my_first_list is my_third_list:', my_first_list is my_third_list)\n", - "\n", - "my_shallow_copy = my_first_list[:]\n", - "print('my_first_list == my_shallow_copy:', my_first_list == my_shallow_copy)\n", - "print('my_first_list is my_shallow_copy:', my_first_list is my_shallow_copy)\n", - "\n", - "my_deep_copy = deepcopy(my_first_list)\n", - "print('my_first_list == my_deep_copy:', my_first_list == my_deep_copy)\n", - "print('my_first_list is my_deep_copy:', my_first_list is my_deep_copy)\n", - "\n", - "print('\\nmy_third_list:', my_third_list)\n", - "print('my_shallow_copy:', my_shallow_copy)\n", - "print('my_deep_copy:', my_deep_copy)\n", - "\n", - "my_first_list[0] = 2\n", - "print('after setting \"my_first_list[0] = 2\"')\n", - "print('my_third_list:', my_third_list)\n", - "print('my_shallow_copy:', my_shallow_copy)\n", - "print('my_deep_copy:', my_deep_copy)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "my_first_list == my_second_list: True\n", - "my_first_list is my_second_list: False\n", - "my_first_list == my_third_list: True\n", - "my_first_list is my_third_list: True\n", - "my_first_list == my_shallow_copy: True\n", - "my_first_list is my_shallow_copy: False\n", - "my_first_list == my_deep_copy: True\n", - "my_first_list is my_deep_copy: False\n", - "\n", - "my_third_list: [1]\n", - "my_shallow_copy: [1]\n", - "my_deep_copy: [1]\n", - "after setting \"my_first_list[0] = 2\"\n", - "my_third_list: [2]\n", - "my_shallow_copy: [1]\n", - "my_deep_copy: [1]\n" - ] - } - ], - "prompt_number": 11 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "### b) Shallow vs. deep copies if list contains other structures and objects\n", - "List modification of the original list does affect \n", - "shallow copies, but not deep copies if the list contains compound objects." - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "my_first_list = [[1],[2]]\n", - "my_second_list = [[1],[2]]\n", - "print('my_first_list == my_second_list:', my_first_list == my_second_list)\n", - "print('my_first_list is my_second_list:', my_first_list is my_second_list)\n", - "\n", - "my_third_list = my_first_list\n", - "print('my_first_list == my_third_list:', my_first_list == my_third_list)\n", - "print('my_first_list is my_third_list:', my_first_list is my_third_list)\n", - "\n", - "my_shallow_copy = my_first_list[:]\n", - "print('my_first_list == my_shallow_copy:', my_first_list == my_shallow_copy)\n", - "print('my_first_list is my_shallow_copy:', my_first_list is my_shallow_copy)\n", - "\n", - "my_deep_copy = deepcopy(my_first_list)\n", - "print('my_first_list == my_deep_copy:', my_first_list == my_deep_copy)\n", - "print('my_first_list is my_deep_copy:', my_first_list is my_deep_copy)\n", - "\n", - "print('\\nmy_third_list:', my_third_list)\n", - "print('my_shallow_copy:', my_shallow_copy)\n", - "print('my_deep_copy:', my_deep_copy)\n", - "\n", - "my_first_list[0][0] = 2\n", - "print('after setting \"my_first_list[0][0] = 2\"')\n", - "print('my_third_list:', my_third_list)\n", - "print('my_shallow_copy:', my_shallow_copy)\n", - "print('my_deep_copy:', my_deep_copy)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "my_first_list == my_second_list: True\n", - "my_first_list is my_second_list: False\n", - "my_first_list == my_third_list: True\n", - "my_first_list is my_third_list: True\n", - "my_first_list == my_shallow_copy: True\n", - "my_first_list is my_shallow_copy: False\n", - "my_first_list == my_deep_copy: True\n", - "my_first_list is my_deep_copy: False\n", - "\n", - "my_third_list: [[1], [2]]\n", - "my_shallow_copy: [[1], [2]]\n", - "my_deep_copy: [[1], [2]]\n", - "after setting \"my_first_list[0][0] = 2\"\n", - "my_third_list: [[2], [2]]\n", - "my_shallow_copy: [[2], [2]]\n", - "my_deep_copy: [[1], [2]]\n" - ] - } - ], - "prompt_number": 13 - }, - { - "cell_type": "heading", - "level": 2, - "metadata": {}, - "source": [ - "Some Python oddity:" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "a = 1\n", - "b = 1\n", - "print('a is b', bool(a is b))\n", - "True\n", - "\n", - "a = 999\n", - "b = 999\n", - "print('a is b', bool(a is b))\n" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "a is b True\n", - "a is b False\n" - ] - } - ], - "prompt_number": 1 - } - ], - "metadata": {} - } - ] -} \ No newline at end of file diff --git a/.ipynb_checkpoints/timeit_test-checkpoint.ipynb b/.ipynb_checkpoints/timeit_test-checkpoint.ipynb deleted file mode 100644 index dd46362..0000000 --- a/.ipynb_checkpoints/timeit_test-checkpoint.ipynb +++ /dev/null @@ -1,628 +0,0 @@ -{ - "metadata": { - "name": "", - "signature": "sha256:5a2264b30b9632e14bd425a887a4455658fbdf9f8102fc5703ad982c3fa09b21" - }, - "nbformat": 3, - "nbformat_minor": 0, - "worksheets": [ - { - "cells": [ - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Sebastian Raschka \n", - "last updated: 04/14/2014 \n", - "\n", - "[Link to this IPython Notebook on GitHub](https://github.com/rasbt/python_reference/blob/master/timeit_test.ipynb)" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "\n", - "# Python benchmarks via `timeit`" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "# Sections\n", - "- [String operations](#string_operations)\n", - " - [String formatting: .format() vs. binary operator %s](#str_format_bin)\n", - " - [String reversing: [::-1] vs. `''.join(reversed())`](#str_reverse)\n", - " - [String concatenation: `+=` vs. `''.join()`](#string_concat)\n", - " - [Assembling strings](#string_assembly) \n", - "- [List operations](#list_operations)\n", - " - [List reversing: [::-1] vs. reverse() vs. reversed()](#list_reverse)\n", - " - [Creating lists using conditional statements](#create_cond_list)\n", - "- [Dictionary operations](#dict_ops) \n", - " - [Adding elements to a dictionary](#adding_dict_elements)" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "\n", - "# String operations" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "## String formatting: `.format()` vs. binary operator `%s`\n", - "\n", - "We expect the string .format() method to perform slower than %, because it is doing the formatting for each object itself, where formatting via the binary % is hard-coded for known types. But let's see how big the difference really is..." - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "import timeit\n", - "\n", - "def test_format():\n", - " return ['{}'.format(i) for i in range(1000000)]\n", - "\n", - "def test_binaryop():\n", - " return ['%s' %i for i in range(1000000)]\n", - "\n", - "%timeit test_format()\n", - "%timeit test_binaryop()\n", - "\n", - "#\n", - "# Python 3.4.0\n", - "# MacOS X 10.9.2\n", - "# 2.5 GHz Intel Core i5\n", - "# 4 GB 1600 Mhz DDR3\n", - "#" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "1 loops, best of 3: 400 ms per loop\n", - "1 loops, best of 3: 241 ms per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n" - ] - } - ], - "prompt_number": 3 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "## String reversing: `[::-1]` vs. `''.join(reversed())`" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "import timeit\n", - "\n", - "def reverse_join(my_str):\n", - " return ''.join(reversed(my_str))\n", - " \n", - "def reverse_slizing(my_str):\n", - " return my_str[::-1]\n", - "\n", - "\n", - "# Test to show that both work\n", - "a = reverse_join('abcd')\n", - "b = reverse_slizing('abcd')\n", - "assert(a == b and a == 'dcba')\n", - "\n", - "%timeit reverse_join('abcd')\n", - "%timeit reverse_slizing('abcd')\n", - "\n", - "# Python 3.4.0\n", - "# MacOS X 10.9.2\n", - "# 2.4 GHz Intel Core Duo\n", - "# 8 GB 1067 Mhz DDR3\n", - "#" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "1000000 loops, best of 3: 1.28 \u00b5s per loop\n", - "1000000 loops, best of 3: 337 ns per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n" - ] - } - ], - "prompt_number": 13 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "## String concatenation: `+=` vs. `''.join()`" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Strings in Python are immutable objects. So, each time we append a character to a string, it has to be created \u201cfrom scratch\u201d in memory. Thus, the answer to the question \u201cWhat is the most efficient way to concatenate strings?\u201d is a quite obvious, but the relative numbers of performance gains are nonetheless interesting." - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "import timeit\n", - "\n", - "def string_add(in_chars):\n", - " new_str = ''\n", - " for char in in_chars:\n", - " new_str += char\n", - " return new_str\n", - "\n", - "def string_join(in_chars):\n", - " return ''.join(in_chars)\n", - "\n", - "test_chars = ['a', 'b', 'c', 'd', 'e', 'f']\n", - "\n", - "%timeit string_add(test_chars)\n", - "%timeit string_join(test_chars)\n", - "\n", - "#\n", - "# Python 3.4.0\n", - "# MacOS X 10.9.2\n", - "# 2.5 GHz Intel Core i5\n", - "# 4 GB 1600 Mhz DDR3\n", - "#" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "1000000 loops, best of 3: 595 ns per loop\n", - "1000000 loops, best of 3: 269 ns per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n" - ] - } - ], - "prompt_number": 16 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "## Assembling strings\n", - "\n", - "Next, I wanted to compare different methods string \u201cassembly.\u201d This is different from simple string concatenation, which we have seen in the previous section, since we insert values into a string, e.g., from a variable." - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "import timeit\n", - "\n", - "def plus_operator():\n", - " return 'a' + str(1) + str(2) \n", - " \n", - "def format_method():\n", - " return 'a{}{}'.format(1,2)\n", - " \n", - "def binary_operator():\n", - " return 'a%s%s' %(1,2)\n", - "\n", - "%timeit plus_operator()\n", - "%timeit format_method()\n", - "%timeit binary_operator()\n", - "\n", - "#\n", - "# Python 3.4.0\n", - "# MacOS X 10.9.2\n", - "# 2.5 GHz Intel Core i5\n", - "# 4 GB 1600 Mhz DDR3\n", - "#" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "1000000 loops, best of 3: 764 ns per loop\n", - "1000000 loops, best of 3: 494 ns per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "10000000 loops, best of 3: 79.3 ns per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n" - ] - } - ], - "prompt_number": 17 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "# List operations" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "## List reversing - `[::-1]` vs. `reverse()` vs. `reversed()`" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "import timeit\n", - "\n", - "def reverse_func(my_list):\n", - " new_list = my_list[:]\n", - " new_list.reverse()\n", - " return new_list\n", - " \n", - "def reversed_func(my_list):\n", - " return list(reversed(my_list))\n", - "\n", - "def reverse_slizing(my_list):\n", - " return my_list[::-1]\n", - "\n", - "%timeit reverse_func([1,2,3,4,5])\n", - "%timeit reversed_func([1,2,3,4,5])\n", - "%timeit reverse_slizing([1,2,3,4,5])\n", - "\n", - "# Python 3.4.0\n", - "# MacOS X 10.9.2\n", - "# 2.4 GHz Intel Core Duo\n", - "# 8 GB 1067 Mhz DDR3\n", - "#" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "1000000 loops, best of 3: 930 ns per loop\n", - "1000000 loops, best of 3: 1.89 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "1000000 loops, best of 3: 775 ns per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n" - ] - } - ], - "prompt_number": 1 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "## Creating lists using conditional statements\n", - "\n", - "In this test, I attempted to figure out the fastest way to create a new list of elements that meet a certain criterion. For the sake of simplicity, the criterion was to check if an element is even or odd, and only if the element was even, it should be included in the list. For example, the resulting list for numbers in the range from 1 to 10 would be \n", - "[2, 4, 6, 8, 10].\n", - "\n", - "Here, I tested three different approaches: \n", - "1) a simple for loop with an if-statement check (`cond_loop()`) \n", - "2) a list comprehension (`list_compr()`) \n", - "3) the built-in filter() function (`filter_func()`) \n", - "\n", - "Note that the filter() function now returns a generator in Python 3, so I had to wrap it in an additional list() function call." - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "import timeit\n", - "\n", - "def cond_loop():\n", - " even_nums = []\n", - " for i in range(100):\n", - " if i % 2 == 0:\n", - " even_nums.append(i)\n", - " return even_nums\n", - "\n", - "def list_compr():\n", - " even_nums = [i for i in range(100) if i % 2 == 0]\n", - " return even_nums\n", - " \n", - "def filter_func():\n", - " even_nums = list(filter((lambda x: x % 2 != 0), range(100)))\n", - " return even_nums\n", - "\n", - "%timeit cond_loop()\n", - "%timeit list_compr()\n", - "%timeit filter_func()\n", - "\n", - "#\n", - "# Python 3.4.0\n", - "# MacOS X 10.9.2\n", - "# 2.5 GHz Intel Core i5\n", - "# 4 GB 1600 Mhz DDR3\n", - "#" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "100000 loops, best of 3: 14.4 \u00b5s per loop\n", - "100000 loops, best of 3: 12 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "10000 loops, best of 3: 23.9 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n" - ] - } - ], - "prompt_number": 14 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "# Dictionary operations " - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "## Adding elements to a Dictionary\n", - "\n", - "All three functions below count how often different elements (values) occur in a list. \n", - "E.g., for the list ['a', 'b', 'a', 'c'], the dictionary would look like this: \n", - "`my_dict = {'a': 2, 'b': 1, 'c': 1}`" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "import random\n", - "import copy\n", - "import timeit\n", - "\n", - "\n", - "\n", - "def add_element_check1(my_dict, elements):\n", - " for e in elements:\n", - " if e not in my_dict:\n", - " my_dict[e] = 1\n", - " else:\n", - " my_dict[e] += 1\n", - " \n", - "def add_element_check2(my_dict, elements):\n", - " for e in elements:\n", - " if e not in my_dict:\n", - " my_dict[e] = 0\n", - " my_dict[e] += 1 \n", - "\n", - "def add_element_except(my_dict, elements):\n", - " for e in elements:\n", - " try:\n", - " my_dict[e] += 1\n", - " except KeyError:\n", - " my_dict[e] = 1\n", - " \n", - "\n", - "random.seed(123)\n", - "rand_ints = [random.randrange(1, 10) for i in range(100)]\n", - "empty_dict = {}\n", - "\n", - "print('Results for 100 integers in range 1-10') \n", - "%timeit add_element_check1(copy.deepcopy(empty_dict), rand_ints)\n", - "%timeit add_element_check2(copy.deepcopy(empty_dict), rand_ints)\n", - "%timeit add_element_except(copy.deepcopy(empty_dict), rand_ints)\n", - " \n", - "print('\\nResults for 1000 integers in range 1-10') \n", - "rand_ints = [random.randrange(1, 10) for i in range(1000)]\n", - "empty_dict = {}\n", - "\n", - "%timeit add_element_check1(copy.deepcopy(empty_dict), rand_ints)\n", - "%timeit add_element_check2(copy.deepcopy(empty_dict), rand_ints)\n", - "%timeit add_element_except(copy.deepcopy(empty_dict), rand_ints)\n", - "\n", - "print('\\nResults for 1000 integers in range 1-1000') \n", - "rand_ints = [random.randrange(1, 10) for i in range(1000)]\n", - "empty_dict = {}\n", - "\n", - "%timeit add_element_check1(copy.deepcopy(empty_dict), rand_ints)\n", - "%timeit add_element_check2(copy.deepcopy(empty_dict), rand_ints)\n", - "%timeit add_element_except(copy.deepcopy(empty_dict), rand_ints)\n", - "\n", - "#\n", - "# Python 3.4.0\n", - "# MacOS X 10.9.2\n", - "# 2.5 GHz Intel Core i5\n", - "# 4 GB 1600 Mhz DDR3\n", - "#" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "Results for 100 integers in range 1-10\n", - "100000 loops, best of 3: 16.6 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "100000 loops, best of 3: 17.6 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "100000 loops, best of 3: 17.9 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "\n", - "Results for 1000 integers in range 1-10\n", - "10000 loops, best of 3: 135 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "10000 loops, best of 3: 125 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "10000 loops, best of 3: 105 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "\n", - "Results for 1000 integers in range 1-1000\n", - "10000 loops, best of 3: 122 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "10000 loops, best of 3: 123 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "10000 loops, best of 3: 104 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n" - ] - } - ], - "prompt_number": 13 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "### Conclusion\n", - "Interestingly, the `try-except` loop pays off if we have more elements (here: 1000 integers instead of 100) as dictionary keys to check. Also, it doesn't matter much whether the elements exist or do not exist in the dictionary, yet." - ] - } - ], - "metadata": {} - } - ] -} \ No newline at end of file diff --git a/.ipynb_checkpoints/timeit_tests-checkpoint.ipynb b/.ipynb_checkpoints/timeit_tests-checkpoint.ipynb deleted file mode 100644 index baf4bed..0000000 --- a/.ipynb_checkpoints/timeit_tests-checkpoint.ipynb +++ /dev/null @@ -1,1706 +0,0 @@ -{ - "metadata": { - "name": "", - "signature": "sha256:ab74fdc9e8ae58388d1fc428599abc86a3d03938d0f51769cef38ab4028d516a" - }, - "nbformat": 3, - "nbformat_minor": 0, - "worksheets": [ - { - "cells": [ - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Sebastian Raschka \n", - "last updated: 04/14/2014 \n", - "\n", - "[Link to this IPython Notebook on GitHub](https://github.com/rasbt/python_reference/blob/master/benchmarks/timeit_tests.ipynb)" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "I am really looking forward to your comments and suggestions to improve and extend this collection! Just send me a quick note \n", - "via Twitter: [@rasbt](https://twitter.com/rasbt) \n", - "or Email: [bluewoodtree@gmail.com](mailto:bluewoodtree@gmail.com)\n", - "
" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "\n", - "# Python benchmarks via `timeit`" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "
\n", - "
" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "# Sections\n", - "- [String operations](#string_operations)\n", - " - [String formatting: .format() vs. binary operator %s](#str_format_bin)\n", - " - [String reversing: [::-1] vs. `''.join(reversed())`](#str_reverse)\n", - " - [String concatenation: `+=` vs. `''.join()`](#string_concat)\n", - " - [Assembling strings](#string_assembly) \n", - " - [Testing if a string is an integer](#is_integer)\n", - " - [Testing if a string is a number](#is_number)\n", - "- [List operations](#list_operations)\n", - " - [List reversing: [::-1] vs. reverse() vs. reversed()](#list_reverse)\n", - " - [Creating lists using conditional statements](#create_cond_list)\n", - "- [Dictionary operations](#dict_ops) \n", - " - [Adding elements to a dictionary](#adding_dict_elements)\n", - "- [Comprehensions vs. for-loops](#comprehensions)\n", - "- [Copying files by searching directory trees](#find_copy)\n", - "- [Returning column vectors slicing through a numpy array](#row_vectors)" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "# String operations" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "
\n", - "
" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "## String formatting: `.format()` vs. binary operator `%s`\n" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "We expect the string .format() method to perform slower than %, because it is doing the formatting for each object itself, where formatting via the binary % is hard-coded for known types. But let's see how big the difference really is..." - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "import timeit\n", - "\n", - "def test_format():\n", - " return ['{}'.format(i) for i in range(1000000)]\n", - "\n", - "def test_binaryop():\n", - " return ['%s' %i for i in range(1000000)]\n", - "\n", - "%timeit test_format()\n", - "%timeit test_binaryop()\n", - "\n", - "#\n", - "# Python 3.4.0\n", - "# MacOS X 10.9.2\n", - "# 2.5 GHz Intel Core i5\n", - "# 4 GB 1600 Mhz DDR3\n", - "#" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "1 loops, best of 3: 400 ms per loop\n", - "1 loops, best of 3: 241 ms per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n" - ] - } - ], - "prompt_number": 3 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "
\n", - "
" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "## String reversing: `[::-1]` vs. `''.join(reversed())`" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "import timeit\n", - "\n", - "def reverse_join(my_str):\n", - " return ''.join(reversed(my_str))\n", - " \n", - "def reverse_slizing(my_str):\n", - " return my_str[::-1]\n", - "\n", - "\n", - "# Test to show that both work\n", - "a = reverse_join('abcd')\n", - "b = reverse_slizing('abcd')\n", - "assert(a == b and a == 'dcba')\n", - "\n", - "%timeit reverse_join('abcd')\n", - "%timeit reverse_slizing('abcd')\n", - "\n", - "# Python 3.4.0\n", - "# MacOS X 10.9.2\n", - "# 2.4 GHz Intel Core Duo\n", - "# 8 GB 1067 Mhz DDR3\n", - "#" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "1000000 loops, best of 3: 1.28 \u00b5s per loop\n", - "1000000 loops, best of 3: 337 ns per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n" - ] - } - ], - "prompt_number": 13 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "
\n", - "
" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "## String concatenation: `+=` vs. `''.join()`" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Strings in Python are immutable objects. So, each time we append a character to a string, it has to be created \u201cfrom scratch\u201d in memory. Thus, the answer to the question \u201cWhat is the most efficient way to concatenate strings?\u201d is a quite obvious, but the relative numbers of performance gains are nonetheless interesting." - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "import timeit\n", - "\n", - "def string_add(in_chars):\n", - " new_str = ''\n", - " for char in in_chars:\n", - " new_str += char\n", - " return new_str\n", - "\n", - "def string_join(in_chars):\n", - " return ''.join(in_chars)\n", - "\n", - "test_chars = ['a', 'b', 'c', 'd', 'e', 'f']\n", - "\n", - "%timeit string_add(test_chars)\n", - "%timeit string_join(test_chars)\n", - "\n", - "#\n", - "# Python 3.4.0\n", - "# MacOS X 10.9.2\n", - "# 2.5 GHz Intel Core i5\n", - "# 4 GB 1600 Mhz DDR3\n", - "#" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "1000000 loops, best of 3: 595 ns per loop\n", - "1000000 loops, best of 3: 269 ns per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n" - ] - } - ], - "prompt_number": 16 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "\n", - "
\n", - "
" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "## Assembling strings\n", - "\n" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Next, I wanted to compare different methods string \u201cassembly.\u201d This is different from simple string concatenation, which we have seen in the previous section, since we insert values into a string, e.g., from a variable." - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "import timeit\n", - "\n", - "def plus_operator():\n", - " return 'a' + str(1) + str(2) \n", - " \n", - "def format_method():\n", - " return 'a{}{}'.format(1,2)\n", - " \n", - "def binary_operator():\n", - " return 'a%s%s' %(1,2)\n", - "\n", - "%timeit plus_operator()\n", - "%timeit format_method()\n", - "%timeit binary_operator()\n", - "\n", - "#\n", - "# Python 3.4.0\n", - "# MacOS X 10.9.2\n", - "# 2.5 GHz Intel Core i5\n", - "# 4 GB 1600 Mhz DDR3\n", - "#" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "1000000 loops, best of 3: 764 ns per loop\n", - "1000000 loops, best of 3: 494 ns per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "10000000 loops, best of 3: 79.3 ns per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n" - ] - } - ], - "prompt_number": 17 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "\n", - "
\n", - "
" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "## Testing if a string is an integer" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "import timeit\n", - "\n", - "def string_is_int(a_str):\n", - " try:\n", - " int(a_str)\n", - " return True\n", - " except ValueError:\n", - " return False\n", - "\n", - "an_int = '123'\n", - "no_int = '123abc'\n", - "\n", - "%timeit string_is_int(an_int)\n", - "%timeit string_is_int(no_int)\n", - "%timeit an_int.isdigit()\n", - "%timeit no_int.isdigit()\n", - "\n", - "#\n", - "# Python 3.4.0\n", - "# MacOS X 10.9.2\n", - "# 2.5 GHz Intel Core i5\n", - "# 4 GB 1600 Mhz DDR3\n", - "#" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "1000000 loops, best of 3: 401 ns per loop\n", - "100000 loops, best of 3: 3.04 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "10000000 loops, best of 3: 92.1 ns per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "10000000 loops, best of 3: 96.3 ns per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n" - ] - } - ], - "prompt_number": 5 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "
\n", - "
\n" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "## Testing if a string is a number" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "import timeit\n", - "\n", - "def string_is_number(a_str):\n", - " try:\n", - " float(a_str)\n", - " return True\n", - " except ValueError:\n", - " return False\n", - " \n", - "a_float = '1.234'\n", - "no_float = '123abc'\n", - "\n", - "a_float.replace('.','',1).isdigit()\n", - "no_float.replace('.','',1).isdigit()\n", - "\n", - "%timeit string_is_number(an_int)\n", - "%timeit string_is_number(no_int)\n", - "%timeit a_float.replace('.','',1).isdigit()\n", - "%timeit no_float.replace('.','',1).isdigit()\n", - "\n", - "#\n", - "# Python 3.4.0\n", - "# MacOS X 10.9.2\n", - "# 2.5 GHz Intel Core i5\n", - "# 4 GB 1600 Mhz DDR3\n", - "#" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "1000000 loops, best of 3: 400 ns per loop\n", - "1000000 loops, best of 3: 1.15 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "1000000 loops, best of 3: 452 ns per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "1000000 loops, best of 3: 394 ns per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n" - ] - } - ], - "prompt_number": 6 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "\n", - "
\n", - "
" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "# List operations" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "
\n", - "
" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "## List reversing - `[::-1]` vs. `reverse()` vs. `reversed()`" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "import timeit\n", - "\n", - "def reverse_func(my_list):\n", - " new_list = my_list[:]\n", - " new_list.reverse()\n", - " return new_list\n", - " \n", - "def reversed_func(my_list):\n", - " return list(reversed(my_list))\n", - "\n", - "def reverse_slizing(my_list):\n", - " return my_list[::-1]\n", - "\n", - "%timeit reverse_func([1,2,3,4,5])\n", - "%timeit reversed_func([1,2,3,4,5])\n", - "%timeit reverse_slizing([1,2,3,4,5])\n", - "\n", - "# Python 3.4.0\n", - "# MacOS X 10.9.2\n", - "# 2.4 GHz Intel Core Duo\n", - "# 8 GB 1067 Mhz DDR3\n", - "#" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "1000000 loops, best of 3: 930 ns per loop\n", - "1000000 loops, best of 3: 1.89 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "1000000 loops, best of 3: 775 ns per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n" - ] - } - ], - "prompt_number": 1 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "
\n", - "
" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "## Creating lists using conditional statements\n" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "In this test, I attempted to figure out the fastest way to create a new list of elements that meet a certain criterion. For the sake of simplicity, the criterion was to check if an element is even or odd, and only if the element was even, it should be included in the list. For example, the resulting list for numbers in the range from 1 to 10 would be \n", - "[2, 4, 6, 8, 10].\n", - "\n", - "Here, I tested three different approaches: \n", - "1) a simple for loop with an if-statement check (`cond_loop()`) \n", - "2) a list comprehension (`list_compr()`) \n", - "3) the built-in filter() function (`filter_func()`) \n", - "\n", - "Note that the filter() function now returns a generator in Python 3, so I had to wrap it in an additional list() function call." - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "import timeit\n", - "\n", - "def cond_loop():\n", - " even_nums = []\n", - " for i in range(100):\n", - " if i % 2 == 0:\n", - " even_nums.append(i)\n", - " return even_nums\n", - "\n", - "def list_compr():\n", - " even_nums = [i for i in range(100) if i % 2 == 0]\n", - " return even_nums\n", - " \n", - "def filter_func():\n", - " even_nums = list(filter((lambda x: x % 2 != 0), range(100)))\n", - " return even_nums\n", - "\n", - "%timeit cond_loop()\n", - "%timeit list_compr()\n", - "%timeit filter_func()\n", - "\n", - "#\n", - "# Python 3.4.0\n", - "# MacOS X 10.9.2\n", - "# 2.5 GHz Intel Core i5\n", - "# 4 GB 1600 Mhz DDR3\n", - "#" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "100000 loops, best of 3: 14.4 \u00b5s per loop\n", - "100000 loops, best of 3: 12 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "10000 loops, best of 3: 23.9 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n" - ] - } - ], - "prompt_number": 14 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "
\n", - "
" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "# Dictionary operations " - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "
\n", - "
" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "## Adding elements to a Dictionary\n" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "All three functions below count how often different elements (values) occur in a list. \n", - "E.g., for the list ['a', 'b', 'a', 'c'], the dictionary would look like this: \n", - "`my_dict = {'a': 2, 'b': 1, 'c': 1}`" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "import random\n", - "import copy\n", - "import timeit\n", - "\n", - "\n", - "\n", - "def add_element_check1(my_dict, elements):\n", - " for e in elements:\n", - " if e not in my_dict:\n", - " my_dict[e] = 1\n", - " else:\n", - " my_dict[e] += 1\n", - " \n", - "def add_element_check2(my_dict, elements):\n", - " for e in elements:\n", - " if e not in my_dict:\n", - " my_dict[e] = 0\n", - " my_dict[e] += 1 \n", - "\n", - "def add_element_except(my_dict, elements):\n", - " for e in elements:\n", - " try:\n", - " my_dict[e] += 1\n", - " except KeyError:\n", - " my_dict[e] = 1\n", - " \n", - "\n", - "random.seed(123)\n", - "rand_ints = [random.randrange(1, 10) for i in range(100)]\n", - "empty_dict = {}\n", - "\n", - "print('Results for 100 integers in range 1-10') \n", - "%timeit add_element_check1(copy.deepcopy(empty_dict), rand_ints)\n", - "%timeit add_element_check2(copy.deepcopy(empty_dict), rand_ints)\n", - "%timeit add_element_except(copy.deepcopy(empty_dict), rand_ints)\n", - " \n", - "print('\\nResults for 1000 integers in range 1-10') \n", - "rand_ints = [random.randrange(1, 10) for i in range(1000)]\n", - "empty_dict = {}\n", - "\n", - "%timeit add_element_check1(copy.deepcopy(empty_dict), rand_ints)\n", - "%timeit add_element_check2(copy.deepcopy(empty_dict), rand_ints)\n", - "%timeit add_element_except(copy.deepcopy(empty_dict), rand_ints)\n", - "\n", - "print('\\nResults for 1000 integers in range 1-1000') \n", - "rand_ints = [random.randrange(1, 10) for i in range(1000)]\n", - "empty_dict = {}\n", - "\n", - "%timeit add_element_check1(copy.deepcopy(empty_dict), rand_ints)\n", - "%timeit add_element_check2(copy.deepcopy(empty_dict), rand_ints)\n", - "%timeit add_element_except(copy.deepcopy(empty_dict), rand_ints)\n", - "\n", - "#\n", - "# Python 3.4.0\n", - "# MacOS X 10.9.2\n", - "# 2.5 GHz Intel Core i5\n", - "# 4 GB 1600 Mhz DDR3\n", - "#" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "Results for 100 integers in range 1-10\n", - "100000 loops, best of 3: 16.6 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "100000 loops, best of 3: 17.6 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "100000 loops, best of 3: 17.9 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "\n", - "Results for 1000 integers in range 1-10\n", - "10000 loops, best of 3: 135 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "10000 loops, best of 3: 125 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "10000 loops, best of 3: 105 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "\n", - "Results for 1000 integers in range 1-1000\n", - "10000 loops, best of 3: 122 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "10000 loops, best of 3: 123 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "10000 loops, best of 3: 104 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n" - ] - } - ], - "prompt_number": 13 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "### Conclusion\n", - "Interestingly, the `try-except` loop pays off if we have more elements (here: 1000 integers instead of 100) as dictionary keys to check. Also, it doesn't matter much whether the elements exist or do not exist in the dictionary, yet." - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "
\n", - "
" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "# Comprehesions vs. for-loops" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Comprehensions are not only shorter and prettier than ye goode olde for-loop, \n", - "but they are also up to ~1.2x faster." - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "import timeit\n", - "n = 1000\n", - "\n", - "#\n", - "# Python 3.4.0\n", - "# MacOS X 10.9.2\n", - "# 2.5 GHz Intel Core i5\n", - "# 4 GB 1600 Mhz DDR3\n", - "#" - ], - "language": "python", - "metadata": {}, - "outputs": [], - "prompt_number": 19 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "## Set comprehensions" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "def set_loop(n):\n", - " a_set = set()\n", - " for i in range(n):\n", - " if i % 3 == 0:\n", - " a_set.add(i)\n", - " return a_set" - ], - "language": "python", - "metadata": {}, - "outputs": [], - "prompt_number": 20 - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "def set_compr(n):\n", - " return {i for i in range(n) if i % 3 == 0}" - ], - "language": "python", - "metadata": {}, - "outputs": [], - "prompt_number": 21 - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "%timeit set_loop(n)\n", - "%timeit set_compr(n)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "10000 loops, best of 3: 136 \u00b5s per loop\n", - "10000 loops, best of 3: 113 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n" - ] - } - ], - "prompt_number": 22 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "## List comprehensions" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "def list_loop(n):\n", - " a_list = list()\n", - " for i in range(n):\n", - " if i % 3 == 0:\n", - " a_list.append(i)\n", - " return a_list" - ], - "language": "python", - "metadata": {}, - "outputs": [], - "prompt_number": 23 - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "def list_compr(n):\n", - " return [i for i in range(n) if i % 3 == 0]" - ], - "language": "python", - "metadata": {}, - "outputs": [], - "prompt_number": 24 - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "%timeit list_loop(n)\n", - "%timeit list_compr(n)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "10000 loops, best of 3: 129 \u00b5s per loop\n", - "10000 loops, best of 3: 111 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n" - ] - } - ], - "prompt_number": 25 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "## Dictionary comprehensions" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "def dict_loop(n):\n", - " a_dict = dict()\n", - " for i in range(n):\n", - " if i % 3 == 0:\n", - " a_dict[i] = i\n", - " return a_dict" - ], - "language": "python", - "metadata": {}, - "outputs": [], - "prompt_number": 26 - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "def dict_compr(n):\n", - " return {i:i for i in range(n) if i % 3 == 0}" - ], - "language": "python", - "metadata": {}, - "outputs": [], - "prompt_number": 27 - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "%timeit dict_loop(n)\n", - "%timeit dict_compr(n)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "10000 loops, best of 3: 121 \u00b5s per loop\n", - "10000 loops, best of 3: 127 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n" - ] - } - ], - "prompt_number": 28 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "
\n", - "
" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "# Copying files by searching directory trees" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Executing `Unix`/`Linux` shell commands:" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "import subprocess\n", - "\n", - "def subprocess_findcopy(path, search_str, dest): \n", - " query = 'find %s -name \"%s\" -exec cp {} %s \\;' %(path, search_str, dest)\n", - " subprocess.call(query, shell=True)\n", - " return " - ], - "language": "python", - "metadata": {}, - "outputs": [], - "prompt_number": 30 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Using Python's `os.walk()` to search the directory tree recursively and matching patterns via `fnmatch.filter()`" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "import shutil\n", - "import os\n", - "import fnmatch\n", - "\n", - "def walk_findcopy(path, search_str, dest):\n", - " for path, subdirs, files in os.walk(path):\n", - " for name in fnmatch.filter(files, search_str):\n", - " try:\n", - " shutil.copy(os.path.join(path,name), dest)\n", - " except NameError:\n", - " pass\n", - " return" - ], - "language": "python", - "metadata": {}, - "outputs": [], - "prompt_number": 33 - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "import timeit\n", - "\n", - "\n", - "def findcopy_timeit(inpath, outpath, search_str):\n", - " \n", - " shutil.rmtree(outpath)\n", - " os.mkdir(outpath)\n", - " print(50*'#')\n", - " print('subprocsess call')\n", - " %timeit subprocess_findcopy(inpath, search_str, outpath)\n", - " print(\"copied %s files\" %len(os.listdir(outpath)))\n", - " shutil.rmtree(outpath)\n", - " os.mkdir(outpath)\n", - " print('\\nos.walk approach')\n", - " %timeit walk_findcopy(inpath, search_str, outpath)\n", - " print(\"copied %s files\" %len(os.listdir(outpath)))\n", - " print(50*'#')\n", - "\n", - "print('small tree')\n", - "inpath = '/Users/sebastian/Desktop/testdir_in'\n", - "outpath = '/Users/sebastian/Desktop/testdir_out'\n", - "search_str = '*.png'\n", - "findcopy_timeit(inpath, outpath, search_str)\n", - "\n", - "print('larger tree')\n", - "inpath = '/Users/sebastian/Dropbox'\n", - "outpath = '/Users/sebastian/Desktop/testdir_out'\n", - "search_str = '*.csv'\n", - "findcopy_timeit(inpath, outpath, search_str)\n" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "small tree\n", - "##################################################" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "subprocsess call\n", - "1 loops, best of 3: 268 ms per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "copied 13 files\n", - "\n", - "os.walk approach\n", - "100 loops, best of 3: 12.2 ms per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "copied 13 files\n", - "##################################################\n", - "larger tree\n", - "##################################################\n", - "subprocsess call\n", - "1 loops, best of 3: 623 ms per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "copied 105 files\n", - "\n", - "os.walk approach\n", - "1 loops, best of 3: 417 ms per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "copied 105 files\n", - "##################################################\n" - ] - } - ], - "prompt_number": 35 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "I have to say that I am really positively surprised. The shell's `find` scales even better than expected!" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "
\n", - "" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "# Returning column vectors slicing through a numpy array" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Given a numpy matrix, I want to iterate through it and return each column as a 1-column vector. \n", - "E.g., if I want to return the 1st column from matrix A below\n", - "\n", - "
\n", - "A = np.array([ [1,2,3], [4,5,6], [7,8,9] ])\n", - ">>> A\n", - "array([[1, 2, 3],\n", - " [4, 5, 6],\n", - " [7, 8, 9]])\n", - "\n", - "I want my result to be:\n", - "
\n", - "array([[1],\n", - " [4],\n", - " [7]])\n", - "\n", - "with `.shape` = `(3,1)`\n", - "\n", - "\n", - "However, the default behavior of numpy is to return the column as a row vector:\n", - "\n", - "
\n", - ">>> A[:,0]\n", - "array([1, 4, 7])\n", - ">>> A[:,0].shape\n", - "(3,)\n", - "" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "import numpy as np\n", - "\n", - "# 1st column, e.g., A[:,0,np.newaxis]\n", - "\n", - "def colvec_method1(A):\n", - " for col in A.T:\n", - " colvec = row[:,np.newaxis]\n", - " yield colvec" - ], - "language": "python", - "metadata": {}, - "outputs": [], - "prompt_number": 83 - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "# 1st column, e.g., A[:,0:1]\n", - "\n", - "def colvec_method2(A):\n", - " for idx in range(A.shape[1]):\n", - " colvec = A[:,idx:idx+1]\n", - " yield colvec" - ], - "language": "python", - "metadata": {}, - "outputs": [], - "prompt_number": 82 - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "# 1st column, e.g., A[:,0].reshape(-1,1)\n", - "\n", - "def colvec_method3(A):\n", - " for idx in range(A.shape[1]):\n", - " colvec = A[:,idx].reshape(-1,1)\n", - " yield colvec" - ], - "language": "python", - "metadata": {}, - "outputs": [], - "prompt_number": 81 - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "# 1st column, e.g., np.vstack(A[:,0]\n", - "\n", - "def colvec_method4(A):\n", - " for idx in range(A.shape[1]):\n", - " colvec = np.vstack(A[:,idx])\n", - " yield colvec" - ], - "language": "python", - "metadata": {}, - "outputs": [], - "prompt_number": 79 - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "# 1st column, e.g., np.row_stack(A[:,0])\n", - "\n", - "def colvec_method5(A):\n", - " for idx in range(A.shape[1]):\n", - " colvec = np.row_stack(A[:,idx])\n", - " yield colvec" - ], - "language": "python", - "metadata": {}, - "outputs": [], - "prompt_number": 77 - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "# 1st column, e.g., np.column_stack((A[:,0],))\n", - "\n", - "def colvec_method6(A):\n", - " for idx in range(A.shape[1]):\n", - " colvec = np.column_stack((A[:,idx],))\n", - " yield colvec" - ], - "language": "python", - "metadata": {}, - "outputs": [], - "prompt_number": 74 - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "# 1st column, e.g., A[:,[0]]\n", - "\n", - "def colvec_method7(A):\n", - " for idx in range(A.shape[1]):\n", - " colvec = A[:,[idx]]\n", - " yield colvec" - ], - "language": "python", - "metadata": {}, - "outputs": [], - "prompt_number": 89 - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "def test_method(method, A):\n", - " for i in method(A): \n", - " assert i.shape == (A.shape[0],1), \"{}, {}\".format(i.shape, A.shape[0],1)" - ], - "language": "python", - "metadata": {}, - "outputs": [], - "prompt_number": 69 - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "import timeit\n", - "\n", - "A = np.random.random((300, 3))\n", - "\n", - "for method in [\n", - " colvec_method1, colvec_method2, \n", - " colvec_method3, colvec_method4, \n", - " colvec_method5, colvec_method6,\n", - " colvec_method7]:\n", - " print('\\nTest:', method.__name__)\n", - " %timeit test_method(colvec_method2, A)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "Test: colvec_method1\n", - "100000 loops, best of 3: 16.6 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "\n", - "Test: colvec_method2\n", - "10000 loops, best of 3: 16.1 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "\n", - "Test: colvec_method3\n", - "100000 loops, best of 3: 16.2 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "\n", - "Test: colvec_method4\n", - "100000 loops, best of 3: 16.4 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "\n", - "Test: colvec_method5\n", - "100000 loops, best of 3: 16.2 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "\n", - "Test: colvec_method6\n", - "100000 loops, best of 3: 16.8 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "\n", - "Test: colvec_method7\n", - "100000 loops, best of 3: 16.3 \u00b5s per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n" - ] - } - ], - "prompt_number": 91 - }, - { - "cell_type": "code", - "collapsed": false, - "input": [], - "language": "python", - "metadata": {}, - "outputs": [] - } - ], - "metadata": {} - } - ] -} \ No newline at end of file diff --git a/CSS/hover.css b/CSS/hover.css new file mode 100644 index 0000000..e04a4aa --- /dev/null +++ b/CSS/hover.css @@ -0,0 +1,26 @@ +.hover_light_dark{ + opacity: 0.5; + filter: alpha(opacity=50); + no-repeat; + } + +.hover_light_dark:hover { + opacity: 1; + filter: alpha(opacity=100); + } + + + +.hover_dark_light{ + opacity: 1; + filter: alpha(opacity=100); + no-repeat; + } + + +.hover_dark_light:hover { + opacity: 0.5; + filter: alpha(opacity=500); + } + + diff --git a/Data/some_soccer_data.csv b/Data/some_soccer_data.csv new file mode 100644 index 0000000..c2cdab8 --- /dev/null +++ b/Data/some_soccer_data.csv @@ -0,0 +1,21 @@ +PLAYER,SALARY,GP,G,A,SOT,PPG,P +"Sergio Agüero + Forward — Manchester City",$19.2m,16,14,3,34,13.12,209.98 +"Eden Hazard + Midfield — Chelsea",$18.9m,21,8,4,17,13.05,274.04 +"Alexis Sánchez + Forward — Arsenal",$17.6m,,12,7,29,11.19,223.86 +"Yaya Touré + Midfield — Manchester City",$16.6m,18,7,1,19,10.99,197.91 +"Ángel Di María + Midfield — Manchester United",$15.0m,13,3,,13,10.17,132.23 +"Santiago Cazorla + Midfield — Arsenal",$14.8m,20,4,,20,9.97, +"David Silva + Midfield — Manchester City",$14.3m,15,6,2,11,10.35,155.26 +"Cesc Fàbregas + Midfield — Chelsea",$14.0m,20,2,14,10,10.47,209.49 +"Saido Berahino + Forward — West Brom",$13.8m,21,9,0,20,7.02,147.43 +"Steven Gerrard + Midfield — Liverpool",$13.8m,20,5,1,11,7.5,150.01 diff --git a/Data/test.csv b/Data/test.csv new file mode 100644 index 0000000..7c8d315 --- /dev/null +++ b/Data/test.csv @@ -0,0 +1,7 @@ +name,column1,column2,column3 +abc,1.1,4.2,1.2 +def,2.1,1.4,5.2 +ghi,1.5,1.2,2.1 +jkl,1.8,1.1,4.2 +mno,9.4,6.6,6.2 +pqr,1.4,8.3,8.4 diff --git a/Data/test_marked.csv b/Data/test_marked.csv new file mode 100644 index 0000000..86dfd82 --- /dev/null +++ b/Data/test_marked.csv @@ -0,0 +1,7 @@ +name,column1,column2,column3 +abc,1.1^,4.2,1.2^ +def,2.1,1.4,5.2 +ghi,1.5,1.2,2.1 +jkl,1.8,1.1^,4.2 +mno,9.4*,6.6,6.2 +pqr,1.4,8.3*,8.4* diff --git a/Images/1_sqlite3_init_db.png b/Images/1_sqlite3_init_db.png new file mode 100644 index 0000000..c85f420 Binary files /dev/null and b/Images/1_sqlite3_init_db.png differ diff --git a/Images/2_sqlite3_add_col.png b/Images/2_sqlite3_add_col.png new file mode 100644 index 0000000..63e1c50 Binary files /dev/null and b/Images/2_sqlite3_add_col.png differ diff --git a/Images/3_sqlite3_insert_update.png b/Images/3_sqlite3_insert_update.png new file mode 100644 index 0000000..9cb1b26 Binary files /dev/null and b/Images/3_sqlite3_insert_update.png differ diff --git a/Images/4_sqlite3_unique_index.png b/Images/4_sqlite3_unique_index.png new file mode 100644 index 0000000..a3ad45a Binary files /dev/null and b/Images/4_sqlite3_unique_index.png differ diff --git a/Images/5_sqlite3_date_time.png b/Images/5_sqlite3_date_time.png new file mode 100644 index 0000000..b28a115 Binary files /dev/null and b/Images/5_sqlite3_date_time.png differ diff --git a/Images/5_sqlite3_date_time_2.png b/Images/5_sqlite3_date_time_2.png new file mode 100644 index 0000000..c077481 Binary files /dev/null and b/Images/5_sqlite3_date_time_2.png differ diff --git a/Images/6_sqlite3_print_selecting_rows.png b/Images/6_sqlite3_print_selecting_rows.png new file mode 100644 index 0000000..26be818 Binary files /dev/null and b/Images/6_sqlite3_print_selecting_rows.png differ diff --git a/Images/7_sqlite3_get_colnames_1.png b/Images/7_sqlite3_get_colnames_1.png new file mode 100644 index 0000000..a400cd4 Binary files /dev/null and b/Images/7_sqlite3_get_colnames_1.png differ diff --git a/Images/7_sqlite3_get_colnames_2.png b/Images/7_sqlite3_get_colnames_2.png new file mode 100644 index 0000000..8b3d95f Binary files /dev/null and b/Images/7_sqlite3_get_colnames_2.png differ diff --git a/Images/8_sqlite3_print_db_info_1.png b/Images/8_sqlite3_print_db_info_1.png new file mode 100644 index 0000000..a400cd4 Binary files /dev/null and b/Images/8_sqlite3_print_db_info_1.png differ diff --git a/Images/8_sqlite3_print_db_info_2.png b/Images/8_sqlite3_print_db_info_2.png new file mode 100644 index 0000000..9aac8e6 Binary files /dev/null and b/Images/8_sqlite3_print_db_info_2.png differ diff --git a/Images/Ipv4_address.png b/Images/Ipv4_address.png new file mode 100644 index 0000000..04028af Binary files /dev/null and b/Images/Ipv4_address.png differ diff --git a/Images/Ipv6_address.png b/Images/Ipv6_address.png new file mode 100644 index 0000000..8929b65 Binary files /dev/null and b/Images/Ipv6_address.png differ diff --git a/Images/MACaddressV3.png b/Images/MACaddressV3.png new file mode 100644 index 0000000..3064c5a Binary files /dev/null and b/Images/MACaddressV3.png differ diff --git a/Images/cython_final_leastsqr.png b/Images/cython_final_leastsqr.png new file mode 100644 index 0000000..036c507 Binary files /dev/null and b/Images/cython_final_leastsqr.png differ diff --git a/Images/cython_vs_chart.png b/Images/cython_vs_chart.png new file mode 100644 index 0000000..a536ef2 Binary files /dev/null and b/Images/cython_vs_chart.png differ diff --git a/Images/ipython_links_ex.png b/Images/ipython_links_ex.png new file mode 100644 index 0000000..6b3dacf Binary files /dev/null and b/Images/ipython_links_ex.png differ diff --git a/Images/ipython_links_format.png b/Images/ipython_links_format.png new file mode 100644 index 0000000..d0627e5 Binary files /dev/null and b/Images/ipython_links_format.png differ diff --git a/Images/ipython_links_overview.png b/Images/ipython_links_overview.png new file mode 100644 index 0000000..16cb19f Binary files /dev/null and b/Images/ipython_links_overview.png differ diff --git a/Images/ipython_links_remedy.png b/Images/ipython_links_remedy.png new file mode 100644 index 0000000..4c200e2 Binary files /dev/null and b/Images/ipython_links_remedy.png differ diff --git a/Images/ipython_links_remedy2.png b/Images/ipython_links_remedy2.png new file mode 100644 index 0000000..ae3abea Binary files /dev/null and b/Images/ipython_links_remedy2.png differ diff --git a/Images/ipython_table_header.png b/Images/ipython_table_header.png new file mode 100644 index 0000000..897ba78 Binary files /dev/null and b/Images/ipython_table_header.png differ diff --git a/Images/lda_overview.png b/Images/lda_overview.png new file mode 100644 index 0000000..8856366 Binary files /dev/null and b/Images/lda_overview.png differ diff --git a/Images/least_squares_perpendicular.png b/Images/least_squares_perpendicular.png new file mode 100644 index 0000000..a0d425f Binary files /dev/null and b/Images/least_squares_perpendicular.png differ diff --git a/Images/least_squares_vertical.png b/Images/least_squares_vertical.png new file mode 100644 index 0000000..1162973 Binary files /dev/null and b/Images/least_squares_vertical.png differ diff --git a/Images/literature.png b/Images/literature.png new file mode 100644 index 0000000..25a5447 Binary files /dev/null and b/Images/literature.png differ diff --git a/Images/literature_small.png b/Images/literature_small.png new file mode 100644 index 0000000..2ffcedf Binary files /dev/null and b/Images/literature_small.png differ diff --git a/Images/logo.png b/Images/logo.png new file mode 100644 index 0000000..64d8efd Binary files /dev/null and b/Images/logo.png differ diff --git a/Images/logo.pxm b/Images/logo.pxm new file mode 100644 index 0000000..b6b42e3 Binary files /dev/null and b/Images/logo.pxm differ diff --git a/Images/matcheat_R_logo.png b/Images/matcheat_R_logo.png new file mode 100644 index 0000000..ce6c192 Binary files /dev/null and b/Images/matcheat_R_logo.png differ diff --git a/Images/matcheat_julia_benchmark.png b/Images/matcheat_julia_benchmark.png new file mode 100644 index 0000000..2af6e46 Binary files /dev/null and b/Images/matcheat_julia_benchmark.png differ diff --git a/Images/matcheat_julia_logo.png b/Images/matcheat_julia_logo.png new file mode 100644 index 0000000..2339fea Binary files /dev/null and b/Images/matcheat_julia_logo.png differ diff --git a/Images/matcheat_matlab_logo.png b/Images/matcheat_matlab_logo.png new file mode 100644 index 0000000..e6993c7 Binary files /dev/null and b/Images/matcheat_matlab_logo.png differ diff --git a/Images/matcheat_matrix.png b/Images/matcheat_matrix.png new file mode 100644 index 0000000..e986078 Binary files /dev/null and b/Images/matcheat_matrix.png differ diff --git a/Images/matcheat_numpy_logo.png b/Images/matcheat_numpy_logo.png new file mode 100644 index 0000000..c56d4c0 Binary files /dev/null and b/Images/matcheat_numpy_logo.png differ diff --git a/Images/matcheat_octave_logo.png b/Images/matcheat_octave_logo.png new file mode 100644 index 0000000..f046a29 Binary files /dev/null and b/Images/matcheat_octave_logo.png differ diff --git a/Images/multiprocessing_scheme.png b/Images/multiprocessing_scheme.png new file mode 100644 index 0000000..9cb130d Binary files /dev/null and b/Images/multiprocessing_scheme.png differ diff --git a/Images/pytest_01.png b/Images/pytest_01.png new file mode 100644 index 0000000..7654293 Binary files /dev/null and b/Images/pytest_01.png differ diff --git a/Images/pytest_02.png b/Images/pytest_02.png new file mode 100644 index 0000000..441156c Binary files /dev/null and b/Images/pytest_02.png differ diff --git a/Images/pytest_02_2.png b/Images/pytest_02_2.png new file mode 100644 index 0000000..a17ce8c Binary files /dev/null and b/Images/pytest_02_2.png differ diff --git a/Images/pytest_03.png b/Images/pytest_03.png new file mode 100644 index 0000000..50bfa9c Binary files /dev/null and b/Images/pytest_03.png differ diff --git a/Images/pytest_04.png b/Images/pytest_04.png new file mode 100644 index 0000000..4fc515c Binary files /dev/null and b/Images/pytest_04.png differ diff --git a/Images/pytest_05.png b/Images/pytest_05.png new file mode 100644 index 0000000..cbdab6b Binary files /dev/null and b/Images/pytest_05.png differ diff --git a/Images/pytest_06.png b/Images/pytest_06.png new file mode 100644 index 0000000..c73b986 Binary files /dev/null and b/Images/pytest_06.png differ diff --git a/Images/pytest_07.png b/Images/pytest_07.png new file mode 100644 index 0000000..853d139 Binary files /dev/null and b/Images/pytest_07.png differ diff --git a/Images/pytest_08.png b/Images/pytest_08.png new file mode 100644 index 0000000..d558eed Binary files /dev/null and b/Images/pytest_08.png differ diff --git a/Images/pytest_09.png b/Images/pytest_09.png new file mode 100644 index 0000000..5ae43b9 Binary files /dev/null and b/Images/pytest_09.png differ diff --git a/Images/pytest_10.png b/Images/pytest_10.png new file mode 100644 index 0000000..77f1bf2 Binary files /dev/null and b/Images/pytest_10.png differ diff --git a/Images/pytest_11.png b/Images/pytest_11.png new file mode 100644 index 0000000..48e9f58 Binary files /dev/null and b/Images/pytest_11.png differ diff --git a/Images/pytest_12.png b/Images/pytest_12.png new file mode 100644 index 0000000..8105eb7 Binary files /dev/null and b/Images/pytest_12.png differ diff --git a/Images/pytest_13.png b/Images/pytest_13.png new file mode 100644 index 0000000..a374b1d Binary files /dev/null and b/Images/pytest_13.png differ diff --git a/Images/python-logo-master-v3-TM-flattened.png b/Images/python-logo-master-v3-TM-flattened.png new file mode 100644 index 0000000..738f6ed Binary files /dev/null and b/Images/python-logo-master-v3-TM-flattened.png differ diff --git a/Images/python_sci_pack_ing.png b/Images/python_sci_pack_ing.png new file mode 100644 index 0000000..e526f62 Binary files /dev/null and b/Images/python_sci_pack_ing.png differ diff --git a/Images/scope_resolution_1.png b/Images/scope_resolution_1.png new file mode 100644 index 0000000..6bf48b0 Binary files /dev/null and b/Images/scope_resolution_1.png differ diff --git a/Images/sqlite_python_logo.png b/Images/sqlite_python_logo.png new file mode 100644 index 0000000..25740d3 Binary files /dev/null and b/Images/sqlite_python_logo.png differ diff --git a/Images/thumb_sqlite3.png b/Images/thumb_sqlite3.png new file mode 100644 index 0000000..52f1504 Binary files /dev/null and b/Images/thumb_sqlite3.png differ diff --git a/README.md b/README.md old mode 100755 new mode 100644 index cc1f5ea..05de8e6 --- a/README.md +++ b/README.md @@ -1,12 +1,220 @@ -python_reference -================ +
A collection of useful scripts, tutorials, and other Python-related things
-Syntax examples for useful Python functions, methods, and modules ++
 -###Links to view the IPython Notebooks
+
-- [Python benchmarks via `timeit`](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/benchmarks/timeit_tests.ipynb?create=1)
-- [Benchmarks of different palindrome functions](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/benchmarks/palindrome_timeit.ipynb?create=1)
-- [A collection of not so obvious Python stuff you should know!](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/not_so_obvious_python_stuff.ipynb?create=1)
+- [// Python tips and tutorials](#-python-tips-and-tutorials)
+- [// Python and the web](#-python-and-the-web)
+- [// Algorithms](#-algorithms)
+- [// Plotting and Visualization](#-plotting-and-visualization)
+- [// Benchmarks](#-benchmarks)
+- [// Python and "Data Science"](#-python-and-data-science)
+- [// Useful scripts and snippets](#-useful-scripts-and-snippets)
+- [// Other](#-other)
+- [// Links](#-links)
+
+
+
+
+
+Python tips and tutorials [back to top]
+
+- A collection of not so obvious Python stuff you should know! [[IPython nb](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/tutorials/not_so_obvious_python_stuff.ipynb?create=1)]
+
+- Python's scope resolution for variable names and the LEGB rule [[IPython nb](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/tutorials/scope_resolution_legb_rule.ipynb?create=1)]
+
+- Key differences between Python 2.x and Python 3.x [[IPython nb](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/tutorials/key_differences_between_python_2_and_3.ipynb?create=1)]
+
+- A thorough guide to SQLite database operations in Python [[Markdown](./tutorials/sqlite3_howto/README.md)]
+
+- Unit testing in Python - Why we want to make it a habit [[Markdown](./tutorials/unit_testing.md)]
+
+- Installing Scientific Packages for Python3 on MacOS 10.9 Mavericks [[Markdown](./tutorials/installing_scientific_packages.md)]
+
+- Sorting CSV files using the Python csv module [[IPython nb](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/tutorials/sorting_csvs.ipynb)]
+
+- Using Cython with and without IPython magic [[IPython nb](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/tutorials/running_cython.ipynb)]
+
+- Parallel processing via the multiprocessing module [[IPython nb](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/tutorials/multiprocessing_intro.ipynb?create=1)]
+
+- Entry point: Data - using sci-packages to prepare data for Machine Learning tasks and other data analyses [[IPython nb](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/tutorials/python_data_entry_point.ipynb?create=1)]
+
+- Awesome things that you can do in IPython Notebooks (in progress) [[IPython nb](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/tutorials/awesome_things_ipynb.ipynb)]
+
+- A collection of useful regular expressions [[IPython nb](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/tutorials/useful_regex.ipynb)]
+
+- Quick guide for dealing with missing numbers in NumPy [[IPython nb](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/tutorials/numpy_nan_quickguide.ipynb)]
+
+- A random collection of useful Python snippets [[IPython nb](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/python_patterns/patterns.ipynb)]
+
+- Things in pandas I wish I'd had known earlier [[IPython nb](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/tutorials/things_in_pandas.ipynb)]
+
+
+
+
-###Links to view the IPython Notebooks
+
-- [Python benchmarks via `timeit`](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/benchmarks/timeit_tests.ipynb?create=1)
-- [Benchmarks of different palindrome functions](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/benchmarks/palindrome_timeit.ipynb?create=1)
-- [A collection of not so obvious Python stuff you should know!](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/not_so_obvious_python_stuff.ipynb?create=1)
+- [// Python tips and tutorials](#-python-tips-and-tutorials)
+- [// Python and the web](#-python-and-the-web)
+- [// Algorithms](#-algorithms)
+- [// Plotting and Visualization](#-plotting-and-visualization)
+- [// Benchmarks](#-benchmarks)
+- [// Python and "Data Science"](#-python-and-data-science)
+- [// Useful scripts and snippets](#-useful-scripts-and-snippets)
+- [// Other](#-other)
+- [// Links](#-links)
+
+
+
+
+
+Python tips and tutorials [back to top]
+
+- A collection of not so obvious Python stuff you should know! [[IPython nb](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/tutorials/not_so_obvious_python_stuff.ipynb?create=1)]
+
+- Python's scope resolution for variable names and the LEGB rule [[IPython nb](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/tutorials/scope_resolution_legb_rule.ipynb?create=1)]
+
+- Key differences between Python 2.x and Python 3.x [[IPython nb](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/tutorials/key_differences_between_python_2_and_3.ipynb?create=1)]
+
+- A thorough guide to SQLite database operations in Python [[Markdown](./tutorials/sqlite3_howto/README.md)]
+
+- Unit testing in Python - Why we want to make it a habit [[Markdown](./tutorials/unit_testing.md)]
+
+- Installing Scientific Packages for Python3 on MacOS 10.9 Mavericks [[Markdown](./tutorials/installing_scientific_packages.md)]
+
+- Sorting CSV files using the Python csv module [[IPython nb](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/tutorials/sorting_csvs.ipynb)]
+
+- Using Cython with and without IPython magic [[IPython nb](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/tutorials/running_cython.ipynb)]
+
+- Parallel processing via the multiprocessing module [[IPython nb](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/tutorials/multiprocessing_intro.ipynb?create=1)]
+
+- Entry point: Data - using sci-packages to prepare data for Machine Learning tasks and other data analyses [[IPython nb](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/tutorials/python_data_entry_point.ipynb?create=1)]
+
+- Awesome things that you can do in IPython Notebooks (in progress) [[IPython nb](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/tutorials/awesome_things_ipynb.ipynb)]
+
+- A collection of useful regular expressions [[IPython nb](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/tutorials/useful_regex.ipynb)]
+
+- Quick guide for dealing with missing numbers in NumPy [[IPython nb](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/tutorials/numpy_nan_quickguide.ipynb)]
+
+- A random collection of useful Python snippets [[IPython nb](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/python_patterns/patterns.ipynb)]
+
+- Things in pandas I wish I'd had known earlier [[IPython nb](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/tutorials/things_in_pandas.ipynb)]
+
+
+
++ + +Python and the web [back to top] + +- Creating internal links in IPython Notebooks and Markdown docs [[IPython nb](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/tutorials/table_of_contents_ipython.ipynb)] + +- Converting Markdown to HTML and adding Python syntax highlighting [[Markdown](./tutorials/markdown_syntax_highlighting/README.md)] + + +
+Algorithms and Data Structures [back to top] + +*This category has been moved to a separate GitHub repository [rasbt/algorithms_in_ipython_notebooks](https://github.com/rasbt/algorithms_in_ipython_notebooks)* + + +- Sorting Algorithms [[Collection of IPython Notebooks](https://github.com/rasbt/algorithms_in_ipython_notebooks/tree/master/ipython_nbs/sorting) + +- Linear regression via the least squares fit method [[IPython nb](http://nbviewer.ipython.org/github/rasbt/algorithms_in_ipython_notebooks/blob/master/ipython_nbs/statistics/linregr_least_squares_fit.ipynb?create=1)] + +- Dixon's Q test to identify outliers for small sample sizes [[IPython nb](http://nbviewer.ipython.org/github/rasbt/algorithms_in_ipython_notebooks/blob/master/ipython_nbs/statistics/dixon_q_test.ipynb?create=1)] + +- Counting points inside a hypercube [[IPython nb](http://nbviewer.ipython.org/github/rasbt/algorithms_in_ipython_notebooks/blob/master/ipython_nbs/geometry/points_in_hybercube.ipynb)] + +- Singly Linked List [[ IPython nbviewer ](http://nbviewer.ipython.org/github/rasbt/algorithms_in_ipython_notebooks/blob/master/ipython_nbs/data-structures/singly-linked-list.ipynb)] + +
+Plotting and Visualization [back to top] + +*The matplotlib-gallery in IPython notebooks has been moved to a separate GitHub repository [matplotlib-gallery](https://github.com/rasbt/matplotlib-gallery)* + +**Featured articles**: + +- Preparing Plots for Publication [[IPython nb](http://nbviewer.ipython.org/github/rasbt/matplotlib-gallery/blob/master/ipynb/publication.ipynb)] + + + + + +
+Benchmarks [back to top] + + +- Simple tricks to speed up the sum calculation in pandas [[IPython nb](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/benchmarks/pandas_sum_tricks.ipynb)] + +
+*More benchmarks can be found in the separate GitHub repository [One-Python-benchmark-per-day](https://github.com/rasbt/One-Python-benchmark-per-day)* + +**Featured articles**: + + + +- (C)Python compilers - Cython vs. Numba vs. Parakeet [[IPython nb](http://nbviewer.ipython.org/github/rasbt/One-Python-benchmark-per-day/blob/master/ipython_nbs/day4_2_cython_numba_parakeet.ipynb)] + +- Just-in-time compilers for NumPy array expressions [[IPython nb](http://nbviewer.ipython.org/github/rasbt/One-Python-benchmark-per-day/blob/master/ipython_nbs/day7_2_jit_numpy.ipynb)] + +- Cython - Bridging the gap between Python and Fortran [[IPython nb](http://nbviewer.ipython.org/github/rasbt/One-Python-benchmark-per-day/blob/master/ipython_nbs/day10_fortran_lstsqr.ipynb)] + +- Parallel processing via the multiprocessing module [[IPython nb](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/tutorials/multiprocessing_intro.ipynb)] + +- Vectorizing a classic for-loop in NumPy [[IPython nb](http://nbviewer.ipython.org/github/rasbt/One-Python-benchmark-per-day/blob/master/ipython_nbs/day16_numpy_vectorization.ipynb)] + +
+ + +Python and "Data Science" [back to top] + +*The "data science"-related posts have been moved to a separate GitHub repository [pattern_classification](https://github.com/rasbt/pattern_classification)* + +**Featured articles**: + +- Entry Point: Data - Using Python's sci-packages to prepare data for Machine Learning tasks and other data analyses [[IPython nb](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/tutorials/python_data_entry_point.ipynb)] + +- About Feature Scaling: Standardization and Min-Max-Scaling (Normalization) [[IPython nb](http://nbviewer.ipython.org/github/rasbt/pattern_classification/blob/master/preprocessing/about_standardization_normalization.ipynb)] + +- Principal Component Analysis (PCA) [[IPython nb](http://nbviewer.ipython.org/github/rasbt/pattern_classification/blob/master/dimensionality_reduction/projection/principal_component_analysis.ipynb)] + +- Linear Discriminant Analysis (LDA) [[IPython nb](http://nbviewer.ipython.org/github/rasbt/pattern_classification/blob/master/dimensionality_reduction/projection/linear_discriminant_analysis.ipynb)] + +- Kernel density estimation via the Parzen-window technique [[IPython nb](http://nbviewer.ipython.org/github/rasbt/pattern_classification/blob/master/parameter_estimation_techniques/parzen_window_technique.ipynb)] + + +
+ +Useful scripts and snippets [back to top] + +- [watermark](https://github.com/rasbt/watermark) - An IPython magic extension for printing date and time stamps, version numbers, and hardware information. + +- [Shell script](./useful_scripts/prepend_python_shebang.sh) For prepending Python-shebangs to .py files. + +- A random string generator [function](./useful_scripts/random_string_generator.py). + +- [Converting large CSV files](https://github.com/rasbt/python_reference/blob/master/useful_scripts/large_csv_to_sqlite.py) to SQLite databases using pandas. + +- [Sparsifying a matrix](https://github.com/rasbt/python_reference/blob/master/useful_scripts/sparsify_matrix.py) by zeroing out all elements but the top k elements in a row using NumPy. + +
+ +Other [back to top] + +- [Python book reviews](./other/python_book_reviews.md) +- [Happy Mother's Day Plot](./other/happy_mothers_day.ipynb) + +
+ +Links [back to top] + + + +- [PyPI - the Python Package Index](https://pypi.python.org/pypi) - The official repository for all open source Python modules and packages. + +- [PEP 8](https://www.python.org/dev/peps/pep-0008/) - The official style guide for Python code. + +- [PEP 257](https://www.python.org/dev/peps/pep-0257/) - Python's official docstring conventions; [pep257 - Python style guide checker](https://pypi.python.org/pypi/pep257) + + +
+ +**// News** + +- [Python subreddit](http://www.reddit.com/r/Python/) - My favorite resource to catch up with Python news and great Python-related articles. + +- [Python community on Google+](https://plus.google.com/communities/103393744324769547228) - A nice and friendly community to share and discuss everything about Python. + +- [Python Weekly](http://www.pythonweekly.com) - A free weekly newsletter featuring curated news, articles, new releases, jobs etc. related to Python. + + +
+ +**// Resources for learning Python** + +- [Dive Into Python](http://www.diveintopython.net) / [Dive Into Python 3](http://getpython3.com/diveintopython3/) - A free Python book for experienced programmers. + +- [The Hitchhiker’s Guide to Python](http://docs.python-guide.org/en/latest/) - A free best-practice handbook for both novices and experts. + +- [Think Python - How to Think Like a Computer Scientist](http://www.greenteapress.com/thinkpython/) - An introduction for beginners starting with basic concepts of programming. + +- [A Byte of Python](https://python.swaroopch.com/) - a free book on programming using the Python language. + +- [Python Patterns](http://matthiaseisen.com/pp/) - A directory of proven, reusable solutions to common programming problems. + +- [Intro to Computer Science - Build a Search Engine & a Social Network](https://www.udacity.com/course/intro-to-computer-science--cs101) - A great, free course for learning Python if you haven't programmed before. + +
+ +**// My favorite Python projects and packages** + +- [The IPython Notebook](http://ipython.org/notebook.html) - An interactive computational environment for combining code execution, documentation (with Markdown and LateX support), inline plots, and rich media all in one document. + +- [matplotlib](http://matplotlib.org) - Python's favorite plotting library. + +- [NumPy](http://www.numpy.org) - A library for multi-dimensional arrays and matrices, along with a large library of high-level mathematical functions to operate on these arrays. + +- [SciPy](http://www.scipy.org) - A library that provides various useful functions for numerical computing, such as modules for optimization, linear algebra, integration, interpolation, ... + + +- [pandas](http://pandas.pydata.org) - High-performance, easy-to-use data structures and data analysis tools build on top of NumPy. + +- [Cython](http://cython.org) - C-extensions for Python, an optimizing static compiler to combine Python and C code. + +- [Numba](http://numba.pydata.org) - A just-in-time specializing compiler which compiles annotated Python and NumPy code to LLVM (through decorators) + +- [scikit-learn](http://scikit-learn.org/stable/) - A powerful machine learning library for Python and tools for efficient data mining and analysis. diff --git a/algorithms/sequential_selection_algorithms.ipynb b/algorithms/sequential_selection_algorithms.ipynb new file mode 100644 index 0000000..9032aa2 --- /dev/null +++ b/algorithms/sequential_selection_algorithms.ipynb @@ -0,0 +1,1419 @@ +{ + "metadata": { + "name": "", + "signature": "sha256:422c66e0088094cc07058647ff0a8c20c5fbb08fad34a18ee957f594d82b1e53" + }, + "nbformat": 3, + "nbformat_minor": 0, + "worksheets": [ + { + "cells": [ + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "*Sebastian Raschka* \n", + "last updated: **03/29/2014** \n", + "[Link to this IPython Notebook on GitHub](https://github.com/rasbt/algorithms_in_ipython_notebooks) \n", + "\n", + "
\n", + "Executed in Python 3.4.0\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "
\n", + "
\n", + "#Sections\n", + "- 1. Introduction
\n", + " - Defining a criterion function for testing
\n", + " - 2. Sequential Forward Selection (SFS)
\n", + " - SFS Code
\n", + " - Example SFS
\n", + "- 3. Sequential Backward Selection (SBS)
\n", + " - SBS Code
\n", + " - Example SBS
\n", + "- 4. \"Plus L take away R\" (+L -R)
\n", + " - +L -R Code
\n", + " - Example +L -R
\n", + " - Example 1: L > R
\n", + " - Example 2: R > L
\n", + "- 5. Sequential Floating Forward Selection (SFFS)
\n", + " - SFFS Code
\n", + " - Example SFFS
\n", + "- 6. Sequential Floating Backward Selection (SFBS)
\n", + " - SFBS Code
\n", + " - Example SFBS
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#Sequential Selection Algorithms in Python\n", + "\n", + "\n", + "## 1. Introduction \n", + "\n", + "[[back to top]](#sections)\n", + "\n", + "One of the biggest challenges of designing a good classifier for solving a ***Statistical Pattern Classification*** problem is to estimate the underlying parameters to fit the model - given that the forms of the underlying probability distributions are known. The larger the number of parameters becomes, the more difficult it naturally is to estimate those parameters accurately (***Curse of Dimensionality***) from a limited number of training samples. \n", + "\n", + "In order to avoid the ***Curse of Dimensionality***, pattern classification is often accompanied by ***Dimensionality Reduction***, which also has the nice side-effect of increasing the computational performance.\n", + "Common techniques are projection-based, such as ***Principal Component Analysis (PCA)***, ***Linear Dimension Analysis (LDA)***, and ***Multivariate Dimension Analysis (MDA)***. \n", + "An alternative to the projection-based approach is the so-called ***Feature Selection***, and in this article, we will take a look at some of the established algorithms to tackle this combinatorial search problem. Note that those algorithms are considered as \"subpoptimal\" in contrast to an ***exhaustive search***, which is often computationally not feasible, though.\n", + "\n", + "Therefore, the goal of the presented ***sequential selection algorithms*** is to reduce the feature space *D = {x_1, x_2, x_n}* to a subset of features *D - n* in order to improve or optimize the **computational performance** of the classifier and to avoid the so-called ***Curse of Dimensionality***. \n", + "The goal is to select a \"sufficiently reduced\" subset from the feature space *D* \"without significantly reducing\" the performance of the classifier. In the process of choosing an \"optimal\" feature subset of size *k*, a so-called ***Criterion Function***, which typically, simply, and intuitively assesses the ***recognition rate*** of the classifier.\n", + "\n", + "F. Ferri, P. Pudil, M. Hatef, and J. Kittler investigated the performance of different ***Sequential Selection Algorithms*** for ***Feature Selection*** on different scales and reported their results in a nice research article: *\"[Comparative Study of Techniques for Large Scale Feature Selection](http://citeseerx.ist.psu.edu/viewdoc/download;jsessionid=02CB16CB1C28EA6CB57E212861CFB180?doi=10.1.1.24.4369&rep=rep1&type=pdf),\" Pattern Recognition in Practice IV, E. Gelsema and L. Kanal, eds., pp. 403-413. Elsevier Science B.V., 1994.* \n", + "Choosing an \"appropriate\" algorithm really depends on the problem - the size and desired recognition rate and computational performance. Thus, I want to encourage you to take (at least) a brief look at their paper and the results they obtained from experimenting with different problems feature space dimensions.\n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "### Defining a criterion function (for testing)\n", + "\n", + "[[back to top]](#sections)\n", + "\n", + "In order to evaluate the performance of our selected ***feature subset*** (typically the recognition rate of the classifier), we need to define a ***criterion function***.\n", + "\n", + "For the sake of simplicity, and in order to get an intuitive idea if our algorithm returns an \"appropriate\" result, let us define a very simple criterion function here. \n", + "The criterion function defined below simply returns the sum of numerical values in a list." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "def simple_crit_func(feat_sub):\n", + " \"\"\" Returns sum of numerical values of an input list. \"\"\" \n", + " return sum(feat_sub)\n", + "\n", + "# Example:\n", + "simple_crit_func([1,2,4])" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "pyout", + "prompt_number": 1, + "text": [ + "7" + ] + } + ], + "prompt_number": 1 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "\n", + "## 2. Sequential Forward Selection (SFS)\n", + "\n", + "[[back to top]](#sections)\n", + "\n", + "The ***Sequential Fortward Selection (SFS)*** is one of the simplest and probably fastest *Feature Selection* algorithms. \n", + "Let's summarize its mechanics in words: \n", + "***SFS*** starts with an empty feature subset and sequentially adds features from the whole input feature space to this subset until the subset reaches a desired (user-specified) size. For every iteration (= inclusion of a new feature), the whole feature subset is evaluated (expect for the features that are already included in the new subset). The evaluation is done by the so-called ***criterion function*** which assesses the feature that leads to the maximum performance improvement of the feature subset if it is included. \n", + "Note that included features are never removed, which is one of the biggest downsides of this algorithm.
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
\n", + "\n", + "**Input:** \n", + "the set of all features, \n", + "- $ Y ={y_1, y_2, ..., y_d} $\n", + "\n", + "\n", + "The ***SFS*** algorithm takes the whole feature set as input, if our feature space consists of, e.g. 10, if our feature space consists of 10 dimensions (***d = 10***).\n", + "
\n", + "\n", + "**Output:** \n", + "a subset of features, \n", + "- $X_k = {x_j \\; | \\; j = 1, 2, ..., k; x_j \u2208 Y}$, \n", + "where $k = (0, 1, 2, ..., d $) \n", + "\n", + "The returned output of the algorithm is a subset of the feature space of a specified size. E.g., a subset of 5 features from a 10-dimensional feature space ($k = 5, \\; d = 10$).\n", + "
\n", + "\n", + "**Initialization:** \n", + "- $ X_0 = \\emptyset, \\; k = 0 $\n", + "\n", + "We initialize the algorithm with an empty set (\"null set\") so that the $k = 0$ (where $k$ is the size of the subset)\n", + "
\n", + "\n", + "**Step 1 (Inclusion):** \n", + "
\n", + "- $ x^+ \\; arg \\; max \\; J(x_k + x), \\quad where \\; x \u2208 Y - X_k $ \n", + "- $ X_k+1 = X_k + x^+ $ \n", + "- $ k = k + 1 $ \n", + "- $ Go \\; to \\; Step 1 $ \n", + "\n", + "We go through the ***feature space*** and look for the feature $x^+$ which maximizes our criterion if we add it to the ***feature subset*** (where $J()$ is the criterion function). We repeat this process until we reach the ***Termination*** criterion.\n", + "
\n", + "\n", + "**Termination:** \n", + "- stop when *k* equals the number of desired features\n", + "\n", + "We add features to the new feature subset $X_k$ until we reach the number of specified features for our final subset. E.g., if our desired number of features is 5 and we start with the \"null set\" (*Initialization*), we would add features to the subset until it contains 5 features.\n", + "\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "\n", + "##SFS Code\n", + "\n", + "[[back to top]](#sections)\n" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "# Sebastian Raschka\n", + "# last updated: 03/29/2014 \n", + "# Sequential Forward Selection (SBS)\n", + "\n", + "\n", + "def seq_forw_select(features, max_k, criterion_func, print_steps=False):\n", + " \"\"\"\n", + " Implementation of a Sequential Forward Selection algorithm.\n", + " \n", + " Keyword Arguments:\n", + " features (list): The feature space as a list of features.\n", + " max_k: Termination criterion; the size of the returned feature subset.\n", + " criterion_func (function): Function that is used to evaluate the\n", + " performance of the feature subset.\n", + " print_steps (bool): Prints the algorithm procedure if True.\n", + " \n", + " Returns the selected feature subset, a list of features of length max_k.\n", + "\n", + " \"\"\"\n", + " \n", + " # Initialization\n", + " feat_sub = []\n", + " k = 0\n", + " d = len(features)\n", + " if max_k > d:\n", + " max_k = d\n", + " \n", + " while True:\n", + " \n", + " # Inclusion step\n", + " if print_steps:\n", + " print('\\nInclusion from feature space', features)\n", + " crit_func_max = criterion_func(feat_sub + [features[0]])\n", + " best_feat = features[0]\n", + " for x in features[1:]:\n", + " crit_func_eval = criterion_func(feat_sub + [x])\n", + " if crit_func_eval > crit_func_max:\n", + " crit_func_max = crit_func_eval\n", + " best_feat = x\n", + " feat_sub.append(best_feat)\n", + " if print_steps:\n", + " print('include: {} -> feature subset: {}'.format(best_feat, feat_sub))\n", + " features.remove(best_feat)\n", + " \n", + " # Termination condition\n", + " k = len(feat_sub)\n", + " if k == max_k:\n", + " break\n", + " \n", + " return feat_sub" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 2 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "### Example SFS:\n", + "\n", + "[[back to top]](#sections)\n", + "\n", + "In this example, we take a look at the individual steps of the [***SFS***](#sfs) algorithmn to select a ***feature subset*** consisting of 3 features out of a ***feature space*** of size 10. \n", + "The input feature space consists of 10 integers: 6, 3, 1, 6, 8, 2, 3, 7, 9, 1, \n", + "and our criterion is to find a subset of size 3 in this ***feature space*** that maximizes the integer sum in this ***feature subset***." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "def example_seq_forw_select():\n", + " ex_features = [6, 3, 1, 6, 8, 2, 3, 7, 9, 1]\n", + " res_forw = seq_forw_select(features=ex_features, max_k=3,\\\n", + " criterion_func=simple_crit_func, print_steps=True) \n", + " return res_forw\n", + " \n", + "# Run example\n", + "res_forw = example_seq_forw_select()\n", + "print('\\nRESULT: [6, 3, 1, 6, 8, 2, 3, 7, 9, 1] ->', res_forw)\n" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "\n", + "Inclusion from feature space [6, 3, 1, 6, 8, 2, 3, 7, 9, 1]\n", + "include: 9 -> feature subset: [9]\n", + "\n", + "Inclusion from feature space [6, 3, 1, 6, 8, 2, 3, 7, 1]\n", + "include: 8 -> feature subset: [9, 8]\n", + "\n", + "Inclusion from feature space [6, 3, 1, 6, 2, 3, 7, 1]\n", + "include: 7 -> feature subset: [9, 8, 7]\n", + "\n", + "RESULT: [6, 3, 1, 6, 8, 2, 3, 7, 9, 1] -> [9, 8, 7]\n" + ] + } + ], + "prompt_number": 3 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "###Result:\n", + "The returned result is definitely what we would expect: the 3 highest values (note that we defined 3 as the number of desired features for our subset) in the input feature list, since our ***criterion*** is to select the numerical values (= features) that yield the maximum mathematical sum in the feature subset." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "## 3. Sequential Backward Selection (SBS)\n", + "\n", + "[[back to top]](#sections)\n", + "\n", + "The ***Sequential Backward Selection (SBS)*** algorithm is very similar to the [***SFS***](#sfs), which we have just seen in the section above. The only difference is that we start with the complete feature set instead of the \"null set\" and remove features sequentially until we reach the number of desired features *k*. \n", + "Note that features are never added back once they were removed, which (similar to [***SFS***](#sfs)) is one of the biggest downsides of this algorithm.
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
\n", + "**Input:** \n", + "the set of all features, \n", + "- $Y = {y_1, y_2, ..., y_d}$ \n", + "\n", + "The ***SBS*** algorithm takes the whole feature set as input, if our feature space consists of, e.g. 10, if our feature space consists of 10 dimensions ($d = 10$).\n", + "
\n", + "\n", + "**Output:** \n", + "a subset of features, \n", + "- $ X_k = {x_j \\;| \\;j = 1, 2, ..., k; x_j \u2208 Y}, \\quad where \\; k = (0, 1, 2, ..., d)$ \n", + "\n", + "The returned output of the algorithm is a subset of the feature space of a specified size. E.g., a subset of 5 features from a 10-dimensional feature space ($k = 5, d = 10$).\n", + "
\n", + "\n", + "**Initialization:** \n", + "- $X_0 = Y, \\; k = d$\n", + "\n", + "We initialize the algorithm with the given feature set so that the $k = d$ (where $k$ has the size of the feature set $d$)\n", + "
\n", + "\n", + "**Step 1 (Exclusion):** \n", + "
\n", + "- $x^- = arg \\; max \\; J(x_k - x), where x \u2208 X_k$ \n", + "- $X_k-1 = X_k - x^-$ \n", + "- $k = k - 1$ \n", + "- $Go \\; to \\; Step 1$ \n", + "\n", + "We go through the ***feature subset*** and look for the feature $x^-$ which minimizes our criterion if we remove it from the ***feature subset*** (where $J()$ is the criterion function). We repeat this process until we reach the ***Termination*** criterion.\n", + "
\n", + "\n", + "**Termination:** \n", + "- stop when $k$ equals the number of desired features\n", + "\n", + "We remove features from the feature subset $X_k$ until we reach the number of specified features for our final subset. E.g., if our desired number of features is 5 and we start with the entire feature space (*Initialization*), we would remove features from the subset until it contains 5 features.\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "\n", + "##SBS Code\n", + "\n", + "[[back to top]](#sections)\n" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "# Sebastian Raschka\n", + "# last updated: 03/29/2014 \n", + "# Sequential Backward Selection (SBS)\n", + "\n", + "from copy import deepcopy\n", + "\n", + "def seq_backw_select(features, max_k, criterion_func, print_steps=False):\n", + " \"\"\"\n", + " Implementation of a Sequential Backward Selection algorithm.\n", + " \n", + " Keyword Arguments:\n", + " features (list): The feature space as a list of features.\n", + " max_k: Termination criterion; the size of the returned feature subset.\n", + " criterion_func (function): Function that is used to evaluate the\n", + " performance of the feature subset.\n", + " print_steps (bool): Prints the algorithm procedure if True.\n", + " \n", + " Returns the selected feature subset, a list of features of length max_k.\n", + "\n", + " \"\"\"\n", + " # Initialization\n", + " feat_sub = deepcopy(features)\n", + " k = len(feat_sub)\n", + " i = 0\n", + "\n", + " while True:\n", + " \n", + " # Exclusion step\n", + " if print_steps:\n", + " print('\\nExclusion from feature subset', feat_sub)\n", + " worst_feat = len(feat_sub)-1\n", + " worst_feat_val = feat_sub[worst_feat]\n", + " crit_func_max = criterion_func(feat_sub[:-1]) \n", + "\n", + " for i in reversed(range(0,len(feat_sub)-1)):\n", + " crit_func_eval = criterion_func(feat_sub[:i] + feat_sub[i+1:])\n", + " if crit_func_eval > crit_func_max:\n", + " worst_feat, crit_func_max = i, crit_func_eval\n", + " worst_feat_val = feat_sub[worst_feat]\n", + " del feat_sub[worst_feat]\n", + " if print_steps:\n", + " print('exclude: {} -> feature subset: {}'.format(worst_feat_val, feat_sub))\n", + " \n", + " # Termination condition\n", + " k = len(feat_sub)\n", + " if k == max_k:\n", + " break\n", + " \n", + " return feat_sub" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 4 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "### Example SBS:\n", + "\n", + "\n", + "[[back to top]](#sections)\n", + "\n", + "\n", + "Like we did for the [***SFS example***](#example_sfs) above, we take a look at the individual steps of the [***SBS***](#sbs) algorithmn to select a ***feature subset*** consisting of 3 features out of a ***feature space*** of size 10. \n", + "The input feature space consists of 10 integers: 6, 3, 1, 6, 8, 2, 3, 7, 9, 1, \n", + "and our criterion is to find a subset of size 3 in this ***feature space*** that maximizes the integer sum in this ***feature subset***." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "def example_seq_backw_select():\n", + " ex_features = [6,3,1,6,8,2,3,7,9,1]\n", + " res_backw = seq_backw_select(features=ex_features, max_k=3,\\\n", + " criterion_func=simple_crit_func, print_steps=True) \n", + " return (res_backw)\n", + " \n", + "# Run example\n", + "res_backw = example_seq_backw_select()\n", + "print('\\nRESULT: [6, 3, 1, 6, 8, 2, 3, 7, 9, 1] ->', res_backw)\n" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "\n", + "Exclusion from feature subset [6, 3, 1, 6, 8, 2, 3, 7, 9, 1]\n", + "exclude: 1 -> feature subset: [6, 3, 1, 6, 8, 2, 3, 7, 9]\n", + "\n", + "Exclusion from feature subset [6, 3, 1, 6, 8, 2, 3, 7, 9]\n", + "exclude: 1 -> feature subset: [6, 3, 6, 8, 2, 3, 7, 9]\n", + "\n", + "Exclusion from feature subset [6, 3, 6, 8, 2, 3, 7, 9]\n", + "exclude: 2 -> feature subset: [6, 3, 6, 8, 3, 7, 9]\n", + "\n", + "Exclusion from feature subset [6, 3, 6, 8, 3, 7, 9]\n", + "exclude: 3 -> feature subset: [6, 3, 6, 8, 7, 9]\n", + "\n", + "Exclusion from feature subset [6, 3, 6, 8, 7, 9]\n", + "exclude: 3 -> feature subset: [6, 6, 8, 7, 9]\n", + "\n", + "Exclusion from feature subset [6, 6, 8, 7, 9]\n", + "exclude: 6 -> feature subset: [6, 8, 7, 9]\n", + "\n", + "Exclusion from feature subset [6, 8, 7, 9]\n", + "exclude: 6 -> feature subset: [8, 7, 9]\n", + "\n", + "RESULT: [6, 3, 1, 6, 8, 2, 3, 7, 9, 1] -> [8, 7, 9]\n" + ] + } + ], + "prompt_number": 5 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### Result:\n", + "The returned ***feature subset*** is similar to the result of the [***SFS***](#example_sfs) algorithm that we have seen above, which is what we would expect: the 3 highest values (note that we defined 3 as the number of desired features for our subset) in the input feature list, since our ***criterion*** is to select the feanumerical values (= features) that yield the maximum mathematical sum in the feature subset." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "## 4. \"Plus L take away R\" (+L -R)\n", + "\n", + "\n", + "[[back to top]](#sections)\n", + "\n", + "\n", + "The ***\"Plus L take away R\" (+L -R)*** is basically a combination of [***SFS***](#sfs) and [***SBS***](#sbs). It append features to the ***feature subset*** *L-times*, and afterwards it removes features *R-times* until we reach our desired size for the ***feature subset***.\n", + "\n", + "**Variant 1: L > R** \n", + "If *L > R*, the algorithm starts with an empty ***feature subset*** and adds *L* features to it from the ***feature space***. Then it goes over to the next step 2, where it removes *R* features from the ***feature subset***, after which it goes back to step 1 to add *L* features again. \n", + "Those steps are repeated until the ***feature subset** reaches the desired size *k*. \n", + "
\n", + "**Variant 2: R > L** \n", + "Else, if *R > L*, the algorithms starts with the whole ***feature space*** as ***feature subset***. It remove s*R* features from it before it adds back *L* features from those features that were just removed. \n", + "Those steps are repeated until the ***feature subset** reaches the desired size *k*. \n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
\n", + "**Input:** \n", + "the set of all features, \n", + "- $Y = {y_1, y_2, ..., y_d}$ \n", + "\n", + "The ***+L -R*** algorithm takes the whole feature set as input, if our feature space consists of, e.g. 10, if our feature space consists of 10 dimensions ($d = 10$).\n", + "
\n", + "\n", + "**Output:** \n", + "a subset of features, \n", + "- $X_k = {x_j \\; | \\; j = 1, 2, ..., k; x_j \u2208 Y}, \\quad where \\; k = (0, 1, 2, ..., d)$ \n", + "\n", + "The returned output of the algorithm is a subset of the feature space of a specified size. E.g., a subset of 5 features from a 10-dimensional feature space ($k = 5, \\; d = 10$).\n", + "
\n", + "\n", + "**Initialization:** \n", + "- $X_0 = Y, \\; k = d$\n", + "\n", + "We initialize the algorithm with the given feature set so that the $k = d$ (where $k$ has the size of the feature set $d$)\n", + "
\n", + "\n", + "**Step 1 (Inclusion):** \n", + "
\n", + "- *repeat L-times:* \n", + " - $x^+ = arg \\; max \\; J(x_k + x), \\quad where \\; x \u2208 Y - X_k $ \n", + " - $X_k+1 = X_k + x^+$ \n", + " - $k = k + 1$ \n", + "- $Go \\; to\\; Step 2$ \n", + "
\n", + "**Step 2 (Exclusion):** \n", + "
\n", + "- *repeat R-times:* \n", + " - $x^- = arg max J(x_k - x), \\quad where \\; x \u2208 X_k$ \n", + " - $X_k-1 = X_k - x^-$ \n", + " - $k = k - 1$ \n", + "- $Go \\; to \\; Step 1$ \n", + "\n", + "In step 1, we go *L-times* through the ***feature space*** and look for the feature $x^+$ which maximizes our criterion if we add it to the ***feature subset*** (where $J()$ is the criterion function). Then we go over to step 2. \n", + "In step 2, we go *R-times* through the ***feature subset*** and look for the feature $x^-$ which minimizes our criterion if we remove it from the ***feature subset*** (where $J()$ is the criterion function). Then we go back to step 1. \n", + "Note that this order of steps only applies if $L > R$, in the opposite case ($R > L$), we have to start with the ***Exlusion*** on the whole ***feature space***, followed by inclusion of removed features.\n", + "
\n", + "\n", + "**Termination:** \n", + "- stop when $k$ equals the number of desired features\n", + "\n", + "We add and remove features from the feature subset $X_k$ until we reach the number of specified features for our final subset. E.g., if our desired number of features is 5 and we start with the entire feature space (*Initialization*), we would remove features from the subset until it contains 5 features.\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "\n", + "##+L -R Code\n", + "\n", + "[[back to top]](#sections)\n" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "# Sebastian Raschka\n", + "# last updated: 03/29/2014 \n", + "# \"Plus L take away R\" (+L -R)\n", + "\n", + "from copy import deepcopy\n", + "\n", + "def plus_L_minus_R(features, max_k, criterion_func, L=3, R=2, print_steps=False):\n", + " \"\"\"\n", + " Implementation of a \"Plus l take away r\" algorithm.\n", + " \n", + " Keyword Arguments:\n", + " features (list): The feature space as a list of features.\n", + " max_k: Termination criterion; the size of the returned feature subset.\n", + " criterion_func (function): Function that is used to evaluate the\n", + " performance of the feature subset.\n", + " L (int): Number of features added per iteration.\n", + " R (int): Number of features removed per iteration.\n", + " print_steps (bool): Prints the algorithm procedure if True.\n", + " \n", + " Returns the selected feature subset, a list of features of length max_k.\n", + "\n", + " \"\"\"\n", + " assert(L != R), 'L must be != R to avoid an infinite loop'\n", + " \n", + " ############################\n", + " ### +L -R for case L > R ###\n", + " ############################\n", + " \n", + " if L > R:\n", + " feat_sub = []\n", + " k = 0\n", + " \n", + " # Initialization\n", + " while True:\n", + " \n", + " # +L (Inclusion)\n", + " if print_steps:\n", + " print('\\nInclusion from features', features)\n", + " for i in range(L):\n", + " if len(features) > 0:\n", + " crit_func_max = criterion_func(feat_sub + [features[0]])\n", + " best_feat = features[0]\n", + " if len(features) > 1:\n", + " for x in features[1:]:\n", + " crit_func_eval = criterion_func(feat_sub + [x])\n", + " if crit_func_eval > crit_func_max:\n", + " crit_func_max = crit_func_eval\n", + " best_feat = x\n", + " features.remove(best_feat)\n", + " feat_sub.append(best_feat)\n", + " if print_steps:\n", + " print('include: {} -> feature_subset: {}'.format(best_feat, feat_sub))\n", + " \n", + " # -R (Exclusion)\n", + " if print_steps:\n", + " print('\\nExclusion from feature_subset', feat_sub)\n", + " for i in range(R):\n", + " if len(features) + len(feat_sub) > max_k:\n", + " worst_feat = len(feat_sub)-1\n", + " worst_feat_val = feat_sub[worst_feat]\n", + " crit_func_max = criterion_func(feat_sub[:-1]) \n", + "\n", + " for j in reversed(range(0,len(feat_sub)-1)):\n", + " crit_func_eval = criterion_func(feat_sub[:j] + feat_sub[j+1:])\n", + " if crit_func_eval > crit_func_max:\n", + " worst_feat, crit_func_max = j, crit_func_eval\n", + " worst_feat_val = feat_sub[worst_feat]\n", + " del feat_sub[worst_feat]\n", + " if print_steps:\n", + " print('exclude: {} -> feature subset: {}'.format(worst_feat_val, feat_sub))\n", + " \n", + " \n", + " # Termination condition\n", + " k = len(feat_sub)\n", + " if k == max_k:\n", + " break\n", + " \n", + " return feat_sub\n", + " \n", + " ############################\n", + " ### +L -R for case L < R ###\n", + " ############################\n", + "\n", + " else:\n", + " # Initialization\n", + " feat_sub = deepcopy(features)\n", + " k = len(feat_sub)\n", + " i = 0\n", + " count = 0\n", + " while True:\n", + " count += 1\n", + " # Exclusion step\n", + " removed_feats = []\n", + " if print_steps:\n", + " print('\\nExclusion from feature subset', feat_sub)\n", + " for i in range(R):\n", + " if len(feat_sub) > max_k:\n", + " worst_feat = len(feat_sub)-1\n", + " worst_feat_val = feat_sub[worst_feat]\n", + " crit_func_max = criterion_func(feat_sub[:-1]) \n", + "\n", + " for i in reversed(range(0,len(feat_sub)-1)):\n", + " crit_func_eval = criterion_func(feat_sub[:i] + feat_sub[i+1:])\n", + " if crit_func_eval > crit_func_max:\n", + " worst_feat, crit_func_max = i, crit_func_eval\n", + " worst_feat_val = feat_sub[worst_feat]\n", + " removed_feats.append(feat_sub.pop(worst_feat))\n", + " if print_steps:\n", + " print('exclude: {} -> feature subset: {}'.format(removed_feats, feat_sub))\n", + " \n", + " # +L (Inclusion)\n", + " included_feats = []\n", + " if len(feat_sub) != max_k:\n", + " for i in range(L):\n", + " if len(removed_feats) > 0:\n", + " crit_func_max = criterion_func(feat_sub + [removed_feats[0]])\n", + " best_feat = removed_feats[0]\n", + " if len(removed_feats) > 1:\n", + " for x in removed_feats[1:]:\n", + " crit_func_eval = criterion_func(feat_sub + [x])\n", + " if crit_func_eval > crit_func_max:\n", + " crit_func_max = crit_func_eval\n", + " best_feat = x\n", + " removed_feats.remove(best_feat)\n", + " feat_sub.append(best_feat)\n", + " included_feats.append(best_feat)\n", + " if print_steps:\n", + " print('\\nInclusion from removed features', removed_feats)\n", + " print('include: {} -> feature_subset: {}'.format(included_feats, feat_sub))\n", + " \n", + " # Termination condition\n", + " k = len(feat_sub)\n", + " if k == max_k:\n", + " break\n", + " if count >= 30:\n", + " break\n", + " return feat_sub\n", + " " + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 6 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "### Example +L -R:\n", + "\n", + "[[back to top]](#sections)\n", + "\n", + "Like we did for the [***SFS***](#example_sfs) example above, let's have look at the individual steps of the ***+L -R*** algorithmn to select a ***feature subset*** consisting of 3 features out of a ***feature space*** of size 10. \n", + "Again, the input feature space consists of the 10 integers: 6, 3, 1, 6, 8, 2, 3, 7, 9, 1, \n", + "and our criterion is to find a subset of size 3 in this ***feature space*** that maximizes the integer sum in this ***feature subset***." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "#### Example 1: L > R\n", + "\n", + "[[back to top]](#sections)\n" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "def example_plus_L_minus_R():\n", + " ex_features = [6, 3, 1, 6, 8, 2, 3, 7, 9, 1]\n", + " res_plmr = plus_L_minus_R(features=ex_features, max_k=3,\\\n", + " criterion_func=simple_crit_func, L=3, R=2, print_steps=True) \n", + " \n", + " return (res_plmr)\n", + " \n", + "# Run example\n", + "res = example_plus_L_minus_R()\n", + "print('\\nRESULT: [6, 3, 1, 6, 8, 2, 3, 7, 9, 1] ->', res)" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "\n", + "Inclusion from features [6, 3, 1, 6, 8, 2, 3, 7, 9, 1]\n", + "include: 9 -> feature_subset: [9]\n", + "include: 8 -> feature_subset: [9, 8]\n", + "include: 7 -> feature_subset: [9, 8, 7]\n", + "\n", + "Exclusion from feature_subset [9, 8, 7]\n", + "exclude: 7 -> feature subset: [9, 8]\n", + "exclude: 8 -> feature subset: [9]\n", + "\n", + "Inclusion from features [6, 3, 1, 6, 2, 3, 1]\n", + "include: 6 -> feature_subset: [9, 6]\n", + "include: 6 -> feature_subset: [9, 6, 6]\n", + "include: 3 -> feature_subset: [9, 6, 6, 3]\n", + "\n", + "Exclusion from feature_subset [9, 6, 6, 3]\n", + "exclude: 3 -> feature subset: [9, 6, 6]\n", + "exclude: 6 -> feature subset: [9, 6]\n", + "\n", + "Inclusion from features [1, 2, 3, 1]\n", + "include: 3 -> feature_subset: [9, 6, 3]\n", + "include: 2 -> feature_subset: [9, 6, 3, 2]\n", + "include: 1 -> feature_subset: [9, 6, 3, 2, 1]\n", + "\n", + "Exclusion from feature_subset [9, 6, 3, 2, 1]\n", + "exclude: 1 -> feature subset: [9, 6, 3, 2]\n", + "exclude: 2 -> feature subset: [9, 6, 3]\n", + "\n", + "RESULT: [6, 3, 1, 6, 8, 2, 3, 7, 9, 1] -> [9, 6, 3]\n" + ] + } + ], + "prompt_number": 7 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "#### Example 2: R > L\n", + "\n", + "[[back to top]](#sections)\n" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "def example_plus_L_minus_R():\n", + " ex_features = [6, 3, 1, 6, 8, 2, 3, 7, 9, 1]\n", + " res_plmr = plus_L_minus_R(features=ex_features, max_k=3,\\\n", + " criterion_func=simple_crit_func, L=2, R=3, print_steps=True) \n", + " \n", + " return (res_plmr)\n", + " \n", + "# Run example\n", + "res = example_plus_L_minus_R()\n", + "print('\\nRESULT: [6, 3, 1, 6, 8, 2, 3, 7, 9, 1] ->', res)" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "\n", + "Exclusion from feature subset [6, 3, 1, 6, 8, 2, 3, 7, 9, 1]\n", + "exclude: [1, 1, 2] -> feature subset: [6, 3, 6, 8, 3, 7, 9]\n", + "\n", + "Inclusion from removed features [1]\n", + "include: [2, 1] -> feature_subset: [6, 3, 6, 8, 3, 7, 9, 2, 1]\n", + "\n", + "Exclusion from feature subset [6, 3, 6, 8, 3, 7, 9, 2, 1]\n", + "exclude: [1, 2, 3] -> feature subset: [6, 3, 6, 8, 7, 9]\n", + "\n", + "Inclusion from removed features [1]\n", + "include: [3, 2] -> feature_subset: [6, 3, 6, 8, 7, 9, 3, 2]\n", + "\n", + "Exclusion from feature subset [6, 3, 6, 8, 7, 9, 3, 2]\n", + "exclude: [2, 3, 3] -> feature subset: [6, 6, 8, 7, 9]\n", + "\n", + "Inclusion from removed features [2]\n", + "include: [3, 3] -> feature_subset: [6, 6, 8, 7, 9, 3, 3]\n", + "\n", + "Exclusion from feature subset [6, 6, 8, 7, 9, 3, 3]\n", + "exclude: [3, 3, 6] -> feature subset: [6, 8, 7, 9]\n", + "\n", + "Inclusion from removed features [3]\n", + "include: [6, 3] -> feature_subset: [6, 8, 7, 9, 6, 3]\n", + "\n", + "Exclusion from feature subset [6, 8, 7, 9, 6, 3]\n", + "exclude: [3, 6, 6] -> feature subset: [8, 7, 9]\n", + "\n", + "RESULT: [6, 3, 1, 6, 8, 2, 3, 7, 9, 1] -> [8, 7, 9]\n" + ] + } + ], + "prompt_number": 8 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### Result:\n", + "The returned ***feature subset*** really is suboptimal in this particular example for L > R ([Example 1: L > R](#example_pLmR1)). This is due to the fact that we add multiple features to our ***feature subset*** and we also remove multiple features from it; we never add back any of the removed features to the investigated ***feature space***. \n", + "\n", + "**Modifying the \"Plus L take away R\" algorithm** \n", + "This algorithm can be tweaked by adding back *r - 1* features to the ***feature subset*** after each ***Exclusion*** step to be assessed by the ***criterion function*** for inclusion in the next iteration. This is a decision that has to be made by considering the particular application, since it decreases the computational performance of the algorithm, but improves the performance of the resulting ***feature subset*** as a classifier." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "## 5. Sequential Floating Forward Selection (SFFS)\n", + "\n", + "[[back to top]](#sections)\n", + "\n", + "The ***Sequential Floating Forward Selection (SFFS)*** algorithm can be considered as extension of the simpler [***SFS***](#sfs) algorithm, which we have seen in the very beginning. In constrast to ***SFS***, the ***SFFS*** algorithm **can** remove features once they were included, so that a larger number of feature subset combinations can be sampled. It is important to emphasize that the removal of included features is **conditional**, which makes it different from the [***+L -R***](#pLmR) algorithm. The ***Conditional Exclusion*** in ***SFFS*** only occurs if the resultin feature subset is assessed as \"better\" by the ***criterion function*** after removal of a particular feature." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
\n", + "**Input:** \n", + "the set of all features, \n", + "- $Y = {y_1, y_2, ..., y_d}$ \n", + "\n", + "The ***SFFS*** algorithm takes the whole feature set as input, if our feature space consists of, e.g. 10, if our feature space consists of 10 dimensions ($d = 10$).\n", + "
\n", + "\n", + "**Output:** \n", + "a subset of features, \n", + "- $X_k = {x_j \\; | \\; j = 1, 2, ..., k; x_j \u2208 Y}, \\quad where \\; k = (0, 1, 2, ..., d)$ \n", + "\n", + "The returned output of the algorithm is a subset of the feature space of a specified size. E.g., a subset of 5 features from a 10-dimensional feature space ($k = 5, \\; d = 10$).\n", + "
\n", + "\n", + "**Initialization:** \n", + "- $X_0 = \\emptyset, \\quad k = 0$\n", + "\n", + "We initialize the algorithm with an empty set (\"null set\") so that the $k = 0$ (where $k$ is the size of the subset)\n", + "
\n", + "\n", + "**Step 1 (Inclusion):** \n", + "
\n", + "- $x^+ = arg\\; max \\;J(x_k + x), \\quad where \\; x \u2208 Y - X_k$ \n", + "- $X_k+1 = X_k + x^+$ \n", + "- $k = k + 1$ \n", + "- $Go \\; to \\; Step 2 $ \n", + "
\n", + "**Step 2 (Conditional Exclusion):** \n", + "
\n", + "- $x^- = arg \\; max \\; J(x_k - x),\\quad where \\; x \u2208 X_k$ \n", + "- $if\\; J(x_k - x) > J(x_k - x):$ \n", + " - $X_k-1 = X_k - x^-$ \n", + " - $k = k - 1$ \n", + "- $Go \\; to \\; Step 1$ \n", + "\n", + "In step 1, we include the feature from the ***feature space*** that leads to the best performance increase for our ***feature subset*** (assessed by the ***criterion function***). Then, we go over to step 2 \n", + "In step 2, we only remove a feature if the resulting subset would gain an increase in performance. We go back to step 1. \n", + "Steps 1 and 2 are reapeated until the **Termination** criterion is reached.\n", + "
\n", + "\n", + "**Termination:** \n", + "- stop when $k$ equals the number of desired features\n", + "\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "\n", + "##SFFS Code\n", + "\n", + "[[back to top]](#sections)\n" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "# Sebastian Raschka\n", + "# last updated: 03/29/2014 \n", + "# Sequential Floating Forward Selection (SFFS)\n", + "\n", + "def seq_float_forw_select(features, max_k, criterion_func, print_steps=False):\n", + " \"\"\"\n", + " Implementation of Sequential Floating Forward Selection.\n", + " \n", + " Keyword Arguments:\n", + " features (list): The feature space as a list of features.\n", + " max_k: Termination criterion; the size of the returned feature subset.\n", + " criterion_func (function): Function that is used to evaluate the\n", + " performance of the feature subset.\n", + " print_steps (bool): Prints the algorithm procedure if True.\n", + " \n", + " Returns the selected feature subset, a list of features of length max_k.\n", + "\n", + " \"\"\"\n", + "\n", + " # Initialization\n", + " feat_sub = []\n", + " k = 0\n", + " \n", + " while True:\n", + " \n", + " # Step 1: Inclusion\n", + " if print_steps:\n", + " print('\\nInclusion from features', features)\n", + " if len(features) > 0:\n", + " crit_func_max = criterion_func(feat_sub + [features[0]])\n", + " best_feat = features[0]\n", + " if len(features) > 1:\n", + " for x in features[1:]:\n", + " crit_func_eval = criterion_func(feat_sub + [x])\n", + " if crit_func_eval > crit_func_max:\n", + " crit_func_max = crit_func_eval\n", + " best_feat = x\n", + " features.remove(best_feat)\n", + " feat_sub.append(best_feat)\n", + " if print_steps:\n", + " print('include: {} -> feature_subset: {}'.format(best_feat, feat_sub))\n", + " \n", + " # Step 2: Conditional Exclusion\n", + " worst_feat_val = None\n", + " if len(features) + len(feat_sub) > max_k:\n", + " crit_func_max = criterion_func(feat_sub) \n", + " for i in reversed(range(0,len(feat_sub))):\n", + " crit_func_eval = criterion_func(feat_sub[:i] + feat_sub[i+1:])\n", + " if crit_func_eval > crit_func_max:\n", + " worst_feat, crit_func_max = i, crit_func_eval\n", + " worst_feat_val = feat_sub[worst_feat]\n", + " if worst_feat_val: \n", + " del feat_sub[worst_feat]\n", + " if print_steps:\n", + " print('exclude: {} -> feature subset: {}'.format(worst_feat_val, feat_sub))\n", + " \n", + " \n", + " # Termination condition\n", + " k = len(feat_sub)\n", + " if k == max_k:\n", + " break\n", + " \n", + " return feat_sub" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 9 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "### Example SFFS:\n", + "\n", + "[[back to top]](#sections)\n", + "\n", + "Since the ***Exclusion*** step in the ***Sequential Floating Forward*** algorithm is ***conditional*** - a feature is only removed if the criterion function asses a better performance after removal - we would never exclude any feature using our [`simple_criterion_func()`](#crit_func\"), which returns the integer sum of a feature set. Thus, let us define another simple criterion function that we use for testing our ***SFFS*** algorithm.\n", + "\n", + "\n", + "Just as we did for the previous examples above, let's take a look at the individual steps of the ***SFFS*** algorithmn to select a ***feature subset*** consisting of 3 features out of a ***feature space*** of size 10. \n", + "Also here, the input feature space consists of the 10 integers: 6, 3, 1, 6, 8, 2, 3, 7, 9, 1, \n", + "and our criterion is to find a subset of size 3 in this ***feature space*** that maximizes the integer sum in this ***feature subset***." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "#### A simple criterion function with a random parameter\n", + "\n", + "[[back to top]](#sections)\n", + "\n", + "The criterion function we define below is also calculates the sum of a subset similar to the [`simple_criterion_func()`](#crit_func\") we used before. However, here we add a random integer ranging from -15 to 15 to the returned sum. Therefore, in some occasions, our criterion function can return a larger sum for a smaller subset - after we removed a feature from the subset after the ***Inclusion*** step - in order to trigger the ***Conditional Exclusion*** step." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "from random import randint\n", + "\n", + "def simple_rand_crit_func(feat_sub):\n", + " \"\"\" \n", + " Returns sum of numerical values of an input list plus \n", + " a random integer ranging from -15 to 15. \n", + " \n", + " \"\"\" \n", + " return sum(feat_sub) + randint(-15,15)\n", + "\n", + "# Example:\n", + "simple_rand_crit_func([1,2,4])" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "pyout", + "prompt_number": 10, + "text": [ + "19" + ] + } + ], + "prompt_number": 10 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "def example_seq_float_forw_select():\n", + " ex_features = [6,3,1,6,8,2,3,7,9,1]\n", + " res_seq_flforw = seq_float_forw_select(features=ex_features, max_k=3,\\\n", + " criterion_func=simple_rand_crit_func, print_steps=True) \n", + " return res_seq_flforw\n", + " \n", + "# Run example\n", + "res_seq_flforw = example_seq_float_forw_select()\n", + "print('\\nRESULT: [6, 3, 1, 6, 8, 2, 3, 7, 9, 1] ->', res_seq_flforw)" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "\n", + "Inclusion from features [6, 3, 1, 6, 8, 2, 3, 7, 9, 1]\n", + "include: 7 -> feature_subset: [7]\n", + "exclude: None -> feature subset: [7]\n", + "\n", + "Inclusion from features [6, 3, 1, 6, 8, 2, 3, 9, 1]\n", + "include: 1 -> feature_subset: [7, 1]\n", + "exclude: 7 -> feature subset: [1]\n", + "\n", + "Inclusion from features [6, 3, 6, 8, 2, 3, 9, 1]\n", + "include: 3 -> feature_subset: [1, 3]\n", + "exclude: None -> feature subset: [1, 3]\n", + "\n", + "Inclusion from features [6, 6, 8, 2, 3, 9, 1]\n", + "include: 9 -> feature_subset: [1, 3, 9]\n", + "exclude: None -> feature subset: [1, 3, 9]\n", + "\n", + "RESULT: [6, 3, 1, 6, 8, 2, 3, 7, 9, 1] -> [1, 3, 9]\n" + ] + } + ], + "prompt_number": 11 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "## 6. Sequential Floating Backward Selection (SFBS)\n", + "\n", + "[[back to top]](#sections)\n", + "\n", + "Just as in the [***SFFS***](#sffs) algorithm, we have a conditional step: Here, we start with the whole feature subset and exclude features sequentially. Only if adding one of the previously excluded features back to a new ***feature subset*** improves the performance (assessed by the criterion function), we add it back in the ***Conditional Inclusion*** step." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
\n", + "**Input:** \n", + "the set of all features, \n", + "- $Y = {y_1, y_2, ..., y_d}$ \n", + "\n", + "The ***SFBS*** algorithm takes the whole feature set as input, if our feature space consists of, e.g. 10, if our feature space consists of 10 dimensions ($d = 10$).\n", + "
\n", + "\n", + "**Output:** \n", + "a subset of features, \n", + "- $X_k = {x_j \\; | \\; j = 1, 2, ..., k; x_j \u2208 Y}, \\quad where \\; k = (0, 1, 2, ..., d)$ \n", + "\n", + "The returned output of the algorithm is a subset of the feature space of a specified size. E.g., a subset of 5 features from a 10-dimensional feature space ($k = 5,\\; d = 10$).\n", + "
\n", + "\n", + "**Initialization:** \n", + "- $X_0 = Y, \\quad k = d$\n", + "\n", + "We initialize the algorithm with the given feature set so that the $k = d$ (where $k$ has the size of the feature set $d$)\n", + "
\n", + "\n", + "**Step 1 (Exclusion):** \n", + "
\n", + "- $x^- = arg max J(x_k - x), \\quad where \\; x \u2208 X_k$ \n", + "- $X_k-1 = X_k - x^-$ \n", + "- $k = k - 1$ \n", + "- $Go \\;to \\;Step 2$ \n", + "\n", + "
\n", + "**Step 2 (Conditional Inclusion):** \n", + "
\n", + "- $x^+ = arg max J(x_k + x), \\quad where \\; x \u2208 Y - X_k$ \n", + "- $if J(x_k + x) > J(x_k + x):$ \n", + " - $X_k+1 = X_k + x^+$ \n", + " - $k = k + 1$ \n", + "- $Go\\; to\\; Step 1$ \n", + "\n", + "In step 1, we exclude the feature from the ***feature space*** that yields the best performance increase of the ***feature subset*** (assessed by the ***criterion function***). Then, we go over to step 2. \n", + "In step 2, we only include one of the removed features if the resulting subset would gain an increase in performance. We go back to step 1. \n", + "Steps 1 and 2 are reapeated until the **Termination** criterion is reached.\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "##SFBS Code\n", + "\n", + "[[back to top]](#sections)\n" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "# Sebastian Raschka\n", + "# last updated: 03/29/2014 \n", + "# Sequential Floating Backward Selection (SFBS)\n", + "\n", + "from copy import deepcopy\n", + "\n", + "def seq_float_backw_select(features, max_k, criterion_func, print_steps=False):\n", + " \"\"\"\n", + " Implementation of Sequential Floating Backward Selection.\n", + " \n", + " Keyword Arguments:\n", + " features (list): The feature space as a list of features.\n", + " max_k: Termination criterion; the size of the returned feature subset.\n", + " criterion_func (function): Function that is used to evaluate the\n", + " performance of the feature subset.\n", + " print_steps (bool): Prints the algorithm procedure if True.\n", + " \n", + " Returns the selected feature subset, a list of features of length max_k.\n", + "\n", + " \"\"\"\n", + "\n", + " # Initialization\n", + " feat_sub = deepcopy(features)\n", + " k = len(feat_sub)\n", + " i = 0\n", + " excluded_features = []\n", + " \n", + " while True:\n", + " \n", + " # Termination condition\n", + " k = len(feat_sub)\n", + " if k == max_k:\n", + " break\n", + " \n", + " # Step 1: Exclusion\n", + " if print_steps:\n", + " print('\\nExclusion from feature subset', feat_sub)\n", + " worst_feat = len(feat_sub)-1\n", + " worst_feat_val = feat_sub[worst_feat]\n", + " crit_func_max = criterion_func(feat_sub[:-1]) \n", + "\n", + " for i in reversed(range(0,len(feat_sub)-1)):\n", + " crit_func_eval = criterion_func(feat_sub[:i] + feat_sub[i+1:])\n", + " if crit_func_eval > crit_func_max:\n", + " worst_feat, crit_func_max = i, crit_func_eval\n", + " worst_feat_val = feat_sub[worst_feat]\n", + " excluded_features.append(feat_sub[worst_feat])\n", + " del feat_sub[worst_feat]\n", + " if print_steps:\n", + " print('exclude: {} -> feature subset: {}'.format(worst_feat_val, feat_sub))\n", + " \n", + " # Step 2: Conditional Inclusion\n", + " if len(excluded_features) > 0 and len(feat_sub) != max_k:\n", + " best_feat = None\n", + " best_feat_val = None\n", + " crit_func_max = criterion_func(feat_sub)\n", + " for i in range(len(excluded_features)):\n", + " crit_func_eval = criterion_func(feat_sub + [excluded_features[i]])\n", + " if crit_func_eval > crit_func_max:\n", + " best_feat, crit_func_max = i, crit_func_eval\n", + " best_feat_val = excluded_features[best_feat]\n", + " if best_feat:\n", + " feat_sub.append(excluded_features[best_feat])\n", + " del excluded_features[best_feat]\n", + " if print_steps:\n", + " print('include: {} -> feature subset: {}'.\\\n", + " format(best_feat_val, feat_sub))\n", + " \n", + " return feat_sub" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 16 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "### Example SFBS:\n", + "\n", + "[[back to top]](#sections)\n", + "\n", + "Note that the ***Inclusion*** step in the ***Sequential Floating Backward Selection*** algorithm is ***conditional*** - a feature is only added back if the criterion function asses a better performance after its inclusion in the ***feature subset***. \n", + "Therefore, we have to be a little bit careful about the ***criterion function***: If we used our [`simple_criterion_func()`](#crit_func\"), which returns the integer sum of a subset, we would trigger an infinite loop and never reach the termination criterion - assuming that our feature space consists of positive integers. The reason is that the [`simple_criterion_func()`](#crit_func\") would always return a larger sum if we include a positive integer (our feature) to the ***feature subset***, thus let us use our [second simple criterion function](#random_crit_func), which contains a (pseudo) random integer between -15 and 15 to the returned sum. \n", + "In order to reduce the number of iterations, we set the number of desired features for the ***feature subset*** (via the argument `max_k`) to 7." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "def example_seq_float_backw_select():\n", + " ex_features = [6,3,1,6,8,2,3,7,9,1]\n", + " res_seq_flbackw = seq_float_backw_select(features=ex_features, max_k=7,\\\n", + " criterion_func=simple_rand_crit_func, print_steps=True) \n", + " return res_seq_flbackw\n", + " \n", + "# Run example\n", + "res_seq_flbackw = example_seq_float_backw_select()\n", + "print('\\nRESULT: [6, 3, 1, 6, 8, 2, 3, 7, 9, 1] ->', res_seq_flbackw)" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "\n", + "Exclusion from feature subset [6, 3, 1, 6, 8, 2, 3, 7, 9, 1]\n", + "exclude: 2 -> feature subset: [6, 3, 1, 6, 8, 3, 7, 9, 1]\n", + "include: None -> feature subset: [6, 3, 1, 6, 8, 3, 7, 9, 1]\n", + "\n", + "Exclusion from feature subset [6, 3, 1, 6, 8, 3, 7, 9, 1]\n", + "exclude: 3 -> feature subset: [6, 3, 1, 6, 8, 7, 9, 1]\n", + "include: 3 -> feature subset: [6, 3, 1, 6, 8, 7, 9, 1, 3]\n", + "\n", + "Exclusion from feature subset [6, 3, 1, 6, 8, 7, 9, 1, 3]\n", + "exclude: 3 -> feature subset: [6, 3, 1, 6, 8, 7, 9, 1]\n", + "include: None -> feature subset: [6, 3, 1, 6, 8, 7, 9, 1]\n", + "\n", + "Exclusion from feature subset [6, 3, 1, 6, 8, 7, 9, 1]\n", + "exclude: 6 -> feature subset: [3, 1, 6, 8, 7, 9, 1]\n", + "\n", + "RESULT: [6, 3, 1, 6, 8, 2, 3, 7, 9, 1] -> [3, 1, 6, 8, 7, 9, 1]\n" + ] + } + ], + "prompt_number": 18 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## 7. Genetic Algorithm (GA)\n", + "\n", + "*to be continued ...*" + ] + } + ], + "metadata": {} + } + ] +} \ No newline at end of file diff --git a/benchmarks/cython_least_squares.ipynb b/benchmarks/cython_least_squares.ipynb new file mode 100644 index 0000000..c71c137 --- /dev/null +++ b/benchmarks/cython_least_squares.ipynb @@ -0,0 +1,1274 @@ +{ + "metadata": { + "name": "", + "signature": "sha256:5d04280c23460c2481423dabf313f3ad28f40fb4ad915967d83e3a08231c8b3d" + }, + "nbformat": 3, + "nbformat_minor": 0, + "worksheets": [ + { + "cells": [ + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[Sebastian Raschka](http://www.sebastianraschka.com) \n", + "last updated: 05/09/2014\n", + "\n", + "- [Link to this IPython Notebook on GitHub](https://github.com/rasbt/python_reference/blob/master/benchmarks/cython_least_squares.ipynb) \n", + "- [Link to the GitHub repository](https://github.com/rasbt/python_reference)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### The code in this notebook was executed in Python 3.4.0" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "I am really looking forward to your comments and suggestions to improve and \n", + "extend this little collection! Just send me a quick note \n", + "via Twitter: [@rasbt](https://twitter.com/rasbt) \n", + "or Email: [bluewoodtree@gmail.com](mailto:bluewoodtree@gmail.com)\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "# Implementing the least squares fit method for linear regression and speeding it up via Cython" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "
\n", + "
\n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#Sections" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "- [Introduction](#introduction)\n", + "- [Least squares fit implementations](#implementations)\n", + " - [1. The matrix approach in (C)Python and NumPy](#matrix_approach)\n", + " - [2. The classic approach in (C)Python, Cython, and Numba](#classic_approach)\n", + " - [3. Using the `numpy.linalg.lstsq` function](#numpy_func)\n", + " - [4. Using the `scipy.stats.linregress` function](#scipy_func)\n", + "- [Generating sample data and benchmarking](#sample_data)\n", + " - [Performance growth rates: (C)Python vs. Cython vs. Numba](#performance1)\n", + " - [Performance growth rates: NumPy and SciPy library functions](#performance2)\n", + "- [Bonus: How to use Cython without the IPython magic](#cython_bonus)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + " \n", + "(Note that this chart just reflects my rather objective thoughts after experimenting with Cython, and it is not based on real numbers or benchmarks.)\n", + "
\n", + "
\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "# Introduction" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Linear regression via the least squares method is the simplest approach to performing a regression analysis of a dependent and a explanatory variable. The objective is to find the best-fitting straight line through a set of points that minimizes the sum of the squared offsets from the line. \n", + "The offsets come in 2 different flavors: perpendicular and vertical - with respect to the line. \n", + "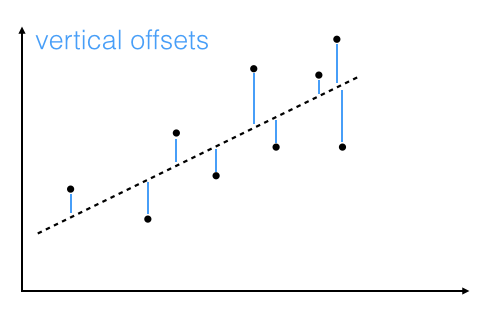 \n", + "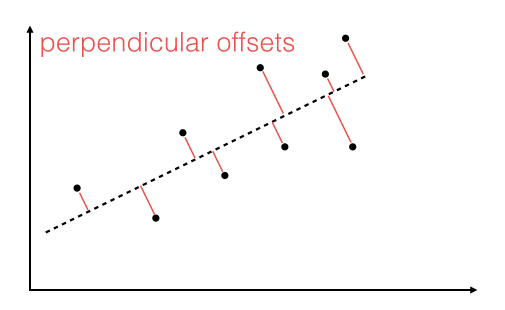 \n", + "\n", + "As Michael Burger summarizes it nicely in his article \"[Problems of Linear Least Square Regression - And Approaches to Handle Them](http://www.arsa-conf.com/archive/?vid=1&aid=2&kid=60101-220)\": \"the perpendicular offset method delivers a more precise result but is are more complicated to handle. Therefore normally the vertical offsets are used.\" \n", + "Here, we will also use the method of computing the vertical offsets.\n", + "\n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "In more mathematical terms, our goal is to compute the best fit to *n* points $(x_i, y_i)$ with $i=1,2,...n,$ via linear equation of the form \n", + "$f(x) = a\\cdot x + b$. \n", + "We further have to assume that the y-component is functionally dependent on the x-component. \n", + "In a cartesian coordinate system, $b$ is the intercept of the straight line with the y-axis, and $a$ is the slope of this line." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "In order to obtain the parameters for the linear regression line for a set of multiple points, we can re-write the problem as matrix equation \n", + "$\\pmb X \\; \\pmb a = \\pmb y$" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "$\\Rightarrow\\Bigg[ \\begin{array}{cc}\n", + "x_1 & 1 \\\\\n", + "... & 1 \\\\\n", + "x_n & 1 \\end{array} \\Bigg]$\n", + "$\\bigg[ \\begin{array}{c}\n", + "a \\\\\n", + "b \\end{array} \\bigg]$\n", + "$=\\Bigg[ \\begin{array}{c}\n", + "y_1 \\\\\n", + "... \\\\\n", + "y_n \\end{array} \\Bigg]$" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "With a little bit of calculus, we can rearrange the term in order to obtain the parameter vector $\\pmb a = [a\\;b]^T$" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "$\\Rightarrow \\pmb a = (\\pmb X^T \\; \\pmb X)^{-1} \\pmb X^T \\; \\pmb y$" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "The more classic approach to obtain the slope parameter $a$ and y-axis intercept $b$ would be:" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "$a = \\frac{S_{x,y}}{\\sigma_{x}^{2}}\\quad$ (slope)\n", + "\n", + "\n", + "$b = \\bar{y} - a\\bar{x}\\quad$ (y-axis intercept)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "where \n", + "\n", + "\n", + "$S_{xy} = \\sum_{i=1}^{n} (x_i - \\bar{x})(y_i - \\bar{y})\\quad$ (covariance)\n", + "\n", + "\n", + "$\\sigma{_x}^{2} = \\sum_{i=1}^{n} (x_i - \\bar{x})^2\\quad$ (variance)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "# Least squares fit implementations" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### 1. The matrix approach in (C)Python and NumPy" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "First, let us implement the equation:" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "$\\pmb a = (\\pmb X^T \\; \\pmb X)^{-1} \\pmb X^T \\; \\pmb y$" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "which I will refer to as the \"matrix approach\"." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### Matrix approach implemented in NumPy and (C)Python" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "import numpy as np\n", + "\n", + "def py_matrix_lstsqr(x, y):\n", + " \"\"\" Computes the least-squares solution to a linear matrix equation. \"\"\"\n", + " X = np.vstack([x, np.ones(len(x))]).T\n", + " return (np.linalg.inv(X.T.dot(X)).dot(X.T)).dot(y)" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 9 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### 2. The classic approach in (C)Python, Cython, and Numba" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Next, we will calculate the parameters separately, using standard library functions in Python only, which I will call the \"classic approach\"." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "$a = \\frac{S_{x,y}}{\\sigma_{x}^{2}}\\quad$ (slope)\n", + "\n", + "\n", + "$b = \\bar{y} - a\\bar{x}\\quad$ (y-axis intercept)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Note: I refrained from using list comprehensions and convenience functions such as `zip()` in\n", + "order to maximize the performance for the Cython compilation into C code in the later sections.\n", + " " + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### Implemented in (C)Python" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "def py_classic_lstsqr(x, y):\n", + " \"\"\" Computes the least-squares solution to a linear matrix equation. \"\"\"\n", + " len_x = len(x)\n", + " x_avg = sum(x)/len_x\n", + " y_avg = sum(y)/len(y)\n", + " var_x = 0\n", + " cov_xy = 0\n", + " for i in range(len_x):\n", + " temp = (x[i] - x_avg)\n", + " var_x += temp**2\n", + " cov_xy += temp*(y[i] - y_avg)\n", + " slope = cov_xy / var_x\n", + " y_interc = y_avg - slope*x_avg\n", + " return (slope, y_interc) " + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 1 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### Implemented in Cython" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Maybe we can speed things up a little bit via [Cython's C-extensions for Python](http://cython.org). Cython is basically a hybrid between C and Python and can be pictured as compiled Python code with type declarations. \n", + "Since we are working in an IPython notebook here, we can make use of the very convenient *IPython magic*: It will take care of the conversion to C code, the compilation, and eventually the loading of the function. " + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%load_ext cythonmagic" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 2 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%%cython\n", + "def cy_classic_lstsqr(x, y):\n", + " \"\"\" Computes the least-squares solution to a linear matrix equation. \"\"\"\n", + " cdef double x_avg, y_avg, var_x, cov_xy,\\\n", + " slope, y_interc, x_i, y_i\n", + " cdef int len_x\n", + " len_x = len(x)\n", + " x_avg = sum(x)/len_x\n", + " y_avg = sum(y)/len(y)\n", + " var_x = 0\n", + " cov_xy = 0\n", + " for i in range(len_x):\n", + " temp = (x[i] - x_avg)\n", + " var_x += temp**2\n", + " cov_xy += temp*(y[i] - y_avg)\n", + " slope = cov_xy / var_x\n", + " y_interc = y_avg - slope*x_avg\n", + " return (slope, y_interc)" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 3 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### Implemented in Numba" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Like we did with Cython before, we will use the minimalist approach to Numba and see how the two - Cython and Numba - compare against each other. \n", + "\n", + "Numba is using the [LLVM compiler infrastructure](http://llvm.org) for compiling Python code to machine code. Its strength is to work with NumPy arrays to speed-up the code. If you want to read more about Numba, please see refer to the original [website and documentation](http://numba.pydata.org/numba-doc/0.13/index.html)." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "from numba import jit\n", + "\n", + "@jit\n", + "def numba_classic_lstsqr(x, y):\n", + " \"\"\" Computes the least-squares solution to a linear matrix equation. \"\"\"\n", + " len_x = len(x)\n", + " x_avg = sum(x)/len_x\n", + " y_avg = sum(y)/len(y)\n", + " var_x = 0\n", + " cov_xy = 0\n", + " for i in range(len_x):\n", + " temp = (x[i] - x_avg)\n", + " var_x += temp**2\n", + " cov_xy += temp*(y[i] - y_avg)\n", + " slope = cov_xy / var_x\n", + " y_interc = y_avg - slope*x_avg\n", + " return (slope, y_interc)" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 4 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### 3. Using the `numpy.linalg.lstsq` function" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "For our convenience, `numpy` has a function that can computes the least squares solution of a linear matrix equation. For more information, please refer to the [documentation](http://docs.scipy.org/doc/numpy/reference/generated/numpy.linalg.lstsq.html)." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "def numpy_lstsqr(x, y):\n", + " \"\"\" Computes the least-squares solution to a linear matrix equation. \"\"\"\n", + " X = np.vstack([x, np.ones(len(x))]).T\n", + " return np.linalg.lstsq(X,y)[0]" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 5 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### 4. Using the `scipy.stats.linregress` function" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Also scipy has a least squares function, `scipy.stats.linregress()`, which returns a tuple of 5 different attributes, where the 1st value in the tuple is the slope, and the second value is the y-axis intercept, respectively. \n", + "The documentation for this function can be found [here](http://docs.scipy.org/doc/scipy-0.13.0/reference/generated/scipy.stats.linregress.html)." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "import scipy.stats\n", + "\n", + "def scipy_lstsqr(x,y):\n", + " \"\"\" Computes the least-squares solution to a linear matrix equation. \"\"\"\n", + " return scipy.stats.linregress(x, y)[0:2]" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 6 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "# Generating sample data and benchmarking" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "#### Visualization" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "To check how our dataset is distributed, and how the least squares regression line looks like, we will plot the results in a scatter plot. \n", + "Note that we are only using our \"matrix approach\" to visualize the results - for simplicity. We expect all 4 approaches to produce similar results, which we will confirm after visualizing the data." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%matplotlib inline" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 7 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "from matplotlib import pyplot as plt\n", + "import random\n", + "\n", + "random.seed(12345)\n", + "\n", + "x = [x_i*random.randrange(8,12)/10 for x_i in range(500)]\n", + "y = [y_i*random.randrange(8,12)/10 for y_i in range(100,600)]\n", + "\n", + "slope, intercept = py_matrix_lstsqr(x, y)\n", + "\n", + "line_x = [round(min(x)) - 1, round(max(x)) + 1]\n", + "line_y = [slope*x_i + intercept for x_i in line_x]\n", + "\n", + "plt.figure(figsize=(8,8))\n", + "plt.scatter(x,y)\n", + "plt.plot(line_x, line_y, color='red', lw='2')\n", + "\n", + "plt.ylabel('y')\n", + "plt.xlabel('x')\n", + "plt.title('Linear regression via least squares fit')\n", + "\n", + "ftext = 'y = ax + b = {:.3f} + {:.3f}x'\\\n", + " .format(slope, intercept)\n", + "plt.figtext(.15,.8, ftext, fontsize=11, ha='left')\n", + "\n", + "plt.show()" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "display_data", + "png": "iVBORw0KGgoAAAANSUhEUgAAAfoAAAH4CAYAAACi3S9CAAAABHNCSVQICAgIfAhkiAAAAAlwSFlz\nAAALEgAACxIB0t1+/AAAIABJREFUeJzs3XdUVMfbwPHv0lm6giAgooBRbNi7GCv2EnvXWOPPnhg1\nGksSxRhjoiYaE0SNsb4xEY0Fe+zGmijGLkZARRERFljYnfeP1Y0ELJRlAedzDkf2lpln7q48e++d\nO6MQQggkSZIkSSqSTIwdgCRJkiRJhiMTvSRJkiQVYTLRS5IkSVIRJhO9JEmSJBVhMtFLkiRJUhEm\nE70kSZIkFWEy0UtGd+jQIcqXL2/sMAqtSpUq8fvvv+drnSNHjuTTTz/N0b7e3t7s3bs3jyN6s/3y\nyy+UKlUKe3t7zp07Z5TPhFRwKeRz9FJ+8fb2JiQkhGbNmhk7FMmIypQpQ0hICE2bNjVI+QcOHKBf\nv378888/Bim/IPLx8eGrr76iffv2mdbNnDmT69ev8+OPPxohMqkgkGf0Ur5RKBQoFApjh6Gn0Wjy\nZJvXJYRAfq+Wnsmrz5YQgtu3b+Pv758n5UlFj0z0ktEdOHCAUqVK6V97e3uzYMECqlatiqOjIz17\n9iQ1NVW/ftu2bQQEBODk5ESDBg3466+/9OuCg4Px9fXF3t6eihUr8uuvv+rXrVy5kgYNGjBhwgSc\nnZ2ZNWtWplhmzpxJ165d6devHw4ODqxatYrHjx/z7rvv4u7ujqenJ9OnT0er1QKg1WqZOHEiLi4u\nlC1bliVLlmBiYqJf36RJE6ZNm0aDBg2wsbHh5s2b/P3337Ro0YLixYtTvnx5Nm3apK9/+/btVKxY\nEXt7ezw9PVmwYAEADx48oF27djg5OVG8eHEaN26c4Xg9uxSemprKuHHj8PDwwMPDg/Hjx6NWq/XH\n2dPTky+//BJXV1fc3d1ZuXJllu/Jhg0bqFWrVoZlCxcupGPHjgAMHDiQ6dOnA/Do0SPatWtHiRIl\nKFasGO3btycqKirLcv9LCKF/z5ydnenRowePHj3Sr+/WrRslS5bE0dGRwMBAIiIiXnisvvzyS1Qq\nFa1btyY6Oho7Ozvs7e25e/dupnpfdJwB5s+fr3+vV6xYgYmJCTdu3AB072dISIh+25UrV9KoUSP9\n67Fjx+Ll5YWDgwM1a9bk8OHD+nXZ/Wxdu3aNwMBAHB0dcXFxoWfPnpnakZqaip2dHRqNhqpVq+Ln\n5wf8+5nYuXMnc+fOZcOGDdjZ2VGtWrXXel+kIkZIUj7x9vYWe/fuzbR8//79wtPTM8N2derUETEx\nMSIuLk5UqFBBLFu2TAghxJkzZ0SJEiXEyZMnhVarFatWrRLe3t5CrVYLIYTYtGmTiImJEUIIsWHD\nBmFjYyPu3r0rhBAiNDRUmJmZiSVLlgiNRiOSk5MzxTJjxgxhbm4utmzZIoQQIjk5WXTq1EmMGDFC\nqFQqcf/+fVG7dm3x3XffCSGEWLp0qfD39xdRUVHi0aNHolmzZsLExERoNBohhBCBgYGidOnSIiIi\nQmg0GhEfHy88PT3FypUrhUajEWfPnhXOzs7i0qVLQggh3NzcxOHDh4UQQsTHx4szZ84IIYSYPHmy\nGDFihEhPTxfp6en6bf57XKdPny7q1asnYmNjRWxsrKhfv76YPn26/jibmZmJGTNmiPT0dLF9+3ah\nVCpFfHx8puOgUqmEnZ2duHr1qn5ZzZo1xYYNG4QQQgwcOFBf7sOHD8XmzZtFcnKyePLkiejWrZvo\n1KlT1h+C/8T71VdfiXr16omoqCihVqvF8OHDRa9evfTbhoaGisTERKFWq8W4ceNEQECAft2LjtWB\nAwcyfJ6y8qJ9d+zYIVxdXcXFixdFUlKS6NWrl1AoFOL69etCCCGaNGkiQkJCMsTXsGFD/es1a9aI\nuLg4odFoxIIFC4Sbm5tITU0VQmT/s9WzZ08xZ84cIYQQqamp4siRIy9sz/Mx/vcYz5w5U/Tr1++l\nx0Mq2uQZvVQgjRkzBjc3N5ycnGjfvj3nzp0DYPny5QwfPpxatWqhUCjo378/lpaWHDt2DICuXbvi\n5uYGQPfu3fHz8+PEiRP6ct3d3Rk1ahQmJiZYWVllWXf9+vXp0KEDAI8fP2bHjh0sXLgQa2trXFxc\nGDduHOvXrwdg48aNjBs3Dnd3dxwdHZkyZUqGy/MKhYKBAwdSoUIFTExM2LlzJ2XKlGHAgAGYmJgQ\nEBBAly5d2LhxIwAWFhZcvHiRhIQEHBwc9GdgFhYWxMTEcOvWLUxNTWnQoEGWsa9du5aPP/4YZ2dn\nnJ2dmTFjRoZ7s+bm5nz88ceYmprSunVrbG1tuXz5cqZyrK2t6dixI+vWrQPg6tWrXL58WX9cAH07\nixUrRufOnbGyssLW1papU6dy8ODBrN/Y//juu+/49NNPcXd3x9zcnBkzZvB///d/+rPagQMHYmNj\no193/vx5njx58tJjJV7j9siL9t24cSODBw/G398fpVKZ5VWfl+nTpw9OTk6YmJgwYcIEUlNTMxzf\n7Hy2LCwsuHXrFlFRUVhYWFC/fv1sxfKMkLeM3ngy0UsF0rNkDbqkk5iYCEBkZCQLFizAyclJ/3Pn\nzh1iYmIAWL16NdWqVdOvu3DhAg8fPtSX9fwtghfx9PTU/x4ZGUlaWholS5bUlzlixAhiY2MBiImJ\nyVDm8/tmVWdkZCQnTpzIEP/atWu5d+8eAD///DPbt2/H29ubJk2acPz4cQA++OADfH19admyJT4+\nPsybNy/L2KOjoyldurT+tZeXF9HR0frXxYsXx8Tk3//2SqVSf2z/q3fv3vpEv3btWn0y/y+VSsXw\n4cPx9vbGwcGBwMBAHj9+/FrJ5datW3Tu3Fl/LPz9/TEzM+PevXtoNBomT56Mr68vDg4OlClTBoVC\nwYMHD156rF7Hi/b97/vp5eX12mUCfPHFF/j7++Po6IiTkxOPHz/WxwvZ+2x9/vnnCCGoXbs2lSpV\nIjQ0NFuxSNIzZsYOQJJex7NOfF5eXnz00UdMnTo10zaRkZEMGzaMffv2Ua9ePRQKBdWqVct0hv2q\nep7fplSpUlhaWvLw4cMMCfKZkiVLZujdnVVP7+fL8/LyIjAwkPDw8Czrr1mzJr/++isajYbFixfT\nvXt3bt++ja2tLV988QVffPEFFy9epGnTptSuXZu33347w/7u7u7cunWLChUqAHD79m3c3d1f2uYX\nad68ObGxsZw/f57169fz1VdfZdmuBQsWcOXKFU6ePEmJEiU4d+4c1atXRwjxyuPt5eVFaGgo9erV\ny7Tuxx9/JCwsjL1791K6dGni4+MpVqyY/v180bF6nQ6fL9q3ZMmS3L59W7/d878D2NjYkJSUpH/9\n/P3/Q4cOMX/+fPbt20fFihUBMsT7/DGDV3+2XF1dWb58OQBHjhyhefPmBAYGUrZs2Ve273kFqQOs\nZBzyjF7KV2q1mpSUFP3P6/Y8fvbHcujQoSxbtoyTJ08ihCApKYnffvuNxMREkpKSUCgUODs7o9Vq\nCQ0N5cKFC9mK779noSVLlqRly5ZMmDCBJ0+eoNVquX79uv4Z5e7du/P1118THR1NfHw88+bNy/SH\n9fky27Vrx5UrV1izZg1paWmkpaXxxx9/8Pfff5OWlsZPP/3E48ePMTU1xc7ODlNTU0DXAfHatWsI\nIbC3t8fU1DTL5NCrVy8+/fRTHjx4wIMHD5g9ezb9+vXL1jF4xtzcnG7duvH+++/z6NEjWrRokaFN\nz9qVmJiItbU1Dg4OxMXFZety94gRI5g6dao+ocbGxhIWFqYv19LSkmLFipGUlJThy93LjpWrqysP\nHz4kISEhyzpftm/37t1ZuXIlly5dQqVSZWpLQEAAmzdvJjk5mWvXrhESEqJ/v588eYKZmRnOzs6o\n1Wpmz579whjg1Z+tTZs2cefOHQAcHR1RKBRZvuev4ubmxq1bt+Tl+zeYTPRSvmrTpg1KpVL/M2vW\nrFc+dvf8+ho1avD999/zv//9j2LFiuHn58fq1asB8Pf3Z+LEidSrVw83NzcuXLhAw4YNsyzndep6\nZvXq1ajVavz9/SlWrBjdunXTn8kNHTqUli1bUqVKFWrUqEHbtm0zJeHny7O1tSU8PJz169fj4eFB\nyZIlmTJlir5n/Jo1ayhTpgwODg4sX76cn376CdD1wG7RogV2dnbUr1+fUaNGERgYmCn+adOmUbNm\nTapUqUKVKlWoWbMm06ZNyzKW19G7d2/27t1Lt27dMrXpWVnjxo0jOTkZZ2dn6tevT+vWrV+7nrFj\nx9KhQwdatmyJvb099erV4+TJkwD079+f0qVL4+HhQaVKlfRXaZ550bEqX748vXr1omzZshQrVizL\nXvcv2jcoKIhx48bRtGlTypUrl2nMh/Hjx2NhYYGrqyuDBg2ib9+++nVBQUEEBQVRrlw5vL29sba2\nznDpP7ufrVOnTlG3bl3s7Ozo2LEjixYtwtvbO8vj+LLj3a1bN0B326ZmzZov3E4qugw6YM7ly5cz\nPBJy48YNPvnkE/r27UuPHj2IjIzE29ubjRs34ujoCMDcuXNZsWIFpqamLFq0iJYtWxoqPEnKczt2\n7GDkyJHcunXL2KFIecTExIRr165l+5K5JBUUBj2jf+uttzh79ixnz57l9OnTKJVKOnfuTHBwMC1a\ntODKlSs0a9aM4OBgACIiItiwYQMRERHs3LmT9957T9/7VpIKopSUFLZv3056ejpRUVHMmjWLLl26\nGDssSZIkvXy7dL9nzx58fX0pVaoUYWFhDBgwAIABAwboBzXZsmULvXr1wtzcHG9vb3x9ffWX8SSp\nIBJCMHPmTIoVK0b16tWpWLEis2fPNnZYUh6Sndmkwi7fet2vX7+eXr16AXDv3j1cXV0BXceZZ48W\nRUdHU7duXf0+np6erz3CliQZg7W1tfwyWsTl5TDIkmQM+ZLo1Wo1W7duzfLZ39fpiPU8X19frl+/\nnucxSpIkSVJB5ePjw7Vr13K0b75cut+xYwc1atTAxcUF0J3FP+tZGhMTQ4kSJQDw8PDI8BzynTt3\n8PDwyFDW9evX9Y/2vIk/M2bMMHoMsv2y7bL9sv2y/fn7k5sT3HxJ9OvWrdNftgfo0KEDq1atAmDV\nqlV06tRJv3z9+vWo1Wpu3rzJ1atXqV27dn6EKEmSJElFksEv3SclJbFnzx6+//57/bLJkyfTvXt3\nQkJC9I/Xge456O7du+uHwfz2229lRxhJkiRJygWDJ3obG5sMYz2DbljIPXv2ZLn91KlTsxzeVNJp\n0qSJsUMwqje5/W9y20G2X7a/ibFDKLQMOmCOISgUCgpZyJIkSZKUK7nJfXIIXEmSJEkqwmSilyRJ\nkqQiTCZ6SZIkSSrCZKKXJEmSpCJMJnpJkiRJKsJkopckSZKkIkwmekmSJEkqwmSilyRJkqQiTCZ6\nSZIkSSrCZKKXJEmSpCJMJnpJkiRJKsJkopckSZKkIkwmekmSJEkqwmSilyRJkqQiTCZ6SZIkSSrC\nZKKXJEmSpCJMJnpJkiRJKsJkopckSZKkIszM2AFIkiRJWYuNjWXHjh2YmJjQtm1bnJycjB2SVAgp\nhBDC2EFkh0KhoJCFLEmSlG03btygdu1AUlLqAOnY2p7nzJkjuLu750v9f/75JzNnzichIYn+/bvQ\nv3/ffKlXylpucp88o5ckSSqAJk78mEePRqDVfgRAauqHTJv2KStWfGvwuv/880/q1GlCSso0wINj\nxz7m8eMERo9+z+B1S3lP3qOXJEkqgKKi7qHV1tC/Tk+vwe3bdw1e78OHDwkMbEVKyhBgAtADlWo1\n8+d/Y/C6AU6cOMHIkWMZO/Z9/v7773yps6iTiV6SJKkAatWqMUrlF8Bj4CFK5Ve0bh1o8HqnTp1N\nQoInYPrcUpN8uWW6f/9+mjZtz7JlJVm8WEmtWo25ePGiwest6mSilyRJKoA+/ngy77zjg6lpCczM\nPBgwoA7jx482eL2XLt1Aq+0P/AB8A/yCiUkPxo4dZvC6P/poHirVV8BkhJhNUtJ4Pv98scHrLepk\nopckSSqAzM3NWb36O1JTVaSkJPHttwsxMTH8n+yGDWtgbb0bCAMOYGLyPs2a+TNx4liD152cnAIU\n178WwpnExGSD1qlSqRg+fCzly9chKKgr169fN2h9xiA740mSJBVgpqamr94olx49esSOHTtQKBSM\nHTuSs2ffY//+NoCCRo0aERa2AYVCYfA4hgzpyaRJE1GpbAAVSuVsBg9eZtA6O3fuw++/W5CSspCr\nVw9Tp04Trlw5T7FixQxab36SiV6SJOkN9s8//1CzZiNUqiqAwMZmKqdOHcLc3BwhBK6urvmS5AHe\ne284arWab78dh5mZGR9/PJ+2bdsapK7k5GQ++mgWu3dvR4gngAVabX3U6gMcOHCALl26GKReY5DP\n0UuSJBnZX3/9RUxMDJUrV6ZkyZL5WnfPnoP5v//zRKOZDYCZ2VR69XrI6tXf5Wsc+UkIQZMmbTlx\nwoLU1J3APcABENjZNWbt2g9p166dkaPMKDe5T96jlyRJMqIxYyZRt24Q3bvPw8+vCrt3787X+v/5\n5y4aTS396/T0WkRGRudrDPnt+vXr/PHHeVJTNwHDgCBgBebmg3Fze0KzZs2MHGHekolekiTJSA4f\nPsyKFT+jUl3k8eO9JCX9TNeuffLlquW6devx9q7C+fOnMDObDyQAj1Eqv6Zly0YGr9+YhBAoFKbo\nUuBXQD9MTafRo4eCP/44iLW1tZEjzFsy0UuSJBnJjRs3UCjqAo5PlzRCpUokMTHRoPXu3buXd9+d\nSGTkIpKSdiNEJAqFM6amJejevQKTJ080aP3G5uPjg79/WSwt3wV2Y2FxET8/d0JCluHg4GDs8PKc\nTPSSJElGUqVKFTSafcDNp0vW4eLijq2trUHr3bhxC8nJE4AmQFU0mp/x8ipPcnIioaFL86WnvzGZ\nmJiwb99WBg1yombN+fTuLThyJBwLCwtjh2YQste9JEmSkQQEBDBv3sd88EFVzMycsLaG7du3GLyX\nu4ODLaamd9Boni25g52dHebm5gattyCxs7Nj6dKFxg4jX8he95IkSUaWkJBAbGwspUqVypezyjt3\n7lC1al0SEjqQnl4Ca+tv+eWXH2nVqpXB65ZyJje5TyZ6SZKkN1B0dDQ//BBCUlIyXbt2platWq/e\nSTIameglSZIkqQiTz9FLkiRJkpQlmeglSZIk6UWEgEWLIC7O2JHkmEz0kiRJkpQVrRZGj4axY6FD\nB93rQkg+XidJkiRJ/5WeDkOHwsqVYGEBkyZBPkwTbAgy0UuSJEnS89Rq6NsXNm0CpRK2bIHmzY0d\nVY4Vzq8nkiRJ0hsvKSmJLl36olQ64uzsxY8//pT7QlNS4J13dEne3h7Cwwt1kgf5eJ0kSZJUSPXo\nMYgtW1JITV0M3MTauiPh4Rtp2LBhzgpMTISOHWHfPiheHHbtgho18jTmnJKP10mSJElvnF27dpGa\nOg9wBmqRkjKY3bv35Kyw+Hho2VKX5N3c4ODBApPkc0smekmSJKlQcnAoBlzWv7a0vEzx4sWyX1Bs\nLDRtCseOgZcXHDoEFSvmXaBGJhN9Ieft7U1ERIRBytZoNIwaNQpfX1/8/PwICQl54bZz586lcuXK\nVKhQgYEDB6JWqwG4desWZmZmVKtWTf/z6NEjAI4ePUqDBg2oWLEiFStWZNKkSXkaf58+ffDw8MDE\nxASVSpVh3fHjx6latSpvvfUWrVq1IjY2Vh/T87F6eHhQ4wXf6lUqFT169MDPz48KFSrw22+/6det\nWbOGKlWqYG5uzjfffJPl/gcOHMDU1PSF6yVJermlSz9HqeyDufk4lMqOeHpeYdCgQdkrJDoaAgPh\n7Fnw89MleV9fwwRsLKKQKYQh50hoaKiYOXPmK7fz9vYWFy5cMEgMq1atEq1atRJCCBEbGys8PT3F\nrVu3Mm23a9cuUaVKFaFSqYQQQgwdOlQEBwcLIYS4efOmcHZ2zrL8CxcuiGvXrgkhhEhNTRUNGzYU\nP/744yvjGjBggDhw4MArt9u/f7+4f/++UCgUIikpSb9co9EIHx8fceTIESGEEJ9++qkYPHhwlmV0\n6tRJLFiwIMt1s2bNEsOGDRNCCHH16lXh5uYmEhMT9W2LiIgQ/fv3F998802mfRMSEkSdOnVE+/bt\nxZIlS17ZFkmSsnbu3Dnx+eefi++++048efIkezvfvClE2bJCgBCVKgkRE2OQGPNCbnLfG3VGP3/+\nfP73v//pX9+7dw83NzdSUlJyXfa9e/do2rQpNWvWpFKlSnz44Yf6dUOHDmXChAn67cqWLcuff/75\n0vKyM03lmjVrqFmzJn5+fnl6drhx40aGDRsGgLOzM506dWLTpk2Ztvvzzz9p1KgR1tbWAAQFBfHT\nT6/u/VqxYkV8fHwAsLCwICAggNu3b79yP4VC8VrHp0mTJri4uGRafvr0aaytralfvz4Aw4cPZ+PG\njZm2u3//PuHh4fTr1y/L8jdu3Mjw4cMB8PX1pWbNmuzYsUPftgoVKmBiYpJlB5oJEyYwadIkihcv\nrl+WnJxM1apVCQsLA2Dfvn1UqFCBpKSkV7ZVMr5z584xYMAIevcewsGDB40dzhujatWqfPDBBwwb\nNgxbW9vX3/HyZWjUCG7c0N2LP3BAd2++CHqjnqMfMmQI/v7+fP755yiVSpYvX06fPn2wsrLKtG23\nbt24du1apuUKhYJjx45haWmZYbmjoyNbt27FxsaGtLQ0goKC2LVrF61atWLx4sXUqVOHLVu2sHjx\nYiZNmkSVKlVeGmtWyeFFYmNjOXXqFPfv36datWo0btyYypUrZ9jm0qVL9O7dO8v9W7Zsybx58zIt\nv337NqVLl9a/9vLy4p9//sm0XY0aNfj+++95+PAhDg4ObNy4kcjISP36hIQEatSogUKhoGfPnrz/\n/vuZyrh//z6bN29m+/btr9Xm7Byf//pvu5ydndFqtcTHx+Po6Khfvnr1alq1apXll4WsynnR8fmv\nHTt28OTJE7p06cLWrVv1X1qsra3ZuHEjLVu2xM3NjSFDhvDLL79gY2OT06ZK+eTs2bM0atSSpKQP\nAGu2bOnBzz+vJCgoyNihFXqJiYkMHPgeO3b8hp2dI4sWBdO9e7fcFfrnn9CiBdy/Dw0bwrZt4OCQ\nNwEXQG9UondycqJDhw6sXr2aIUOG8MMPP7Bv374st83qzPVl0tPTef/99zl27BhCCO7evcu5c+do\n1aoVVlZWbNy4kRo1atCmTRtGjBiRZRnbt2/no48+AiAuLg61Ws2vv/4KwOjRoxk8eHCW+7377rsA\nlChRgrZt23LgwIFMib5ChQqcPXs2W216XW+//TajRo2iZcuWWFlZ0axZM3bv3g2Au7s7UVFRODs7\nExsbS4cOHXByctLHDPDkyRM6dOjA+++/T9WqVbOs45NPPmHz5s2ALsEePnxY/+191apVr/zilBOh\noaFZfgHKjfj4eCZPnsyePbqewUKIDF9a3nrrLWbPnk39+vX5+uuvX3g8pILliy++JSnpQ0D3JVal\nKsGsWV/JRJ8HBg0axbZtaaSmXkSlus7Age9QurQXderUyVmBJ09CUBA8eqRL9r/8AkX8y/QblehB\nlzD79OmDi4sL/v7++kvH/9W1a1euX7+e5bpjx45lugrw5ZdfEh8fz8mTJ7GwsGD48OEZbglcvHgR\nBwcH7t69i0ajwdTUNFO5bdq0oU2bNoAueUVGRvLxxx+/sk3PJwohRJaXtSMiIujTp0+W+7do0YLP\nP/8803IvLy9u3bql74wWGRlJmTJlsixjzJgxjBkzBtBd0q74tMeqhYUFzs7OALi4uNCnTx+OHDmi\nT/QqlYp27doRFBTE+PHjX9jG6dOnM336dAAGDRrEoEGDaNy48Qu3f5nSpUtnuOLw4MEDTExMMpzN\nHz9+nEePHunfj6w8Oz7PLr9HRkbStGnTTNs9/35cuHCBu3fvUrt2bX3d27Zt49GjR0ybNg3Q3Vpw\ndXV9rasDUsGQkqIGnr9sbKfvkGoIT5484eHDh3h4eGBubm6wegqCnTt3kJp6DnAD3EhNHUh4+O6c\nJfrff4e2bXXPy3foABs2QBZXdIucPOgjkK/yIuS3335blCpVSmzdujUPItKZOHGiGD9+vBBCiDt3\n7ghXV1cxa9YsIYQQN27cEF5eXuLatWtiwIABYvLkya8s73U745UuXVoMHTpUCCHE/fv3hYeHR551\nzlu5cqVo1aqV0Gq14v79+y/sjCeEEDFPO7HExcWJ6tWriy1btuhjUqvVQgghkpKSRPPmzcWiRYuE\nEEIkJyeLpk2big8//DBbcQ0cOPC1OuMJIYRWqxUKhULfSU6IfzvjHT58WAghxCeffJKpM97QoUNf\nGdfMmTP1x/7KlSvC1dU1Qz1C6DoOvqyz3cCBAzN01tu8ebOoXr26iIuLExUrVhQ7dux4rXZKxhUe\nHi6UypICfhawQyiVviIkJNQgdS1a9K2wsLAVSqWnKFGitPjrr78MUk9B4e5eTsABoZtGTghr6y45\n68C6c6cQ1ta6Qnr1EuLp36XCIje5741M9GvWrBHe3t55EM2/IiMjRe3atUWlSpVEUFCQ6NOnj5g1\na5ZQq9Widu3aYt26dUIIXbLz9/cXO3fufGl5K1eu1H9ReBlvb28xdepUUaNGDeHr65tlD++c0mg0\nYuTIkcLHx0f4+PiI77//Xr9u2bJl4uOPP9a/rly5sqhYsaIoV66cWLx4sX755s2bRaVKlUTVqlWF\nv7+/+PDDD4VWqxVCCLFkyRJhamoqqlWrJgICAkRAQICYM2fOK+MaOHCgOHjw4Cu369y5s/D09BQm\nJibCw8NDBAUF6dcdPXpUVK5cWfj5+YmWLVuK+/fv69epVCrh4OAgLl++nKnMgIAA/ZeapKQk0a1b\nN+Hr6yveeustERYWpt9u7dq1wtPTU9jY2AgnJyfh6ekpLl26lGVbnr1nN2/eFKVKlRJXr14VQuh6\n7nt5eYmoqKhXtlUyvi1btogaNZqKqlUbix9+WGGQOs6cOfP0C8WNp4kvVHh6vmWQugqKX375RVhb\nlxBmZhNF0rhoAAAgAElEQVSFtXUn4etbJfu96zdvFsLcXJfkhwwRIj3dMMEaUG5y3xs5BO6QIUOo\nUKECEydOzKOoJEmSDC80NJTRo/eTlLT66RKBqakVCQmPUCqVeV7ftWvXOH/+PN7e3i8cTyI/nD59\nmt27d+Po6Ejfvn2z17t+zRoYOBA0Gt10swsXQjaeaiooCvQQuPHx8XTt2pUKFSrg7+/PiRMniIuL\no0WLFpQrV46WLVsSHx+v337u3Ln4+flRvnx5wsPD8zSW6Ohoypcvz/Xr1xk1alSeli1JkmRoZcuW\nBY4BCU+X/I6trZP+0da89NNP66hatT6DB/9I48ZdGDducp7X8bpq1KjB5MmTGTFiRPaS/PLl0L+/\nLslPm1Zok3yu5dFVhRfq37+/CAkJEUIIkZaWJuLj48UHH3wg5s2bJ4QQIjg4WH8v9OLFi6Jq1apC\nrVaLmzdvCh8fH6HRaDKUlw8hS5IkFUharVaMGDFOKJWewsGhubCxcRbh4eF5Xk9KSoqwsrIX8OfT\nWwRxQqksJU6fPp3ndRnMggVCf2P/6QBehVlucp9BL90/fvyYatWqcePGjQzLy5cvz8GDB3F1deXu\n3bs0adKEv//+m7lz52JiYqIfbCYoKIiZM2dSt25d/b5y9jpJkt50586dIyYmhqpVq+Lu7p7n5UdH\nR+PrW43k5Hv6Zfb27Vi1agidOnXK8/rylBDwyScwY4bu9ZIlUASu4BbYS/c3b97ExcWFQYMGUb16\ndYYOHUpSUhL37t3D1dUVAFdXV+7d032YoqOj8fT01O/v6elJVFSUIUOUJEkqdAICAmjdurVBkjzo\n/i7b2loB654uOUda2omCP66DEDBpki7Jm5hAaGiRSPK5ZdDn6NPT0zlz5gxLliyhVq1ajBs3juDg\n4AzbvGo406zWzZw5U/97kyZNaNKkSV6FLEmS9Ma7ceMGtrYOxMb2BYZhYSFYuXLlC8fRKBC0Wl1S\nX7YMzMxg7VrolssR9IzowIEDHDhwIE/KMmii9/T0xNPTk1q1agG6QWjmzp2Lm5sbd+/exc3NjZiY\nGEqUKAGAh4dHhkFC7ty5g4eHR6Zyn0/0kiRJUt7RaDQ0bdqOqKhRwHBgDxYWgwgMbGTs0F4sPR0G\nD4YffwRLS/j5Z93AOIXYf09iZ82aleOyDHrp3s3NjVKlSnHlyhUA9uzZQ8WKFWnfvj2rVq0CdCPA\nPbvn06FDB9avX49arebmzZtcvXpVP4KYJEmSZHhRUVHExSUixBjAEmiLqWkAZ86cMXZoWVOroWdP\nXZK3sYHt2wt9ks9rBh8Cd/HixfTp0we1Wo2Pjw+hoaFoNBq6d+9OSEgI3t7e+pnD/P396d69O/7+\n/piZmfHtt99maxY3SZIkKXecnJxIT08A7gCegIr09GsvnNzJqJKT4Z13YMcO3aQ0O3ZAvXrGjqrA\neSMHzJEkSSpo1Go1oaGh3LwZSb16dejYsaPRYgkOXsAnnyxCo2mHuflhOnSoxZo13+fqxCsxMZF3\n3x3Nrl27cHQsztKln9O6deucB/nkCbRvDwcPgrMzhIdDtWo5L6+Ay03uk4lekiTJyDQaDYGBbTh7\nVqBSNcLGZi1jxvRgzpyZRovp0KFDnD17ljJlytCuXbtcX13t0qUv27drSE2dB/yNUtmXY8f25Gzm\nyUePoHVrOHECSpaEPXvA3z9X8RV0MtFLkiQVYlOmTGHevE0IcRkwBe5hbl6Gx48fGmTUO2OwtnYg\nJeU6oJvN0tx8HHPmePL+++9nr6D796FlSzh/HkqXhr174QWzkBYlBfY5ekmSJOnlQkNXEhy8BCHK\noEvyAC4oFBaoVCpjhpanbGwcgFv61+bmN3FwcMheIVFREBioS/LlysGhQ29Eks8tmeglSZKMRKvV\nMm7cdGAl8BewGrgNjMXHx49ixYoZM7w89eWXc1AqO6JQTMfKqhvu7rfo1avX6xdw8yY0agR//w1V\nqujmli9VKttxpKSkkJycnO39CjOZ6CVJkoxk9Oj3SUhQAWWBHcBSoBqwik2bVhapp4769+/Lzp3r\nmTbNhODghpw5c/j1J6j5+29dkr95E2rXhv374enoqq9Lo9HQv/9wbG0dsbNzomvXfqjV6hy0pPCR\n9+glSZKM4NSpU9Sp0wCtdhz/JvkY4F2aN2/E7t3bjBtgQXHunO6efGwsNG4M27aBnV22i5k7dz6f\nfrodlSoMMMXauitjx9Zi7tycD0STn+Q9ekmSpFwKDw/HxyeA4sW96NNnqEHvjx87dozGjYPQagHe\nB7oB/YH3aNKkJjt3bjFY3YXK8ePw9tu6JB8UpHtOPptJPiEhga+++orly39CpXoPsAOUJCePZu/e\nowYJu6CRiV6SpDfeX3/9RadOfbhxYw5xcQfYvDmeAQNGGqy+GTO+IDl5LrohZrsC1VAo+lOsmAUb\nN67H1NT0FSVkX0JCAt26DaREibJUqdKAkydP5nkdeerAAWjeHOLjoXNn+PVXUCqzVURCQgIBAfWZ\nMuUYt25ZAb/r15maHsPbO/MQ60WRwUfGkyRJKuh27dqFRtMbaANASsq3/PZbOYPVp1KlAMWBhcAX\nwExcXBI4fvyQwUag69SpD0ePOpOauovY2D9o2rQtERGn8fLyMkh9ubJ9u27Eu5QU6NMHVq7UTVST\nTatXr+bu3QqkpGwA7gF1USjOYGvrgLX1JRYs+P1VRRQJ8oxekqQ3np2dHWZmt59bchulMvv3gV/X\n0KE9USonA4eBmiiVD1m8+DODzQ6nVqs5eHAXqanfAX5Ab6AF+/fvN0h9ufLzz9Cpky7JDxsGq1fn\nKMmD7oxerX52TF2BXVhaXmDFisFcvnyWUjnotV8YyUQvSdIbr3fv3ri6XsXSshcKxSyUyo588cWn\nBqtvwIB+fPnlB/j5vU+5clNYsmQm3bsbbkpVMzMzzMzM0Z3VAggUiqjX7/WeX1avhu7dIS0NJkzQ\nTTlrkvM01apVKywsVgH7gNtYWX1Ex47v0LVrVxwdHfMs7IJO9rqXJElCd/b3/fffExsbR1BQiwxT\nhBYFc+bM57PPvkelGoyV1Sl8fP7h1KmDWFlZGTs0naVL4b33dL/PmKH7yYPHC8PCwvjf/yaTkBBP\nmzZt+OGHRSizea+/IJBD4EqSJGWDEIL169ezf/9RvL09GDt2NDY2NsYOK0+Fh4czY8YCUlPVjBzZ\nlyFDBrN161b27z+Ep6cbI0aMKDhtnj8fJk369/fsDov7BpCJXpIkKRsmTZrGN9+EoVINxtLyGD4+\nNzl9+veCc3abA7/99htjx07jyZME6tatzu7dv5Oc/DXggFI5ni++mMDIkcOMHWZGQsDMmTB7tu71\nt9/CSMM97VCYyUQvSZL0mtRqNTY29qSn3wZKAAJb24b89NOHdOjQwdjh5cipU6cIDGyLSrUKKIOp\naVs0mlHA+Kdb7KFixRlcuHDEiFH+hxAwcSIsXKi7Dx8aCv37GzuqAis3uU8+XidJ0hslLS0NUABO\nT5coUChKFNrxzyMiIujRoz8q1SAgCACNpgHw/IA/Ksxy2HPdIDQa3f345cvB3BzWrdM9TpcNycnJ\nLF68hCtXImnYsCYDBgwoUkMG56UC9M5LkiQZno2NDQ0aNOX48aGkpk5EoTiGiclxAgOXGju0bLt0\n6RI1azYiObkREAnsBdKAeigU7yOEGeCIUjmbGTOWGDVWvfR0GDgQfvoJrKxg82bd3PLZkJaWRuPG\nrblwwYmUlCasW/cNJ06cZ+nShYaJuZCTl+4lSXrjJCQkMHz4eA4dOoanpwfff/8llStXNnZY2RIT\nE0OFClV5/DgQmI9uMhwPwBa4yOTJ/+POnYekpKgZMqQXrVq1yvMYhBCo1WosLS1fb4fUVOjZUzfK\nna0tbN0KOXi6Yf/+/XToMIHExNPonhKPx9zckwcPorG3t892eYWBHOtekiTpJdLT0zl+/DiHDh0i\nOTkZe3t71q0L4c6dCI4f313okjzA5MmzePKkJqAFQoB26Ka6PY6p6UQuXrzFjz8uZ9OmlTlO8klJ\nSXTu3AdLS1ucnNwJCQnVrwsJCUWpdESptKNmzSbcu3fvJSUBKhV06KBL8o6OsGdPjpK8rigVJibF\n+TeF2WFiYkFKSkqOyivq5KV7SZKKtKSkJBo1CuLq1UcoFFYUK5bM8eN7cXNzM3ZouXLrVjRabS9g\nBnAdGIeu7wFoNM24fj0813UMGTKGHTu0qNX/oFbfZsyYdpQt6421tTVjxkwjJeU4UI7z56fQpUt/\njhzZlXVBCQnQrh0cOgQuLrB7N1StmuO46tevj7n5CBSKrxHibSwsllK5chWDDR9c2MkzekmSirRP\nPgnm0qVSJCb+yZMnfxAd3Y4xYyYbO6wcu3v3LleuXKFp07oolaFAOGCPbprbJCAdS8vvqVevRq7r\nCg/fTWrqHHQdF6uSnDyE3bv3cvToUdLTuwIVAFPS06fzxx+Hsi4kLk43Oc2hQ+DhAb//nqskD+Dk\n5MTRo3tp0GAXnp696NBBRXj4L7Iz3gvIM3pJkoq0ixevk5LSmmfnNWlpbbh06SPjBpUDQgiGDx/L\n6tU/YmbmiIuLDUFBAWzZUhEhBJ6efty9WxKFwoxatWqzcOE3ua7Tyak4cXERQBlAYGkZgYtLXVxd\nXTE3/xW1WgOYAqdwcsriCsm9e9CiBfz1F5QpA3v36v7NA+XKlePQoe15UlZRJxO9JElFWt26Vdm7\ndy3Jyd0AcywtV1GrVu7OKPObEIIRI0YSEvI7Wu0tUlPtSUmZTalSJ0hKSgDA0tKSuLg40tLSKFGi\nRJ6c3S5bNp+OHXuh0fQATmFpGcn9+7707NmTatVWc+5cfYR4CyF2sGrVjxl3/ucf3Zn8lStQvrzu\nnrzHmzEtbEEje91LklSkpaWl0alTb/btO4BCYU7FiuXYuzesUPXOXrbse0aPnkJ6+lhg+tOlkTg5\nNSAu7o5B646IiODTTz/j55/3oVaPw9z8BsWK7ebcuaOcPHmShw8f0rBhQ/z8/P7d6fp1aNYMIiMh\nIAB27YISJQwaZ1EnB8yRJEl6AXNzc7Zt20hUVBTp6el4eXlhkosZ0fLb4cOH+fDDz0hPHwjsBiYB\nlkAYZcv6Grz+GzdusHlzOGr1NqAOaWkQH9+fdevWMX78+Mw7RETozuRjYqBuXd3c8k5OmbeT8o1M\n9JIkFXkKhQJPT09jh5FtBw8epHXrbiQnFwcaA7fRdYBzwtLyFmvWGHZI27Vr1zF06Iekpgp087nr\npKe7kZiYlHmHM2egVSt48ED36FxYGNjZGTRG6dXkpXtJkoqM9PR0fvjhBy5dukbNmlXp27dvoe6J\n3apVV8LD26DrVT8O3aN0F7GyWsnevdupX7++QesvX74Oly9/CmwFLgNfADdQKody9Ohuqj7fe/7o\nUWjTBh4/1v37f/8H1tYGje9NIi/dS5L0xtNqtbRp05UjR56gUrXExmYx+/cfZ8WK3Pc+N5a0tHTA\nHEgGWgHf4uqaxM6dBwgICDB4/enp6YASXYKfCrTGycmMjRvXZkzy+/bpBsNJSoKuXXXD21pYGDw+\n6fUUnhtVkiRJL3HmzBmOHr2ISrUT+JCkpD2sXbuWu3fvGju0HHvvvb6YmIwFlgN2QCRduwYZNMnH\nxsZy7tw5EhIS+N//BmJjMxzYDwSgVKrZuXMjzZs3/3eHbdt0Z/BJSTBggG6CGpnkCxSZ6CVJKhKS\nkpIwNXVBdwYMYIeZmT2JiYnGDCtXTE1NsbDwAQ4CXwFHWbFitcFuX3733Q94eZUjMLAfHh4+VKpU\ngXnz/ke1asE0aLCWbds2ULt27X932LgROnfWjWH/3nuwYgUUpFnyJEBeupckqYioXr06lpbRmJh8\njVbbBjOzlbi7F8Pb29vYoeVYXFwcpqb+/HtO5ktqqoq0tDQs8vis+dq1a4wfP5WUlD9ISfEFDtC5\nczcePLjDqFEjMu8QGgpDhoBWC5MmQXAwFOL+EEWZPKOXJKlIsLOz48iR3dSqtQ1n55YEBl7kwIHf\nCtY87K9p+/btdO8+iB079qHR/IZu+tl4zMwmUbNmozxP8gCXL1/GwqI68OyRvSZotVbExMRk3njJ\nEhg8WJfkZ8+WSb6Ak73uJUmSXiImJoaDBw+iVCpp1arV60/JmkO6R9omoVJ9hEJxHyurL7G1dSIh\n4QF16jRm06ZQShhg8JnLly9TrVpjkpP/ALyAY9jYtOPBgyisrKz+3TA4GKZM0f2+YAFMmJDnsUiZ\n5Sb3yUQvSZL0AufOnSMwMAgh6iPEPcqU0XL8+F6USqXB6ixXrhZXr84FdB3eFIqpTJig5Ysvgg1W\n5zMLFy5m6tSZWFj4kp5+nY0bV9G2bVvdSiFg+nT47DPd2fuyZTBsWLbKT0xMRKPR4ODgYIDoizY5\nH70kSZIBvPvuOBIS5vDkyWYSEw9z9aoHS5YY9nG9tDQ1YKt/LYQtKSlqg9b5zPjxo7ly5Rzbt3/J\nrVuXMib58eN1Sd7UFH78MVtJXqPR0L//MJycSuDi4kGrVp1RqVQGaoX0XzLRS5IkZeH+/fvcvn0b\nqPd0iYKUlLpERkYbtN533+2FldUgdNPPrkOp/Ir+/XsatM7nlSpVigYNGvw7t7tGA0OHwtdf6x6b\n27QJ+vTJVplffbWYn3++THr6PdLS4vj9dzMmTfrYANHn3JEjR+jYsQ9t2/Zk165dxg4nT8lEL0mS\n9B+DB4/A3b0scXGJwGdAOnAPpTKUJk0MNxrd+vUb+eyzYDSaWExNu1Kp0tds3bo+4yNt+SktDfr2\nhZAQ3Sh3YWG6x+myaf/+E6hUQ9GNBWBBSsp7/P77yTwPN6eOHDlCy5adCQtryPbtLenSZRDbtm0z\ndlh5RiZ6SZKk58ycOYvQ0M1oNNfQaq8CEYASMzNvxo3rTteuXQ1S782bNxk8eBQpKQdJS3uARhNK\nTEwUjRs3Nkh9r5SSohvlbv163Xj1O3fqxrHPAV/fUlhY/A7o7jGbmh6ibNlSeRhs7nz55XeoVDOA\nkcBgVKqFBAd/a+yw8kzhe+5EkiTJgBYu/AZoD7g9XfIHYE5CwmOs83Ds9vT0dL7+egknTpynYkUf\n/P3fwty8NsnJz4aWfQeVahT37t3DI7/ncU9Kgk6ddHPIOznpppmtVSvHxc2YMYXffnube/caAdZY\nW19j0aKDeRdvLmm1gn8HWgIwQ6PRGiucPCcTvSRJEnD+/Hl69RpGQkIC8DsQDzgC27C1dcnTJA/Q\nrdsAwsPvolL1JCwsjLJlfyE9/R7wECgO/AmkULx48dcqT6vVcvHiRdLS0qhUqVLOn7V//BjatoUj\nR3RzyO/eDVWq5Kysp5ycnPjzz2Ps37+f9PR0AgMDC1TP+9GjB7FrV2+Sk5WAJUrlRCZM+NLYYeUZ\n+XidJElvvEePHlG2bEXi4+cC+4BjwGOgNHCR0NBvGDhwYJ7VFxUVhY9PFVJTzwG9gb8AFf7+Vbh1\n6y5mZlVJSztJSMgSevXq8cryUlJSaNGiE2fPXsHExIqSJa04fHgXLi4upKen8+TJExwdHV89k9+D\nBxAUBKdPg6cn7N0L5crlRZMLvPDwcObMWUJ6uoZx4wbTtes7xg4pA/kcvSRJUg6lpKTQvXsftm27\nhRCngRTgfWAlJUq4sGhRMD16vDrZZseNGzeoXLkRKlV9wB1YCDzEyqoxn3zyLuXKlaNSpUqULVv2\ntcqbNesz5s07RXLyJsAUc/OJdO78CB8fDz7/fAEKhSleXj7s2fMrZcqUybqQmBho0QIuXgQfH91l\n+0I8fHBRIxO9JElSDnXrNoCwsFuo1beBvwFL4CEWFmWIjr752pfOX8fOnTuZOnUeycnJJCQ8Ijr6\nIXAC8Hm6xVwmTHjEggWfZ6vczp378euvzYCBT5ccpnjxXjx8mPi0fD/gc/z9N3Px4onMBURGQvPm\ncO0a+PvrLte7u+eskZJByAFzJEmSckCr1fLrrxtQq8OAOsDbwFQsLOozevToPE3y69ato3373pw9\n+z/+/juYR4/MsLIyRzcFLIAGa+tDlCnjle2yq1f3x9r6ZyANEJiZrScu7i7QBSgHKICJXLp0Co1G\nk3Hnq1ehUSNdkq9WDQ4elEm+iJFn9JIkvbGEEFhbO5Ca+hfgCazD3PwT3nuvNQsXLnz1Pe3XdObM\nGerUeZv09GnAB0+XHqFUqaEkJDxCiGpotTFUqlScAwd+y/Z4+qmpqbRu3ZUTJ85hYmKFp6c9V678\niVZbEd0ZvSVwCBubLiQmxv6744ULujP5e/egfn347TdwdMyTNhcEp06dYvv2HTg42DNgwAAcC3Hb\ncpP7ZK97SZLeWAqFgqlTpzBvXmtUqlGYm5/HzU3B7Nmz8yzJA0ye/Bnp6XWAhOeWPsbOzpHTpw9y\n9OhRbG1tCQwMzNFse5aWluzdG8aVK1dQq9VUqFCBgIAGXLyYBlRDd1a/h6+//vrfnU6fhpYtIS4O\nmjaFLVvA1vYFNRQ+W7dupWfPIaSkDMLcPIIFC5Zy/vwxnJycjB1a/hOFTCEMWZKkAm79+vWiX79h\nYvLkaeLhw4e5KissLEyUKuUvHBxKiu7dB4rNmzcLX98aApYLcBEwQ8AiYWJSXPz888951ILM7ty5\nI6pXbyxMTEyFnV1xsWLFin9XHjokhL29ECBEu3ZCJCfnef33798Xbdt2E2XLVhOdO/cV0dHReV7H\ny3h7VxYQLnQD9QthadlHzJ8/P19jyEu5yX3y0r0kSW+U06dP8+mnC0lMTGbIkB706NE9z8o+c+YM\nDRsGkZy8FiiJQtEcU1NXTE3TSE3VAPOBdZia7mPixEHMmzc3z+p+ESFExqsTe/ZAx46gUkH37rBm\nDZibv7iAHLh79y7e3pVJTW0LDAF+pXTp7fz995mMU94aUPHiXsTF7edZR0eFYgZTpmj57LNP8qX+\nvCY740mSJL2Gv/76i8DAIH79tTZ79rzD4MGTWbFiZZ6Vv2vXLtTq/uimmN2EEE1JTz9LauoFwAdz\n84G4up5k/vyp2UrySUlJDB06Bj+/mjRt2oHLly+/9r4ZknxYmG4wHJUKBg2CtWvzPMkDDBgwktRU\nE2AF0BCYz927ppw+fTrP63qR9u3bYm09EfgHOIyV1XLatAnKt/oLEnmPXpKkN8by5StJShoNjAFA\npSpJcPAkBg8emCflOzg4YGFxmuRk0CWYQHQ93gFm4OU1kmvXsp/s3nmnHwcPWpCS8g3Xrx+nXr2m\nXL587t8Z5v4jPj6egwcPYm5uzttvv60b1W/9et0ENRoNjB4NX30FJq93rqdSqYiOjsbd3R2lUvnK\n7S9fvg5oADVgBWjRaBIxN8CXihdZuvRLtNpxbNlSCxsbexYu/JoGDRrkW/0FiTyjlyTpjaHVasn4\nZ88kT28F9uvXDze3v7Gy6gncBb5EN8qeCkvLxTRqVCfbZapUKvbs2U5KymqgDkKMJS2tJvv3789y\n+8jISN56K4B+/b6hZ885VKlSD9XixdC7ty7JT5mim3L2NZP8zp07KVHCi4CAFri4eLJlS9gr9wkI\nqIxC4QZ0AlYC7+Dqakn16tVft9m5Zm1tzerV3/H48V2io6/k6S2aQidPegnko0IYsiRJBcTZs2eF\nUuksYJmAn4VS6SuWLv0uT+t4/PixWLBggahQobowMXEU4CbAVtSp87ZISEjIdnmpqanCzMxSwIOn\nHcu0wta2sfjll1+y3L51627C1HS2ftsJpnWFvkfaZ59lq+74+HhhY1NcwKGnRZwQSmVxcf/+/Zfu\nd/fuXeHjU0WYm7sJExMX4e1dQTx69ChbdUsZ5Sb3yUv3kiS9MQICAti7dyszZy4gKSmZIUM+ZsCA\nfnlah729PTY2NkRG2qDVRgNWmJlNw97+EnZ2dtkuz8LCgpEjRxMS0gqVaigWFscpWfIxrf4zZey1\na9do3bor165FAqMAmMJc5miO6zb4+msYMyZbdd+4cQNTU3d099kBamNu7sPVq1dfeNsAwNXVlYiI\nP4iIiMDCwoIKFSrk6eOKUjbl4ReOfFEIQ5Yk6SW++WaZKFmynHBxKSOmTZslNBqNsUPKtWHDRgv4\nUn8iDRdEyZLlclyeVqsV338fInr0GCymTJku4uPjM60vU6aSUCgWChgvoIuYwwdCgNCA2NWjV47q\nvX//vrCychRw+Wk7bggrq2Lizp07OW5Lbh08eFD4+VUXTk6eokuXvuLx48dGiyU/5Sb3FbqsKRO9\nJBUdGzZsFEqlj4CTAi4IpbKmmDv3C2OHlWuLFy8RSmUzASkChDA1/VS8/Xb7XJWp0WiESqXKtDwy\nMlK0a9ddKBSWAoRQ8EQsoowQINQgvm3cTKSnp+e43h9+CBXW1s7CwaGpsLZ2EYsXL815I3Lp6tWr\nwsbGWcAvAm4KS8sBomXLzkaLJz8V6ERfunRpUblyZREQECBq1aolhBDi4cOHonnz5sLPz0+0aNEi\nw72bOXPmCF9fX/HWW2+JXbt2ZQ5YJnpJKjI6deorIOS5M989okqVRsYOK9fS0tJEUFAXYWNTWtjb\nBwhPz3IiMjIyx+X98MMKYWVlJ0xNLUSVKvVFVFSUEEKIuLg44eJSWpiYzBBgI0w4L1YwUAgQKSjE\nqY8/zpP23Lx5U+zatUtcv349T8rLqWXLlglr60HPfV5UwtTUPFdfZAqL3OQ+g9+jVygUHDhwgGLF\niumXBQcH06JFCyZNmsS8efMIDg4mODiYiIgINmzYQEREBFFRUTRv3pwrV65g8pq9QyVJKlyKFbPH\nxOQ2Wu2zJZE4OtobM6Q8YWZmxvbt/8fFixdRqVRUrlxZ94hbDpw8eZIxYz4iJeUPwI+LF2fQuXM/\nTpzYy549e0hJqYhWOxNzyvAjtelBKkkoWNi4BR/NnJkn7fH29sa7AExZa2+v+7yAQPfY4j9YWNjI\nHPEK+XJ0xH8eXwkLC2PAgAEADBgwgF9//RWALVu2/D979xkfVfE1cPy3NduSUNIoodfQQeklgBTp\nHRA/SjQAACAASURBVEUQRaog2MEOSlP0ryKK8AARRDoISBEEDIQivQQQQq8hEAglfct5Xuwag4Ak\n2U1C8H5f6d07M2fWjzl7507h2WefRafTUaJECcqUKcOuXbtyIkSFQpEL3n33dby9v0erfQW1eiRm\n80g+/fR9t+o8efIkn376KV988QWXLl3yUKSZp1KpqFy5MrVr185ykgfYsWMHdnsXoDygxm5/h717\ntwKg0WiAVLxIZilL6EkKt4CDn07kvfBfH7sJcJ06daJYsVsYDF2BjzGZWvLpp+Meu356Wo480T/1\n1FNoNBoGDRrEgAEDiImJITAwEHDOzoyJiQHg8uXL1K1bN61s0aJFc/V/VIVCkb1Kly7NoUO7mD17\nDlarjZ49w6lUqVKW6rJarXTr9hwrV64C+qDRWBk79kn27t1KqVKlPBt4DipUqBBa7QJSUmw4/2Tv\nokCBQgC0aNGCwj6j+C6+LM3kItfR8k27jox+++1cjTm7GI1Gdu8OZ8aMGVy+HEPz5tNp2bJlbof1\nyMv2RL9t2zYKFSrEtWvXaNGiBRUqVLjrc5VK9a+/xu732eh0w1GhoaGEhoZ6KlyFQpHDihUrxgcf\nuPcUD/Dhh2NZtWonzv3kh2K3w61bH/DJJ5MIC5vqdv25pWvXrkybNpddu2oDFbHb1zFnzlwAvO12\nIgv5ort0gut6AyuGDuODSRNzN+BsZjabGTFiRG6Hke3Cw8MJDw/3SF3ZnugLFXL+8vT396dz587s\n2rWLwMBArly5QlBQENHR0QQEBABQpEgRLly4kFb24sWLFClS5J46R3vovZNCoXh8rFq1EYcjACib\ndk2kHNeuncxynadOnaJ37yGcPBlFpUqV+fHHqQQHB3sg2gc7ceIEx44do0yZMlSsWBGNRsP69T8T\nFhbGhx9+xpUrcfTuPZBl076k8bhx6Pbvh+BgCm7cSL+yZR/egCJP+OdD7JgxY7JcV7a+o09MTOTO\nnTuA81CG9evXU6VKFTp06MDs2bMBmD17Np06dQKgQ4cOLFiwgNTUVM6cOcOJEyeoXbt2doaoUCge\nE4GBfkAw8CFwCjiCVjuG7t3bZKm+hIQEGjRowa5drYmN3UhEhC/ly9ehR48X6Nv3JQICyhAcXIkZ\nM2Z5rA/Tps2gWrUG9O49lVq1mjJp0leAc2Tzk0++4MqVlxFJQX99EgHde8L+/VCmDGzdCkqSVzyI\n5yb/3+v06dNSrVo1qVatmlSqVEnGjx8vIs7ldc2bN7/v8rpx48ZJ6dKlpXz58vLrr7/eU2c2h6xQ\nKLLJnTt37rsO3FMiIyPFYvEXjaaGgK+oVBZ5990PxeFwZKm+7du3i49PLdcyrvkCRV1LAasLlBLY\nKbBF9PpgWbx4idvxx8bGujanOeFq84IYDAXl7NmzEh0dLQaDn4BIcc7ISUqJgNwKDhbJ4XPeFbnD\nndynnEevUCiyVUJCAp079+b339ch4qB//8F8993/smVJ1IULF1i1ahVarZYuXbpQsGDBLNcVGRlJ\n3brtSEyMApoCHwFlgDrAj8DTrjt/oHXr1axdu9it2CMjI2nQoCd37hxNu+brW49ffvmMJ554gnz5\n/CmRuoINvEAwF9mn9sK26mdqP/30v9SqeFwo59ErFIpH1uuvv0tEhBc2203s9iv8+OMfTJv2f9nS\nVnBwMEOGDGHAgAFuJXmAypUr06xZXUymlsBV4BjQCNAD0enuvIRO5/6f0pIlSwKxwAbXlR3YbCcp\nX748RqORWa+OIIKWBHOR7WoTkzt05cnWefd8davVypIlS/j+++85evTowwsoskx5olcoFNmqfPna\nREV9DdRzXZlB9+7bWLQoLDfDyhC73U5YWBhhYT+yffte4DvgFZzzmIcBycAU9u6N8MgRrOHh4XTs\n2JPUVCE11Ur+/PmpX78OP7zclwK9ekFcHBcqVODI2LG06tIlz64ft1qtNGnShsjIRByOisBK5s+f\nQYcOHXI7tEeWO7lPOb1OoVBki6SkJL799jsSE+NRqSIQqQcIev02SpUqmtvhZYhGo6F///4EBATQ\ns+fXJCc/D5iBF4H/odNBWNh0j52zHhoaSkTEb9Su3RSH4y2uX29P/JpP8FrdFhwO6NCB4IULCTYY\nPNJeblm0aBGHDqWSkBCBc2B5Ky+99BzXrimJPjsoiV6hUHhcamoqDRq05M8//UhO7gB8gl6/Hi+v\nFIKC4nnnna9yO8RMCQgIQKM5ByQBXYHK6HQ1iI6+8MBXBPHx8fTpM5CtW/8gKKgQ06ZNon79+v/a\nzvXr12ncuAUpKcWBd2nJOn62r8KEg/j27bEsWQI6nae7l+OuXr2K1VqVv98eV+fmzZjcDOmxpryj\nVygUHrdx40ZOnEghOXkpMBHYg822hbCwVzl4cDu+vr65HWKm1KlThzZtGmCx1MfL62VMppaMHz/+\ngUk+KSmJ4sWrsHx5IrGxazh8eDgtWnTk9OnTiAg2m+2+5X755RdSUioBCXRiCb/QHhNJzFLrufPd\nd49Fkgdo1KgRGs1i4CCQilb7IXXrNs3tsB5byhO9QqHwuMTERFSqQP5+liiDRqPnqaeecmvf99yi\nUqlYuPAHfvnlF86dO0fNms/RoEGDB97/4YdjuXHjInAY51B/BVJTV/HRR2NYtmwlycnx1K3blBUr\n5uHn55dWzuFwoFL50YvrzKYHWoSv8GN3zw70K5o3XndkxBNPPMG0aV/w8svNSUy8xRNPhLJ06U+5\nHdY9oqKiWLJkKVqthl69elE0j/43UCbjKRQKj7t69SrlylXj1q2PgYbo9V9Rs+YpduzY8NCyj4PG\njdsTEREO7Me5JE9QqWqg010hNXUjUBad7k0aNjzLpk0r08pdvXqVCSXL80XiTdTAOJUvv9avweYt\nGx/bE9rsdrvrcJ5Hy759+2jcuBXJyc+hVidjMq1g375tuXZugrK8TqFQPFICAgKIiFjPE0/MJyio\nI+3aJbFmjXvrzPOSKlXKotFUBVoBnwI90OlO43D0AioBeqzW0WzfHn5XuYCffuJLV5L/NrgsjjGj\nCN+84bFN8sAjmeQB3nrrYxISxmK3f4XV+j137gxk7NjPczusLFGG7hUKRbaoUqUKu3dvyu0wcsW4\ncR8SHt6SM2fspKRMQKXS4O3ty507ewAHzmes/RQo4DzFExEYOxY+/ND571OmMHTo0FyKPvdYrVaW\nLl3KlStXaNSoEbVq1cq1WOLibgF/P707HKW5du1ErsXjjsf3Z6JCoVBkwu3bt9mxYwcnTrj/xzxf\nvnzs37+VMWOGotNZsNvnc/36bGy2o3h51cNk6ofJ9AyzZn3jTPIjRzqTvFoNYWHwH0zyNpuNpk3b\n0b//t4wadZLGjdvx44+5996+e/e2mEwfAFHAIUym8fTs2S7X4nGLe7vv5rw8GLJCoXjE7du3T/Ln\nLyw+Pk+I0Rgogwe/muU98tPr1Km3wAzX3vUisFJKlqwk06ZNk2PHjonY7SJDhjg/1GpFFi3yQG/y\npiVLlojFUlfA5vquDorFUtAj/x2ywm63y8iRH0j+/EXFz6+4TJr0Za7E8Rd3cp8ydK9QKNyyf/9+\nNm/ejJ+fHz169ECv1+d2SA8VHR3N+PGfEx0dS/v2TzF69GfExU0CegG3+PHH+nTo8CtPu7mPvNls\nQKW6zt9zqG5RpEhxBg4cCDYbvPgizJkDXl6wdCm0betmz/Ku2NhYHI4Q4K939hVJTLyF3W5Hq835\nVKVWq5k48WMmTvw4x9v2NGXWvUKhyLLFi5fQt+9QHI7uaLVHqVDBzvbtvz3Syf7GjRuEhNTi+vVO\n2GyVMZm+JCnpOCK3AefSP73+FSZOLMVrr73mVluRkZHUq9eMhISXASMm0/9YvXoRofXrQ69ezuRu\nNsPKldCsmfudy8OOHj3Kk0+Gkpi4DKiJVvsRNWvuY+fOjbkd2iNBmXWvUChyxeDBr5KUtJKUlCkk\nJGzg2DENixYtyu2w/tWSJUu4c+dJbLYvgZdITFwJmID5rjuuo9Oto3Llym63VaVKFXbt2sywYfEM\nGhTN77+vIrROHejUyZnkfX3ht9/+80keICQkhAULZlKwYC+02gLUrn2IlSvn5XZYjwVl6F6hUGTZ\n7duxOJeLAaix2UK4fv16bob0UFarFYfDku6KNxqNlXz5xpCS8gWpqdEMHjyEFi1aeKS9kJAQvvnm\nC+e/3LkDbdpAeDj4+cH69VCjhkfayYz4+HgiIiJQqVQ0adLkkdnEqH379sTGts/tMB47yhO9QqHI\nsgYNnkKnGwXEAztRqxfTpEmTLNUlIpw/f56oqCjsdrtH40yvffv26HSrgalABEZjL/r0eYHz54+x\ndet8Tp2K5PPPx91TzuFwcODAAXbu3ElKSspdny1YsJDKlRtQsWJdpk+fcf+G4+KgRQsID0cKFeKd\n+k0o3LYX1as3Ztu2bZ7v6ANER0dToUJNevacSI8en1C5cu1H/seZwk2emQ+Yc/JgyArFYys2Nlaa\nNm0nWq1BChQoIosWLc5SPVarVTp2fFYMBn8xm4tLSMiTcu3aNQ9H+7cDBw5IaGh7CQmpJ2+99b6k\npqb+6/1JSUnSsGErMZtLibd3FSlZsrJER0eLiMiKFSvEZAoWWCOwUUymsjJr1g93VxATI1KtmgiI\no3hxaVa8okBtgR0Cc8VgKCCHDx++p90//vhDRo16T8aNGy8xMTEe6fszz/QTrXaka2a7Q3S6oTJo\n0AiP1K3IPu7kvjyXNZVEr1A8fr788msxmZoJJLmSzwjp1u35LNVltVrlzJkzcuvWLRERmTkzTAoX\nLi9+fsXl9dffEZvNluk6P/lkvBgMHQSsAg7RakdKp07PiYhI27bPCMxKt4RuhVSv3kjat39GKldu\nIO88P0Ds5cs7PyxXTtbPnClgETiVrsxwGTt23F1trlq1SozGAIEPRafrLwEBxeXKlStZ+k7Se+KJ\n5gK/pmt7kTRv3tnteh/k9u3bEh4eLnv37s21pXKPA3dynzJ0r1Aoct3u3ZEkJnYHDIAKq7U3e/ce\nynQ9x44do1ixClSq1Ah//yK89NJAXnnlIy5f/oHY2HV8/30EH31077D8wxw6FEVycnuc05pU2Gyd\nOHLkOOBcQgdx6e6+yJEjkaxZE0L84WEM+HEe6uPHkcqV+bxDV/p9NNHVz/RlYvHyunulwquvfkhS\n0g/AGKzW/+PGjdZMnTot07H/U+PGtTEYvgdSgESMxhk0bvyk2/Xez/HjxylVqjIdOoyiceNutGvX\nI1tfyyjuT0n0CoUi11WpUhajcQ3gPL5Vo1lJxYrlMl1P+/bPcOXKmyQmXiA19SizZy8jMfFNoC5Q\nnsTEz1i06JdM11urViWMxiU4k6Og18+jRg3nrPxRo4ZjNk8AxgKfode/g05XgzL27kTwJiUlgd0q\nNcOrPsFH323n4sUhrn52A74FXkWj+YU+ffrc1WZCQjwQnPbvNlsxbt+Oz3Ts/zRu3IeEhqrR6/3R\n6QJo0yaAd9550+16/+n06dN07vw816+/ye3bO0hI+JPw8KuEhYV5vC3FQ3hwZCFH5MGQFQrFQyQn\nJ7vegZcRH5+aEhxcQS5evJipOmw2m6hU6nQ7q4loNDVEpRog0EqgkEBlCQmpnen4UlNTpXXrLmI0\nBomXV4CYTEWkefOOsmXLFhEROXjwoAwc+Ir06/eyfPHFF1LfVF1i8BcBCaeB5NcYRK+3CMS4Yvuf\ngE5UqnxisfhLRETEPW0OH/62mEzNBY4JbBKTqZBs3rw5wzFbrVYZM2a8NGzYVnr3HnDP93njxg2J\ni4vL9HfxMA6HQ4YOfUMMBn8BH4Ez6V4TjJU33xzp8Tb/C9zJfXkuayqJXqF4PNntdtm7d69s375d\nEhMTs1SHv39xgcUCMwW+Er2+uKhU3gIfCZwXmCI+PkFp7+8zw+FwyOTJk8VgKCrwk8B0MZn8Zfv2\n7XfdlxQeLjfVGhGQtVSRgsb60rfvIDEafV0xOJOewdBFJk6c+MA5A6mpqTJs2Bvi719SihevLAsX\nLpJbt27JoUOH5MaNGw+Nt3fv/q55D8tFqx0lgYElsyWxpxcTEyOtW7cXjaaEQJxAW4EPBBwCN8Vs\nrik//fRTtsbwuFISvUKhUIjIzz//LCqVRaCNQE/Ras1iMAS7Eo0zwfr61svUk3F6tWo1E1iR7gn1\nS3n22Zf+vuH330XMZhGQg6XLSpe2PeV///tabDabjBz5gZjNNQV+Eo3mHfHzC5arV69muO21a9eK\n2VxQvL0risGQT+bOnffAe1NSUkSj0QvcTovVYnlaFi5cmKV+Z8SdO3ckOLi8qNWNBYa42r0gUEnA\nT7y88kn//sOUCXlZ5E7uUzbMUSgUj40tW3ag0fTFZpsCgM32GXb7x8BtwBdIwWaLxtvbO1P1bt68\nmd9+20hMTAygSvdJAlFRR5k8eTI9fXwIHDIEkpPhueeo+sMPLE23R/uECWMoXrwoK1euoHBhP8aM\n2Y6/v3+G2o+Pj6dbt94kJKwAGgBHGDCgCU2aNKJo0aIZ7EX2bh++YcMGbt4sisMxEhgG3ACKAq8R\nHDyJnTt/p1ChQtnWvuLBlESvUCgeG5cuXcNmq5fuSkN8fApgs4WSkNAZs/k3mjWrQ/Xq1TNcZ1jY\nbIYNe4/ExJfQar1RqV5C5GvgIjCBw4fbs/2NlQy2ufZkHzgQpk51HjmbjkqlYsiQgQwZMjDT/Tp/\n/jxqdUGcSR6gEnp9CCdOnLhvotfr9fTs2YflyzuTmDgcrXYnZvOftGrVKtNtZ5TzR4QGaAF0AcoB\nFgoUsLFq1Rolyeci5VAbhUKRI5KTk/nyy685fPgkdepUY+jQIWg0mocXzIQff/yJwYMnkJi4CvDF\naOzFoEHVqFu3BgcOHKJ8+bL06dMnU+0WKFCEuLhVQA2cM+6fpFQpDfHxt7l8uR3POaoQxotocLC4\nWGm6nz0BKtXDqs2U27dvExRUnKSk34HqwGmMxtr8+edeihcvft8yNpuNceM+47fftlGiRGE+/XQ0\nRYoU8Whc6d26dYsKFWpy7dqz2O0N8PKaRIMGBlauXIzZbM62dv8r3Ml9SqJXKBTZKiEhgXfeGU1Y\n2HySkqpht7fHZFpAmzbFWLx4jkfbEhHGjBnPZ59Nwmaz0qNHL2bN+tat0/QMBm9SUs4D+QHQ64cx\nblwJtm49QKEVGqbi7MNonmdZ5VMcitz6r/WdPHmSAwcOEBwcTJ06dTIcx8KFi+nXbwg6XTlSU6OY\nNGkcQ4cOynK/3HHgwAFmzvwRtVrFgAEvpB0AdOnSJV5//X3Onr1E06Z1+fjj9x/pkwzzEiXRKxSK\nR5LD4aBGjYYcOWLCbj8LHMc5vJuIwVCMkycPZstT5l9/I1QeeLLu1KkXv/4qpKRMBDagUr0OxDNK\n68V4axIAb/IBU00RvP56Mz755IMH1rVgwSL69RuKTtcQu/0Afft24dtvv8hwLNHR0Zw4cYKSJUsS\nHBz88ALZICIigtatu5CYOBywYzZ/S0TEemrkwuE8/yVu5b6szwHMHXkwZIXiP+vFFwcJFBD4XeDJ\ndLPV7WIyFZVTp07ldogPdefOHenZ80XJl6+waDT5RMX/5CM++KsjMlxnFqMxnwwcOFysVusD60lN\nTRWDwUfggKvoTTGbS8gff/yR6ZhsNpv83//9n7z66psSFhYmdrvdnS5miN1ul8GDX3WtjZ9218qD\nzp17Z3v7/3Xu5D5lZzyFQpHm2rVr7NmzxyOnme3du5f581fi3O61JnATGAPsRasdTqlSRShRooTb\n7TgcDs6ePcvVq1fdrut+LBYLCxbM4tixfWg1KiZxkdF8gh01gww1CF34I4mJcUyb9jVa7YPnN8fF\nxeFc6FTNdcUXjaY658+fz1Q8IkKXLr0ZMWIOX31VkGHDptG794AMlY2MjKRjx140atSOqVOnZ+oJ\nccqU75gzZwdQB0i/WiCA+PikTPVBkcM89nMjh+TBkBWKPOGHH34UozG/+PhUF5OpgCxb9rNb9S1d\nulS8vdsLtBDoKfB/AmVFp/OXLl16S2xsrNsxnzt3ToKCSotG4ydarbf07t0/255ukxMTZbpaKwKS\ngk66ME/M5rKydevWDJW32+0SGFhSYLbrSfigmEz+EhUVlak4Dh8+7DotL8lVT7wYDP5y5syZfy13\n4sQJsVj8RaX6UmCZmExVZOzYTx/a3r59+6Ro0fKug3jCBH4QKCewxbVjX0lZuHBRpvqgyDx3cl+e\ny5pKolcoPO/SpUtiNBYQOOJKHnvEZCqQpZ3Url69Kl269JFixUJEo/EV2CUwSqC2GAwF5fr16x6J\nOSUlRXx8gl2bszgE7ohO94RMn/5/HqlfxHkM78yZM2XmtGmS2K2bCEgiSCevVmKxVJEePV7I1AYw\nBw8elMDAkuLllV8MBh+ZPz/zG9j88ccf4uNTI93QuYjFUk4iIyP/tdyYMR+LRvNqunKHxN+/5L+W\niY+PlwIFigjME3hb4CXXd/29QEkxmQrLjBmzMt0HRea5k/uUdfQKhYJTp06h15cnKSnEdaUWGk0Q\n58+fJ1++fBmux2az0bjx05w61QirNQyVahwqVSMMhnyYzV6sW7eeAgUKuB2vzWajV6++3L6dAAzG\nuYmNBau1D9u27WHAgP5ut3HhwgVq1WqILeEJwlJ2YbRfxGE2c/6LL2glwoBixXj66aczNeGvatWq\nXL58ktjYWPLnz49Op8t0XFWqVMFsvk18/CQcjs5oNPMpWFBDuXL/fgiQM870Q/WOh8Z+4sQJrNZ8\nwLNAK6AxUA+TyQ+DIZWdOzdTpkyZTPdBkbOUd/QKhYJSpUqRmnocOOa6sg+bLZpixYplqp5jx45x\n8eJNrNb/AbURWY7ZXJolS2Zx5cppatas6ZF4R478kF9+OQhYgF9dV+3ASsqWzXjMEyZMxNs7EIPB\nl+7d+5KQkJD22fvvjyPpek/mJSbQ0X6ROAyMrNmAY0FBJCUlYTKZsjSrX61WExAQkKUkD2Aymdi6\ndT11627Ez+8pGjbcSUTEuocuY+vV61mMxnmoVJ8DSzCZnuO114b8a5mAgABSUy8DV4ECwFp0uj+Z\nMKElx48fUJJ8XuG5gYWckQdDVijyhFmzZovBkF98fGqKyVRAFi9emuk6oqKixGAIEkh0DQ+nitlc\nQg4ePOjRWAMDywhsFvATCBKoJ1BSvL0LS3JycobqeP31N10rAvYKXBOttpP06tU/7fP2TdpJOBVF\nQGLwl6p8IwUKlBCLpbro9a+IyVRSPv54okf7ld0OHz4snTv3ltDQDjJt2owMvXZ4//2PxWQqISbT\nS2I2l5Y33ng3ByJV/JM7uU9ZR69QKNLExMRw7tw5SpUqhZ+fX6bKXr16lUaNWhMVFYlzsLAzRmMC\ntWs72LTpF9Rqzw0glixZjbNnvwL8gFeAQ5QoEcSePREULFjwoeXj4uLw9y+C3f42MNp19TT58jUh\nLu4C3LjBlRo1CDp/nosU4imWcc4wFLs9Bqs1CjAB0ej15YiJuZCp1xt50datWzly5Ajly5cnNDQ0\nt8P5T3In9ylD9wqFIk1gYCC1a9fOdJK32WxUqFCHqKgngSTgOGr1Zp55JpB165Z5NMkDfPrp+xiN\nzwFr0GqrUrCgme3bN2YoyYNz4xmt1gxEpbt6DJPJDDExEBpK0PnzxPr40lwXx2ldM0JDi2A0lseZ\n5AEKodXm4+bNmx7t26OoYcOGDBo0SEnyeZQyGU+hULht+fLlxMXFAh/h/LNSAoejH8HBOry8vDze\nXo8e3QkI8GfJkpX4+uZj6NCdmTo0pUSJEuj1KlJSdgKdcZ6yFsbUdz+Fxo0hKgoqVMBvwwaOFS4M\nQGxsLKVLVwaWAy1Rqf6PAgVMnD59mrlz5xIYGEifPn0wGAwe769C4Q4l0SsUCrc5j2/1BXYBnQAH\nsIVChZ7LtjZDQ0Oz/IRpMplYu/Zn2rbtQmLiRlQqGzPfHUmHSZPg3DmoXh3WrYOAgLRDaf39/Vm3\nbjk9e/bj8uUeVKhQkx49nqd9+74kJz+HwbCV77//kR07Nij7u/+DiLBt2zYuXbpEzZo1KVu2bG6H\n9J+ivKNXKBRu279/P/XqNSclRQW0BI5jMkVz48bZbHmivx+bzcbq1auJi4ujUaNGlC5dOu2z3377\njZ9/Xk3+/D4MHz6UwMDAtDJXr17F/9o1dE8/DdHRULcurFkD+fP/a3sigtmcn6SkP4AKgGCxNGHW\nrFfo3r17Nvb0b8uXL2fgwFe5dSuWxo2fYtGiMPI/JO6cJiK8+OLLLFmyAY2mGlbrZubM+Z5u3brm\ndmh5irLXvUKhyHU//TRfjEZfUanUUqpUiJw7dy7H2k5NTZV69Z4Si6WOmM3PicnkJxs2bBARkZde\nGihQUGCSqFSDxN+/mMTExPxdeO9eET8/5y4yoaEit29nqE2r1SpqtVYgJW0TGpPpBZk+fXp2dPEu\nDodDFi9eLF5efgJbBeJErx8kzZt3yPa2M2vz5s1iNpcVuOP6nvaK0eibI/vzP07cyX15LmsqiV6h\neHQ5HA5JTU3N0Tbj4+OlevXaArUF7K5kslaKFq0gGzduFPAV2J6WjNXqPjJp0iRn4W3bRHx9nR88\n/bRYb9+Wd98dLRUq1JEGDVrLrl277mkvJSUlbVlao0atRacbInBFYK2YTH5y/PjxbO2vzWaTdu16\niE6XT2BAup3ubotWa8jWtrNi7ty5YrH0SBenQ3Q6s9y8eTO3Q8tT3Ml9yqx7hULhMSqVKssbwWTV\nwIGvEhmZDDTi74VETxIbe5nVq9cBOiAg7X6HI4iEhETYtAlatoRbt6BrV1i+nFffGc1XX/3OsWOf\ns21bD5o2bcOJEycAuH79Og0btsJotGA0+jBlylSWL/+JZs1iMJsrERz8OitWzH/oDnXumjlzJps2\nxWC1fgGc5u/d7o7h7Z2xVQc5qVatWtjtvwORAKhU0wkKKoqPj0/uBvZf4sEfHDkiD4asUOSq2UZO\nXwAAIABJREFUpKQk6ddvqPj7l5QyZWrKr7/+mtsh3eXOnTsyePCrUr16E3nmmX5y5cqVTJX38ysh\n8JNAUYEoAZuo1cMlNLSdjB8/QdTqqgLNBfYLfCfgI520RklWqZyPmM8/L+I6XtZi8Rc4l/b0qdMN\nk88++0xERFq16iI63csCqQInxGQqJps2bfL49/Eww4e/ITBRIFmgvqtvQ8RoDJQFCzK/d35OmDt3\nnhgMPqLX+0rRouXk6NGjuR1SnuNO7stzWVNJ9ApF5vTuPUAMhg4CxwVWidHoJwcOHMjtsETEOdTf\noEFL8fLqJbBBdLo3pXjxEElMTMxwHRUqPCmwzHXQikVAK8HBFSQmJkZu3LghRYuWE42mskBhAYv0\noLek4jyFboaXSeLSHbKTP38RgcNpid7Lq6989dVXIiJisfgJRKd9plK9I6NHj/H4d/IwYWFhYjLV\nFYgXSBSVqqOULh0ie/bsyfFYMsNqtUpsbGymDgFS/M2d3KcM3SsUj7nly5eTnPwdUA5oi9X6PGvX\nrs3tsAC4dOkSe/fuJyVlNtAcq/Uzbtwws3PnzgzX8f33kzCZBmIw7MNkakSpUhU4cmQXAQEB5M+f\nn8jInXz99WBef/0ZBukNzGMeOmx8xlu84VWNg5GRaXW9995bmEydgeloNG/h7b2JZ599FgA/vyBg\nj+tOB0bjPgoVCvLYd5FRzz//PJ06VcJgKIHFUpXixU8SHr6OWrVqpd1z9epVXnhhCA0btuW998aQ\nmpqa43H+k1arpWDBglk6H0DhHmUdvULxmDOZLMTHXwKKAKDVXsRiKZm7QbloNBpE7ICNv/8cpaLR\naO57v81mY926dWlL6IoXL06TJk3Yv38b69evx2KpR/fu3TGbzWll8uXLx9ChQ7k9fjw+qbEAfMDH\njOV1TLaKdy1He+ONEQQHF2bp0rUEBORn1KgdBAQ43+/PmjWZdu16oFI9jUp1ivLltfTt2zdD/YyO\njmbEiJFcvHiNli0b8t57b2d5LoNareann2Zw/vyHxMfHU7Zs2bvqSkhI4Mknm3D58tPYbIPYt28a\nhw/3ZcWK+VlqT/EY8ODIQo7IgyErFLnqhx/miMlUROAT0eufl6JFy2XpnPns4HA4pG3b7mI0Pi0w\nX7y8XpBKlWpLSkrKPfempKS4ltA9KRZLTzGb/WTz5s0Za2jChLTD20fqA0SjeVvM5lrSs+eLmRpK\nPnnypMycOVOWLl163xjv59ixY6LR/DVDfoloNE2lc+fnMtxmZq1Zs0a8vRulm+WeKDqd+ZH5b67I\nGndyn/JEr1A8hmJjYzl48CBBQUH07duH4sWDWb16HQULhjBo0Fe5eghLREQE3377A2q1ihEjBrBs\n2VwmTvycbduWEhJSijFjvr7vznJz587l4EEHiYk7AA2wkr59h3LmTOQ996YRgQ8+gHHjQKVCpk6l\nYZEi+Bw6RNmyI+nWrVumhpJLly5910Y8GdGlSy/s9jLANECF3d6GlSsLcvPmzfv+d1ixYgVjx36D\n3W5nxIh+9O3bJ1PtyT2bqihD5f95nvu9kTPyYMgKRY7asmWLWCz+4uvbWIzGwjJo0IhHZgLUxo0b\nxWQKEJgs8KWYTH6ybdu2DJX95JNPRK0ele5JNUZMpgIPLuBwiIwY4bxZoxGZO9dDvcgci6WgQMN0\ncSeLWm2SGzdu3HPv2rVrxWgs5JpcuEpMppIyZ07m4o6Pj5dixSqITveawHIxGttK+/Y9PdUdRS5x\nJ/fluaypJHqF4t/5+xcXWO1KKrfEbC4vv/32W6bquHnzphw9elTi4+M9GlvTph0FfkiX9L6Vjh17\nZais80dCcYEzAnbRal+Tpk3b3/9mm02kf39nI3q9yLJlHuxF5tSo0cg14/9tgTUCT0uVKnXue2+H\nDr0E/i/d97Nc6tRpmek2Y2Ji5IUXhkiDBm3k3XdHZ/g1g+LR5U7uU4buFYrHiM1mIzb2AtDadcUH\nkYacOnWKp556KkN1zJu3gP79h6DV+iMSx88/z89w2YexWm2AOd0VM6mptgfca2XBggXExMTQsGFD\nmjVrxtixbzBqVCXsdjvVq9dlwYJF9ysIffvC/PlgNMLPP0OrVh6JPysWLJhB/frNuXXrJ+z2MEqV\nKsSOHdvve69erwMS0l2JR6fL/J/pgIAAwsK+y1rAisePB39w5Ig8GLJCkaNKlKgkMMv1RHhRTKZg\n2b59e4bKXrhwQYzGggKHXOV/F4vFz2NP9gsXLnI9la8UWCYmUxFZtWrVPffdvHlTAgPLiEpVW1Sq\nYWIwBMnMmWEi4twCNiEh4f4NJCWJdOzofBz29hbJ6GS9bHbnzh3Ztm2bREZG/utrlF27donJ5Ccw\nSWCyGI0Bsn79+hyM9G63b98Wm82Wa+0r/uZO7stzWVNJ9ArFv4uMjBR//+JisZQSvd5bxo+flOGy\nGzZsEF/fJumGjkUsltLy559/eiy+efPmS61azeSJJ5rLsvsMqTscDqlS5QmBGvL33vVHxGDw/fe5\nBvHxIi1aOIPOn1/kPvvU5wW7d++W3r0HSM+e/SQ8PDxXYjh37pxUqFBLtFqjeHlZZMaMWbkSh+Jv\n7uQ+5ZhaheIxlJqaytmzZ/Hz86NAgQIZLnf69GkqV65DUtJeoBhwCKOxCVeunMuxvclv3LhBQEBh\n7PZngTDXVRtgICioFCkpifTs2Z3Jkz/7e/34rVvQrh1s3QoBAfDbb1C1ao7E+ziqXr0hhw+3xm5/\nD4jCZGrKli2/3LUpjyJnuZP7sn1nPLvdTo0aNWjfvj3g/J+4RYsWlCtXjpYtW3Lz5s20eydMmEDZ\nsmWpUKEC69evz+7QFIrHll6vp1y5chlO8mfOnCE8PByj0cj48R9hNNbC17cxRmMzZs2alqMHkGg0\nGlQqNbAK+B24BXRDpcrPlSuziYvbwuzZh3nrrQ+cBa5fh+bNnUm+aFGIiFCSvBscDgeRkX9gt4/E\nuTSvPA5He/7444/cDk2RRdme6L/++mtCQkLS1qpOnDiRFi1aEBUVRfPmzZk4cSIAR48eZeHChRw9\nepRff/2Vl19+GYfDkd3hKRT/eV98MZlKlWrTqdMHlClThWLFinDkyC6WLRvNiRMHeeaZHjkaj6+v\nL1279sTLqwjQGwhEowlH5G2gHlCKpKTPWbr0F7hyBUJDYe9eKF3ameSz+fS4nGS1Wvnhhx8YO3Ys\nGzduzJE21Wo1+fIFAX8l9lQ0mr0UKVIkR9pXZAMPvT64rwsXLkjz5s1l06ZN0q5dOxERKV++fNrp\nVNHR0VK+fHkRERk/frxMnDgxrWyrVq1kx44d99SZzSErFHmGw+GQS5cuydmzZ7O8Tj4qKkqMRn+B\n86534XvEaMz34MluOcRqtcqECZOkRo0GUqRIJSlWrLxoNIPTzR34WZqXrSlSpozzQkiIyKVLuRqz\np9lsNmnUqLWYzaGiVo8Uk6mETJr0ZY60vXr1ajGZ/MTbu6dYLFWkTZtuYrfbc6Rtxf25k/uy9Yn+\ntddeY9KkSajVfzcTExNDYGAgAIGBgcTExABw+fJlihYtmnZf0aJFuXTpUnaGp1DkWVarlY4dn6VU\nqSpUrFiXOnWacfv27UzXc/r0afT6qkCw60ot1GpfoqOjPRpvZmm1WkqXLsHx4+e5dOl9zp8fgt3+\nIzpdX9TqUVQx9OOXWxfh5EmoUQM2b4bChXMkth9+mEPBgsEYjb507dqHhISEhxfKgg0bNrB/fwwJ\nCRtwOCaSmLiZd999F6vVmi3tpdemTRsOHfqDqVPbs2zZF/zyy8K7/o4r8pZsW0e/atUqAgICqFGj\nBuHh4fe9R6VS/ev2kw/6bPTo0Wn/HBoaSmhoqBuRKhR5z6BBL7N69RUcjkuAjkOHBvDqq+8wa9a3\nmaqnTJkyJCfvAzYCzYDfUasTH4lh2kmTppGYOBno5LqSRMWKSxnc0Jv+izTorl6F+vVh9WpsFguv\nDnuDOXPmotPpef/9t3jtteEej2nLli28/PI7JCWtBEqwZs1Q+vcfzvz5Mz3eVlxcHCpVKZzb/QIU\nRURFUlJSlg/EyYysbPer8Jzw8PAH5s7MyrZEv337dlauXMmaNWtITk7m9u3b9OnTh8DAQK5cuUJQ\nUBDR0dFpJ0MVKVKECxcupJW/ePHiA//YpE/0CsV/zaZNm5g9+2ccjv8BBgBSUl5g5853MlVPbGws\nHTv2AkxAZzQaA0ajsHz5QgwGg8fjziznD/3083TMtA4IYsiC+XDjBjRrBitWgMXCR++NISxsL4mJ\nu4DbvP9+F4oUKUSPHt09GtO6db+RlNQfcM4+T06eyLp1DT3axl8aNGiAw/EKsBxogFb7OZUqVc/R\niZGK3PPPh9gxY8ZkvTIPvkJ4oPDw8LR39G+99Vbau/gJEybIyJEjRUTkyJEjUq1aNUlJSZHTp09L\nqVKl7vveMYdCVigeWZ069RZoJ9Ar3Trzt6RLlz6Zqqdr1z6i0w0XcAgkisEQKhMnfpZNUWfesmXL\nXKfuzRGYJs298onVbHa+k2/Xzrk5jkv58rUFItK9w58mPXv283hMX375pRgMPdK1s1ZKlKji8Xb+\nEhERISVKVBaTKb80btxGoqOjs60txaPNndyXY1vg/jUMP2rUKHr06MHMmTMpUaIEixY5t7AMCQmh\nR48ehISEoNVq+e677zJ1qpRC8V+h1WqAJjif9KoBWvT6C0yZ8i+nuN3Hvn2RWK3/h3MJlZHk5J4c\nPLgnU3XcunWLCRMmcfbsZZo1q8+AAS957P/bzp07s3ChjsmTf+DJ27F8fDAZTUIy9OgBc+dCuuHr\nggXzAycA59O1VnsCf3/3T+i7du0affu+zO7deyhWrDjfffcpRYrMIjq6I1ZrCbTaeUyd+qPb7TxI\nw4YN//10PoUiIzz4gyNH5MGQFQqP2r59uxiNfgJfC4wUvb6ALFmyJNP1tGnTXTSaD11PplYxGtvL\n+PGfZrj85cuXXSfRPSswXfT66jJs2BuZjuOhVqxwHkwDIi++6Dyw5h927twpZrOfaLWviJdXX/Hz\nC5aLFy+61axzh766rlPgTohKNU3y5y8sZ8+elenTp8vnn38uhw4dEhHnKoFt27bJxo0b5c6dOxlu\nY9euXbJgwQI5cuSIW7EqHn/u5L48lzWVRK9QOJN95869pX37ZzO0F/pfS/H+Wtoq4lz+GhxcXnx8\naorZXEYaNmwpycnJGY6hVq0GArVdQ/8icF00Gq9Mn5RmtVolLi7u/ksE5893HjELIsOGifzLEq/j\nx4/Lp59+Kl9++eVd/cyKixcvysyZM0Wvz5+ufyI+Pi3u2Zs/MTFRnnwyVCyWSuLjU18KFSot586d\ne2gbb7zxrphMxcTbu6uYTIEybdoMt2L+p507d0qtWqESHFxJBgx4RRITEz1avyJnKYleoVA8UHx8\nvDRp0kYMhoLi5ZVf2rbtnpaMExMTZdu2bbJnz55Mr5PW680CbdO9r04RtVqfqTX406fPFL3eLDqd\nRcqUqSZnzpz5+8MZM0RUKmflo0Y5z5fPAb///ruYzX7i7d1KwEvguqt/NrFYKsvmfxyU88kn48Vg\n6CJgExDRaD6RVq26PrD+mJgYadq0jahU+dPVHSVeXj5y+/Ztt+O32WyyYMECMRjyC8wW2C8GQyfp\n2jVzczgUjxYl0SsUigcaNuwNMRh6CqQKJInR2EY++mhslupyOBwyefK3UqlSfdFq8wkECPxP4A+B\nzlKx4hMZrmvPnj1iMhUSOC7gELV6olSq5Dqn/auv/j5VZ9y4LMWaVUFBpQTWupp/Q6CswHgxGltL\ngwYt7znN7dlnXxL4Pt0Pnt1SsmT1+9admpoq5crVELW6q0DoXYcHmc3F5NSpU27FnpqaKo0bPy1e\nXoUEnktX/03Rag1Z3lhJkfvcyX3KDggKxWNux479JCf3A3SAgaSkvmzbti9LdU2ZMpVRo77jyJGP\nsdleBuJRqb5HpepE/vx72bJlbYbr2rlzJyLtgHKACofjDY4e3Y1j7Fh49VXnTV99Be++m6VYs0JE\nuHr1PBDqujIJrbYkTZpsZNKkdmza9AsajeauMvXr18BkmofzHHkHev1Mateucd/6jx8/zuXLd3A4\nvgYOAztcnyzGYHDctWlYZkVFRfHWW2+xe7eVlJSPgfQbKF1Dp8v9JZOKXOK53xs5Iw+GrFDkql69\nXhKt9g3Xk51DvLxekldeeTNLdVWsWFdgU7onxbflyScbyJw5cyQp3XK3jFixYoVYLDUEkl11bZb/\nGSzOilUq59D9v3j//Y/Ez6+UBAaWkQkTJnnsabVq1fqiVk90vZs/KyZTsGzduvWB99tsNnnmmRfF\nyyufGI1BUqNGQ7lx48Z97z1x4oQYjYVcfV4tUEDAKPnzF5E9e/ZkOeZ33hktRmOA6PXFBcYK3Bao\nIPCiwFdiMpWViRM/z3L9itznTu7Lc1lTSfQKxcOlpqbKvn375ODBg3L58mUpXjxEvL3riLd3TalQ\noZbcvHkzS/VWq9ZYYHlaolepxsjAga9kqS673S4dOjwjFkuI+Fi6yndag7hecovMmycizlcFx44d\nk127dt317r9Nm44CRQV2CewTgyFEpkyZmqU4/unMmTNSsmRl8fIqIDqdSb744usMlbty5YqcO3fu\nX+c6OBwOadOmm5hMLQSmisHQWho1uvd1QGbs3r1bTKaiAlcFFgmECMQKXBeVqoEULlxOli5dmuX6\nFY8Gd3Kfch69QvGYuX79Og0btuLixQREUqlSpSSrVy/iwIEDaDQa6tati5eXV5bqXrlyJT169CMl\npSWgwWRay+7dWwgJCQEgNTWVsLAwzp49T/36ddOOp34QEeH3DRsoMXYspbZsAb0eFi2Cjh1xOBz0\n6TOQn39eg04XiMFwk4iIdVy9epUmTbricHwJ9HLV9At1637Ljh2/Zqlf94vr2rVr+Pj4eHyXQKvV\nyjfffMvevUeoXr0CI0a8gl6vz3Q9s2f/SFjYEu7cieX48QASEn4GBHgH+BKDwZfSpUuycePKtPNF\nFHmXW7nPIz81clAeDFmhyFHPPTdAdLqhrqFnmxgM3eTddz/ySN1z5swVLy9/0WieEa22vDRv3i5t\nyNxqtUq9ek+JydRSYLSYTOUePukvJUWkRw/nk7zJJJJuqeC8efPEbH5SIN41ejBZatZsLDNnzhSN\npqLAp+leIUyRp57q7JE+5gVTpkwVk6ms6wn+XQEfgVPy18l++fMXlgsXLiiT7x4j7uS+PJc1lUSv\nUPy7qlUb/eM9+lx5+ukebtdrs9nEYPAWOOyqN1nM5hDZsGGDiIisX7/e9c7d5vr8smi1hgeuzT93\n/LjsLRwsApJsMIhjy5a7Pv/ggw8FPkjXj0vi7R0gf/zxhxgMQQJ+rlnxb4tKZXLrHbenpKamyscf\nj5eWLbvJK6+8KXFxcdnSTokSVQW2pvtu2olGYxKLpbTky1dIdu7cmS3tKnKPO7lPmXWvUDxmqlcP\nQa9fiPNAGCtG41Jq1arkdr2JiYnYbDYgxHXFC5WqStqRtrdu3UKlCubv09YCUat1JCYm3lPXnvBw\nosqHUPPyBa5joYmtKKNWrb/rnkqVQjCbVwN3AFCr51O+fAh16tThww/fQK9PRqebjcHwPTNmfMPG\njRv54IOP2LMnc9v4elL37n2ZMGEz69d3Zfr0OOrVe4qUlBS36xURLl68mPZd37vNcA0GDerPnj2r\niY4+Te3atd1uU/EY8dzvjZyRB0NWKHLUjRs3pHLlOmI2lxKTKVgaNWqd6RnxD1KmTDVRqz8T52E6\nO8Vo9JOoqCgREYmOjhZv7wCBnwTOiU73mtSo0fDeSuLiZKfOuaXtZYKkEpECF0Svt9w11OxwOKRz\n516uYeliAmZ5442RaZ/HxMRIZGSknD17VgICSohe309UqvfEZAqQNWvWeKS/mXH16lXR630FEtNW\nOHh715KNGze6Ve+dO3ekXr2nxGDwFy+vAtKuXQ+ZPPnbdEP334jZ7CeHDx/2UE8UjyJ3cl+ey5pK\nolcoHs5ms8nhw4flzz//dOs9bVxcnPTq1V/KlKklbdp0l4iICKlQ4QlRq7Xi7e0nP//881337969\nWypWrC358hWWFi06y9WrV++u8No1cVSvLgJyDpOUIcqVFK+KRmO8J9ayZWsIfCJwQOCkmEzFJCIi\n4q57PvpojGi1g9INY6+ScuUyvnGPp0RHR4uXV35xbkz015a59TO0RfG/GThwuHh59RawinPDo1by\nyScTZPbsH6Vp047Svv2zj8RrC0X2cif3KbPuFQrFfTkcDmrXbkpkZAVSU/uh0awjIGAOUVEH0Ol0\n6PX6zJ1Ud/kytGgBR49ySq2hqcOHC7wPVAY+oHXrQqxdu/yu9rVaHSLJODf7AYNhMJMmVWbYsGGE\nh4fz3XezOXBgPydO9MQ52xzgMIULd+PSpWMe+ibuz2q18u2333Ho0HFq1Ahh8OBBtGjRiZ07vUlO\n7o9Wu4FChZbz5597MZvNWW6nWrXGHDo0BmjquvITTz+9kjVrFnqkH4q8wZ3cp7yjVygU93Xx4kWO\nHj1OaupUoA52+4ckJASwa9cuvLy8Mpfkz52Dxo3h6FGoVImLP80l1ihotVNQq/sQEuJg5crFAMyd\nOw9//+JYLAXQ6/MB61yVxKPRbKV06dKsX7+etm2fYfHiJzlxogEwCdgCnMRofI2uXTt49Lv4J4fD\nQevWXXj33dWEhVVk1KhldO3ah9WrF/HSS0WpUWMcXbteZefO391K8gDly5dCq/1r2aDg5bWOkJDS\n7ndC8d/hoVGFHJMHQ1Yo8qQrV66Il1e+tOVtYBeLpfI9Q+d/WbFihbRt+4x069ZX9u7d+/cHx4+L\nFC3qHMuuVUskNlZERM6fPy/Lli2Tbdu2pQ3Zb9261bX//U6Ba6LTPSVarbf4+jYVk6mY9O07WBwO\nhzRq1NY1F+Cv4fq+YjQGSsGCxWTIkNckNTU1276Xffv2ScGCRVyz/lNc7SeJ0RgkJ06c8Hh70dHR\nUrx4RfHxqS0WS1WpXLmORw6/UeQt7uQ+bW7/0FAoFI+mwMBAOnbsyKpVbUlMfA6DYT0BASqOHDmC\nv78/5cuXT7t3/vwF9O//NomJHwM3Wbu2Fdu2baCaWu0cro+JgYYNYdUq8PUFIDg4mODg4Lva/O23\nDSQlvQg4Z41brWFYLDVZvPgd/P39qV69uuu6DTCmK9mQFi2srFjxUzZ+IxAWNpt+/V7G+ZpgEfDX\nRjdeaDTeJCUlZbluq9WKTqe753pQUBBHj+5h165daRse3e8+heKBPPiDI0fkwZAVijzLZrPJV19N\nlm7d+krRomXFbK4tJtMLYjL5ydq1a0VEZNGixWI0FhZYk+4Je6yM7dRdJH9+54UWLUTi4x/a3uTJ\nk11Hvv5VzwYJDq54z30//TRPTKYSAisFFovJVEh+/fVXj/c/vf3794uXV0HX1rtJAhUFPhQ4IBrN\n21K6dNUsjSTs379fgoMriEqlloCAErJt27ZsiF6R17mT+/Jc1lQSvUKR8+bMmSNmcxPXsjoR2CRB\nQaVlyZKlYjIFC1QS2JCWoBsxWBJ1Oue/dOggksHlfXfu3JGyZauJydRedLoRYjT6y6pVq+57748/\n/iS1ajWT2rVbyIoVK2TPnj0ybtw4+fbbb+XOnTue7L6IiEydOlUMhucFvAVOClwU6CAqVT5p1Ki1\nXL58OdN1JiYmSsGCRQXmur7bFeLtHSDXr1/3ePyKvM2d3KcM3SsUeYSIMGtWGFu27KJs2WK89toI\ntyd6ZVRMTAypqTX4e/5uTW7cuMKUKbNJTPwM5xGtg4HPaUkEP/M9Rivw7LMwezZkcKjZYrGwf/82\n5s2bx82bN2nZ8jeqVat233t79+5F797Ove6XL19Oo0ZtSE19Hr1+L1988T0HDmzD29vbzZ7/rVCh\nQmi1R4BPgYZAI2Anw4cP4quvJmapzlOnTpGaagGec13pgFo9gSNHjtCoUSOPxK1Q5LnH4zwYskLh\nEYMHvyom0xMCU8Rg6CFVq9aTlJQUt+u1Wq1y+PBhiYqKeuCa+x07driOVz0iYBWt9jVp1OhpadGi\ni8As15P8bOlICUn+67G+f38RN05lywiHwyFjx34qKpWPwO9pIwpGY1f55ptv0u47ePCgPPVUJ6la\ntbF89NFYsVqtmW7LbrdL69ZdxGKpLkZjG9HpLDJx4kS34ndOePQViHbFfkOMxkA5fvy4W/UqHj/u\n5L48lzWVRK/4L4qPjxet1ijOY1nrC+QTtdpfpk+f7la9165dk4oVnxCLpbQYjYWlZctOD/zxMHNm\nmJhM+USt1kqdOs0lJiZGNm3aJEajv8A30os+Yv0r044YIeLGRj0HDhyQl14aKi++OER27Nhx33vs\ndrsMHz5C9PpyrhnwF9MSvVo9Sj7++BMRETl79qx4eweISjVFYKOYTI1lyJBXsxSX3W6XNWvWSFhY\nmBw7dizL/Utv9OjxYjIVF5Opn5jNZWT48Lc9Uq/i8aIkeoXiMXf9+nXR6SwCJQW+Ebgm8J34+ARJ\nfAYmuT1Ijx4viF7/ijhPuksRo/FpGTfu0wfe73A47plwtmXLFvm+Zj2x/5Vl338/Lck7HA5Zs2aN\nfP/997J79+4MxbRnzx4xmfwExgv8P3tnHhZl9b7xe/aZd4YBZFUBE0TEfV9zScP9p7lruaWpaS5l\nkdpimaW2mZq54JqaW6amiAuaqKm57xsiGIobIoIyMMDM/ftjRoKvoCCoYedzXV7F+57znOeckvs9\n2/N8R0ly486dO3OUeTC7Vii8CCwg0J9AdwLXCOyhTufBv/76iyQ5ffp0ajSDsh3wi6MkORdglApP\nVFQUa9duRoPBjdWqvfzQR8LevXs5Z86ch/opEDxACL1A8IJjtVpZrVp9Ar7ZBIt0cKie70xlBw4c\n4IIFC7g7W5a48uXrENiXzeZ8du7ct2DOTZ36j0PZlrKtVit79HiTen1l6nQDKUml+NPG0LK+AAAg\nAElEQVRPcx5rrmvXfgR+yObTYjZr9n85ymzZsoUGQ1UCIwgE2+/69yPgSLW6BJct+4WbN29maGgo\nv//+e+p0vbPZu0gHB7eC9bEQhITMp1xuJPANgWuUyX6km1uZQn2gCf57FEb7xGE8gaAYIJPJsHRp\nCKpXbwyrNQmAI4D7yMy8AWdn58fWnzLle0ycOA0yWXMAX2Hw4B6YOnUSKlUKQEzMWmRk1AdggU63\nETVq1MufUyTw5ZfA+PG2n2fOBN55J+v1/v37ERq6BykpJ2G78x6N996rigED+kGr1eZpNjXVbO/f\nA5yQlpaeo0x8fDyAQABjADQEcA2AAgaDGps2rcHAgSNw86YBMpkGWm00tNpMpKePhcUSCEn6DsHB\n7+Wvj4Vky5YtGDnyY1itngCCAQDkcJjNi3D69GnUq5fPsRYICkMRfnA8E4qhywJBkTF48Ejq9VUp\nl4+hXl+d/fq9/dg6u3btolJpyLaHbTvwdf78ed64cYN+flXp4FCZen1ZNm7cOs/88TmwWsng4Acb\n4uSiRQ8VWbt2LY3G/8uxAqHVuvDGjRuPNL1hwwZKkheBTQTCKUnluGjRzznKREVF2Zf3dxG4Qpms\nHT08vBkdHc3hw9+nWj3Evh1BKpUfs23bLhw8eAQ7dHidCxcuLlSin/xw8OBBVq/emDqdO4FP7WcI\nkuzjcJ+SVIrnzp17qj4IXiwKo32PrTl9+nTeuXPniRsoaoTQC/7LWK1Wrlu3jl9++SXXrFnzWMHa\nsmULtVpn2tK8/iO4jo6NGBERQZJMS0vjoUOHeOLECVoslsc7YbGQQ4fSrqLkqlU5/Nu7dy9/++03\n7tu3zy7GOwlkUiabRh+fCvkS2ZUrV7FKlZdZsWIDzpu3INcyYWFhdHHxolyuYPXqLzM2NpYk2bJl\nVwIrsvU3nNWrN318v4qIv//+mwaDG4GlBN6yby8MI1CdwMeUyyuzV68BT/1jQ/Bi8VSF/qOPPqKf\nnx+7devGzZs3P/f/OYXQC4oLVquV0dHRPHfuHDOf8jWzvPD1rUZgHW3R3JbbZ7mb6eDgztv2mPMF\nIiOD7NvXpqAaDblxY9Yrq9XKnj3fpF5fjkbj/1GSXDlp0iSWKFGaMpmc5cvXfCqx4P/3d9KkSd9Q\nklrY9+3N1Gq7cMSI4CJvNy8WLFhAvf4N+0fGVfvY9yXwGpVKHT/++OPn/ntUUPx4qkJP2k64bt68\nmT169KCfnx/HjRvHqKioJ260MAihFxQH0tPT2bZtV+p0HtTry7By5XpPJqyFxMXFh7YobkcJ+BNQ\nUKcrwV27dhXcmNlMduliE3m9nty+PcfrsLAw6vWVCZjsIhdBZ+dSJPlE99ZJW3jYLVu28Pr16/mu\nk5GRwe7d+1Gl0lOtdmDLlq/RZDI9UftPwooVK2gwvJq1dQAcpFyu4ieffMpjx449Mz8ELxaF0b58\npamVy+Xw9PSEh4cHFAoFEhMT0bVrVwQHBz+towMCQbFm8OCh2Lw5HqmpfyMlJQYXLtTBO+8U7O+L\nxWIptB9t27aGVjsGQGkAy6HTeWDz5rVo0qRJwQylpgKvvQb89pstKc22bUCLFjmKxMbGgqyHf5LN\nvIy7d28gIyMDSmXBzv2SxFtvjUCjRh3Qo8e3KFeuCiIiIvJVV6lUYtWqxYiPj8P165exdes66HS6\nx1V7ItLT07F3717s27cP6em2A4MdOnRAqVIJ0GheB/AdJKk3vvjiS0yc+EVWUh6B4JnyuC+BadOm\nsWbNmgwKCuKqVauy7tBaLBb6+vo+8RfGk5IPlwWC58qOHTuoULgQmJdtn3g//f1r56v+5s2bs5a7\nAwPr8NKlS0/sS0pKCrt3709JcqaLizcXLfqZGRkZDA0N5dKlS3n58uXHG0lOJps1s3XExYXMnoI2\nG9u3b6da7W5fQSBlsuksX77GE/kdHh5Ovb4CgWT7+G2li4vXE9l6Wty5c4cBATXp4FCNDg5VGRhY\nO+s8U3JyMidPnsJhw97lunXrnrOngheBwmjfY2uOHz8+z18GZ86ceeKGnxQh9IJ/O927v0mgA4HX\nCGTYl3A/YMeOrz+2bkxMjP0AWwSBDMrl37Js2cpFtqdrNpvZoMGrNBjq0GDoQb3e9dHL+HfukPXr\n20S+ZEkyj7/zly9fppubD9XqagQ0BAwsXdr/iffk586dS0kakO1DyUKZTEGz2cwJEyaxdOlAvvRS\nVf7889Insl8YrFYrz58/z44de2Q73W+lWj2YQ4aMeub+CP4bPFWh/7chhF7wb6dPn8EEviIQRKAc\ngUpUq13yld1s9erVdHDolE3grFSrjUW2v79gwQJKUnMCmXb7v/OllyrnXvjWLZorViQBXtPoOPO9\n4Dz32jt16k2FYoLdZhoVikEcNGj4E/t54MAB+xW7v+0257Fs2cr8+uvvKUk1CRwhEEFJ8s4zu93T\nIDExkX5+1SiXuxDwoC1N7oP/Vuv58svtnpkvgv8WhdG+fO3RCwSC/PPee29Dr/8BQAsAXaDRXMfK\nlfNQsmTJh8rGxcXh44/HY9SoD7B37164u7uDPAcgzV4iCoAFRqOxwH7Yfjfk5Nq1a0hLqwtAYX9S\nH7duXXu4clwcLI0aQX32LC6gBOqaf8SHcw9h8OCRubYVG3sNFkt9+08aWCwtcPny9QL7/IC6deti\n4sRgqNWVoNf7wMNjEjZuXIklS36DyfQ9gJoAmsJkGodly9Y+cTsFITMzE1Wr1sGlSzJYrbEA+gNY\nBCADQAa02mWoXz/3THsCwfNECL1AUMTUqFEDe/ZsQ+/el9C9+y2Ehf2KTp06PVTu6tWrqFq1Hr7+\nOgkzZiShadM2CAmZh6ZNK8NgqA+dbhAkqSlmzJgGVT7TvALArFlzYTC4QqXSok2brrh3717Wu4YN\nG0KrXQEgBoAVSuUU1KvXKKeBmBigcWMoLl7EabkRTXAWVzEQJtM6LF26MNdDgi1aNIJONw22dLVJ\nkKSZCApq9FC5gjB69EjEx8fh1KlduHLlAipVqgSDQQ/gnw8Imew6jMZnk6r39OnTuHEjEUAnABKA\n8QDuAnCDVuuFevVM+OKLT56JLwJBgSi6hYVnQzF0WSDIlXHjPqFCMdIe3KUkgS8I9Kanpy9/+eUX\nzp49+7GJYK5fv87w8PCsKGvbt2+nJPkQOEvgPjWaPuzaNWfs+qlTZ1CtlqhQaFi7dlPeunXrn5fn\nzpGlS5MA43196a1vk21pOpEKhTrXmABms5nduvWlQqGmQqHmgAHDnkrsgJ07d1KncyXwOeXy0TQa\n3XOcAzhy5Ahff/0tdu3aj9v/5/pfYTl+/Dg1Gg8CtQncs4/JVwwMrMXLly+Lu/GCp0phtE9mN1Bs\nkMlkuS5JCgTFjZEj38ePP7oCWAxgIQDbDFij6YtJk6pj9OjRj6y/detWdOnSG0plZaSnn8eIEYMg\nl1swZYoawGf2Upfh7NwYd+5cyVHXYrHAbDZDkqR/Hp44AQQFAfHxQJMmuPPzzwis1wwJCb1hsdSG\nJE1Dz56VsGDBT3n6ZDabQRK3b9+GwWCAk5NTQYflsRw5cgTLl6+GRqPGoEEDULZsWQDA0aNH0bhx\nK5hMYwDoodN9gV9/nY927doVSbuZmZmoXbspTp1KgtV6A4ADVKoknDt3CH5+fkXShkCQF4XSvqL5\n1nh2FEOXBYJcCQ0NpVzuTEAiEJM1c5bLx/Lzzyc8sm5mZiYNBhcCe+z1blOSfDh69GhqtZ2zBWv5\nneXK5eOK219/kU5ONgdatSJTUkiSsbGxfP31t/jyy+341VffPHaWfu3aNQYG1qZO506VSs/33hv7\n0Ew3IyODP/wwnb16DeSXX05mamrq4/3LB717DyLwbbYViF9Zp86rRWL7AUlJSRw+/H3WqtWMvXr1\ney5BkAT/TQqjfSJ7nUDwHLBarRg16mNYrV1gOxj3NoBpAGKg1S5E+/Zhj6yflJSE9PQMAC/bn7hA\noaiLKlWqoEyZCFy92hpWqw9ksvUICVn9aGciImBt3x7ylBSsgwzBZ2Kw9MQJNGjQAN7e3ggJmQaL\nJX8HAnv3fhsXL76KzMxJAO4gJKQZGjWqjS5dumSV6dHjTWzZEgeTqQd0ui0IDW2PP//cCoVCkbfh\nfGA2Z+CfYD0AICEzM7NQNv8Xo9GIH3/8rkhtCgRPnSL84HgmFEOXBYKHaNiwhf2+eQKBNAKjCLjT\nxaUsN23a9Nj6VquVbm4+BNbYZ6+XqNN58NSpUzSZTFy6dClnzZrFCxcuPNpQWBitWi0JcClqUoEk\nAmvp4ODOa9eusX//t6lUaqlUSmzZstNjQ8k6O5cmcDnbrPoLjhkzLuv9uXPnqNW6ZAuTm0mDIYAH\nDhzI17g9ij/++IM6nQeBVQRCKUl+XLhwcaHtCgT/BgqjfcVONYXQC4o74eHhdpF/ibbMbg/uyzfl\nolzSvebFoUOHWKJEaRoMZanRGDlr1tyCObJmDa0qFQlwDjSUwZIl0EZjczZt2pxKZV3a0quaqdV2\n5bBhox9psmrVRgTm2+2kU5KaMyQkhJcuXWK5ctWoUGgJOBM52qrD3bt3F8z3PAgLC2P9+i1Zq1Zz\nIfKCF4rCaJ84jCcQPGUyMzMxduxnWLVqPQwGA+rWrYAlS34DMBPAh7Bd1zoBb+8kXLx4HBqN5rE2\nSWLlypU4fPgoXFyc8fbbb6NEiRL5d2rpUqB/f8BqxVQ0wPs4CeAigJIAUqFQ+IDUwmr9CkBfe6Vd\nqFz5Y5w69WeeZk+ePImmTVuDrAiLJQ516pTD1q1rUbVqA0RG9oTV+h6AOgBqAxgEhSIMnp6/IDLy\neM6DgXmQkZGBhQsX4tKly6hTpya6du0KmUyW/34LBMWUwmifEHqB4Cnz7rtjMG/eQZhMPwC4ArX6\nDVgsLrBY3AC8DiAcMtkOXL4cCR8fn3zZ7NSpFzZvPgOzuTskKRzNmrkjNHT1Q6IXGxuLS5cuwd/f\nH15eXraHc+YAQ4cCAL7Tl0Bwyh4AGwCEAGgDhWIbyHhYrQMA3LM/l0Eun4i2bc9g48aVj/QtISEB\nBw8ehNFoRIMGDZCZmQmdTg+r1Qxb6I4EKBSN4eycjtq1ayAkZCq8vb0f22er1YpXX+2AAwfSYDI1\ng16/GoMGtcUPP0zJ15jlZk8uF6FEBMUDcepeIHgGWK1Wzp4dwrp1g9i8eUfu27cvX/Xc3X3t99of\n7Ft/Qm/vQKrVXlQofKlQOPHXX3/Ntx/jxo23n9S/mxVyVq/35ZH/STYzZ8486nQudHRsTJ3OxRYX\n/ttvHzjBSSVK0dMzgErlGPsp/U1UqcqwV69eVCq97OcHqhJ4mUAjKpWOjImJKciQkbSNm4ODK4ED\n9qZTaTBU4ebNmwtkZ8+ePTQYAmnLH2C7aaBS6ZmYmFggO7dv32bjxm0olytpMLhy8eIlBaovEDwP\nCqN9xU41hdALnhfffz+dklSRwEYC8ylJrvnKL16mTOVse/GkSjWEX3wxkdu3b+evv/7Kq1ev5tuH\nmzdvUq02EPDJdoWONBjqMyIiIqvc1atXqdOVIHDRXuYMJyo0WSI/TNaBwEGqVK9To3GlXv8SNZoS\nbNu2C5csWUKNxpXAJwROEhhAoASrVGn0RONGkuvXr6dO50qDoTs1mvL09CzP/v3fZnR09GPr/vzz\nUjo6elIuV1KhqJ/tg8lKnc69QONHki1adKBK9Y79EOQJSlJJ/vXXX0/aNYHgmSCEXiB4BpQpU4XA\n/mxC8znfffeDx9ZbsWIlJakUga+pVA6nq6s3r1+/nqPM/fv3+dFHn/G113rz66+/yzN5zKlTp2gw\nBBCoQuAz+/37GXR09GRSUlJWub1799LRsW6WIH6H0SRAi0zGwdqAbH3IpEZTgrt37+bnn0+kJHnS\nwaELlUonymSuBLwJvEyttiK//356ocYvMjKSPXu+QY3Gm8AiyuXj6eRU8pFCvXfvXkpSSQJH7af5\nHQnMJRBDpXIMAwNr02KxFMgPjcaBwJ1sH17v8euvvy5U3wSCp01htE/coxcI8ontnnd61s8yWTqU\nysff/e7Zswc8PNzx228b4ejojOHDD8DT0zPrfWZmJho3bo2zZ71gNrfCtm2/YN++I1i/fvlDtnx9\nfaFS3QPwCYBNAOZALk/Hli1hOe65lytXDhkZlyDDYczCfLyNuUgHcHLMGCyfuRmAFbb9chPIDHh4\neGDy5O9gNp8AUAbATSiV5VGihCNksusYOnQA3ntvxBON2wP8/f0RHr4HZnMYgMqwWgGT6QaWL1+O\n4ODgXOvs3LkTZnMfADXsT9ZDJusEJ6eJqFmzFpYt21jgfXZnZ3fcuHECQDMAVqjVJ+HuXvXJOyYQ\n/Nspwg+OZ0IxdFnwgjBv3gJKUlkCP1Mm+4Z6vWtWjPnCsHfvXhoMlbNdOTNRoymRZ1rbw4cPs2RJ\nP8rlKrq4eOV5NW3NytX8RaEmAZoA/vXZ50xPT2eNGi9Tq+1GYA4lqRH79BnEkydP0sGhQraZPuno\n2CDf197u3r3LJUuWcMGCBYyLi8vx7sSJE2zSpB0DAupSo8m+nUAqlSM5efLkPO3OmTOHktQu2zbF\ndpYuHZAvn/IiNDSUOp0rdbq3aDA0Zu3aTWk2mwtlUyB42hRG+4qdagqhFzxPVq/+lW3b9mCPHm/y\n5MmTD71PTU1ldHT0YwPLZCciIoJGY11mX07X6Tx5+fLlR9YzmUx5J1JJSyM7dSIBZkoSU7IF4UlJ\nSeGECV+yV6+BnDlzFi0WC1NSUujo6Ml/8qtHUK93ZXx8/GP9v3XrFkuX9qde356S1JOOjp48c+YM\nSVsIXQcHdwKzCPxJhaI8FYrqBLYRmE293pWRkZFZthYvXsJSpQLo4uLDkSODmZyczMqV61GvD6JG\nM5SS5FYk+efPnDnDWbNmceXKlULkBcUCIfQCwb+ArVu30mBwoV7vTb2+RL4FKSUlhd7eAVQqPyaw\nhxrNQNau3fTJs6GlpNji1QO2+PX5PGi2b98+OjuXolrtSAcHV4aHh+er3qhRwfbDbbYmZbLpbNGi\nI0ly9uzZ1On6ZfuIuU25XMvq1ZuxRYuOPHr0aJadrVu3UpK8COwlcIGS1JTBwZ8wNTWVS5Ys4fTp\n03nq1KmCj4dA8AIghF4geM7cvXuXBoMrgV12QdtHSXLJ14yYJOPi4vjaa28wMLA++/V7m3fv3s3x\nPj09PX+HzpKSyCZNbKrq5kYeP57jdUZGBkePHsfSpSvQ378WN2zYkOO9xWJhfHx8gVLMdu7cl8CC\nbGK+h4GB9UmSCxYsoCR1zvbub2q1xlw/YgYNGk7g+2xlD/Gll6rl2w+B4EWmMNonokUIBEXApUuX\nIJeXAtDE/qQBVCpfXLx48bF1SSI6Ohp9+nTG1q2rsXjxbDg6OgIAUlJS0LZtN+h0Bmi1BowfPzHv\noBl37gCvvgrs3g2UKmX7Z7VqOYp8+OGnmDNnH+LiVuHixYno0eMt7N+/P+u9XC6Hq6trgRLMtGnT\nFJL0I4DrAO5Bp5uMVq2aAgA6d+4MR8cTUCpHApgPSWqHDz8MzjWanbOzEQpFbLYnf8NodMi3HwKB\nIA+K7HPjGVEMXRb8B7h16xa1WqdsB81iqNWWeOwdb6vVyi5d+lCvL0+jsQP1eldu3749633//kOp\n1fYgkEogjpJUkStXruT69evZunU3duz4hi0hzI0bZJUqtqlw2bLkpUu5tufhUY7A6Wyz5gn84IMx\nheq71Wrlhx9+QpVKR4VCzR49+jMtLS3r/c2bNzlqVDC7devPxYuXZM3mjx07xt9//z0rCE9cXBxd\nXb2pUr1FmWwcJcktx1gIBP9lCqN94nqdQFAEuLm5YejQgZg5szrU6gqwWmMxZcpElC5d+pH1QkND\nsWXLSaSknACgBbAdPXsOQHz83wCAHTt2Iy3tF/u7UjCZhmD27AU4dOg8TKYvASTj7NbWOOnuAG1s\nLBAQAGzfDjwId/s/SJIetpl3JQCAUnkdDg6lCtV3mUwGk+keLBaCJM6cOY+EhASUKmWz6+7ujmnT\nvslR5733xiEkZBmUyqrIzDyIJUvmokuXzjh16iAWL/4ZJlMqOnfehurVqxfKN4FAgOI3PS6GLgv+\nA7z//kfU6/2o1Q6kRlOGgwYNz1e9mTNnUqsdkm2GnU65XJG1H1+3bgv+kw3OSrW6Dz08yhMIJUD6\nIooxcLZVrlaNvHkzh32r1crjx4/zjz/+4J07d7hmzRrqdJ4EvqJS+Q5dXb3zvMaXX954oz+BEgQi\n7dfgxrFu3eZ5lj906BAlyTtb0Joj1OkcmZ6eXig/BIIXmcJo31Pbo09LS0O9evVQvXp1VKxYEePG\njQMA3LlzB0FBQShfvjxatmyJu3fvZtWZPHky/P39UaFCBWzbtu1puSYQFJqzZ8+iceO28PWtjq5d\n38BPP4UgJeUA0tLmw2w+jKVLl+PKlSuPtVO7dm3I5aEAYgAQcvkMVKhQKysIzNy538HB4SPo9b1g\nMAShTJmTcHFxAaBEBZzDbjTBS0hElKs7sHMn4O6eZZsk3njjLTRs2AGvvTYeZctWxEsvvYQtW1Zh\n5Mg7GDfOFdu2/Y5Ro8ahevWmGDZsNFJSUgo0DgkJCVi5cgWAXgD8AcgAfIwjR/bmWefvv/+GUlkT\ngLP9SU1YrQokJiYWqG2BQJBPiu5742FSUlJI2k761qtXj3v27GFwcHBWuMkpU6ZwzBjb/uCZM2dY\nrVo1pqenMyYmhn5+frmeMn7KLgsEj+XmzZt0cipJmWwmgcNUqVpSocgeVpY0Gqvy8OHD+bI3Y8Ys\nqtV6ajQl+NJLlXjpf/bXr169ysWLF3PVqlVMSUnhokU/s4G2FG/BSAKMkKt4IJe97DVr1lCvr0Eg\nxe7Xcvr5/XOK/f79+/TyKk+lchyBHdRqe7Jp07YFutYXFRVFjcaFQH3+k2xmO52cSj9Ubv/+/UxK\nSmJkZCR1OrdsZwVW0s3Np8ChbAWC/xKF0b5nopopKSmsXbs2T58+zYCAAN64cYMkef36dQYE2KJc\nTZo0iVOmTMmq06pVK+7fv/9hh4XQC54zK1eupINDx2zCHk9AR2CVXexW0Nm5FO/du5dvm2lpabx5\n82b+RHbfPpoliQT4l4sHd2/dmmuxr7/+mkrl6Gx+3qVarc96v23bNhqNjXJsG2g0zll/P/NDRkYG\nvb0DCFSz/+lCQM81a9aQtG0dDB/+AXU6dxqNtejkVJJHjhzhkiXLqNUaqdN50tXV+6HMewKBICeF\n0b6ner3OarWievXq8PDwwCuvvIJKlSrh5s2b8PDwAAB4eHjg5s2bAIBr1679ky8bgJeXF+Li4p6m\newLBE6HVakHeAfDPNTeFIhOlS38CmUwDb+8J2L59IwwGQ456UVFRqF79ZWi1RpQvXxPHjh3LeqfR\naODu7p7rtbMc/PEHEBQEtckEdOmCetdi0bhly1yLVqtWDRrNRgDxAACZbBEqVPjnup1CoQBpztaP\nTJCWAl2tUyqV2LVrM2rUMEKlioSHx2GsX/8LunTpAgAIDw/HokUbkZp6AcnJh3H37lR07twHffq8\ngTt3buDChYO4fj0aNWvWzHebAoGgYDzVU/dyuRzHjx9HUlISWrVqhZ07d+Z4L5PJHvmLLa93n3/+\neda/N2vWDM2aNSsKdwWCfNGyZUuULv0FLl/uC7O5PvT6BRg0aDR++GEKLJbchTIjIwPNmrXFtWvD\nQG7ExYub0Lx5O8TEnIWTk1Ou7WzYsAEhISug12sxduxI1Lh2DejSBTCbgb59gQULAGXef4VbtWqF\nESNex9Sp5aBSlYCTkwq//RaW9b5Ro0YoXZqIjn4L6ektIEmL0br1/8HV1bVA41G2bFkcPbo713fn\nz59HZmYLAA/62AlXrvQGSeh0Onh7exeoLYHgv0JERAQiIiKKxljRLSw8mi+++ILffvstAwICslJ0\nXrt2LWvpfvLkyTmSW7Rq1SrXHNHP0GWBIE+SkpI4fvwE9ukzmIsWLX7skvv58+dpMPj+T9KYRty5\nc2eu5W2pbW3pXIGp7K1xoFWptFUcOpQswH52fHw8IyMjcz3VfvfuXb777ods3bobv/xySpGffN++\nfTv1+nIEbtv7/TN9fasUaRsCwX+BwmjfU1PN+Ph4JiYmkrQl32jcuDG3b9/O4ODgrL34yZMnP3QY\nz2w2Mzo6mr6+vrn+8hRCLyiO3Lx5kxqNo30/nwRSKEnePHHiRK7lK1VqSGAzAbIfFjETMpvIBweT\nBYyBb7FYuHbtWk6dOpV79uwpiu4UiA8++JhabQkajZXp4pJ3nwUCQd78K4X+5MmTrFGjBqtVq8Yq\nVarwm2++IUkmJCSwRYsW9Pf3Z1BQUNbHAEl+9dVX9PPzY0BAALds2ZK7w0LoBc+QpKQk9u07hH5+\nNRkU1OmhE/EF4YMPPqZeH0Cl8gPq9TXZq9eAPFcCAgPrE9jBYZiZtQSwtmbdAot8eno6X365JbXa\nalSrh1OSfPjttz88cR+elCtXrvDYsWNZN3EEAkHBKIz2yewGig0ymSzvWN8CQRFCEo0bt8bhwyVh\nNg+DXL4TLi6zEBl5Is999ccRFhaGkydPwt/fH507d87zHEpIyHxcGT4WEzMSAABjVXp02rMD9erV\ny3dbqampqFWrEc6duwUgCrboerFQqQJx9248JEl6oj4IBIJnT2G0T4TAFQjyICEhAYcOHUB6+m0A\nSlitdWE2h2Pv3r1o167dE9ls3bo12rRp8+jT9SQGX/kbyEiAFcAP5auiTciMAok8AEyf/iOiolQA\nqsEm8gDgDYVCQnJyshB6geA/gsheJxDkgVqtBpkBwGR/QpBJUKvVBbaVnJyMRo2CoFJpIUnO+P77\n6bkXJIHRo4EvvwQUCsiXLcP7F06gadOm+W4rLi4OISEh2LhxGzIyOgA4BGCTvR9T4OnpAfdsEfQE\nAsGLjVi6FwgewcCB72DlymMwmfpBo9kFf/9oHDmyO99in5iYiDfffAcbNmwGabfYDeoAACAASURB\nVATwGwBnaLVB+PXXGWjfvv0/hS0W4O23gfnzAbUaWLkS6NSpQP6eOXMGDRu2QGZmS2RmnkJ6uhXA\nFAAjAVyG0eiJ48d3o2zZsgWyKxAIni+F0T4xoxf8J7h8+TIaNWqFEiW8UK9eC0RFReWr3rx5P+Lb\nb/uiV69DGDOmAvbv355vkSeJVq06IzTUAHIvgDEAOgJwQlraEISHR/xTOCMD6NPHJvI6HbBhQ4FF\nHgBGjfoY9+59ApNpCdLTj0AmU0Au7wqtNgk1atTHokXTsXDhIsyaNQtpaWmwWq2YOnUGmjbtgF69\nBiImJqbAbQoEgn83YkYveOExm80oV64qrl0bAKu1J+TytXB3n4lLl0491X1qW6pWP6SnJwB4EESn\nDYC3ASzF5Ml1MHbsGCAtDejZE/j9d8DBAQgNBZo0KVBbW7duxebN27Fixe+4dWs+gAf1F6Ft2y0I\nCZmKwYPfQVjYPgCDIZcfQOXKKWjSpCEWLvwTJtOHUChOw2ici3PnjmZFrxQIBP8OxIxeIHgEFy5c\nQFKSHFbrGABlYLW+B5PJiNOnT+coZzabER8fX2QfklqtFlZrOoAk+xMrgFgA70Ol2gJHRyPSExOB\nDh1sIu/sDOzYUWCRnzVrLjp3HoLp011w+7YeMtl4AHcBXIVePx2dO7fG3r17ERa2BcA+AF/Cat2G\ns2ctmDPnJ5hMawF0hsUyHmlpzbB+/foi6b9AIPh3IIRe8MJjNBqRkZEA4L79SSoyMm7BaDRmlZk9\nOwRGowu8vQPg61sF0dHR+bKdnJyMN94YBB+fymjUqDXOnDmT9U6v12PYsOHQ61sAmArbbD4WQAKs\n1sH44v2VOOtTFggPt6WXjYgA6tQpcP/GjfsMJlMogLGwWg9ALr8GudwTGk0gRo58DQMG9EdY2A4A\nFgA+9loyWCw+sMW5z34DoGhXzKKiojBjxgzMmzcPSUlJj68gEAiKnkLc338uFEOXBc8Rq9VKq9XK\n3r0HUa+vQ+AL6vUN2KVLn6xgNcuXL6dC4UogigApl3/HwMA6+bLftGlbajR9CRynTDaLjo6evHnz\nZo72ly1bxiFDRrJ9+44E1ATOsARu8xBqkQBTXFzICxeeuI8ajYFAQlZoXY1mKKdOncqbN2+yWbP2\n1Omc6ODgTsCPwGACVwisJyCxXr0mlKQGBH6nXP4lnZxKZoWoPnv2LGfOnMlly5YxNTW1wH7t37+f\ner0rNZohlKTOLF3an7dv337ifgoE/2UKo33FTjWF0AvyQ0ZGBgcOfIdqtZ46nSPHjfuMS5cu5Ycf\njuPPP/+clfv8/PnzVKv1BPpni0OfQZlMzszMzEe2ce/ePSqV2mx52ElJasdPP/2USUlJOcpGRUVR\nq3UloKQH/uZJVCYBRskc+MtXXxWqr1279qVW25nAOQJrKEmu3LZtGyXJk8Bwe9jdVQT0BLwJONr/\nfEmt1pnjxn3KRo3asmvXvoyKiiJJbtmyhZLkSq12EPX6FqxcuR5NJlOB/KpZsymBZVljo1IN4scf\njy9UXwWC/ypC6AWC/+Gjjz6nJDW3i1wsJaka581b8FC5Dz8cR6CrPZd6ql2U/qCLi1dWmQMHDvCd\nd97j6NEfMjIykiT5yy/L2aHD65TJVNni11sJVKNOV5FubmWyRJMkw8LC6OgYRG+8zEg4kgBPowzL\napx5/vz5QvXVZDKxf/+h9PAox4oV6zE8PJwlS/oSUBHIzBJarbYzVSpnAnMJXLYn1qnLvXv3PmTT\n2zuQwNasful07Th79uwC+eXjU5nA0WwfUNM4cOA7heqrQPBfpTDaJ/boBS8koaE7YDJ9DMAVgDdM\nptHYuHHHQ+WsVgIIAFAVtghy/weZ7P+wfPkCAMAff/yBV15pj59+csUPPyhQq1YjBAePxaBBE7Bh\nQwuQdQE0APA+gOYAVEhNPYqEhBEYMGBkVjvlypWDd9ph7EEU/JGEo5DjFdzGtNWLERAQUKi+6nQ6\nLFo0CzduXMSZM3/Bw8MD9+4pAagBXH7QUwB/Qy7PBFAFQBkAR5GRcQnlypV7yGZiYry9HADIYDZX\nxa1b8QXyq127IOh04wEkADgPSfoR7du/+iRdFAgEhaEIPzieCcXQZcFzoGXLzpTJpmfNJpXKDzhk\nyMiHyp08eZKS5EpgNoFJVKu9ciwv16/fksDyLDsy2QSq1c4ETtufxdiXwV+yL4s3IHCPwCmWKlWe\nN2/etG0TnDrF+0YjCfAvhZ6lpBIMDd2Uwxer1cpdu3ZxxYoVvHjx4hP3PTY2llqtC4Hv7T59QKA+\nq1VryA0bNlCvd6GDgz8lyZm//bY2Vxvt2nWnWj2QQAqBk5Sk0ty9e3eB/EhNTWWvXgOo0Rjo4ODG\nqVOnP3GfBIL/OoXRvmKnmkLoBfnh9OnTNBo9qNP1oyR1o7t7GcbFxeVa9sCBAwwK6sz69Vtx9uyQ\nHBnlKld+mcD2bMvPs6lUGu0Cn0HAh8Db9mV7C4FuBMZSLg+iXK6lRlOCbT3KMNPJiQSY2qgR/9q+\nnfHx8Tl8sFqt7NnzTer1AXRw6EpJcuX69evz1Ver1cpvvpnKsmWr0d+/Nn/5ZTkHDRpBvb4agT5U\nqUqzbt2XaTabSdrOFpw9e5bJycl52kxMTOSrr3akQqGmg4MbFyxYlC9fBALB06Ew2icC5gheWK5e\nvYrQ0FAolUp06tQJLi4uBbYxdeoMfPrpAphMcwHchyS9iVdfbYDt26/DZOoFYAKAXwA8WJJeBZls\nKGxx8Q+gIeIRhlfhiDSgXTtgzRpAq83RBkn8+OOPGDNmFtLSjgHQATgIvb4NIiK24syZMyhfvjwa\nNGiQq4/Tp8/ERx+FwGQKyfJx9eo5SE1NxalTp1GhQgB69OgBubzgO3UkH52ARyAQPBMKpX1F8qnx\nDCmGLguKMVarlVOmfJdjtpyZmcnPP/+Kvr5VCXjYT+xbCKQTaM369RtTr+/HFgjnfUgkwNWQMS2X\nGbTFYmFQUEcqlS72Q4EPVg6slMlU1OlK0WB4g5JUhsHBn+TqY9WqjQlsy1Z3Drt16/+0h0YgEDxD\nCqN94jCeQPAIZDIZxox5H9HRxxEZeQgdO3ZA1659MWnSJNy+fRMuLhoAuwB4AygJne4oBg7sizaZ\nO7AJ7aCHCYvwCvooHdC991AsXrwk66ucJF55pRXCw08jMzMMwG4AZ+3tzgCpRWrqAdy/vwwm0xHM\nnBmCixcvPuSjXi8B+OegnEwWD4NB97SHRiAQFBOE0AsEBeCtt0Zi82Yr0tOvIDl5K0ymDDRvXhEe\nHnrI5WYolf7YNexdrEiPgwbpmKPywUD8hQzrYGzY8CqGD/8WEydOBgBs2LABf/11DkALAHUBfA+g\nPgAtSpb8CXq9CwAve8suUKvL4fr16w/5VK9eJQCDAUwCMBaSNB3BwSOe/mAIBIJigRB6gSCf3Lhx\nAxs3/g6z+VUAegDVkJY2BIGBfkhKug2r9Ri63huAnzNMUJKI6toVR/u1hEbbEVbrtwD6IyVlHb79\n9gcAQHR0NKzW+gDCAPwNoDeAT+HvH4iLF49DrU4HsAa2MLXhsFguomLFijl8Onz4MEJCVgD4GbZZ\n/UF4epZEYGBgkfX7wIEDCArqjAYNWmPOnHnijIxAUMxQPm8HBILnze7du3HgwAF4eXmhe/fuUCgU\nSEtLQ1paGpycnAAAx44dQ7NmbWAy1QQwG8BiAFuh0ZyFQuEJtdoXg9LCMAOjAABfaDzRdswYVN63\nD+SpbK3pYbFkAgBq1qwJtXoGMjOHAKgEQAOl0opNmw5CkiRs2/Y72rXrhjt3+sDBwRnr1v0KV1dX\nAEBqairOnTuHzZs3g2wPoIv9TyZiYrSwWCxQKBQoLKdOnULz5u1hMn0FwBOnTo2DyZSK0aNHPrau\nQCD4l1BUBwWeFcXQZcFz4sqVK+zUqTerVm3MYcNGMyUl5aEyU6fOoCT5UKV6j3p9Q7Zo0YFjx46n\nUqmlSqVnnTrNePv2bVat2ojAQvthNwuBNlSrK7JcuaqMiYnheJXuwUk4jsQo6vUuTEhI4OXLl2kw\nuBGYSeAPSlJTDh78z33+iROnUKXSU6FwpVZbgrNmzcrhn9Vq5b1793Jc+btw4QI9PMrSwaEy1WpH\nKpWBBNLszUewRInSRTaG778/hsD4bAf9/qKPT+Uisy8QCPJHYbSv2KmmEHpBfkhOTmbJkn5UKD4l\n8AeVynrU671YqVJDLl68hCSZnp5OlUqy34kngXRqtT7UagMI3CSQSaVyAB0dvQkYCERmE7wpbNGi\nNe8lJ5PjxpEALQCHadwoSSW4cWNoli8nTpxg8+Yd6OHhR1/fGhw0aHhW4pvLly/TwcGNMtkIAjMp\nST5cuHDxI/tWo0bjbMGA7lKhKEmt1p8ODp0pSa7cvHlzocbu+vXrPHfuHM1mMz/4YCxlsnHZ+r2H\nL71UtVD2BQJBwRFCLxD8D5s2baKDQzO7OP1KoAyBLQS2UpJe4ooVK5mYmEiVymAPdvMg8UoFApOy\nrrgB9QkMJdCLtsxvmQSuU6+vyDWrV5MjRtgqKhRMXbiQ58+f5/379x/yZ8CAYZSkxgRWUKUaRS+v\n8kxOTuYnn4ynQvFuNiHdzTJlHj1j1utdCFzPVucj9u3bjytXrmR0dPQTj5nVauXo0eOo0TjSYPBj\nqVLluHXrVur1rpTJviWwjJLkyzlzQp64DYFA8GQURvvEYTxBsSIxMRGtW3eBVmuEu3tZrF+/Ptdy\ntv3pNNgOsv0CYDKAVgBawmT6GnPnLoejoyP8/QOhUHwGIAlAGGSyK9Bq/wSQCaAngFMAPgLwE4Bo\nAAbI5WXw7oiu6BwWBvz4I6BW49bs2eixPhzdug3CuHETkJqamuVLeno6fv55PkymjQB6IiNjGpKS\nymDbtm1IT8+AxeKQzXMjMjIyHjkG/v6BkMnW2H+6D71+C9q1a4sePXrA29sbsbGxSElJKeDIAmFh\nYZg7dx3M5ku4fz8KN24Mw4cfTsT+/X+ga9czaN36d8yfPwlDhgwqsG2BQPAcKcIPjmdCMXRZUIQE\nBb1GtXoQbfnX91CS3Hns2LGHyqWmptLfvzrV6rcIvExgVrYZ8Fy2bt2VJHn16lXWq9eCarWeXl4B\n3Lp1K+vUaUat1pdAZQI17CsCtr15na45Z02fTvbo8SAvLe+tXUt39zJUKL4g8Ae12s5s06ZLDl8U\nCrU9brytmoNDO65atYpHjx61x9pfRmAnJak2x4+f+MgxOH/+PN3dX6LRWJU6nQf79h1Cq9XKEydO\n0N29DCWpFDUaB86ZM69AYztp0iQqFMHZximBGo1DgWwIBIKnQ2G0r9ipphD6/y7bt2+nLfXq3Swx\nUquHc+rUqSRtOegvXbrEW7dukbTFax81KpgNGrSgSuVEYAqBrylJrrmmZn1Aeno6hwwZQpVqKIF9\nBNwIdCFQgY3rNKWlXTtb40YjuWcP161bRweHltkEMo1KpS5HLPkuXfpQp2tLYCsVigl0c/PhnTt3\nSJIRERGsX78lK1VqyEmTvrUlwXkMKSkpPHz4cFbyG6vVylKlyhH42e7DRUqSJ0+cOJHv8V29ejX1\n+prZPkh+ZkBArXzXFwgETw8h9IIXnhMnTthnvu4EDmTtoev1QVy8eDFjY2NZtmxl6vXeVKuNHDky\nOMdJ9cOHD3PAgGEcMGAYDx06lGc7sbGxXLduHefOnUtJ8rIf1IulTNadlV+qQEvz5jY1L1GCV9av\nZ/36r1KvL0G5vHa2vf4kKpXaHKf8zWYzg4M/Ya1azdmpU29evny5SMcnKSmJSqWU7WODNBh6ccmS\nJbmWP3jwIEeOfJ8ffDCWUVFRJG0fC716DaAkedHRsQGdnUvx+PHjReqnQCB4MoTQC154Jk78kgrF\nBwRW0RZf/l0CjVm1agOmpqayQYMgKhQT7GKbQL2+Mn/77bcCtbFt2zbq9a40GttTry/HmjUbUaWS\nqFY7skbZykytVcumoJ6eTD10iCVL+lEu/47AOQJeBN4ksJSS1Jj9+g15SiORO1arlQ4OrgT+zDqN\nr9f7cs+ePQ+V3bFjByXJjcBEyuUf0sHBnRcuXMiyc/LkSUZERDAxMfGZ9kEgEOSNEHrBC8/UqVOp\n0fSxi9ghAsNpNLrRZDJx5MhgAjoCcdlmtOP5ySefFqiNEiW8COyw10+hwVCZv//+OxMuXKC1Rg2b\nYW9vMjKShw8fpoNDlRz72TKZKxs2bMXvv5/GzMzMpzQSeRMWFkZJcqWjY0tKkhffeef9XMvVrfsq\ngRVZvstkn3Hw4BHP2FuBQFAQCqN94tS9oFjQt29fODn9CaXybQA7IUnrMW3at1iyZBnmz98JoCps\noWQBwAy9fgfKlfPLl+3Lly8jMLAO7tyJA9DU/lSC2VwDYfPnw7FjR8iOHQPKlQP+/BPw94fRaERG\nRjwAk728BqQFer0ao0ePKpKodAWlTZs2iIw8juXLR2Hv3o2YOfO7rHe0fdQDAFJSTAA8sr3zQHKy\n6X/NCQSCFwSRj15QbIiPj8ePP/6EhIQkvPZaW2i1WrRu3QMm00QA9WC7PlcGMtlltGvXGOvXL89T\ncK1WK06ePImUlBT07DkQcXH9Qf4KoC+AUQCi4YM6+AMm+CEN1sBAyHfsAEqWBABs2bIFr73WG2az\nF4CuAEIBBEAuX4HU1PtQq9WP7IvFYsGKFSsQExODmjVrol27dkU1TA/1c+TIYISEzAYAvPnmIPj5\nlcGECUthMs0FcA+S1B9r1oSgTZs2T8UHgUBQeAqjfULoBcWSK1euIDCwJlJSmgFwBhAC4C6Aj1C3\n7jn89dcfkMlkudZNS0tDUNBrOHo0EqmpySCTAZgBRAHoAOAa/JGGHdDDG4k4Kjfi/LQv8foIW0a4\nEydOoGHDIJhM/QFsstepCKAllMoyuHXrOm7dugUfHx/odA+niyWJ//u/HoiIiIPJ1BSStBYjRvTA\n5MkTiniUgO++m4bPPlsNk+l3AHJIUieMG9cOCoUcc+cuhVqtxuefv4/XX+9V5G0LBIKio1DaV9h9\ng2dNMXRZ8BRYsWIFHRy62O/TVyTwKoEgOjmVzDpYlhcTJ06iTteRwNcEOhBwIHDKvmdtYmV48Doc\nSYC78TJdVQP4ww8/kCSPHDnCN954gwrFaAKJBDwJ6Ak4Uqn04auvtqNWa4ssZzR65HoYbv/+/dTr\n/QmY7W3eolptyPPw25kzZ/jDDz9wwYIFuUbdexTNmnUg8Fu2swQb2aBB6wLZEAgEz5/CaJ/YoxcU\nS5ydnUFeAmAEcBBAZygUEThz5jDKly//yLonT0YiNbU9bKlhW8CWja4FgH6oDX/sQjw8kYRtaIzW\neB/3Fb8jKCgIH374KRo37ojffjsBiyUawI8AygOIBHAEMpkOu3btRlraX7h/PwrJyYvQvn23hyLd\n3b17F0qlD4AHy/uuUCodkJyc/JCv27dvR506TTF27EWMGLEW1as3wv379/M9TqVLu0OhOJ71s1x+\nHKVLu+e7vkAgeAEowg+OZ0IxdFlQhFitVqalpTEzM5NNmrShXt+UCsWHlKQy/O67afmy8fXX31GS\nWhGYS6CuPQDPMTZGVSbZp77r4UUN3CiTGbl27VqePHmSklSKwG37TN6Xtvj5O7LNlhdTqfTNcZdd\nkkoyNjY2R/u3b9+mo6MngaUEblKh+IJ+flVzDZTj61uNwKasuAFabbesAEH54e+//6arqzclqSsl\nqQdLlChdqHj4AoHg+VAY7RP56AXFhvnzF2DEiPdhNqcgIKA6Nm5cgb179yIuLg716y9E8+bN82Xn\nvfdGIiJiP3buHI/MTAsyMz3QEgqsQyokAMuhQj/oYJHdwYYNq9G+fTts2rQJKlUVAC52K4cA+AI4\nC8DWrkJxFuQdADcAeAI4DDIV7u45Z9AuLi7YuTMMr78+GLGxo1C1ak2sXh0KufzhBbbExATY9v8B\nQIa0tIqIj0/I95j5+Pjg3Lmj2LBhAwCgffsZD/kjEAhebMRhPEGxYMWKFXj99cEA/gBQC8DXKF9+\nDS5cOPJE9kgiJiYG6enp+CmoA767GgMNMjEfPhiCVwD5GqxZsxSdOnUCAMTGxiIwsBZMpi329tfA\n0fEdWCxERsZrkMlMMBh2o1+/3pg1awHU6kBkZJzGhAnj4O7uBn9/fzRo0KDAfvbo8SY2bDAjLW0W\ngL8hSe0RGroEr7zyyhP1WyAQFE/EYTzBC4+ra2kC3bMti1spl6sfeTjt3r177Nq1Lw0GN5YqVZ7r\n1q17uNAvv9Ail5MAp6ElZehHlUrPZcuWPVR07dp11OmcqNW60sXFi4cOHWJMTAynT5/On376KSvG\nfmRkJMPDw/nRR59RkkrTYHidklSGY8d+VuB+37t3jx069KJaraeTU0nOn7+wwDYEAkHxpzDaJ2b0\ngmKBWi0hI8MXwBEAGgCnoFTWh9l8L9clbwDo2rUvQkMzYDaPADASQBTq1q2FtWuXoHTp0sC8ecCQ\nIQCJUx06YujtdGh1Ggwc2B1HjpyAyZSG3r27o2HDhlk2MzIykJCQADc3t0cGxbFdrysPs/k0AC8A\nt6HVVsTp0/vh55e/QD4CgUDwgMJon9ijFzx30tPTcePGDXh6euYZaKZmzYY4cOAOgLoAqgH4HcHB\n7+Yp8gCwefMmmM07ALwCYBiARTh48Bf4+FTEaHk6vs1MsxWcPBlVxo7FnwAuXbqEmjUb4f79/rBa\nS2LRotcwadI4+Pn5oXbt2nBxcUF6ejosFstjhV6tLmkPqAMArtBo/HD9+nUh9AKB4NlSRKsKz4xi\n6LLgEWzdupUGgyslqRQNBldu27Yt13JXrlxh+fI1qFTqqVCoOXLk6Mfa9vDwJfCK/YT8gyV/Cz+G\nU9ax+CV1G+WoM3z4aMpkH2VtDwBBlMtL0WhsQ63WmVqtAyWpFB0c3Lhjx448205JSaGTU0kCa+y2\nttHBwZ0JCQkFGyCBQCCgSGojKKbcuXOHer0rgd12MdxFvd41z8AxVquVt27dYmpqar7sr1q1moAT\ngZIE0ghYOQWjSYCZkLM/JtPVtUyOOv36vU3gh6zgMkAlAib7z1sJlLL/+w4aDG5MSkrKs/2DBw/S\n3f0lKpVaOjuXZERERL7HRiAQCLJTGO0TAXMEz42LFy9CofAB0Nj+pAkUCi9ERUXlWl4mk8HNzQ1a\nrTbHc7PZjNjYWJjN5hzPu3fvhm7d2gPQQoY2mIkmGIOpyADQC8uxGFXg5FQiR52+fbtBp/sGwGYA\nuwDUBvAgjO0rsF2dswJoDrncHdHR0Xn2r06dOrhxIxqJifFISIhD06ZN8ywrEAgETwsh9ILnhpeX\nF9LTY2CLUAcAl5Ge/je8vLweVS0H4eHhcHX1QrlytWEwuOLtt4ciMzMz631IyI9Q4CYWYjfewZ9I\nA9AJdfArdkGn64+ffpqSw17z5s2xbNlMVKgwASVL/g6Vaks2/34EUAW2vzaXkJ4eZzvU9whkMhkM\nBkOecfcFAoHgqVOEKwvPhGLosuARTJs2kzqdO43GNtTp3Dljxqx8101MTLQv/e+yL6fvJmBgmzZd\naLVaSZK3rl7lr7Bdn7sHic3xO4FR1GqNPHnyZJ62zWYzZ82axebNW1Gp1FGnc6ejYylqNC52X904\ne3ZIofsvEAgE+aEw2ieu1wmeOxcuXEBkZCTKly+PgICAh95fuXIFy5cvR2amBd27d4O/vz8AYO/e\nvWjS5E1YrZHZSteCWh2NixdPwMfNDezaFbKwMNyFHG1RHvtRA8AGLF78E/r165erP5mZmWjatC2O\nHZMhNfVlSNIyDBz4f5g27RtERkbi4sWLqFChQpYfRUlcXBwSEhLg7++fa+Y7gUDw30QEzBEUezIy\nMhgXF0ez2Zzj+cWLF+no6EmVaigVindpMLjx6NGjJMn33gu2Z46Lts/oYwg4U6crxYtHj5KvvGI7\neOfszA5e5QnIqdMZOW/evEf6Eh4eToOhOoFMu90bVKl0NJlMBeqTxWJhVFQUY2JislYYHsX7/9/e\nnQdEVb0NHP8OMwPMgOAOCCKGIiIIKq65YC6E5p7mXtpiWpbtZmbaT8W0zD1zqUxLLTX33dxywdwV\nLU1REUVxIZEBZmDO+8fQlK9LiwgyPJ+/nLuc+zy3iWfuveee88YQ5eJSQhUrVlWVLu2vjh49+q+O\nJ4RwXPdT++Q9elHg9uzZQ0xMRzIyLICZefO+oGNH29Cz//vfx6Sl9cdqHQbAzZtBvPXWCAYM6M3G\njTuATkAEUBM4ClQnokIagf37Q1wc+Pig3biRZSEhmM1m9Hr93z4vv3HjBk5OfsAf78mXQaPRYzKZ\n/vFV9o0bN2jevB3x8SdRKptHH63LypXf4eLicsft169fz/Tpi8jKOklWVinS0mbRsWPv/zzErxBC\n/EE644kCZTabefzxDly7NpmMjEtkZGykV68XOHfuHABXr/6O1VrxL3s8wu7d+3j66Y85fjwLuAws\nBSyAjvByyWzTmdHExUGFCrB9O4TYJoVxdnb+R53iHn30UWyT1swFzqDXv0lISBglS5b8mz3/9MYb\nQzl8OACT6SwZGefYsUMxevS4W7ZRSvHtt/Pp3PkZPvxwNBZLS/6cNKc7p04d/cfHE0KIu5FCL/LF\nwYMHWbVqFefPn79l+YULF8jKcgI65C6phV5fg/j4eAC6dGmN0TgaOAz8irPzu5jNBm7e/Ins7J+A\n60BX3N1TaFKpLHvdrOiOHoWgIFuR/w+j0Hl5ebF582pCQz+jRIlGPPbYWdav/+Ef/UhYsmQJPXo8\nz9Kla8nK6ontroAzGRnd2L370C3bjh07nuef/5BFixqya1cpzOaVwB9z0i/F3//2/gpCCPGv5d0T\nhPxRCEMu8l5++U1lNPopT89oZTSWVitXrrSvS09PV66uHgric5+Hpyij0cf+fNpqtapx4z5VpUsH\nqBIl/HJ7wb+au+1pBRuVVuuijq1apawVK9pGvAsLUyo5Od/zHDVqjDIa5fUbaAAAIABJREFUKymY\npjSaGgoG5I6ul6NcXHqoN98ccsv2Hh5eCn6xj8Kn1YYqvb6U8vSsrUqUKGfviyCEEPdT+6TXvXig\ndu7cScuWPUlPPwB4Artxc2vNjRsp9nHqv/56Hv37v45OV5fs7AO88spzxMYOv2N7u3fvpmnTDmRm\nhmCb4KYMIU4X2F/KDZeUFKhdG9auhX9xm/1esX/77SKMRlcGDHiBgICAO253/fp1YmKeJC4uDtgN\nhAIpaDRhODuXRK+HSpVKsn37Wtzd3e37ubmVwmQ6DNjexXd27s/rrxenbdu2VKtWDQ8Pj/vOQQjh\nGO6n9kmhFw/Ut99+S79+y7h5c6F9mV5fjKFD38JoNNKlSxf8/f357bffOHLkCBUrViQiIuKebTZs\nGMWOHaeBQ1TnHBtoRFnSuB4WRomffoI8KJCrV6/mySf7kJExCCenqxQr9g0HDuykYsWKt23bsWMv\nVq1yw2xeDOwHygOg0w2gXz8nevfuTc2aNdHpbu372r//a3z99UFMphHAr7i7v8ehQ7t55JFH7jt+\nIYRjkUIvHlrx8fHUqdMMk2k7UBn4Bo2mH1ptL5ycwNX1B37+eRtBQUG37XvlyhUOHDhAqVKlqFGj\nhv0ZucHgQWZmb+rQi7U8TglSWQv0MpYk8WrSbUPk/hfVqzfkyJG3gHYAODm9wyuvKD79dOxt2/r4\nBJGcvAz4HDgCjARO4Ob2Jvv377hjbmB7X/+DD0axdOk6SpcuyYQJ/6NGjRr3HbsQwvHcT+2Tznji\ngapWrRoTJozGxaUWRqMvzs4DUepFsrM/w2z+jLS0QQwdOvq2/fbs2UNgYCidO8fSuHEnunbta/+S\nKwVNWMpGmlOCVJZQk3aUx4SRixcv5knc6ekmwMv+2Wr1Ii3NdMdty5cvj0azDRgHNECjeRJ//4/Z\ntGnlXYs8gE6nY9SoD4iP38nWrSulyAshHggp9OKBe/75vly9epHjx3dRq1YdoLF9nVKVuHIl9bZ9\nunTpy40bk/n99x9JTz/GqlWHWLp0KQAfN3uMNVygGDeZhwdd+AUzA9FoTPj4+ORJzE8/3Rmj8RVg\nL7AOo/FjevTodMdtv/xyEp6eI/DwaI+7+xpq1Ajkl1/2ULdu3TyJRQgh7scDLfSJiYk0bdqUatWq\nERoayqRJkwC4du0aLVq0ICgoiJYtW5Ka+ucf+tjYWCpXrkxwcDDr169/kOGJfDRlynSqVo0gLm4r\nWu0QIB74CaPxf3Tu3Oq27S9cOA08nvvJgNnchFOnTsHixby0YTUGFF+7etAbKy5Gf4zGj1i48Os8\nuW0PMHToO7z1Vjv8/fsQFDSUr7+eTNOmTe+4bbVq1Thx4hBz5jzP4sWj2b17kwxfK4R4eNxfh/97\nu3jxojpw4IBSSqm0tDQVFBSkjh07pt566y310UcfKaWUGjNmjHrnnXeUUkrFx8er8PBwZTabVUJC\nggoMDFQ5OTm3tPmAQxYPwOLFi5XRWFnBSQXXlJNTeQUuCtyVwVBS7d69+7Z9wsLqK41mvH0IWje3\nQHXk7beVcrJNUKNef10pq1UlJiaqHTt2qJSUlALITAgh8sf91L4HekXv7e1t70Ht7u5O1apVSUpK\nYvny5fYJRZ5++mn7Ldlly5bRrVs39Ho9AQEBVKpUiT179jzIEEUeyszM5OjRoyQnJ9+yfOXKTZhM\nLwOVgJtYrWnAFiCNjIzZPP54h9vmkv/hh7n4+n6Om5s/zs6V+bZJCKFjx4LVCh98AB9/DBoNfn5+\nNGjQgNKlS9v3VUqxYsUKJk2axE8//fSg0xZCiIdavj2jP3PmDAcOHKBu3bpcunQJLy9bRycvLy8u\nXboE2EZJ++tc5H5+fiQlJeVXiOI+xMfH4+8fTIMGTxIQUJUhQ0bY1/n6lkGvPwJcAZpjK/j1cte2\nx2JxsQ95+4dHHnmEIUNe5bHHGrDk0Tq0Xb3CtmLcOBg+HO4ySp1Siu7dn6Nbt6G8/favREf34KOP\nxudxtrceLzZ2HKVK+VOypB/vvTcCq9X6wI4nhBD/Vr5ManPz5k06derExIkTKVas2C3rNBrNPYcW\nvdO64cOH2/8dFRVFVFRUXoUq/qN27XqQkvI+8CyQwqRJ9Tlx4hd++eUcZcuWpFSp46Sk1CMnpw6w\nGVvRLw2cIDv7mv2HH9iKZ82ajTh48DrDqEhrNgFgmTQJ/cCB94xj7969rFixmfT0o4AReJdhw6oy\nYMDzt3338sIXX3zFyJFzMJnWAjomTOhOyZLFeeONV/P8WEKIomPLli1s2bIlT9p64IXeYrHQqVMn\nevXqRfv27QHbVXxycjLe3t5cvHiRsmXLAuDr60tiYqJ93/Pnz+Pr63tbm38t9KLgKaVISDgK9Mxd\nUoaMjKYsX74Li2Uax4//jIfHXkqUcOPKlUFAFaAGUB2tdidTpky4ZRS42bO/4ODBOMbxIm8yhRyc\neEFbji5BQUT/TSwpKSlotZWwFXkAP3S6YqSmpnL58mXi4+MJCAigevXqeZL7woWrMJneA2wT55hM\nI/j++0lS6IUQ9+X/X8SOGDHi7hv/jQd6614pxbPPPktISAiDBg2yL2/bti1z5swBYM6cOfYfAG3b\ntmXBggWYzWYSEhI4efIkderUeZAhijyg0Wjw9a0MLMtdcgOrdT0Wy0igMVbrG1gsDShf3gedbjnw\nAbAYZ+fLDBzYl+ee63NLe5s2bGMa8CZTMKPnKRbypdX/tuf4d1KrVi2s1oPASiALjWYipUp5sHnz\nVsLC6tGr1+fUrx/D0KH/y5Pcy5QpjpPT6b+ci1OULOmZJ20LIUSeyJPugHexfft2pdFoVHh4uIqI\niFARERFqzZo16urVq6pZs2aqcuXKqkWLFur69ev2fUaNGqUCAwNVlSpV1Nq1a29r8wGHLP6jPXv2\nKE9Pb+XpWU8ZDN5KozEquJzba14pd/doNW3aNFWhQlVVrFiEcnMLVA0bRqvMzMxbG7JY1P6w6kqB\nMqFVMUxQME7pdB7q6tWr/yiW7du3Kx+fSsrJSadCQuqoQ4cOKVdXTwVHc+O5rAwGb/vEOffjxIkT\nysPDS+n1Lyqd7mXl7l5GHTp06L7bFUKIv7qf2idD4Io8k5qaytGjRylTpgxTp85i9uytmEwD0Ov3\n4u29kfj4n9Hr9Rw8eBAXFxfCw8PtE9sAkJUF3bvDkiWkazS003jxo9Lg5JTF0qVzeOKJJ245XnZ2\nNu+99yHff7+C4sU9+fTTETRp0sS+XimFRqMhISGBsLAmpKf/2eHP07Ml8+e/RkxMzH3nnZiYyIIF\nC7BarXTu3FnGqhdC5DkZ614UOKUU58+fx2w2U7FiRTQaDdOnz2Dduu1UqODD+++/c8srcLcxmaBT\nJ9vMc8WLk7FkCStSUsjMzKR58+aUK1futl1eeeUtZs/+GZNpLHAGo/Eldu3adNvzd7PZjLd3Ra5f\nn4Zt7PoDGI0tOX58H/7+/nl6HoQQ4kGQQi8KVHZ2Np069WL9+o04OblSubI/mzevpESJEv+sgbQ0\naNMGtm6FMmVg/Xr4mxnsAEqWLM/165uxva5nm3hm2DA3Pvhg2G3bxsXFERPTkcxMK5DBl1/OwN+/\nPGazmdq1a2M0Gm/bRwghHhb3U/vy5fU64dg+/XQSGzakkJmZCDhz/PjLDBjwJvPnz/77na9dg5gY\n2LMHypWDTZsgOBj489b73bi4uALX7J+12qsYDKXuuG3dunW5fPksycnJuLm5ER3dkePHL+Pk5I6H\nRyq7dm26ZQwHIYRwFDKpjbgvixYtZvDg/5GR0Q1wBZwwm3uzb9/hv9/50iVo2tRW5CtWhO3bOaXX\n07NnX4oXL49Wq8fLq+Jd3yX93//exWjsDExCq30NT8/19hEX70Sn0+Hn58ekSVM5cqQ0N28e5saN\nOC5e7Er//m/+l/SFEOKhJ1f04h/bv38/n3wyjawsC/369SQgIIDevV/Eau0OrAaeAbTodCuoWrXy\nvRs7fx6aNYMTJ6BKFdi4kdNmMzVqNCAtTQu8BbzE5cubadOmC7/+evC25/TPPdeXcuW8WbRoJaVK\nefLaa7tvGXjnbuLjT5GZGQ1oAcjJieHXX1f/+xMihBCFgBR68Y8cOHCARo2iMZkGA+6sWfM0L73U\nE52uCfAx8ARQDVB4e8Nnn22+a1s5J06gadECp3PnIDzc9ky+bFmmvz2Emzc7AkuA13K3jsbJKZJ9\n+/bdsUNeq1ataNXq9tnv7qVu3XBWrZqPydQDcMHZ+SsiI8P/VRtCCFFYyK178besVisDB76NyfQW\n8AbQD5NpCsuXb8JqPQxYgfXABzg7n+f48TsXZYDvPxxJcpVgnM6d46CrkcS5cyF3ZESTKROlvIF0\n4I9X4TLIyTlhHz3xTrGdPHmSs2fP/uOOKq+++jKPP+6Di4sfBoMf1arFM3XquH9+QoQQohCRQi/+\n1jPP9Gf37l/5c1hZAAOuru507NgUo7EGbm7dMBoH8eWXs3F3d79jO/HffEPUBx/gi2IzUTTJeoMn\ner5oX9+zZxeMxmlAV6AB0Adn55q0bt3ojiMkXr9+nVq1GhMR0Yzg4Dq0bt0Zi8Xyt/nodDoWL57H\n6dNHOHZsJ3v3bv3nbwgIIUQhI4Ve3NP58+f5/vvF5OTMAEYCC4FVuLq+xMCBz+Dq6oLFcgGLZRNl\nypSkadMmd2znzPz5lO/dmzJYWY0frZjPDfU+R4/G2Wd7q1evHkuWfE2NGr/h7+9BmzapfPfdGBYs\n+OKOve8HDnyHY8eqYTKdITPzHFu2pPPxxxP+cW7lypUjICDg1kF7hBDCwcgzenFP6enpaLUeQDTw\nFTAROECDBtVxcdEzb94GLJalQH2SkmLp2fNFNm1adksbv//wA6W798AdxSKa0R1fLDwNvEeJEj63\nFNro6Giio/9u6hqb/fuPYDaPw/Z71YWMjKfYvXsD48aNZ+/eo0REBPP666/i4uJi38dkMpGamoq3\nt7cUeCFEkSAD5oh7ys7OpkqVmpw925acnKexTVwzAReXkmi1lzCZtIA3kAbMolSp3ly5cta+/+/f\nfotrj564oJhDb55lNjkAeGAwGFi8eN5/Hoa2U6feLF/uQ3b2GMCKq2s3ypU7zsWL5cjIeBKDYTl1\n6uTw448rcXJyYvz4Sbz77hCcnIx4eZVm06YVBAYG3ucZEkKIB09GxhMP1IULF2jUKIbTpy9im152\nGvAKcBXYCrgAY4CFREaW4Oeff7Tt+P33ZD/1FDqlmEZZXuYCCi1wDSenchw+vI9q1arZj7N69Wr6\n9h3I9euXqFevCYsWfUWZMmXs681mM5mZmfYpbZOTk6lfvznXrjljtZp45JGSnDiRQGbmmdyYLLi5\nVWHHjh9IT0+nRYuumEzbgQpoNOMJCfmOo0d3P+jTJ4QQ9+1+ap/cuxT3ZLFYGDHiIxITfwM6AOuA\n48Bu4ElsBRWgAxpNAnPnTrN9/Oor6NoVnVKMpQ8v8QiK7sBkNJpGuLuXZuTI8aSmpgLwyy+/0Lnz\n01y6NAuzOYldu4Jo27a7PY7hw0fh5uZJqVI+1K4dxZUrV/D29ub48b2sWTOFzZvnMXfudLRaI+Cc\nu5cOJyd3zGYz+/btw2p9AqgAgFIvcfz4XvnRKIRweFLoxT0NGTKCefPisVh+BH4AXgKeBprnfjYB\nCphD48YNCQ4OhqlToU8fsFqZXq4ig6kNbASqA+NRSs+NGz+wZIkTLVq0RynFtm3bgDZAU8ATi2Us\ne/ZswWw28/jjbRgxYhrZ2afIzk7j0KHq9OzZDwAnJyfq169PZGQkISEhlC9fAr3+dWAPOt0QSpe2\nUr16dSpUqIBWuxPIyM1sM2XK+N9ziF0hhHAEUujFPS1duhaTaSRQF9gGfAs8CrgDQUAAEIhe/znf\nfPM5fPQRvPyybedPPqHR+hUULzGSYsXaA19g+2EQB9TGbJ7O0aPxXLx4kZIlS+Lk9Cu2d/IBTuDq\nWowpUz5j06ZDwPNAOcAJi+VNdu/exaOPRmMwuGM0ejJ16nR0Oh3btq2hbdurBAb2p1Wrs+zcuREX\nFxfatGlDTEw4bm7V8fBojbt7bxYs+CK/TqMQQhQYeUYv7ujKlSssXbqUUaMmc+bMm0AvIAnbVfkJ\noH7uv3XodCvZuGElTTZtgpEjURoN8S+/TNVPP0Wr1XLt2jWGDRvG55/vJDs7GziI7TfmTZydy3Hh\nQgIeHh40avQ4R49aMZsj0Onm0759c/bu/YWTJ6sDF4FVufstwGh8HYulIxbLBCABo7EZq1Z9TVRU\n1F1zUkqxe/duUlJSiIyMvOugPkII8bCRzngiTyUmJlKz5qOYTA3IyUklK2s7tiv649hmizsKlAHm\noNePYP63M+i0YwdMmEA20M+lLt/pzdSu7cuMGZ8ycOC77Nu3h6tXG2C1JgFeQDNgJr161eLrr2cA\nts52CxcuJCEhgalTv+TGjXpkZp7G1gHwJHADKItO9xM6HWRmnshtC5ycBtOqVTyNGzcmJiaG0NDQ\nfD1nQgjxIEmhF3lGKUXr1p1Yt64KVmts7tJ2aDRxKPU9ts54U4EmODsfonePVszUmmHWLLKArgxj\nKSOAbNzcGqPVniQ9fRA5OaFAb+Bd4CRa7Sbq1XuEbds23vY++5gxH/HBB8cwm+cAl4HaQHlcXbPR\n639j3boVdOv2PGfPjgVaAVY0mih0OhPQEL3+G5YvX0CzZs3y45QJIcQDJ73uRZ5QStG9+7OsW7cT\nq/Wvk7yYUCoWaIRtdLzReHruYd3KGczISoVZs8BgoKPWmaX8Md2rjqysWmRleZKT8x7QDtiORvMR\nYWEnGDr0ebZsWX/HQWuuXUvFbP7j/faywFKKFTvBl18OIiHhV+rXr8+XX07GaHwaN7deODvXRamz\nWCw7sFgmYDLN5qWXBj+4EyWEEIWIFHpht27dOlas2IPVOhj4BLgApKDVnsz99x9cqV09gqipU9F8\n+y24u8PatVyNbIRWOwZbL/zf0GoXodXqcz8DVESns7Bt2wqGD3+Po0ePMn/+fPbt23dLHE88EYPR\n+DmwCziPwfABnTt3oWvXrpQqVQqApk2bcvjwbiZPfozoaF/gOf581S+I69evPZiTJIQQhY0qZAph\nyIXG559/rozGvgqsCoYoKKZArzp16qbc3csojeZtBR+oUoZS6lqdOkqBUiVKKBUXp5RSKikpSVWv\nXl/pdK7K2dlNTZ48VYWG1lUuLj0UzFRGY0PVq9cLSimlxo79VBmNPqpYsc7KaPRTw4ePviWWefO+\nUV5egcrDw0v16vWCysjIuGvcGzduVEajn4K9Ci4qg6Gd6tv3pQd3ooQQIp/dT+2TZ/TCbt++fTRq\n1IaMjG1AJWAilSp9xa5dG+jb9yV2744joGRxNrgoPA8ftk0vu2EDVK9+Szvp6em4urqi1Wq5efMm\nY8Z8zIkTZ2nUKJKXXupPSkoKFSoEk5W1H5iObaKcVCZPHsXLL7/0n2KfPftL3nnnAzIy0mnfvgOz\nZk3GYDDc5xkRQoiHg3TGE3li4cLv6dnzmdxX4PR4eXmxadNyOnbsxZkzDXE3N2Wd5gUi1RWUnx+a\njRuhSpV/fZyDBw/SuHEP0tLCsD0SmAEk4+LSlbVrF9zzFTkhhCiKpDOeuG9nz56lT5/+ZGfvxPYa\n21QyM03cuHGDCxeyKGF+ly0MI1Jd4bRGx+mvvvpPRR7Ax8eH9PTz2EbLmwwEA1FkZb3GkiUr8iol\nIYQQSKEXueLj49HrawHh2Dq1PY3FYhvsxs9qYhtNCOMoxwimhUsprP7+/+k4V69e5Z13BuPk9BgQ\nCCTa1+l0iXh6uudBNkIIIf4g89ELACpUqEB29hEgBdtgOEewWm/SxNeXH7Mv4kMW+wmgnUsAJSp7\nYjab/1G7WVlZ6PV6nJyciI+Pp1Gjlty86UJ29kBsV/LPAP2A83h6bmDAgD0PKEMhhCia5IpeAFCt\nWjVef70/RmM4Hh4xGAyPsWjEe7jHxOBjzuJsOT9eqeZNcs5Ozp51p3bt5gwfPvqu7V27do2GDaNx\nc/PAYCjGJ59M5JlnBpKaOgyLZRTwJVAL28Q4S9FolgCQkZFx1zaFEEL8e9IZT9ziyJEjnDlzhppK\n4du3L1y9Sk5UFGtffJEOvfpisWwC6gGXMBgi2Lt3EyEhIbe107p1FzZuLI3ZPAk4j9H4GHq9md9/\n/xGoDAwDxuZuHQV8h5PT50RF7WDTpmX5k6wQQhQS0hlP5JmwsDDalCyJb69ecPUq5pYtCTt7haee\n/RSLpQq2KWovAl7o9WGcOXPmlv1TUlI4fvw4O3b8hNn8LranQwGYTL0oU6Y0ev00bAPoDEKrDQA6\nYxtW1xOrtQWnTiXkY7ZCCOH4pNALAC5cuMCkSZNYMmAA1hYt4MYN6NyZNwKqcCqpIenpu4D92Iay\nHQocxmLZT1BQkL2NESNiKV++EnXrtiMtLQP4OXeNFYNhLy+80IOQkJ9xcSmDXu9PkyaBGAyngTTA\nirPzDGrXrpnPmQshhGOTW/eCU6dOERnZiKY3Q5ifvRkXrPzeoQOe339P8+jObNrUDduVN8A6NJqe\naLUmnJy0ZGdnUKtWI4YOHUS3bq9gMu0GvIF30WgmYzS2RaM5S8WKOUREhJOQkESdOqG8997bWCwW\ngoJqcuPGNcAZd3c3TpzYh4+PT4GdCyGEeBjJrXtxX4YOHc3jvzdiYfZWXLAyhQY8k+MKWi2NG9fG\nYJgFmIAsDIYZtG7dBL3eE7P5J6zW0fz8cwo9e/bDam2IrcgD/A+lTEye3IIZM14mNTWVBQvc+Omn\nZ/nss4O88MKrDBz4DhkZT2F7xe4Q2dm1+OKLOQV1GoQQwiFJoRfUOLifb9T36MkmlsEMpBFLl6/H\n3b08ZcuWICamDHq9F87OZWnUyErDhrXJyekMfA6sAN4nLa0TWVnL+fO9+FX4+DxCnz59KFasGKmp\nPlgs44EOZGQsYcmS7/jxx5+wWDoDpYEAMjM7sm/fsYI5CUII4aDkPfoiJjk5mVGjxpGUlEKbNs14\n5kYqb/9yEIAhvEEs5YHhwFTS0zPo3/8VVq6cz6xZk7FarZQqVYpvvvkGnW4lZvMebPPFewJd0Oni\n0WiqYzCEYLX+xvTpM0lJScm93aT5SxQalNJy/Xo2Gs23KFUPyMZgWEKtWo/m7wkRQggHJ8/oi5Br\n164REhLJ1avtyM4OY5h+CCMslwBY2Tyap3buxWTKwfaOe/vcvabSuPFKtm5dY2/HYrHQpEkrdu3a\nAvwOGAFwd2/D8OFRBAcH8/77Yzh+/Fes1izatGnL7t17SE5uQ05OFLaJbDyAMUAobm4+KJVB3brV\nWbNmMS4uLgghhPiTPKMX/8iSJUtIS6tFdvZ4RvEbIyyXsAJq5kye2LCWs2d/QaPRANa/7GXF2dn5\nlnb0ej3btq2hQYMmODu3B9ah1X6IwXCYPn36MH/+Mo4dCyYz8yJm8wWWLfuNMmVKEBq6A632RSAS\n+Aooh4uLnqVLp7J37zo2blwuRV4IIfKYFHoHdPbsWb766iuWLFlCVlaWfbnFYgGrGxN5lSHEko2W\np7XO8OyzAJQuXZrWraOAF4Cvgc9xchrK8OFv33YMnU7H5s2ref31+kRGjqN9+9/4+edtlCxZkri4\n/WRlPQdoATeys/ty8KCOEyeS8PDQoNOZgR24uj5NnTp1aNasGVWrVsXJSb6OQgiR1+TWvYP57rvv\n6NHjeZRqjk6XRHCwll27NmIwGEg8c4YtlavSKzuTLPT0cg7BrXt9vvzyM/v+SimGDh3KggWrKVbM\njYkTR9GkSZN/FUNMTGc2bKhOTs772O4OdANCgccpVepJmjaN4sSJBOrXr8nHH4/E3V0mshFCiHuR\n+egFYBv0pnz5alits4BOgEKni+HTT5/g5X79oFcvWLiQDCctA/2qUPKpjowaNQy9Xp+ncZw7d456\n9R7j2rWSZGXdwDZJzjogmeLFG3H9elKeHk8IIRydPKMv4qxWKzNnzqRjx6ewWsH2DBxAQ3Z2PX45\neBg6dYKFC8HDA8PWLUw/dQiNRkPlyrUIDW3A+vXr8ywef39/fv31AJMnP4+LSzK22emOYjQ+S48e\nXfPsOEIIIf6eXNE7gF69XmDx4iNkZHgDx4EmwBQgCSN12eSWSb30G1CyJKxfD7Vq8dprg5kxYxcm\n03ggEaPxBbZuXU1kZOQ9j/Vvbdu2jUGDhpGa+judOrUmNnY4Op281SmEEP+G3LovwpKTkylfPojs\n7AvANWzPwgOAX/AAVuFKQ9LA2xs2bIDQUADKln2ElJRVQNXcloYxeLCV2NiRBZCFEEKIe5Fb90XY\nwYMHyc7WA26AP/A2cIVS1OFHytKQNM476ehbKZxt167Z9zMYjECK/bNOdxk3N0M+Ry+EEOJBk0Jf\niMXFxbFs2TI0GmfgFeAIoMObLLayk1okcRItDazj+fKnJ4mJeZKdO3cCEBv7HgZDd+ATtNpX8PRc\nRd++fVi+fDkTJ05k27ZtzJkzl9Kl/TEaS/DUU30wmUwFmK0QQoj/Qm7dF1LDh49m3LjpKBVKRsYu\noBoQjz9l2UQSlUjnpIuBxllTSaZP7l6T6Nr1EPPnz+bKlStERNTj8uV0IIcqVSoSHFyFNWsOk53d\nEFiE1QoWy0rAH1fXAXTuXIqvv/68oFIWQogiS57RFzHnz5+ncuVwMjOPAV7AEGAy1V29WJF5Gn8U\nlurVidF4sunQG9g6580GNtCsmY6NG1fSo8dzfP+9OxbLp4DC2TkGpeKxWE5gG9L2FaAEMCL3qKdw\nc3uUmzeT8z9hIYQo4uQZfRGzf/9+rNay2Io8wGjqGMqwyzkFfxQ8+ij6bdt4YchAXF37ARWBnUAk\n27btYeHC7/j558NYLG2xTTbjhNkcjlIV+GPcetvz/qN/OeoJ0tOzWLVqVT5lKYQQIi/IFX0hc/r0\naSIi6pGWZgbmAm2oxXjWa96kpFLQvDksXQpubmRkZFCqlC8ZGVGNMUnHAAARyUlEQVTAktwWduDm\n1onMTAs5OTGAH/ADcB2NxoRS84EWwKfAqNx/BwDfAB0pU2YDSUm/5PkgO0IIIe5OruiLkPHjp5Ce\n/jywBhhAQ5z5kTcoqRQp9euTk1vkATp37k1GRioQ+JcWKpCefoOcnG3YrvI3A4uAeSiVCbwGFAOW\nYpt+tjRQDtgIOHP9umLevHn5lK0QQoj7JYW+kElLM2G1egH1acEs1mGb8HWRrhxBh9N5rFVnzGYz\nb745hFWrlgDuwBxgA3Aa24Q1Ltg67+mBWUAY0DJ32x8AM7AbF5fywHzgCjAJWIrV2ooLFy7kZ8pC\nCCHugxT6QqZHj44YjWNpx4esoA1GLMyiKU9lnyM1fR87d6YxcOBAPvtsOeAMjARygD5ATeAAtufw\nCwAFXPxL6+FoNM2Akbi4dKN8+QwiIsLRaLYDQcAiXF1X8Oijj+ZnykIIIe6DPKMvhHYNHEjtKVPQ\nARPR8RoJKPxy13bH2XktZvO7wA1gBRCN7db7MWy3/H8BPsQ2YE4x4HXgQu62FwgOrk737h0ZNGgQ\nmZmZtGrVhQMHdqHVahk3biyvvPJS/iYshBBFnLxeV5TMnAn9+oFSxDq5McRqBHoD44CVQA+gA7bh\ncJcC84AvgTNABrY54gOAg0AMtiv6hthG1msFPIZe34OqVQ+zf/92tFotABkZGbi4uMic8UIIUQCk\nM15RMWECvPACKMUQJyNDrNOxzVS3DCgOdAWGAtOB34Ha2Ir8YeAZoCzwG7ZOeDGAN5CK7UdBKaAz\n8B4WyzR+++08p0+fth/aYDBIkRdCiEJI/nIXBkrByJHw2msAvOpkINZaEbgJTARM2Ap+CWy36w3A\nJqAD3t7nqF07HGfnj4HG/Pme/AhsPwJaYrttPxRoCwwGLFitWTg7O+dXhkIIIR6QB1ro+/bti5eX\nF2FhYfZl165do0WLFgQFBdGyZUtSU1Pt62JjY6lcuTLBwcF5Oj96oWexwIYNWDUaXtBVYJK1N9AM\neB9YDvQHdvLqq0/i7j4DjWYkMAuDYTqTJo1lz57NrFixBNu79JdyG90EVAE+o0cPX4KCKuDqegH4\nEoOhHU2bNsLf3z//cxVCCJGnHmih79OnD2vXrr1l2ZgxY2jRogUnTpygWbNmjBkzBoBjx46xcOFC\njh07xtq1axkwYABWq/VBhld4ODszvFYD2mpcmZndE2iH7fn7WGAfWu1sund/igkTJhAXt4U+fS7y\n1FN7+eGHL+jc+UkArly5hu0WfQBQAZgGdMDLqzzz5s3k4MHdvP12OO3b/8j77zdj2bL5aDSaAklX\nCCFEHlIPWEJCggoNDbV/rlKlikpOTlZKKXXx4kVVpUoVpZRSo0ePVmPGjLFvFx0drXbt2nVbe/kQ\n8kPnyJEjymDwUTBTQYSC3xV8ocBHabUe6qWXXldms/mu+1+6dEkZDCUVPKPAT4GbAj+l0birHTt2\n5GMmQggh/ov7qX26/P5hcenSJby8bGO0e3l5cemS7VbyhQsXqFevnn07Pz8/kpKS8ju8h9K5c+dw\ndg4jI+NZ4BDwCOBM8eJWPv10It26dbttSNrt27fz2Wdz0GqdaNasAc7Oj5CRMRvblfw6dLo4Fi/+\nhgYNGuR/QkIIIfJNvhf6v9JoNPe8PXy3dcOHD7f/OyoqiqioqDyO7OESGhqKxbIf2A9MBrzRaGLJ\nyanLK69M56OPphAX9yMeHh4AbNiwgXbtepKRMRTIYvHiN3Ifg/wKvAxEodc3loFvhBDiIbVlyxa2\nbNmSJ23le6H38vIiOTkZb29vLl68SNmyZQHw9fUlMTHRvt358+fx9fW9Yxt/LfRFgb+/P3PnzqBn\nz+YopScnx0pOzmDS0oYCioSEPowePY7Y2A+ZO3ceb701koyM8djeqYeMDD21ay/h6NGGODtXwmz+\njVmzplGqVKkCzUsIIcSd/f+L2BEjRtx947+R76/XtW3bljlz5gAwZ84c2rdvb1++YMECzGYzCQkJ\nnDx5kjp16uR3eA+tjh07kJR0mlGjBlOiREms1qa5azRkZTXmxImzvPPO+wwY8AmXLxuwjVv/B3d8\nfPw4deooq1d/yunT8XTv3rUAshBCCJHfHugVfbdu3di6dStXrlyhfPnyfPjhhwwePJguXbowe/Zs\nAgIC+O677wAICQmhS5cuhISEoNPpmDZtmvT6/oukpCRq147i+vXKmM3uwHigDpCJ0fgVjz7akcGD\nB5OdfQ7bBDavYZu0JgujcRj9+8/Gx8cHHx+fAsxCCCFEfpMhcAuBM2fOEBZWk5s3G2N7re4m0Bw4\njF6voWXLGIKDH2H8+AkolYZtdrq5ODkNpUKFkowf/4H9zokQQojCR4bAdXAvvvgmN2+GAzVyl7gD\n8ylevATLly9m8+atjB/vjFKB2IbB3YdGc5Nixczs3LlGirwQQhRhckVfCFSqVItTp54FYrHNMFcB\nJ6fn6dLFkzNnzrN7dy+gJ5AONKNYsSSqVw9l+vRxhIaGFmToQggh8oBc0Tu4Bg0icXHZCwzHNhmN\nD6GhF5g5cyJpaTfBPkWtG9CHmJgW/PTTGinyQgghpNAXBpMnj6VGjXO4uLyNTvc7tWpFMGDA0wB0\n794eo/Ft4AiwE6Mxlh495Fa9EEIIG7l1X0gopVi0aBFPPz2AnJyn0OkS8fY+zb592/nkk0nMnDkX\nZ2dnhg17g+ee61vQ4QohhMhD91P7pNAXIpUq1eDUqf8BTwDg4tKNUaMieeONNwo2MCGEEA+UPKMv\nIlJTrwFV7Z+zsqpy+fLVggtICCHEQ08KfSESHd0CV9ch2KabPYjROJPHH29R0GEJIYR4iEmhL0Rm\nzJhIq1bOuLgEULx4KyZN+pCmTZv+/Y5CCCGKLHlGL4QQQjzk5Bm9EEIIIe5ICr0QQgjhwKTQCyGE\nEA5MCr0QQgjhwKTQCyGEEA5MCr0QQgjhwKTQCyGEEA5MCr0QQgjhwKTQCyGEEA5MCr0QQgjhwKTQ\nCyGEEA5MCr0QQgjhwKTQCyGEEA5MCr0QQgjhwKTQCyGEEA5MCr0QQgjhwKTQCyGEEA5MCr0QQgjh\nwKTQCyGEEA5MCr0QQgjhwKTQCyGEEA5MCr0QQgjhwKTQCyGEEA5MCr0QQgjhwKTQCyGEEA5MCr0Q\nQgjhwKTQCyGEEA5MCr0QQgjhwKTQCyGEEA5MCr0QQgjhwKTQCyGEEA5MCr0QQgjhwKTQCyGEEA5M\nCr0QQgjhwKTQCyGEEA5MCr0QQgjhwKTQCyGEEA5MCr0QQgjhwKTQCyGEEA5MCr0QQgjhwKTQCyGE\nEA5MCr0QQgjhwKTQCyGEEA5MCr0QQgjhwKTQCyGEEA7soSv0a9euJTg4mMqVK/PRRx8VdDgPnS1b\nthR0CAWqKOdflHMHyV/y31LQIRRaD1Whz8nJ4eWXX2bt2rUcO3aM+fPnc/z48YIO66FS1L/sRTn/\nopw7SP6S/5aCDqHQeqgK/Z49e6hUqRIBAQHo9Xq6du3KsmXLCjosIYQQotB6qAp9UlIS5cuXt3/2\n8/MjKSmpACMSQgghCjeNUkoVdBB/WLx4MWvXrmXmzJkAzJs3j7i4OCZPnmzfplKlSpw6daqgQhRC\nCCHyXWBgIL/99tt/2leXx7HcF19fXxITE+2fExMT8fPzu2Wb/5qoEEIIURQ9VLfuIyMjOXnyJGfO\nnMFsNrNw4ULatm1b0GEJIYQQhdZDdUWv0+mYMmUK0dHR5OTk8Oyzz1K1atWCDksIIYQotB6qZ/RC\nCCGEyFsP1a37v/r++++pVq0aWq2W/fv337IuNjaWypUrExwczPr16+3L9+3bR1hYGJUrV+bVV1/N\n75AfqKIwkFDfvn3x8vIiLCzMvuzatWu0aNGCoKAgWrZsSWpqqn3d3b4HhVViYiJNmzalWrVqhIaG\nMmnSJKBonIPMzEzq1q1LREQEISEhvPvuu0DRyP2vcnJyqFGjBm3atAGKVv4BAQFUr16dGjVqUKdO\nHaBo5Z+amsqTTz5J1apVCQkJIS4uLu/yVw+p48ePq19//VVFRUWpffv22ZfHx8er8PBwZTabVUJC\nggoMDFRWq1UppVTt2rVVXFycUkqpmJgYtWbNmgKJPa9lZ2erwMBAlZCQoMxmswoPD1fHjh0r6LDy\n3LZt29T+/ftVaGiofdlbb72lPvroI6WUUmPGjFHvvPOOUurO34OcnJwCiTuvXLx4UR04cEAppVRa\nWpoKCgpSx44dKzLnID09XSmllMViUXXr1lXbt28vMrn/4ZNPPlHdu3dXbdq0UUoVre9/QECAunr1\n6i3LilL+vXv3VrNnz1ZK2f4fSE1NzbP8H9pC/4f/X+hHjx6txowZY/8cHR2tdu3apS5cuKCCg4Pt\ny+fPn6/69euXr7E+KDt37lTR0dH2z7GxsSo2NrYAI3pwEhISbin0VapUUcnJyUopWyGsUqWKUuru\n3wNH0q5dO7Vhw4Yidw7S09NVZGSkOnr0aJHKPTExUTVr1kz9+OOP6oknnlBKFa3vf0BAgLpy5cot\ny4pK/qmpqapixYq3Lc+r/B/aW/d3c+HChVteuftjUJ3/v9zX19dhBtspygMJXbp0CS8vLwC8vLy4\ndOkScPfvgaM4c+YMBw4coG7dukXmHFitViIiIvDy8rI/wigquQO89tprjBs3DienP/8sF6X8NRoN\nzZs3JzIy0j6WSlHJPyEhgTJlytCnTx9q1qzJ888/T3p6ep7lX6C97lu0aEFycvJty0ePHm1/RiVs\n/wMI23m417lwlPN08+ZNOnXqxMSJEylWrNgt6xz5HDg5OXHw4EF+//13oqOj2bx58y3rHTn3lStX\nUrZsWWrUqHHXMd0dOX+AHTt24OPjQ0pKCi1atCA4OPiW9Y6cf3Z2Nvv372fKlCnUrl2bQYMGMWbM\nmFu2uZ/8C7TQb9iw4V/v8/8H1Tl//jx+fn74+vpy/vz5W5b7+vrmSZwF7Z8MJOSovLy8SE5Oxtvb\nm4sXL1K2bFngzt8DR/jvbbFY6NSpE7169aJ9+/ZA0TsHnp6etG7dmn379hWZ3Hfu3Mny5ctZvXo1\nmZmZ3Lhxg169ehWZ/AF8fHwAKFOmDB06dGDPnj1FJn8/Pz/8/PyoXbs2AE8++SSxsbF4e3vnSf6F\n4ta9+ssbgG3btmXBggWYzWYSEhI4efIkderUwdvbGw8PD+Li4lBKMXfuXPsfysKuKA8k1LZtW+bM\nmQPAnDlz7P9N7/Y9KMyUUjz77LOEhIQwaNAg+/KicA6uXLli71GckZHBhg0bqFGjRpHIHWx3MRMT\nE0lISGDBggU89thjzJ07t8jkbzKZSEtLAyA9PZ3169cTFhZWZPL39vamfPnynDhxAoCNGzdSrVo1\n2rRpkzf552WHgry0ZMkS5efnp1xdXZWXl5d6/PHH7etGjRqlAgMDVZUqVdTatWvty/fu3atCQ0NV\nYGCgGjhwYEGE/cCsXr1aBQUFqcDAQDV69OiCDueB6Nq1q/Lx8VF6vV75+fmpL774Ql29elU1a9ZM\nVa5cWbVo0UJdv37dvv3dvgeF1fbt25VGo1Hh4eEqIiJCRUREqDVr1hSJc3D48GFVo0YNFR4ersLC\nwtTYsWOVUqpI5P7/bdmyxd7rvqjkf/r0aRUeHq7Cw8NVtWrV7H/jikr+Sil18OBBFRkZqapXr646\ndOigUlNT8yx/GTBHCCGEcGCF4ta9EEIIIf4bKfRCCCGEA5NCL4QQQjgwKfRCCCGEA5NCL4QQQjgw\nKfRCCCGEA5NCL4QQQjgwKfRCCCGEA5NCL4S4p59//pnw8HCysrJIT08nNDSUY8eOFXRYQoh/SEbG\nE0L8rffff5/MzEwyMjIoX74877zzTkGHJIT4h6TQCyH+lsViITIyEoPBwK5duwr1lKBCFDVy614I\n8beuXLlCeno6N2/eJCMjo6DDEUL8C3JFL4T4W23btqV79+6cPn2aixcvMnny5IIOSQjxD+kKOgAh\nxMPt66+/xsXFha5du2K1WmnQoAFbtmwhKiqqoEMTQvwDckUvhBBCODB5Ri+EEEI4MCn0QgghhAOT\nQi+EEEI4MCn0QgghhAOTQi+EEEI4MCn0QgghhAOTQi+EEEI4sP8Diwf1C+duoqkAAAAASUVORK5C\nYII=\n", + "text": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "#### Comparing the results from the different implementations" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "As mentioned above, let us now confirm that the different implementations computed the same parameters (i.e., slope and y-axis intercept) as solution of the linear equation." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "import prettytable\n", + "\n", + "params = [appr(x,y) for appr in [py_matrix_lstsqr, py_classic_lstsqr, numpy_lstsqr, scipy_lstsqr]]\n", + "\n", + "print(params)\n", + "\n", + "fit_table = prettytable.PrettyTable([\"\", \"slope\", \"y-intercept\"])\n", + "fit_table.add_row([\"matrix approach\", params[0][0], params[0][1]])\n", + "fit_table.add_row([\"classic approach\", params[1][0], params[1][1]])\n", + "fit_table.add_row([\"numpy function\", params[2][0], params[2][1]])\n", + "fit_table.add_row([\"scipy function\", params[3][0], params[3][1]])\n", + "\n", + "print(fit_table)" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "[array([ 0.95181895, 107.01399744]), (0.9518189531912674, 107.01399744459181), array([ 0.95181895, 107.01399744]), (0.95181895319126764, 107.01399744459175)]\n", + "+------------------+--------------------+--------------------+\n", + "| | slope | y-intercept |\n", + "+------------------+--------------------+--------------------+\n", + "| matrix approach | 0.951818953191 | 107.013997445 |\n", + "| classic approach | 0.9518189531912674 | 107.01399744459181 |\n", + "| numpy function | 0.951818953191 | 107.013997445 |\n", + "| scipy function | 0.951818953191 | 107.013997445 |\n", + "+------------------+--------------------+--------------------+\n" + ] + } + ], + "prompt_number": 12 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "# Performance growth rates: (C)Python vs. Cython vs. Numba" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Now, finally let us take a look at the effect of different sample sizes on the relative performances for each approach." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "import timeit\n", + "import random\n", + "random.seed(12345)\n", + "\n", + "funcs = ['py_classic_lstsqr', 'cy_classic_lstsqr', 'numba_classic_lstsqr']\n", + "\n", + "orders_n = [10**n for n in range(1, 7)]\n", + "perf1 = {f:[] for f in funcs}\n", + "\n", + "for n in orders_n:\n", + " x_list = ([x_i*np.random.randint(8,12)/10 for x_i in range(n)])\n", + " y_list = ([y_i*np.random.randint(10,14)/10 for y_i in range(n)])\n", + " for f in funcs:\n", + " perf1[f].append(timeit.Timer('%s(x_list,y_list)' %f, \n", + " 'from __main__ import %s, x_list, y_list' %f).timeit(1000))" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 14 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "from matplotlib import pyplot as plt\n", + "\n", + "labels = [ ('py_classic_lstsqr', '\"classic\" least squares in reg. (C)Python'),\n", + " ('cy_classic_lstsqr', '\"classic\" least squares in Cython'),\n", + " ('numba_classic_lstsqr','\"classic\" least squares in Numba')\n", + " ]\n", + "\n", + "plt.rcParams.update({'font.size': 12})\n", + "\n", + "fig = plt.figure(figsize=(10,8))\n", + "for lb in labels:\n", + " plt.plot(orders_n, perf1[lb[0]], alpha=0.5, label=lb[1], marker='o', lw=3)\n", + "plt.xlabel('sample size n')\n", + "plt.ylabel('time per computation in milliseconds [ms]')\n", + "#plt.xlim([1,max(orders_n) + max(orders_n) * 10])\n", + "plt.legend(loc=4)\n", + "plt.grid()\n", + "plt.xscale('log')\n", + "plt.yscale('log')\n", + "max_perf = max( py/cy for py,cy in zip(perf1['py_classic_lstsqr'],\n", + " perf1['cy_classic_lstsqr']) )\n", + "min_perf = min( py/cy for py,cy in zip(perf1['py_classic_lstsqr'],\n", + " perf1['cy_classic_lstsqr']) )\n", + "ftext = 'Using Cython is {:.2f}x to '\\\n", + " '{:.2f}x faster than regular (C)Python'\\\n", + " .format(min_perf, max_perf)\n", + "plt.figtext(.14,.75, ftext, fontsize=11, ha='left')\n", + "plt.title('Performance of least square fit implementations')\n", + "plt.show()" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "display_data", + "png": "iVBORw0KGgoAAAANSUhEUgAAAnYAAAIECAYAAACUvmMzAAAABHNCSVQICAgIfAhkiAAAAAlwSFlz\nAAALEgAACxIB0t1+/AAAIABJREFUeJzs3XdcFMf/P/DXHnA06UUFRKoISBWwI1VjL2lqNGqiH6OJ\n+eg3McnHfBSMMcbkl6omGjVqJPqJmogoKjbOgsbeC1JVUEBRUIrAwfz+2NzCUg89ysH7+Xj4eHhz\nu7MzN7vc3HtnZjnGGAMhhBBCCFF7kpYuACGEEEIIUQ3q2BFCCCGEtBHUsSOEEEIIaSOoY0cIIYQQ\n0kZQx44QQgghpI2gjh0hhBBCSBtBHTvSZsnlcrz11lswNzeHRCLB0aNHW7pIamnbtm1wdHSEpqYm\n3nrrrVq3mTJlCsLDw5u5ZKR62xw5cgQSiQT37t1rdF729vb44osvmqCUNdnZ2WHJkiXNcqzWSiKR\nYPPmzS1dDNIGUceOtKgpU6ZAIpFAIpFAS0sLdnZ2mDlzJh49evTCef/555/YsmULdu/ejaysLPTp\n00cFJW5fysvL8dZbb2HcuHG4e/cufvjhh1q34zgOHMc1a9mioqIgkbTfP2G1tU3fvn2RlZWFzp07\nAwCOHz8OiUSCO3fuNJjf2bNnMXfu3KYuNoCWOV9eREZGxnP/OAwLC8PUqVNrpGdlZeHll19WRfEI\nEdFs6QIQEhgYiK1bt0Iul+Ps2bOYPn067t69i927dz9XfqWlpZBKpUhKSoK1tTV69+79QuVT5Nce\n3bt3D4WFhRgyZIjQWagNYwy01nnjVVRUAMBzdVDrahtLS8sa2yrTNmZmZo0uQ3ujynO8tnYiRBXa\n789d0mpoaWnB0tISVlZWGDlyJP79739j3759KCkpAQD873//g7e3N3R1dWFvb48PPvgARUVFwv5B\nQUGYNm0aFixYACsrK3Tt2hXBwcFYuHAhUlNTIZFI4ODgAAAoKyvDJ598AhsbG2hra8Pd3R1btmwR\nlUcikWD58uWYMGECjI2N8eabb2LDhg3Q0tKCTCaDh4cH9PT0EBISgqysLMTHx8Pb2xsdOnRAeHi4\n6DZYWloaxo4dC2tra+jr68PT0xNRUVGi4wUFBWH69OlYvHgxOnfuDDMzM0yePBmFhYWi7f744w/0\n7NkTurq6MDc3x9ChQ5GXlye8v3z5cnTv3h26urro1q0bvvjiC5SXl9f72f/9998IDAyEnp4eTE1N\n8cYbb+DBgwcAgA0bNqBr164A+M53YyMWDbXbgQMHEBQUBDMzMxgbGyMoKAhnzpwR5bF27Vq4urpC\nV1cXZmZmGDhwIDIzMyGTyfDmm28CgBDxres2MQB88cUXcHR0hI6ODiwtLfHSSy/h2bNnos/OxsYG\n+vr6eOmll/Dbb7+Jbmkq2r+q2qI406dPh5OTE/T09ODo6IhPP/0UpaWlwvuRkZFwdnbG1q1b0b17\nd2hrayMpKQkFBQX497//LZTB19cXO3bsqLM+dbWNTCYTyp2eno7AwEAA/G1WiUSCkJCQOvOsfnvU\nzs4OCxcuxMyZM2FsbIxOnTrh559/xrNnz/Duu+/C1NQUNjY2WLlypSgfiUSCH3/8ES+//DI6dOgA\nGxsb/Pjjj3UeF+Cvy8jISDg4OEBXVxc9evTAL7/8UiPfFStW4PXXX0eHDh1gZ2eHHTt24PHjxxg/\nfjwMDQ3h6OiIv/76S7RfdnY2pkyZAktLSxgaGqJ///44duyY8L7iMzt48CACAwOhr68Pd3d37Nu3\nT9jG1tYWABAcHCz6e9LQ9T1lyhQcPnwYGzduFM5TxflS/Vbs/fv3MW7cOJiYmEBPTw/BwcE4d+5c\no8oJNHyuk3aAEdKCJk+ezMLDw0Vp33zzDeM4jhUUFLD169czExMTFhUVxdLS0tjRo0eZp6cnmzRp\nkrD9wIEDmYGBAZs5cya7ceMGu3r1Knv06BH78MMPmb29PcvOzmYPHz5kjDH24YcfMjMzM7Z9+3aW\nlJTEvvjiCyaRSNihQ4eE/DiOY2ZmZmzlypUsNTWVJSUlsfXr1zOJRMKCg4PZ6dOn2fnz55mzszPr\n378/CwwMZKdOnWIXL15k3bt3Z6+//rqQ15UrV9jKlSvZ5cuXWWpqKlu+fDnT1NRk8fHxovIbGxuz\n//u//2OJiYls//79zNTUlC1YsEDY5tdff2VaWlrs888/F+q4YsUKoV4RERGsa9euLDo6mqWnp7M9\ne/YwW1tbUR7V3b9/nxkYGLA33niDXb16lR0/fpx5enqywMBAxhhjxcXF7MyZM4zjOLZr1y6WnZ3N\nSktL62zHsLAw4bUy7bZjxw62bds2duvWLXb9+nU2bdo0ZmpqynJzcxljjJ09e5ZpamqyTZs2sTt3\n7rArV66wdevWsYyMDFZaWspWrlzJOI5j2dnZLDs7mz158qTWsv3555/M0NCQ7d69m929e5ddvHiR\n/fDDD6y4uJgxxlh0dDTT1NRk3333HUtKSmLr1q1jlpaWTCKRsMzMTKE+mpqaonzv3r3LOI5jR44c\nYYwxVlFRwT799FN2+vRpdvv2bRYTE8M6d+7MIiIihH0iIiKYnp4eCwoKYqdPn2ZJSUns6dOnLCgo\niAUHB7OEhASWlpbGfvnlFyaVSkXnZVV1tU18fDzjOI5lZmay8vJyFhMTwziOY2fPnmXZ2dns8ePH\ndZ4PdnZ2bMmSJcLrrl27MmNjY/bdd9+xlJQU9vnnnzOJRMIGDx4spC1dupRJJBJ2/fp1YT+O45ip\nqSlbsWIFS0pKYj/88APT1NRkO3furPNYkydPZl5eXuzAgQMsPT2d/fHHH8zY2JitW7dOlG+nTp3Y\nb7/9xlJSUtisWbOYvr4+GzRoENu4cSNLSUlhs2fPZvr6+sI5VFRUxFxdXdkrr7zCzp07x1JSUtiS\nJUuYtrY2u3HjBmOMCZ+Zl5cXi4uLY8nJyWzq1KnM0NBQ+LwuXLjAOI5jO3bsEP09aej6zs/PZ4GB\ngWzcuHHCeaq4hjiOY7///rtw7gQEBDAfHx+WkJDArly5wl5//XVmYmIiHEuZcjZ0rpP2gTp2pEVV\n7xBcu3aNOTg4sD59+jDG+C+X1atXi/Y5cuQI4ziO5eXlMcb4jpGLi0uNvCMiIpiTk5PwurCwkGlr\na7Off/5ZtN2YMWNYSEiI8JrjODZt2jTRNuvXr2ccx7FLly4JaV9//TXjOI6dP39eSPvuu++Yubl5\nvXUeNWoUmz59uvB64MCBzNvbW7TNzJkzhc+AMca6dOnCZs+eXWt+hYWFTE9Pj8XFxYnSN27cyIyN\njessx3//+1/WpUsXVlZWJqRdunSJcRzHjh49yhhjLC0tjXEcxxISEuqtU/V2VKbdqisvL2cmJibC\nl91ff/3FjIyM6uywbdq0iXEcV2+5GGPs22+/Zd26dRPVs6p+/fqxiRMnitI+/PBDoYPEmHIdu7qO\n7ezsLLyOiIhgEomE3b17V0iLj49nOjo6LD8/X7Tv1KlT2ejRo+vMu7a2qdqxY4yxY8eOMY7j2O3b\nt+vMR6G2jt2YMWOE1xUVFczQ0JCNHDlSlGZiYsJWrFghpHEcx958801R3hMmTGADBgyo9VipqalM\nIpGwxMRE0T6LFi0SXRccx7G5c+cKrx88eMA4jmPvv/++kPb48WPGcRyLjY1ljPHtZmNjw+RyuSjv\n4OBgNmfOHMZY5We2Y8cO4f3s7GzGcRzbv38/Y0y5tlaofn2HhYWxqVOn1tiuasfu4MGDjOM4obPJ\nGGMlJSWsc+fO7LPPPlO6nA2d66R9oFuxpMXJZDIYGBhAT08PHh4ecHJywu+//44HDx7gzp07mDt3\nLgwMDIR/Q4cOBcdxSE5OFvLo2bNng8dJTk5GaWmpcHtKITAwENeuXROlBQQE1Nif4zh4eHgIrzt2\n7AgA8PT0FKXl5uYKY3GKiorwySefoEePHjAzM4OBgQH27NkjGszOcRy8vLxEx+rcuTOys7MBADk5\nOcjIyMCgQYNqrde1a9dQXFyMsWPHij6nd955B0+ePEFubm6d+/Xu3RuampVDbT09PWFkZITr16/X\nuo8ylG23tLQ0TJo0Cc7OzjAyMoKRkRHy8/OFz2bQoEFwcHCAvb09xo8fjzVr1tRZl/q8/vrrKCsr\nQ9euXTF16lRERUWhoKBAeP/GjRvo27evaJ9+/fo9V93XrFmDXr16oVOnTjAwMMD8+fNrTFzo2LEj\nbGxshNdnzpxBaWkprK2tRZ/X77//LjrHm1v185LjOFhYWIjOd47jYGlpKdy+V6g+Ualv3741rjGF\ns2fPgjGGnj17iuq/dOnSGvWvWh5zc3NoaGiIymNsbAypVIqcnBwA/GeblZUFY2NjUd7Hjx+vkbe3\nt7fwf0tLS2hoaAjXYF2Uub6Vce3aNZiZmaF79+5CmlQqRa9evWp8bvWVs6FznbQPNHmCtLjevXtj\n48aN0NTUhJWVldDRUPyx+vHHHxEcHFxjP2trawD8l4u+vr5Ky1RbfhKJRDSTT/F/DQ2NGmmMMXAc\nh3nz5iEmJgbfffcdXFxcoKenhw8++AD5+fmivKtPzuA4ThhY3xDFdtu3b0e3bt1qvG9iYlLrfhzH\nNcmEB0V5Gmq34cOHw9LSEj/99BO6dOkCLS0t9O/fXxiTpq+vj7NnzyIhIQEHDx7EqlWr8NFHH+HQ\noUPw9fVVujxWVla4efMm4uPjcfjwYSxevBgff/wxTp06Jepg1ae2yQ1lZWWi19u2bcN7772HZcuW\nYeDAgTA0NMTWrVvx6aefirarfm5VVFTAyMgIZ8+erXGMlp60U31cIcdxtaYpe67WRrHvyZMnoaen\nVyPv+spTVxkVeVZUVMDV1RXR0dE19qt+rNo+64bqpez1/bwUf0eULacqznWi/ihiR1qcjo4OHBwc\nYGtrK4oedezYEV26dMHNmzfh4OBQ45+2tnajjuPk5ARtbW0cOXJElH7kyBFRJE6Vjh07hokTJ+KV\nV16Bh4cH7O3tkZiY2KilHiwtLWFjY4O4uLha33d3d4eOjg5SUlJq/ZzqmnHp7u6Ov//+W9RBuXTp\nEvLz89GjR4/GVbQKZdotNzcXN27cwCeffILw8HBhIoEi0qIgkUgwYMAALFq0COfOnUPnzp2FyS6K\nLzhlOqdSqRSDBw/GsmXLcOXKFRQVFWHnzp0AADc3NyQkJIi2r/7a0tIS5eXlovKdP39etM3Ro0fh\n4+ODOXPmwMfHB46OjkhLS2uwbP7+/sjLy0NxcXGNz+pFv4wVn1FDk2hU7eTJk6LXJ06cgLu7e63b\nKqLtt2/frlF/e3v7FyqHv78/UlNTYWBgUCPvTp06KZ1PXZ+jMte3VCqFXC6vN393d3fhmlAoKSnB\nqVOnGn0t1neuk/ZB7SJ22dnZGDt2LKRSKaRSKTZv3kzT9NuwJUuW4O2334aJiQlGjhwJLS0t3Lhx\nA/v27cOqVasAKL/Uhp6eHt5//30sWLBAuKW0fft2xMTE4ODBg01SfhcXF0RHR2Ps2LHQ19fHt99+\ni/v374u+VJQpf0REBGbOnImOHTvi5ZdfRkVFBeLj4zF+/HiYmZlh/vz5mD9/PjiOQ2hoKORyOa5c\nuYKLFy/iyy+/rDXP9957Dz/88AOmTJmC+fPn4/Hjx5g1axYCAwOf+1akQkPtZmJiAgsLC/zyyy9w\ncHDAw4cP8dFHH0FXV1fIY+fOnUhLS8OAAQNgYWGBc+fO4e7du3BzcwMA4Ut/586d6NevH/T09GqN\ntK5btw6MMfj7+8PY2BiHDh3C06dPhXw++OADvPrqqwgICMCQIUNw/PhxREVFib6ce/XqBQMDA3zy\nySf4z3/+g5SUFHz22Wei43Tv3h2//vorYmJi4O7ujt27d9c7s1UhJCQEYWFhGDt2LL766it4eHjg\n8ePHOHHiBHR1dTFt2rTGN8A/unbtColEgtjYWLz22mvQ1taGkZFRrdtWPwdrOyeVTYuNjcXKlSsx\naNAg7Nu3D1u3bsX27dtr3cfJyQlvvfUWpk+fjq+++gq9e/dGYWEhzp07J5wXz+uNN97Ad999h2HD\nhmHJkiVwdnZGdnY2Dh8+DDc3N4waNUqpfMzNzdGhQwfExcXB1dUV2traMDExUer6tre3R3x8PFJT\nU2FoaAhjY2PRD1gACA0NRUBAACZMmICVK1fC0NAQixcvRmlpKWbOnKl0fRs610n7oHYROwsLCyQk\nJCA+Ph4TJkzAmjVrWrpI5AU0tFDpxIkTsXXrVuzevRu9evVCQEAAFi1aJIpk1JVHbelLlizB9OnT\nMWfOHHh4eGDz5s34/fffa71lWFt+jU377rvvhOVXwsLC0KVLF7zyyis1bulWz6d62ttvv40NGzZg\n+/bt8PHxwcCBAxEXFyd8Qfz3v//Ft99+izVr1sDb2xsDBgzADz/8UG/Ew9LSEvv370dGRgb8/f0x\nYsQIobPbUB1rq3PV7RpqN4lEgm3btiElJQWenp546623MHfuXNF6bKampti1axeGDBkCFxcXfPLJ\nJ1iwYIGw2Ku/vz/+/e9/Y8aMGejYsSNmz55da9lMTU2xfv16BAcHw83NDd9//z3WrFkjtPno0aPx\nzTff4KuvvoKXlxe2bNmCZcuWiTofJiYm2LJlC/7++294eXlhyZIl+Prrr0V1njFjBiZNmoSpU6fC\n19cXZ86cQWRkZINtDQAxMTEYO3Ys5s6dC1dXVwwfPhx79+6Fk5NTg597fWkdO3bE0qVL8eWXX8LK\nygpjxoxROq/nOd8VFi5ciIMHD8Lb2xtffvklvv76a1Enqvo+v/zyC+bOnYslS5bA3d0dYWFh2LRp\nExwdHessrzIUEXo/Pz9MnToVLi4uePnll3H27FnY2dnVW4eqJBIJVq5cia1bt6JLly5ClFGZ6/uD\nDz6Aubk5vLy8YGlpiRMnTtR6jOjoaHTv3h3Dhg1DQEAAcnJycODAAZiamipdzobOddI+cKwpBtk0\nk+XLl0MqlWLGjBktXRRCSBsik8kQEhKCjIwMWFlZtXRx1IpEIkFUVBQmTJjQ0kUhpF1Su1uxAD8O\n6F//+hfy8vJqLGhKCCGEENJetdit2BUrVsDPzw86Ojo1nqP36NEjjBkzRlhdvPqTAby8vHDq1Cl8\n/vnnWLx4cXMWmxDSTqjTs0wJIUShxSJ21tbWWLBgAeLi4lBcXCx6791334WOjg5ycnJw4cIFDBs2\nDF5eXnBzc0NZWZkwvd3Q0FB47BQhhKhKUFBQs88kbSteZOkTQsiLa/ExdgsWLEBGRgbWr18PACgs\nLISpqSmuXbsmDByePHkyrKyssHTpUpw+fRrz5s2DhoYGtLS0sG7dulqXBLC2thY9s5MQQgghpLVy\ndHRUyaLkLT4rtnq/8tatW9DU1BTNBvPy8hJW3w4ICMCRI0dw+PBhxMXF1bnO071794RlJFryX0RE\nRIvn1Zj9lNm2vm2e573a0lX5ubWGtmur7dda204d2+9F266+99Xt2lN1OdrLtdda2k/drj1ltm2O\nay8lJUUl/aoW79hVH8dSUFAAQ0NDUZqBgQGePn3anMVSmaCgoBbPqzH7KbNtfds8z3u1paenpzdY\njqamyrZ7kfxac/u11rYD1K/9XrTt6ntf3a49gP52NvReXdu3hvZTt2tPmW2b49pTlRa/Ffvf//4X\nmZmZwq3YCxcuoH///igsLBS2+X//7//h6NGjiImJUTrfpnpcEmkeU6ZMwYYNG1q6GOQ5UNupN2o/\n9Ubtp75U1W9pdRG7bt26QS6Xi+4zX7p06bkecRQZGQmZTPaiRSQtYMqUKS1dBPKcqO3UG7WfeqP2\nUz8ymQyRkZEqy6/FInbl5eUoKyvDokWLkJmZiTVr1kBTUxMaGhoYP348OI7D2rVrcf78eQwfPhwn\nT56Eq6ur0vlTxI4QQggh6kLtI3aLFy+Gnp4eli1bhqioKOjq6mLJkiUAgJ9++gnFxcWwtLTExIkT\nsWrVqkZ16oj6o0ir+qK2U2/UfuqN2o+02Dp2kZGRdYYeTUxMlHp4NiGEEEIIqdTikyeaCsdxiIiI\nQFBQUJPOPiGEEEIIeV4ymQwymQyLFi1Sya3YNt2xa6NVI4QQQkgbo/Zj7AipD40TUV/UduqN2k+9\nUfuRFhtjRwghhBDS3iUm3sb+/ap56gTQxm/F0hg7QgghhLRWiYm38Z//bMfVq0+RlERj7OpFY+wI\nIYQQ0lpVVAAffHAYFy+GgDHgyBEaY0faMBonor6o7dQbtZ96o/ZTD7m5wK+/AjduSKDqGBSNsSOE\nEEIIaQaMAadPAwcPAmVlgERSAQAwMFDdMehWLCGEEEJIE8vLA3buBNLSKtMePbqNR4+S4egYis8+\no1uxDYqMjKSwNCGEEEJaDGPA+fPATz+JO3UdOwLz53fFoEEPcPbsZJUdjyJ2pFWSyWQ0m1lNUdup\nN2o/9Ubt17o8fQrExABJSZVpHAf07w8MHAhoalZNV02/hcbYEUIIIYSoEGPA1avAnj1AcXFlurk5\nMHo0YGPTdMemiB0hhBBCiIoUFgKxscD16+L03r2B0FBAS6v2/ShiRwghhBDSity8CezaxXfuFIyN\n+SidnV3zlKFNT54g6osmvagvajv1Ru2n3qj9WkZxMbBjB/C//4k7dX5+wMyZ9Xfqbicm4vDKlSor\nS5uO2EVGRtIjxQghhBDSZJKT+QkST55UphkaAiNHAk5O9e97OzER2xctwtPMTJWVh8bYEUIIIYQ0\nUkkJsH8/cO6cON3LC3jpJUBXt+E8Dq9ciZCUFODmTXB799IYO0IIIYSQ5paeDkRH84sOK+jrA8OH\nA66uSmYil0Ny9Spw44ZKy0Zj7EirRONE1Be1nXqj9lNv1H5Nq6wM2LcP2LBB3KlzdQVmzWpEpy4j\nA1i9GhW3b6u8jBSxI4QQQghpQEYGH6V7+LAyTUcHGDYM6NGDX3i4QXI5EB8PnDgBMAZHBwccungR\noebmKisnjbEjhBBCCKmDXA4cOQIcP84vPKzg7MxPkDAwUDKjjAz+YbEPHlSmSaW47eyMlJwchL73\nHo2xI4QQQghpKllZ/DIm2dmVaVIpPznCx+f5onQCe3tg1Cg8e5iNG+cOqqzMbXqMXWRkJI03UFPU\nbuqL2k69UfupN2o/1aioAI4eBdasEXfq7O35sXS+vkp26v4ZS4eEhMpOnVTK3799800kPszGolWL\nsDFmo8rK3qYjdpGRkS1dBEIIIYSokQcP+LF0VZeW09ICwsKAgADVROlgbAwA2HN6Dyp6V0CvQA/Y\nrJry0xg7QgghhLR7FRXAqVPAoUN8v0zBxgYYMwYwM1MyozrG0iE8nH8UxT89w8SHiZi3Zh6edOZX\nNj4y9QiNsSOEEEIIeVGPH/NRuqqrj2hoAMHBQN++gESZgWtKRumKyoqwL3kfLmdfhrxcXkdmz69N\nj7Ej6ovGiagvajv1Ru2n3qj9Gocx4OxZ4OefxZ26zp2Bf/0L6N9fyU5dA2PpFJ26mw9v4qczP+Fy\n9mUAgIODAyRpEvSw7KGyOlHEjhBCCCHtTn4+/4zXlJTKNIkEGDAACAzkI3YNakSUbm/SXlzJuSLa\nPcQnBNN9pyPhcoIKasSjMXaEEEIIaTcYAy5fBvbuBZ49q0y3sODH0llZKZmRkmPprj+4jthbsSgs\nKxQ2M5AaYHi34XAxdxHSVNVvoYgdIYQQQtqFggJg927g5s3KNI7jx9EFBwOayvSKlIzSFZYWYk/S\nHlx7cE20u3cnbwx2HAxdLV0V1Kgm6tiRVkkmkyEoKKili0GeA7WdeqP2U2/UfnW7dg2IjQWKiirT\nTE2B0aMBW1slM1EiSscY46N0SbEoKqs8mKG2IUZ0GwFnM2fVVKgObbpjFxkZiaCgIDrJCSGEkHaq\nqAjYswe4elWcHhDAr00nlSqRiZJRuoLSAuxJ2oPrD66Ldvfp5IPBToOho6lTI2uZTKbSSS80xo4Q\nQgghbdKtW/wEiYKCyjQjI74v5uCgZCZKRumu5lzF3uS9NaJ0I11GwsnUqcHD0Bg7QgghhJBalJQA\n+/YBFy6I0318gMGDAZ2agbOalIzSPS15itikWNx8eFO0e8/OPTHIcRC0NbVfsDaNQx070irROBH1\nRW2n3qj91Bu1H5CaygfY8vMr0zp0AEaOBLp1UzITJaN0V3KuYG/SXhTLi4XNjLSNMNJlJBxNHVVT\noUaijh0hhBBC1F5pKXDwIHD6tDi9Rw9g6FBAT0+JTBoRpdt9azcScxNFu/tZ+SHcIbzZo3RV0Rg7\nQgghhKi1u3eBHTuAR48q0/T0+Ac/uLsrmYmSUbpL2ZewL3kfnskrF8Ez1jHGKJdRsDexf+460Bg7\nQgghhLRrdQXYXFyAESP4W7DPnUm1KN2TkifYlbgLSY+SRLsHWAcgzCEMUg1lptc2PerYkVaJxomo\nL2o79Ubtp97aU/vdu8dH6aoG2LS1gSFDAC8v4cEP9VMySncx6yL2Je9DSXmJsJmJjglGdR8FO2M7\nldVJFahjRwghhBC1UV4OHDsGHD0KVFRUpjs48AE2IyMlMlEySpf/LB+7bu1C8qNk0e69rHsh1CG0\n1UTpqqIxdoQQQghRCzk5fJTu/v3KNC0tYNAg0eNZ66dklO78/fPYn7JfFKUz1TXFKJdR6GrcVXWV\n+geNsSOEEEJIu1BRAZw8CRw+zEfsFGxt+UeCmZoqkYmSUbq8Z3mISYxB6uNUYRMOHHrb9EaIfQi0\nNLRUVKumQR070iq1p3EibQ21nXqj9lNvbbH9cnOB6Gh+5quCpiYQGgr06gVIJEpkomSU7tz9c9if\nsh+l5aXCZma6ZhjdfTS6GHVRXaWaEHXsCCGEENLqMMavSXfwIFBWVpluZQWMGQNYWCiRiZJRusfF\njxGTGIO0vDRhEw4c+nTpg2C74FYfpauqTY+xi4iIQFBQUJv79UIIIYS0ZXl5fIAtrbKfBYkECAoC\n+vdXbZTuzL0zOJh6UBSlM9czx+juo2FjaKOyOtVFJpNBJpNh0aJFKhlj16Y7dm20aoQQQkibxBj/\nfNe4OP55rwodO/Jj6Tp3ViITJaN0j4ofISYxBul56cImHDj0s+2HILsgaEqa96YmTZ4gbVpbHCfS\nXlDbqTd/k3VSAAAgAElEQVRqP/Wmzu339CkQEwMkVVn/l+P4CN3Agfy4ugYpGaU7nXkaB1MPoqyi\n8h6vhZ4FRncfDWtDa9VVqgVQx44QQgghLYYx4OpVYM8eoLi4Mt3MjB9LZ6PM3VAlo3S5RbnYmbgT\nd/LvCJtIOAn6demHgXYDmz1K1xToViwhhBBCWkRhIRAbC1y/Lk7v3Zuf9aqlzJwFJaJ0FawCpzJO\n4VDaIcgr5MJmlvqWGN19NKwMrFRToRdAt2IJIYQQorZu3gR27eI7dwrGxvxYOjs7JTJQMkr3sOgh\ndt7cibtPKtdLkXASDLAdgMCugdCQaKimQq0EdexIq6TO40TaO2o79Ubtp97Uof2ePQP27gUuXRKn\n9+zJP0FCW1uJTJSM0v2d8TcOpx0WRek6deiEUS6j0NlAmZkY6oc6doQQQghpFsnJ/ASJJ08q0wwM\n+ACbk5MSGSgZpXtQ+AA7E3ci40mGsImEkyCwayAG2A5oc1G6qmiMHSGEEEKaVGkpsH8/cPasON3T\nExgyBNDVVSITJaN0J+6egCxdJorSde7QGaO6j0KnDp1UU6EmQGPsCCGEENLq3b7NPxLs8ePKNH19\nYPhwwNVViQyUjNLlFOZg582dyHyaKWyiwWlgoN1A9OvSr01H6aqijh1pldRhnAipHbWdeqP2U2+t\nqf3KyoDDh4G//xb3x1xd+U6dvr4SmSgZpUu4kwBZugzlrFzYzMrACqNcRqFjh46qq5QaoI4dIYQQ\nQlQqMxPYsQN4+LAyTUcHGDoU8PDgFx6ul5JRuuyCbETfjMb9gvvCJhqcBoLsgtDPth8knDLPHmtb\naIwdIYQQQlSivBw4cgQ4dkzcH3N2BkaMAAwNlchEiShdeUU5jt85jqO3j4qidNYG1hjVfRQs9S1V\nV6lmQmPsCCGEENJqZGXxUbrs7Mo0qRR46SXAx0d1UbqsgixE34xGVkGWsImmRBPBdsHo06VPu4zS\nVUUdO9IqtaZxIqRxqO3UG7WfemuJ9quoAI4f5yN15ZXBM9jZ8YsN/9Mfq5+SUbpjd47h6O2jqGAV\nwmY2hjYY3X00zPXMVVYndUYdO0IIIYQ8l4cP+ShdZuVEVGhq8v2xgADVRenuP72P6JvRyC6sDAdq\nSjQRYh+C3ja9232Uriq1G2N3+vRpzJkzB1paWrC2tsZvv/0GTc2a/VMaY0cIIYQ0Dcb42a6HDvF9\nMwUbGz5KZ65M8EyJKJ28Qo6jt4/i+J3joiidrZEtRrmMgpmemeoq1cJU1W9Ru45dVlYWTExMoK2t\njfnz56Nnz554+eWXa2xHHTtCCCFE9R4/5telu327Mk1DAwgOBvr2BSQNBc+UjNLde3oP0TejkVOY\nI2yiJdFCqEMoAqwD2lyUrt1OnujUqXLVaC0tLWhotI8FB9sbGuejvqjt1Bu1n3pryvZjDDh3jn+C\nRGlpZXqnTsCYMUBHZZaLUzJKdyT9CBLuJoiidF2NumJU91Ew1TVVXaXaILXr2Cncvn0bBw4cwMKF\nC1u6KIQQQkib9uQJ3x9LSalMk0iAAQOAwEA+YlcvJaN0mU8yEX0zGg+KKjt+WhIthDmEIcA6AFyD\ng/ZIi92KXbFiBTZs2ICrV69i/PjxWL9+vfDeo0eP8Pbbb+PAgQMwNzfH0qVLMX78eOH9J0+eYMSI\nEVi7di2cnZ1rzZ9uxRJCCCEvhjHg8mVg717g2bPKdAsLPkpnZaVEJkpG6eLT4nHi7gkwVH532xnb\nYaTLyHYRpVP7W7HW1tZYsGAB4uLiUFxcLHrv3XffhY6ODnJycnDhwgUMGzYMXl5ecHNzg1wux7hx\n4xAREVFnp44QQgghL6agANi9G7h5szKN44A+fYCQEH72a72UjNLdzb+LnYk78bCo8jEVUg0pwh3C\n4WflR1G6RmrxyRMLFixARkaGELErLCyEqakprl27BicnJwDA5MmTYWVlhaVLl2LTpk2YO3cuPDw8\nAAAzZ87Ea6+9ViNfitipNxrno76o7dQbtZ96U1X7Xb/Od+qKiirTTEz4KJ2trRIZKBGlKysvQ3x6\nPE7ePSmK0tkb22Oky0iY6Jq8cD3UidpH7BSqV+LWrVvQ1NQUOnUA4OXlBZlMBgCYNGkSJk2apFTe\nU6ZMgZ2dHQDA2NgY3t7ewgmvyI9et87XFy9ebFXlodf0ml7T6/bwurgY+OorGdLSADs7/v30dBlc\nXICZM4MglTaQn1wO2fffA1evIuif719ZejrQqROC5swBjI0hk8mQXZCNHIsc5BbnIv1iOgCgW89u\nGOQ4CE8Tn+LSqUut4vNoyteK/6enp0OVWl3E7tixY3jttddw/37lA33XrFmDzZs3Iz4+Xul8KWJH\nCCGEKO/WLWDXLuDp08o0Q0P+rqmjoxIZKBmlO5R2CKcyTomidI4mjhjhMgLGOso8pqJtarMRuw4d\nOuDJkyeitPz8fBgYGDRnsQghhJB2oaQEiIsDzp8Xp/v4AIMHAzo6DWSg5Fi623m3sTNxJx4VPxI2\n0dbQxmCnwfDp5ENj6VRE0tIFqN6Q3bp1g1wuR3JyspB26dIl9OjRo9F5R0ZGikKeRH1Qu6kvajv1\nRu2n3hrbfmlpwE8/iTt1HToA48fzfbIGO3UZGcDq1UBCQmWnTioFhg0D3nwTMDZGaXkp9ibtxfqL\n60WdOidTJ8zynwXfzr7tulMnk8kQGRmpsvxaLGJXXl6OsrIyyOVylJeXo6SkBJqamtDX18fYsWOx\ncOFCrF27FufPn8euXbtw8uTJRh9DlR8UIYQQ0laUlQEHDwKnTonT3d35PpmeXgMZKBmlS3uchpjE\nGDx+9ljYREdTB4MdB8O7k3e77tApBAUFISgoCIsWLVJJfi0WsVu8eDH09PSwbNkyREVFQVdXF0uW\nLAEA/PTTTyguLoalpSUmTpyIVatWwdXVtUnLI5FIUFR1+g8Ac3Nz3Llzp8597t27h5CQEJWX5cyZ\nMwgPD4eTkxMCAgIQGhqKY8eO1bvPpUuXsG3bNlFabXVSpenTpyMhIUHp7e/du4fg4GAYGxvD39+/\n3m2nTJkCV1dX+Pj4wMfHBwcOHBDee/bsGWbOnIlu3brB09MTM2bMaHTZIyMjUVZW1uj9du7cCT8/\nP3h4eKBHjx749ttv69z2ww8/hIODAyQSCa5fvy5679atW+jTpw9cXFzQt29fUYRaWdHR0XBzc0PP\nnj1x69atRu9/5MgR0ef6IvLz8/HVV18BqBwgHBQUhNjYWJXk31I2bNiAV199VSV5ZWdno0+fPsLr\nsrIyLFy4EC4uLvDy8oKvry8+/PBDyOVy7N69G++++65of4lEAi8vL3h7e8PX1xeHDx9u8JjVz/Mp\nU6Zg5cqV9e6jaD+inpRpv7t3gVWrxJ06XV3glVeAV19VolOnZJQu9lYsNl7aKOrUdTPrhln+s+DT\nmW69NpUWi9hFRkbWGVEzMTHBjh07mrdAtWjopLOyslLqj2tjXLlyBcOHD0dUVBTCw8MBAKmpqcIs\n0bpcuHABsbGxNb6EmnICyZo1axq1fYcOHfD5558jPz8fERER9W7LcRz+/PNPuLm51Xjvo48+gp6e\nntCZycnJqbFNQz777DPMmzcPWlpajdqvc+fO2L17Nzp16oQnT56gZ8+eCAgIQP/+/WtsO2bMGMyZ\nMwcDBgyo8d4777yD2bNnY8KECfj9998xY8YMHDp0qFFlWb16NRYvXlzrs5KVER8fj8LCQuE8a4yK\nigpIJJW/Cx8/foyvv/4aH330kZDWnH+0q5dHVZ6nDnWVZdmyZXjnnXeE11OnTkVJSQnOnz8PfX19\nlJeX49dff0VJSQmGDx+OTz/9FBkZGbCxsRH2OXnyJPT09BATE4PXX38dD6oOUq9F9fOcvkjbN7kc\nkMnE/TEA6NYNGDmSvwXbYAZKROlSH6ciJjEGec/yhE10NHUwxGkIPDt60nnYxFp8jF1TUtUYO8YY\nZs2aBVdXV3h7ewtf4unp6TA3Nxe2k0gkWLp0KQICAuDo6Ii//vpLeO/PP/+Eq6srfH198cUXX9QZ\nTVu2bBmmTZsm+rJ1cHDA2LFjsW3bNgwfPlxILykpgZWVFe7cuYOFCxfi4MGD8PHxwZw5c4Rtfvzx\nx1rLs2/fPvj6+sLLywthYWFI+ec5MTKZDN7e3njnnXeE6MDNqqtTVlE1IvPLL7/Azc0NPj4+8PLy\nQmJiYo3tDQ0N0a9fP+g1+HOQj8rV1iktKCjApk2bsHjxYiHN0tISAHDz5k3Y2toKUdZFixaJnlii\noIiE9O3bFz4+Pnjy5Amys7MxZswYeHl5wdPTE5s2baq1XAEBAcLzig0NDeHq6lpnVLdfv36iL2UF\nxcLbirKNGzcO58+fR25urtJ1mDt3Lo4fP46PPvoIoaGhAIA33ngD/v7+8PT0xNixY5GXx/9RTUxM\nRJ8+feDt7Q0PDw988803uHr1KlavXo3ffvsNPj4+QrRtz5496N+/P/z8/NC3b1+c+ucnvUwmg6en\nJ9566y34+Phg3759NT7TvLw8+Pj4CGtMAnxUcMCAAXB0dMR//vMfIf2bb75BQEAAfH190bdvX1y6\ndEl4r77rqKrIyEi8+uqrGDx4MNzd3ZGXl1dn+QHg008/hbOzM3r37o2PP/5YiBpXj8pVfV31HMzK\nykJISAj8/PzQo0cPfPzxx3WWJT8/X1RWuVyOzZs345VXXgEAJCUlITo6GmvXroW+vj4AQENDA9On\nTxdejx07Fr/99lutdQ8LC0Nubi4yMjJgZWWFrKws4b33338fS5cuxXvvvQeAP899fX2FMl29ehWh\noaHo1q0bJk+eLOynuAYcHR1rXAN2dnaIiIhA3759YW9v32DUj7Scur7z7t8HfvkFOH68sk+mrQ2M\nHs2Pp2uwU6dElK5EXoLdt3bjt0u/iTp1LmYueNf/XXh18qJOXS1UPcYOrI1qbNU4jmOFhYWiNHNz\nc3b79m12/vx55urqKqTn5eUxxhhLS0tj5ubmojxWrlzJGGMsISGBWVtbM8YYy8rKYmZmZiw5OZkx\nxth3331X6/EYY8zNzY3t3Lmz1jLK5XLWtWtXlpaWxhhj7LfffmNjx45ljDG2YcMG9sorr9SoU23l\nyc7OZhYWFuzGjRuMMcbWrVvHevXqxRhjLD4+nmlpabGLFy8yxhhbsmQJe+ONN2otT1BQEIuNjWWM\nMWZkZMSysrIYY4yVlpayoqKiWvdRHMPPz6/O9xljrFOnTszDw4N5eHiwWbNmCZ/5xYsXmaOjI5s3\nbx7z8/NjQUFB7Pjx48J+mzZtYr1792ZxcXHMxcWFPX36tNb8q3/+r732Glu4cCFjjLH79+8zKysr\ndvXq1XrLeOPGDWZhYcHu379f73Z2dnbs2rVrwuuzZ88yd3d30TZubm7swoULjapD1c+fMcYePnwo\n/P/TTz9ln3zyCWOMsffff58tXbpUeE/xWUZGRrJ58+YJ6cnJyaxPnz7syZMnjDHGrl69ymxtbRlj\nfJtpaGiwv//+u9aypKenC9dCfHw8Y4yxgQMHsnHjxjHGGMvPz2fm5ubCNfDgwQNh3wMHDrDevXsL\nr+s6b6uLiIhgtra2LDc3t8Hyx8TEMC8vL1ZUVMQqKirY2LFjmb+/P2OMsfXr14uunaqvq/7/2bNn\nrKCggDHGn+MhISFs3759tZalutOnTzMfHx/h9R9//MG8vb1r3VZh//79LCQkRPS5KI6/fv16oW6f\nfPIJW7RoEWOMsadPnzJLS0vh861+nk+ePJkNGDCAlZSUsNLSUubu7s4OHDjAGKu8BuLj44VrQHHe\n2tnZCedKeno669ChQ61/v0jLU1x/CnI5Y/HxjC1axFhEROW/jRsZ++dPQf3Kyhjbv5+xyEhxBhs2\nMPb4sbBZcm4y+/bEtywiPkL49+WxL9mlrEusoqJCRbVr21TVJWvTETtV4DgOjo6OKCsrw1tvvYWo\nqKh6b2+OGzcOANCrVy/cu3cPpaWlOHXqFHx9feH4z0JAU6dOfa6yaGhoYMaMGVi1ahUAYOXKlUL0\nqa4y1VUeLy8vdO/eHQA/7ubixYsoLCwEAGHMj2K/lKpPfa5DSEgI3nzzTaxYsQIZGRnQ1dV9rjoq\nnD17FpcvX8bZs2fBGBOiD+Xl5UhNTYWvry/OnDmDZcuWYezYsXj6z8JLEydOhIuLC8aMGYMtW7ag\nQ4M/Q3mHDh0Sxup16tQJQ4cOrXfdxPv372P06NH4+eefhQieqjSmDlXbfePGjfDz84Onpye2bNki\nRMEGDhyItWvXYuHChYiPj4eRkVGt+8fFxSElJQWBgYHw8fHBxIkTUV5eLtzuc3Z2Rq9evRosh2KM\nD8dxQuRLEd1UnEtnz55FYGAgPDw88MEHH9QYalDbeVsdx3EYNmwYTE1N6y1/Tk4O4uPj8frrr0NX\nVxccx2Hy5MmNHqYgl8vx4YcfwtvbG35+frh69aoo0li1LNWlpaXB2tq6UceztrZGamqqKE0RZf7j\njz8QHR0NgI+Wrl+/HuXl5YiKisLgwYNFdxKq4jgOo0ePhlQqhZaWFnx9fYVjKK6BoKAg4RqoOtRE\n0SZdu3aFiYkJMjIyGlUf0jyqjrHLyQHWruVvv1ZU8GlaWnyQbdIkoMqfgtopEaV7Jn+GmMQYbLq8\nCfkllZHq7ubd8W7Au3TrtQW0+Dp2rYWFhQUePnwI23+elSKXy5Gfnw8LCwvo6Ojg2rVrkMlkOHjw\nID7++GNcuHCh1nx0/pkbrqGhIeTTGL6+vjh16hRGjhxZ6/v/+te/4OPjgxEjRiA/P7/ByRvPUx6d\nKvPbNTQ0lNrnr7/+wpkzZ3D48GEEBwdj1apVeOmll2rdVpmLXPElKJVKMXPmTIwaNQoAYGtrC01N\nTeFLJiAgAObm5khKSoKvry9KS0tx7do1mJiYiG5PKaPqFz1jrM5y5uTkIDw8HB9//PFzjW/r0qUL\nMjMzhWOUl5fj3r176NKlCwA0qg6KMh47dgyrVq3CyZMnYWZmhs2bNwtjIMeOHYu+ffsiLi4OX375\nJX799Vds2rSp1o7NSy+9hI0bN9Z6LGU7yVXVdi6VlpbilVdewfHjx+Ht7Y179+7VuGVd23krlUpr\n5K+4bdlQ+asv/Fn1/5qamqhQfOuBHwZQm2+//RZ5eXk4ffo0pFIpZsyYIWzLcVyNslQ/flU+Pj5I\nSkpCXl4ejI1rX5C1tsVKFWPsqrKxsYGfnx+io6Px008/NTj2VVtbW/h/9eu7vmvgef4ukJZRUQGc\nPAkcPgyUl1em29ryt17r+P1RScmxdMmPkhGTGIMnJZVrz+pp6WGo81C4W7hTh66FtOmIXWPG2IWH\nh2P16tXC619++QV9+vSBjo4OHj58iMLCQgwaNAhLly6FkZFRjV/S9enVqxfOnz8v7FPXFycAzJs3\nD2vWrBENpE9LSxPGGZmZmSEsLAzjx48XzZozMjKqMa6nLr1798alS5eEcXAbN26Er69vvV9M9Skv\nL0dKSgr8/f3x8ccfY9CgQfVO9mgoUlJUVITdu3cL2/7vf/+Dj48PAH6mcnBwsDCb89atW8jJyREe\nQTdv3jz4+/tj//79eOedd5CZmVnrMQwMDIQxaAA/ZknxhZiVlYW9e/fW2mnOzc1FeHg4Zs+e3ajI\na9U6W1pawtvbG5s3bwYAbNmyBb6+vjAzM2tUHarKy8uDkZERTE1NUVJSgl9//VV4Lzk5GZaWlpg8\neTIWLlyIM2fOAKh5zgwaNAj79u0TzeBVbNsQQ0NDFBUVoby8XHTN1dbWz549Q3l5udCZ++mnn5Q6\nRnXV866v/EFBQdi+fTuKi4tRUVGBTZs2CV86Tk5OuHz5MkpLS1FaWort27fXerz8/Hx07twZUqkU\nmZmZ2LlzZ731rMrOzk7Ujs7Ozhg5ciRmzJiBgoICAPx1tG7dOiFynpGRAXt7e6U+i9mzZ2POnDmQ\nSqWiqGr187w+imtAJpPVew2Q1ikx8TZWrjyMt9/+HpMmHcaWLbeFTp2mJjBoEDBlihKdOiWjdDtv\n7kTU5ShRp87Nwg2z/Gehh2UP6tQ1gqrH2LX5jp2yU/e///57pKenw8vLCz4+PoiLixMGD9+9exfh\n4eHw9vaGl5cXhg4dit69ewMQ/xKvfiIrXnfs2BGrVq3C0KFD0bNnTzx8+BBaWlq1TiLw9PTErl27\nsHTpUjg5OcHT0xPTpk0T3e57++238fjxY9HA59DQUBQWFsLb21uYPFFXeSwsLLBp0yZMmDABXl5e\n2Lx5M6KiooRtqtepoQu0vLwcU6dOhaenJ7y9vZGVlVXrEiSKL/PXXnsNly9fRpcuXfDZZ58B4G/N\nDRs2DADfsfq///s/eHl5wcPDA8nJyaIv/1WrVuGLL76Ap6cnxo8fj6ioKBgaGiI6OhpHjx7F999/\nDzc3N0RERGD8+PGiaIzCBx98gJCQEPj6+uLJkyf48ccfcenSJXh5eWHQoEFYtmxZrUvsfPnll0hO\nTsaqVauEpVgUHfWqdQD4QeyK6FxYWJhoUsGqVauwfPlyuLi4YOXKlcLt9cbUoaohQ4bA0dER3bp1\nQ1BQEHr27Cm027Zt2+Dp6QlfX1+8//77+OGHHwDws3bPnDkjTJ5wcnJCVFQU3n77bXh7e8PNzU0U\n/anvPDA1NcUbb7wBDw8PzJ49u959DA0N8dlnn8Hf3x9+fn7o0KGDUtdRddXPzfrKP2LECAwePBie\nnp7o06cPrK2tYWhoCID/oRMWFgZ3d3eEh4fDzc1NyLfqMd5//30kJCTAw8MD06ZNQ1hYWJ1lqc7b\n2xuZmZmiCVMbN26Es7MzevbsCQ8PD3h6eiIxMVGIqJ04caLGMeoSGBgIXV1dzJo1S5Re9TxXdOLr\nykdxDbz99tv1XgOk9UlMvI0NG5Jx8WIIzp/3RmZmCC5eTMbDh7dhZQXMmAH07QvUO3FcLgcOHADW\nrRM/EszeHpg1C/D3BzgOt3JvYeXplbiQVXnXSk9LD6+6vYrX3F9DB2njI/vtXVBQkEo7di3+rNim\n0tqeFVtQUCDcylq/fj3Wr1+Po0ePPlden3/+ObKzs7F8+XJVFpGQNk1xDVZUVGDatGmwsbERflg0\nhzlz5sDHx0f0g6w+3t7e2L17d60zq6tLS0tD//79kZKSIrplStqHr746jBMnQlA1OMtxgJfXYXzz\nTQj+GdFQNyWe8VpcVox9yftwKfuSaFd3C3cMdR4Kfenz3fEhldrMs2Lbix9//BHbtm2DXC6HmZlZ\no9eAU3B3d4dUKkVcXJyKS0hI2/bmm28iPT0dxcXF8PPzE6251xz+85//YPTo0Up17GJjY+tcLqe6\nhQsXYv369fj222+pU9fOMAacPg3IZBJUXT1LXx/o3h3o0kVSf6dOybF0iQ8TsevWLhSUFlQeQ0sf\nw7oNg5tFzbVGScuiiB1plWQyGa2Ar6ao7dQbtZ96ePgQiIkB7twBTp8+jKIifjykVCpD795BkEgA\nS8vDmDWrjnGSSkTpisqKsC95Hy5nXxbt6mHpgSHOQ6Cn1fCapER5FLFTgmKMHf2RIoQQ0hYoZrzG\nx/MBNwBwcHBEUtIh9OgRitxcfixdSckhhIY61cxAySjdjQc3EJsUK4rSdZB2wPBuw9HdvHtTVrHd\nkclkKnmYggJF7AghhBA1kJPDB9mqTpSXSIABA4COHW9DJktBaakEUmkFQkMd4eLSVZyBklG6PUl7\ncDXnqmhXz46eGOI0BLpaL7ZGKambqvot1LEjhBBCWrHycn71kSNHxOvSde7MB9kaXCNdySjd9QfX\nEXsrFoVlhcImBlIDDO82HC7mLiqsEakN3YolbRqN81Ff1Hbqjdqvdbl/nw+yVV2rXEMDCArilzCp\nPjmiRvspEaUrLC3EnqQ9uPbgmigv707eGOw4mKJ0aoY6doQQQkgrI5cDR48Cx49XPg4MAGxs+CCb\nhYUSGTQQpWOM4VrOVexJ2oOisspptYbahhjRbQSczZxVWynSLOhWLCGEENKKZGYC0dHiIJumJhAa\nCvTq1cBCw4BSUbqC0gLE3orFjYc3RLv6dPLBYKfB0NGkpXOaG92KVQLNiiWEEKIuysoAmaxmkK1r\nV2DkSOCfpw7W6nZiIlLi4iC5cQMVd+7A0d4eXc3N+TerRemuZl/BnqQ9KJYXC/sbahtipMtIOJnW\nMpOWNKlmmxU7adIkpTLQ1tbG2rVrVVYgVaGInXqjcT7qi9pOvVH7tYw7d/ggW25uZZpUCoSFCU/z\nqtPtxEQkL1+O0NRUyO7dQ5CxMQ7J5XDy90fXCROEKN3TkqeITYrFzYc3Rfv37NwTgxwHQVtTu4lq\nR5TR5BG7rVu3Yv78+XUeRFGAb775plV27AghhJDWrrQUOHSIf4JE1a9bBwc+SvfPhNW6yeVIWb4c\noVfFy5OEmpvjsLU1uvr7gzGGy1mXsC95nyhKZ6RthJEuI+Fo6qjCGpGWVmfEztHRESkpKQ1m4OLi\ngsTERJUX7EVRxI4QQkhrlpbGPz3i8ePKNG1tYPBgwMen/igdAGEwnmzXLgQ9e8anaWjwvUIrK8hM\nTNBz5tvYdWsXbuXeEu3qZ+WHcIdwitK1IrSOXQOoY0cIIaQ1KikB9u8Hzp0Tpzs7AyNGAIaGDWQg\nl/OD8RISAMZw+PRphBQV8eE9FxdAVxeMMfymU4r7A4zwTP5M2NVYxxijXEbB3sRe5fUiL0ZV/ZaG\n5tbUKjU1Fenp6S98cELqosqBpKR5UdupN2q/ppWUBKxcKe7U6eoCY8YAEyYo0anLzARWr+bXQfmn\nE+Do4oJDdnaAlxdk2dkokZfgx4wLuGL9VNSpC7AOwCz/WdSpa+OU6tiNGzcOJ06cAACsX78e7u7u\ncHNzo7F1hBBCiBKKi/klTH7/HXjypDLd1RV4913Ay6uBW69yOXDwILB2rXgZEzs7dF24EJIJr2Ph\n3RtYkXYREy/vxmVXA3Sw5WfFmuiYYIr3FAx1HgqphrRpKkhaDaVuxVpYWCAzMxNSqRQ9evTA6tWr\nYeN/7AEAACAASURBVGxsjFGjRiE5Obk5ytloHMchIiKCljshhBDSom7eBHbvBgoKKtP09IBhwwA3\nN+XH0tVYl+6fKbOJKbew+sBqpJmk4fEzfsCePFkObzdvDPMfhlCHUOrQtWKK5U4WLVrUfGPsjI2N\nkZeXh8zMTAQEBCDznycQGxgY4OnTpy9ciKZAY+wIIYS0pMJCYO9eoNqEVfToAQwZAujrN5CBXM4/\nIDYhQfz4CTs7fl06ExMwxvDxLx/jnM45lLPKB8nqauqir7wvFkxZoLL6kKbVrAsUe3l5YenSpUhP\nT8ewYcMAABkZGTAyMnrhAhBSG1pLS31R26k3ar8Xxxhw/TqwZw/fuVPo0AEYPhzo3l2JTO7d46N0\nOTmVaVpa/NMj/lnY7mHRQ8QkxuDyg8sot+E7dXk389AjoAfsje1hkGOg2ooRtaBUx27dunVYsGAB\npFIpvvrqKwDAyZMn8cYbbzRp4QghhBB1UlAAxMYCN8RP6oK3N7+Mia5uAxnUFaXr2pWP0pmaoryi\nHCfvnoQsXQZ5hRySf4bL62npwdzUXHh6hFRCt1/bI1ruhBBCCHlBjAGXLwP79vETJRQMDfklTJyd\nlcikrihdWBgQEABwHO4/vY+YxBjcL7gvbJJ7Pxe5Gblw8nOChOM7eSVJJZgSPAUuTi4qqiFpas2+\njt2xY8dw4cIFPH36VDg4x3GYP3/+CxeiKVDHjhBCSHN48gTYtYtfyqQqPz/+zql2Q2sAl5fzUbrj\nx+uM0skr5DiSfgQJdxNQwSq36dyhM0Z1H4X8rHwcOn8IpRWlkEqkCPUNpU6dmmnWMXazZ8/G1q1b\nMWDAAOg2GEcm5MXROB/1RW2n3qj9lMcYcOECEBfHLzqsYGzMPw7MwUGJTO7f56N02dmVadWidHfy\n72DnzZ3ILa58kKymRBPBdsHo06UPJJwEnZw6wcXJhdqPKNexi4qKwrVr12BlZdXU5SGEEEJavbw8\n/nFgqamVaRzH98VCQ/nVSOqlRJSuRF6CQ2mHcDrztGjXrkZdMdJlJMz0zFRXIdJmKHUr1tPTE4cP\nH4a5uXlzlEkl6FYsIYQQVWMMOHOGXyu4tLQy3cyM74/Z2iqRSV1RutBQoFcvgOOQlJuE3bd2I78k\nX9hEW0Mb4Y7h6Nm5J7gGF78j6qZZx9idOXMGX3zxBSZMmICOHTuK3gsMDHzhQjQF6tgRQghRpdxc\nPkp3+3ZlGscBffoAwcF836xe5eXA0aPAsWPiKJ2tLTB6NGBqiqKyIsQlx+FS9iXRrt3MumGY8zAY\n6dAyY21Vs46xO3fuHPbs2YNjx47VGGN39+7dFy5EU4mMjKQnT6gpGieivqjt1Bu1X00VFcDffwOH\nD/OrkShYWPBROhsbJTLJyuKjdFlZlWlVonQMwPWca9iTtAeFZZWL3+lp6WGI0xD0sOyhVJSO2k/9\nKJ48oSpKdew+/fRT7N69G+Hh4So7cHOIjIxs6SIQQghRYw8eADt3AhkZlWkSCdC/PxAYCGg29C1a\nX5Ru1CjAzAxPS54iNikWNx/eFO3qYemBl5xegr60oUdUEHWmCEAtWrRIJfkpdSvW1tYWycnJkDY4\nGrT1oFuxhBBCnld5OXDiBCCT8f9X6NSJ74917qxEJvVF6QICwDgOF7IuYH/KfjyTPxM2MdQ2xDDn\nYXAxp+VK2pNmHWO3YcMGnD59GgsWLKgxxk4ikbxwIZoCdewIIYQ8j6wsPkp3v3INYGhoAAMHAv36\n8f+vV3k5H6E7erTOKN2j4kfYlbgLaXlpol39rPwQ5hAGHU0d1VWIqIVm7djV1XnjOA7lVX/KtCLU\nsVNvNE5EfVHbqbf23H513TW1tub7Y5aWSmRSW5ROU1MYS1fBAacyTuFw2mGUVZQJm5jqmmKky0jY\nGdu9UB3ac/upu2adPJFadaEeQgghpI2p7Wlempr8bNc+ffhxdfWqK0rXpQvfKzQ3R05hDnbe3InM\np5nC2xw49O3SF0F2QdDSaGhaLSENo2fFEkIIabfkcn4cXUICv0adgq0t//QIpZZvzc7me4VV791W\nidLJUYHjd47j2O1jKGeVd7k66nfEqO6jYGVAi/+TZojYLViwAIsXL24wg4iICJXN5CCEEEKay927\n/Fi6hw8r0xRP8/L3VzJKd/w4H6WrOiypSpQu40kGYhJjkFNYGQrU4DQw0G4g+nXpBw1JQwP2CGmc\nOiN2HTp0wOXLl+vdmTGGnj17Ii8vr0kK9yIoYqfeaJyI+qK2U2/tof3KyoBDh4BTp8RROnt7Pkpn\nYqJEJnVF6UJCgN69UcrkiE+Lx98Zf4Oh8iBdDLtgpMtIWOhbqK5CVbSH9murmjxiV1RUBCcnpwYz\n0NbWfuFCEEIIIc0hPZ1/esSjR5Vp2tpAeDjQsyf/JIl61RWls7Hhnx5hbo7Ux6nYlbgLj589Ft7W\nkmghzCEM/tb+kHCtczUJ0jbQGDtCCCFtXkkJ/3zXM2fE6U5OwIgRgJEyT+pqIEr3rKIU+1P24/z9\n86LdHE0cMcJlBIx1jF+8IqTNatZZsYQQQoi6Sknho3T5+ZVpOjrASy8BXv+fvfuOjvI6Ez/+naLe\nOxISEqo0U2x6Fb2YYhwHG8fENnZibxwf/36bdn6OicHeTc5uNol343WctRPjEve1Te8gquggQIBA\nEqoIBOoSKtN+f7xoRkMdaQp60fM5xyfMHc373jfXY66ee+/zDHEgSmc2K1G6nTtvG6U7e/Us686t\no6GtwXYPvS+zUmcxJGaIQ+XAhHAFidiJbkn2iaiXjJ263U/j19ICmzbBsWP27RkZMHcuBAU5cJHK\nSiVKd/Gira1DHpRG4zU2nN9A7pVcu48NiBrAnLQ5BHoHOv8gnXA/jV9PIxE7Byxfvtxag00IIUTP\nce4crFkDDbYAGv7+MHs2DBrkRJSud2945BEskZGcuHyCjfkbaTY2W98O9A5kTtocBkQNcO0DiftW\nVlYWWVlZLrueROyEEELcN65dg40b4cakDgMHwpw5EBDgwEXuEqWrbatn7bm15Ffn231sWK9hzEiZ\ngZ+Xn/MPInocj0bsKisr8fPzIygoCKPRyEcffYROp2PJkiXdtlasEEKInuX0aVi3DpqabG2BgcqE\nboAjATSzWclUnJV12yjdoYuH2Fq4lTZTm/XtUN9Q5qXPIyU8xWXPIkRXORSxGzlyJH/9618ZNmwY\nv/rVr1i7di1eXl5kZmby1ltveaKfnSYRO3WTfSLqJWOnbmocv8ZGWL9emdh1NGQIzJypLMHe1ZUr\nSpSu3FbuC51OidKNHcvVlmpW562mpK7E+rYGDaPiRzGl7xS8dd6ueRgnqXH8hMKjEbvz588zdOhQ\nAD755BP27dtHUFAQAwYM6LYTOyGEEPc3iwVOnoQNG6DZts2NoCAlhUl6ugMXMZth3z7YsePmKN2C\nBZgiI9hXupesoiy7cmBR/lHMz5hPQkiC6x5ICBdwKGIXGRlJWVkZ58+f54knniA3NxeTyURISAiN\njY2e6GenScROCCHuX/X1yrJrXp59+4MPwowZSjqTu7pLlK6i6TKr8lZxqfGS9W2tRsuEPhOYkDgB\nvfa+Pn8oPMyjEbtZs2axaNEiqqqqePzxxwE4ffo08fHxTndACCGEcJTFAsePK2lMWlps7aGhSpQu\nxZFtbreL0sXFwSOPYIgIY2fRdvaV7sNsMdveDopjQcYCYgJjXPdAQriYQxG7lpYWPvzwQ7y9vVmy\nZAl6vZ6srCwuXbrEE0884Yl+dppE7NRN9omol4ydunXn8autVVKYFBTYt48cCVOnKqXB7up2UbrM\nTBg3juL6Ulbnraaqucr6tl6rZ0rfKYyOH93ty4F15/ETd+bRiJ2vry8vvPCCXZv8iyOEEMITLBY4\nfBi2bIE222FUwsNh/nxISnLgImYzZGcrUTqj0dZ+PUrXGh7C1vwNHLpoX3MsKTSJ+RnzCfcLd8mz\nCOFut43YLVmyxP4Hr2dztFgsdqVRPvroIzd2r+skYieEEOpXXa2UAysqsrVpNDB6tFKi1cvLgYtc\nvapE6crKbG0donTnawpYc24N9a311rd9dD7MSJnBg7EPSjkw4RFuj9ilpKRY/2W+evUqH374IfPm\nzSMxMZHi4mLWrl3L008/7XQHhBBCiBuZzXDwIGzbBgaDrT0yEhYsgARHDqPeJUp3LSyQjXnfceKy\nfTbj9Ih05qbPJdgn2DUPI4QHObTHbsaMGSxbtowJEyZY2/bs2cMbb7zB5s2b3drBrpKInbrJPhH1\nkrFTt+4wflevwqpVUFpqa9NqYdw4mDRJKQLh0EVuFaWbNAnL2LHkVp9lw/kNNBls2Yz9vfyZkzaH\ngVEDVRul6w7jJ7rGo3vs9u/fz+jRo+3aRo0aRXZ2ttMdEEIIIcB2WDUryz7AFhOjROni4hy8yP79\nsH27/UViY+GRR6gP9WPdma/Iq7LPkzI4ZjCzUmfh7+VINmMhui+HInaTJk1ixIgRvPnmm/j5+XHt\n2jVef/11Dhw4wK5duzzRz06TiJ0QQqjH5ctKlK5jeVatFiZOhAkTlGDbXd0q1NchSne0MofNBZtp\nNbVa3w72CWZu+lzSIxzJZiyE+3g0Yrdy5UqefPJJgoODCQsLo6amhuHDh/Ppp5863QEhhBA9l8kE\ne/bArl03p5RbsECJ1t3VXaJ01cFerD71CUW1RXYfGxE3gmnJ0/DRO5InRQh1cChi166kpISLFy8S\nGxtLYmKiO/vlNInYqZvsE1EvGTt18+T4VVQo2+AuX7a16fXKYdWxY5WI3V1VVSkXuTFKN3Ei5nFj\n2V9xiO0XtmM02yZ8EX4RzM+YT2Jo9/57rCvk+6deHo3YtfP19SU6OhqTyURhYSEAycnJTneiM+rr\n65k2bRpnzpzhwIEDDBgwwKP3F0II4RyjEXbuhL17lWBbu/h4JUoXFeXARcxmOHBAOTbbMUrXqxcs\nXMjlAFids5LyBlsiYq1Gy9iEsUxKnISXzpE8KUKoj0MRu40bN/Lcc89RUVFh/2GNBlPH2LkHGI1G\namtr+cUvfsHPf/5zBg4ceMufk4idEEJ0P2Vlyja4K1dsbV5eSk66UaOciNJptTBpEsaxo9ldto/d\nJbvtyoH1CuzF/Iz5xAU5cgJDCM/zaMTuJz/5CcuWLeOHP/wh/v739sSQXq8nMjLynvZBCCFE5xgM\nSjq57GylkkS7pCSlekS4I4Ud7hSle+QRSv0MrD72Pleu2WaNOo2OzKRMxiaMRad15ASGEOrm0MSu\ntraWF154QbV5fYT6yD4R9ZKxUzd3jF9xsRKlq662tXl7w/TpMHy4UknirqqqlIuUlNjarh+bbRs7\niu0lOzlw9gAWbLPGhOAEFvRbQKR/zwkGyPdPOFTN+LnnnuPvf/+7S2/89ttvM3z4cHx9fXn22Wft\n3quurmbhwoUEBgaSlJTEZ599dstryERTCCG6r7Y2WL8ePvjAflKXkgI/+QmMGOHApM5iUU68vvuu\n/aSuVy/48Y8pHNKHd47+lf1l+62TOm+dN3PS5rB02NIeNakTAhzcYzd+/HgOHjxIYmIivXr1sn1Y\no+lyHrtvv/0WrVbLpk2baG5u5oMPPrC+t3jxYgD+9re/cezYMR5++GH27dtnd1Di2WeflT12QgjR\nTRUWKjVea2ttbT4+MHMmDBvmYJSuulrZS3eLKF3z6OFsLtrGsUvH7D6SGp7K3PS5hPqGuuZBhPAQ\nV81bHJrYrVy58radcLZe7LJlyygrK7NO7JqamggPDyc3N5fU1FQAnn76aeLi4vjd734HwJw5c8jJ\nySExMZEXXnjhln1o71tSUhIAoaGhDB061BqizsrKApDX8lpey2t57cLXLS3wxz9mce4cJCUp7xcV\nZREfD7/8ZSbBwQ5cb8cOOHOGzJoaMBjIKipS3h81Ch55hA8PbGZ/2X5iBilJ7oqOF+Gt8+al77/E\n4JjB7Ny5s9v8/yGv5fXtXrf/uej6v98ffvih5yZ27vTaa69RXl5undgdO3aM8ePH09Rkq9/3xz/+\nkaysLFavXu3wdSVip25ZWVnWL4FQFxk7dXNm/M6fhzVroL7e1ubnB7NnwwMPdCJKt2qVsjGvnVYL\nEybQOPpB1hdu4vSV03YfGRg1kNlpswn0DuxSv+8n8v1TL4+eirVYLHzwwQd8/PHHlJeXEx8fz1NP\nPcWzzz7r9D63Gz/f2NhIcHCwXVtQUBANDQ1O3UcIIYR7NDfDxo2Qk2Pf3r8/PPwwBDoy37JY4OBB\n2LpVOULbLiYGy4IF5Ggr2XTkXZqNzda3Ar0DeTjtYfpH9XfNgwhxH3BoYvfb3/6Wjz76iJ/97Gf0\n6dOHkpISfv/733Px4kVee+01pzpw4+w0MDCQ+o6/7gF1dXUEBQU5dR+hLvIbp3rJ2KlbZ8fvzBlY\ntw4aG21tAQEwZw7cZgv0ze4QpasdOZg1+espqCmw+8iDsQ8yPXk6fl5+nerv/U6+f8Khid17773H\nzp077cqIzZw5kwkTJjg9sbsxYpeeno7RaCQ/P9+6xy4nJ4dBgwZ1+trLly8nMzNT/kUXQggXa2pS\nTrzm5tq3P/CAsvTqUMrTO0TpzAvmc8hcxrajf6XN1GZ9K8w3jHkZ80gOS3bNgwhxj2VlZdntu3OW\nQ3vsoqOjuXDhAgEBAda2xsZGkpOTqays7NKNTSYTBoOBFStWUF5eznvvvYder0en07F48WI0Gg3v\nv/8+R48eZe7cuWRnZ9O/v+Phdtljp26yT0S9ZOzU7W7jZ7Eok7n16+HaNVt7UBDMnQsZGQ7eqKZG\nidJd3zgOKFG68eO5MnwAq/PXUVpvqyyhQcPo+NFM7jsZb513p56pJ5Hvn3q5at6ideSHZs2axVNP\nPcXZs2dpbm7mzJkz/PCHP2TmzJldvvGbb76Jv78///Zv/8Ynn3yCn58f//qv/wrAO++8Q3NzM9HR\n0Tz11FO8++67nZrUCSGEcL2GBvjiC/j6a/tJ3bBhSl46hyZ17VG6d96xn9RFR2Na+iy7UvS8e+w9\nu0ldlH8Uzz34HDNTZ8qkToi7cChiV1dXx8svv8wXX3yBwWDAy8uLRYsW8ec//5nQ0O6ZK0ij0fD6\n66/LUqwQQjjJYlEORmzcCC0ttvaQEJg3D67vmrm7O0TpLg5LZVX+Oi43Xba9pdEyMXEi4/uMR691\naOeQEKrTvhS7YsUKz6c7MZlMXL16lcjISHS67l1zT5ZihRDCeXV1SgqT/Hz79uHDlZJgPj4OXMRi\ngUOHlL10bbb9ckRHY5j3MFlt59hXus+uHFjvoN7Mz5hPTGCMax5EiG7Oo0uxH374ITk5Oeh0OmJi\nYtDpdOTk5PDxxx873QEhbsWVG0mFZ8nYqVv7+FkscOSIsmLacVIXFgZPP63sp3NoUldTAx9+qGzK\na5/UXT/xWrRoBn+5uIq9pXutkzovrRczU2by3IPPyaSuC+T7JxyKbS9btozjx4/btcXHxzNv3jyW\nLFnilo4JIYS4N2pqlHJgFy7Y2jQaGDUKpkwBb0e2uVkscPgwbNliH6WLiqJ13hy2tORy+NQndh/p\nG9qXeRnzCPcLd82DCNEDObQUGxYWxtWrV+2WX41GIxEREdTV1bm1g10lS7FCCOG4vLxitmwpID9f\nS0GBmcTEFCIjlRRXERGwYAH06ePgxW43Mxw/nnODYllbuJH6Vlu+Uh+dDzNTZzKs1zCnk94LoVYe\nrTzRv39/vv76ax5//HFr27ffftvtT6pKHjshhLi7vLxi/vu/8ykunkr77+rHj29j2DCYPz+RzEzw\n8nLgQneI0l2bM4MNzSc4eWa33UcyIjJ4OP1hgn2CEaInuid57Pbs2cOcOXOYPn06ycnJFBQUsHXr\nVtavX8/48eNd1hlXkoidukkuJvWSsVOX5mb42c+2c+bMFCwWqK3NIjQ0k4AAGDt2O7/+9RTHLlRb\nq5x4vSFKZxk7llMDItlQtIVrBluOlACvAOakzWFA1ACJ0rmQfP/Uy6MRu/Hjx3Py5Ek+/fRTysrK\nGDlyJP/5n/9JQkKC0x0QQgjheWYzHDsG27ZBYaGWjn+fJCYq/wQEOHC+rv2UxebNN0XpGmZPZU3T\nUc6d32v3kSExQ5iZOhN/L0fKUwghOqPT6U4uX75MXFycO/vkEhKxE0KIWysthQ0b4OJF5fXBg9u5\ndm0KYWFKTrr2IkPR0dv5yU/uELGrrVX20hUW2tquR+mOZASxpXgHraZW61shPiHMTZ9LWkSaG55K\nCHXzaMSupqaGl156ia+//hq9Xs+1a9dYvXo1Bw8e5F/+5V+c7oQQQgj3a2hQUsnl5Ni3Dx6cQnX1\nNmJjp9K+Ktrauo2pU2+Tefh2UbrISGpmTmJV0xGKCovsPjKy90im9p2Kj96RHClCiK5yKI/diy++\nSHBwMMXFxfhcT1w0ZswYPv/8c7d2zlnLly+XnD4qJeOmXjJ23Y/JBPv2wdtv20/q9HrIzIQ330zk\nZz9LJSZmO1evvkV09HaeeSaVjIzEmy9WWwsffwxr19omdRoN5rFj2DvnAf67YhVFtUXWH4/wi+DZ\noc8yJ22OTOo8QL5/6pOVlcXy5ctddj2HlmIjIyOpqKjAy8uLsLAwampqAAgODqa+vv4un743ZClW\n3WQDsHrJ2HUv+flKKbCrV+3b+/eHmTPhxqqQtx0/iwWOHlWidK225VUiI6mcPpbvGg9zseGitVmr\n0TIuYRyTkiZJOTAPku+ferlq3uLQxC41NZVdu3YRFxdnndiVlJQwY8YMzp4963Qn3EEmdkKInqym\nBjZtghv/Ex0VBbNnQ3JyJy5WV6fspSsosLVpNJhGj2JXXy27K/Zjtpitb/UK7MWCjAXEBsU69xBC\n9CAe3WP3/PPP89hjj/Ev//IvmM1msrOzefXVV3nhhRec7oAQQgjXMRhg925l6dVotLX7+CjLriNH\ngsOlvu8Qpbs4eQTfNh3mysUr1ma9Vk9mUiZj4seg03bveuJC3K8cithZLBb+67/+i7/+9a8UFRXR\np08fXnzxRV555ZVum39IInbqJssJ6iVjd29YLHD6tDIHu7Eg0LBhMHUqBAbe/TrW8btNlM4wcjjb\nEs0cqDxqre8K0CekD/Mz5hPpH+maBxJdIt8/9fJoxE6j0fDKK6/wyiuvOH1DIYQQrnX5spK+pKjI\nvr13b2XZNT7+7tcozsujYOtWTpw+jXnDBlIMBhKDO1SDiIigKHMo3zUdobay1trsrfNmevJ0hscN\n77a/6AvRkzgUsdu+fTtJSUkkJydTUVHBr371K3Q6Hb/73e/o1auXJ/rZaRqNhtdff11Kigkh7lvN\nzZCVBYcOKQmH2wUEwLRpMHQoODLXKs7LI3/lSqZaLHDuHFRXs81oJHXoUBKjomgd8SAbE1o5VnXK\n7nOp4anMS59HiG+Iax9MiB6kvaTYihUrPHd4ol+/fmzevJk+ffqwePFiNBoNvr6+XL16ldWrVzvd\nCXeQpVghxP2qY9WIa7YqXWi1yh66zEzw9XX8etvffpspOTlKomGTydYeHU38qy+w+toxGtsare1+\nej9mp83mgegHJEonhIt4dCn24sWL9OnTB4PBwKZNm6z57GJj5cSTcA/ZJ6JeMnbudWPViHbJycqy\na1RUJy9YWYl2925lPRfIqq0lMzQUQ1wMOXFe7Krdbffjg6IHMTt1NgHeAU48hXAX+f4JhyZ2wcHB\nXLp0idzcXAYOHEhQUBCtra0YDAZ3908IIQTQ2AhbttxcNSIkRMlH17+/Y8uuVgYD7NoFe/dibmjg\nWvM1ahpqqGhrZXekF9faSigzhhN0/ceDvIN4OP1h+kX2c9UjCSHcwKGJ3csvv8zIkSNpbW3lrbfe\nAmDv3r3079/frZ0TPZf8xqleMnauZTLBgQOwc6d9xhG9HsaPh3HjwMurkxctLFQqR1RXAxAWEcY/\nzp9kWFII2iAtBeYr7Kgwoh+dTBDwUOxDTE+Zjq++E+u74p6Q759waI8dQF5eHjqdjtRUpXbguXPn\naG1t5YEHHnBrB7tK9tgJIdSus1Uj7qqpSclafOKEXfOWinN82beWy6fz8TaaadVpqB0QSowmkd+/\n8Hv6hvV17kGEEHfl0T12ABkZGXav09PTnb65ELcj+0TUS8bOeXeqGjFrFqSkdPKCFgscP64kuWtu\ntjX7+HBhZBrvHDxOTa9r0CuO2rO1hPYLJSk4gaHNQ2VSpzLy/RO3ndj169fPWi4sISHhlj+j0Wgo\nKSlxT89cYPny5ZLuRAihGi6tGtHu6lVl2fWGJHc1afGs62skv+0UraY2a7uvzpcHYx8k2CcYv0q/\nLj+LEMIx7elOXOW2S7G7d+9mwoQJ1pveTnedNMlSrBBCLVxVNcKO0Qh79yoHJDqkMGkN8mfPAyHs\n8b5krRxx9eJVcvNySRueRmxgLBqNhtbzrTwz+RkyUjNudwchhAu5at7i8B47tZGJnRBCDSorlfQl\nFy7Yt3emasRNiothzRq7zXlmDZxNDWNt70auYctooNVoGdl7JLHGWPae2EubuQ1vrTdTH5wqkzoh\nPMjtE7tly5bd9ibt7RqNhjfeeMPpTriDTOzUTfaJqJeMnWNcVTXipotu2QJHj9o1Xw7RszZDQ6m/\nfYqq1PBUZqXOsqvvKuOnbjJ+6uX2wxOlpaV3zCjePrETQgjhOLNZOcewdatrqkYAylruqVPKEdqm\nJmtzo8bI7lQvDsYasGht/72O8ItgVuos0iLSnHsYIUS3I0uxQgjhIberGtG3r7LsGh3dhYvW1MC6\ndUpulOsMJgO5kWY2pmlo8fe2tvvqfZmUOImRvUei03b2FIYQwp3cHrErLCx06ALJyclOd0IIIe5n\nLq8aAcqBiOxsJXPx9SpAFouFMurZ1F9HWZzttIUGDQ/FPcTkpMlSCkyI+9xtI3ZarfbuH9ZoMHU4\nbdWdSMRO3WSfiHrJ2Nm4pWoEQFmZcjjien1XgJqWWvbFGjnaPxSTt+139qTQJGalzqJXYC+HlCzL\n+AAAIABJREFULi3jp24yfurl9oidueNuXiGEEJ1SUKAsu7qsagQos8Nt25QTF9f/Amg2NJOrq2Ln\nQ8E0RNkOQYT6hjIjZQb9I/vLfmghepD7eo/d66+/LgmKhRAe5fKqEe3OnIH166GhAQCj2Uhx00V2\np+gp7hdrPRzhpfViQuIExsSPwUvXlXCgEMKT2hMUr1ixwr3pTmbOnMmmTZsArImKb/qwRsOuXbuc\n7oQ7yFKsEMKTDAbYs0fJCeyyqhGgZCzesME6U7RYLFxuusyRgAZOjkigJdB2hHZIzBCmJk8l2CfY\nuYcRQnic25dif/jDH1r//Nxzz922E0K4g+wTUa+eNnZ3qhoxdKiSk67TVSNAyYty6JCy9NqmlPyq\nb63ndHMpR4dEUZmUaj1x0TuoN7PTZhMf3JVsxvZ62vjdb2T8xG0ndj/4wQ+sf37mmWc80RchhFAV\nt1SNALh0STkcUV4OQKuxlcKaQo711lKYmY7RR1liDfIOYlryNAbHDJZftIUQQCf22O3atYtjx47R\ndD35ZXuC4ldffdWtHewqWYoVQriLW6pGgBKZy8qC/fvBbMZkNlFWX8YZzVXOjE6lLiYEAL1Wz5j4\nMUxInIC3zvvO1xRCqILbl2I7evnll/nyyy+ZMGECfn5+Tt9UCCHUyC1VI9qdP68kGq6txWKxcOXa\nFfLri8jrH03pA8Mw65QUVAOiBjA9eTphfmFOP48Q4v7jUMQuLCyM3Nxc4uLiPNEnl5CInbrJPhH1\nul/HrqxMOZTq0qoRoGQv3rhRKQkGNLY1cr7qPEVhGs6NSac5xB+AmIAYZqfNJik0qesP4YD7dfx6\nChk/9fJoxC4hIQFvbwn3CyF6nsZGJUJ3/Lh9u1NVI0A5dXH0qFKSoqWFNlMbF2ouUGq4Sv6IFC6l\nxIBGg7+XP1P6TuHB2AfRau6eOF4I0bM5FLE7dOgQv/3tb3nyySeJiYmxe2/ixIlu65wzJGInhHDG\nnapGjBunVI7oUtUIgCtXlMMRJSWYLWbK68spqi2iPDmSguEpGPy80Wq0jOo9iklJk/DVd3V9Vwih\nFh6N2B05coT169eze/fum/bYlZaWOt0JIYToTu5UNWLGDAjr6vY2oxF27YK9e7EYjVQ3V5NfnU+1\nH5ybPoiaOOXCaeFpzEydSaR/5F0uKIQQ9hyK2EVERPD5558zffp0T/TJJSRip26yT0S91Dx2bqsa\nAUpOlLVroaqKprYmCmoKqGqpoWRQAsVDEjHrdUT6RzIzZSZpEWlOPYcz1Dx+QsZPzTwasQsICGDS\npElO30wIIbojt1WNAOX47ObNcPw4BpOB4rpiyuvLqY0K4tyM4TSFBeCr92VS4iRG9h6JTtvVGwkh\nhIMRu5UrV3Lw4EGWLVt20x47rbZ7buaViJ0Q4m7cVjWi/eInTsCmTViamqhorOBCzQWa9RYKH0rm\nYnosGo2Wh+IeYnLSZAK8A5x+HiGEerlq3uLQxO52kzeNRoPJZHK6E+6g0Wh4/fXXyczMlLC0EOIm\nbqsaAVBVpeSkKyykprmG/Op8mgxNVCZFkT8ylTZ/H5JCk5iVOotegb2ceg4hhLplZWWRlZXFihUr\nPDexKyoquu17SUlJTnfCHSRip26yT0S9uvvY3alqxNSpMGxYF9OXgHKUdu9e2LWL5uYGCmoKuHrt\nKi0BPpwfnU5VQgShvqHMSJlB/8j+3bIMWHcfP3FnMn7q5dE9dt118iaEEI5ya9UIgJISWLsW46WL\nlNSVUFpXilljoWxAPEXD+qLz9WNKn/GMiR+Dl66reVKEEOLOHK4VqzYSsRNCtHNb1QiAlhbYuhXL\noUNcbrpMYU0hbaY2GiKCyBubTmNEEENihjA1eSrBPsFOPYcQ4v7l0T12aiQTOyGE26pGgO3kxYYN\n1F0tJ786n4a2Bkx6HRce7Et5v97EhcQzO2028cHObNgTQvQEHl2KFcLTZJ+IenWHsXNr1QiA2lpY\nt47WM6corCnkctNlAK4mRHB+VBre4VE8kjyNwTGDu+U+ujvpDuMnuk7GT8jETghxX3Fb1QhQNurt\n349p21ZKqwopqVNKgrX6e5M/Mo2avr0Y22cc4/uMx1sn9bWFEJ7n0FJsYWEhv/71rzl+/DiNjY22\nD2s0lJSUuLWDXSVLsUL0LLerGhEZqeyjc6pqBMDFi1hWr+ZKwQkKawppMbZg0cDF9DguPJRMeu/B\nTE+eTpifMzNHIURP5dGl2CeffJLU1FT++Mc/3lQrVggh7iW3Vo0AZS13xw4adm0lv+o8da1KJuOm\n0ADyxqbj1zedp9JmkxSa5MxjCCGESzgUsQsODqampgadU/919CyJ2Kmb7BNRL0+NnVurRrTLy6N1\n9XcUFR+norECALNOS9HQJKqGpDMldTrDYoeh1XTPCjxdId89dZPxUy+PRuwmTpzIsWPHGD58uNM3\nFEIIZ7m1agRAfT3mdesoP7iVotoiTBalwk51XBj5YzIY0n8yTyZNwlfvTOI7IYRwPYcidi+99BJf\nfPEFjz76qF2tWI1GwxtvvOHWDnaVROyEuP+0tMCOHW6qGgFgNmM5dIira7+ksOI0zcZmANp8vSgY\nkUrw8HHMTJtFpH+kcw8ihBA38GjErqmpiblz52IwGCgrKwPAYrGo7hi/EEKdLBY4dsyNVSMALl+m\n4etPuXBqN9XN1dbmitRe1E0cybSB80iLSHPyJkII4V6SoFh0S7JPRL1cPXZurRoBYDDQum0zJRs+\n52JdGRaU/25cC/GnePwDDBuzkBFxI9Bp1bPH2Bny3VM3GT/1cnvErqioyFojtrCw8LYXSE5OdroT\nQghxI7dWjbjOnH+esn+8S2nxCQxmg9Km1VAyOJGoGQv5Yeo0ArwDnLuJEEJ40G0jdkFBQTQ0NACg\n1d76xJdGo8FkMrmvd06QiJ0Q6uT2qhEATU1c/vpDSvaso8nQZG2ujQmhedY0Jo/4Pr0Cezl5EyGE\ncFyPrhX7q1/9iuzsbJKSkvj73/+OXn9z4FEmdkKoj1urRgBYLNTv38WFL/9KTY1tbdfgo6dy7BCG\nzV5K/6gBsn9YCOFxrpq3qC75Uk5ODhcvXmTXrl3069ePr7/++l53SbhBVlbWve6C6KKujF1NDXz+\nOXz8sf2kLjISliyBxx93flLXeqmc3N//kmN/XW43qatKicP/lV/w+JJ/Y0D0wB4/qZPvnrrJ+AnV\n1YrNzs5m5syZAMyaNYsPPviAJ5544h73SgjRFW6vGgFYDAYK1nzIpQ1fYTTY1nabA33RzZ3PjMlL\nCPIJcu4mQgjRTahuYldTU0NsbCygVMSorq6+yyeEGsmpLvVyZOw8UjUCuHgymwsf/xeGygrbvTUa\nmkcOY/Cil4mP6Ov8Te4z8t1TNxk/cc+WYt9++22GDx+Or68vzz77rN171dXVLFy4kMDAQJKSkvjs\ns8+s74WGhlJfXw9AXV0d4eHhHu23EMI5lZXw0Ufw1Vf2k7q4OHj+eXjkEecndfW1l8n+7//HuT/8\nP7tJXVuvKKL/72vMfvEPMqkTQtyXOj2xM5vNdv90Ve/evVm2bBlLly696b2XXnoJX19fKisr+cc/\n/sE//dM/cfr0aQDGjh3L1q1bAdi0aRPjx4/vch9E9yX7RNTrdmPX0qIcjHj3XftSYAEBMH8+/OhH\nzpcCMxjbOLzxAw79v6dpPZRtbTd7exHwyCIy3/yYgYOn9vh9dHci3z11k/ETDi3FHjlyhJ/+9Kfk\n5OTQ0tJibXcm3cnChQsBOHz4sLWaBShVLr755htyc3Px9/dn3LhxLFiwgI8//pjf/e53DBkyhJiY\nGCZOnEhiYiK//OUvu3R/IYRneKJqhMVi4ez5bIo/+W98S8rpuC3PZ9BQ+i35v4TFJDp3EyGEUAGH\nJnZPP/008+fP529/+xv+/v4u7cCNR3vPnTuHXq8nNTXV2jZkyBC730L+/d//3aFrP/PMM9Yky6Gh\noQwdOtS6/6D9evK6e75ub+su/ZHXjr/OzMy0vk5NzWT9eti3T3mdlKT8fEtLFqNGwaxZzt+voraM\nlb/9Jd65Z3goKgSA45dq0QeGsPCXv6H3yKnKz5+50C3+/+nurzuOX3foj7yW8btfX7f/uaioCFdy\nKI9dcHAwdXV1blm+WLZsGWVlZXzwwQcA7N69m0WLFlFRYdsX89577/Hpp5+yY8cOh68reeyE8Ky8\nvGK2bi3AYNBiMpnx8Unh6lX7KJkrq0Y0tTWxN/sLDN99Q0BNo7Vdr/MiZvI8Uh77EVpfP+duIoQQ\nHuL2kmIdLVy4kE2bNjFr1iynb3ijGx8iMDDQejiiXV1dHUFBko6gJ8nqEK0T3V9eXjErV+bj5TWV\nw4ezaG2dQmvrNoYOhcjIRJdWjTCZTRws3E3pNyuJOlOE9/X/hGjQEJn8AClPv4JvYorzD9VDyXdP\n3WT8hEMTu+bmZhYuXMiECROIiYmxtms0Gj766COnOnBjFDA9PR2j0Uh+fr51OTYnJ4dBgwZ1+trL\nly+3hqaFEO6jROqmcuIEXLwIoaGg10/lwoXtjB+fyMyZzicYtlgsnK86x8GtHxG16xDR19qs74UG\nRZO08BlCM2cpm/eEEEIlsrKy7JZnneXQUuzy5ctv/WGNhtdff71LNzaZTBgMBlasWEF5eTnvvfce\ner0enU7H4sWL0Wg0vP/++xw9epS5c+eSnZ1N//79Hb6+LMUK4RlmM7zyShanTmXS8Svn7w/DhmXx\nr/+a6fQ9rjRdYduxb9Bv2kxkaZXtHl7+9HloKr2eeE6ZTQohhEp5dCn2dhM7Z7z55pu88cYb1tef\nfPIJy5cv5ze/+Q3vvPMOS5cuJTo6msjISN59991OTeqEEJ5RXQ3ffgsFBWbrpE6rhb59oXdv6NWr\n6ymRAJoNzWQVbufitu9IOlqIzqicwtdr9STE9Sd+0fPoHhjs/IY9IYS4TzgUsQPYsWMHH330EeXl\n5cTHx/PUU08xZcoUd/evyyRip26yT6R7s1jgyBHYtEkpC3b1ajHHj+cTHj4Vf/8s+vXLpLV1G888\nk0pGRufTjJgtZo5cPMLBw6vosyuHoKoG63txQXH0mbwQ39lzwU8OR7iafPfUTcZPvTwasXv//fd5\n9dVXef755xk1ahQlJSU8+eSTvPHGG/z4xz92uhPuInvshHC9hgZYvRrOn7e1RUcn8uKLUFu7nbNn\nTxAdbWbq1K5N6gprCtl8Zi0Bew8x4HQZmuv/oQv1DSU5fRTBj/0A+vRx1eMIIcQ9dU/22KWlpfH1\n118zZMgQa9uJEyd49NFHyc/Pd1lnXEkidkK4Xm4urF0Lzc22tqgoePRRuF7Cucuqm6vZXLCZyuN7\nSd9/Ht9GJRm6r96X5KgMomY9imb8eNDp7nIlIYRQH1fNWxya2EVERFBRUYG3t7e1rbW1lbi4OKqq\nqu7wyXtHJnZCuE5zM6xfDydP2to0Ghg9GqZOBb1Dsf9bazW2srtkN4fzskg+kEd00RUAdBodfUL6\nED9kArr5CyAiwsmnEEKI7stV8xaH8gKMGzeOf/7nf6apqQmAxsZGfv7znzN27FinOyDErbgyLC2c\nU1AAf/mL/aQuJASeflpJNnzjpM7RsbNYLBy/dJw/H/gvCrd8yfBv91sndTEBMYxMyyTxmVfQPfOs\nTOo8SL576ibjJxz6Pfvdd9/liSeeICQkhPDwcKqrqxk7diyfffaZu/vnFNljJ0TXGQywZQscPGjf\nPnQozJrlXH3X0rpSNuRvoK7kPOnZ5wiprAMg2CeY1PBUgkdOUGaNLi5hKIQQ3c092WPXrrS0lIsX\nLxIXF0dCQoLLOuEOshQrRNeVlSlpTDrutAgIgHnzoF+/rl+3vrWeLQVbyK3IITGnmIRTJWjNFrx1\n3qSEpRCd0A/NvHmQnOz8QwghhIq4fY+dxWKxVoUwm2+fi0rbTbO8y8ROiM4zmWDnTti9G7tkwxkZ\nMH++MrnrCoPJwL7Sfewp2UNAeSUZ+87h19CMVqMlITiBhLAk9BMnwYQJztccE0IIFXJ7upPg4GAa\nGpTcUfrb7IzWaDSYTCanOyHEjSQXk+dduQLffAMVFbY2Hx9l2XXoUMdzAHccO4vFQu6VXLYUbOFa\n3VVSDuXTq+AyAFH+UaSEp+DbN00JBUZHu/iJRFfId0/dZPzEbSd2ubm51j8XFhZ6pDNCCM+zWGD/\nfti2DYxGW3tSEjzySNcrdVU0VLAhfwMltcX0KrjMoEMFeLUaCPQOJDU8ldDQXjBtGjz0kFSOEEII\nF3Foj91//Md/8POf//ym9j/+8Y/88z//s1s65qz2OrZyeEKI26urg+++gwsXbG16vZLCZPTozs23\n8vLz2HpkK42GRgprCvGO8CY+wI/07HOEXarFS+tF37C+xAbGohk0SAkFBgW5/qGEEEJF2g9PrFix\nwnN57IKCgqzLsh2FhYVRU1PjdCfcQfbYCXF7Fgvk5MCGDdDaamvv1UtJNtzZVdG8/DxW7ljJlZgr\nXKi5gMVkpH9WNbOaAwgO8KN3cG+SQpPQh0fCww9DWpprH0gIIVTOIyXFtm/fjsViwWQysX37drv3\nCgoKCA4OdroDQtyK7BNxn6YmpXrEmTO2No1GObcwaVLXCjtsOLSBM03HMH9ZgFdhPcONZiKCvDDp\ndYxIn4i/dwCMGQOZmdAh0bnofuS7p24yfuKOE7ulS5ei0WhobW3lueees7ZrNBpiYmL485//7PYO\nCiFcJy9PqfN6Pdc4AOHhsHAhdDWDUWVTJduPrKfXydM8XtvGhYprjAzwJrvJC+ID8U9MVQ5HOFtz\nTAghxF05tBS7ZMkSPv74Y0/0x2VkKVYIm9ZW2LgRjh2zbx8xAqZP73oQ7eTlk6w99Q0XX/uAZ+pq\n0ZnBW+eNn94Pi07Lt71T+NmqzdBN0yIJIUR34ZGl2HZqm9QJIWyKi5Vkw7W1tragIFiwAFJTu3ZN\nk9nEpvMbKN25hgePXyCkyYy2yYJ/cADeOm+uBftR6R9C/OARMqkTQggPcmhiV1dXx/Lly9m5cydV\nVVXWhMUajYaSkhK3dtAZUlJMvWSfiPOMRti+HbKz7ZMNDxqknF/w8+vadetb69m48W38s/aSUaOs\n6foEBpLsHUGN2cAWvZahMQkMiu2LuXf3rlAjbibfPXWT8VMfV5cUc2hi99JLL1FaWspvfvMb67Ls\n73//e773ve+5rCPusHz58nvdBSHuiUuXlGTDlZW2Nl9fmDtXmdh1VXHeQU598keiSy9Z2yL9I0kY\nN5zDVdVMTUggrbiYEUlJbGttJXXqVCeeQggh7n/tAagVK1a45HoO7bGLiorizJkzREZGEhISQl1d\nHeXl5cybN4+jR4+6pCOuJnvsRE9kNsPevZCVpZQHa5eSoiy9dvUgu6W+ntNfvM2VfVus4T8NGpJi\n0ukzezGasWMpvnCBgm3b0La1Yfb2JmXqVBIzMpx/KCGE6AHcXiu2o8jISCoqKvDy8iI+Pp5Tp04R\nHBxMSEjILfPbdQcysRM9TXW1speutNTW5uUFM2bA8OFdLO7Q1kbbrh2cWfV3ahou266r9yZ16iJi\n5j0BgYHOd14IIXo4V81bHNrVPHjwYHbt2gXA+PHjeemll3jxxRfJkN/GhZu4cr/B/c5igcOH4S9/\nsZ/UxcfDiy8qJ187Pakzm+HIEep//68c+/wtu0kdaekMef1dYhY/f8tJnYydusn4qZuMn3Boj917\n771n/fN//ud/8uqrr1JXV8dHH33kto4JIe6uoQFWrYL8fFubVqvkAR4/vgsHUi0W5WJbtnCp8ATn\nqs5htiiHpRrDAwme+xjjMpeg03Yhi7EQQgi3c2gp9sCBA4waNeqm9oMHDzJy5Ei3dMxZshQr7ne5\nuUoFieZmW1tUlFISrEu5gC9dgs2bMRfkc77qPBWNFQC0+vtQOjyNsQ+/yIAYJ05eCCGEuC2P5rGb\nNm3aLffSzZo1i+rqaqc74S6S7kTcj5qbYf16OHnS1qbRwOjRMHUq6B36VndQX6/kRcnJocXQTG5l\nLg1tDZj0Okoe6EPLiGF8f+iTRPpHuvQ5hBBCuD7dyR0jdmazGYvFQmhoKHV1dXbvFRQUMG7cOCo7\n5lPoRiRip26Si+nWCgqUpdf6eltbaCg88ggkJXXyYq2tyhHa7GwwGKhurub0ldMYLCYq0mMpGppE\neuKDzM+Yj7fO8dIUMnbqJuOnbjJ+6uWRiJ2+w6/++hvCAFqtll//+tdOd0AIcXdtbbB1Kxw8aN8+\ndCjMng0+Pp24mNkMR4/Cjh3Q1ITFYqG4rpii2iKq4iMoGJ5MS1gQM1NmMrL3SDRdOk4rhBDiXrhj\nxK6oqAiAiRMnsnv3butMUqPREBUVhb+/v0c62RUSsRP3i7IyJY1JVZWtLSAA5s2Dfv06cSGLBc6f\nh82b4epVAAwmA2eunqHYr42CESnUxoYR7BPM9wd8n4QQqRohhBCe4tE8dmokEzuhdiYT7NwJu3fb\nlwTr10+Z1AUEdOJiFRXKhO7CBWtTQ2sDR68VcHpwLJeTo0GjoW9oXx4b8BgB3p25uBBCCGd5dGK3\nZMmSW3YA6LYpT2Rip249fZ/IlStKSbCKClubj4+y7DpkSCfy0tXVWQ9GtLNYLFxsq2JLXAsl/WMx\n65XUJRP6TGBy38loNZ3NkWKvp4+d2sn4qZuMn3p59FRsSkqK3Q0vXbrE//7v//KDH/zA6Q4IIWws\nFti/H7ZtA6PR1p6UpByQCA118EItLbBnj3KxDhcyYeFgnJntfYIx+CmnXH31vjzS7xH6RXZmXVcI\nIUR31OWl2MOHD7N8+XLWrl3r6j65hETshNrU1sJ338H1ra2Akrpk6lQllYlDUTqTCY4cUYrFXrtm\n91Zjcjxfx9VSpG+0tsUExPD4oMcJ9wt3yTMIIYTomnu+x85oNBIWFia1YoVwksWirJRu2KBkIGnX\nq5eSbDg62sGL5OXBli32pywA4uIoGJ7CV9cO0WJssTYPiRnC3PS5eOm8XPMgQgghusyjS7Hbtm2z\nS3nQ1NTE559/zsCBA53ugBC30lP2iTQ1KdUjzpyxtWk0MGECTJoEOkcqd5WXKwcjiovt20NCME+d\nwo6AK+wu3W1t1ml0zEmbw4OxD7ollUlPGbv7lYyfusn4CYcmds8995zdXwABAQEMHTqUzz77zG0d\ncwWpPCG6s7w8WL1amdy1Cw9XonTx8Q5coLZW2YzXsQQFgK8vTJhA07BBfH3uOy6U2k7ChviEsGjg\nInoH93bNQwghhHCKRytPqJksxYruqrUVNm6EY8fs20eMgOnTwftuRR5aWpQcKAcO2J+w0GqVi0ya\nRKmhiq9Of0V9q61ERWp4Ko/2fxR/r+6bf1IIIXoqjy7FAtTW1rJu3TouXrxIXFwcc+bMISwszOkO\nCNGTFBcryYZra21tQUGwYAGkpt7lwyYTHD6sJLe74WAE/fvDtGlYwsM5WH6QTQWbMFvMAGjQMClp\nEhMTJzqdykQIIUT35lDEbvv27Tz66KNkZGSQmJhIcXExZ8+e5X//93+ZNm2aJ/rZaRKxU7f7bZ+I\n0aikk8vOtk82PGgQPPww+Pnd4cMWC5w9q9QUu/FgRO/eMHMm9OlDm6mNNXlrOFlpW5r10/vxaP9H\nSYtIc+0D3cH9NnY9jYyfusn4qZdHI3YvvfQS//M//8OiRYusbV999RU//elPOXv2rNOdEOJ+dumS\nkmy4stLW5uenTOgGDbrLh8vKlIMRJSX27aGhMG0aDBwIGg1Xr13ly9wvqWyy3SQuKI5FAxcR6uto\n8jshhBBq51DELjQ0lKqqKnQdjugZDAaioqKo7bim1I1IxE7ca2Yz7N2rpJQzmWztqakwfz4EB9/h\nwzU1ysGIU6fs2319YeJEGDlSSXIHnL5ymu/Ofkebqc36Yw/FPsTstNnotQ7vthBCCHEPeTRit2TJ\nEt5++21eeeUVa9tf/vKXW5YaE0JAdbWyl6601Nbm5QUzZsDw4XdINtzcbDsY0XE2qNMpByMmTgR/\n5fCDyWxi24Vt7CvdZ/0xvVbP3PS5DO011A1PJYQQortzKGI3btw4Dh48SHR0NL1796a8vJzKykpG\njRplTYOi0WjYtWuX2zvsKInYqZta94lYLErhh02bwGCwtcfHw8KFEBFxmw+aTHDokHIwornZ/r0B\nA5Rl13BbdYiG1ga+Pv01xXW23HVhvmE8PuhxegX2cuETdZ5ax04oZPzUTcZPvTwasfvRj37Ej370\no7t2SIierKEBVq2C/Hxbm1YLmZkwfrzy55tYLEp24q1blTBfR/HxysGIhAS75uLaYr46/RWNbbbS\nYBkRGSzsvxBfva/rHkgIIYTqSB47IVzg1ClYt84+2BYVpSQbjo29zYdKS5WDER3XawHCwpQI3YAB\ndmu2FouF7LJsthZutUtlMqXvFMb3GS+/XAkhhIp5PI/drl27OHbsGE3X0+RbLBY0Gg2vvvqq050Q\nQq2am2H9evviDxoNjB4NU6dazzfYq65WInSnT9u3+/kpe+hGjLjpg63GVlblreL0Fdtn/L38eWzA\nYySHJbvwiYQQQqiZQxO7l19+mS+//JIJEybgd8eEW0K4hhr2iRQUwHffKUuw7UJD4ZFHICnpFh9o\nboZdu+DgwZsPRowapRSIvcX3q7Kpki9OfUFVsy2HXXxwPIsGLiLY505Ha+8NNYyduD0ZP3WT8RMO\nTew++eQTcnNziYuLc3d/hOj22tqUgNvBg/btQ4fC7Nng43PDB4xG5Yd37VLKgXU0aJAS2rtNFZeT\nl0+yOm81BrPtJMbI3iOZmTITnVZ3y88IIYTouRzaYzd48GC2b99OZGSkJ/rkEhqNhtdff53MzEz5\n7UW4TFmZksakYwGIgACYNw/69bvhhy0WyM1V8tHV1Ni/16ePkvskPv6W9zGZTWwq2MTBctvs0Uvr\nxfyM+TwQ84CLnkYIIcS9lpWVRVZWFitWrHDJHjuHJnaHDh3it7/9LU8++SQxMTF2702cfxizAAAg\nAElEQVScONHpTriDHJ4QrmQyKZlIdu+2LwnWr58yqQsIuOEDJSXKwYiyMvv28HCYPl354G0OO9S3\n1vNl7peU1ds+G+EXweODHic6INpFTySEEKI78ejhiSNHjrB+/Xp279590x670htP9AnhAt1pn8iV\nK0pJsIoKW5uPj7LsOmTIDfOzqiplnfbMGfuL+PkpeU+GD1f21N1GYU0hX5/+mmuGa9a2AVEDWJCx\nAB/9jWu83VN3GjvReTJ+6ibjJxya2P36179m7dq1TJ8+3d39EaLbsFhg/35lJdVotLUnJSkHJEI7\nlmC9dk0J6R06pNQSa6fTKUdkJ0xQyoHd9l4W9pTsYfuF7VhQfmPTarRMS57GmPgxkspECCGEQxxa\niu3Tpw/5+fl4e3t7ok8uIUuxwhm1tcqJ16IiW5ter5xzGD26Q5TOaFTKf+3effPBiAceUD5gNwO8\nWYuxhW/PfEteVZ61LdA7kO8P+D6JoYmueSAhhBDdmqvmLQ5N7FauXMnBgwdZtmzZTXvstLdMp3/v\nycROdIXFAjk5sGEDtLba2mNjlZJg0dEdfvDUKSWcV1trf5HEROVgRO/ed73fpcZLfHHqC2pabIcr\nEkMSeWzAYwT5BLngiYQQQqiBRyd2t5u8aTQaTB3zcXUjMrFTt3uxT6SpCdasgbNnbW0ajbKKOmlS\nh61xxcVKMdiLF+0vEBGhHIzIyLjtwYiOjl86ztpzazGabeu8YxPGMrXvVFWnMpE9Puom46duMn7q\n5dHDE4WFhU7fSIjuLC8PVq9WJnftwsOVkmDWjCRXryoHIzrO/AD8/ZWDEQ89dMeDEe2MZiMbzm/g\nSMURa5u3zptH+j3CgKgBzj+MEEKIHqtTtWLNZjOXL18mJiam2y7BtpOInXBEayts3AjHjtm3jxih\nBN+8vVFmezt3wuHD9gcj9Hplw9348Xc8GNFRbUstX+Z+ycUGW7Qvyj+Kxwc9TqS/evJECiGEcC2P\nRuzq6+v56U9/yueff47RaESv1/PEE0/w5z//mZCQEKc7IcS9UFysJBvuuEUuKAgWLIDUVMBggD3X\nD0Z03HAHMHgwTJly14MRHZ2vOs83Z76h2dhsbXsg+gHmZczDW6eeg0lCCCG6L4fCbi+//DJNTU2c\nOnWKa9euWf/35Zdfdnf/RA+VlZXltmsbjUru4JUr7Sd1gwbBT34CqSkWOHEC3n5bWXrtOKlLSoIf\n/1hZo3VwUmexWMgqyuLTk59aJ3VajZY5aXN4tP+j992kzp1jJ9xPxk/dZPyEQxG7jRs3UlhYSMD1\n9Prp6emsXLmS5ORkt3ZOCFerqFCidJWVtjY/P3j4YWVix4ULyqyvYzZigMhIZW02Pd2hgxHtrhmu\n8c2Zb8ivzre2BfsE8/0B3ychJMHJpxFCCCHsObTHLikpiaysLJKSkqxtRUVFTJw4kZKSEnf2r8tk\nj53oyGyGvXshK0spD9YuNRXmz4fg1itKdC4vz/6DAQG2gxGd3FdaXl/Ol7lfUtdaZ23rG9qXxwY8\nRoD3jTXIhBBC9GQe3WP3/PPPM336dH72s5+RmJhIUVERf/rTn/jRj37kdAeEcLfqaiVK17H6nZeX\nkmpueL9GNDuz4OjRmw9GjBmjHIzw6VwpL4vFwtGKo6w/vx6TxTaLnNBnApP7Tkar6d4Hj4QQQqiX\nQxE7s9nMypUr+cc//kFFRQVxcXEsXryYpUuXdttSRxKxUzdX5GKyWODIESXlnMFga4+Ph4VzDUSc\ny4Y9e6CtzfamRqMUgJ08GbpwMMhgMrDu/DqOXzpubfPV+7Kw30IyIjOceRzVkDxa6ibjp24yfurl\n0YidVqtl6dKlLF261OkbOqu+vp5p06Zx5swZDhw4wIABkvdL3KyhAVatgnzb1ja0WsicZGF8UA7a\nT7dDfb39h/r2VcJ4sbFdumd1czVfnPqCy02XrW29AnuxaOAiwv3Cu3RNIYQQojMciti9/PLLLF68\nmLFjx1rb9u3bx5dffslbb73l1g7eyGg0Ultbyy9+8Qt+/vOfM3DgwFv+nETseq5Tp2DdOmi2ZRUh\nKgq+/1Ah0cc3w6VL9h+IilImdKmpnToY0VHe1Ty+PfstLUZbvdihvYbycNrDeOm8unRNIYQQPYdH\nS4pFRkZSXl6OT4e9Ri0tLSQkJHDlyhWnO9EVzz77rEzshJ3mZli/Hk6etLVpNDCxXyUTW7egKzxv\n/4HAQGXJddiwTh+MaGe2mNlxYQe7S3Zb23QaHXPS5vBg7IPddquCEEKI7sVV8xaH/jbTarWYO24s\nR9l3JxMn4S6dzcVUUADvvGM/qYvya+TF3muYfPYv9pM6Ly+l+OvLL3fptGu7prYmPs752G5SF+ob\nynMPPsdDcQ/12Emd5NFSNxk/dZPxEw79jTZ+/Hhee+016+TOZDLx+uuvM2HChE7d7O2332b48OH4\n+vry7LPP2r1XXV3NwoULCQwMJCkpic8++8z63p/+9CcmT57MH/7wB7vP9NS/OIVNW5uy7Prxx8q+\nOgCdqY1pXjt5ofW/iCk7opyiACV8N2yYMqGbPLnTp107Kq0r5a9H/sqF2gvWttTwVH780I+JC4pz\n5pGEEEKILnNoKba0tJS5c+dSUVFBYmIiJSUlxMbGsmbNGhISHE+y+u2336LVatm0aRPNzc188MEH\n1vcWL14MwN/+9jeOHTvGww8/zL59+257OEKWYkVZmZLGpKpKea2xmEmqy2G2z3ai/Rrsfzg5WdlH\n16uXU/e0WCwcLD/IpoJNmC3KLzoaNExKmsTExImSykQIIUSXeHSPHShRuoMHD1JaWkpCQgKjRo1C\n28UlrGXLllFWVmad2DU1NREeHk5ubi6pqakAPP3008TFxfG73/3ups/PmTOHnJwcEhMTeeGFF3j6\n6advfjCZ2N23TCbYuVMp4do+xGHVBYxr2szgmMt4d6zQFR2tTOhSUrp8MKJdm6mNNXlrOFlpW+/1\n0/vxvQHfIzU81alrCyGE6Nk8mu4EQKfTMWbMGMaMGeP0TW/s+Llz59Dr9dZJHcCQIUNuu1dg/fr1\nDt3nmWeesVbLCA0NZejQodb8Pu3Xltfd8/Vbb711y/EaODCTb76B7OzrryP7k1GyBWPLVhrCwNs7\nSfn5y5dh2DAyn38etFqn+7Nq4yq2X9hOWP8wAIqOFxHhF8H/efr/EOobes///+pOrzt+b7tDf+S1\njF9Pei3jp57X7X8uKirClRyO2LnSjRG73bt3s2jRIio61Od87733+PTTT9mxY0eX7iERO3XLysqy\nfglAiczt3w/btoHRCN6tDfQt2kG/5mP072fB1/f6D3p5wbhxMHYs9qG7rjt95TTfnf2ONpMtkfFD\nsQ8xO202eq3Dvxv1GDeOnVAXGT91k/FTL49H7Fzpxo4HBgZSf0Oy2Lq6OoKCgjzZLdGNdPwPU20t\nfPcdFBUpByOSSveRWLaX1EQD8enXV1jbD0ZMngwu+vfGZDaxtXAr2WXZ1ja9Vs/c9LkM7TXUJfe4\nH8lfKuom46duMn7irhM7i8XChQsX6NOnD3q9a+aBN55mTU9Px2g0kp+fb12OzcnJYdCgQU7dZ/ny\n5WRmZsq/6CplsUBODmzYAG0tZmIvHafvhe2EezfSfxgEBFz/wdRUmD4dYmJcdu+G1ga+Pv01xXXF\n1rYw3zAeH/Q4vQKdO4AhhBBCtMvKyrJbnnXWXZdiLRYLAQEBNDY2dvmwRDuTyYTBYGDFihWUl5fz\n3nvvodfr0el0LF68GI1Gw/vvv8/Ro0eZO3cu2dnZ9O/fv0v3kqVYdcrLK2br1gJyck5gNA7G3y+Z\ndK2BlMItBDRVkpgIiYnXU8/FxNgORrhQcW0xX53+isa2RmtbRkQGC/svxFfve4dPCpClILWT8VM3\nGT/18thSrEajYdiwYeTl5XV5ktXuzTff5I033rC+/uSTT1i+fDm/+c1veOedd1i6dCnR0dFERkby\n7rvvOn0/oS55ecWsXJlPQ8NUjhzRkujTj+SrfyI1tJWIiFD6PwjBwShLrVOmwJAhXU4ufCsWi4Xs\nsmy2Fm61S2Uype8UxvcZL3kThRBCdHsOHZ547bXX+OSTT3jmmWdISEiwzio1Gg1Lly71RD87TSJ2\n6vOHP2xn99be6M+tpW/TaULaLhHg25e0uAZmz+6Lzs9bORgxZozLDka0azW2sipvFaevnLa2BXgF\n8L0B3yM5LNml9xJCCCFu5NHDE3v27CEpKYmdO3fe9F53ndiB7LHrVsxmaGqy/dPYaP2zqb6RghNN\nXPt0K2OvlDIdMxosaDRwgOMYfaPQjXxIORgRGOjyrlU2VfLFqS+oaq6ytsUHx7No4CKCfYJdfj8h\nhBCincf32KmVROw8wGi0m6BZ/3yrtuZmWzbhDmpq4Px5uHYN9uduYJbBD4BjllomhYTSEhDBNxnh\nvPbde255hJOXT7I6bzUGs8HaNqr3KGakzECn1bnlnvc72eOjbjJ+6ibjp14eT3dSVVXFunXruHTp\nEr/85S8pLy/HYrEQHx/vdCdEN2GxQGvrLaNqt/xza2uXb9XaCgUFUFlpawv280VjrCEgIAwdflTG\nDqHRq4a0/rEueDh7JrOJTQWbOFh+0NrmpfVifsZ8Hoh5wOX3E0IIITzBoYjdzp07+d73vsfw4cPZ\nu3cvDQ0NZGVl8Yc//IE1a9Z4op+dJhG76ywWJRzm6GTNaHRPPzQa8PPD7B9I4eUAThQEcE0bSJtX\nAAbvACwBgfg0fck8UzlF5Y0YLTp0Ogt9+4ZxYkA6U37yE5d1pa6ljq9Of0VZfZm1LcIvgscHPU50\nQLTL7iOEEEI4yqO1YocOHcp//Md/MG3aNMLCwqipqaGlpYX/z959xzV19X8A/9ywwooEJMjeskSp\nG22RIeB6XK1Wq22Bum1dbZ/2p0WgVqvtU2nF1lnFhbb6wmr7VFSmYgeKo25FBRTFgYMNgZzfH3mI\nXtkYCMHv+/XipTn33pNzc5JwON8zbGxscO/ZLpd2pEM37Kqr6x2vViscWloqH9/WGgQC+WJyBgby\nfxv6v74+snMF+P13fi8dAHh6ylcteXj7MrJiYxGgo6M4llRRAaeQENi6uCilyNcfXceeC3tQKi1V\npLmbumOUyyjoaOo0cCUhhBDSeto0FJuTk4PBgwfz0rS0tFBdXf3CBWhNajV5QipteIza8+PVWouW\nVv0NtOfThML/bfvQsKIi4PAvwD//8NNNTYFhwwB7e/ljQxcXICQEyUlJ+OfCBXR3d4dTQIBSGnWM\nMaTnpiP5RjIY5B8cASdAoEMg+lv1p6VMlIjG+Kg3qj/1RvWnfpQ9eaJJDTs3NzckJCRgyJAhirSk\npCR4erbvsUiRkZGqe3LGgPLypodAKysbz7OlhMKm9aoZGCh1GRGZDMjIAFJS+MPxtLUBX1+gXz9A\n47n5CbYuLrB1cYFAiV9OZdIy/HLpF1wuuKxIM9A2wDj3cbA1slXKcxBCCCEtUdMBFRUVpZT8mhSK\n/euvvzBixAgMGzYMu3fvxttvv41ff/0V+/btQ9++fZVSEGVrlVCsTNb4eLVn01qrR5PjAD29JodA\noaSt4JojJwf4/Xfg7l1+erdu8rCrqI1WEckvzsdP537Co/JHijTbTrZ4w/0NGOrQXsSEEELahzYd\nYwcAeXl52L59O3JycmBjY4PJkye36xmxHMchafVqOA4e3HAor6qq7vFqdTXaSkvrXLJDKTQ0mh4C\n1dVV6o4LylRcDBw+LN/j9VmdO8vDrg5tuNbv6fzT+O3Kb6iSPZ0QMsB6AALsA2gpE0IIIe1Kmzfs\nAEAmk+HBgwcwNTVt92OSOI4DmzcPSSUlcAoKgq2pad2NtfLy1iuEtnbTQ6A6Ok0ar9Ze1Rd21dKS\nh137968ddm3Ii4wTqZJV4cDVA8i8k6lI09HQwSjXUXA3dW9RnqTpaIyPeqP6U29Uf+qrTSdPPHr0\nCHPmzMHPP/8MqVQKLS0tjBs3DqtWrYKxsfELF6K1RO7dC18jI1zLy4Ntnz7KyVRXt+mNNS0t5Txn\nO5ebC/z3v7XDrh4e8rBrp05tV5bH5Y/x8/mfcbvotiLNVM8Ub3Z7E531OrddQQghhJAmUMnOE6NH\nj4ampiaWLFkCGxsb5ObmYvHixaisrMS+ffuUVhhl4jgObNAgAECqUAjf/v3rPlEgqH+82vNpenrN\n63bq4IqLgcRE4PRpfrqJiTzs6ujYtuW5WnAV8RfjUVb1dNawp8QT/3L5F7Q1lLu3LCGEEKJMbRqK\n7dSpE+7cuQM9PT1FWmlpKczNzfHkyZMXLkRr4DgObMQIQFsbyaam8B83rv7xamocAlUFmQw4cQJI\nTuZHsrW0gEGD5GHXtpyvIWMypGWn4UjOEcVSJhqcBoKdgtHHok+7HzZACCGEtGko1tXVFdnZ2XB3\nfzo+KScnB66uri9cgFbVq5digVsoaYHbl93Nm/Kwa34+P93dHQgOVl7YtanjREqlpYi/GI+sh1mK\nNJGOCOPcx8G6k7VyCkOahcb4qDeqP/VG9Uea1LDz9/dHUFAQ3nnnHVhbWyM3Nxfbt2/H22+/jU2b\nNoExBo7jEBYW1trlbZZkiURpC9y+7EpK5LNd6wq7Dh0KODm1fZnyCvPw8/mf8aTiaa+xvZE93nB/\nA/ra+m1fIEIIIUTFmhSKrWn9PxvSqmnMPSslJUW5pXsBHMchIiJCfXaeaKcaCrv6+ADe3m2/TB5j\nDJl3MnHg6gFUs6drBb5m8xr87P0g4NrnUjCEEELI82omT0RFRbX9cifqpEPvFdtGbt6ULzJ85w4/\n3c1NHnY1Mmr7Mkmrpfjtym84c/fpQnlCTSHGuI6BS2fqmSWEEKKelNVuoa4NUktJCbBvH/Djj/xG\nnbExMGkS8Oabrd+oq2vq98Oyh9h4ciOvUdfFoAum9ZpGjbp2RJnT9knbo/pTb1R/pO33miLtlkwG\nZGYCSUn8sKumpjzsOmCASnYnAwBcenAJey/uRUX109WPvbp4YbjzcGhpvBzrBRJCCCGNoVAsAQDc\nuiWf7fp82NXVFRgyRDVhV0C+lEnyjWSk56Yr0jQ4DQxzHoae5j1pKRNCCCEdQpsud0I6rtJS+SLD\nJ0/y08Vi+WzXrl1VUy4AKKkswZ4Le3Dj8Q1FmpHQCOM9xsPC0EJ1BSOEEELaqSY37C5evIjdu3fj\n7t27+P7773Hp0iVUVlaie/furVk+0kpkMnljLikJKHu6UQM0NYHXXgMGDlRN2PVy1mUkZiYi42QG\nCo0KYWlric4W8q3AnIydMNZtLPS09BrJhagSraOl3qj+1BvVH2nS5Indu3fDx8cHeXl52Lp1KwCg\nqKgICxYsaNXCkdaRlwds3Aj89hu/UefiAsyeLd89QlWNutiUWJzSPYUz3Bk8Mn+E0xdOo+B2AXzt\nfDHJcxI16gghhJAGNGmMnaurK3bt2gUvLy+IxWI8evQIUqkU5ubmePDgQVuUs9loHbvaSkvlPXQn\nTwLP1rpYLB9Hp+p1nL/c9iUytDLwqPyRIk1ToIlXq15FZFik6gpGCCGEtBKVrGNnYmKC+/fvQyAQ\n8Bp2lpaWuHfv3gsXojXQ5ImnGJM35hITa4ddX31VHnbVUuHE0lJpKVKzU7Hqp1Uot3o6HddQ2xAe\nEg90edAF8ybMU10BCSGEkFbWpuvY9ezZE9u2beOl/fTTT+jbt+8LF4C0rtu35WHXX3/lN+qcnYFZ\nswBfX9U16mRMhoy8DMT8HYOMvAwI/vd25MBBK1cLr5i/AqGmENoCbdUUkLQIraOl3qj+1BvVH2nS\nSKqYmBgEBgbixx9/RGlpKYKCgnDlyhUcOnSotctHWqisTB52zczkh12NjJ7OdlXlSiHXH11HQlYC\n7pU87fF1cHBA9o1suPVxw/3H9yHgBKi4WoEAvwDVFZQQQghRI01ex66kpAS//fYbcnJyYGNjg+HD\nh8PQ0LC1y9diL2soljHg1Cl52LW09Gm6hoY87Prqq6oNuz4se4hD1w7h0oNLvHSxUIxgp2DgEZB8\nKhmVskpoC7QR0DMALk60qwQhhJCOTVntFlqguAO5c0e+yPCtW/x0Z2d5L52xsWrKBQAVVRU4mnsU\nf978E9WsWpGuraENH1sf9LfqD00BLatICCHk5dSmCxTn5OQgKioKp06dQnFxMa8QV65ceeFCkBdT\nVgYkJwMnTtQOu9bMdlVV2JUxhjN3zyDxeiKKK4t5x7y6eCHAPgCGOrV7fmktJvVFdafeqP7UG9Uf\naVLDbty4cXBzc8OSJUsgFApbu0ykiRgDTp8GDh+uHXYdOFC+0LAqw643n9zEgawDuF10m5duJbLC\nUKehsBRZqqhkhBBCSMfUpFBsp06d8PDhQ2hoaLRFmZSio4di79wBfv8duHmTn+7kJA+7mpioplwA\nUFhRiMTrifjn7j+8dENtQwQ6BsJT4kl7vBJCCCHPaNNQ7IgRI5CWlgZ/f/8XfsK2FBkZ2eEWKC4v\nl4ddjx/nh107dZKHXV1dVRd2lVZL8eetP3E05yikMqkiXVOgiQHWA/CqzavQ1qClSwghhJAaNQsU\nK0uTeuwePHgAb29vdO3aFRKJ5OnFHIdNmzYprTDK1NF67BgDzpyRh11LSp6ma2gAAwbIw67aKmoz\nMcZw8cFFHLp2CI/LH/OOuZu6I9AhEGJdcbPypHEi6ovqTr1R/ak3qj/11aY9dmFhYdDW1oabmxuE\nQqHiySmc1jby8+Vh19xcfrqjozzs2rmzasoFAPnF+UjISkD242xeupm+GYY4DYG92F41BSOEEEJe\nQk3qsTM0NEReXh5EIlFblEkpOkKPXXk5kJICZGTww64ikTzs6uamurBrSWUJUrJTkHk7EwxPC6en\npQd/e3/0NO8JAdekjU0IIYSQl16b9th1794dBQUFatWwU2eMAf/8Axw6VDvs6u0N+PioLuxaLavG\n8dvHkZqdivKqp/u6CjgB+lr2xSDbQdDV0lVN4QghhJCXXJMadv7+/ggODkZoaCjMzMwAQBGKDQsL\na9UCvmzu3pUvMvx82NXBARg2TLVh16yHWUjISsCD0ge8dEexI4Y4DYGpvqnSnovGiagvqjv1RvWn\n3qj+SJMadkePHoWFhUWde8NSw045ysuB1FR52FUme5ouEgHBwYC7u+rCrgWlBTh47SCuFPAXozbW\nNcYQpyFwNnam8ZaEEEJIO0BbiqkYY8DZs/Kw6zObekAgkIddBw1SXdi1vKocR3KO4O9bf/O2AdPR\n0MEgu0Hoa9mXtgEjhBBClKDVx9g9O+tV9mwX0nMEAhog31J378pnu+bk8NPt7eVhV1PlRTabRcZk\nOJ1/GknXk1AifTrIjwMn3wbMIQAG2gaqKRwhhBBC6lVvw04kEqGoqEh+kmbdp3Ech+rq6jqPkfpV\nVMjDrn//zQ+7GhrKw64eHqoLu+Y+ycWBqwdwp/gOL91aZI2hzkNhYWjRJuWgcSLqi+pOvVH9qTeq\nP1Jvw+78+fOK/1+/fr1NCtPRMQacOwccPFg77Nq/vzzsqqOjmrI9KX+CxOuJOHvvLC9dpCNCoEMg\nukm60Tg6QgghpJ1r0hi7//znP/joo49qpa9cuRILFixolYK9qPY2xu7ePXnYNTubn25nJw+7PrOh\nR5uSVktx7OYxHMs9VmsbsIHWAzHQZiBtA0YIIYS0MmW1W5q8QHFNWPZZYrEYjx49euFCtIb20rCr\nqADS0oC//qoddg0KArp1U03YlTGGC/cv4NC1Q3hS8YR3zMPUA4GOgTASGrV9wQghhJCXUJssUJyc\nnAzGGKqrq5GcnMw7du3atXa/YHFkZCR8fX1VMt6AMeD8eXnY9dk2sUAA9OsH+PqqLux6p+gOErIS\nkPOEP2uji0EXDHUaClsjW9UU7Bk0TkR9Ud2pN6o/9Ub1p35SU1ORmpqqtPwabNiFhYWB4zhUVFTg\nvffeU6RzHAczMzPExMQorSCtITIyUiXPe/++POx64wY/3dYWGD5cdWHXksoSJN9Ixsk7J2ttAxZg\nH4BXzF+hbcAIIYSQNlTTARUVFaWU/JoUin377bexbds2pTxhW1FFKLaiAjhyBPjzT37Y1cBAHnb1\n9FRN2LVaVo2MvAykZqeiorpCkS7gBOhn2Q+D7AZBqCls+4IRQgghBEAbj7FTR23ZsGMMuHBBHnYt\nLHyaLhAAffvKw65CFbWbrhZcRUJWAgrKCnjpzsbOCHYKRmc9Fe5RRgghhBAAbTTGjjTuwQN52PX5\nFWFsbORh1/9trdv25Sp9gINZB3H14VVeuomuiXwbMBNn1RSsiWiciPqiulNvVH/qjeqPUMOuhSor\nn4Zdn12jWV9fHnbt3l01YdfyqnKkZafh77y/IWNP48E6GjrwtfNFX8u+0BBotH3BCCGEENLqKBTb\nTIwBFy8CCQn8sCvHycOufn6qCbvKmAyn7pxC0o0klEpLn5YLHHqa94S/vT/0tfXbvmCEEEIIaRSF\nYlXgwQPgwAHg2jV+uo2NfJHhLl1UU66cxzk4kHUA+cX5vHTbTrYY4jQE5obmqikYIYQQQtoUNeya\noLISOHoU+OOP2mHXwECgRw/VhF0flz/G4WuHcf7+eV56J51OCHIMgrupu9puA0bjRNQX1Z16o/pT\nb1R/hBp2DWAMuHRJHnZ98szmDBwH9OkD+PurJuxaWV2JY7nHcOzmMVTJqhTpWgItvGrzKgZYD4CW\nhlbbF4wQQgghKkVj7OpRUCAPu2Zl8dOtreWzXVURdmWM4dy9czh8/TAKKwp5xzwlnhjsMBidhJ3a\nvmCEEEIIeSE0xq6VSKXy2a7Ph1319ORhVy8v1YRdbxfdRkJWAnKf5PLSzQ3MMdR5KGw62bR9oQgh\nhBDSrlDD7n8aCrv27i0Pu+rqtn25iiuLkXQ9CafzT/O2AdPX0sdgh8Ho0aVHh9wGjMaJqC+qO/VG\n9afeqP4INexQf9jVykoedjVXwaTSKlkV/r71N47kHOFtA6bBaaCfVT8Msh0EHdQ4tIAAACAASURB\nVE2dti8YIYQQQtqtl3qMnVQqn+167FjtsOvgwcArr7R92JUxhisFV3Dw2kE8LHvIO9bVpCuCHYNh\nomfStoUihBBCSKuiMXYvgDHg8mV52PXx46fpqg673i+5j4SsBFx7xF8or7NeZwxxGgInY6e2LxQh\nhBBC1IbaDc7KyMjAgAEDMGjQILz11luoqqpq/KJnPHwIxMUBu3bxG3WWlsDUqfLQa1s36sqkZUjI\nSsCaE2t4jTqhphBDnIZgZu+ZL12jLjU1VdVFIC1EdafeqP7UG9UfUbseOxsbG6SkpEBHRwcLFy7E\nvn378Prrrzd6nVQKpKfLw67PtgX19ICAAKBnz7YPu8qYDJm3M5GSnVJrG7BeFr3gZ+dH24ARQggh\npMnUeoxdREQEXnnlFYwePbrWsWdj1TVh10ePnj0ub8wFBMgbd23txqMbSMhKwN2Su7x0OyM7DHEa\ngi4GKtqfjBBCCCFtTllj7NS2YZeTk4OJEyfi6NGj0NDQqHWc4zisWJEEgcARxcW2vGMWFvKQq6Vl\nW5X2qUdlj3D4+mFcuH+Bl24kNEKQYxDcOrup7TZghBBCCGkZZTXs2nSM3erVq9G7d28IhUKEhoby\njj18+BBjxoyBgYEB7OzssHPnTsWx6Oho+Pn54ZtvvgEAFBYW4p133sGWLVvqbNTVSEjwxy+/ZOHB\ngxwA8rFz//oXMGVK2zfqKqsrkXwjGd8f/57XqNMSaMHf3h+z+8xW671dlY3Giagvqjv1RvWn3qj+\nSJuOsbO0tER4eDgOHjyIsrIy3rHZs2dDKBTi3r17OHXqFIYPH44ePXrA3d0d8+fPx/z58wEAVVVV\nmDBhAiIiIuDs7Nzg88lkgKZmAG7cSEZwsK1Kwq6MMZy9dxaHrx1GUWUR71h3s+4Y7DAYIh1R2xaK\nEEIIIR2SSkKx4eHhuHXrFjZv3gwAKCkpgbGxMc6fPw8nJ/nsz3fffRcWFhb48ssveddu27YN8+fP\nh6enJwBg5syZGD9+fK3n4DgOZmbvQiSyg4VFNkaP9oKXl5diRe6av2pa8/H9kvt4Yv4EtwpvIft0\nNgDAzssOFoYWMLpjBImBpE3LQ4/pcUsfd+rUCYWF/P2JCSGENI9YLEZ8fLzicWpqKrKzswEAW7Zs\nUd8xdp999hny8vIUDbtTp07h1VdfRUlJieKclStXIjU1Ffv372/Rc3Ach6lTGczNATOzZMya5a+U\nsjdFUUURkm7ItwF7loG2gXwbMLMeFHIlakVZYz8IIeRl1tB3qVovUPx8o6a4uBgiET8caWhoiKIi\nfuiyuSwsgIqKJAQEtM0acFWyKvx16y8cyTmCyupKRboGpwFva2+8ZvMabQPWRLTfISGEENJ8KmnY\nPd8iNTAwqBXmefLkCQwNDV/oeSSSZAQEOMHFxbbxk18AYwyXCy7jYNZBPCp/xDvm2tkVQY5BMNY1\nbtUyEEIIIYS0ix67rl27oqqqCllZWYoxdmfOnEG3bt1e6Hnu3TuCO3cErdqwu1dyDwlZCbj+6Dov\n3VTPFEOchsDR2LHVnrsjo946QgghL4PU1FSlzmZu0zF21dXVkEqliIqKQl5eHjZs2ABNTU1oaGhg\n4sSJ4DgOGzduxMmTJzFixAj8+eefcHNza9FztfaYoDJpGVKyU3Di9gnImEyRrqupCz97P/S26A0B\np3Y7thFSJxpjRwghL64txti1actjyZIl0NPTw4oVK7B9+3bo6upi6dKlAIAffvgBZWVlkEgkmDx5\nMtauXdviRl1rkjEZMvIysOrvVcjIy1A06jhw6GvZFx/0+wB9LftSo+4F0VpMRNWys7MhEAjwxx9/\ntNlzCgQCxMXFtUrevr6+mDp1aqvkTVrG3t4ey5YtU3UxlGbMmDH46quvmn1dYGAg1qxZ0wolqltk\nZGSjy6WpszZtfURGRkImk/F+Fi9eDEA+BXjv3r0oLi5GdnY2JkyY0JZFa5Lrj65j7Ym1+P3q7yir\neroOn72RPWb0noFhzsOgp6WC/ckIIfUKCQlRLIguEAhw5MgRFZeofvn5+U3a+7qGQCBAWloaYmNj\nYW9v3+C5HMe1+Wz8L774otFyvcxOnDihWKNV3aWnpyM9PR0ffPABLz03NxczZ86Eg4MDhEIhrKys\nMGTIEOzbt09xTkREBKKiolBa+nTP9Jo/rGp+jIyM0L9//2atlHHr1q12/5lvDSoZY9dWIiMj4evr\n+8LjtR6WPcSha4dw6cElXrpYKEawUzBcTFxo+RIlozF26uXy5RwkJl6DVCqAlpYMgwc7vtDYVmXm\np4oGTUtJJJJmX6Mu96aupFIptLS0WiVvExOTVsm3Nctcn++++w5vvfUWdHV1FWmnT5+Gv78/HBwc\nEB0dDQ8PD1RXVyMpKQnz58+Hn58fRCIRXn31VYhEIvz000+1dqXav38/+vbti4cPH2LFihV4/fXX\ncezYMfTt27fJZWvvw0iUPcauQ8cLaxp2LVVRVYHE64n4PuN7XqNOW0MbAfYBmN13Nlw7u9IXK3mp\nXb6cg9jYLNy/74/Hj31x/74/YmOzcPlyTrvIr6Ev9Xv37iE0NBRdunSBrq4uXF1dFetr1mXRokVw\nd3eHvr4+bGxsMHPmTN6M/sLCQoSGhsLc3BxCoRA2Njb48MMPFcfT09MxcOBAiEQiiEQieHl54dCh\nQ4rjz4dii4uLMW/ePNjY2EAoFMLe3r7Wou0vIiYmBq6urtDV1UXXrl2xbNkyVFdXK47HxcWhX79+\nMDIygqmpKUaMGIGrV6/y8li2bBkcHR0hFAohkUgwZMgQlJeXIzY2FosXL0ZOTo6i1+Xzzz+vsxxS\nqRQLFiyAtbU1hEIhLCwsMHHiRMVxxhjCw8MhkUhgaGiICRMmIDo6mtd4qSu8lp6eDoFAgNzcXADA\n48ePMXnyZNja2kJPTw+urq5YuXIl75qQkBAEBgYiJiYGdnZ2EAqFqKiowN27dxESEgKJRKJojBw9\nerTJ91AXOzs7xXCkmscRERGYO3cuTExM0KVLFyxYsIBXJ8+r6dmKi4vDsGHDYGBgoIiE7dq1C15e\nXtDV1YW9vT0+/PBDXq9YWVkZpk2bBiMjIxgbG2POnDlYuHBhs8OUxcXF2L9/P8aMGaNIY4zh3Xff\nhbW1NTIyMjBq1Cg4OTnBxcUFs2bNwrlz56Cvr684f8yYMdi+fXutvI2NjSGRSODq6ooNGzZAR0cH\n+/btQ1paGjQ0NHDr1i3e+Vu3boWRkRFKS0thY2MDAPDz84NAIICDgwPv3P3798PV1RUGBgbw8/ND\nVlYW7/jvv/+OXr16QSgUwszMDLNnz+a9fjXvlfXr18PW1hadOnXCqFGjcO/evWa9fr6+voiMjGzW\nNQ3p0D12LcUYw5m7Z5B4PRHFlcW8Yz3MemCww2AY6rzYUiykYbSOnfpITLwGHZ0A8P/gDMA//ySj\nT5/m97JlZFxDaWkAL83XNwBJSckt6rWr7w+vsrIyDBo0CPr6+oiLi4OjoyOuXbuGBw8e1JuXnp4e\nNmzYAGtra2RlZWH27NmYM2cOYmNjAcgXXz916hT2798Pc3Nz3Lx5ExcuyPeGrqqqwsiRIxEWFoat\nW7cCAM6dOwe9evY5ZIxhxIgRuHXrFlavXo3u3bsjLy8Ply49/SOz5t5a0isZGRmJ2NhYfPfdd/Dy\n8sKFCxcwY8YMlJeXKxpglZWVWLx4Mdzd3VFYWIjFixdj+PDhOH/+PLS0tBAfH48VK1YgLi4OPXr0\nQEFBAdLS0gAAEyZMwOXLl7Fjxw6cOHECAHi/yJ8VExOD3bt3Y8eOHXBwcEB+fj5vbOOqVasQHR2N\nNWvWwNvbG3v37kVUVFSte27sNaioqICnpyc++ugjiMVipKenY8aMGTA2NkZISIjivIyMDIhEIvz6\n668QCASoqqqCn58fPDw8kJCQACMjI+zatQuBgYE4ffo0XF1dG72HutRVbzExMfj000+RkZGBkydP\nYtKkSejWrRvCwsIazOuTTz7BV199hTVr1oAxhtjYWCxYsAAxMTEYOHAgbt68iffffx/3799XvP8+\n+eQT7N+/H9u3b4eLiws2b96MNWvWwNTUtMHnet4ff/yBqqoq9OnTR5F25swZnD17Ftu3b4dAULsP\n6fn3fb9+/bBq1aoGexs1NDSgoaEBqVSKQYMGoWvXrti0aZOiIQsAGzZswKRJk6Cnp4eTJ0+iZ8+e\niI+Px4ABA3h7y9+5cwdr167Fzp07oaGhgbCwMISFhSnCtv/88w9GjhyJuXPnYufOnbh+/TqmT5+O\noqIixesHAMePH4dEIsGBAwdQWFiIt956Cx999BHvnDbHOqiW3trNJzfZ+hPrWURKBO9nQ+YGdvPJ\nTSWXktQnJSVF1UUgz2jo8xQdncIiIhgbNIj/ExwsT2/uT3BwSq28IiLkz6NMGzduZEKhkOXl5dV5\n/MaNG4zjOHbs2LF684iPj2c6OjqKx6NGjWIhISF1nvvw4UPGcRxLTU2tNz+O49iOHTsYY4wlJiYy\njuNYZmZmU26nUb6+vmzq1KmMMcZKSkqYnp4eO3jwIO+cLVu2MCMjo3rzKCgoYBzHsT/++IMxxtjK\nlStZ165dmVQqrfP8JUuWMDs7u0bLNnfuXObv71/vcUtLS/bZZ5/x0t544w2mpaWleBwREcGcnJx4\n5xw9epRxHMdycnLqzXvOnDksMDBQ8fjdd99lYrGYlZSUKNI2b97MrKysWFVVFe9aPz8/Nm/evCbd\nQ13s7OzY0qVLFY9tbW3ZqFGjeOcMHTqUTZw4sd48at6nX3zxBS/d1taWrVu3jpeWlpbGOI5jjx8/\nZsXFxUxHR4dt2rSJd07//v2Zs7Nzs+4jJiaGmZiY8NJ++uknxnEcO3XqVJPyyMzMZBzHsatXr/Lu\nKz09nTHGWFlZGYuIiGAcxynetytXrmS2trZMJpMxxhi7ePEi4ziOnT59mjHG2M2bNxnHcSwtLY33\nXBEREUxTU5M9ePCAV16BQMAqKioYY4xNnjyZ9evXj3fdvn37mEAgYLm5uYwx+XvFzMyMVVZWKs5Z\nsWIFMzc3r/c+G/ouVVaTrMOHYpsaty6sKET8xXhsPLkReUV5inRDbUOMcR2D9155D1Yiq1YqKXke\n9dapDy0tWZ3pGhp1pzdGIKj7Om3tluVXn8zMTHh4eMDCwqLJ18THx8PHxweWlpYwNDTE5MmTIZVK\nkZ+fDwCYNWsW9uzZA09PT8ybNw8JCQmKULBYLMaUKVMQHByMYcOGYcWKFbhy5UqD5ROLxejZs+eL\n3Wgdzp8/j7KyMowdOxaGhoaKnxkzZqCwsBAFBQUA5GOkxowZAwcHB4hEItjayntMc3LkYfE333wT\nUqkUtra2CA0Nxfbt21FcXFzv89YnNDQUZ8+ehZOTE2bOnIn4+HhIpVIA8vD27du3MWDAAN41AwcO\nbPbYKZlMhuXLl8PLywumpqYwNDTEunXrFKHaGm5ubrwepePHjyM/Px9GRka81ys9PV0RvmvoHpqK\n4zh4eXnx0szNzXH37t1Gr312zNn9+/eRm5uL+fPn88o7bNgwcByHrKwsZGVlobKyEv379+fl079/\n/2a/rnVtKNDcPGp2n3r8+DEvPSgoCIaGhjAwMMAPP/yAb7/9FkFBQQDke8rfu3cPBw8eBABs3LgR\nvXv3Ro8ePRp9PgsLC94YR3NzczDGFGHUCxcuwMfHh3eNj48PGGOKXngAcHV15fUwNrW+npWamkqh\n2KZqygtVJavCnzf/xNHco7xtwDQFmvC28sZrtq9BW0O7FUtJiHobPNgRsbFJ8PV9Gj6tqEhCSIgT\nXFyan9/ly/L8dHT4+bXG1oDN+eXz999/Y/z48Vi4cCG++eYbiMVi/Pnnn3j33XdRWSn/7ggKCkJu\nbi4OHjyI1NRUTJ48GZ6enkhKSoJAIMD69esxd+5cHDp0CIcPH0Z4eDhWr16NadOmKf3eGiKTyRvJ\ne/bsQdeuXWsdF4vFKC0tRVBQEHx8fBAbGwszMzMwxuDh4aG4XwsLC1y6dAkpKSlITk7GkiVL8Mkn\nn+Dvv/+GlVXT/xDu0aMHbty4gcOHDyMlJQVz585FeHg4/vrrrybnIRAIatXn8w2rb775BsuXL8e3\n336LV155BYaGhli5ciX++9//8s57Pkwok8ng5uaGX375pdbz1pzb0D00ZxclbW3+7xuO4xT11ZBn\nw9w1569atQp+fn61zrW0tFSE9JUxRtzIyKjWFqAu//vwnz9/vlZjtS5PnjxR5PWs2NhY9OrVSzEO\n8FnGxsZ44403sGHDBgQEBGDr1q1NXj6mrtcZAO+1bsr3w/Nh45asRVczyTMqKqpZ19WnQ/fYNYQx\nhgv3L2B1xmok3UjiNercOrthdp/ZCHAIoEaditA6durDxcUWISFOkEiSYWSUCokk+X+NupbNYlV2\nfvXp3bs3Lly4gLy8vMZPhnwgfufOnfH555+jT58+cHJyws2bN2udJxaLMWHCBKxduxb//e9/kZaW\nhosXLyqOe3h4YP78+fj999/x3nvvYf369XU+X69evfDo0SNkZma27AYb4OHhAaFQiGvXrsHBwaHW\nj0AgwMWLF/HgwQMsXboUPj4+cHFxwcOHD2v90tLW1kZwcDBWrFiBs2fPorS0VLGUhba2doMD/5+l\nr6+P0aNH47vvvsOJEydw8eJFHDlyBCKRCJaWljh27Bjv/GPHjvEaJRKJBPfu3eP9Yj558iTvmiNH\njmDo0KEICQlBjx494ODggCtXrjTauOnTpw+uX78OQ0PDWq9Vly5dGr2HtmZmZgZra2tcunSpzvrV\n0dGBk5MTtLW1a40D/Ouvv5rd2HN2dsajR494vbVeXl7w9PTEihUr6nwPFBcX89JzcnKgo6OjmPBQ\nw9LSEg4ODrUadTWmT5+OX3/9FWvXrkV5eTlvwkpN462p78FneXh41Kq7tLQ0cBwHDw8PRVp7nDzZ\noXvs6nO3+C4SshJw4/ENXrpEX4KhTkNhL6Z1lwhpDhcXW6U2vJSdX10mTpyIr776CiNHjsRXX30F\nBwcHXL9+HQUFBRg/fnyt811dXXH//n1s2rQJvr6+SE9Pr7Wo6qJFi9C7d2+4u7tDIBBg+/btMDQ0\nhI2NDbKysrBhwwaMHDkSVlZWuH37No4ePYpevXrVWb6AgAC89tprePPNN7Fy5Up4enri9u3buHTp\nEt57771m3y9jTNEoMzAwwMKFC7Fw4UJwHIeAgABUVVXh7NmzOH36NJYvXw5bW1vo6Ohg1apVWLBg\nAbKzs/Hpp5/yfpH9+OOPYIyhT58+MDIyQlJSEoqKiuDu7g5AvgBvfn4+/vrrLzg5OUFfX5+3HEaN\nr7/+GpaWlujRowf09PSwc+dOaGpqKnoTP/zwQ4SHh8PV1RX9+vXD/v37kZSUxMvD398fpaWlWLx4\nMUJDQ3Hy5En88MMPvHNcXV2xbds2pKamwsLCAlu3bkVGRgbEYnGDr92kSZMQHR2N4cOHY+nSpXB2\ndsbdu3eRnJwMd3d3jBo1qtF7qK9OGnr8IpYuXYr33nsPYrEYI0eOhJaWFi5evIiEhASsXbsW+vr6\nmD59Oj777DOYmZnB2dkZW7ZswcWLF2FmZqbIZ+/evfi///s/JCcn1ztswdvbG5qamjh+/DivhzA2\nNhYBAQHo168fwsPD4e7ujurqaqSlpeGrr77CqVOnFCHYv/76C97e3rV60hozcOBAuLi44OOPP8a7\n777L67ns3LkzDAwMcPDgQbi5uUFHR6fRuq7x8ccfo2fPnliwYAGmTZuG7OxsfPDBB5g8eTKvN1qZ\ndaY0Shmp1w7VdWsllSXst8u/sciUSN7EiOVHl7OMWxmsWlatgpIS0v511K+K/Px89s4777DOnTsz\noVDI3Nzc2JYtWxhj8sHbAoGAN3kiPDycmZmZMX19fTZ8+HC2c+dOJhAIFIPzlyxZwrp168YMDAxY\np06dmK+vr+L6O3fusLFjxzIrKyumo6PDLCws2LRp01hhYaEi/2cnTzDGWFFREfvggw+Yubk509bW\nZvb29mzFihUtutdnJ0/U2LhxI/Py8mJCoZCJxWLWv39/tnbtWsXxPXv2MGdnZyYUClnPnj1ZWloa\n09TUVLxG8fHxbMCAAUwsFjM9PT3m6enJG4wvlUrZW2+9xYyNjRnHcSwqKqrOsq1bt4716tWLiUQi\nZmBgwPr27cv279+vOC6TydjChQtZ586dmb6+Phs3bhyLjo5mmpqavHw2bdrEHBwcmK6uLhs2bBjb\ntWsXr36ePHnCxo8fz0QiETMxMWHvv/8+Cw8PZ/b29oo8QkJCeJMpahQUFLCZM2cyS0tLpq2tzSwt\nLdnYsWMVA/Ubu4e6PD954vnHjDE2ZcoU5ufnV28edb1Pa/zyyy/M29ub6enpMZFIxLy8vNiSJUsU\nx8vKyti0adOYSCRiRkZGbNasWWzu3LnM09NTcc7mzZt5r2F9xo0bxz744INa6dnZ2Wz69OnMzs6O\naWtrMwsLCxYUFMR27drFO8/JyYn9+OOPTbqv53377beM4zh24sSJWse2bt3K7O3tmaampqKeIyMj\na00QOXr0aK37/P3331mvXr2Yjo4OMzU1ZbNmzWKlpaWK43W9V7Zt28YEAkG9ZW3ou1RZ37Ntulds\nW+I4DhEREfD19cVrPq/hxO0TSMlOQXlVueIcASdAH4s+8LXzha5W7b8iCSFytFcsaW9iY2MxderU\nZk9QIA3z9/eHiYkJdu/e3azrjh07htGjRyMnJ6feJXzqc/ToUbzxxhvIzs6us0e3Mf/+97+RlJTU\nKsMWlK2u79KaBYqjoqKU8j3boRt2jDFce3gNCVkJuF96n3fcUeyIYKdgSPSbv9I7aX20jl37Qg07\n0t5Qw+7FnTt3DpmZmfD29kZlZSW2bduGr7/+GgkJCYqZp80xduxYeHt74+OPP27WdYGBgXj99dcx\nY8aMZl335MkTXLlyBUFBQYiJicHkyZObdb0qNPRdqqzv2Q49xi40OhT6ZvrobNFZkWasa4xgx2B0\nNenaLgc9EkIIaRr6Dn8xHMdh7dq1mDt3Lm/mb0sadYB8OaCWOHz4cIuuGzVqFDIyMjBx4kS1aNS1\nlQ7dYzdo8yBUZVXBy90LFtYWGGQ7CP2s+kFT0KHbs4QoHfXYEULIi6MeOyXQdNKE9IEUc96YAwNt\nA1UXhxBCCCGk1XTodexEOiL0NO8JN4kbNerUDK1jRwghhDRfh+6xe5LwBA9feQgnC+WvWE8IIYQQ\n8qJqZsUqS4ceYxeREoGKqxUI8QuBi1ML9jYihACgMXaEEKIMNMbuBUnuSRDgF0CNOkIIIYS8FDp0\nj10HvbWXAq1j177Q54kQQl5cW/TYdejJE4QQ0lLZ2dkQCAS1NklvTQKBAHFxca2St6+vL6ZOndoq\neZOWsbe3x7Jly1RdjEa15vuSKB817Ei7RL11RFlCQkIQGhoKQP4L6siRIyouUf3y8/Px+uuvN/l8\ngUCAtLQ0xMbGwt7evsFzOY5r8wV9v/jii0bL9TI7ceIE5s+fr5S8/vzzT4wZMwZdunSBrq4unJyc\n8Pbbb+PUqVNNzmPKlCnw8/NTSnmI6lDDjhDSoamiQdNSEokEOjo6zbpGXe5NXbXmlmUmJiYt2hv1\neZs3b4aPjw+EQiHi4uJw6dIl/PTTT7Czs8PcuXOVUFKiTjp0wy4yMpLWQ1NTVG/q5XLWZXz/0/f4\ndte3+P6n73E563K7ya+hMSv37t1DaGioopfD1dUVmzdvrvf8RYsWwd3dHfr6+rCxscHMmTNRWFio\nOF5YWIjQ0FCYm5tDKBTCxsYGH374oeJ4eno6Bg4cCJFIBJFIBC8vLxw6dEhx/PmQV3FxMebNmwcb\nGxsIhULY29vjyy+/bOlLUUtMTAxcXV2hq6uLrl27YtmyZaiurlYcj4uLQ79+/WBkZARTU1OMGDEC\nV69e5eWxbNkyODo6QigUQiKRYMiQISgvL0dsbCwWL16MnJwcCAQCCAQCfP7553WWQyqVYsGCBbC2\ntoZQKISFhQUmTpyoOM4YQ3h4OCQSCQwNDTFhwgRER0dDS0tLcU5kZCScnZ15+aanp0MgECA3NxcA\n8PjxY0yePBm2trbQ09ODq6srVq5cybsmJCQEgYGBiImJgZ2dHYRCISoqKnD37l2EhIRAIpFAJBLh\n1VdfxdGjR5t8D3Wxs7PD0qVLeY8jIiIwd+5cmJiYoEuXLliwYAGvTp53+/ZtzJw5E1OnTsXOnTvh\n7+8PW1tb9OrVC0uWLMGvv/4KQB4FmT59Ou9axhgcHR3xxRdfICoqCps2bUJaWpqivrZu3ao498mT\nJ3j77bchEolgbW2N5cuX8/IqKirC9OnTIZFIIBQK0adPH95WYTVDG3bv3o0RI0ZAX18fjo6O2LJl\nS4Ov0csgNTUVkZGRSsuvQ8+KVeYLRQip2+Wsy4hNiYWO89OeptiUWISgZcsMKTu/+nq0ysrKMGjQ\nIOjr6yMuLg6Ojo64du0aHjx4UG9eenp62LBhA6ytrZGVlYXZs2djzpw5iI2NBQB89tlnOHXqFPbv\n3w9zc3PcvHkTFy5cAABUVVVh5MiRCAsLU/zCPHfuHPT09Op8LsYYRowYgVu3bmH16tXo3r078vLy\ncOnSpVr31pJeycjISMTGxuK7776Dl5cXLly4gBkzZqC8vFzRAKusrMTixYvh7u6OwsJCLF68GMOH\nD8f58+ehpaWF+Ph4rFixAnFxcejRowcKCgqQlpYGAJgwYQIuX76MHTt24MSJEwAAfX39OssSExOD\n3bt3Y8eOHXBwcEB+fj5vbOOqVasQHR2NNWvWwNvbG3v37kVUVFSte27sNaioqICnpyc++ugjiMVi\npKenY8aMGTA2NkZISIjivIyMDIhEIvz6668QCASoqqqCn58fPDw8kJCQACMjI+zatQuBgYE4ffo0\nXF1dG72HutRVbzExMfj000+RkZGBkydPYtKkSejWrRvCwsLqzOPnn39G893umQAAIABJREFUZWUl\nPvvsszqPd+rUCQAwY8YMTJs2DStXrlTUQ3JyMnJzczFlyhQYGhri6tWryM7OVuz5WnMtAERFRWHp\n0qX4/PPPceDAAbz//vvo27cv/P39AQBhYWHIzMzEjh07YGNjgzVr1mDEiBH4559/4OLy9HP76aef\nYsWKFVi1ahV+/PFHTJkyBQMGDKjVKH+Z+Pr6wtfXF1FRUcrJkHVQHfjWCGlzDX2eVu9azSJSItig\nzYN4P8OWDGMRKRHN/hm6ZGitvCJSItj3P32v1HvauHEjEwqFLC8vr87jN27cYBzHsWPHjtWbR3x8\nPNPR0VE8HjVqFAsJCanz3IcPHzKO41hqamq9+XEcx3bs2MEYYywxMZFxHMcyMzObcjuN8vX1ZVOn\nTmWMMVZSUsL09PTYwYMHeeds2bKFGRkZ1ZtHQUEB4ziO/fHHH4wxxlauXMm6du3KpFJpnecvWbKE\n2dnZNVq2uXPnMn9//3qPW1pass8++4yX9sYbbzAtLS3F44iICObk5MQ75+jRo4zjOJaTk1Nv3nPm\nzGGBgYGKx++++y4Ti8WspKREkbZ582ZmZWXFqqqqeNf6+fmxefPmNeke6mJnZ8eWLl2qeGxra8tG\njRrFO2fo0KFs4sSJ9eYxc+bMBuusRnl5OTM1NWUbN25UpE2YMIGNHj1a8fi9995jvr6+ta7lOI7N\nnTuXl+bm5sb+7//+jzHG2NWrVxnHcezAgQO8c3r27MnCwsIYY08/T9HR0Yrj1dXVzNDQkK1fv77R\n8ncUDX2XKqvd0qFDsYSQ1idldY9Bqkb94aOGyCCrM71SVtmi/OqTmZkJDw8PWFhYNPma+Ph4+Pj4\nwNLSEoaGhpg8eTKkUiny8/MBALNmzcKePXvg6emJefPmISEhQREKFovFmDJlCoKDgzFs2DCsWLEC\nV65cabB8YrEYPXv2fLEbrcP58+dRVlaGsWPHwtDQUPEzY8YMFBYWoqCgAABw+vRpjBkzBg4ODhCJ\nRLC1tQUA5OTkAADefPNNSKVS2NraIjQ0FNu3b0dxcXGzyxMaGoqzZ8/CyckJM2fORHx8vGJsW2Fh\nIW7fvo0BAwbwrhk4cGCzl4aQyWRYvnw5vLy8YGpqCkNDQ6xbt04Rqq3h5ubG60k9fvw48vPzYWRk\nxHu90tPTkZWV1eg9NBXHcfDy8uKlmZub4+7du/Vewxhr0uugo6ODkJAQbNiwAQBQUFCAX375pckz\npZ8vl4WFBe7duwcAil5pHx8f3jk+Pj44f/58vfkIBAJIJJIG7480HzXsSLtEY+zUhxanVWe6BjRa\nlJ+gnq8lbYF2i/JrSHMaBn///TfGjx8PX19f/PLLLzh16hTWrl0LxhgqK+WNzqCgIOTm5mLRokUo\nLy/H5MmT4e/vD5lM3lhdv349MjMzERgYiLS0NHTr1g3r169X+n01pqY8e/bswZkzZxQ/586dw9Wr\nVyEWi1FaWoqgoCBoaGggNjYWx48fx/Hjx8FxnOJ+LSwscOnSJWzatAkSiQRLliyBi4sLbt261azy\n9OjRAzdu3MB//vMfaGtrY+7cufDy8kJRUVGT8xAIBLXq8/mG1TfffIPly5dj3rx5SExMxJkzZzBl\nyhRUVFTwzns+PC6TyeDm5sZ7rc6cOYNLly4pGkrKuAcA0Nbmv885jlPUV11cXV1RWFiIvLy8RvOe\nPn06jh8/jrNnz2Lbtm2QSCQYOnRoi8oFoMFyAXV/vpp7f6T5OvQYO0JI6xvcazBiU2Lh6+yrSKu4\nWoGQCS0cY2dVe4xdxdUKBPgFKKO4Cr1798bmzZuRl5cHS0vLRs9PT09H586deRMAfv7551rnicVi\nTJgwARMmTEBoaCi8vb1x8eJFeHh4AAA8PDzg4eGB+fPnY+bMmVi/fj2mTZtWK59evXrh0aNHyMzM\nRK9evV7gTmvz8PCAUCjEtWvXMGTIkDrPuXjxIh48eIClS5cqxkj98ccftX5Za2trIzg4GMHBwViy\nZAnMzMywb98+zJ49G9ra2g0O/H+Wvr4+Ro8ejdGjR2PhwoUwNzfHkSNHMHz4cFhaWuLYsWO8Rsix\nY8d449MkEgnu3bsHmUwGgUD+x8HJkyd5z3HkyBEMHTqUN57uypUrjY7N69OnD7Zt2wZDQ0OYmpq2\n6B5ay7hx4/Dpp5/iiy++wJo1a2odf/ToEcRiMQDA0dER/v7+2LBhA1JSUhAWFsa79+bU17PX1by3\n09LSeHV05MgRpb93SeOoYUfaJVrHTn24OLkgBCFIOpmESlkltAXaL7SVn7Lzq8/EiRPx1VdfYeTI\nkfjqq6/g4OCA69evo6CgAOPHj691vqurK+7fv49NmzbB19cX6enptX6RLlq0CL1794a7uzsEAgG2\nb98OQ0ND2NjYICsrCxs2bMDIkSNhZWWF27dv4+jRo/X+4gsICMBrr72GN998EytXroSnpydu376N\nS5cu4b333mv2/T4bsjMwMMDChQuxcOFCcByHgIAAVFVV4ezZszh9+jSWL18OW1tb6OjoYNWqVViw\nYAGys7Px6aef8n6h//jjj2CMoU+fPjAyMkJSUhKKiorg7u4OQL4Ab35+Pv766y84OTlBX1+/zuU9\nvv76a1haWqJHjx7Q09PDzp07oampia5duwIAPvzwQ4SHh8PV1RX9+vXD/v37kZSUxMvD398fpaWl\nWLx4MUJDQ3Hy5En88MMPvHNcXV2xbds2pKamwsLCAlu3bkVGRoai4VOfSZMmITo6GsOHD8fSpUvh\n7OyMu3fvIjk5Ge7u7hg1alSj91BfnTT0uCksLCywevVqTJ8+HY8fP8bUqVPh4OCAhw8fYt++fUhN\nTVVMaAHkvXaTJk2CTCbDlClTeHk5ODhgz549uHDhgmL2b109dTVlrSmvo6Mjxo0bh1mzZmHdunWK\nyRMXLlzArl27Gix/S+6ZNEIpI/XaoQ58a4S0uY76ecrPz2fvvPMO69y5MxMKhczNzY1t2bKFMSYf\n7C0QCHiTJ8LDw5mZmRnT19dnw4cPZzt37mQCgUAxOH/JkiWsW7duzMDAgHXq1In5+voqrr9z5w4b\nO3Yss7KyYjo6OszCwoJNmzaNFRYWKvJ/dvIEY4wVFRWxDz74gJmbmzNtbW1mb2/PVqxY0aJ7fXby\nRI2NGzcyLy8vJhQKmVgsZv3792dr165VHN+zZw9zdnZmQqGQ9ezZk6WlpTFNTU3FaxQfH88GDBjA\nxGIx09PTY56enmzTpk2K66VSKXvrrbeYsbEx4ziORUVF1Vm2devWsV69ejGRSMQMDAxY37592f79\n+xXHZTIZW7hwIevcuTPT19dn48aNY9HR0UxTU5OXz6ZNm5iDgwPT1dVlw4YNY7t27eLVz5MnT9j4\n8eOZSCRiJiYm7P3332fh4eHM3t5ekUdISAhvMkWNgoICNnPmTGZpacm0tbWZpaUlGzt2LDt9+nST\n7qEuz0+eeP4xY4xNmTKF+fn5NZgPY4ylp6ez0aNHM4lEwnR0dJiDgwObMGEC+/vvv3nnSaVSJpFI\n2IgRI2rl8fDhQzZs2DDWqVMnxnGcop6ff18yxtjgwYNZaGio4nFhYSGbPn06MzU1ZTo6OqxPnz7s\n8OHDiuN1fZ4YY8zJyane90VH1NB3qbK+Z2mvWNIu0V6x7Qt9nkh7Exsbi6lTp7bqAsIdUUFBAayt\nrfHTTz/hX//6l6qL89KhvWJfEC1QTAghhMjXUczPz8eiRYtgZWVFjbp2RNkLFFOPHSGkUfR5Iu1N\nbGwspk2bppihSxqWmpoKf39/ODg4YNu2bfD29lZ1kV5KbdFjRw07Qkij6PNECCEvjkKx5KVFIXRC\nCCGk+ahhRwghhBDSQVAolhDSKPo8EULIi6NQLCGEEEIIaTJq2JF2icbYEUIIIc1HDTtCCCGEkA6C\nxtgRQhpFnydCCHlxNMaOEEJUJDs7GwKBAH/88UebPadAIEBcXFyr5O3r64upU6e2St6kZezt7bFs\n2TJVF0MpIiMj4ezsrOpiEFDDjrRTNMaOKEtISAhCQ0MByBtOR44cUXGJ6pefn4/XX3+9yecLBAKk\npaUhNjYW9vb2DZ7LcRw4jnvRIjbLF1980Wi5XmYnTpzA/PnzXzgfgUAATU1NnDt3jpdOr//LSVPV\nBSCEqL+cy5dxLTERAqkUMi0tOA4eDFsXl3aRnyoaNC0lkUiafY263Ju6kkql0NLSapW8TUxMlJaX\njo4OPv74Yxw4cEBpeRL11KF77CIjI6nnR035+vqqugikiXIuX0ZWbCz879+H7+PH8L9/H1mxsci5\nfLld5NfQmJV79+4hNDQUXbp0ga6uLlxdXbF58+Z6z1+0aBHc3d2hr68PGxsbzJw5E4WFhYrjhYWF\nCA0Nhbm5OYRCIWxsbPDhhx8qjqenp2PgwIEQiUQQiUTw8vLCoUOHFMefD8UWFxdj3rx5sLGxgVAo\nhL29Pb788ssWvQ51iYmJgaurK3R1ddG1a1csW7YM1dXViuNxcXHo168fjIyMYGpqihEjRuDq1au8\nPJYtWwZHR0cIhUJIJBIMGTIE5eXliI2NxeLFi5GTkwOBQACBQIDPP/+8znJIpVIsWLAA1tbWEAqF\nsLCwwMSJExXHGWMIDw+HRCKBoaEhJkyYgOjoaF6Dq65QYHp6OgQCAXJzcwEAjx8/xuTJk2Fraws9\nPT24urpi5cqVvGtCQkIQGBiImJgY2NnZQSgUoqKiAnfv3kVISAgkEglEIhFeffVVHD16tMn3UBc7\nOzssXbqU9zgiIgJz586FiYkJunTpggULFvDqpD4ffPABDh8+jMTExHrPacprFBsbCy0tLaSmpsLT\n0xN6enrw9/dHfn4+UlJS4OXlBQMDAwQGBuL27du1niMuLg4ODg7Q1dVFUFAQcnJyFMdu3LiBsWPH\nwtLSEvr6+ujevTu2b9/e6L11dKmpqYiMjFRafh26x06ZLxQhpG7XEhMRoKMDPPNHVACA5H/+gW2f\nPs3PLyMDAaWlvLQAX18kJyW1qNeuvh6tsrIyDBo0CPr6+oiLi4OjoyOuXbuGBw8e1JuXnp4eNmzY\nAGtra2RlZWH27NmYM2cOYmNjAQCfffYZTp06hf3798Pc3Bw3b97EhQsXAABVVVUYOXIkwsLCsHXr\nVgDAuXPnoKenV+dzMcYwYsQI3Lp1C6tXr0b37t2Rl5eHS5cu1bq3lvRKRkZGIjY2Ft999x28vLxw\n4cIFzJgxA+Xl5YoGWGVlJRYvXgx3d3cUFhZi8eLFGD58OM6fPw8tLS3Ex8djxYoViIuLQ48ePVBQ\nUIC0tDQAwIQJE3D58mXs2LEDJ06cAADo6+vXWZaYmBjs3r0bO3bsgIODA/Lz83ljG1etWoXo6Gis\nWbMG3t7e2Lt3L6Kiomrdc2OvQUVFBTw9PfHRRx9BLBYjPT0dM2bMgLGxMUJCQhTnZWRkQCQS4ddf\nf4VAIEBVVRX8/Pzg4eGBhIQEGBkZYdeuXQgMDMTp06fh6ura6D3Upa56i4mJwaeffoqMjAycPHkS\nkyZNQrdu3RAWFtZgXp6enggJCcHHH3+MkydP1vtaNOV9IpPJ8Pnnn2PTpk3Q1NTEm2++iXHjxkEg\nEGD9+vXQ0dHBhAkTsGDBAuzatUtx3Z07d7B27Vrs2bMHMpkM77//PsaOHYvMzEwAQElJCQYPHoyo\nqCgYGBjgv//9L0JDQ2FlZfVS/zHv6+sLX19fREVFKSW/Dt2wI+orNTX1pf6gqxOBVFp3ehN6Geq8\nTiarO72yskX5PdsDJ3sm77i4OGRnZ+PatWuwsLAAANja2jaY16JFixT/t7GxwbJlyzBx4kRFwy43\nNxevvPIK+vyvQWtlZQVvb28AQFFRER4/fox//etfcHR0BADFv3VJTk7GkSNHcOLECfTs2ROAvEdn\n4MCBinNqenJ8fHzw7rvvNvxCPKO0tBRff/019u7di6CgIMW9L1myBHPnzlU07J5t7ADy17Jz5844\nceIEvL29kZOTgy5duiA4OBiampqwsrJCjx49FOfr6+tDQ0Oj0RBzbm4uunbtCh8fHwDy1613796K\n419//TXmz5+Pt99+GwDw8ccfIyMjA/v27ePl09iMQjMzM3zyySeKx7a2tsjIyEBcXBzvXjU0NLBt\n2zZFozs2NhZFRUXYtWsXNDQ0AAALFy5EYmIi1q1bh+jo6Ebvoal8fHzw73//G4D8/bF582YkJiY2\n2rDjOA5LliyBs7MztmzZUqvuajRl1iVjDN9++y26d+8OAJg2bRr+/e9/IzMzE6+88goAYPr06bze\nRkD+voqNjYWDgwMAYNu2bXBxcUFycjL8/f3RrVs3dOvWTXH++++/j8TERMTFxdH3vRJ16FAsIaT1\nyeoZfyT73y/AZucnqPtrSaat3aL86pOZmQkPDw9Fo64p4uPj4ePjA0tLSxgaGmLy5MmQSqXIz88H\nAMyaNQt79uyBp6cn5s2bh4SEBMUvUrFYjClTpiA4OBjDhg3DihUrcOXKlQbLJxaLFY06ZTp//jzK\nysowduxYGBoaKn5mzJiBwsJCFBQUAABOnz6NMWPGwMHBASKRSNHwrQmvvfnmm5BKpbC1tUVoaCi2\nb9+O4uLiZpcnNDQUZ8+ehZOTE2bOnIn4+HhI//cHQ2FhIW7fvo0BAwbwrhk4cGCzl4aQyWRYvnw5\nvLy8YGpqCkNDQ6xbt04Rhqzh5ubG60k9fvw48vPzYWRkxHu90tPTkZWV1eg9NBXHcfDy8uKlmZub\n4+7du0263tzcHB9++CHCw8Px/+3deVAUZ/oH8O8gDAPMjI6iyFEKCIoIiwmia0wIikexxotdD7Ie\nsAYsrzVuVkMgoC5SCSagbnklbowoOkazVsqrSi0Vr1JYK0BZEYJoQDciKBgOUQLD+/vDZX4Og3JI\nZmj4fqqmyu5+++2n+3GGZ97unn769Gmbtt00Dl9fX/20g4MDAOgLvcZ5ZWVlBjno27evvqgDAE9P\nT9jb2+tHrWtqahAdHQ0fHx/06dMHKpUKJ06cMDr+9Go4YkedEr+9Sceg8eNxZvduBD+XszO1tfAI\nDwfacep00I8/PuvP2tqwv+DgDojWUFsKg4yMDMyaNQsxMTFITk6GRqPBlStXsGDBAvz6v9HEiRMn\n4s6dOzh58iTS09Mxd+5c+Pr64syZM/rTWCtWrMCpU6dw+vRpxMXFYcuWLYiKiurwfXuZxpHLb7/9\nFoMHDzZartFoUFNTg4kTJyIwMBC7d++Gg4MDhBAYNmyYfn+dnJyQl5eHc+fO4ezZs0hISMCHH36I\njIwMuLi4tDoePz8//PTTTzh9+jTOnTuHFStWIC4uDlevXm11HxYWFkb5bFpYJScn49NPP8WmTZvw\n2muvQaVSISUlBcePHzdo1/T0eENDA4YOHYrvvvvOaLuNbV+2DyqVqtX7IW/yBUYmkxmMNLdk9erV\n2LlzJ5KTk41Ou7bmGDW2e37dxn/3eO7LWuM8IUSrLwNYtWoVjhw5go0bN2LIkCGwtbXFBx98YHCd\nKr06jtgR0SsZOGQIPMLDcbZfP6T36oWz/frBIzy83XexdnR/LzJixAjcuHEDP//8c6vaX7p0Cfb2\n9vjHP/6BgIAAeHh44O7du0btNBoN5syZgx07duD48eM4f/48cnNz9cuHDRuGlStX4sSJE1i4cCG+\n/PLLZrfn7++PR48e6a9P6kjDhg2DQqHArVu34O7ubvSysLBAbm4uHj58iMTERAQGBmLIkCEoLy83\nKgzkcjkmTZqEpKQkXL9+HTU1NfpTpHK5vFUX/gPPTttOnz4dmzdvxrVr15Cbm4sLFy5ArVbD2dkZ\nly9fNmh/+fJlg4KiX79+KC0tNSiCvv/+e4N1Lly4gJCQEISHh8PPzw/u7u7Iz89vsTAJCAjA7du3\noVKpjI5V//79W9wHU7Kzs8O6deuwYcMGo5G+1hyjV/HgwQPcvn1bP52fn4+HDx/C29sbwLPjP3fu\nXPzpT3+Cr68v3Nzc8GM7b4qiF+OIHXVKvMZOWgYOGdKhhVdH99ecsLAwbNiwAVOnTsWGDRvg7u6O\n27dvo6ysDLNmzTJq7+XlhQcPHmDXrl0ICgrCpUuXsH37doM2sbGxGDFiBLy9vWFhYYG0tDSoVCoM\nGDAABQUF2LlzJ6ZOnQoXFxfcu3cPFy9ehL+/f7PxBQcH46233sLs2bORkpICX19f3Lt3D3l5eVi4\ncGGb91cIoS/KlEolYmJiEBMTA5lMhuDgYNTX1+P69evIzs7Gp59+ioEDB8La2hr//Oc/8be//Q2F\nhYWIjo42KIK++uorCCEQEBCAXr164cyZM6iqqtL/IXdzc8P9+/dx9epVeHh4wM7ODjY2NkaxffbZ\nZ3B2doafnx9sbW2h1WphaWmpH01sPL3o5eWFUaNG4ciRIzhz5oxBH+PGjUNNTQ3i4+MRERGB77//\nHtu2bTNo4+Xlhb179yI9PR1OTk7Ys2cPMjMzodFoXnrs/vznP2Pjxo2YPHkyEhMT4enpiZKSEpw9\nexbe3t6YNm1ai/vwopy8bLq9Fi5ciM2bN+Orr74yuL6xNcfoVdja2iIiIgIpKSkQQmD58uV47bXX\nMG7cOADPjv93332H0NBQ2NnZISUlBcXFxXB0dOywGAiA6KK68K51C+fOnTN3CPScrvp+un//vpg/\nf76wt7cXCoVCDB06VKSmpgohhPjpp5+EhYWFuHz5sr59XFyccHBwEHZ2dmLy5MlCq9UKCwsLUVRU\nJIQQIiEhQfj4+AilUil69uwpgoKC9OsXFxeL0NBQ4eLiIqytrYWTk5OIiooSlZWV+v5lMpnYt2+f\nfrqqqkosX75cODo6CrlcLtzc3ERSUlK79jUoKEhERkYazPvXv/4lhg8fLhQKhdBoNOL3v/+92LFj\nh375t99+Kzw9PYVCoRCvv/66OH/+vLC0tNQfo8OHD4s33nhDaDQaYWtrK3x9fcWuXbv069fV1Yl3\n331X9O7dW8hkMrFu3bpmY/viiy+Ev7+/UKvVQqlUipEjR4ojR47olzc0NIiYmBhhb28v7OzsxMyZ\nM8XGjRuFpaWlQT+7du0S7u7uwsbGRvzhD38QBw4cMMhPRUWFmDVrllCr1aJPnz5i2bJlIi4uTri5\nuen7CA8PFxMmTDCKsaysTCxevFg4OzsLuVwunJ2dRWhoqMjOzm7VPjTH1dVVJCYmvnBaCCHee+89\nMXbs2Jf20/T/jRBCHD9+XMhkMoN9a80x+vrrr4WVlZXBOnv37hUWFhYG8xr/7+t0OiGEEGvXrhWe\nnp5i3759wtXVVSgUCjF+/HhRWFioX+fu3bti0qRJws7OTjg6Ooq1a9eKhQsXtrh/XcnLPks76nOW\nz4olohbx/USdze7duxEZGdnmGxSIzInPiiUiIiKiVmNhR50SnxhCRC3h49SIjLGwIyIiyQkPD9f/\n7AoR/T9eY0dELeL7iYjo1fEaOyIiIiJqNRZ21CnxGjsiIqK2Y2FHRERE1EVI7hq7kpIShIaGQi6X\nQy6XY//+/ejTp49RO14TRNRxevfujUePHpk7DCIiSdNoNCgvL292WUfVLZIr7BoaGmBh8WygMTU1\nFcXFxYiOjjZqx8KOiIiIpKLb3jzRWNQBQGVlZYvP+CNp4jV20sXcSRvzJ23MH0musAOAnJwcjBo1\nClu2bEFYWJi5w6HfQHZ2trlDoHZi7qSN+ZM25o9MWtht2bIFI0aMgEKhQEREhMGy8vJyzJgxA0ql\nEq6urtBqtfplGzduxNixY5GcnAwA8PPzQ0ZGBtavX4+EhART7gKZyC+//GLuEKidmDtpY/6kjfkj\nkxZ2zs7OiIuLw1/+8hejZUuXLoVCoUBpaSn27duHxYsX48aNGwCAlStX4ty5c/jggw8MHvisVqtR\nW1trsvjboyOHxdvbV1vWa03bl7Vpz7LOeuqgo+PqivnrrLkDpJe/V83dy5ZL7b0H8LOzpWXdJXev\n0l9H5k9K7z2TFnYzZszAtGnTjO5iffz4MQ4fPoyEhATY2tpizJgxmDZtGvbu3WvUR3Z2Nt5++22M\nGzcOKSkpWL16tanCbxd+OLW8rLn5hYWFLcbxW+uKH04ttemIPy6dIXeA9PLXWQq7rpi/7vLeAzpH\n/qT23mtNWykVdma5K/bjjz/Gzz//jK+//hoAkJWVhTfffBOPHz/Wt0lJSUF6ejqOHDnSrm14eHjg\n1q1bHRIvERER0W9p0KBBKCgoeOV+LDsgljaTyWQG09XV1VCr1QbzVCoVqqqq2r2Njjg4RERERFJi\nlrtimw4SKpVKVFZWGsyrqKiASqUyZVhEREREkmaWwq7piN3gwYNRX19vMMqWk5MDHx8fU4dGRERE\nJFkmLex0Oh2ePn2K+vp66HQ61NbWQqfTwc7ODqGhoYiPj0dNTQ0uXbqEo0ePYt68eaYMj4iIiEjS\nTFrYNd71mpSUhLS0NNjY2CAxMREAsG3bNjx58gT9+vXD3LlzsWPHDgwdOtSU4RERERFJmuSeFfsq\nKisrMX78eOTm5iIjIwPe3t7mDonaIDMzE++//z6srKzg7OyMPXv2wNLSLPf/UBuVlJQgNDQUcrkc\ncrkc+/fvN/rZI+r8tFotVqxYgdLSUnOHQm1QWFiIgIAA+Pj4QCaT4eDBg7C3tzd3WNRK6enpWL9+\nPRoaGvDXv/4V06dPf2n7blXY1dfX45dffsGqVavw97//HcOGDTN3SNQG9+/fh0ajgbW1NWJiYuDv\n748//vGP5g6LWqGhoUH/nOfU1FQUFxcjOjrazFFRW+h0OsycORN37tzBtWvXzB0OtUFhYSFWrVqF\nQ4cOmTsUaqMnT55g9uzZ+Pe//w0rK6tWrSPJZ8W2l6WlJb+lSFj//v1hbW0NALCyskKPHj3MHBG1\nVmNRBzwbOddoNGaMhtpDq9Vi1qxZRje/kTRcvnwZgYGBiI2NNXco1AZXrlyBjY0NpkyZgtDQUJSU\nlLS4Trcq7KhrKCoqwunTpzFlyhRzh0JtkJOTg1GjRmHLli0ICwsdOHdqAAAJYUlEQVQzdzjUBjqd\nDocOHcLs2bPNHQq1g5OTE27duoULFy6gtLQUhw8fNndI1EolJSUoKCjAsWPHEBkZibVr17a4jiQL\nuy1btmDEiBFQKBSIiIgwWFZeXo4ZM2ZAqVTC1dUVWq222T74rdN8XiV/lZWVmD9/PlJTUzliZwav\nkjs/Pz9kZGRg/fr1SEhIMGXY9D/tzV9aWhpH6zqB9uZPLpfDxsYGABAaGoqcnByTxk3tz51Go8GY\nMWNgaWmJcePG4YcffmhxW5K88tzZ2RlxcXE4efIknjx5YrBs6dKlUCgUKC0tRVZWFiZPngw/Pz+j\nGyW60aWFnU5781dfX485c+ZgzZo18PT0NFP03Vt7c1dXV6e/PkStVqO2ttYc4Xd77c1fbm4usrKy\nkJaWhps3b+L999/Hpk2bzLQX3Vd781ddXQ2lUgkAuHDhAq8vN4P25i4gIADJyckAgOzsbAwaNKjl\njQkJ+/jjj0V4eLh+urq6WsjlcnHz5k39vPnz54vo6Gj9dEhIiHBychKjR48Wu3fvNmm8ZKit+duz\nZ4/o06ePCAoKEkFBQeKbb74xecz0TFtzl5GRIQIDA8XYsWPFxIkTxd27d00eM/2/9nx2NgoICDBJ\njPRibc3fiRMnhL+/v3jrrbfEggULhE6nM3nM9Ex73ntbt24VgYGBIigoSNy+fbvFbUhyxK6RaDLq\nlp+fD0tLS3h4eOjn+fn5IT09XT994sQJU4VHLWhr/ubNm8cfre4k2pq7kSNH4vz586YMkV6iPZ+d\njTIzM3/r8KgFbc1fSEgIQkJCTBkivUB73ntLlizBkiVLWr0NSV5j16jp9R7V1dVQq9UG81QqFaqq\nqkwZFrUS8yddzJ20MX/SxvxJlylyJ+nCrmnlq1QqUVlZaTCvoqICKpXKlGFRKzF/0sXcSRvzJ23M\nn3SZIneSLuyaVr6DBw9GfX09CgoK9PNycnLg4+Nj6tCoFZg/6WLupI35kzbmT7pMkTtJFnY6nQ5P\nnz5FfX09dDodamtrodPpYGdnh9DQUMTHx6OmpgaXLl3C0aNHeV1WJ8P8SRdzJ23Mn7Qxf9Jl0ty9\n+j0eprdmzRohk8kMXuvWrRNCCFFeXi6mT58u7OzsxMCBA4VWqzVztNQU8yddzJ20MX/SxvxJlylz\n162eFUtERETUlUnyVCwRERERGWNhR0RERNRFsLAjIiIi6iJY2BERERF1ESzsiIiIiLoIFnZERERE\nXQQLOyIiIqIugoUdERERURfBwo6IqInw8HDExcV1aJ+LFy/G+vXrO7RPIqKmLM0dABFRZyOTyYwe\n1v2qtm/f3qH9ERE1hyN2RETN4NMWiUiKWNgRUaeSlJQEFxcXqNVqeHl54ezZswCAzMxMjB49GhqN\nBk5OTli+fDnq6ur061lYWGD79u3w9PSEWq1GfHw8bt26hdGjR6NXr16YM2eOvn16ejpcXFzwySef\noG/fvnBzc8P+/ftfGNOxY8cwfPhwaDQajBkzBtevX39h25UrV8LBwQE9e/bE7373O9y4cQOA4end\nKVOmQKVS6V89evTAnj17AAB5eXmYMGEC+vTpAy8vLxw6dOiF2woKCkJ8fDzefPNNqNVqTJo0CWVl\nZa080kTUFbGwI6JO48cff8TWrVtx7do1VFZW4tSpU3B1dQUAWFpaYvPmzSgrK8OVK1dw5swZbNu2\nzWD9U6dOISsrC1evXkVSUhIiIyOh1Wpx584dXL9+HVqtVt+2pKQEZWVluHfvHlJTUxEVFYWbN28a\nxZSVlYWFCxdi586dKC8vx6JFizB16lT8+uuvRm1PnjyJixcv4ubNm6ioqMChQ4fQu3dvAIand48e\nPYqqqipUVVXh4MGDcHR0RHBwMB4/fowJEyZg7ty5ePDgAQ4cOIAlS5YgNzf3hcdMq9Vi9+7dKC0t\nxa+//orPP/+8zcediLoOFnZE1Gn06NEDtbW1+OGHH1BXV4cBAwbA3d0dAPD6669j5MiRsLCwwMCB\nAxEVFYXz588brL969WoolUp4e3vD19cXISEhcHV1hVqtRkhICLKysgzaJyQkwMrKCoGBgZg8eTK+\n+eYb/bLGIuzLL7/EokWLEBAQAJlMhvnz58Pa2hpXr141il8ul6Oqqgq5ubloaGjAkCFD0L9/f/3y\npqd38/PzER4ejoMHD8LZ2RnHjh2Dm5sbFixYAAsLCwwfPhyhoaEvHLWTyWSIiIiAh4cHFAoFZs2a\nhezs7DYccSLqaljYEVGn4eHhgU2bNmHt2rVwcHBAWFgYiouLATwrgt555x04OjqiZ8+eiI2NNTrt\n6ODgoP+3jY2NwbRCoUB1dbV+WqPRwMbGRj89cOBA/baeV1RUhOTkZGg0Gv3rv//9b7Ntx44di2XL\nlmHp0qVwcHDAokWLUFVV1ey+VlRUYNq0aUhMTMQbb7yh31ZGRobBtvbv34+SkpIXHrPnC0cbGxuD\nfSSi7oeFHRF1KmFhYbh48SKKioogk8nw4YcfAnj2cyHe3t4oKChARUUFEhMT0dDQ0Op+m97l+ujR\nI9TU1Oini4qK4OTkZLTegAEDEBsbi0ePHulf1dXVmD17drPbWb58Oa5du4YbN24gPz8fn332mVGb\nhoYGvPvuuwgODsZ7771nsK23337bYFtVVVXYunVrq/eTiLo3FnZE1Gnk5+fj7NmzqK2thbW1NRQK\nBXr06AEAqK6uhkqlgq2tLfLy8lr18yHPn/ps7i7XNWvWoK6uDhcvXsTx48cxc+ZMfdvG9pGRkdix\nYwcyMzMhhMDjx49x/PjxZkfGrl27hoyMDNTV1cHW1tYg/ue3Hxsbi5qaGmzatMlg/XfeeQf5+flI\nS0tDXV0d6urq8J///Ad5eXmt2kciIhZ2RNRp1NbW4qOPPkLfvn3h6OiIhw8f4pNPPgEAfP7559i/\nfz/UajWioqIwZ84cg1G45n53runy56f79++vv8N23rx5+OKLLzB48GCjtv7+/ti5cyeWLVuG3r17\nw9PTU38Ha1OVlZWIiopC79694erqCnt7e6xatcqozwMHDuhPuTbeGavVaqFUKnHq1CkcOHAAzs7O\ncHR0xEcffdTsjRqt2Uci6n5kgl/3iKibSU9Px7x583D37l1zh0JE1KE4YkdERETURbCwI6Juiacs\niagr4qlYIiIioi6CI3ZEREREXQQLOyIiIqIugoUdERERURfBwo6IiIioi2BhR0RERNRF/B91bdtd\n40wV6QAAAABJRU5ErkJggg==\n", + "text": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "# Performance growth rates: NumPy and SciPy library functions" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Okay, now that we have seen that Cython improved the performance of our Python code and that the matrix equation (using NumPy) performed even better, let us see how they compare against the in-built NumPy and SciPy library functions. \n", + "\n", + "Note that we are now passing `numpy.arrays` to the NumPy, SciPy, and (C)Python matrix functions (not to the Cython implemtation though!), since they are optimized for it." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "import timeit\n", + "import random\n", + "random.seed(12345)\n", + "\n", + "funcs = ['cy_classic_lstsqr', 'py_matrix_lstsqr',\n", + " 'numpy_lstsqr', 'scipy_lstsqr']\n", + "\n", + "orders_n = [10**n for n in range(1, 7)]\n", + "perf2 = {f:[] for f in funcs}\n", + "\n", + "for n in orders_n:\n", + " x_list = [x_i*random.randrange(8,12)/10 for x_i in range(n)]\n", + " y_list = [y_i*random.randrange(10,14)/10 for y_i in range(n)]\n", + " x_ary = np.asarray(x_list)\n", + " y_ary = np.asarray(y_list)\n", + " for f in funcs:\n", + " if f != 'cy_classic_lstsqr':\n", + " perf2[f].append(timeit.Timer('%s(x_ary,y_ary)' %f, \n", + " 'from __main__ import %s, x_ary, y_ary' %f).timeit(1000))\n", + " else:\n", + " perf2[f].append(timeit.Timer('%s(x_list,y_list)' %f, \n", + " 'from __main__ import %s, x_list, y_list' %f).timeit(1000))\n" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 35 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "labels = [ ('cy_classic_lstsqr', '\"classic\" least squares in Cython (numpy arrays)'),\n", + " ('py_matrix_lstsqr', '\"matrix\" least squares in (C)Python and NumPy'),\n", + " ('numpy_lstsqr', 'in NumPy'),\n", + " ('scipy_lstsqr','in SciPy'),\n", + " ]\n", + "\n", + "plt.rcParams.update({'font.size': 12})\n", + "\n", + "fig = plt.figure(figsize=(10,8))\n", + "for lb in labels:\n", + " plt.plot(orders_n, perf2[lb[0]], alpha=0.5, label=lb[1], marker='o', lw=3)\n", + "plt.xlabel('sample size n')\n", + "plt.ylabel('time per computation in milliseconds [ms]')\n", + "#plt.xlim([1,max(orders_n) + max(orders_n) * 10])\n", + "plt.legend(loc=4)\n", + "plt.grid()\n", + "plt.xscale('log')\n", + "plt.yscale('log')\n", + "plt.title('Performance of least square fit implementations')\n", + "plt.show()" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "display_data", + "png": "iVBORw0KGgoAAAANSUhEUgAAAnYAAAIECAYAAACUvmMzAAAABHNCSVQICAgIfAhkiAAAAAlwSFlz\nAAALEgAACxIB0t1+/AAAIABJREFUeJzs3Xdc09f+P/DXJ4RN2ENwgIADQcFtnbj3tgpWW2312nF7\n29723tufrQVrrR3f1ttWe9uqdbYqpe5aN3EvXBUUZCgKArL3SnJ+f3wkgxlJIPnA+/l49PFoTpKT\nk5ykvPv+vM85HGOMgRBCCCGECJ7I0AMghBBCCCH6QYEdIYQQQkgrQYEdIYQQQkgrQYEdIYQQQkgr\nQYEdIYQQQkgrQYEdIYQQQkgrQYEdabVkMhlefvllODs7QyQS4cyZM4YekiD99ttv8PHxgVgsxssv\nv1znYxYtWoSxY8e28MhIzbk5ffo0RCIRHj9+/Mx9de7cGZ9++mkzjLI2Ly8vrF69ukVey1iJRCL8\n+uuvhh4GaYUosCMGtWjRIohEIohEIpiamsLLywuvvfYacnNzde77999/x86dO3Ho0CFkZGTgueee\n08OI2xa5XI6XX34ZISEhePToEb755ps6H8dxHDiOa9Gx7dixAyJR2/1PWF1zM3jwYGRkZMDd3R0A\ncO7cOYhEIjx8+LDR/qKjo/HOO+8097ABGOb7oovU1NQm/8/hmDFjsHjx4lrtGRkZmD17tj6GR4gG\nsaEHQMjw4cMREREBmUyG6OhoLF26FI8ePcKhQ4ea1F9lZSXMzMyQkJCA9u3bY9CgQTqNr7q/tujx\n48coKSnBxIkTlcFCXRhjoL3On51CoQCAJgWo9c2Nq6trrcdqMzdOTk7PPIa2Rp/f8brmiRB9aLv/\nu0uMhqmpKVxdXeHh4YFp06bhrbfewpEjR1BRUQEA2LVrF4KCgmBpaYnOnTvj3XffRWlpqfL5wcHB\nWLJkCVasWAEPDw94enpi5MiR+Oijj5CcnAyRSARvb28AQFVVFd5//3106NAB5ubm8Pf3x86dOzXG\nIxKJ8N1332H+/Pmwt7fHiy++iC1btsDU1BRSqRQ9e/aElZUVRo0ahYyMDERFRSEoKAg2NjYYO3as\nxmWw+/fvY9asWWjfvj2sra3Rq1cv7NixQ+P1goODsXTpUqxatQru7u5wcnLCSy+9hJKSEo3H7d69\nG3379oWlpSWcnZ0xadIk5OfnK+//7rvv0L17d1haWqJr16749NNPIZfLG/zsL126hOHDh8PKygqO\njo544YUXkJWVBQDYsmULPD09AfDB97NmLBqbt+PHjyM4OBhOTk6wt7dHcHAwrl69qtHHxo0b4efn\nB0tLSzg5OWHEiBFIS0uDVCrFiy++CADKjG99l4kB4NNPP4WPjw8sLCzg6uqKCRMmoLy8XOOz69Ch\nA6ytrTFhwgRs27ZN45Jm9fyrqyuLs3TpUvj6+sLKygo+Pj744IMPUFlZqbw/PDwcXbp0QUREBLp3\n7w5zc3MkJCSguLgYb731lnIMffr0wd69e+t9P/XNjVQqVY77wYMHGD58OAD+MqtIJMKoUaPq7bPm\n5VEvLy989NFHeO2112Bvb4927drhf//7H8rLy/HGG2/A0dERHTp0wPr16zX6EYlE+PbbbzF79mzY\n2NigQ4cO+Pbbb+t9XYD/XYaHh8Pb2xuWlpYICAjATz/9VKvfdevWYd68ebCxsYGXlxf27t2LvLw8\nhIaGwtbWFj4+PtizZ4/G8zIzM7Fo0SK4urrC1tYWQ4cOxdmzZ5X3V39mJ06cwPDhw2FtbQ1/f38c\nOXJE+ZhOnToBAEaOHKnx35PGft+LFi3CqVOnsHXrVuX3tPr7UvNSbHp6OkJCQuDg4AArKyuMHDkS\n165de6ZxAo1/10kbwAgxoJdeeomNHTtWo+2rr75iHMex4uJitnnzZubg4MB27NjB7t+/z86cOcN6\n9erFFi5cqHz8iBEjmEQiYa+99hq7e/cui4mJYbm5uey9995jnTt3ZpmZmSw7O5sxxth7773HnJyc\nWGRkJEtISGCffvopE4lE7OTJk8r+OI5jTk5ObP369Sw5OZklJCSwzZs3M5FIxEaOHMmuXLnCrl+/\nzrp06cKGDh3Khg8fzi5fvsxu3rzJunfvzubNm6fs6/bt22z9+vXsr7/+YsnJyey7775jYrGYRUVF\naYzf3t6e/fOf/2Tx8fHs2LFjzNHRka1YsUL5mJ9//pmZmpqyTz75RPke161bp3xfYWFhzNPTk+3b\nt489ePCAHT58mHXq1Emjj5rS09OZRCJhL7zwAouJiWHnzp1jvXr1YsOHD2eMMVZWVsauXr3KOI5j\nBw8eZJmZmayysrLeeRwzZozytjbztnfvXvbbb7+xe/fusTt37rAlS5YwR0dHlpOTwxhjLDo6monF\nYrZ9+3b28OFDdvv2bbZp0yaWmprKKisr2fr16xnHcSwzM5NlZmaywsLCOsf2+++/M1tbW3bo0CH2\n6NEjdvPmTfbNN9+wsrIyxhhj+/btY2KxmK1du5YlJCSwTZs2MVdXVyYSiVhaWpry/YjFYo1+Hz16\nxDiOY6dPn2aMMaZQKNgHH3zArly5wlJSUtiBAweYu7s7CwsLUz4nLCyMWVlZseDgYHblyhWWkJDA\nioqKWHBwMBs5ciQ7f/48u3//Pvvpp5+YmZmZxvdSXX1zExUVxTiOY2lpaUwul7MDBw4wjuNYdHQ0\ny8zMZHl5efV+H7y8vNjq1auVtz09PZm9vT1bu3YtS0pKYp988gkTiURs/PjxyrY1a9YwkUjE7ty5\no3wex3HM0dGRrVu3jiUkJLBvvvmGicVitn///npf66WXXmKBgYHs+PHj7MGDB2z37t3M3t6ebdq0\nSaPfdu3asW3btrGkpCT2+uuvM2trazZu3Di2detWlpSUxN58801mbW2t/A6VlpYyPz8/NmfOHHbt\n2jWWlJTEVq9ezczNzdndu3cZY0z5mQUGBrKjR4+yxMREtnjxYmZra6v8vG7cuME4jmN79+7V+O9J\nY7/vgoICNnz4cBYSEqL8nlb/hjiOY7/88ovyuzNgwADWu3dvdv78eXb79m02b9485uDgoHwtbcbZ\n2HedtA0U2BGDqhkQxMbGMm9vb/bcc88xxvg/Lj/++KPGc06fPs04jmP5+fmMMT4w6tatW62+w8LC\nmK+vr/J2SUkJMzc3Z//73/80Hjdz5kw2atQo5W2O49iSJUs0HrN582bGcRy7deuWsu3LL79kHMex\n69evK9vWrl3LnJ2dG3zP06dPZ0uXLlXeHjFiBAsKCtJ4zGuvvab8DBhjrGPHjuzNN9+ss7+SkhJm\nZWXFjh49qtG+detWZm9vX+84PvzwQ9axY0dWVVWlbLt16xbjOI6dOXOGMcbY/fv3Gcdx7Pz58w2+\np5rzqM281SSXy5mDg4Pyj92ePXuYnZ1dvQHb9u3bGcdxDY6LMca+/vpr1rVrV433qW7IkCFswYIF\nGm3vvfeeMkBiTLvArr7X7tKli/J2WFgYE4lE7NGjR8q2qKgoZmFhwQoKCjSeu3jxYjZjxox6+65r\nbtQDO8YYO3v2LOM4jqWkpNTbT7W6AruZM2cqbysUCmZra8umTZum0ebg4MDWrVunbOM4jr344osa\nfc+fP58NGzasztdKTk5mIpGIxcfHazxn5cqVGr8LjuPYO++8o7ydlZXFOI5j//jHP5RteXl5jOM4\n9scffzDG+Hnr0KEDk8lkGn2PHDmSvf3224wx1We2d+9e5f2ZmZmM4zh27Ngxxph2c12t5u97zJgx\nbPHixbUepx7YnThxgnEcpww2GWOsoqKCubu7s48//ljrcTb2XSdtA12KJQYnlUohkUhgZWWFnj17\nwtfXF7/88guysrLw8OFDvPPOO5BIJMp/Jk2aBI7jkJiYqOyjb9++jb5OYmIiKisrlZenqg0fPhyx\nsbEabQMGDKj1fI7j0LNnT+VtNzc3AECvXr002nJycpS1OKWlpXj//fcREBAAJycnSCQSHD58WKOY\nneM4BAYGaryWu7s7MjMzAQBPnjxBamoqxo0bV+f7io2NRVlZGWbNmqXxOb366qsoLCxETk5Ovc8b\nNGgQxGJVqW2vXr1gZ2eHO3fu1PkcbWg7b/fv38fChQvRpUsX2NnZwc7ODgUFBcrPZty4cfD29kbn\nzp0RGhqKDRs21PteGjJv3jxUVVXB09MTixcvxo4dO1BcXKy8/+7duxg8eLDGc4YMGdKk975hwwYM\nHDgQ7dq1g0QiwfLly2stXHBzc0OHDh2Ut69evYrKykq0b99e4/P65ZdfNL7jLa3m95LjOLi4uGh8\n3zmOg6urq/LyfbWaC5UGDx5c6zdWLTo6Gowx9O3bV+P9r1mzptb7Vx+Ps7MzTExMNMZjb28PMzMz\nPHnyBAD/2WZkZMDe3l6j73PnztXqOygoSPnvrq6uMDExUf4G66PN71sbsbGxcHJyQvfu3ZVtZmZm\nGDhwYK3PraFxNvZdJ20DLZ4gBjdo0CBs3boVYrEYHh4eykCj+j9W3377LUaOHFnree3btwfA/3Gx\ntrbW65jq6k8kEmms5Kv+dxMTk1ptjDFwHId//etfOHDgANauXYtu3brBysoK7777LgoKCjT6rrk4\ng+M4ZWF9Y6ofFxkZia5du9a638HBoc7ncRzXLAseqsfT2LxNmTIFrq6u+P7779GxY0eYmppi6NCh\nypo0a2trREdH4/z58zhx4gR++OEH/Pvf/8bJkyfRp08frcfj4eGBuLg4REVF4dSpU1i1ahX+85//\n4PLlyxoBVkPqWtxQVVWlcfu3337D3//+d3z++ecYMWIEbG1tERERgQ8++EDjcTW/WwqFAnZ2doiO\njq71GoZetFOzrpDjuDrbtP2u1qX6uRcvXoSVlVWtvhsaT31jrO5ToVDAz88P+/btq/W8mq9V12fd\n2PvS9vfdVNX/HdF2nPr4rhPho4wdMTgLCwt4e3ujU6dOGtkjNzc3dOzYEXFxcfD29q71j7m5+TO9\njq+vL8zNzXH69GmN9tOnT2tk4vTp7NmzWLBgAebMmYOePXuic+fOiI+Pf6atHlxdXdGhQwccPXq0\nzvv9/f1hYWGBpKSkOj+n+lZc+vv749KlSxoByq1bt1BQUICAgIBne6NqtJm3nJwc3L17F++//z7G\njh2rXEhQnWmpJhKJMGzYMKxcuRLXrl2Du7u7crFL9R84bYJTMzMzjB8/Hp9//jlu376N0tJS7N+/\nHwDQo0cPnD9/XuPxNW+7urpCLpdrjO/69esajzlz5gx69+6Nt99+G71794aPjw/u37/f6Nj69++P\n/Px8lJWV1fqsdP1jXP0ZNbaIRt8uXryocfvChQvw9/ev87HV2faUlJRa779z5846jaN///5ITk6G\nRCKp1Xe7du207qe+z1Gb37eZmRlkMlmD/fv7+yt/E9UqKipw+fLlZ/4tNvRdJ22D4DJ2mZmZmDVr\nFszMzGBmZoZff/2Vlum3YqtXr8Yrr7wCBwcHTJs2Daamprh79y6OHDmCH374AYD2W21YWVnhH//4\nB1asWKG8pBQZGYkDBw7gxIkTzTL+bt26Yd++fZg1axasra3x9ddfIz09XeOPijbjDwsLw2uvvQY3\nNzfMnj0bCoUCUVFRCA0NhZOTE5YvX47ly5eD4ziMHj0aMpkMt2/fxs2bN/HZZ5/V2eff//53fPPN\nN1i0aBGWL1+OvLw8vP766xg+fHiTL0VWa2zeHBwc4OLigp9++gne3t7Izs7Gv//9b1haWir72L9/\nP+7fv49hw4bBxcUF165dw6NHj9CjRw8AUP7R379/P4YMGQIrK6s6M62bNm0CYwz9+/eHvb09Tp48\niaKiImU/7777Lp5//nkMGDAAEydOxLlz57Bjxw6NP84DBw6ERCLB+++/j//3//4fkpKS8PHHH2u8\nTvfu3fHzzz/jwIED8Pf3x6FDhxpc2Vpt1KhRGDNmDGbNmoUvvvgCPXv2RF5eHi5cuABLS0ssWbLk\n2SfgKU9PT4hEIvzxxx+YO3cuzM3NYWdnV+dja34H6/pOatv2xx9/YP369Rg3bhyOHDmCiIgIREZG\n1vkcX19fvPzyy1i6dCm++OILDBo0CCUlJbh27Zrye9FUL7zwAtauXYvJkydj9erV6NKlCzIzM3Hq\n1Cn06NED06dP16ofZ2dn2NjY4OjRo/Dz84O5uTkcHBy0+n137twZUVFRSE5Ohq2tLezt7TX+BxYA\nRo8ejQEDBmD+/PlYv349bG1tsWrVKlRWVuK1117T+v029l0nbYPgMnYuLi44f/48oqKiMH/+fGzY\nsMHQQyI6aGyj0gULFiAiIgKHDh3CwIEDMWDAAKxcuVIjk1FfH3W1r169GkuXLsXbb7+Nnj174tdf\nf8Uvv/xS5yXDuvp71ra1a9cqt18ZM2YMOnbsiDlz5tS6pFuzn5ptr7zyCrZs2YLIyEj07t0bI0aM\nwNGjR5V/ID788EN8/fXX2LBhA4KCgjBs2DB88803DWY8XF1dcezYMaSmpqJ///6YOnWqMtht7D3W\n9Z7VH9fYvIlEIvz2229ISkpCr1698PLLL+Odd97R2I/N0dERBw8exMSJE9GtWze8//77WLFihXKz\n1/79++Ott97CsmXL4ObmhjfffLPOsTk6OmLz5s0YOXIkevTogf/+97/YsGGDcs5nzJiBr776Cl98\n8QUCAwOxc+dOfP755xrBh4ODA3bu3IlLly4hMDAQq1evxpdffqnxnpctW4aFCxdi8eLF6NOnD65e\nvYrw8PBG5xoADhw4gFmzZuGdd96Bn58fpkyZgj///BO+vr6Nfu4Ntbm5uWHNmjX47LPP4OHhgZkz\nZ2rdV1O+79U++ugjnDhxAkFBQfjss8/w5ZdfagRRNZ/z008/4Z133sHq1avh7++PMWPGYPv27fDx\n8al3vNqoztD369cPixcvRrdu3TB79mxER0fDy8urwfegTiQSYf369YiIiEDHjh2VWUZtft/vvvsu\nnJ2dERgYCFdXV1y4cKHO19i3bx+6d++OyZMnY8CAAXjy5AmOHz8OR0dHrcfZ2HedtA0ca44imxby\n3XffwczMDMuWLTP0UAghrYhUKsWoUaOQmpoKDw8PQw9HUEQiEXbs2IH58+cbeiiEtEmCuxQL8HVA\nf/vb35Cfn19rQ1NCCCGEkLbKYJdi161bh379+sHCwqLWOXq5ubmYOXOmcnfxmicDBAYG4vLly/jk\nk0+watWqlhw2IaSNENJZpoQQUs1gGbv27dtjxYoVOHr0KMrKyjTue+ONN2BhYYEnT57gxo0bmDx5\nMgIDA9GjRw9UVVUpl7fb2toqj50ihBB9CQ4ObvGVpK2FLlufEEJ0Z/AauxUrViA1NRWbN28GAJSU\nlMDR0RGxsbHKwuGXXnoJHh4eWLNmDa5cuYJ//etfMDExgampKTZt2lTnlgDt27fXOLOTEEIIIcRY\n+fj46GVTcoOviq0ZV967dw9isVhjNVhgYKBy9+0BAwbg9OnTOHXqFI4ePVrvPk+PHz9WbiNhyH/C\nwsIM3tezPE+bxzb0mKbcV1e7Pj83Y5i71jp/xjp3Qpw/XeeuofuF9tvT9zjaym/PWOZPaL89bR7b\nEr+9pKQkvcRVBg/sataxFBcXw9bWVqNNIpGgqKioJYelN8HBwQbv61mep81jG3pMU+6rq/3BgweN\njqO56XPudOnPmOfPWOcOEN786Tp3Dd0vtN8eQP/tbOy++h5vDPMntN+eNo9tid+evhj8UuyHH36I\ntLQ05aXYGzduYOjQoSgpKVE+5v/+7/9w5swZHDhwQOt+m+u4JNIyFi1ahC1bthh6GKQJaO6EjeZP\n2Gj+hEtfcYvRZey6du0KmUymcZ351q1bTTriKDw8HFKpVNchEgNYtGiRoYdAmojmTtho/oSN5k94\npFIpwsPD9dafwTJ2crkcVVVVWLlyJdLS0rBhwwaIxWKYmJggNDQUHMdh48aNuH79OqZMmYKLFy/C\nz89P6/4pY0cIIYQQoRB8xm7VqlWwsrLC559/jh07dsDS0hKrV68GAHz//fcoKyuDq6srFixYgB9+\n+OGZgjoifJRpFS6aO2Gj+RM2mj9isH3swsPD6009Ojg4aHV4NiGEEEIIUTH44onmwnEcwsLCEBwc\n3KyrTwghhBBCmkoqlUIqlWLlypV6uRTbqgO7VvrWCCGEENLKCL7GjpCGUJ2IcNHcCRvNn7DR/BEK\n7AghhBBCWolWfSmWauwIIYQQYsyoxk5LVGNHCCGEEKGgGjvSqlGdiHDR3AkbzZ+w0fwRCuwIIYQQ\nQloJuhRLCCGEEGJgdClWC+Hh4ZSWJoQQQojRkkql9Z7E1RSUsSNGSSqV0mpmgaK5EzaaP2Gj+RMu\nytgRQgghhBANlLEjhBBCCDEwytgRQgghhBANFNgRo0SLXoSL5k7YaP6EjeaPtOrAjlbFEkIIIcSY\n0apYLVGNHSGEEEKEgmrsCCGEEEKIBgrsiFGiS+jCRXMnbDR/wkbzRyiwI4QQQghpJajGjhBCCCHE\nwPQVt4j1MBZCCCGEENIE8fEpOHEiSW/9tepLsbTdiXDRvAkXzZ2w0fwJG82fsMTHp2Dlykhs3XpG\nb3226oydPveFIYQQQgjRp8OHkyCXvwsrKwBYqZc+qcaOEEIIIaSF3bsHvPeeFIWFwQCA06epxo4Q\nQgghRFDKyoAjR4BbtwCZTKH3/lt1jR0RLqoTES6aO2Gj+RM2mj/jFh8PfP89H9QBgLe3D0SikwgI\n0N9rUMaOEEIIIaQZqWfp1I0c6YklS4ALF07p7bWoxo4QQgghpJnExwOHDgFFRao2GxtgyhSge3dV\nG+1jRwghhBBipOrL0vXsCUyciKcrYfWPauyIUaI6EeGiuRM2mj9ho/kzDvHxwPr1mkGdjQ0QEgLM\nnt18QR3QyjN24eHhCA4ORnBwsKGHQgghhJBWrqwM+PNP4K+/NNt79QImTKg7oJNKpXoNyKnGjhBC\nCCFER/HxwMGDQHGxqq2uWrr6UI0dIYQQQoiBNZSlmzgRsLRs2fFQjR0xSlQnIlw0d8JG8ydsNH8t\nq7qWTj2oq66lmzWr5YM6gDJ2hBBCCCHPxNiydOqoxo4QQgghREv11dJNnQp069b0fqnGjhBCCCGk\nhRhzlk4d1dgRo0R1IsJFcydsNH/CRvPXPOLi6q6lCw01XC1dfShjRwghhBBSB6Fk6dRRjR0hhBBC\nSA1xcfwZr/qupasP1dgRQgghhOhZaSl/xquQsnTqqMaOGCWqExEumjtho/kTNpo/3cTFAd9/L4xa\nuvpQxo4QQgghbVppKV9Ld/u2ZntgIH/GqxACumqtusYuLCwMwcHBCA4ONvRwCCGEEGKEWrqWriap\nVAqpVIqVK1fqpcauVQd2rfStEUIIIURHxpal01fcQjV2xChRnYhw0dwJG82fsNH8aae6lk49qJNI\n+Fq6mTOFdem1JqqxI4QQQkibYGxZuuZAl2IJIYQQ0urVVUsnkQBTprRMLV1jaB87QgghhJBGtIUs\nnTqqsSNGiepEhIvmTtho/oSN5k/T3bv8Ga81a+nmzxd+LV19KGNHCCGEkFalrWXp1FGNHSGEEEJa\njbt3+Vq6khJVm0TC70vXtavhxtUYqrEjhBBCCHmqtBQ4fBiIidFsDwoCxo9v3Vk6dVRjR4wS1YkI\nF82dsNH8CVtbnb/qWjr1oK66lm7GjLYT1AGUsSOEEEKIQFGWrjaqsSOEEEKI4Ai1lq4+VGNHCCGE\nkDaHsnQNoxo7YpTaap1Ia0BzJ2w0f8LW2uePaukaRxk7QgghhBg1ytJpT3A1dleuXMHbb78NU1NT\ntG/fHtu2bYNYXDs+pRo7QgghRPju3AH++KN2Ld20aUCXLoYbl77pK24RXGCXkZEBBwcHmJubY/ny\n5ejbty9mz55d63EU2BFCCCHC1VCWbsIEwMLCMONqLvqKWwRXY9euXTuYm5sDAExNTWFiYmLgEZHm\n0NrrRFozmjtho/kTttYyf3fu1F1L98ILfC1dawvq9EmwNXYpKSk4fvw4PvroI0MPhRBCCCF6UFLC\nn/FaM0vXuzdfS9caA7qU+HgknTiht/4MlrFbt24d+vXrBwsLCyxevFjjvtzcXMycORM2Njbw8vLC\nzp07Ne4vLCzEiy++iK1bt1LGrpUKDg429BBIE9HcCRvNn7AJef7u3AG+/14zqLO15bN006e33qAu\nccsWjIqN1VufBsvYtW/fHitWrMDRo0dRVlamcd8bb7wBCwsLPHnyBDdu3MDkyZMRGBiIHj16QCaT\nISQkBGFhYejSmqomCSGEkDaopISvpasZ27TmLF21pGPHMDotDXj4UG99GixjN3PmTEyfPh1OTk4a\n7SUlJdizZw9WrVoFKysrDBkyBNOnT8f27dsBADt37sSVK1ewatUqjBw5EhEREYYYPmlmraVOpC2i\nuRM2mj9hE9r8VWfp1IO61p6lUyovh+jyZb0GdYAR1NjVXAFy7949iMVi+Pr6KtsCAwOVX9aFCxdi\n4cKFWvW9aNEieHl5AQDs7e0RFBSkTFNX90e3jfP2zZs3jWo8dJtu0226Tbf1d7usDCguDkZsLPDg\nAX+/l1cwevcGLC2lSEsDunQxnvHq/XZBAXDqFDZfvowtRUXQJ4Nvd7JixQqkpqZi8+bNAICzZ89i\n7ty5SE9PVz5mw4YN+PXXXxEVFaV1v7TdCSGEEGJ86tqXztaWP+O1TVRYJSQAkZFARQVSsrORePMm\nRnt7g9u+vXWcFVvzTdjY2KCwsFCjraCgABKJpCWHRQghhBA9asu1dAAAxoDz54GTJ/l/B+Dp7g6M\nG4dTqanA05IzXYn00osOOI7TuN21a1fIZDIkJiYq227duoWAgIBn7js8PFyZAiXCQvMmXDR3wkbz\nJ2zGOn/11dItWNAGaukAoKoK+P134MQJZVAHOzvg5Zdx39YWZ5480dtLGSxjJ5fLUVVVBZlMBrlc\njoqKCojFYlhbW2PWrFn46KOPsHHjRly/fh0HDx7ExYsXn/k1wsPD9T9wQgghhGilzWfpAL6ebtcu\nQK3EDJ6ewNy5gLU1gt3dERwcjJUrV+rl5QxWYxceHo6PP/64VttHH32EvLw8vPzyyzh+/DicnZ3x\n2WefISRAG3DXAAAgAElEQVQk5Jn6pxo7QgghxHBiY/lautJSVZutLX/Gq9r6yNYtJQWIiNAsKOzf\nnz8TrcY+vG32rFhtUWBHCCGEtDzK0j0VHc1/EAoFf1skAiZNAvr1q/Phbfas2GdBNXbCRfMmXDR3\nwkbzJ2yGnr/YWP6M1zZbSwcAcjlw6BD/T3VQZ20NLFpUZ1AnlUr1Wjpm8FWxzYlq7AghhJDmV1LC\nX3a9c0ezvU8fYNy4NhLQAUBxMX/pVX3TYXd3ICSEXyxRh+Dg4NZRY9fc6FIsIYQQ0vyolu6px4/5\nRRLqW7b17Ml/EKamjT5dX3FLq87YEUIIIaR5UJZOze3bwP79gEzG3+Y4YMwYYPBg/t9bENXYEaNE\n8yZcNHfCRvMnbC01f9W1dOpBXXUt3bRpbSioUyiA48f5PeqqgzoLC/6w2yFDtArqqMbuGVCNHSGE\nEKI/lKVTU1bGB3RqByrA2RkIDQWcnLTuhmrstEQ1doQQQoj+UC2dmqwsvp4uJ0fV1rUrMHs2YG7e\npC6pxo4QQgghzY6ydDXExwN79gAVFaq24cOBkSObVE8XnxiPE9dO6G14rbrGjggX1fkIF82dsNH8\nCZs+54+xumvp7OyAhQvbWC0dwH8gZ8/ymbrqoM7UFHj+eWDUqCYHdVuitiDdJb3xB2upVWfswsPD\nldeuCSGEEKIdytLVUFnJr3pV33nZ3p7fn65duyZ3e+LaCdwvuI/oddF6GCSPauwIIYQQAkCVpTt8\nWLOWzs6Oz9D5+BhubAaTnw/s3AlkZqravLz4TJ21tU5df7DxA1wyvQQ5k+P04tNUY0cIIYQQ/Sgu\n5gO6mlm6vn2BsWPbYJYOAB484E+SUI9yBwzgD701MWlyt4wxnH90HjczbkLeQa77ONVQjR0xSlTn\nI1w0d8JG8ydsTZk/xoCYGOD77+uupZs6tQ0GdYwBV64A27apgjoTEz5tOWmSTkFdlbwKe+7uwYnk\nE+js3RmyRBnMTZq2krYulLEjhBBC2qjiYr6W7u5dzfa+fflauibu3CFsMhmfurx+XdVmYwPMmwd0\n7KhT1wXlBdgVswvpxfxiCWcPZ0ySTIJlkSWO4ZhOfVejGjtCCCGkjaFaunoUFfGXXh89UrV5ePCL\nJGxtder6YcFD7I7ZjZKqEmVbP49+mOg7ESYiE9rHThu0KpYQQgjRRFm6eqSlAbt3A4WFqrbAQGDK\nFH5bEx1ce3wNhxMOQ874ejoRJ8KkLpPQz6MfpFKpXksg6s3YLVy4UKsOzM3NsXHjRr0NSF8oYyds\nUqmUAnKBorkTNpo/YWto/ihL14Bbt4CDB1XnvXIcH+UOGtSk/emqyRVyHE06iitpV5RtVqZWmOs/\nF172XhqPbfaMXUREBJYvX17vi1QP4KuvvjLKwI4QQgghPMrS1UOhAI4fBy5eVLVZWgJz5ugc6ZZW\nlSIiNgIP8h8o29rZtENIQAjsLex16rsh9WbsfHx8kJSU1GgH3bp1Q3x8vN4HpivK2BFCCGnrKEvX\ngLIyIDISUI91XFyA0FDA0VGnrjOKM7ArZhfyy/OVbf4u/pjefTrMTMzqfI6+4hZaPEEIIYS0QpSl\na8CTJ/zRYLm5qrbu3YGZM3X+YO5k3cHeu3tRpahSto3qPArDOg0D18BlXX3FLU3axy45ORkPHjzQ\n+cUJqQ/tpSVcNHfCRvMnbFKpFIwBt2/zZ7yqB3V2dsCLL/L70rXpoC4uDti4UTOoGzGC385Ehw+G\nMYao+1GIiI1QBnXmJuYIDQjFcM/hDQZ1+qTVqtiQkBD84x//wODBg7F582a8/vrr4DgO3377LZYs\nWdLcYySEEEJIA+LjU3DiRBJu3foL27YpYGXlA2dnT+X9lKUDf136zBkgKkrVZmbGZ+n8/HTqukJW\ngb1xexGXHadsc7R0RGhAKFysXRp8bnxyMk6on0GrI60uxbq4uCAtLQ1mZmYICAjAjz/+CHt7e0yf\nPh2JiYl6G4w+cRyHsLAw2u6EEEJIqxYfn4ItWxKRnz8aCQn8wk6Z7CSCgnzh4+OJ6dMBb29Dj9LA\nKiuBvXs1U5gODvz+dG5uOnWdW5aLXTG78KTkibLNx8EHc3rMgaWpZYPPjU9Oxsrt25GQnY3odeta\nrsbO3t4e+fn5SEtLw4ABA5CWlgYAkEgkKCoq0nkQzYFq7AghhLQFX399ChcujEJ2tma7n98p/Pe/\no9p2lg4A8vKAnTv5urpqnTsDzz8PWFnp1HVyXjJ+i/0NZbIyZdtzHZ7DWJ+xEHGNV7t9e+AALvr4\nIL2yEqf79Gm5GrvAwECsWbMGH3/8MSZPngwASE1NhZ2dnc4DIKQuVOcjXDR3wkbzJyyxsUBUlEgZ\n1OXnS2Fuzu+r262biIK65GTgp580g7pBg/hDcHUI6hhjuJR6CTv+2qEM6sQiMWZ0n4HxvuO1Cuqe\nVFbidGEh0isrmzyOumhVY7dp0yasWLECZmZm+OKLLwAAFy9exAsvvKDXwRBCCCGkcaWl/BYmMTGA\nXK5Qtjs5Af37A2IxYGamaKCHVo4x4PJl4Ngxfq86ADAx4VeOBAXp1LVMIcMf9/7AjYwbyjaJmQTz\nAuahg20HLYbGcL24GH/m5KCsekNkPaLtTgghhBABiYvjD0koeXrkaHZ2CuLiEtGjx2jl9msVFSex\naJEvunXzrL+j1komAw4dAm7eVLVJJPyq1w6NB14NKaoowu7Y3UgtTFW2tZe0R0hACCTmkkafXy6X\n42BODmKfTl72o0f4KyEBfkOH4qfu3Vt2H7uzZ8/ixo0bKCoqUgZNHMdh+fLlOg+iOVBgRwghpDUp\nKwP+/BP46y/N9t69gc6dU3DuXBIqK0UwM1Ng9GifthnUFRXx572mqgIvdOjAB3WSxgOvhqQVpmFX\nzC4UVarWFgS6BWJqt6kQixq/AJpaXo7IrCzkq2XpXM3MEFRcjJv37uGNadNaLrB78803ERERgWHD\nhsHSUnOFx/bt23UeRHOgwE7Y6LxK4aK5EzaaP+N07x6fpVNfryiR8KdHdOmiamvT85eaygd16h9S\nUBAwZQp/bVoHf2X+hQPxByBT8EEZBw7jfcdjYPuBje5PxxjDhcJCnMzLg0ItLuknkWC8oyNMRXw9\nXrOfFatux44diI2NhYeHh84vSAghhBDtlJcDR48CN25otgcGAhMm8MeaEvAf0KFDgFzO3xaJgPHj\ngQEDAB02BlYwBU4kn8CFRxeUbZZiS8zpMQc+jo2fx1Ysk2FvdjaSylSrZi1EIkxzdkYPa+smj6sh\nWmXsevXqhVOnTsHZ2blZBtEcKGNHCCFEyJKSgP37gcJCVZu1NV//37274cZlVBQKfoHEpUuqNktL\nfisTHTfvK6sqQ+SdSCTlqc6SdbFyQWjPUDhaNn6WbFJZGfZmZaG4OtgE0MHcHHNcXGBvalrr8S16\nVuzVq1fx6aefYv78+XCrsZHf8OHDdR5Ec6DAjhBCiBBVVADHjwPR0ZrtAQHApEk6b73WepSWAr/9\nBty/r2pzdQVCQ/nNh3WQVZKFXTG7kFOWo2zr5tQNs/xmwVzc8B4ycsYQlZeH84WFGnHIUDs7jHRw\ngEk9GcQWvRR77do1HD58GGfPnq1VY/fo0SOdB9FcwsPD6eQJgWrTdSICR3MnbDR/hnX/Pp+ly89X\ntVlZAZMnA/7+jT+/zcxfZiawaxe/+XA1Pz/+eDAzM526vpdzD7/f+R0V8gpl23DP4RjpNbLRerr8\nqir8np2NR+XlyjZrExPMcnGBTz3XzaVSqV73j9QqsPvggw9w6NAhjB07Vm8v3BLCw8MNPQRCCCGk\nUZWVwIkTwJUrmu1+fnztfzOVYwnTnTvAvn38h1Zt5Ehg+HCd6ukYYzj38BxO3T8FBj5zZioyxYzu\nM+Dv2nhUfaekBAeys1GuUO0f6GNpiZnOzrBpYPFGdQJq5cqVTR67Oq0uxXbq1AmJiYkw0zEKbkl0\nKZYQQogQPHzIxym5uao2S0v+smtAgE6xSuvCGCCVAqdPq9rMzIBZs3QuOqySV2F//H7EPIlRttmZ\n2yG0Zyja2bRr+LkKBY7m5iJabTWuiOMwyt4eQ+zsGs3yVWvRGrstW7bgypUrWLFiRa0aO5FIq1PJ\nWhwFdoQQQoxZVRVw6hRf96/+56prV36BhI7brrUuFRXAnj1AfLyqzdERCAnh6+p0UFBegF0xu5Be\nnK5s87TzxFz/ubA2azhVmlVZicisLGSqZQ/txWLMdnFBRwuLZxpHiwZ29QVvHMdBrrbaw5hQYCds\nbaZOpBWiuRM2mr+WkZoK7N0L5Khq82FhwW9hEhjY9Cxdq5y/3Fxg504gK0vV5uMDzJmj834vKfkp\niIiNQElVibKtv0d/TPCdABORSb3PY4zhRnEx/szNRZXapdce1taY5uQEC5P6n1ufFl08kZycrPML\nEUIIIW2dTMZfTTx/XjNL5+vLbzZsa2uwoRmnpCR+5avaYgQMHgyMGcPvVaeD6MfROJxwGArGB2Yi\nToRJXSahn0e/Bp9XoVDgYHY2YkpUwaCY4zDB0RF9JRKtL702FzorlhBCCGkBjx/zWTr1xJO5Ob+P\nbu/eVEungTH+GvWxY6oIWCzmr1EHBurUtVwhx5HEI7j6+KqyzdrUGnP958LTvuFj2NIqKhCZlYW8\nqiplm4uZGea4uMBNx3UI+opb6g13V6xYoVUHYWFhOg+CEEIIaa3kcr6WbuNGzaCuc2fgtdeAPn0o\nqNNQVcWvJjl6VBXU2doCixfrHNSVVJZg261tGkGdu407/tb3bw0GdYwxXCgowKb0dI2gro9Egr+5\nu+sc1OlTvRk7Gxsb/FXzpOEaGGPo27cv8tU33DESlLETtlZZJ9JG0NwJG82ffmVk8Fm6zExVm6kp\nMG4c0K+f/gM6wc9fYSG/P93jx6q2jh2BefMAGxudus4ozsCumF3IL1fFLAGuAZjebTpMTWqfBFGt\nRC7HvuxsJJSWKtvMRSJMdXJCgI5jUtfsNXalpaXw9fVttANz84Z3YCaEEELaGrkcOHeO35lDrbYe\nnp7AjBk6H4zQOj16BOzeDRQXq9r69OH3fWlgHzhtxD6Jxb64fahS8Nk2DhxGdR6FoZ2GNlgTd7+s\nDHuys1Ekkynb2j89FsyhjmPBjAHV2BFCCCF69OQJfyVRPelkagqMHg0MHEiXXet0/Trwxx98RAzw\nCyMmTAD699d50+GoB1E4k3JG2WZuYo7ZPWajq1PXep+nYAzS/HycLSjQiCUG29lhdAPHgumiRVfF\nEkIIIaRhCgVw4QIQFaWKTwD+SuKMGYCTk+HGZrTkcr6WTv3IDSsrYO5cwMtLp64rZBXYc3cP4nNU\ne985WjoiNCAULtYu9T6vQCbD71lZeFjjWLAZzs7oIoCDeo1zd2HS5unz3DzSsmjuhI3mr2mys4Gf\nf+aPBasO6sRivpZu8eKWC+oENX8lJcD27ZpBnZsb8Le/6RzU5ZblYuP1jRpBnY+DD5b2WdpgUBdX\nUoIfHj/WCOo6W1riVQ8PQQR1QCvP2IWHhyvPYCOEEEL0TaHgd+U4dYrfo65a+/Z8ls6l/hiibcvI\n4BdJqC++9PcHpk/njwnTQVJuEiLvRKJMVqZsG9xxMMZ4j4GIqzufJVMocCwvD1cKC5VtIo5DsL09\nhtrZQdSM18+lUqleA3KqsSOEEEKaICcH2L+fP+u1mokJEBwMDBmi8/65rVdsLF+EWL1tCMcBo0YB\nQ4fqXE93KfUSjiUdAwP/918sEmNq16kIbFf/NinZT48Fy1A7Fszu6bFgnZ7xWDBdtGiN3ZMnT2Bp\naQmJRAKZTIZt27bBxMQECxcuNNqzYgkhhJDmwBh/9fDECVVsAgDu7nyWrsaR6qQaY3xq8+xZVZu5\nOTB7Nn9Arg5kChkO3TuEmxk3lW0SMwlCAkLQ3rZ9PcNhuFVcjMO5uahUW7rs9/RYMMsmHAtmDLSK\nyqZMmYLExEQAwAcffICvvvoKa9euxT//+c9mHRxpuwRVJ0I00NwJG81fw/LygK1bgT//VAV1IhGf\npVuyxPBBndHOX3k5f96relDn5MR/aDoGdUUVRdhyc4tGUNfBtgP+1vdv9QZ1FQoF9mZnY192tjKo\nE3McJjs5Ya6Li2CDOkDLjF1CQgKCgoIAADt27MCFCxcgkUjQo0cP/Pe//23WARJCCCGGxhhw7Rp/\nwpXaFTu4ufFZOnd3w43N6OXk8EFddraqzdcXmDMH0PFSZ2phKnbH7EZRZZGyLahdEKZ0nQKxqO4Q\nJ72iAr9lZSFXLd3qbGqKOS4uaNcK9ubVqsbO2dkZqampSEhIQEhICGJjYyGXy2FnZ4di9Y0EjQjV\n2BFCCNGHggLgwAH+PPpqIhFfEjZ8uM5757ZuiYlAZCSfsas2ZAi/qZ+OpVy3Mm7h4L2DkCn4VSsi\nToRxPuMwsP3AOjcdZozhcmEhjuflQa4WH/SWSDDR0RFmBi4ta9EauwkTJmDu3LnIycnBvHnzAAB3\n7txBhw4ddB4AIYQQYowYA27eBI4cASoqVO0uLnyWrn3dV/kIwH94Fy7whYjVwYpYzK967dlTp64V\nTIHjScdxMfWiss1SbInn/Z+Ht4N3nc8pfXos2D21Y8HMRCJMcXJCLz0eC2YMtMrYlZeXY+vWrTAz\nM8PChQshFoshlUqRkZGBkJCQlhjnM6OMnbAJ/rzDNozmTtho/nhFRXyWLiFB1cZxwODBwMiRxpul\nM4r5q6riP7zbt1VttrZASAjg4aFT12VVZYi8E4mkPFX61NXaFSEBIXC0dKzzOQ/KyvB7jWPB3J8e\nC+ZkRMeCtWjGzsLCAsuWLdNoM/gXhxBCCNEzxvh45PBhzauHTk58lq5jR8ONTRAKCvj96dLTVW2d\nOvEnSeiYGcsqycLOmJ3ILctVtnV37o6Z3WfCXFy7Nk7BGM7k5+N0jWPBnrOzw2h7e4hb6a4e9Wbs\nFi5cqPnAp9erGWMa1663bdvWjMNrOsrYEUIIeRbFxcChQ0BcnGb7oEF8SZgRJXeM08OHwO7d/IkS\n1fr1AyZO5Df400F8djz23N2DCrnqmvgIzxEI9gqus56u8OmxYClq0bnV02PBuhrpCRLNnrHz8fFR\nfljZ2dnYunUrpk6dCk9PT6SkpODQoUN46aWXdB4AIYQQYmgxMXyWTq0ECw4OfEmYjqdbtQ3R0fwe\nMNXnqYlEwKRJfGCnA8YYzj48i6j7UcpNh01FppjpNxM9XHrU+Zz40lLsy85GmdqBvV4WFpjl4gJb\nY72Grkda1diNGzcOK1aswLBhw5Rt586dw8cff4xjx4416wCbijJ2wmYUdSKkSWjuhK2tzV9JCR/Q\nxcZqtvfvD4wdq/PpVi2uxedPLucDuuhoVZu1NX/p1dNTp64r5ZXYH7cfsVmqybG3sEdIQAja2bSr\n9XiZQoETeXm4pHYsGPf0WLBhzXwsmD60aI3dpUuXMGjQII22gQMH4uLFi/U8gxBCCDFud+/yl17V\nrxza2fFZOu+6F1cSdSUlQEQEkJKianN35xdJ2Nnp1HV+eT52xexCRnGGss3L3gtz/efCyrT2pdSc\nqipEZmUhXW35su3TY8E8W/BYMGOgVcZuxIgR6N+/P1atWgVLS0uUlpYiLCwMly9fxpkzZ1pinM+M\nMnaEEELqUlbGZ+nUF20CQJ8+wPjx/ClXpBHp6fwiiYICVVtAAB8V61iMmJKfgt2xu1FapbouPqD9\nAIz3GQ8TUe1avb+Ki3EoJ0fjWLBuVlaY7uwMKwGdINGiGbstW7Zg/vz5sLW1hYODA/Ly8tCvXz/8\n+uuvOg+AEEIIaSn37vE7cajvrW9rC0ybxh+GQLQQEwPs3686U43j+NUlQ4bw/66D6MfROJxwGArG\nB2kmnAkmdZmEvh59az22UqHA4Zwc3FSbTBOOwzhHRwyQSOpcVNEWaJWxq/bw4UM8fvwY7u7u8NTx\n2nlzo4ydsLW1Op/WhOZO2Frr/JWX8xsN37yp2R4UBEyYoPPJVkajWedPoQBOnQLOnVO1WVgAs2cD\nXbro1LVcIcefiX8i+rGqVs/a1BrzAuahk12nWo/PeHosWI7asWBOT48FcxdoyrVFM3bVLCws4Orq\nCrlcjuTkZACAdwsXIhQWFmLMmDG4e/cuLl++jB496l4VQwghhAD8qVYHDgBqNfWwsQGmTgW6dTPc\nuASlvBz4/XfNHZudnfl6OmdnnbouqSxBRGwEUgpUtXruNu4ICQiBnYVmrR5jDFeLinA0N1fjWLBA\nGxtMcnKCeSvdm+5ZaJWxO3LkCF555RWkq284CD66lKstJ24JMpkM+fn5+Ne//oX33nsP/v7+dT6O\nMnaEENK2VVQAx44B165ptvfsyW+tZqTbmRmf7Gxg504gJ0fV1rUrMGuWzqnO9KJ07IrZhYIKVa1e\ngGsApnebDlMTzVq9Mrkc+7OzEVfjWLDJTk4IbAXHgrVoxu7111/HihUr8OKLL8LKwL8EsVgMZx3/\n74AQQkjrlpzMl4Gp1/ZbWwOTJwN0oecZ3LvHZ+rUD8sdNow/V03H7FjMkxjsj9uPKgV/OZUDh9He\nozGk45Ba9XEPy8vxe1YWCtSOBWtnZoY5Li5wFtqeNM1Mq1nJz8/HsmXLDB7UkbZDKpUaegikiWju\nhE3o81dZCfzxB7Btm2ZQ16MH8PrrrT+o09v8MQacPctn6qqDOlNTYM4cfqGEDkEdYwwnk08i8k6k\nMqgzNzFHaM9QDO00VCOoqz4WbHNGhkZQN9DWFkvc3Smoq4NWM/PKK6/g559/1usLr1u3Dv369YOF\nhQUWL16scV9ubi5mzpwJGxsbeHl5YefOnXX20VZXvBBCCKktJQX43/+Aq1dVbZaWfCzy/PN8xo5o\nobKSz9KdPMkHeAC/L90rr/BbmuigQlaBXTG7cPbhWWWbk6UTlvZdiq5OXTUeWySTYXtmJk7l5Skv\nUVqamCDE1RUTnZxa7VmvutKqxm7o0KG4cuUKPD090a6dardnjuOavI/d3r17IRKJcPToUZSVlWHz\n5s3K+0JDQwEAmzZtwo0bNzB58mRcuHBBY6HE4sWLqcaOEEIIqqr4GOTyZVUcAvALI6ZO1fns+bYl\nP5/fny5DtTEwPD35kyR0jIxzSnOwK2YXskqzlG2+jr6Y02MOLMSatXoJpaXYm52NUrU6/k4WFpjt\n4gK7VnosWIvW2C1ZsgRLliypcxBNNXPmTABAdHQ0UlNTle0lJSXYs2cPYmNjYWVlhSFDhmD69OnY\nvn071qxZAwCYNGkSbt26hfj4eCxbtqzeM2sXLVoEr6eH/Nnb2yMoKEi5DLw6XU236Tbdptt0W7i3\nHz0CvvxSisJCwMuLv//xYykGDABCQoLBccY1XqO+7eUFRERAeueO6nb//pBaWABXr+rUf1phGh45\nPkK5rBwPbj4AALww9QWM9h6NM6fPKB8vZwxfHzyI2JISeD098erB5csItLbGoilTIOI44/m8dLxd\n/e8PHjyAPj3TPnbN4cMPP0RaWpoyY3fjxg0MHToUJWpnvHz99deQSqU4cOCA1v1Sxk7YpFKp8kdA\nhIXmTtiEMn8yGRAVBVy4oJml69KFz9LZ2hpubIbUpPljjD/r9c8/+b3qAMDEBJg0Cehbe2PgZ+ua\n4VLqJRxLOgYGfqLEIjGmdZuGXm69NB6b+/RYsMdqCzUkYjFmOTujs6WlTuMQghbN2DHGsHnzZmzf\nvh1paWno0KEDFixYgMWLF+tc51bz+cXFxbCt8YuUSCQoKirS6XUIIYS0DmlpwL59QJbqih7MzfmN\nhoOCdD78oG2Ry/nz1dT3hLGx4S+9dqq9MfCzkClkOBh/ELcybynbbM1tERIQAg+Jh8Zjbz89FqxC\n7ViwLlZWmOHsDGsBHQtmDLQK7D799FNs27YN7777Ljp16oSHDx/iyy+/xOPHj/Hhhx/qNICa0amN\njQ0K1XeRBFBQUACJRKLT6xBhEULGgNSN5k7YjHn+ZDLg9Gng/HlVYgkAvL35I0p1PHe+VXim+Ssu\nBnbvBh49UrV5eADz5un8YRZWFGJ3zG6kFaUp2zradsS8gHmwMVMVPVYqFPgzNxc31JI3JhyHMQ4O\nGGRrS4skm0CrwG7Dhg04ffq0xjFi48ePx7Bhw3QO7GpOWteuXSGTyZCYmAjfpwf33bp1CwFNWIkT\nHh6O4OBgo/4PFSGEkMalp/NZusxMVZuZGTBuHH+1kP7+P6PHj/lFEuqJlF69+OvYpqb1P08LqYWp\n2B2zG0WVqmCtd7vemNx1MsQiVdiRWVmJyKwsZFVWKtscnx4L5iHQY8GaQiqVatTd6UqrGjtXV1fc\nv38f1morYoqLi+Ht7Y0nT5406YXlcjmqqqqwcuVKpKWlYcOGDRCLxTAxMUFoaCg4jsPGjRtx/fp1\nTJkyBRcvXoSfn5/2b4xq7ARNKHU+pDaaO2EztvmTy/nt1M6c0czSeXnxWToHB4MNzShpNX9//cWf\nsVa9LxzHAWPHAs89p3OEfDPjJg7GH4Sc8atZRZwI433GY0D7AcpEDmMM0U+PBZOp/Z3uaWODKW34\nWDB9xS1afXoTJkzAggULEBcXh7KyMty9excvvvgixo8f3+QXXrVqFaysrPD5559jx44dsLS0xOrV\nqwEA33//PcrKyuDq6ooFCxbghx9+eKagjhBCiPBlZgIbNwJSqSqoMzXljwN76SUK6p6ZQsGfsbZn\njyqos7AAXngBGDxYp6BOwRQ4kngE++L2KYM6S7ElFvZaiIEdBiqDujK5HL9lZeGPnBxlUGcqEmG6\nszNmOTu32aBOn7TK2BUUFODNN9/E7t27UVVVBVNTU8ydOxffffcd7O3tW2Kcz4zjOISFhdGlWEII\nERiFgq+jk0r5jF21Tp2AGTMAR0eDDU24ysqAyEggKUnV5uIChIQATk66dV1Vht/u/IbkvGRlm6u1\nK0IDQuFgqYq+Hz09Fixf7QQJNzP+WDAXMzOdxiBk1ZdiV65cqZeM3TNtdyKXy5GdnQ1nZ2eYGPkq\nFbMrdEAAACAASURBVLoUSwghwpOVxdfSpalq7iEW86dYDRyo8/GkbVNWFn80WG6uqq1bN2DWLH45\nsQ6elDzBzts7kVeep2zzc/bDTL+ZMDPhgzXGGM4VFCAqPx8Ktb/L/W1tMc7BAaY0qQBa+FLs1q1b\ncevWLZiYmMDNzQ0mJia4desWtm/frvMACKmLPgtJScuiuRM2Q81fdZbuxx81g7r27YFXX+XLv+jv\nf+NqzV98PLBhg2ZQN2IEn6nTMaiLy47DxusbNYK6YK9gzPWfqwzqip8eC3YyL08Z1FmIRJjn6orJ\nTk4U1DUDrVbFrlixAjdv3tRo69ChA6ZOnYqFCxc2y8AIIYS0DTk5fJZOfdcNExNg5Ei+9Iv+9jcB\nY/yqk1OnVG2mpsDMmYDa8ZxN65rhTMoZRD2IUraZmZhhZveZ8HNR1cMnPj0WrETtenpHCwvMdnaG\nvY4rb0n9tLoU6+DggOzsbI3LrzKZDE5OTigoKGjWATYVXYolhBDjxhh/vuvJk/x5r9Xc3fn4w9XV\ncGMTtMpKPlJ+ejQYAMDeHggNBdzcdOtaXol9cftwJ0vVt4OFA0ICQuBmw/ctZwyn8vJwXi0+4DgO\nQ+3sEGxvDxPam6ZOLXryhJ+fHyIjIzFv3jxl2969e41+pSrtY0cIIcYpL4+PPVJSVG0iEX+VcOhQ\nPmNHmiAvj9+fTn3Dv86dgeefB6ysdOo6vzwfO2/vRGaJqu/O9p3xvP/zsDLl+86rqsLvWVlIVTsW\nzMbEBLNcXODdBo4FawqD7GN37tw5TJo0CWPHjoW3tzeSkpJw4sQJHD58GEOHDtXbYPSJMnbCZmx7\naRHt0dwJW3PPX/WxpMeP84mlam5ufJauXbtme+lWLSU+Hkm7d+OvP/9ELwcH+Hh7w9PZmV9xMm6c\nzpHyg/wHiIiNQGlVqbJtYPuBGOczDiYivu/YkhIcyM7WOBbM19ISM5ydYSPWKo/UprVoxm7o0KG4\nffs2fv31V6SmpmLAgAH45ptv0LFjR50HQAghpG3Iz+f3xU1W7YoBkQgYNgwYPpyydE2VEheHxNWr\nMTo1FaKyMgSbm+PkX38Bb78Nz4kTdeqbMYbox9H4M/FPKBgfsJlwJpjcdTL6uPcBAFQpFDiSm4tr\naseCiTgOox0cMJiOBWtxz7zdSWZmJjw8PBp/sIFRxo4QQowDY8CNG8DRo4DaFTq4uPBZOgH8STFe\n+fk49fe/Y1RqqqrNzAzw98epLl0w6vXXm9y1XCHH4YTDuJZ+TdlmY2aDef7z0NGOT+w8eXos2BO1\n9Ku9WIw5Li7oYGHR5Ndui1o0Y5eXl4c33ngDkZGREIvFKC0txYEDB3DlyhV88sknOg+CEEJI61RY\nyGfpEhNVbRwHDBkCBAfze9SRJmCMPxrs8GGIsrNV7RIJEBAAmJtDpH6t+xkVVxYjIjYCDwseKts8\nJB6Y5z8PdhZ2YIzhenExjuTmokrt0qu/tTWmOjnBgtKvBqPVIvJXX30Vtra2SElJgfnTfW+ee+45\n7Nq1q1kHp6vw8HDaU0ugaN6Ei+ZO2PQ1f4wBN28C33+vGdQ5OwOvvAKMGUNBXZOVlgIREcDevUBF\nBRTV+8F4ekLq4KDcn07RxNMc0ovSseHaBo2grqdrTywOWgw7CzuUy+WIzMrCwexsZVAn5jhMdXbG\nHBcXCuqekVQqRXh4uN760+pndfLkSaSnp8NUbd8ZFxcXPHnyRG8DaQ76/KAIIYRop6gIOHSI3xu3\nGscBgwYBo0bx26mRJkpIAPbvB4qLlU0+vXvjZH4+Rru4AA8eAABOVlTAd/ToZ+4+5kkM9sftR5WC\n33+GA4cx3mMwuONgcByH1PJyRNY4Fsz16bFgrm34WDBdVO/esXLlSr30p1WNna+vL86cOQMPDw84\nODggLy8PDx8+xLhx4xAXF6eXgegb1dgRQkjLYgyIiQEOH+aPJq3m6AhMnw54ehpubIJXWQkcO8Yv\nKVbXrx8wbhxS7t9H0smTEFVWQmFmBp/Ro+HZrZvW3SuYAqfun8K5h+eUbRZiC8z2m40uTl3AGMOF\nwkKNEyQAoK9EggmOjnSChB60aI3dkiVLMGfOHHzyySdQKBS4ePEili9fjmXLluk8AEIIIcJXUsJn\n6e7e1WwfMIC/7ErJHB2kpgJ79mgeC2ZjA0ybBnTtCgDw7NbtmQI5deWycuy5uwf3cu4p25ytnBES\nEAJnK2cUy2TYl52NRLVo3VwkwjRnZ/hbWzftPZFmo1XGjjGGb7/9Fj/++CMePHiATp064dVXX8Vb\nb71ltMuYKWMnbLQXmnDR3AlbU+bvzh0+qCtVbXEGe3s+S9e5s37H16bI5cDp0/zRYOp/z/z8gKlT\n69xw+FnnL6c0BztjdiK7VLUAo4tjF8zuMRsWYgskl5VhT1YWitWOBWtvbo45Li5woGvqetWiGTuO\n4/DWW2/hrbfe0vkFCSGEtA6lpfxl15gYzfZ+/YCxY3U+Y75ty8ris3Tp6ao2c3Ng0iSgVy++aFFH\nibmJiLwTiXJZubJtaKehGNV5FBg4nMzLw7mCAo1gY4idHUY5ONCxYEZMq4zdqVOn4OXlBW9vb6Sn\np+M///kPTExMsGbNGrQz0m3COY5DWFgYHSlGCCHNIC6Oz9Kp1fDD1pbP0vn4GG5cgscYcOUKfzSH\n2gIFeHkBM2bwqVCdX4LhwqMLOJF8Agx8CCAWiTG923T0dOuJ/Koq/J6djUflqoDP2sQEM52d4avj\nsWSktuojxVauXKmXjJ1WgV337t1x7NgxdOrUCaGhoeA4DhYWFsjOzsaBAwd0HkRzoEuxhBCif2Vl\nwJEjwK1bmu29ewPjxwO0J60OCgv5A3TVj+YwMQFGjwaee04vWboqeRUO3juIvzL/UrbZmtsiJCAE\nHhIP3C0pwf7sbJSr7U3nbWmJWXQsWLPTV9yiVWBna2uLwsJCVFVVwc3NTbmfnbu7O3JycnQeRHOg\nwE7YqE5LuGjuhK2h+UtI4DcbVjs5ChIJX+71tIafNEX1cuI//gDUsmRo144/msPNTeuuGpq/wopC\n7IrZhcdFj5Vtnew6Ya7/XJiLrXAsLw9XCwuV94k4DiPt7THUzs5o6+lbkxatsbO1tUVGRgZiY2Ph\n7+8PiUSCiooKVFVV6TwAQgghxq28nN9p4/p1zfZevYCJEwFLS8OMq1UoK+OvacfGqtqa4WiORwWP\nsDt2N4orVdfO+7j3waQuk5AnU2D7/2fvzqOjrtNE/7+/VZXKUklVVkISskACISCLgOwiElQUXMA7\njtraTessp7X9zZ3uvjPn2O2I9u3pO/fO9DinnRlnbFvHdlpbRbRFFBUIO7IKghAIIQshgZClUtlr\n+f7++FKVqpCESlKpJXle53BMPqnlk3yt5KnP53meT20tl3sdC/ZgWhrZsgQbcfz6P+aZZ55h/vz5\ndHV18dJLLwGwd+9eioqKRnRyYuySFZ/IJdcusvW+fuXlWj9cq7VnzGTSVummTg3u3EadsjLth+u9\nBJqUpK3S5eQM6SH7ev0dqz3G5rObcapaZatO0bGqYBXzMuZxvK2NLb2OBSsymbgvJYVYOUEiIvm1\nFQtQWlqKXq+noKAAgLNnz9LV1cWMGTNGdIJDJVuxQggxdP31w73pJq0wU3Loh8Fu14ojDh70HZ8z\nR0tUDFA5sUt1sbVsK1/VfOUZi4uK46HpD5FhzmFzQwPfeFW/GBSFu5KTmZeQIFuvIRDUHLtIJIFd\nZJM8rcgl1y6ylZSUkJe3nA8/hObmnvG4OFi9GqZPD93cRoWaGq2NiXd+usmkNRseYoNhb+7XX7u9\nnfdOvceF5guer6Wb0nn4pofp0MXxfn09jV7pVGnXjgVLl07SITPiOXZTp071HBeWnZ3d7ySqqqr6\n/Fo42LBhg7Q7EUIIP5SWVvLZZ+f54osTuFwuJk3KJzVVOwOsqEgL6uLjQzzJSOZ0ao2Gd+0Cr21P\nCgu1oC6AJzhcabvC29+8TVNnk2esKLWIB6Y+wNG2Tr5sqsXpFUDMuXYsmFGOBQsJd7uTQOl3xW73\n7t3ceuutniftT7gGTbJiJ4QQ/iktreSll8qoqCj2nPHqcGxj4cICvvvdXGbMCEinjbGroUFbpaup\n6RkzGrXKk9mzA/LDLS0r5csjX1Jtq+bby9+SMzGH1MxUAG7Pu525E5bwUUMD57yOB4nW6ViTksIM\nidjDgmzF3oAEdkIIcWNtbfA//+d2zp1b4TOekgKLF2/nxz9e0c89xQ2pqpak+PnnWl6dW06OViCR\nlBSQpyktK+WNHW9Qm1ZLRXMFAI4yB7fcdAt/ufIviTbl8sHVq9i8Gh5nXjsWLFmOBQsbI74V+9xz\nz/X7JO5xRVF48cUXhz0JIXqTPK3IJdcuMqgqHDum5fBXV/dswdlsJdxyy3LS00Gvl625IbPZtIrX\nsrKeMb0ebr8dFi+GAG57fvLVJ5w1n6WhuYHmM80kTk0kfmo8afZ0avXp7Lp82edv+WKLhWI5FmzU\n6jewq66uHrAqxh3YCSGEiCz19fDxx+BOkdbptJyvceMgM1PriwtgNLr6eQQxoFOntN507n1t0H64\n69b1/HAD5FzDOUoqS2jJ7GksnBSTxKS0mXzTmkm3VwVM3LVjwSZLSfOoJluxQggxRtjtWu7+vn1a\nLr9bd3cljY1ljB9f7Bnr6trG+vUFFBbmhmCmEaqzE7ZsgRM9x3WhKNpxYCtWBKzZMGhHg31R/gUH\naw5ycM9B2idouXMTzBOwJM2gVEklus3MLdPmATDx2rFgCXIsWNga8Ry7cu+z6gYwadKkYU9iJEhg\nJ4QQPcrKtBOrmnoKJdHptAMOli2D8vJKtm07T3e3DqPRRXFxvgR1g1Ferp3z6nUkFxaLlkuXlxfQ\np6q11fLB6Q+ob68H4Oqlqxw6dJzo9Em0xYyjxWUkIc7C4ltWkJaS6jkWTCe7bGFtxAM7nR/7/4qi\n4PR+2xdGJLCLbJKnFbnk2oWX1lb47DPtKFJvOTmwZo22Q+hNrt8g2e2wbRscOOA7Pns2rFoFATyS\nS1VV9lXvY/uF7Z5TJADMXSkcPA8n8/NoOnGSuFkzMZeWc8ecOfxg7lxy5ViwiDDixRMul+RWCCFE\npFJVOHIEvvzS91z5mBi44w7tkANZwBmm2lqtjUl9fc9YXJx23lqAj9y0dlrZdGaTp+oVwKBEcfPE\nu3h3dyl1i6aRChgS60hMyyQ1M5fx5eUS1I1Bo3qzXRoURy65ZpFLrl3oXb6sFUdcvOg7PnOmdmLV\nQL1w5fr5weWCPXugpMS32fDkyXD//QHv5Hzyykk2n91Mp0OL0FXAYMojYdwSDjkNXPZqY5I0bx4F\nsbFkRkdL5B4hgtag+K677mLr1q0AnkbF191ZUdi1a1fAJhNIshUrhBhrurth507Yv9833khO1rZd\nwzQlOrI0NsKmTVBd3TMWFaVtuwZ4GbTL0cWWc1s4fvk4AC6gnniikueQGJ/nyZk7uHMn7bNmkRYV\nRV5MDCa9HoBxJ0/y1L33Bmw+YmSN+Fbsd7/7Xc/HTz75ZL+TEGIkSJ5P5JJrFxpnz2oFmd7nu+r1\nsHQp3Hqr/wWZcv36oapw9Chs3apF0G4TJmhtTJKTA/p0VdYqPjj9Ac2dzThRqCWeq4bxTEydjiXa\n4rmdXlFYM2MGZ8+eJXH+fCoOHMC0cCFdR45QPGdOQOckIkO/L/XvfOc7no/Xr18fjLkIIYQYJJsN\nPv0Uvv3WdzwvT1ulS00NybRGl9ZW+OMftejZTaeD5cu1yDmAzYadLic7K3eyu3I33SjUYKEGM8mm\nTG5KLsCg0/5sG3U65iUksNBsxpyXR2lSEttOnuTqhQuMi4+neM4cCmWJdkzyu4/drl27OHbsGG1t\nbUBPg+Jnn312RCc4VLIVK4QYzVwuOHQItm+Hrq6e8bg4uPNOmDVLUqwC4vRpLWHR64xV0tK0NiaZ\nmQF9qob2Bj44/QHltjqqMVNLAopiZErKFMaZ0gCtyfBCs5lbEhKIvbblKkaHEd+K9fbMM8/w7rvv\ncuuttxIbGzvsJxVCCDF0tbVarHHpku/47NlaUCcHCwRAZ6fWJ+brr33HFy6E4mItry5AVFXlaO1R\nNp77kgtqHHVMQEUhMTqRqalTiTFEYzEYWGKxcHN8PFEBXCEUo49fK3ZJSUmcOnWKzAC/OxlJsmIX\n2STPJ3LJtRs5XV2wYwd89ZWW8uWWmqptuwaiD65cP6CiQiuQsFp7xsxmeOCBgFegtNvbeePbzexo\nqqeeOEBBQWFi4kSyLdmkG40stViYbjL5dbarXL/IFdQVu+zsbIxG47CfTAghxNCcOaMVR3gfbGAw\naIURS5YE9LSqscvh0Pa29+/3jZxnzoR77gl4s+EddaX8Z9l+6pwKoPWgiTPEUZRWxDRzKkstFqbE\nxkqhohgUv1bsDh06xN///d/z6KOPkp6e7vO1ZcuWjdjkhkNW7IQQo4HVqhVHnDnjOz5pEqxeDSkp\noZnXqFNXpzUbvnKlZyw2VlsKnT49YE/jUlW+sbXwWvlXnGj23UvPSsjizsxpLE9KISc6WgK6MSao\nK3ZHjhxhy5Yt7N69+7ocu2rvXj5CCCECwuXStlx37PDtrmEyaU2GZ8yQ4oiAcLlg3z7tB+19RGZ+\nvtZs2GwOyNM4XC5OtLXx6ZVq9tWdos3e5vmaUR/FfVkz+B9ZhYyPjg7I84mxy68Vu5SUFN555x3u\nuOOOYMwpIGTFLrJJnkjkkms3fDU1WnFEXZ3v+Ny5sHKltpA0UsbU9Wtq0nLpqqp6xqKitDPXbrkl\nIJFzl8vFUZuNfVYr3zZVUN50ARXtb5MOlXkJFv6/omImxFlu8Ej+GVPXb5QJ6oqdyWTitttuG/aT\nCSGE6F9np5bideiQb4rXuHHajmBOTujmNqqoqlbt+umnvsuhWVlaG5MANP9rczo52NLCQZuN5u52\nzlwtpamzCQA9LnKVdtbnL+DWrHmy5SoCyq8VuzfeeIODBw/y3HPPXZdjpwvTsmtZsRNCRApV1dql\nffqp1nDYLSoKbrsNFi3STpEQAdDWpjUbLi3tGdPpYNkyrRJlmD/oZrud/S0tHG1txe5yUd9eT+nV\nszhUB0acTMDK3PgE/nTaWlLjpHu06BGouMWvwK6/4E1RFJzeOQlhRFEUnn/+eZYvXy7L0kKIsNXc\nDJ98AufO+Y4XFGjFEUlJoZnXqFRaqgV1bT35baSkaEeCZWUN66GvdHez12rlm7Y2XKqKw+WkrLGM\nurY6YrCTQwvjaeW2nKUsz1uOXieRutCUlJRQUlLCCy+8ELzArqKiot+v5QWicdIIkBW7yCZ5IpFL\nrp1/nE44cABKSsBu7xmPj4e774Zp00JTHDEqr19Xl3bG69GjvuPz52v5dMNoNlzd2ckeq5VSr5Mp\nrF0tnK4/jcHZQg5W0mgjMdrCuqJ15CbmDvm5/DEqr98YEdQcu3AN3oQQIhJVV8PmzXD5cs+YosC8\nedqhBgFslyaqqrQCiaamnrGEBK3itaBgSA+pqirnOzrYbbVS2dnZM45KRXMlVmspU7CSRAcKMDN9\nJvdMvocYg1xYMfL8Pis20siKnRAi3HR0wLZtcPiw73h6Otx7L0yYEJp5jUpOp9bCZO9e30qU6dO1\nSpQhlBa7VJVv29rYY7VS5110AXQ4OmhqPIGp4wIWtMN7YwwxrJ68mhnpM4b1rYixIagrdkIIIYZO\nVeHkSe3oUe/0rqgouP12WLBAiiMC6soVrdmwd7+YmBgtafGmmwa9x+1wufi6tZW9LS00ee+bAwoQ\nb7+CvW4nma6e7dhcSy5ri9aSGJM4nO9EiEGTwE6EJckTiVxy7Xw1NmrFEefP+45PmaKdUpUYZn/3\nI/r6qaqWuPjll77NhidN0rZeLYPrFdfpdHLYZuNASwutvQoFo3Q6psdG0VS3h8rGb3Fn6ekUHSsm\nrmBx9mJ0SvC7RkT09RMBIYGdEEKMAKdT2wXctUs7gtTNbNaKI6ZOlZMjAqq5GT78ELyL/QwGraPz\nggWD+mG3OhwcaGnhkM1Gl8vl87VYvZ75CQmkOa+y9ez72Lp7+tOkxqWyrmgdmQmZw/1uhBgyv3Ls\nysvL+elPf8rXX39Na2trz50VhSrvjt1hRHLshBChUlmpFUfU1/eMKYoWX9x+O8ipUQGkqnDiBGzZ\nolW/umVkaG1M0tL8fqhGu519Vitft7bi6PX3w2wwsMhsZpYpll0V2zlw8YDP12/JvIU78+8kSj/0\nClsxtgU1x+7RRx+loKCAX/3qV9edFSuEEELT3q7tAvbuqpGRoRVHZMpCTmC1t2tnr50+3TOmKFqj\n4dtu8ztxsa6ri70tLZxsa7vuD2tqVBRLLBZmxsdzte0K/3Xs91xu6ylnNkWZuH/q/UxJmRKQb0mI\n4fJrxc5sNtPU1IQ+grJ7ZcUuskmeSOQai9fOvWi0dasWa7gZjbBihdYuLUwP6blOxFy/c+fgo4/A\naxeJ5GTtSLDs7BveXVVVqrq62GO1cs77ol2TGR3NrRYLhXFxKMBXNV/xZfmXOFw9++qTkydz/9T7\niTfGB+I7CoiIuX7iOkFdsVu2bBnHjh1j3rx5w35CIYQYTa5e1YojLlzwHS8q0nLpzObQzGvU6u6G\nzz+/vmfMvHlw551aND0AVVU529HBHquVaq8edG75sbEstVjIi4lBURRsXTY+PPMh55t6ql8MOgN3\n5d/FvEw551WEH79W7J5++mn+8Ic/sG7dOp+zYhVF4cUXXxzRCQ6VrNgJIUaSwwF79sDu3b4FmBaL\nVu1aWBi6uY1aFy9qbUwaG3vG4uPhvvu0MuMBOFWVk21t7LVaudKrB52iKBTFxbHUYiHTKwHydP1p\n/lj6RzocHZ6xjPgM1hWtI83kf+6eEP4I6opdW1sba9aswW63c/HiRUB71yPvVIQQY9GFC1pxREND\nz5hOBwsXwvLlN1w0EoPldMLOnVoU7f2Hr6hIazZsMvV7V7vLxdHWVvZZrVi9y5MBvaIwKz6eJRYL\nKV7HinU7u/n03KccqzvmGVNQWJKzhNvzbpdzXkVYk5MnRFiSPJHINZqvXVubtgt4/LjveFaWVhwx\nfnxo5hVIYXf96uu1Vbra2p6x6GhtWXTmzH7bmHQ4nRy02fiqpYX2Xj3ojDod8xISWGg2Yzb4rm9c\nbLnIB6c/oLGjZ1XQEm1hbdFa8hLzAvZtjZSwu37CbyO+YldRUeE5I7a8vLzfB5g0adKwJyGEEOFM\nVeHYMfjiC+1YMLfoaK1N2ty5kVMcETFUFb76Sisz9l5py8uDBx7ot7Nzy7UedIdtNrp79aCL0+tZ\naDZzS0ICsb2KAV2qi92Vu9lZuROX2nO/m8bdxOrJq4mNko4QIjL0u2KXkJCAzaY1XtT18xtLURSc\nvd4JhQtZsRNCBEJ9vbbtWlnpOz59OqxapZ0nLwLMatUqXr0XFfR6KC6GRYv6XKW72t3NvpYWjre2\n4uz1uz/RYGCxxcLN8fFE9fH3rKmjiQ9Of0B1S7VnLFofzeopq5kxboakHYmgCFTcEpFbsX/7t3/L\n/v37ycvL47e//S0Gw/ULjxLYCSGGw27XUrr27vUtjkhM1I4cnTw5dHMbtdyH6n7yCXhXrI4fr7Ux\n8Srec7t0rWXJ6fb2637njzMaWWqxMN1kQt9HcKaqKscvH2fLuS10O3sKKnIsOawrWifnvIqgClTc\nEnGbB8ePH+fSpUvs2rWLqVOn8v7774d6SmIElJSUhHoKYohGw7U7fx7+/d+148DcQZ1OB0uXwtNP\nj+6gLmTXr6MD3n8fNm7sCeoURfuh/9mf+QR1qqpS3tHBm3V1/OelS3zbq7FwdkwMj6an84PMTGbG\nx/cZ1HXYO3jv2/f48MyHnqBOp+gonljM+tnrIzaoGw2vPzE8EXdW7P79+7nrrrsAWLVqFa+//joP\nP/xwiGclhBgNWlu1JsPffOM7np2tFUeMGxeaeY16ZWXa1qut59xVEhO1VbrcXM+QS1U5097OHquV\nS97Hh10z+VrLktyYmAGfrrypnA/PfEhLV4tnLCU2hXVF68gyZw3/+xEihCIusGtqaiIjIwPQTsRo\n9O5nJEYNqeqKXJF47VQVjhzR8vS9dwBjYuCOO2DOnEGdIR/Rgnr97HatIuXgQd/xOXPgrrs8h+o6\nXC5OXOtB12C3+9xUURRuMplYYjYz/gaH8DpcDrZf2M6+6n0+43Mz5nJXwV0Y9ZHfpyYSX38isEIW\n2L388su88cYbnDx5kkceeYTXX3/d87XGxkaefPJJvvjiC1JTU/nlL3/JI488AkBiYiItLdq7LKvV\nSnJyckjmL4QYHS5f1oojqqt9x2fO1A4yiA+f06JGl5oarY2JdzNAk0lbGp06FYAul4ujNhv7W1po\n6dWDzqAo3JyQwGKzmSSvHnT9udJ2hY3fbvQ55zUuKo77Cu9jaurUwHxPQoSBQQd2rl7l4/1VzN5I\nVlYWzz33HFu3bqXDu38A2kkXMTExXLlyhWPHjrF69WpmzZrFtGnTWLx4Mb/61a94/PHH2bp1K0uX\nLh3S84vwJr2YIlekXLvubq3n7f794P1rLTlZK47Izw/d3EJpxK+f06lVpeza5fuDLyzUTpAwmWhz\nOjnY0sJBm42OXp0XonU65pvNLEhIIL6PwrneVFXlYM1Bvij/wuec14LkAh6Y+kBYnfMaCJHy+hMj\nx6/A7siRI/zwhz/k+PHjdHrtUwyn3cnatWsBOHz4sOc0C9BOufjggw84deoUcXFxLFmyhPvvv5/f\n/e53/PKXv2TWrFmkp6ezbNkycnNz+Zu/+ZshPb8QYuw6d04rvGxu7hnT62HJErj1VvBjAUgMxdWr\nsGmTtlrnZjRqh+rOnk2zw8H+hgaOtrZi77WIEK/Xs8hiYW58PDF6/05+sHXZ+Kj0I8oayzxjBp2B\nO/Pv5JbMW6SNiRiV/Arsvve973Hffffx2muvERcXF9AJ9C7tPXv2LAaDgYKCAs/YrFmzfCp95Q/y\nNQAAIABJREFU/u///b9+Pfb69es9TZYTExOZPXu2552M+/Hk8/D83D0WLvORz/3/fPny5WE1H+/P\n585dzqefwpYt2ud5edrXOzpKWLQIVqwIr/mOmuu3YwecOcPyxkaw2ympqNC+vmwZrF3LpsOHOfnR\nR6izZ+NSVSoOHAAgb+FCkqOiiD5xgoKYGJasWOH381dZq6hLraPd3k7F19rzLVy6kHVF6/j20Lfs\nPLczLH7egf48nF9/8rnv5+6PK669HgLFrz52ZrMZq9U6Iu9unnvuOS5evOjJsdu9ezcPPfQQtV7H\nx7z66qv8/ve/Z8eOHX4/rvSxE0K4uVxw+DBs2wbexZSxsVoe3ezZY6c4IuhsNq3itaxn1Qy9Hm6/\nneo5c9hjs1Ha3n7d3cYbtR5000wmdIO4ON3ObraWbeVI7RHPmILCouxFrJi4AoMu4moGxRgx4keK\neVu7di1bt25l1apVw37C3np/E/Hx8Z7iCDer1UqCtHcfU0q8VutEZAm3a1dbqxVHeO/+gRbM3XHH\ngOfHj0kBvX6nTmk/fK88anXcOM6vWcPuqCgqL1++7i55MTEstVjIj40d9GJCTUsNH5z+gIaOnoIM\nc7SZtVPXMjFp4tC/jwgSbq8/EXx+BXYdHR2sXbuWW2+9lXSvJpGKovDmm28OawK9X7hTpkzB4XBQ\nVlbm2Y49fvw4N91006Afe8OGDZ6laSHE2NLdDTt2wIEDWjsTt9RUWLNGO3JUjJDOTtiyBU6c8Ay5\nFIVvFy9mz9Sp1Dkcvsd5AFOv9aCbcIMedH1xqS72VO2hpKLE55zX6WnTWTNljZzzKsJaSUmJz/bs\ncPm1Fbthw4a+76woPP/880N6YqfTid1u54UXXqCmpoZXX30Vg8GAXq/nkUceQVEUfvOb33D06FHW\nrFnD/v37KSoq8vvxZStWiLHrzBktrvBe/DcYtMKIJUu0j8UIKS+HDz/0/PAdisLX6ensXbyYpl47\nLzpFYabJxBKLhTSjcUhP19TRxKYzm6iyVnnGovXR3DP5Hmamz5QCCRExIv6s2A0bNvDiiy9eN/Z3\nf/d3NDU18cQTT3j62P2f//N/Bn26hAR2Qow9Vit8+qkW2HmbOFFbpUtJCc28xgS7XUtivFb40KnT\ncTghgQOFhbROnuwTTUfpdMyNj2eRxYJliFG2qqqcuHyCLee20OXsSZzMNmezrmgdSbFJw/t+hAiy\noAd2O3bs4M0336SmpoYJEybw2GOPseJahVI4ksAuskmeSOQKxbVzueCrr7St1+6es9wxmbQDDGbM\nkOIIfw3p+tXWas2G6+tp1es5YDZzKDmZrsJCSEvz3CxWr2d+QgILzGbi/GxZ0pcOewebz27mVP0p\nz5hO0XFb7m3cmnsrOkU35MeOdPK7M3IFtXjiN7/5Dc8++yx/9md/xoIFC6iqquLRRx/lxRdf5C/+\n4i+GPYmRIjl2Qox+NTVafr5XIT2gnUp1xx1a5asYIS4X7NkDJSU06nTsS0nh6/h4HCkpWsPha9ur\nZoOBRWYzcxMSMOqGF3RdaLrApjObfM55TY5NZl3ROiaYJwzrsYUIhZDk2E2ePJn333+fWbNmecZO\nnDjBunXrKPMuYQ8jsmInxOjW1aXt/B065FsckZamnUqVkxO6uY0JjY3wwQfUXbnCXrOZkyYTql6v\nHdmRmQlAalQUSywWZsbHox/mkqnD5WDHhR3sq96HSs8Fn5Mxh1UFq0bFOa9ibAvqVmxKSgq1tbUY\nvZJbu7q6yMzMpMH7nL8wIoGdEKOTqsLp01ounc3WM24wwG23weLFWps0MUJUFfXwYap27mRPXBzn\n3E3rzWbtjNfYWLKio1lqsVAYFzeoHnT9qW+rZ+PpjdS11nnG4qLiuHfKvRSl+V9UJ0Q4C1Tc4tea\n+JIlS/jRj35EW1sbAK2trfzkJz9h8eLFw57ASPrXjz+mtLw81NMQQxDIZWkRXCN57Zqb4fe/h3ff\n9Q3qCgrgqae0qlcJ6oZnoOun2myUvvsuvz1yhNdTU7WgTlG03jGzZ5OfnMz3xo/nzzIyKBpkY+E+\nn+/aOa//ceQ/fIK6/KR8fjDvBxLU9UF+dwq/cuxeeeUVHn74YSwWC8nJyTQ2NrJ48WLefvvtkZ7f\nsLz35ZfsPXqU5x9/nMJJk0I9HSHEEDmdWrFlSYlWfOkWHw+rVsH06VIcMZKcqsrJb75h79GjXAFw\n95qLi0MpKqIoPZ2lFguZ0dEBe87W7lY+OvMR5xrPecYMOgMrJ61kQdYCaWMiRo2Q5Ni5VVdXc+nS\nJTIzM8nOzg7YJEaCoijc8fXXRCsKaSdP8j/uuguzXo/ZYPD8N0GvJ2qYibxCiJFVXa0VR3gfUqAo\nMG8eFBf3xBgi8OwuF0cbGth3+DDWXqdE6LOymDVjBktSUkiJigro85ZeLeWPpX+kzd7mGUs3pbOu\naB3p8ekD3FOIyDXiVbGqqnreEblcWifvrKwssrKyfMZ0YRwYdbtcdAN2u52DvY4pc4vV668L+Mx6\nPQleH0frdPLuUIgg6+jQiiOOHPEtjkhP14ojJkgBZECVlpfz5alT2AHV5SI1L49aoP3bb30O2DVG\nRTFv9mwWFhZiDnCn525nN5+f/5zDlw77jC+asIjiScVyzqsQfuj3VWI2m7FdS2Ix9PPiVRQFZ69j\nYcKRfoAIuMPppMPp5LJ386tejDqdzypf7yDQbDAQJ8FfQEkvpsg13GunqnDyJGzdCq2tPeNRUXD7\n7bBggeTRBVppeTm/PXwYde5cju3ejWvWLLq++ILZikLqta7OcU4nC1NTuWXlSmJH4IDdS7ZLbPx2\no885rwnGBNYWrWVSkqTS+Et+d4p+A7tTp3oaP5ZHaAHCQrMZ26FD3H3LLaQkJ9PidNLicGC79t8W\npxOXH8ue3S4XV10urnon9/SiV5R+g74Er/8GokJMiNGqsRE++QTOn/cdnzIF7rkHEhNDM69Io6oq\nXS4XbS4X7U4n7S4XbU6n5+N2p9Pna9t27+ZKQQEdZ87QdvEipqgoYnNzufD11xRYLCzu7OTm224j\nasaMgM/VpbrYW7WXHRU7fM55nZY2jTVT1hAXFRfw5xRiNOs3sMvxagL1/vvv85Of/OS62/zqV7/i\nRz/60cjMLAC++dnP+NN772XN9Ol9fl1VVdqcTk/A1+J0+gR97v/aXa4+7+/Nqao0Oxw0Oxz93kZR\nFOK9tn4T+tgGlrw/jbzjjFxDuXZOJ+zbBzt3gvdLKCEB7r4biorGdnGEU1V9gzKvjz1BW6+v+fOm\n1c3a1kZHbS1JDgdJEyaA3U5HaysTGht5ZupU9A89pLUzCbDmzmY2nd5EpbXSM2bUG7m74G5mj58t\nuyBDIL87I09IiicSEhI827LekpKSaGpqCthkAilQSYiqqtLpcvUZ8HkHgx0B3JKO65Xjd93qn+T9\niVGkslIrjqiv7xlTFJg/H1asgAAWWoYFVVXp9grUBlpJc3+t0483l4OcBLS1af1jrFY+3bqVuDlz\niHK5MHV3k9XSQkpnJ82VlfzLa6+NSFR94vIJPjn7ic85rxPME1hXtI7k2OSAP58Q4S4oR4pt374d\nVVVxOp1s377d52vnz5/HPALv4MKNoijE6vXE6vWMM/bf2by7j+Cv9+dtLpdfF6392i9zf/P++sv/\ni+S8P8kTiVz+XruODvjiCzh61Hc8I0Mrjrh2eEHYc6kqHYNYSWt3OnEEoXm6UafDpNcTp9MRB8RZ\nrZjq64mrqyOurg5TRwdxLhdxTifZZ85QVl1N9OzZVFRUkDplCl2XLzM3OzvgQV2no5NPzn7CN1e+\n8YzpFB3LcpexLHfZmD7nNRDkd6cYMLB74oknUBSFrq4unnzySc+4oiikp6fz61//esQnGCmMOh0p\nOt2AZf9OVcU2wJavOxgMZN5ff1u+7v/GS96fCDJVhRMntOKI9vaecaNRW6GbPx9CmY1gd7n8Xklr\nd7no8PMN23AoiqIFaO5ATa/H5PW5Sa/3/djhwHDxopasWFmpHag7QJrIxKgoljY1se3oUa52dDAu\nOppik4ma9MC2FqlormDT6U1Yu6yesaSYJNYVrSPbEt4ttISIFH5txT7++OP87ne/C8Z8AiZSjxTr\nK++vr/w/f/L+/NE776+//D+D5P2JAGho0LZdL1zwHZ86Vculs1gC+3zuVAp/V9LaXK6AvbYGYlCU\n64Mxd8DWR/AWo9MN/AasrQ2qqrR/lZVQW+vbI6YvCQmQmwu5uVR2d1O2eTPFXvve27q6KFi/ntzC\nwmF/v06Xkx0VO9hbtdfnnNebx9/MqoJVRBtG2X67EEMQ1LNiI1GkBnb+cP+x8gn6+lj9C2ReTty1\nQC+hj1U/93+jJfgT/XA4YM8e2L1bK5Rws1i0ald/Ywd3EUHvYKyvlbQ2p5OOQRYRDFWsvytp175m\nHO5rxWrVArjKSi2Y805Q7E9KCuTkeII5EhN9tlkrS0s5v20buu5uXEYj+cXFAQnqrrZfZeO3G6lt\nrfWMxRpiubfwXqalTRv24wsxWgQlx87NarWyYcMGdu7cSUNDg6c5saIoVFVVDXsSI2XDhg0sX758\n1OUbeOf9pQ8y7693MNjqZ9GHO++vbpB5f72DQX/z/iRPJPK4G9ye/uYbimbMYOX06RROmsSFC9oq\nXUNPezJ0Opi/QGXJbSp2vZOLnTdeSWt3OukKwmqaXlH8Xklzfz6i6Qyqqv3wvAO55uaB76MoWifn\n3NyeYC4+fsC75BYWkltYSElJCSsC8NpTVZXDlw7z+fnPsbt6UkYmJU3igakPYI4e/TnaoSC/OyNP\nSKpiH3vsMaqrq/nrv/5rz7bs//t//48HH3wwbNudjOYVu0Dyzvvrr+WLv3l//nDn/fW35VtXVcX+\nM2coPXnSJzgIlL7+n+g90td32vt+fd5mhB6nz8ce6vcxQo9zrrycd77+GuPcuVR+9RUZ8+fTvP8Q\nqd03cdWWg13v1P7pXMSlOJk83YXe5MQZhNdo9CBW0kx6PUZFCW3RkculnZ/mHci1tQ18H71eqzZx\nB3I5OUM+ay0QgUFbdxsflX7E2YazPVNU9KyctJKFExZGbFFXJJDALnIFdSs2LS2N06dPk5qaisVi\nwWq1UlNTw7333svR3iVtYUICu8BxeeX99bXl6/7vcCv9rlZX83VpKYZ583BvVDmOHOHmwkJSe51N\n7HdAJP8PBMXBnTu5OrmQpuZOXC4Fu11FVWOIOVpK1uTbAC32mDRRiz+G+nddpyjE+rmSZtLridXp\nwj8/1OHQihvc+XHV1T5HePUpKgqys3sCuQkTtLEwcLbhLB+d+cjnnNdxpnE8WPSgnPMqxACCuhWr\nqiqWa1nNCQkJNDc3k5GRwblz54Y9ARH+dIpCgsFAgsHQb1OxvvL++sr/Gyjvr7y8HMO8eQC4b6Wb\nO5ey48e1pqkibLW0d1J3uRNVTaS9XcujU9VmFGcnAGlpUFAA0b0yB6J0Or9X0uJ0OmIiuIWPR1eX\nFry5A7kbVKwCEBvrmx83fnzYnatmd9r5/PznHLp0yGd84YSFrJy0Us55FSJI/HqlzZw5k127dlFc\nXMzSpUt5+umnMZlMFAYgsVaMDoPJ+7uu19+1j09FRWFXFOyqSvPhwyReC/KcI/SHvK8AofdIX8/c\n51ivx/LnfsF8HL/vN8THabvchsuSTEc7dJ85jHHyfHTONPRXjvL97yRSmNN38DYmTllxV6y6t1X9\nqVg1m30DubS0oB29MZStvFpbLRtPb+Rq+1XPWLwxnrVT15KfnB/gGYqByFas8Cuwe/XVVz0f/8u/\n/AvPPvssVquVN998c8QmJkYno05HqtFIah9fazSbqbdYcAEXTCbyrq0Sj0tK4ge5uZ7bDTloifSV\nnjBls8HWb7JoOH6OhNlz6WyIIyYhnvjyI6zOyeJ7M5JCPcXgGoGK1XDlUl3sq97Hjgs7cKo9hVhT\nU6dyX+F9cs6rECHgV47dV199xYIFC64bP3jwIPPnzx+RiQ2X5NhFntLyct44epTouXM9Y11HjrB+\nzpyAFlCIwDl9Gv74R9i5czv11jyaY04RZYKMVChKnM60ggqeempFqKc5cnpXrFZWaoHdQIZQsRqO\nrJ1WNp3ZREVzhWfMqDeyqmAVN4+/Wd5ICTFIQc2xW7lyZZ9nxa5atYrGxsZhT2KkjNZ2J6NV4aRJ\nrAe2nTxJN2AEiiWoC0tdXfDZZ3DsmPb5pEn52I6XsTDzXiZO1NqZdHVto7i4ILQTDbQQV6yGi5NX\nTrL57GY6HZ2esayELNYVrSMlLiWEMxMi8gS13Ynr2lE5iYmJWHu9Cz1//jxLlizhypUrAZtMIMmK\nXWSTPJHwVV0NH3wATU09YxYLzJxZSWnpeb799gTTps2kuDifwsLc/h8oEgy3YjU3F7KywqZi1R8D\nvfY6HZ1sObeFE5dPeMYUFM85r3pdeBV0jEXyuzNyBWXFzmAw9PkxgE6n46c//emwJyCEiAxOJ+za\npf3z/t0zYwasXg0xMbkUF+dSUqKL3D8s7opV92rcKKlYDYTK5ko2ndlEc2dPc+SkmCTWFq0lx5IT\nwpkJIbwNuGJXUVEBwLJly9i9e7cnklQUhbS0NOLiwjcxVlbshAichgZtla6mpmcsJkYL6GbMCN28\nhi3CKlZDwelyUlJRwp6qPT7nvM5Kn8U9k++Rc16FCBA5K/YGJLATYvhUFY4e1fLp7D2nQpGXBw88\noBVvRhTvitXKSrh69cb3SUnxLXSIkIrVQGhob2Dj6Y1csl3yjMUYYrh3yr1MHzc9hDMTYvQJavHE\n448/3ucEAGl5IkaE5ImEXlubVvFaWtozptfDihWwaJFWINGXsLl2qqoFbu4VuTFUsTocJSUl3Hbb\nbRytPcpnZZ/5nPM6MXEia4vWyjmvYSxsXn8iZPwK7PLz830iybq6OjZu3Mh3vvOdEZ2cECI0zp2D\njz6C1taesbQ0WLcOMjJCN68BuVxQV+e7tTqYitXcXK3oIcIrVoeqtKyUL498yfETx/mvY/+FKd1E\naqbWcVKv6CmeVMyiCYukjYkQYW7IW7GHDx9mw4YNbN68OdBzCgjZihVi8Ox2+PxzOOR7KhQLFsDK\nlWFW3DmUilWjUTtXNUIrVkdKaVkpb+x4A1uWjbMNZ+l2duMoczB72myKCop4cNqDjI8fH+ppCjGq\nhTzHzuFwkJSU1Gd/u3AggZ0Qg3PpklYg4Z12Fh+v5dIVhEM7OqlYDThbl43zTed5+Z2XqUyu9Nl2\nBZhum85LT71ElF6CXyFGWlBz7LZt2+az/N7W1sY777zD9OmSPCtGhuSJBI/LBXv3wo4d2sduU6fC\nfffBYIvfA3btpGI14BwuB5XNlZxvOs/5xvNcbrsMwMXWi9gTtaCu+Uwz46aPozClkPyYfAnqIoz8\n7hR+BXZPPvmkT2BnMpmYPXs2b7/99ohNLBDk5InIU1layvkvv+TE6dO4Tp0if+VKcgsLQz2tUau5\nWVulq6rqGTMa4e67YfbsIMdEzc2+hQ5SsTpsqqpytf0qZY1lnG86T2Xz9atyADq0SpgoXRTJscnM\ny5yHUW/E2GoM9pSFGHOCevJEJJOt2DDkdGrbad3d2r9eH1eePUvZxx9T7C631OnY5nBQsHo1ufn5\nYDBo22juf4P9XP7ge6gqnDgBW7b4pqVNmKAVSCQnB2ECw6lYdQdzY6xi1R8d9g7Km8o533SessYy\nWrpa+r2tXtGTY8khyhbFgRMHSJqW5HkT33Wui/W3r6ewQN5YCREMQc+xa25u5pNPPuHSpUtkZmZy\nzz33kJSUNOwJjBQJ7IZJVbX8pT4CsBt+3N/XnM4Bn3L7wYOsaG+/ftxkYsUttwz/expOUDjQ54F4\nrCAGnR0dsHkznDrVM6bTwbJl2r/+2pgMi1SsjhiX6uJiy0XON57nfNN5alpqfBoJ95Yal0p+Uj75\nyfnkJeZh1GurcqVlpWw7uo1uVzdGnZHiOcUS1AkRREHNsdu+fTvr1q2jsLCQ3NxcKisreeqpp9i4\ncSMrV64c9iREAKjqjQOrwQRj3d2+CVdBoPN6vpLmZpZf636ru0FA6Den84bBZcjodIEPOPsIQKsv\n6flihx5bmx6LzoCq02NO0nPPvXoycwxg6+O+fkZ63tvoM6dMIf+mm8g1GLQgTipWA6q5s1nbXm08\nz4XmC3Q6Ovu9bYwhhklJkzzBXGJM312lCwsKKSwolBytCCfXT/gV2D399NP853/+Jw899JBn7L33\n3uOHP/whZ86cGbHJjWouV2BXw7q7Q/0d3ZhOp/3xNhohOvq6j111dVjrarnUdJkLaifjXR1kJKXj\nmpAN8+ZpK4ju4Mz7Y38+D9eAzs3l0v7Zr89/CtTDX7igxVeTvMYzMqDABPoPB7izotwwaKysr6ds\n3z6KjUZ0ly+z/NQptr37LsyeTW5qat+PKxWrfut2dlPRXOEJ5ho6Gvq9rYJCljmL/KR8CpILyDJn\noVNGYhlWCBGO/NqKTUxMpKGhAb3XL1273U5aWhrNzc0D3DN0FEVh28svBy753r0tGYgArKvrxm0a\nwoHB0Hcg1k9gdsOP+8hz67B30NTZRFNHE9s/38KZ37zGrala7k+0IZo9DQp3/PUG7rj73uF9L6o6\nuEBwOEHkUD4fQa2tcPq0785nVBQUFkJ/Mddg+bWNbjb7FjpIxWq/VFWlrrXOU71aZa3Cqfb/5sQc\nbaYguYD8pHwmJU0iNio2iLMVQgRC0I8Ue/nll/mrv/orz9i///u/93nUWDhZUVPDtldegQcfJDc7\ne2gBmPvjcF/xAe2vdX+B1VCCsQCsnrhUF7YuG422K54ArrGj0fNxh6PDc9uDFQex3ZVA1bfNRDsd\ndOm7ab49kVNHXuF8Wg1ZCVlkmbPITMhknGnc4FYhFEULVA1+/S8fXKqqLakFOKhUHU5KTzk4fs6J\nGu9EZ3KiczkYn+ZkzmwnsUY/n8uPXzS6vrbtY2PRjR+vNcKTitUbau1u9eTJnW88T5u9/xzEKF0U\neYl55Cfnk5+UT2pcqpwIIYQA/FyxW7JkCQcPHmTcuHFkZWVRU1PDlStXWLBggeeXiaIo7Nq1a8Qn\n7C9FUVBvuw0IYPJ9ICnKjQOrwQRjUVEjlPV+Y3anvc+grbGjkebO5gFXGrwd2HOAzglarlDzmWYS\np2q5QDEXY1i4dKHPbaN0UWQkZJCVoAV6WeYskmKS5I/bNS0t8OGHUF7eM2YwwJ13wi23DDK+cged\nAwSR23/7W1ZcvQqqSsnFiywvKgKjke3jxrHiqacC/v2NBg6Xg2prtacVSV1r3YC3Tzela6tyyfnk\nWHIw6EbmTYrkaEU2uX6RK6grdn/+53/On//5n99wQuEqIMn3Op1/QZa/wVhUVMSsXqiqSru9/bqg\nzf2xrXvop49E6aJIik0iKSaJ+sR6upO7iTZEc6HmAvGmeGzdNhSu/znZXXaqrFVUWXsasMUaYj0r\neu7VvXjj2GuH8e238PHHWvWrW0aG1sYkLW0ID6jTaf8GKGTI/9M/Zdsbb1AcHQ02GxiNbOvqoqC4\neAhPODqpqkpDRwPnG7U2JBXNFX32lHOLi4rzFDzkJ+WTEJ0QxNkKISLV6O5jt3Il6PVsT0xkxR13\nDC8YG+V90JwuJ9Yua5+rbk2dTXQ7h16cYYoykRSbRHJsMkkxST4fxxvjPW8K3OdVRk+O9ty361wX\njyx7hIRxCdTYaqhpqeGS7RLWrhv0PLvGEm3xrOhlJWSRkZBBjGF0tszo6oJPP4Wvv+4ZUxRYsgRu\nv33k6xIqS0s5v20buu5uXEYj+cXFY765dKejU+spd22Ltbmz/5xknaIjx5LjCeYy4jPC+g2zECKw\ngt7HbteuXRw7doy2axnYqqqiKArPPvvssCcxEhRFQX3+eW3VYP36Mf8HBqDL0dXvqpu1y4pLHVp7\nE52iIzEm8bqgzb0SF22IvvGDXONvL63W7lZPkOcO+Lzz9fqjoJASl+JZ0ctKyCI9Pn3EtrWCpaoK\nNm2CpqaeMYtFW6XLzQ3dvMYal+riku2Sp3q1xlYz4OsqOTbZU/SQl5g3qNeKEGJ0CWpg98wzz/Du\nu+9y6623EhvrW231u9/9btiTGAmKorDtX/91TK0aqKqKrdvW76pbu/36qkV/Reuj+111s8RYAt5O\nYbB5Iqqq0tTZpAV6LTXU2GqotdUOuNXlplf0pMen+xRnpMalRkSLCKcTdu6E3bt9axxmzoR77glN\nP9+xluNj7bR6TnkobyofsKdctD6aiUkTPa1IkmLDr8n7WLt+o41cv8gV1By7t956i1OnTpGZmTns\nJwym0Zi07XA5aOpo6nPVramzCYdr6K0zzNHmPlfdkmOTiTXEhvW2kKIoJMcmkxybzE3jbgK01ZP6\ntnrPil6NrYYrbVeuW0Fxqk4u2S5xyXaJQ5cOAWDUG8lMyPTJ17NEW8LqZ9DQABs3wqVLPWMxMbBm\nDdx0U+jmNdrZnXYqmis8wdzV9v7PtFVQyEzI9OTJTTBPQK+TXn1CiJHj14rdzJkz2b59O6mBanoV\nBIqi8Pzzz7N8+fKIeveiqiodjg5PoNbY0ejzsa3LNuBxQQMx6AwkxiT2ueqWGJNIlH70d/m3O+3U\ntdb55OsN1OzVmynK5JOvl5mQicloGuEZX09V4cgR2LrVt5/xxIlaZxGLJehTGtVUVeVK2xVP9Wpl\nc+WAld4JxgRP9eqkpEnERcUFcbZCiEhTUlJCSUkJL7zwQvC2Yg8dOsTf//3f8+ijj5Kenu7ztWXL\nlg17EiMhnM+KdakuWrpargva3B8PtJVzI3FRcf2uuiUYE8JqxSlcdNg7PLl67q1cfyt9E2MSffL1\nMhIyPGdvjoS2NvjoIzh7tmdMr4fiYli0aFTX9wRVW3cb5U3lnmCutbu139sadAZyLbmeYC4tLk1e\nZ0KIQQtqjt0rr7zCX/3VX5GQkHBdjl11dfWwJzESQh3YdTu7+111a+5sHnKhgoKCJcYNNbGAAAAg\nAElEQVRCUsy1wK1XADdaKj5DnSfS0tXiU5xxyXbJr4BbQSHNlOaTr5duSg/I9tvZs1pQ532CxLhx\nWoHE+PHDfviACfW1Gwqny0l1S7WnFUlta+2Atx9nGuepXs215I6q1e5IvH6ih1y/yBXUHLuf/vSn\nbN68mTvuuGPYTzhaqKpKm72t31W3gd7h30iULsoTtHkHcO4tU8nRGXnmaDPmNDNFaUWAdr0bOxp9\n8vXqWuuuy2lU0bbtrrRd4VjdMUBb0RkfP96nmXJKbIrfqzrd3fD553D4sO/4woXaSt0A7eVEP9zF\nNu7q1QvNFwZs6RNriPXkyeUn52OONgdxtkII4T+/VuxycnIoKyvDaBy5LaZAC0Tk63Q5ae5s7nPV\nramjya+Ky/7EG+OvC9rcH5uiTLKVEwGcLidX2q745OtdabviVw5ktD76umbKfW2VX7qkFUg0eKUB\nJiRouXT5+YH+jka3TkcnFc0VnmCuqbOp39vqFB0TzBM8rUgyEjIiokpaCBG5groV+8Ybb3Dw4EGe\ne+6563LsdCE6xupG/P0BdTo6+111s3Zah1yooFf0Wm+3PlbdkmKTRjQPS4ROt7ObWlutT77eQAGE\ntwRjgmdFL8OURcU3mezfHYv3MaxFRXDvvRAn+fg35FJd1NpqPdWrF1suDpgCkRSTRH6y1oYkLzFv\n1KQ1CCEiQ1ADu/6CN0VRcAbiuK4RoCgKL7/zMsVzisnIzuh31c2fprb9iTHE9LvqZo42yzv8YRhN\neSLt9vbrmikPdMB7RwecOQNWK8SSTAJZJBuyuO/2TFYuyMBoCO+911Beu5auFs8pD+VN5QP2bjTq\njUxMnOjZYk2OTZaVckbXa28skusXuYKaY1fufZJ4BPnE8Ql/ePMPzCyaSWrm4Fu1KChab7c+Vt2S\nY5OJMcTIHwJxQ3FRcUxOmczklMmAlt9l7bL6NFO+ZLtEl6Oby5fh3Dmt8TBAB41EmRuxFH3DQTsc\n3qtjnGmcT77eONO4Mfsmwu7Uzgx2V69eabsy4O0z4jM81avZ5mzJVxVCjDqDOivW5XJx+fJl0tPT\nw3YL1k1RFG57/TYATBdN3LL0lj5vZ9AZ+l11S4xJjPijpkRkaG1z8c5HDRw+W0MLNdi4RJtSR06u\nk9zcgduYROmiyEjI8MnXS4pJGpVvOlRVpb693lO9WmmtHLApd7wx3nPKw6SkSSHpOyiEEP4I6opd\nS0sLP/zhD3nnnXdwOBwYDAYefvhhfv3rX2OJgG6oep2eCeYJfQZw3ofQCxEK58/Dhx/qsNnSGE8a\n45lNcjLcv9aBwXLZJ1/vavvV6/I+7S5t1arKWuUZizXEXtdMOSE6IdjfWkC029spbyr3bLG2dLX0\ne1u9oic3MddTvZpuSpfXtxBiTPFrxe573/sera2t/PKXvyQnJ4eqqiqeffZZ4uLiePPNN4Mxz0FT\nFIUfb/0xMYYYMq9m8tRDo+94sdFsLOSJOBzw5Zdw4IDv+Ny5cNdd0FcRepejy3P8mTtfz9pl9ev5\nzNHm65opj0SBwHCvndPlpMZW46levWS7NGARU2pcqqd6NTcxVwqThmksvPZGM7l+kSuoK3afffYZ\n5eXlmEzaNsaUKVN44403mDRp0rAnMJLijfF0neui+PbiUE9FCB+XL2ttTK54pYTFxcH990NhYf/3\nizZoh8hPTJroGWvtbvXJ16tpqemzKKilq4WWrhZOXz3tGUuNS/XJ1xsfPz4k6QdNHU2e6tULTRfo\ncnb1e9sYQwyTkiZ5gjlLTPjvGgghRLD4tWKXl5dHSUkJeXl5nrGKigqWLVtGVVVV/3cMIUVR+Nc/\n/CvFc4opLBjgL6UQQaSq2grdl1/2FEgATJ6sBXXx8YF4DpXmzmafZsq1tlq/+i7qFT3p8ek++Xqp\ncakBL87ocnRR0VzhCeYaOxr7va2CwgTzBE8rksyEzDFbLCKEGL2C2u7kf//v/81//dd/8eMf/5jc\n3FwqKir453/+Zx5//HGee+65YU9iJIT6SDEhemtpgU2b4MKFnrGoKLjzTpg3b2TPeXWpLurb6n2a\nKV9uu+zX0XZGvZGM+AzPFm6WOQtLtGVQuWuqqlLbWuvJk6u2VuNU+2+VZIm2eKpXJyZOJDYqtt/b\nCiHEaBDUwM7lcvHGG2/w3//939TW1pKZmckjjzzCE088EbaJyRLYRbbRlidy6hR8/DF0eh03m5EB\nDz4IqYPvxBMQdqedutY6n3y9ho6GG98RrYWLd75eZkImF6su8uWRLzl96jRF04tYOGMhhmQDZY1l\nlDeVD9i7L0oXRV5inieYG8yRayKwRttrb6yR6xe5gppjp9PpeOKJJ3jiiSeG/YTD1dLSwsqVKzl9\n+jRfffUV06ZNC/WUhOhXZyd8+ikcP94zpiiwdCksXw76ELZRi9JHkW3JJtuS7RnrsHdQ21rrk69n\n67Zdd992ezvnGs9xrvEcAFcvXeXM2TMkTUuiSd/EBccF3v7928yeNrvfHpLj48d7WpFkW7KltZAQ\nQgSAXyt2zzzzDI888giLFy/2jO3bt493332Xl156aUQn2JvD4aC5uZn/9b/+Fz/5yU+YPn16n7eT\nFTsRapWV2tZrc3PPWGIirF0Lubmhm9dgtXS1XNdMudPR6XObg3sO0j7h+lMevHtImqJMnlMe8pPz\niTcGIKFQCCFGiaBuxaamplJTU0N0dLRnrLOzk+zsbOrr64c9iaH4/ve/L4GdCEtOJ5SUwJ49WrGE\n26xZcPfdEBPhR5CqqkpjR6NPvt7GLRuvC+wUFDIaMnjqT54iPymf8fHjZXtVCCH6EfStWJfLN8na\n5XJJ4CRGTKTmiVy9Ch98AJcu9YzFxsKaNdDPe5CIoygKKXEppMSlMDN9JgDtZ9upSKrA1m2j8utK\npsydQmJMIhlxGSzNWRriGYvBiNTXntDI9RN+9QxYunQpP/vZzzzBndPp5Pnnn+fWW28d1JO9/PLL\nzJs3j5iYGL7//e/7fK2xsZG1a9cSHx9PXl4eb7/9tudr//zP/8ztt9/OP/3TP/ncR979i3ChqnDo\nEPzHf/gGdRMnwg9+MHqCuv7cOe9OjFVGMhMyyUjIICUuBcd5B8VzpIekEEIEk19bsdXV1axZs4ba\n2lpyc3OpqqoiIyODjz/+mOzs7Bvd3WPTpk3odDq2bt1KR0cHr7/+uudrjzzyCACvvfYax44dY/Xq\n1ezbt6/f4gjZihXhorUVPvoIzp3rGdPrYeVKWLhwZNuYhJPSslK2Hd1Gt6sbo84oPSSFEGIQgppj\nB9oq3cGDB6muriY7O5sFCxag0w2tSehzzz3HxYsXPYFdW1sbycnJnDp1ioKCAkA7xiwzM5Nf/vKX\n193/nnvu4fjx4+Tm5vKXf/mXfO9737v+G5PATgRBaSn88Y/Q5tXJY9w4rY1Jenro5iWEECKyBDXH\nDkCv17No0SIWLVo07CftPfGzZ89iMBg8QR3ArFmzKCkp6fP+W7Zs8et51q9f7zktIzExkdmzZ3ty\nD9yPLZ+H5+cvvfRSWF+vL74o4dAh6O7WPq+o0L7+yCPLKS6GPXtKOH06fOYbzM+9X7fhMB/5XK7f\nWPpcrl/kfO7+uKKigkDye8UukHqv2O3evZuHHnqI2tpaz21effVVfv/737Njx44hPYes2EW2kpIS\nz4sg3NTUaAUSDV69fBMStDYmYX58clCE87UTNybXL7LJ9YtcQV+xC6TeE4+Pj6elpcVnzGq1kpCQ\nEMxpiTASjr+YXC7YvRt27tQ+dps2De69V6t+FeF57YT/5PpFNrl+4oaBnaqqXLhwgZycHAyGwMSB\nvatZp0yZgsPhoKyszLMde/z4cW666aZhPc+GDRtYvny5/I8uhq2xUWs2XF3dMxYdDffcAzNnjp0C\nCSGEEIFVUlLisz07XDfcilVVFZPJRGtr65CLJdycTid2u50XXniBmpoaXn31VQwGA3q9nkceeQRF\nUfjNb37D0aNHWbNmDfv376eoqGhIzyVbsZEtXLYTVBW+/lo7Fqy7u2c8J0fbek1KCt3cwlW4XDsx\nNHL9Iptcv8gVqLjlhpGaoijcfPPNlJaWDvvJfv7znxMXF8c//MM/8NZbbxEbG8svfvELAP7t3/6N\njo4Oxo0bx2OPPcYrr7wy5KBOiEBob4d339VambiDOp0OVqyA9eslqBNCCBF+/Cqe+NnPfsZbb73F\n+vXryc7O9kSViqLwxBNPBGOegyYrdmI4zp+HDz8Em61nLCUF1q2DrKzQzUsIIcToFNTiiT179pCX\nl8fOnTuv+1q4BnYgOXZi8Ox22LYNDhzwHZ83D+68E4zG0MxLCCHE6BT0HLtIJSt2kS0UeSJ1dVob\nkytXesZMJrj/fpgyJahTiWiS4xPZ5PpFNrl+kSvo7U4aGhr45JNPqKur42/+5m+oqalBVVUmTJgw\n7EkIEUqqCvv3ayt1TmfP+JQpcN99EB8furkJIYQQg+HXit3OnTt58MEHmTdvHnv37sVms1FSUsI/\n/dM/8fHHHwdjnoMmK3bCH1arlkt34ULPWFQU3HUXzJ0rbUyEEEIER1DPip09ezb/+I//yMqVK0lK\nSqKpqYnOzk5ycnK44r1vFUYksBM3cvIkbN4MnZ09Y5mZWoFEamro5iWEEGLsCVq7E4DKykpWrlzp\nMxYVFYXTe98qDG3YsCGgCYkieEbyunV2arl077/fE9QpCixbBk8+KUHdcMlrLrLJ9Ytscv0iT0lJ\nCRs2bAjY4/mVY1dUVMRnn33GqlWrPGPbtm1jxowZAZvISAjkD0qMDhUV2gkSVmvPWGKitkqXkxOy\naQkhhBij3N07XnjhhYA8nl9bsQcOHGDNmjXcc889vPfeezz++ON8/PHHfPTRR8yfPz8gEwk02YoV\n3pxO2LED9u7ViiXcZs+Gu+/WjgcTQgghQiWoOXYANTU1vPXWW1RWVpKTk8Njjz0W1hWxEtgJt/p6\nbeu1trZnLDYW1qyB6dNDNy8hhBDCLeiBHYDL5eLq1aukpaWhhHm5oAR2kS0QvZhUFQ4dgs8/B4ej\nZ3zSJHjgATCbhzdH0TfpoxXZ5PpFNrl+kSuoxRNNTU08/vjjxMbGMn78eGJiYnjsscdobGwc9gRG\nkhRPjF2trfDf/w1btvQEdQYDrFoFjz8uQZ0QQojwEOjiCb9W7B544AEMBgM///nPycnJoaqqir/7\nu7+ju7ubjz76KGCTCSRZsRu7zpyBP/4R2tt7xtLTtQKJ9PTQzUsIIYToT1C3Yi0WC7W1tcTFxXnG\n2tvbycjIwOpdXhhGJLAbe7q74bPP4OhR3/HFi2HFCm3FTgghhAhHQd2KnTp1KhUVFT5jlZWVTJ06\nddgTEKIvg91Cv3gRXnnFN6gzm+G734U775SgLpgk/SGyyfWLbHL9hF9/7lasWMGdd97Jd7/7XbKz\ns6mqquKtt97i8ccf57e//S2qqqIoCk888cRIz1cIHy4X7Nql/XO5esanT9eqXmNjQzc3IYQQItj8\n2op1V9h4V8K6gzlvO3bsCOzshkFRFJ5//nlP4z8x+jQ2am1MLl7sGYuOhtWrYcYMOedVCCFE+Csp\nKaGkpIQXXngh+O1OIonk2I1eqgrHjmn5dN3dPeO5ubB2rXaShBBCCBFJgppjJ0Sw9Zcn0t4O776r\nVb26gzqdDlauhO99T4K6cCA5PpFNrl9kk+snJKVcRIyyMvjwQ61HnVtqqtbGJDMzdPMSQgghwoVs\nxYqwZ7fDl1/CV1/5jt9yi1bxGhUVmnkJIYQQgRKouEVW7ERYq63VCiTq63vGTCbtSLDJk0M3LyGE\nECIc+Z1jd/r0aV588UWefvppAM6cOcOJEydGbGJibNu+vYS9e+E3v/EN6goL4amnJKgLZ5LjE9nk\n+kU2uX7Cr8DuvffeY9myZdTU1PDmm28CYLPZ+NGPfjSikxNjT2lpJf/4j9v5xS++5he/2M7ly5WA\ntt16773w8MPaip0QQgghrudXjt3UqVN55513mD17NklJSTQ1NWG328nIyODq1avBmOegSR+7yHP6\ndCX/8A9lXLxYjMOhjTkc27jrrgKeeiqXlJTQzk8IIYQItJD0sUtJSaG+vh6dTucT2GVlZXHlypVh\nT2IkSPFEZKmshL/92+3U1a3wGc/NhXnztvPMMyv6uacQQggR+YLax27OnDn87ne/8xn7wx/+wPz5\n84c9ATG2tbTAxo3w+uvQ3Nzzv2N7ewk33wwTJ4LTKe0WI4nk+EQ2uX6RTa6f8Ksq9te//jV33HEH\nr732Gu3t7dx5552cPXuWzz//fKTnJ0YphwP279fOeLXbtTGdzoVOBzk54HSCxaKNG42u/h9ICCGE\nEB5+97Fra2tj8+bNVFZWkpOTw+rVq0lISBjp+Q2ZbMWGJ1WFs2e148Camny/lpRUSWVlGWZzsWes\nq2sb69cXUFiYG+SZCiGEEMETqLhFGhSLoLl6VQvoysp8x9PT4e67IS9Pq4rdtu083d06jEYXxcX5\nEtQJIYQY9YIa2FVWVvLCCy9w7NgxWr3Oc1IUhbNnzw57EiNBArvw0dUFO3fCgQPg8tpVjY2F22+H\nefO08169lZSUSDVzhJJrF9nk+kU2uX6RK6gnT/zJn/wJRUVF/PznPycmJmbYTyrGBlWF48e148C8\nz3dVFJg7F1asgLi40M1PCCGEGG38WrGzWCw0Njai1+uDMaeAkBW70KqpgS1btP96y8nRtl0zMkIz\nLyGEECIcBXXFbs2aNf8/e/cdFcXVN3D8u0svi2BZEBQQUBBEsKBGjYBgi71FTDACggV9LLHEQFSs\nT9SIMda4xmDPo0aNSV4rAgZ7jy12xWiwRo2dsu8fG1aW3gQW7+ccz3HvzNy5szPL/vZWEhISaN1a\nu+YSi4qKEhMUl7KnTyE2Fk6c0Ew3M4M2baBePVWNnSAIgiAIbyYoLikFqrG7f/8+7733HnXq1EEu\nl785WCJh+fLlJVaYkiRq7EpXWhocOqTqS/fq1Zt0HR1o0QJatgR9/YLnJ/qJaC9x77SbuH/aTdw/\n7VWqNXYhISHo6+tTt25dDA0N1SeXiKoXAdUo1+3bVaNeM3NxgbZtoXLlsimXIAiCILxrClRjJ5PJ\nuHXrFmZmZqVRphIhauzevocPYccOuHBBM71qVWjfHpycyqZcgiAIgqBtSrXGrn79+jx48ECrAjvh\n7Xn9Gn77DfbvVzXBZjAwAB8faNJE1QQrCIIgCELpKlBg17p1a9q1a0dwcDCWlpYA6qbYkJCQt1pA\nofxQKuHMGdi1S7XGa2YNGoCfH5ialsy5RD8R7SXunXYT90+7ifsnFCiw++2337C2ts5xbVgR2L0b\nkpNV05ckJWmm29iopi+pUaNsyiUIgiAIwhtiSTEhT8+fw549cOyYqsYug4mJavoSDw8xfYkgCIIg\nFNdb72OXedRreuZ1oLKQZl0LSqgQ0tPh6FGIi4MXL96kS6XQrBm0agViERJBEARBKF9yjcoyD5TQ\n1dXN8Z+enl6pFFIoXdevw7ffqppeMwd1Tk4QHq6awuRtB3UlOVmjULrEvdNu4v5pN3H/hFxr7M6e\nPav+/9WrV0ulMELZevwYdu6ETLceAAsL1fQldeqIZldBEARBKM8K1Mfuq6++YsyYMdnSo6Oj+fTT\nT99KwYpL9LEruJQU1dQliYmq/2fQ01M1ub73HugWaJiNIAiCIAhFUVJxS4EnKP7nn3+ypVtYWPD3\n338XuxBvgwjs8qdUwh9/qCYZfvRIc5u7u2pwhJi6UBAEQRDevlKZoHjPnj0olUrS0tLYs2ePxrYr\nV66U+wmLo6Ki8PHxEXP65ODePdi2DbK2sltZqaYvsbMrm3JlKI9zMVWuXLnc/pARBEEQyj8LCwse\nPnyokRYfH1+ifSPzrLGzt7dHIpGQlJSEra3tm4MkEiwtLfn888/p0qVLiRWmJIkau5y9fAnx8XD4\nsGrkawZjY2jdGho2VI18LWvlMbATz5QgCIJQHHl9j5RqU2y/fv1YtWpVsU9WmsSXsKb0dDh5EmJj\n4dmzN+kSCXh5ga8vGBmVXfm0gXimBEEQhOIoN4GdNhJfwm/cvKlqdr19WzPd3l7V7PrvKnFCPsQz\nJQiCIBRHaQR2YqxjBfbPP7B7N5w6pZleqZJqLjpX1/I7fUl5bIoVBEEQhPJOBHYVUGoqHDoECQnw\n+vWbdF1daNECWrZUTWUiCIIgCELFIppiK5hLl2D7dnjwQDO9bl1VLZ2FRdmUqyJ4V58pQRAEoWSU\nRlNsORj/KJSEBw9g7VpYs0YzqKtWDT75BPr0EUGdUHjXr19HKpWyf//+UjunVCpl7dq1byVvHx8f\nwsLC3kreQtHUqlWLGTNmlHUx8vU2n8uCGjVqFOHh4WVaBiFniYmJ2Nvb8+rVq7IuigjstN2rV6p+\ndIsWwcWLb9INDVXLgA0eDA4OZVe+ohLrHZasoKAggoODAdUX1N69e8u4RLlLTk6mZ8+eBd5fKpWS\nkJBATEwMtWrVynNfiUSCpJQ7lk6bNi3fcr3Ljh49yqhRo0okrwMHDtC9e3esrKwwMjLCycmJfv36\nceLEiQLnERoaiq+vb4mUpyRdu3YNhULBF198UdZFEXLQsmVLnJycWLBgQVkXRfSx01ZKJZw+Dbt2\nqQZJZJBIoEED8PMDE5OyK9+76MKFG+zefYWUFCl6eun4+zvi7Fy8mZ5LKs+yCGiKSi6XF/oYbbk2\nbZWSkoLeW+qYW6VKlRLJ5/vvv2fgwIH06tWLtWvX4ujoyP3799myZQsjRowo1z9mCmLRokX4+flh\nbW1d1kUpl16/fo2+vn629Lf57GYVEhJCZGQkn376aZn+TRI1dlro9m1Yvhw2bdIM6mrWhLAw6NJF\n+4M6bRsRe+HCDWJiLnPvXmsePfLh3r3WxMRc5sKFG+Uiz7z6bdy9e5fg4GB1LYeLiwvff/99rvtH\nRkbi6uqKiYkJtra2DBkyhCdPnqi3P3nyhODgYKpXr46hoSG2traMHj1avT0xMZEWLVpgZmaGmZkZ\nnp6e7Ny5U709a5PX06dPGTlyJLa2thgaGlKrVi3++9//Fvo9yM38+fNxcXHByMiIOnXqMGPGDNLS\n0tTb165dS9OmTTE3N6datWp06tSJS5cuaeQxY8YMHB0dMTQ0RC6X0759e16+fElMTAwTJ07kxo0b\nSKVSpFIpU6ZMybEcKSkpfPrpp9SsWRNDQ0Osra3p27evertSqWTChAnI5XJkMhkBAQHMnTtX40sr\nKiqK2rVra+SbmJiIVColKSkJgEePHhEYGIidnR3Gxsa4uLgQHR2tcUxQUBBt2rRh/vz52NvbY2ho\nyKtXr7hz5w5BQUHI5XLMzMxo2bIlv/32W4GvISf29vZMnz5d4/WkSZMYMWIEVapUwcrKik8//VTj\nnmR1+/ZthgwZQlhYGOvWraN169bY2dnRqFEjpk6dys8//wyo/q4MGjRI41ilUomjoyPTpk1j8uTJ\nLF++nISEBPX9WrlypXrfx48f069fP8zMzKhZsyZffvmlRl7//PMPgwYNQi6XY2hoiJeXF7t27VJv\nz+jasGHDBjp16oSJiQmOjo6sWLEiz/cIYM2aNXTv3l0jLaNrwdSpU6levTpVqlShf//+PMs0WWnG\nvcxs9erVSDPNRJ/x3GzYsAEnJydMTEzo2bMnT58+ZcOGDTg7O2NmZkbv3r01PusZec+dOxcbGxtM\nTEz48MMP1Sv0xMfHo6ury59//qlx/pUrV2Jubs6LFy9yvNZr167Ro0cPdZ7169dn9erV2a49NDSU\nCRMmUL16dezt7dWfs7Vr1/LBBx9gamrKxIkTAQgLC8PJyQljY2McHR2JjIzk9b+jC69evYpUKuXA\ngQMa59i7dy+6urrcvHkTgGXLllG3bl2MjIyoUqUK3t7e3Lp1S71/586duXnzJomJiTleV2kRNXZa\n5Nkz2LMHjh9X1dhlkMnA3x/q1y+/05dUdLt3X8HAwA/NFmQ/fv99D15eRau1O3z4Cs+f+2mk+fj4\nERu7p9C1drn9enzx4gXe3t6YmJioazmuXLnC/fv3c83L2NgYhUJBzZo1uXz5MkOHDmX48OHExMQA\n8MUXX3DixAm2bt1K9erVuXnzJufOnQMgNTWVLl26EBISov7CPHPmDMbGxjmeS6lU0qlTJ/78808W\nLFhA/fr1uXXrFn/88Ue2aytKrWRUVBQxMTHMmzcPT09Pzp07x+DBg3n58qU6AHv9+jUTJ07E1dWV\nJ0+eMHHiRDp27MjZs2fR09Nj06ZNzJw5k7Vr1+Lh4cGDBw9ISEgAICAggAsXLrBmzRqOHj0KgEku\nv7rmz5/Phg0bWLNmDQ4ODiQnJ2v0bfzmm2+YO3cuixcv5r333mPz5s1Mnjw52zXn9x68evUKd3d3\nxowZg4WFBYmJiQwePJjKlSsTFBSk3u/w4cOYmZnx888/I5VKSU1NxdfXFzc3N7Zv3465uTk//PAD\nbdq04eTJk7i4uOR7DTnJ6b7Nnz+f8ePHc/jwYY4fP87HH39MvXr1CAkJyTGP9evX8/r161ybKStV\nqgTA4MGDGThwINHR0er7sGfPHpKSkggNDUUmk3Hp0iWuX7/Opk2bNI4FmDx5MtOnT2fKlCls27aN\nYcOG0aRJE1q3bg2oamyOHTvGmjVrsLW1ZfHixXTq1Inff/8dZ2dndT7jx49n5syZfPPNN3z33XeE\nhobSvHnzbEF5hosXL5KcnEzTpk2zbdu4cSMhISEkJCRw48YNAgICsLOzUz+/Bf1c/PXXX6xcuZIt\nW7bw8OFDevXqRY8ePdDT02Pjxo08efKEnj17MmPGDI2A9vDhw5iYmLBz507u379PWFgYAwYMYNOm\nTfj4+FC7dm2WL1+uDrAAFAoFH3/8MUa5zIr/7Nkz/P39mTx5Mqampvz6668EBwdTo0YNjR/969ev\nJzAwkLi4ONLS0tQ/YD/77DNmzZrF4sWLAdXfEUtLS9atW4elpSWnTp1i0KBB6ASL1c8AACAASURB\nVOnpERUVhYODA23btkWhUPDee+9plLNdu3bUrFmTY8eOMWTIEL7//nu8vb15/Pgxhw8f1ii3TCbD\nzc2NPXv28P777+f7nr81ygqqIl1aaqpSeeCAUvnf/yqVkya9+TdlilK5a5dS+fJlGRfwLYiLiyvr\nImST1zM1d26cctIkpdLbW/Nfu3ZxGvesMP/atYvLlt+kSapzlZRly5YpDQ0Nlbdu3cpx+7Vr15QS\niUS5b9++XPPYtGmT0sDAQP26a9euyqCgoBz3ffjwoVIikSjj4+NzzU8ikSjXrFmjVCqVyt27dysl\nEony2LFjBbmcfPn4+CjDwsKUSqVS+ezZM6WxsbFyx44dGvusWLFCaW5unmseDx48UEokEuX+/fuV\nSqVSGR0draxTp44yJSUlx/2nTp2qtLe3z7dsI0aMULZu3TrX7TY2NsovvvhCI61Xr15KPT099etJ\nkyYpnZycNPb57bfflBKJRHnjxo1c8x4+fLiyTZs26tf9+/dXWlhYKJ89e6ZO+/7775U1atRQpqam\nahzr6+urHDlyZIGuISf29vbK6dOnq1/b2dkpu3btqrFPhw4dlH379s01jyFDhuR5zzK8fPlSWa1a\nNeWyZcvUaQEBAcpu3bqpXw8YMEDp4+OT7ViJRKIcMWKERlrdunWVn3/+uVKpVCovXbqklEgkym3b\ntmns07BhQ2VISIhSqXzzeZo7d656e1pamlImkymXLl2aa7l//vlnpUQiUT59+lQj3dvbW+np6amR\nNmTIEOV7772nft2/f3+lv7+/xj6rVq1SSiQS9etJkyYpdXV1lQ8ePFCnDR06VKmjo6O8f/++Om3E\niBHKxo0ba+Qtk8mUT548Uaft3LlTKZFIlFeuXFEqlarPh52dnTI9PV2pVCqV58+fV0okEuXJkydz\nvd6cdO3aVf3Zzbh2Z2dnjX0y3t9p06blm190dLSydu3a6tebNm1SmpiYqK/l77//VhobGyu3bNmi\n3l6pUiWNa81Jly5dlB999FGu2/P6HimpuEU0xZZzV6/CkiWqKUxevnyTXqcOhIerauoMDMqufIKK\nnl56juk6OjmnF4RUmvOx+vpFzzOrY8eO4ebmVqh+O5s2baJVq1bY2Nggk8kIDAwkJSWF5ORkAMLD\nw9m4cSPu7u6MHDmS7du3q39JW1hYEBoaSrt27fjggw+YOXMmFzOP+smhfBYWFjRs2LB4F5qDs2fP\n8uLFC3r06IFMJlP/Gzx4ME+ePOHBv8PLT548Sffu3XFwcMDMzAw7O1Vt6Y0bqibxPn36kJKSgp2d\nHcHBwaxevZqnT58WujzBwcGcPn0aJycnhgwZwqZNm0hJSQFUzdu3b9+mefPmGse0aNGi0NMjpKen\n8+WXX+Lp6Um1atWQyWR8++236qbaDHXr1tWoST1y5AjJycmYm5trvF+JiYlcvnw532soKIlEgqen\np0Za9erVuXPnTq7HKJXKAr0PBgYGBAUFoVAoAHjw4AFbtmwp8EjprOWytrbm7t27AOpa6VatWmns\n06pVK86ePZtrPlKpFLlcnuf1PX78GMhe2yuRSPDw8NBIy++9yo2NjQ2VK1dWv7a0tMTKykqjD6Sl\npaX6ejO4uroik8nUrzOe0Yz345NPPuHu3bvs2LEDUDVnNm7cOFu5M3v+/Dnjx4+nXr16VKlSBZlM\nxv/93/9le0YbNWqU4/FNmjTJlqZQKGjatClWVlbIZDIiIiI08uvcuTOVKlVizZo1gKq52tzcnM6d\nOwPQtm1bHBwcqFWrFn379kWhUKj/RmQmk8l49OhRrtdWGkRTbDn16BHs2AHnz2umV6kC7dqpAruK\nTNv62Pn7OxITE4uPz5um01evYgkKciJTC0yhXLigytPAQDNPPz+n4hZXQ2ECg0OHDvHhhx8SERHB\nnDlzsLCw4MCBA/Tv31/dX6Vt27YkJSWxY8cO4uPjCQwMxN3dndjYWKRSKUuXLmXEiBHs3LmTXbt2\nMWHCBBYsWMDAgQNL9Lryk56uCpA3btxInRw+UBYWFjx//py2bdvSqlUrYmJisLS0RKlU4ubmpr5e\na2tr/vjjD+Li4tizZw9Tp07ls88+49ChQ9SoUaPA5fHw8ODatWvs2rWLuLg4RowYwYQJEzh48GCB\n85BKpdnuZ9bAas6cOXz55Zd8/fXXNGjQAJlMRnR0NL/++qvGflmbx9PT06lbty5btmzJdt6MffO6\nhsxf/vnJ2gleIpGo71dOXFxcePLkCbdu3cLGxibPvAcNGsScOXM4ffo0sbGxyOVyOnToUKRyAXmW\nC3L+fBX2+szNzQFVE2XW4C6/vAryTADZBhhIJJIc07KWM7+/H1WqVKFXr14oFAr8/PxYuXJlvtPb\njB07lq1btzJ37lycnZ0xNjZm9OjR6gA3oyy5dWvImr5hwwaGDRvGzJkz8fb2xszMjPXr1xMZGane\nR1dXlwEDBqBQKBg8eDDLli0jODhY3RfRxMSEo0ePsm/fPnbv3s2SJUsYN24csbGxGj88Hz9+jEUZ\nzy0mauzKmZQUiIuDBQs0gzp9fWjTBoYMqfhBnTZydrYjKMgJuXwP5ubxyOV7/g3qij4q9m3kmVXj\nxo05d+6cRgfgvCQmJlK1alWmTJmCl5cXTk5O6o7FmVlYWBAQEMCSJUv49ddfSUhI4HymB9rNzY1R\no0bxf//3fwwYMIClS5fmeL5GjRrx999/c+zYsaJdYB7c3NwwNDTkypUrODg4ZPsnlUo5f/489+/f\nZ/r06bRq1QpnZ2cePnyY7ctMX1+fdu3aMXPmTE6fPs3z58/56aef1Nvy6vifmYmJCd26dWPevHkc\nPXqU8+fPs3fvXszMzLCxsWHfvn0a++/bt0+j/5RcLufu3bsaX77Hjx/XOGbv3r106NCBoKAgPDw8\ncHBw4OLFi/n2w/Ly8uLq1avIZLJs75WVlVW+1/A29e7dGwMDA6ZNm5bj9ozO/ACOjo60bt0ahULB\nd999R0hIiMa1F+Z+ZT7Ozc0NQN2/MsPevXtxd3cv8LXkJKPvXUYtcWFYWlpyO8tC4VmfieI4f/48\n/2QaxZfRp9LV1VWdNmjQIH7++WeWLFnCy5cv8x1Q89tvvxEYGEivXr1wd3enVq1aXLhwocgjTffu\n3UuDBg0YOXIkDRo0wNHRkWvXrmXLLzQ0lFOnTrFkyRJOnz5NaGioxnapVMr777/P5MmTOXbsGNWr\nV882t+GNGzdy/KFYmrSuxu7w4cOMHDkSPT09bGxsWLlyJbq6WncZ2SiVcO4c7NwJmX6UAODhoWpy\nLcQPXq2njWvFOjvblWjQ9bbyzKxv377MmjWLLl26MGvWLBwcHLh69SoPHjzgww8/zLa/i4sL9+7d\nY/ny5fj4+JCYmKjuoJwhMjKSxo0b4+rqilQqZfXq1chkMmxtbbl8+TIKhYIuXbpQo0YNbt++zW+/\n/ZZrk4qfnx/vv/8+ffr0ITo6Gnd3d27fvs0ff/zBgAEDCn29mZvsTE1NiYiIICIiAolEgp+fH6mp\nqZw+fZqTJ0/y5ZdfYmdnh4GBAd988w2ffvop169fZ/z48RpfCN999x1KpRIvLy/Mzc2JjY3ln3/+\nUX+x1apVi+TkZA4ePKgecZhTp/HZs2djY2ODh4cHxsbGrFu3Dl1dXfWXxOjRo5kwYQIuLi40bdqU\nrVu3Ehsbq5FH69atef78ORMnTiQ4OJjjx4+zaNEijX1cXFxYtWoV8fHxWFtbs3LlSg4fPpxvLcPH\nH3/M3Llz6dixI9OnT6d27drcuXOHPXv24OrqSteuXfO9htzuSV6vC8La2poFCxYwaNAgHj16RFhY\nGA4ODjx8+JCffvqJ+Ph4jYBr0KBBfPzxx6Snp2f78nZwcGDjxo2cO3dOPfo3p5q6jLJmlNfR0ZHe\nvXsTHh7Ot99+qx48ce7cOX744Yc8y5/fNdepUwcrKysOHTqkETAVpAna39+fmTNnsmjRItq1a8ee\nPXvYsGFDnscUhkQi4ZNPPmHatGk8ePCAoUOH0rVrVxwyTaDaokULnJ2dGTt2LP3798+1pi2Ds7Mz\nW7ZsoUePHpiYmBAdHc1ff/2l8QOioM3voHrmly9fztatW3Fzc+OXX35h8+bN2Y63tbWlffv2jBw5\nEn9/f+zt7dXbtm7dytWrV3n//fepVq0ax44d4+bNm+qAHlSjos+dO1fm311aV2Nna2tLXFwcCQkJ\n2Nvbq38Va7M7d2DFCtiwQTOos7aGAQOge/d3K6gTSo+RkREJCQnUq1ePgIAAXF1d+c9//sPLTB06\nMwcxHTt2JDIykoiICOrXr8/69euZPXu2xj5GRkZMnDiRxo0b4+XlxZkzZ9i2bRsymQxTU1MuX75M\nQEAAzs7O9OrVixYtWuQ5qeevv/7KBx98wODBg3FxcaFfv3459m0piKwjBL/44guio6NRKBR4enry\n/vvvM2/ePPWEwlWrVmX16tXs2rWLevXqMW7cOObMmaMxVUTlypX5/vvv8fX1xdXVla+//hqFQqGe\n5LZ79+707t2bjh07IpfLmT17do5lq1SpEtHR0TRv3pz69evz008/8eOPP6pra0aMGMHw4cMZNWoU\nDRo04NChQ0ycOFHjy6lOnTooFArWrVuHu7s7MTExzJgxQ+OaJ0yYgLe3N127dqV58+Y8fvyY4cOH\na+yT00hKAwMDEhISaNy4McHBwTg7O9OzZ0+OHj2q/gLM7xpyuyd5vc6tPFkNGDCAhIQEdY2Qi4sL\nvXv35uLFi9ne827dumFubk779u2zNd0OGDAALy8vmjdvjlwuzzMoy1quZcuW0a5dOwIDA/H09OTA\ngQP88ssvGoFtbteXn8DAQDZv3pzn+XNK8/PzY9q0acyYMQNPT0/i4+OZOHFivve7oGlNmjShZcuW\ntGnThg4dOuDh4cHy5cuzlT80NJTXr18XqMvF3LlzsbOzw9fXF39/f2rWrEmvXr3yLXNGelaDBg2i\nX79+BAcH07BhQ44cOUJUVFSO+4aFheVYTgsLC37++Wc6dOiAs7Mz48ePZ8KECeqJ30EV/NWsWTNb\nP8vSptVrxU6aNIkGDRrQrVu3bNu0YV3PFy9Uza5HjmhOX2JsrKqh8/QEqdaF3hWXNjxTwrslJiaG\nsLCwQg9QeNc9ePCAmjVr8r///U/dOb68u379OvXq1ePChQv59iMsLUFBQdy6dUtjrr7cZPRHexvd\nKkrSokWLmDp1Kjdv3ix0a6Cfnx8dOnRgzJgxue4j1orNw40bN9i1a5fWfCgzS0+Ho0dh/nw4fPhN\nUCeVQrNmMHw4NGwogjpBEISSlJqaSnJyMpGRkdSoUUOrvj/s7e0ZOHCgVqyrm9njx485cuQICoWi\nxJaOexuePXvGH3/8waxZsxg6dGihg7rExESuXr3K8OHD31IJC65UQ4cFCxbQuHFjDA0NNaovAR4+\nfEj37t0xNTXF3t6edevWqbfNnTsXX19f5syZA6iG/n/yySesWLECHR2d0ryEYktKgqVL4Zdf4Pnz\nN+kODqp1Xdu3V63z+q4Ta8UKQsGI5dQKLjExEWtra3bv3l2g1R7Km+joaBYuXFjWxVArSBN5165d\n8fb2pkePHgQGBpZSyQpv6NCheHh44O7uztixYwt9fMuWLbl27Vqu/TFLU6k2xW7evBmpVMqOHTt4\n8eKFxrJFGaNkvvvuO06cOEHHjh3Zv3+/RkdReDNz/ZgxY9SzfeekvDWbPXmiWtf19GnNdHNz1fQl\nLi5i1YjMyuPgifL2TAmCIAjapTSaYsukj92ECRP4888/1YHds2fPqFy5MmfPnsXJSTVHV//+/bG2\nts62JuSqVasYNWqUevj4kCFDchy9J5FI6N+/v7pTr7m5OZ6enupgIaNG6G2/btnShwMHYMWKeFJT\nwd5etf3mzXjc3WHoUB/09EqvPOJ10V/7+vqKwE4QBEEoMolEQlxcnPp1fHw8169fB2DFihXaG9h9\n8cUX3Lp1Sx3YnThxgpYtW2osXBwdHU18fDxbt24t0jnKunZFqYSLF1WTDD98qLnNzQ3atoVMSxAK\nWqCsnylBEARBu5VGjV2ZTACXtU3+6dOnmJmZaaTJZDKNSQ+1yf37qiXA/l1lR83SEjp0gExT4wi5\nKI9NsYIgCIJQ3pVJYJc1IjU1NeXJkycaaY8fPy7UEjTlwatXkJAABw+qRr5mMDICX19o3FiMdBUE\nQRAE4e0pFzV2derUITU1lcuXL6v72J06dYp69eoV6zxRUVH4+Pi89ZofpRJOnYLduyHz2t8SCTRq\nBK1bq+amEwpO1NYJgiAI74L4+PgSnQmiVPvYpaWlkZKSwuTJk7l16xYKhQJdXV10dHTo27cvEomE\nZcuWcfz4cTp16sSBAweoW7dukc5VWv2hbt2Cbdvgzz81021tVc2u1au/9SIIpUT0sRMEQRCKo8JN\nUDx16lSMjY2ZOXMmq1evxsjIiOnTpwOq2Z5fvHiBXC4nMDCQJUuWFDmoKw1Pn8JPP4FCoRnUmZlB\nz54QHCyCuuIQ89i9e2rVqvVWJ1+VSqXZFuwWys7169eRSqXqReOLa9SoUYSHhxf6uNDQUD777LMS\nKUNBxMTEoKenV2rnK01BQUG0adOmrIvxzivVwC4qKor09HSNfxMnTgRU67Bt3ryZp0+fcv36dQIC\nAkqzaAWWlgYHDqhWjThx4k26jg60agXDhoG7u5iTTihfgoKC1JOCS6VS9u7d+9bPOW3aNPWaqwVx\n9OjRQs1MHx8fr16zNfP1lSf+/v7lslzlga2tLcnJyTRp0qTYeV27dg2FQsEXX3yhkf7gwQPGjRuH\ni4sLRkZGWFpa4u3tzapVq0hLSwNU028tWrSIW7duaRwrlUrV/0xNTfH09MxxDdS86OrqsnLlyuJd\nnJbJb8JiqVSKrq4uZ86c0Ugv7N+L4oiJidG4v1ZWVnTu3DlbmbRVmfSxKy0l3cfuyhVVs+v9+5rp\nLi6q6UsqVy6R0whoZx+7C5cvsPvYblKUKehJ9PBv5I+zk3O5yLMgM8SXldevX6Ovr0+VKlWKlU95\nvT5tl3F/SppUKkUul5dIXosWLcLPzw9ra2t12s2bN2nZsiX6+vpMmTKFBg0aoKenx759+/jqq6/w\n8PCgfv362NnZ8d5777F06VImT56ske/ChQvp2bMnT548Yfny5YSGhlKpUiV69uxZoHK9i903CnK9\nBgYGjB07lm3btpVCiXKmo6OjDuavXr3KiBEjaN++PefPny/1gZsl3ceuQo/RzAjsiuvvv+GHH2DV\nKs2grmpVCAyEgAAR1L3rLly+QExcDPcs7/HI6hH3LO8RExfDhcsXykWeuf2xzWgOW7duHe3atcPE\nxARXV1cSExNJSkqiffv2mJqa4ubmRmJiosaxYWFhODk5YWxsjKOjI5GRkbx+/RpQ/SKeOHEiN27c\nUP8qnjJlCqBa83LChAmEh4dTtWpVvL291ekZXTMuX75MpUqV+Prrr9XnO3/+PCYmJixbtqxQ15ib\np0+fMmLECGrUqIGJiQkNGzZk8+bNGvtERkbi6uqKiYkJtra2DBkyRGME/5MnTwgODqZ69eoYGhpi\na2vL6NGjAVUt4p49e1ixYoX6PcitpvTPP/+kZ8+eVKtWDSMjIxwdHfnqq6/U2x8+fEifPn0wNTXF\nysqKCRMm0L9/f41mLx8fH8LCwjTyzVoLcvz4cTp06IClpSUymYwmTZqwY8cOjWNyuz/Hjh2jbdu2\nyGQy5HI5PXv2JCkpqcDXkFXWptiM1xs2bKBTp06YmJjg6OhYoKW/1qxZQ/fu3TXSwsPDSUlJ4fjx\n4/Tt2xcXFxccHR355JNPOH78uHqgHkD37t1ZvXp1tnwrVaqEXC7HycmJGTNmULt2bTZv3sy1a9eQ\nSqUcOHBAY/+9e/eiq6tLUlIS9vb2pKWlERwcjFQqzbb85f79+2nYsCEmJiY0btyYo0ePamw/ePAg\nrVq1wtjYmMqVK/Pxxx9z79499faoqChq167N1q1bcXFxwdTUFF9fXy5nnWcri127duHj40OVKlUw\nNzfHx8eHI0eOaOwjlUpZvHgx/fr1w8zMjJo1a/Lll19q7JPTM1nQz+B//vMfdu3axe7du3PdJ+P6\nMktMTEQqlaqfu4xm7fj4eNzd3TE2NqZ169YkJycTFxeHp6cnpqamtGnThtu3b2c7h1wuRy6X06xZ\nM+bOncvt27c5ePAgUVFRuLi4ZNs/JCQEf3//Al1jYfj4+BAVFVVi+VXoGrviev0aEhNh/35ITX2T\nbmAAPj7QpImqCVYoedo2j93uY7sxqG1A/PX4N4l68PsPv+PV0qtIeR5OPMzzGs/h+ps0n9o+xB6P\nLXStXX61WRMmTCA6OpoFCxbw2WefERAQQO3atRk5ciTz588nIiKCjz76iKtXr6Krq4tSqcTS0pJ1\n69ZhaWnJqVOnGDRoEHp6ekRFRREQEMCFCxdYs2aN+gvL1NRUfb5vvvmG0aNHc/DgQVL//XBlrlV0\ncnJi8eLFhISE4O3tTd26denTpw+dO3cmNDQ023UVtkZSqVTSuXNnJBIJ69evx9raml27dhEQEMC2\nbdvUyxUaGxujUCioWbMmly9fZujQoQwfPpyYmBhANdn6iRMn2Lp1K9WrV+fmzZucO3dOfY3Xrl3D\n2tqaefPmAaouJzkJDw/n5cuXxMbGYm5uztWrV0lOTlZvHzBgAGfPnuWXX35BLpfz3//+l61bt9K0\naVON9yK/9+Cff/6hb9++REdHo6enx4oVK+jSpQtnzpzR+BLNen/OnTuHj48PY8aMYcGCBepBcG3a\ntOH333/HwMAgx2u4c+dOge9JhvHjxzNz5ky++eYbvvvuO0JDQ2nevHm2L/kMFy9eJDk5WeO9ePjw\nIdu2bWPKlCk51r7o6OhgnGmqgqZNm3Lt2jWSkpKwtbXNtWwGBga8fv2aWrVq0bZtWxQKBe+99556\nu0KhoF27dtja2nL06FGqV69OdHQ0ffr00cgnPT2diIgI5s+fT9WqVRk1ahQffvghly5dQkdHh+Tk\nZNq2bUuXLl1YvHgxjx49Ijw8nF69epGQkKDO56+//mLJkiWsW7cOHR0dQkJCCAkJybOrxbNnzxg2\nbBgeHh6kpqYSHR1N+/btuXTpEpUz1VBMnjyZ6dOnM2XKFLZt28awYcNo0qSJ+rNRkGcyN+7u7gQF\nBTF27FiOHz+e63NbkM90eno6U6ZMYfny5ejq6tKnTx969+6NVCpl6dKlGBgYEBAQwKeffsoPP/yQ\naz6G/y7SnpKSQlhYGNOnT2fv3r20atUKUH12NmzYUOjm+LIgArscKJVw9izs3Kla4zWzBg3Azw8y\nfUcJAinKlBzT00grcp7ppOeY/jr9daHzyrwuc3p69nyHDx9Oly5dAIiIiKBJkyaMHj2arl27Aqqa\nq4YNG3Lx4kVcXV2RSCRMmzZNfbytrS2XL19m8eLFREVFYWhoiImJCTo6Ojk2tzVp0kTdvzY3H330\nEbt37yYgIIDmzZvz7NkzFAqFeruPj4+6n1Tm6yuIhIQEDh48yJ07d9STo4eFhXHgwAHmz5+v/vKK\njIzUuMYZM2bQt29fdWCXlJREgwYN8PJSBe81atRQf9GbmZmhr6+PkZFRvk2OSUlJdO/enfr166vP\nleHy5cv89NNP6poWgOXLlxepP1JG7VuGqVOn8vPPP7NhwwYiIiLU6VnvT1BQEJ06dWLSpEnqtFWr\nVlG5cmV27NhBly5d8ryGwvjPf/5Dr1691OWbP38+8fHxeQZ2Wc93+fJl0tPTs601npuMpScvXryo\nkU9GDVRqaioxMTGcOXOGYcOGATBo0CD69evHvHnzkMlkPHr0iE2bNqkH6FStWhV4U+uXmVKp5Ouv\nv8bT0xNQ1U41a9aMq1evUrt2bRYuXIi5uTkxMTHo6qq+pletWoWnpyeJiYm0bNkSgFevXrFq1Sp1\nN4Zx48bRt2/fPJvPu3XrpvH622+/5ccff2T79u189NFH6vSAgAAGDBgAqH54LFiwgN27d9O6deti\nP5MSiYSpU6dSu3ZtVqxYQVBQUI77FaQGMOO9zHjuBg4cyLhx4zh27BgNGjQAVPcqozUgJ/fu3WPS\npEmYmZnRpEkTqlatygcffIBCoVAHdmvXrsXY2DhbzXB5VOGbYgvbbp2cDDExsHGjZlBnYwOhodC1\nqwjqSoM21dYB6ElyHuWmQ9GrdKW5fDz1pSXf38nDw0P9f0tLSwD1H8rMaXfv3lWnKRQKmjZtipWV\nFTKZjIiICI2mudxIJJICd5jPqB1atWoVa9euLbG+L0eOHOH169fY2Nggk8nU/9asWaPRlLVp0yZa\ntWql3i8wMJCUlBR1bVp4eDgbN27E3d2dkSNHsn379iL1qRo5ciQzZsygWbNmjB8/nt9++029LaMG\nsHnz5uo0PT09dTBZGPfu3SM8PJy6detiYWGBTCbj7NmzGvctp/tz5MgRNm/erPFeVa1alVevXnHp\n0qV8r6EwMoIdeNMPL6+av8ePHwNgYmKiTivsPcgI7h89eqSRHhoaikwmw8jIiNGjR/P5558zcOBA\nADp37kylSpVYs2YNAKtXr8bc3JzOnTvnez6JRKLxmav+7xQKGdd59uxZmjVrpg7qQPV5rFSpEmfP\nnlWnWVtba/RNrV69OkqlUuNzmtW1a9fo168ftWvXplKlSlSqVInHjx9n++xmvg8Z58rItySeyerV\nqzN69GgmTJjAy5cvC3xcVhKJRL12POT+9+vBgwcaz0VaWpr6Wba0tOTq1av8+OOP6oB80KBB/Pjj\nj+rnS6FQ0L9/f417UlLi4+NFU2xBFeaNev4c4uLg6FFVjV0GExNo0wY8PMRIVyF3/o38iYmLwae2\njzrt1aVXBAUEFXkAxYUaqj52BrUNNPL08/UrbnGzyTz9QkbzR05pGbV9GzZsYNiwYcycORNvb2/M\nzMxYv369Rg1XXjJ/Cefl0qVL/PXXX0ilUi5dulSgZp6CSE9Pp1KlStn6uH4IfQAAIABJREFUNQHq\nmo5Dhw7x4YcfEhERwZw5c7CwsODAgQP0799f3Zewbdu2JCUlsWPHDuLj4wkMDMTd3Z3Y2Fj1iN2C\nCAoKon379mzfvp24uDg6dOhA9+7dWbVqVa7HZA1epFJptrSUFM2a5KCgIP78809mz55NrVq1MDQ0\nJCAgQH09GbLeH6VSySeffML48eOzlSOj+a4o15CTrDVNEokkx1rmDObm5oCqiTGj3LVr10YqlXL2\n7NlsNVQ5yfjyzsgrw4wZM+jatSumpqbZat10dXUZMGAACoWCwYMHs2zZMnV/uvxIpVKNZsasn6+C\nDrrI6b3KnE9OOnXqhFwuZ9GiRdSsWRM9PT1atmyZ7RnIqcYvr3yh8AH1uHHjUCgUzJkzJ1uza0Ge\n54z9cnovM/dpzEhTKpUa20+dOoVEIkEul2d75tu3b49cLmflypW8//77HD9+nHXr1hXq+goqY5Bn\n1sE7RVWha+wKIj0dDh9WTV9y5MiboE4qhebN4T//AU9PEdSVNm2bx87ZyZkg3yDkd+WYJ5sjvysn\nyLfoQd3byrOk7N27lwYNGjBy5EgaNGiAo6Mj165d09hHX19f3VRaFM+ePSMgIIC+ffsye/Zshg4d\nypUrV4pbdAAaN27Mo0ePePHiBQ4ODhr/atSoAag6aletWpUpU6bg5eWFk5MTN2/ezJaXhYUFAQEB\nLFmyhF9//ZWEhATOnz8PqN6D1MwddPNgZWVFUFAQK1asYNmyZaxZs4anT5+qmxP37dun3vf169fZ\nOrzL5fJsU3Zk7b/022+/ER4eTqdOnXBzc8PKyqpA72njxo05depUtvfKwcFBIxjK7Rrepowm2hs3\nbqjTKleuTIcOHViwYEG25SpBFSA8f/5c/Trj2Dp16mjsZ2lpiYODQ65N6aGhoZw6dYolS5Zw+vRp\njf6fUPTPgJubGwcPHtQIZE6dOsXjx4+LtSLTgwcPOH/+POPHj6dNmza4uLhgYGCQZw1fhszPUUGf\nyfyYmJgwefJkZs2ala1WVi6Xc/fuXY1g8vjx44XKPz8ODg7UqlUrxx+aUqmUsLAwFAoFCoUCb2/v\nXLsDlDcVusYuP9evq6YvyVrL7+ioWjXi3xpZQSgQZyfnEg+63kaeJcHFxYXly5ezdetW3Nzc+OWX\nX7KNKHVwcCA5OZmDBw/i5OSEiYkJRkZGuf6qz5o+fPhwlEolCxYswNjYmN27d9O3b1/2799f7OYQ\nPz8//P396dGjB7NmzcLd3Z2///6b/fv3Y2RkRGhoKC4uLty7d4/ly5fj4+NDYmIiixcv1sgnMjKS\nxo0b4+rqilQqZfXq1chkMnU/rVq1ahEXF8fVq1cxMzPD3Nw8x7IPGzaMjh07UqdOHV6+fMmmTZuw\ntbXF1NQUJycnunTpwtChQ/n222+Ry+V8+eWX2QImf39/hgwZwsaNG/H09GTjxo0kJiZqBF7Ozs6s\nXr2aFi1akJqaysSJE0lPT9d473O6Pxn9LgMDAxkxYgRVq1bl+vXr/PTTT4wYMYJatWrleQ3FkV8t\nUJ06dbCysuLQoUMafeoWLVpEixYtaNSoEVOmTMHDwwN9fX0OHjzIV199xcqVK9XNdQcPHsTe3r7Q\n/QJtbW1p3749I0eOxN/fX91XL0OtWrXYs2cP7du3R09PT93Ml59hw4Yxb948goKCiIiI4O+//yY8\nPJxWrVrRokWLQpUxMwsLC6pVq8bSpUtxcHDg/v37jBs3DiMjo3yPVSqV6ntR0GeyIAYMGMC8efP4\n7rvvNALo1q1b8/z5cyZOnEhwcDDHjx9n0aJFhc6/OAYMGMDkyZO5ePFiofvxlqV3ssbu8WPYsEHV\nly5zUGdhAX37qqYwEUFd2dK2PnbaLKeRZ/mlZXQcDw4OpmHDhhw5coSoqCiNfbp160bv3r3p2LEj\ncrmc2bNn55p31vT169ezdu1afvjhB/XoxZiYGG7fvl3g5t78bN26lR49ejBq1Cjq1q1Lp06d2LZt\nm3oajI4dOxIZGUlERAT169dn/fr1zJ49W6OcRkZGTJw4kcaNG+Pl5cWZM2fYtm2bui/g6NGjqVq1\nKh4eHlhaWua5ysLIkSNxd3fH29ubFy9eaMzxtXz5cjw9PenUqRM+Pj7UrFmT7t27awQ9/fv3Z+jQ\noQwdOhQvLy9u3brF8OHDNcr7/fffk56eTpMmTejRowcffPABXl5eOTZlZebi4sL+/ft5+vQp7dq1\nw83NjYEDB/Ly5UuNkb55XUNOsp6roM9iVoGBgdl+WNSsWZPjx4/TrVs3oqKiaNSoES1atEChUDBk\nyBDc3NzU+27evJnAwMB8z5OTsLAwXr9+re57l9mcOXM4duwY9vb26r5fuV1T5jS5XM7OnTv5888/\n8fLyonPnztSvX5+NGzdq7F/Y9ytjOpkrV65Qv359QkJCGDVqlLqPX16yni+3Z7KwpFIps2bN4sWL\nFxr516lTB4VCwbp163B3dycmJoYZM2YU+ZkpyHFZWVlZ0bFjR2QymXpAjzYo1bViS5NEImHSpEka\nExSnpKimLklMVP0/g56eatWI996Dt9AvUqgg3sXJRoXyKygoiNu3b7Nz586yLkqZu379OvXq1ePC\nhQvY2NiU2rGgqhmcOnUqN2/efCsd64Wy1aRJE95//33mzJlTIvnl9D2SMUHx5MmTS+Q7pkIHdhmX\nplTCH3/Ajh2QZdAT7u6qwRH/DooSyonyOI+dCOyE8iQoKIhbt26xa9eusi5KufDpp5/y6tUrFi5c\nWKjjwsLCqFKlSrYJePPz7Nkzbt68Sfv27QkNDc22nJmg3e7fv88vv/xCWFgYly5dytbMXlR5fY+U\n1HdMhf95ce+eqh/d1aua6VZWqn50dnZlUy5BEITiKM/LxJWF6OjoIh2XeW7Ewhg6dCjr1q2jbdu2\njB07tkh5COWXXC6ncuXKzJ8/v8SCutJSoWvswsNjkUodqVLlTfRmZKSaYLhhQ9XIV0EoKFFjJwiC\nIBSHqLErprNnW5OaGounJ1SrZoeXF/j6qoI7QRAEQRCEiqbC11np6vrx+PEVBg+GDz4QQZ220LZ5\n7ARBEAShPKjQNXY3b0ZRr54PDRtKyTTSXBAEQRAEoVzIGBVbUip0H7svvlCiowNy+R7Cw1uXdZEE\nLSf62AmCIAjFURp97Cp0U6yODrx6FYufn2NZF0UQBEEQBOGtq9CBnVy+h6AgJ5ydxZwm2kb0sRME\nQRCEwqvQfexE86sgqIjJbAVBEN4NFbrGTtBe5W3VCW03f/58jXUmi8rHxwepVMrixYs10hMTE5FK\npSQlJRX7HPm5fv06UqlU/c/c3JxmzZqxdevWt35uQRCE8k4EdoLwDpDJZFSqVKnY+UgkEgwNDZk8\neTJPnz4tgZIV3datW0lOTubgwYPUrVuXnj17cvjw4TItkyAIQlmr0IFdVFSU6KulpbTxvt24cIE9\nCxcS//XX7Fm4kBsXLpSbPIOCgmjTpk2210uXLsXOzo5KlSrRtWtX7t69m29ePXv2xMDAIM+1NePj\n45FKpdy+fVsjXVdXl5UrVwJvat7WrVtHu3btMDExwdXVlcTERJKSkmjfvj2mpqa4ubmRmJiY7RyV\nK1dGLpfj4uKCQqHAwMCAn376iYSEBHR0dPjzzz819l+5ciXm5ua8ePEi32sUBEEoLfHx8URFRZVY\nfhU+sBNNekJpuHHhApdjYmh97x4+jx7R+t49LsfEFCu4K8k8c1pX9MiRIyQkJLBt2zZ27NjB6dOn\nGTNmTL55GRoaMn36dObOncutW7cKXY6sJkyYwNChQzl58iQuLi4EBATQv39/hgwZwokTJ3B1deWj\njz4iNTU113x1dHTQ0dEhJSUFb29v6tSpw/LlyzX2USgUfPzxxxiJWcoFQShHfHx8SjSwq9CDJwTt\npW0B+ZXdu/EzMIBMNY1+wJ7ff8fOy6toeR4+jN/z5xppfj4+7ImNxc7ZuVB5KZXKbPMjGRoaEhMT\ng56eHgCDBw/m66+/zjcviURCYGAgX3/9NZGRkcTExBSqLFkNHz6cLl26ABAREUGTJk0YPXo0Xbt2\nBSAyMpKGDRty8eJFXF1dNa4J4OXLl3z55Zf8888/+Pv7AzBw4EDmzZvHhAkTkEgk/PHHH+zbt48F\nCxYUq6yCIAjlXYWusROE0iJNSck5PS2t6Hmmp+ec/vp1kfPMzMXFRR3UAVSvXp07d+4U+PjZs2ez\nevVqTp06VaxyeHh4qP9v+e8SMfXr18+WlrWZuG3btshkMkxNTVm0aBFff/01bdu2BaB///7cvXuX\nHTt2ALBs2TIaN26scS5BEISKSAR2QrmkbX3s0jMFSBrpOjpFz1Oa88czXV+/yHlmppelzIWd9dzX\n15cOHTowduzYbE2s0n/Lnjm/tLQ00nMIVjOXIyOfnNKyHhsTE8OpU6e4e/cud+/eZfjw4eptlStX\nplevXigUClJSUli5ciUDBw4s8LUJgiBoK9EUKwglwNHfn9iYGPwyNSHHvnqFU1AQFLLZVJ3nhQuq\nPA0MNPP08ytmaVVy6u9WWLNmzaJ+/fp4ZWlulsvlANy6dQsbGxsATp48WaJLstnY2ODg4JDr9kGD\nBuHr68uSJUt4+fIlffv2LbFzC4IglFcisBPKJW3rY2fn7AxBQeyJjUX6+jXp+vo4+fkVui/c284z\ns6IEWVn76tWtW5cBAwYwd+5cjf2cnJyws7MjKiqKuXPncu/ePSIiIkokmCyoFi1a4OzszNixY+nf\nvz8mJialdm5BEISyIgI7QSghds7OJRZ0lXSeWUfF5jRKNiO9MPkATJkyhbVr12qk6+rq8r///Y/w\n8HAaNGiAs7Mz8+fPx9fXN9/zFSStoAFiaGgoo0aNEs2wgiC8MyTKkmwbKUcK219IKF/i4+PLXa2d\neKa0z7hx44iNjeXYsWNlXRRBEIQ8v0dK6jumQtfYZcxjV94CBEEQ3q7Hjx9z8eJFFAoF8+fPL+vi\nCIIg5Co+Pr5EBwyKGjtBKCDxTGkPHx8fDh8+TN++ffnuu+/KujiCIAhA6dTYicBOEApIPFOCIAhC\ncZRGYCfmsRPKJW2bx04QBEEQygMR2AmCIAiCIFQQoilWEApIPFOCIAhCcYimWEEQBEEQBKHARGAn\nlEuij50gCIIgFJ4I7ARBEARBECoI0cdOEApIPFOCIAhCcYg+doIglIigoCDatGlT1sUAVJMHi7Vb\nBUEQ3g4R2AnlkuhjV7Lmz5/Pxo0bi53PgwcPGD58OA4ODhgaGiKXy2nVqhU//PBDgfPYsmUL0dHR\n6tdBQUFIpVKkUil6enrY29szZMgQHj58WOzyCoIgvGsq9FqxglCaLly9yu6zZ0kB9AB/NzecHRzK\nRZ4ymaxY5cjQs2dPnjx5wtKlS3F2dubevXscOnSoUEGYubl5trRWrVqxfv16UlNTOXr0KGFhYdy8\neZNffvmlRMotCILwrqjQNXZRUVGi5kdL+fj4lHURCuXC1avEHD/OvXr1eFSvHvfq1SPm+HEuXL1a\nLvLM2hSb8Xrp0qXY2dlRqVIlunbtyt27d3PN49GjR+zdu5dp06bh7+9PzZo1adiwIUOGDCE8PFxj\n34ULF+Lq6oqhoSGWlpb06tVLvc3Hx4ewsDCN/fX09JDL5VhbW9OlSxdGjBjB9u3befnyJT4+Pgwa\nNEhjf6VSiaOjI9OnTy/0eyEIglCexMfHExUVVWL5Vegau5J8owQhL7vPnsWgUSPiHz16k+joyO97\n9+IlkRQpz8N79/LcwwMy5enTqBGxZ84UutZOIpEgyVKOI0eOIJfL2bZtG0+ePOGjjz5izJgxrFy5\nMsc8TE1NkclkbNmyBR8fH4yNjXPcb9KkSURHRzNz5kzatm3Ls2fP2LZtW55lyfra0NCQ9PR00tLS\nGDx4MAMHDiQ6OhoTExMA9uzZQ1JSEgMGDCjU+yAIglDe+Pj44OPjw+TJk0skvwpdYydoL22raU3J\nJT2tiEEdQHoux74uQl5KpTLbaCtDQ0NiYmJwdXWlWbNmDB48mN27d+eah66uLitWrGDz5s1YWFjg\n5eXFyJEjiYuLU+/z7NkzZs2axeTJkwkPD8fJyQkPDw/Gjx+fb/kynDt3joULF9KsWTNMTEzo3r07\nhoaGGv34li1bRqdOnbCysirsWyEIglChicBOEEqAXi7pOsUYui7N5Vj9IueoycXFBT29NyWvXr06\nd+7cyfOYbt26cevWLbZv307Pnj05d+4cfn5+DBs2DICzZ8/y6tUr2rZtW6iyxMfHI5PJMDY2xt3d\nHScnJ9asWQOAgYEBQUFBKBQKQDWAY8uWLdmacwVBEIQK3hQraC9t62Pn7+ZGzLFj+DRqpE57dewY\nQa1a4VyrVpHyvKBUEnP8OAZZ8vRr2LDY5QU0gjoo+BxK+vr6+Pr64uvry/jx45k+fToTJkxg3Lhx\nRS5Ls2bNWLFiBbq6ulhbW6Orq/mnadCgQcyZM4fTp08TGxuLXC6nQ4cORT6fIAhCRSUCO0EoAc4O\nDgQBsWfO8BpVrZpfw4bFGhX7NvLMLGu/tqJycXEB4N69e+oBEzt27KBevXoFzsPQ0BCHPK7L0dGR\n1q1bo1AoiIuLIyQkpMTKLwiCUJGIwE4ol+Lj47Wu1s7ZwaHEgq63mWeGws5w/uDBA3r27ElISAj1\n69fH3NycM2fO8Pnnn+Pg4ICnpyc6OjqMHj2aqKgojIyM8Pf358WLF2zbtk3dzy6n/n4FMWjQID7+\n+GPS09MJDQ0t9PGCIAjvAhHYCcI7IOtI1JxGpmak50Ymk9GiRQsWLlzI5cuXefHiBdWrV6ddu3ZE\nRkaio6MDwNSpU6lWrRrffPMNo0aNwsLCAm9v70KXJatu3bphbm5OkyZNsLGxKdB1C4IgvGvEWrGC\nUEDimSpbDx48oGbNmvzvf/+jc+fOZV0cQRCEQhNrxQqC8M5LTU0lOTmZyMhIatSoIYI6QRCEPIjA\nTiiXtG0eO+HtSUxMxNramt27d7NixYqyLo4gCEK5JvrYCYJQrvn4+JCenl7WxRAEQdAKoo+dIBSQ\neKYEQRCE4hB97ARBEARBEIQCE4GdUC6JPnaCIAiCUHgisBMEQRAEQaggtK6P3Z07d+jRowf6+vro\n6+uzdu1aqlSpkm0/0R9KKGmVK1fm77//LutiCIIgCFrKwsKChw8f5ritpOIWrQvs0tPTkUpVFY0r\nVqzgr7/+Ui9VlJkI7ARBEARB0Bbv7OCJjKAO4MmTJ1hYWJRhaYS3RfSx017i3mk3cf+0m7h/gtYF\ndgCnTp2iadOmLFiwgL59+5Z1cYS34OTJk2VdBKGIxL3TbuL+aTdx/4RSDewWLFhA48aNMTQ0JDg4\nWGPbw4cP6d69O6amptjb27Nu3Tr1trlz5+Lr68ucOXMA8PDw4NChQ0ybNo2pU6eW5iUIpeTRo0dl\nXQShiMS9027i/mk3cf+EUg3sbGxsmDBhAiEhIdm2DR06FENDQ+7evcuaNWsYMmQI586dA2DUqFHE\nxcUxevRoUlJS1MeYmZnx6tWrUit/UZRktXhR8yrMcQXZN699irKtvDYdlHS5KuL9K6/3DrTv/hX3\n3uW1Xds+eyD+dua37V25d8XJryTvnzZ99ko1sOvevTtdu3bNNor12bNnbNq0ialTp2JsbEyLFi3o\n2rUrq1atypbHyZMn8fb2pnXr1kRHRzNu3LjSKn6RiD9O+W/LKf369ev5luNtq4h/nPLbpyS+XMrD\nvQPtu3/lJbCriPfvXfnsQfm4f9r22SvIvtoU2JXJqNgvvviCW7du8f333wNw4sQJWrZsybNnz9T7\nREdHEx8fz9atW4t0DicnJ65cuVIi5RUEQRAEQXibHB0duXz5crHz0S2BshSaRCLReP306VPMzMw0\n0mQyGf/880+Rz1ESb44gCIIgCII2KZNRsVkrCU1NTXny5IlG2uPHj5HJZKVZLEEQBEEQBK1WJoFd\n1hq7OnXqkJqaqlHLdurUKerVq1faRRMEQRAEQdBapRrYpaWl8fLlS1JTU0lLS+PVq1ekpaVhYmJC\njx49mDhxIs+fPycxMZGff/6Zfv36lWbxBEEQBEEQtFqpBnYZo15nzpzJ6tWrMTIyYvr06QAsWrSI\nFy9eIJfLCQwMZMmSJdStW7c0iycIgiAIgqDVtG6t2OJ48uQJ/v7+nD9/nkOHDuHq6lrWRRIK4fDh\nw4wcORI9PT1sbGxYuXIlurplMv5HKKQ7d+7Qo0cP9PX10dfXZ+3atdmmPRLKv3Xr1jFixAju3r1b\n1kURCuH69et4eXlRr149JBIJ69evp2rVqmVdLKGA4uPjmTZtGunp6QwfPpxu3brluf87Fdilpqby\n6NEjxo4dy5gxY3BzcyvrIgmFkJycjIWFBQYGBkRERNCoUSN69uxZ1sUSCiA9PV29zvOKFSv466+/\nGD9+fBmXSiiMtLQ0evfuTVJSEkePHi3r4giFcP36dcaOHcuGDRvKuihCIb148YI+ffrw448/oqen\nV6BjtHKt2KLS1dUVv1K0mJWVFQYGBgDo6emho6NTxiUSCiojqANVzbmFhUUZlkYoinXr1vHhhx9m\nG/wmaId9+/bRqlUrIiMjy7ooQiEcOHAAIyMjOnfuTI8ePbhz506+x7xTgZ1QMdy4cYNdu3bRuXPn\nsi6KUAinTp2iadOmLFiwgL59+5Z1cYRCSEtLY8OGDfTp06esiyIUgbW1NVeuXGHv3r3cvXuXTZs2\nlXWRhAK6c+cOly9f5pdffiEsLIyoqKh8j9HKwG7BggU0btwYQ0NDgoODNbY9fPiQ7t27Y2pqir29\n/f+3d6chUfx/HMDfq+bt2lrhFamlJVIpiYJZZkX9WLQDodLII0olD6gHHWJqoRFRgT2w80mFuB4P\ntUAh0QxKE9YIj1YLrCisLHRXa111fg/6O//W459a/9VZ3y9YcGZn5vuZeaN8nGMXKpVq0m3wv865\n8yf59ff3IyEhAffu3eMZuznwJ9kFBgaisbERBQUFyM/PN2XZ9B+zza+4uJhn6+aB2eZnbW0NOzs7\nAEBMTAxevHhh0rpp9tkpFAqEh4fDysoK27ZtQ2tr62/HkuSd556ensjJyUF1dTW+f/9u9F56ejps\nbW3x6dMnqNVqREVFITAwcMKDEgvo1sJ5Z7b5DQ8PIzY2Fnl5efDz85uj6he22WZnMBjE+0Pkcjn0\nev1clL/gzTa/9vZ2qNVqFBcXo7OzE8ePH0dhYeEc7cXCNdv8dDodHB0dAQCPHz/m/eVzYLbZhYSE\n4OrVqwCAlpYWrFq16veDCRJ29uxZISkpSZzW6XSCtbW10NnZKc5LSEgQzpw5I04rlUrBw8NDCAsL\nE+7evWvSesnYTPO7f/++sGTJEiEyMlKIjIwUysrKTF4z/TTT7BobG4WIiAhh69atws6dO4V3796Z\nvGb6r9n87RwTEhJikhppajPN7+HDh0JwcLCwefNmITExURgZGTF5zfTTbH73ioqKhIiICCEyMlJ4\n8+bNb8eQ5Bm7McK4s24ajQZWVlbw9fUV5wUGBqKurk6cfvjwoanKo9+YaX7x8fH80Op5YqbZhYaG\nor6+3pQl0v8wm7+dY5qamv7f5dFvzDQ/pVIJpVJpyhJpCrP53UtLS0NaWtq0x5DkPXZjxt/vodPp\nIJfLjeY5OTlBq9WasiyaJuYnXcxO2piftDE/6TJFdpJu7MZ3vo6Ojujv7zea19fXBycnJ1OWRdPE\n/KSL2Ukb85M25iddpshO0o3d+M539erVGB4eRldXlzjvxYsXWLt2ralLo2lgftLF7KSN+Ukb85Mu\nU2QnycZuZGQEP378wPDwMEZGRqDX6zEyMgIHBwfExMQgNzcXg4ODePLkCSorK3lf1jzD/KSL2Ukb\n85M25iddJs3uz5/xML28vDxBJpMZvc6fPy8IgiB8/fpV2Lt3r+Dg4CB4eXkJKpVqjqul8ZifdDE7\naWN+0sb8pMuU2S2o74olIiIiMmeSvBRLRERERBOxsSMiIiIyE2zsiIiIiMwEGzsiIiIiM8HGjoiI\niMhMsLEjIiIiMhNs7IiIiIjMBBs7IiIiIjPBxo6IaJykpCTk5OT81W0eO3YMBQUFf3WbRETjWc11\nAURE841MJpvwZd1/6saNG391e0REk+EZOyKiSfDbFolIitjYEdG8cunSJSxfvhxyuRz+/v6ora0F\nADQ1NSEsLAwKhQIeHh7IzMyEwWAQ17OwsMCNGzfg5+cHuVyO3NxcvH79GmFhYVi8eDFiY2PF5evq\n6rB8+XJcvHgRy5Ytg4+PD0pKSqasqaqqCkFBQVAoFAgPD8fLly+nXPbEiRNwdXWFs7Mz1q9fj7a2\nNgDGl3d37doFJycn8WVpaYn79+8DADo6OrBjxw4sWbIE/v7+qKiomHKsyMhI5ObmYtOmTZDL5fjn\nn3/Q29s7zSNNROaIjR0RzRuvXr1CUVERmpub0d/fj5qaGnh7ewMArKyscO3aNfT29uLp06d49OgR\nrl+/brR+TU0N1Go1nj17hkuXLiE5ORkqlQpv377Fy5cvoVKpxGV7enrQ29uLDx8+4N69e0hJSUFn\nZ+eEmtRqNY4cOYI7d+7g69evSE1Nxe7duzE0NDRh2erqajQ0NKCzsxN9fX2oqKiAi4sLAOPLu5WV\nldBqtdBqtSgvL4e7uzu2b9+OgYEB7NixA4cOHcLnz59RWlqKtLQ0tLe3T3nMVCoV7t69i0+fPmFo\naAhXrlyZ8XEnIvPBxo6I5g1LS0vo9Xq0trbCYDBgxYoVWLlyJQBgw4YNCA0NhYWFBby8vJCSkoL6\n+nqj9U+dOgVHR0cEBARg3bp1UCqV8Pb2hlwuh1KphFqtNlo+Pz8fixYtQkREBKKiolBWVia+N9aE\n3b59G6mpqQgJCYFMJkNCQgJsbGzw7NmzCfVbW1tDq9Wivb0do6OjWLNmDdzc3MT3x1/e1Wg0SEpK\nQnl5OTw9PVFVVQUfHx8kJibCwsICQUFBiImJmfKsnUwmw+HDh+EKxs8YAAAC2UlEQVTr6wtbW1vs\n378fLS0tMzjiRGRu2NgR0bzh6+uLwsJCnDt3Dq6uroiLi8PHjx8B/GyCoqOj4e7uDmdnZ2RnZ0+4\n7Ojq6ir+bGdnZzRta2sLnU4nTisUCtjZ2YnTXl5e4li/6u7uxtWrV6FQKMTX+/fvJ11269atyMjI\nQHp6OlxdXZGamgqtVjvpvvb19WHPnj24cOECNm7cKI7V2NhoNFZJSQl6enqmPGa/No52dnZG+0hE\nCw8bOyKaV+Li4tDQ0IDu7m7IZDKcPn0awM+PCwkICEBXVxf6+vpw4cIFjI6OTnu7459y/fbtGwYH\nB8Xp7u5ueHh4TFhvxYoVyM7Oxrdv38SXTqfDgQMHJh0nMzMTzc3NaGtrg0ajweXLlycsMzo6ioMH\nD2L79u04evSo0VhbtmwxGkur1aKoqGja+0lECxsbOyKaNzQaDWpra6HX62FjYwNbW1tYWloCAHQ6\nHZycnGBvb4+Ojo5pfXzIr5c+J3vKNS8vDwaDAQ0NDXjw4AH27dsnLju2fHJyMm7evImmpiYIgoCB\ngQE8ePBg0jNjzc3NaGxshMFggL29vVH9v46fnZ2NwcFBFBYWGq0fHR0NjUaD4uJiGAwGGAwGPH/+\nHB0dHdPaRyIiNnZENG/o9XpkZWVh2bJlcHd3x5cvX3Dx4kUAwJUrV1BSUgK5XI6UlBTExsYanYWb\n7HPnxr//67Sbm5v4hG18fDxu3bqF1atXT1g2ODgYd+7cQUZGBlxcXODn5yc+wTpef38/UlJS4OLi\nAm9vbyxduhQnT56csM3S0lLxkuvYk7EqlQqOjo6oqalBaWkpPD094e7ujqysrEkf1JjOPhLRwiMT\n+O8eES0wdXV1iI+Px7t37+a6FCKiv4pn7IiIiIjMBBs7IlqQeMmSiMwRL8USERERmQmesSMiIiIy\nE2zsiIiIiMwEGzsiIiIiM8HGjoiIiMhMsLEjIiIiMhP/Ah/1RvhmnkuIAAAAAElFTkSuQmCC\n", + "text": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "# Bonus: How to use Cython without the IPython magic" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "IPython's notebook is really great for explanatory analysis and documentation, but what if we want to compile our Python code via Cython without letting IPython's magic doing all the work? \n", + "These are the steps you would need." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### 1. Creating a .pyx file containing the the desired code or function." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%%file ccy_classic_lstsqr.pyx\n", + "\n", + "def ccy_classic_lstsqr(x, y):\n", + " \"\"\" Computes the least-squares solution to a linear matrix equation. \"\"\"\n", + " x_avg = sum(x)/len(x)\n", + " y_avg = sum(y)/len(y)\n", + " var_x = sum([(x_i - x_avg)**2 for x_i in x])\n", + " cov_xy = sum([(x_i - x_avg)*(y_i - y_avg) for x_i,y_i in zip(x,y)])\n", + " slope = cov_xy / var_x\n", + " y_interc = y_avg - slope*x_avg\n", + " return (slope, y_interc)" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "Writing ccy_classic_lstsqr.pyx\n" + ] + } + ], + "prompt_number": 11 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### 2. Creating a simple setup file" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%%file setup.py\n", + "\n", + "from distutils.core import setup\n", + "from distutils.extension import Extension\n", + "from Cython.Distutils import build_ext\n", + "\n", + "setup(\n", + " cmdclass = {'build_ext': build_ext},\n", + " ext_modules = [Extension(\"ccy_classic_lstsqr\", [\"ccy_classic_lstsqr.pyx\"])]\n", + ")" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "Writing setup.py\n" + ] + } + ], + "prompt_number": 12 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
\n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "####3. Building and Compiling" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "!python3 setup.py build_ext --inplace" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "running build_ext\r\n" + ] + }, + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "cythoning ccy_classic_lstsqr.pyx to ccy_classic_lstsqr.c\r\n" + ] + }, + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "building 'ccy_classic_lstsqr' extension\r\n", + "creating build\r\n", + "creating build/temp.macosx-10.6-intel-3.4\r\n", + "/usr/bin/clang -fno-strict-aliasing -Werror=declaration-after-statement -fno-common -dynamic -DNDEBUG -g -fwrapv -O3 -Wall -Wstrict-prototypes -arch i386 -arch x86_64 -g -I/Library/Frameworks/Python.framework/Versions/3.4/include/python3.4m -c ccy_classic_lstsqr.c -o build/temp.macosx-10.6-intel-3.4/ccy_classic_lstsqr.o\r\n" + ] + }, + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "\u001b[1mccy_classic_lstsqr.c:2040:28: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function '__Pyx_PyObject_AsString'\r\n", + " [-Wunused-function]\u001b[0m\r\n", + "static CYTHON_INLINE char* __Pyx_PyObject_AsString(PyObject* o) {\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mccy_classic_lstsqr.c:2037:32: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function\r\n", + " '__Pyx_PyUnicode_FromString' [-Wunused-function]\u001b[0m\r\n", + "static CYTHON_INLINE PyObject* __Pyx_PyUnicode_FromString(char* c_str) {\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mccy_classic_lstsqr.c:2104:26: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function '__Pyx_PyObject_IsTrue'\r\n", + " [-Wunused-function]\u001b[0m\r\n", + "static CYTHON_INLINE int __Pyx_PyObject_IsTrue(PyObject* x) {\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mccy_classic_lstsqr.c:2159:33: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function '__Pyx_PyIndex_AsSsize_t'\r\n", + " [-Wunused-function]\u001b[0m\r\n", + "static CYTHON_INLINE Py_ssize_t __Pyx_PyIndex_AsSsize_t(PyObject* b) {\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mccy_classic_lstsqr.c:2188:33: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function '__Pyx_PyInt_FromSize_t'\r\n", + " [-Wunused-function]\u001b[0m\r\n", + "static CYTHON_INLINE PyObject * __Pyx_PyInt_FromSize_t(size_t ival) {\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mccy_classic_lstsqr.c:1584:32: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function '__Pyx_PyInt_From_long'\r\n", + " [-Wunused-function]\u001b[0m\r\n", + "static CYTHON_INLINE PyObject* __Pyx_PyInt_From_long(long value) {\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mccy_classic_lstsqr.c:1631:27: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1mfunction '__Pyx_PyInt_As_long' is not\r\n", + " needed and will not be emitted [-Wunneeded-internal-declaration]\u001b[0m\r\n", + "static CYTHON_INLINE long __Pyx_PyInt_As_long(PyObject *x) {\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mccy_classic_lstsqr.c:1731:26: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1mfunction '__Pyx_PyInt_As_int' is not\r\n", + " needed and will not be emitted [-Wunneeded-internal-declaration]\u001b[0m\r\n", + "static CYTHON_INLINE int __Pyx_PyInt_As_int(PyObject *x) {\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m" + ] + }, + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "8 warnings generated.\r\n" + ] + }, + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "\u001b[1mccy_classic_lstsqr.c:2040:28: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function '__Pyx_PyObject_AsString'\r\n", + " [-Wunused-function]\u001b[0m\r\n", + "static CYTHON_INLINE char* __Pyx_PyObject_AsString(PyObject* o) {\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mccy_classic_lstsqr.c:2037:32: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function\r\n", + " '__Pyx_PyUnicode_FromString' [-Wunused-function]\u001b[0m\r\n", + "static CYTHON_INLINE PyObject* __Pyx_PyUnicode_FromString(char* c_str) {\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mccy_classic_lstsqr.c:2104:26: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function '__Pyx_PyObject_IsTrue'\r\n", + " [-Wunused-function]\u001b[0m\r\n", + "static CYTHON_INLINE int __Pyx_PyObject_IsTrue(PyObject* x) {\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mccy_classic_lstsqr.c:2159:33: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function '__Pyx_PyIndex_AsSsize_t'\r\n", + " [-Wunused-function]\u001b[0m\r\n", + "static CYTHON_INLINE Py_ssize_t __Pyx_PyIndex_AsSsize_t(PyObject* b) {\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mccy_classic_lstsqr.c:2188:33: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function '__Pyx_PyInt_FromSize_t'\r\n", + " [-Wunused-function]\u001b[0m\r\n", + "static CYTHON_INLINE PyObject * __Pyx_PyInt_FromSize_t(size_t ival) {\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mccy_classic_lstsqr.c:1584:32: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function '__Pyx_PyInt_From_long'\r\n", + " [-Wunused-function]\u001b[0m\r\n", + "static CYTHON_INLINE PyObject* __Pyx_PyInt_From_long(long value) {\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mccy_classic_lstsqr.c:1631:27: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1mfunction '__Pyx_PyInt_As_long' is not\r\n", + " needed and will not be emitted [-Wunneeded-internal-declaration]\u001b[0m\r\n", + "static CYTHON_INLINE long __Pyx_PyInt_As_long(PyObject *x) {\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mccy_classic_lstsqr.c:1731:26: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1mfunction '__Pyx_PyInt_As_int' is not\r\n", + " needed and will not be emitted [-Wunneeded-internal-declaration]\u001b[0m\r\n", + "static CYTHON_INLINE int __Pyx_PyInt_As_int(PyObject *x) {\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m" + ] + }, + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "8" + ] + }, + { + "output_type": "stream", + "stream": "stdout", + "text": [ + " warnings generated.\r\n", + "/usr/bin/clang -bundle -undefined dynamic_lookup -arch i386 -arch x86_64 -g build/temp.macosx-10.6-intel-3.4/ccy_classic_lstsqr.o -o /Users/sebastian/Github/python_reference/benchmarks/ccy_classic_lstsqr.so\r\n" + ] + } + ], + "prompt_number": 13 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### 4. Importing and running the code" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "import ccy_classic_lstsqr\n", + "\n", + "%timeit py_classic_lstsqr(x, y)\n", + "%timeit cy_classic_lstsqr(x, y)\n", + "%timeit ccy_classic_lstsqr.ccy_classic_lstsqr(x, y)" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "100 loops, best of 3: 2.9 ms per loop\n", + "1000 loops, best of 3: 212 \u00b5s per loop" + ] + }, + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "\n", + "1000 loops, best of 3: 207 \u00b5s per loop" + ] + }, + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "\n" + ] + } + ], + "prompt_number": 20 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "
\n", + "
" + ] + } + ], + "metadata": {} + } + ] +} \ No newline at end of file diff --git a/benchmarks/palindrome_timeit.ipynb b/benchmarks/palindrome_timeit.ipynb index ef0e33f..afe4a0a 100644 --- a/benchmarks/palindrome_timeit.ipynb +++ b/benchmarks/palindrome_timeit.ipynb @@ -1,7 +1,7 @@ { "metadata": { "name": "", - "signature": "sha256:6ea19109869c82ee989c8ea0599ec49401e74246a542ad0b7b05f6ef464bda19" + "signature": "sha256:ab859015961b19530bb3c16fe1edf4e7f9baca715bdf8187c724e8c0c4cfc8ad" }, "nbformat": 3, "nbformat_minor": 0, @@ -12,8 +12,20 @@ "cell_type": "markdown", "metadata": {}, "source": [ - "Sebastian Raschka 04/2014\n", + "[Sebastian Raschka](http://sebastianraschka.com) \n", + "last updated: 05/06/2014\n", "\n", + "- [Link to this IPython Notebook on GitHub](https://github.com/rasbt/python_reference/blob/master/benchmarks/palindrome_timeit.ipynb) \n", + "- [Link to the GitHub repository](https://github.com/rasbt/python_reference) \n", + "\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ "#Timing different Implementations of palindrome functions" ] }, @@ -27,11 +39,9 @@ "\n", "# All functions return True if an input string is a palindrome. Else returns False.\n", "\n", - "\n", - "\n", - "####\n", + "#############################################################\n", "#### case-insensitive ignoring punctuation characters\n", - "####\n", + "############################################################\n", "\n", "def palindrome_short(my_str):\n", " stripped_str = \"\".join(l.lower() for l in my_str if l.isalpha())\n", @@ -56,9 +66,9 @@ "\n", "\n", "\n", - "####\n", + "############################################################\n", "#### functions considering all characters (case-sensitive)\n", - "####\n", + "############################################################\n", "\n", "def palindrome_reverse1(my_str):\n", " return my_str == my_str[::-1]\n", @@ -71,15 +81,12 @@ " return True\n", " if my_str[0] != my_str[-1]:\n", " return False\n", - " return palindrome(my_str[1:-1])\n", - "\n", - "\n", - "\n" + " return palindrome(my_str[1:-1])" ], "language": "python", "metadata": {}, "outputs": [], - "prompt_number": 10 + "prompt_number": 1 }, { "cell_type": "code", @@ -107,7 +114,7 @@ "stream": "stdout", "text": [ "case-insensitive functions ignoring punctuation characters\n", - "100000 loops, best of 3: 15.3 \u00b5s per loop" + "10000 loops, best of 3: 15.5 \u00b5s per loop" ] }, { @@ -115,7 +122,7 @@ "stream": "stdout", "text": [ "\n", - "100000 loops, best of 3: 19.9 \u00b5s per loop" + "10000 loops, best of 3: 19.5 \u00b5s per loop" ] }, { @@ -123,7 +130,7 @@ "stream": "stdout", "text": [ "\n", - "100000 loops, best of 3: 13.5 \u00b5s per loop" + "100000 loops, best of 3: 11.7 \u00b5s per loop" ] }, { @@ -131,7 +138,7 @@ "stream": "stdout", "text": [ "\n", - "100000 loops, best of 3: 8.58 \u00b5s per loop" + "100000 loops, best of 3: 8.24 \u00b5s per loop" ] }, { @@ -139,7 +146,7 @@ "stream": "stdout", "text": [ "\n", - "100000 loops, best of 3: 9.42 \u00b5s per loop" + "100000 loops, best of 3: 9.38 \u00b5s per loop" ] }, { @@ -150,7 +157,7 @@ "\n", "\n", "functions considering all characters (case-sensitive)\n", - "1000000 loops, best of 3: 508 ns per loop" + "1000000 loops, best of 3: 507 ns per loop" ] }, { @@ -166,7 +173,7 @@ "stream": "stdout", "text": [ "\n", - "1000000 loops, best of 3: 480 ns per loop" + "1000000 loops, best of 3: 581 ns per loop" ] }, { @@ -177,15 +184,120 @@ ] } ], - "prompt_number": 11 + "prompt_number": 2 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "import timeit\n", + "\n", + "funcs_s = ['palindrome_short', 'palindrome_regex', 'palindrome_stringlib', \n", + " 'palindrome_stringlib2', 'palindrome_isalpha'] \n", + "\n", + "funcs_ins = ['palindrome_reverse1', 'palindrome_reverse2', 'palindrome_recurs']\n", + "\n", + "times_s, times_ins = [], []\n", + "\n", + "for f in funcs_s:\n", + " times_s.append(min(timeit.Timer('%s(test_str)' %f, \n", + " 'from __main__ import %s, test_str' %f)\n", + " .repeat(repeat=3, number=1000)))\n", + " \n", + "for f in funcs_ins:\n", + " times_ins.append(min(timeit.Timer('%s(test_str)' %f, \n", + " 'from __main__ import %s, test_str' %f)\n", + " .repeat(repeat=3, number=1000)))" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 13 }, { "cell_type": "code", "collapsed": false, - "input": [], + "input": [ + "%pylab inline" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 8 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "import matplotlib.pyplot as plt\n", + "\n", + "labels = [('cy_lstsqr', 'Cython implementation'), \n", + " ('lin_lstsqr_mat', 'numpy matrix equation'),\n", + " ('numpy_lstsqr', 'numpy.linalg.lstsq()'), \n", + " ('scipy_lstsqr', 'scipy.stats.linregress()')] \n", + "\n", + "matplotlib.rcParams.update({'font.size': 12})\n", + "\n", + "fig = plt.figure(figsize=(10,8))\n", + "for f in funcs_s:\n", + " plt.plot(orders_n, times_n[lb[0]], alpha=0.5, label=lb[1], marker='o', lw=3)\n", + "plt.xlabel('sample size n')\n", + "plt.ylabel('time per computation in milliseconds [ms]')\n", + "plt.xlim([1,max(orders_n) + max(orders_n) * 10])\n", + "plt.legend(loc=2)\n", + "plt.grid()\n", + "plt.xscale('log')\n", + "plt.yscale('log')\n", + "plt.title('Performance of least square fit implementations for different sample sizes')\n", + "plt.show()" + ], "language": "python", "metadata": {}, "outputs": [] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "import matplotlib.pyplot as plt\n", + "\n", + "x_pos = np.arange(len(times_s))\n", + "plt.bar(x_pos, times_s, align='center', alpha=0.5, color=\"green\")\n", + "plt.xticks(x_pos, funcs_s, rotation=90)\n", + "plt.ylabel('time per computation in ms')\n", + "plt.title('Performance of different case-sensitive palindrome functions')\n", + "plt.grid()\n", + "plt.show()\n", + "\n", + "x_pos = np.arange(len(times_ins))\n", + "plt.bar(x_pos, times_ins, align='center', alpha=0.5, color=\"blue\")\n", + "plt.xticks(x_pos, funcs_ins, rotation=90)\n", + "plt.ylabel('time per computation in ms')\n", + "plt.title('Performance of different case-insensitive palindrome functions')\n", + "plt.grid()\n", + "plt.show()" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "display_data", + "png": "iVBORw0KGgoAAAANSUhEUgAAAZsAAAFrCAYAAAAHCiThAAAABHNCSVQICAgIfAhkiAAAAAlwSFlz\nAAALEgAACxIB0t1+/AAAIABJREFUeJzt3XlYVOX7P/D3gCg7KCCLyCIoooEobqQIfkwNE6SPaago\nuIHmGlZSmaKZW5pppKkVaimplWtIfDTRXHBJzXJDyEFBcCdBBJzh+f3Bj/NlZGBAOTOch/t1XVwX\nZ+bMmfuegbnn3M8zz8gYYwyEEEKIiPR0HQAhhBD+UbEhhBAiOio2hBBCREfFhhBCiOio2BBCCBEd\nFRtCCCGiaxTF5vbt2+jTpw/Mzc3x7rvv6jocnXvy5AmCg4NhaWmJN998U+P+qampaN26tbD90ksv\n4ciRIwAAxhjGjh2LFi1aoGfPngCAtWvXwtbWFubm5nj48KE4SZBa2bJlCwYOHFjt9b///jvat2+v\nxYg027hxI/z9/YVtMzMzyOXy5zpWYGAgvvnmm3qKrH7NmTMHNjY2cHBw0Or9Tp48GQsXLtTqfQIA\nWAPl7OzMjIyMmKmpKbO1tWWRkZGssLDwuY61YMECNnTo0HqOULo2b97MunfvzpRKZa32P3ToEHN0\ndFR73ZEjR5ijoyMrKipijDFWWlrKjIyM2F9//VVv8daFs7MzO3jwoE7uWwpkMhnLzMzUdRg1SkhI\nYL17966XYwUGBrJvvvmmXo5Vn7KyspiRkRG7d++eqPdTn4/li2qwZzYymQz79u1DQUEBzp49izNn\nztS5GjPGUFZWhqysLHh6ej5XHAqF4rlu15BlZWWhXbt20NN78ac/KysLLi4uMDIyAgDk5eWhuLj4\nuR/vsrKyF4pHJpOB0eeUa0SPTzmlUqmz+75x4wasrKxgZWWlsxi0TsfFrlouLi4q71DfeecdNnjw\nYMYYYydOnGB+fn7M0tKSderUiaWmpgr7BQQEsA8//JD16tWLGRkZsfDwcGZgYMCaNm3KTE1N2cGD\nB1lJSQmbMWMGc3BwYA4ODmzmzJmspKSEMVb+Lr5Vq1Zs6dKlzM7Ojo0ePZrFxcWxN954g4WHhzMz\nMzPm5eXF0tPT2aJFi1jLli2Zk5MTS0lJEWL49ttvmaenJzMzM2Nt2rRh69atE66rOP6KFStYy5Yt\nmb29PUtISBCuLyoqYjExMczZ2ZlZWFiw3r17sydPnmjM+1mXLl1iAQEBzNLSknXs2JHt2bOHMcbY\n3LlzWdOmTZmBgQEzNTVl3377bZXbFhUVsYiICNa8eXPWoUMHtmzZMpUzG2dnZ3bgwAH29ddfM0ND\nQ6avr89MTU3ZiBEjmImJCZPJZMzU1JT169ePMcbY5cuX2SuvvMJatGjBPDw82Pbt24VjRUREsEmT\nJrGgoCBmYmLCDh48yHJycth///tfZmNjw1xdXdnq1auF/efNm8eGDRvGxowZw8zMzFjHjh3ZmTNn\nGGOMhYeHMz09PeGM+NNPP1X72OzatYt16tSJmZubMzc3N5acnKzxebt79y577bXXmKWlJWvRogXz\n9/dnZWVljDFWY7zPevLkCRs1ahSzsrJilpaWrFu3buz27duMMcby8/PZuHHjmL29PWvVqhWbM2eO\ncPaZkJDAevXqxd555x3WvHlz5urqyvbv3y8cNyEhgbVp04aZmZkxV1dXtmXLFuHyine2/v7+TCaT\nMRMTE2Zqasq2b9+ucta6ZMkS9sYbb6jEO336dDZ9+nSN8T1r3rx5bOjQoezNN99kZmZmrEuXLuzP\nP/8Url+8eDFzc3NjZmZmrEOHDmznzp0quVR+N175bCwiIoK99dZb7LXXXmNmZmasR48eKmdqKSkp\nzMPDg1lYWLCpU6eygIAA4cwmISGBvfzyy+ztt99mVlZW7KOPPmL//vsvGz16NLOxsWHOzs5s4cKF\nwvNaeX9LS0vm5ubGjh07xr799lvWunVr1rJlS7Zp0ybhvouLi9msWbOYk5MTs7W1ZZMmTRL+dyv7\n3//+x4yMjJienh4zNTVlY8eOVds9qHyWXtPfPWOM3bhxg73++uvMxsaGWVlZsalTp7LLly+zZs2a\nCf+fzZs3Fx7DOXPmCLddv349c3d3Zy1atGAhISHs1q1bKo/9V199xdq2bcssLS3ZlClThOuuXbvG\n+vTpwywsLJi1tTV788031f4tVGjQxebAgQOMsfIHsmPHjmzu3LksOzubWVlZCf9o//vf/5iVlZVw\nOhoQEMCcnZ3ZpUuXmFKpZE+fPmWRkZHso48+Eo790UcfMT8/P3b37l129+5d9vLLLwvXHzp0iDVp\n0oTFxsay0tJS9uTJEzZv3jxmaGjIUlJSmEKhYGPGjGHOzs5s0aJFTKFQsA0bNjBXV1fh+L/88gv7\n559/GGOMHT58mBkbG7OzZ8+qHH/evHlMoVCwpKQkZmxszPLz8xljjL311lusb9++7NatW0ypVLIT\nJ06wkpKSavO+e/dulceutLSUubm5scWLF7OnT5+y3377jZmZmbGrV68yxhiLi4tjo0ePrvaxnz17\nNuvTpw97+PAhu3nzJuvYsSNr3bq1ynNT8U+wceNGlRcGuVzOZDKZ8CJUWFjIHB0d2caNG5lSqWTn\nzp1j1tbW7NKlS4yx8j98CwsLdvz4ccZYeaHr0qUL+/jjj9nTp0/ZP//8w9q0acN+/fVXxhgTnov9\n+/ezsrIy9v7777OePXuqjU2dkydPMgsLC+FvKycnh125cqXa5+3cuXOMMcZiY2PZpEmTmEKhYAqF\ngh09epQxxphSqawx3md99dVXLDg4mD158oSVlZWxs2fPskePHjHGGAsNDWWTJk1iRUVF7M6dO6x7\n9+5CwUtISGAGBgbs66+/ZmVlZWzt2rXMwcFBeIzNzc1Zeno6Y4yxvLw8dvHiReF21b1wM6baIpXL\n5czY2JgVFBQwxhhTKBTM3t6enTx5UmN8z5o3bx4zMDBgP/30E1MoFGz58uXM1dWVKRQKxhhjO3bs\nYLm5uYwxxrZt28ZMTExYXl6expgjIiKYlZUVO336NFMoFGzUqFEsLCyMMVb+hsDMzEy4z5UrV7Im\nTZqoFJsmTZqw+Ph4plQq2ZMnT9jo0aNZaGgoKywsZHK5nLVr167K/hs3bmRlZWVszpw5rFWrVmzq\n1KmstLSUpaSkMDMzM/b48WPGGGMzZ85kQ4YMYQ8fPmQFBQUsODiYvf/++2ofn9TUVJXioq7YVP5b\nrunvXqFQMG9vbxYTE8OKiopYcXExO3bsGGOs6v8nY0zl9fDgwYPM2tqanTt3jpWUlLBp06axPn36\nqDz2wcHB7N9//2U3btxgNjY2wt92WFgYW7RoEWOMsZKSEuE+q9Ngi42zszMzNTVllpaWzNnZmU2Z\nMoU9efKELVmypMoL5cCBA4V3GIGBgWzevHkq10dGRqpUcjc3N5V3hb/++itzcXFhjJU/6U2bNhXO\ndBgrf6IHDBggbO/Zs4eZmpoK74AePXrEZDIZ+/fff9XmEhoaylatWiUc38jISOUdYcuWLdnJkyeZ\nUqlkRkZG7MKFC1WOoSnvyo4cOcLs7OxULhsxYgSLi4sT8gkPD1cbK2Osyovl+vXrVf4RKv8TPPvC\ncP36dZVi88MPPzB/f3+V40dFRbH58+czxspfPCIiIoTr0tLSmJOTk8r+ixYtYmPHjhVi79+/v3Dd\nxYsXmZGRkdrY1ImKimIxMTHVXl9Z5edt7ty5bMiQISwjI0NlH03xPuvbb79lL7/8cpXnOC8vjzVr\n1kzlnfDWrVtZ3759GWPlj7O7u7tw3ePHj5lMJmO3b99mhYWFzNLSkv3000/C2FmFuhQbxhjr3bs3\n27x5M2Os/CzBzc2tVvE9a968eczPz0/YLisrY/b29uz3339Xu7+Pjw/bvXu3xpgjIyPZxIkTheuS\nkpJY+/btGWOMbdq0SeU+GWPM0dFRpXhUfq4UCgVr2rQpu3z5snDZunXrWGBgoLB/27ZthesuXLjA\nZDIZu3PnjnCZlZUV+/PPP1lZWRkzMTFReWyPHz+u8ia0smcf99oUm+r+7o8fP85sbGzUnmWqG7Op\nXGzGjRvHZs+eLVxXWFjIDAwMWFZWFmOs/LGvXESGDx/Oli5dyhhjbMyYMSwqKoplZ2erzfFZDXrM\nZvfu3Xj48CHkcjni4+NhaGiIrKws7NixA82bNxd+jh07hry8POG2lWdOqXPr1i04OzsL205OTrh1\n65awbWNjg6ZNm6rcpmXLlsLvRkZGsLa2hkwmE7YBoLCwEACwf/9+9OzZE1ZWVmjevDmSkpJw//59\n4fZWVlYq4yXGxsYoLCzEvXv3UFxcDDc3tyox1ybvyvk9+xg4OzsjJyenxseluts7OTnV6nbqZGVl\n4eTJkypxb926Fbdv3wZQ/jw7Ojqq7H/r1i2V/RcvXow7d+4I+9ja2gq/Gxsbo7i4uNZjPdnZ2Wof\nX6Dm5+3dd9+Fu7s7BgwYADc3NyxdurRW8ZqamsLMzAzm5ubIzs7G6NGjMXDgQISFhaFVq1aYPXs2\nFAoFsrKy8PTpU9jb2wvHmTRpEu7evSvEZ2dnp5I3UP43Z2Jigm3btuGrr76Cg4MDBg8ejKtXr9bq\n8XjWyJEjkZiYCADYunUrRo0aJeSpKb5nVX5eK57n3NxcAMDmzZvRuXNn4Vh///23yv9ITSo//0ZG\nRsL/3a1bt1TuE6j6WlB5+969e3j69GmV14LK/yfP3hdQ/vrw7P3fvXsXRUVF8PX1FXIKCgrCvXv3\napVTbVT3d3/z5k04Ozs/1xhsbm6uSv4mJiawsrJSeQye/bsrKCgAACxbtgyMMXTv3h0vvfQSEhIS\naryvJnWOTsecnJwwevRorF+/vtp9KopAdRwcHCCXy4VB7Bs3bqhMP3z29pqOV1lJSQmGDh2K77//\nHkOGDIG+vj5ef/31Wg3KWltbw9DQEBkZGfD29la5rjZ5V3BwcMDNmzfBGBNiz8rKqvUUV3t7e9y4\ncUPl8XleTk5OCAgIQEpKSrX7VH58nZyc4OrqivT0dI37Ps/1rVu3RkZGRpXLNT1vpqamWL58OZYv\nX46LFy/iP//5D7p166Yx3ooXwsrmzp2LuXPnIisrC4MGDYKHhwcGDRqEZs2a4f79+8/1ojFgwAAM\nGDAAJSUl+PDDDzFx4kRhenpdvPHGG5g1axZycnKwa9cupKWlASh/3Ooa382bN4Xfy8rKkJ2dDQcH\nB2RlZSEqKgq//fYb/Pz8IJPJ0Llz5xeeuODg4IDdu3cL24wxlRgA1b8Pa2trGBgYVHkteLZg1Ya1\ntTWMjIxw6dIl2Nvb1/n2JiYmKCoqEraVSmWNhbyy1q1b48aNG1AqldDX11e5rravhRUeP36M+/fv\no1WrVhrv19bWVng9OnbsGF555RUEBASgTZs2avdvsGc21QkPD8fevXuRkpICpVKJ4uJipKamqlTi\nZ/9on90eMWIEFi5ciHv37uHevXtYsGABRo8eXe191uWfoLS0FKWlpbC2toaenh72799f4wttZXp6\nehg3bhxiYmKQm5sLpVKJEydOoLS0tFZ5V+jZsyeMjY2xbNkyPH36FKmpqdi3bx/CwsJqFcfw4cOx\nePFi5OfnIzs7G1988UWt83/W4MGDkZ6eju+//x5Pnz7F06dPcfr0aVy5cgVA1ce2e/fuMDMzw7Jl\ny/DkyRMolUr8/fffOHPmjNr9n2Vra4vMzMxqrx8/fjwSEhLw22+/oaysDDk5Obh69arG523fvn3I\nyMgAYwzm5ubQ19eHvr6+xniflZqair/++gtKpRJmZmYwMDCAvr4+7OzsMGDAAMTExKCgoABlZWXI\nzMysVcG4c+cOdu/ejcePH8PAwAAmJiZVXnRq+/jY2NggMDAQkZGRaNOmDTw8PACUvwGpa3x//PEH\ndu7cCYVCgc8//xyGhobo2bMnHj9+DJlMBmtra5SVlSEhIQF///23xjyBmp//QYMG4eLFi8J9rl69\nWu2ZfwV9fX0MHz4cH374IQoLC5GVlYWVK1ciPDy8VrFUpqenh4kTJ2LmzJlCkcjJyan1/367du1Q\nXFyMpKQkPH36FAsXLkRJSUmtbtu9e3fY29sjNjYWRUVFKC4uxvHjxwGUP9/Z2dl4+vSpsD8rHz4B\nUP5amJCQgD///BMlJSX44IMP0LNnz2q7GZUf/x07diA7OxsAYGlpCZlMVuMbEckVG0dHR+zevRuL\nFi1Cy5Yt4eTkhBUrVqg8COrOTCpfNmfOHHTt2hXe3t7w9vZG165dMWfOnFrfvrp9gPIPoK1evRrD\nhw9HixYtkJiYiCFDhtR428qWL18OLy8vdOvWDVZWVnj//fdRVlZWbd7q2kcGBgbYu3cv9u/fDxsb\nG0ydOhXfffcd2rVrV20+lc2bNw/Ozs5wdXXFq6++ijFjxlS7v6bHxtTUFCkpKfjhhx/QqlUr2Nvb\n4/3330dpaana2+vp6WHfvn04f/482rRpAxsbG0RFReHRo0e1ur/3338fCxcuRPPmzfHZZ59Vibdb\nt25ISEjA22+/DUtLSwQGBuLGjRsan7eMjAz0798fZmZmePnllzFlyhQEBARojPdZeXl5GDZsGCws\nLNChQwcEBgYKb3Q2b96M0tJSdOjQAS1atMCwYcOEF8ua8i4rK8PKlSvRqlUrWFlZ4ffff8fatWvV\n3i4uLg4RERFo3rw5fvzxR7XHHTlyJA4ePIiRI0eqXF5TfM+SyWQYMmQItm3bhhYtWmDLli34+eef\noa+vjw4dOmDWrFnw8/ODnZ0d/v77b/Tu3VvltpVjevb36h4Ha2tr7NixA7GxsbC2tkZGRkaNxwWA\nL774AiYmJmjTpg38/f0xatQojB07VuN9qbN06VK4u7ujZ8+esLCwQP/+/as94332WBYWFlizZg0m\nTJgAR0dHmJqaqrT8aopFX18fe/fuRUZGBpycnNC6dWts374dANCvXz907NgRdnZ2wlBA5WP169cP\nH3/8MYYOHQoHBwdcv34dP/zwQ7X5Vr7tmTNn0LNnT5iZmWHIkCFYvXo1XFxcqs+Xvei5aw2Sk5Mx\nc+ZMKJVKTJgwAbNnz66yz/Tp07F//34YGxtj48aN6Ny5M27evIkxY8bgzp07kMlkiIqKwvTp0wEA\nDx48wJtvvil8vmP79u2wtLQUKwVCyHOYP38+MjIy8N133+k6FNJAiHZmo1QqMXXqVCQnJ+PSpUtI\nTEzE5cuXVfZJSkpCRkYGrl27hvXr12Py5MkAyt+Zr1y5EhcvXkRaWhq+/PJLoe2yZMkS4R1Dv379\nsGTJErFSIIQ8JxHfwxKJEq3YnDp1Cu7u7nBxcYGBgQHCwsJUBu8AYM+ePYiIiAAA9OjRA/n5+bh9\n+zbs7Ozg4+MDoLwN4+npKYxNVL5NREQEdu3aJVYKhJDnpKlVSxof0Waj5eTkqPQcHR0dcfLkSY37\nZGdnq0zxk8vlOHfuHHr06AGgfFHNiuttbW2FKbSEkIZj3rx5ug6BNDCindnU9l3Ns6fblW9XWFiI\nN954A6tWrYKpqana+6B3T4QQ0vCJdmbTqlUrlTnuN2/erDJ//dl9srOzhfndT58+xdChQxEeHo7Q\n0FBhH1tbW+Tl5cHOzg65ubkqH7aszN3dvcYpnoQQQlR16tQJ58+fF+XYop3ZdO3aFdeuXYNcLkdp\naSm2bduGkJAQlX1CQkKwefNmAEBaWhosLS1ha2sLxhjGjx+PDh06YObMmVVus2nTJgDApk2bVApR\nZZmZmcJ8ct5+5s2bp/MYKD/Kj/Lj7+fPP/8UoRqUE+3MpkmTJoiPj8fAgQOhVCoxfvx4eHp6Yt26\ndQCA6OhoDBo0CElJSXB3d4eJiYmw3MGxY8fw/fffw9vbG507dwYALF68GK+++ipiY2MxfPhwfPPN\nN8LU58bmeb9ISiooP2mj/Ig6oi5XExQUhKCgIJXLoqOjVbbj4+Or3K53797VrnXVokULHDhwoP6C\nJIQQIjrJrSBAgMjISF2HICrKT9ooP6KOqCsI6BJ9YyMhhNSNmK+bklv1mZQv5hgYGKjrMBAbF4u8\n/OoXOnxeedl5sHO007xjHdlZ2mFJnO5XnGgoz59YKD+iDhUb8tzy8vPgEupS/wc+D7j41P9x5bvk\n9X5MQkjt0JiNBPH+rkqMQtOQ8P78UX5EHSo2hBBCREfFRoJSU1N1HYKo5Oflug5BVLw/f5QfUYeK\nDSGEENFRsZEg3nvGNGYjbZQfUYeKDSGEENFRsZEg3nvGNGYjbZQfUYeKDSGEENFRsZEg3nvGNGYj\nbZQfUYeKDSGEENFRsZEg3nvGNGYjbZQfUYeKDSGEENFRsZEg3nvGNGYjbZQfUYeKDSGEENFRsZEg\n3nvGNGYjbZQfUYeKDSGEENFRsZEg3nvGNGYjbZQfUYeKDSGEENFRsZEg3nvGNGYjbZQfUYeKDSGE\nENFRsZEg3nvGNGYjbZQfUYeKDSGEENFRsZEg3nvGNGYjbZQfUYeKDSGEENFRsZEg3nvGNGYjbZQf\nUYeKDSGEENFRsZEg3nvGNGYjbZQfUYeKDSGEENFRsZEg3nvGNGYjbZQfUYeKDSGEENFRsZEg3nvG\nNGYjbZQfUYeKDSGEENFRsZEg3nvGNGYjbZQfUYeKDSGEENFRsZEg3nvGNGYjbZQfUYeKDSGEENFp\nLDbvvvsuHj16hKdPn6Jfv36wtrbGd999p43YSDV47xnTmI20UX5EHY3FJiUlBebm5ti3bx9cXFyQ\nmZmJTz/9VBuxEUII4YTGYqNQKAAA+/btwxtvvAELCwvIZDLRAyPV471nTGM20kb5EXWaaNohODgY\n7du3h6GhIdauXYs7d+7A0NBQG7ERQgjhhIwxxjTtdP/+fVhaWkJfXx+PHz9GQUEB7OzstBHfc5PJ\nZKhFauQFRM6MhEuoi67DqDX5Ljk2fr5R12EQ0mCJ+bqp8cxGoVDg6NGjkMvlQktNJpMhJiZGlIAI\nIYTwR+OYTXBwMDZt2oQHDx6gsLAQhYWFKCgo0EZspBq894xpzEbaKD+ijsYzm5ycHFy4cOG5Dp6c\nnIyZM2dCqVRiwoQJmD17dpV9pk+fjv3798PY2BgbN25E586dAQDjxo3DL7/8gpYtW+Kvv/4S9o+L\ni8PXX38NGxsbAMDixYvx6quvPld8hBBCtEPjmc2AAQPw66+/1vnASqUSU6dORXJyMi5duoTExERc\nvnxZZZ+kpCRkZGTg2rVrWL9+PSZPnixcN3bsWCQnJ1c5bkUL79y5czh37lyjLDS8z/Onz9lIG+VH\n1NFYbF5++WW8/vrrMDQ0hJmZGczMzGBubq7xwKdOnYK7uztcXFxgYGCAsLAw7N69W2WfPXv2ICIi\nAgDQo0cP5OfnIy8vDwDg7++P5s2bqz02DfwTQoi0aCw2MTExSEtLQ1FREQoKClBQUIBHjx5pPHBO\nTg5at24tbDs6OiInJ6fO+6jzxRdfoFOnThg/fjzy8/M17s8b3nvGNGYjbZQfUUdjsXFyckLHjh2h\np1e3ZdRq+8HPZ89SNN1u8uTJuH79Os6fPw97e3vMmjWrTnERQgjRPo0TBFxdXdG3b18EBQWhadOm\nAGo39blVq1a4efOmsH3z5k04OjrWuE92djZatWpV43Fbtmwp/D5hwgQEBwdXu29kZCRcXFwAAJaW\nlvDx8RH6rRXvTqS4HRgY2CDiycvOgwtcAPzf2UjFeMuLbLv4uNTr8Sq287LLW7RiPR613W4ozx/l\nR/lV/C6XyyE2jR/qjIuLK9/xmTOOefPm1XhghUIBDw8PHDx4EA4ODujevTsSExPh6ekp7JOUlIT4\n+HgkJSUhLS0NM2fORFpamnC9XC5HcHCwymy03Nxc2NvbAwBWrlyJ06dPY+vWrVUTawAf6oyNi0Ve\nfp7mHRsQO0s7LIlbUqt96UOdhPBFpx/qrCg2dT5wkyaIj4/HwIEDoVQqMX78eHh6emLdunUAgOjo\naAwaNAhJSUlwd3eHiYkJEhIShNuPGDEChw8fxv3799G6dWssWLAAY8eOxezZs3H+/HnIZDK4uroK\nx2uI8vLzRHkxlp+XizZjS75LLspx6xSDiPk1BKmpqcI7TB5RfkQdjcXmRQQFBSEoKEjlsujoaJXt\n+Ph4tbdNTExUe/nmzZvrJzhCCCFaQ1+eJkE8v+sH+M+P93fFlB9Rh4oNIYQQ0Wlso925cwcbNmyo\nshDnt99+K3pwRD3exzR4z4/3nj/lR9TRWGyGDBmCPn36oH///sJnbejL0wghhNSFxmLz5MkTLF26\nVBuxkFri+V0/wH9+vL8rpvyIOhrHbAYPHoxffvlFG7EQQgjhlMZi8/nnnyM4OLjOC3ES8fC+dhjv\n+VX+9DaPKD+ijsY2WmFhoTbiIIQQwrFqi83ly5fh6emJs2fPqr2+S5cuogVFasb7mAbv+fHe86f8\niDrVFpvPPvsMGzZsQExMjNrZZ4cOHRI1MEIIIfyotths2LABAPUnGyLeP4fCe368f06D8iPq0AoC\nhBBCREfFRoJ4ftcP8J8f7++KKT+iDhUbQgghoqtVscnJycGxY8dw5MgRHD58GEeOHBE7LlID3j+H\nwnt+vI+DUn5EHY2fs5k9eza2bduGDh06QF9fX7i8T58+ogZGCCGEHxqLzc6dO3H16lU0a9ZMG/GQ\nWuB9TIP3/Hjv+VN+RB2NbTQ3NzeUlpZqIxZCCCGc0lhsjIyM4OPjg6ioKEybNg3Tpk3D9OnTtREb\nqQbvYxq858d7z5/yI+pobKOFhIQgJCREWEWAMUbfZ0MIIaRONBabyMhIlJSUID09HQDQvn17GBgY\niB4YqR7vYxq858d7z5/yI+poLDapqamIiIiAs7MzAODGjRvYtGkTAgICRA+OEEIIHzSO2cTExCAl\nJQVHjhzBkSNHkJKSgrffflsbsZFq8D6mwXt+vPf8KT+ijsZio1Ao4OHhIWy3a9cOCoVC1KAIIYTw\nRWMbzdfXFxMmTEB4eDgYY9iyZQu6du2qjdhINXgf0+A9P957/pQfUUdjsVm7di2+/PJLrF69GgDg\n7++Pt956S/TACCGE8ENjG83Q0BCzZs3Czz//jJ9//hlvv/02rSagY7yPafCeH+89f8qPqFPtmc2w\nYcOwY8ePLCzsAAAgAElEQVQOvPTSS1U+VyOTyXDhwgXRgyOEEMKHaovNqlWrAAC//PILGGMq19GH\nOnWL9zEN3vPjvedP+RF1qm2jOTg4AADWrFkDFxcXlZ81a9ZoLUBCCCHSp3HMJiUlpcplSUlJogRD\naof3MQ3e8+O950/5EXWqbaOtXbsWa9asQWZmJry8vITLCwoK0KtXL60ERwghhA/VFpuRI0ciKCgI\nsbGxWLp0qTBuY2ZmBisrK60FSKrifUyD9/x47/lTfkSdaouNhYUFLCws8MMPPwAA7ty5g+LiYjx+\n/BiPHz+Gk5OT1oIkhBAibRrHbPbs2YO2bdvC1dUVAQEBcHFxQVBQkDZiI9XgfUyD9/x47/lTfkQd\njcVmzpw5OHHiBNq1a4fr16/j4MGD6NGjhzZiI4QQwgmNxcbAwADW1tYoKyuDUqlE3759cebMGW3E\nRqrB+5gG7/nx3vOn/Ig6GtdGa968OQoKCuDv749Ro0ahZcuWMDU11UZshOhUbFws8vLzdB1GrdlZ\n2mFJ3BJdh0GIWhqLze7du2FoaIiVK1diy5YtePToEebNm6eN2Eg15OflXL/7byj55eXnwSW0/uMQ\nKz/5Lnm9H/N5pKamcv3un/f8xKKxjbZgwQLo6+vDwMAAkZGRmD59OpYtW6aN2AghhHCCVhCQoIbw\nrl9MlJ+08f6un/f8xEIrCBBCCBFdtWc2I0eOxN69exESEoJ9+/Zh79692Lt3L/744w9s2bJFmzGS\nZ/D+ORTKT9p4/xwK7/mJReMKAkuXLoVMJhO+VoBWECCEEFJXGmejDR48WPi9uLgY169fh4eHBy5e\nvChqYKR6vPf8KT9p431Mg/f8xKKx2Pz1118q22fPnsWXX34pWkCEEEL4o3E22rO6dOmCkydPihEL\nqSXee/6Un7TxPqbBe35i0Xhms2LFCuH3srIynD17Fq1atarVwZOTkzFz5kwolUpMmDABs2fPrrLP\n9OnTsX//fhgbG2Pjxo3o3LkzAGDcuHH45Zdf0LJlS5WzqwcPHuDNN99EVlYWXFxcsH37dlhaWtYq\nHkIIIbqh8cymoKAAhYWFKCwsRGlpKQYPHozdu3drPLBSqcTUqVORnJyMS5cuITExEZcvX1bZJykp\nCRkZGbh27RrWr1+PyZMnC9eNHTsWycnJVY67ZMkS9O/fH+np6ejXrx+WLGl8y3Pw3vOn/KSN9zEN\n3vMTi8Yzm7i4OADAv//+C5lMBnNz81od+NSpU3B3d4eLiwsAICwsDLt374anp6ewz549exAREQEA\n6NGjB/Lz85GXlwc7Ozv4+/tDLpdXOe6ePXtw+PBhAEBERAQCAwMbZcEhhBAp0Xhmc/r0aXh5ecHb\n2xteXl7o1KlTrVZ9zsnJQevWrYVtR0dH5OTk1HmfZ92+fRu2trYAAFtbW9y+fVtjLLzhvedP+Ukb\n72MavOcnFo1nNuPGjcOaNWvg7+8PADh69CjGjRuHCxcu1Hi7is/laFLxddN1vV3FvjXtHxkZKZxZ\nWVpawsfHRzgFrviDEXM7LzsPLii//4oXmIoWSkPdrsBjfnnZ/7eCM+VH27QN4Xd1XaT6JmPPvto/\no3Pnzjh37pzKZV26dMHZs2drPHBaWhri4uKEcZfFixdDT09PZZLApEmTEBgYiLCwMABA+/btcfjw\nYeHMRS6XIzg4WGWCQPv27ZGamgo7Ozvk5uaib9++uHLlStXEZLIqhUzbImdGirJqsJjku+TY+PnG\nWu0rtfzqkhvAf36EPEvM102NbbSAgABER0cjNTUVqampmDx5MgICAnD27NkaC07Xrl1x7do1yOVy\nlJaWYtu2bQgJCVHZJyQkBJs3bwZQXpwsLS2FQlOdkJAQbNq0CQCwadMmhIaGakySEEKIbmlso50/\nfx4ymQzz588HUN72kslkOH/+PADg0KFD6g/cpAni4+MxcOBAKJVKjB8/Hp6enli3bh0AIDo6GoMG\nDUJSUhLc3d1hYmKChIQE4fYjRozA4cOHcf/+fbRu3RoLFizA2LFjERsbi+HDh+Obb74Rpj43Ng3l\n+17EQvlJG+/f98J7fmLRWGxeZDAsKCgIQUFBKpdFR0erbMfHx6u9bWJiotrLW7RogQMHDjx3TIQQ\nQrRPY7F5+PAhNm/eDLlcDoVCAaC8r7d69WrRgyPq8fyuGKD8pI73d/285ycWjcVm0KBB8PPzg7e3\nN/T09IQ2GiGEEFJbGotNSUkJPvvsM23EQmqJ954/5SdtDWFMIzYuFnn5eZp3fA552Xmwc7Sr9+Pa\nWdphSRy/H1DXWGxGjhyJ9evXIzg4GM2aNRMub9GihaiBEULI88rLzxNv2vp5cVqh8l3yej9mQ6Kx\n2BgaGuLdd9/FJ598Aj298pnSMpkM//zzj+jBEfV4flcMUH5Sp+uzGrHx/vyJpVarPmdmZsLa2lob\n8RBCCOGQxg91tm3bFkZGRtqIhdQS72trUX7SxvvaYbw/f2LReGZjbGwMHx8f9O3bVxizoanPhBBC\n6kJjsQkNDUVoaKgw3ZmmPuse7z1jyk/aaMyGqKOx2ERGRqKkpATp6ekAyhfCNDAwED0wQggh/NA4\nZpOamop27dphypQpmDJlCtq2bSt8eRnRDd57xpSftNGYDVFH45lNTEwMUlJS4OHhAQBIT09HWFiY\nxq8YIIQQQipoPLNRKBRCoQGAdu3aCWukEd3gvWdM+UkbjdkQdTSe2fj6+mLChAkIDw8HYwxbtmxB\n165dtREbIYQQTmg8s1m7di08PT2xevVqfPHFF+jYsSPWrl2rjdhINXjvGVN+0kZjNkQdjWc2SqUS\nM2fOxKxZs4TtkpIS0QMjhBDCD41nNv/5z3/w5MkTYbuoqAivvPKKqEGRmvHeM6b8pI3GbIg6tfqK\nAVNTU2HbzMwMRUVFogZFCBGXmEvwi4X3Jfh5p7HYmJiY4I8//oCvry8A4MyZM7RWmo7x/n0olJ/4\nxFyCX6z8GsoS/A3h+ZMijcXm888/x/Dhw2Fvbw8AyM3NxbZt20QPjBBCCD80Fptu3brh8uXLuHr1\nKgDAw8MDTZs2FT0wUj3e31VRftJG+RF1NBYbAGjatCm8vLzEjoUQQginNM5GIw0P7/P8KT9po/yI\nOjUWG8YYbt68qa1YCCGEcErjmU1QUJA24iB1wHvPmPKTNsqPqFNjsZHJZPD19cWpU6e0FQ8hhBAO\naTyzSUtLg5+fH9q0aQMvLy94eXnB29tbG7GRavDeM6b8pI3yI+ponI3266+/aiMOQgghHNN4ZuPi\n4oKbN2/i0KFDcHFxgYmJCRhj2oiNVIP3njHlJ22UH1FHY7GJi4vDsmXLsHjxYgBAaWkpwsPDRQ+M\nEEIIPzQWm507d2L37t0wMTEBALRq1QoFBQWiB0aqx3vPmPKTNsqPqKOx2DRr1gx6ev+32+PHj0UN\niBBCCH80Fpthw4YhOjoa+fn5WL9+Pfr164cJEyZoIzZSDd57xpSftFF+RB2Ns9HeffddpKSkwMzM\nDOnp6fj444/Rv39/bcRGCCGEE7VaG83Lywv+/v7o06cPLcjZAPDeM6b8pI3yI+poLDZff/01evTo\ngZ9//hk//fQTevTogW+++UYbsRFCCOGExjbasmXLcO7cOVhZWQEA7t+/Dz8/P4wfP1704Ih6vPeM\nKT9po/yIOhrPbKytrWFqaipsm5qawtraWtSgCCGE8EVjsXFzc0PPnj0RFxeHuLg49OzZE23btsWK\nFSvw2WefaSNG8gzee8aUn7RRfkQdjW00Nzc3uLm5QSaTAQCGDBkCmUyGwsJC0YMjhBDCB43FJi4u\nTgthkLrgvWdM+Ukb5UfUoa+FJoQQIjoqNhLEe8+Y8pM2yo+oQ8WGEEKI6DQWm6tXr6Jfv37o2LEj\nAODChQtYuHCh6IGR6vHeM6b8pI3yI+poLDYTJ07EokWL0LRpUwDlS9ckJiaKHhghhBB+aCw2RUVF\n6NGjh7Atk8lgYGAgalCkZrz3jCk/aaP8iDoai42NjQ0yMjKE7R9//BH29va1OnhycjLat2+Ptm3b\nYunSpWr3mT59Otq2bYtOnTrh3LlzGm8bFxcHR0dHdO7cGZ07d0ZycnKtYiGEEKI7Gj9nEx8fj6io\nKFy5cgUODg5wdXXFli1bNB5YqVRi6tSpOHDgAFq1aoVu3bohJCQEnp6ewj5JSUnIyMjAtWvXcPLk\nSUyePBlpaWk13lYmkyEmJgYxMTEvlrmE8d4zpvykjfIj6tRqBYGDBw/i8ePHKCsrg5mZWa0OfOrU\nKbi7u8PFxQUAEBYWht27d6sUmz179iAiIgIA0KNHD+Tn5yMvLw/Xr1+v8baMsbrkSAghRMc0ttEe\nPnyIVatWYc6cOfjggw8wbdo0TJ8+XeOBc3Jy0Lp1a2Hb0dEROTk5tdrn1q1bNd72iy++QKdOnTB+\n/Hjk5+drjIU3vPeMKT9po/yIOhrPbAYNGgQ/Pz94e3tDT08PjDFhnbSa1GYfoO5nKZMnT8bcuXMB\nAB999BFmzZpV7ffrREZGCmdHlpaW8PHxQWBgIAAgNTUVAETdzsvOgwvK77/iD7TiFLyhblfgMb+8\n7DzK7/9v52XnAecbVvyUX93+Putju+J3uVwOscmYhlf7Ll264OzZs3U+cFpaGuLi4oQB/MWLF0NP\nTw+zZ88W9pk0aRICAwMRFhYGAGjfvj0OHz6M69eva7wtAMjlcgQHB+Ovv/6qmphMpvN2W+TMSLiE\nuug0hrqS75Jj4+cba7Wv1PKrS24A3/lJLTeA8tMGMV83NbbRRo4cifXr1yM3NxcPHjwQfjTp2rUr\nrl27BrlcjtLSUmzbtg0hISEq+4SEhGDz5s0AyouTpaUlbG1ta7xtbm6ucPudO3fS11QTQogEaGyj\nGRoa4t1338Unn3wCPb3y2iSTyfDPP//UfOAmTRAfH4+BAwdCqVRi/Pjx8PT0xLp16wAA0dHRGDRo\nEJKSkuDu7g4TExMkJCTUeFsAmD17Ns6fPw+ZTAZXV1fheI2J/Lyc6xkxlJ+0UX5EHY3FZsWKFcjM\nzHyub+cMCgpCUFCQymXR0dEq2/Hx8bW+LQDhTIgQQoh0aGyjtW3bFkZGRtqIhdQS7++qKD9po/yI\nOhrPbIyNjeHj44O+ffuiWbNmAMrbaKtXrxY9OEIIIXzQWGxCQ0MRGhqqclltpzUTcfDeM6b8pI3y\nI+poLDaRkZFaCIMQQgjPqi02w4YNw44dO9ROLZbJZLhw4YKogZHq8f6uivKTNsqPqFNtsVm1ahUA\nYN++fVU+5ENtNEIIIXVR7Ww0BwcHAMCaNWvg4uKi8rNmzRqtBUiq4n1tJspP2ig/oo7Gqc8pKSlV\nLktKShIlGEIIIXyqto22du1arFmzBpmZmSrjNgUFBejVq5dWgiPq8d4zpvykjfIj6lRbbEaOHImg\noCDExsZi6dKlwriNmZkZrKystBYgIYQQ6au2jWZhYQEXFxf88MMPcHZ2FsZrqNDoHu89Y8pP2ig/\noo7GMRtCCCHkRVGxkSDee8aUn7RRfkQdKjaEEEJER8VGgnjvGVN+0kb5EXWo2BBCCBEdFRsJ4r1n\nTPlJG+VH1KFiQwghRHRUbCSI954x5SdtlB9Rh4oNIYQQ0VGxkSDee8aUn7RRfkQdKjaEEEJER8VG\ngnjvGVN+0kb5EXWo2BBCCBEdFRsJ4r1nTPlJG+VH1KFiQwghRHRUbCSI954x5SdtlB9Rh4oNIYQQ\n0VGxkSDee8aUn7RRfkQdKjaEEEJER8VGgnjvGVN+0kb5EXWo2BBCCBEdFRsJ4r1nTPlJG+VH1KFi\nQwghRHRUbCSI954x5SdtlB9Rh4oNIYQQ0VGxkSDee8aUn7RRfkQdKjaEEEJER8VGgnjvGVN+0kb5\nEXWo2BBCCBEdFRsJ4r1nTPlJG+VH1KFiQwghRHRUbCSI954x5SdtlB9Rh4oNIYQQ0VGxkSDee8aU\nn7RRfkQdKjaEEEJEJ2qxSU5ORvv27dG2bVssXbpU7T7Tp09H27Zt0alTJ5w7d07jbR88eID+/fuj\nXbt2GDBgAPLz88VMoUHivWdM+Ukb5UfUEa3YKJVKTJ06FcnJybh06RISExNx+fJllX2SkpKQkZGB\na9euYf369Zg8ebLG2y5ZsgT9+/dHeno6+vXrhyVLloiVQoOVl5Gn6xBERflJG+VH1BGt2Jw6dQru\n7u5wcXGBgYEBwsLCsHv3bpV99uzZg4iICABAjx49kJ+fj7y8vBpvW/k2ERER2LVrl1gpNFjFhcW6\nDkFUlJ+0UX5EHdGKTU5ODlq3bi1sOzo6Iicnp1b73Lp1q9rb3r59G7a2tgAAW1tb3L59W6wUCCGE\n1BPRio1MJqvVfoyxWu2j7ngymazW98OT/Dy+x6koP2mj/IhaTCQnTpxgAwcOFLYXLVrElixZorJP\ndHQ0S0xMFLY9PDxYXl5ejbf18PBgubm5jDHGbt26xTw8PNTef6dOnRgA+qEf+qEf+qnlT6dOneqt\nBjyrCUTStWtXXLt2DXK5HA4ODti2bRsSExNV9gkJCUF8fDzCwsKQlpYGS0tL2NrawsrKqtrbhoSE\nYNOmTZg9ezY2bdqE0NBQtfd//vx5sVIjhBBSR6IVmyZNmiA+Ph4DBw6EUqnE+PHj4enpiXXr1gEA\noqOjMWjQICQlJcHd3R0mJiZISEio8bYAEBsbi+HDh+Obb76Bi4sLtm/fLlYKhBBC6omMsVoMmhBC\nCCEvgFYQIIQQIjoqNhLRr1+/Wl0mVXfu3Kly2dWrV3UQiTju3buHadOmoXPnzujSpQtmzJiB+/fv\n6zqsF/Lvv/8iNjYW4eHh2Lp1q8p1b731lo6iEse+ffuwdOlSzJ8/HwsWLMCCBQt0HZLkULFp4J48\neYL79+/j7t27ePDggfAjl8urfG5Jyvz9/bFt2zYAAGMMK1asqHbyhxSFhYWhZcuW+Pnnn/Hjjz/C\nxsYGb775pq7DeiFjx44FAAwdOhSJiYkYOnQoiovLP/B44sQJXYZWr6Kjo7F9+3Z88cUXAIDt27cj\nKytLx1FJD43ZNHCff/45Vq1ahVu3bsHBwUG43MzMDFFRUZg6daoOo6s/ubm5iIqKgqGhIW7fvo32\n7dvjs88+g6mpqa5DqxcvvfQS/v77b5XLvLy88Ndff+koohfXqVMn/Pnnn8L2J598gqSkJOzevRv9\n+/dXWetQyiqeJ29vb1y4cAGFhYV49dVXcfToUV2HJil0ZtPAzZw5ExkZGZgzZw6uX78u/Fy4cIGb\nQgMA9vb2GDhwII4fPw65XI7IyEhuCg0ADBgwAImJiSgrK0NZWRm2bduGAQMG6DqsF1JaWoqysjJh\n+8MPP8TEiRMREBCABw8e6DCy+mVkZAQAMDY2Rk5ODpo0aYK8PFofrc5E+wQPqVdiftiqIejXrx8L\nDw9nDx8+ZBcuXGDdunVjs2bN0nVYL8zExISZmpoyU1NTJpPJmL6+PtPX12cymYyZmprqOrwX8s47\n77CUlJQql+/fv5+5u7vrICJxLFiwgD148ID9+OOPzNbWltna2rI5c+boOizJoTaaRLzzzjvo2bMn\nhg4dyuUSPTt37sTrr78ubCsUCixevBgfffSRDqMiRFVxcTGKi4thaWmp61Akh4qNRJiamqKoqAj6\n+vowNDQEUL423KNHj3QcWf35/fffkZGRgbFjx+Lu3bsoKChAmzZtdB3WC7ly5Qrat2+Ps2fPqr2+\nS5cuWo6o/t27dw/z58/H0aNHIZPJ4O/vj7lz58LKykrXodWbY8eOQS6XQ6lUCpeNGTNGhxFJDxUb\n0iDExcXhjz/+wNWrV5Geno6cnBwMHz4cx44d03VoL2TixInYsGEDAgMD1Z6RHjp0SAdR1a9XXnkF\nAQEBCA8PB2MMW7duRWpqKg4cOKDr0OpFeHg4/vnnH/j4+EBfX1+4vGJ2GqkdKjYSsnv3bhw5cgQy\nmQwBAQEIDg7WdUj1puKbWn19fYVZTBWzf0jDxuNMu8o8PT1x6dIlLtvX2iTa2mikfsXGxuL06dMY\nNWoUGGNYvXo1jh8/jsWLF+s6tHrRrFkz6On93+TIx48f6zCa+vPTTz/V+CL13//+V4vRiKNipl3F\n54Z27Ngh+Zl2lb300kvIzc1V+egBqTs6s5EILy8vnD9/XjiNVyqV8PHx4ebd46effoqMjAykpKTg\n/fffx7fffouRI0di+vTpug7thURGRtZYbCoWn5UiU1NTIbfHjx8LbxbKyspgYmKCgoICXYb3wio6\nB4WFhTh37hy6d++OZs2aASgfL92zZ48uw5McKjYS4e3tjUOHDgmDrvfv30ffvn25ajOlpKQgJSUF\nADBw4ED0799fxxGRxiw1NVVlu6Kwsv//ZY4BAQE6iEq6qNhIRGJiImJjYxEYGAgAOHz4MJYsWYKw\nsDDdBkZqZcWKFVXOcCwsLODr6wsfHx8dRfViGsNMuwq5ubk4deoU9PT00K1bN9jZ2ek6JMmhYiMh\nt27dwunTpyGTydC9e3eu/uDNzMyqXGZhYYFu3bphxYoVkp8CPXLkSJw5cwbBwcFgjOGXX36Bl5cX\nsrKy8MYbb2D27Nm6DrHOGsNMOwD4+uuvsWDBAvTt2xdA+RnP3LlzMX78eB1HJi1UbCQkJycHcrkc\nCoVC+Ofu06ePjqOqH3PmzEHr1q0xYsQIAMAPP/yAzMxMdO7cGV999VWVlobU+Pv7Y//+/cISPIWF\nhRg0aBCSk5Ph6+uLy5cv6zhCUp127drhxIkTKi1sPz8/pKen6zgyaaHZaBIxe/ZsbNu2DR06dFCZ\n689LsdmzZ4/K+FNUVBR8fHywdOlSLmbc3b17F02bNhW2DQwMcPv2bRgbGwsf0pWaxjDTDgCsra1V\n1ukzNTWFtbW1DiOSJio2ErFz505cvXpVmA3DG2NjY2zbtg3Dhg0DAPz4448qKyVI3ahRo9CjRw+E\nhoaCMYa9e/di5MiRePz4MTp06KDr8J7L3r17G0WxcXNzQ8+ePTFkyBAA5Z938/b2FsbhYmJidByh\nNFAbTSKCgoKwfft2tWMbPMjMzMSMGTOQlpYGAOjZsyc+//xztGrVCn/88Qd69+6t4whf3OnTp3Hs\n2DHIZDL06tULXbt21XVIpBbi4uIAVJ2NVmHevHm6CEtyqNg0cNOmTQNQPjng/Pnz6Nevn8pc/9Wr\nV+syPFIHSqUSeXl5KmNuTk5OOo7qxfE4047UP2qjNXC+vr6QyWRgjCE4OFj4nYfWUmVXr17FW2+9\nhby8PFy8eBEXLlzAnj17MGfOHF2HVi+++OILzJ8/Hy1btlQZc+PhQ7l//PGH2pl2X331lWRn2gGo\ncTko+lBn3dGZjQQ9ePAAN2/eRKdOnXQdSr3p06cPPv30U0yaNAnnzp0DYwwvvfQSLl68qOvQ6oWb\nmxtOnTrF1UrIFXidaadpBmTFZ95I7dCZjUQEBgZiz549UCgU8PX1hY2NDXr16oWVK1fqOrR6UVRU\nhB49egjbMpkMBgYGOoyofjk5OcHc3FzXYYiCx5l2ABWT+kbFRiLy8/Nhbm6Or7/+GmPGjMH8+fPh\n5eWl67DqjY2NDTIyMoTtH3/8Efb29jqMqH65urqib9++eO2114QXZl5mMvE4066y9PR0fPDBB7h4\n8SKKi4sBlD93//zzj44jkxYqNhKhVCqRm5uL7du3Y+HChQD4mBJcIT4+HlFRUbh69SocHBzg6uqK\nLVu26DqseuPk5AQnJyeUlpaitLSUq3G3jz76CK+++qow027dunXCTDsensOxY8di/vz5iImJQXJy\nMhISElS+RI3UDo3ZSMSOHTvw8ccfo1evXli7di0yMzPx3nvv4aefftJ1aC9MqVRi9uzZWL58OQoL\nC1FWVsZty4lXvM60A8rXeDt79qzKd/RUXEZqj85sJGLYsGHCBx6B8gHnyoVm8eLFeP/993UR2gvT\n19fH0aNHwRhT+aQ2D2bMmIFVq1apndnEy4wmnmfaAYChoSGUSiXc3d0RHx8PBwcHbr5vSZvozIYT\nnTt3Fr7hUoomTZqEW7duYdiwYTA2NgZQ/mIs9U+h//HHH/D19cXhw4fx7L8aL8vU8zzTDgBOnToF\nT09P5Ofn46OPPsKjR4/w3nvvoWfPnroOTVKo2HBC6sWmui8Zk/KXi1VQKBQYM2YMtm7dqutQRNG3\nb1+kpKRwNXuwOkqlEoWFhbCwsNB1KJJDbTTSIGzcuLHG66XcJmzSpAlu3LiBkpISLte243mmHQCM\nGDEC69atg76+Prp164Z///0XM2bMwHvvvafr0CSFig2RhO3bt0u22ADlL8i9e/dGSEiISpuQhxdk\nnmfaAcClS5dgbm6OLVu2ICgoCEuWLEGXLl2o2NQRFRtOVJ48QBoeNzc3uLm5oaysDIWFhboOp15V\nLFTJK4VCgadPn2LXrl2YMmUKDAwMuCqm2kLFRiI0rR32wQcf6DhCUpMOHTpg+PDhKpdt375dR9HU\nj8Yw0w4AoqOj4eLiAm9vb/Tp0wdyuZzGbJ4DTRCQCN7XDtNE6hMg1MUv9Zwaw0w7dRhjUCqVaNKE\n3qvXBT1aEsH72mGaSLVNuH//fiQlJSEnJwfTp08XXpQLCgok//z5+vpCoVBg3bp1XM60++677zB6\n9GiVr1CoeP54GW/TJj1dB0Bqh/e1w65evYp+/fqhY8eOAIALFy4Iy/IA0m0TOjg4wNfXF4aGhvD1\n9RV+QkJC8Ouvv+o6vBdWeaYdb4qKigCUvzGo+CksLERhYSEKCgp0HJ30UBtNIjIzMxEVFYXjx4+j\nefPmwtphLi4uug6tXvDeJnz69KlwJvPgwQNkZ2fD29tbx1HVj9GjR+PKlStczrSrDSlPy9cmaqNJ\nhJubGw4ePIjHjx+jrKyMu6+H5r1N2L9/f26/IoLnmXa1IfVp+dpCxUYiHj58iM2bN0Mul0OhUADg\n62uheW8T8vwVETzOtCP1j8ZsJGLQoEHIysqCt7c3unbtKvT+eREfH4/o6GhcuXIFDg4OWLlyJdau\nXQ3D/kkAAAn9SURBVKvrsOpN5a+IeO211wDw8xURixcvrtVlpHGjMxuJKCkpwWeffabrMETDe5tw\n7ty5GDhwIHr16oXu3bsjMzMTbdu21XVYL4TnmXak/tEEAYlYvnw5zM3NERwcrLK+VosWLXQYVf3h\nvU2oiRQHmf/880+cO3cOc+fOxccffywUG3Nzc/Tt2xfNmzfXcYTasWjRIsnOltQmKjYSER8fjw8/\n/BCWlpbQ0yvvfvL01bR+fn7w8/ODl5cX9PT0hPW1IiIidB2aVkj5A548z7QDNK/eQWqHio1EuLq6\n4vTp07C2ttZ1KKJo7N98KOViExgYyO1MO4D/afnaQhMEJKJt27YwMjLSdRiiGTlyJNavX4/c3Fw8\nePBA+CENX8VMu59//hljxozBqVOncODAAV2HVW94n5avLTRBQCKMjY3h4+ODvn37CmM2PI1pGBoa\n4t1338Unn3zCZZuQZ5Vn2lWs+sDLTDuA/2n52kLFRiJCQ0MRGhqqskYTT//QK1asQGZmJrdtQk2k\nuvYbwOdMu8ri4+MRFRUlTMuvWL2D1A2N2UhISUkJ0tPTAQDt27fn6lR+wIAB2LlzJ0xMTHQdiiga\n8yCzFGfaqcPrtHxtoWIjEampqYiIiICzszMA4MaNG9i0aRM3y7iHhobi4sWL3LYJG/Mgs5QnPwA0\nLb++UBtNImJiYpCSkgIPDw8AQHp6OsLCwriZwcV7m5AGmaVr0KBB8PPzg7e3t8q0fFI3VGwkQqFQ\nCIUGANq1aye8y+JBZGQk121CGmSWLt5X79AWKjYS4evriwkTJiA8PByMMWzZsgVdu3bVdVj1hvc2\nIQ0yS1fFtHxeV+/QFhqzkYiSkhLEx8fj2LFjAAB/f3+89dZbKn/8UtalSxckJiZy2yas0BgHmaW+\nnAvvq3doCxUbCVAoFHjppZdw5coVXYciGm9vb1y4cEHjZVLF8yAz7zPteF+9Q1toBQEJaNKkCTw8\nPJCVlaXrUERT0SZMTU3FoUOHMGHCBK7ahDx/RcTEiROxaNEiNG3aFADg5eWFxMREHUdVf3hfvUNb\naMxGIh48eICOHTuie/fuwmdRZDIZ9uzZo+PI6sdXX32F+Ph44Z1+RZuQFzwPMvM+04731Tu0hYqN\nRHz88ce6DkE0CoUCnTp1wpUrVzBr1ixdhyMKngeZeZ9px/u0fG2hMRvSIAwZMgSrV68WZqPxhudB\n5szMTERFReH48eNo3ry5MNPOxcVF16HVG56n5WsLFZsGztTUtNp3UTKZDI8ePdJyROLw9/fHuXPn\nuG0TNoZBZl5n2vE+LV9bqI3WwBUWFgIA5syZAwcHB4SHhwMAtmzZglu3bukytHrFc5sQ4HuQmeeZ\ndgD/q3doC53ZSATvU4N5x/Pab7x/yyr979UPOrORCBMTE3z//fcYMWIEAOCHH36AqampjqN6cY2l\nTcjzIDPPM+0A/lfv0BY6s5GI69evY8aMGTh+/DgAoFevXli1ahU3g7DVtQl5aq/xOsi8fPlymJub\ncznTDuB/9Q5toWJDGgTeWxU8DzLzPNOuMazeoS3URpOIO3fuYMOGDVUGYb/99lsdR1Y/eG0TVuB5\nkJnnb1mtvHoHr9PytYWKjUQMGTIEffr0Qf/+/VXePfJi69atmDFjBmbOnAmgvE24detWHUdVf3j+\nigieZ9oB/K/eoS3URpMIHx8fnD9/XtdhkOc0duxY6Ovrqwwyl5WVcXFmyvNMO6C8BapOYGCgVuOQ\nOio2EjFnzhz4+fnhtdde03UoouC9TcjzIPPGjRsBoMpMO16mPpP6QcVGIkxNTVFUVISmTZsKs5h4\nmhrs5+eHPn36wNfXV6VNOHToUB1H9uIawyAzjzPtGsu0fG2hMRuJqFhJgFdPnjzB0qVLdR2GKHgf\nZOZ1pl1jWb1DW+jMpoG7fPkyPD09q5211KVLFy1HJA7e24Q8r/3G+7es8j4tX1vozKaB++yzz7Bh\nwwbExMSoPaU/dOiQDqKqf59//rnwBVw8tgl5+nDqs3ieaQfwPy1fW+jMhhDyQnieaQfwv3qHtlCx\naeB++umnGj9P89///leL0dQ/3tuEjWGQmeeZdqT+ULFp4CIjI2ssNgkJCVqMpv5NnDgRGzZsQGBg\nINdtQl7XfmsMM+14n5avLVRsCNECngeZef+WVZ6n5WsTTRCQkH379uHSpUsoLi4WLps7d64OI3px\nvLcJK/A8yMz7ci48T8vXJio2EhEdHY0nT57gt99+w8SJE7Fjxw706NFD12G9sL179zaKYsPz2m9S\nbwVqMnjwYPzyyy/cTsvXFmqjSYSXlxf++usvofVSWFiIV199FUePHtV1aIRwjffVO7SFzmwkomJV\nXSMjI+Tk5MDKygp5eXk6jqp+8dgmrMDjIHNjmGkH8L96h7ZQsZGIwYMH4+HDh3jvvffg6+sLoHwm\nFy94bRNW4PErInhfzoX3afnaRm00iXjy5AnWrFmDo0ePQiaToXfv3pg8eTI33yPCe5uQ56+I4HWm\nXWOZlq8tdGYjEWPGjIG5uTmmT58Oxhi2bt2KMWPGYMeOHboOrV7w3ibkeZCZ15l2GzZsAFD999mQ\nuqEzG4no0KEDLl26pPEyqVqwYAGmTZuG3377DVOmTAFQ/s6Sl5lOPA8y87qcS2OZlq8tVGwkIjw8\nHFOmTIGfnx8AIC0tDV9++SW+++47HUdWP3hvExLp4X31Dm2jYiMR7du3R3p6Olq3bg2ZTIYbN27A\nw8MDTZo0gUwmk3x/fNiwYTA3NxcWc9y6dSv+/fdfybcJG8MgM48z7Uj9o2IjEXK5vMbrpd6y4LVN\n2BgGmRvDci48T8vXFpogIBFSLyaadOnSBSdOnFBpE1ZM8ZayxjDIzPtyLrxPy9cWOrMhDQKvbcLG\nMMjM+7es8j4tX1vozIY0CMnJyboOQRSNYe033r9llfdp+dpCxYY0CLy2CTdu3KjrEETH+3IuvK/e\noS3URiNES3gbZG4MM+0AmpZfX6jYEKIF1Q0yf/PNN7oO7bk1hpl2AL/T8rWNig0hWkCDzNLF67R8\nbaMxG0K0gMdB5sYw0w7gd1q+tlGxIUQLeBxkbgwz7QDgzJkz6NWrV5Vp+V5eXpKelq9t1EYjRAto\nkFm6eF+9Q1uo2BCiBbwPMvM2047UP2qjEaIFFy9eVBlQ/s9//oMOHTroMKL6Q8u5kNrQ03UAhDQG\nFYPMFXgaZD5+/Dg2b96MFi1aYN68eUhLS8PVq1d1HRZpYOjMhhAt4HmQmceZdqT+UbEhRAt4XfsN\n4HOmHal/NEGAEPJCaKYdqQ0qNoSQF8L7TDtSP6jYEEJeCC3nQmqDZqMRQl4IzzPtSP2hMxtCyAvh\n9VtWSf2iYkMIeSG0nAupDSo2hBBCREdjNoQQQkRHxYYQQojoqNgQQggRHRUbQgghoqNiQwghRHT/\nDzH9gT2pJYOrAAAAAElFTkSuQmCC\n", + "text": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 1, + "metadata": {}, + "source": [ + "4 Simple Tricks To Speed up the Sum Calculation in Pandas" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "I wanted to improve the performance of some passages in my code a little bit and found that some simple tweaks can speed up the `pandas` section significantly. I thought that it might be one useful thing to share -- and no Cython or just-in-time compilation is required! " + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "In my case, I had a large dataframe where I wanted to calculate the sum of specific columns for different combinations of rows (approx. 100,000,000 of them, that's why I was looking for ways to speed it up). Anyway, below is a simple toy DataFrame to explore the `.sum()` method a little bit." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "import pandas as pd\n", + "import numpy as np\n", + "\n", + "df = pd.DataFrame()\n", + "\n", + "for col in ('a', 'b', 'c', 'd'):\n", + " df[col] = pd.Series(range(1000), index=range(1000))" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 2 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "df.tail()" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "html": [ + "
| \n", + " | a | \n", + "b | \n", + "c | \n", + "d | \n", + "
|---|---|---|---|---|
| 995 | \n", + "995 | \n", + "995 | \n", + "995 | \n", + "995 | \n", + "
| 996 | \n", + "996 | \n", + "996 | \n", + "996 | \n", + "996 | \n", + "
| 997 | \n", + "997 | \n", + "997 | \n", + "997 | \n", + "997 | \n", + "
| 998 | \n", + "998 | \n", + "998 | \n", + "998 | \n", + "998 | \n", + "
| 999 | \n", + "999 | \n", + "999 | \n", + "999 | \n", + "999 | \n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Let's assume we are interested in calculating the sum of column `a`, `c`, and `d`, which would look like this:" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "df.loc[:, ['a', 'c', 'd']].sum(axis=0)" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "pyout", + "prompt_number": 4, + "text": [ + "a 499500\n", + "c 499500\n", + "d 499500\n", + "dtype: int64" + ] + } + ], + "prompt_number": 4 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Now, the `.loc` method is probably the most \"costliest\" one for this kind of operation. Since we are only intersted in the resulting numbers (i.e., the column sums), there is no need to make a copy of the array. Anyway, let's use the method above as a reference for comparison:" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "# 1\n", + "%timeit -n 1000 -r 5 df.loc[:, ['a', 'c', 'd']].sum(axis=0)" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "1000 loops, best of 5: 1.37 ms per loop\n" + ] + } + ], + "prompt_number": 5 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Although this is a rather small DataFrame (1000 x 4), let's see by how much we can speed it up using a different slicing method:" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "# 2\n", + "%timeit -n 1000 -r 5 df[['a', 'c', 'd']].sum(axis=0)" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "1000 loops, best of 5: 986 \u00b5s per loop\n" + ] + } + ], + "prompt_number": 6 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Next, let us use the Numpy representation of the `NDFrame` via the `.values` attribue:" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "# 3\n", + "%timeit -n 1000 -r 5 df[['a', 'c', 'd']].values.sum(axis=0)" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "1000 loops, best of 5: 687 \u00b5s per loop\n" + ] + } + ], + "prompt_number": 7 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "While the speed improvements in #2 and #3 were not really a surprise, the next \"trick\" surprised me a little bit. Here, we are calculating the sum of each column separately rather than slicing the array." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "[df[col].values.sum(axis=0) for col in ('a', 'c', 'd')]" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "pyout", + "prompt_number": 8, + "text": [ + "[499500, 499500, 499500]" + ] + } + ], + "prompt_number": 8 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "# 4\n", + "%timeit -n 1000 -r 5 [df[col].values.sum(axis=0) for col in ('a', 'c', 'd')]" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "1000 loops, best of 5: 64.4 \u00b5s per loop\n" + ] + } + ], + "prompt_number": 9 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "In this case, this is an almost 10x improvement!" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "One more thing: Let's try the Einstein summation convention [`einsum`](http://docs.scipy.org/doc/numpy/reference/generated/numpy.einsum.html)." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "from numpy import einsum\n", + "[einsum('i->', df[col].values) for col in ('a', 'c', 'd')]" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "pyout", + "prompt_number": 10, + "text": [ + "[499500, 499500, 499500]" + ] + } + ], + "prompt_number": 10 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "# 5\n", + "%timeit -n 1000 -r 5 [einsum('i->', df[col].values) for col in ('a', 'c', 'd')]" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "1000 loops, best of 5: 55.7 \u00b5s per loop\n" + ] + } + ], + "prompt_number": 11 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "heading", + "level": 3, + "metadata": {}, + "source": [ + "Conclusion:" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Using some simple tricks, the column sum calculation improved from 1370 to 57.7 \u00b5s per loop (approx. 25x faster!)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "heading", + "level": 3, + "metadata": {}, + "source": [ + "What about larger DataFrames?" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "So, what does this trend look like for larger DataFrames?" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "import timeit\n", + "import random\n", + "from numpy import einsum\n", + "import pandas as pd\n", + "\n", + "def run_loc_sum(df):\n", + " return df.loc[:, ['a', 'c', 'd']].sum(axis=0)\n", + "\n", + "def run_einsum(df):\n", + " return [einsum('i->', df[col].values) for col in ('a', 'c', 'd')]\n", + "\n", + "orders = [10**i for i in range(4, 8)]\n", + "loc_res = []\n", + "einsum_res = []\n", + "\n", + "for n in orders:\n", + "\n", + " df = pd.DataFrame()\n", + " for col in ('a', 'b', 'c', 'd'):\n", + " df[col] = pd.Series(range(n), index=range(n))\n", + " \n", + " print('n=%s (%s of %s)' %(n, orders.index(n)+1, len(orders)))\n", + "\n", + " loc_res.append(min(timeit.Timer('run_loc_sum(df)' , \n", + " 'from __main__ import run_loc_sum, df').repeat(repeat=5, number=1)))\n", + "\n", + " einsum_res.append(min(timeit.Timer('run_einsum(df)' , \n", + " 'from __main__ import run_einsum, df').repeat(repeat=5, number=1)))\n", + "\n", + "print('finished')" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "n=10000 (1 of 4)\n", + "n=100000 (2 of 4)" + ] + }, + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "\n", + "n=1000000 (3 of 4)" + ] + }, + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "\n", + "n=10000000 (4 of 4)" + ] + }, + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "\n", + "finished" + ] + }, + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "\n" + ] + } + ], + "prompt_number": 23 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%matplotlib inline" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 24 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "from matplotlib import pyplot as plt\n", + "\n", + "def plot_1():\n", + " \n", + " fig = plt.figure(figsize=(12,6))\n", + " \n", + " plt.plot(orders, loc_res, \n", + " label=\"df.loc[:, ['a', 'c', 'd']].sum(axis=0)\", \n", + " lw=2, alpha=0.6)\n", + " plt.plot(orders,einsum_res, \n", + " label=\"[einsum('i->', df[col].values) for col in ('a', 'c', 'd')]\", \n", + " lw=2, alpha=0.6)\n", + "\n", + " plt.title('Pandas Column Sums', fontsize=20)\n", + " plt.xlim([min(orders), max(orders)])\n", + " plt.grid()\n", + "\n", + " #plt.xscale('log')\n", + " plt.ticklabel_format(style='plain', axis='x')\n", + " plt.legend(loc='upper left', fontsize=14)\n", + " plt.xlabel('Number of rows', fontsize=16)\n", + " plt.ylabel('time in seconds', fontsize=16)\n", + " \n", + " plt.tight_layout()\n", + " plt.show()\n", + " \n", + "plot_1()" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "display_data", + "png": "iVBORw0KGgoAAAANSUhEUgAAA1cAAAGpCAYAAABhxcywAAAABHNCSVQICAgIfAhkiAAAAAlwSFlz\nAAALEgAACxIB0t1+/AAAIABJREFUeJzs3Xd4VFX+x/H3SSihJDSpkhCKVEEIRYUICazYUNdVWYpS\nbKyKLq4FAV2IKOiKLBZsiwKKKLpi/+EqQghBpYsg0oQQJRQFpHfO748zGVMmPZlJ+byeJ0+45965\n99yZM2S+c875HmOtRURERERERAomKNAVEBERERERKQ0UXImIiIiIiBQCBVciIiIiIiKFQMGViIiI\niIhIIVBwJSIiIiIiUggUXImIiIiIiBQCBVciIqWEMWaGMeasMSYi0HXxF2NMvDHmbKDrISIiAgqu\nRESy5QlW0v6cNsb8aoz5yhjTP9D186FYLV5ojAk3xjxpjFlpjNlvjDlpjNltjPnSGHOvMSasEC5T\nrO65KBhjOhtj3jLGbDfGHDfGHDDGbDHGfGKMedAYUznQdRQRESgX6AqIiJQAFojz/Ls80Aq4Fog1\nxnSy1t4fsJplZgJdgVTGmNuAF4AKwHfAW8B+oCbQDZgCPArUDlQdSwJjzE3ATFw7XAC8DxwDIoFo\n4EpP2dYAVVFERDwUXImI5IK19rG028aYnsCXwAhjzHPW2u2BqVnxZIwZCLwK7ANuttbO83HMhcBU\nf9etJPH0SE0FzgCXWWsX+jjmImCvv+smIiKZaVigiEg+WGsXABtxPUWdAIwxfzbGzDLGbDLGHPb8\nrDDG3GOMydSjlGaOVCNjzDBjzFpjzDFjzC5jzCtZDZkzxvzJGLPYGHPEGLPXGPOBMaZlVnU1xgwx\nxrxvjNlqjDnqGVKW6AmAfB3fxBjzqmfY2VHPNb43xrxkjKmZ03NjjAkFnsP1tPTzFVh5nsOlQFcf\nj+9ljPncGLPPMwRuozFmYm6HEHru96wxZnAW+88aYxZmKBvnKe9hjOnved2OGGNSjDHPGGMqeI77\nkzFmkTHmoGeY45u+nhNjTJIxZpsxprIx5mljTLLnXjYbYx7KzX14nA+EAut8BVYA1tpvrbUH0lw7\n0nMv07O4/0zz1IwxMZ7HjDXGdPI8/7977vF9Y0y457imxpg5nqGxR40xC40x7Xxco64xZpLntTvs\nOc8GY8x0Y0zjPNy/iEiJop4rEZH8Sw2YUuf8TMT1MHwD7ACqAb2AZ4HOwKAszvM00Bv4GPgc6Anc\nDjTzPP6PCxpzAzAHOO75vRO4BPga+D6L878IrAPiPcefgxtK9qYxpoW19p9pzl8fWI77QP8Z8B4Q\nAjQBbgKex/VGZecGoAbwjbV2fnYHWmtPZri/YcBLwCHPtfcAscBI4GpjTLe0gUQOspuLldW+e4Ar\ngA+AhcBlwH1AbWPMx8As4FPgZdzQxoFALdzzmfH85YEvgPq45/I0cB3wpDEmJGNvaBZ+8/xuYIyp\nbK09movHpK1DXvd1xj3X8biex3a4Orc1xlwHJAA/ADNwwxL/AnxpjGlirT0C3t62Jbg28wXwEe69\nEglcg3tdt+XhPkRESg5rrX70ox/96CeLH+AscMZH+Z88+04D4Z6yxj6OM7gPomeBLhn2pZYnAQ3T\nlAcDizz7Oqcpr4ob/nUCiMpwrsmpdQUiMuzzVa/ywHzgJNAgTfk9nvPc4+MxlYCQXDxnr3nO8Vge\nn+tGnnv7HWieYd9UzzlfyVAen/H1AYZ4jh2UzWu6IEPZOE/5fqBFmvIKuMD0DC6ovCTDa/uF53EX\nZDhfkqf8U6BimvLanmvsB8rl8nlZ6jnXauAuoD1QIZvjIz3Hv57Ffl/PWYznMWeB/hn2TfOU/w6M\nyrDvEc++e9OUXe0pe8bHtcsBVQv6vtSPfvSjn+L6o2GBIiI5M57hUuOMMU8YY/6L62GywBRr7c8A\n1tpM38Zbay1uiBy43ilfHrPW/pLmMWeA1CFdndMcdy2uR2i2tXZVhnOMAw76OnkW9TqF69EqR/re\nsdQejeM+HnPMWpup3If6nt+/ZHtUZjfhgr4XrLWbMuwbAxwGbkodoldEnrPWbkzdsK5nbQ4ukPrY\nWrs4zT6L68kC18OTkcUFHSfSPOZXXA9lNaB5Lut0Ay4gugCXIGQVcNgYs9QY85BnGGZhWWytfTtD\n2UzP773Akxn2veH5fYGPc/lqQ6ettYcLVkURkeJLwwJFRHJnrOe3xfU6LAJes9bOTj3AGFMLeBA3\nRKwJkDE99rlZnHuFj7LUwKRGmrIoz+9FGQ+21h40xnwHdM+4z7h1r0bigqhwXA9UWg3S/PtjYAIw\n1RhzGa5nJtFauz6Luhem1PtbkHGHtfZ3Y8xq3BDIlmQ9BLKgfL0WOz2/V/rYl+L53dDHvgPWWl8Z\n/H72/K7hY18mnuC9p2de3aVAR6ALLvDuDNxljImx1ibl5nw5yO7+v/MElGn5uv943LDYh40xUcA8\nINHzeK1JJiKlmoIrEZGcWWttcHYHGGOq4+YqReKGcc3ADSM7jfsQ/XegYhYP/91H2WnP77TXreb5\nvTuL8+zyUa8mwDKgOm6+zOfAAdwwt8bA4LT1stYmG2O64HrCLsfNqcEY8zMwyVr7fBbXTiv1w7iv\ngCM7qfe3M4v9OzMcVxR8zec6nYt95X3s8/W6pn1Mtm0qI2vtBmBD6rYxpgXwOnAx8G/c3KiCytP9\nW2tPG5erpXyaskPGZTCMw82xusyz6zdjzIvA49ba0xnPJSJSGii4EhEpHLfhAqtxNnPa9otxwVVB\npX64rZvF/no+yv6BW1dqiLX2jbQ7jFsEOVNGPc+H+H7GmGDccK8/4eZiPWuMOWKtfT2Hei4GhuJ6\nyv6Zw7Fppd5ffeBHH/vrZzguK6m9I5n+xnmC4FLBWrvRGHMzsAWX9CNVlvfvUeTPgbV2B+49gTGm\nNS5Jy9249hBE3tqFiEiJoTlXIiKFo5nn9/s+9vUopGukDkuLybjDGFMNl+gg47CtZp6yPNfLWnvG\nWrvKWvsvoL+n+Npc1PO/uF67i40xvbI7MMP8qdR5ZDE+jquOu79j+A680trv+R3hY1+nHB5b0qTO\nX0qb6j/1/sMzHuxJZ5/buV6Fwlq73lr7Am5II+SuDYmIlEgKrkRECkdq0oi0PQgYYzoAowrpGh/h\nPjgPMMZ0zLBvHOBrHahtuA/eGet1GZ6ehQzlUZ5ALaPUXrEcU4F7Ehbc69mcY4zxmcjD06O3NE3R\nLOAUcI8xpmmGw8fj0sPP8iTjyM5yXO/NAGOMd36ZZz2qf+VU/+LEs2bVvb7W+DJuPN4Yz2ZCarm1\n9hBu+GC0MaZVmuODcVklQ4q4zq2NMb56V3PdhkRESioNCxQRKRxv4JJZTDHGxOKGap0HXIXrNepX\n0AtYa48YY+7AZa9bbIyZg5tnFQ20wX3AzpjQ4kXcEL33PFkOd+IWpr0MeBf4a4bjBwF3GGMSga24\nYK4pLr32cWBKLus62xPYvAB87km28Y3nfLVw84TaAb+mecx2Y8wIXNr1VcaYd3HrPPUALsL1WI30\ncbl0CzRba3cZY94Cbga+M8b8Hy7wvAKXDKR9bu6hmKiOe87/ZYxZgltj6hBQBzfUrjFuDt79GR73\nNC4l/hLP634cF2AHA2vwnd2vsPQGnjbGfA1sxq1V1hDXY3XGUzcRkVJJwZWISCGw1u40xlyCS1Ud\njQtefgTuBL7Cd3BlyX6hV1/Xed8Yczkue2Ff3IfmBFzwMQqXTS/t8Ws9wd7juECvHPAdLvnBATIH\nV7Nxazt1xWWlq4TLXDgbt25RrrMGWmtfM8b8DxiOGxI2AKiCC7DWASNwCRnSPuYlY8wW4AHgelzG\nxWRcj9MEa23GdPNZPYe344KO/ri1obbjFnOe5OOesztPTvuyktMCvrk933rca9Ub9xr3xc2hO4IL\n4N8CnrXW7k13AWune3q2/oELmPfhej7H4IL9vN5PXnyOG5LYHZfQIgyXVfB/wGRr7bdFeG0RkYAy\nmbOqFvEFjXkd9wd+j7W2rY/9A4GHcN9EHgLutNYWVcpdERERERGRQhGIOVfTcel9s7IV6G6tbYcb\nY/+qX2olIiIiIiJSAH4Prjyr2+/PZv831trUNLtLyfs6KSIiIiIiIn5X3LMF3gr8X6ArISIiIiIi\nkpNim9DCMwH7FqBboOsiIiIiIiKSk2IZXBlj2gH/AS631vocQmiM8W8mDhERERERKRGstSbnowpf\nsQuujDERwFzgJmvtluyO9XemQymbhgwZwowZMwJdDSkj1N7EX9TWxJ/U3sSf3EoUgeH34MoY8zZu\nQchzjDE/49ZqKQ9grX0F+CdQA3jJ88ScstZ28Xc9RVJFRkYGugpShqi9ib+orYk/qb1JWeH34Mpa\n2z+H/bcBt/mpOiIiIiIiIoWiuGcLFAm46tWrB7oKUoaovYm/qK2JP6m9SVmh4EokB+3btw90FaQM\nUXsTf1FbE39Se5OywpTUpBDGGFtS6y4iIiIiIkXDGKNsgYUpkBlCRESk9NCXeCIikhelMrgC/UEU\nEZGCKe1f1MXHxxMTExPoakgZofYmZYXmXImIiIiIiBSCUjnnyjPO0s81EhGR0kR/S0RESqZAzrlS\nz5WIiIiIiEghUHAlIiJSBsXHxwe6ClKGqL1JWaHgSkREREREpBAouCrm+vTpw9ChQ73bR48e5YYb\nbqB69eoEBQWRnJyc7vj4+HiCgoLYt29foVw/9XxBQUFceeWVBT5fZGQkQUFBBAcHs2fPnkKoYcky\nfPhwYmNjs9wfExPjfb5XrVoFpH8Nrr76an9VNc9uvfVWxo0bV6jnbNy4MZMnTy7Uc+bWiRMnCA8P\n57vvvgvI9UWKmjK3iT+pvUlZoeCqmDPGpEsH/Prrr7N48WKWLFnCrl27aNiwoV/qsX79et5+++0C\nn8cYw9ixY9m5cye1a9cGICkpiaCgvDXFIUOGEBcXl6fHxMTEMHPmzFwdGx8fT+PGjfN0/hkzZmQb\nOKVK+3pmvI8PP/yQZcuWpTu+W7du7Ny5k759+xbb1NAbN27k/fff57777ivU865YsYI777yzUM+Z\n1rhx4zj33HOpXLkysbGxrF+/3ruvYsWK3HfffYwZM6bIri8iIiKli4KrEmbLli20atWKNm3aUKdO\nnTwHJflVp04dqlWrVijnCg0NpU6dOgUKFDIGnUX1mKKQNvtYxjpVr16dc845J93x5cuXp27duoSE\nhBTbzGUvvvgiV199daG1kVS1atWiUqVKhXrOVE899RSTJ0/mhRdeYPny5dSpU4dLL72Uw4cPe48Z\nMGAAX375Jdu2bSuSOogEkubAiD+pvUlZoeCqGDl69ChDhgwhNDSUevXqMXHixHT7Y2JieO6550hI\nSCAoKIiePXvm6rxz586lbdu2hISEEBERwYQJE9LtP3nyJKNHjyYyMpKQkBCaNm3K888/X2j3lVf7\n9u2jf//+hIeHU7lyZc4//3xmzJiR6bj8BBppH3PgwAHuvPNOGjRoQKVKlWjdujXvvvtuQaqezpkz\nZ3jggQeoWbMmNWvW5L777uPMmTPZ1qkgDhw4wM0330zdunWpVKkSTZs25dlnn/XuDwoKYu7cueke\nExkZyTPPPJPumJdffplrrrmGKlWq0KJFC+Lj40lOTqZ3795UrVqVqKgovv/++3TnmTNnDtdcc026\nslmzZtG5c2fCwsKoW7cuffv2JSUlxbt//Pjx1K9fn19//dVb1r9/fzp27Mjp06d91u+VV16hefPm\nVKpUidq1a3P55Zf7fE5zYq1lypQpjBo1iuuuu442bdowc+ZMDh06xOzZs73H1atXj86dO/POO+/k\n+RoiIiJS9pQLdAUCZdiwwj/nK68U7PEPPPAA8+fPZ+7cuTRo0IC4uDgSEhK4/vrrAfjggw944IEH\n2LhxI3PnzqVChQo5nnPlypX07duXRx99lIEDB7Js2TKGDRtGWFgYw4cPB2Dw4MEkJiby3HPP0aFD\nB3755Re2b9+e7Xnj4+Pp2bMn8fHxdO/evWA3TvqhcsePH6dTp06MGjWKsLAwvvzyS4YNG0ZERES6\ngDI/vVCpj7HWcuWVV3LgwAFmzJhBixYt2LRpE0ePHs33+TP2Qj3zzDNMmzaNadOm0a5dO1544QVm\nz55Nx44dfdapoB555BHWrVvHZ599Rt26ddm6dWu6wCU3dQZ4/PHHeeaZZ5g8eTJjx46lf//+tGnT\nhr///e9MnTqVESNGMHjwYFavXg3Ahg0b2LNnD507d053nlOnTjF+/HhatmzJr7/+ysiRI+nfvz+L\nFi0CYMyYMXz55ZfccsstfPLJJ7zxxht8/PHHrF69mnLlymWq34oVKxg+fDhvvPEG0dHR7N+/n4UL\nF3qv99Zbb/G3v/0t2/t99dVX6d+/P9u2bWP37t307t3buy8kJITu3bvz9ddfc8cdd3jLu3TpwqJF\nixg1alS25xYpaTQHRvxJ7U3KijIbXBU3hw8f5vXXX2f69OlceumlAEyfPj3dnKoaNWpQqVIlypcv\nT506dXJ13smTJxMTE8PYsWMBaNasGZs3b+app55i+PDhbN68mTlz5vD55597P2hGRkYSHR2d7Xmr\nVKlCy5YtqVy5cn5uN53IyMh0vQ8NGjTg/vvv927ffvvtLFiwgLffftsbXE2fPj3P10n7QXz+/Pl8\n++23rF+/nhYtWgDQqFEj7/6YmBi2bt2ap/MPHjyYwYMHe7enTJnCyJEjueGGGwB49tln+d///pfu\nMfm5j6wkJycTFRVFp06dAAgPD8/XeQYPHsxf//pXAEaPHs3bb7/NyJEjvck0HnroIWJjY9m3bx81\na9Zk8+bNAERERKQ7T9pELJGRkbz44ou0bt2alJQUGjRoQFBQELNmzeKCCy7goYce4pVXXmHy5Mk0\nb948y/urUqUKV199NVWrViU8PJx27dp591977bVcfPHF2d5b6vtm165dANStWzfT/rS9a+Cex48+\n+ijb84qIiIhAGQ6uCtrLVNh++uknTp48me7DYZUqVWjbtm2Bzrthwwb69OmTrqxbt27ExcVx+PBh\nVq9eTVBQUK4SMaTVuXPndJP/C9OZM2d48sknmTNnDikpKZw4cYKTJ0/muY7ZWb16NfXr1/cGVoXt\nwIED7Nq1K93raYzhwgsv5Oeffy6Sa955553ccMMNrFy5kksvvZSrr746X72KaQOW1GAkbTtMLduz\nZw81a9bk4MGDVKxYMdP8v1WrVhEXF8eaNWvYt2+fd/hjcnIyDRo0AFxA9uyzzzJkyBD69OnDsGy6\nlHv37k2jRo1o3Lgxl112Gb179+Yvf/kLVatWBaBq1arefxdExp68sLAwDhw4UODzihQ38fHx6k0Q\nv1F7k7JCc66KucKYj5PVOYpDcgdfJk2axOTJkxk5ciQLFixgzZo1/PnPf+bEiROBrlqBFWVCissv\nv5zt27fzwAMP8Ntvv3HVVVdxyy23ePcbYzJd/9SpU5nOU758+XSPyars7NmzAFSrVo0TJ054twGO\nHDnCZZddRtWqVZk1axYrVqzg888/B9wcv7QWLVpEcHAwycnJmfalVbVqVVatWsW7775LREQEEydO\npGXLluzcuRNwwwJDQ0Oz/UnNeFmvXj0Adu/ene4au3fv9u5LdfDgQapXr55lvURERERSKbgqJpo2\nbUr58uX55ptvvGVHjhxh3bp1BTpvq1atWLJkSbqyxMREwsPDqVKlCu3bt+fs2bMsWLCgQNcpTImJ\niVxzzTUMHDiQdu3a0bhxYzZu3FiowWBUVBQ7d+5kw4YNhXbOtKpVq0b9+vXTvZ7WWpYtW1akQW2t\nWrW46aabmD59OtOmTWPmzJneAKp27drphrzt3r3bG5gURLNmzQDSrbm2YcMG9u7dy4QJE4iOjqZ5\n8+aZAhlwyVZmz57NwoULOXDgQI7zmoKDg4mNjWXChAl8//33HDlyhM8++wxwwwLXrFmT7U/q0MbG\njRtTr149vvjiC++5jx8/TmJiIl27dk13ze3bt2c5VFGkJFMvgviT2puUFWV2WGBxU7VqVW699VZG\njhxJ7dq1qV+/Po899li63gBfRo0axfLly5k/f77P/ffffz+dO3cmLi6O/v37s3z5ciZPnuzNRNi8\neXP69u3LbbfdxrPPPpsuocVNN92U5XWXLVvGoEGDePPNNzMlMiioFi1aMGfOHJYsWUKtWrV4/vnn\nSUpKombNmoV2jV69enHhhRdy/fXX8+9//5vzzjuPLVu2cPToUa699tpCucbf//53Jk6cSPPmzTn/\n/PN58cUX2bVrF+eee26hnD/ja//Pf/6Tjh070rp1a06fPs3cuXO9QTtAz549mTp1Kl27diUoKIjR\no0cTEhJS4Hq0aNGC2rVrs2zZMiIjIwE33K9ixYo8//zz3HXXXfz44488+uij6R63Y8cObr/9diZO\nnEh0dDRvvvkmPXv25Morr6RXr15A+p6+Tz/9lJ9++onu3btTs2ZNFi5cyKFDh2jVqhWQt2GBxhhG\njBjBhAkTaNmyJeeddx6PP/44oaGhDBgwIN2xy5Yty5QJUURERIqnNLnJAkI9V8XIpEmTiI2N5brr\nrqNXr160a9cu05yZjNnddu3alSnxQtr9HTp04L333uP999+nbdu2jB49mlGjRnH33Xd7j3njjTcY\nMGAA9957L61atWLo0KEcPHgw27oePXqUzZs3c+zYMW/ZuHHjCmXdrUceeYQuXbpwxRVX0KNHD0JD\nQxk4cGC2j8nrtY0xzJs3j27dunHTTTfRunVr7rvvPp/D5OCPhY7feOONXF/j/vvvZ+jQodx2221c\ndNFFADneR051Tivjax8SEsKYMWNo37490dHRHDlyhE8++cS7/5lnnqFJkybExMTQt29fbr/99lwl\nRvHV05a2zBhDv379+Pjjj71ltWvXZubMmXz44Ye0adOG8ePH8+9//zvTAsodO3ZkxIgRAERHR/Pw\nww8zZMgQ9u/fn+k6NWrU4KOPPuLSSy+lVatWTJ48mddee41u3brleA++PPTQQ9x3333cfffddO7c\nmd27d/PFF19QpUoV7zG7d+9m5cqV9OvXL1/XECnOtO6Q+JPamxS1Xbtg9mwYOTKw9TDFdVHSnBhj\nbHZziUrqfRU3qSnXf/31V2rVqpXtsYMHD2bPnj3Mmzcvy2MaN27M8OHD02UDLAy5uXZBLFy4kKuu\nuor169d7e2eKSlJSEk2aNGHFihVERUV5y4cMGcLevXvTBUzFxcaNG7nwwgtJSkoqVfOTnnnmGRYs\nWOAdeihlS2n/W6IEA+JPam9SFKyFH36ABQvc71Svvmqw1gYkuYB6riRbqT0HkZGR3vW2fLHWsnDh\nwlwtPjxmzBhCQ0P57bffCqWOebl2fs2bN4+HH364yAOrK664gvPPPz9dj83ixYupWrUqs2fPLrZJ\nSFq0aMENN9yQbtHiku7EiRNMmTKFJ554ItBVESkS+qAr/qT2JoXpxAlYtAjGjYPnn3eBVYUK0L27\nKwsk9VxJto4fP+5NglClSpVM6wLlVXJyMqdPnwZcwFYYwwhLk5SUFI4fPw5Aw4YNqVChQqG/BiKS\nO/pbIiJSvOzdCwsXQmIipM5MqVkTYmIgOhpSR/Z7/v8OyDfSCq5ERER8KO1/SzRMS/xJ7U3yy1rY\nvNkN/VuzBlJzvTVrBj17QocOkPG7+kAGV8oWKCIiIiIixcqpU7BiBXz1Ffz8sysLDoYLL4RevaBR\no8DWLyvquRIREfFBf0tERPzvwAE3nyohAQ4dcmVhYW4+VffuUK1azudQz5WIiIiIiJRZ27e7XqoV\nK+DMGVcWHu56qTp1As+yncWegisREZEySHNgxJ/U3sSXs2dh9WoXVP30kysLCoKoKDefqlkzKKaJ\nkrOk4EpERERERPzmyBGX8S8+Hvbtc2WVKsEll7jMfzksrVqsac6ViIiID/pbIiJSuFJSXNa/pUvh\n5ElXVq+e66W66CKoWLFwrhPIOVdaZKgYiYmJISgoiKCgIJYtW5brx82YMYPQ0NAirFnhOn36NC1b\ntmTRokU+9yclJREUFMSqVav8XLOc9enTh6FDh3q3jx49yg033ED16tUJDg5m+/bt3vW7goOD2bNn\nT6FdOygoiLlz56bbDgoKKvLXPhCvx5o1awgPD/eu+ZWdjz76iPPOO4/y5ctzyy23+KF2uZeb92bj\nxo2ZPHlyga+V0/uqNJs0aRKNGzfOcv/111+faYHr+Ph473vo6quvLuoqioiUWdbC99/DlCkQFweL\nF7vAqk0buPdet+hvjx6FF1gFmoKrYsQYwy233MKuXbuIiorK9eP69evHtm3birBmhWvGjBmcc845\n9OjRw1sWFBREcnIyABEREezatYsLLrig0K45bty4dEFRfhljMGkG/77++ussXryYJUuWsHPnTsLD\nwzHGMHbsWHbu3Ent2rULfM2s7Nq1iylTphTZ+QPpggsuoEOHDjz//PM5Hnvrrbdy4403kpycnOkD\ndEmwYsUK7rzzzgKfJ6f3VW7k530yZMgQ4uLicnVsaqCeF/Hx8dkGTr5kvI8xY8YwYcIEjh496i3r\n1q0bO3fupG/fvune02VJfHx8oKsgZYjaW9lz/Lhb8HfsWJg6FX78ESpUcIFUXJwLrNq0KXlzqnKi\nOVfFTOXKlalTp06eHhMSEkJISEgR1ajwvfDCC9xzzz1Z7g8KCsrxOThx4gQHDx7MdfBSVB+etmzZ\nQqtWrWjTpk268tDQ0Dy/jnlVp04dwsLCivQagTRo0CAeeughHnzwwSyP2b9/P/v27aN3797Ur18/\n39c6efIkFSpUyPfjC6JWIQ0sz+l9lRv5eZ9k/MKhOMhYn6ioKGrXrs27777LkCFDAChfvjx169Yl\nJCSEI0eOBKCWIiKl02+/uaF/X38Nx465slq1IDYWunWDypUDW7+ipp6rEmDHjh3069ePmjVrUrNm\nTfr06cOWLVu8+zMOPRo3bhxt27blnXfeoWnTpoSFhXHdddexd+9e7zFr166lV69eVKtWjdDQUNq3\nb+/9Vil1uMy+1BmGZB4alnrM559/TlRUFJUrV6Z79+7s2LGDBQsW0K5dO0JDQ7nmmmvYv3+/9zzr\n16/n++8kOT2GAAAgAElEQVS/55prrsnyfnMzDG3Xrl00bNiQP//5z8ydO5eTqQN3s5CfeRNHjx5l\nyJAhhIaGUq9ePSZOnJhuf0xMDM899xwJCQkEBQXRs2fPbM+3YcMGrrnmGqpXr05oaChdu3Zl3bp1\n3vqNHz+e8PBwQkJCaNeuHR9//HGe65xR165deeCBB9KVHTx4kEqVKvHhhx8CMGvWLDp37kxYWBh1\n69alb9++pKSkZHnO3LQPcK/1VVdd5T3vgAED2L17t3d/dm0Q4Morr+SXX37hm2++ybIeqYFJz549\nCQoKIiEhAYC5c+fStm1bQkJCiIiIYMKECekeGxkZSVxcHLfccgs1atTg5ptvzvJ+Z86c6T1XvXr1\nvB/OAZKTk7nuuusICwsjLCyM66+/nh07dmR5Ll8iIyN55plnvNtBQUH85z//4cYbb6Rq1ao0bdqU\nt956K9tz5OZ9dfbsWW699VaaNGlC5cqVad68OU8//XS690Z+5xelfdzJkycZPXo0kZGRhISE0LRp\n01z1QObFv/71L+rVq0doaCiDBw/m8OHDWdYn1bXXXsvbb79dqPUo6ZS5TfxJ7a10sxY2boSXXoJH\nHnHZ/44dg/POg7/9DR5/HC69tPQHVlCGe66GfTKs0M/5ytWvFPo5jx49SmxsLNHR0SQkJFChQgWe\nfvpp/vSnP/Hjjz9SqVIln49LSkrivffe46OPPuLw4cP069ePMWPG8PLLLwMwYMAAOnTowEsvvUS5\ncuVYu3Ztvnq/xo0bx/PPP09YWBgDBgygb9++VKxYkddee42goCBuvPFG4uLivMPXEhISCA8Pz9Tj\nlNdvvhs1asQ333zDm2++yV133cUdd9xBv379GDRoEF26dMl0fH6+XX/ggQeYP38+c+fOpUGDBsTF\nxZGQkMD1118PwAcffMADDzzAxo0bmTt3brY9HykpKURHR3PJJZcwf/58atasyfLlyznjWchhypQp\nTJo0iVdeeYVOnTrx5ptv8pe//IWVK1cWaHjkzTffzBNPPMHTTz/tvf/333+fypUrc9VVVwFw6tQp\nxo8fT8uWLfn1118ZOXIk/fv3L9DcnZ07d9K9e3duv/12Jk+ezKlTpxg9ejTXXnst3377LZBzG6xc\nuTJt2rRh0aJFXHzxxZmu0a1bN3744QfatGnD3Llz6dq1KzVq1GDlypX07duXRx99lIEDB7Js2TKG\nDRtGWFgYw4cP9z5+8uTJPProozzyyCNZBhWvvPIKI0aMYOLEifTp04fDhw+zcOFCwAUr1157LVWq\nVCE+Ph5rLcOHD+fPf/4zy5cvz/Vz5attPvbYYzz11FM89dRTTJs2jVtuuYXu3bsTHh7u8xy5eV+d\nPXuWhg0b8t5771G7dm2WLl3KHXfcQa1atbxz1fLbC5X2MYMHDyYxMZHnnnuODh068Msvv5CUlOTz\n2Pyc/9133+XRRx/lhRdeIDY2lnfffZcnn3ySc845J93xGa/TuXNnnn32Wc6ePZvnoYkiIuLbqVOw\nbJnrqfrlF1dWrhx07uzWp8riz1apVmaDq5LinXfeAdzcnlQvv/wydevW5dNPP+XGG2/0+bjTp0+n\n69G64447mD59und/cnIyDz74IM2bNwegSZMm+arf+PHj6datGwB/+9vfuOeee1i1ahXt27cH3Aet\n//73v97jN2/eTKNGjTKdJzXIyIuoqCiioqKYNGkS//vf/3jzzTeJjY0lIiKCQYMGMWjQIM4991wA\nxo4dm6dzHz58mNdff53p06dz6aWXAjB9+nQaNmzoPaZGjRpUqlSJ8uXL5zgEcOrUqYSGhvLee+9R\nrpx726V9zidNmsSDDz5Iv379ALyB3KRJk3jzzTfzVPe0+vbty4gRI1i4cKG3Z+2tt97ixhtvpLxn\nNb60c1MiIyN58cUXad26NSkpKTRo0CBf133ppZdo3759ut6+mTNnUqtWLVasWEGnTp1y1QYjIiLY\ntGmTz2uUL1/eG0zUrFnT+xpMnjyZmJgY72verFkzNm/ezFNPPZUuuIqJicnUq5fR+PHjue+++xgx\nYoS3LLVtf/XVV6xdu5atW7cSEREBwOzZs2nWrBkLFizIsSczO4MGDWLAgAHeOjz77LMsXrzYW5ZR\nbt5X5cqVSzc3KiIigpUrV/L22297g6u8vk+AdP+vbN68mTlz5vD555/Tu3dvwLWp6Oho77/z+l6P\niYlh69at3u0pU6YwZMgQbr/9dgBGjx7NwoUL+Sl1gZQs7iMiIoKjR4+yY8eOLIPUskbrDok/qb2V\nLr//DosWQUICpA4eCAtzadQvucT9u6wqs8FVUfQyFYWVK1eybdu2TBnHjh07lu4DR0aNGjVK95j6\n9euny1z3j3/8g9tuu42ZM2fSq1cvrr/+elq0aJHn+rVr187779QPt23btk1Xlva6Bw8epEqVKnm6\nRps2bbyT8rt3785nn32Wbn9wcDBXXnklV155Jb/99htDhw5lzJgxbN68OV1Qmhc//fQTJ0+eTNdj\nUqVKlXT3lherV68mOjraG1ildfDgQXbu3OkNUlNFR0fzf//3f/m6XqpatWpx+eWX89Zbb9GzZ09S\nUlKIj49n3Lhx3mNWrVpFXFwca9asYd++fd5enOTk5HwHVytXriQhISFTuzXG8NNPP9GpU6dctcHQ\n0FAOHDiQp2tv2LCBPn36pCvr1q0bcXFxHD58mKpVq2KMoVOnTtmeZ8+ePaSkpNCrVy+f+3/88Uca\nNGjgDazAZf5r0KAB69evL1BwlfZ9FRwcTO3atbPNPJnb99XLL7/MtGnTSE5O5tixY5w6dYrIyMh8\n1zOj1atXExQURGxsbKGdM6MNGzZwxx13pCu76KKL0g2V9iV1fuKBAwcUXImI5NO2ba6XauVKSP2u\nrFEjl0q9UyfXa1XW6Sko5s6ePUv79u2ZM2dOpn01atTI8nGpvRKpjDGcPXvWuz127FgGDhzIvHnz\n+N///kdcXBwvv/wyQ4cO9Q6ZSTtU6tSpUzleJ3UYTnBwcJbXrVatGhs2bMiy3r58/vnn3uv7GgZp\nrWXJkiXMmjWL9957j9DQUEaNGsWtt96ap+vkRn7npORnvRxrbaEkCrjpppu4/fbbefHFF3nnnXeI\niIjw9iQcOXKEyy67jN69ezNr1izq1KnDr7/+yiWXXJLlPLbctA9rLX369GHSpEmZHp8ahGfXBlMd\nPHgwX4lBslsDL1Veg/y8KOjrltP7N6PcvK/mzJnDfffdxzPPPEPXrl0JCwvjhRde4IMPPihQXUuK\ngwcPAlC9evUA16T4UC+C+JPaW8l15gysWuXmUaUmpw4Kgo4d3dC/Jk1KX8a/gtDA82KuY8eObNmy\nhVq1atGkSZN0P9kFV7nRrFkz7rnnHj799FNuvfVWpk2bBuAdapU2qcF3331XoGulvWZeUkMDhIeH\ne+85bUa4TZs28c9//pOmTZtyxRVXcPz4cd577z2SkpJ44okn8j3UEaBp06aUL18+XTKFI0eOeBNQ\n5FWHDh1ITEz0GaSGhYXRoEEDEhMT05UnJiZmykKYH6lr+Hz66ae89dZb6YaWbdiwgb179zJhwgSi\no6Np3rx5uqQTvuSmfURFRbFu3ToiIiIytduqVat6j8uqDabavn075513Xp7ut1WrVixZsiRdWWJi\nIuHh4XkKqOrUqcO5557L/Pnzs7xOSkoK27dv95Zt3bqVlJQUWrdunac6F1Ru3leJiYlceOGF3HXX\nXbRv354mTZqwZcuWQs301759e86ePcuCBQsK7ZwZtWrVKlOSk2+//TbH+9i+fTuVK1fOd2+siEhZ\nc/gwzJsHY8bAtGkusKpSBS67DJ54Au64A5o2VWCVkYKrYm7gwIHUrVuXa6+9loSEBLZt20ZCQgIP\nPPBAjsNgsnLs2DHuvvtuFi1aRFJSEkuXLk33Qb5Zs2aEh4czbtw4Nm/ezBdffMHjjz9eKPdzySWX\n8PPPP/Prr78W6DzJycm0bt2ar7/+mnHjxrF7925mzJhRoKFYaVWtWpVbb72VkSNHMn/+fH744Qdu\nueWWbHsPsnPXXXdx+PBh+vbty4oVK9iyZQtvv/02a9asAeDBBx9k0qRJvPPOO96gMTExMcc5QRl9\n8MEHtGzZMl3gExISwvXXX8/48eNZvXo1N910k3dfREQEFStW5Pnnn2fr1q189tlnPProo9leIzft\n4+677+bAgQP89a9/ZdmyZWzdupX58+czbNgwDh8+zPHjx7Ntg+CSuaxfv57u3bvn6Tm4//77WbRo\nEXFxcWzatIm33nqLyZMn89BDD+XpPODWR5oyZQpTpkxh06ZNfPfdd94Ffy+99FLatWvHwIEDWbly\nJStWrGDgwIF07NixSIfF+ZKb91WLFi1YtWoVn3/+OZs3b2b8+PHe7IqFpXnz5vTt25fbbruNuXPn\nsm3bNhYvXsysWbMK7Rp///vfmTlzJtOmTWPz5s1MnDgxV4uuL1u2jG7duimZRRpad0j8Se2t5Nix\nA958Ex5+GD78EPbvh/r1YeBAmDgR/vIXqFkz0LUsvvRXppirVKkSCQkJNGnShBtvvJFWrVoxZMgQ\nfv/9d2qmadlpv7XNKuNXalm5cuX4/fffGTJkCC1btuQvf/kLXbt29X5oLF++PO+88w5bt27lggsu\nIC4ujokTJ2Y6Z3bXyKoubdq0oW3btnz00UfZ3ndO30LXrl2bpKQk5s+fz6BBg6icx9yeM2bMyHGB\n1UmTJhEbG8t1111Hr169aNeuXaYP+rnNrtagQQMSEhI4efIksbGxREVFMXXqVO/wr3vvvZcHH3yQ\nhx56yPv8pKYTz4sDBw6wefNmTp8+na78pptu4vvvvycqKoqWLVt6y2vXrs3MmTP58MMPadOmDePH\nj+ff//53tq91btpH/fr1WbJkCUFBQVx++eWcf/75DB8+nJCQECpWrEhwcHC2bRDgs88+Izw83Gem\nwKzqBq6X8L333uP999+nbdu2jB49mlGjRnH33Xfn/on0+Nvf/sbUqVP5z3/+Q9u2bbniiitYv369\nd/9HH31E7dq1iY2NpWfPnjRo0MCb4j6r+hWF3Lyvhg0bRt++fRkwYABdunQhOTmZ+++/P9vz5uZ9\nktEbb7zBgAEDuPfee2nVqhVDhw71DsnzJSgoiMceeyzX5+/bty/jxo1jzJgxREVF8cMPP/CPf/wj\nx8d98skn9O/fP1N5cVujS0QkEM6ehTVr4N//hsceg8RElwmwbVsYMcItBNy9O1SsGOiaFn8mv3NI\nAs0YY7ObV1ES7ysmJoa2bdsW+powxc20adOYPn16pqFb/jR27Fjmzp3LmjVrCv2b7MaNGzN8+PAc\nP7gWhhkzZnDPPfdw6NChIr+Wv1199dV0794920WE5Q9F8b4qyvcJwLZt22jWrBmJiYk5BtEFsXLl\nSq644gqSkpIyfREzZMgQ9u7dyyeffJLpcSX1b4mISG4dPw5LlsDChZA6+KFiReja1S36W7duYOuX\nX57/vwPy7Zl6rooRYwyvvvoqoaGhrFy5MtDVKTJDhw5l7969BVpHqaDmzZvH1KlTi2yI0JgxYwgN\nDeW3334rkvODG7p45513lspv3r///nu+++477rnnnkBXpcQoivdVUb9P5s2bx+DBg4s0sAKYMGEC\njzzySLrAavHixVStWpXZs2eXyveQiEh29uyBOXNg5Eh4910XWJ1zDtx4Izz1FPTrV3IDq0BTz1Ux\nkpKSwvHjxwFo2LBhtovSSvGVnJzsHZYXGRlZZB9MU1PxBwUFFWo6bZGy4Pjx4965iVWqVKGuj08R\nJfVvSW5p3SHxJ7W3wLMWNmxwqdTXrnXbAC1auFTq7dq5LIClQSB7rpSKvRhRFqvSIe26R0WpINkQ\nRcq6kJAQvYdEpEw4eRKWLnVBVWq+q/LloUsXF1Q1bBjY+pU26rkSERHxQX9LRKQk278f4uNh8WI4\ncsSVVasGMTFwySUQGhrI2hUt9VyJiIiIiEiBWOvWo/rqK7fwb+oKMo0bu16qqCgop0//RUpPr4iI\nSBmkOTDiT2pvRev0aRdMffUVJCW5sqAg6NQJevUCjYL2n1IbXCn7k4iIiIiUZocOQUICLFoEBw64\nsipV3JpUPXpAjRqBrV9ZVCrnXImIiIiIlFa//OJ6qZYvd4v9AjRo4HqpunSBsp5wukzNuTLGvA5c\nBeyx1rbN4pjngCuAo8AQa+1qP1ZRRERERKRYOXsW1qxxWf82bXJlxsAFF7j5VC1auG0JrEBks58O\nXJ7VTmPMlUAza+15wB3AS/6qmIgv8fHxga6ClCFqb+IvamviT2pv+Xf0KHz5JTz6KLz8sgusQkJc\nL9Vjj8Fdd0HLlgqsigu/91xZaxcbYyKzOeQaYKbn2KXGmOrGmLrW2t3+qJ+IiIiISKDt3u16qb75\nBk6ccGW1a7teqq5dXYAlxU9A5lx5gqtPfA0LNMZ8Aky01n7t2Z4PjLTWrsxwnOZciYiIiEipYS38\n+KObT7Vu3R/lrVq5oOr8810WQMlemZpzlUsZnwyfUdSQIUOIjIwEoHr16rRv396b5jO1+1nb2ta2\ntrWtbW1rW9vaLs7bF18cw9Kl8Prr8ezdCw0axFC+PISFxdOhA9x4Y/Gqb3HbTv13Umoe+gAqjj1X\nLwPx1tp3PNsbgB4ZhwWq50r8JT4+3vsmFilqam/iL2pr4k9qb77t2wcLF0JioptbBS59ekwMREdD\n1aoBrV6JpZ6r9D4GhgPvGGMuAn7XfCsRERERKQ2shZ9+cvOpVq92WQDBLfTbsydERUFwcGDrKPnn\n954rY8zbQA/gHGA3MBYoD2CtfcVzzAu4jIJHgKHW2lU+zqOeKxEREREpEU6fhhUr3Hyq5GRXFhwM\nHTu6zH+emS5SCALZc6VFhEVEREREisjBg5CQAIsWuX+DG+7Xowd07w7Vqwe2fqVRIIOroEBcVKQk\nSTtZUqSoqb2Jv6itiT+VxfaWnAwzZsCoUfDJJy6watgQBg2CJ5+Ea65RYFUaFcc5VyIiIiIiJc7Z\ns/Ddd27o35YtrswYaN/eDf077zwt9lvaaVigiIiIiEgBHD3qMv4tXOgyAAJUqgTdukFsLJxzTmDr\nV9YoW6CIiIiISAmzc6cLqL75Bk6edGV167qA6uKLISQksPUT/1NwJZIDrc0h/qT2Jv6itib+VJra\nm7Xwww8ulfoPP/xR3rq1S6V+/vka+leWKbgSEREREcnBiROuh2rBAtjtWYG1QgW46CIXVNWvH9j6\nSfGgOVciIiIiIlnYu9cN/UtMhGPHXFnNmhATA9HRUKVKQKsnPmjOlYiIiIhIMWEtbN7seqnWrHFZ\nAAGaNXO9VB06QJAWNBIfFFyJ5KA0jROX4k/tTfxFbU38qaS0t1OnYMUKl0r9559dWXDwH0P/GjUK\nbP2k+FNwJSIiIiJl2oEDsGgRJCTAoUOuLCwMund3P9WqBbZ+UnJozpWIiIiIlElJSW7o34oVcOaM\nK4uIcL1UnTtDOXVDlEiacyUiIiIi4gdnzsB337mhfz/95MqCgiAqygVVzZoplbrkn6biieQgPj4+\n0FWQMkTtTfxFbU38qTi0tyNH4PPPYcwYePVVF1hVrgy9e8Pjj8OwYXDeeQqspGDUcyUiIiIipVZK\nihv6t3QpnDzpyurVc71UF10EFSsGtn5SumjOlYiIiIiUKtbC2rUuqPrxxz/Kzz/fBVWtW6uHqjTT\nnCsRERERkQI6fhy+/tot+rtnjyurUAG6doXYWNdjJVKUNOdKJAfFYZy4lB1qb+IvamviT0Xd3n77\nDd59Fx5+GObMcYFVrVpwww3w1FPQv78CK/EP9VyJiIiISIljLWzc6Ib+ff+92wZo3twN/bvgApcF\nUMSfNOdKREREREqMU6dg2TKXSn3HDldWrpxbl6pXLwgPD2z9JPA050pEREREJBu//w7x8bB4MRw+\n7MqqVYMePeCSSyAsLKDVEwE050okR5qXIP6k9ib+orYm/lSQ9rZtG7z2GoweDfPmucCqUSO45RaY\nMAGuukqBlRQf6rkSERERkWLlzBlYtcoN/du2zZUFBUHHjm7oX5MmSqUuxZPmXImIiIhIsXD4MCQk\nwKJFbhggQJUqbthfjx5Qs2Zg6yclg+ZciYiIiEiZtWOHy/q3dKlLWAHQoIFbm+rCC6FixcDWTyS3\nFFyJ5CA+Pp6YmJhAV0PKCLU38Re1NfEnX+3t7FlYu9YN/du48Y/ytm3d0L+WLTX0T0oeBVciIiIi\n4jfHj8OSJbBwIfz6qyurWBG6dnU9VXXrBrZ+IgWhOVciIiIiUuT27HEB1ddfuwAL4JxzXEDVrRtU\nqhTY+knpoTlXIiIiIlLqWAsbNrihf+vWuW2AFi3c0L+2bV0WQJHSQsGVSA40L0H8Se1N/EVtTYrS\nyZMuOcWCBZCSAikp8TRqFEOXLtCzJzRsGOgaihQNBVciIiIiUij274f4eFi8GI4ccWXVqkHjxnD3\n3RAaGtDqiRQ5zbkSERERkXyzFrZudb1Uq1a5LIDgAqqePSEqCsrp63zxI825EhEREZES5fRpWLnS\nBVVJSa4sOBg6d3ZBVZMmAa2eSEBoCqFIDuLj4wNdBSlD1N7EX9TWJL8OHYLPPoPRo+H1111gVbUq\nXHEFPPEE3HZb5sBK7U3KCvVciYiIiEiOfv7Z9VItXw6nTrmyc891vVQXXgjlywe2fiLFgeZciYiI\niIhPZ8/CmjUuqNq0yZUZA+3auaCqRQu3LVKcaM6ViIiIiBQbR4/CkiUu899vv7mykBC32G9MDNSp\nE8jaiRRfmnMlkgONExd/UnsTf1FbE19274a334aHH4b//tcFVrVrw1//Ck89BX375i+wUnuTskI9\nVyIiIiJlmLWwfr0b+rdu3R/lrVq5oX/nnw9B+jpeJFc050pERESkDDpxAr791gVVu3a5svLl4aKL\nXFDVoEFg6yeSX5pzJSIiIiJ+sXevm0uVmOjmVgHUqOHmUkVHu7TqIpI/6uQVyYHGiYs/qb2Jv6it\nlS3WwubN8Mor8Mgj8MUXLrBq2hRuv92tT3X55UUXWKm9SVmhnisRERGRUur0aVixAr76CpKTXVlw\nMHTpAr16QWRkQKsnUupozpWIiIhIKXPwICxaBAkJ7t/geqV69IDu3aF69cDWT6Qoac6ViIiIiBTY\n9u0uQcXy5XDmjCsLD3cJKjp3dgkrRKToKLgSyUF8fDwxMTGBroaUEWpv4i9qa6XH2bPw3Xdu6N+W\nLa7MGGjf3g39O+88tx1Iam9SVii4EhERESmBjhxxGf/i42HfPldWqZLL+BcTA+ecE8jaiZRNmnMl\nIiIiUoLs3AkLF8I338DJk66sbl2IjYWLL4aQkMDWTyTQNOdKRERERLJkLfzwgxv6t379H+WtW7uh\nf23aBH7on4gouBLJkcaJiz+pvYm/qK2VDCdOuB6qBQtg925XVqECXHSRS1JRv35g65dbam9SVii4\nEhERESlmfvvNzaVKTIRjx1xZzZpuLlV0NFSpEsjaiUhW/D7nyhhzOTAFCAamWWufyrD/HGAWUA8X\n/E2y1s7wcR7NuRIREZFSw1rYvNkN/fv+e5cFEKBZM9dL1aEDBAUFto4iJUEg51z5NbgyxgQDG4E/\nATuA5UB/a+2PaY4ZB1S01o7yBFobgbrW2tMZzqXgSkREREq8U6fculQLFsDPP7uy4GC3LlXPntCo\nUWDrJ1LSBDK48vf3H12ALdbaJGvtKeAd4NoMx+wEwjz/DgP2ZgysRPwpPj4+0FWQMkTtTfxFbS3w\nfv8dPv4YRo2CmTNdYBUWBn36wJNPwtChpSewUnuTsiJXc66MMd2AGtbaTz3btYCpQBvgC+Aha+2Z\nXJzqXODnNNu/ABdmOOY/wAJjTAoQCvTNTR1FRERESoKkJNdLtWIFnPF8eoqIcL1UnTtDOc2IFymx\ncjUs0BizGJhvrY3zbL8OXA98BVwGPGWtfSwX57keuNxae7tn+ybgQmvtPWmOeQQ4x1o7whjTFPgS\nuMBaeyjDuTQsUEREREqEM2dg9WoXVP30kysLCoL27V1Q1ayZUqmLFJaSsM5VS+ApAGNMBeAG4D5r\n7WvGmBHAMCDH4Ao3zyo8zXY4rvcqra7AEwDW2p+MMduAFsCKjCcbMmQIkZGRAFSvXp327dt703ym\ndj9rW9va1ra2ta1tbQdqe968eNauhX37Yti/H1JS4qlYEfr3jyEmBtaujWfHDjjvvOJRX21ruyRu\np/47KSmJQMttz9UxoLe1drExJhpIAOpZa/cYY3oA86y1lXNxnnK4BBW9gBRgGZkTWkwGDlhr44wx\ndYGVQDtr7b4M51LPlfhFfHy8900sUtTU3sRf1NaKVkqK66VauhROnnRl9eq5XqqLLoKKFQNbP39T\nexN/Kgk9VylAe2AxcDmwzlq7x7OvBnA0Nyex1p42xgwH/odLxf6atfZHY8wwz/5XgAnAdGPMGlzC\njYcyBlYiIiIixY21sHatC6p+/PGP8vPPd0FV69Ya+idS2uW252o8MAIXFF0FjLXW/suzLw7Xq3Vx\nUVbUR53UcyUiIiIBd/w4fP01LFwIezxfPVeoAF27Qmys67ESEf8pCT1XccBx4GJgIjA5zb72wHuF\nXC8RERGRYu3XX11AtWSJC7AAatVyAVW3blA5xwkTIlLa+HUR4cKknivxF40TF39SexN/UVvLH2th\n40Y39O/77902QPPmbujfBRe4LICSntqb+FNJ6LkSERERKbNOnXLJKRYsgB07XFm5ctCliwuqwsOz\nf7yIlA1Z9lx5UqBbIDXqy6qbyADWWtuk8KuXNfVciYiISFH7/XeIj4fFi+HwYVdWrRr06AGXXAJh\nYQGtnoj4UFx7rhZl2O4J1AWWAHs8/+4G7MItJiwiIiJSKmzd6nqpVq1yCwADREa6XqqOHV2vlYhI\nRln+12CtHZL6b2PMHUAXoKu19pc05eG4DIJfF2EdRQJK48TFn9TexF/U1jI7c8YFU199Bdu2ubKg\nILUICZkAACAASURBVBdM9eoFTZoolXp+qb1JWZHb710eAkanDawArLU/G2PG4dam+k8h101ERESk\nyB065Ib9LVrkhgECVKnihv3FxECNGgGtnoiUILld5+oY8Fdr7cc+9l0LzLHWhhRB/bKrk+ZciYiI\nSL798osb+rdsmUtYAdCggRv6d+GFbq0qESl5AjnnKrfB1SrgCG6x4GNpyisDXwCVrbVRRVZL33VS\ncCUiIiJ5cvYsrF3rhv5t3PhHedu2buhfy5Ya+idS0hXXhBZpPQj8H7DdGPN/wG6gHnAlEOb5LVIq\naZy4+JPam/hLWWtrx465xX7j493ivwAhIdC1q1v0t06dgFav1Ctr7U3KrlwFV9bar4wx7YFHgO64\nwGonLpnF49baDUVXRREREZH82bMHFi6Er7+G48dd2TnnuICqWzeoVCmw9ROR0iVXwwKLIw0LFBER\nEV+shQ0b3NC/devcNkCLFm7oX9u2LgugiJROJWFYoIiIiEixdvIkLF3qklSkpLiy8uWhSxeXpKJh\nw8DWT0RKv1wHV8aYGKA/EA6kzQxoAGut7Vm4VRMpHjROXPxJ7U38pTS1tf373VyqxYvhyBFXVr06\n9Ojh0qmHhga0ekLpam8i2clVcGWMGQa8BOwDNgEni7JSIiIiItmxFrZudUP/Vq92WQABGjd2Q/86\ndIByGp8jIn6W21Tsm4DlwFBrbbEIrDTnSkREpOw5fRpWrnRD/5KSXFlwMERFuaF/TZoEtHoiUgyU\nhDlX5wJ3FpfASkRERMqWQ4cgIQEWLYIDB1xZ1apu2F9MjBsGKCISaLnNlbMK0HdBUibFx8cHugpS\nhqi9ib+UlLb2888wcyY8/DB8/LELrM49F26+GZ58Ev78ZwVWJUFJaW8iBZXbnqt7gNnGmE3W2kVF\nWSEREREp286ehTVr3NC/TZtcmTFwwQVu6F+LFm5bRKS4ye2cq5+BMCAUOALsx5MlkD+yBUYUYT19\n1UlzrkREREqRo0dhyRK36O/eva4sJMQt9hsbC7VrB7Z+IlIylIQ5V1/lsF9RjoiIiOTL7t2ul+qb\nb+DECVdWu7brpera1QVYIiIlQa56rooj9VyJv2htDvEntTfxl0C3NWth/XoXVK1b90d5q1YuqGrb\nVkP/SpNAtzcpW0pCz5WIiIhIgZ04Ad9+64KqXbtcWfnycNFFLqhq0CCw9RMRKYhc91wZY9oBY4Ee\nQA3cgsLxwGPW2rVFVcFs6qOeKxERkRJi716Ij4fERDe3CqBGDZdG/ZJLoEqVQNZOREqTQPZc5Tah\nRWdgEXAM+BjYDdQDrgZCgB7W2hVFWE9fdVJwJSIiUoxZC1u2uF6q775zWQABmjZ1vVQdOrgFgEVE\nClNJCK7m47IF9rLWHkpTHgrMBw5aay8tslr6rpOCK/ELjRMXf1J7E38pyrZ2+jQsX+6CquRkVxYc\nDJ06uaAqMrJILivFmP5vE38qCXOuLgIGpQ2sAKy1/9/encdZWd4H//9cMyyyySIIoiguuKIgoiyy\nDJgmJl1s0sbEZjNJmzS/ps/TX/t6qqZpkzRpYvr8+iRdniYmsUljk5q1JmbRWOCACiKERWQRQVEW\nRRAURNaZ6/fHdcY5jjPMGTjnPufM+bxfr/Pi3Nd9z32+g1+B71zX9b33hxC+CHy75JFJkqSasm8f\nLFwIixal9wCDBsGsWenlw34l9XTFzlztBz4QY/xxB+feAfx7jHFQGeI7XkzOXEmSVAWeeSbNUi1b\nBs3NaWzMmDRLdfXVqWGFJGWlVpYFDiYtC9xXMD6Q9AwslwVKklRHWlrSPqp589K+Kkit0ydMgOuu\ng3HjbKUuqTJqobi6hraGFj8DngPOAN4G9AeaYoyPljHOjmKyuFImXCeuLJlvysqJ5tqBA6njXy4H\ne/aksX79YMaM1Plv+PBSRqmewj/blKWq33MVY3w0hDAF+Bvgetpasc8HPluJVuySJCk7zz2Xlv49\n8ggcOZLGRo5MS/+mTYO+fSsbnyRVg6Kfc1VtnLmSJKm8YoS1a9PSv3Xr2sYvvTQt/bvsMpf+Sao+\nVT9zFUI4HRgaY3yig3MXAXtijLtKHZwkScre4cOweDEsWAA7d6axPn3SDNWcOXDGGZWNT5KqVUOR\n1/0r8OednPsz4P+WJhyp+uRyuUqHoDpivikrHeXa7t3wgx/ALbfA3XenwmrYMHjHO+D22+EP/sDC\nSifGP9tUL4p9ztW1wMc7OfcrLK4kSapJMcKTT6alf489lroAAlxwQVr6N3EiNBT7o1hJqnPFdgs8\nBPxmjHFeB+feBPw8xpjpVlb3XEmSdOKOHk3PpZo/H7ZuTWONjem5VHPnwjnnVDY+STpRVb/nCtgO\nTCU906q9a0it2SVJUpV76SVYtCi99u9PY6eeCrNmwezZ6b0k6cQUO9H/A+C2EMJvFQ7mj28Dvl/q\nwKRq4TpxZcl8U7ls2QJ33gmf+AT8/OfwxBM5zj4bPvhB+MIX4Ld/28JK5eOfbaoXxc5cfRaYBfw0\nhPAcaSbrLGAUsAT4THnCkyRJJ6q5GVauTEv/Nm9OYw0NMGkSzJwJ73mPrdQlqZSKfs5VCKEP8F7g\nzcBpwG7gfuA/YozHyhZh5/G450qSpA4cOAAPPgi5HOzdm8b694cZM6CpCU47rZLRSVJ5VXLPlQ8R\nliSph9ixI3X9W7o0NawAGDUqNaiYOhX6Ztp6SpIqoxYaWgAQQpgAzCTNXN0RY3w+hDAO2Blj3FeO\nAKVKy+VyNDU1VToM1QnzTd0VI6xZk5b+rV/fNj5+fCqqLr2046V/5pqyZL6pXhRVXIUQ+gLfAd6R\nH4rAvcDzwBeBjcCt5QhQkiS90aFDsHhxKqp27UpjffvCtGmpqBo5srLxSVI9KvY5V/8f8GHgT4AH\ngJ3A5BjjihDCHwF/EmOcWNZI3xiTywIlSXVn1y5YsAAefjgVWADDh6e9VNdem/ZWSVI9q4VlgTcB\nfx1j/G4Iof3XbAHGljIoSZLUJkZ44ok0S/XYY+kY4MIL0yzVhAmpC6AkqbKKLa5OA9Z1cq4BcIus\neizXiStL5psKHT2amlPMnw/bt6exXr3gmmtSUTVmzInf21xTlsw31Ytii6stwHRgfgfnrgaeKFVA\nkiTVu717YeHC1E79lVfS2ODBMHs2zJoFgwZVNj5JUseK3XN1G/BXwEeBHwMHgMnAEOCHwKdjjP9U\nxjg7isk9V5KkHuWpp9Is1YoV6QHAAGPHplmqq65Ks1aSpOOr+udc5fdZ/QdwI3AE6AMcAk4B/hN4\nb9aVjsWVJKknOHYsFVPz58PTT6exhgaYNCkVVeed13ErdUlSx6q+uHrt4hBmAtcDpwMvAvfFGHPl\nCa3LWCyulAnXiStL5lv92L8/LfvL5eDll9PYgAEwc2bq/Dd0aHk/31xTlsw3ZakWugUCEGN8EHiw\nTLFIktTjbduWZqkefTQ1rAAYPTrNUk2ZAn36VDY+SdKJK3ZZ4EXAkBjj0vxxP+BTwGXAr2KM/1z0\nB4ZwPfBloBH4Rozxix1c0wR8CegN7I4xNnVwjTNXkqSa0NKSWqjPn59aqre64opUVF18sUv/JKlU\nqn5ZYAjhAWBljPEv88f/B/g48DhwBfBnMcZ/KeI+jaTOgm8CtgPLgJtijOsLrhkCPAy8Jca4LYQw\nPMa4u4N7WVxJkqrawYPpYb8LFsDu/N9kp5wC06fDnDlw+umVjU+SeqJKFlfFPnLwCmAxvFYgvR+4\nNcY4Cfgs8EdF3ucaYFOMcUuM8ShwN3BDu2v+APhRjHEbQEeFlZSlXC5X6RBUR8y3nuGFF+Duu+HW\nW+EHP0iF1YgRcOONcPvt8K53Vb6wMteUJfNN9aLYPVeDgdYi50pgGPCD/PFC4H8VeZ8zga0Fx9uA\nKe2uGQf0DiEsAAYB/xhjvKvI+0uSVBExwoYNMG8erFnTNn7RRXDddXD55akLoCSp5yq2uNpJKnoe\nAn4D2BxjbC2SBgLHirxPMev4egOTgOuA/sCSEMIjMcYn21948803M3bsWACGDBnCxIkTX+tE0/oT\nEo89PtnjpqamqorH4559bL7V3vEDD+RYtw5eeqmJHTtgx44cvXrBO97RxJw5sGlTjr17oaGhOuL1\n2GOPPe5px63vt2zZQqUVu+fqn4F3kp519UHgjhjjJ/LnbgVuzC8R7Oo+U0kPHL4+f3wb0FLY1CKE\ncAvQL8b46fzxN0gt33/Y7l7uuZIkVczevZDLpXbqBw6ksSFDoKkptVMfOLCS0UlS/aqFPVe3AfcC\nbwF+AvxdwbkbgF8VeZ/lwLgQwtgQQh/gXcBP213zE2BGCKExhNCftGxwXZH3l0qu8KciUrmZb9Ut\nRti8Gb72NfjEJ+C++1Jhde658Id/CJ//PLz1rbVRWJlrypL5pnpR1LLAGOMrdNK0IsY4rdgPizEe\nCyF8HLif1Ir9zhjj+hDCR/Pn74gxbggh3Ac8BrQAX48xWlxJkirm2DFYvjy1Un/mmTTW2AhXX51a\nqZ93XmXjkyRVh6KWBVYjlwVKkspt/35YtAgWLoSXX05jAwemZX9NTWkZoCSpulRyWWCxDS0kSaob\nW7emrn/LlqVZK4Azz0xd/665Bnr3rmx8kqTqZHEldSGXy73WlUYqN/OtclpaYPXqtPRv48Y0FgJM\nmJCKqgsvTMc9hbmmLJlvqhcWV5Kkuvbqq/Dww7BgAbz4Yho75RS49lqYMyc9/FeSpGK450qSVJee\nfz7NUi1ZAkeOpLHTT08NKqZNSwWWJKn2uOdKkqQMxAjr1qWi6vHH28YvuSQVVZdf3rOW/kmSslV0\ncRVCaAJuAsYAhT/PC0CMMc4tbWhSdXCduLJkvpXH4cPwyCOpqHr++TTWuzdMnZqKqtGjKxtfJZhr\nypL5pnpRVHGVfw7VV4A9wEbgSDmDkiSpFF58EXI5eOihtLcKYOjQ1EZ95kwYMKCS0UmSepqi9lyF\nEDYCy4APxhirorByz5UkqSMxwqZNaZZq1arUBRDg/PNT17+JE9MDgCVJPVMt7Lk6E/hYtRRWkiS1\nd+xYei7V/Pnw7LNprLERpkxJS//Gjq1oeJKkOtBQ5HUrgPPKGYhUrXK5XKVDUB0x37pv3z649164\n7Tb41rdSYTVoEPzmb8IXvgAf+pCFVUfMNWXJfFO9KHbm6k+B74YQNsYYF5YzIEmSivHMMzBvHixf\nDs3NaWzMmDRLdfXVqWGFJElZKnbP1VbgVGAQcADYS75LIG3dAs8uY5wdxeSeK0mqMy0tsHJlWvq3\naVMaa2iACRNSUTVunK3UJane1cKeq3ldnLfKkSSVzYEDqeNfLgd79qSxfv1gxozU+W/48EpGJ0lS\nUtTMVTVy5kpZ8dkcypL59nrPPZdmqR55BI7kWyqNHJlmqaZNg759KxtfLTPXlCXzTVmqhZkrSZIy\nESM8/ngqqtataxu/7LJUVF12mUv/JEnVqdOZqxDC+4GfxxhfDCF8gC6W/sUYv12G+DrlzJUk9SyH\nDsGSJbBgAezcmcb69EkzVHPmwBlnVDY+SVJtqOTM1fGKqxZgaozx0fz744oxFtvWvSQsriSpZ9i9\nOxVUDz8MBw+msWHDUkE1Ywb071/Z+CRJtaValwWeB+woeC/VJdeJK0v1km8xwpNPplbqq1enY4AL\nLoDrroOJE1MXQJVPveSaqoP5pnrRaXEVY9zS0XtJkk7U0aPw6KNpP9W2bWmsV6/0XKq5c+HsTB/q\nIUlSadktUJJUdi+9BAsXwqJF8MoraezUU2HWLJg9O72XJKkUqnVZoCRJJ2XLlrT079e/hubmNHb2\n2Wnp3+TJadZKkqSewr/WpC64TlxZ6gn51twMK1akpX9PPZXGGhpg0qRUVJ1/vq3Uq0FPyDXVDvNN\n9cLiSpJUEgcOwIMPQi4He/emsf79U8e/piY47bRKRidJUvm550qSdFJ27EhL/5YuTQ0rAEaNSrNU\nU6ZA376VjU+SVF9qZs9VCGEEMBUYBvws/4DhfsCRGGNzOQKUJFWfGGHNmlRUbdjQNj5+fOr6d+ml\nLv2TJNWfooqrEEIA/jfwp0BvIAJXAy8C9wAPA39bphilinKduLJU7fl26BAsXpz2U+3alcb69oVp\n01JRNXJkZeNT8ao919SzmG+qF8XOXN0G/AnwGeABYGnBuXuB92FxJUk91gsvwIIFqbA6dCiNDR+e\n9lJde23aWyVJUr0ras9VCOEp4Bsxxs+HEHoBR4DJMcYVIYS3Av8RY8x0q7J7riSpvGKEJ55Is1SP\nPZaOAS68MM1STZiQugBKklRNamHP1ZnAkk7OHQEGlCYcSVKlHT2amlPMnw/bt6exXr3gmmtSUTVm\nTGXjkySpWhX7M8cdwOWdnLsCeLo04UjVJ5fLVToE1ZFK5tvevXDPPXDLLXDXXamwGjwYfud34Pbb\n4QMfsLDqSfyzTVky31Qvip25+j7wNyGEFRTMYIUQLgL+Avh6GWKTJJVZjPD006nr34oV0NKSxseO\nTa3UJ01Ks1aSJKlrxe656g/cD1wLPAOcQ5qtGgMsBt4SYzxcxjg7isk9V5J0go4dS8XUvHmwZUsa\na2hIxdTcuXDeebZSlyTVpkruuSr6IcL5RhY3AdcDpwO7gfuA78QYj5Utws7jsbiSpG7avx8efBBy\nOXj55TQ2YADMnJk6/w0dWsnoJEk6eTVRXFUbiytlxWdzKEvlyrdt29Is1bJlqWEFwOjRaZZqyhTo\n06fkH6kq559typL5pizVQrfA1wkhvKERRoyx5eTDkSSVSktLaqE+f35qqQ5pqd8VV6Si6uKLXfon\nSVIpdWfP1aeAdwJn8caiLMYYG0sf3nFjcuZKkjpw8CA8/HB66O/u3WnslFNg+nSYMwdOP72y8UmS\nVE61MHP1f4H3APcCd5OebVXIKkeSKmznzlRQLV4Mh/MthkaMSAXV9OnQr19l45MkqacrdubqReBv\nY4z/WP6QiuPMlbLiOnFlqbv5FiOsX5+W/q1Z0zZ+8cVp6d/ll6cugFJ7/tmmLJlvylItzFwdAdaV\nMxBJUvGOHIFHHkkzVTt2pLHevVNzijlz4KyzKhufJEn1qNiZq78HTosxfrj8IRXHmStJ9WjPntRG\n/aGH4MCBNDZkSGqjPnMmDBxYyegkSaq8qm/FHkLoDdwJjCI9THhv+2tijP9W8uiOH5PFlaS6ECNs\n3pyW/q1cmboAApx7Llx3XXrwb2OmLYUkSapetVBcTQF+Qnp4cIdijJmu6re4UlZcJ64sFebbsWOw\nfHkqqp55Jp1vbISrrkr7qc49t3Jxqvb5Z5uyZL4pS7Ww5+pfgReBPwKe4I3dAiVJJbJvHyxaBAsX\npveQlvvNmgWzZ6dlgJIkqfoUO3N1EPj9GOPPyx9ScZy5ktTTbN0K8+bBsmVp1grgzDPT0r9rrkkN\nKyRJ0vHVwszVRmBAOQORpHrU0gKrVqWlf08+mcZCgIkT09K/Cy9Mx5IkqfoVu0/qVuCTIYSx5QtF\nqk65XK7SIagHevVVeOAB+OQn4Y47UmF1yikwalSOz34WPvYxuOgiCyuVj3+2KUvmm+pFsTNXnwBG\nAE+EEDby+m6BAYgxxlmlDk6Seprnn0+zVEuWpGdVAZx+epqlmjYtPbtqxIjKxihJkk5MsXuuckAk\nFVIdiTHGOSWMq0vuuZJUK2KEtWtTUbV2bdv4JZek/VTjxztDJUlSqVR9K/ZqZHElqdodPpxmoubP\nTzNWAH36wJQpaaZq9OjKxidJUk9UyeIq02dTSbXIdeLqrhdfhB/+EG65Bb773VRYDR0K73gH3H47\nvPe9nRdW5puyYq4pS+ab6kWne65CCLOAlTHG/fn3xxVjXFTMB4YQrge+DDQC34gxfrGT664GlgA3\nxhh/XMy9JalSYoRNm9Is1apVqQsgwPnnp6V/EyemBwBLkqSeq9NlgSGEFmBqjPHR/PvjiTHGLv/Z\nEEJoJD2E+E3AdmAZcFOMcX0H1z0AvAp8M8b4ow7u5bJASRV39CgsX56eT7V1axprbITJk1NRdc45\nlY1PkqR6U63PuZoLrC94XwrXAJtijFsAQgh3AzcUfE6rPwV+CFxdos+VpJLatw8WLoRFi9J7gEGD\nYNYsmD0bBg+ubHySJCl7nRZXMcZcR+9P0pnA1oLjbcCUwgtCCGeSCq65pOLK6SlVVC6Xo6mpqdJh\nqEo880yapVq+HJqb09iYMWmWavJk6N375O5vvikr5pqyZL6pXhT1nKsQwlPA22OMqzs4dznwkxjj\neUXcqphC6cvArTHGGEIIdN7+XZIy0dICK1emomrz5jTW0ABXXpmKqgsusJW6JEkq/iHCY4G+nZw7\nJX++GNuBMQXHY0izV4WuAu5OdRXDgbeGEI7GGH/a/mY333wzY8emjx4yZAgTJ0587acirV1pPPb4\nZI+bmpqqKh6Pszu++uomHnoI/v3fc+zfD6NHN9GvHwwdmmPiRLjhhtJ/vvnmsccee+yxx907bn2/\nZcsWKq3Yhwi/1tyig3N/DHw+xjisiPv0IjW0uA7YATxKBw0tCq7/JnBvR90CbWghqVyeey7NUi1d\nCkeOpLGRI9Ms1dSp0LezHzVJkqSKq8qGFiGE/xf484Khe0MIR9pd1g8YBtxdzIfFGI+FED4O3E9q\nxX5njHF9COGj+fN3dCd4KQu5XO61n5Co54oRHn88tVJft65t/LLL0gN/L7ssm6V/5puyYq4pS+ab\n6sXxlgU+DczLv38/qW367nbXHAbWAt8o9gNjjL8EftlurMOiKsb4wWLvK0kn4tAhWLIEFiyAnTvT\nWJ8+MG0azJkDZ5xR2fgkSVLtKHZZ4LeAv40xPlX2iIrkskBJJ2P37lRQPfwwHDyYxoYNSwXVjBnQ\nv39l45MkSSemkssCiyquqpHFlaTuihE2bkxL/1avTscA48alpX8TJ6YugJIkqXZV5Z4rSYnrxGvf\n0aPw6KOpqNqW70/aqxdcfXUqqs4+u7LxFTLflBVzTVky31QvLK4k9VgvvQQLF8KiRfDKK2ns1FNh\n9myYNSu9lyRJKhWXBUrqcbZsSa3Uf/1raG5OY2efnVqpT56cZq0kSVLP5LJASTpJzc2wYkVa+vdU\nvvVOQwNcdVVa+nf++dm0UpckSfXLrdtSFwqf/q3q88or8Mtfwl/9FXzjG6mw6t8f3vxm+Nzn4CMf\ngQsuqJ3CynxTVsw1Zcl8U71w5kpSTdq+Pc1SLV2aGlZAeibV3LkwZQr07VvZ+CRJUv1xz5WkmtHS\nAmvWpKJqw4a28csvT0XVJZfUzgyVJEkqD/dcSdJxHDqUHva7YAHs2pXG+vaFadNSUTVyZGXjkyRJ\nAvdcSV1ynXjlvPACfO97cMst8P3vp8Jq+HB45zvh9tvhppt6XmFlvikr5pqyZL6pXjhzJamqxAhP\nPJFaqa9Zk44BLrwwtVK/4orUBVCSJKnauOdKUlU4cgQefTTtp9q+PY317g1XX52W/o0ZU9n4JElS\nbXDPlaS6tXcv5HLw4INw4EAaGzwYmppg5kwYNKiS0UmSJBXPxTVSF1wnXnoxpudRff3r8IlPwH33\npcJq7Fj48Ifh85+Ht72tPgsr801ZMdeUJfNN9cKZK0mZOXYMVqxI+6m2bEljDQ0weXLaT3XuubZS\nlyRJtcs9V5LKbv/+tOwvl4OXX05jAwakZX9NTTB0aCWjkyRJPYl7riT1SNu2pVmqZcvg6NE0Nnp0\nmqW65hro06ey8UmSJJWSxZXUhVwuR1NTU6XDqBktLfDYY6mo2rgxjYWQWqjPnQsXX+zSv+Mx35QV\nc01ZMt9ULyyuJJXEq6/C4sWwYAHs3p3GTjkFpk+HOXPg9NMrG58kSVK5uedK0knZuTM9m2rJEjh8\nOI2NGJFmqaZPTwWWJElSVtxzJammxAjr16eias2atvGLL05F1eWXpy6AkiRJ9cTiSuqC68TbHD4M\nS5emouq559JY794wZUoqqs48s7Lx9QTmm7JirilL5pvqhcWVpC7t2ZPaqD/0UHrYL8CQIamN+syZ\nMHBgJaOTJEmqDu65ktShGGHz5jRLtXJl6gIIcN55aZZq0iRobKxsjJIkSe2550pS1Th2DJYvT63U\nn302jTU2pudSzZ0L555b2fgkSZKqlVvOpS7kcrlKh5CJffvgZz+D226Db34zFVYDB8Lb3gaf/zx8\n+MMWVlmol3xT5ZlrypL5pnrhzJVU5559Ni39W7YszVpBakxx3XVptqp378rGJ0mSVCvccyXVoZYW\nWLUqFVVPPpnGQoAJE9LSvwsvTMeSJEm1xj1XkjLx6qup49+CBakDIEC/fnDttTBnDgwfXtn4JEmS\napl7rqQu9IR14s8/D9/9LtxyC/zoR6mwOv10ePe74fbb4Z3vtLCqFj0h31QbzDVlyXxTvXDmSuqh\nYoS1a9PSv7Vr28YvvTQt/Rs/3qV/kiRJpeSeK6mHOXwYHnkkFVXPP5/G+vSBKVNSUTV6dGXjkyRJ\nKif3XEk6aS++mPZSPfQQHDyYxoYOTXupZsyAAQMqG58kSVJP554rqQvVvE48Rti4Eb76VfjkJ+GB\nB1Jhdf758JGPpOdTveUtFla1pJrzTT2LuaYsmW+qF85cSTXo6FFYvhzmzYOtW9NYY2Na+nfddXDO\nOZWNT5IkqR6550qqIS+/DAsXwqJFsH9/Ghs0CGbPhlmzYPDgysYnSZJUae65knRczzyTZqmWL4fm\n5jQ2ZkyapZo8GXr3rmx8kiRJsriSupTL5Whqasr8c1taYOXKVFRt3pzGGhpg0qTU9e+CC2yl3hNV\nKt9Uf8w1Zcl8U72wuJKqzIED8OCDkMvB3r1prF+/1PFvzhw47bSKhidJkqROuOdKqhI7dqRnUy1d\nCkeOpLFRo9Is1dSp0LdvZeOTJEmqBe65kupUjLBmTSqq1q9vG7/ssrSf6tJLXfonSZJUKyyupC6U\nY534oUOwZEl66O/OnWmsTx+YNi0t/TvjjJJ+nGqI+xKUFXNNWTLfVC8srqQM7d6dZqkWL04PDiBj\n+wAAGuNJREFU+wUYNiwt/bv2Wujfv7LxSZIk6cS550oqsxhh48ZUVK1enY4Bxo1LRdXEiakLoCRJ\nkk6ee66kHujoUXj00VRUbduWxnr1gquvTvupxoypbHySJEkqLX9eLnUhl8t16/qXXoJ77oFbb4Vv\nfzsVVqeeCr/92/CFL8DNN1tYqXPdzTfpRJlrypL5pnrhzJVUIk8/nWapfv1raG5OY+eck5b+TZ6c\nZq0kSZLUc7nnSjoJzc2wYgXMm5eKK0j7p668Mi39O+88W6lLkiRlyT1XUo155RV48EFYuBD27k1j\nAwbAjBnQ1JQ6AEqSJKm+uOdK6kLhOvHt2+Guu9J+qnvuSYXVGWfAe96T9lO94x0WVjo57ktQVsw1\nZcl8U72oyMxVCOF64MtAI/CNGOMX251/D/CXQAD2Ax+LMT6WeaAS0NKSWqjPnw8bNrSNX3552k91\nySUu/ZMkSVIF9lyFEBqBJ4A3AduBZcBNMcb1BddMA9bFGF/OF2KfjjFObXcf91yprA4dgocfhgUL\nYNeuNNa3L0yfDnPmwMiRlY1PkiRJb1Rve66uATbFGLcAhBDuBm4AXiuuYoxLCq5fCpyVZYCqby+8\nkAqqxYtTgQUwfHgqqKZPh/79KxufJEmSqlMliqszga0Fx9uAKce5/sPAL8oakepejGnJ3/z5sGZN\nOga46CI49dQcH/pQEw3uUFQGcrkcTU1NlQ5DdcBcU5bMN9WLShRXRa/lCyHMAT4EXNvR+Ztvvpmx\nY8cCMGTIECZOnPja/7itGyc99vh4x9OnN7F0Kdx5Z44XX4TRo5vo3RsGDsxx1VXwznc2kcvBokXV\nEa/HHnvscamOW1VLPB737ONW1RKPxz3ruPX9li1bqLRK7LmaStpDdX3++DagpYOmFlcAPwaujzFu\n6uA+7rnSCdu7F3K51E79wIE0NngwNDXBzJkwaFAlo5MkSdKJqrc9V8uBcSGEscAO4F3ATYUXhBDO\nJhVW7+2osJJORIzw1FNp6d+KFakLIMDYsemBv5MmQS+f/CZJkqQTlPk/JWOMx0IIHwfuJ7VivzPG\nuD6E8NH8+TuAvwGGAl8Jqcf10RjjNVnHqp7h2LFUTM2bB62zxQ0NMHlyKqrOPff4rdRzudxr089S\nuZlvyoq5piyZb6oXFfk5fYzxl8Av243dUfD+D4E/zDou9Sz798OiRbBwIbz8chobMABmzYLZs2Ho\n0MrGJ0mSpJ4l8z1XpeKeK3Vm69a09G/ZMjh6NI2NHp1mqa65Bvr0qWx8kiRJKp9623MllVxLC6xe\nnYqqjRvTWAgwYQLMnZtaqh9v6Z8kSZJ0siyuVNNefRUefjh1/tu9O42dckp62O+cOXD66Sf/Ga4T\nV5bMN2XFXFOWzDfVC4sr1aSdO9Ms1ZIlcPhwGhsxIs1STZ+eCixJkiQpS+65Us2IEdatS0XV44+3\njV9ySSqqxo9PXQAlSZJUv9xzJR3H4cOwdGlqpf7882msd2+YMiUVVWeeWdn4JEmSJAB/zq+qtWcP\n/OhHcOut8J3vpMJq6FB4+9vh9tvhfe/LprDK5XLl/xApz3xTVsw1Zcl8U71w5kpVJUbYvDkt/Vu5\nMnUBBDjvvDRLNWkSNDZWNkZJkiSpI+65UlU4dgyWL09L/559No01NsJVV6XnU40dW9HwJEmSVCPc\nc6W6tW8fLFwIixal9wADB8KsWTB7NgwZUtn4JEmSpGK550oV8eyz8K1vwW23wc9+lgqrs86C978/\n7ae64YbqKaxcJ64smW/KirmmLJlvqhfOXCkzLS2walVa+rdpUxoLASZOTEv/xo1Lx5IkSVItcs+V\nyu7VV+Ghh2DBgtQBEKBfP7j2WpgzB4YPr2x8kiRJ6jncc6Ue6bnnUkG1ZAkcOZLGRo5MBdW0aXDK\nKZWNT5IkSSoliyuVVIywdm1qpb52bdv4pZemVurjx9fe0r9cLkdTU1Olw1CdMN+UFXNNWTLfVC8s\nrlQShw+nGar582HnzjTWpw9MnZqKqjPOqGx8kiRJUrm550onZfduyOXSnqqDB9PYsGHQ1AQzZsCA\nAZWMTpIkSfXGPVeqKTHCk0+mWarVq1MXQIALLkizVFdeCQ02+ZckSVKdsbhS0Y4ehWXLUlG1dWsa\na2yEKVNSK/VzzqlsfOXiOnFlyXxTVsw1Zcl8U72wuFKXXn4ZFi6ERYtg//40duqpMGtWeg0eXNn4\nJEmSpGrgnit1asuWNEu1fDk0N6exs89OS/8mT4bevSsaniRJkvQG7rlS1WhuhlWrYN482Lw5jTU0\nwKRJqai64ILaa6UuSZIkZcG2AwLgwAG47z74q7+Cr30tFVb9+8Ob3wyf+xx89KMwblx9Fla5XK7S\nIaiOmG/KirmmLJlvqhfOXNW5HTvS0r+lS+HIkTQ2alSapZo6Ffr2rWx8kiRJUq1wz1UdihHWrElF\n1fr1bePjx6ei6tJL63OGSpIkSbXPPVfKxKFDsHgxLFgAL7yQxvr0gWnTUlE1alRl45MkSZJqmcVV\nD9fcnB74u3JlWvp38GAaP+00mDMHrr027a1S53w2h7Jkvikr5pqyZL6pXlhc9UBHjsC6danr32OP\npWYVrS68MM1STZiQugBKkiRJKg33XPUQr76aCqlVq2Dt2rbmFJCW+115ZXo21VlnVS5GSZIkqdzc\nc6UT8tJLsHp1WvK3cWPbg34Bxo5NBdXEie6lkiRJkrJgcVVjdu5Ms1MrV8LTT7eNNzTAxRenYmri\nRBg6tHIx9jSuE1eWzDdlxVxTlsw31QuLqyoXIzz7bCqoVq1Kz6Vq1bs3XHZZKqauuAIGDKhcnJIk\nSVK9c89VFWppgU2b0uzUqlWwZ0/buf79UyE1cWJ6HpUP+ZUkSZLauOdKHD2aHui7cmVqTPHKK23n\nhgxpW+534YXQ2Fi5OCVJkiR1zOKqgg4ehDVr0uzU44/D4cNt50aObGtIMXYshIrU3gLXiStb5puy\nYq4pS+abuhJjpDk209zSzLGWY697NcdmjjYfpTl2cK7d9c2xuesPKyOLq4y9/HJbh78nnnh9h79z\nzknF1JVXpg5/FlSSJEkqhdbipZjipH0h09zSzNGWo50WPse7X3eKo57APVcZ2LWrrcPfU0+lJhWQ\nOvyNG9e25G/YsMrGKUmSpBMTY+z2LEtnxUmxhUyxxVHrddWusaGRXg29Xns1hkZ6N/amMbQbb3dd\n67Wt728cf6N7rnqSGGHbtraCavv2tnO9e6dGFK0d/gYOrFyckiRJtaIlthRdSFSikKmF4qWr4qSj\nQqaj6zu7pndD79dd293iKPSAZVsWVyXS0gKbN7e1TN+9u+1cv35w+eVpud9ll9nhr9a4TlxZMt+U\nFXNN7bXEli6Lk/ZFR7HFyWNLH+PCqy58w327Uxy1xJZK/xZ1qTvFSftrOypOOitmTqQ46inFS7Wz\nuDoJx46lDn+rVqV9VPv3t50bPBgmTEgzVBddBL38nZYkqW7FGInEbs2ynGgh093lYq3XlrN42fH8\nDnY9u+uk7hFCOKFZls6Kk2ILmWKLo4bQYPEi91x116FDr+/wd+hQ27kRI9o6/J13ng0pJEnKSozx\ndTMvXRUS7YuUjq4rdmlZscVRtf+bqyE0dFqcFFt0dHZd+2u7Wxz1auhFIFi8qCg+56rK7d/f1uFv\nw4Y0Y9VqzJi2gmr0aAsqSVLPVNhprJhCorvFycksF2t9VbvW4qWYGZTuFDInsuG/o89vCA2V/i2S\nap7FVSd2725rSLF5c1uHvxBe3+Fv+PDKxqnyc1+CsmS+1a/jtUnuaClYdwqZjtoqr1u2jvOuPK+o\ne7Z+frVrPzNSjg33xe6T6eh+9Vy8+Geb6oXFVV6Mqatfa0OKrVvbzvXqBZdckoqpCRNg0KDKxSlJ\nOjGtbZKLLSROdLlYR4VMMbM8WRcvO/bu4NCuQ11fWKC7syzFFjKd7Wkpdpan9XqXjEmqtLrecxVj\neu7UypWpoNpVsM/ylFNg/Pi05G/8+HQsSepca5vkUs+ydHffS2efXwszLye64T6LtsoWL5JqhXuu\nMnTsGDzxRCqoVq+Gffvazg0alGamrrwSLr7YDn+SqktHbZKrqZCplTbJnRUnpe4c1t3iyOJFkmpf\nXZQPhw+nzn6rVqVOfwcPtp0bPrxt/9T550ND/S6HVidcJ14fCjuNdWdGpLPlXcUuF2t/z00rNjFm\nwpgOP7/ai5fCNsldFSftC5Qs2irbJvn1/LNNWTLfVC96bHH1yitpZmrVqvQsqqNH286deWZbh7+z\nzrLDn5SF1s36xc6ydLTvpRzLxQqvr4Zl0nsO7uGUAx2vQ27faaw7G+6LLXo62yfTVXHUWrxIklTP\netSeqxhh8WJ45BHYtAlaWlqvTc+dai2oRoyoQMBSmbVvk1zqWZbOru9OIVPtWpdmlaqtcXeXlh1v\nBqfeO41JklQs91yVwHPPwV13pbbpAI2NcNllbUv+Tj21svGp9nW3TXKpl4sd75612Ca5q0LiZAqZ\nzpaNHa+QsXiRJEknK/PiKoRwPfBloBH4Rozxix1c80/AW4FXgZtjjCs7u19zM9x/P/z856lZxeDB\n8Pa3p4KqX79yfRcqh/ZtkrsqJE50lqW7hcyWVVsYNX5UTRQvJzIj0tnm/GKXi3WnOHK/S9fcl6Cs\nmGvKkvmmepFpcRVCaAT+BXgTsB1YFkL4aYxxfcE1bwMuiDGOCyFMAb4CTO3ofs88A9/+Nmzblo5n\nzIDf+z3o37/M30iNOl6b5HIsFyu2OGp9Vetm/eeefI4Rl6a1pN0pTk62c1h3iyOLl55h1apV/gNE\nmTDXlCXzTfUi65mra4BNMcYtACGEu4EbgPUF1/wO8O8AMcalIYQhIYSRMcad7W92++1pX9WIEfDe\n96b26Vlrv0m/owdHHm+sowdVdnfseEVR4bXVWry0Kuw0lkXnsGKLoy8s+wKf+c3P2GlMmXjppZcq\nHYLqhLmmLJlvqhdZF1dnAlsLjrcBU4q45izgDcXV3sYNTLm2hVmzmznUu5kVz6WZmVIUKd0ZqxWt\nnca62zmsmOvaX9vd4qhXQy8CoSqLl9aYJUmSpOPJurgqtjVh+39hd/h1DdO/xIZBsGHVyQV1sjpq\nWdzdsfbFSFfXti9Uihlzs/6J2bJlS6VDUB0x35QVc01ZMt9ULzJtxR5CmAp8OsZ4ff74NqClsKlF\nCOGrQC7GeHf+eAMwu/2ywBBCbfaQlyRJklRW9dKKfTkwLoQwFtgBvAu4qd01PwU+DtydL8Ze6mi/\nVaV+wyRJkiSpI5kWVzHGYyGEjwP3k1qx3xljXB9C+Gj+/B0xxl+EEN4WQtgEHAA+mGWMkiRJknQi\nMl0WKEmSJEk9lR0O1GOEEMaEEBaEENaGEB4PIfyP/PiwEMIDIYSNIYRfhRCGFHzNbSGEJ0MIG0II\nby4YvyqEsCZ/7h8LxvuGEL6XH38khHBOwbkP5D9jYwjh/QXj54YQlua/5u4QQu/y/24oKyGExhDC\nyhDCvflj801lkX80yQ9DCOtDCOtCCFPMN5VDPnfW5vPku/ncMNdUEiGEfwsh7AwhrCkYq9r8CiH8\nU358dQjhyi6/wRijL1894gWMAibm3w8EngAuAf4e+Mv8+C3A7fn3lwKrgN7AWGATbbO5jwLX5N//\nArg+//7/Af41//5dwN3598OAzcCQ/GszMDh/7vvAjfn3XwH+uNK/V75Kmnd/DnwH+Gn+2HzzVa5c\n+3fgQ/n3vYDB5puvMuTZWOApoG/++HvAB8w1XyXMsZnAlcCagrGqzC/gbcAv8u+nAI90+f1V+jfY\nl69yvYB7gDcBG4CR+bFRwIb8+9uAWwquvw+YCpwBrC8Yfzfw1YJrpuTf9wJ25d/fBHyl4Gu+mv+6\nAOwCGvLjU4H7Kv1746tkOXYW8N/AHODe/Jj55qscuTYYeKqDcfPNV6lzbRjph5ND83lwL/Ab5pqv\nEufZWF5fXFVlfgF3AO/qKM7OXi4LVI8UUkfKK4GlpP8JWjtO7gRG5t+PJj2kutU20kOs249vz49D\nwUOuY4zHgJdDCKcd517DSB0vWzq4l2rfl4D/BbQUjJlvKodzgV0hhG+GEFaEEL4eQhiA+aYSizHu\nAf4BeJbU2fmlGOMDmGsqr2rNr9Gt9yr4mrOO941YXKnHCSEMBH4E/M8Y4/7CczH92CGrLi52i+nB\nQgi/BbwQY1zJGx98DphvKqlewCTSUpdJpG66txZeYL6pFEII5wN/RppZGA0MDCG8t/Aac03lVIX5\n1f7v+ON+jcWVepT8BsQfAXfFGO/JD+8MIYzKnz8DeCE/vh0YU/DlZ5F+IrGd1/9UonW89WvOzt+r\nF2mt7osd3GtMfmwPMCSE0FBwr+0n+W2qOkwHfieE8DTwn8DcEMJdmG8qj23AthjjsvzxD0nF1vPm\nm0psMrA4xvhi/qf+PwamYa6pvKr1786OPv+4uWdxpR4jhBCAO4F1McYvF5z6KWkzLvlf7ykYf3cI\noU8I4VxgHPBojPF5YF9InbgC8D7gJx3c6/eBefn3vwLeHFI3r6Gk9en353/6sgB4ZwefrxoWY/xE\njHFMjPFc0prt+THG92G+qQzyebI1hHBhfuhNwFrSfhjzTaW0AZgaQuiXz5E3Aesw11Re1fp350+B\n9wOEEKaSlg+2Ll/sWKU3tPnyVaoXMIO092UVsDL/up60lva/gY35/7GGFHzNJ0idZzYAbykYvwpY\nkz/3TwXjfUkdZZ4EHgHGFpz7YH78SeADBePnkvZ+PUnqutS70r9Xvkqee7Np6xZovvkqV55NAJYB\nq0mzCYPNN19lyrW/JBXva0hdKnuba75KmF//SdrPd4S0n+mD1ZxfwL/kP2M1MKmr78+HCEuSJElS\nCbgsUJIkSZJKwOJKkiRJkkrA4kqSJEmSSsDiSpIkSZJKwOJKkiRJkkrA4kqSJEmSSsDiSpJ6iBDC\nzSGElhDC3hDCkHbneuXPfaoCcX06/9lV/XdOCKEhhPDlEMJzIYTmEMKPKx2TJKm2VPVfdJKkEzIY\nuKWTc5V6uGEtPFTx94H/AXwRmE56kKokSUWzuJKknudXwJ+GEE6vdCAFQllvHkLfEtzmkvyv/xhj\nXBpj3NSNz28MITSWIAZJUg2zuJKknudz+V8/ebyLWpfrdTD+rRDC0wXHY/PL+v44hHB7COH5EMK+\nEMJdIYT+IYSLQggPhBD2hxCeDCG8r5OPvDSEsCCEcCCEsCOE8JkQwuuKrhDCiBDCV0MI20IIh0II\n60MIf9TumtbljzNDCD8IIewFHunie70+hLAkhPBqCOGlEMJ/hRAuLDi/BWhdMtmcv//7j3O/lhDC\n50IIt+Z/rw4D4/Pn3htCWB1COBhC2BVC+HYIYVTB1/5zCOHJdvf7df6e5xeM/V0I4fmC47eEEBbn\n498fQtgQQvjr433fkqRsWVxJUs/zHPAvwEdCCGd3cW1ny/U6Gr8NGAm8D/gb4F3AN4D/An4C/C7w\nGPCtEMKlHXz9PaRZtRuA7wJ/nb8PACGEU4GHgOtJhc7bgHuBr4QQPt7B/b4DbAZ+j86XQRJCuB74\nObAPuBH4GKkQeiiEMDp/2e8C38q/n5p//aKze+bdDLwV+PN8rM+FED4CfBtYC7wduBV4C7AwhDAg\n/3XzgfNDCGPy8Q0FJgKvAnML7j8XWJC/5jzgp/nv90bgt4H/A/TvIkZJUoZ6VToASVLJRdK+oY+S\nipQPH+fazpbrdTT+ZIzxg/n3D4QQZgLvBt4bY/wupBkY4HdI+5f+tt3Xfy3G+Pf59/+dL6b+IoTw\npRjjPuB/AmcD42OMm/PXzc835/hUCOFfY4yFM20/iDHeepzvrdXngE3AW1u/PoSwBNgI/AXwFzHG\nVSGEHQAxxkeLuGerN8cYD+fv2Qh8FlgQY/yD1gtCCBuAB4EPAf8MLCT9N2oC7gJmAy8DPwbmAF8P\nIQwErgK+mb/NJKA38LEY4yv5sVw34pQkZcCZK0nqgWKMe4F/AN5fuPztJP2y3fET+V/vL/jcl4AX\ngLM6+Prvtzv+HjCQ/HI60ozVI8CWfHfDXiGEXqTZrtOA9rNh/9VVwPnZoiuB7xUWZjHGLcDDpMLm\nRN3XWljlXQSMIM2ovSbG+DDwTOtnxRj3AKuB6/KXzCUVSv9NKq4AZpF+ALogf7wSOAp8L4Twe1W2\nn06SlGdxJUk915eAPaQZpFJ069vb7vjIccZP6eDrd3ZyfGb+19NJBcjR/D1aX98nxX9au69/roiY\nh5Jm4Tq6dicwrIh7dKb9PYd1Mt76WUMLjhfQVkjNyR8vAEaGEC7Jj22PMT4JkJ/Jewvp7+27SEsQ\nl4QQZp1E/JKkErO4kqQeKsZ4APgC8E7Snp72DkF6Bla78dMoT+v0Ue2OR+Z/3Z7/dTdpNmlyB6+r\ngV+3+/piYtybv679Z7fG82IR9+hM+8/fk//1jE4+a0/BcQ4YE0KYRpqRmx9j3AmsJ81kvbbf6rUP\nizEXY3wrqdX+m4BjwM9DCO2LTklShVhcSVLP9q+k4uXvOjj3TP7Xy1sH8vubppcplhvbHb8b2A+s\nyR/fR2qHvjXGuKKD1yt0U77A/DVwY+FDjEMI55C+z9wJfB+d2UCaoXp34WAIYTppL1nhZy0Emkmz\nirtijGvz4/NJDTom0K64ahVjPBpjXAD8b2AAMLZk34Ek6aTY0EKSerAY45EQwt8CX+vg9C9IjRS+\nHkL4FGkp31+SCp6TeS5VZ1/7h/kCZzlpiduHgU/FGPfnz3+J1IHwwRDCl0gNJwYAFwMzYoy/e4Lx\n/DWpW+DPQghfIe3z+gxpVusfTvCebxBjbAkh/A1wRwjhLtLeqzNJhe1G4N8Krt0XQlhB2ndVuBdt\nAfAnpFmx+a2DIYQ/BmaS/pttA4aTujduBx4v1fcgSTo5zlxJUs/S0VK5bwJPtj8XY3wZ+C2ghfQP\n/L8D/pH0D/xiltzFTq5rP9Z63Q3Ab5Datv8B8NkY42cL4tlHmk36Bam1+n3AnaS24/M7uGdRYoz3\nA78JDCE10fgKqVX6jBjj84WXdue+nXzW10mt6i8ntZ7/Iqnhx+wY48F2l7f+Ps/vYOyZGOMzBeOr\nSIXmF/L3+2dSW/a57ZpqSJIqKMRYjmX1kiRJklRfnLmSJEmSpBKwuJIkSZKkErC4kiRJkqQSsLiS\nJEmSpBKwuJIkSZKkErC4kiRJkqQSsLiSJEmSpBKwuJIkSZKkErC4kiRJkqQS+P8BCETaKHSkNQEA\nAAAASUVORK5CYII=\n", + "text": [ + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Another question to ask: How does this scale if we have a growing number of columns?" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "import timeit\n", + "import random\n", + "from numpy import einsum\n", + "import pandas as pd\n", + "\n", + "def run_loc_sum(df, n):\n", + " return df.loc[:, 0:n-1].sum(axis=0)\n", + "\n", + "def run_einsum(df, n):\n", + " return [einsum('i->', df[col].values) for col in range(0,n-1)]\n", + "\n", + "orders = [10**i for i in range(2, 5)]\n", + "loc_res = []\n", + "einsum_res = []\n", + "\n", + "for n in orders:\n", + "\n", + " df = pd.DataFrame()\n", + " for col in range(n):\n", + " df[col] = pd.Series(range(1000), index=range(1000))\n", + " \n", + " print('n=%s (%s of %s)' %(n, orders.index(n)+1, len(orders)))\n", + "\n", + " loc_res.append(min(timeit.Timer('run_loc_sum(df, n)' , \n", + " 'from __main__ import run_loc_sum, df, n').repeat(repeat=5, number=1)))\n", + "\n", + " einsum_res.append(min(timeit.Timer('run_einsum(df, n)' , \n", + " 'from __main__ import run_einsum, df, n').repeat(repeat=5, number=1)))\n", + "\n", + "print('finished')" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "n=100 (1 of 3)\n", + "n=1000 (2 of 3)" + ] + }, + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "\n", + "n=10000 (3 of 3)" + ] + }, + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "\n", + "finished" + ] + }, + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "\n" + ] + } + ], + "prompt_number": 35 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "from matplotlib import pyplot as plt\n", + "\n", + "def plot_2():\n", + " \n", + " fig = plt.figure(figsize=(12,6))\n", + " \n", + " plt.plot(orders, loc_res, \n", + " label=\"df.loc[:, 0:n-1].sum(axis=0)\", \n", + " lw=2, alpha=0.6)\n", + " plt.plot(orders,einsum_res, \n", + " label=\"[einsum('i->', df[col].values) for col in range(0,n-1)]\", \n", + " lw=2, alpha=0.6)\n", + "\n", + " plt.title('Pandas Column Sums', fontsize=20)\n", + " plt.xlim([min(orders), max(orders)])\n", + " plt.grid()\n", + "\n", + " #plt.xscale('log')\n", + " plt.ticklabel_format(style='plain', axis='x')\n", + " plt.legend(loc='upper left', fontsize=14)\n", + " plt.xlabel('Number of columns', fontsize=16)\n", + " plt.ylabel('time in seconds', fontsize=16)\n", + " \n", + " plt.tight_layout()\n", + " plt.show()\n", + " \n", + "plot_2()" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "display_data", + "png": "iVBORw0KGgoAAAANSUhEUgAAA1cAAAGpCAYAAABhxcywAAAABHNCSVQICAgIfAhkiAAAAAlwSFlz\nAAALEgAACxIB0t1+/AAAIABJREFUeJzs3Xd8VUX+//HXJIQWQhHpBEJTirTQEZCyYC8sioqCwYar\ngq4rILoKiIKF9YdtXRtlFRYXRWX1qyCGEEBUOiIgQaqEKk16m98fc+9NDzch996U9/PxyCOcOefM\nmXPvhNxP5jNzjLUWERERERERuTBhoW6AiIiIiIhIYaDgSkREREREJA8ouBIREREREckDCq5ERERE\nRETygIIrERERERGRPKDgSkREREREJA8ouBIRKSKMMZONMeeMMbVC3ZZgMcYkGGPOhbodIiJSNCi4\nEhG5AJ5gJfXXGWPMXmPMt8aY20Pdvkzkq4cbGmOijTEvGGOWGWMOGGNOGWN2G2O+McYMMcaUzYPL\n5Kt7DgRjTBtjzFRjzFZjzAljzCFjzEZjzP+MMUONMaVD3UYRkaKgWKgbICJSCFhgtOffEUAj4Eag\nmzGmtbX2byFrWUYm1A3wMsbcC7wBFAdWAlOBA8BFwOXABOBpoFKo2lgQGGPuBKbg+mE88AlwHIgB\nOgHXeMo2haiJIiJFhoIrEZE8YK19NvW2MaY78A3wqDHmNWvt1tC0LH8yxtwBvAPsB/pba7/K5Jh2\nwJvBbltB4hmRehM4C1xprZ2XyTHtgd+D3TYRkaJIaYEiIgFgrY0HfsGNFLUGMMbcZIz50BizwRhz\nxPO11Bgz2BiTYUQp1Ryp2saYQcaYn4wxx40xu4wxb2eVMmeM+ZMxZoEx5qgx5ndjzKfGmIZZtdUY\nE2eM+cQYs8kYc8yTUrbQEwBldnxdY8w7nrSzY55rrDbGvGWMueh8r40xJgp4DTfScltmgZXnNfwB\n6JjJ+T2MMV8bY/Z7UuB+McaM8zeF0HO/54wxd2Wx/5wxZl66slGe8iuMMbd73rejxphkY8w/jDHF\nPcf9yRgz3xhz2JPm+EFmr4kxZosxZrMxprQx5mVjzDbPvSQZY4b5cx8elwFRwJrMAisAa+331tpD\nqa4d47mXSVncf4Z5asaYrp5zRhpjWnte/4Oee/zEGBPtOa6eMeYjT2rsMWPMPGNMs0yuUcUYM97z\n3h3x1LPeGDPJGFMnB/cvIpKvaORKRCRwvAGTd87PONwIw2JgB1AO6AG8CrQBBmRRz8tAL2AW8DXQ\nHbgPqO85P+WCxtwMfASc8HzfCXQGvgNWZ1H/P4E1QILn+ItxqWQfGGMutdY+k6r+asAS3Af6L4EZ\nQEmgLnAn8DpuNCo7NwMVgMXW2rnZHWitPZXu/gYBbwF/eK69B+gGDAeuN8ZcnjqQOI/s5mJltW8w\ncDXwKTAPuBL4K1DJGDML+BD4AvgXLrXxDqAi7vVMX38EMAeohnstzwC9gReMMSXTj4ZmYZ/ne3Vj\nTGlr7TE/zkndhpzua4N7rRNwI4/NcG1uaozpDSQCPwOTcWmJfwa+McbUtdYeBd9o2yJcn5kDfI77\nWYkBbsC9r5tzcB8iIvmHtVZf+tKXvvSVyy/gHHA2k/I/efadAaI9ZXUyOc7gPoieA9qm2+ct3wLU\nTFUeDsz37GuTqrwMLv3rJBCbrq5XvG0FaqXbl1m7IoC5wCmgeqrywZ56BmdyTimgpB+v2fueOp7N\n4Wtd23NvB4FL0u1701Pn2+nKE9K/P0Cc59gB2byn8enKRnnKDwCXpiovjgtMz+KCys7p3ts5nvOa\np6tvi6f8C6BEqvJKnmscAIr5+br84KlrBfAg0AIons3xMZ7jJ2axP7PXrKvnnHPA7en2vecpPwiM\nSLfv7559Q1KVXe8p+0cm1y4GlLnQn0t96Utf+grVl9ICRUQunPGkS40yxjxvjPkYN8JkgQnW2u0A\n1toMf4231lpcihy40anMPGut/S3VOWcBb0pXm1TH3YgbEZpmrV2ero5RwOHMKs+iXadxI1rFSDs6\n5h3ROJHJOcettRnKM1HN8/23bI/K6E5c0PeGtXZDun1PAUeAO70pegHymrX2F++GdSNrH+ECqVnW\n2gWp9lncSBa4EZ70LC7oOJnqnL24EcpywCV+tulmXEDUHLdAyHLgiDHmB2PMME8aZl5ZYK39T7qy\nKZ7vvwMvpNv3b8/35pnUlVkfOmOtPXJhTRQRCR2lBYqI5I2Rnu8WN+owH3jfWjvNe4AxpiIwFJci\nVhdIvzx2jSzqXppJmTcwqZCqLNbzfX76g621h40xK4Eu6fcZ99yr4bggKho3ApVa9VT/ngWMBd40\nxlyJG5lZaK1dm0Xb85L3/uLT77DWHjTGrMClQDYk6xTIC5XZe7HT831ZJvuSPd9rZrLvkLU2sxX8\ntnu+V8hkXwae4L27Z15dT6AV0BYXeLcBHjTGdLXWbvGnvvPI7v5XegLK1DK7/wRcWuwTxphY4Ctg\noed8PZNMRAo0BVciIhfOWmvDszvAGFMeN1cpBpfGNRmXRnYG9yH6EaBEFqcfzKTsjOd76uuW83zf\nnUU9uzJpV13gR6A8br7M18AhXJpbHeCu1O2y1m4zxrTFjYRdhZtTgzFmOzDeWvt6FtdOzfthPLOA\nIzve+9uZxf6d6Y4LhMzmc53xY19EJvsye19Tn5Ntn0rPWrseWO/dNsZcCkwEOgD/Dzc36kLl6P6t\ntWeMW6slIlXZH8atYDgaN8fqSs+ufcaYfwLPWWvPpK9LRKQgUHAlIhIc9+ICq1E247LtHXDB1YXy\nfritksX+qpmUPYZ7rlSctfbfqXcY9xDkDCvqeT7E32aMCcele/0JNxfrVWPMUWvtxPO0cwEwEDdS\n9sx5jk3Ne3/VgHWZ7K+W7riseEdHMvwO9ATBhYK19hdjTH9gI27RD68s798j4K+BtXYH7mcCY0xj\n3CItD+H6Qxg56xciIvmG5lyJiARHfc/3TzLZd0UeXcObltY1/Q5jTDncQgfp07bqe8py3C5r7Vlr\n7XJr7UvA7Z7iG/1o58e4UbsOxpge2R2Ybv6Udx5Z10yOK4+7v+NkHnildsDzvVYm+1qf59yCxjt/\nKfVS/977j05/sGc5e3/neuUJa+1aa+0buJRG8K8PiYjkSwquRESCw7toROoRBIwxLYEReXSNz3Ef\nnPsZY1ql2zcKyOw5UJtxH7zTt+tKPCML6cpjPYFaet5RsfMuBe5ZsGCIZ/MjY0ymC3l4RvR+SFX0\nIXAaGGyMqZfu8DG45eE/9CzGkZ0luNGbfsYY3/wyz/OoXjpf+/MTzzOrhmT2jC/j8vGe8mwmesut\ntX/g0gc7GWMapTo+HLeqZMkAt7mxMSaz0VW/+5CISH6ltEARkeD4N24xiwnGmG64VK0GwLW4UaPb\nLvQC1tqjxpj7cavXLTDGfISbZ9UJaIL7gJ1+QYt/4lL0ZnhWOdyJezDtlcB/gVvTHT8AuN8YsxDY\nhAvm6uGW1z4BTPCzrdM8gc0bwNeexTYWe+qriJsn1AzYm+qcrcaYR3HLri83xvwX95ynK4D2uBGr\n4ZlcLs0Dmq21u4wxU4H+wEpjzP/hAs+rcYuBtPDnHvKJ8rjX/CVjzCLcM6b+ACrjUu3q4Obg/S3d\neS/jlsRf5HnfT+AC7HBgFZmv7pdXegEvG2O+A5JwzyqriRuxOutpm4hIgaTgSkQkCKy1O40xnXFL\nVXfCBS/rgL8A35J5cGXJ/kGvmV3nE2PMVbjVC/viPjQn4oKPEbjV9FIf/5Mn2HsOF+gVA1biFj84\nRMbgahru2U4dcavSlcKtXDgN99wiv1cNtNa+b4yZDTyMSwnrB0TiAqw1wKO4BRlSn/OWMWYj8DjQ\nB7fi4jbciNNYa2365eazeg3vwwUdt+OeDbUV9zDn8Zncc3b1nG9fVs73AF9/61uLe6964d7jvrg5\ndEdxAfxU4FVr7e9pLmDtJM/I1mO4gHk/buTzKVywn9P7yYmvcSmJXXALWpTFrSo4G3jFWvt9AK8t\nIhJQJuOqqQG+oPulPwH317H3rLUvZnFcG9xfMW+11n7iKduCe07LWeC0tbZtUBotIiIiIiJyHkEd\nufLkc7+BW1lqB7DEGDPLWrsuk+NexP11KzULdLXW7g9Ge0VERERERPwV7AUt2gIbrbVbPBOOp5P5\nqkCDcatJ7c1kn8mkTEREREREJKSCHVzVIOXJ8+Dy9GukPsAYUwMXcL3lKUqdt2iBucaYpcaY+wLZ\nUBERERERkZwI9oIW/kzwmgA8Ya21nsm2qUeqLvdMCq8EfGOMWW+tXRCQloqIiIiIiORAsIOrHaR9\naGE0bvQqtVbAdBdXcTFwtTHmtLV2lrV2J4C1dq8x5lNcmmGa4MoYE9wVOkREREREpMCx1ub5dKNg\nB1dLgQbGmBjcsqu34pbB9bHW1vX+2xgzCfiftXaWMaY0EG6t/cMYE4lbdnZ0ZhcJ9gqIUvjFxcUx\nefLkUDdDihj1OwkF9TsJNvU5CQXPQE6eC2pwZa09Y4x5GPcsi3DgfWvtOmPMIM/+t7M5vSow0/NC\nFAOmWmvnBLrNIgAxMTGhboIUQep3EgrqdxJs6nNSmAT9IcLW2q+Ar9KVZRpUWWsHpvr3JqBFYFsn\nIiIiIiKSO8FeLVCkQCpfvnyomyBFkPqdhIL6nQSb+pwUJgquRPzQooUGTSX41O8kFNTvJNjU56Qw\nMYVt8QdjjC1s9yQiIiIiInnHGFMoVgsMqUCtCiIiIpIV/cFPRKToKFLBFeiXnIiIBE9B/KNeQkIC\nXbt2DXUzpAhRn5PCRHOuRERERERE8kCRmnPlya0McotERKSo0u8dEZH8KVBzrjRyJSIiIiIikgcU\nXImIiIhPQkJCqJsgRYz6nBQmCq5ERERERETygIKrAuq6665j4MCBvu1jx45x8803U758ecLCwti2\nbVua4xMSEggLC2P//v15cn1vfWFhYVxzzTV5Umd+MmrUKN/9/eMf/wh1czJ15swZGjZsyPz58/Os\nzi1bthAWFsby5cvzrM6cWLVqFdHR0Zw4cSIk1xcRtGqbBJ36nBQmCq4KKGNMmiV+J06cyIIFC1i0\naBG7du2iZs2aQWnH2rVr+c9//pMndY0aNYoaNWpQunRpunXrxtq1a/Ok3vR27dpFv379aNSoEcWK\nFUsTpHoNHTqUnTt3UrNmzXy7lPLkyZO5+OKLueKKK/Kszlq1arFr1y6aN2+eZ3WmduDAAfr370/5\n8uUpX748AwYM4NChQ779zZs3p2XLlrz++usBub6IiIjImTOBq1vBVSGxceNGGjVqRJMmTahcuTJh\nYcF5aytXrky5cuUuuJ4XX3yRV155hTfeeIMlS5ZQuXJlevbsyZEjR/KglWmdPHmSSpUqMWLECNq1\na5dp8BQZGUmVKlUIDw/P8+vnlTfeeCPTwPBChIWFUbly5YDdd79+/Vi5ciWzZ8/m66+/Zvny5fTv\n3z/NMQMGDOCtt94KyPVF5Pw0/0WCTX1OguXgQZg1C0aMCNw1FFwVAMeOHSMuLo6oqCiqVq3KuHHj\n0uzv2rUrr732GomJiYSFhdG9e3e/6p05cyZNmzalZMmS1KpVi7Fjx6bZf+rUKZ588kliYmIoWbIk\n9erVC8iIgrWWCRMmMGLECHr37k2TJk2YMmUKf/zxB9OmTcvyPG9qYnx8PO3atSMyMpI2bdqwYsWK\nbK9Xu3ZtXn31VQYMGMBFF110QW3/6aef6NGjB+XKlSMqKooWLVr4fklkloqZPu3Oe8zXX39NbGws\npUuXpkuXLuzYsYP4+HiaNWtGVFQUN9xwAwcOHPDVs3btWlavXs0NN9yQpj1PPPEEDRs2pHTp0tSp\nU4fhw4dz8uRJ3/6ePXvSs2dP3/aRI0do0KABgwcPzrR9p0+fZsiQIdSoUcPXT0bk8n+kdevWMXv2\nbN555x3atWtH+/btefvtt/niiy/YsGGD77hrrrmG3377jcWLF+fqOiIiIiJe1sKvv8K778KTT8KX\nX8Lhw4G7XrHAVV0wDRoUmHrffjv35z7++OPMnTuXmTNnUr16dUaPHk1iYiJ9+vQB4NNPP+Xxxx/n\nl19+YebMmRQvXvy8dS5btoy+ffvy9NNPc8cdd/Djjz8yaNAgypYty8MPPwzAXXfdxcKFC3nttddo\n2bIlv/32G1u3bs223oSEBLp3705CQgJdunTx6/42b97M7t276dWrl6+sZMmSdOnShe+++477778f\ngLi4OObPn8/mzZvTnP/kk0/y0ksvUbVqVR555BHuuOOOgKUUptevXz9atmzJW2+9RbFixfjpp58o\nWbJkjusZNWoUr7/+OmXLlqVfv3707duXEiVK8P777xMWFsYtt9zC6NGjmTBhAgCJiYlER0dTqVKl\nNPWUKVOGSZMmUaNGDX7++WceeOABSpQowbPPPgvAv//9b5o1a8b48eN5/PHHGTJkCCVLlmT8+PGZ\ntuu1117js88+46OPPiImJobt27enCYQeeOABpk6dmu29rVu3jpo1a7J48WLKlClDhw4dfPs6duxI\nZGQkixcv5pJLLgGgdOnSNGnShPnz56c5VkSCQ/NfJNjU5yQQTp+GpUshPh68SxGEhUGrVtCtG7zz\nTmCuq+Aqnzty5AgTJ05k0qRJvhGHSZMmpZlTVaFCBUqVKkVERASVK1f2q95XXnmFrl27MnLkSADq\n169PUlISL774Ig8//DBJSUl89NFHfP31176gJyYmhk6dOmVbb2RkpG/kxF+7du0CoEqVKmnKK1eu\nTHJysm+7evXq1K9fP8P5Y8aM8c07euaZZ+jUqRPJyclUr17d7zbk1rZt2xg6dKgvMKhbt26u6hkz\nZgyXX3454AKWwYMHs3z5clq0aAG4QPfjjz/2HZ+UlETt2rUz1PP3v//d92/vKNM//vEPX3BVrVo1\n3nvvPW699VYOHTrEtGnTWLJkCSVKlMjy/i655BLf+16zZs00Ac+YMWMYNmxYtvdWrVo1wL3P6YNB\nYwyVK1f29YHUbU8dxImIiIj448ABmD8fFiwA7+ySMmWgc2e44gqoUCGw11dwlc6FjDAFwq+//sqp\nU6fSfKCNjIykadOmF1Tv+vXrue6669KUXX755YwePZojR46wYsUKwsLC6NatW47qbdOmTZ6OGqWe\nD5U+bdGrWbNmvn97P8jv2bOH6tWrU6ZMGV8d/fv355///GeetQ3gscce495772XKlCn06NGDPn36\ncOmll+a4ntT34A2QU7/HlStXZs+ePb7tw4cPExkZmaGejz/+mAkTJvDrr79y5MgRzp49y7lz59Ic\nc+ONN3L77bfz/PPP8/LLL2fbl+Li4ujZsyeXXHIJvXr14pprruHqq6/2vaaVKlXKEDDlhaioqDQL\nXYhI8CQkJGgkQYJKfU4ulLWwcSPMmwcrVoD3o0+tWtC9O7RuDRERwWmL5lwVUNbagNUR7NXxqlat\nCsDu3bvTlO/evdu3LzsRqX5avG33BhSrV69m1apVrFq1yjd6k5dGjhzJ2rVruemmm/juu+9o1qwZ\nkyZNAvAtKpL6dT59+rTf95B6UQljTJogqVy5chkW+/j++++5/fbbufrqq/niiy9YuXIlzz33HKdO\nnUpz3IkTJ1iyZAnFihUjKSkp2/tr2bIlW7ZsYdy4cZw7d4677rqLnj17+u7pgQceICoqKtuv3377\nDXDv8969e9PUb61lz549Gd7nw4cPUyHQf1oSERGRAu30aVi0CJ5/HsaPh2XLXHnr1jBsmJtj1aFD\n8AIr0MhVvlevXj0iIiJYvHgxMTExABw9epQ1a9bQoEGDXNfbqFEjFi1alKZs4cKFREdHExkZSYsW\nLTh37hzx8fFceeWVF3IL51WnTh2qVq3KnDlzaNWqFeACgIULF2Y5F8hfuU3Ty4n69eszePBgBg8e\nzIMPPsh7773HwIEDfSM6ycnJVKxYEYCVK1fm2TVnzJiRpmzRokXUqFGDp556yle2ZcuWDOcOHTqU\n06dPM2fOHK688kquvfZarr/++iyvVaZMGfr06UOfPn2Ii4ujffv2/Prrr9SvXz9HaYEdOnTgyJEj\nLF682DcSu3jxYo4ePUrHjh3TnLN161ZfmqSIBJdGECTY1Ockp/bvd6l/CxempP5FRUGXLu6rfPnQ\ntU3BVT5XpkwZ7rnnHoYPH06lSpWoVq0azz77bIZUr/RGjBjBkiVLmDt3bqb7//a3v9GmTRtGjx7N\n7bffzpIlS3jllVd8KxFecskl9O3bl3vvvZdXX301zYIWd955Z5bX/fHHHxkwYAAffPABbdq08ese\njTE8+uijjB07loYNG9KgQQOee+45oqKi6Nevn9/3lBPeIOfQoUOEhYWxcuVKihcvTuPGjbM9r0eP\nHrRr146xY8dy/PhxHn/8cfr27Uvt2rXZvXs3CxcupH379oALgKKjoxk1ahQvvPACmzdv5rnnnrvg\ntgN07tyZBx98kL179/qCuEsvvZQdO3Ywbdo02rdvz+zZs5k+fXqa87766iveeecdFi5cSJs2bRg1\nahT33nsvq1evzjDnDdzcvOrVq9O8eXMiIiKYOnUq5cqV8835y0laYKNGjbjqqqsYNGgQ77zzDtZa\nBg0axPXXX5/mDwXHjh1j7dq1fi+IIiIiIoWftZCU5FL/Vq5MSf2rXdul/rVqFdwRqqwouCoAxo8f\nz9GjR+nduzeRkZEMHjyYY8eOpTkm/UOFd+3axaZNmzIc49WyZUtmzJjByJEjGTt2LFWrVmXEiBE8\n9NBDvmP+/e9/8/TTTzNkyBD27dtHzZo1eeyxx7Jt67Fjx0hKSuL48eO+slGjRp03IBw2bBjHjx/n\noYce4sCBA7Rv3545c+akmVd0vnvKriy92NhY37HWWv73v/8RExOTof70Nm3a5FtIolixYhw8eJC4\nuDh27txJxYoVuf76632jbREREUyfPp0HH3zQ93DccePGZRgl8uce0r+/TZo0oWnTpnz++efce++9\nAFx33XUMHTqURx99lOPHj3PllVfy7LPP+t7TvXv3cvfdd/P000/7At8nnniC2bNnc/fdd/Pll19m\nuHbZsmV5+eWXSUpKwhhDbGwsX331Va5WRASYNm0agwcP9o2G3njjjbzxxhtpjvnyyy+Jjo7WSoEi\nIaL5LxJs6nOSnVOn4McfXVDlmWlAeDi0aeOCqjp1IMgzWrJl8mLuTn5ijLHZzSUqbPcbKt4l1/fu\n3etLecvKXXfdxZ49e/jqq6+C1Lq8ExMTw5AhQ84bVIbCe++9x6RJkzKkdxZ0119/PV26dGHo0KGh\nborIBSuIv3f0QVeCTX1OMvP77ympf0ePurKyZd2qf3mR+uf5/znPwzKNXEmueEc3YmJi6NWrF598\n8kmmx1lrmTdvHvHx8cFs3gUbO3Ys48aNSzMCl98MHDiQ8ePHM3/+fN9S9AXd6tWrWblyZYb5ZCIS\nPPqQK8GmPide3tS/+HhYtSol9S8mxj2bqnVrKJbPoxeNXEmunDhxwvcMqsjIyEzn6xRkBw4c4MCB\nAwBUrFiRcuXKhbhFIlIQ6feOiMj5nToFP/zgUv927HBl4eFuHpU39S+vBWrkSsGViIhIgBTE3ztK\n0ZJgU58run7/HRIS3HLqqVP/vKv+BfJv20oLFBERERGRAs1a+OUXN0q1enVK6l+dOm6UKjY2/6f+\nZUcjVyIiIgGi3zsiIs7Jkympf56ZJYSHu3lU3bu7eVXBpJErEREREREpUPbtS0n98z5JqFw5uOIK\nt/Jf2bIhbV6eU3AlIiIiPpr/IsGmPlf4WAvr16ek/nkH8OvVc6v+tWxZsFP/slNIb0tERERERILp\n5En4/nsXVO3c6cqKFXMP/O3WDWrXDm37giEs1A2Q8+vatSthYWGEhYXx448/+n3e5MmTiYqKCmDL\n8taZM2do2LAh8+fPz3T/li1bCAsLY/ny5UFu2fldd911DBw40Ld97Ngxbr75ZsqXL094eDhbt24l\nJiaGsLAwwsPD2bNnT55dOywsjJkzZ6bZDgsLC/h7H4r3Y9WqVURHR3PixInzHvv555/ToEEDIiIi\nuPvuu4PQOv/587NZp04dXnnllSC1KLQ2bdpE5cqVOXz4cKibkqlZs2bRqlWrDOXen+mwsDD2798f\ngpYFhkYQJNjU5wq+PXvgv/+F4cNh2jQXWJUvDzfeCC+8AHFxRSOwghAEV8aYq4wx640xScaY4dkc\n18YYc8YY0yen5xY2xhjuvvtudu3aRWxsrN/n3XbbbWzevDmALctbkydP5uKLL07zQNywsDC2bdsG\nQK1atdi1axfNmzfPs2uOGjUqTVCUW8YY34OVASZOnMiCBQtYtGgRO3fuJDo6GmMMI0eOZOfOnVSq\nVOmCr5mVXbt2MWHChIDVH0rNmzenZcuWvP766+c99p577uGWW25h27ZtvPrqq0FoXd5aunQpf/nL\nX0LdjKB45plnuP/++ymbKvH+p59+4oorrqB06dLUrFmTMWPGBOz6jzzyCG3atKFkyZLUyeRhKjfc\ncANnzpzJ8HDrZcuWZfkAdRGRws5aWLsW3ngDnnkGvv0Wjh+H+vXhvvtg7Fi45hooQH/nzxNBTQs0\nxoQDbwB/AnYAS4wxs6y16zI57kXg65yeW1iVLl2aypUr5+ickiVLUrJkyQC1KO+98cYbDB48OMv9\nYWFh530NTp48yeHDh/0OXlIHRHlp48aNNGrUiCZNmqQpj4qKyvH7mFOVK1dO8yG1sBkwYADDhg1j\n6NChWR5z4MAB9u/fT69evahWrVqur3Xq1CmKFy+e6/MvRMWKFfO8zlDeT1b27NnDjBkzWLt2ra/s\n8OHD9OzZk65du7J06VLWrVvHwIEDiYyM5LHHHsvzNlhriYuLY/Xq1XzzzTeZHtO/f3/efPNNbrnl\nFl9ZxYoVqVChQp63J9Q0/0WCTX2uYDlxAhYvdotU7NrlyiIiUlL/atUKafNCLtgjV22BjdbaLdba\n08B04MZMjhsMfAzszcW5RcaOHTu47bbbuOiii7jooou47rrr2Lhxo29/+tSjUaNG0bRpU6ZPn069\nevUoW7YsvXv35vfff/cd89NPP9GjRw/KlStHVFQULVq0ICEhAXD/+aVPf0mfGuY95uuvvyY2NpbS\npUvTpUs7UocbAAAgAElEQVQXduzYQXx8PM2aNSMqKoobbriBAwcO+OpZu3Ytq1ev5oYbbsjyfv1J\nQ9u1axc1a9bkpptuYubMmZw6dSrb1zA3SyQfO3aMuLg4oqKiqFq1KuPGjUuzv2vXrrz22mskJiYS\nFhZG9+7ds61v/fr13HDDDZQvX56oqCg6duzImjVrfO0bM2YM0dHRlCxZkmbNmjFr1qwctzm9jh07\n8vjjj6cpO3z4MKVKleKzzz4D4MMPP6RNmzaULVuWKlWq0LdvX5K9a6dmwp/+Ae69vvbaa3319uvX\nj927d/v2Z9cHAa655hp+++03Fi9enGU7vIFJ9+7dCQsLIzExEYCZM2fStGlTSpYsSa1atRg7dmya\nc2NiYhg9ejR33303FSpUoH///lne75QpU3x1Va1albi4ON++bdu20bt3b8qWLUvZsmXp06cPO7yP\nnPdTTEwM//jHP3zbYWFhvPvuu9xyyy2UKVOGevXqMXXq1GzriIuL4/rrr+fFF1+kZs2a1PL8xjvf\ne+t9L+Pj42nXrh2RkZG0adOGFStWpKl/4sSJ1KpVi8jISHr37s1bb71FWFjaXyv/+9//aNWqFaVK\nlaJu3br8/e9/5/Tp0779H3/8MfXr16devXq+sqlTp3LixAmmTJlC48aN6dOnD8OHDz9vmmRuXiOA\n1157jYceeogGDRpk+X/CDTfcQGJiIju9kwhERIqYPXvgo49c6t/06S6wqlABbroJxo2Du+5SYAXB\nD65qANtTbf/mKfMxxtTABU1veYq8v+nOe25RcuzYMbp160bp0qVJTEzk+++/p1q1avzpT3/i+PHj\nWZ63ZcsWZsyYweeff86cOXNYsWIFTz31lG9/v379qFGjBkuWLGHVqlWMHj06V6Nfo0aN4vXXX+eH\nH37gwIED9O3bl+eee47333+fhIQE1qxZw+jRo33HJyYmEh0dnWHEKacjS7Vr12bx4sXUqVOHBx98\nkOrVq/Pwww9nOVctfTqfPx5//HHmzp3LzJkz+fbbb1mxYoXvwzvAp59+ysCBA+nYsSO7du1KMx8q\nveTkZDp16kR4eDhz585l1apVPPLII5w9exaACRMmMH78eF5++WXWrFlD7969+fOf/8yqVaty1Ob0\n+vfvz/Tp09N8kPzkk08oXbo01157LQCnT59mzJgxrF69mi+++IJ9+/Zx++23X9B1d+7cSZcuXWjW\nrBlLlizh22+/5ciRI9x4Y8rfSc7XB0uXLk2TJk2ynJt3+eWX8/PPPwMumNq1axcdOnRg2bJl9O3b\nl5tvvpk1a9bwwgsvMG7cON54440057/yyis0btyYZcuWZQi+vN5++20eeOAB7rnnHtasWcPXX3/t\nS1c9d+4cN954I3v37iUhIYF58+aRnJzMTTfdlKPXKrO++eyzz9K7d29Wr17Nrbfeyt1338327duz\nqMGZP38+a9asYc6cOXz77beA/+/tk08+yUsvvcTy5cupWLEid9xxh2/f4sWLue+++xg8eDCrVq3i\n2muvZeTIkWnaPHv2bO68806GDBnC2rVrmThxIh9//DFPPvmk75jExETatGmT5rqLFy+mc+fOlChR\nwlfWq1cvkpOT2bp1a7b3m5vXyB8NGjSgfPnyWfa7wkQjCBJs6nP5l7WwZg28/jo8/TTEx7uRqwYN\nYNAgl/p39dVFL/UvO8FeLdCfYYIJwBPWWmvcb2nvb+qgPIVx0P8GBaTet69/O0/rmz59OuD+cuz1\nr3/9iypVqvDFF1+kSV1J7cyZM2lGtO6//34mTZrk279t2zaGDh3KJZdcAkDdunVz1b4xY8Zw+eWX\nA/DAAw8wePBgli9fTosWLQC46667+Pjjj33HJyUlUTuTmY7eICMnYmNjiY2NZfz48cyePZsPPviA\nbt26UatWLQYMGMCAAQOoUcPF5SNHjsxR3UeOHGHixIlMmjSJnj17AjBp0iRq1qzpO6ZChQqUKlWK\niIiI86YAvvnmm0RFRTFjxgyKedYkTf2ajx8/nqFDh3LbbbcBMHr0aBITExk/fjwffPBBjtqeWt++\nfXn00UeZN2+eb2Rt6tSp3HLLLURERACkmYsWExPDP//5Txo3bkxycjLVq1fP1XXfeustWrRokWa0\nb8qUKVSsWJGlS5fSunVrv/pgrVq12LBhQ6bXiIiI8AXpF110ke89eOWVV+jatavvPa9fvz5JSUm8\n+OKLPPzww77zu3btmmFUL70xY8bw17/+lUcffdRX5u3b3377LT/99BObNm3yjRRNmzaN+vXrEx8f\nf96RzOwMGDCAfv36+drw6quvsmDBAl9ZZkqVKsXEiRN97yv4/96OGTPGNwfymWeeoVOnTr5jXnvt\nNa688kpfemb9+vVZsmQJ7777ru/8559/nmHDhnHXXXcBbpGOF154gf79+/Pyyy8DLoX2mmuuSdPm\nXbt2+V47rypVqvj2ZfZ/xYW8Rv4wxhAdHU1SUtIF1SMiUhCcOAHffedS/7zJJRER0LatS/2Ljg5p\n8/K1YAdXO4DUb0c0bgQqtVbAdM9fPy8GrjbGnPbzXMClwsR4HvNcvnx5WrRoUej+KrJs2TI2b96c\nYcWx48ePs2nTpizPq127dppzqlWrlmbluscee4x7772XKVOm0KNHD/r06cOll16a4/Y1a9bM92/v\nh9umTZumKUt93cOHDxMZGZmjazRp0sS32EWXLl348ssv0+wPDw/nmmuu4ZprrmHfvn0MHDiQp556\niqSkpDRBaU78+uuvnDp1ig4dOvjKIiMj09xbTqxYsYJOnTr5AqvUDh8+zM6dO31BqlenTp34v//7\nv1xdz6tixYpcddVVTJ06le7du5OcnExCQgKjRo3yHbN8+XJGjx7NqlWr2L9/v2+Ua9u2bbkOrpYt\nW0ZiYmKGfmuM4ddff6V169Z+9cGoqCgOHTqUo2uvX7+e6667Lk3Z5ZdfzujRozly5AhlypTBGEPr\n1q2zrWfPnj0kJyfTo0ePTPevW7eO6tWrpwkO6tSpQ/Xq1Vm7du0FBVepf67Cw8OpVKnSeVeevOyy\ny9IEVuD/e5v6et65a3v27KF69er88ssvGdJ427Ztmya4WrZsGUuWLOGFF17wlZ07d44TJ06we/du\nqlSpwuHDhylTpkyaei5kLmR2r9HVV1/NwoULARdU/vTTTzmqu2zZsjnud6l501u9v4/y67a3LL+0\nR9uFfzt93wt1e4rydqNGXZk3D2bMSODUKahevSsXXQQVKiTQtClcfXX+am9OtleuXMnBgwcBl8kV\nKMEOrpYCDYwxMUAycCuQJhfFWuv7M7UxZhLwP2vtLGNMsfOd6zV58uRcNzCvR5gC5dy5c7Ro0YKP\nPvoow77sJlin/5BljOHcuXO+7ZEjR3LHHXfw1VdfMXv2bEaPHs2//vUvBg4c6JtLkTqVLPXciayu\n4/2gFB4enuV1y5Urx/r167Nsd2a+/vpr3/VLlSqVYb+1lkWLFvHhhx8yY8YMoqKiGDFiBPfcc0+O\nruOP3MzdAvc65PRca22eLMRx5513ct999/HPf/6T6dOnU6tWLTp16gTA0aNHufLKK+nVqxcffvgh\nlStXZu/evXTu3DnLeWz+9A9rLddddx3jx4/PcL43CM+uD3odPnw4VwuDZPVap349cxrk58SFvm/n\n+/nNTOnSpdNs5+S9zezn+HzXS81ay6hRozIdSb/44osB97N/5MiRNPuqVq3KLu8saQ/vvLyqVatm\ne83sXqP333/ft4x/+uP8cfjwYcqXL5/j87y8v+Tz+3b6DyWhbo+2ta3twG5bCxUruqDqP/9xZRdf\n3JVLLoHu3aF5cwgLyz/tze12+rIpU6YQCEENrqy1Z4wxDwOzgXDgfWvtOmPMIM/+LCObrM4NRrvz\no1atWjF9+nQqVqxIuXLl8rTu+vXrM3jwYAYPHsyDDz7Ie++9x8CBA32pVsnJyb4FA1auXJln10y/\nzPH5RGcxJr1hwwY+/PBDPvzwQ/bu3UufPn2YMWPGBY0YeNWrV4+IiAgWL17sGx09evQoa9asoUGD\nBjmur2XLlnz44YecPn06w4e9smXLUr16dRYuXEi3bt185QsXLsywCmFuXH/99QB88cUXTJ06NU3a\n1Pr16/n9998ZO3asLwXLu8hGVvzpH7Gxsfz3v/+lVq1amY7WeWXVB722bt2aYUTvfBo1asSiRYvS\nlC1cuJDo6OgcBVSVK1emRo0azJ07N9PRq0aNGvnmBnlfu02bNpGcnEzjxo1z1OZAyM17m5mGDRtm\nmMuYfjs2NpZ169Zlm15cv379DPOoOnTowPDhwzl58qRv3tU333xDjRo1sk0JPJ/cjriCCxS3b9+e\nq5/zgib9BxCRQFOfC43jx1NS/7xJEMWLp6T+pZrxIDkQFuwLWmu/stZeaq2tb60d5yl7O7PAylo7\n0Fo7M7tzi6o77riDKlWqcOONN5KYmMjmzZtJTEzk8ccfT7NiYE4cP36chx56iPnz57NlyxZ++OGH\nNB/k69evT3R0NKNGjSIpKYk5c+bw3HPP5cn9dO7cme3bt7N3797zH5yNbdu20bhxY7777jtGjRrF\n7t27mTx5cp4EVgBlypThnnvuYfjw4cydO5eff/6Zu+++O0d/zU/twQcf5MiRI/Tt25elS5eyceNG\n/vOf//gWrBg6dCjjx49n+vTpbNiwgWeeeYaFCxeed05Qep9++ikNGzZMsyJcyZIl6dOnD2PGjGHF\nihXceeedvn21atWiRIkSvP7662zatIkvv/ySp59+Ottr+NM/HnroIQ4dOsStt97Kjz/+yKZNm5g7\ndy6DBg3iyJEjnDhxIts+CG4xl7Vr19KlS5ccvQZ/+9vfmD9/PqNHj2bDhg1MnTqVV155hWHDhuWo\nHoCnnnqKCRMmMGHCBDZs2MDKlSt9K9n17NmTZs2acccdd7Bs2TKWLl3KHXfcQatWrdIEyaGSm/c2\nM0OGDGHOnDmMHz+epKQk3n//fT777LM0o3PPPPMM06ZNY+TIkaxZs4b169fz8ccfM3x4ymMKO3fu\nzJIlS9LU3a9fP0qXLk1cXBw///wzM2fO5MUXXwzIMuzg5n2tXLmS5ORkTp06xapVq1i5cmWakdcN\nGzZw8OBBOnfuHJA2iIgEy86dboRq+HD34N89e6BiRejTxz3wt39/BVYXIujBleSNUqVKkZiYSN26\ndbnlllto1KgRcXFxHDx4kIsuush3XOoPOlmtjOctK1asGAcPHiQuLo6GDRvy5z//mY4dO/o+NEZE\nRDB9+nQ2bdpE8+bNGT16NOPGjctQZ3bXyKotTZo0oWnTpnz++efZ3vf50qoqVarEli1bmDt3LgMG\nDMiQEnU+kydPTvPg4syMHz+ebt260bt3b3r06EGzZs0yfND3dxXC6tWrk5iYyKlTp+jWrRuxsbG8\n+eabvlGsIUOGMHToUIYNG+Z7fbzLiefEoUOHSEpK4syZM2nK77zzTlavXk1sbCwNGzb0lVeqVIkp\nU6bw2Wef0aRJE8aMGcP/+3//L9v32p/+Ua1aNRYtWkRYWBhXXXUVl112GQ8//DAlS5akRIkShIeH\nZ9sHAb788kuio6PTzHvLTPq2tmzZkhkzZvDJJ5/QtGlTnnzySUaMGMFDDz3k/wvp8cADD/Dmm2/y\n7rvv0rRpU66++uo0z2n6/PPPqVSpEt26daN79+5Ur17dt8R9Vu0LhMz6YW7e28zK2rdvz7vvvstr\nr71G8+bN+fzzzxk2bFiGFf6+/PJL5s2bR7t27WjXrh0vvfRSmtGnPn368Ouvv6b5o1DZsmX55ptv\nSE5OpnXr1gwePJjHH3+cv/71r75jvMv8//vf/879C+Rx3333ERsby4QJE9i1axctW7akVatWaZZd\nnzVrFl26dLmg0a+CIvX8F5FgUJ8LvHPnYPVqePVVGDXKjVadPAmXXgp/+Qs89xz06gUBzIwvMkxu\n54rkV8YYm928ioJ4v127dqVp06a8/vrroW5KQL333ntMmjQpQ+pWMI0cOZKZM2eyatWqDM/ruVB1\n6tTh4Ycf5m9/+1ue1puZyZMnM3jwYP7444+AXyvYrr/+erp06ZLtQ4QlNP76178SHx+f40cF3Hnn\nndSuXZvnn3/e73PmzZvHtddey9q1a30puoFiraV58+Y8/fTTGeaPJSQk0L17d/bt25fmD1teBfH3\nToIe6CpBpj4XOMeOpaT+eZODiheHdu1c6l+NIvtQI9//z3n+l04FVwVAt27d+O677yhevDgJCQm0\natUq1E0KiLNnz9KkSRPefvtt3/LPwda2bVvGjx+f45Qzf9SpU4edO3cSERHB5s2bfRP681qZMmU4\ne/YsERERHD58OCDXCJXVq1dz7bXXkpSUlKvnr0neevnll+nZsydlypRh7ty5PPbYY4wbN45HHnkk\nR/Vs2rSJ9u3bs3HjRsqWLevXOcOGDaNMmTI888wzuWl6jsyaNYvRo0ezbNmyNOVNmjRh8+bNnDx5\nkr179xaa4EpECr6dO2HePFi8GLxrFV18MXTtCh07aoQKFFz5rTAGV8nJyb4VrmrWrEnx4sVD3CLJ\njW3btvnS8mJiYvJ8ZMzLuxR/WFhYwP+iL0XbbbfdRkJCAocOHaJu3boMGjSIIUOGhLpZQbN9+3bf\nvKw6depkmUpZEH/viEjBc+4c/PSTC6rWpVryrVEjN0rVtCkE6KNHgaTgyk+FMbgSEZGCqSD+3lGK\nlgSb+tyFOXYMFi1yqX/79rmy4sWhQwc3UlUEpormSqCCq2A/50pERERERC5QcjLEx8MPP6Sk/lWq\nlJL6l8M1vSSPaORKREQkQPR7R0Ty0rlzsGqVS/375ZeU8saNXerfZZcp9c9fGrkSERERESmCjh6F\nhQth/nz4/XdXVqJESupftWohbZ6kouBKREREfDT/RYJNfS5rv/3mRql+/DFt6l/37i6wKlUqtO2T\njIpccBWMB3eKiIiIiOSGN/UvPh42bEgpb9LEBVVNmoA+zuZfRWrOlYiIiIhIfnTkSErq3/79rqxk\nSTdC1a0bVKkS2vYVNppzJSIiIiJSyGzfnpL653l0HlWqpKz6V7JkSJsnOaTgSsQPygeXUFC/k1BQ\nv5NgK4p97tw5WLHCBVVJSSnll13mUv8aN1bqX0Gl4EpEREREJAj++CMl9e/AAVdWsqQboeraVal/\nhYHmXImIiIiIBNC2bW6UasmSlNS/qlXdXKr27ZX6FwqacyUiIiIiUkCcPZuS+rdxoyszBpo2dal/\njRop9a8wUnAl4oeimA8uoad+J6GgfifBVtj63B9/wIIFLvXv4EFXVqpUSupf5cohbZ4EmIIrERER\nEZELtHVrSurfmTOurFq1lNS/EiVC2z4JDs25EhERERHJhTNnUlL/fv3VlRkDzZq5oKphQ6X+5Vea\ncyUiIiIikg8cPpyS+nfokCsrVQo6dXKpfxdfHNLmSQiFhboBIgVBQkJCqJsgRZD6nYSC+p0EW0Hq\nc1u2wMSJMGIEzJrlAqvq1eGOO+DFF+HmmxVYFXUauRIRERERycKZM7B8OcTHw+bNrswYaNHCpf5d\neqlS/ySF5lyJiIiIiKRz6BAkJrr0P2/qX+nSLvXviis0QlXQac6ViIiIiEiAbd7sRqmWLXPPqgKo\nUcONUrVtq1X/JHsKrkT8UNiewSEFg/qdhIL6nQRbfuhzZ87A0qVu1b8tW1xZWBi0bOke+NuggVL/\nxD8KrkRERESkSDp4MCX17/BhVxYZmZL6V7FiaNsnBY/mXImIiIhIkWEtbNrkRqmWL09J/atZ041S\ntWkDxYuHto0SeJpzJSIiIiKSS6dPp6T+bd3qysLCIDbWBVX16yv1Ty6cgisRP+SHfHApetTvJBTU\n7yTYAt3nDh50D/tdsAD++MOVlSkDnTu71L8KFQJ2aSmCFFyJiIiISKHiTf2Lj4cVK1JS/6KjU1L/\nIiJC20YpnII+58oYcxUwAQgH3rPWvphu/43As8A5z9dQa228Z98W4DBwFjhtrW2bSf2acyUiIiJS\nBJ0+DUuWuNS/bdtcWepV/+rVU+qfOIGacxXU4MoYEw78AvwJ2AEsAW631q5LdUyktfao599NgU+t\ntfU925uBVtba/dlcQ8GViIiISBFy4EBK6t+RI65MqX+SnUAFV2F5XeF5tAU2Wmu3WGtPA9OBG1Mf\n4A2sPMoA+9LVob83SNAlJCSEuglSBKnfSSio30mw5bbPWQtJSfDOO/Dkk/DVVy6wqlUL4uLghRfg\nppsUWElwBXvOVQ1ge6rt34B26Q8yxtwEjAOqAb1S7bLAXGPMWeBta+27AWyriIiIiOQzp0/Djz+6\n1L/tnk+V4eHQurVL/atbV6l/EjrBTgvsA1xlrb3Ps30n0M5aOziL4zvj5mVd6tmuZq3daYypBHwD\nDLbWLkh3jtICRURERAqZ/ftTUv+OevKcoqKgSxf3Vb58aNsnBUthec7VDiA61XY0bvQqU9baBcaY\nYsaYitba3621Oz3le40xn+LSDBekPy8uLo6YmBgAypcvT4sWLXxLfHqHnrWtbW1rW9va1ra2tZ2/\nt6+4oitJSfCvfyWwcSNUq+b2nzuXQMuWcP/9XSlWLP+0V9v5d3vlypUcPHgQgC1bthAowR65KoZb\n0KIHkAz8SMYFLeoBm6y11hgTC8yw1tYzxpQGwq21fxhjIoE5wGhr7Zx019DIleS5hIQE3w+oSLCo\n30koqN9JsGXW506dcql/8fGwY4crCw+HVq2gWzeoU0epf3JhCsXIlbX2jDHmYWA2bin2962164wx\ngzz73wb6AAOMMaeBI8BtntOrAjON+0kqBkxNH1iJiIiISMH1+++QkACLFqWk/pUtm5L6V65cSJsn\ncl5Bf85VoGnkSkRERKTgsBY2bHCjVKtXw7lzrrxOHbdARWwsFAv2RBYp9ArFyJWIiIiICMDJk/DD\nD26kKnXqX7t2LqjyTJ8XKVAUXIn4QXMQJBTU7yQU1O8k0PbtS0n9O3YMkpMTaNSoK1dc4R76W7Zs\nqFsoknsKrkREREQkoKyFX35JSf3zzuCoWxdatID77lPqnxQOmnMlIiIiIgFx8iR8/70bqUpOdmXF\nirkH/nbrptQ/CR3NuRIRERGRAmHv3pTUv+PHXVm5cij1Two9BVciftAcBAkF9TsJBfU7yS1rYd06\nmDcPfvopJfWvXj23QEXLlm7BivTU56QwUXAlIiIiIrl24oRL/Zs3D3btcmXFikGbNi71r3bt0LZP\nJJg050pEREREcmzPHpf69913Kal/FSq41L9OnSAqKqTNE8mW5lyJiIiISEhZC2vXpqT+edWv71L/\nWrTIPPVPpKhQcCXiB+WDSyio30koqN9JZk6cgMWLXVC1e7cri4iAtm1d6l90dO7rVp+TwkTBlYiI\niIhkavfulNS/EydcWYUK0LWrS/0rUyaUrRPJfzTnSkRERER8rIWff3ajVGvWpJRfcokbpWrRAsLC\nQtc+kbygOVciIiIiEjDHj7vUv4SEtKl/7dq5oKpmzZA2T6RA0N8dRPyQkJAQ6iZIEaR+J6Ggflf0\n7NoF//kPPPEEfPSRC6wuugj+/Gd48UXo3z+wgZX6nBQmGrkSERERKWKsdav9zZvnVv/zuvRSN0rV\nvLlS/0RyQ3OuRERERIqIY8fc4hQJCbB3rysrXjwl9a9GjZA2TyRoNOdKRERERHJl5043SvX993Dy\npCurWNEFVB07QmRkaNsnUlgouBLxg57BIaGgfiehoH5XeJw7l5L6t25dSnnDhu6Bv02b5o/UP/U5\nKUwUXImIiIgUIseOwaJFLvVv3z5XVrw4tG/vRqqqVw9p80QKNc25EhERESkEkpNTUv9OnXJlF1+c\nkvpXunRo2yeSn2jOlYiIiIik4U39i4+H9etTyhs1cql/l12WP1L/RIoKBVciflA+uISC+p2Egvpd\nwXD0qEv9mz8/JfWvRImU1L9q1ULbvpxQn5PCRMGViIiISAGxY4dL/fvhh5TUv0qVoGtXpf6J5Aea\ncyUiIiKSj507B6tWuaDql19Syhs3Tkn9M3k+c0SkcNOcKxEREZEi5OhRWLjQrfq3f78rK1kyJfWv\natWQNk9EMqEpjiJ+SEhICHUTpAhSv5NQUL8Lvd9+gw8+gOHDYeZMF1hVrgy33govvAC33164Aiv1\nOSlMNHIlIiIiEmLnzsHKlS71b8OGlPLLLnOjVE2aKPVPpCDQnCsRERGREDlyxKX+zZ+fNvWvY0e3\nSEWVKiFtnkihpTlXIiIiIoXE9u3u2VRLlsDp066sShU3StWhgwuwRKTgUXAl4gc9g0NCQf1OQkH9\nLnDOnnWpf/HxsHFjSnnTpi6oaty4aKb+qc9JYaLgSkRERCSA/vgDFiyAxEQ4cMCVlSqVkvpXuXJI\nmycieSjoc66MMVcBE4Bw4D1r7Yvp9t8IPAuc83wNtdbG+3Ou5xjNuRIREZGQ27rVLVCxdGlK6l/V\nqu7ZVO3aKfVPJJQCNecqqMGVMSYc+AX4E7ADWALcbq1dl+qYSGvtUc+/mwKfWmvr+3Ou5xwFVyIi\nIhISZ8/C8uUuqPr1V1dmjEv9694dGjYsmql/IvlNoIKrYD/nqi2w0Vq7xVp7GpgO3Jj6AG9g5VEG\n2OfvuSKBomdwSCio30koqN/lzuHD8OWX8OST8N57LrAqVQr+9CcYMwYeeggaNVJglRn1OSlM/Jpz\nZYy5HKhgrf3Cs10ReBNoAswBhllrz/pRVQ1ge6rt34B2mVzvJmAcUA3olZNzRURERIJly5aU1L8z\nZ1xZtWpugYr27aFEiZA2T0SCzN8FLV4A5gJfeLZfBq4GvgUeAA7h5kmdj1/5etbaz4DPjDGdgQ+M\nMQ39bKdIQGgVIwkF9TsJBfW78ztzJiX1b9MmV2YMNG/uUv8uvVQjVDmhPieFib/BVUPgRQBjTHHg\nZuCv1tr3jTGPAoPwL7jaAUSn2o7GjUBlylq7wBhTDLjIc5xf58bFxRETEwNA+fLladGihe8H1zv0\nrO5VtWUAACAASURBVG1ta1vb2ta2trWdk+2jR8HariQmwrp1bn/9+l25/HKIiEigXDlo2DD/tFfb\n2tZ2yvbKlSs5ePAgAFu2bCFQ/FrQwhhzHOjlCXY6AYlAVWvtHmPMFcBX1trSftRTDLcoRQ8gGfiR\njAta1AM2WWutMSYWmGGtrefPuZ7ztaCF5LmEhATfD6hIsKjfSSio32W0ZYt7NtXSpW7BCoDq1d0o\nVdu2Sv27UOpzEgqBWtDC35GrZKAFsAC4Clhjrd3j2VcBOOZPJdbaM8aYh4HZuOXU37fWrjPGDPLs\nfxvoAwwwxpwGjgC3ZXeun+0XERER8Zs39S8+HjZvdmVhYdCihQuqLrlEqX8ikpG/I1djgEdxgc21\nwEhr7UuefaNxo1odAtlQf2nkSkRERHLr0CH3sN/ERLcCIEBkJHTqBFdcARUrhrZ9IpI3Qj1yNRo4\nAXTAreL3Sqp9LYAZedwuERERkaCw1o1OzZsHy5alpP7VqJGS+le8eGjbKCIFQ1AfIhwMGrmSQFA+\nuISC+p2EQlHqd2fOuHlU8+a5eVWQkvrXrRs0aKDUv2AoSn1O8o9Qj1yJiIiIFAoHD7q0vwUL0qb+\nde7sUv8uuii07RORgivLkStjzGbcc6m8EV1Ww0EGsNbaunnfvJzTyJWIiIikZ617JlV8PKxYkZL6\nFx3tRqnatoWIiNC2UUSCJxQjV/PTbXcHqgCLgD2ef18O7MI9TFhEREQkXzl92qX+xcfDtm2uLCwM\nWrVyQVX9+kr9E5G8k2VwZa2N8/7bGHM/0BboaK39LVV5NG4Fwe8C2EaRkFM+uISC+p2EQmHpdwcO\nwPz5LvXvyBFXVqZMSupfhQqhbZ+kKCx9TgT8n3M1DHgydWAFYK3dbowZBYwF3s3jtomIiIj4zVrY\nuNEtULFiBZw758pr1XKr/rVurdQ/EQksf59zdRy41Vo7K5N9NwIfWWtLBqB9OaY5VyIiIkXL6dOw\nZIlL/du+3ZWFhUFsrAuq6tZV6p+IpBWoOVf+BlfLgaO4hwUfT1VeGpgDlLbWxuZ143JDwZWIiEjR\ncOAAJCTAwoUpqX9RUdCli/sqXz6kzRORfCzUS7EPBf4P2GqM+T9gN1AVuAYo6/kuUmgpH1xCQf1O\nQiG/9ztv6l98PKxcmZL6V7u2G6Vq1UqpfwVNfu9zIjnhV3Blrf3WGNMC+DvQBRdY7cQtZvGctXZ9\n4JooIiIiRd2pU/Djj24+1W+eGeDh4W4J9W7doE4dpf6JSOj5lRZYkCgtUEREpPD4/Xe36t/ChXD0\nqCsrW9al/XXurNQ/EcmdUKcFioiIiASFtZCU5FL/Vq1KSf2LiUlJ/SumTzAikg/5/V+TMaYrcDsQ\nDaReGdAA1lrbPW+bJpJ/KB9cQkH9TkIhlP3u1Cn44QeX+rdjhyvzpv517+5S/6Tw0f91Upj4FVwZ\nYwYBbwH7gQ3AqUA2SkRERIqO3393q/4tWpQx9a9LFyhXLqTNExHxm79LsW8AlgADrbX5OrDSnCsR\nEZH8z1r45Rc3SrV6dUrqX506bpQqNlapfyISOKGec1UD+Et+D6xEREQkfzt5MiX1LznZlYWHQ7t2\nLqiKiQlp80RELkiYn8ctB+oGsiEi+VlCQkKomyBFkPqdhEKg+t2+ffDxx/DEEzB1qgusypWDG26A\nF16Au+9WYFVU6f86KUz8HbkaDEwzxmyw1s4PZINERESkcLAW1q9PSf3zZu3Xq+eeTdWypVL/RKRw\n8XfO1XagLBAFHAUO4FklkJTVAmsFsJ1+05wrERGR0Dp5Er7/3gVVO3e6smLFoE0bF1TVrh3a9omI\nhHrO1bfn2a9oRkREpIjbs8et+vfdd3D8uCsrXx6uuMI98DcqKqTNExEJOL9GrgoSjVxJIOgZHBIK\n6ncSCjntd9bC/2fvzuPkvso7339OVe97t9RSq7tLkiXZwrtsjCQvwsJwiYHcwJ3kZsINISYMMAlk\nkrnzmgxM7gRCMrkh92YguUyIA4QkhIRJcglhDYZIbcu2bNnY8iovki2pN7WW3vfqqmf+ONVdXfpp\nKUlV9auq/r5fL73cv1Onuo/sY6mffp7znEOH/IW/zz+fLv3bssU3qNi2zTesEDkf/VknYQg7cyUi\nIiKyZHYW9u/3maoTJ/xYZWW69G99URwWEBEprKwzV865m4BPAncDrfgLhXuAT5vZc/la4KVS5kpE\nRCR/Tp70Z6kefdQHWACtrb707667VPonIqUhX5mrbBtavAl4EJgBvgUMAR3A/wrUAHeb2ZO5Xtzl\nUHAlIiKSW2bwwgs+qHr++fT41VenS/8i2V7uIiJSBMIOrn6E7xb4VjObWDbeCPwIGDez/yXXi7sc\nCq4kH1QPLmHQvpMwLN93s7M+Q9XTA0ND/vXKSti+3Zf+xWKhLVPKiP6skzCEfeZqJ/D+5YEVgJlN\nOOc+A/xVrhcmIiIi4Rga8lmq/fvTpX9tbbB7ty/9q68PdXkiIkUr28zVBPCLZvaNc7z2r4C/NLOi\nqLJW5kpEROTSLZb+7dnj/7nommt86d/NN6v0T0TKRzGUBTbjywLHl4034O/AUlmgiIhICZqZ8Rmq\nvXt9swqAqqp06V93d7jrExHJh7CDq+2kG1p8BxgE1gHvBOqA3WZ2INeLuxwKriQfVA8uYdC+k3wa\nHPRnqfbvh7k5P7ZqFbS29vArv7JbpX9SMPqzTsIQ6pkrMzvgnNsB/BZwL+lW7HuA3ymmVuwiIiJy\nbsmk7/a3dy+8+GJ6fOtWX/p3003w0EM6UyUicrmyvucqZ1/QuXuBzwFR4Etm9pmzXv954DcAB0wA\nv2xmz6ZeOwqMAwkgbmbbz/H5lbkSERFZZno63fXv1Ck/VlUFO3f60r/OzlCXJyJScKFmrpxza4BW\nM3v5HK9tBYbN7FQWnycKfB54G9APPOGc+5aZHVo27TXgzWY2lgrE/gzfrRDA8CWIw9msW0REZCUb\nHEx3/Zuf92OrV/uuf3feCXV1oS5PRKTsZNuK/U+AM8BHzvHarwOrgJ/N4vNsBw6b2VEA59zXgXcD\nS8GVme1fNv9x4OyjtDmPMEUuRvXgEgbtO7kcySQ895wPqg4t+9Hltdf6LNWNN16465/2nRSa9pyU\nk2yDqzuBj53ntQeA/57l5+kCepc99wE7LjD/g8D3lj0b8CPnXAK438y+mOXXFRERKWvT0/DII770\n7/RpP1ZVBbff7jNVKv0TEcm/bIOrVmD0PK9N4DNX2cj6MJRz7i3AL+EDu0V3mtmgc64d+KFz7iUz\n25ft5xS5XPqJmoRB+06yMTDg76Z6/PF06V97uw+o7rjj0kv/tO+k0LTnpJxkG1z14889/cs5XtuO\nb82e7eeJLXuO4bNXGZxzNwFfBO41s5HFcTMbTP3zlHPuH1NfOxBc3XfffWzcuBGAlpYWtm3btvQ/\nbk9PD4Ce9axnPetZzyX7nEzCn/95D089BYmEf31goIcNG+DDH97NDTfAQw/1cOBAcaxXz3rWs57D\nfj548CCjoz5XdPToUfIl23uufh9fFvhzZvadZeM/Cfwt8AUz+40sPk8F8DLwVmAAOAC8d3lDC+fc\nenyL9/eZ2WPLxuuAqJlNOOfq8eWIv21mD5z1NdQtUHKup6dn6X9QkULRvpOzTU2lS//OnPFj1dXp\n0r916678a2jfSaFpz0kYQu0WCPwO8GbgW865QXwGqhvoAPYDv53NJzGzBefcx4Af4Fuxf9nMDjnn\nPpJ6/X78XVqtwBecc5Buud4BfCM1VgF87ezASkREpBz19fkGFQcOZJb+3XOPD6xqa8Ndn4iIeFnf\nc+WcqwLeB7wdf8bqND5I+mszW8jbCi+RMlciIlIOkkl45hl/nuqVV9Lj11/vg6rrrwen/rkiIpcl\nX5mrgl8inG8KrkREpJRNTsLDD8ODD8Jw6lbHmhrfnGL3bli7NtTliYiUhbDLAhcXcTOwC5+5ut/M\nTjjnrgaGzGw814sTKRaqB5cwaN+tLL296dK/eNyPrV3r76a6/XYfYBWC9p0UmvaclJOsgivnXDXw\nNeBfpYYM+DZwAvgM8Arw8XwsUEREpFwlk/D00z6oevXV9PgNN/jSv+uuU+mfiEgpybZb4P+Lv9D3\no8APgSHgNjN7yjn3IeCjZrYtryvNksoCRUSk2E1MpEv/RlIXjqj0T0SkcMIuC3wv8F/M7G9S7dSX\nOwpszOWiREREytHx4z5L9cQT6dK/jg5f+rdzZ+FK/0REJD+yDa5WAS+e57UIUJ2b5YgUJ9WDSxi0\n78pDIgEHD/quf4cP+zHn4MYbfenftdcWV+mf9p0UmvaclJNsg6ujwB34y33P9ib8xcAiIiKSMjEB\n+/bBQw+lS/9qa9Olf2vWhLo8ERHJg2zPXH0C+E3gI8A3gCngNqAF+AfgU2b2x3lcZ9Z05kpERMJ0\n7Fi69G8hdQvkunXp0r9q1XqIiIQu1HuuUues/hr4WWAeqAJmgRrgb4H3FUtEo+BKREQKLZGAp57y\nQdWRI37MObjpJh9UveENxVX6JyKy0hXFJcLOuV3AvcAa4Azwz2bWk+tFXQkFV5IPqgeXMGjfFb/x\ncV/69+CDMDbmx2pr4a67fOnf6tWhLu+yaN9JoWnPSRjC7hYIgJntA/blehEiIiKl5OhR36Dixz9O\nl/51dvos1Y4dKv0TEVmpsi0L3Aq0mNnjqeda4JPA9cADZvb/5XWVl0CZKxERyYeFBV/6t2cPvP66\nH4tE0qV/W7eq9E9EpFSEnbn6PPA08Hjq+b8CHwOeBz6bCmg+n+vFiYiIhG1szHf827cvXfpXV5cu\n/Vu1KtTliYhIEYlkOe8m4FEA51wUeD/wcTO7Ffgd4EP5WZ5Icejp6Ql7CbICad+F6/XX4ctfhk98\nAr7zHR9YdXXB+94Hv//78NM/XZ6BlfadFJr2nJSTbDNXzcDp1Me3AG3A36eeHwT+Y47XJSIiUnAL\nC/Dkk77r39GjfiwSgVtu8Rf+Xn21Sv9EROT8sj1zdQx/l9VXUnde/ZKZXZ167SeBvzazlvwuNTs6\ncyUiIpdqdDRd+jc+7sfq633p3913l2eGSkRkJQv7zNW3gP/bOXc98AHg/mWv3QC8luuFiYiI5JOZ\nL/3bs8c3qkgk/Hh3t89SvelNUFUV7hpFRKS0ZBtcfQJ/YfBPAP+Eb2ix6N3AAzlel0hR0R0cEgbt\nu/xYLP3bsweOHfNjkQjceqsPqrZsWdmlf9p3Umjac1JOsgquzGyS8zStMLPbc7oiERGRPBgd9Zf9\n7tsHExN+rKEBdu3ypX+treGuT0RESl9WZ65Kic5ciYjIIjN47TWfpXr66XTpXyyWLv2rrAx3jSIi\nUnhhn7kSEREpGfE4PPGE7/p3/Lgfi0Tgttv8hb+bN6/s0j8REcmPbO+5ElnRdAeHhEH77tKNjMA3\nvwkf/zj85V/6wKqhAd75Tvi934MPfUhnqi5G+04KTXtOyokyVyIiUtLM4PBhn6V6+mlIJv34+vW+\n9O+221T6JyIihaEzVyIiUpLicThwwAdVvb1+LBpNX/i7aZMyVCIicm46cyUiIgIMD6e7/k1N+bHG\nRnjzm/2vlqK40l5ERFairIMr59xu4L1ADH/n1dJLgJnZPbldmkjx0B0cEgbtuzQzePVVn6U6eDBd\n+rdxo29QcdttUKEfF+aE9p0UmvaclJOs/ipyzn0E+AIwDLwCzOdzUSIiIgDz8770b88e6O/3Y9Eo\nbN/ug6qrrlLpn4iIFI+szlw5514BngA+YGZFHVjpzJWISOk7cwZ6euCRR9Klf01N6dK/5uZQlyci\nIiUu7DNXXcAvF3tgJSIipcsMXnnFZ6mefTZd+nfVVb5Bxa23qvRPRESKW7b3XD0FbMrnQkSKme7g\nkDCslH03NwcPPQS/8zvw3/6bP1PlHOzYAZ/4hL+zavt2BVaFslL2nRQP7TkpJ9n+VfWrwN84514x\nswev5As65+4FPgdEgS+Z2WfOev3ngd/AN8qYwGfMns3mvSIiUjpOn06X/k1P+7HmZrj7bti1y5cB\nioiIlJJsz1z1Ak1AIzAFjJDqEki6W+D6LD5PFHgZeBvQjz/H9V4zO7Rszu3Ai2Y2lgqmPmVmO7N5\nb+r9OnMlIlKkzODll9Olf4t/XG/a5Ev/brlFGSoREcm/sM9c/ctFXs82mtkOHDazowDOua8D7waW\nAiQz279s/uNAd7bvFRGR4jQ3B4895jNVAwN+rKLCt1B/y1t8S3UREZFSl1VwZWb35ejrdQG9y577\ngB0XmP9B4HuX+V6RnNEdHBKGcth3p06lS/9mZvyYSv+KWznsOykt2nNSTgpdfJF1vZ5z7i3ALwF3\nXup7RUQkPGbw0ku+9O+559Klf5s3p0v/otFw1ygiIpIP5w2unHPvB75rZmecc7/IRYIbM/urLL5e\nPxBb9hzDZ6DO/to3AV8E7jWzkUt5L8B9993HxlSNSUtLC9u2bVv6ichiRxo961nPei7258WxYlnP\nxZ4feKCHF1+EkZHdnDgBAwM9RKPwnvfs5i1vgddf72FyEqLR4livnvWs5+J43r17d1GtR8/l+Xzw\n4EFGR0cBeOXIK+TLeRtaOOeSwE4zO5D6+ILMLHLRL+ZcBb4pxVuBAeAAwYYW64E9wPvM7LFLeW9q\nnhpaiIgU0MmT0NMDjz6aLv1rbfWlf3fdBY2NoS5PRERWKDNjdHaU42PHM36Nzo7yZz/1ZwVvaLEJ\nH8QsfnzFzGzBOfcx4Af4dupfNrNDzrmPpF6/H/gtoBX4gnMOIG5m28/33lysS+RiepZlD0QKpZj3\nnRm8+CLs3QvPP58u/duyxZf+bdum0r9SVcz7TsqT9pzkgplxZuZMIJCamJsIzK2pqMnbOs4bXC12\n5Tv74ytlZt8Hvn/W2P3LPv43wL/J9r0iIlI4s7Owf78PqoaG/Fhlpb/k9y1vgVjswu8XERG5UmbG\nyamTgUBqOj4dmFtfVc/65vUZv9rr2vlj/jgva8vqnqtSorJAEZHcGxpKl/7Nzvqx1lbYvduX/jU0\nhLk6EREpV0lLcmLyREYQ1TvWy+zCbGBuU3VTIJBqq20jVQ2XIex7rkREZIUxgxdeSJf+LbrmGp+l\n2rYNIhc9bSsiIpKdheQCgxODGYFU33gf84n5wNzW2tZAINVc3XzOQKqQFFyJZEH14BKGsPbdzIwv\n/evpySz927HDB1Xd3Rd8u5Q4/XknhaY9tzLFE3H6J/ozAqn+8X4WkguBuavrVmcEUbHmGE3VxXlR\nooIrEREB4MQJn6V67LF06V9bW7r0r74+1OWJiEiJmluYo2+8LyOQGpgYIGnBhuRrG9ZmBlJNMeqr\nSucvIJ25EhFZwcz8Rb979/ruf4u2bvVZqptvVumfiIhkbyY+Q+94b0YgdWLyBGd/fx5xEToaOgIZ\nqXx28luuKM5cOefagZ1AG/Cd1AXDtcC8mSVyvTgREcmPmRnfnGLvXjh1yo9VVaVL/7q6wl2fiIgU\nv6n5qUDHvpNTJwPzopEonU2dxJpibGjZwPrm9XQ1dlFdUR3CqvMrq+DK+ZNh/w/wq0AlYMCbgDPA\nN4FHgE/naY0ioVM9uIQhH/tucDBd+jc358dWrfIB1R13qPRP9OedFJ72XGkYnxsPBFJnps8E5lVE\nKuhu6s7ISHU2dlIZrQxh1YWXbebqE8BHgd8Gfgg8vuy1bwO/gIIrEZGilEz6bn979sChZVevv+EN\n/sLfG29U6Z+IiHhmxujsaCCQGp0dDcytilYRa45lZKTWNawjGlm5t8hndebKOfca8CUz+z3nXAUw\nD9xmZk85594B/LWZrcrzWrOiM1ciIt70dLr07/RpP1ZVBTt3+kxVZ2e46xMRkXCZGWdmzgQCqYm5\nicDcmoqaQOvztQ1ribjS/Olc2GeuuoD953ltHlAhiYhIkRgYSJf+zaeuBlm9Ol36V1cX7vpERKTw\nzIyTUycDgdR0fDowt76qfqlT32JGqr2uPfQ7pEpBtsHVAHAjsPccr90EvJ6zFYkUIdWDSxguZd8l\nk77r35498NJL6fFrr/WlfzfcoNI/yY7+vJNC057LvaQlOTF5IiOI6h3rZXZhNjC3sbqRDc0bMjJS\nbbVtCqQuU7bB1d8Bv+Wce4plGSzn3FbgPwBfzMPaRETkIqam4JFH4MEH06V/1dVw++3+fqp160Jd\nnoiI5NlCcoHBicGMQKpvvI/5xHxgbmttayAj1VzdrEAqh7I9c1UH/AC4EzgGbMBnq2LAo8BPmNlc\nHteZNZ25EpGVoL/fl/49/ni69K+93Zf+3X67Sv9ERMpRPBGnf6I/I5DqH+9nIbkQmLu6bnXgDqmm\n6qYQVl2c8nXmKutLhFONLN4L3AusAU4D/wx8zcyC/0VDouBKRMpVMgnPPOODqpdfTo9fd1269E8/\nfBQRKQ9zC3P0jfdlBFIDEwMkLRmYu7ZhLbGmGOub17OhZQOxphj1VWqJcCGhB1elQsGV5IPqwSUM\ni/tuagoefhh6emB42L9WU5Pu+tfREeoypczozzspNO05mInP0DvemxFInZg8wdnf00ZchI6GDmLN\nqUCqeQOx5hg1FTUhrbx0hd0t8OzFBI5Fm50jjBYRkct26hR89au+9C8e92Nr1qRL/2prw12fiIhc\nuqn5qUDHvpNTJwPzopEonU2dGRmprsYuqiuqQ1i1ZOtSzlx9EvjfgW6CQZmZWVHcFqbMlYiUsmQS\nDh70pX+vvJIev+EGH1Rdf71K/0RESsX43HggkDozfSYwryJSQXdTd0ZGqrOxk8poZQirXhnCzlz9\nd+DngW8DX8ffbbWcohkRkSswOelL/x58MLP07447fNe/tWtDXZ6IiFyAmTE6OxoIpEZnRwNzq6JV\nxJpjGRmpdQ3riEaKIk8hVyjbzNUZ4NNm9kf5X9KVUeZK8kH14JIvvb3+bqonnkiX/q1d67NU8XgP\nb3/77lDXJyuP/ryTQiu1PWdmnJk5EwikJuYmAnNrKmqWOvUtZqTWNqwlEjxhIwUWduZqHngx119c\nRGQlSiR86d+ePXD4cHr8xht9UHXddb70r6cntCWKiAg+kDo5dTIQSE3HpwNz66vql+6QWsxItde1\n6w6pFSbbzNUfAKvM7IP5X9KVUeZKRIrVxATs2wcPPQQjI36stjZd+rdmTajLExFZ0ZKW5MTkiYwg\nqnesl9mF2cDcxurGpU59ixmptto2BVIlJNRW7M65SuDLQAf+MuGRs+eY2Z/nenGXQ8GViBSbY8d8\ng4onn0yX/nV0+LupduzwZ6tERKRwFpILDE4MZgRSfeN9zCfObisArbWtgYxUc3WzAqkSF3ZwtQP4\nJ/zlwedkZkVRPKrgSvKh1OrBJXyJBDz1lA+qjhzxY87BTTf50r83vOHiXf+07yQM2ndSaPnec/FE\nnP6J/oxsVN94HwvJhcDc1XWrM85IrW9eT1N1U97WJuEJ+8zVnwBngA8BLxPsFigiIsD4eLr0bzTV\nJKq2Fu66C+6+G9rbw12fiEg5m1uYo2+8LyMjNTg5SCKZCMxd27B2KRu1+Ku+qj6EVUs5yTZzNQP8\njJl9N/9LujLKXIlIGI4d8w0qnnwSFlI/DF23zmepdu6Eat35KCKSUzPxGXrHezMyUicmT5C0ZMa8\niIvQ0dCRkY2KNcWordRN7CtZ2JmrVwCF8iIiyywswNNP+6Dqtdf8mHNw883+PNXWrbrwV0QkF6bm\npwId+05OnQzMi0aixJpiGRmp7qZuqiv0Ey4pjGwzV+8APgP8lJkdzfeiroQyV5IPOoMgyy2W/j34\nIIyN+bG6OrjzTt/1b/Xq3Hwd7TsJg/adFNrZe258bjwjG3V87Dinp08H3lcRqaC7qTsjI9XV2EVl\ntLKAq5dSFXbm6j8D7cDLzrlXyOwW6AAzszfnenEiIsXk6NF06V8iVb7f2emzVNu3q/RPRORSmBmj\ns6McHj7MxMsT9I73cmz0GKOzo4G5VdEqYs2ZGal1jeuoiGT7raxIYWSbueoBDB9InYuZ2VtyuK7L\npsyViOTSwoLv+rdnD7z+uh+LRHzXv3vugWuuUemfiMjFmBlnZs4EMlLjc+OBuTUVNYGOfR0NHURc\nUTSmljIRaiv2UqLgSkRyYWzMd/x76CFfBghQX5/u+rdqVbjrExEpVmbGyamT6UAq1XRian4qMLe+\nqj7jDqn1zetZU79Gd0hJ3oVdFiiyoukMwspg5rNTe/fCj3+cLv3r6kqX/lVVFW492ncSBu07uRRJ\nS3Ji8kQgIzW7MBuY21jdyIbmDRkZqVW1q3jwwQfZvXN34RcvkgfnDa6cc28GnjazidTHF2RmD2Xz\nBZ1z9wKfA6LAl8zsM2e9/gbgK8AtwG+a2R8ue+0oMA4kgLiZbc/ma4qIXMjCgj9HtXevP1cFvvTv\n1lt9K/Wrr1bpn4hIIplgYGIgIyPVO9bLfCJ4/WlLTQsbWjZkZKRaalqUkZKyd96yQOdcEthpZgdS\nH1+ImVn0ol/MuSj+EuK3Af3AE8B7zezQsjntwAbgPcDIWcHV68AbzWz4Al9DZYEikpXRUV/2t29f\nZunfrl2+9K+tLdz1iYiEJZ6I0z/Rn5GR6hvvYyG5EJi7qm5VICPVVN0UwqpFshdGWeA9wKFlH+fC\nduDwYjt359zXgXcv+zqY2SnglHPuXef5HPqRh4hcNjN/J9WePf6OqsXSv1jMZ6m2b4dKdfEVkRVk\nbmGOvvG+jIzUwMQAiWQiMHdN/ZqlAGrxV32VrkIVWXTe4MrMes718RXqAnqXPfcBOy7h/Qb8yDmX\nAO43sy/maF0iF6QzCKUvHvelf3v2wPHjfiwSgTe+0QdVW7YUX+mf9p2EQfuuvM0uzGZcxNs71suJ\nyRMkLbNIyTnHusZ1GUFUrClGbWVtztekPSflJKuGFs6514D/zcyeOcdrNwL/ZGabsvhUV1qvd6eZ\nDaZKB3/onHvJzPZd4ecUkTI2MpLu+jc56ccaGtKlf62t4a5PRCRfpuanMgOp8V6GJocC8yIupwFw\ncQAAIABJREFUQndTd0Yg1d3UTXWFLu8TuVTZdgvcCJzv/7Ca1OvZ6Adiy55j+OxVVsxsMPXPU865\nf8SXGQaCq/vuu4+NG/2SWlpa2LZt29JPRHp6egD0rGc9l/Hz3Xfv5sgR+MIXenj1VVi3zr++sNDD\nrbfChz+8m8rK4lnv+Z4Xx4plPXrWs56L9/l7D3yPoakh1t6wlt6xXvY9uI+xuTE6b+wEYOC5AQDW\n37yerqYupl6ZYk39Gt5977vpauzikX2PwCjs3lb49e/evTv0f396Lv/ngwcPMjrqL6g+uti9Kg+y\nvUR4qbnFOV77t8DvmdlFj3475yrwDS3eCgwABzirocWyuZ8CJhYbWjjn6oBoqnthPfAA8Ntm9sBZ\n71NDC5EVKh6HJ57wpX+9qQLkSMR3/bvnHti0qfhK/0RELoWZMTY3xrHRYxl3SI3MjATmVkWrAhmp\ndY3rqIjoJh6Rgje0cM79e+D/XDb0befc2b02a4E24OvZfDEzW3DOfQz4Ab4V+5fN7JBz7iOp1+93\nznXguwg2AUnn3K8B1wFrgG+kWnhWAF87O7ASyZeeZdkDKT4jI9DTAw8/nC79a2yEN7/Z/2ppCXV5\nl037TsKgfVc8zIzhmWGOjR3LuENqfG48MLemoiajW9/65vV0NHQQcZEQVn5ptOeknFzoRxevA/+S\n+vj9+IDn9Flz5oAXgC9l+wXN7PvA988au3/ZxyfILB1cNAlsy/briEh5M4PDh32W6uBBSKbOYm/Y\n4LNUb3yjuv6JSOkwM05NnwpkpKbmpwJz6yrrAh371tSv0R1SIkUg27LAvwA+bWav5X1FV0hlgSLl\nbX4eDhzwF/72pU5sRqPprn9XXaXSPxEpbklLMjQ5FMhIzS7MBuY2VDWwoWVDRiC1qnaVAimRK5Sv\nssCsgqtSouBKpDydOQMPPuhL/6ZSP8htavJlf7t2lW7pn4iUt0QyweDkYEZGqnesl/nE2SctoKWm\nJZCRaqlpUSAlkgdhXCIsIimqBw+HGbz6qi/9e+aZdOnfxo3p0r+KMv5TTPtOwqB9d/niiTgDEwMZ\nGam+8T4WkguBuavqVgUCqabqphBWHT7tOSknZfxtiYiUqsXSvz17oL/fj0WjsGNHuvRPRCRMcwtz\n9E/0Z2SkBiYGSCQTgblr6tcEAqn6qvoQVi0i+aayQBEpGmfO+K5/jzySWfp3992+/K9pZf5QV0RC\nNrswS+9Yb0ZG6sTkCZKWzJjnnKOjoSMjiIo1xaitrA1p5SJyPioLFJGyZAYvv+wbVDz7bLr076qr\nfOnfrbeWd+mfiBSXqfkpesd7MzJSQ5NDgXkRFwncIdXd1E11RXUIqxaRYqFvWUSyoHrw3Jubg8cf\n90HVwIAfWyz9u+cef65qpdO+kzCspH03MTfB8bHjHBs7ttSx7/T02bfOQEWkgq6mroxAqquxi8qo\n7nvIhZW056T8KbgSkYI6fTpd+jc97ceam33p365dKv0TkdwzM8bmxnwgNXps6Q6pkZmRwNyqaFUg\nI7WucR0VEX3LJCIXpzNXIpJ3ZvDSS+nSv8X/RTdv9g0qbrlFpX8ikhtmxvDMMMfHjmf8Gp8bD8yt\nqagh1hzLCKQ6GjqIuEgIKxeRQtKZKxEpOXNz8NhjPqgaHPRjFRXwpjf5oGrDhnDXJyKlzcw4NX0q\nEEhNzU8F5tZV1gU69q2pX6M7pEQkpxRciWRB9eCX5uRJX/r36KMwM+PHWlrSpX+NjaEur2Ro30kY\ninXfJS3J0ORQIJCaXZgNzG2oamBDy4aMQGpV7SoFUkWqWPecyOVQcCUiOWEGhw75u6mefz5d+rdl\ni29QsW2bb1ghInIxiWSCwcnBjCCqd6yX+cR8YG5LTUsgI9VS06JASkRCoTNXInJFZmdh/36fqTpx\nwo9VVvrSv3vugVgs1OWJSJGLJ+IMTAxkBFL9E/3EE/HA3FV1qwKBVFO1uuCIyKXTmSsRKSonT/qz\nVI8+6gMsgNZW2L0b7rxTpX8iEjSfmKdvvC8jkBqYGCCRTATmrqlfEwik6qvqQ1i1iEj2FFyJZEH1\n4J4ZvPhiuvRv0dVXp0v/ImqylTPadxKGXO272YXZpbujFn+dmDxB0pIZ85xzrGtclxFExZpi1FbW\nXvEapDTozzopJwquROSiFkv/9u6FoSE/VlkJ27f7rn8q/RNZ2abmp5bujlr8NTQ5FJgXcZHAHVLd\nTd1UV1SHsGoRkdzTmSsROa+hIR9Q7d+fLv1ra/Olf3fdBfWq0BFZcSbmJgId+05Pnw7Mq4hU0NXU\nlRFIdTV2URmtDGHVIiKZdOZKRArCDF54wZf+vfBCevyaa3zp3803q/RPZCUwM8bmxgKB1MjMSGBu\nVbQqkJFa17iOioi+zRCRlUV/6olkYSXUg8/MpEv/Tp70Y1VV6dK/7u5w17cSrYR9J8XBzBieGeb4\n2HG++8Pv0ry1meNjxxmfGw/MramoIdYcywikOho6iDj91EUuj/6sk3Ki4EpkhRsc9G3U9++HuTk/\ntmpVuuufSv9EyouZcWr6VCAjNTU/BcBA3wCdrZ0A1FXWBTr2ralfozukRETOQ2euRFagZNJ3+9u7\n13f/W7R1qy/9u+kmlf6JlIOkJRmaHMq8jHe8l5n4TGBuQ1UDG1o2ZARSq2pXKZASkbKkM1cicsWm\np/29VD09cOqUH6uqgp07felfZ2eoyxORK5BIJhicHMwMpMZ6mU/MB+Y21zSzoTkzkGqpaVEgJSJy\nhRRciWSh1OvBBwfTXf/mU99nrV6dLv2rqwt1eXIepb7vJH/iiTgDEwMZgVT/RD/xRDwwd1XdqsAd\nUs01zef93Np3Umjac1JOFFyJlKlkEp57zgdVhw6lx6+91mepbrxRpX8ipWA+MU/feF9GIDUwMUAi\nmQjMba9vz8hIxZpjNFQ1hLBqEZGVSWeuRMrM9DQ88ogv/Tudunqmqgpuv90HVevWhbo8EbmA2YVZ\nescyL+M9MXmCpCUz5jnn6GjoCFzGW1epNLSISDZ05kpELmhgwN9N9fjj6dK/9nZf+nfHHSr9Eyk2\n0/HpQMe+ocmhwLyIi9DV1JWRkepu6qa6ojqEVYuIyIUouBLJQrHWgyeT8OyzPqh6+eX0+HXX+SzV\nDTeo9K+UFeu+k0s3MTcRCKROT58OzKuIVNDV1JWRkepq7KIyWlmwtWrfSaFpz0k5UXAlUoKmptKl\nf2fO+LHqal/6t3u3Sv9EwmJmjM2NBQKpkZmRwNzKaCXdTd0ZGal1jeuoiOivZhGRUqUzVyIlpK/P\nN6g4cCCz9O+ee3xgVVsb7vpEVhIzY3hmOBBIjc+NB+bWVNQQa45lZKQ6GjqIOKWWRUTCoDNXIitU\nMgnPPONL/155JT1+/fU+qLr+etDVNCL5ZWacmj4VCKSm5qcCc2sra1nfvD4jI7Wmfo3ukBIRWQEU\nXIlkIYx68MlJePhhePBBGB72YzU1vjnF7t2wdm1BlyMh0DmEcCQtydDkUOZlvOO9zMRnAnMbqhp8\nINWSDqRW1a4q6UBK+04KTXtOyknBgyvn3L3A54Ao8CUz+8xZr78B+ApwC/CbZvaH2b5XpBz09qZL\n/+Kp+0DXrvUNKm6/3QdYIpIbiWSCwcnBzEBqrJf5xHxgbnNNcyAj1VLTUtKBlIiI5FZBz1w556LA\ny8DbgH7gCeC9ZnZo2Zx2YAPwHmBkMbjK5r2peTpzJSUnmYSDB33p36uvpsdvvNEHVdddp9I/kSu1\nkFygf7w/I5Dqn+gnnogH5rbVtmVkpGJNMZprmkNYtYiI5EO5nLnaDhw2s6MAzrmvA+8GlgIkMzsF\nnHLOvetS3ytSahZL/3p6YCTVTKymBu6805f+rVkT5upEStd8Yp6+8b6MQGpgYoBEMhGY217fnpGR\nijXHaKhqCGHVIiJS6godXHUBvcue+4AdBXivyBXJdT14b6/PUj3xRLr0r6PDZ6l27lTpn3g6h5Cd\n2YVZesd6MwKpE5MnSFoyY55zjo6GjoyMVHdTN3WVumF7Oe07KTTtOSknhQ6urqReT7V+UtISiXTp\n3+HDfsw5X/p3zz1w7bUq/RO5mOn4dKBj38mpk5xdDh5xkaXLeBczUt1N3VRXVIe0chERWQkKHVz1\nA7FlzzF8Biqn773vvvvYuHEjAC0tLWzbtm3pJyI9PT0AetZzwZ6npwF289BD8MIL/vXNm3dzxx1Q\nWdlDaytcd13xrFfPxfO8OFYs6yn08/ce+B5DU0N03NDB8bHj7HtwH2NzY3Te2AnAwHMDAMRujtHV\n1MXUK1OsbVjLu+99N12NXTyy7xEYhd3biuP3o2c96/ncz7t37y6q9ei5PJ8PHjzI6OgoAEePHiVf\nCt3QogLflOKtwABwgHM0pUjN/RQwsayhRVbvVUMLKRbHjvmuf088AQsLfmzdunTpX7V+gC4C+Duk\nxubGAhmpkZmRwNzKaCXdTd0ZGal1jeuoiOhmERERyV5ZNLQwswXn3MeAH+DbqX/ZzA455z6Sev1+\n51wHvhNgE5B0zv0acJ2ZTZ7rvYVcv6xcPcuyBxeSSMBTT/mg6sgRP+Yc3HyzD6re8AaV/kn2st13\npcTMGJ4ZDgRS43PjgbnVFdXEmmIZZ6Q6GjqIuEgIK185ynHfSXHTnpNyUvAf9ZnZ94HvnzV2/7KP\nT5BZ/nfB94oUg/Fx2LfPX/g7NubHamvhrrtg925YvTrU5YmEwsw4NX2K3rFejo0dWwqkpuanAnNr\nK2uX7o5azEi117crkBIRkZJS0LLAQlBZoBTS0aO+QcWPf5wu/evs9FmqHTtU+icrR9KSDE0O0Tve\ny7FRH0j1jvcyE58JzG2oakgHUqmM1KraVbqMV0RECqYsygJFysHCgi/927MHXn/dj0UisG2bD6q2\nblXpn5S3RDLB4ORgRkaqd6yX+cR8YG5zTXMgI9VS06JASkREypKCK5Es9PT0cOutvuPfQw+lS//q\n69MX/q5aFeoSpQwVwzmEheQC/eP9GRmp/ol+4ol4YG5bbVtGRirWFKO5pjmEVcuVKIZ9JyuL9pyU\nEwVXIhfx+uvwve/B3/2db1gB0NXl76bavh2qqsJdn0iuzCfm6Rvvy8hIDUwMkEgmAnPb69szMlKx\n5hgNVQ0hrFpERKR46MyVyDksLPhzVHv2+HNV4Ev/br7ZB1VXX63SPyltswuz9I71ZmSkTkyeIGnJ\njHnOOdbWr83ISHU3dVNXWRfSykVERK6czlyJFMDoqO/699BDvgMg+NK/u+6Cu+9W6Z+Upun49NK5\nqMWM1Mmpk5z9g6iIi9DV1JWRkepu6qa6Qp1ZREREsqHgSlY8M1/6t2ePb1SxWPrX3e2zVG96Ezz6\naA+rVu0OcZWyEl3OOYSJuYmlTn3Hx45zbPQYp6dPB+ZFI9HMQKplA12NXVRGK3O0eilVOv8ihaY9\nJ+VEwZWsWAsL8OSTPqg6dsyPRSJw660+qNqyRaV/UrzMjLG5saWM1PGx4xwbO8bIzEhgbmW0ku6m\n7oyM1LrGdVRE9FeAiIhILunMlaw4o6P+st99+2Biwo81NMCuXb70r7U13PWJnM3MGJ4ZDmSkxufG\nA3OrK6qJNcUyMlIdDR26jFdERGQZnbkSuQJm8NprPkv19NPp0r9YLF36V6lqKCkCZsap6VNL2ajF\nX5Pzk4G5tZW1S0HUYkaqvb5dgZSIiEhIFFxJWYvH4YknYO9eOH7cj0UicNtt/sLfzZuzK/1TPbjk\nQ9KSDE0OLWWjFn/NxGcAGHhugM4bOwFoqGrICKTWN69ndd1qXcYrOac/76TQtOeknCi4krI0MpIu\n/ZtM/cC/oQHe/Gb/S6V/UmiJZILBycGMjFTveC9zC3OBuc01zT6AGlnPu970LtY3r6e1plWBlIiI\nSJHTmSspG2Zw+LDPUj39NCRT1/WsX+9L/267TaV/UhgLyQX6x/szMlJ9433EE/HA3LbatkBGqrmm\nOYRVi4iIrBw6cyVyHvE4HDjgg6reXj8WjfpzVG95C2zapK5/kj/ziXn6xvsyMlL9E/0kkonA3Pb6\n9owgKtYUo7G6MYRVi4iISD4ouJKSNTzsS/8efjhd+tfYmC79a2nJ3ddSPbgAzC7M0jvWm5GRGpwY\nJGnJjHnOOToaOjIDqeYYdZV1l/T1tO8kDNp3Umjac1JOFFxJSTGDV1/1WaqDB9Olfxs3+izVbbdB\nhXa15MB0fDrjDqnjY8cZmhri7LLjiItkXMa7vnk93U3d1FTUhLRyERERCYvOXElJmJ/3pX979kB/\nvx+LRuGNb/RB1VVXqfRPLt/E3ETGHVLHx45zaupUYF40EqWrMRhIVUZ1mE9ERKSU6MyVrEhnzkBP\nDzzyCExN+bGmpnTpX7PO/cslGp0dDWSkhmeGA/Mqo5V0N3VnBFKdjZ1URPTHpoiIiJybvkuQomMG\nr7zis1TPPpsu/bvqKt/179ZbC1/6p3rw0mNmDM8MBzJSY7NjgbnVFdXEmmIZgdS6xnWhX8arfSdh\n0L6TQtOek3Ki4EqKxtwcPP64z1QtL/3bscMHVRs3hrk6KWZmxqnpUxnZqONjx5mcnwzMra2sDbQ+\nX1O/JvRASkREREqfzlxJ6E6fTpf+TU/7seZmuPtu2LXLlwGKLEpakqHJoYxs1PGx48zEZwJzG6oa\nAoHU6rrVuoxXRERkhdOZKykrZvDyy+nSv8V4eNMmn6W65RZ1/RMfSA1MDGRkpHrHe5lbmAvMba5p\nDgRSrTWtCqRERESkYPTtqxTUYunf3r0wMODHKirSF/5u2BDu+s5H9eD5t5BcoH+8PyMj1TfeRzwR\nD8xtq20LBFLNNeXX3UT7TsKgfSeFpj0n5UTBlRTEYunfww/DTKp6q6UlXfrX2Bjq8qTA4ok4veO9\nGRmp/ol+EslEYG57fXvmZbxNMRqrtWFERESk+OjMleSNGbz0ki/9e+65dOnf5s3p0r9oNNw1Sv7N\nLszSN96XcT5qcGKQpCUz5jnnWFu/NjOQao5RV1kX0spFRESkXOnMlZSMuTl47DEfVJ044cdKofRP\nrtx0fDrQsW9oaoizf+ARcRG6moKX8dZU1IS0chEREZErp+BKcubkSV/69+ij6dK/1lZf+nfXXaVd\n+qd68KCJuYlAx75TU6cC86KRaCCQ6mrqoipaFcKqS4v2nYRB+04KTXtOyomCK7kiZvDii75BxfPP\np0v/tmzxpX/btqn0rxyMzY5lBFHHx44zPDMcmFcZraS7qTsjkOps7KQioj9qREREpPzpzJVcltlZ\n2L/fB1VDQ36sshK2b/elf7FYuOuTy2NmjMyOBAKpsdmxwNzqimpiTbGMQKqjoYNoRNG0iIiIFDed\nuZKiMDSULv2bnfVjra2we7cv/WtoCHN1cinMjNPTpwOB1OT8ZGBubWVtoPX5mvo1RFwkhJWLiIiI\nFKeCB1fOuXuBzwFR4Etm9plzzPlj4B3ANHCfmT2dGj8KjAMJIG5m2wu17pXMDF54IV36t+iaa3yW\nats2iJT599ilXg+etCQnp04GAqmZ+ExgbkNVQyCQWl23WpfxhqDU952UJu07KTTtOSknBQ2unHNR\n4PPA24B+4Ann3LfM7NCyOe8EtpjZ1c65HcAXgJ2plw3YbWbBwx6Sc7OzPkPV05NZ+rdjhw+qurtD\nXZ6cR9KSDE4MZgRRveO9zC3MBeY2VTexoWVDRiDVWtOqQEpERETkMhT0zJVz7nbgk2Z2b+r54wBm\n9vvL5vwpsNfM/kfq+SXgbjMbcs69DtxmZmcu8DV05uoKDQ35LNX+/enSv7a2dOlffX2oy5NlFpIL\nDEwMZARSfeN9xBPxwNy22rZARqq5pjmEVYuIiIiEq1zOXHUBvcue+4AdWczpAobwmasfOecSwP1m\n9sU8rnVFMfMlf3v2+O5/i7Zu9Vmqm28u/9K/YhdPxAOX8fZP9JNIJgJz2+vbMy/jbYrRWF3CvfBF\nRERESkChg6tsU0rniyLvMrMB51w78EPn3Etmti9Ha1uRZmZ86d/evXAqdUVRVVW69K+rK9z1FYtC\n14PPLswGAqnBiUGSlsyY55yjo6EjM5BqjlFXWVewtUr+6ByChEH7TgpNe07KSaGDq35geZPuGD4z\ndaE53akxzGwg9c9Tzrl/BLYDgeDqvvvuY+PGjQC0tLSwbdu2pf9pe3p6AFb889atu9m7F/7hH3qI\nx6GzczerVkFbWw/XXw/veEdxrTfs50X5+PyzC7Ncte0qjo8d54F/eYCTUyepvboWM2PguQEAOm/s\nJOIizB+ZZ23DWt7+1rezvnk9rz39GlWuit23pj/fIIOh//vSc26eDx48WFTr0fPKeF5ULOvRs571\nrOdcPB88eJDR0VEAjh49Sr4U+sxVBfAy8FZgADgAvPccDS0+ZmbvdM7tBD5nZjudc3VA1MwmnHP1\nwAPAb5vZA2d9DZ25Oo9kMl36d+hQevwNb/AX/t54o0r/8m1yfjLQse/U1KnAvGgkSldjV0ZGqqup\ni6poVQirFhERESkvZXHmyswWnHMfA36Ab8X+ZTM75Jz7SOr1+83se865dzrnDgNTwAdSb+8AvpHq\nYlYBfO3swErObWLCN6d48EE4fdqPVVXBzp2+9K+zM9z1laux2bFAIDU8E2x0WRmtpLupOyOQ6mzs\npCKia+hERERESklBM1eFoMyVZwYvvwz79sHTT0Mi1fNg9WofUN1xB9TpWE7Wenp6llLLZzMzRmZH\nAoHU2OxYYG51RTWxplhGINXR0EE0Es3z70BK0YX2nUi+aN9JoWnPSRjKInMl+Tc+7htUPPxwukFF\nJOK7/d11F9xwg0r/roSZcXr6dCCQmpyfDMytrawNtD5fU7+GiNN/ABEREZFypMxVGTDz7dMffhie\neSadpWpr8wHVHXdAa2u4ayxFSUtycupkIJCaic8E5tZX1bOhOfMy3tV1q3UZr4iIiEgRUuZKAkZH\nfZbqkUfSZ6kiEdi2DXbtguuuU5YqW0lLMjgxmBFE9Y73MrcwF5jbVN3EhpYNS/dHbWjZQGtNqwIp\nERERkRVOwVWJSSZ9lmrfPnj2Wf8M/izVXXfB7bdDS0u4ayx2C8kFBiYGMgKpvvE+4ol4YG5bbRvr\nm9cz+tIo73r7u9jQvIHmmuYQVi0rkc4hSBi076TQtOeknCi4KhEjI+mzVMOphnPRKNx6q89SXXst\nKHESFE/EA5fx9k/0k0gmAnPb69vTF/Gmmk40VjcC0DPVw01rbyr08kVERESkhOjMVRFbvJdq3z7/\nz8UsVXu7D6huvx2amsJdYzGZW5ijd7w3I5AanBgkacmMec451tavTQdSzT6QqqtU+0QRERGRlUBn\nrlaQ4WF/juqRR3zGCnyW6rbbfFC1dauyVNPxaXrHMgOpoakhzg6sIy5CV1NXRkYq1hyjpqImpJWL\niIiISLlScFUkEgl47jlf9vf8874DIMDatT6g2rkTGhvDXWNYJucnAx37Tk2dCsyLRqKZgVRzjO6m\nbqqiVVe8BtWDSxi07yQM2ndSaNpzUk4UXIXs9Ol0lmosdedsRUX6LNXVV6+sLNXY7FggkBqeGQ7M\nq4xW0t3UnZGR6mrqoiKiLS0iIiIi4dCZqxAkEv4+qn374NChdJaqoyOdpWpoCHeN+WZmjMyOBAKp\nsdmxwNyqaFXG/VGx5hjrGtYRjURDWLmIiIiIlDqduSoDp075sr9HH4XxcT9WWZnOUm3ZUp5ZKjPj\n9PTpQCA1OT8ZmFtbWZsZSDXFWNuwlojThV0iIiIiUtwUXOXZwkJmlmpRZ6cPqHbsgPr68NaXa2bG\n0NRQIJCaic8E5tZX1bOheUNGRqq9rr0oL+NVPbiEQftOwqB9J4WmPSflRMFVngwN+SzV/v0wMeHH\nqqp8x7+77oJNm0o/S5W0JIMTg4HLeGcXZgNzm6qb2NCyIeMOqbbatqIMpERERERELofOXOVQPA4H\nD/os1csvp8e7u32Wavt2qCvRq5QWkgsMTAwEAql4Ih6Y21rbupSRWrxDqqWmJYRVi4iIiIgE6cxV\nERsc9Fmqxx6DydQxoqoqH0zt2gUbNpRWliqeiNM33pcRSPVP9JNIJgJz2+vbM85HrW9eT2P1Cu0Z\nLyIiIiIrmoKryxSPw1NP+SzVq6+mx9evT2epakrgntq5hTl6xzMv4x2cGCRpycDctQ1rAxmpusoS\nTcVdItWDSxi07yQM2ndSaNpzUk4UXF2igQEfUD32GExP+7GaGnjTm9JZqmI1HZ+mdywzkBqaGuLs\nMsqIi2RextsUI9Yco6aiBKJFEREREZGQ6MxVFubn4cc/9kHVkSPp8Y0bfUB1223Fl6Va7Np3ZPgI\nR0aOcGT4CCcmTwTmRSNROhs7MzJS3U3dVEWrQli1iIiIiEj+6cxVCPr6fED1+OMwk+okXlPj26fv\n2gWxWLjrWy6eiHN09OhSIHVk5AhT81MZcyqjlXQ1dmV07etq6qIiom0gIiIiInKl9F31Webm4Mkn\nfVD1+uvp8auuSmepqqvDW9+i8bnxpSDq8PBhjo8dDzScaK5pZkvbFja1bmJL2xa6m7oVSF0m1YNL\nGLTvJAzad1Jo2nNSTvSddsrx477j3+OPw2zqmqbaWti5099L1d0d3trMjIGJgYys1KmpUxlznHN0\nN3WzuW0zm1s3s7ltM6tqV+keKRERERGRAlnRZ65mZ+GJJ3yW6tix9PiWLT6geuMbfUv1QptbmOP1\n0dc5MnyE10Ze48jIEWbiMxlzqiuq2dS6ic2tm9nUuolNrZuorawt/GJFREREREqMzlzliJkPpB5+\nGA4c8GWAAPX16SxVZ2dh1zQyM5KRleod6w20Qm+rbVvKSm1p20JXUxcRFynsQkVERERE5LxWTHA1\nM+ODqX37oLc3PX7NNf4s1S23QGVl/teRtCT94/0cHj68FFANzwxnzIm4COub17OlbctSQNVa25r/\nxcl5qR5cwqB9J2HQvpNC056TclLWwZWZb0qxb59vUjE/78cbGuD2232WqqMjv2uYic8K8MDiAAAO\n4ElEQVQslfgdGTnC6yOvM7swmzGntrJ2qcRvc9tmrmq5iuqKIuiaISIiIiIiWSvLM1dTU8bjj/ug\nqr8//drWreksVUUewkoz48zMGV4bec1npoaP0D/RH7ikt72+famD3+bWzaxrXKcSPxERERGRAsnX\nmauyDK4++lEjHvfPjY3pLNXatbn9Wolkgt7x3oyW6GOzYxlzopHoUonfYnaquaY5twsREREREZGs\nqaHFJYjH4dprfZbq5ptzl6Wamp9a6t53ZPgIR0ePMp+Yz5hTX1W/VN63uXUzG1s2UhktwGEuySvV\ng0sYtO8kDNp3Umjac1JOyjK4+t3fhfb2K/scZsap6VMcGfYZqddGXmNgYiAwb23D2qUOfptaN9HR\n0KG7pUREREREVqCyLAu8nN/TQnKBY6PHMlqiT8xNZMypjFayoXnDUlZqU+smGqsbc7V0EREREREp\nAJUF5tjE3ES68cTIEY6NHmMhuZAxp7G6cSkrtbltM+ub11MRWbH/ykRERERE5AIKHik45+4FPgdE\ngS+Z2WfOMeePgXcA08B9ZvZ0tu89FzPjxOSJjKzU0ORQYF5nY+dSVmpz22ba69pV4ieA6sElHNp3\nEgbtOyk07TkpJwUNrpxzUeDzwNuAfuAJ59y3zOzQsjnvBLaY2dXOuR3AF4Cd2bx30XxinqOjR5cC\nqddGXmNqfipjTlW0iqtar1pqib6pdRN1lXX5+q1LiTt48KD+4JeC076TMGjfSaFpz0k5KXTmajtw\n2MyOAjjnvg68G1geIP0U8JcAZva4c67FOdcBXJXFewH49X/+dRLJRMZYS01LRlYq1hQjGonm+vcn\nZWp0dDTsJcgKpH0nYdC+k0LTnpNyUujgqgvoXfbcB+zIYk4X0JnFewFfBhhrjmW0RG+rbVOJn4iI\niIiI5E2hg6ts2/hdURT02Xs/S01FzZV8CpEMR48eDXsJsgJp30kYtO+k0LTnpJwUtBW7c24n8Ckz\nuzf1/AkgubwxhXPuT4EeM/t66vkl4G58WeAF35saL6/e8iIiIiIiknPl0Ir9SeBq59xGYAD418B7\nz5rzLeBjwNdTwdiomQ05585k8d68/EsSERERERG5mIIGV2a24Jz7GPADfDv1L5vZIefcR1Kv329m\n33POvdM5dxiYAj5wofcWcv0iIiIiIiLnU9CyQBERERERkXIVCXsBImFwzsWcc3udcy845553zv27\n1Hibc+6HzrlXnHMPOOdalr3nE865V51zLznn3r5s/I3OuedSr/1RGL8fKS3Ouahz7mnn3LdTz9p3\nklepa03+wTl3yDn3onNuh/ad5FNqD72Q2i9/45yr1p6TXHPO/blzbsg599yysZzts9S+/R+p8cec\ncxsutiYFV7JSxYF/b2bXAzuBjzrnrgU+DvzQzK4B/iX1jHPuOvw5v+uAe4E/cene/l8APmhmV+PP\nBd5b2N+KlKBfA14k3UFV+07y7Y+A75nZtcBNwEto30mepM7Hfwi41cxuxB/n+Dm05yT3voLfM8vl\ncp99EDiTGv8skNFI71wUXMmKZGYnzOxg6uNJ/GXUXSy7xDr1z/ekPn438LdmFk9dZH0Y2OGcWwc0\nmtmB1Ly/WvYekQDnXDfwTuBLpK+d0L6TvHHONQO7zOzPwZ9hNrMxtO8kf8bxP8Ssc85VAHX4ZmTa\nc5JTZrYPGDlrOJf7bPnn+v+Bt15sTQquZMVL/YTtFuBxYK2ZDaVeGgLWpj7uxF9cvWj55dbLx/tT\n4yLn81ngPwLJZWPad5JPVwGnnHNfcc495Zz7onOuHu07yRMzGwb+EDiOD6pGzeyHaM9JYeRyn3UB\nveB/MAWMOefaLvTFFVzJiuaca8D/JOLXzGxi+Wvmu72o44vkjHPuJ4GTZvY057ksXftO8qACuBX4\nEzO7Fd+J9+PLJ2jfSS455zYDvw5sxH/j2uCce9/yOdpzUghh7DMFV7JiOecq8YHVV83sm6nhIedc\nR+r1dcDJ1Hg/EFv29m78Tzn6Ux8vH+/P57qlpN0B/JRz7nXgb4F7nHNfRftO8qsP6DOzJ1LP/4AP\ntk5o30me3AY8amZnUj/t/wZwO9pzUhi5+Du1b9l71qc+VwXQnMrMnpeCK1mRUgcYvwy8aGafW/bS\nt4BfTH38i8A3l43/nHOuyjl3FXA1cMDMTgDjqc5bDviFZe8RyWBm/9nMYmZ2Ff5w9x4z+wW07ySP\nUvul1zl3TWrobcALwLfRvpP8eAnY6ZyrTe2Vt+Gb+GjPSSHk4u/UfzrH5/oZfIOMCyroJcIiReRO\n4H3As865p1NjnwB+H/g759wHgaPAzwKY2YvOub/D/+WwAPyKpS+J+xXgL4BafDeufy7Ub0JK3uIe\n0r6TfPtV4GvOuSrgCPABfAc37TvJOTN7xjn3V8CT+POlTwF/BjSiPSc55Jz7W+BuYLVzrhf4LXL7\nd+qXga86514FzuB/MHrhNekSYRERERERkSunskAREREREZEcUHAlIiIiIiKSAwquREREREREckDB\nlYiIiIiISA4ouBIREREREckBBVciIiIiIiI5oOBKRKSEOOfuc84lnXMjzrmWs16rSL32yRDW9anU\n1y7qv1eccxHn3Oecc4POuYRz7hsF+Jqfcs4l8/11REQkfEX9l6CIiJxXM/CfzvNaWBcYlsLFiT8D\n/DvgM8AdwG8U6OuWwr8bERG5QgquRERK0wPArzrn1oS9kGVcXj+5c9U5+DTXpv75R2b2uJkdzsHn\nzEZe/92IiEhxUHAlIlKafjf1z//rQpPOV5LmnPsL59zry543psr6/q1z7vedcyecc+POua865+qc\nc1udcz90zk045151zv3Ceb7kdc65vc65KefcgHPut51zGYGFc67dOfenzrk+59ysc+6Qc+5DZ81Z\nLH/c5Zz7e+fcCPDYRX6v9zrn9jvnpp1zo865f3TOXbPs9aPAYslkIvX533+Bz1fhnPtPzrkXnXMz\nzrmTzrnvO+e2LpuzNfV1RlJfd79z7icuss7Ff9e/eNb47tT4m5eN9Tjn9jnn3uGceya1jh8753Y4\n5yqdc3+QKnE845z7inOu7hxf58POuU+n/nuMOOe+5ZzrOutr/x/OuadT/33HnHPPOuc+fKHfh4iI\nBCm4EhEpTYPA54EPO+fWX2Tu+UrSzjX+CWAt8AvAbwH/GvgS8I/APwHvAZ4F/sI5d9053v9NfFbt\n3cDfAP8l9XkAcM41AQ8D9+IDnXcC3wa+4Jz72Dk+39eAI8BPc/4ySJxz9wLfBcaBnwV+GbgBeNg5\n15ma9h7gL1If70z9+t75PifwdXwQ+53U7+dDwAvAutTX7Ez9Xm4EPpr6uqPAd1PruZhsSgUN2IIv\nY/yv+LLGGvx/iy8Dq4D3A58Gfp508LjcJ4BNwAeAXwNuB/568UXn3F3AV4G9qd/nTwNfxJeeiojI\nJagIewEiInJZDP8N90fw31B/8AJzz1eSdq7xV83sA6mPf+ic2wX8HPA+M/sbAOfcj4Gfwn+j/+mz\n3v9nZvYHqY9/lAqm/oNz7rNmNo7/5n49cIOZHUnN25NqzvFJ59yfmNnyTNvfm9nHL/B7W/S7/M/2\n7i/EqiqK4/h31SSCPjgzVPOg1GM9BAZFJUNFkBFj9AcSiYggn4IKFHqTMUUKwoYgGsiHnsewCGya\nsSgtQpGsoAiaogwpHdEZzQozcPWw9onLnnP/zpnicn8fuJw55+x99j73zsNdrH3WhR+A+4v+ZnYY\nmAG2Alvd/Ssz+xXA3Y82upiZ3QM8Ajzr7q/VnHq35u8twCrgNnf/MfWbBL4lAqGpFubdjAEDwB3u\nfjyNcUWax5C7r0/tPkgZr0dZGIT+5O6P19zb1cDLZjbk7qeIIPOcu2+p6fNhBXMXEek5ylyJiHQp\nd58HdgNP1C5/W6T3s/3v0na6ZtxzwGlgdUn/vdn+BLCSyCJBZKyOAMfTsrs+M+sjsl2DQJ4Ne6fZ\nhM1sBXAzMFEbmKVg5DPgrmbXKLGeCGD3NGhzJ3C4CKzSmJeJjNdaM1vZwbhlZorAKlnwmdQcL/tM\n8uzcN2lbZDyPAv1pCegGy6pQiohI6xRciYh0tzFgjsggVVGRbj7bv9Tg+PKS/rN19otnfK4hgp2/\n0zWK115i/oNZ/5MtzLmfyPCUtZ0lMj/tGgTm3P2vBm0G6ox5Ks2nv4Nxy7TzmfTZwnL4c9l+cU/L\nAdz9EyLjtQZ4Gzidnq+7aVGzFhHpQQquRES6mLv/AbxIfDleW9LkIkRxhuz4IEtTHnwo2782bX9J\n2zNENumWktetwLGsfytznE/t8rGL+Zxt4Rq5M8CAmZUFkIWzpOevSsZ0FgY/hYtpuyw7ngeW/xl3\n3+fudxPLHB8m7msqL0YiIiKNKbgSEel+rxPBy66Scz+n7b9ZiLTsa90SzWVjtr8JuAB8nfaniHLo\nJ9z9i5LX7+0OmALMY8DG2qyNmV1H3OfBDu5jmsg+bW7Q5hBwexqnGPNKoghIo3uZJbJHeWZopIN5\nVsrd/3T394A3iACrk6yfiEjPUkELEZEu5+6XzGwH8YU4NwmcB/aY2SixFOx5IuBZTFaiXt/NKcD5\nHLiPKLQx6u4X0vkxIvj41MzGiIITK4AbgGF3f6jD+WwjqgXuN7Nx4jmvF4js0e52L+buB81sH/CK\nma0hKuldRTxntd/dD6V7eZIoJjFKvKdPE9X96gZK7u5mNgE8ZWYzxHswQv1nw5b698N2EMs1PyaW\nOa4mfmj5S3fvJOsnItKzlLkSEek+ZUvl3gS+z8+5+3lgA3CZeK5pF/Aq8UW61VLgZe3yY0W7B4F7\niWp2jwE73X1nzXx+I7JJk0RVuymipPgDwEdNxqg/SfdpIkBZRRTRGCfKpg+ninjN7qfMJmA7UcK9\nKH1+I1BUHDwJDKdxxoG30vgj7n6gyZjPEc83bScKYCwDnilp1858221bOAJcTwSLB4CXiP+P/z2T\nJiLSbcx9KZbci4iIiIiI9BZlrkRERERERCqg4EpERERERKQCCq5EREREREQqoOBKRERERESkAgqu\nREREREREKqDgSkREREREpAIKrkRERERERCqg4EpERERERKQCCq5EREREREQq8A/8v0nMEi/XJgAA\nAABJRU5ErkJggg==\n", + "text": [ + "
" + ] + } + ], + "metadata": {} + } + ] +} \ No newline at end of file diff --git a/benchmarks/timeit_tests.ipynb b/benchmarks/timeit_tests.ipynb index baf4bed..1099508 100644 --- a/benchmarks/timeit_tests.ipynb +++ b/benchmarks/timeit_tests.ipynb @@ -1,7 +1,7 @@ { "metadata": { "name": "", - "signature": "sha256:ab74fdc9e8ae58388d1fc428599abc86a3d03938d0f51769cef38ab4028d516a" + "signature": "sha256:a4749ce2a9f843d9846081abaa9265690ebabfaf5a0aa18877f65945b1f56805" }, "nbformat": 3, "nbformat_minor": 0, @@ -12,10 +12,11 @@ "cell_type": "markdown", "metadata": {}, "source": [ - "Sebastian Raschka \n", - "last updated: 04/14/2014 \n", + "[Sebastian Raschka](http://sebastianraschka.com)\n", + "last updated: 05/07/2014 \n", "\n", - "[Link to this IPython Notebook on GitHub](https://github.com/rasbt/python_reference/blob/master/benchmarks/timeit_tests.ipynb)" + "- [Link to this IPython Notebook on GitHub](https://github.com/rasbt/python_reference/blob/master/benchmarks/timeit_tests.ipynb) \n", + "- [Link to the GitHub repository](https://github.com/rasbt/python_reference/blob/master/benchmarks/timeit_tests.ipynb) \n" ] }, { @@ -24,7 +25,7 @@ "source": [ "
\n", "I am really looking forward to your comments and suggestions to improve and extend this collection! Just send me a quick note \n", - "via Twitter: [@rasbt](https://twitter.com/rasbt) \n", + "via Twitter: [@rasbt](https://twitter.com/rasbt) \n", "or Email: [bluewoodtree@gmail.com](mailto:bluewoodtree@gmail.com)\n", "
" ] @@ -33,11 +34,16 @@ "cell_type": "markdown", "metadata": {}, "source": [ - "\n", - "\n", "# Python benchmarks via `timeit`" ] }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "- Code was executed in Python 3.4.0" + ] + }, { "cell_type": "markdown", "metadata": {}, @@ -66,7 +72,13 @@ " - [Adding elements to a dictionary](#adding_dict_elements)\n", "- [Comprehensions vs. for-loops](#comprehensions)\n", "- [Copying files by searching directory trees](#find_copy)\n", - "- [Returning column vectors slicing through a numpy array](#row_vectors)" + "- [Returning column vectors slicing through a numpy array](#row_vectors)\n", + "- [Speed of numpy functions vs Python built-ins and std. lib.](#numpy)\n", + " - [`sum()` vs. `numpy.sum()`](#np_sum)\n", + " - [`range()` vs. `numpy.arange()`](#np_arange)\n", + " - [`statistics.mean()` vs. `numpy.mean()`](#np_mean)\n", + "- [Cython vs. regular (C)Python](#cython)\n", + "- [Numba vs. Cython vs. regular (C)Python & NumPy](#numba)" ] }, { @@ -118,7 +130,7 @@ "cell_type": "markdown", "metadata": {}, "source": [ - "We expect the string .format() method to perform slower than %, because it is doing the formatting for each object itself, where formatting via the binary % is hard-coded for known types. But let's see how big the difference really is..." + "We expect the string `.format()` method to perform slower than %, because it is doing the formatting for each object itself, where formatting via the binary % is hard-coded for known types. But let's see how big the difference really is..." ] }, { @@ -127,21 +139,16 @@ "input": [ "import timeit\n", "\n", - "def test_format():\n", - " return ['{}'.format(i) for i in range(1000000)]\n", + "n = 10000\n", "\n", - "def test_binaryop():\n", - " return ['%s' %i for i in range(1000000)]\n", + "def test_format(n):\n", + " return ['{}'.format(i) for i in range(n)]\n", "\n", - "%timeit test_format()\n", - "%timeit test_binaryop()\n", + "def test_binaryop(n):\n", + " return ['%s' %i for i in range(n)]\n", "\n", - "#\n", - "# Python 3.4.0\n", - "# MacOS X 10.9.2\n", - "# 2.5 GHz Intel Core i5\n", - "# 4 GB 1600 Mhz DDR3\n", - "#" + "%timeit test_format(n)\n", + "%timeit test_binaryop(n)" ], "language": "python", "metadata": {}, @@ -150,8 +157,8 @@ "output_type": "stream", "stream": "stdout", "text": [ - "1 loops, best of 3: 400 ms per loop\n", - "1 loops, best of 3: 241 ms per loop" + "100 loops, best of 3: 4.01 ms per loop\n", + "1000 loops, best of 3: 1.82 ms per loop" ] }, { @@ -162,7 +169,86 @@ ] } ], - "prompt_number": 3 + "prompt_number": 131 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "funcs = ['test_format', 'test_binaryop']\n", + "\n", + "orders_n = [10**n for n in range(1, 6)]\n", + "times_n = {f:[] for f in funcs}\n", + "\n", + "for n in orders_n:\n", + " for f in funcs:\n", + " times_n[f].append(min(timeit.Timer('%s(n)' %f, \n", + " 'from __main__ import %s, n' %f)\n", + " .repeat(repeat=3, number=1000)))" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 132 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%pylab inline" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 7 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "import matplotlib.pyplot as plt\n", + "\n", + "labels = [('test_format', '.format() method'), \n", + " ('test_binaryop', 'binary operator %')] \n", + "\n", + "matplotlib.rcParams.update({'font.size': 12})\n", + "\n", + "fig = plt.figure(figsize=(10,8))\n", + "for lb in labels:\n", + " plt.plot(orders_n, times_n[lb[0]], alpha=0.5, label=lb[1], marker='o', lw=3)\n", + "plt.xlabel('sample size n')\n", + "plt.ylabel('time per computation in milliseconds [ms]')\n", + "plt.xlim([1,max(orders_n) + max(orders_n) * 10])\n", + "plt.legend(loc=2)\n", + "plt.grid()\n", + "plt.xscale('log')\n", + "plt.yscale('log')\n", + "plt.title('Performance of different string formatting methods')\n", + "max_perf = max( f/b for f,b in zip(times_n['test_format'],\n", + " times_n['test_binaryop']) )\n", + "min_perf = min( f/b for f,b in zip(times_n['test_format'],\n", + " times_n['test_binaryop']) )\n", + " \n", + "ftext = 'The binary op. % is {:.2f}x to {:.2f}x faster than .format()'\\\n", + " .format(min_perf, max_perf) \n", + "plt.figtext(.14,.75, ftext, fontsize=11, ha='left')\n", + "\n", + "\n", + "plt.show()" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "display_data", + "png": "iVBORw0KGgoAAAANSUhEUgAAAnIAAAIECAYAAACdVcNJAAAABHNCSVQICAgIfAhkiAAAAAlwSFlz\nAAALEgAACxIB0t1+/AAAIABJREFUeJzsnXlczdn/x1/3tq9KKRKtpL0ou9xIYxlDvr7WqNDIzDBj\nJssYJDPMDD/rMBgzlH0wtrGTVkK2UJSiZCktKgm3bu/fH3376NZt1b0tzvPxuA99zuec93mf9+fc\nz317n/f5fHhERGAwGAwGg8FgNDv4ja0Ag8FgMBgMBqN+MEeOwWAwGAwGo5nCHDkGg8FgMBiMZgpz\n5BgMBoPBYDCaKcyRYzAYDAaDwWimMEeOwWAwGAwGo5nCHDlGs6O4uBhTpkyBrq4u+Hw+IiIiGlul\nZsmBAwdgZmYGeXl5TJkypdbtlixZgk6dOlV5XJXssLAw2NjYQFFREQMGDGiYQTRDgoKCoKCgILP+\n6nudmzJ8Ph979uxpbDWkgkAggK+vr1Rke3t7Y9CgQVKRzWg8mCPHkAre3t7g8/ng8/lQUFCAsbEx\nZsyYgZycnA+W/c8//2Dv3r04fvw40tPT0atXrwbQ+ONCJBJhypQpGDduHNLS0rBu3bp6y5ozZw6u\nXLlSo+wZM2bAyckJjx49wqFDhz54DA2Bubk5AgMDP1iOvLw8duzYUau648aNw7Nnzz64z9rQkNdZ\nmuzatQt8fuWfIzc3N/j4+FQqT09Px3/+8x9ZqCY1qhozj8cDj8eTWr/SlM1oHOQbWwFGy8XFxQX7\n9+9HcXExrl27Bl9fX6SlpeH48eP1kicUCqGoqIgHDx6gffv26Nmz5wfpVybvY+TZs2d4/fo1hgwZ\ngnbt2n2QLDU1NaipqVUrm4iQlJSEH374Ae3bt693X0QEkUgEefmGuXU11I8aj8dDTc9WL9NdWVkZ\nysrKDdJvTTTUdS4qKpJpFLEm9PT0GluFZgt7B0ALhBgMKeDl5UVubm5iZcuWLSM5OTl6+/YtERHt\n3buX7O3tSVlZmYyNjenbb7+l169fc/X79+9PU6dOpYULF1K7du2obdu2JBAIiMfjcR8TExMiIhIK\nhTRv3jxq3749KSoqkpWVFe3Zs0esfx6PR+vXr6fx48dTq1ataOzYsbR9+3aSl5en0NBQsrGxIRUV\nFXJ1daXnz5/ThQsXyN7entTU1MjNzY2ePn3KyXr48CF5eHiQgYEBqaqqkq2tLe3cuVOsv/79+9O0\nadNo6dKl1LZtW2rdujVNnjyZCgoKxOrt27ePunbtSsrKyqSjo0NDhgyhly9fcufXr19PFhYWpKys\nTJ06daJly5ZRcXFxtfaPjo6mfv36kYqKCmlra9OECRPoxYsXRES0fft2MRvyeDwKDw+XKOfNmzfk\n5+dHrVq1Im1tbZoxYwbNnz+fzM3NuToBAQHccUXZfD6fwsLCKvUXHBxMREQPHjygUaNGkZaWFmlr\na5O7uzvduXOHk13++jg4OJCioiKdPn2ahEIhBQQEkImJCSkrK5O1tTVt2bKl0vX+/fffydPTkzQ0\nNMjQ0JB+/vlnsetTUa/U1FSJdrh79y65u7uTlpYWqampkaWlJXe9jYyMKo25Kt1PnTrFlVcc48WL\nF8nR0ZFUVVWpW7duFBMTI6bD+fPnycbGhpSVlcnBwYEiIiKIx+PRrl27JOpc3XU+ceIEde3alZSU\nlEhPT4+++OILse9e2fd3/fr1ZGRkRHJycvTmzRvi8Xj022+/0ZgxY0hNTY2MjIzo0KFDlJOTQ+PG\njSMNDQ0yNTWlf/75R0yXBQsWkKWlJamqqlKHDh3Iz8+P8vLyiIgoNDS0kp7e3t7k7e1dpf4Vx13T\ntSYiysrKotGjR5Oamhq1bduWAgMDJd6nyvPo0SPi8Xi0Z88ecnd3J1VVVbK0tKTIyEhKTU2lTz75\nhNTU1MjKyooiIyPF2lY3tyWN2cfHh4iIBAJBre4bK1euJBMTE1JUVCQzMzNau3at2Pns7GzuOunr\n69PChQtp8uTJYuONjIyk3r17k4aGBmloaJC9vT2dOXOmSnswmibMkWNIBS8vLxo0aJBY2apVq4jH\n41FBQQFt376dtLW1adeuXfTo0SOKiIggOzs7mjRpEle/f//+pKGhQTNmzKB79+7R3bt3KScnh/z9\n/cnExIQyMjIoKyuLiIj8/f1JR0eHDh48SA8ePKDly5cTn8+nkJAQTh6PxyMdHR3auHEjPXz4kB48\neEDbt28nPp9Prq6udPXqVbpx4wZ16tSJ+vbtSy4uLnTlyhW6desWdenShcaOHcvJunPnDm3cuJFu\n375NDx8+pN9++4370S6vv5aWFn377beUkJBAZ8+epdatW9OiRYu4Otu2bSMFBQX66aefuDFu2LCB\nG1dAQAAZGRnRkSNHKCUlhU6ePEkdO3YUk1GR58+fk4aGBk2cOJHu3r1LUVFRZGdnRy4uLkRU6pzF\nxMQQj8ejf//9lzIyMkgoFEqU9c0335Cenh4dO3aMEhISyN/fnzQ1NalTp05cnYCAAO64Ktnp6enc\nj21GRga9efOG0tPTSV9fn7744gu6e/cuJSYm0syZM0lHR4cyMzOJiLjr06NHDwoLC6NHjx5RZmYm\neXl5kb29PZ07d45SUlLo77//Ji0tLfrrr7/Erre+vj79+eef9PDhQ9q4cSPxeDxuTuTk5JCJiQnN\nmTOHMjIyKCMjg0QikUQ72Nra0sSJE+nevXv06NEjOnXqFB0/fpyIiDIzM0leXp7Wr1/PyalOd0mO\nHJ/Pp/79+1NUVBTdv3+fhgwZQiYmJpzD/uTJE1JRUSFfX1+6d+8ehYSEUNeuXYnH49Hu3bsl6lzV\ntYiNjSU5OTluXp46dYo6duwo9t3z8vIiTU1NGjVqFN2+fZvu3r1LxcXFxOPxqG3btrRjxw5KTk6m\nL774gtTU1Mjd3Z2Cg4MpOTmZZs6cSWpqapSdnc3J++mnnygqKopSU1MpJCSEunTpQl5eXkRU+p+w\nsmtTZr/8/HzKy8sjFxcXGjduHFdeNk8rjruma01ENHz4cLKwsKCwsDCKi4sjHx8f0tLSqnSfKk+Z\nI2dmZkZHjx6lxMRE8vDwoPbt25NAIKAjR45QYmIijR49mjp06EBFRUVERDXO7arGTFS7+8aGDRtI\nRUWFtm7dSklJSbR582ZSVlYWm/8jR46kTp06UWhoKMXFxZGnpydpampy4y0qKiJtbW367rvvKCkp\niZKSkujIkSOVHFJG04c5cgypUPF/unFxcWRqakq9evUiotIoRsUISnh4OPF4PMrNzSWi0huahYVF\nJdnlI0BERK9fvyYlJSXatGmTWD0PDw8aMGAAd8zj8WjatGlidcqiFrGxsVzZypUricfj0Y0bN7iy\nNWvWkK6ubrVjHjFiBPn6+nLH/fv3JwcHB7E6M2bM4GxARNShQweaOXOmRHmvX78mVVXVSv9DDg4O\nJi0trSr1WLhwodiPChFRbGws8Xg8ioiIIKL3P1AXL16sUk5BQQEpKyvTn3/+KVbu5ORUyZErfz2q\nkl3xxzcgIIB69uwpVqekpEQsulB2faKiorg6Dx8+JD6fTwkJCWJtAwMDxezN4/Ho66+/FqtjaWlJ\n33//PXdsbm5OgYGBVdqgjFatWlFQUFCV5+Xl5bkoYxmSdC8rr+jI8Xg8unnzJld25coV4vF4lJiY\nSESlES0TExMqKSnh6pw+fbpaR45I8rXw9PSkHj16iNU7evQo8fl8evz4MRGVfn+1tbXFonREpTad\nPXs2d5yZmUk8Ho9mzZrFlb18+ZJ4PB6dOHGiSr0OHTpESkpK3PHOnTuJx+NVqufm5sZFqirqUdGR\nq+5aJyYmEo/HowsXLnDni4qKqEOHDrVy5NatW8eVlTnHq1ev5spu3rxJPB6P4uLiiKh2c7uqMdfm\nvmFoaEjz5s0TqzN79mwyNTUlotJoII/Ho/Pnz3PnhUIhtW/fnhtvTk4O8Xg8CgsLq3L8jOYB2+zA\nkBphYWHQ0NCAqqoqbG1tYW5ujt27dyMzMxOPHz/G7NmzoaGhwX2GDh0KHo+HpKQkTka3bt1q7Ccp\nKQlCoRAuLi5i5S4uLoiLixMr6969e6X2PB4Ptra23LG+vj4AwM7OTqwsOzubyy8pLCzE/PnzYWNj\nAx0dHWhoaODkyZN4/PixmFx7e3uxvtq1a4eMjAwAwIsXL/DkyRO4u7tLHFdcXBzevHmDUaNGidnJ\nz88P+fn5yM7OrrJdz549xfLI7Ozs0KpVK8THx0tsI4nk5GS8e/cOvXv3Fivv06dPg+TZxMTE4Pr1\n62Jj09TURGpqqtgcAABnZ2fu72vXroGI0K1bN7G2P//8c6V2Dg4OYscGBgZ48eJFnXX19/fHtGnT\n4OrqisDAQNy8ebPWbcvrXhUV50pZPlvZXImPj4ezs7NYTl99c0Tj4+MlfleISGx+WFpaQlVVtVL7\n8nrq6upCTk5O7LuipaUFRUVFMTsfOnQILi4uaN++PTQ0NODp6YmioiKkp6fXawySqO5al42rvM3k\n5eXh5ORUK9nlx1zV/QEA119d5nZFarpv5Ofn4+nTpxKvYUpKCt6+fcuNt/x3V0FBQWwuamtrY9q0\nafjkk08wdOhQ/Prrr0hMTKyVPRhNC7bZgSE1evbsieDgYMjLy8PAwIBzLMpuSOvXr4erq2uldmXJ\n8DweTyyJviGQJI/P54v9QJb9LScnV6mMiMDj8TBnzhwcO3YMa9asgYWFBVRVVfHdd98hLy9PTHbF\nzRQ8Hg8lJSW10rWs3sGDB9G5c+dK57W1tSW2q03ifVOAiODm5oYNGzZUOteqVSvubzk5OTE7ltkl\nOjq6kqNRcfPCh9i/PAsXLsTEiRNx+vRpXLhwAcuXL8fcuXPx448/Vtuuou5VUdUcLNO1oXca1mZ+\nSHLiAEjc9FCxrLydr1y5gjFjxmDBggVYtWoVtLW1ER0dDS8vLwiFwnpoLxlJdq54rSvasbbfk/Lj\nK5Mhqaysv9rO7apoqHlbkYrj/eOPP/D111/j7NmzOHfuHBYtWoQNGzbg888//+C+GLKDReQYUkNZ\nWRmmpqbo2LGjWHRIX18fHTp0wP3792Fqalrpo6SkVKd+zM3NoaSkhPDwcLHy8PBwsUhbQxIZGQlP\nT0+MHj0atra2MDExQUJCQp1+cPX09GBoaIgzZ85IPG9tbQ1lZWUkJydLtJOkRxeUtbt8+TKKioq4\nstjYWOTl5cHGxqbW+pmZmUFRUREXL14UK7948WKDOBZOTk64e/cu2rdvX2lsOjo6VbYri9KmpqZW\namdiYlInHRQVFSESiWpV18TEBDNmzMCBAwcQGBiITZs21UtOfbCyskJMTIzYj/nly5frJcva2rrS\nsxfDw8PB4/FgbW39QXpKIioqCrq6uli6dCmcnZ1hbm6OtLQ0sTpljktFR0NRURHFxcX16rf8HLWy\nsgIAXLp0iSsrLi7G9evX6yW7Jmozt6sac01oamrC0NBQ4v3O1NQUysrK3HjLf3eFQiFiYmIqybO2\ntsbs2bNx8uRJTJ06FX/88Ued9GE0Ps3Okbt69Sp69+6N/v37Y8KECfX+kjMal2XLlmH9+vVYvnw5\n7t69i4SEBBw5cgR+fn5cHSrN4axRlqqqKmbNmoVFixbh4MGDSExMxPLly3Hs2DEsWLBAKvpbWFjg\nyJEjiImJQXx8PD7//HM8f/5cTN/a6B8QEIAtW7bgp59+wr179xAXF4cNGzYgOzsb6urqWLBgARYs\nWIDff/8dCQkJiIuLw759+zB//vwqZX711VfIz8+Ht7c34uLiEBUVhUmTJsHFxQV9+vSp9RjV1NTg\n5+eHhQsX4t9//0VCQgLmzp2LxMTEBon4ffXVVxCJRBgxYgSioqKQkpKCqKgo/PDDD4iOjq6ynbm5\nOaZMmQJfX1/s2rULSUlJiI2NxbZt27BixYpq+6x4TUxMTBAVFYW0tDRkZWVJHNfr16/x5ZdfIjQ0\nFI8ePcLNmzdx+vRpMafHxMQEFy5cwPPnz5GVlVUPa1TPF198gYyMDMyYMQP37t1DaGgofvjhBwB1\nj9bNmTMHN27cwLfffov79+/j9OnTmDlzJjw9PWFoaNjgunfp0gWZmZnYtm0bHj58iB07dog5wQA4\nB/zo0aPIzMzE69evufLr16/j4cOHyMrKqtP9vvy17tSpE4YPH44vv/wSERERiI+Px/Tp05Gfny+V\n56rVZm5XNeba3De+//57/Pbbb/jzzz/x4MEDbNmyBZs3b+bud+bm5vjss8/w5ZdfIiwsDPHx8Zg2\nbRoKCgo4GUlJSZg3bx4uXryI1NRUREdHIzIyUirOPEO6NDtHrmPHjggNDUV4eDiMjY1x9OjRxlaJ\nIYGaHmrp6emJ/fv34/jx4+jRowe6d++OwMBAsR+SqmRIKl+2bBl8fX3xzTffwNbWFnv27MHu3bsl\nLt1KklfXsjVr1sDIyAiurq5wc3NDhw4dMHr06ErLYxXlVCybOnUqgoKCcPDgQTg6OqJ///44c+YM\nF8FcuHAhVq9eja1bt8LBwQH9+vXDunXrqo086enp4ezZs3jy5AmcnZ0xfPhw2NnZ4eDBgzWOsSK/\n/PILRo4ciUmTJqFHjx7Iz8/Hl19+Watx1oSenh6io6Ohq6uLUaNGoUuXLvD09ERaWhoMDAyqlfXH\nH39g9uzZWLZsGaytreHm5oadO3fCzMys2j4r6hoYGIjc3FxYWFhAX1+/UqQIKM2lys3NxdSpU2Fl\nZYXBgwejXbt2Ym8WWLVqFa5fvw5jY2MuX6o6O9TGXuXLDAwMcOzYMVy6dAmOjo6YPXs2fvrpJwCo\n8Zl0FWXb2tri2LFjiIiIgIODAyZPnozhw4dj8+bNYm0aysEZNmwYfvjhByxYsAB2dnbYv38/Vq5c\nKSbf2dkZX3/9NaZPnw59fX3MnDkTAPDdd99BV1cX9vb20NPTE4uo1UTFMWzfvh02NjYYMmQIBgwY\nAENDQ7i7u9fZfrUpq83crmrMtblvzJgxA0uXLsXy5cthbW2NlStX4tdffxV7ePK2bdvg4OCATz/9\nFAKBAB06dICHhwd3Xl1dHUlJSRg3bhwsLCwwevRo9OnTR+JyMKNpw6PmkExTBQEBAXB0dMTIkSMb\nWxUGg8GQKRERERAIBLhz5w6LotQDkUiELl26YOTIkVi5cmVjq8Ng1Jtm68ilpqZi/PjxiIyMFEtK\nZzAYjJbIpk2bYG9vDwMDA8THx2P27NnQ0dGpU5TqYyYyMhIZGRlwdHTEq1evsGbNGuzbtw83btxg\njjCjWdNoS6sbNmyAk5MTlJWVK71LLycnBx4eHlBXV4exsTH27t0rdj4/Px+TJ09GcHAwc+IYDMZH\nwePHjzF+/Hh06dIFX3zxBfr3748TJ040tlrNBpFIhGXLlsHBwQEDBgxASkoKQkNDmRPHaPY0WkTu\n8OHD4PP5OHPmDN68eYPt27dz58aPHw8A+Ouvv3Dz5k0MGzYMly5dgpWVFYqLi/HZZ5/B398fAwYM\naAzVGQwGg8FgMJoG0n/mcPUsXLiQvL29ueOCggJSVFSkBw8ecGWTJ0+m+fPnExHRjh07SEdHhwQC\nAQkEAvr7778lyjUwMCAA7MM+7MM+7MM+7MM+Tf5jb29fLz+q0XetUoWAYGJiIuTl5WFubs6V2dvb\nc0/onzRpErKyshAaGorQ0FCMGTNGotxnz55x27hl8QkICJBp+9rUr65OVedqWy6p3ofaQJb2rqsM\nadm7LraszTVoyjZv6Dle3/PM3vWvz+4pDSeD3VNa9hyvj71jY2Pr5Uc1uiNXcZt1QUEBNDU1xco0\nNDTw6tUrWapVZwQCgUzb16Z+dXWqOlfbckn1UlJSatSpofhQe9dVhrTsXdW52pTJ0t6S+pd2+5rq\n1/c8s3f967N7SsPJYPeUlj3HZWnvRt+1unDhQjx9+pTLkbt58yb69u3LPRwRAP7v//4PEREROHbs\nWK3lNpfXFLUkvL29ERQU1NhqfDQwe8sWZm/Zw2wuW5i9ZUtFe9fXb2lyEbnOnTujuLhY7MXCsbGx\ndXq1UBlLlixBWFjYh6rIqCXe3t6NrcJHBbO3bGH2lj3M5rKF2Vu2lNk7LCwMS5YsqbecRovIiUQi\nFBUVITAwEE+fPsXWrVshLy8POTk5jB8/HjweD3/++Sdu3LiBTz/9FNHR0bC0tKy1fBaRYzAYDAaD\n0VxodhG5H3/8Eaqqqvj111+xa9cuqKioYNmyZQCA33//HW/evIGenh48PT2xefPmOjlxjMaBRT9l\nC7O3bGH2lj3M5rKF2Vu2NJS95RtESj1YsmRJlaFEbW1tHD58WLYKMRgMBoPBYDQzGn2zg7SoLkTZ\nunVrvHz5UsYaMRjSQ1tbGzk5OY2tBoPBYDDqSX2XVhstIicLlixZAoFAUGnL78uXL1n+HKNFUXHT\nEIPBYDCaB2FhYR+0zPpRRuTYRghGS4PNaekTFhbWIM8aY9QeZnPZwuwtWyrau9ltdmAwGAwGg8Fg\nfBgsIsdgtADYnGYwGIzmDYvIMRgMBoPBYHxktGhH7mN5s8OdO3fQvXt3qKiowNTUtLHVqRceHh5Y\nsWIFdzxo0CBs2rSpETWSTFhYGPh8Pp49e9bgslNSUsDn83Hp0qUGl834cD6Ge0lTg9lctjB7y5Yy\ne3/omx1avCP3MSRuzp07F1paWkhISEBMTExjqyORJ0+egM/nIyIiotK5qKgoREVFYebMmVxZQEAA\nAgMDUVhYKEs1xTA3N0dgYGCj9c9gMBiMlo9AIPggR65FP36kviQkpOL8+WQUFfGhoFACNzczWFgY\nNTmZZSQlJcHLywsdO3astwwigkgkgry8dKeEpPX/devWYcKECVBRUeHK+vbtC01NTfz999/w8fGR\nqk5VwR7pwSjPx/CfwqYGs7lsYfaWLQ1l7xYdkasPCQmpCApKQmbmAOTmCpCZOQBBQUlISEhtUjKB\n90txycnJWLx4Mfh8PpYuXfq/PhMwbNgwaGhoQENDA5999hmSk5O5tkFBQVBQUEBYWBgcHR2hrKyM\n8+fPQyAQYNq0aVi4cCH09PSgra2NxYsXg4gQEBCAtm3bQk9PDwsXLhTTZc+ePejRowe0tLTQpk0b\nfPrpp3jw4AF3vszJdHV1BZ/P55aACwoKcOzYMXh4eFQan4eHB3bt2lWtDby9vTFo0CD89ttvMDQ0\nhIaGBvz8/CASibBhwwYYGRmhdevWmD59OoqKisTa/vbbb+jSpQtUVFTQuXNnLF++HCKRCEDpFyw5\nORmBgYHg8/mQk5PD48ePubbx8fFwcXGBmpoarK2tcfr0aTHZNdkfAPbv3w9zc3OoqKigT58+uH37\ndrVjZTAYDAajIiwiV4Hz55OhpDQQ4qkCA3H79gU4O9cvgnb1ajIKCweKlQkEAxEScuGDonIdO3bE\n8+fP4ezsDE9PT3zzzTdQU1PDmzdv4O7ujs6dOyMiIgJEBH9/fwwePBjx8fFQUFAAAJSUlGD+/PlY\nu3YtjIyMoK6uDgA4ePAgZsyYgUuXLiEyMhJTp07F1atX4eDggKioKFy6dAne3t7o27cvBg8eDAAQ\nCoVYvHgxrKyskJ+fj8WLF2PYsGGIi4uDgoICbty4ga5du+LQoUPo3bs35OTkAACXLl1CcXExnJ2d\nK42vR48eWL9+PYqKijidJdv3KgwNDRESEoIHDx7gv//9L1JSUtC2bVucPXsWycnJGD16NBwdHeHn\n5wegdNk9KCgI69atg4ODA+Lj4+Hn54e3b99i6dKlOHz4MLp164bRo0fD398fAKCrq4uHDx8CAPz9\n/bFixQqYmZlh2bJlGDt2LFJTU6GlpVUr+9+8eRMTJkzAvHnz4O3tjbt37+Lrr7+u91xgSB/2jC3Z\nw2wuW5i9ZUtD2Zs5chUoKpIcpBSJ6h+8LCmR3FYo/LCAKJ/Ph76+PuTk5KCurg49PT0AwF9//YWs\nrCzcvHkTrVu3BgDs27cPxsbG2LdvHyZNmgSgdJlz1apV6NOnj5hcU1NT/PzzzwBK88RWrVqF58+f\nc1Enc3NzrF69GiEhIZwj5+3tLSZj+/bt0NXVxbVr19CrVy/o6uoCKH09WpmeAJCYmAhtbW2oqalV\nGp+xsTHevXuH1NRUmJubV2kHFRUVbN26FfLy8rCwsMDAgQNx9epVPH36FAoKCrCwsIC7uztCQkLg\n5+eHwsJCrFy5EocPH4a7uzsAwMjICD/++CO+/vprLF26FNra2pXsWp4lS5ZwbX/55RcEBQUhJiYG\ngwYNwp49e6q0/99//w1PT0+sWrUKvXr1wrJlywAAnTp1wrNnz8TyBBkMBoPBqIkW7chV9Yqu6lBQ\nKJFYLicnubw28PmS2yoq1l9mdcTFxcHa2ppzIgBAT08PFhYWiI+PF6tbMRLG4/Fgb28vVta2bVu0\na9euUllmZiZ3fOvWLQQGBiI2NhZZWVlcLlxqaip69epVpa55eXnQ0NCQeE5TUxMAkJubW2V7ALC0\ntBTL7dPX14eFhYVYFE9fXx/3798HUGqfN2/eYNSoUWJ5cCKRCO/evUN2djZ0dHSq7dPBwYH7W09P\nD3JycsjIyODkV2X/uLg4AKVLs4MGDRKTWdGhZjQtWKRC9jCbyxZmb9lSZu8PfUVXi3fk6oqbmxmC\ngkIgELxfCn33LgTe3uawsKifHgkJpTKVlMRlDhxYdZTpQ5G0qaBimZycHBQVFSvVq7iMyePxJC5t\nlpSUOqKFhYVwd3eHi4sLgoKCoK+vDyKCtbU1hEJhtXpqaWnh1atXEs/l5eVxdaqj4gYNHo8nsaxM\n37J/Dx48iM6dO1eSp62tXW1/ACTarUwuULP92QN8GQwGgwGACzjV9ykJbLNDBSwsjODtbQ49vQvQ\n0gqDnt6F/zlx9c9lk4bM6rCxsUF8fDyys7O5soyMDCQmJsLGxqZB+igfybp37x6ysrKwbNkyuLi4\nwMLCAjk5OWKOSpnjU7aZoIxOnTrh5cuXKCgoqNRHamoqlJSUatyNW9fdpdbW1lBWVkZycjJMTU0r\nffh8PqdzRX1rQ23sb2VlVel5cRcvXqxzXwzZwZ6xJXuYzWULs7dsaSh7t+iIXH2xsDBqcCeroWRu\n2LABGze2WKy+AAAgAElEQVRuxL1797iyipGdCRMmYOnSpRg7dixWrlyJkpIS+Pv7w9DQEGPHjq1W\nPhFVkldTmZGREZSUlLB+/Xp8++23SElJwfz588UcLF1dXairq+PMmTOwtLSEkpIStLW10atXL8jL\nyyMmJgaurq5ifVy+fBm9evWSGP2qqEtdUFdXx4IFC7BgwQLweDwMHDgQxcXFuHPnDm7duoVffvkF\nAGBiYoKoqCikpaVBRUWlxuXWMmpj/9mzZ8PZ2RkLFy7E5MmTERcXh9WrV9dpHAwGg8FgsIhcMyM7\nOxuJiYliZRUjUsrKyjh79iyUlJTg4uICgUAADQ0NnD59WmzJUVIki8fjVSqvqUxXVxe7du3CuXPn\nYGNjg7lz52LVqlVcZAso3ZixceNG7N+/Hx06dEC3bt0AABoaGhgxYgQOHz5cSZfDhw/D09OzWnvU\nR18AWLhwIVavXo2tW7fCwcEB/fr1w7p162BiYsLVCQwMRG5uLiwsLKCvr4+0tDROVnXUxv5du3bF\nnj17sG/fPtjZ2WHFihVYs2YNe3ZdE4blD8keZnPZwuwtWxrK3jxqoYk61eUgsfykpsXFixcxcuRI\npKamQlVVFQAQGRmJ0aNHIyUlRexBwQzJsDnNYDAYzZv63sdZRI7R6PTp0wf9+vXDxo0bubKlS5ci\nMDCQOXGMJgPLH5I9zOayhdlbtrAcuVpQn8ePMBqHQ4cOiR2fO3eukTRhMBgMBkN2fOjjR9jSKoPR\nAmBzmsFgMJo3bGmVwWAwGAwG4yODOXIMBoNRC1j+kOxhNpctzN6ypaHszRw5BoPBYDAYjGYKy5Fj\nMFoAbE4zGAxG86a+9/EWvWuVwWAwGAxG9SQkpOL48WQIhXxoapbAzc1Maq+QZDQ8bGmVwWAwagHL\nH5I9zObSJyEhFVu2JOH8+QE4fBh48mQAgoKSkJCQ2tiqtXhYjtxHikAggK+vb7V1vL29MWjQIBlp\nxGAwGIzmyrFjyYiPH4g3bwChELh9G1BUHIiQkOTGVo1RS1q0I7dkyZIW9z86Se8Rrchvv/2GgwcP\nykijlstPP/0k9u5VaXDhwgVYWlpCU1MTHh4eyMvLEzs/evRorFy5Uqo6MGoHe7C47GE2ly45OcDF\ni3y8fVt6rK0tgIkJwOMBQmGLdg+aBGXzOywsDEuWLKm3nBZ9pcre7PCxoaGhgVatWkm9H6FQKPU+\npEFRUVGT6LOkpATjxo3D1KlTce3aNWRlZWHZsmXc+YMHDyItLQ3+/v6yVJXBYHwEZGcD27cDQmEJ\nAIDPB2xsAF3d0vOKiiWNqN3HhUAgYI5cQ5OQlICNf2/E2n1rsfHvjUhISmhSMkUiEebPn482bdqg\nVatWmD59Ot69e8edr7i0Wnb8xx9/wMjICK1atcKIESPw4sULrs6jR48watQotG/fHmpqarCzs8Ou\nXbvE+hUIBJg2bRoWLVoEAwMDGBkZITAwEF26dKmk45QpU+Dm5lblGIqKijB//nwYGhpCSUkJ1tbW\n2Lt3r1gdPp+P9evX4z//+Q/U1dVhaGiI9evXi9UpKCjA119/DUNDQ6ipqaFr1644fPgwdz4lJQV8\nPh979uzB0KFDoa6ujsWLFwMAfH19YW5uDlVVVZiZmeGHH37gnNOgoCAsXrwYqamp4PP54PP5WLp0\nKQDg1atXmD59OvT09KCsrAxnZ2exV4pV12d5srOzkZWVhVmzZqFz586YMGEC4uPjAQA5OTn47rvv\nsH379hojsAzZ0NKi+80BZnPpkJlZ6sS9egWYmpqhpCQENjbAq1dhAIB370IwcKBZ4yr5EcBy5KRE\nQlICgkKDkKmfidy2ucjUz0RQaNAHOV4NKZOIcPDgQbx8+RJRUVHYvXs3jhw5gu+//56rI2n5NSYm\nBuHh4Th16hTOnDmDO3fuiEV6Xr9+DTc3N5w+fRp3797F559/Dh8fn0oTbf/+/cjOzsaFCxdw/vx5\nTJs2DcnJyYiIiODqvHr1CgcOHMD06dOrHMeCBQvw559/Yt26dYiLi4Onpyc8PT1x4cIFsXqBgYEY\nMGAAbt26hblz5+K7777DsWPHOFsMHz4cd+7cwf79+xEXF4cZM2Zg3LhxleTMmzcPkyZNQlxcHPz8\n/EBE0NfXx969e3H//n2sXbsW27dvx/LlywEA48aNw7x582BoaIj09HSkp6dz9poyZQrOnTuH3bt3\nIzY2Fn369MGnn36KhISEKvuUZAtdXV0YGBjg5MmTKCoqwtmzZ+Hg4AAAmDVrFnx9fWFlZVWlDRkM\nBqOuvHgBBAUBBQWlx+3aGWHxYnN06XIB6uq3oKd3Ad7e5mzXajOCPUeuAhv/3ohM/UyEpYSJlas9\nUYNzX+d66XI16ioKDQvFygTGAui90MMXY76okyyBQIDHjx8jOTmZc9a2bt2KWbNmIScnByoqKvD2\n9sbTp0+5KJG3tzdOnz6NtLQ0KCgoAABWrFiBtWvX4tmzZ1X2NXLkSOjp6eGPP/7g+k5PT8f9+/fF\n6o0YMQKamprYuXMnAGDLli1YvHgxnj59Cnn5yk+4KSwsROvWrbF27Vr4+flx5aNGjUJeXh5CQkIA\nlEbkJk2ahODgYK7OxIkTkZaWhoiICISFhWHIkCHIyMiApqYmV2fKlCl4+fIlDh8+jJSUFJiamuLH\nH3/EDz/8UK1t16xZg02bNiExMRFAaY7cX3/9hUePHnF1kpKS0LlzZ5w8eRKDBw/myrt16wYHBwf8\n9ddfdeozOjoa3377LZ4/fw6BQICNGzciLCwMixYtQkhICGbPno3IyEjY2trijz/+gJ6enkQ57Dly\nDAajJtLTgR07gML//RwpKgITJwJGzGdrErB3rTYQRSQ5f0oEUb1llkByroGwpH45Zt27dxeLuPXu\n3Rvv3r1DcnLVu4y6dOnCOXEA0K5dO2RkZHDHhYWFmD9/PmxsbKCjowMNDQ2cPHkSjx8/FpPTrVu3\nSrKnT5+Of/75h0vU37p1K7y8vCQ6cUCpMyQUCuHi4iJW7uLigri4OLGyXr16iR337t2bqxMTEwOh\nUIj27dtDQ0OD++zevRtJSUli7bp3715Jj61bt6JHjx5o27YtNDQ0sGDBgkrjrUjZ0mdtdJfUZ0V6\n9eqF6OhopKSkICgoCMXFxZg5cya2bduGn3/+GSKRCElJSbCwsMCsWbNqlMdgMBiSePYMCA5+78Qp\nKQGTJjEnriXAHLkKKPAUJJbLQa7eMvlVmFmRr1gvefXx2Ms7cUBlz3/OnDnYvXs3t9P31q1bGDp0\nqFjuHY/Hg5qaWiXZgwcPhp6eHnbs2IFbt27hxo0bNT4ipSEoKSlBq1atEBsbK/a5d+8eTp06JVa3\not4HDhzAV199hfHjx+PUqVO4desWFi9eXO8NHJKuiSRb1cR3332HiRMnwsHBASEhIZgwYQJ4PB4m\nTZqE8+fP10s3RsPA8rVkD7N5w/DkSWkk7s2b0mNl5VInrkMH8XrM3rKloezN3uxQAbdubggKDYKg\nk4Are/fgHbzHecPC3KJeMhMMS3PklDopickc6DqwXvJiYmJQUlICPr/UQbx06RKUlJRgZlZ1cmpN\nCfORkZHw9PTE6NGjAZQ6SQkJCWjXrl2N+vD5fPj6+mLr1q24f/8++vfvj06dOlVZ39zcHEpKSggP\nDxfLAQsPD4etra1Y3ejoaLHl10uXLsHa2hoA4OTkhNzcXLx584Yrqy0RERFwdHTEN998w5WVX0IF\nAEVFRYhE4pHYsn7Cw8MxZMgQMXmSopV14fz587hy5Qpu3LgBoPQalDmWQqEQJSVsFxmDwagbaWnA\nrl1A2f/JVVRKnTgDg8bVi9FwMEeuAhbmFvCGN0JuhEBYIoQiXxEDXQfW24mThszs7Gx8+eWX+Prr\nr5GcnIzFixfDz88PKioqVbapKYpnYWGBI0eOYNSoUVBTU8Pq1avx/PlztG3bVkxGVXKmTp2KwMBA\nJCYmYvv27dX2paqqilmzZmHRokVo06YN7OzscPDgQRw7dqxS1OnEiRPYuHEj3N3dcfr0aezfv597\nRt7AgQPh5uaGUaNGYcWKFbC1tcXLly9x6dIlqKioYNq0aVXq0KVLF2zbtg3Hjh2DtbU1jh8/Lrbb\nFQBMTU2Rnp6Oy5cvw9zcHGpqajAzM8N///tffPHFF9iyZQs6duyITZs2IT4+Hvv27at23NVRUFCA\nGTNmYM+ePVz01MXFBRs2bECXLl2wevXqj/JROk0JZn/Zw2z+YaSmArt3lz7oFwBUVYHJk4Fyt3Ux\nmL1lS4PZm1oo1Q2tOQ9bIBDQ1KlTac6cOaSjo0MaGhrk6+tLb9++5ep4e3vToEGDqjwmItq5cyfx\n+XzuOC0tjT755BNSU1Ojdu3a0ZIlS2jq1Knk6uoq1revr2+Vuo0cOZJ0dXVJKBTWOI6ioiKaP38+\ntW/fnhQVFcna2pr27t0rVofH49G6deto5MiRpKqqSgYGBrRmzRqxOm/evKH58+eTiYkJKSoqUtu2\nbWnIkCEUGhpKRESPHj0iPp9PFy9erNT/9OnTqXXr1qSpqUkTJ06kDRs2iNmkqKiIJkyYQK1btyYe\nj0eBgYFERJSfn0/Tp0+nNm3akJKSEjk7O9O5c+e4dlX1WR1fffUVzZkzR6wsOzubhg8fThoaGtS/\nf39KS0ursn1zntMMBqPhefiQ6KefiAICSj8rVhBlZDS2VozqqO99nO1aZTQY3bt3R79+/bBq1aoG\nkcfn87Fr1y5MmDChQeS1ZNiclj5hYWEsYiFjmM3rR3IysHcvUFxceqyuDnh5AW3aVN+O2Vu2VLR3\nfe/jLXpptezNDmxiSpesrCwcP34cN2/exP79+xtbHQaDwfhoefAA+Pvv906cpmapE6ej07h6Maom\nLCzsgzY+sIgc44Ph8/lo3bo1fvrpJ7GNCQ0hl0Xkageb0wwGIyEB2L8fKNuj1apVqRPXunXj6sWo\nHSwix2g0pLWbku3SZDAYjNoRHw8cPAiU3Ta1tEqdOG3txtWLIX3Yc+QYDAajFrBnbMkeZvPacfeu\nuBPXujXg41N3J47ZW7aw58gxGAwGg/GRc/s2cPgwULYip6NTGokr99ZCRguH5cgxGC0ANqcZjI+P\nW7eAo0ffO3Ft2pQ+J05Do3H1YtQPliPHYDAYDMZHwvXrwPHj7504Pb1SJ05dvXH1YsiejzJHTltb\nGzwej33Yp8V8tFlGs9Rh+UOyh9lcMjExwL//vnfi2rYFvL0/3Ilj9pYtLEfuA8jJyWlsFVok7GGS\nsoXZm8H4+Lh8GTh9+v2xgUHpu1OreUMjo4XzUebIMRgMBoPR3Lh0CTh79v2xoSHg6QkoKzeeToyG\no75+y0cZkWMwGAwGozkRGQmEhLw/7tCh1IlTUmo8nRhNg48yR44hHVh+hWxh9pYtzN6yh9m8lPBw\ncSfOyEg6Thyzt2xhOXIMBoPBYLRgiIDQUCAi4n2ZiQkwfjygqNh4ejGaFixHjsFgMBiMJgYRcP48\ncPHi+zIzM2DcOEBBofH0YkgPliPHYDAYDEYLgKh0U0N09PuyTp2AsWMBeSn8aickJeDo5aMQkQia\nippw6+YGC3OLhu+IIRVYjhyjwWD5FbKF2Vu2MHvLno/R5kTAqVPiTpyFhXSduM1nNyOEQnD4/mGk\n6aQhKDQICUkJDd8ZQ4yGmt8t2pFbsmTJR3kjYDAYDEbzgwg4cQK4evV9maUlMGaMdJw4ADh6+Sji\nNeLxTvQOQpEQtzNuQ9FcESE3QmpuzGgQwsLCsGTJknq3ZzlyDAaDwWA0MiUlpW9ruHnzfZmNDeDh\nAcjJSafPnDc58Fnrg7x2eQAAPo8PGz0btFZpDa10LXwz7hvpdMyQCMuRYzAYDAajGVJSAhw9CsTG\nvi+zswNGjgT4Ulo3yy7MRnBsMIpERQBKnThbPVtoq5S+7k+Rz7bFNhda9NIqQ7awZWzZwuwtW5i9\nZc/HYPOSEuDQIXEnzsFB+k5c0K0g5L/Lh6mpKUqSS2CrZ4u8hNLI3LsH7zCw60DpdM7gYM+RYzAY\nDAajGSMSAf/8A8THvy/r1g349FOAx5NOn1mFWQi+FYxXwlcAgHaG7TCyy0gkJiUiPiceei/0MNB1\nINu12oxgOXIMBoPBYMiY4mLg4EHg/v33Zc7OwNChsnPiFPgKmGg3EcZaxtLpkFEnWI4cg8FgMBjN\ngOJiYP9+IDHxfVnPnsAnn0jPict8nYng2GAUCAsAMCeuJcFy5BgNxseQz9KUYPaWLczesqcl2ryo\nCNi3T9yJ69NHtk6copwiPO08KzlxLdHeTRmWI8dgMBgMRjOiqAjYuxd4+PB9Wb9+wIAB0nXigm4F\n4XXRawClTtxE24kw0jKSTocMmcNy5BgMBoPBkDJCIbBnD5CS8r5MIAD695eeE/fi9QsE3woWc+I8\n7TzRsVVH6XTI+CBYjhyDwWAwGE2Qd++A3buBx4/flw0YALi4SK9P5sR9PLAcOUaDwfIrZAuzt2xh\n9pY9LcHmb98CO3eKO3GDBknXicsoyBBbTlWSU8Iku0k1OnEtwd7NCZYjx2AwGAxGE+bNm1In7tmz\n92WDB5fuUJUW6QXp2BG7A4VFhQBKnThPO090aNVBep0yGhWWI8dgMBgMRgNTWAjs2AGkp78vGzoU\n6N5den1KcuIm2U+Coaah9DplNBgsR47BYDAYjCbA69elTlxGxvuy4cNL39ogLdIL0hF8Kxhvit8A\nAJTllTHJbhLaa7aXXqeMJgHLkWM0GCy/QrYwe8sWZm/Z0xxtXlAABAW9d+J4PGDECOk6cc9fPW8Q\nJ6452rs5w3LkGAwGg8FoQuTnA8HBQHZ26TGPB3h4AHZ20uvz+avn2BG7Q8yJm2w/GQYaBtLrlNGk\nYBE5AD179oSjoyOsra0hLy8PR0dHODo6YsqUKQgPD4ezs/MH92FsbIz48m9GLsewYcPw6NGjD+6j\nsREIBACAH3/8ETY2NujVqxcel9uqNWzYMDws/yTMCvj6+uLixYu17u/Zs2dwdXWFlpZWjddo5MiR\ncHBwgKOjI/r06YOYmBgAQEpKCne9HR0dYWxsDB0dnVrrUMaSJUtQVFRU53ZHjx6Fk5MTbG1tYWNj\ng9WrV1dZ19/fH6ampuDz+YiPj+fsXZ7AwEDufF05cuQIrKys0K1bNySWf+x8LQkPD8e5c+fq3E4S\neXl5WLFihViZQCDAiRMnGkR+bdi0aRMsLS3RrVs3FBQUSLS3NJFkTyJCv3798OTJEwDAmDFjuLnc\nEpG1zT+EvLzSSFyZE8fnA//5j3SduGevniE49n0kTkVe5YOcuOZk75ZAg9mbWij1GVpKSgrp6uqK\nlYWGhpKTk9MH62NsbEx37979YDm1obi4WCb9SCIvL48sLCyopKSEduzYQf7+/kREFBQURMuWLWvw\nvqKioujEiRM1XqO8vDzu76NHj5Ktra3Eet988w3NnDmzzrrweDwqKCioc7srV67Q8+fPOR3Nzc0p\nMjJSYt2oqChKS0sjY2NjiouLq3T++vXrNGTIEDIxMZF4viYGDx5MBw8erHO7MgICArjrXVdEIpHY\n8aNHjyp9FwUCAR0/frze+tUVS0tLunbtWp3bVRxLfZFkz6NHj5K3tzd3fOXKFRo8eHCD9MeoPy9f\nEq1dSxQQUPoJDCSqx1ewTjzJe0I/R/5MAaEBFBAaQL9E/kLP8p9Jt1OGVKmvS8YicuWgKnaLFBcX\nw8/PD/b29nBwcMD9+/e5c8HBwejZsyecnJwwcODAaiMZu3btgpOTEzp16oSNGzdy5eWjdQKBAHPn\nzkW/fv1gZmaG77//nqu3atUqdO/eHV27dkXv3r0RGxvLnePz+QgMDET37t0RGBgIW1tbXLt2jTu/\nevVqTJ8+vZJOIpEI/v7+sLW1ha2tLebMmYOSkhIAgLe3N3x9fdGnTx9YWFjg888/rzbqFBYWBjk5\nOYhEIgiFQhQUFEBJSQnZ2dnYtm0b5s6dW2XbsrGXRVz++OMPWFlZwdHREfb29khISKhUX1NTE336\n9IGqqmq1csvqlpGbmws9Pb1KdYRCIXbv3o0pU6YAAO7fv4+OHTtyUcXAwECMHz++Ursvv/wSANC7\nd284OjoiPz8fGRkZ8PDwgL29Pezs7LBz506JenXv3h1t27bldLS0tBSLYpanT58+MDR8v/usfH7F\nu3fv8NVXX2HTpk1i87i2Y5g9ezaioqIwd+5cDBw4EAAwceJEODs7w87ODqNGjUJubi4AICEhAb16\n9YKDgwNsbW2xatUq3L17F1u2bMGOHTvg6OjIRdNOnjyJvn37wsnJCb1798aVK1c43e3s7DBlyhQ4\nOjri9OnTlWyam5sLR0dH9O3blysPDw+v13fj559/Rvfu3WFmZoZDhw5JtG95xo4di+TkZHh6esLT\n0xMA8P3338POzg729vYYNWoUMjMzAQBBQUFwc3PDqFGjYGtrizt37oDP52P58uXo3r07TE1Ncf78\necydOxeOjo6wtbXl7iHp6ekYMGAAnJycYGNjg3nz5gEA7ty5I9GeW7duFbt+3bt3R2JiIp4+fVrj\nmJojzSFnKycH2L4dePmy9FhODhgzBrCykl6fT/OfYuftnXhb/BbA+0hcO412HyS3Odi7JdFg9m5Q\nd1IG5OXlkbOzM6mrq1cbdajP0CRFAUJDQ0lBQYFu3bpFRETLli2jiRMnEhFRREQEDRs2jN69e0dE\nRCdPnqQ+ffpIlG1sbExTp04lIqKMjAwyMDCgO3fucOfKxiIQCGjcuHHcWHV1dSkpKYmIiDIzMzl5\n586do549e3LHPB6PVqxYwR1v3ryZfHx8iIiopKSEOnXqRLdv366k1++//05ubm5UVFREQqGQBg4c\nSJs2bSIiIi8vL7K3t6fXr19TcXExubu704YNG6q0X2hoKCfTwcGBhgwZQhkZGTRlypQqo0zlEQgE\ndOLECSIiatWqFaWnpxMRkVAopMLCwmr7rU3UdOrUqdSxY0cyMDCge/fuVTp/4MABcnR0FCvbuXMn\n9ezZk86cOUMWFhb06tUribJ5PB69fv2aOx4zZgwtXryYiIieP39OBgYGNUZk7927R23atOEidFVR\nNl/K7E1ENHfuXPr999/Fztd1DOXtT0SUlZXF/f3DDz/Q/PnziYho1qxZ9PPPP3PncnNziYhoyZIl\nNGfOHK48KSmJevXqRfn5+UREdPfuXerYsSMRlV4zOTk5unz5skRdJEXH+/fvX+/vxsaNG4mI6OLF\ni9S+fXuJfVakvB3v3LlDurq63JxctGgRjR07loiItm/fTurq6vTw4UOxPsuux4EDB0hVVZWz7YoV\nK8jT05OIiN6+fctFcoVCIQ0YMIBOnz5NRJXtKRKJSFNTUyy6TEQ0YcIE2rFjR63G1NwoP8ebIllZ\nRKtWvY/ELV1KlJAg3T7T8tJoecRysUjc81fV3zNqS1O3d0ujor3r65I1u80OqqqqOHnyJObMmSOz\n58RZWFjA3t4eANCjRw/8+++/AIB///0XsbGx6NGjB4DSiF5Z1EISU6dOBQDo6elh2LBhCA0NhY2N\nTaV6//3vfwG8j9AkJyfDzMwM165dw/Lly/Hy5Uvw+fxK0T8vLy/ub09PTyxduhQvX77ElStX0LZt\nW9ja2lbqKyQkBD4+PpCXL50KPj4+OHz4MPz8/MDj8TB27Fgu4uXl5YV//vmHi0BVpGy9f8aMGZgx\nYwYAICIiAnJycrCysoKPjw9evXqFMWPGYMyYMVXaCQAGDBiAyZMnY/jw4Rg2bBhMTEyqrV8b/vzz\nTwClkdFRo0ZVyiPbtm0bF40rw9PTE+fPn4eHhweioqKgrq5eq75CQkKwZs0aAEDbtm0xdOhQhIaG\nwtraWmL958+fY+TIkdi0aRMXoauJMntHR0fj+vXr+PXXX7lz5b8bdRlD+XbBwcHYs2cPhEIhXr9+\nDQsLCwBA//79MXfuXBQWFsLV1RWurq4S2585cwbJyclwKfcIe5FIxEWyOnXqxH13qtOjDB6PV+/v\nxrhx4wCUfn+fPXsGoVAIRUXFKu1QkdDQUHh4eEBfXx8AMH36dO6eAAB9+/atNEfHjh0LAHB0dISc\nnByGDh0KAOjatSsXFSwuLoa/vz+io6NBREhPT0dsbCw++eQTEJGYHbKyslBSUiIWXQYAQ0PDanNP\nmzNNOWcrK6t0Y8OrV6XH8vLAuHGAubn0+nyS/wQ7Y3finegdAEBVQRWT7SejrXrt7hk10ZTt3RJp\nKHs3O0dOXl4eurq6Mu1TWVmZ+1tOTg7FxcXc8ZQpUxAYGFgrOeVvykQEXhVvSpbUn1AoxOjRoxEV\nFQUHBwc8e/ZMbJkNgNgPtJqaGiZMmIBt27YhPDy8SuerJr0qnqsLQqEQixYtwpEjR7B69Wq4urpi\n4sSJsLe3x4gRI6CkpFRl20OHDiEmJgYXLlyAq6srNm/ejMGDB0usW5Udq8LT0xOff/45cnJy0Lp1\nawDA06dPERERgd27d1caQ1xcHLS1tZFe/smetaC21/vFixcYNGgQ5s2bh//85z916gModZbv3bvH\nORJPnjzBJ598wi351WUMZTpGRkZi8+bNiI6Oho6ODvbs2YOtW7cCAEaNGoXevXvjzJkz+OWXX7Bt\n2zbs3LlT4vwYPHgwgoODJfZVW6e4PPX9bpS1k5OTA1DqQNXFkav4oM6KY5U0lvJ9lp/r5e8hq1ev\nRm5uLq5evQpFRUVMnz4db9++rbVeknRjSJ8XL0qduNelb8CCggIwfjxgaiq9PtPy0rDr9i4xJ87L\n3gv66vrS65TRLGA5ch/A8OHDsWPHDi4/RSQS4fr16xLrEhGCgoIAAJmZmTh16pRYJKNi3Yq8ffsW\nIpGI+4H6/fffa9Tvyy+/xNq1a3Hjxo0qHQQ3NzcEBwejuLgYRUVFCA4OxqBBgzg9Dhw4gMLCQhQX\nF2Pnzp1c/pQkKq73//rrr5g2bRq0tbVRWFjIlRcVFUEoFFYpRyQSITk5Gc7Ozpg3bx7c3d1x69at\nKnhUMYsAACAASURBVOvX9CP2+vVrpKWlccf//vsvDAwMOCcOKI0+ffrpp9DW1hZrO2fOHDg7O+Ps\n2bPw8/OrMhdJQ0NDLBrr5ubGOT7p6ek4deoUBgwYUKlddnY2Bg0ahJkzZ8LHx6facZSHiDh7z5s3\nD0+fPsWjR4/w6NEjGBoa4uzZs3Bzc6vTGMqTm5uLVq1aoXXr1nj37h22bdvGnUtKSoKenh68vLyw\nePFibtdkq1atkJeXx9Vzd3fH6dOnxSKftd1hqampicLCQohEokrjrkh9vht1xdXVFUeOHEHG/x4O\ntnXrVri7u3+w3Ly8PLRr1w6Kiop4+vQpjh49yp2raE9dXV3weDy8KgsB/Y8nT57AVJoeRCPSFHO2\n0tNLd6eWOXGKisDEiS3DiWuK9m7JNJS9G82R27BhA5ycnKCsrFzpBywnJwceHh5QV1eHsbEx9u7d\nK1FGXSMxtaGiTB6PJ1ZW/rhfv35YtmwZPvvsMy7xu2zZVZLcNm3acEnfCxYsqHKZTdK4NDU1sXTp\nUjg7O8PJyQnq6uqV9KqIsbExLC0tMWXKFG7ptCKff/457Ozs4OjoiK5du8LBwQG+vr6cTGdnZ7i7\nu8PKygpGRkb4/PPPAQBbtmxBQECARJlA6Y/95cuXMWnSJAClTuXGjRthZ2eHyZMnQ0NDo8q2IpEI\nPj4+sLOzg4ODA9LT06vcqGFoaIgxY8bg9u3b6NChA5YuXQoAuHbtGoYNGwYAKCgowJgxY7hxbty4\nUewHEyh15Couqx45cgQRERFYu3YtrKysEBAQgPHjx3ObQcrz3XffYcCAAejatSvy8/Oxfv16xMbG\nwt7eHu7u7vj1119haWlZqd0vv/yCpKQkbN68mXsESlkEq/wYAGDWrFno0KEDnj59Cjc3t0r6SqIu\nYyjPkCFDYGZmhs6dO0MgEKBbt27cHDtw4ADs7OzQtWtXzJo1C+vWrQMAeHh4ICYmhkvONzc3x65d\nuzB16lQ4ODjAysqKc26B6r+/rVu3xsSJE2Frayu22aEhvhtV9fvvv/9yc78i1tbW8PX1xaBBg2Bv\nb487d+5w4654j6ipz/L1Z82ahYsXL8LW1hbTpk3jnG+gsj35fD5cXFwQHR0tJvvy5ctV/qeQ0bA8\nf14aiSv7f6mSEuDpCRgbS6/Px3mPsfP2++VUNQU1eDt4s0gcg6PR3rV6+PBh8Pl8nDlzBm/evMH2\n7du5c2W7sv766y/cvHkTw4YNw6VLl2BVbhuQj48P/P39q3WGPvblhvz8fFhaWuLatWto167uu5l8\nfHzg5ORU7bIsg8GQHUeOHMHRo0e5++XVq1exePHiSrt+GQ3P06fAzp1A2cq3snKpE2coxdeYPs57\njF23d0EoKl3BUFNQg5eDF/TUKu+6ZzR/6uu3NFpEzsPDAyNGjKj08NXXr1/j0KFD+PHHH6Gqqoo+\nffpgxIgRYo9vGDp0KM6ePQtfX98q828+djZv3gxra2v4+/vXy4krQxpRTwaDUT9GjhyJ5ORkbnn8\n//7v/7goNEN6pKWVvju1zIlTUQEmT5auE5eamyrmxKkrqsPbwZs5cYxKNPpmh4reZ2JiIuTl5WFe\nbuuPvb292FryyZMnayXb29sbxv+LeWtpacHBwYHbJVImr6Ued+nSBTt37vwgeV5eXnWqf+vWLXzz\nzTdNYvwfwzGz98dp74iICISFheHBgwfYv39/o+sjzeOyssbUJzUV+PHHMBQXA8bGAqiqAubmYUhM\nBAwMpNP/vuP7cP7heRjalXqKGXcz4GDmgDZqbaQ63rKypnL9W/rxrVu3kJubi5SUFHwIjba0Wsai\nRYvw5MkTbqkgMjISY8aMwfPnz7k6W7duxZ49exAaGlpruWxpVfaEhYVxE5UhfZi9ZQuzt+xpbJun\npAC7dwNlz0FXUyuNxOlLMT0tJTcFu2/vRlFJaadlkThdVek/raGx7f2xUdHe9fVbmlxETl1dHfn5\n+WJleXl51SbHM5oG7AYgW5i9ZQuzt+xpTJs/fAjs3fveiVNXB7y8gDZtpNfno5ePsOfOHs6J01DU\ngJeDl0ycOIDNcVnTUPbmN4iUD6BiDlbnzp1RXFyMpKQkriw2Nlbig3MZDAaDwWhokpKAPXveO3Ea\nGoC3t+ydOFlF4hjNm0Zz5EQiEd6+fYvi4v9n786jqrqyxI9/HzPIKCiKMjiLE6Y0pmLU4BCTStQE\nYxwZ1HQ63Um5KpWq7qxOYsSkq+rXvWrqVP9+K11JOwDOGstojMaIzyFqTFRAEVFEQREHUEBApvfe\n749bvOczmjzgncu0P2uxynsE9nHXDW7vPWefBkwmE7W1tZhMJrp06cLMmTN57733qK6u5tChQ2zf\nvt3axqIpkpOT7d79C7Uk1/qSfOtL8q2/1sh5bq72JK6x73tAACxaBCr70OffzmfNqTXfK+KCfYJ/\n5CudS+5xfTXm22g0kpyc3Ozv02qFXOOu1P/4j/8gLS0Nb29vfvOb3wBaQ8+7d+/SvXt34uPj+eij\njx7Yg+vHJCcny6NiIYQQDsnJgQ0boLEPdWCg9iTunt7hTpd/O5+1p9bSYNYqR39P/1Yp4kTriY2N\nbVEh1+qbHVSRzQ5CCCEclZ0NW7ZAY6/soCCtiAsIUBfzwq0LrDu97ntFXFdvhZWjaLPa7WYHIYQQ\nojWdOgWffgqNf4cGB2sbG/z91cXMu5XH+tPrrUVcgGcASSOTpIgTTfbQQs7RNWmenp588sknTpuQ\nMzW+WpXXq/qQrev6knzrS/KtPz1ynpEB27bZiriQEK2IU9ko4UFF3MKRCwnyDvqRr1RL7nF9Nebb\naDS2aH3iQwu5jRs38vbbbz/0MV/jI8A//OEPbbqQE0IIIR7kxAnYvt1WxHXvrvWJ8/VVF/N86XnW\nn16PyaItxAv0CiQpJqnVizjRehofOC1fvrxZX//QNXL9+vXjwoULP/oNBg0aRG5ubrOCqyRr5IQQ\nQjzMd9/Bjh226x49ICFBa/qryrnSc2w4vcGuiFs4ciGBXoHqgop2o7l1i2x2EEII0al88w188YXt\numdP7Umct7e6mFLEiR/T3LqlWe1H8vPzW3w2mOh4pAeRviTf+pJ8609Fzo8csS/ievXS1sSpLOJy\nS3Ltirggr6A2WcTJPa4vZ+XboUJu7ty5HD58GICVK1cydOhQhgwZ0mbXxgkhhBD3O3QIdu+2XYeH\na69TvbzUxTxbcpaN2RvbfBEn2i+HXq1269aNoqIiPDw8GDZsGP/zP/9DYGAgzz//vN1RWm2JwWBg\n2bJlsmtVCCEE+/fDvn2268hImD8fPD3VxTxbcpZN2Zu+V8QFeClsTifancZdq8uXL1e3Ri4wMJCy\nsjKKiooYM2YMRUVFAPj5+XHnzp2mz1oHskZOCCGExQJGo1bINerTB+bNAw8PdXFzbuaw6cwmzBat\nw3BX764sHLkQf0+FzelEu6Z0jVxMTAy/+93veP/993nuuecAuHLlCgEqW16LdkfWV+hL8q0vybf+\nWppziwX27rUv4vr1057EqSziztw80y6LOLnH9aXrGrn//d//JSsri5qaGj744AMAjhw5woIFC5wy\nCSGEEMKZLBb48kttXVyjAQO0J3Hu7urinrl5hs1nNluLuGDv4HZRxIn2S9qPCCGE6FAsFti1S2sz\n0mjQIHjpJXBTeDBl9o1stuRs+V4R5+ep8JgI0WEoP2v14MGDnDx5kjt37liDGQwG3n777SYH1Ysc\n0SWEEJ2LxQKff641/G0UHQ2zZoGrq7q4p2+c5tOcT61FXIhPCEkxSVLEiR/V0iO6HHoit2TJEjZu\n3Mj48ePxvq/ZTmpqarODqyRP5PQn5/TpS/KtL8m3/pqac4tFO3LrxAnb2NChMHOmFHGOkHtcX/fn\nW+kTubS0NLKzswkLC2tyACGEEEI1sxm2bYPMTNvYiBHwwgvg0qzW9445df0Un+Z8igXtL+BuPt1I\nGpmEr4fCA1uFuIdDT+RGjBhBeno6ISEheszJKeSJnBBCdA5mM2zdCqdO2cZGjoQZM9QWcVnXs9ia\ns1WKOOEUSs9a/fbbb/ntb3/L/PnzCQ0Ntfu9CRMmNDmoHqSQE0KIjs9kgk8/hexs29hPfgLTp4PB\noC7u/UVc9y7dSYxJlCJONJvSPnLHjx9n586d/PM//zMLFiyw+xCikfQg0pfkW1+Sb/39WM5NJti8\n2b6Ie/RR9UVc5rXM7xVxSTHt/0mc3OP6cla+HVoj984777Bjxw6eeuoppwQVQgghWqKhATZuhHPn\nbGM//Sk8/bTaIi7jWgbbzm6zFnGhXUJJjEmki0cXdUGF+AEOvVqNiIggLy8PD5WtsJ1MXq0KIUTH\nVF8PGzbAvUd9jx0LTz2ltog7WXySz3I/syvikkYm4ePuoy6o6DSUvlp9//33eeONNyguLsZsNtt9\ntGXJycnyqFgIITqQ+npYt86+iBs/Xv8irodvDynihFMYjUaSk5Ob/fUOPZFzeci2H4PBgMlkanZw\nleSJnP6kB5G+JN/6knzr7/6c19XB2rVw6ZLtc2Jj4ckn1RZxJ4pP8FnuZ9brHr49SIxJ7HBFnNzj\n+tK1j1x+fn6Tv7EQQgjhLLW1sGYNFBbaxiZNAtWNE45fPc72c9ut1z19e5IYk4i3u/cPfJUQ+pGz\nVoUQQrRpNTWQlgZXrtjGnnoKnnhCbVwp4oSenL5GbunSpQ59g2XLljU5qBBCCOGIu3chNdW+iHv6\nafVF3HdXv7Mr4sL8wqSIE23SQ5/I+fr6kpWV9YNfbLFYGDVqFGVlZUom1xLyRE5/sr5CX5JvfUm+\n9bdrl5GCgliKi21jzz4LY8aojftt0bd8fv5z63VjEefl5qU2cCuTe1xfytfIVVdX079//x/9Bp6e\nnk0OKoQQQjxMbm4Bn39+gW3bsujSxUzfvv0ICYlk+nQYNUpt7GNFx9h5fqf1updfLxJiEjp8ESfa\nL1kjJ4QQos3IzS3g44/zyMmZTFWVNmYy7eWXv+zPzJmRSmN/c+Ubvsj7wnrd27838SPipYgTulDa\nR669kj5yQgjRvuzYcYHsbFsRBzB06GSuXbugNK4UcaK1tLSPXIcv5OR9v36kaNaX5Ftfkm/1ysrg\n0CEX7t5tvDYSHQ09ekBdnbq/ro5eOWpXxIX7h5MwovO9TpV7XF+N+Y6NjW1RIedQHzkhhBBCpdJS\nWL0aamu1E4MMBoiKgtBQ7fc9PNScJHTk8hF2X9htvQ73Dyd+RDyebrL+W7QPskZOCCFEq7pxA1JS\noLISSkoKyMrKIyZmMsHB2u/X1u5l4cL+DBrk3DVyhy8f5ssLX1qvIwIiWDB8gRRxolU0t25xqJC7\nceMG3t7e+Pn50dDQQEpKCq6uriQkJDz0+K7WJoWcEEK0fcXFWp+46mrt2t0dHnusgHPnLlBX54KH\nh5nJk/tJESc6PKWbHaZNm0be308ofuedd/jDH/7An/70J958880mBxQdl6yv0JfkW1+Sb+e7ckV7\nndpYxHl6Qnw8TJkSyWuvTWLkSHjttUlOL+K+LvzaroiLDIiU16nIPa43Z+XboTVy58+fZ+TIkQCk\npaVx+PBh/Pz8GDJkCH/+85+dMhEhhBCdR0GBdnZqXZ127eUFCQnQq5fauIcKD/FV/lfW68iASBaM\nWICHq4fawEIo4tCr1ZCQEK5cucL58+eZO3cu2dnZmEwmAgICqKys1GOeTSavVoUQom26cAHWr4f6\neu3axwcSE7XdqSodLDjI3ot7rddRgVHMHz5fijjRJjj9ZId7PfPMM8yePZvS0lLmzJkDwJkzZ+jd\nu3eTAwohhOi8cnNh40YwmbRrPz+tiOvWTW3cAwUHSL+Ybr3uE9iHecPnSREn2j2H1sh98sknPPfc\nc/zDP/wDb7/9NgClpaUt6nsiOh5ZX6Evybe+JN8tl50NGzbYiriAAFi06OFFnLNyvv/S/u8VcfIk\n7vvkHteXrmvkvLy8ePXVV+3GpNGuEEIIR2Vmwt/+Bo1vjoKCICkJAgPVxt1/aT/7Lu2zXvcN6su8\nYfNwd3VXG1gInTx0jVxCQoL9JxoMAFgsFuuvAVJSUhROr/kMBgPLli0jNjZWik4hhGhFx4/Djh22\nIi4kRHud6u+vNq7xkhHjJaP1ul9QP+YOmytFnGhTjEYjRqOR5cuXO7ePXHJysrVgKykpYfXq1Uyf\nPp3IyEgKCgrYsWMHSUlJfPjhhy37Eygimx2EEKL1HT0Ku3bZrkNDtSKuSxd1MS0WC8ZLRvYX7LeO\nSREn2jqlDYGnTp3K0qVLGT9+vHXs0KFDvP/++3z55Zc/8JWtRwo5/RmNRnn6qSPJt74k30138CDs\ntW0SJSxMazHi7e3Y1zcn5w8q4vp37c+coXOkiPsRco/r6/58K921evToUX7605/ajT322GMcOXKk\nyQGFEEJ0bBYL7NsHBw7YxiIiYP58rV+curgW9l3ax4ECW+D+Xfszd9hc3FzkaHHRMTn0RO7JJ5/k\n0Ucf5YMPPsDb25vq6mqWLVvGN998w4F7/0ttQ+SJnBBC6M9igT174PBh21ifPjBvHngo3CRqsVhI\nv5jOwcKD1rEBXQcwZ9gcKeJEu6D0iK5Vq1bx9ddf4+/vT/fu3QkICODQoUOsXr26yQGFEEJ0TBYL\n7NxpX8QNGKA9iVNdxO29uNeuiBsYPFCKONEpOFTI9enThyNHjnDhwgU+++wz8vLyOHLkCH369FE9\nP9GOSA8ifUm+9SX5/mFmM2zbBt9+axuLjoY5c8C9mUvTHMm5xWLhq/yvOFR4yDo2MHggs4fOliKu\nieQe15eufeQaeXl50b17d0wmE/n5+QD07dvXKRMRQgjRPplMsHUrnD5tGxs+HOLiwMWhxwXNY7FY\n2JO/h8OXbY8ABwUP4qWhL0kRJzoNh9bI7dq1i5dffpni4mL7LzYYMDW26G5jZI2cEEKo19AAmzfD\n2bO2sUcegenT9S/iBocM5qUhL+Hq4qousBCKKF0j99prr7F06VIqKysxm83Wj7ZaxAkhhFCvvh7W\nr7cv4saMgRkz1BdxX174Uoo4IXCwkCsrK+PVV1/Fx8dH9XxEOybrK/Ql+daX5NteXR2sXQt5ebax\nsWPhZz+Dew7/aZEH5dxisbD7wm6OXLG1v4oOiZYizgnkHteXs/LtUCH38ssvs2LFCqcEFEII0b7V\n1EBqKly8aBuLjYWnnnJeEfcgFouFXXm7OHrlqHVsSLchzBoyS4o40Wk5tEZu3LhxHDt2jMjISHr0\n6GH7YoNB+sgJIUQnUl0NaWlw9aptbMoUGDdObVyLxcIXeV9wrOiYdWxItyG8GP2iFHGiQ1B6RNeq\nVaseGjQpKanJQfUghZwQQjhXZaX2JO76ddvYz34Gjz2mNu6Dirih3YYyM3qmFHGiw1BayLVHUsjp\nT87p05fkW1+dPd8VFZCSAiUl2rXBANOmwahR6mIajUaefPJJdp7fybdXbQ3qhnUfxszombgYFO6o\n6IQ6+z2uN2edterQfwUWi4UVK1YwceJEBg4cyKRJk1ixYoUUSkII0QmUlcHKlbYizsVF6xGnsogD\n7e+ez89/LkWcED/AoSdyv/nNb0hJSeFXv/oVERERFBYW8qc//YkFCxbw7rvv6jHPJjMYDCxbtozY\n2Fj5F4YQQjRTaSmsXq09kQOtiJs1C4YMURczNy+XPd/tIeN6BlfvXKVv376EhIUwvPtw4qLjpIgT\nHYrRaMRoNLJ8+XJ1r1ajoqLYv38/kZGR1rGCggLGjx9PYWFhk4PqQV6tCiFEy9y4ob1OrazUrt3c\nYPZsGDhQXczcvFxW7VvFpaBLFFdqTegb8hqYPX42P5/6cyniRIel9NVqdXU1ISEhdmPBwcHU1NQ0\nOaDouKQHkb4k3/rqbPkuLoZVq2xFnLs7zJ+vtogD+PK7L7kYdJHiymLKzpYB0CumFy5lLlLEKdbZ\n7vHWpmsfuWeeeYb4+HjOnj3L3bt3ycnJITExkaefftopkxBCCNF2XLmivU6trtauPTwgPh5UH61t\ntpg5fu041yqvWcdCu4QyOGQwDZYGtcGFaKccerVaXl7OkiVL2LBhA/X19bi7uzN79mz+8pe/EBgY\nqMc8m0xerQohRNMVFMCaNdrJDQBeXpCQAL16qY1rMpvYkrOFVVtXUd1bqyB7+PZgUPAgDAYD3W90\n57XZr6mdhBCtSJf2IyaTiZKSEkJCQnB1bdu9e6SQE0KIprlwQTs7tb5eu/bxgcREuKcPvBIN5gY2\nZm/kXOk5Sq6WkHEmg4hHIhjQdQAGg4Ha87UsnLiQQf0HqZ2IEK1I6Rq51atXk5mZiaurK6Ghobi6\nupKZmUlqamqTA4qOS9ZX6Evyra+Onu/cXO3s1MYiztcXFi1SX8TVmepYe2ot50rPARASFsLCiQt5\nouEJSveV0v1GdynidNLR7/G2xln5dnPkk5YuXUpGRobdWO/evZk+fToJCQlOmYgQQojWkZ0NW7aA\n2axdBwRoT+KCg9XGrW2oZe2ptRSUF1jHJkROYGLURAwGgzSoFcIBDr1aDQoKoqSkxO51akNDA8HB\nwZSXlyudYHPJq1UhhPhxmZnwt79B44/LoCBISgLVy59rGmpIy0rjSsUV69ikPpOYEDlBbWAh2iil\nr1ajo6PZvHmz3djWrVuJjo5uckAhhBBtw/Hj9kVcSIj2OlV1EVddX83qjNV2RdzT/Z6WIk6IZnCo\nkPvP//xPXnnlFV588UX+5V/+hZkzZ/Lyyy/z+9//XvX8RDsi6yv0JfnWV0fL99GjsH27rYgLDdWK\nOH9/tXEr6ypZlbHK2uwX4LkBz/F4+OPf+9yOlvO2TvKtL137yI0bN45Tp04xevRoqqurGTNmDNnZ\n2YwbN84pkxBCCKGfQ4dg1y7bdVgYLFwIXbqojVtRW8HKkyu5UXUDAAMGnh/0PI/2elRtYCE6sCa3\nH7l+/TphYWEq5+QUskZOCCHsWSxgNML+/bax8HBYsEDrF6dSWU0ZqzNWc7vmNgAuBhfiBscxPHS4\n2sBCtBNK18jdvn2b+fPn4+3tTf/+/QH47LPPePfdd5scUAghhP4sFtizx76I69NHa/aruogrrS5l\nxckV1iLO1eDKS0NekiJOCCdwqJD7p3/6J/z9/SkoKMDT0xOAxx9/nPXr1yudnGhfZH2FviTf+mrP\n+bZYYOdOOHzYNjZggHZ2qoeH2tg3q26yMmMlFbUVALi5uDFn2Byiu/34Zrn2nPP2SPKtL137yO3d\nu5fi4mLc3d2tY926dePGjRtOmYQQQgg1zGZtU8PJk7ax6Gh48UVwc+hvgOa7VnmNlMwUquu1I7fc\nXdyZN3wefYMUH9oqRCfi0Bq5/v37c+DAAcLCwggKCuL27dsUFhYydepUzp49q8c8m0zWyAkhOjuT\nCbZuhdOnbWPDh8MLL4DqUxaLKopIzUqlpqEGAA9XDxYMX0BkYKTawEK0U0rXyP3DP/wDs2bNIj09\nHbPZzJEjR0hKSuLVV19tckAhhBDqNTTApk32Rdwjj0BcnPoirrC8kJTMFGsR5+XmRWJMohRxQijg\nUCH31ltvMWfOHH7+859TX1/PokWLeP7553njjTdUz0+0I7K+Ql+Sb321p3zX18P69XDvC5MxY2DG\nDHBx6Kd+8128fZHUzFRqTbUA+Lj7kBSTRG//3k3+Xu0p5x2B5Ftfuq6RMxgM/OIXv+AXv/iFU4IK\nIYRQo64O1q2DixdtY2PHwlNPgcGgNvb50vNsyN5Ag7kBAF8PXxJjEunepbvawEJ0Yg6tkUtPTycq\nKoq+fftSXFzMW2+9haurK7/73e/o0aOHHvO089Zbb3HkyBGioqJYsWIFbg9YsStr5IQQnU1NDaxZ\nA5cv28aefBJiY9UXcTk3c9h8ZjMmiwkAf09/kmKSCPYJVhtYiA5C6Rq51157zVosvfnmmzQ0NGAw\nGPjHf/zHJgdsqczMTK5evcqBAwcYPHjw986AFUKIzqi6GlJS7Iu4KVNg4kT1RdzpG6fZdGaTtYgL\n9Apk0chFUsQJoQOHCrmrV68SERFBfX09u3fv5n/+53/46KOP+Prrr1XP73uOHDnC008/DcAzzzzT\nKnMQDybrK/Ql+dZXW853ZSWsXg1Xr9rGfvYz0OMUxYxrGWw5swWzxQxAsHcwi0YuIsg7qMXfuy3n\nvCOSfOtL1zVy/v7+XLt2jezsbIYOHYqfnx+1tbXU19c7ZRJNcfv2bXr27Gmd161bt3SfgxBCtBUV\nFdqTuJIS7dpggGnTYNQo9bG/u/odO87tsF538+lGYkwifp5+6oMLIQAHn8gtWbKEMWPGMH/+fF57\n7TUAvv76a6Kjf7wz98P893//N6NHj8bLy4tFixbZ/d6tW7eIi4vD19eXqKgo1q1bZ/29wMBAKiq0\nDuHl5eV07dq12XMQzhUbG9vaU+hUJN/6aov5LiuDlSvti7i4OH2KuKNXjtoVcT18e7Bw5EKnFnFt\nMecdmeRbX87Kt0NP5N566y1eeOEFXF1drWet9u7dm08++aTZgXv16sXSpUvZvXs3d+/etfu9119/\nHS8vL27cuMHJkyd57rnniImJYciQIYwdO5Y//vGPJCQksHv3bsbp8e5ACCHamNJS7Ulcebl27eIC\ns2bBkCHqYx8sOMjei3ut1738ehE/Ih5vd2/1wYUQdhzuKDRo0CBrEQcwcOBAhg9v/oHHcXFxPP/8\n8wQH2y+Graqq4tNPP+WDDz7Ax8eHJ554gueff57U1FQAYmJiCA0NZcKECeTk5PDiiy82ew7CuWR9\nhb4k3/pqS/m+cUN7EtdYxLm5wdy56os4i8XCvov77Iq4iIAIEmMSlRRxbSnnnYHkW1/K18gNHjzY\nevxWeHj4Az/HYDBQWFjYogncv9X23LlzuLm52RWNMTExdn/g//zP/3Toey9cuJCoqChAeyU7cuRI\n66PMxu8n1867zsjIaFPz6ejXku/Ome/iYkhONlJbC1FRsbi7Q58+Rq5ehYED1cW3WCzUR9Rz2uhz\nHgAAIABJREFU+PJhLmVcAmBi7ETmDZ/H4YOHlfx5G7WF//87w3WjtjKfjn6dkZGB0Wjk0qVLtMRD\n+8gdPHiQ8ePH2wV9kMaJNdfSpUu5cuUKK1eutMadPXs2xcXF1s/5+OOPWbt2Lfv27XP4+0ofOSFE\nR3PlCqSlaf3iADw8YMECiFR88pXFYuGLvC84VnTMOjag6wBmD52Nu6u72uBCdBLNrVse+kSusYiD\nlhdrP+T+Sfv6+lo3MzQqLy/Hz092QQkhOq+CAq3Zb12ddu3lBfHx0LvpJ181idliZse5HZwoPmEd\niw6J5sUhL+Lm4tAyayGEQg/9r3Dp0qUPrQ4bxw0GA++//36LJmC4r1PlwIEDaWhoIC8vz/p6NTMz\nk2HDhrUojlDPaDQqLfqFPcm3vloz3xcuaGenNnZ88vGBxERQfbCO2WJma85WTt04ZR0b1n0YcYPj\ncHVxVRscucf1JvnWl7Py/dBC7vLly98rsu7VWMg1l8lkor6+noaGBkwmE7W1tbi5udGlSxdmzpzJ\ne++9xyeffMKJEyfYvn07R44caXKM5ORkYmNj5cYUQrRbubmwcSOYtEMT8PWFpCTo1k1tXJPZxJac\nLZy5ecY6NrLHSGYMmoGLwUVtcCE6EaPR+INL2H6MQ2etqpCcnPy9p3nJycm899573L59m8WLF7Nn\nzx5CQkL4P//n/zB37twmfX9ZIyeEaO+ys2HLFjBrhyYQEKA9iQtWfPJVg7mBjdkbOVd6zjr2aNij\nPDvg2Rb9A14I8XDNrVseWsjl5+c79A369u3b5KB6kEJOCNGeZWXB1q3Q+GMsKEh7EhcYqDZunamO\n9afXk3/b9nfA470fZ2q/qVLECaFQc+uWhz4f79+//49+DBgwoEWTFh1LSx4Ni6aTfOtLz3wfP25f\nxIWEwKJF6ou42oZa1mStsSviJkROaLUiTu5xfUm+9eWsfD90jZy58Vm+EEII3Rw9Crt22a5DQyEh\nQVsbp1JNQw1pWWlcqbhiHZvUZxITIieoDSyEaJFWWyOnmsFgYNmyZbLZQQjRbhw6BF99ZbsOC9OK\nOG/FJ19V11eTmplKcaWtf+fT/Z7m8fDH1QYWQlg3Oyxfvty5a+Sefvppdu/eDdj3lLP7YoOBAwcO\nNDmoHmSNnBCivbBYwGiE/fttY+HhWrNfLy+1sSvrKknJTOFG1Q3r2HMDnuPRXo+qDSyEsOP0hsCJ\niYnWX7/88ssPDSpEI+lBpC/Jt75U5dtigT174PBh21ifPjBvnnZyg0oVtRWszlhN6d1SAAwYmDFo\nBo/0fERtYAfJPa4vybe+lPeRW7BggfXXCxcubHEgIYQQ9iwW2LkTvv3WNta/P8yZA+6KT74qqylj\ndcZqbtfcBsDF4ELc4DiGhw5XG1gI4VQOr5E7cOAAJ0+epKqqCrA1BH777beVTrC55NWqEKItM5th\n+3Y4edI2Fh0NL74IbopPviqtLmV15moqarXjEF0NrswaMovobtFqAwshHsrpr1bvtWTJEjZu3Mj4\n8ePxVr3q1onkZAchRFtkMmntRU6fto0NHw4vvACuik++ull1k9WZq6msqwTAzcWN2UNnMzB4oNrA\nQogH0uVkh6CgILKzswkLC2t2IL3JEzn9yfoKfUm+9eWsfDc0aKc15OTYxh55BKZPBxfFJ19dq7xG\nSmYK1fXVALi7uDNv+Dz6BrXNxu5yj+tL8q2v+/Ot9IlceHg4HqpX3QohRAdXXw8bNkBenm3s0Ufh\n2WdB9d6xoooiUrNSqWmoAcDD1YMFwxcQGRipNrAQQimHnsh9++23/Pa3v2X+/PmEhoba/d6ECW2z\nWaQ8kRNCtCV1dbBuHVy8aBsbOxaeekp9EVdYXsiarDXUmmoB8HLzIn5EPL39e6sNLIRwmNIncseP\nH2fnzp0cPHjwe2vkLl++3OSgQgjRmdTUwJo1cO+PyyefhNhY9UXcxdsXWXtqLfXmegB83H1IGJFA\nT7+eagMLIXTh0IqMd955hx07dlBSUsLly5ftPoRoJOf06Uvyra/m5ru6GlJS7Iu4KVNg4kT1Rdz5\n0vOsObXGWsT5eviycOTCdlPEyT2uL8m3vpSftXqvLl268OSTTzoloJ5k16oQojVVVWlF3PXrtrGf\n/Qwee0x97LMlZ9mUvQmTxQSAv6c/STFJBPsEqw8uhHCYLrtWV61axbFjx1i6dOn31si5qN5m1Uyy\nRk4I0ZoqKrQirqREuzYYYNo0GDVKfezTN07zac6nmC1mAAK9AkmKSSLIO0h9cCFEszS3bnGokHtY\nsWYwGDCZTE0Oqgcp5IQQraWsDFavhtvaoQkYDBAXByNGqI+dcS2DbWe3YUH7+RfsHUxiTCIBXgHq\ngwshmq25dYtDj9Py8/Mf+HHhwoUmBxQdl6yv0JfkW1+O5ru0FFautBVxLi7w0kv6FHHfXf2Ov539\nm7WI6+bTjYUjF7bbIk7ucX1JvvWl6xq5qKgopwQTQoiO7OZN7UlcpXZoAq6u2rmpA3U4NOHolaPs\nyttlve7h24OEEQl08eiiPrgQotU4fNZqeyOvVoUQeiouhtRUbZcqaIfez50L/fqpj32w4CB7L+61\nXvfy60X8iHi83dvPkYpCdHZK+8gJIYR4uCtXIC1N6xcH4OEBCxZApOJDEywWC8ZLRvYX7LeORQRE\nsGD4AjzdPNUGF0K0CW1zy6mTJCcnyzt/HUmu9SX51tfD8l1QoO1ObSzivLwgMVGfIm5P/h67Iq5P\nYB/iR8R3mCJO7nF9Sb711Zhvo9FIcnJys79Ph34i15LECCHEj8nP147dqtf67eLjoxVxPXqojWux\nWPgi7wuOFR2zjg3oOoDZQ2fj7uquNrgQwqka+90uX768WV/v0Bq5/Px83nnnHTIyMqhsXMWL9j63\nsLCwWYFVkzVyQgiVzp2DjRuhoUG79vXVirju3dXGNVvM7Di3gxPFJ6xj0SHRvDjkRdxcOvS/zYXo\n0JSukZs/fz79+/fnj3/84/fOWhVCiM7mzBnYvBnMWr9dAgK0Ii5Y8aEJZouZrTlbOXXjlHVsWPdh\nxA2Ow9XFVW1wIUSb5NATOX9/f27fvo2ra/v5QSFP5PRnNBrlODQdSb711ZjvrCzYuhUaf7wEBUFS\nEgQGqo1vMpvYkrOFMzfPWMdG9hjJjEEzcDF0zOXOco/rS/Ktr/vzrbQh8IQJEzh58mSTv7kQQnQk\nx4/bF3EhIbBokfoirsHcwIbsDXZF3KNhj/L8oOc7bBEnhHCMQ0/kXn/9dTZs2MDMmTPtzlo1GAy8\n//77SifYXPJETgjhDLm5BXz11QXOn3fh3Dkzffv2IyQkktBQSEjQ1sapVGeqY/3p9eTfzreOPd77\ncab2m4rBYFAbXAihG6Vr5Kqqqpg2bRr19fVcuXIF0HZNyQ8RIURHlptbwKpVeVy/Ppn8v9dRGRl7\nmToVkpIi8fFRG7+2oZa1p9ZSUF5gHZsQOYGJURPl568QAnCwkFu1apXiaaiRnJxs3dYr1JP1FfqS\nfKu3Z88FioomU1gIZWVGAgNj6dp1Mv7+6fj4qG0UV9NQQ1pWGlcqrljHJvWZxITICUrjtiVyj+tL\n8q2vxnwbjcYW9fB7aCF36dIl6xmr+fn5D/s0+vbt2+zgqkkfOSFEc5nNkJnpwr0dlgIDYfhwUN1L\nvbq+mtTMVIori61jT/d7msfDH1caVwihP2V95Pz8/Lhz5w4ALi4P/qFlMBgwmUzNCqyarJETQjSX\nyaRtalixIp3q6kmA1lpkyBBwdYXu3dN57bVJSmJX1lWSkpnCjaob1rHnBjzHo70eVRJPCNE2OH3X\namMRB2A2mx/40VaLOCGEaK76eli/Hk6fhr59+9HQsJfu3WHoUK2Iq63dy+TJ/ZTErqitYOXJldYi\nzoCB5wc9L0WcEOKhZN+6cBo5p09fkm/nq6mBtDQ4f167DgmJJD6+P08+mc6tW3+me/d0Fi7sz6BB\nzl8fV1ZTxsqTKym9WwqAi8GFmdEzeaTnI06P1V7IPa4vybe+nJVvOc9FCCGAqiqtiCu2LUtjwgSY\nODESgyESo9FF2ULw0upSVmeupqK2AgBXgyuzhswiulu0knhCiI7DoT5y7ZGskRNCOKq8HFJToaTE\nNjZ1Kowdqz72zaqbrM5cTWWddo61m4sbs4fOZmDwQPXBhRBthtI+ckII0VGVlkJKilbMARgMMH06\n/OQn6mNfq7xGSmYK1fXVALi7uDNv+Dz6BrXdbgBCiLalyWvk7t/wIEQjWV+hL8l3y127BitW2Io4\nV1d46aUHF3HOzndRRRGrMlZZizgPVw/iR8RLEXcPucf1JfnWl7Py7VAhd/z4cR5//HF8fHxwc3Oz\nfri7uztlEkIIobfCQli1SlsbB+DuDvPmaS1GlMcuLyQlM4WahhoAvNy8SIxJJDJQbZNhIUTH49Aa\nuWHDhjFjxgzi4+Pxue9MmsamwW2NrJETQjxMXh5s2KC1GgHw8oIFCyA8XH3si7cvsvbUWurNWnAf\ndx8SRiTQ06+n+uBCiDaruXWLQ4Wcv78/5eXl7epsPynkhBAPkp0Nn36qNf0F6NIFEhKgRw/1sc+X\nnmdD9gYazA0A+Hr4khiTSPcu3dUHF0K0aU5vCHyvuLg4du/e3eRv3tqSk5Plnb+OJNf6knw33fHj\nsHmzrYgLCIDFix0r4lqa77MlZ1l/er21iPP39GfRyEVSxP0Aucf1JfnWV2O+jUZji44UdWjX6t27\nd4mLi2P8+PGEhoZaxw0GAykpKc0OrpqctSqEaPT117Bnj+06JAQSE8HfX33s0zdO82nOp5gt2gax\nQK9AkmKSCPIOUh9cCNGmKTtr9V4PK4gMBgPLli1rVmDV5NWqEALAYoH0dDh40DYWFqatievSRX38\njGsZbDu7DQvaz6Ng72ASYxIJ8ApQH1wI0W4oXSPXHkkhJ4SwWGDnTvj2W9tYZCTMnw+enurjf3f1\nO3ac22G97ubTjcSYRPw8/dQHF0K0K0rXyAHs27ePRYsWMXXqVBYvXkx6enqTg4mOTdZX6Evy/cNM\nJm1Tw71F3MCBEB/fvCKuqfk+euWoXRHXw7cHC0culCKuCeQe15fkW1+69pH75JNPmDNnDj179mTm\nzJn06NGD+fPn89e//tUpkxBCCGeqr9fai5w6ZRsbPhzmzNH6xal2sOAgu/J2Wa97+fUiKSaJLh46\nvMsVQnQqDr1aHTBgAJs3byYmJsY6lpWVxcyZM8nLy1M6weaSV6tCdE61tbB2LRQU2MYefRSefVY7\nfksli8WC8ZKR/QX7rWMRAREsGL4ATzcd3uUKIdotpWvkgoODKS4uxsPDwzpWW1tLWFgYpaWlTQ6q\nBynkhOh8qqogLQ2Ki21j48fDpEn6FHF78vdw+PJh61jfoL7MHTYXD1ePH/hKIYRQvEbuiSee4M03\n36Tq72fZVFZW8utf/5qxY8c2OaDouGR9hb4k3/bKy2HlSvsibupUmDzZOUXcD+XbYrHwRd4XdkXc\ngK4DmDdsnhRxLSD3uL4k3/rSdY3cRx99RFZWFgEBAXTv3p3AwEAyMzP56KOPnDIJIYRoidJSWLEC\nSkq0a4MBZswAPf6tabaY2X5uO8eKjlnHokOimTtsLu6uch61EEKtJrUfuXz5MlevXiUsLIxwPQ4l\nbAF5tSpE53DtGqSmaq9VAVxdYeZMGDpUfWyzxczWnK2cumHbVTGs+zDiBsfh6uKqfgJCiA7D6Wvk\nLBaL9WxVs9n80G/g4uJwBxNdSSEnRMdXWKhtbKip0a7d3bWdqf37q49tMpvYkrOFMzfPWMdG9hjJ\njEEzcDG0zZ+LQoi2y+lr5PzvObfGzc3tgR/ueuzjF+2GrK/QV2fPd16e9iSusYjz8oKEBHVF3L35\nbjA3sCF7g10R92jYozw/6Hkp4pyos9/jepN868tZ+X7oWavZ2dnWX+fn5zslmBBCOEN2ttbs12TS\nrrt00Yq4Hj3Ux64z1bH+9Hryb9t+Lj7e+3Gm9ptqfYshhBB6cWiN3O9//3t+/etff2/8j3/8I2++\n+aaSibWUvFoVomM6cQK2b9eO3wIICIDERAgOVh+7tqGWtafWUlBua1I3IXICE6MmShEnhGgRpX3k\n/Pz8uHPnzvfGg4KCuH37dpOD6sFgMLBs2TJiY2OJjY1t7ekIIZzg8GH48kvbdUiI9iQuQOH587l5\nuXx1/CuqG6o5WXySrmFdCQkLAWBSn0lMiJygLrgQosMzGo0YjUaWL1/u/EIuPT0di8XC9OnT2bFj\nh93vXbhwgX//93+n4N726W2IPJHTn9FolKJZR50p3xYLpKfDwYO2sZ49tXNTuyg89So3L5dV+1bh\n0teF9H3pePb3pCGvgZFDRrJg/AIeD39cXXDRqe7xtkDyra/7893cuuWha+QAFi9ejMFgoLa2lpdf\nftkuWGhoKH/5y1+aHFAIIZrCYoGdO+Hbb21jkZEwb562wUGlr45/BX0g41oGdxvu4oknbv3d8Lnj\nI0WcEKJNcOjVakJCAqmpqXrMx2nkiZwQ7Z/JBH/7G5yytWlj4EB46SWt1Yhqv035LQdcDlDTUGMd\nGxQ8iOiqaN6Y+4b6CQghOg0lT+QatbciTgjR/tXXw6ZNcO6cbWz4cHjhBa3pr2o3qm7wbdG31PTU\nijgDBgaHDCbUNxSPu3LslhCibXCo4VF5eTm//OUv+clPfkJkZCTh4eGEh4cTERGhen6iHZEeRPrq\nyPmurYU1a+yLuNGjIS5OnyLuSsUVVp5cSa/IXjTkNeBicCHwWiChvqHUnq9l8k8mq5+E6ND3eFsk\n+daXrmetvv7665w4cYL33nuPW7du8Ze//IWIiAjeeENeLQghnKuqClavhkuXbGPjx8Nzz4EeB8nk\n384nJTOFuw13CQkLYcywMcRaYulV3YvuN7qzcOJCBvUfpH4iQgjhAIfWyHXr1o2cnBxCQkIICAig\nvLycoqIipk+fzokTJ/SYZ5PJGjkh2p+KCkhJgZIS29hTT8ETT+gTP+dmDpvPbMZk0ToNd3HvQvyI\neHr69dRnAkKITkvpGjmLxULA3xs1+fn5UVZWRs+ePTl//nyTAwohxIOUlmpHbpWVadcGA0ybBqNG\n6RP/ZPFJPsv9DAvaD9IAzwASYhII8QnRZwJCCNEMDr2oGDFiBAcOHABg3LhxvP766/zTP/0TgwbJ\n6wVhI+sr9NWR8n3tGqxcaSviXF1h1iz9irgjl4+wLXebtYgL9g5m8SOL7Yq4jpTv9kJyri/Jt750\nXSP38ccfExUVBcB//dd/4eXlRXl5OSkpKU6ZhBCi87p8GVatgspK7drdXesRN3So+tgWi4X0i+ns\nvrDbOtbTtyeLH1lMgJfC4yKEEMJJHFoj98033/DYY499b/zYsWOMGTNGycRaStbICdH25eXBhg1a\nqxHQGvzOnw96bIi3WCzsPL+Tb6/aOg1HBkQyb/g8vNwUdxoWQoj7tMpZq127duXWrVtNDqoHKeSE\naNvOnIEtW7Smv6AdtZWQAD16qI9tMpv429m/ceqGrdPwwOCBvDTkJdxddeg0LIQQ92lu3fKDr1bN\nZjOmv/+UNZvNdh/nz5/Hzc2hvRKik5D1Ffpqz/k+cUJr9ttYxAUEwOLF+hRx9aZ6NmRvsCvihncf\nzpyhc36wiGvP+W6vJOf6knzry1n5/sFK7N5C7f6izcXFhXfeeccpkxBCdB6HD8OXX9quQ0K0J3EB\nOixJq2moYd2pdRSUF1jHHg17lGcHPIvBYFA/ASGEcLIffLV66e8dOSdMmMDBgwetj/wMBgPdunXD\nx8dHl0k2h7xaFaJtsVhg3z74+wZ4AHr2hPh47bWqalV1VaRmpXKt8pp1bELkBCZGTZQiTgjR6pSu\nkWuPpJATou2wWOCLL+DYMdtYZKS2O9VLh30F5TXlpGSmUHq31Dr2dL+neTz8cfXBhRDCAUobAick\nJDwwICAtSISV0WgkNja2tafRabSXfJtMsG0bZGXZxgYMgNmztVYjqpVUl5CSmUJFbQUABgzMGDSD\nR3o+0qTv017y3ZFIzvUl+daXs/LtUCHXr18/u0rx2rVrbNmyhQULFrR4AkKIjqu+HjZvhtxc29iw\nYRAXpzX9Ve3qnaukZaVRXV8NgKvBlVlDZhHdLVp9cCGE0EGzX61+9913JCcns2PHDmfPySnk1aoQ\nrau2Ftatg78vtQVg9Gh49llwcagVectcKrvEulPrqDXVAuDh6sHcYXPpG9RXfXAhhGgi3dfINTQ0\nEBQU9MD+cm2BFHJCtJ7qakhLg6tXbWPjxsHkydoZqqqdKz3HxuyNNJgbAPB282bBiAX09u+tPrgQ\nQjSDkj5yjfbu3Ut6err1Y/v27SQlJTFUjzN07lNRUcGYMWPw8/PjzJkzuscXDyc9iPTVVvNdUaGd\nm3pvEffUUzBlij5FXNb1LNafXm8t4vw8/Fj0yKIWF3FtNd8dmeRcX5JvfenSR67Ryy+/bLc9v0uX\nLowcOZJ169Y5ZRJN4ePjw86dO/mXf/kXeeImRBtz6xakpEBZmXZtMMC0aTBqlD7xjxUdY+f5ndbr\nIK8gEmMSCfIO0mcCQgihs3bbfmTRokX8+te/fuhTQXm1KoS+rl+H1FSorNSuXVxg5kxtc4NqFouF\ng4UHSb+Ybh0L7RJK/Ih4/Dz91E9ACCFaSGn7EYCysjI+//xzrl69SlhYGM8++yxBQfKvXCEEXL4M\na9ZATY127e6utRcZMEB9bIvFwpcXvuTIlSPWsd7+vVkwfAHe7t7qJyCEEK3IoTVy6enpREVF8eGH\nH/Ltt9/y4YcfEhUVxVdffdWkYP/93//N6NGj8fLyYtGiRXa/d+vWLeLi4vD19SUqKsrute2f/vQn\nJk6cyB/+8Ae7r5Fu7G2LrK/QV1vJ94UL2uvUxiLO01M7rUGPIs5sMbMtd5tdEdcvqB+JMYlOL+La\nSr47E8m5viTf+tJ1jdzrr7/OX//6V2bPnm0d27RpEz//+c85e/asw8F69erF0qVL2b17N3fv3v1e\nDC8vL27cuMHJkyd57rnniImJYciQIfzyl7/kl7/85fe+n7w6FaJ1nTkDW7ZoTX9BO2orPl47eku1\nBnMDm89s5myJ7WfQkG5DmBk9EzcXh182CCFEu+bQGrnAwEBKS0txvaeDZ319Pd26daOscVVzEyxd\nupQrV66wcuVKAKqqqujatSvZ2dn0798fgKSkJMLCwvjd7373va9/9tlnyczMJDIykldffZWkpKTv\n/8EMBpKSkoiKirL+GUaOHGntotxYCcu1XMt1867Pn4erV2OxWODSJSNdusDy5bGEhKiP/+XeL0m/\nmI5Xf+18r0sZlxjQdQD/lvBvuBhc2kR+5Fqu5Vquf+i68deN59qvXr1aXR+5JUuW0L9/f37xi19Y\nxz788EPOnz/PX/7ylyYHfffddykqKrIWcidPnmTcuHFUVVVZP+ePf/wjRqORzz77rMnfH2SzgxAq\nHTkCu3fbroODITERAgLUx66ur2ZN1hqK7hRZx8aGj+Wpvk/JcgshRLultI/ciRMn+PWvf02vXr0Y\nM2YMvXr14le/+hUnT55k/PjxjB8/ngkTJjRpsveqrKzE39/fbszPz6/NNhsWD3bvvzKEeq2Rb4sF\n0tPti7iePWHxYn2KuIraClaeXGlXxE3uM1mXIk7ub/1JzvUl+daXs/Lt0EKSV155hVdeeeUHP6cp\nP0Tvrzh9fX2pqKiwGysvL8fPT9oGCNFWWCzwxRdw7JhtLCIC5s8HLy/18W/dvUVKZgplNdpyDgMG\nnhv4HKPDRqsPLoQQbZRDhdzChQudGvT+om/gwIE0NDSQl5dnXSOXmZnJsBY2oEpOTiY2Ntb6Xlqo\nJXnWl575Nplg2zbIyrKNDRigtRhxd1cf/3rldVKzUqms05rUuRhcmBk9k2HddWhS93dyf+tPcq4v\nybe+7l0z15Kncw43BD5w4AAnT560rmOzWCwYDAbefvtth4OZTCbq6+tZvnw5RUVFfPzxx7i5ueHq\n6sq8efMwGAx88sknnDhxgmnTpnHkyBGio6Ob9weTNXJCOEVDA2zaBLm5trFhwyAuDu7Z/6TM5fLL\nrDm1hpoGrb+Jm4sbc4bOYUCwDv1NhBBCJ0rXyC1ZsoSXXnqJgwcPkpOTQ05ODmfPniUnJ6dJwT74\n4AN8fHz4j//4D9LS0vD29uY3v/kNAP/v//0/7t69S/fu3YmPj+ejjz5qdhEnWoesr9CXHvmurYW0\nNPsibtQo7cQGPYq4vFt5pGSmWIs4LzcvEmMSW6WIk/tbf5JzfUm+9aXrGrm0tDSys7MJCwtrUbDk\n5GSSk5Mf+HtBQUFs3bq1Rd9fCOE81dVaEXf1qm1s3DiYPFk7Q1W17BvZfJrzKSaL1qSui3sXEmIS\n6OHbQ31wIYRoJxx6tTpixAjS09MJCQnRY05OIa9WhWi+igrt3NSbN21jU6ZohZweThSfYHvudixo\n/w0HeAaQGJNIsE+wPhMQQgidKT1r9X//93955ZVXmD9/PqGhoXa/15S2I3qTzQ5CNN2tW9qRW429\nvg0GeO45GK3T5tCvC79mT/4e63WITwiJMYn4e/r/wFcJIUT7pMtmh48++ohf/OIX+Pn54e1tf37h\n5cuXmx1cJXkipz+j0ShFs45U5Pv6de1JXKW2ORQXF209XAs3kDvEYrGw9+JeDhUeso6F+YURPyIe\nH3cf9RP4EXJ/609yri/Jt77uz7fSJ3LvvPMOO3bs4KmnnmpyACFE+3D5MqxZAzXavgLc3GDOHK3N\niGpmi5md53fy3dXvrGNRgVHMGzYPTzdP9RMQQoh2yqEnchEREeTl5eHh4aHHnJxCnsgJ4bgLF2D9\neqiv1649PbVGv5GR6mObzCa2nt3K6RunrWODggcxa8gs3F11aFInhBBtgNL2I++//z5vvPEGxcXF\nmM1muw8hRPt25gysXWsr4rp0gYUL9Sni6k31rDu9zq6IGxE6gtlDZ0sRJ4QQDnCokFvPguzCAAAg\nAElEQVS8eDEfffQRvXr1ws3NzfrhrkdL9xZITk6Wvjg6klzryxn5PnlSa/Zr0jp8EBAAixZp56eq\nVtNQQ2pWKnm38qxjj/V6jLjBcbi66NCkronk/taf5Fxfkm99NebbaDQ+tDWbIxxaI5efn9/sAK2p\nJYkRoqM7cgR277ZdBwdDYqJWzKlWWVdJWlYa1yqvWcdio2J5MvLJJp3bLIQQ7V1jd43ly5c36+sd\nPqILwGw2c/36dUJDQ3FxcehhXquRNXJCPJjFAkYj7N9vG+vRAxIStNeqqpXVlJGSmcKtu7esY8/0\nf4af9v6p+uBCCNFGKV0jV1FRQWJiIl5eXvTq1QsvLy8SExMpLy9vckAhROuxWGDXLvsiLiJCWxOn\nRxF3s+omK06usBZxLgYXXhj8ghRxQgjRTA6ftVpVVcXp06eprq62/u+SJUtUz0+0I7K+Ql9NzbfJ\nBFu3wjff2Mb699eexHl5OXduD1JUUcTKjJVU1FYA4GpwZfbQ2YzsMVJ9cCeQ+1t/knN9Sb71petZ\nq7t27SI/P58uf/8n+8CBA1m1ahV9+/Z1yiSEEGo1NGibGnJzbWNDh2rNfl112Fdw8fZF1p1eR52p\nDgAPVw/mDZtHn6A+6oMLIUQH5tAauaioKIxGI1FRUdaxS5cuMWHCBAoLC1XOr9kMBgPLli2TI7pE\np1dbq/WIu3jRNjZqlHbslh5LXc+WnGXzmc00mBsA8HH3YcHwBfTy76U+uBBCtHGNR3QtX768WWvk\nHCrk/v3f/53Vq1fzq1/9isjISC5dusSf/vQnEhISWLp0abMmrppsdhACqqu10xqKimxjTzwBU6Zo\nZ6iqlnktk2252zBbtJ6T/p7+JIxIoFuXbuqDCyFEO6J0s8Pbb7/Nv/3bv7Fp0yZ+9atfsWXLFt56\n6y3efffdJgcUHZesr9DXj+W7ogJWrrQv4qZMgaee0qeI++bKN2w9u9VaxHX17sriRxa32yJO7m/9\nSc71JfnWl65r5FxcXFi8eDGLFy92SlAhhFq3bkFKCpSVadcGg/YqdfRo9bEtFgv7C/ZjvGS0joV2\nCSUhJgFfD1/1ExBCiE7EoVerS5YsYd68eYwdO9Y6dvjwYTZu3Mif//xnpRNsLnm1Kjqr69chNRUq\nK7VrFxeIi4Phw9XHtlgs7MrbxTdFtq2x4f7hLBixAC83HbbGCiFEO9XcusWhQi4kJISioiI8PT2t\nYzU1NYSHh3Pz5s0mB9WDFHKiM7pyRVsTd/eudu3mBrNnw8CB6mObzCY+y/2MzOuZ1rH+Xfsze+hs\nPFw91E9ACCHaMaVr5FxcXDCbzXZjZrNZCiVhR9ZX6Ov+fOfna69TG4s4T0+tR5weRVyDuYGN2Rvt\nirih3YYyb9i8DlPEyf2tP8m5viTf+nJWvh0q5MaNG8e7775rLeZMJhPLli1j/PjxTpmEKsnJyXJj\nik4hJ0d7ElentWnDx0c7rSEyUn3s2oZa0rLSyC21Nakb1XMULw55EVcXHZrUCSFEO2Y0Glt0NrxD\nr1YvX77MtGnTKC4uJjIyksLCQnr27Mn27dsJDw9vdnCV5NWq6CwyMmDbNu34LQB/f0hMhJAQ9bGr\n66tJy0rj6p2r1rFxEeOY3GcyBj22xgohRAehdI0caE/hjh07xuXLlwkPD+exxx7DRY9uos0khZzo\nDI4e1c5ObRQcrL1ODQxUH7uitoKUzBRKqkusY1P6TmFcxDj1wYUQooNRXsi1N1LI6c9oNMopGjrI\nzS1gz54LfPVVFnV1I+jbtx8hIZH06AHx8eCrQ4eP0upSUjJTKK8tB8CAgWkDpzEqbJT64K1E7m/9\nSc71JfnW1/35VrrZQQjRNuTmFrBqVR5Hjkzi0qWRVFdPIiMjD3f3AhYu1KeIu1Z5jRUnV1iLOFeD\nK7OGzOrQRZwQQrRV8kROiHbkv/4rnYMHJ1Fie5tJ164wcWI6S5ZMUh6/sLyQtafWUtNQA4C7iztz\nhs2hf9f+ymMLIURHpuyJnMViIT8/n4aGhmZNTAjhHOXlcOiQi10R160bDBsGJpP6h+vnS8+Tmplq\nLeK83LxIjEmUIk4IIVqRQz/9hw0b1qY3Noi2QVq9qHP5Mvz1r1BVZevn6OZmZMgQ7eQGDw/zD3x1\ny52+cZp1p9dRb64HwNfDl0UjFxEe0DZ3rasg97f+JOf6knzrS7c+cgaDgUceeYTc3Nwf+1QhhAIZ\nGbBqFVRVQd++/TCb9zJwIPTurZ2hWlu7l8mT+ymL/93V79hyZgtmi1YsBnoFsviRxYT6hiqLKYQQ\nwjEOrZF79913SUtLY+HChYSHh1vf4xoMBhYvXqzHPJvMYDCwbNkyYmNjZReOaJfMZvjqKzh82Dbm\n4wOjRxdw9uwF6upc8PAwM3lyPwYNUtP591DhIb7K/8p63c2nGwkxCfh7+iuJJ4QQnY3RaMRoNLJ8\n+XJ17UcaC6EHNfjct29fk4PqQTY7iPasthY2b4bz521j3bvDvHkQFKQ+vsVi4av8r/j68tfWsV5+\nvVgwYgE+7j7qJyCEEJ2M9JG7jxRy+pMeRM5x6xasWwc3b9rGBg2CmTO181Mbqcq32WJmx7kdnCg+\nYR3rE9iHucPm4unm+QNf2bHJ/a0/ybm+JN/6clYfOTdHP7G0tJTPP/+ca9eu8a//+q8UFRVhsVjo\n3bt3k4MKIR7s4kXYuNF28D3AuHEwebK2Hk61BnMDn+Z8ypmbZ6xjg0MGM2vILNxcHP5xIYQQQicO\nPZHbv38/L774IqNHj+brr7/mzp07GI1G/vCHP7B9+3Y95tlk8kROtDfffQc7d2pr4wDc3GDGDBgx\nQp/4daY6NpzewIXbF6xjI3uMZMagGbgYZNe6EEKopPTV6siRI/n973/PlClTCAoK4vbt29TU1BAR\nEcGNGzeaNWHVpJAT7YXJBLt3w7FjtjFfX5g7V9uZqoe79XdZe2otlysuW8d+2vunPN3v6QeujRVC\nCOFcSo/oKigoYMqUKXZj7u7umEymJgcUHZf0IGq6u3dhzRr7Iq5nT/jHf/zxIs5Z+b5Te4dVGavs\niriJUROliLuP3N/6k5zrS/KtL936yAFER0eza9cuu7G9e/cyfPhwp0xCiM6opAQ+/hjy821jQ4fC\n4sXgr1N3j9t3b7MyYyXXq65bx37W/2c8GfWkFHFCCNEOOPRq9ejRo0ybNo1nn32WTZs2kZCQwPbt\n29m2bRtjxozRY55NJq9WRVuWlwebNmltRhpNnAgTJuizqQHgRtUNUjNTuVN3BwAXgwsvDH6BEaE6\nLcoTQghhpbz9SFFREWlpaRQUFBAREUF8fHyb3rEqhZxoiywWOHoUvvxS+zWAuzvExcGQIfrN40rF\nFdZkreFug7Y91s3FjZeGvMSgkEH6TUIIIYSVLn3kzGYzJSUldOvWrc2/dpFCTn/Sg+iHNTTA55/D\nyZO2sYAAbVNDz55N/37NzXf+7XzWn15PnakOAE9XT+YNn0dUYFTTJ9GJyP2tP8m5viTf+nJWHzmH\n1sjdvn2bhIQEvL296dGjB15eXsTHx3Pr1q0mB9RTcnKyLN4UbUJVFaSk2BdxvXvDK680r4hrrpyb\nOazJWmMt4nzcfUgamSRFnBBCtBKj0UhycnKzv96hJ3IvvPACbm5ufPDBB0RERFBYWMh7771HXV0d\n27Zta3ZwleSJnGgrrl/XTmooK7ONxcTA9Olarzi9ZFzLYNvZbVjQ/rvw9/QnMSaREJ8Q/SYhhBDi\ngZS+Wg0ICKC4uBgfH9sZi9XV1fTs2ZPy8vImB9WDFHKiLTh7Fj79FOq0B2AYDDBlCowdq9+mBoCj\nV46yK8+28zzYO5iEmAQCvQL1m4QQQoiHUvpqdfDgwVy6dMlurKCggMGDBzc5oOi45DW2jcUCBw/C\n+vW2Is7TUzv0/oknnFPEOZJvi8VC+sV0uyKuh28PFj+yWIq4JpL7W3+Sc31JvvXlrHw79GJn0qRJ\nTJ06lcTERMLDwyksLCQtLY2EhARWrFiBxWLBYDCwePFip0xKiPasvh4++wxOnbKNBQVpRVz37vrN\nw2Kx8EXeFxwrsnUbjgiIYP7w+Xi5eek3ESGEEMo49Gq1cVfFvTtVG4u3e+3bt8+5s2sBebUqWsOd\nO9pTuKIi21hUFMyeDfesTFDOZDbxt7N/49QNWzU5oOsAZg+djburu34TEUII4RBd2o+0J1LICb1d\nvaptarhzxzY2ahQ8+yy4uuo3j3pTPZvObOJc6Tnr2LDuw4gbHIeri44TEUII4TCla+SEcERnXl9x\n+jSsWGEr4lxctAJu2jR1RdyD8l3TUENaVppdETc6bDQzo2dKEddCnfn+bi2Sc31JvvWl6xo5IcSD\nWSywbx8cOGAb8/LSXqX27avvXKrqqkjLSqO4stg6Nj5iPJP6TGrzDbyFEEI0j7xaFaKZ6upg61bI\nybGNhYRomxqCg/WdS3lNOalZqZRUl1jHpvabytjwsfpORAghRLM0t26RJ3JCNENZmbap4do121j/\n/jBrlvZETk8l1SWkZqZSXqv1dDRgYPqg6fyk50/0nYgQQgjdObxGLicnh/fff5/XX38dgLNnz5KV\nlaVsYqL96SzrKwoL4eOP7Yu4n/4U5s/Xt4gzGo0U3ylm5cmV1iLO1eDKS0NfkiJOgc5yf7clknN9\nSb715ax8O1TIbdq0iQkTJlBUVERKSgoAd+7c4c0333TKJIRoLzIyYPVq7exU0DYyzJgBzzyjbXDQ\nQ25eLv93w/9lxRcrePXDVykoKADAw9WD+cPnM6TbEH0mIoQQotU5tEbu/7d351FNX3n/wN9hCXuQ\nRUGwiIqgPu5bH+pSwLowKlbGWu2pVu1Uj9rO2JmuY1Uc7dPTmdZ2TtU69bF1q6jtY4/iigpRXNFf\ngVpFNhV3UFFWCUn4/v5ICUawQkhuSPJ+ncM55ib53sv7xPjx+733frt164atW7eib9++8PHxwf37\n96FWq9G+fXvcvXv3aW+3CM6RI1OqrQUOHQJOnKhvc3cHXn4Z6NhR3Dhy8nOwPnU9KoIrcP7OedRK\ntdDka/Dfvf4bC2IXoIOig7jBEBGRyZh1jtydO3fQu3fvBu0Ook5BEFlQdTXwf/8H5OXVtwUE6BY1\ntBF8l6vks8m43fY2rhRfgQTdX3j3CHf4qfxYxBER2aEmVWL9+/fHpk2bDNq2bduGwYMHm2VQZJ1s\ncX5FSQmwbp1hERcRAcyaJb6Iu1F2A0euHsHlB5chQcKDiw/g6uSKfoH94CZ3EzsYO2SLn+/WjpmL\nxbzFErqP3FdffYWRI0di3bp1qKqqwqhRo5Cbm4vk5GSTDMJcEhISEBUVpb/FGFFzXL4MbN8OPHxY\n3zZsGBATY5qb3jdVjbYGqZdTcer6KVTVVOnb3Zzc0C+wH1ycXCB3kIsbEBERmYxSqWxRUdfkfeQq\nKyuxe/duFBYWIiQkBGPHjoWXl5fRHZsb58hRS5w5A+zbp5sbBwBOTrpFDY3MMDCrgpICJOUm4UH1\nAwDA3Zt38Uv2L+g6sCs6KDpAJpNBlafCjOgZiAiLEDs4IiIyGd5r9TEs5MgYWi2wf7+ukKvj6amb\nDxccLG4cVeoqHMg/gKyiLIP2Lj5dEOEYgbMXzqKmtgZyBzlG9B/BIo6IyMqZtZArLCzE0qVLkZGR\ngYqKCoNOc3Nzf+edlsNCTjylUmnVl7EfPtRdSr18ub4tKAiYMgVQKMSMQZIk/Fr8K/bn70elulLf\n7ubkhjFhY9A7oLf+dlvWnre1Yd7iMXOxmLdYj+dt1lWrL730Erp3745ly5bBVfS29UQC3LkDJCbq\nFjfU+a//Al58EXB2FjOG0upS7MnbY3DDewDo1a4XxoSNgYfcQ8xAiIjIajTpjJy3tzdKSkrg6Ogo\nYkwmwTNy1FR5ecCPPwIqVX1bdDQwfLiYRQ2SJOHMzTM4dOkQarQ1+nZvF2+MDR+LcL9w8w+CiIgs\nyqxn5MaNG4cjR44gJiam2R0QtVaSBJw6BSQn6/4M6M6+TZwI9BB0c4Q7lXewK2cXrpVd07fJIMOg\n4EEY0WkEXJxcxAyEiIisUpPOyN29exeRkZEIDw9Hu3bt6t8sk+Hbb7816wCNxTNy4lnT/AqNBtiz\nB8jIqG/z9tYtaggMFNB/rQbHrh5DWmEatJJW397WvS3iIuLwjPczTz2GNeVtC5i3eMxcLOYtltA5\ncrNmzYJcLkf37t3h6uqq70wmcjMtIhOprAS2bQOuXq1ve+YZ3e22PD3N3/+10mvYlbMLd6ru6Nsc\nZY4Y1nEYhoYMhZNDk/5aEhERNe2MnJeXF27cuAGFqKV7JsAzctSY27d1ixpKS+vb+vYFxo3T7RVn\nTiqNCimXU5B+I11/ey0A6KDogLiIOLTzaPc77yYiIltm1jNyvXv3xr1796yqkCN6XHY28NNPQM1v\n6wlkMmDkSCAy0vyLGvLu5WF37m6UquorSLmjHC90fgEDgwbCQcb7FhMRUfM1qZCLiYnB6NGjMXPm\nTAQEBACA/tLqrFmzzDpAsh6tdX6FJAFpaUBKSn2biwvwxz8C4WZeEFpZU4n9+ftxrvicQXtX364Y\nFz4O3q7eRh+7teZtq5i3eMxcLOYtlqnyblIhl5aWhqCgoEbvrcpCjloztRrYuRP49df6Nh8f4JVX\ngLZtzdevJEn4pegXHCg4gCp1/f1R3Z3dERsWi57tenKOKRERtRhv0UU2q6wM2LoVuHmzvi00FJg8\nGXB3N1+/D6ofICknCQX3Cwza+wT0weiw0XB3NmPnRERklUw+R+7RVam1dXcOb4SDA+f2UOtz44au\niCsvr28bOBCIjQXMta91rVSL9BvpOHzpMNS1an17G9c2GBc+DmG+YebpmIiI7NYTq7BHFzY4OTk1\n+uMs6t5FZBWUSqWlhwAAOHcO+O67+iLOwQEYO1a3MtVcRVxRRRHW/bwO+/P364s4GWSI7BCJeYPm\nmaWIay152wvmLR4zF4t5i2WqvJ94Ru78+fP6P1+6dMkknRGZkyQBqanA0aP1bW5uwEsvAZ07m6dP\nTa0GRwuP4tjVY6iV6s9cB3gEIC4iDsGKYPN0TEREhCbOkfvss8/wzjvvNGhfsWIF/vrXv5plYC3F\nOXL2paYG2LEDuHixvs3fX3enBj8/8/RZ+KAQSblJuFt1V9/mKHPE86HPY8gzQ+DoYD33JiYiIssy\ntm5p8obA5Y9ONvqNj48P7t+/3+xORWAhZz8ePNBt8ltUVN8WFgZMmgS4upq+v2pNNQ5dOoSzN88a\ntId4hyAuIg7+7v6m75SIiGyaWTYETklJgSRJ0Gq1SHl0Ey4ABQUF3CCYDFhiD6KrV3W326qsrG+L\njNRt9GuOdTg5d3OwJ28PylRl+jYXRxeM7DISA9oPELqlCPd8Eot5i8fMxWLeYgnZR27WrFmQyWRQ\nqVR4/fXX9e0ymQwBAQH46quvWjyA5kpPT8eCBQvg7OyM4OBgbNy4EU7mvrcStUoZGcDu3YD2t3vO\nOzrqFjT062f6vipqKrAvbx/O3zlv0B7hF4Gx4WOhcOF/aoiISLwmXVqdNm0aNm3aJGI8T3X79m34\n+PjAxcUFf//73zFgwAD88Y9/bPA6Xlq1XbW1wMGDwMmT9W0eHrqb3oeEmLYvSZKQeTsTBwoOoFpT\nXd+fswf+0PUP6NG2Bzf2JSKiFjPrvVZbSxEHAIGBgfo/Ozs7w9Fc+0lQq1RdDfz4I5CfX98WEKBb\n1NCmjWn7KnlYgqScJFx+cNmgvV9gP4zqMgpuzm6m7ZCIiKiZrHY338LCQhw8eBDjx4+39FDoN+be\ng+jePeB//9ewiOvWDXj9ddMWcbVSLY5fPY6vz3xtUMT5uPpgep/pmNBtQqso4rjnk1jMWzxmLhbz\nFstUeQst5FauXImBAwfC1dUVM2fONHiupKQEEydOhKenJ0JDQ5GYmKh/7osvvkB0dDQ+//xzAEBZ\nWRmmT5+ODRs28Iycnbh0SVfE3a3f6QPDhukup8rlpuvnVvktrP1/a3Hw0kGDjX2HPDME8wbNQ2cf\nM21IR0REZASh91r96aef4ODggAMHDuDhw4f47rvv9M9NnToVALBu3TpkZGRg7NixOHHiBHr06GFw\nDI1Gg7i4OLzzzjuIiYl5Yl+cI2c7zpwB9u3TzY0DACcnYMIEoFcv0/Wh1qpxpPAITlw7YbCxb6Bn\nIOIi4hDkFWS6zoiIiB5j1n3kTG3RokW4fv26vpCrrKyEr68vzp8/j7Aw3a2MXnvtNQQFBeGTTz4x\neO+mTZvw9ttvo9dv/4rPnTsXkydPbtAHCznrp9UC+/frCrk6Xl7AlClAsAlvmHD5/mUk5Sah5GGJ\nvs3JwQlRoVGI7BDJjX2JiMjszLrYwdQeH2hubi6cnJz0RRwA9OnTp9Hrx9OmTcO0adOa1M+MGTMQ\nGhoKAGjTpg369u2r37Ol7th8bLrHmZmZWLBggUmOt3+/Ekol4Oqqe3zlihJ+fsBf/xoFhcI041Vp\nVFA9o8LPt37GlcwrAIDQvqEIbRMKn9s+0FzSwDHEUVh+zX1syrz5+OmPmbf4x3VtrWU8tv64rq21\njMfWH2dmZuLBgwe4cuUKWqJVnJFLS0vD5MmTcevWLf1r1q5diy1btiA1NdWoPnhGTjylUqn/oLbE\nnTu6OzWU1J8gQ8+eusupzs4tPjwA4MKdC9ibtxcVNRX6NlcnV4zqMgr9AvtZxZYipsqbmoZ5i8fM\nxWLeYj2et1WfkfP09ERZWZlBW2lpKby8vEQOi1rIFF8AeXm67UVUqvq2mBjdwgZT1FblqnLszduL\n7LvZBu092vZAbFgsvFys5zPHL1yxmLd4zFws5i2WqfK2SCH3+NmO8PBwaDQa5Ofn6y+vZmVloWfP\nnpYYHlmAJOk2+D14UPdnQHf2LT4e6N7dFMeX8POtn5FckAyVtr5K9JJ74Q9d/4DubU3QCRERkWAO\nIjvTarWorq6GRqOBVquFSqWCVquFh4cH4uPjsXjxYlRVVeHYsWNISkpq8ly4J0lISDC49k/mZWzW\nGg2wcyeQnFxfxHl76/aHM0URd6/qHjZkbUBSbpJBETeg/QDMHzzfaos4frbFYt7iMXOxmLdYdXkr\nlUokJCQYfRyhZ+SWLVuGf/zjH/rHmzdvRkJCAhYvXozVq1dj1qxZaNeuHfz9/bFmzRp0b+G/4i0J\nhsSoqNDd9P7atfq2Z57R7Q/n6dmyY2trtThx7QSOFB6Bplajb/dz88P4iPEIbRPasg6IiIhaKCoq\nClFRUVi6dKlR77fIYgcRuNih9bt9W7eoobS0vq1vX92N751a+F+Mm+U3sfPiThRVFunbHGQOGPLM\nEAzvOBzOjiZaNUFERGQCVrXYgSg7G9ixA1Drbp4AmQwYORKIjGzZooYabQ2UV5Q4ee0kJNT/hQjy\nCkJcRBwCPQN/591ERETWRegcOdE4R06spmQtScDRo7rLqXVFnIsL8MorwHPPtayIKygpwNdnvsaJ\nayf0RZyzgzNGdxmNP/X/k80Vcfxsi8W8xWPmYjFvsaxyjpxonCPXuqjVukUNv/5a3+brC0ydCrRt\na/xxq9RVSC5IRubtTIP2zj6dMT58PHzcfIw/OBERkRlxjtwTcI5c61JWBmzdCty8Wd/WqRPw0kuA\nu7txx5QkCefvnMe+vH2oVFfq292c3DA6bDT6BPSxio19iYiIOEeOWq0bN3RFXHl5fdugQcCYMYCj\nkbcxLa0uxZ68Pci9l2vQ3rNdT4wJGwNPeQuXvBIREVkBm54jR2I1Nr/i3Dngu+/qizgHB2DsWN2P\nMUWcJEk4c+MMVp9ZbVDEKVwUmNpzKib1mGQ3RRzns4jFvMVj5mIxb7FMlbdNn5FLSEjQX3smsSQJ\nSEkB0tLq29zcgMmTdZdUjXGn8g525ezCtbJrBu2DgwdjRKcRcHFyacGIiYiIxFMqlS0q6jhHjkxO\npQJ++gm4eLG+zd9ftzLV17f5x9PWanHs6jEcLTwKraStP6a7P+Ii4hDiHWKCURMREVkO58hRq/Dg\ngW6T36L6fXgRFgZMmgS4ujb/eNfLrmNXzi4UVxbr2xxljhgaMhTDOg6DkwM/wkREZL84R45MZutW\nJb75xrCIe+453Zm45hZxNdoa7Mvbh3U/rzMo4jooOmDOwDmI7hRt90Uc57OIxbzFY+ZiMW+xOEeO\nWo2cnEJ8+20BUlN/gb9/LTp37oKAgI4YNw7o16/5x8u7l4fdubtRqqq/d5fcUY4RnUZgUPAgOMj4\n/w8iIiKAc+SohbKzC7F0aT5u3x6hb5PJDmPx4jBER3ds1rEqaypxoOAAfin6xaA9zDcM48LHoY1r\nG5OMmYiIqLXhHLlGcNWq+X3/fYFBEefhAfTqNQLZ2SlNLuQkScK54nPYn78fVeoqfbu7szvGhI1B\nr3a9uLEvERHZpJauWrXpa1R1hRyZj5+fAwJ/u4WpJCnRv79uPlxNTdM+Wg+qH+D7c99jR/YOgyKu\nd0BvzB80H70DerOIewLOZxGLeYvHzMVi3mLV5R0VFcV7rZLlyOW1CA8HFArdtiN1m/zK5bW/+75a\nqRbpN9KRcjkFNdoafbu3izfGR4xHmG+YOYdNRERkEzhHjlokJ6cQ69fnw8Wl/vKqSnUYM2aEISKi\n8UurxZXF2JWzC9fLruvbZJDh2Q7PIqZTDOSOcrOPm4iIqDUxtm5hIUctlpNTiMOHC1BT4wC5vBYj\nRnRptIjT1GqQVpiGtKtpqJXqz9i182iHuIg4dFB0EDlsIiKiVsPYusWm58iRGBERHTFvXgz69gXm\nzYtptIi7WnoVa86uwZHCI/oizlHmiOjQaMwZMIdFnBE4n0Us5i0eMxeLeYvFfeTIKqg0Khy6dAhn\nbp4xaA/xDsH48PFo69HWQiMjIiKyfjZ9aXXJkiXcfsSCcu7mYE/eHpSpyvRtLo4ueKHzCxgYNJCr\nUYmIyO7VbT+ydOlSzpF7FOfIWU5FTQX25e3D+TvnDdrD/cIxtutYeLt6W2hkRK0WOwMAABHDSURB\nVERErRPnyJHFpaamIvN2JlalrzIo4jycPTCpxyRM7TmVRZwJcT6LWMxbPGYuFvMWi3PkqNXIyc/B\nzlM7ceD4AbgEu6Bz587wD/IHAPQN7ItRXUbB3dndwqMkIiKyPby0Si2SnZeN/9n1P7jhf0O/GlWT\nr8HwfsPxevTr6OLbxcIjJCIiav14r1WyiC1Ht+Ca3zXgkc9ep/6dEKgKZBFHRERkZpwjRy3i6+4L\nXzdfAEB1XjUGtB+ALr5dUCv7/Vt0UctxPotYzFs8Zi4W8xaLc+SoVZA7yBHuF47iymJo/DXwcvHS\ntxMREZF5cY4ctUhOfg7Wp66HS1cXfZsqT4UZ0TMQERZhwZERERFZD86Ra0RCQgI3BDaziLAIzMAM\nHP75MGpqayB3kGNE9AgWcURERE1QtyGwsXhGjkxGqVSyaBaIeYvFvMVj5mIxb7Eez5sbAhMRERHZ\nGZ6RIyIiIrIwnpEjIiIisjMs5MhkuAeRWMxbLOYtHjMXi3mLZaq8WcgRERERWSnOkSMiIiKyMM6R\nIyIiIrIzLOTIZDi/QizmLRbzFo+Zi8W8xeIcOSIiIiI7Z9Nz5JYsWcJbdBEREVGrVXeLrqVLlxo1\nR86mCzkb/dWIiIjIxnCxA1kc51eIxbzFYt7iMXOxmLdYnCNHREREZOd4aZWIiIjIwnhplYiIiMjO\nsJAjk+H8CrGYt1jMWzxmLhbzFotz5IiIiIjsHOfIEREREVkY58gRERER2RkWcmQynF8hFvMWi3mL\nx8zFYt5icY4cERERkZ3jHDkiIiIiC+McOSIiIiI7w0KOTIbzK8Ri3mIxb/GYuVjMWyzOkSMiIiKy\nczY9R27JkiWIiopCVFSUpYdDRERE1IBSqYRSqcTSpUuNmiNn04Wcjf5qREREZGO42IEsjvMrxGLe\nYjFv8Zi5WMxbLM6RIyIiIrJzvLRKREREZGG8tEpERERkZ1jIkclwfoVYzFss5i0eMxeLeYvFOXJE\nREREdo5z5IiIiIgsjHPkiIiIiOwMCzkyGc6vEIt5i8W8xWPmYjFvsThHjoiIiMjOcY4cERERkYVx\njhwRERGRnWEhRybD+RViMW+xmLd4zFws5i0W58gRERER2TnOkSMiIiKyMM6RIyIiIrIzLOTIZDi/\nQizmLRbzFo+Zi8W8xeIcOSIiIiI7Z3Vz5IqKihAfHw+5XA65XI4tW7bAz8+vwes4R46IiIishbF1\ni9UVcrW1tXBw0J1I3LBhA27duoUPPvigwetYyBEREZG1sJvFDnVFHACUlZXBx8fHgqOhR3F+hVjM\nWyzmLR4zF4t5i2XXc+SysrLw7LPPYuXKlZg6daqlh0O/yczMtPQQ7ArzFot5i8fMxWLeYpkqb6GF\n3MqVKzFw4EC4urpi5syZBs+VlJRg4sSJ8PT0RGhoKBITE/XPffHFF4iOjsbnn38OAOjTpw9Onz6N\n5cuXY9myZSJ/BfodDx48sPQQ7ArzFot5i8fMxWLeYpkqb6GFXHBwMBYtWoRZs2Y1eG7+/PlwdXVF\ncXExvv/+e8ydOxcXLlwAALz99ttITU3F3/72N6jVav17FAoFVCqVsPH/npaeIm3u+5vy+t97zZOe\na2q7pU/Bm6L/5hzDXHk/6bmmtonU2j7jxj7PvI1/Pb9TTHcMfqfY9mdcZN5CC7mJEydiwoQJDVaZ\nVlZWYseOHVi2bBnc3d0xZMgQTJgwAZs2bWpwjMzMTDz//POIiYnBihUr8N5774ka/u+y5Q9kY+2N\nve7KlStPHZOp8EtXbN6N9W/u97e2Qs7e837aa/idwu+U5rLlz7jIvC2yavWjjz7CjRs38N133wEA\nMjIyMHToUFRWVupfs2LFCiiVSuzatcuoPsLCwlBQUGCS8RIRERGZU58+fYyaN+dkhrE8lUwmM3hc\nUVEBhUJh0Obl5YXy8nKj+8jPzzf6vURERETWwCKrVh8/Cejp6YmysjKDttLSUnh5eYkcFhEREZFV\nsUgh9/gZufDwcGg0GoOzaFlZWejZs6fooRERERFZDaGFnFarRXV1NTQaDbRaLVQqFbRaLTw8PBAf\nH4/FixejqqoKx44dQ1JSEqZNmyZyeERERERWRWghV7cq9dNPP8XmzZvh5uaGjz/+GACwevVqPHz4\nEO3atcOrr76KNWvWoHv37iKHR0RERGRVrO5eqy31/vvv4+TJkwgNDcW3334LJyeLrPewG2VlZXjh\nhReQnZ2N06dPo0ePHpYekk1LT0/HggUL4OzsjODgYGzcuJGfcTMqKipCfHw85HI55HI5tmzZ0mB7\nJTKPxMRE/OUvf0FxcbGlh2LTrly5gkGDBqFnz56QyWTYvn07/P39LT0sm6ZUKrF8+XLU1tbiz3/+\nM1588cXffb1V3qLLWFlZWbh58yaOHj2Kbt264ccff7T0kGyeu7s79u7di0mTJhl1M2BqnpCQEKSm\npuLIkSMIDQ3Fzp07LT0km9a2bVscP34cqampeOWVV7B27VpLD8kuaLVa/PDDDwgJCbH0UOxCVFQU\nUlNTkZKSwiLOzB4+fIgVK1Zg3759SElJeWoRB9hZIXfy5EmMHj0aADBmzBgcP37cwiOyfU5OTvyL\nL1BgYCBcXFwAAM7OznB0dLTwiGybg0P9V2hZWRl8fHwsOBr7kZiYiMmTJzdYOEfmcfz4cQwfPhwL\nFy609FBs3smTJ+Hm5obx48cjPj4eRUVFT32PXRVy9+/f129polAoUFJSYuEREZlHYWEhDh48iPHj\nx1t6KDYvKysLzz77LFauXImpU6daejg2r+5s3Msvv2zpodiFoKAgFBQU4OjRoyguLsaOHTssPSSb\nVlRUhPz8fOzevRtvvPEGEhISnvoeqyzkVq5ciYEDB8LV1RUzZ840eK6kpAQTJ06Ep6cnQkNDkZiY\nqH+uTZs2+v3qSktL4evrK3Tc1szYzB/F/z03XUvyLisrw/Tp07FhwwaekWuiluTdp08fnD59GsuX\nL8eyZctEDtuqGZv55s2beTbOCMbmLZfL4ebmBgCIj49HVlaW0HFbK2Pz9vHxwZAhQ+Dk5ISYmBic\nP3/+qX1ZZSEXHByMRYsWYdasWQ2emz9/PlxdXVFcXIzvv/8ec+fOxYULFwAAzz33HA4dOgQAOHDg\nAIYOHSp03NbM2MwfxTlyTWds3hqNBlOmTMGSJUvQtWtX0cO2WsbmrVar9a9TKBRQqVTCxmztjM08\nOzsbGzduRGxsLPLy8rBgwQLRQ7dKxuZdUVGhf93Ro0f5vdJExuY9aNAgZGdnA9DdW75Lly5P70yy\nYh999JE0Y8YM/eOKigpJLpdLeXl5+rbp06dLH3zwgf7xu+++Kw0bNkx69dVXJbVaLXS8tsCYzGNj\nY6WgoCApMjJSWr9+vdDxWrvm5r1x40bJz89PioqKkqKioqRt27YJH7M1a27ep0+floYPHy5FR0dL\no0aNkq5duyZ8zNbOmO+UOoMGDRIyRlvS3Lz37t0rDRgwQBo2bJj02muvSVqtVviYrZkxn+9Vq1ZJ\nw4cPl6KioqRLly49tQ+r3pdAeuwMT25uLpycnBAWFqZv69OnD5RKpf7xP//5T1HDs0nGZL53715R\nw7M5zc172rRp3Ei7BZqb9+DBg3HkyBGRQ7Q5xnyn1ElPTzf38GxOc/OOjY1FbGysyCHaFGM+3/Pm\nzcO8efOa3IdVXlqt8/gciYqKCigUCoM2Ly8vlJeXixyWTWPmYjFvsZi3eMxcLOYtloi8rbqQe7zS\n9fT01C9mqFNaWqpfqUotx8zFYt5iMW/xmLlYzFssEXlbdSH3eKUbHh4OjUaD/Px8fVtWVhZ69uwp\nemg2i5mLxbzFYt7iMXOxmLdYIvK2ykJOq9WiuroaGo0GWq0WKpUKWq0WHh4eiI+Px+LFi1FVVYVj\nx44hKSmJc4ZMgJmLxbzFYt7iMXOxmLdYQvNu6YoMS1iyZIkkk8kMfpYuXSpJkiSVlJRIL774ouTh\n4SF17NhRSkxMtPBobQMzF4t5i8W8xWPmYjFvsUTmLZMkbu5FREREZI2s8tIqEREREbGQIyIiIrJa\nLOSIiIiIrBQLOSIiIiIrxUKOiIiIyEqxkCMiIiKyUizkiIiIiKwUCzkiIiIiK8VCjojoMTNmzMCi\nRYtMesy5c+di+fLlJj0mEZGTpQdARNTayGSyBje7bqmvv/7apMcjIgJ4Ro6IqFG8eyERWQMWckTU\nqnz66afo0KEDFAoFunXrhpSUFABAeno6IiMj4ePjg6CgILz11ltQq9X69zk4OODrr79G165doVAo\nsHjxYhQUFCAyMhJt2rTBlClT9K9XKpXo0KEDPvnkE7Rt2xadOnXCli1bnjim3bt3o2/fvvDx8cGQ\nIUNw7ty5J7727bffRkBAALy9vdG7d29cuHABgOHl2vHjx8PLy0v/4+joiI0bNwIALl68iJEjR8LP\nzw/dunXDDz/88MS+oqKisHjxYgwdOhQKhQKjR4/GvXv3mpg0EdkCFnJE1Grk5ORg1apVOHv2LMrK\nypCcnIzQ0FAAgJOTE/7973/j3r17OHnyJA4fPozVq1cbvD85ORkZGRk4deoUPv30U7zxxhtITEzE\n1atXce7cOSQmJupfW1RUhHv37uHmzZvYsGEDZs+ejby8vAZjysjIwOuvv461a9eipKQEc+bMQVxc\nHGpqahq89sCBA0hLS0NeXh5KS0vxww8/wNfXF4Dh5dqkpCSUl5ejvLwc27dvR/v27TFixAhUVlZi\n5MiRePXVV3Hnzh1s3boV8+bNQ3Z29hMzS0xMxPr161FcXIyamhp89tlnzc6diKwXCzkiajUcHR2h\nUqlw/vx5qNVqhISEoHPnzgCA/v37Y/DgwXBwcEDHjh0xe/ZsHDlyxOD97733Hjw9PdGjRw/06tUL\nsbGxCA0NhUKhQGxsLDIyMgxev2zZMjg7O2P48OEYO3Ystm3bpn+uruj65ptvMGfOHAwaNAgymQzT\np0+Hi4sLTp061WD8crkc5eXlyM7ORm1tLSIiIhAYGKh//vHLtbm5uZgxYwa2b9+O4OBg7N69G506\ndcJrr70GBwcH9O3bF/Hx8U88KyeTyTBz5kyEhYXB1dUVkydPRmZmZjMSJyJrx0KOiFqNsLAwfPnl\nl0hISEBAQACmTp2KW7duAdAVPePGjUP79u3h7e2NhQsXNriMGBAQoP+zm5ubwWNXV1dUVFToH/v4\n+MDNzU3/uGPHjvq+HlVYWIjPP/8cPj4++p/r1683+tro6Gi8+eabmD9/PgICAjBnzhyUl5c3+ruW\nlpZiwoQJ+Pjjj/Hcc8/p+zp9+rRBX1u2bEFRUdETM3u0UHRzczP4HYnI9rGQI6JWZerUqUhLS0Nh\nYSFkMhnef/99ALrtO3r06IH8/HyUlpbi448/Rm1tbZOP+/gq1Pv376Oqqkr/uLCwEEFBQQ3eFxIS\ngoULF+L+/fv6n4qKCrz88suN9vPWW2/h7NmzuHDhAnJzc/Gvf/2rwWtqa2vxyiuvYMSIEfjTn/5k\n0Nfzzz9v0Fd5eTlWrVrV5N+TiOwLCzkiajVyc3ORkpIClUoFFxcXuLq6wtHREQBQUVEBLy8vuLu7\n4+LFi03azuPRS5mNrUJdsmQJ1Go10tLSsGfPHrz00kv619a9/o033sCaNWuQnp4OSZJQWVmJPXv2\nNHrm6+zZszh9+jTUajXc3d0Nxv9o/wsXLkRVVRW+/PJLg/ePGzcOubm52Lx5M9RqNdRqNc6cOYOL\nFy826XckIvvDQo6IWg2VSoUPP/wQbdu2Rfv27XH37l188sknAIDPPvsMW7ZsgUKhwOzZszFlyhSD\ns2yN7fv2+POPPg4MDNSvgJ02bRr+85//IDw8vMFrBwwYgLVr1+LNN9+Er68vunbtql9h+riysjLM\nnj0bvr6+CA0Nhb+/P959990Gx9y6dav+EmrdytXExER4enoiOTkZW7duRXBwMNq3b48PP/yw0YUV\nTfkdicj2yST+d46I7IxSqcS0adNw7do1Sw+FiKhFeEaOiIiIyEqxkCMiu8RLkERkC3hplYiIiMhK\n8YwcERERkZViIUdERERkpVjIEREREVkpFnJEREREVoqFHBEREZGV+v+4w6N5N8cjyQAAAABJRU5E\nrkJggg==\n", + "text": [ + "
\n", - "
" - ] + "outputs": [], + "prompt_number": 117 }, { - "cell_type": "markdown", + "cell_type": "code", + "collapsed": false, + "input": [ + "%pylab inline" + ], + "language": "python", "metadata": {}, - "source": [ + "outputs": [] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "import matplotlib.pyplot as plt\n", + "\n", + "labels = [('reverse_func', 'copy.deepcopy(my_list).reverse()'), \n", + " ('reversed_func', 'list(reversed(my_list))'),\n", + " ('reverse_slizing', 'my_list[::-1]'),\n", + " ] \n", + "\n", + "matplotlib.rcParams.update({'font.size': 12})\n", + "\n", + "fig = plt.figure(figsize=(10,8))\n", + "for lb in labels:\n", + " plt.plot(orders_n, times_n[lb[0]], \n", + " alpha=0.5, label=lb[1], marker='o', lw=3)\n", + "plt.xlabel('sample size n')\n", + "plt.ylabel('time per computation in milliseconds [ms]')\n", + "#plt.xlim([1,max(orders_n) + max(orders_n) * 10])\n", + "plt.legend(loc=2)\n", + "plt.grid()\n", + "plt.xscale('log')\n", + "plt.yscale('log')\n", + "plt.title('Performance of different list reversing approaches')\n", + "\n", + "max_perf = max( f/s for f,s in zip(times_n['reverse_func'],\n", + " times_n['reverse_slizing']) )\n", + "min_perf = min( f/s for f,s in zip(times_n['reverse_func'],\n", + " times_n['reverse_slizing']) )\n", + "\n", + "ftext = 'my_list[::-1] is {:.2f}x to '\\\n", + " '{:.2f}x faster than copy.deepcopy(my_list).reverse()'\\\n", + " .format(min_perf, max_perf)\n", + "plt.figtext(.14,.75, ftext, fontsize=11, ha='left')\n", + "\n", + "plt.show()" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "display_data", + "png": "iVBORw0KGgoAAAANSUhEUgAAAnYAAAIECAYAAACUvmMzAAAABHNCSVQICAgIfAhkiAAAAAlwSFlz\nAAALEgAACxIB0t1+/AAAIABJREFUeJzsnXlczdn/x1/3tpfbqk3RqlKRpcWgulRI1jBfe9nGxMxg\nmPkZNOqLmBnL12C2GNkNZuwUbciaKCSVUmJkjySt798fd/pMn26lcmlxno/HffB5n3Pe530+n889\nve8573OOgIgIDAaDwWAwGIxmj7CxDWAwGAwGg8FgyAbm2DEYDAaDwWC0EJhjx2AwGAwGg9FCYI4d\ng8FgMBgMRguBOXYMBoPBYDAYLQTm2DEYDAaDwWC0EJhjx2j2lJaWYtKkSWjdujWEQiFOnTrV2CY1\nS/bs2QMLCwvIy8tj0qRJdS4XFBSE9u3b13hdk+7Y2FjY29tDUVERffr0kU0j3hNV2xgWFgYFBYVG\ntKjpkJWVBaFQiLNnzza2KS0Gf39/eHl5NbYZjGYCc+wY7wV/f38IhUIIhUIoKCjA1NQUAQEBePr0\n6Vvr/vPPP7Fz504cPnwYubm5+Oijj2Rg8YdFWVkZJk2ahFGjRiEnJwdr1qxpsK6vvvoKFy5ceKPu\ngIAAODo64vbt2/jrr7/eug2ywNLSEsHBwXXKKxAIuP+PGjUKf//9d53r8fT0xMSJE+ttX3OgXbt2\nyM3NhbOzc2Ob0mIQCAS8943BqA35xjaA8eHg5uaG3bt3o7S0FJcuXcLUqVORk5ODw4cPN0hfcXEx\nFBUVkZ6eDiMjI3Tv3v2t7KvQ9yHy999/o6CgAN7e3jA0NHwrXWpqalBTU6tVNxHh1q1bWLBgAYyM\njBpcFxGhrKwM8vKy6crq88ez8t7uysrKUFZWlokNtVFeXg4AEAob5zd5Xb4jQqEQenp678mixuN9\nPgt2jgCjPrARO8Z7Q0FBAXp6emjTpg0GDx6MmTNnIjw8HEVFRQCAXbt2oXPnzlBRUYGZmRnmzJmD\nV69eceXFYjGmTJmCwMBAtGnTBiYmJujduze+/fZbZGZmQigUwtzcHABQUlKCefPmwdjYGEpKSrCz\ns8POnTt59giFQqxduxZjxoyBpqYmJkyYwE2pxcbGomPHjlBVVUWfPn2Qm5uLmJgYdO7cGa1atYKX\nlxdvhOb27dvw9fWFkZER1NTU0KlTJ2zbto1Xn1gsxtSpU7F48WIYGhpCR0cHfn5+KCgo4OX7448/\n0K1bN6ioqKB169YYMGAA8vLyuPS1a9fCxsYGKioqsLKyQkhICMrKymq99+fPn4ebmxtUVVWhra2N\nsWPH4tGjRwAk04gmJiYAJM53bdPZr1+/RkBAADQ1NaGtrY3p06dzz6+CytOUVXXLycnh5MmTkJOT\nQ1lZGSZMmAChUIgtW7YAAG7duoXhw4dDS0sL2tra6NevH65fv87prvx8unTpAmVlZURFRaGkpARB\nQUEwNzeHiooK7O3t8dtvv0k9759//hnjx4+Huro62rZti+XLl/OeT0ZGBoKDg7nR5Tt37tR6X6va\nVcGLFy8wceJEGBoaQllZGe3atcOcOXMASEavo6OjsXnzZq6emu53xb3cvXs3bGxsoKSkhPT0dLx8\n+RIzZ86EsbEx1NTU0LVrV+zbt48r17NnT0ybNk1KX4cOHfDtt99y1/X5zhkaGsLU1BQAcODAAXTp\n0gVqamrQ0tKCi4sLEhMTAUhPxVZc79mzBwMHDoSamhosLCywefNmnm23b99G3759oaKiAlNTU/z6\n66/cd6Y2pk6dCktLS6iqqsLCwgILFixAcXGx1D3csWMH93707dsX2dnZDcpT9Vncv38fo0aNgpaW\nFlRVVdG7d28kJCTUy0YAiIyMhKurK9TU1KCpqQmxWIzMzEwunYjw22+/wcTEBBoaGhgyZAgePnzI\n03HixAn07NkTqqqqMDY2xqRJk3izIsnJyejXrx+0tLTQqlUr2NraSvVTjBYAMRjvAT8/P/Ly8uLJ\nVq5cSQKBgF6+fEmbNm0iLS0t2rZtG92+fZtOnTpFnTp1ovHjx3P53d3dSSQSUUBAAKWkpND169fp\n6dOnNHfuXDIzM6MHDx7Q48ePiYho7ty5pKOjQ3v37qX09HQKCQkhoVBIUVFRnD6BQEA6Ojq0fv16\nyszMpPT0dNq0aRMJhULq3bs3Xbx4kS5fvkzt27enXr16kZubG124cIESExPJxsaG/vOf/3C6rl27\nRuvXr6erV69SZmYmrV27luTl5SkmJoZnv6amJn355ZeUmppKx48fJ21tbQoMDOTy/P7776SgoEBL\nlizh2rhu3TquXYsWLSITExPav38/ZWVl0dGjR6ldu3Y8HVW5f/8+iUQiGjt2LF2/fp3i4uKoU6dO\n5ObmRkREhYWFFB8fTwKBgA4dOkQPHjyg4uLianXNmjWL9PT06ODBg5Samkpz584ldXV1at++PZdn\n0aJF3HVNunNzc0kgENBPP/1EDx48oMLCQsrNzSV9fX2aPn06Xb9+ndLS0ujzzz8nHR0devToERER\n93xcXFwoNjaWbt++TY8ePSI/Pz9ycHCgEydOUFZWFv3xxx+kqalJGzdu5D1vfX192rBhA2VmZtL6\n9etJIBBw78TTp0/JzMyMvvrqK3rw4AE9ePCAysrKqr0PixYtIktLS+5606ZNJC8vz11//vnn5ODg\nQBcvXqScnBw6e/YsbdiwgYiInj9/Tm5ubjRq1Ciunpru96JFi0hVVZXEYjFdvHiR0tPTKT8/n8Ri\nMfXu3ZvOnDlDt2/fpt9++40UFRW5tvz222+kpaVFRUVFnK4LFy6QQCCg9PR0zuaGfOfu379PCgoK\n9MMPP1BWVhbdvHmTdu7cSdeuXSMiotu3b5NAIKAzZ87wrs3NzWnPnj2UkZFB8+fPJ3l5eUpLSyMi\novLycnJwcKDu3btTfHw8JSYm0oABA0hDQ4OmTp1a7b2pKLdgwQK6ePEiZWdn08GDB8nQ0JAWLVrE\nu4dqamrk6upKCQkJFB8fTy4uLtS1a9d656nuWTg7O1OXLl3ozJkzdO3aNfrPf/5DWlpa3He2Ljae\nOHGC5OTkaPbs2XT16lVKTU2lsLAwSk1NJSJJ/6mhoUFjxoyh5ORkOnfuHJmZmfGeVVRUFKmqqtK6\ndevo1q1bFB8fT7179yZ3d3cuT8eOHWns2LGUkpJCt2/fpmPHjtHhw4drvL+M5glz7BjvBT8/P/L0\n9OSuk5OTydzcnD766CMiIjIxMaFff/2VV+bkyZMkEAgoLy+PiCR/ZKytraV0V/0jW1BQQEpKSvTz\nzz/z8g0bNoz69OnDXQsEApoyZQovz6ZNm0ggEFBSUhIn++GHH0ggENDly5c52erVq6l169a1tnnI\nkCG8P0ru7u7UuXNnXp6AgADuHhARtW3blj7//PNq9RUUFJCqqipFRETw5Js3byZNTc0a7Vi4cCG1\nbduWSkpKOFlSUhIJBAI6deoUEUn/Ma6Oly9fkrKyMuegVODo6Cjl2FV+HjXpFggEtH37dl657t27\n8/KUl5eThYUF/e9//yOif59PXFwclyczM5OEQiH3R7CC4OBg3v0WCAQ0c+ZMXp4OHTrQN998w11b\nWlpScHBwjfegpjZWdeyGDBlC/v7+NZb39PSkiRMn1qkeoVBIOTk5nCwmJoaUlZXp+fPnvLwTJ06k\noUOHEhHRs2fPSEVFhfbs2cOlz5gxg3r06MFdN/Q7d/nyZRIIBJSVlVWtzTU5dqtXr+bylJWVkUgk\not9++42IiI4fP04CgYAyMjK4PE+fPiVVVdVaHbvqWLVqldT7WFV3WloaCQQCio6Orleeqs8iMjKS\nBAIBpaSkcLKioiIyNDSk//73v3W2sVevXjRo0KAa8/v5+ZG+vj7vB8B3331HhoaG3LW7uzvvXSYi\nys7O5vVnGhoaFBYWVmM9jJYBm4plvDdiY2MhEomgqqqKjh07wtLSEtu3b8ejR49w584dzJ49GyKR\niPsMGDAAAoEAt27d4nR069btjfXcunULxcXFcHNz48nd3NyQnJzMk1UX4C0QCNCxY0fuWl9fHwDQ\nqVMnnuzJkydc7MurV68wb9482NvbQ0dHByKRCEePHuVN5QkEAjg4OPDqMjQ0xIMHDwAADx8+xN27\nd9G3b99q25WcnIzCwkL4+vry7tOnn36KFy9e4MmTJzWW6969Oy8OrVOnTtDQ0MCNGzeqLVMdGRkZ\nKCoqQo8ePXjynj17yiQGKD4+HgkJCby2qaurIzs7m/cOAICTkxP3/0uXLoGI0K1bN17ZZcuWSZXr\n3Lkz77pNmzZS01myYPr06di7dy86duyIWbNmITw8vMH3SF9fH8bGxtx1fHw8iouLYWRkxGvv9u3b\nufZqampi8ODB2Lp1KwBJaMKuXbswYcIEAHir75yDgwP69esHe3t7+Pr64scff8Tdu3ff2I7K974i\nDq/i3b9x4wZat27NhVIAgJaWFqytrd+oNzQ0FC4uLjAwMIBIJML8+fOlptB1dXV5utu3b4/WrVvz\n+oO65Kn6LJKTk6GjowMbGxtOpqioCBcXF165N9l4+fLlGr/3FdjY2PCm+yv3HYDkvVi9ejXvedrZ\n2UEgECA9PR0AMHfuXEyZMgW9e/dGcHAwrly5UmudjOYJWzzBeG90794dmzdvhry8PNq0acM5GhWd\n048//ojevXtLlasIrhcIBLygfFlQnT6hUMgLoq/4v5ycnJSMiCAQCPDVV1/h4MGDWL16NaytraGq\nqoo5c+bg+fPnPN1VA88FAgEXhP0mKvLt3bsXVlZWUulaWlrVlhMIBM0i+JqI4OnpiXXr1kmlaWho\ncP+Xk5Pj3ceK+3Lu3DmoqqryylVdDPE2978+9O3bF3fu3EFERARiY2Mxbtw4dOzYEVFRUfUOtq/6\njpaXl0NDQwOXLl2Sylu5fRMmTMCwYcPw+PFjxMXFoaCgAKNGjeJ0AA37zgmFQhw7dgzx8fGIjIzE\nn3/+iXnz5mHPnj3w8fGpsR1vuvfVLVx503u7Z88efPbZZ/juu+/g7u4OdXV17N69GwsWLKi1XEOp\na/9T0S/I0saq2+lU/V4TEebNm4fx48dLla34cbpw4UKMHTsW4eHhiI6ORkhICL7++mssXry4XrYw\nmjbMsWO8N5SVlXm/iCvQ19dH27ZtcfPmTUyePPmt67G0tISSkhJOnjwJW1tbTn7y5EneSJwsOX36\nNMaNG4cRI0YAkPzhTE1NrdcKUz09PRgbGyMiIgIDBw6USrezs4OysjIyMjLQv3//Ouu1s7PDpk2b\nUFJSwv1xSEpKwvPnz2Fvb19nPRYWFlBUVMSZM2fQoUMHTn7mzBmZbMXg6OiIsLAwGBkZQUlJqc7l\nKkaUsrOza3Us6oKiouIbF6LUFS0tLYwaNQqjRo3CxIkT8dFHHyElJQV2dnZQVFREaWlpg/Q6OTkh\nLy8PhYWFsLOzqzFf3759oa2tjV27diE6OhqDBg3iHGRZfOecnJzg5OSEb775Bt7e3ti0aVOD77+t\nrS0ePXqEzMxMro949uwZ0tLSeKOzVTl16hS6dOmCWbNmcbLbt29L5auqOy0tDY8fP+b1D3XJUxU7\nOzs8efIEKSkp3HeiqKgIFy5cwGeffVZnG7t164aIiAiuTHW86Tvm6OiI69evV9vHVsbMzAwBAQEI\nCAjA8uXLsWLFCubYtTCapWN38eJFzJo1CwoKCjAyMsKWLVtktt0Bo3FYunQpJk+eDC0tLQwePBgK\nCgpISUlBeHg4fvnlFwCSX6R1GXlSVVXFF198gcDAQOjq6qJTp07Yu3cvDh48iMjIyHdiv7W1Nfbv\n3w9fX1+oqalh1apVuH//PgwMDLg8dbF/0aJFCAgIgL6+PoYPH47y8nLExMRg9OjR0NHRwfz58zF/\n/nwIBAJ4eHigtLQU165dQ2JiIm+FZ2U+++wzrFmzBv7+/pg/fz6ePXuG6dOnw83NDT179qxzG9XU\n1PDpp59i4cKF0NfXh5WVFTZu3Ii0tDSZbG/x2WefYePGjRgyZAgWLlwIY2Nj3L17F8eOHcPAgQNr\n3J/Q0tISkyZNwtSpU/H999+je/fuKCgoQEJCAh4/foyvv/66xjqrPhMzMzPExcUhJycHKioq0NHR\naZDTumDBAjg6OsLW1hZCoRDbtm2DSCRCu3btuHpiYmKQmZkJdXV1aGpq1rkP69OnDzw9PeHr64vv\nv/8eHTt2xLNnz3D27FmoqKhgypQpAAB5eXmMGTMGP/30EzIzM/Hnn3/y9DT0O3fu3DlERkaiX79+\nMDAwQHp6Oq5evcrVW1cq6/Xy8oKDgwPGjx+PNWvWQEFBAQsWLICCgkKt99/Gxga///47Dh48CDs7\nOxw+fJi3OrgCVVVVTJw4EatWrQIR4fPPP0eXLl14G2PXJU9VPDw84OzsjDFjxmD9+vVQV1fH4sWL\nUVxcjICAgDrbGBgYCG9vb8yePRsTJ06EkpISzp07hx49enCj82/qO/773/+ib9++mDNnDsaPHw+R\nSIT09HTs3bsX69atQ1lZGb7++muMGDECpqamyMvLQ3h4eK0/DhjNk2YZY9euXTvExMTg5MmTMDU1\nxYEDBxrbJMYbeNMGm+PGjcPu3btx+PBhuLi4wNnZGcHBwbx4lpp0VCdfunQppk6dilmzZqFjx47Y\nsWMHtm/fXu20U3X66itbvXo1t/2Kp6cn2rZtixEjRkhN6VbVU1U2efJkhIWFYe/evejSpQvc3d0R\nERHB/dFfuHAhVq1ahdDQUHTu3Bmurq5Ys2YNzMzMamyPnp4ejh8/jrt378LJyQmDBg3inN03tbEq\ny5cvx9ChQzF+/Hi4uLjgxYsXmDFjRp3a+Sb09PRw7tw5tG7dGr6+vrCxscG4ceOQk5ODNm3a1Krr\nt99+w+zZs7F06VLY2dnB09MTW7duhYWFRa11VrU1ODgYeXl5sLa2hr6+PnJycupUrqpdKioq+Pbb\nb+Ho6AgnJydcv34dx44dg0gkAgDMmTMHrVu3hoODA/T19Ws8paGmd/7gwYPw9fXF7Nmz0aFDBwwc\nOBDHjh2DpaUlL5+fnx9u3rwJTU1NeHt789Ia+p3T0NDA+fPnMWTIEFhZWWHy5MkYN24cAgMDq70X\n1V1XJ9u3bx/U1NTg6uqKwYMHw8fHB9bW1rXuDzht2jSMHz8eEydORNeuXREfH4+goCAp3YaGhpg2\nbRpGjBgBV1dXtGrVSmpT7DflqelZ7N+/HzY2NvDx8YGzszMePnyIEydOQFtbu842enl54ejRo7hw\n4QK6d+8OFxcXbN26lZu+rq3vq0AsFiM6OhpXr16Fm5sbHBwc8OWXX0JdXR0KCgqQl5dHXl4eJk+e\nDFtbW/Tv3x+GhobYsWNHjfeX0TwRUHMIvqmFRYsWoUuXLhg6dGhjm8JgMBgMGZGfnw9jY2OEhIRg\nxowZDdYTFBSE7du3cwsIGpqHwWguNOv5y+zsbJw4cYK34SaDwWAwmh+HDh2CnJwcOnTogIcPHyI4\nOBhycnL4+OOPG9s0BqNZ0ahTsevWrYOjoyOUlZWlzk18+vQphg0bhlatWsHU1FTq1IAXL15gwoQJ\n2Lx5M2+1IoPBYDCaH69evcJXX30Fe3t7DBo0CAAQFxcHXV3dt9Jbl3NW2VmsjJZEo07F7tu3D0Kh\nEBERESgsLMSmTZu4tNGjRwMANm7ciCtXrsDHxwdnz56Fra0tSktLMXjwYMydO7fWwFYGg8FgMBiM\nD4r3tBFyrSxcuJC3S/vLly9JUVGRO/qGiGjChAk0b948IiLasmUL6ejokFgsJrFYTH/88YeUzjZt\n2hAA9mEf9mEf9mEf9mGfJv/R0rKQiU/VJFbFUpVBw7S0NMjLy/NWeDk4OHA7eY8fPx6PHz9GTEwM\nYmJiqo3B+Pvvv7ml+k35s2jRomZRR0N11KdcXfK+KU9t6Q1Na0qfd22nrPQ3RI+s35W65GvIO8He\nFdnWwfqWpvFhfUv98r7t+1JQQDh2jNC/fwzc3Qnu7oRnzzJk4lM1icUTVWMbXr58CXV1dZ5MJBIh\nPz//fZr1XhCLxc2ijobqqE+5uuR9U57a0mtLy8rKemPdTYF3/b7ISn9D9Mj6XalLvoa8L+xdkW0d\nrG9pGrC+pX55G/q+FBcDmppirFkDFBUBAoHsT75pEtudLFy4EPfu3eNi7K5cuYJevXqhoKCAy7Ni\nxQqcOnUKBw8erJPO5nKMEqNp4O/vj7CwsMY2g9EMYO8Koz6w94UBAGVlwOXLwMmTwMuX/8ofP85G\nZuYtWFl5YM0a2fgtTWIqtuqInZWVFUpLS3kHUSclJdXr+CNAsjdRbGysLExktHD8/f0b2wRGM4G9\nK4z6wN6XDxsi4Pp1YP164MgRvlOnqwt89pkJRo16hPR0P5nV2agjdmVlZSgpKUFwcDDu3buH0NBQ\nyMvLQ05ODqNHj4ZAIMCGDRtw+fJlDBw4EOfOneOdUVkbbMSOwWAwGAxGY0AEZGYCkZHA/fv8NHV1\noHdvwMEBEFYaXpOV39KoI3aLFy+GqqoqvvvuO2zbtg0qKipYunQpAOCnn35CYWEh9PT0MG7cOPzy\nyy91duoYjPrCRnYZdYW9K4z6wN6XD49794AtW4CtW/lOnYoK4OUFfP450KUL36mTJY26eCIoKAhB\nQUHVpmlpaVV7mDODwWAwGAxGU+PJEyAqCrhxgy9XUABcXIBevYBajj6WGU1iVey7IigoCGKxWGp1\nira2Np49e9Y4RjEYjA8eLS0tPH36tLHNYLwH3sfqZEbjkp8PxMYCV64A5ZUWuQqFkpE5sRgQiWou\nHxsbK9OR3SaxKvZdUNtcNYu/YzAYjQnrgxiM5s/r18CZM8D580BJCT/N1hbo0wdo3bru+mTVL7To\nETsGg8FgMBqT2NhYNmrXwigtBS5eBE6fBgoL+WlmZoCnJ2Bk1Di2AcyxYzAYDAaDwXgj5eVAUhIQ\nEwO8eMFPMzCQOHQWFkCVHdzeOy3asaspxo7BYDAYjPcB+/vT/CECUlMlCyMePeKnaWlJplzt7Rvu\n0LEYuzrCYuwYDEZThfVBDEbzIDtbshddTg5frqYGuLsD3boBcnKyqYvF2DEYDAaD0cRhMXbNkwcP\nJCN0aWl8uZIS0KMH8NFHgKJi49j2JprEkWKMloepqSm32fSHzIEDB9CxY8fGNgNisRhTp07lrv39\n/eHl5fVWOuPi4mBqaoqioqK3Na9ZU/UZb9u2Dd27d29EixgMRkPJywP27QN++YXv1MnJAd27A198\nIRmpa6pOHcBG7GokNTUbkZEZKCkRQkGhHJ6eFrC2Nmky+po6AoFA6gzgD43y8nJ8/fXXWLx4cWOb\nIvU81q5di/LKGy69AXl5efz++++YMGECJ+vVqxcsLS2xbt06zJkzR6b2Nheqe8ZjxozB4sWL8eef\nf2L48OGNaB2jKcBG65oHBQWSVa7x8UBZ2b9ygQDo1ElyBJimZuPZVx9a9IhdUFBQgwISU1OzERZ2\nC48e9UFenhiPHvVBWNgtpKZmN8gOWetjNA+OHTuGJ0+ewNfXt7FNkUIkEkFDQ6PO+WuK/Zg0aRLW\nrVtX77iQ0tLSeuWXJeXl5fVyamujumcsFArh5+eHH3/8USZ1MBiMd0dxMXDyJPDjj5L96Co7dVZW\nwKefAsOGvVunLjY2tsZTuBpCi3fsGvJrKTIyA0pKHoiNBfc5d84DX36ZgaAg1Psze3YGzp3j61NS\n8kBUVEaD27Z+/XrY2tpCWVkZ+vr6GDFiBAAgPz8f06ZNg56eHpSVleHk5IQTJ05w5bKysiAUCrF9\n+3Z4eHhAVVUVFhYW+OOPP7g8YrEY06ZN49VHRLCwsKh2ejUpKQk9evSAsrIyrKyssHv3bqk8L1++\nxMyZM2FsbAw1NTV07dpV6si4Bw8ewN/fH3p6elBXV0evXr1w+vRpLj02NhZCoRCHDx+Gs7MzVFRU\n0LFjR8TExPD0ZGRkYMSIEdDR0YGamhocHBxw5MgRLv3o0aPo1q0bd+9mzJiBV69ecekV05SrV6+G\nkZER1NTU8PHHH3OnlcTGxkJeXh53797l1btlyxZoamqi8J+NjbZt24aBAwdCXv7fgfGgoCC0b98e\ne/bsgaWlJdTU1DB8+HC8fPkSe/bsgbW1NdTV1TFy5Ei8+Gc9fV3rqw9Vp2KTk5PRr18/aGlpoVWr\nVrC1tcW2bdsASKbVy8rKMHHiRAiFQshVihQeNGgQcnJyEBcXV2NdFc/t6NGj6NWrF1RUVLBx40YA\nkpFDGxsbqKiowMrKCiEhISj7p2ddsGABbGxspPQFBATA1dWVu05ISEDfvn0hEomgp6eH4cOH486d\nO1x6xT3fvXs3bGxsoKSkhPT09FrbDNTtna3uGQPA0KFDcfr0aeRUjbhmfHCws2KbJmVlkr3o1qyR\nbF9SOaKkbVtg4kRgzBhAX//d2yIWi5lj964pKan+tpSVNex2lZdXX664uGH6Fi1ahHnz5uGzzz7D\n9evXcfz4cTg6OgKQjKCcOHEC27dvR1JSEnr27ImBAwciNTWVp+Prr7/GlClTkJSUhDFjxmDs2LFI\nTEwEAHz66afYuXMnCgoKuPzR0dG4c+cOJk+ezNNTWFiIAQMGQFtbG/Hx8diyZQtWrFiBhw8fcnmI\nCIMGDcK1a9ewe/duJCcnIyAgAKNGjUJ0dDSnp3fv3igoKEB4eDgSExMxYMAAeHl54ebNm7w6v/zy\nSwQFBSExMREuLi4YNGgQcnNzAQC5ubno0aMHXrx4gUOHDiE5ORkhISGcM3L16lUMHjwYYrEYV69e\nxebNm3H48GF8+umnvDouXryIkydP4vjx4zh69CgSExO5tovFYrRv3x6///47r0xoaCjGjh0LFRUV\nAMCpU6fg4uIi9fzu37+PLVu2YP/+/Th27BhOnz4NX19fhIWFYe/evZwsJCSkXvXVh6pTs6NHj4au\nri7OnTuH69evY9WqVdDS0gIAXLp0CXJyclizZg1yc3Nxv9Kp1iKRCHZ2dtxzrI05c+bgm2++wc2b\nNzFw4EAEBQVh5cqV+O6773Dz5k2sWbMGv/76K4KDgwFInM+0tDRcvHiR01FUVITdu3fDz88PAHDj\nxg2IxWKECb1UAAAgAElEQVT07NkTCQkJiImJgZycHLy8vHixf3///Td+/vlnbN26FSkpKTAyMqq1\nzXV5Z4Gan3GHDh2goaFRp/vCYDDeH0TAtWvAunXA0aOSKdgKdHWBUaOASZMAk+YcKUUtlNqa9qZm\nr1sXRYsWEbm78z8DBkjk9f14e0dJ6Vq0iGj9+qh6t+vly5ekrKxMK1eulEpLT08ngUBAx44d48m7\ndu1KkyZNIiKi27dvk0AgoG+//ZaXp0ePHjR+/HgiInr9+jXp6urShg0buPRRo0bR0KFDpeoMDQ2l\nVq1aUV5eHie7fv06CQQCWrp0KRERxcTEkLKyMj1//pxXduLEiZzOTZs2kbGxMZWWlvLy9O7dm2bN\nmsXpEQgE9Pvvv3PppaWlZGJiQoGBgUREtHDhQjI0NKRXr15J2UpENG7cOHJxceHJDhw4QEKhkO7c\nuUNERH5+fiQSiejFixdcnuPHj5NAIKCMjAwiIlq1ahWZmJhQeXk5ERGlpKSQQCCgxMREIiLKz88n\ngUBAhw8f5tW1aNEikpeXpydPnnCyGTNmkJycHD1+/JiTzZw5kxwdHbnrN9X3JsRiMU2dOpW79vPz\nI09PT+5aQ0ODwsLCaiwvLy9PmzdvrjZt8ODBNGbMmBrLVjy3bdu2cbKCggJSVVWliIgIXt7NmzeT\npqYmd929e3eaMWMGd71nzx5SUVHh3iU/Pz8aNWoUT8fr169JVVWV9u/fT0SSey4UCiknJ4eXr7Y2\n1+WdrekZV9CpUydasGBBtWktuOtlMJok5eVE6elEv/wi/Td65Uqiy5eJysoa10ZZ9Qts8UQ1eHpa\nICwsCmKxBycrKoqCv78lrK3rry81VaJPSYmvz8PDst66kpOTUVRUhL59+0ql3bhxAwDg5ubGk7u5\nueHcuXM82UcffcS77tmzJ6KiogAASkpK8Pf3R2hoKCZPnownT55g//79+PPPP6ut09bWlhevZWdn\nx7uOj49HcXExjKqcsVJcXAwrKysuT25uLjSrBDIUFRVBTU2tRtvl5OTg7OzMtT0hIQE9evSocRTr\nxo0b8PDw4Mnc3NxARLhx4wbatm0LALC1tYWo0qnNPXr04Mqbm5tjwoQJWLBgASIiItC/f39s2LAB\njo6OcHBwAAA8f/4cAHg6KjAyMoK2tjZ3ra+vDwMDA+jo6PBklUc9/fz8aq3vbZk7dy6mTJmCsLAw\niMViDB48GF26dKlTWZFIxE1T14azszP3/+TkZBQWFsLX15c3clhWVoaioiI8efIEOjo68PPzQ2Bg\nINasWQM5OTls2bIFQ4YMgbq6OgDJe5ORkSF1n4uKinDr1i3uWl9fH8bGxnVuc13e2dqeMQCoq6sj\nLy/vjfeFwWC8W+7dk+xFd/s2X66iAri6Ak5OgIJC49j2LmCOXTVYW5vA3x+IiopGcbEQiorl8PCw\nbPAqVlnrqy9Uh8D2qnmmTZuGlStX4tq1a4iKioKenh68vb0bpL+8vBwaGhq4dOmSVJriP2vGy8vL\n0aFDB+zfv18qj6qqar1sf5M9DbkfVdHR0cGIESMQGhoKDw8PbNmyhZs6BcA5qPn5+VJlFar0IAKB\noFpZ5QB/bW3tWut7WxYuXIixY8ciPDwc0dHRCAkJqfOK3ufPn3NTmLVR2UGvaNvevXs5R6kyFfr+\n85//YNasWTh8+DB69OiBiIgIHDhwgMtHRJgwYQLmzZsnpaOy81z1xwFQe5vr8s7W9owByX2p+kOF\n8eHB9rFrPB4/BqKjgX9+93MoKEi2LunZE1BWbhzb3iUt2rF7myPFrK1NZOp4yUpfxYKJiIgI2Nvb\n89Ls7OwAACdPnuQ5YadOnUK3bt14ec+dO4f+/ftz12fPnuXKA4CFhQX69OmD0NBQxMTEYNKkSdVu\nX2JnZ4fQ0FA8f/6cG6VLTk7mRjMAwNHREXl5eSgsLOTVURknJyds3boVIpEIurq6td6Dc+fOcUH1\npaWluHjxIhdz5ejoiNDQULx69apah9DOzg6nTp3iyU6ePAmBQMCzLSUlBfn5+dxozNmzZwFI7n8F\n06ZNQ+/evfHLL7/g9evXGD16NJempqYGQ0NDZGfLbuVzbfU1hKrP08zMDAEBAQgICMDy5cuxYsUK\nzrFTVFTkFjVUJTs7m1u8U1fs7OygrKyMjIwM3ntYFS0tLQwaNAhbt25FdnY2tLW10a9fPy7d0dER\nSUlJMDc3r1f9FdTU5rq8s7U9YyJCTk5OtU4rg8F4t7x4IVnpeuWK5HzXCoRCoGtXyT50NQy0Nwqy\nPlKsxQZ61Na05t7shQsXUqtWrWj9+vWUmppKiYmJtGzZMiIi+vjjj8nU1JQiIiIoJSWFvvjiC1JS\nUqLU1FQi+jfGztjYmHbs2EGpqakUGBhIQqGQrly5wqtnz549pKioSPLy8nT37l0iIvrrr7/I2tqa\n7t27R0REr169IkNDQxo4cCAlJSXRuXPnyNnZmVRVVbkYOyIiLy8vsrKyov3791NGRgZdunSJfvzx\nRwoNDSUiSVyUvb09OTk50fHjx+n27dt0/vx5CgkJ4WKlKmK1rK2t6ejRo3Tjxg2aMmUKqamp0f37\n94mI6P79+6Snp0eenp505swZyszMpEOHDnFxh1evXiV5eXmaPXs2paSk0LFjx6ht27Y0YcIEzlY/\nPz9SV1enoUOH0vXr1+nkyZPUvn37amMM7e3tSUlJiT755BOptNGjR/P0EknivSwtLXmyxYsXk6mp\nKU+2bNkyMjY2rld9teHu7k5TpkzhtbEixi4/P5+mT59O0dHRlJmZSZcvXyZ3d3dyc3Pj8tvZ2dG4\ncePo77//pkePHnHyFy9ekJycHJ08eZKTrV27lmxsbLjriudW8c5Ubre6ujqtX7+ebt68SdevX6ed\nO3fS//3f//HyHTx4kBQVFcnW1pbmzp3LS0tJSSGRSERjx46lixcvUmZmJkVHR9PMmTMpMzOTiKq/\n5y9fvnxjm9/0zhJV/4yJiJKTk0kgEFB2drZUGlHz74MYjKZIYSHRiRNES5ZIx9Ht3k1UKYy5SSKr\nfqHF9i4t2bEjIlqzZg1ZW1uToqIi6evr08cff0xEkj+006ZNI11dXVJSUiInJyc6ceIEV67Csdu2\nbRuJxWJSVlYmc3Nz2rlzp1QdJSUlpKenRwMHDuRkmzZtIqFQyPuDdeXKFfroo49ISUmJLC0tadeu\nXWRqaspz7AoLC2nevHlkZmZGioqKZGBgQN7e3hQTE8PlefLkCQUEBJCRkREpKiqSkZER+fr6cgsE\nKhyEQ4cOUbdu3UhJSYns7OwoMjKSZ3daWhoNGzaMNDQ0SFVVlTp37sxbUHL06FGuvK6uLk2fPp23\n2KLC6VmxYgUZGhqSqqoqjRgxgp4+fSp1j/73v/+RQCCgS5cuSaUdOXKEdHR0qKSkhJMFBQVR+/bt\nefmWLFlCZmZmPNny5cupbdu29aqvNqounvD39ycvLy8ikjjVY8aMITMzM1JWViY9PT0aNWoU58wT\nEYWHh1OHDh1IUVGRhEIhJ9+2bZuUUxoUFMTLExMTQ0KhUMqxIyLasGEDde7cmZSVlUlLS4u6d+9O\nv/zyCy9PxXsoFArp6tWrUjquXbtGQ4YMIS0tLVJRUSFLS0uaNm0aPXv2jLOn6j2vS5vr8s5W94yJ\nJM/U1dVVytYKWkIfxGA0FYqLieLiiJYvl3bowsKIKn2tmzSy6hcE/yhrcdR2mO6HfAB3VlYWzM3N\nERcXxy0IqIknT56gbdu2+OOPPzBo0KD3ZGHNxMbGok+fPrh79y7atGnzzurx9/fHvXv3ePv/1cTX\nX3+NqKgoJCQkSKUREWxtbREUFIT//Oc/MrGttvoaAw8PD3h7e2Pu3LmNbUqjUN0zLi8vh42NDUJC\nQmqcov6Q+6APDRZj9+4oLwcSEyV7w/6z7SeHoSHg6QmYm0tOj2gOyKpfaNExdoyGUVpaisePHyMo\nKAjGxsZNwqlrajx//hxpaWkIDQ3F2rVrq80jEAjw3XffYeHChW/t2NWlvvdNXFwcMjMz8cUXXzS2\nKY1Gdc94x44d3OIaBoMhe4iAmzclCyMePeKnaWsDffoAdnbNx6GTNWzE7gMjKysLFhYWOH36dI0j\ndhUjY+bm5ti6davU1iiNRWxsLDw8PJCTk/NOR+wmTpyIe/fu4fjx4zXmEYvFuHjxIkaPHs2dovAu\nqa2+kJAQLFu2rNpyAoGAO8GC0XT4kPsgBuNtyMqSbF1S5SAetGolWRTRtStQ6XCcZoWs+gXm2DEY\nzZxnz57Vuo9cQ1eMMt4drA9iMOrHgwcShy49nS9XUpJsW9K9O/DPTkTNFjYVWwfeZrsTBqO5oKWl\nVad95BgMxvuHxdi9Hc+eSc5yvXZNMgVbgZwc4Ows2WD4DVudNnlkvd0JG7FjMBiM9wzrgz4cmGPX\nMAoKgFOngEuXgMpbaAoEQKdOQO/eQEvb/5tNxb4B5tgxGIymCuuDGIzqKSoCzp0Dzp4Fiov5aVZW\ngIcHoK/fOLa9a9hULIPBYDAYjBZBWZlkdO7UKcloXWXatgW8vIB27RrHtuYGc+wYDAaDwXhHsKnY\n2iECrl+XbF1SdQ2Yrq5kLzorqw9365KGwBw7BoPBYDAY7xUiICNDstI1N5efpqEhiaHr1Elyviuj\nfrAYOwaDwXjPsD6I8SFz967EocvK4stVVCSrXJ2dAfkPcNhJVv0C84WbGf7+/vDy8gIg2c6lffv2\nMtG7Zs0aDBgwQCa6mhuxsbEQCoX4+++/efLZs2dj+vTpjWTVvwiFQmzfvp27NjMzQ0hIyFvpXLJk\nidRpGGVlZejQoQOOHTv2VroZDAajOh4/Bv74A9iwge/UKShIHLqZM4EePT5Mp06WsNtXA6m3UhGZ\nEIkSKoGCQAGe3TxhbWndJPQJ/gk2+Oqrr+p1nJOlpSXGjx+PRYsW8eQvXrzA4sWLcfTo0QbZ0xK5\nffs2QkNDkZaW1timAPj3mQPApUuXoFrHjZvi4uLg5uaGrKwstKsUeTxr1iy0a9cOly5dgqOjIwBA\nTk4OCxYswP/93//B29tbtg1gMD5QWIyd5BzX2FjJua7l5f/KhULJSRHu7oBI1GjmtTha9IhdUFBQ\ngzb9S72VirCYMDzSf4Q8gzw80n+EsJgwpN5KbZAdstZXMVSrpqYGbW3tOpcT1BB9unXrVhgZGcHZ\n2bnGsiUlJfUzUsYUV133/o756aef4OHh8U6PLmsoOjo6UFFRqVeZqsP7rVq1wogRI6TOnR0+fDiy\ns7MRExPz1nYyGIwPm8JCyZTrjz8Cly/znTo7O2DGDGDgQObUxcbGIigoSGb6Wrxj15BfSpEJkVBq\nr4TYrFjuc07hHL7c9SWCYoPq/Zm9azbOKZzj6VNqr4Soy1ENaleFg1Z1Kvbu3bsYPnw4dHV1oaKi\nAgsLC6xYsQKA5KzRjIwMBAcHQygUQigU4s6dOwCAbdu2YdiwYbw6KqZ8165dC1NTUygrK6OoqAgP\nHjyAv78/9PT0oK6ujl69euH06dMAgPLycrRr107q3NKioiJoaWnh999/52Rr166FjY0NVFRUYGVl\nhZCQEJRV2oXS1NQUgYGBmD59Olq3bg13d3cAwIYNG9ChQweoqKhAR0cH7u7uuHfvHlcuISEBffv2\nhUgkgp6eHoYPH861s3LdxsbGUFNTQ//+/aXSAWD79u1S90QsFmPKlClYuHAh9PT0oKWlhW+//RZE\nhEWLFsHAwAB6enpYuHAhVyYoKAg2NjZS+idNmgRPT08peV0wNTXF0qVLuesDBw6gS5cuUFNTg5aW\nFlxcXJCYmIisrCy4ubkBkEzfCoVC9OnThys3bNgw7N27l+c0q6iooH///ti2bVuDbGMwGHw+xNG6\nkhLgzBlgzRogLg4oLf03zdwc+OQTYORIQEen8WxsSojFYubYvWtKqPrRqTKUVSt/E+Uor1ZeXC7b\nUajp06cjPz8fUVFRSE1NxcaNG2FsbAwA2LdvH0xNTTF37lzk5uYiNzcXxsbGKCgoQEJCAlxcXKT0\nXbx4EbGxsTh06BCuXr2K0tJS9O7dGwUFBQgPD0diYiIGDBgALy8v3Lx5E0KhEOPHj8fWrVt5eg4c\nOICioiKMHDkSgMTZWblyJb777jvcvHkTa9aswa+//org4GBeuR9//BEGBgY4f/48Nm3ahISEBAQE\nBGDBggVIS0vDyZMn4efnx+W/ceMGxGIxevbsiYSEBMTExEBOTg5eXl4oKiribPnyyy8xd+5cJCUl\n4eOPP8ZXX33FG81MS0tDbm5utfdk7969KCsrw9mzZ7Fq1SosWbIE3t7eKCoqQlxcHFasWIGQkBCE\nh4cDAKZOnYqMjAycOnWK05Gfn489e/Zg2rRp9Xq+FQgEAs7e3NxcjBw5EmPHjsWNGzdw/vx5zJ49\nG/Ly8mjXrh0OHDgAAIiPj0dubi7++usvTo+LiwsKCwtx/vx5nn4XFxdER0c3yDYGg/HhUl4uGZlb\nuxY4cQJ4/frfNENDYPx4YMIEoAlOhLQoWIxdNSgIFKqVy0GuQfqENfjPikLZnlh8584dDBs2DJ06\ndQIAXkyVlpYW5OTk0KpVK+jp6XHy27dvo7S0lJe3Ajk5OWzdupWL5woLC0N+fj527doFOTnJvZg/\nfz6ioqLw66+/YvXq1Rg/fjyWLVvGi93asmULhg0bBpFIhFevXuGHH37Avn370LdvXwCAiYkJFi9e\njJkzZ+K///0vV7+zszO+/fZb7nrfvn1QU1PDkCFDIBKJ0LZtW9jb23Pp33//PQYOHMiLIdy6dSu0\ntbURERGBwYMH44cffsCoUaMwa9YsAJK4w5SUFKxcuZIrUxFXV909MTc350YkLS0tsXLlSty/f59z\n5CwtLbFq1SpERUWhf//+MDIywoABAxAaGsqNnu3YsQOqqqpSI4IN4f79+ygtLcXIkSNhYmICALC2\n/jd2s+IMWV1dXd5zBwBtbW2IRCKkpaVxtgGSEcHs7GyUlpZCnkUxMxhvxYcQY0cE3LwJREVJFkhU\nRlsb6NNHMvXK9qJ7P7Beuxo8u3kiLCYM4vZiTlaUXgT/Uf4NWvCQaiyJsVNqr8TT59HbQxbmcsya\nNQvTpk3DsWPHIBaL4ePjA1dX11rLPH/+HAAgqibIoUOHDrwg/YpRH80qB/QVFRVx+WxsbODs7Iyt\nW7fC0dERDx8+xPHjx3Ho0CEAQHJyMgoLC+Hr68sbJSsrK0NRURGePHkCHR0dCAQCqZi/vn37wtzc\nHGZmZvDy8kKfPn3g6+sLnX/G8+Pj45GRkSHVlqKiIqSnpwMAUlJSMHbsWF56z549eY5dxT1RU1Pj\n5RMIBHBwcODJDAwMYGhoKCV79OgRdz1t2jSMGDEC69atg4aGBkJDQ+Hn5ycTp8nBwQH9+vWDvb09\nvLy8IBaL4evry43Uvgl1dXXk5eVJyQAgLy8PrVu3fmsbGQxGyyUrSxJHd/cuX96qlWRRRNeugFzD\nxkQYDYQ5dtVgbWkNf/gj6nIUisuLoShUhEdvjwavYpW1vprw9/dH//79ER4ejpiYGHh7e2PYsGFS\nU6OVqXDS8vPzpdKqrrwsLy9Hhw4dsH///lrzTpgwAcHBwVi5ciV27NgBXV1dbnSu/J/o2b1798LK\nykpKT8UIEyDtWKmpqeHSpUs4c+YMIiMj8csvv+Drr79GVFQUunbtCiLChAkTMG/ePCm9OvUI5qi4\nJwUFBVI2KCjwR3MFAoGUDPi3nQDQv39/6OnpYcuWLXB1dcXly5exc+fOOttTG0KhEMeOHUN8fDwi\nIyPx559/Yt68edizZw98fHzeWP758+dSjnqFY1tVzmAw6k9LHa3LzZU4dLdu8eVKSkDPnkD37oCi\nbCelGHWEOXY1YG1pLVPHS9b6asLAwAD+/v7w9/eHt7c3xowZg59//hmtWrWCoqIib4ECIJl2k5eX\nR3Z2Nuzs7GrV7eTkhK1bt0IkEkFXV7fGfKNGjcKXX36J8PBwbNmyBWPHjuVG5+zs7KCsrIyMjAz0\n79+/3u0TCoVwdXWFq6srgoODYWtri507d6Jr165wdHREUlISzM3Nayxva2uLM2fOICAggJOdOXOG\nl6diQUp2djZsbW3rbWPV1cdCoRBTp05FaGgobt68CXd3d5ntP1iBk5MTnJyc8M0338Db2xubNm2C\nj48PFP/pWas+dwB48uQJXr58KeVgZ2dnc+8Fg8FgVObZMyAmBrh2TTIFW4GcnGRjYVdXoI67MTHe\nEaznbobUtDP1Z599Bh8fH1hZWeH169f466+/0K5dO7Rq1QqAZGVkXFwccnJyuFWlampqcHR0xIUL\nF964QfHYsWOxevVq+Pj4YOnSpWjfvj0ePHiA6Oho2NraYsiQIQAksVs+Pj4IDAxEUlISb8SwVatW\nmD9/PubPnw+BQAAPDw+Ulpbi2rVrSExMxPLly2ts48GDB5GZmQlXV1fo6uoiISEBOTk5nPM1f/58\nODs7Y9y4cZg5cyZat26NrKwsHDhwADNnzoSZmRnmzJmDkSNHwtnZGd7e3oiLi5NaAWplZQUDAwNc\nuHCB59gRkZRddZVNnjwZwcHBSEtLw6ZNm2q9z2+isu6zZ88iKioK/fr1g4GBAdLT03H16lVMmTIF\ngCR+USgU4siRI/j444+hpKQEDQ0NAMCFCxegrKyM7t278/SfP3++xY4yMBjvm5YSY1dQAJw6BVy6\nBFT+nSgQAA4OgFgMsEH+pgFbFdvMqLwisvL/K5g1axY6duwId3d3FBYW8k4RCA4ORl5eHqytraGv\nr4+cnBwAwLhx47Bv374a66lASUkJJ0+ehKOjIyZOnAhra2sMHz4cly5dgqmpKS+vn58fkpKS0KVL\nF6mRwIULF2LVqlUIDQ1F586d4erqijVr1sDMzIxXf1W0tLRw6NAheHt7w9raGvPmzUNgYCAmTpwI\nQBLfd/bsWbx8+RL9+vWDnZ0dPvnkE7x+/ZqbVhw6dChWrlyJ77//Hg4ODti5cye+++47qfrqek/q\nKjMwMICPjw9EIhFGjBgh1bb6UFm3pqYmzp8/jyFDhsDKygqTJ0/GuHHjEBgYCADQ19fHsmXLsHz5\ncrRp04a3YGPfvn0YMWIEN6oHAIWFhYiIiMC4cePeykYGg9EyKCqSbC68Zg1w4QLfqbO2BgICgKFD\nmVPXlGBnxTKQn58PMzMzHDlypNotPj5EsrKyYG9vj9TUVBgZGclEp7OzM1xdXXkLNRqL/Px8mJiY\n4Pjx49zqZUCyiviHH37A1atXG9G6lg/rgxhNndJSICFBMkpXUMBPa9cO8PSU/MuQHbLqF5hjxwAg\n2TMuIiICR44caWxTmgxffvklioqKsH79+rfS8/jxYxw+fBhTp05Fenq61OhmY7B06VJcu3YNu3bt\n4mRlZWWwt7fH6tWrGxT/yKg7rA9iNFWIJPFz0dFAlQXz0NMDPDwAKyu2dcm7gDl2b4A5doymglAo\nhLa2NpYsWYJPP/2Ul1YR51cdbm5uzNFuobA+6MOhucTYEUlWuEZFSVa8VkZDA+jdG+jUSXK+K+Pd\nIKt+gS2eYDDeMZW3PqnKxo0b8bry9uyVqO95sAwGg9EQ7t6VbF2SlcWXq6gAbm6AkxPAFsk3H1r0\no6o4K7Y5/FpifJi0YWfrMBgtmqb89+fxY8kIXUoKX66gAHz0EdCjB6Cs3Di2fUjExsYiNjZWZvrY\nVCyDwWC8Z1gfxGhMXryQrHS9coW/F51QCHTrJhmlq+YwIsY7hk3FMhgMBoPRxGlKMXaFhUBcnGTb\nktJSfpq9veRMV23txrGNITuYY8dgMBgMRgumpETizMXFAVVDei0sJCtdWVRIy4FNxTIYDMZ7hvVB\njPdBeblkujU2Fqh6HHibNpK96Go5gZHxnmFTsQwGg8FgMKQgkiyIiI6WLJCojLa2ZITO1pbtRddS\nYTvSMN6IUCjE9u3buWszMzOEhIS8lU5/f38IhUIIhUL89NNPb2tinXF0dOTqPXPmzHurl8FgfJjI\ncrVjXbh9G9iwAdi9m+/UtWoFDBwIzJgB2Nkxp64lw0bsaiA7NRUZkZEQlpSgXEEBFp6eMLG2bjL6\n3jeVzye9dOkSVFVV61QuLi4Obm5uyMrKQrsq58+4ublh9+7dEMlo+dXSpUsRHh6OpKQkvHz5Enfv\n3pXaTuTEiRPIyMiAs7NztefRMhgMRnMkN1eyF92tW3y5khLQqxfg4gJUOhaa0YJhjl01ZKem4lZY\nGDyUlDhZVFgY4O/fIGdM1voaGx0dnXqXqS5uQEFBAXp6erIwCQBQXFyMoUOHYtCgQZg3b161ebS0\ntNC6dWuZ1clgMBi18a5XxD57JplyvXaNL5eXB5ydJU5dHX+HM1oIzLGrhozISIkTVmkI3QNA9NWr\nMHFyqr++ixfh8eoVT+YhFiM6Kqrejp1YLIalpSUMDAzw22+/oaSkBJ9//jmCg4MRFBSEX3/9FeXl\n5fjkk0+wZMkSBAUFYdeuXbh58yZPz6RJk3Dnzh1ERkbWuz2mpqaYOnUqFixYAAA4cOAAgoKCkJaW\nBkVFRVhZWeHXX3+FpqYm3NzcAEimbyvsj46OrneddSE4OBjA+5/6YDAYjPfNy5fAqVNAQgJQVvav\nXCAAOncGxGLJUWCMDw8WY1cNwpKS6uWVvz310VfDkVLC4uIG6du7dy/Kyspw9uxZrFq1CkuWLIG3\ntzeKiooQFxeHFStWICQkBOHh4Zg6dSoyMjJw6tQprnx+fj727NmDadOmNah+gUDATWPm5uZi5MiR\nGDt2LG7cuIHz589j9uzZkJeXR7t27XDgwAEAQHx8PHJzc/HXX3/VqjsoKAjCKocRisVi9O7du0G2\nMhgMRmMi6x+aRUVATAzw44/AxYt8p87aGggIAIYMYU7dhwwbsauGcgWF6uVycg3TV8OpyeUNDHgw\nNzfHsmXLAACWlpZYuXIl7t+/j/DwcE62atUqREdHo3///hgwYABCQ0O50bMdO3ZAVVUVw4YNa1D9\nlYMcCbkAACAASURBVLl//z5KS0sxcuRImJiYAACsK41CamlpAQB0dXXrNO2qq6sLGxsbnszExITF\nwzEYjA+a0lLg0iXJKF2VCSC0ayfZuqRKGDPjA4U5dtVg4emJqLAweFSKjYgqKoKlv7/kJ1F99aWm\nSvRVjrErKoKlh0e9dQkEAjg4OPBkBgYGMDQ0lJI9fPgQADBt2jSMGDEC69atg4aGBkJDQ+Hn5wd5\nGZzq7ODggH79+sHe3h5eXl4Qi8Xw9fWFsbFxg/TNmDEDM2bM4Mk2b97Mu7azs8OdO3cASKaFr1UN\nLmEwGIwmwtvG2JWXS+LnYmKAvDx+mp6exKFr356tcmX8C3PsqsHE2hrw90d0VBSExcUoV1SEpYdH\ngxc6yFqfQpURRYFAICUDgPJ/poD79+8PPT09bNmyBa6urrh8+TJ27tzZoLqrIhQKcezYMcTHxyMy\nMhJ//vkn5s2bhz179sDHx0cmdVQlPDwcJf9Ml1fXbgaDwWjuEElWuEZGAg8e8NM0NCTHf3XsKDnf\nlcGoDHPsasDE2lqmK1Zlre9NVJ66FAqFmDp1KkJDQ3Hz5k24u7ujffv2Mq3PyckJTk5O+Oabb+Dt\n7Y1NmzbBx8cHiv9MN5c1MD6xOtq2bSszXQwGg/EuachZsXfvAidOANnZfLmqKuDmBjg6Sla9MhjV\nwXz9ZgYRSW0dUhfZ5MmTcfPmTWzcuBGffPLJW9tQwdmzZ7F48WJcvHgRd+7cQVRUFK5evQo7OzsA\nkvg4oVCII0eO4OHDh3jx4kWtutetW4cOHTrwZBMmTICfn98b7bpz5w4SExNx65+NnJKTk5GYmIhn\nz57Vt4kMBoPx3nn0CNi1S7LBcGWnTkFB4tB98QXQvTtz6hi10ywduxcvXsDZ2RkikQg3btxobHPe\nK5VXpNZHZmBgAB8fH4hEIowYMeKtbahAU1MT58+fx5AhQ2BlZYXJkydj3LhxCAwMBADo6+tj2bJl\nWL58Odq0aYOhQ4dWq6eCJ0+eIC0tjSfLyclBTk7OG+369ttv0bVrV3zyyScQCATo168funXrhkOH\nDjW0qQwGg/FW1GW07vlz4MAB4KefgMo7UwmFgJMTMHOmZOpVWfnd2cloOQioGZ5EXVpairy8PHz1\n1VeYO3cuNzpUmdoO0/1QD+B2dnaGq6srVq5c2dimwN/fH/fu3cOJEyfee91ZWVkwNzdHXFwcevTo\n8d7rZzA+1D6IwaewEDh9WrJtSWkpP83eXuLMaWs3jm2M94+s+oVmOaArLy/PTg+oB48fP8bhw4dx\n5coV7N69u7HNASB5gWNjYyESibB69WpMmTLlvdTr5uaGy5cvs+1TGAzGe6G6GLuSEuDCBSAuDnj9\nmp/fwkKy0rXKRgcMRp1plo4do37o6elBW1sba9euhampKS/N29sbcXFx1ZZzc3PDkSNH3olN33//\nPTdd+z6d9J07d6KoqAgAYGRk9N7qZTAYjPJy4MoVyaFG+fn8tDZtAC8v4J9DehiMBtOojt26desQ\nFhaG69evY/To0di0aROX9vTpU0yePBknTpxA69atsWzZMowePVpKBxt5eTPlNZx8AQAbN27E66o/\nGf9BRUXlXZkEXV1d6OrqvjP9NcGcOQaD8T4Ri8UgAlJSgKgo4MkTfrqOjmTK1daW7UXHkA2N6tgZ\nGRkhMDAQERERKCws5KXNmDEDysrKePjwIa5cuQIfHx84ODjA1taWl4/Fqbwdbdq0aWwTGAwGo8WR\nmpqNyMgM5OYKcetWOXR0LNC6tQmXLhIB7u5Aly5AAw81YjCqpUksnggMDMTdu3e5EbuCggJoa2sj\nOTkZlpaWAAA/Pz+0adOGO0prwIABSEpKwv+zd+dxUVf7/8BfA8i+CoIiArK4sIMLqKgICiqpaYtm\nmmbdrKvV7d6WR5aCdX/lbb+VPbqpubZZXytLEhMYF8QNEBUV2fd932GG8/vjEx/4wAAzMjMM+H4+\nHj4ezPnMfD5nhsPHM+9zzvs4ODhg8+bNvdJh0OIJQoimonvQyJaWlovduzOQnx+C7GwxzM2DIJHE\nwMfHBXZ2DggMBPz9uTQmhHQaUYsner6Ru3fvQkdHh+/UAdzWVd03U46KihrwvBs3buTnlJmbm8PH\nx2fQ27sQQogydJ9U33lvo8fD/3FlJfDSS/+HoiI/mJsDAFBTI4ZIpA0gEy++6IBLl8SIj9eM+tLj\noXvc+XNOTg6USSMjdufOncOjjz6K4uJi/jl79uzBt99+i7i4OLnO2V/Pd/To0ZS0lhAyZCwsLFBV\nVTXU1SBKVFcHnDnDLY64cEGMlpYg/tjYsYCjIzB2rBj/+EeQ7BOQ+96IjtgZGxv32qGgtrYWJiYm\nSrke3VAJIYQoQ3Mzl7bk0qWuXHRaWtyCNSsrbpWrkRFXrqvb90I2QpRFI3ae6LmyddKkSZBIJPzW\nUACQkpICDw8Phc4bGRkpCHkS0hdqJ0Re1FYIALS1ccmF//tfID5emGB43jxneHjEwMMDKC8XAwBa\nW2MQEuI8NJUlGk0sFiMyMlJp5xvSiJ1UKkV7ezskEgmkUilaW1uho6MDIyMjrFq1Cjt27MDevXuR\nlJSE3377DQkJCQqdX5kfFCGEECKVAomJwNmzQEOD8JitLRASAjg5OeDuXSAmJhYVFddhbd2BkBAX\nTJ7sIPuk5L4WFBSEoKAg7Ny5UynnG9I5dpGRkXjrrbd6le3YsQPV1dXYtGkTn8du165dWLNmjdzn\nplVnhBBClIUx4MYNIC4O6DlF29KS69BNnUq56Mi9U1a/ZUiHYiMjI9HR0SH4t2PHDgDc5OKff/4Z\nDQ0NyMnJUahTp0nEYjFmzJgBACgqKkJwcHC/z8/NzcWePXsEZY6Ojpg6dSqOHDmi8PXb2tqwePFi\nmQmBc3Nz4ePjAz09Pdy6davXa3/77Te8+uqrCl2vs66+vr7w9fUV7AV74sQJTJs2DV5eXggKCupz\nJdD+/fvh7e0NX19feHh44P333+ePbdiwgT+3r68vtLW18fvvvytUx19//RVXrlxR6DUANzdz6dKl\nmDJlCry8vPDQQw+hoqJCrvfX3+fSnVQqxZYtW+Di4gJXV1fs27ePP6aM915ZWYnZs2fD19f3nvYM\nrq2txXvvvafw6/py4MABpKenCx4/8sgjSju/JtLS0kJTU9NQV0MpXnnlFfzwww8qv05QUBC/C05E\nRMSAWyP2bFcnTpzAli1b7unajAF37wJffgkcOybs1JmaAsuXA1u2qC/BcPfPfPfu3YL7IyEAADZC\nAWAREREsLi5uSOsRFxfHpk+fPqjnOzo6stTU1Hu6vkQiYTExMezatWvMyspK5nMGc355z1VVVcWs\nrKxYeno6Y4yxI0eOsMWLF8s8R11dHf9zfX09s7e3Z0lJSb2el5KSwiwtLVlbW5tCddywYQP7/PPP\nBWXytJOqqip25swZ/vErr7zCnnrqKf5Yf+9P3s/44MGDLCwsjDHGWHl5ObOzs2M5OTm9nnev7/37\n779n4eHhCr2mu+zs7D7b0UAkEkmvsvnz57Pff/+df3zgwAH28MMP33P91GGw9xSRSMQaGxuVU5kh\nVFpayqZOnaqWawUFBbETJ07I/fye7Yoxxry8vFh+fv6Ar21vb+d/zs1lbN8+xiIihP927WIsPp4x\nef78ZLUXWX8L8uj5mbe0tDAnJyfW3Nx8T+cjmiEuLo5FREQwZXXJNGLxhKpERkbyeWMGoqWlhXfe\neQczZ86Ek5MTTp8+jVdffRW+vr7w9PTEnTt3AAAPPPAAfvrpJ/51x44dQ1hYmFzXyMnJ4fdFbWpq\nwiOPPAJ3d3f4+PjwEcktW7bg1q1b8PX1xaOPPqrAu5VNW1sbwcHBMDMzU/i13aMnaWlpmDVrFnx8\nfODp6dlvtIfJCCVnZGTAxsaGz024ZMkSREdHy1yh3H31c8Nfk1jMOxNCdbN3716sW7cOo0aNQkdH\nB8LCwvDpp58CAG7dugVHR0cUFRUJXhMdHY3ffvsNu3btgq+vLx8F/e677+Dp6QlPT09s2rQJjY2N\nva5nYWGBefPm8Y/9/f2Rm5sr9/uT9bn0dPToUTzzzDMAuD10H3zwQfz4449Kee9xcXF49dVXER8f\nD19fX5w/fx7fffcdAgIC4OfnBz8/P8TGxgLgtqH7+9//jqlTp8LHxwdz584FwLXPmpoa+Pr6IjAw\nEABQXFyMRx55BP7+/vDy8uKTiANcpPL111+Hv78/nn32WUF99u/fj8TERLzwwgvw9fVFTEwMAKCu\nrg5r1qyBh4cHAgMDUVpaCgC4ceMG5s2bh2nTpsHd3R3//e9/+XNt3LgRzz33HEJCQjBp0qReCcu7\n+/333zFjxgz4+PjAz88PN27cAACcPHkSfn5+8Pb2xsKFC5GZmQmAi7p7e3tjw4YN8PDwwHPPPYfb\nt28DkO9+cOzYMT5a+/bbbwuOXbp0CcHBwZg+fTqmT58uyM8ZFRWFwMBATJ8+HbNnz8alS5dk1sff\n35+vDwB8/fXX8PHxgY+PD2bOnIny8nIAwKFDh+Dl5QVvb2+sWrWKLz9w4AAWLVqEFStWwN3dHSEh\nIXzb8fT0xNWrV/lzf/TRR9i8eTMA4PDhw3jwwQf5Y5GRkVizZg3Cw8Ph6uqKRx99FFevXsWCBQvg\n4uLCR/+vXLkCT09Pwefg7e2Nixcv9vk7627jxo3YvXs3AC767uXlxd+nz5w506tddbbpVatW4dCh\nQzLPGRQUhJdeegmzZs3Cgw8+iJIS4LHH/oPZs/3x5pvT8N13y9HQUAqgCZ98Mgbr1lVi9mwuwfDL\nL7/MTynq6/fZee9/5ZVXMG3aNOzdu1dm3YH+/556fuZ6enqYN28ejh07JtdnRzRTUFCQctcEKKV7\nqIEUfWsikYh98cUXjDHGfvzxR2ZoaMh/Q3zvvffYunXrGGOMnTx5ki1YsIB/XXBwMDt+/Hif5+0e\ngese7Th27BgfmWGMsZqaGsYYY2KxeMCI3ZUrV9jSpUv5xzt27GBffvllv++vv0hLX9Gk/fv389GT\nF154gb377rv8serq6j7P5enpyTw9Pdnf//53/n3V1NQwS0tLduXKFcYYY59++ikTiUQsOTlZ5nmO\nHz/O3N3dmZ6eHvv44497HW9tbWVWVlYsJSWFLysrK2MTJ05kZ8+eZZ6eniwqKkrmuTdu3Mh2797N\nP46KimIeHh6svr6eMcbYE088wV577TWZr+0klUpZSEgI++yzz+R6f319Lj15enqyq1ev8o/fe+89\n9sILLyjtvfeMiFVWVvI/37lzh9nZ2THGGEtKShJEBjrrm5OT06sdLVy4kJ09e5avW2BgIPvzzz/5\n971lyxaZdWGsdyRm//79zMLCghUUFDDGGPvb3/7G3njjDcYYF71tbW3lf3Zzc2N37txhjHFR2Llz\n57LW1lbW1tbG3N3d+Tp0l5aWxsaOHcsyMjIYY4y1tbWx+vp6VlpaysaMGcNu377NGGNs3759zN/f\nnzHG/Q2LRCL+PR48eJD/Gx3oflBSUsIsLS3Z3bt3GWPc77MzYlddXc18fX1ZcXExY4yxoqIiZmdn\nx2pra1lGRgabNWsWH72+efMms7e3H7A+cXFxzMXFhZWWljLGGGtsbGQtLS3sxo0bzNbWlpWUlDDG\nGNu+fTtbvXo1/5kbGBjwddy5cyffRr788kv25JNPMsYY6+joYK6uruz69euMMcbCw8MF7zUiIoK5\nurqyuro6JpVKmbe3NwsNDWVtbW2ssbGRWVtb8597QEAAH/0+e/Ys8/Pz6/W76q57O+n+9+vt7c0u\nXrzI16/z85IV4Tt16hQLDg7u8/wrVqxg5eVS9tNPjK1ceZj5+T3DduzoYBERjD3wwBdswYLHWX09\nY08//TT79NNPGWNcdM/W1pbl5ub2+/vMzs5mIpGIHT16lL9mX3Xv7++p52fOGGNfffUV27RpU7+f\nHxkelNUlG9ERO0WtXr0aAPj5S0uXLgUA+Pn58alXQkNDUVxcjDt37uD27dvIysrCAw88oPC1fHx8\ncPv2bWzduhU//fQTdHV1AcgX1Zk+fTo/3wQAdu7cyX+LVpX58+dj79692LFjB+Li4mRG0ADg/Pnz\nuH79Oq5evQrGGLZu3QoAMDMzww8//ICXXnoJM2bMQHl5OczNzaGjI3th9rJly3Dz5k2kp6dj9+7d\nuHDhguD4L7/8AgcHB3h5efFlY8aMwddff43g4GCEhYVhyZIlfb6f7p/z6dOn8dhjj8HY2BgA8Mwz\nz+D06dP9fh7PP/88TE1N5X5/586dk/m53IvBvPee7SsjIwOhoaHw8PDAmjVrUFJSgrKyMjg7O6O9\nvR2bNm3CkSNH+Nf1fH1jYyPEYjEfHfH390dJSQkf4QaAJ554ot/30/Occ+bMwfjx4wEAAQEBfOSs\nsbERmzZtgpeXFwIDA1FUVISUlBQA3KTjBx98ELq6uhg1ahT8/Pz413X3559/Ijw8HM7OXNqJUaNG\nwdjYGJcuXYK3tzemTJkCgIsKXbt2jY/curi48FHLdevW4caNG2hoaBjwfnDp0iX4+fnB1dUVAPC3\nv/2NP3bhwgVkZ2djyZIl8PX1xdKlS6GlpYX09HRER0cjMzMT8+bNg6+vL9atWwepVMpH2WTVp76+\nHidOnMCGDRtgbW0NADA0NISenh7i4uIQHh4OGxsbAMDmzZsFbXzu3Ll8HZ9++mk+yrVu3TpER0ej\nuroa0dHRGDt2LB9ty87O5n9Pnb+DxYsXw8TEBFpaWvDy8kJYWBhGjRoFQ0NDTJ48mf+dvPDCC/ji\niy8AcPPE7nX+W3BwMP7xj3/ggw8+wK1btwTR/p7tavz48cjKypJ5HqkUmDJlLb74Qgs3bgBpaceR\nnX0aX33lh0OHfJGZ+QXa23NhbMy1jQMHDgAA/vjjD0ydOhX29vZ9/j47/+/Q19cXzB+VVfeB/p56\nfuYDvS9yf9KIBMWq0jkUK+9wrL6+PgBu+FJPT48v19bWhuSvJEUikQhbt27F7t27IRKJ8Oyzz/bK\nwyePiRMn4tatWzh9+jT++OMPbNu2jR8SGoytW7ciPj4eADes13mzHqxVq1Zh9uzZiI6Oxq5du/D1\n11/j8OHDvZ7XedPR1dXFc889hxUrVvDHQkJCEBISAgAoLS3F+++/z/8H25cJEyZg4cKFSEhIwOzZ\ns/nyr7/+Gps2ber1/KSkJIwZMwb5+fn9nrf770wkEglujAN1rl9++WVkZmbit99+E5T39/7s7OwA\nyP5curO3t0dOTg6mTZsGgFvgMnHiRMFzBvveu3vsscfw8ccfY/ny5WCMwdDQEC0tLbC2tkZqairE\nYjFOnz6N1157DcnJyb1e39HRAS0tLVy9ehXafexk3tlh7kvPv5/Ov0OAmyLR+be3bds22Nra4tCh\nQ9DS0kJYWBhaWlr45/b1N9vzWrJ+v4r8DXfPYzfQ/aC/8zLG4OXlxQ/BdXfp0iUsXrwYBw8elLte\nndfr6/11L+/5nL6OGRkZYe3atfj6669x5syZATtgPX8Hff1OHn74Ybz++utITk6GWCzmO0ry6qzj\nRx99hNTUVMTExOCRRx7BP//5Tzz99NP8e+5O1mfT0sLloCsoAPLzjdH9dvnoo9uxc+dG/NUX5s2Z\nMwf19fW4efMmDhw4gCeffJI/Juv3KRaLMXr0aBh1Zin+i6y6r169esC/p54oA8TwJxaLlZofc0RH\n7BSZY6eIDRs24JdffsHRo0f5m4iiCgsLIRKJsGLFCnz00UcoLy9HdXU1TE1NUVtbe891+/zzz5Gc\nnIzk5GSldeoALrJjbW2NDRs2YMeOHbh8+XKv5zQ1NfF1Z4zh+++/h6+vL3+8pKQEANcZ2LZtG557\n7jkYGBj0Ok/3aE9FRQXi4uIwc+ZMvqygoADnz5/H448/Lnjd5cuXsXv3bly/fh3l5eX43//+J/O9\nmJqaoqamhn+8cOFCxMXFoaGhAYwx7N27F6GhoTJfu23bNiQlJeHnn3/GqB47ePf1/gb6XLp75JFH\nsGfPHjDGUF5ejl9//RUPP/yw0t57T7W1tfx+yvv27UNraysA7nNvbGxEaGgo3n33XZiZmSErKwum\npqZoamqCVCoFwM2HnDt3rmAeUH5+Pj8vbiA9fxcD1dXOzg5aWlq4efMmzp07J9frugsNDUVUVBQf\nRWltbUVDQwMCAgKQkpKCtLQ0AMDBgwfh5+fH/2ecmZmJ8+fPAwBiYmLg5eXFd1j7ux/4+/sjOTmZ\nv97evXv5Y7Nnz0Z6errght65Wjs0NBQnT54UrFbvvpK7e32+/fZbeHl5wcTEBOHh4Th06BDKysoA\ncHNUW1tbsWDBAkRFRfG/lz179gjaeHx8PF/H/fv3819QAG5e5SeffIKkpCQ89NBDfLmjoyMKCgr4\nx/J0LjqfM2rUKGzatAnLly/HunXrBJ15RaSlpcHd3R0vvPAC1q1bx88HlNWuCgoK+C9J7e1ch+6/\n/+WSDDPWVTcHB+CZZ5YjMXE39PS4c7S2tuL69ev8uTZs2IAPPvgA586d4z+TWbNm9fn7lLfuA/09\n9fzMO9+Xk5OTQp8b0SzKnmM3oiN2ipD17a77z90fGxsbY8mSJWhpaYGlpeWA55X1Df769et4/fXX\nAXApLrZt24axY8dizJgxmDx5Mjw9PTF16lSZy/qvXr2KiIgIwfJ/W1vbPodjZ8yYgcLCQtTU1GDC\nhAlYsmQJvvrqq37r3bPuP/74I7755hvo6upCJBLxE/W7Ky0txUMPPQSpVAqpVAp3d3d+uAXg9gSO\nj49HW1sbwsLCsGvXLv5YeHg43n77bfj5+eGrr77CqVOn+I7Tyy+/zA87Adwk8OXLlwsWhNTU1ODx\nxx/HwYMHYWVlhW+++QYBAQGYNWuWYMgSANavX4+NGzfixx9/xL/+9S+sW7cOmzdvxqxZs/jP6803\n3+z1/lJTU7Fr1y5MnjyZjx46OTnh//7v//p9fwN9Lt3f+/r163Hp0iW+Ux4REQEHh66kpoN97z3b\n4yeffIIHH3wQFhYWWLx4Mb+4Jz8/H3/7298gkUggkUiwdOlSBAQEAAAef/xxeHp6YvTo0Th//jy+\n+eYbvPTSS/y1TExMsH//fn7Yrz/PPPMM/vWvf+H999/HBx98IPPvpfPxm2++ifXr12Pfvn2YNGkS\n5s+f3+u99fcY4IYw9+zZg9WrV0MqlUJbWxuHDh2Cu7s7Dh8+jLVr10IikcDa2lqQXsjT0xN79+7F\nc889ByMjI8Ek/J73g+5/n9bW1vjqq6+wbNkyGBgY4KGHHuLrZWFhgePHj+OVV17BP/7xD7S1tcHZ\n2RnHjx+Hi4sLjhw5gqeeegrNzc1oa2tDYGAgnzqpr/rMnz8fr7/+OhYuXAgtLS3o6enh999/h7u7\nO3bt2oVFixZBJBLB2dmZ7/yLRCLMmTMHL7/8MtLT0zFu3DhBNL4zVU9AQIBg6sSCBQtw8eJFLF++\nvNfvSp7fyVNPPYWdO3fiueee6/V7GkjneV5//XWkp6dDR0cHFhYWfHqg7u3qww8/RHBwMC5cuIDg\n4IW4epXb0/WDD3zx+ON/wNh4LABg9GgR1q0DnJ0BkWgdGKvg21hHRwe2bNnCt/EnnngCEydOxKZN\nm/hOaV+/z99++w05OTm9Pou+6t7f31PPzxzghvQXLlyo8GdIRq4hTVCsSqoMT0skEnh7e+PQoUP8\nkJkqTZw4kb85q+r8J06cgJubm0rOT8hwJhaL8corr/QZfVH3/WCg+ijqwIEDOHHihMzV1wC3Snnq\n1Km4evUqxo0bx5eXlZUhKChIZg5MeRw5cgQ//PBDrykNqsAYMHWqD9au/R0dHXaCY6NHA8HBgLu7\n5icX7vmZt7a2ws3NDampqfcc9SSaY0QkKB6OOr9Nh4WFqeUmDnAT49esWXNPCYr705mgWCKR9BpW\nvN/Q/p+kLz0jUd3bylDcD2RFxlR1vi+//BLu7u54+eWXBZ06ALC2tkZ4ePiAyYJlCQsLw1tvvaXy\n5LqMARkZwNatJ2BiMkfQqTMxAR54gEsu7OGhuk6dMu8tPT/zvXv34tlnn6VOHREY0RG7iIgIhRZP\nDMaKFSuQl5cnKHNwcMAvv/yi8muTwROLxWppJ2T4o7aiGlFRUXjjjTd6lb/77rtYvHixwufLzwdO\nnwb+SjXJ09cHAgMBf38uD52qUXshA+lcPLFz506lROxGdMduhL41QgghfSgrA2JigL/WwfBGjQIC\nAoDZswEZa7YIGXLK6rfQ4glCCCHDXk0NEBcHXL/ODcF20tICpk0D5s3jhl8JGelojh0hoDl2RH7U\nVjRLQwPwxx/AZ58BKSnCTp2nJ7B1KxAePnSdOmovRN0oYkcIIWTYaWkBLlwALl4E2tqExyZN4la6\njh07NHUjZCjRHDtCCCHDRns7cOUKl1i4uVl4zN4eCAnhkgwTMtzQHDs5KLqlGCGEEM3U0QFcuwaI\nxUBdnfCYjQ3XoXN11fxcdIT0pOwtxfqM2K1fv16uE+jp6Qm2ydEUFLEjiqCUBERe1FbUizHg9m0g\nNhaoqBAes7AAFizg8tBpaeiMcWovRF4qj9gdPXoU27Zt6/MinRX48MMPNbJjRwghZHjLyuJy0RUV\nCcuNjblVrtOmAdraQ1M3QjRVnxE7Z2dnZGZmDniCyZMn8xtnaxKK2BFCyPBUWMh16LKzheV6el3J\nhXV1h6ZuhKiKsvottHiCEEKIRigv54Zcb98WluvocJ25OXMAQ8OhqRshqjake8VmZWUhJydn0Bcn\nRFNQrikiL2oryldbC/z6K/DFF8JOXWdy4RdeABYtGp6dOmovRN3k6titWbMGFy5cAADs378f7u7u\ncHNzo7l1hBBC7lljIxAdDXz6KZCcLEwu7O4ObNkCLFsGmJoOXR0JGW7kGoodM2YMCgsLoaurCw8P\nD/zvf/+Dubk5VqxYgYyMDHXUU2EikQgRERGU7oQQQjRMayuQkMD9a20VHnNx4VKXjBs3NHUjbkfQ\ncwAAIABJREFURN06053s3LlTfXPszM3NUVNTg8LCQsycOROFhYUAABMTE9TX1w+6EqpAc+wIIUSz\nSCTA1avA2bNAU5PwmJ0dsHAh4Og4JFUjZMipNUGxt7c33n33XeTk5CA8PBwAUFBQADMzs0FXgBBN\nQLmmiLyorSiuowO4fh2Ii+Pm03U3ZgwXoZs8eWQmF6b2QtRNro7dvn37sH37dujq6uK9994DACQk\nJODxxx9XaeUIIYQMX4wBd+5wK13Ly4XHzMy45MJeXpqbXJiQ4YjSnRBCCFG67GwuF91fM3d4RkZd\nyYV1RvSmloQoRu17xZ47dw7Jycmor6/nLy4SibBt27ZBV4IQQsjIUFQExMQAPfPb6+kBs2cDAQHc\nz4QQ1ZCrY/f888/j6NGjmDt3LgwMDFRdJ0LUjubBEHlRW5GtooKbQ5eaKizX0QFmzADmzh2eeegG\ni9oLUTe5OnZHjhxBamoqbG1tVV0fQgghw0hdHXDmDJeHrqOjq1wkAnx8gKAgbj4dIUQ95Jpj5+Xl\nhdjYWFhZWamjTkpBc+wIIUR1mpqA8+eBy5e5NCbdublxCyPGjBmauhEyHKl1r9grV67gnXfewdq1\na2FjYyM4Nm/evEFXQhWoY0cIIcrX1gZcvAjEx/dOLuzkxKUuGT9+aOpGyHCm1sUTiYmJiIqKwrlz\n53rNscvPzx90JVQlMjKSdp4gcqF5MERe92tbkUqBxERu2LWxUXjM1pZLLuzkNDR102T3a3sh8uvc\neUJZ5IrYWVpa4vvvv8eiRYuUdmFVo4gdUQTdfIm87re20tEB3LzJLYyorhYes7ICgoOBqVNHZnJh\nZbjf2gu5d2odirW3t0dGRgZ0dXUHfUF1oY4dIYTcO8aAu3e51CVlZcJjpqbcHDpvb0ouTIiyqLVj\nd+DAAVy+fBnbt2/vNcdOS0P/qqljRwgh9yY3l0su3HOmjaEhl7ZkxgxKLkyIsqm1Y9dX500kEkEq\nlQ66EqpAHTuiCBouIfIayW2lpISL0KWnC8t1dYFZs7gEw5RcWDEjub0Q5VLr4omsrKxBX4gQQohm\nqqri5tDduCEs19YGpk/ntgAzMhqauhFCFEN7xRJCyH2qvp5b5ZqU1Du5sLc3l1zY3HzIqkfIfUVZ\n/ZY+J8ht375drhNEREQMuhKEEELUp7mZm0P36afA1avCTt2UKcBzzwEPPkidOkKGoz4jdsbGxrh+\n/Xq/L2aMYdq0aaipqVFJ5QaDInZEETQPhshrOLeV9nbg0iVux4iWFuExR0cuF52d3ZBUbcQazu2F\nqJfK59g1NTXBxcVlwBPo0UxaQgjRaFIpN9x69iw3/NrduHHcbhHOzpSLjpCRgObYEULICMVYV3Lh\nqirhMUtLLrmwmxt16AjRBGpdFUsIIWT4YAzIyOBSl5SUCI+ZmHCLInx8uFWvhJCRRTOzCxOiZsrc\np4+MbJreVvLygAMHgG++EXbqDAyARYuAF14Apk2jTp26aHp7ISPPiI7YRUZGIigoiCauEkJGvNJS\nIDYWSEsTlo8aBQQEAHPmAPr6Q1M3QkjfxGKxUr8A0Bw7QggZxqqru5ILd7/laWl1JRc2Nh66+hFC\n5KPWOXZlZWUwMDCAiYkJJBIJDh06BG1tbaxfv15j94olhJCRrKGBW+WamMiteu0kEgGensCCBYCF\nxdDVjxAyNOTqlT3wwAPIyMgAALzxxhv48MMP8fHHH+Of//ynSitHiLrQPBgir6FuKy0t3JDrp58C\nly8LO3WTJgHPPgusWkWdOk0x1O2F3H/kitilp6fDx8cHAHDkyBFcuHABJiYmcHNzwyeffKLSChJC\nCOGSC1+5Apw7x+0c0Z29PZdc2N5+aOpGCNEccs2xs7KyQkFBAdLT07FmzRqkpqZCKpXCzMwMDQ0N\n6qinwmiOHSFkJOjoAJKTuT1d6+qEx2xsuA6diwvloiNkuFPrHLvFixfj0UcfRWVlJVavXg0AuHXr\nFuxo7xlCCFEJxoBbt7hh18pK4TELCy65sIcHdegIIUJyRexaWlpw8OBB6OrqYv369dDR0YFYLEZJ\nSQnWrFmjjnoqjCJ2RBG0nyORl6rbCmNAVhZw+jRQXCw8ZmwMzJ8P+PlRHrrhgu4tRF5qjdjp6+tj\n8+bNgjJqqIQQolwFBdxuEdnZwnJ9fS4Pnb8/oKs7NHUjhAwPfUbs1q9fL3ziX/F+xhj/MwAcOnRI\nhdW7dxSxI4QMF+XlXIfuzh1huY5OV3JhA4OhqRshRD1UHrFzdnbmO3AVFRU4ePAgli1bBgcHB+Tm\n5uL333/Hhg0bBl0BQgi5X9XUAGIxkJLSO7mwnx+XXNjUdMiqRwgZhuSaYxcaGort27dj7ty5fNn5\n8+fx1ltv4dSpUyqt4L2iiB1RBM2DIfJSRltpbOTSlly5IsxDB3ALIhYsACwtB3UJoiHo3kLkpdY5\ndhcvXkRAQICgzN/fHwkJCYOuACGE3C9aW4GEBODCBaCtTXjMxQUICQHGjRuauhFCRga5Inbz58/H\njBkz8Pbbb8PAwABNTU2IiIjApUuXcPbsWXXUU2EUsSOEaAqJpCu5cFOT8NiECVyHztFxSKpGCNEQ\nao3YHThwAGvXroWpqSksLCxQXV2N6dOn49tvvx10BQghZKTq6ODmz4nFQG2t8Ji1NdehmzSJctER\nQpRHrohdp7y8PBQVFWHcuHFwcHBQZb0GjSJ2RBE0D4bIS562whi3wjU2llvx2p25OTeHztOTWyRB\nRja6txB5qTVi10lfXx/W1taQSqXIysoCADg5OQ26Eop67bXXkJCQAEdHR3z99dfQ0VHobRBCiMpk\nZ3PJhQsLheVGRtwq12nTuDQmhBCiCnJF7E6ePImnnnoKxT3SoItEIkh7LulSsZSUFHzwwQc4fPgw\n3nnnHTg5Ocnc/YIidoQQdSoq4jp0f33n5enpcXnoAgIouTAhpG/K6rfINRDw97//Hdu3b0dDQwM6\nOjr4f+ru1AFAQkICwsLCAHB72MbHx6u9DoQQ0qmiAjh6FPjqK2GnTkcHmD0bePFFLlJHnTpCiDrI\n1bGrqanB5s2bYWhoqOr6DKi6uhomJiYAAFNTU1RVVQ1xjchIIBaLh7oKZJjobCt1dcDx48AXXwC3\nbnUdF4m45MLPPw+EhgIacNskQ4juLUTd5OrYPfXUU/j666+VeuHPP/8c06dPh76+Pp588knBsaqq\nKqxcuRLGxsZwdHTEd999xx8zNzdHXV0dAKC2thajR49War0IIUSWtLRc7N4di+++u4atW2OxfXsu\nkpK4la+d3NyALVuA5csBM7Ohqysh5P4l1xy7wMBAXL58GQ4ODhg7dmzXi0Wie85j9/PPP0NLSwvR\n0dFobm7G/v37+WOPPfYYAGDfvn1ITk5GeHg4Lly4ADc3N6SkpOCjjz7CwYMH8c4778DZ2RmrV6/u\n/cZojh0hREnS0nKxb18GyspCkJ/P7RYhkcTAx8cFVlYOcHbmUpfY2g51TQkhw5VaV8U+/fTTePrp\np2VW4l6tXLkSAHD16lUUFBTw5Y2NjTh27BhSU1NhaGiIOXPmYMWKFTh8+DDeffddeHt7w8bGBvPm\nzYODgwNeffXVPq+xceNGOP6V9dPc3Bw+Pj78svPO8Dg9psf0mB7391giAXbs+D9kZPjhr1kgqKkR\nA9BGVVUm/vlPB+TliXH3LmBrO/T1pcf0mB4Pj8edP+fk5ECZFMpjpwpvvvkmCgsL+YhdcnIyAgMD\n0djYyD/no48+glgsxvHjx+U+L0XsiCLEYjH/R0cIwA2xXrsGiMXAqVNitLQEAeA6dba2QZg4EXBx\nEeOll4KGsppEw9G9hchLrRE7xhj279+Pw4cPo7CwEHZ2dli3bh2efPLJQUXtgN5Rv4aGBpiamgrK\nTExMUF9fP6jrEEKIPBgDUlOBuDigspIr09LiJtLp63NbgPn5cYsk9PQ6+jkTIYSon1wdu3feeQeH\nDh3Cv/71L9jb2yMvLw/vv/8+ioqK8Oabbw6qAj17p8bGxvziiE61tbX8SlhCVIG+URPGgPR0ICYG\nKC0VHvPwcEZZWQzs7UOgpRUEAGhtjUFIiIva60mGF7q3kIGkZaThdOJppZ1Pro7dnj17cObMGcE2\nYmFhYZg7d+6gO3Y9I3aTJk2CRCJBRkYGXFy4m2ZKSgo8PDwUPndkZCSCgoLoD4sQ0q+cHK5Dl58v\nLNfXBwIDgZkzHZCdDcTExKKtTQu6uh0ICXHB5MmavbUiIUSzpWWkYccXO3Cr4NbAT5aTXB27pqYm\nWFlZCcosLS3R0tJyzxeWSqVob2+HRCKBVCpFa2srdHR0YGRkhFWrVmHHjh3Yu3cvkpKS8NtvvyEh\nIUHha0RGRt5z/cj9hebB3J+KirgOXWamsHzUKG6niNmzAQMDrmzyZAdMnuzwV1sJVn9lybBE9xbS\nF8YYDsQdQJVPFSy9LIEflXNeLXmetHjxYqxbtw537txBc3Mzbt++jSeeeILfAeJevP322zA0NMR/\n/vMfHDlyBAYGBvh//+//AQC++OILNDc3w9raGuvWrcOXX36JqVOn3vO1CCGku/Jy4IcfuN0iunfq\ntLUBf39ut4iQkK5OHSGEKFNpQyn2X9uP5NJktBTWwPR04cAvkpNcq2Jra2vx/PPP44cffkB7eztG\njRqFRx99FJ999hnMzc2VVhllEolEiIiIoKFYQgivuppb5Xr9OjenrpNIBPj4APPnAxp6SyOEjAAt\nkhaIc8S4XHgZHawDF36OhdblOzAra8XRrDqlrIpVKN2JVCpFRUUFrKysoK2tPeiLqxKlOyGEdKqv\nB86dAxITueTC3bm7AwsWAD1mmxBCiNIwxnCj7AZOZZ5CQ1sDX162PwaPp+TBlgFO1/OU0m+Rayj2\n4MGDSElJgba2NmxsbKCtrY2UlBQcPnx40BUgRBN0TxhJRo7mZuD0aeDTT4HLl4WdOldXYPNm4JFH\nFOvUUVshiqD2Qsoay3Dg2gEcu31M0KnzaDLGiioT+OrZQLddV2nXk2vxxPbt23Ht2jVBmZ2dHZYt\nW4b169crrTKEEKIMbW3AxYtAfDzQ2io8Zm/PzZ9zoAWthBAVapW0QpwjxqXCS+hgXTkvzUQGWFVo\nCvu0EsRJAUMDQxgaGAK3M5RyXbmGYi0sLFBRUSEYfpVIJLC0tERtba1SKqJsNBRLyP1HIgGuXuWG\nXbttXgMAGDcOCA4GXFy4OXWEEKIKjDHcLLuJU5mnUN/WtbmClkgLQXDE7MRy6NRx5bkVFchITUXI\nlCkQffml+naemDp1Kn766SesXr2aL/v55581fqUq5bEj5P7Quf3XmTNAz++aVlbcHDo3N+rQEUJU\nq6yxDFHpUcipyRGUOxnZYXm+Icyv3xWUO8yZgwRPT2xQYMvUgcgVsTt//jyWLl2KRYsWwcnJCZmZ\nmTh9+jSioqIQGBiotMooE0XsiCIo19TwJGv7r05mZkBQEODtDWjJNZtYPtRWiCKovdwfWiWtOJN7\nBhcLLgqGXU10TRA+yh2T4+9AVFPT9QJ9fWDJEsDLi//Gqda9YgMDA3Hjxg18++23KCgowMyZM/Hf\n//4XEyZMGHQFCCFEUYwBGRlccuGSEuExIyNg3jxg2jRAR647HCGE3BvGGFLLUxGdEd1r2DVgjB8W\nZEoxKvGi8EWTJgHLlgEq2ipV4XQnpaWlsLW1VUlllIkidoSMTLm5XIcuL09Yrq8PzJnDJRjWVd4C\nM0IIkam8sRxR6VHIrskWlDuYOWCZnies/oznkmd2khGl605Z/Ra5OnbV1dXYsmULfvrpJ+jo6KCp\nqQnHjx/H5cuX8e9//3vQlVAF6tgRMrIUFQGxsVykrjtZ238RQoiqtEnbcCbnDBIKEgTDrsa6xgib\nsAAeN0ohunxZ+CI5onTK6rfINfPk2WefhampKXJzc6GnpwcAmDVrFr7//vtBV0CVIiMjKYcQkQu1\nE81VXg4cPcpt/9W9U6etDcycqf7tv6itEEVQexk5GGNILUvF55c/R3x+PN+p0xJpIcAuAFttlsHz\n53hhp05fH1i5EnjssT47dWKxWKl728sVsbOyskJxcTFGjRoFCwsLVP8VWjQ1NUVdXZ3SKqNMFLEj\niqAJzpqnpobb/islpff2X97e3MKIodj+i9oKUQS1l5GhoqkCUelRyKrOEpTbm9kj3GERbC7dBC5d\nEr5o0iTggQcAU1O5rqHWoVgXFxecPXsWtra2fMcuLy8PoaGhuHPnzqAroQrUsSNkeGpoAM6elb39\nl5sbl7pkzJihqRsh5P7SJm3D2dyzSMhPgJR13ZCMRhkh1DkUXi1mEB0/DlRVdb1IXx9YvJj7BqpA\njiW1rop9+umn8fDDD+Pf//43Ojo6kJCQgG3btmHz5s2DrgAhhADc9l/x8dyX3vZ24TEXFy658DBY\nt0UIGQEYY7hdcRvRGdGobe1KjimCCDPHz8SC8XOgfyYeuPyLcEjB1ZWbSydnlE4V5IrYMcbw6aef\n4n//+x9ycnJgb2+PZ599Fi+++CJEGprxkyJ2RBE0XDJ02tq4zlx8PNDSIjymidt/UVshiqD2MvxU\nNlXij4w/kFElXKk1wXQCwieFY2xlK/Drr0qJ0nWn1oidSCTCiy++iBdffHHQFySEEIDb/isxkRt2\n7bn919ixXIeOtv8ihKhLu7QdZ3PP4kL+hV7DroucF8F7tBtEsbHcN1ENi9J1J1fELjY2Fo6OjnBy\nckJxcTFee+01aGtr491338XYsWPVUU+FiUQiRERE0JZihGiYjg5uQYRY3Hv7L0tLbsiVtv8ihKgL\nYwx3Ku7gZMbJXsOuM8bPwALHBTAoLgd++aV3lC4sDPDxGdQNSywWQywWY+fOnepbPDFlyhScOnUK\n9vb2eOyxxyASiaCvr4+KigocV+L+ZspEQ7GEaBbGgFu3uFx06tr+ixBC+lPVXIU/0v9AelW6oNzO\n1A7hruEYp2/FZURXQ5ROratiO9OatLe3w8bGhs9nN27cOFT2vENrCOrYEUXQPBjV6dz+KzYWKC4W\nHhuO239RWyGKoPaimdql7Tifdx7n884Lhl0NRxlikdMi+Iz1gSg/n5tL172fo6fHzaUbZJROFrXO\nsTM1NUVJSQlSU1Ph7u4OExMTtLa2or3n0jVCCOmmv+2/Zs/mdoyg7b8IIerCGMPdyrv4I+MP1LTU\n8OUiiDDddjqCJwbDADrAqVPAxYvCKJ2LC7B8ucbMpeuLXB27559/HjNnzkRrays++eQTAEB8fDym\nTp2q0soRoi70jVq5iou5Dp2s7b/8/bk9XYfr9l/UVogiqL1ojqrmKpzMOIm7lXcF5eNNxiN8Ujhs\nTWy5b6FqjNKpglxDsQCQlpYGbW1tuLi4AADu3r2L1tZWeHp6qrSC94qGYglRv4oKbsj11i1hubY2\nN9w6d26/WyUSQojStUvbEZ8fj/N55yHpkPDlhqMMsdBpIXzH+kIkkXA3L1lRumXLuInAKqbWOXbD\nEXXsiCJoHszg1NQAZ84A167J3v5r/nzAwmLo6qdM1FaIIqi9DK27lXfxR/ofqG6p5stEEGGa7TQE\nTwyG4ShDID+fW/HaM0oXFgb4+qotSqfyOXZTpkzhtwubMGFCn5XI6zl5RoNERkZSuhNCVKihATh3\nDrh6lbb/IoRojurmapzMOIm0yjRBua2JLcJdwzHedDy3xU109JBG6YCudCfK0mfE7ty5c5g7dy5/\n0b5oaqeJInaEqE5zM3DhAnc/pO2/CCGaQtIhQXxePM7lnRMMuxroGHDDruN8oSXS0pgoXXc0FDsA\n6tgRonz9bf81YQK3W4Sj45BUjRByn0uvTMcfGX+gqrkribAIIviN80OIUwg37NreDsTFAQkJwiid\nszO34lVNUTpZVD4Uu3379j4v0lkuEonw1ltvDboShAw1mgfTv87tv86d44Zfuxs7lovQuboOiwVj\ng0ZthSiC2ovq1bTU4GTGSdypuCMotzWxxVLXpbAzteMKOvPSVVR0PWmIo3Sq0GfHLj8/H6J+3mRn\nx44QMnINtP3XggWAu/uIuR8SQoYRSYcEF/Iv4FzuObR3dM0JMdAxQIhTCPzG+XHDrhocpVMFGool\nhPTCGHD7Nrf6v/uXW4DLzRkUxKV0ou2/CCFDIaMqA1HpUYJhVwDcsOvEEBjpGnEFBQXcXLqeUbrQ\nUMDPT6O+lap8KDYrK0uuEzg5OQ26EoQQzcAYkJnJJRfuuf2XoSG3/df06cNn+y9CyMhS21KLkxkn\ncbvitqB8nPE4LHVdiglmf2XxuM+idN31GbHTkuOruEgkgrRnjgMNQRE7ogiaB8MlXI+J4bYB605P\nj9spwt+f+/l+R22FKILai3JIOiRIyE/A2dyzgmFXfR19BE8MxnTb6dywKyA7Sqery82l07AoXXcq\nj9h1dHQM+uSEEM1XXMwNuaanC8tHjQJmzgQCA4fv9l+EkOEvsyoTUelRqGyuFJT7jPXBIqdFXcOu\nEgkXpbtwQRilc3LionTm5mqs9dAZ0XPsIiIiKEExIX2oqODugampwnItLW77r3nzaPsvQsjQqW2p\nRXRmNG6VC/coHGs8Fktdl8LezL6rsK8oXWgod0PT0Cgd0JWgeOfOnarNYxcWFobo6GgA4BMV93qx\nSISzZ88OuhKqQEOxhMjW3/ZfXl7cwoiRsv0XIWT4kXZIkVCQgDM5ZwTDrnraegieGIwZ42d0DbuO\noCidyodin3jiCf7np556qs9KEDIS3A/zYPrb/mvqVC51ibX10NRtOLkf2gpRHmovismqzkJUehQq\nmoTL8b1tvLHIeRGMdY27CgsLuShdeXlX2TCJ0qlSnx27xx9/nP9548aN6qgLIUQFWlq6tv9qaxMe\nc3bmkguPHz80dSOEEACoa61DdEY0UsuFc0NsjGyw1HUpHMwdugolEi65Znz8sI/SqYLcc+zOnj2L\n5ORkNDY2AuhKULxt2zaVVvBe0VAsud+1tQGXLwPnz8ve/is4GJg4cWjqRgghADfserHgIs7knkGb\ntOubp562HhZMXICZ42d2DbsCIzpKp/Kh2O6ef/55HD16FHPnzoUBLY8jRKNJJEBSEnD2bO/tv2xs\nuP1c75ftvwghmiu7OhtR6VEobyoXlHvZeGGR0yKY6HVbvdVXlG7iRGDFivs+StedXBE7CwsLpKam\nwtbWVh11UgqK2BFFjIR5MB0dwPXr3L2vpkZ4bPRobg6dhwd16AZrJLQVoj7UXnqra63DqcxTuFl2\nU1BubWSNcNdw4bAr0HeUbtEiLmP6CLmpqTViN2HCBOjq6g76YoQQ5evc/isuTnjfA7jtv+bP57b/\n0tYemvoRQgjADbteKrwEcY6417BrkGMQZo6fCW2tbjcqitLdE7kidleuXME777yDtWvXwsbGRnBs\n3rx5KqvcYFDEjox0ndt/xcYCRUXCY4aGwNy5wIwZtP0XIWTo5dTk4MTdE72GXT2tPRHqHCocdgXu\nmyhdd2qN2CUmJiIqKgrnzp3rNccuPz9/0JUghCimv+2/Zs8GAgJo+y9CyNCrb63HqcxTuFF2Q1A+\nxnAMwieFw9HcUfgCiYRLtBkfz80v6eToyEXpKMnmgOSK2FlaWuL777/HokWL1FEnpaCIHVHEcJkH\nU1LCdeh6bv+lo8Pt5TpnDhetI6ozXNoK0Qz3a3vpYB24VMANu7ZKW/lyXW1dBDkGwX+8v3DYFeCG\nHn75BSgr6yob4VG67tQasTMyMsL8+fMHfTFCyL2prOTm0N0UzjWm7b8IIRontyYXUelRKG0sFZR7\nWHsg1DkUpnqmwhdQlE6p5IrYHThwAJcvX8b27dt7zbHT0tLq41VDiyJ2ZCSore3a/qv7/Y62/yKE\naJqGtgb8mfknUkpTBOVWhlZY6roUThZOvV8kK0o3ahQXpZsxY8RH6bpTVr9Fro5dX503kUgEac+9\niTSESCRCREQEgoKC7sswOBneGhu57b+uXOm9/deUKVxyYdr+ixCiCTpYB64UXkFsdmyvYdf5DvMR\nYBfQe9iVonQ8sVgMsViMnTt3qq9jl5OT0+cxR0fHQVdCFShiRxShKfNg+tv+y8mJSy5M238NLU1p\nK2R4GOntJa82Dyfunug17Oo+xh2hzqEw0zfr/SKK0smk1jl2mtp5I2SkaG8HLl3ivrw2NwuP2dlx\nHTra/osQoika2hpwOus0rpVcE5RbGlhiqetSOI927v0iiYTbEuf8eWGUzsGBi9KNHq3iWt8f5N4r\ndrihiB0ZDqRSIDFR9vZf1tZch27SpPv2CywhRMN0sA5cLbqK2OxYtEi6NqEepTUK8x25YVcdLRkx\no+Ji4Oefe0fpFi4EZs6kmxzUHLEjhChXRwdw4wa30pW2/yKEDAf5tfk4kX4CJQ0lgnK3MW4Icw6T\nPewqlXJz6ShKpzbUsSME6psHwxhw5w63WwRt/zU8jfQ5U0S5RkJ7aWxrxOms00guSRaUWxpYYonr\nEriMdpH9wuJibi5dabf5dxSlUznq2BGiBowBWVlccuG+tv+aPp275xFCiCboYB1ILEpETHZMr2HX\neQ7zMGvCLNnDrlIpN7/k3DmK0g0BuebYZWVl4Y033sC1a9fQ0G0ikEgkQl5enkoreK9ojh3RFPn5\nXIeu5+JyPT1g1izuH23/RQjRJAV1BThx9wSKG4oF5VOtpiLMJQzm+uayX0hRunum1jl2a9euhYuL\nCz766KNee8USQmQrKeGGXO/eFZbr6HD3t8BA2v6LEKJZGtsaEZMdg6TiJEH5aIPRWOKyBK6WrrJf\nSFE6jSFXxM7U1BTV1dXQHkYTfyhiRxShzHkw/W3/5efHbf9lair7tUTzjYQ5U0R9hkt76WAdSCpO\nQkxWDJolXTmXdLR0MM9hHmZPmC172BXoO0oXEsJtYk1ROrmoNWI3b948JCcnY/r06YO+ICEjVX/b\nf3l6ctt/0ZdWQoimKawrxIn0EyiqF04AnmI1BWHOYbAw6GMXiL6idPb2wIMP0g1viMgJ7JYsAAAg\nAElEQVQVsduyZQt++OEHrFq1SrBXrEgkwltvvaXSCt4ritgRdWls5FbyX7nC5d/sbsoULnVJjy2W\nCSFkyDW1NyEmixt2Zej6/9JC3wJLXJdgkuWkvl9cUsJF6Uq6pT6hKN2gqDVi19jYiAceeADt7e0o\nKCgAADDGIKJfHLmPtbQACQncP1nbfwUHc7tGEEKIJmGMIak4CaezTvcadg20D0SgfWDfw65SKReh\nO3u2d5RuxQrA0lLFtScDoZ0nCIFi82Da24HLl7koXc/tv8aP576wOjkpv45EMwyXOVNEM2haeymq\nL8KJuydQWF8oKJ9kOQlLXJb0PewK9B+lmzmTm0hM7pnKI3Y5OTn8HrFZWVl9nsCJ/gcj9wmpFEhK\n4ubRydr+KzgYmDyZRiAIIZqnub0ZMdkxSCxKFAy7muubY4nLEky2mtz3iylKN6z0GbEzMTFBfX09\nAECrj164SCSCVCpVXe0GgSJ2RFk6t/8Si4HqauExC4uu7b/oyyohRNMwxpBckozTWafR1N7El+to\n6WDOhDkItA/EKO1+MqPLitLp6HTNpaMbn9Ioq98y7IZi6+rqsHDhQty+fRuXLl2Cm5ubzOdRx44M\nVn/bf5mYcNt/+frS9l+EEM1UXF+ME+knUFBXICh3He2KJa5LMNqgn1WrfUXpJkzgVrxSlE7p1Lp4\nQpMYGhoiKioKr7zyCnXciNJ0nwfTuf1XbCxQKJyGAgMDbvuvGTNo+6/7labNmSKabSjaS3N7M2Kz\nY3G16GqvYdfFLosx2XJy/4sfKUo3rA27jp2Ojg6srKyGuhpkhOpr+y9dXWD2bNr+ixCiuRhjuFZy\nDX9m/SkYdtUWaWOO/RzMtZ/b/7CrVMqtCjtzpneUbsUKgP7vHRaGXceOEGVKS8vF6dOZqKzUwsGD\nsTAycoaVlQN/nLb/Ij1RtI4oQl3tpbi+GFHpUcivyxeUu4x2wRKXJbA0HGDotLSUi9IVd9sblqJ0\nw5Jaf1Off/45pk+fDn19fTz55JOCY1VVVVi5ciWMjY3h6OiI7777jj/28ccfY8GCBfjwww8Fr6E8\nemQw7tzJxWefZeDMmWDExgYhOzsY165loKIiF1pawPTpwAsvAKGh1KkjhGimFkkLotKj8FXiV4JO\nnZmeGVa7r8bjno/336mTSrkI3VdfCTt1EyYAzz7LDVNQp25YUThi19E9PIu+V8zKMn78eGzfvh3R\n0dFo7pEAbMuWLdDX10dZWRmSk5MRHh4Ob29vuLm54aWXXsJLL73U63w0x47ci5YWbpXrhx9moqQk\nBABQUyOGuXkQdHRC0Noai61bHWg3HCITzbEjilBVe2GMIaU0BX9m/onG9ka+XFukjdkTZmOuw1zo\nauv2f5K+onTBwUBAAHXohim5OnaJiYnYunUrUlJS0NLSwpcrmu5k5cqVAICrV6/yO1gA3M4Wx44d\nQ2pqKgwNDTFnzhysWLEChw8fxrvvvtvrPEuXLkVKSgrS0tKwefNmbNiwQe46kPsTY9xCiMRE4OZN\nLslwTY3wpmVpCUycCNjZaVGnjhCisUoaShCVHoW82jxBubOFM5a4LoGV4QBz4aRSID6ei9R1/z/c\nzo5b8Upz6YY1uTp2GzZswPLly7Fv3z4YKmFMqmek7e7du9DR0YGLiwtf5u3tDbFYLPP1UVFRcl1n\n48aNfJJlc3Nz+Pj48N+cOs9Nj0f2Y3//IFy/Dnz3nRg1NYCjI3c8J0eM6uokGBkFwdoaGDMGMDIS\nw9g4CLq6HRpTf3qseY+DgoI0qj70WLMfK7O9BAQGIC47Dj+c+AEA4OjjCAAoSy3DzPEzsW7+OohE\nov7PV1oK8X/+A1RVIeiv/x/F+fmAry+CNm0CtLQ06vMbyY87f87puVpvkOTKY2dqaora2lqlzWnb\nvn07CgoKsH//fgDAuXPn8Oijj6K4Wzh4z549+PbbbxEXF3dP16A8dvcvxrjVrYmJQGoqIJH0fo6N\nDWBpmYsrVzJgZBTCl7e2xmDjRhdMnuzQ+0WEEDIEGGO4Xnodf2b9iYa2rm1vtEXamDVhFuY5zBt4\n2JWidBpPrXnsVq5ciejoaCxevHjQFwR6R+yMjY1RV1cnKKutrYWJiYlSrkfuD83NQEoK16HrmVAY\n4PLOeXgA06Zxe7qKRA7w9gZiYmJx69Z1uLl5ISSEOnWkf2KxmP/mTchABtteShtKEZUehdzaXEG5\nk4UTlrouHXjYFQDKyoCff+49l27BAlocMQLJ1bFrbm7GypUrMXfuXNjY2PDlIpEIhw4dUviiPSN/\nkyZNgkQiQUZGBj8cm5KSAg8PD4XP3V1kZCQfCicjE2NAXh7Xmbt1S3Z0buxYrjPn6Qno6wuPTZ7s\ngMmTHSAWa1E7IYRojFZJK8Q5YlwqvIQO1rVo0VTPFGHOYXAb4zbwKFpHR1deOorSaSyxWCwYnh0s\nuYZiIyMjZb9YJEJERITcF5NKpWhvb8fOnTtRWFiIPXv2QEdHB9ra2njssccgEomwd+9eJCUl4YEH\nHkBCQgKmTp0q9/l71o2GYkeupibg2jUgKQmoqOh9XFeX68hNmwaMGwdQZhxCyHDAGMPNspuIzowW\nDLtqibQwy24W5jvOH3jYFeCidL/8AhQVdZVRlE6jDcu9YiMjI/HWW2/1KtuxYweqq6uxadMm/Pnn\nn7CyssKuXbuwZs2ae74WdexGHsa4HSESE4Hbt4VfQDvZ2nKdOQ8P2iGCEDK8lDWWISo9Cjk1OYLy\nieYTsdR1KcYYjRn4JB0d3Fw6sbh3lG7FCm6lGNFIau/YxcXF4dChQygsLISdnR3WrVuH4ODgQVdA\nVahjN3I0NnZF5yorex/X0xNG5+4FzZsi8qK2QhQhT3tplbTiTO4ZXCy4KBh2NdE1QZhLGNzHuMu3\neJGidMOaWhdP7N27F9u2bcPTTz8Nf39/5OXlYe3atXjrrbfwzDPPDLoSqkJz7IYvxoDsbC46d+eO\n7Ojc+PFd0TldOUYmCCFEkzDGkFqeiuiMaNS31fPlWiItBNgFYL7DfOjpyDH00FeUbvx4bi4dRek0\n2pDMsXN1dcVPP/0Eb29vvuz69etYtWoVMjIylFYZZaKI3fDU0AAkJ3PRuerq3sf19ABvb8DPj1sU\nQQghw1F5Yzmi0qOQXZMtKHc0d8RS16WwNrKW70RlZcCvv3IZ2Dtpa3NRutmzKUo3jKh1KNbS0hLF\nxcXQ7RYWaW1tha2tLSpljY1pAOrYDR+MAZmZXHQuLY378tnThAlcdM7NjaJzhJDhq03ahjM5Z5BQ\nkCAYdjXWNUaYcxg8rD3kG3alKN2Io9aO3fLly2Fvb4///Oc/MDIyQkNDA15//XXk5OTgt99+G3Ql\nVIE6dpqvvr4rOldT0/u4vj4XnZs2DbCW88vrvaJ5U0Re1FaIIjrbC2MMt8pvITozGnWtXXlbtURa\n8B/vjyDHIPmGXQEuUecvv1CUboRR6xy7L7/8EmvWrIGZmRlGjx6NqqoqzJ49G999992gK6BKNMdO\n83R0dEXn7t6VHZ2zt++Kzo0apf46EkKIMlU0VSAqPQpZ1VmCcgczByx1XQobY5s+XtlDRwdw4QIQ\nF9c7Srdiheq/AROVGJI5dp3y8/NRVFQEW1tbTJgwQWmVUAWK2GmWurqu6Fxtbe/jBgaAjw83d45G\nEAghw11aRhpOXjmJO5V3kFeTh4lOE2FlyyUENtY1RqhzKDytPeXfqpOidCOeyodiGWN8g+uQFVb5\ni5aGNibq2A29jg4gPZ2LzqWnc3PpenJ05KJzU6dyq/IJIWS4u51+Gx+c+AAFlgVolbYCACQZEvi6\n+SJ8ZjiCHIOgr6M/wFn+0leUztaWm0tHUboRQ+VDsaampqiv55Zf6/TxP65IJIJUVh4Kcl+rreUi\nc8nJXKSuJ0PDruicpuxoQ/OmiLyorZC+dKYv2XlsJ8ptygEpUHOnBuZTzGHpbgnbdlssdlFgz/W+\nonRBQcCcORSlIzL12bFLTU3lf87KyurraYQA4L5IdkbnMjJkR+cmTuSic1OmUHSOEDJyMMaQVpmG\n2OxYlDWWob69KyedtkgbU6ymwMbIBgalBvKdsDNKJxYLN8CmKB2RQ5//vdrb2/M///TTT3j55Zd7\nPeejjz7CP//5T9XUTAlo8YTqVVdzkbnkZG6Va09GRoCvLxedGz1a/fWTF7URIi9qK6QTYwyZ1ZmI\nzY5FUX3Xbg9a0IKOlg4mmE7A+EXjoaPF/VerqyVHrqaKCi5KV1DQVUZRuhFtSBZPmJiY8MOy3VlY\nWKBaVhZZDUBz7FRHKuXyzSUmAllZsqNzzs5cdG7yZO6eRAghI0lOTQ5is2ORV5snKNfV1oWt1Bap\nd1JhNMWIL29Nb8XGBRsx2WWy7BN2dAAJCdxcOorS3ZfUku4kNjYWjDFIpVLExsYKjmVmZsLU1HTQ\nFSDDR1UVN3fu2jVuh4iejI27onMWFuqv32DQvCkiL2or97f82nzE5cT1Sl2io6WDmeNnYs6EOTDS\nNUKaXRpikmJw6+YtuHm4IWRBSN+dur6idPPnc1E6+nZMFNBvx27Tpk0QiURobW3FU089xZeLRCLY\n2Njgs88+U3kFydCSSrm9Wjujcz2JRF3RuUmT6P5DCBmZiuuLEZcTh7uVdwXl2iJtTLOdhrn2c2Gi\nZ8KXT3aZjMkukyG27ueLAEXpiArINRS7fv16HD58WB31URoaih2cysqu6FxjY+/jJiZd0Tlzc/XX\njxBC1KGssQziHDFuld8SlGuJtOAz1gfzHObBXP8eboIUpSM9qHVLseGIOnaKk0iA27e56FxOTu/j\nIhHg6spF51xdaQ4vIWTkqmyqxJncM7hRegMMXf+XiCCCp40n5jvMh6WhpeIn7ugALl4EYmOFUbpx\n47gonY2cu1CQEUetW4rV1tYiMjISZ86cQWVlJZ+wWCQSIS8vb4BXDx1aFSufigquM5eSAjQ19T5u\naspF5nx9ATMz9ddPHWjeFJEXtZWRraalBmdzz+JayTV0MGFyfrcxbghyDIK1kfxDpIL2UlEB/Por\nkJ/f9QSK0t33lL0qVq6O3ZYtW5Cfn48dO3bww7Lvv/8+HnroIaVVRBUiIyOHugoaq729KzqXm9v7\nuEjEzZmbNg1wcaHoHCFkZKtvrce5vHNILEqElAkT70+ynIQFjgswzmTcvZ2conSkH50BqJ07dyrl\nfHINxY4ZMwa3b9+GlZUVzMzMUFtbi8LCQixbtgxJSUlKqYiy0VCsbGVl3Ny5lBSgubn3cTOzrugc\nLXomhIx0jW2NiM+Px+XCy5B0SATHnCycsMBxASaYKb43+v9v787Do6rv/YG/ZyaTTPZ93yEkYU3Y\nRbYsoKK0Kr210lsEacVfRVrt9fbWKltpr+1z3XqFbtSqiMTi0/a2KopIMmwCAROQNRtZIED2fZnM\ncn5/HDPJJAPMJDNnlrxfz8Nj5pwzcz6J30w+810+3+qSElR8/jnkLS0wXLqE8cHBSOzfakehABYt\nAhYsYC8dGUk6FCsIAgK/HoPz9/dHa2sroqOjUVZWNuoAyP60WuDCBbF3bvAIQD+5XKw3N3MmMG4c\ne+eIyP31aHtw/NpxnLh2An36PpNzCYEJyE7KRnJw8oheu7qkBOVvvYXchgagshIwGHCwuhrIzETi\n1KnspSO7siixmzZtGg4fPozc3FwsWLAA69evh6+vL9LSblGTh5xCXZ2YzH31FdDbO/x8cLDYO5eZ\nKa5yHcs4b4osxbbi2jQ6DU7WnsQXV79Ar870jTHGPwY5yTkYHzweMplsxPeo2LsXuefPA52dULe2\nIisoCLlKJfI9PJD4gx+wl47syqLEbufOncavf/vb3+LnP/852trasGvXLrsFRiPT1zfQOzd4FX0/\nuVzcq7W/d24U711ERC5Dq9fi1PVTOFpzFN1a01ViEb4RyEnOQVpo2qgSOnR0AAcOQH7smOmnaT8/\nID0d8rg4JnVkdxYldo2NjZg7dy4AIDIyEm+++SYAoLCw0H6RkVVu3hzondNohp8PCRnonfPzkz4+\nZ8ceGLIU24pr0Rl0KLpRhMPVh9HZZ7plTqh3KLKTszE5fPLoEjq9XlwccegQ0NcHQ/98FrkcWdOn\nA/HxgFwOg6cFe8USjZJFid2SJUvM7hV73333obm52eZB2Yq7lzvRaIDz58WE7vr14ecVCmDiRLF3\nLimJvXNENHboDXqcrTuLQ1WH0KZpMzkXpApCVlIWpkVOg1w2yknF5eXAp5+KpUy+Nn7cOBy8dg25\n6emASgUAOKjRICU3d3T3Irdk63Int10VazAYIAgCgoKC0NZm+otRUVGB+fPno76+3mbB2JI7r4q9\nfl1M5s6dE4dehwoNFZO5jAzA13f4eRqO86bIUmwrzs0gGHC+/jzUVWo095h2PPh7+mNx0mJMj5oO\nhXyUQ6ItLcD+/eKei4OFhwPLlqFaq0XFwYP46uJFTJs0CeNzc5HIeel0G5KsivXw8DD7NQDI5XK8\n8MILow6ALKPRiIncl18CN24MP69QAJMmiQldYiJ754hobBEEAZcaL6GgsgAN3Q0m53yVvliYuBAz\no2dCqVCO7kZaLXD0KHDsmGlNOi8vIDsbmD0bUCiQCCAxLQ1yfhAgid22x67q632lFi1ahCNHjhgz\nSZlMhvDwcPj4+EgS5Ei4Q4+dIJj2zmm1w68JCxvonXPi/x1ERHYhCALKmsuQX5mPm503Tc55e3hj\nfsJ8zImdA0/FKOe3CYLYO/fpp8CQESxkZgJLlnACM40K94q9A1dO7Hp7B3rnbt4cft7DA5g8WUzo\n4uPZO0dEY48gCKhsrUR+ZT6utZuWAPBSeGFe/DzcFXcXVB6q0d+soQH45BPgyhXT4zExwP33A3Fx\no78HjXmSFihetWqV2QAAsOSJjQiCWJ7kyy/FciXmeuciIsRkbto0wNtb+hjdGedNkaXYVhyvpq0G\n+ZX5qGqtMjmulCsxN24u7o6/Gz5KGwxhaDSAWg2cPCluC9bPx0fsoZs+/Y6frNleSGoWJXbjx483\nySRv3ryJv/3tb/j3f/93uwY3FvT0iCVKvvxS3O5rKKVyoHcuLo69c0Q0dtW216KgqgDlzeUmxxUy\nBWbHzsaChAXw87TBcKggiG/MBw4AnYNKpMhk4hy67Gx+uianNeKh2NOnT2PLli346KOPbB2TTTjz\nUKwgiFt79ffO6XTDr4mMHOidU9lgJIGIyFXVddahoKoAlxtNV6DKZXLMiJ6BhQkLEagKtM3NbtwA\n9u0bvv9iYqI47MqtwMhOHD7HTqfTITg42Gx9O2fgjIldd/dA71xDw/DzSiUwdaqY0MXEsHeOiMa2\nxu5GqKvUOF9/3uS4DDJkRGVgceJiBHsH2+Zm3d3AwYNAUZH46btfQABwzz3i0AnflMmOJJ1jd/Dg\nQZOq3F1dXXj//fcxefLkUQfg7gQBqK4Wk7lLl8z3zkVFAbNmiUmdl5f0MRLnwZDl2Fbsr6WnBYeq\nD+HszbMQYPqHbkrEFGQlZSHMJ8w2NzMYgNOngYICcW5MP4UCmDcPWLQIGMWOEWwvJDWLErvvf//7\nJomdr68vMjMzkZeXZ7fAbMGRO090dQFnz4oJXVPT8POengO9c9HR/CBIRNSuacfh6sMoulEEg2Aw\nOZcelo7spGxE+tlwKLS6WlztOrT8wIQJwH33idXeiexM0p0nXJkjhmIFAaiqGuid0+uHXxMTIyZz\nU6awd46ICAA6+zpxtOYoTl8/DZ3BdFgjJSQF2UnZiA2Itd0N29vFhRHnzpkeDwkRE7rUVNvdi8hC\nkg7FAkBrays+/vhjXL9+HTExMbj//vsRHGyjuQ0urqsLOHNGTOjMbZ3r5SUugpgxQ+ydIyIioFvb\njS+ufoGT105CazCt8ZQYmIic5BwkBiXa7oZ6PXDiBHDokOl+jEolsHAhcPfdYqFQIhdmUY9dfn4+\nVqxYgbS0NCQmJqK6uhqXL1/G3/72NyxZskSKOK1m7x47QRBrVX75pViM3GAYfk1cnNg7N3nyqKZo\nkAQ4D4YsxbYyer26Xpy4dgLHrx6HRq8xORcXEIec5BwkByWbTAEatfJycdh16NyYyZPFxRGBNlpV\nOwTbC1lK0h679evX409/+hMeeeQR47EPPvgATz/9NC4P3QDZzXV0iL1zRUXiHtBDqVRi79zMmVwV\nT0Q0WJ++D4W1hThWcww9uh6Tc1F+UchJzsGEkAm2TehaWsRtwEpKTI9HRADLlgHJyba7F5ETsKjH\nLigoCE1NTVAoFMZjWq0W4eHhaG1ttWuAI2XLHjuDYaB3rqTEfO9cfPxA75xylHtMExG5E51Bh9PX\nT+NI9RF0abtMzoX7hCM7ORsTwybaNqHTaoGjR4Fjx0zLEahUQFaWWGh40N80IkeTfEux7du348c/\n/rHx2O9//3uzW425k/b2gd45c/mrSiXu/Txjhvjhj4iIBugNehTfLMbh6sNo17SbnAvxDkFWUham\nREyBXCa33U0FQVy9tn8/0NZmem76dHErMF9f292PyMlY1GM3f/58FBYWIiIiArGxsaitrUV9fT3m\nzp1r/IQlk8lw+PBhuwdsqZFmvgaDOBXjyy+BsjLzvXOJiWLv3MSJ7J1zF5wHQ5ZiW7kzg2DAV3Vf\nQV2lRmuv6afiQK9ALE5ajIzIDCjkNu4xa2gQd42orDQ9HhsrDrvGxdn2fhZgeyFLSdpj98QTT+CJ\nJ564Y0CurK0NKC4W/w39kAeI2wL2986Fh0sfHxGRsxMEARcaLqCgsgBNPaaLFPw8/bAocRFmRM+A\nh9zGK097ewG1GigsNP007usL5OaKPXUu/jeKyFJjuo6dwSD2yvX3zpm7PClpoHeOq+CJiIYTBAEl\nTSUoqCxAXVedyTkfpQ8WJCzA7JjZUCpsPMQhCGIl+AMHxLpT/eRycQ5ddjY32yaXIXkdu8OHD6O4\nuBhdX//yCIIAmUyGn//856MOQmqtrWLPXFGRuMp1KB8f8QPejBksPE5EdCuCIKCipQL5lfm43nHd\n5JzKQ4W74+/G3Ni58PKwQzX269fFYddr10yPJyWJw64sS0BjlEWJ3YYNG7B3714sXLgQ3t7e9o7J\nLvR6oLRU7J2rqDDfOzdunNg7l5bG3rmxhvNgyFJsK6Kq1irkV+ajpq3G5LinwhN3xd2FeXHz4K20\nw9+Lri7g4EHx0/ngN/KAALEe3eTJTjXsyvZCUrMofdm9ezcuXLiAmJgYe8djUzt25GP27PHo6EhE\ncTHQ2Tn8Gl/fgd65kBDpYyQiciXX2q8hvzIfV1qumBz3kHtgTuwczI+fD19PO6w6NRiA06eB/Hxx\nTl0/hULcMWLhQlaCJ4KFc+ymTZuG/Px8hIWFSRGTTchkMqSlbYZS6Y/Fi/8NYWGm29KMHz/QO8dS\nRkREt3ej4wYKqgpQ2lRqclwhU2BmzEwsTFgIfy9/+9y8ulocdq0znb+H1FRxb1d+KicXplaroVar\nsXXrVpvMsbMosTt16hT++7//G9/97ncROWTewqJFi0YdhD3IZDIsXix+a76++Zg9Owd+fgO9c9zm\nlojozuq76qGuUuNiw0WT43KZHJlRmViUuAhBqiD73Ly9XVwYce6c6fGQEDGhS021z32JHEDSxRNf\nfvkl9u3bhyNHjgybY3f16tVRB2FvwcFyPPooMGECe+fIPM6DIUuNlbbS3NMMdZUa5+rOQcDAHxsZ\nZJgaORWLExcj1MdOq8t0OuDECeDwYaCvb+C4UgksWgTMm+cyE6HHSnsh52HRb8YLL7yAjz76CEuX\nLrV3PDaVmAhERwMJCQakpzs6GiIi59fa24rD1Ydx5uYZGATTCu2TwichKykLEb523GqnrEzc27XJ\ntA4epkwBli4FAgPtd28iN2DRUGxCQgLKy8vh6UITU2UyGTZvFqDRHMSaNSlIS0u885OIiMaoDk0H\njtQcwZfXv4Re0JucSw1NRXZSNqL9o+0XQHOzmNCVms7hQ0QEcP/9YhkTIjdmq6FYixK7t99+G4WF\nhdi4ceOwOXZyuQ33+LMhmUyGHTsOIjd3PJM6IqJb6OrrwrGrx1BYWwidQWdyblzwOGQnZSM+MN5+\nAfT1AUePAseOiXWp+qlUYoHh2bPFgsNEbk7SxO5WyZtMJoNerzd7ztFs9QOisYHzYMhS7tJWerQ9\nOH7tOE5cO4E+fZ/JufiAeOQk5yA5ONl+AQgCcPEi8Nlnpvs4ymTiKrfcXLEelYtzl/ZC9ifp4okr\nV67c+SIiInJ6Gp0GJ2tP4ourX6BX12tyLsY/BjnJORgfPN6++3/X1wOffAJUVpoej40Vh11jY+13\nbyI3Z9VesQaDAXV1dYiMjHTaIdh+7LEjIhqg1Wtx6vopHK05im5tt8m5CN8I5CTnIC00zb4JXW8v\noFYDhYViweF+vr7AkiVAZqZT7RpBJCVJe+za29vx9NNP4/3334dOp4OHhwceffRRvPHGGwjkCiUi\nIqelM+hQdKMIh6sPo7PPdPudUO9QZCdnY3L4ZPsmdIIAnDkDfP65uCVYP7kcmDMHyMoS59QR0ahZ\n1O22YcMGdHV14fz58+ju7jb+d8OGDfaOj0gSarXa0SGQi3CVtqI36FF0owhvnHwD+8r2mSR1Qaog\nPJT+ENbPWY8pEVPsm9TV1gJ//jPwz3+aJnXJycD/+39ioWE3Tupcpb2Q+7Cox+7TTz/FlStX4Pv1\nRNbU1FS8/fbbGDdunF2DIyIi6xgEA87Xn4e6So3mnmaTc/6e/lictBjTo6ZDIbdztfauLuDgQaC4\nWOyx6xcYCNxzDzBpEoddiezAojl2SUlJUKvVSBpUR6iqqgqLFi1CTU2NPeMbMc6xI6KxRBAEXGq8\nhILKAjR0N5ic81X6YmHiQsyMngmlQmnfQAwG4NQpoKBAnFPXT6EA5s8HFiwAXKgmKpFUJJ1j94Mf\n/ABLly7Ff/zHfyAxMRFVVVV47bXX8MQTT4w6ACIiGjlBEFDWXIaCygLc6LxhciArwWEAACAASURB\nVM7bwxvzE+ZjTuwceCokSKaqqoB9+8RVr4OlpQH33ivu8UpEdmVRj53BYMDbb7+N9957Dzdu3EBM\nTAxWrlyJtWvX2nduxiiwx46swVpTZClnaSuCIKCytRL5lfm41n7N5JyXwgvz4ufhrri7oPKQYP5a\ne7tYj+78edPjoaHiHLoJE+wfg5NylvZCzk/SHju5XI61a9di7dq1o76hLRQWFuKZZ56BUqlEbGws\ndu3aBQ8X2RCaiGi0atpqkF+Zj6rWKpPjSrkSc+Pm4u74u+Gj9LF/IDodcPw4cPgwoNUOHPf0BBYt\nAu66C+B7M5GkLOqx27BhA1auXIm7777beOyLL77A3r178frrr9s1QHNu3ryJ4OBgeHl54ec//zlm\nzpyJb33rWybXsMeOiNzN9Y7ryK/MR3lzuclxhUyB2bGzsSBhAfw8/aQJprRU3Nu12XSBBqZOBZYu\nBQICpImDyE1IuqVYWFgYamtr4eXlZTzW29uL+Ph4NDQ03OaZ9rd582ZMnz4dDz30kMlxJnZE5C7q\nOutQUFWAy42XTY7LZXLMiJ6BhQkLEaiSqKZoc7OY0JWWmh6PjBR3jUjk3txEIyH5UKxhcJVwiPPu\nHJ04VVdX48CBA9i0aZND4yDXx3kwZCkp20pjdyPUVWpcqL8AAQPvtzLIkBGVgcWJixHsHSxJLOjr\nA44cAb74Ahi8R7hKBeTkALNmiQWHyQTfW0hqFv0WLliwAC+++KIxudPr9di8eTMWLlxo9Q23b9+O\nWbNmQaVS4fHHHzc519zcjIcffhh+fn5ISkpCXl6e8dxrr72G7OxsvPLKKwDE3TAee+wxvPPOO1Ao\n7FyPiYhIQi09Lfi/y/+HHYU7cL7+vElSNyViCtbPWY+H0h+SJqkTBHFRxPbtYmLXn9TJZMDMmcCG\nDeLuEUzqiJyCRUOxV69exfLly3Hjxg0kJiaipqYG0dHR+PDDDxEfH2/VDf/xj39ALpdj//796Onp\nwVtvvWU8t3LlSgDAm2++ieLiYjzwwAP44osvMGnSJJPX0Ol0+OY3v4nnnnsOOTk55r8xDsUSkYtp\n17TjcPVhFN0ogkEwHSVJD0tHVlIWovyipAuorg745BOxjMlgcXHisGtMjHSxELk5SefYAWIvXWFh\nIa5evYr4+HjMnTsX8lF8Qtu4cSOuXbtmTOy6uroQEhKCCxcuICUlBQCwevVqxMTE4KWXXjJ57rvv\nvotnn30WU6dOBQD88Ic/xCOPPGL6jTGxIyIX0dnXiaM1R3H6+mnoDDqTcykhKchOykZsQKx0AfX2\nigWGT50SCw738/UVF0ZkZHDXCCIbk3SOHQAoFArMmzcP8+bNG/VNAQwLvrS0FB4eHsakDgAyMjLM\n7rO3atUqrFq16o73WLNmjXG3jKCgIGRmZhrnOvS/Lh/zMQC8/vrrbB98bNHj/q9t8Xpz5s/BF1e/\nQN6HedALeiRlJgEAqs5UIdI3Ek99+ykkBiVCrVajDGX2//4WLwaKi6H+4x8BjQZZX79/qqurgYkT\nkbVhA6BSOdX/D2d/bMv2wsfu9bj/66qhPeKjZHGPna0N7bE7cuQIHnnkEdy4MVA5fefOndizZw8K\nCgqsfn322JE11Gq18ZeO6HZs0VZ6db04ce0Ejl89Do1eY3Iu1j8WueNykRyULG0B+NpacdeI2lrT\n4+PGAcuWAeHh0sXiRvjeQpaSvMfO1oYG7+fnh/b2dpNjbW1t8Pf3lzIsGqP4xkuWGk1b6dP3obC2\nEMdqjqFH12NyLsovCjnJOZgQMkHahK6rC/j8c6C42PR4YKC4DdjEiRx2HQW+t5DU7pjYCYKAyspK\nJCQk2HR3h6FvXKmpqdDpdCgvLzcOx549exZTpkwZ8T22bNmCrKws/mIRkUPpDDqcvn4aR6qPoEvb\nZXIu3Ccc2cnZmBg2UdqETq8X59Cp1eKcun4eHsD8+cCCBYBSKV08RGOUWq02GZ4drTsOxQqCAF9f\nX3R2do5qsUQ/vV4PrVaLrVu3ora2Fjt37oSHhwcUCgVWrlwJmUyGP//5zygqKsLy5ctx/PhxTJw4\n0er7cCiWrMHhErKUNW1Fb9Cj+GYxDlcfRrvGdEQixDsEWUlZmBIxBXLZ6N9brVJZKa52ra83PZ6e\nLvbSBUtUG28M4HsLWUqyoViZTIbp06ejpKRkRAnWUNu2bcMvfvEL4+Pdu3djy5Yt2LRpE373u99h\n7dq1iIiIQFhYGP7whz/Y5J5ERFIyCAZ8VfcV1FVqtPa2mpwL9ArE4qTFyIjMgEIucQ3Otjbgs8+A\nCxdMj4eGivPoBi1eIyLXZNHiiRdffBG7d+/GmjVrEB8fb8wqZTIZ1q5dK0WcVmOPHRFJTRAEXGi4\nAHWVGo3djSbn/Dz9sChxEWZEz4CHXOLpzTqduGPEkSOAVjtw3NMTWLwYuOsugIXeiRxK0sUTR48e\nRVJSEg4dOjTsnLMmdgDn2BGRNARBQElTCQoqC1DXVWdyzkfpgwUJCzA7ZjaUConnrAmCuKfrp58C\nLS2m56ZNE2vScYEakUNJPsfOVbHHjqzBeTBkqcFtRRAEVLRUIL8yH9c7rptcp/JQ4e74uzE3di68\nPLykD7SpSUzoyspMj0dFicOuiYnSxzQG8b2FLCV5uZOmpiZ8/PHHuHnzJn7605+itrYWgiAgLi5u\n1EEQEbmaqtYq5Ffmo6atxuS4p8ITd8XdhXlx8+Ct9JY+sL4+4PBh4PjxgX1dAcDbG8jJEfd3tcFC\nOCJyThb12B06dAjf+ta3MGvWLBw7dgwdHR1Qq9V45ZVX8OGHH0oRp9XYY0dEtlRSXoLPv/wcDd0N\nuNJ8BQHRAQiLCTOe95B7YE7sHMyPnw9fT1/pAxQE4Px54MABYHBNUJlMTOZycgAfH+njIiKLSLpX\nbGZmJl5++WUsWbIEwcHBaGlpQW9vLxISElA/dLm8k2BiR0S2cvL8SezYvwNtMW1o07QBAHTlOmRO\nykRkbCRmxszEwoSF8Pdy0Hy1ujpx14jqatPj8fHA/fcD0dGOiYuILCbpUGx1dTWWLFlickypVEI/\nuJvfCXHxBFmK82BoMINgQG17LUqaSlDSWIKPP/sY3XHdgAZovdyKoPQgKFOU0DXpsGHFBgSpghwT\naE8PUFAgFhoe/AfBz09cGDFtGneNcDC+t9Cd2HrxhEWJ3cSJE/Hpp5/ivvvuMx47ePAgpk6darNA\n7GHLli2ODoGIXESfvg8VzRUoaSpBaVMpurXdxnMGGEyujfSNRGJQImKaYhyT1BkM4hZgBw8C3QNx\nQi4XS5csXgx4OWDBBhFZrb8DauvWrTZ5PYsSu1dffRXLly/H/fffj97eXqxbtw4ffvgh/vnPf9ok\nCCJH4yfqsald047SplKUNJagsrUSOoPO7HVKKOHr7YswnzCELg2Fp8ITAOAp95QyXNG1a+Kw63XT\nVbgYN05c7RoeLn1MdEt8byGpWVzupLa2Frt370Z1dTUSEhLwve99z6lXxHKOHRENJQgC6rrqUNJY\ngpKmkmElSgbz8/RDWmga0sLS0NfYh/cOvwevCQO9YJoyDdZkr0FaSpoUoQOdncDnnwNnzpgeDwoS\ntwFLT+ewK5ELk3TxRD+DwYDGxkaEh4dLu1n1CDCxI2twHoz70hl0qG6tNs6X61/8YE6kbyTSwtKQ\nFpqGGP8Yk/e5kvISHCw6iIvnL2LSlEnInZErTVKn1wOFhYBaDWg0A8c9PIAFC4D58wGlxIWPyWJ8\nbyFLSbp4oqWlBT/60Y+wd+9eaLVaKJVKfPvb38b//u//IiQkZNRB2AsXTxCNTd3abpQ1laGkqQQV\nzRXQ6DVmr5PL5EgKSjL2zN1uvlxaShrSUtKgjpDwD/WVK8AnnwANDabH09PFXrrgYGniICK7ccjO\nEw899BA8PDywbds2JCQkoKamBps2bUJfX5/TzrNjjx3R2NLU3WTslatpq4EA87//Kg8VJoRMQFpY\nGlJCUqDyUEkcqQXa2oD9+4GLF02Ph4WJ8+jGj3dMXERkN5IOxQYGBuLGjRvwGVTcsru7G9HR0Whr\nu/WwhiMxsSNybwbBgGvt14zz5Rq7G295bbAqGOlh6UgLS0N8QDwUcifd8F6nA44dA44eBbTageOe\nnkBWFjB3LqBw0tiJaFQkHYpNT09HVVUVJk2aZDxWXV2N9PT0UQdA5Aw4D8Y13K4kyWAyyBAXEGec\nLxfmE2azecF2aSuCAJSWinu7trSYnsvIAJYsAfwdVPyYRoXvLSQ1ixK7nJwc3HPPPXjssccQHx+P\nmpoa7N69G6tWrcJf/vIXCIIAmUyGtWvX2jteIhpjBpckudJyBXrBfGF0pVyJ8SHjkRaahgmhE+Dn\n6SdxpCPU1CTOoysvNz0eFSXuGpGQ4Ji4iMglWTQU2/9pY/An3v5kbrCCggLbRjcKMpkMmzdv5uIJ\nIhcjCAJudt40zpe70Xnjltf6e/ojNTQVaWFpSA5KhlLhQqtDNRrg8GHgxAlx5Ws/b28gNxeYMUMs\nOExEbq1/8cTWrVulL3fiSjjHjsh16Aw6VLVWGefLtWvab3ltlF+UmMyZKUniEgQBOHcOOHAA6OgY\nOC6TATNnAjk5wKD5zEQ0Nkg6x47I3XEejPQGlyQpby5Hn77P7HUKmUIsSRKWhtTQVMfty/q1UbWV\nmzfFYdfqatPjCQniatfo6FHHR86F7y0kNSZ2RCQZS0uSeHt4Y0LoBKSFpmF8yHjnLElijZ4eID8f\nOH1a7LHr5+cH3HMPMHUqd40gIpvgUCwR2Y1BMOBq21XjKtbblSQJ8Q4xFgpOCEyAXOYG88sMBqCo\nSEzquget4JXLgbvuAhYvBry8bv18IhozOBRLRE5Jo9OgoqUCJY0lKGsuc0hJEqdw9Sqwbx9wY8ji\nj/HjxWHXsDDHxEVEbs3ixO7SpUv44IMPUFdXhx07duDy5cvo6+vDtGnT7BkfkSQ4D2Z02jXtxoUP\nlS2Vty1JkhKSgtTQVKSGpsLX01fiSEfvjm2ls1NcGHH2rOnxoCDgvvuAtDQOu44hfG8hqVmU2H3w\nwQd46qmnsGLFCuzZswc7duxAR0cHnn/+eXz++ef2jpGInIy1JUn6e+WSg5PhIXfTgQK9HigsBNRq\nsZRJPw8PYOFC4O67AaULlWMhIpdk0Ry79PR0vP/++8jMzERwcDBaWlqg1WoRHR2NxsZbz5lxJNax\nI7Ita0uS9M+Xi/aLdq8hVnOuXBFXuzY0mB6fOBG4916xt46IyAyH1LELDQ1FQ0MD5HK5SWIXGxuL\n+vr6UQdhD1w8QTR63dpu464PFS0VLlOSRDKtrcD+/cClS6bHw8LEXSPGjXNMXETkciRdPDFjxgy8\n++67WL16tfHYX//6V8yZM2fUARA5A86DGdDY3WjslbvadtWikiQpISnw8hgbqzvVajWy5s8Hjh0D\njh4FdLqBk15e4krXuXMBhcJxQZLT4HsLSc2ixO6NN97A0qVL8eabb6K7uxv33HMPSktL8dlnn9k7\nPiKys8ElSUoaS9DU03TLa0O8Q5Aelo7U0FT3KUlioeqSElQcOICvvvgChu3bMT4qComDV7ZmZABL\nlgD+/o4LkojGPIvr2HV1deGjjz5CdXU1EhIS8MADD8Dfid/AOBRLdGuDS5KUNpWiR9dj9joZZIgP\njDfOlwv1DnX/+XJmVJ8/j/LXXkNuYyPQ1gYAOKjTISUzE4lTp4rDrvHxDo6SiFyZrfIWFigmGiPa\netuMvXJVrVW3LEniqfDE+ODxSAtLw4SQCS5ZksRmbtwAioqQ/8c/Iqd9yGIRpRL5M2Yg56WXxILD\nRESjIOkcu+rqamzduhXFxcXo7Ow0CaK0tHTUQRA5mjvOgxEEATc6bxjny93svHnLa8dMSRJL9PYC\n586JO0Z8XVxY3jewaETd2oqsSZOA5GTIw8OZ1NFtueN7Czk3i969v/3tb2PixInYtm0bVCoX37OR\nyI3pDDpUtlQat/BiSRILCQJQXQ0UFwMXLpguiABgkMsBb28gKkqsUZeaKh739HREtEREt2TRUGxg\nYCCam5uhcKFVXhyKpbGiq68LZc1lFpUkSQ5ORlqoWJIkUBUocaROqLMTOHNGTOiazCwa8fAAJk1C\ndUAAyg8eRO6gfV0PajRIWbMGiWlpEgZMRO5K0qHY5cuX49ChQ8jJyRn1DaW0ZcsWFigmtyMIApp6\nmiwuSZIamoq0sDSMDx4/ZkqS3JbBAJSXi0OtpaXi46GiooAZM4CpUwFvbyQCQHw88g8ehLyvDwZP\nT6Tk5jKpI6JR6y9QbCsW9dg1NjZi3rx5SE1NRURExMCTZTL85S9/sVkwtsQeO7KGs8+DMQgG1LTV\nGJO55p7mW14b6h1qnC8XHxg/pkqS3FZzs9gzd+YM0NEx/LyXl5jIzZgBREffcj9XZ28r5FzYXshS\nkvbYrV27Fp6enpg4cSJUKpXx5mN6Tg6RnWl0GpQ3l6OkqQRlTWUWlyQJ8wkze92YpNOJu0IUFQGV\nleavSUwUk7lJk7iXKxG5PIt67Pz9/VFbW4uAgAApYrIJ9tiRK2rtbTVu4cWSJKNw86bYO/fVV0CP\nmYTY1xfIzASmTxe3/yIicjBJe+ymTZuGpqYml0rsiFyBNSVJArwCjL1ySUFJY7skiTkazUCZkuvX\nh5+XyYAJE8RkLjWVW34RkVuy6C9DTk4O7r33Xjz++OOIjIwEAONQ7Nq1a+0aIJEUpJwHo9VrUdla\nadz1oaPPzHyvr0X7RRvny0X5RXH6w1CCAFy9KiZzFy4AWu3wa4KCxKHWzEzABh9OOWeKrMH2QlKz\nKLE7cuQIYmJizO4Ny8SO6M66+rrEIdamElQ0V0BrMJOAgCVJLNbZCZw9Kw63NjYOP69QABMnigld\ncvItF0IQEbkbbilGZAeCIKCxu9G4hde19mu3LEnio/TBhJAJLElyJwYDUFEh9s6VlJgvUxIRAcyc\nKa5u9fGRPkYiohGy+xy7wateDebeQL8m53Y6RACsK0kS5hMm1pdjSZI7a20Ve+aKi4Gh+7UCgKfn\nQJmSmBj2zhHRmHbLxC4gIAAdX9d68vAwf5lMJoNeb37VHpErGek8mF5dLyqaKywqSZIQmIC0MHGI\nlSVJ7kCnAy5fHihTYu5TbEKCuBBi8mQxuZMI50yRNdheSGq3TOwuXLhg/PrKlSuSBEPkClp7W429\nctWt1bctSZISkoK00DRMCJ0AHyWHBu+ovl5M5s6eNV+mxMdnoExJeLj08REROTmL5ti9/PLLeO65\n54Ydf/XVV/GTn/zELoGNFufYka0IgoDrHdeN8+XquupueS1LkoyARiOuaC0qAq5dG35eJgPGjxeH\nWtPSWKaEiNySrfIWiwsUd5jZgic4OBgtLS2jDsIeZDIZtr+/HUtmLkFaCvdzJOsMLklS0lSCzr7O\nW17LkiQjIAhiEtdfpqSvb/g1gYEDZUoCuTqYiNybJAWK8/PzIQgC9Ho98vPzTc5VVFQ4fcHiXf/a\nhf3H9uNHa36ElHEpAMS5Tv1/ePu/luHrx2a+HnrtSJ830nuQfZWUl+DzLz/HpQuXkJyWjORxyejz\n77tjSZJxweOM8+UCvJz798CpdHWJu0EUFQENDcPPKxRAevpAmRInXJzFOVNkDbYXuhO1Wg21Wm2z\n17ttj11SUhJkMhlqamqQkJAw8CSZDJGRkXj++efxzW9+02bB2JJMJsPitxYDAHyv+WL2gtkOjmhk\npEg6pX6es8R2tfoq9p/eD4/xHigvKofneE/oynXInJSJsBjTxQ0+Sh/jKtZxweNYksQaBgNw5cpA\nmRJzC67Cw8Vkbto0cbsvJ8Y/1GQNtheylKRDsatWrcK777476ptJaXBip7qmwl0L7nJwRORsCo8W\nojuue9jx/g8CYT5hxvlycQFxLElirdZW4MwZsUxJW9vw856ewJQpYkIXG8syJUQ0pkm6V6yrJXX9\nglXBECDA39sfSUFJEAQBAgTjD87c1/1FZM19LfXzyL4MGF6fMdArEEmhSdgwZwNCfUIdEJWL0+vF\nXrmiIrGYsLk3qbg4MZmbPBnwYs8nEZEtufWSvYyoDGjKNFjz4BqXXEDhKgmoqz6v5WIL2oLaxK8v\nt2DqnKlQKpSIUEQwqbNWQ8NAmZLu4b2g8PEBMjLEMiUREdLHZ0McWiNrsL2Q1Nw6sYuoj0Budq5L\nJnUATOaKgaNUNhd1TxTeLngbXhO8IPOWQalQQlOmQW52rqNDcw19fQNlSq5eHX5eJgPGjRsoU3KL\nQudERGQ73CuWxrSS8hIcLDqIPkMfPOWeyJ3huh8EJCEIQG2tmMydP3/rMiX9RYSDgqSPkYjIBUm6\neMIVMbEjsqHu7oEyJfX1w8/L5QNlSsaNc8oyJUREzkzSxRNE7o7zYMwQBHGf1qIi4NIl82VKwsLE\nZC4jw+nLlNgK2wpZg+2FpMbEjohMtbeLJUqKi8WSJUMplQNlSuLiWKaEiMiJcCiWiMTeuNJSsXeu\nvNx8mZLYWDGZmzKFZUqIiGyMQ7FENHqNjWLP3Jkz4nZfQ3l7i7tBzJgBREZKHx8REVmFiR0Rxtg8\nmL4+4OJFsXeupsb8Nf1lStLTWaZkiDHVVmjU2F5IanzHJhoLBAG4cUNM5s6dAzSa4dcEBAyUKQkO\nlj5GIiIaNc6xI3JnPT1imZLiYuDmzeHn5XKxePCMGcD48SxTQkTkIJxjR0TmCQJQVTVQpkSnG35N\naOhAmRI/P8lDJCIi+3C5xK6urg4rVqyAp6cnPD09sWfPHoSGcl9PGh23mAfT0SEugigqAlpahp9X\nKoFJk8SELiGBZUpGyC3aCkmG7YWk5nKJXXh4OI4dOwYAeOedd7Bz50787Gc/c3BURA6i1wNlZWIy\nV1ZmvkxJTMxAmRKVSvoYiYhIMi49x+6NN96Ap6cnnnzyyWHnOMeO3FpT00CZks7O4edVqoEyJVFR\n0sdHRERWGdNz7M6ePYt169ahtbUVp06dcnQ4RNLQasUyJcXF4hw6c5KTxVWtEyeKQ69ERDSmSLoE\nbvv27Zg1axZUKhUef/xxk3PNzc14+OGH4efnh6SkJOTl5RnPvfbaa8jOzsYrr7wCAMjIyMDJkyfx\ny1/+Etu2bZPyWyA3pVarHR3Crd24AXz8MfDKK8A//jE8qfP3BxYuBH70I2D1arGnjkmd3Th1WyGn\nw/ZCUpO0xy42NhYbN27E/v370dPTY3Ju/fr1UKlUqK+vR3FxMR544AFkZGRg0qRJePbZZ/Hss88C\nALRaLZRf/9EKCAiAxlw9LiJX19sr1psrKhITu6HkcmDCBHGodcIElikhIiIADppjt3HjRly7dg1v\nvfUWAKCrqwshISG4cOECUlJSAACrV69GTEwMXnrpJZPnnjp1Cs899xwUCgWUSiXefPNNxMXFDbuH\nTCbD6tWrkZSUBAAICgpCZmamcXVS/6coPuZjp3ksCMhKTgaKiqD++GNAr0fW1+1X/XUvXdaMGcCM\nGVC3twM+Ps4VPx/zMR/zMR9b/Lj/66qv39/feecdm8yxc0hi9+KLL6K2ttaY2BUXF2PBggXoGrRX\n5auvvgq1Wo1//etfI7oHF0+Qy+joAM6eFefONTUNP+/hMVCmJDGRZUqIiNyQSy+ekA35w9TZ2YmA\ngACTY/7+/ujo6JAyLBrD1Gq18dOUJAwGsTxJcTFQWio+HioqSkzmpk4FvL2li41uS/K2Qi6N7YWk\n5pDEbmhG6ufnh/b2dpNjbW1t8Pf3lzIsIvtrbh4oU2Lug4tKJSZyM2YA0dHSx0dERC7NKXrsUlNT\nodPpUF5ebpxjd/bsWUyZMmVU99myZQuysrL4aYnuyK5tRKcTt/YqKgIqK81fk5goJnOTJnFFq5Pj\n+wlZg+2F7kStVpvMuxstSefY6fV6aLVabN26FbW1tdi5cyc8PDygUCiwcuVKyGQy/PnPf0ZRURGW\nL1+O48ePY+LEiSO6F+fYkcPdvCkmc+fOAUNWgQMQ92jNzBTrznFbPCKiMc1WeYukid2WLVvwi1/8\nYtixTZs2oaWlBWvXrsWBAwcQFhaGX//613j00UdHfC8mdmQNm82D6e0Fzp8XE7rr14efl8lMy5Qo\nFKO/J0mKc6bIGmwvZCmXXDyxZcsWbNmyxey54OBg/OMf/5AyHCLbEASgpkacO3fhgrhDxFDBwWLP\nXGYmMGShEBERka249F6xtyOTybB582bOsSP76ewUy5QUFd26TMnEiWLvXFISy5QQEdEw/XPstm7d\n6npDsVLiUCzZhcEAVFSIyVxJifkyJZGRYjI3bRrLlBARkUVcciiWyFndcR5MS8tAmZIhpXkAAF5e\nYpmS6dOBmBj2zrkxzpkia7C9kNSY2BHdik4HXL4s9s5duWL+moSEgTIlnp7SxkdERDSEWw/Fco4d\njUhdndg7d/as+TIlvr5ARoaY0IWFSR8fERG5Dc6xsxDn2JElqktKUPH555B3d8PQ2Ijxfn5I1OuH\nXyiTASkpYjKXmsoyJUREZFOcY0c0UoIAdHSg+uRJlO/ejdzeXqjLypATEICDOh2QmYnE/p64oKCB\nMiWBgY6Nm5wC50yRNdheSGpM7Mh9GQzioofGRqChQfzX2Cj+02hQUViI3O7ugWsB5Hp4IL+qColZ\nWWJCN24cF0IQEZHL4FAsuT6dTqwjNzh5a2gQj5kbVv2a+sQJZPX2Dhzw9QWio6FOTUXWT38qQeBE\nREQiDsVaYMuWLVw84U40GtPErf/rlhZxeNUa3t4whIaKPXU+PuIwq78/IJPB4Odnn/iJiIiG6F88\nYSvssSPnIghAV9dA8jY4ievosP71/P2B8HBx9erg//r6orq0FOVvv41cLy+oq6qQlZSEgxoNUtas\nQWJamu2/N3ILnDNF1mB7IUuxx45cmyAAbW2myVv/f82VGLkdmUxc5BAePjyJU6lu+bTEtDRgzRrk\nHzyIrxobYYiIQEpuLpM6IiJyWeyxI/vS68Wh0qHz3xobAa3WutdSKIDQoELHbgAAETtJREFU0IHE\nrT95Cw0FlEr7xE9ERCQB9tiRc9FqB1acDk7imprM76d6O56ew4dOw8OB4GBALrdP/ERERG6AiR1Z\np6dnePLW0CAOq1r7ScPHx/z8t4AAyUuMcB4MWYpthazB9kJSc+vEjqtiR0gQgM5O8ytQOzutf73A\nwOHJW1iYWF6EiIhoDOOqWAtxjp0FDAagtXX44oXGRmBwfTdLyGRASMjw5C0sDPDysk/8REREboJz\n7MhyOh3Q3Dw8eWtsFM9Zw8NDXKwwdAVqSIh4joiIiByGf4ndSV+f+flvLS3WL2Dw8hq+eCEsTCwr\n4oYLGDgPhizFtkLWYHshqTGxc0Xd3ebnv7W1Wf9afn7mV6D6+XGPVCIiIhfDOXbOShCA9nbzOzD0\nb1xvjf4CvkOTOG9v28dOREREVuEcO3dhMAwU8B3aA9fXZ91ryeWmBXz7/xsaKtaGIyIiIrfm1omd\nU5U70elMC/j2/7epSdydwRpKpfnyISEh4u4MZDXOgyFLsa2QNdhe6E5sXe7E7RM7yfX2mi8f0tJi\nfQFfb+/hixfCw8W6cJz/RkRE5PL6O6C2bt1qk9fjHLuREASgq8v8CtSODutfz9/f/ApUX18mcERE\nRGMA59hJQRDElabm5r/19Fj3WjKZuNepuSFUlco+8RMREdGYwsQOEOe4NTebH0LVaq17LYVioIDv\n4OQtNFScG0dOifNgyFJsK2QNtheSmlsndvk7dmD8kiVITEsTD2i15pO3pibrC/h6epovHxIc7JYF\nfImIiMj5ufccu3XrcLCtDSkzZyJRqRSHVa39dn18hi9eCAsDAgI4/42IiIhswlZz7Nw7sVu8GACQ\n7+uLnNmzb/+EwEDzK1B9fCSIloiIiMYyLp6wgry/TpxcLg6VmluBygK+YxrnwZCl2FbIGmwvJDW3\nTuy2NDQgKz4ehlmzgKeeEgv4erj1t0xEREQuxNYFit17KHbzZhzUaJCyZs3AAgoiIiIiJ8OhWAvk\nR0QgJTeXSR0RERGNCW5dlyPnqaeY1JFFbNkNTu6NbYWswfZCUnPrxI6IiIhoLHHvOXbu+a0RERGR\nm7FV3sIeOyIiIiI3wcSOCJwHQ5ZjWyFrsL2Q1JjYEREREbkJzrEjIiIicjDOsSMiIiIiE0zsiMB5\nMGQ5thWyBtsLSY2JHREREZGbcOs5dps3b0ZWVhaysrIcHQ4RERHRMGq1Gmq1Glu3brXJHDu3Tuzc\n9FsjIiIiN8PFE0Q2xHkwZCm2FbIG2wtJjYkdERERkZvgUCwRERGRg3EoloiIiIhMMLEjAufBkOXY\nVsgabC8kNSZ2RERERG6Cc+yIiIiIHIxz7IiIiIjIBBM7InAeDFmObYWswfZCUmNiR0REROQmOMeO\niIiIyME4x46IiIiITDCxIwLnwZDl2FbIGmwvJDUmdkRERERuwmXn2OXl5eHHP/4x6uvrzZ7nHDsi\nIiJyFWN6jp1er8cHH3yAhIQER4dCRERE5DRcMrHLy8vDI488AplM5uhQyE1wHgxZim2FrMH2QlJz\nucSuv7fuO9/5jqNDITdy5swZR4dALoJthazB9kJSkzSx2759O2bNmgWVSoXHH3/c5FxzczMefvhh\n+Pn5ISkpCXl5ecZzr776KrKzs/Hyyy/jvffeY28d2Vxra6ujQyAXwbZC1mB7IalJmtjFxsZi48aN\nWLt27bBz69evh0qlQn19Pd577z388Ic/xMWLFwEAP/nJT1BQUIDnnnsOFy9exK5du7Bs2TKUlZXh\nmWeekfJbsDkpuultcY+RvoY1z7Pk2jtdc7vz7jAkYu/vwVavP5LXsXVbseQ6d24vfG+x7tqx3FYA\nvrdYe60ztxdJE7uHH34YDz74IEJDQ02Od3V14e9//zu2bdsGHx8fzJ8/Hw8++CDefffdYa/x61//\nGvv378cnn3yC1NRUvP7661KFbxd887XuWnv9MlVVVd3x3s6Ab77WXWuP9sK2Ytt78L3FOfC9xbpr\nnTmxc0i5kxdffBG1tbV46623AADFxcVYsGABurq6jNe8+uqrUKvV+Ne//jWie6SkpKCiosIm8RIR\nERHZ0/jx41FeXj7q1/GwQSxWGzo/rrOzEwEBASbH/P390dHRMeJ72OKHQ0RERORKHLIqdmgnoZ+f\nH9rb202OtbW1wd/fX8qwiIiIiFyaQxK7oT12qamp0Ol0Jr1sZ8+exZQpU6QOjYiIiMhlSZrY6fV6\n9Pb2QqfTQa/XQ6PRQK/Xw9fXFytWrMCmTZvQ3d2No0eP4sMPP8SqVaukDI+IiIjIpUma2PWvev3N\nb36D3bt3w9vbG7/61a8AAL/73e/Q09ODiIgIfO9738Mf/vAHTJw4UcrwiIiIiFyaQ1bFOkp7ezuW\nLFmCS5cu4eTJk5g0aZKjQyInVlhYiGeeeQZKpRKxsbHYtWsXPDwcst6InFxdXR1WrFgBT09PeHp6\nYs+ePcPKOhENlZeXhx//+Meor693dCjkpKqqqjB79mxMmTIFMpkMe/fuRVhY2G2f43Jbio2Gj48P\n9u3bh3/7t38btoCDaKiEhAQUFBTg0KFDSEpKwj//+U9Hh0ROKjw8HMeOHUNBQQG++93vYufOnY4O\niZxc//aYCQkJjg6FnFxWVhYKCgqQn59/x6QOGGOJnYeHh0U/FCIAiIqKgpeXFwBAqVRCoVA4OCJy\nVnL5wFtpe3s7goODHRgNuYK8vDxuj0kWOXbsGBYtWoQXXnjBouvHVGJHNBLV1dU4cOAAvvGNbzg6\nFHJiZ8+exdy5c7F9+3asXLnS0eGQE+vvrfvOd77j6FDIycXExKCiogKHDx9GfX09/v73v9/xOS6Z\n2G3fvh2zZs2CSqXC448/bnKuubkZDz/8MPz8/JCUlIS8vDyzr8FPSWPHaNpLe3s7HnvsMbzzzjvs\nsRsDRtNWMjIycPLkSfzyl7/Etm3bpAybHGSk7WX37t3srRtjRtpWPD094e3tDQBYsWIFzp49e8d7\nueRM8NjYWGzcuBH79+9HT0+Pybn169dDpVKhvr4excXFeOCBB5CRkTFsoQTn2I0dI20vOp0Ojz76\nKDZv3owJEyY4KHqS0kjbilarhVKpBAAEBARAo9E4InyS2Ejby6VLl1BcXIzdu3ejrKwMzzzzjMvv\ne063N9K20tnZCT8/PwDA4cOHMXny5DvfTHBhL774orBmzRrj487OTsHT01MoKyszHnvssceEn/3s\nZ8bHy5YtE2JiYoR58+YJb7/9tqTxkmNZ21527dolhIaGCllZWUJWVpbw17/+VfKYyTGsbSsnT54U\nFi1aJGRnZwv33HOPcPXqVcljJscZyd+ifrNnz5YkRnIO1raVffv2CTNnzhQWLlworF69WtDr9Xe8\nh0v22PUThvS6lZaWwsPDAykpKcZjGRkZUKvVxsf79u2TKjxyMta2l1WrVrFI9hhlbVuZM2cODh06\nJGWI5ERG8reoX2Fhob3DIydibVtZtmwZli1bZtU9XHKOXb+h8xM6OzsREBBgcszf3x8dHR1ShkVO\niu2FLMW2QtZgeyFLSdFWXDqxG5r5+vn5ob293eRYW1sb/P39pQyLnBTbC1mKbYWswfZClpKirbh0\nYjc0801NTYVOp0N5ebnx2NmzZzFlyhSpQyMnxPZClmJbIWuwvZClpGgrLpnY6fV69Pb2QqfTQa/X\nQ6PRQK/Xw9fXFytWrMCmTZvQ3d2No0eP4sMPP+Q8qTGO7YUsxbZC1mB7IUtJ2lZstdJDSps3bxZk\nMpnJv61btwqCIAjNzc3CQw89JPj6+gqJiYlCXl6eg6MlR2N7IUuxrZA12F7IUlK2FZkgsKAbERER\nkTtwyaFYIiIiIhqOiR0RERGRm2BiR0REROQmmNgRERERuQkmdkRERERugokdERERkZtgYkdERETk\nJpjYEREREbkJJnZEREOsWbMGGzdutOlr/vCHP8Qvf/lLm74mEdFQHo4OgIjI2chksmGbdY/W73//\ne5u+HhGROeyxIyIyg7stEpErYmJHRE7lN7/5DeLi4hAQEID09HTk5+cDAAoLCzFv3jwEBwcjJiYG\nGzZsgFarNT5PLpfj97//PSZMmICAgABs2rQJFRUVmDdvHoKCgvDoo48ar1er1YiLi8NLL72E8PBw\nJCcnY8+ePbeM6aOPPkJmZiaCg4Mxf/58nDt37pbXPvvss4iMjERgYCCmTZuGixcvAjAd3v3GN74B\nf39/4z+FQoFdu3YBAC5fvoylS5ciNDQU6enp+OCDD255r6ysLGzatAkLFixAQEAA7r33XjQ1NVn4\nkyYid8TEjoicRklJCXbs2IHTp0+jvb0dn332GZKSkgAAHh4e+O1vf4umpiYcP34cBw8exO9+9zuT\n53/22WcoLi7GiRMn8Jvf/AZPPPEE8vLyUFNTg3PnziEvL894bV1dHZqamnD9+nW88847WLduHcrK\nyobFVFxcjO9///vYuXMnmpub8eSTT+Kb3/wm+vr6hl27f/9+HDlyBGVlZWhra8MHH3yAkJAQAKbD\nux9++CE6OjrQ0dGBvXv3Ijo6Grm5uejq6sLSpUvxve99Dw0NDXj//ffx1FNP4dKlS7f8meXl5eHt\nt99GfX09+vr68PLLL1v9cyci98HEjoichkKhgEajwYULF6DVapGQkIBx48YBAGbMmIE5c+ZALpcj\nMTER69atw6FDh0ye/9Of/hR+fn6YNGkSpk6dimXLliEpKQkBAQFYtmwZiouLTa7ftm0blEolFi1a\nhAceeAB//etfjef6k7A//elPePLJJzF79mzIZDI89thj8PLywokTJ4bF7+npiY6ODly6dAkGgwFp\naWmIiooynh86vFtaWoo1a9Zg7969iI2NxUcffYTk5GSsXr0acrkcmZmZWLFixS177WQyGR5//HGk\npKRApVLhkUcewZkzZ6z4iRORu2FiR0ROIyUlBa+//jq2bNmCyMhIrFy5Ejdu3AAgJkHLly9HdHQ0\nAgMD8cILLwwbdoyMjDR+7e3tbfJYpVKhs7PT+Dg4OBje3t7Gx4mJicZ7DVZdXY1XXnkFwcHBxn/X\nrl0ze212djaefvpprF+/HpGRkXjyySfR0dFh9ntta2vDgw8+iF/96le4++67jfc6efKkyb327NmD\nurq6W/7MBieO3t7eJt8jEY09TOyIyKmsXLkSR44cQXV1NWQyGf7rv/4LgFguZNKkSSgvL0dbWxt+\n9atfwWAwWPy6Q1e5trS0oLu72/i4uroaMTExw56XkJCAF154AS0tLcZ/nZ2d+M53vmP2Phs2bMDp\n06dx8eJFlJaW4n/+53+GXWMwGPDd734Xubm5+MEPfmByr8WLF5vcq6OjAzt27LD4+ySisY2JHRE5\njdLSUuTn50Oj0cDLywsqlQoKhQIA0NnZCX9/f/j4+ODy5csWlQ8ZPPRpbpXr5s2bodVqceTIEXz8\n8cf49re/bby2//onnngCf/jDH1BYWAhBENDV1YWPP/7YbM/Y6dOncfLkSWi1Wvj4+JjEP/j+L7zw\nArq7u/H666+bPH/58uUoLS3F7t27odVqodVqcerUKVy+fNmi75GIiIkdETkNjUaD559/HuHh4YiO\njkZjYyNeeuklAMDLL7+MPXv2ICAgAOvWrcOjjz5q0gtnru7c0PODH0dFRRlX2K5atQp//OMfkZqa\nOuzamTNnYufOnXj66acREhKCCRMmGFewDtXe3o5169YhJCQESUlJCAsLw3/+538Oe83333/fOOTa\nvzI2Ly8Pfn5++Oyzz/D+++8jNjYW0dHReP75580u1LDkeySisUcm8OMeEY0xarUaq1atwtWrVx0d\nChGRTbHHjoiIiMhNMLEjojGJQ5ZE5I44FEtERETkJthjR0REROQmmNgRERERuQkmdkRERERugokd\nERERkZtgYkdERETkJv4/dDERlOXTSFkAAAAASUVORK5CYII=\n", + "text": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ "\n", "## Creating lists using conditional statements\n" ] @@ -736,31 +1282,24 @@ "input": [ "import timeit\n", "\n", - "def cond_loop():\n", + "def cond_loop(n):\n", " even_nums = []\n", - " for i in range(100):\n", + " for i in range(n):\n", " if i % 2 == 0:\n", " even_nums.append(i)\n", " return even_nums\n", "\n", - "def list_compr():\n", - " even_nums = [i for i in range(100) if i % 2 == 0]\n", + "def list_compr(n):\n", + " even_nums = [i for i in range(n) if i % 2 == 0]\n", " return even_nums\n", " \n", - "def filter_func():\n", - " even_nums = list(filter((lambda x: x % 2 != 0), range(100)))\n", + "def filter_func(n):\n", + " even_nums = list(filter((lambda x: x % 2 != 0), range(n)))\n", " return even_nums\n", "\n", - "%timeit cond_loop()\n", - "%timeit list_compr()\n", - "%timeit filter_func()\n", - "\n", - "#\n", - "# Python 3.4.0\n", - "# MacOS X 10.9.2\n", - "# 2.5 GHz Intel Core i5\n", - "# 4 GB 1600 Mhz DDR3\n", - "#" + "%timeit cond_loop(n)\n", + "%timeit list_compr(n)\n", + "%timeit filter_func(n)" ], "language": "python", "metadata": {}, @@ -769,8 +1308,8 @@ "output_type": "stream", "stream": "stdout", "text": [ - "100000 loops, best of 3: 14.4 \u00b5s per loop\n", - "100000 loops, best of 3: 12 \u00b5s per loop" + "10 loops, best of 3: 18.1 ms per loop\n", + "100 loops, best of 3: 15.3 ms per loop" ] }, { @@ -778,7 +1317,7 @@ "stream": "stdout", "text": [ "\n", - "10000 loops, best of 3: 23.9 \u00b5s per loop" + "10 loops, best of 3: 22.8 ms per loop" ] }, { @@ -789,7 +1328,91 @@ ] } ], - "prompt_number": 14 + "prompt_number": 119 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "funcs = ['cond_loop', 'list_compr',\n", + " 'filter_func']\n", + "\n", + "orders_n = [10**n for n in range(1, 6)]\n", + "times_n = {f:[] for f in funcs}\n", + "\n", + "for n in orders_n:\n", + " test_list = list([i for i in range(n)])\n", + " for f in funcs:\n", + " times_n[f].append(min(timeit.Timer('%s(n)' %f, \n", + " 'from __main__ import %s, n' %f)\n", + " .repeat(repeat=3, number=1000)))" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 120 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%pylab inline" + ], + "language": "python", + "metadata": {}, + "outputs": [] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "import matplotlib.pyplot as plt\n", + "\n", + "labels = [('cond_loop', 'explicit for-loop'), \n", + " ('list_compr', 'list comprehension'),\n", + " ('filter_func', 'lambda function'),\n", + " ] \n", + "\n", + "matplotlib.rcParams.update({'font.size': 12})\n", + "\n", + "fig = plt.figure(figsize=(10,8))\n", + "for lb in labels:\n", + " plt.plot(orders_n, times_n[lb[0]], \n", + " alpha=0.5, label=lb[1], marker='o', lw=3)\n", + "plt.xlabel('sample size n')\n", + "plt.ylabel('time per computation in milliseconds [ms]')\n", + "#plt.xlim([1,max(orders_n) + max(orders_n) * 10])\n", + "plt.legend(loc=2)\n", + "plt.grid()\n", + "plt.xscale('log')\n", + "plt.yscale('log')\n", + "plt.title('Performance of different conditional list creation methods')\n", + "\n", + "max_perf = max( f/c for f,c in zip(times_n['filter_func'],\n", + " times_n['cond_loop']) )\n", + "min_perf = min( f/c for f,c in zip(times_n['filter_func'],\n", + " times_n['cond_loop']) )\n", + "\n", + "ftext = 'the list comprehension is {:.2f}x to '\\\n", + " '{:.2f}x faster than the lambda function'\\\n", + " .format(min_perf, max_perf)\n", + "plt.figtext(.14,.75, ftext, fontsize=11, ha='left')\n", + "\n", + "plt.show()" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "display_data", + "png": "iVBORw0KGgoAAAANSUhEUgAAAnYAAAIECAYAAACUvmMzAAAABHNCSVQICAgIfAhkiAAAAAlwSFlz\nAAALEgAACxIB0t1+/AAAIABJREFUeJzsnXlcjdkfxz/3at+UciPtK0qLVGjkkr2msQ0hRAzFDMY6\nBhUyWUf2ZZB9GT+GwdhSEdlCQ5GKspOdRKnv74+mR0/d6t5cSs779bqv133Oc873fM95vs9zv/d7\nlkdARAQGg8FgMBgMxhePsKoVYDAYDAaDwWDIB+bYMRgMBoPBYNQQmGPHYDAYDAaDUUNgjh2DwWAw\nGAxGDYE5dgwGg8FgMBg1BObYMRgMBoPBYNQQmGPHkDvv37/H4MGDoaenB6FQiOPHj1e1Sl8kf/75\nJywsLKCgoIDBgwdLXS4kJARWVlZlHpclOyYmBnZ2dlBSUkLbtm3l04ivAH9/f7Rv377M47IwMzPD\nrFmzPqVqEsnIyIBQKMSpU6fKzBMZGQlFRUXuOCYmBkKhEPfu3fscKlYp0l6/LxGxWIyhQ4d+Etk1\nud++NJhj95Xi7+8PoVAIoVAIRUVFmJqaIjAwEE+fPv1o2f/73/+wdetW7Nu3Dw8ePECLFi3koPHX\nRX5+PgYPHgxfX1/cvn0bERERlZY1fvx4nDlzpkLZgYGBaNasGW7evIldu3Z9dBvkgaWlJUJDQ6ta\njQoRCATc98WLF2Pnzp3c8ZAhQ9CmTZtSZc6fP48xY8Z8Fv0+Fnd3dzx48AD169eXKn9Zba5ObNq0\nCUJh6Z/AktfvS6SstgkEAp6typtPKZshPQpVrQCj6vDw8MCOHTvw/v17nD9/HkOHDsXt27exb9++\nSsnLzc2FkpISUlNT0aBBAzRv3vyj9CuS9zVy7949ZGdno3PnzlL/mJaFuro61NXVy5VNREhLS8Ov\nv/6KBg0aVLouIkJ+fj4UFOTzaPlSfiiK7/OuqakpVRldXd1PpY7cUVRUhEgkqmo1UFBQAAASnRZ5\nIe31Y5SGve+gesAidl8xRQ9rAwMD+Pj4YNSoUTh48CDevXsHANi2bRscHR2hqqoKMzMzjB07Fm/e\nvOHKi8ViDBkyBFOnToWBgQFMTEzQpk0bTJs2DTdu3IBQKIS5uTkAIC8vD5MmTYKhoSGUlZVha2uL\nrVu38vQRCoVYvHgx+vbtC21tbQwYMIAbEoqJiUGTJk2gpqaGtm3b4sGDB4iOjoajoyM0NDTQvn17\n3jDRzZs30b17dzRo0ADq6uqwt7fHpk2bePUVDUvMmDED9evXh66uLgYOHIjs7Gxevu3bt8PZ2Rmq\nqqrQ09NDly5d8Pz5c+784sWL0bBhQ6iqqsLa2hqzZs1Cfn5+uX1/+vRpeHh4QE1NDXXq1EG/fv2Q\nlZUFoHAYzMTEBECh813ecPbbt28RGBgIbW1t1KlTB0FBQdz1K6L4UGxJ2bVq1UJsbCxq1aqF/Px8\nDBgwAEKhEBs2bAAApKWloUePHtDR0UGdOnXQsWNHXLlyhZNd/Po4OTlBRUUFUVFRyMvLQ0hICMzN\nzaGqqgo7OzusWrWq1PVevnw5+vfvDy0tLRgZGSE8PJx3fdLT0xEaGspFl2/dulVmn5Z3naS1v/L0\nAYCnT5+id+/e0NDQQL169TB16tRSP2bFh6RCQkKwdu1axMbGcm0o6ltTU1OEhYVx5V69eoVhw4ZB\nJBJBRUUFLi4uOHLkCHe+aAj1zz//hLe3N9TV1WFhYYH169fz6o+IiICTkxM0NTVRv3599OnTBw8e\nPCiz36Sh5FBsXl4efv75ZxgZGUFFRQUGBgbo06dPhW2WREJCAjp16oTatWtDU1MTbm5uOHv2LCfL\nysoKO3bsQMOGDaGsrIzU1FS8fv0ao0aNgqGhIdTV1dG0aVPs3r2bJ/fXX39F48aNoa6uDmNjYwQG\nBuLly5dcewYMGAAAnI5FUxIkDSnOmzcP5ubmUFZWhqWlZakIuqmpKYKDgzFq1Cjo6uqiXr16+Pnn\nn8t9DhRdz61bt6Jjx45QV1dH48aNERcXh1u3bqFTp07Q0NCAra0t4uLieGXLuy/LaxtQ6HxV9Myr\nqL3S3AdxcXFwd3eHlpYWtLS04OjoiMOHD5fZHww5QoyvkoEDB1L79u15afPnzyeBQECvX7+mdevW\nkY6ODm3atIlu3rxJx48fJ3t7e+rfvz+Xv3Xr1qSpqUmBgYF09epVunLlCj19+pTGjRtHZmZm9PDh\nQ3r8+DEREY0bN450dXVp586dlJqaSrNmzSKhUEhRUVGcPIFAQLq6urR06VK6ceMGpaam0rp160go\nFFKbNm3o7NmzdOHCBbKysqJvvvmGPDw86MyZM3Tp0iVq2LAh9e7dm5N1+fJlWrp0Kf37779048YN\nWrx4MSkoKFB0dDRPf21tbfr5558pJSWFDh8+THXq1KGpU6dyedauXUuKioo0c+ZMro1Llizh2hUc\nHEwmJib0119/UUZGBh04cICMjY15Mkpy//590tTUpH79+tGVK1coLi6O7O3tycPDg4iIcnJy6Ny5\ncyQQCOjvv/+mhw8fUm5urkRZo0ePJpFIRHv37qWUlBQaN24caWlpkZWVFZcnODiYOy5L9oMHD0gg\nENCyZcvo4cOHlJOTQw8ePCB9fX0KCgqiK1eu0PXr1+nHH38kXV1dysrKIiLiro+bmxvFxMTQzZs3\nKSsriwYOHEgODg505MgRysjIoO3bt5O2tjatWbOGd7319fXpjz/+oBs3btDSpUtJIBBwNvH06VMy\nMzOj8ePH08OHD+nhw4eUn58vsR8quk7S2l95+hARde3alaysrCg6OpqSkpLIz8+PtLS0ePdS8Xvr\n9evX1K9fP3J3d+fakJOTQ0REpqamFBYWxpXr2bMnmZmZ0eHDh+natWs0atQoUlJSomvXrhER0c2b\nN0kgEJC5uTn9+eeflJ6eTpMnTyYFBQW6fv06JyciIoKioqIoIyOD4uPjqWXLltS6dWvufJGckydP\nSuzLouuqoKDAHUdHR5NAIKC7d+8SUeGzwtDQkGJjY+n27dt07tw5ioiIqLDNJbly5QqpqalR3759\nKSEhgdLT02nHjh0UHx9PRIW2q6amRmKxmM6ePUupqan06tUrEovF1KZNGzp58iTdvHmTVq1aRUpK\nSrxrNXPmTIqLi6PMzEyKioqihg0b0sCBA4mIKDc3l7u+RTq+fPmy1PUjIlqyZAmpqqrS6tWrKS0t\njVasWEEqKio8WzYxMSEdHR2aPXs2paWl0Y4dO0hRUZGXpyRF18HCwoL27NlD169fp27dulGDBg1I\nLBbTX3/9RdevX6eePXuSkZER5eXlERFVeF+W1zZpnnnStLei+yAvL490dHRo7NixlJaWRmlpafTX\nX3/RiRMnyuwPhvxgjt1XysCBA6ldu3bccVJSEpmbm1OLFi2IqPBBtXLlSl6Z2NhYEggE9Pz5cyIq\nfEjY2NiUkh0cHEyWlpbccXZ2NikrK9Py5ct5+bp160Zt27bljgUCAQ0ZMoSXZ926dSQQCCgxMZFL\nmzt3LgkEArpw4QKX9vvvv5Oenl65bf7uu+9o6NCh3HHr1q3J0dGRlycwMJDrAyIiIyMj+vHHHyXK\ny87OJjU1NTp06BAvff369aStrV2mHlOmTOE9qImIEhMTSSAQ0PHjx4lIuh/f169fk4qKCv3xxx+8\n9GbNmpVy7Ipfj7JkCwQC2rx5M69c8+bNeXkKCgrIwsKCFi5cSEQfrk9cXByX58aNGyQUCiklJYVX\nNjQ0lNffAoGARo0axcvTqFEj+uWXX7hjS0tLCg0NLbMPiqjoOklrf+Xpk5qaSgKBgI4ePcqdz83N\npQYNGpRy7IrfWwEBASQWi0vpVdyxK5L9zz//8PI0bdqUBg8eTEQfrtvvv//Onc/PzydNTU1atWqV\nxLYTEV24cIEEAgHdu3ePJ+djHLtRo0bx+q4kZbW5JH5+fqXuweIEBweTUCik27dv83RRUVGhFy9e\n8PIOGjSIunbtWqasXbt2kbKyMne8ceNGEggEpfKVvH6GhoY0ceJEXp4xY8aQubk5d2xiYkLfffcd\nL0/nzp2pT58+ZepTdB2KHGIi4v50LViwgEu7ePEiCQQCSkpKIiLp7suy2ibNM6+i9kpzHzx9+pQE\nAgHFxMSU2X7Gp4MNxX7FxMTEQFNTE2pqamjSpAksLS2xefNmZGVl4datWxgzZgw0NTW5T5cuXSAQ\nCJCWlsbJcHZ2rrCetLQ05ObmwsPDg5fu4eGBpKQkXpqrq2up8gKBAE2aNOGO9fX1AQD29va8tCdP\nnnDDAW/evMGkSZNgZ2cHXV1daGpq4sCBA7yhPIFAAAcHB15d9evXx8OHDwEAjx49wp07d9ChQweJ\n7UpKSkJOTg66d+/O66fhw4fj5cuXePLkSZnlmjdvzpuHZm9vj9q1ayM5OVliGUmkp6fj3bt3aNmy\nJS/d3d1dLnNdzp07h4SEBF7btLS0kJmZybMBAHBxceG+nz9/HkQEZ2dnXtnffvutVDlHR0fesYGB\nAR49eiSTnhVdJ1nsrzx9iq5N8f5WVFTktb2yFMmWVUehUAiRSMTZLFB4X3fs2BHGxsbQ0tJCq1at\nAACZmZkfrWcRgwYNwuXLl2FpaYnAwEDs2rULeXl5MstJSEiAp6dnuXn09fVhaGjIHZ87dw65ublo\n0KABz742b97Ms69du3bBw8ODy+fn54e8vDyZhqVfvnyJu3fvSrwuGRkZePv2LYDCZ0lJ2yn+LCmP\n4s+gsp5tADg7lOW+LElFzzxp2ivNfaCjo4MhQ4agY8eO6NKlC2bPno3r169X2BcM+cAWT3zFNG/e\nHOvXr4eCggIMDAw4R6PoJl+0aJHElW1Fk+sFAgFvUr48kCRPKBTyJtEXfa9Vq1apNCKCQCDA+PHj\nsXfvXvz++++wsbGBmpoaxo4dixcvXvBkl1ycIRAIuAnaFVGUb+fOnbC2ti51XkdHR2I5gUDwRUwy\nJiK0a9cOS5YsKXWudu3a3PdatWrx+rGoX+Lj46GmpsYrV3IxxMf0/6dA0mKdivT5lNdSkuzy+uzW\nrVvo0qULBg4ciJCQEOjp6eH27dto164dcnNz5aaXg4MDbt68iSNHjiA6OhqjRo3C1KlTcfr0aZkW\nH0hzL5R8JhQUFKB27do4f/58qbxFfXPmzBn06tULkydPxvz586Gjo4P4+HgMHDhQrv0gqe4ipLXl\n4tvKFN0fktKKZEl7X8pbz4ooeR1XrVqFUaNG4fDhwzhy5AimTp2KJUuW4IcffvjouhjlwyJ2XzEq\nKiowNzeHsbExL3qkr68PIyMjXLt2Debm5qU+ysrKMtVjaWkJZWVlxMbG8tJjY2N5kTh5cuLECfj5\n+aFnz55o0qQJzMzMkJKSItMqS5FIBENDQxw6dEjieVtbW6ioqCA9PV1iP5W1cs/W1hanT5/mRTgS\nExPx4sUL2NnZSa2fhYUFlJSUcPLkSV76yZMn5bKatFmzZrhy5QoaNGhQqm3lregsiuJmZmaWKmdm\nZiaTDkpKShUuRKnoOn2M/RXvx8aNGwMAr79zc3Nx7ty5j26Dra0tp1Nxjh8/LtM9cu7cObx9+xYL\nFy5EixYtYGVl9dELJ8pCXV0dXbt2RUREBM6fP4+rV69yi3ykaTNQaCtRUVEyOccuLi54/vw5cnJy\nStlXUWQvLi4Oenp6mD59OlxcXGBpaYnbt2/z5BQ5OOXVraWlBUNDQ4m2Y25uDhUVFan1lhfS3JfS\ntE0S0rRXlvvA1tYWY8aMwYEDBxAQEFBqARXj0/DFRezOnj2L0aNHQ1FREQ0aNMCGDRvktrUC4wNh\nYWEICAiAjo4OfHx8oKioiKtXr+LgwYNYsWIFgMKHhjQPDjU1Nfz000+YOnUq6tatC3t7e+zcuRN7\n9+7F0aNHP4n+NjY2+Ouvv9C9e3eoq6tjwYIFuH//PurVq8flkUb/4OBgBAYGQl9fHz169EBBQQGi\no6PRp08f6OrqYvLkyZg8eTIEAgE8PT3x/v17XL58GZcuXSq1orKIkSNHIiIiAv7+/pg8eTKePXuG\noKAgeHh4wN3dXeo2qqurY/jw4ZgyZQr09fVhbW2NNWvW4Pr163LZmmLkyJFYs2YNvvvuO0yZMgWG\nhoa4c+cO/vnnH3h7e5e5P6GlpSUGDx6MoUOHYs6cOWjevDmys7ORkJCAx48fY8KECWXWWfKamJmZ\nIS4uDrdv34aqqip0dXUlOq0VXafK2l9xfSwtLeHj44MRI0Zg5cqVEIlECA8Px+vXr8u1I3Nzc+zc\nuRPJyckQiUTQ0tKCkpISr4yFhQW+//57BAUFYeXKlTA2Nsby5cuRnJyMbdu2VahjEVZWVhAIBJg3\nbx769u2LxMREzJgxo9zylWHu3Llo0KABHBwcoKamhq1bt0JBQYGLXJfV5pJMmDABbm5u6NevH8aO\nHQttbW1cuHABRkZGZW6X1LZtW7Rr1w7du3fHnDlz0KRJEzx79gynTp2CqqoqhgwZgoYNGyIrKwtr\n166FWCxGXFwcli9fzpNT9Cdjz549cHd3h5qamsQRg19++QVjx46FlZUVWrdujWPHjmHFihVYtmwZ\nl+dzRuCluS/Laps0z7yK2lvefVBEWloaVq9eDR8fHxgaGuLevXs4ceKEVFN3GB/PFxexMzY2RnR0\nNGJjY2Fqaoo9e/ZUtUpfJBVtVOnn54cdO3Zg3759cHNzg6urK0JDQ3lzXcqSISk9LCwMQ4cOxejR\no9GkSRNs2bIFmzdvlmoT07LqKC/t999/57ZfadeuHYyMjNCzZ89SQ7ol5ZRMCwgIQGRkJHbu3Akn\nJye0bt0ahw4d4v5MTJkyBQsWLMDq1avh6OiIVq1aISIiotzIlEgkwuHDh3Hnzh24uLjg22+/5ZyN\nitpYkvDwcHTt2hX9+/eHm5sbXr58iREjRkjVzooQiUSIj4+Hnp4eunfvjoYNG8LPzw+3b9+GgYFB\nubJWrVqFMWPGICwsDLa2tmjXrh02btwICwuLcussqWtoaCieP38OGxsb6Ovrl4q6FFHRdaqs/ZXU\nZ+3atXB0dIS3tzfEYjGMjIzQrVu3cvs7ICAALi4uaNmyJUQiEeeoley3P/74Ax07doSfnx8cHR0R\nHx+Pffv28Yb5K7J7e3t7LF68GCtXroStrS0WLFiAhQsXVur6l1emdu3aWLBgAVq2bAl7e3vs2bMH\n//vf/7htdcpqc0ns7OwQExODrKwstG7dGk5OTvj999+561bWM2bv3r3o3r07xowZg0aNGsHb2xv/\n/PMPLC0tAQBeXl749ddfMXnyZNjb22PHjh2YO3cuT5aLiwtGjRqFYcOGQV9fHz/++KPEOgMDAzF9\n+nTMmjULtra2mDt3LmbPno1BgwaV25/SbAZcmWebNPeltG2rbHvLug+K0NDQQFpaGnx9fWFjY4Oe\nPXvC3d1d4vAxQ/4I6EuY7FMGwcHBcHJyQteuXataFQaDwWAwGIwq54uL2BWRmZmJI0eO4Ntvv61q\nVRgMBoPBYDCqBVXm2C1ZsgTNmjWDiooKL8QLFO5q3a1bN2hoaMDU1LTUDvEvX77EgAEDsH79et7K\nSAaDwWAwGIyvmSpbddCgQQNMnToVhw4dQk5ODu/ciBEjoKKigkePHuHixYvw8vKCg4MDGjdujPfv\n38PX1xfBwcHcfA4Gg8FgMBgMBqr+lWJTpkwhf39/7vj169ekpKREqampXNqAAQNo0qRJRES0YcMG\n0tXVJbFYTGKxmLZv3y5RroGBAQFgH/ZhH/ZhH/ZhH/ap9h8LCwu5+FVVPseOSqzduH79OhQUFLjV\nTUDhZphFu6/3798fjx8/RnR0NKKjo9GrVy+Jcu/du8ct7a7On+Dg4C+ijsrKkKWcNHkrylPe+cqe\nq06fT62nvORXRo68bUWafJWxCWYr8q2DPVuqx4c9W2TL+ynsJT09XS5+VZU7diWXXr9+/RpaWlq8\nNE1NTbx69epzqvXZEIvFX0QdlZUhSzlp8laUp7zz5Z3LyMiosO7qwKe2F3nJr4wceduKNPkqYy/M\nVuRbB3u2VA/Ys0W2vJ/KXuRBlW93MmXKFNy9exfr1q0DAFy8eBHffPMNsrOzuTzz5s3D8ePHsXfv\nXqnlfimvbWJUD/z9/REZGVnVajC+AJitMGSB2QtDWuTlt1S7iJ21tTXev3/Pe5lxYmKiTK9aKiIk\nJAQxMTEfqyLjK8Df37+qVWB8ITBbYcgCsxdGRcTExCAkJERu8qosYpefn4+8vDyEhobi7t27WL16\nNRQUFFCrVi306dMHAoEAf/zxBy5cuABvb2/Ex8ejUaNGUstnETsGg8FgMBhfCl98xG7GjBlQU1PD\n7NmzsWnTJqiqqiIsLAwAsGzZMuTk5EAkEsHPzw8rVqyQyaljMGSFRXYZ0sJshSELzF4Yn5sq28cu\nJCSkzNCjjo4Odu/e/XkVYjAYDAaDwfjCqfLFE5+K8kKaderUwbNnzz6zRgxG1aCjo4OnT59WtRoM\nBoPBKAd5DcVWWcTucxASEgKxWFxqafGzZ8/Y/DvGV0PJBUoMBoPBqD7ExMTIdcj+q4zYsYUVjK8J\nZu/yJSYm5rPsEceoGTB7YUjLF794gsFgMBgMBoMhX1jEjsGo4TB7ZzAYjOoPi9gxGAwGg8FgMHjU\naMeOvXmi8oSEhMDKyoo7joyMhKKiotTlZc1fnNu3b8PT0xMaGhqoVatWpWSUh6mpKbdnIoMhK+yZ\nwpAFZi+MipD3mydqvGPHJq1WnuKrKX19fXHv3j2py5bMv2nTJgiF0pnbrFmz8PjxYyQmJuL+/fvS\nKywlAoGArRRlMBgMRrVALBbL1bGr0dudVJaUlEwcPZqOvDwhFBUL0K6dBWxsTKqdzE9N8bF+FRUV\nqKioSF1W1vzFSU1NhYuLCywsLCpVvoj3799DQYGZOEO+sD+LDFlg9sL43NToiF1lSEnJRGRkGrKy\n2uL5czGystoiMjINKSmZ1Ubm4sWL0bBhQ6iqqsLa2hqzZs1Cfn4+AGDHjh1QVlbGuXPnuPwbNmyA\nmpoarly5AqDwpdTt27fH77//jgYNGkBdXR29evUqd9NmSUOrCQkJ6NSpE2rXrg1NTU24ubnh7Nmz\npfLHxMRgwIABAAChUAihUIjBgwdLrEcoFOLYsWNYu3YtL9/9+/fh6+sLHR0dqKmpoU2bNkhISODK\nxcTEQCgU4sCBA/jmm2+gqqqKNWvWSNWfr169wrBhwyASiaCiogIXFxccOXKElyclJQVeXl7Q1NSE\npqYmfHx8kJ6eXqp/oqKiYGtrC1VVVTRv3hyJiYlS6cBgMBgMhjxg4YwSHD2aDmVlT/CnRXji33+P\nwcWlchG2s2fT8eaNJy9NLPZEVNQxmaN2ISEhiIyMREREBBwdHZGcnIzhw4fj7du3mD59Onr16oWj\nR4+iT58+uHjxIu7fv4+RI0diwYIFsLOzK6bTWairq+Pw4cN4/Pgxhg4dioCAAOzatUsqPZKSkuDh\n4YGuXbsiOjoa2traSEhIQEFBQam87u7uWLJkCUaOHIkHDx4AAFRVVSXKvX//Prp37w5zc3PMnz8f\nqqqqICJ07doVeXl52L9/P7S0tDBz5ky0b98eqamp0NXV5cqPHTsW8+bNg52dndRz/AYPHoyEhARs\n3rwZxsbGWL58Oby9vfHvv//CxsYGOTk56NChA6ytrXH8+HEQEcaNG4dOnTohOTmZq6egoAATJ07E\nihUroK2tjcmTJ8PLywtpaWmVjl4yqh9sXzKGLDB7YXxumGNXgrw8yUHM/PzKBzcLCiSXzc2VTeab\nN28wd+5c7N69Gx06dAAAmJiYYMaMGRg1ahSmT58OAIiIiICLiwuGDBmC69evo3379hg+fDhPFhFh\n48aN0NTUBAAsXboUHTt2xI0bN2Bubl6hLuHh4bC2tsbmzZu5tLLKKSoqQktLCwAgEonKlauvrw8l\nJSWoqqpyeaOionDu3DkkJyejYcOGAAqjkKampli2bBmmTp3KlZ8yZQq8vLwq1L+ItLQ0/O9//8OB\nAwfQvn17AMDChQtx4sQJzJkzB2vWrMGWLVvw+PFjXLx4EXXq1AEAbNu2Daampti2bRv69+8PoLBP\n586di1atWgEANm7cCCMjI2zZsqXMCCWDwWAwGPKkRjt2Zb1SrDwUFUtHnACgVi3J6dIgFEouq6Qk\nm8ykpCTk5OSge/fuvMn/+fn5ePfuHZ48eQJdXV2oqqpi+/btcHBwQP369REdHV1KVuPGjTmnDgBa\ntmwJAEhOTpbKsUtISECXLl1k0r+yJCUlQVdXl3PqAEBJSQlubm5ISkri5XV1deW+d+7cGXFxcdzx\nq1evSslOTk4GAHh4ePDSPTw8EB8fz9Vva2vLOXVAoYNqY2PDlS+iRYsW3HdtbW00atSoVB7Glw2L\nvjBkgdkLoyLk/UqxGu/YyUq7dhaIjIyCWPxh6PTduyj4+1vCxqZyeqSkFMpUVubL9PS0lElO0TDn\nzp07YW1tXeq8jo4O9/3EiRMQCAR48eIFHj16BG1tbV7ej90EsTpsektEpVa3qqurc9/XrFmDt2/f\nykW2pLZK0/6q7iMGg8FgVG+KAlChoaFykccWT5TAxsYE/v6WEImOQVs7BiLRsf+cusqvYJWXTFtb\nW6ioqCA9PR3m5ualPkXbiVy5cgVjx47FmjVr4OnpCV9fX+Tm5vJkXb16lRfBOnXqFIDCSJ40ODs7\nIyoqSmrHRUlJCUDlHB1bW1s8efIEV69e5dLevXuHM2fO8OYNlsTAwIDXP2XJBoDY2Fhe+vHjxznZ\ntra2SE5OxpMnT7jzDx8+xPXr10vVXxTlA4Dnz5/j2rVrUvcp48uA7UvGkAVmL4zPTY2O2FUWGxsT\nuW9FIg+ZGhoamDx5MiZPngyBQABPT0+8f/8ely9fxqVLlxAeHo63b9+iT58+6NatGwYMGIBvv/0W\nDg4OmDBhAhYuXMjJEggEGDBgAGbOnIknT55gxIgR+O6776QahgWACRMmwM3NDf369cPYsWOhra2N\nCxcuwMjICM2bNy+V38zMDACwZ88euLu7Q01NjRddKw4R8RxAT09PuLq6om/fvli6dCm0tLQwY8YM\n5ObmIjA7TMLTAAAgAElEQVQwUJYu5OQXYWFhge+//x5BQUFYuXIlt3giOTkZ27ZtAwD069cPM2bM\nQO/evTF37lwUFBRg3LhxMDQ0RO/evTlZAoEAEydOxPz586GtrY1ff/0VWlpa6Nu3r8w6MhgMBoNR\nGVjE7gtjypQpWLBgAVavXg1HR0e0atUKERERnOM0ZswY5OTkYMWKFQAKh2e3bNmCZcuW4Z9//uHk\nuLq64ptvvkH79u3RuXNnODg4YO3atdx5SZv4Fj+2s7NDTEwMsrKy0Lp1azg5OeH333/n7RtXPL+L\niwtGjRqFYcOGQV9fHz/++GOZbZRU919//YWGDRvCy8sLrq6uePToEY4cOcKb9ybtpsMl8/3xxx/o\n2LEj/Pz84OjoiPj4eOzbt48b7lZRUcHhw4ehrKwMDw8PiMViaGpq4uDBg7z2CoVCzJo1C8OGDYOL\niwsePXqE/fv3sxWxNQw2Z4ohC8xeGJ8bAdXQSUDlzQGrDvPDqhJ/f3/cvXu31F5tjMoTGRmJoUOH\nIi8vr6pVKcXXbu8MBoPxJSCvZzWL2DEYDIYMsDlTDFlg9sL43NRoxy4kJITdVBJg70r9NLA+ZTAY\nDIasxMTEyPVdsWwolsGo4TB7ZzAYjOoPG4plMBgMBoPBYPBgjh2DwWDIAJvewZAFZi+Mzw1z7BgM\nBoPBYDBqCGyOHYNRw2H2zmAwGNUfNseOwWAwGAwGg8GDOXYMBoMhA2zOFEMWmL0wPjfMsfvC8Pf3\nR/v27bnjkJAQWFlZVaFGXxZisRhDhw6tajUAFL4/d9asWVWtBoPBYDBqEDXasauJGxSX3Fx4/Pjx\nOHPmjNTlLS0tERoa+ilU+yKoTpsznz9/HmPGjKlqNRgywt79yZAFZi+MipD3BsUKFWf5cpFnR1UX\niIg3uVJdXR3q6upSl68uTo28ycvLg6KiYlWrIRO6urpVrQKDwWAwqhixWAyxWCy3oEuNjthVlpS0\nFCzdvhQLty3E0u1LkZKWUi1lAqWHYu/cuYMePXqgbt26UFVVhYWFBebNmweg0HjS09MRGhoKoVAI\noVCIW7dulSl7+/btcHZ2hqqqKvT09NClSxc8f/4cQKEjNWnSJBgaGkJZWRm2trbYunUrr7xQKMSS\nJUvQu3dvaGhowNTUFLt378azZ8/Qp08faGlpwcLCArt27eLKZGRkQCgUYvPmzfD09ISamhosLCyw\nffv2Unm2bNmCLl26QENDA9OmTQMAbNu2DY6OjlBVVYWZmRnGjh2LN2/e8PQiIsyYMQP169eHrq4u\nBg4ciOzsbF6eiuQUDemWJycpKQkdO3aEjo4ONDQ00LhxY2zatIk7b2pqirCwMO741atXGDZsGEQi\nEVRUVODi4oIjR46Uaveff/4Jb29vqKurw8LCAuvXry/zGjLkT00bBWB8Wpi9MD43zLErQUpaCiKj\nI5Gln4Xn9Z4jSz8LkdGRH+WIfQqZZREUFIRXr14hKioKKSkpWLNmDQwNDQEAu3fvhqmpKcaNG4cH\nDx7gwYMH3LmSrFu3Dv3790f37t1x8eJFxMbGwsvLC/n5+QCAyZMn448//kBERASSkpLg5+cHPz8/\nHDt2jCcnLCwM3t7e+Pfff+Hl5YX+/fvD19cXnTt3xqVLl+Dl5YUBAwbg6dOnvHITJkzAkCFDkJiY\niL59+6Jfv364dOkSL8/EiRPRv39/JCUlYdiwYYiMjERQUBDGjx+Pq1evYsOGDTh69CiGDx/OlSEi\n7Ny5E8+fP0dsbCy2bduGffv2Yfbs2VweaeQAqFBOnz59ULduXcTHx+PKlStYsGABdHR0uPMlh4UH\nDx6MI0eOYPPmzUhMTIS7uzu8vb2RksK3k0mTJsHf3x+XL1+Gr68vhgwZgtTUVInXkcFgMBhfF2wf\nuxIs3b4UWfpZiMmI4aWr31GHyzculdLlbNxZvDHkR43EpmKIHokQ1CtIJln+/v64e/cuF8kJCQnB\n5s2buR92R0dHdOvWDcHBwRLLW1lZoX///lyEqyyMjY3RtWtXLFq0qNS5N2/eoE6dOli4cCHP2ene\nvTtevHiBqKgoAIURu9GjR2PBggUAgMePH0MkEuHHH39EREQEAOD58+eoU6cO9u3bhy5duiAjIwPm\n5uaYOnUqLyzt7u4OCwsLbNiwgcszY8YM/Prrr1weU1NTTJ48GT/88AOXdvz4cYjFYjx79gy1a9eG\nWCzGixcvcPHiRS5PUFAQLl26hFOnTslVjra2NiIiIjBw4ECJfWxmZoahQ4di8uTJSEtLg7W1NQ4c\nOIBOnTpxeZydneHo6Ig1a9Zw7V6wYAFGjx4NACgoKIC2tjbmz59f5qIQto8dg8FgVH/YPnafiDzK\nk5iej/xKyyxAgcT03ILcSsssi9GjR2PWrFlo3rw5Jk2ahBMnTsgs49GjR7hz5w46dOgg8XxaWhpy\nc3Ph4eHBS/fw8EBSUhIvzcHBgfuup6eHWrVqwd7enkvT1taGkpISHj16xCvXokUL3rG7u3sp2a6u\nrtz3rKws3Lp1C2PGjIGmpib36dKlCwQCAdLS0iTqBAD169fHw4cP5SoHAMaNG4chQ4agTZs2CA0N\n5TmBJUlOTgYAqfrU0dGR+y4UCiESiXj1MhgMBuPrhTl2JVAUSJ6AXwu1Ki1TWEY3KwmVKi2zLPz9\n/ZGZmYnhw4fj/v376Ny5M/r37y/3eqRF0oKGkmkCgQAFBZKd3yIk/YspvmikqPyiRYuQmJjIff79\n91+kpqbCzs6Oq0tJid/vxeuXlxwAmDJlCq5fv45evXrhypUraN68OaZOnVpuO6Vpd0X1Mj4tbM4U\nQxaYvTA+NzV6VWxlaOfcDpHRkRBbibm0d6nv4O/rDxtLm0rJTDEsnGOnbKXMk+nZxvNj1ZVIvXr1\n4O/vD39/f3Tu3Bl9+/bF8uXLoaGhASUlJW6eXFmIRCIYGhri0KFD8Pb2LnXe0tISysrKiI2NRePG\njbn02NhYNGnSRC5tiI+P5w1Jnjp1Cra2tmXm19fXh5GREa5du4aAgIBK1ysvOUWYmZkhMDAQgYGB\nCA8Px7x58zBjxoxS+YraFhsbi86dO3Ppx48fh7Oz80frwWAwGIyvA+bYlcDG0gb+8EfUhSjkFuRC\nSagEzzaelXbqPpXMshg5ciS8vLxgbW2Nt2/fYteuXTA2NoaGhgaAQkcjLi4Ot2/fhqqqKnR1dSVu\ngRIcHIzAwEDo6+ujR48eKCgoQHR0NPr06QNdXV389NNPmDp1KurWrQt7e3vs3LkTe/fuxdGjR+XS\njrVr16Jhw4ZwdnbGpk2bcPr0aSxdurTcMmFhYQgICICOjg58fHygqKiIq1ev4uDBg1ixYgWA0tvF\nfCo5r1+/xsSJE9GzZ0+Ympri+fPnOHjwIM85LV7ewsIC33//PYKCgrBy5UoYGxtj+fLlSE5OxrZt\n28rVl82f+7ywfckYssDshfG5YY6dBGwsbeTudMlLZsmVlJI23B09ejRu374NNTU1tGjRAv/88w93\nLjQ0FD/88ANsbGzw7t073Lx5E8bGxqXqCQgIgKqqKubMmYOZM2dCQ0MDLVq04IZ1w8LCuMURWVlZ\nsLKywubNm9GmTZuPbiMAhIeHY9WqVTh9+jQMDAywefNm3twySc6on58fNDU1MXv2bISFhUFBQQHm\n5ubo0aMHr1zJsiXT5CFHUVERz58/R0BAAO7fvw8tLS20bduW23pGUhv++OMPjB8/Hn5+fnj58iXs\n7e2xb98+WFtbl9vumro3IYPBYDBkh62KZVQrilZ+xsXFoWXLllWtTo2A2bt8iYmJYVEYhtQwe2FI\nC1sVKwU18ZViDAaDwWAwag7yfqUYi9gxqhUZGRmwsLDAiRMnWMROTjB7ZzAYjOqPvJ7VzLFjMGo4\nzN4ZDAaj+sOGYhkMBqMKYNM7GLLA7IXxuWGOHYPBYDAYDEYNgQ3FMhg1HGbvDAaDUf1hQ7EMBoPB\nYDAYDB7MsWMwGAwZYHOmGLLA7IXxuWFvnmAwGAwGg8GoIjJTUpAup9dxAmyOHYNR42H2zmAwGNWT\nzJQUpEVGwvPtWwgWLmRz7L5G/P390b59+89Sl1AoxJYtW2QuJxaLMXTo0I+u/88//4SFhQUUFBQw\nePDgj5b3sYSEhMDKyqqq1WAwGAxGDSH9n3/geesWcPGi3GQyx+4LQ9LL56sb8tAxPz8fgwcPhq+v\nL27fvo2IiAg5aVcxcXFxEAqFuHXrFi99/PjxOHPmzGfTg1E9YXOmGLLA7IVRJqmpEMbEAHfvylUs\nm2MngaLxbmFeHgoUFWHRrh1MbGyqhUwi+iqG1e7du4fs7Gx07twZ9evXrxIdSvazuro61NXVq0QX\nBoPBYNQQsrOBgweBy5dRkJsrd/EsYleCovHutllZED9/jrZZWUiLjERmSkq1klnEhQsX0LlzZ+jr\n60NTUxOurq44dOgQL4+pqSmmTZuGwMBAaGtro169eli+fDnevn2LESNGoE6dOjA0NMTSpUtLyX/8\n+DF69OgBDQ0NGBoaYtGiRfy2ZWaiU6dOUFNTg7GxMRYvXlxKxpYtW+Dm5gZtbW3UrVsX3t7eSE1N\nLbNNkZGRMDExAQB4eHhAKBQiNjYWkZGRUFRU5OW9c+cOhEIhjh8/DqDw37FQKMTRo0fh4eEBdXV1\n2Nra4uDBg7xyjx49wqBBg1CvXj2oqqqiYcOGWLduHTIzM+Hh4QEAMDMzg1AoRNu2bQFIHopdv349\nGjduDGVlZRgZGWHq1KnIz8/nzhcNS8+YMQP169eHrq4uBg4ciOzs7DLbz6jeiMXiqlaB8QXB7IXB\nQQT8+y+wdClw+TIAwMLcHFECAdCokdyqYRG7EqQfPQpPZWWgWPjcE8Cxf/+FiYtL5WSePQvPN294\naZ5iMY5FRX10JPDVq1fo06cPFixYAEVFRaxfvx4+Pj64cuUKzwlZvHgxgoODceHCBWzduhUjR47E\nnj170KlTJ5w/fx47duzATz/9hLZt26JRMQMLDQ3F9OnTMXv2bBw4cABjx46FqakpfHx8QETo1q0b\nFBUVERsbCyUlJYwfPx4XLlzg1Z2bm4tp06ahcePGePnyJaZNmwYvLy8kJSWVctQAwNfXF3Z2dnB1\ndcXevXvh6uoKHR0d3Lx5U+p+GTduHObMmQMLCwuEhYWhd+/eyMzMhLa2NnJyctC6dWuoq6tjy5Yt\nsLCwQHp6Oh4/fgwjIyPs2bMH3333Hc6dOwcjIyMoKSlJrGP//v0ICAhAWFgYevTogQsXLmD48OEQ\nCASYPn06l2/nzp0YPHgwYmNjkZmZCV9fX5iYmPDyMBgMBqMG8/w5sH8/UCKoYdK2LRAQgGPx8XKr\nijl2JRDm5UlOLxaFkVlmQYHkdDmEYFu3bs07njFjBv7++2/8+eefmDx5Mpfepk0bjB49GgAwefJk\nzJkzB8rKylzaxIkTMWfOHBw7dozn2Hl7e2PEiBEAgJ9++glnzpzBvHnz4OPjg6ioKFy6dAnXr1+H\npaUlgMLonLGxMU8nf39/3vG6deugp6eH8+fPo0WLFqXapKKiAj09PQBAnTp1IBKJZO6XkJAQdOjQ\nAQAQHh6OyMhInDt3Du3bt8eWLVuQkZGB9PR0GBgYAAAXIQQAHR0dAEDdunXLrTs8PBw9e/bExIkT\nAQCWlpZ48OABJk2ahGnTpkFBofD2MjU1xfz58wEA1tbW6N27N44ePcocuy+UmJgYFoVhSA2zl68c\nIuDsWSAqCij+m1+7NuDtDVhZwQSAiZMT8N9v7cdSo4diQ0JCZJ64WiAhggQABbVqVVqPAqHkbi4o\nIxIkC1lZWQgKCkKjRo2go6MDTU1NJCUl8Sb+CwQCODg48I7r1q0Le3t7XppIJEJWVhZPfknHq2XL\nlkhKSgIAJCcnQ09Pj3PqAEBPTw82JaKQly5dQrdu3WBubg4tLS3OicrMzPzI1peNo6Mj910kEqFW\nrVp4+PAhACAhIQG2tracU1dZkpOTuWHbIjw8PPD27Vukp6dzacX7HgDq16/P6cJgMBiMGkpWFrB2\nLfDPPx+cOoEAcHUFgoKA/0a2YmJiEBISIrdqa3TErjIdZdGuHaIiI+FZ7B9W1Lt3sPT3Byo5bGqR\nklIoU1mZL9PTs1LyiuPv7487d+5g7ty5MDMzg4qKCnx9fZFbIhpYcshTIBBITCsoI7ooC8UXHbx5\n8wYdOnSAh4cHIiMjoa+vDyKCra1tKR0rQijBQc4rI8Iqafi0eNs+1wIUgUBQShd59TOjamDRF4Ys\nMHv5CsnPB+LigOPHC78XUbcu4OMDGBnxstevb4a6deX3m1CjI3aVwcTGBpb+/jgmEiFGWxvHRCJY\n+vt/1Fy4TyGziBMnTiAoKAje3t6wtbVFvXr1eNGijyW+xLj/qVOnYGtrCwBo3LgxHj9+jLS0NO78\n48ePkVJsUcjVq1fx+PFjhIWFwcPDAzY2Nnj69GmlHCuRSIT8/Hw8evSIS7tw4YLMcpo1a4bk5GTc\nLWOJeZEjll/B8LutrS1iY2N5abGxsVBTU4OFhYXMejEYDAbjC+fOHWDlSiA6+oNTJxQCrVsDw4aV\ncupSUjIRGZmGS5fayk2FGh2xqywmNjZycbo+tUwAsLGxwaZNm+Du7o73799j2rRpKCgo4DlOkpwo\nadP279+PpUuXokOHDjh48CB27NiBnTt3AgDatWsHBwcH+Pn5YfHixVBUVMTEiRN5ESoTExMoKytj\n0aJF+Pnnn5GRkYFJkyZVap87Nzc3aGpqYtKkSfjll1+Qnp5eqXlqffr0wZw5c+Dj44M5c+bA3Nwc\nN27cwJMnT9CrVy+YmJhAKBRi//796NWrF5SVlVG7du1Scn755Rd8++23mD17Nrp164ZLly4hNDQU\nY8eO5ebXfS3b03xNsDlTDFlg9vKVkJsLHDsGnDlTOK+uiAYNCqN0+voSi+3enY6kJE+8fCk/VVjE\n7guj5Oa/69atQ0FBAVxdXdG9e3d06dIFLi4uvDySnChp06ZNm4ajR4/C0dER4eHhmDt3Lr777jvu\n/F9//YXatWvDw8MDPj4+8Pb2RtOmTbnzenp62LRpE44cOQI7OztMmDAB8+fPlzisWpE+Ojo62Lp1\nK06fPg0HBweEhYVh7ty5pfJV5DSqqqoiNjYWdnZ28PX1RePGjfHjjz/i7du3AAB9fX389ttvCA8P\nh4GBAbp168bJLS67c+fOWLt2LdavX48mTZrg559/xogRIxAcHMzTRZJ+1X2TaQaDwWBISXo6sGwZ\ncPr0B6dOURHo1AkICJDo1OXnA7GxQGysUK5OHcDeFctg1HiYvTMYDMYn4M0b4NAhIDGRn25hUbji\n9b8dFkpy5w6wdy/w6BFw9uwxvHnTFgIBEBMjn2c1G4plMBgMBoPBkBYiICmpcLVr8c3mVVULo3T2\n9oWrX0sgabTW3NwCN25EwdbWE/J6+xxz7BgMBkMG2Jwphiwwe6lhvHwJ7NsHXL/OT7ezK3TqNDQk\nFktPB/7+u3Cf4iIUFQE/PxM8eZGODXt/kpuKzLFjMBgMBoPBKA8i4Px54OhR4N27D+laWoCXV5nb\noeXkFI7WXrrETy8arX30JAV7Lh+BRXcNYJl8VGVz7BiMGg6zdwaDwfgIHj8unBRXbON/AICLC+Dp\nCaiolCpCBCQnAwcOlB6t7dgRcHAoHK1dsm0JLqtfRvqzdEQNjGJz7BgMBoPBYDA+Cfn5wMmThctX\ni+9rqqtbuIVJsVdRFufly0KH7to1frqtLdC584fR2mc5z3Dq7inc070nV7WZY8dgMBgywOZMMWSB\n2csXyr17wJ49QPHXPwqFgLt74WbDCqXdJyLgwgXg8GH+aK2mZuGwa9FobQEV4Ozds4i6EYXnOc9L\nyflYmGPHYDAYDAaDARQuXY2JAeLj+RsNGxgURunq1ZNY7MmTwsURGRn89GbNgHbtPozWZmVnYU/K\nHtx5eQcAYG5ujsTkRJg0lRz9qwxf5Ry7OnXq4NmzZ59ZIwajatDR0cHTp0+rWg0Gg8Go3ty4Ueid\nFfcPFBWBNm2A5s0LI3YlyM8v9AFjYoD37z+k6+oC334LmJr+l68gH3G34nA88zjy6cOwrkhdBFsl\nWySlJGFE7xFymWP3VTp2DAaDwWAwGAAKl64ePgxcvMhPNzMr9M7q1JFY7P79wjUV9+9/SBMKgZYt\nC0drFRUL0+6+vIs9KXvwKPvDe85rCWrBw8QD3xh/g1rCWgDk57ewoVgGA2weDEN6mK0wZIHZSzWG\nCLh6tXClw+vXH9JVVAqXrjo6StxoOC+vcD3FqVNAQcGH9Pr1C0dr69f/L19+HqIzohF/Ox6EDw6b\noZYhfGx8IFIXfZJmMceOwWAwGAzG18WrV8D+/aWXrjZuDHTpUuZGwxkZhaO1T558SFNQKBytbdHi\nw2jtzWc3sTdlL569/TCsqyhUhKe5J1wbuEIoqPh96ZXl00n+wggJCUFeXh537O/vj6VLl36UzJiY\nGLi4uAAA7t27h7Zt25abPzMzE6tXr/6oOqsDISEhGD9+/Gepy8vLCzdv3vxoOUX/qFeuXImFCxfK\nVLZfv35o0KABhEIh3rx5U2a+ESNGoFGjRnB0dMQ333yDhIQE7py/vz+MjIzg5OQEJycn/PbbbzK3\nITIyEqmpqTKXIyK0aNECjo6OcHBwQLt27ZCWliYx76ZNm2Bvbw9FRUWJ98fixYvRqFEj2Nvbw8nJ\nSWZdUlNT4eTkBGdnZ2zdulXm8vK+hxYuXIisrCzuOCQkBPv375eb/JJI0t/U1BTJyckfJfdT3JOy\n6FWZNjx58gQtW7aEk5MT5s+fXxkVy2XPnj04d+4cd5yQkAA/Pz+518OiddUMIiAhAViyhO/UaWoC\nvr5Ar14Snbq3bwsdushIvlNnagoEBhYulhUKgbfv3+LvlL+xPnE9z6kz1zFHkEsQmhs2/6ROHcAi\ndhzTp0/H+PHjofjfoLhAQvj1YzAwMMCxY8fKzXPz5k2sWrUKQ4cOlWvd8ub9+/dQkLDUuwh59115\nyPtHdtiwYTKXGTp0KBYuXAh9ff1y83Xp0gWLFi1CrVq1sH//fvTu3ZtzoAQCAX755RcEBQVVSm+g\n0LGrW7curKysZConEAhw+PBhaGpqAgAWLVqEn3/+GXv37i2V18nJCdu3b0d4eHip67xr1y7s3LkT\n58+fh7q6Os8hkpZdu3bB3d0dS5Yskbks8HH3UH5+PmrVqsVLi4iIQPv27VG3bl0An962Jekvj3k3\nn0JvWfSqTP1Hjx5FnTp1sG/fPpnLSsPu3bvh4uLC/fl2dnbGpk2bPkldjGpCWUtXnZ2B9u0lbjQM\nFPp/+/cXBvmKUFYGOnQAmjb9MFqb8jgF+67vw6vcDxlVFFTQ0aIjHOs5frbfRhaxQ2EkBQBatmyJ\npk2b4sWLFwCAK1euwNPTE9bW1hg4cCCX/+XLlxgyZAjc3Nzg4OCA0aNHo6D4QLsEMjIyoKenBwB4\n8+YNvv/+e9ja2sLR0RG+vr6cHsnJyXByckKvXr0kyvntt99gb28PR0dHuLu7c+mzZ89GkyZN0KRJ\nEwwePBjZ/211HRISAl9fX3h5ecHKygq9evXC+fPn0aZNG1haWmLChAmcDLFYjDFjxsDNzQ1WVlb4\n9ddfS51r0aIFunbtytXp5uYGZ2dn+Pj44GGx/X7u3r0LLy8vNGrUCN7e3sjJyQEA5ObmYvz48XBz\nc4OjoyMGDBjA6erv74/AwECJfb5q1So0btwYTk5OcHBwwPX/3tNXPBKQlpYGT09PODg4wNnZGYcO\nHeLKC4VC/Pbbb3B1dYWFhQV27drF69eY/96+XDyycerUKTg7O8PJyQl2dnbYtm2bxGsiFou5H/7y\n8PLy4hyH5s2b486dO7zzkn4kc3Jy4ODgwDlZx44dQ6NGjbg+K2LdunVISEjATz/9BCcnJxw7dgwF\nBQUYN24cZxfjx48v006LnDoAePHiBUQiyXM/bG1t0ahRIwiFwlL6zp8/H6GhoVBXVwcArk+kbcPm\nzZuxcOFC/Pnnn3BycsKNGzcwf/58uLq6omnTpmjZsiUSExMByHYPpaSkoEuXLnB1dYWjoyMiIyO5\nOoVCIUJDQ+Hq6orp06fz9AkLC8O9e/fQs2dPODk54erVqwCACxcuSLTtqKgo7hlib2+P7du3c7LE\nYjEmTJiAVq1awcLCAr/88ovE/i3rGbBjxw60bNkSZmZmvEhpeW0ri8uXL8PDwwPOzs6wtbVFREQE\nd87f3x/Dhw+Hp6cnTE1NMXr0aBw5cgStWrWCmZkZFi1axJO1adMmNGvWDFZWVjy9Tpw4gSZNmsDe\n3h4//vgjz1bGjRvH6duuXTvcKrmbP4Do6GhMmDABJ0+ehJOTE+Li4iAWi3l/5MRiMQ4cOFBh/969\nexc9evSAg4MDHBwcEB4ejsOHD+Pvv/9GeHg4nJycsHHjRt4ICwBs2LAB9vb2cHBwQPfu3bk/KpGR\nkejQoQN8fX1hZ2eHb775hvfsK0mMvN7szqg8+flAXBywfDnfqatTB/D3L1wgIcGpe/0a2LED2LaN\n79Q1bAiMHFnoDwoEQHZuNnYm78TWK1t5Tl0jvUYY4TICTvWdPmvAA/SF8eLFC3JxcSENDQ1KSkoq\nM5+sTRMIBJSdnc0dDxw4kFq1akXv3r2j3NxcsrW1pSNHjhARUUBAAG3cuJGIiPLz88nX15dWr15d\nSmZ0dDQ1a9aMiIhu3rxJenp6RES0a9cu6tixI5fv+fPnREQUExPD5ZdEZGQktWjRgl6/fk1ERE+f\nPiUiogMHDpCdnR29evWKiIgGDBhAEydOJCKi4OBgsrKyopcvX1J+fj45ODhQhw4dKDc3l7Kzs0kk\nElFaWhoREYnFYurYsSPl5+fT69evqUmTJrRv3z7u3HfffUf5+flERLRx40b64YcfqKCggIiIli1b\nRv369ePV+eLFCyIi6tChA9c/M2bMoJkzZ3JtmjBhAv36669l9vnRo0eJiKh27dr04MEDIiLKzc2l\nN+ei7uEAACAASURBVG/eEBGRqakpZweurq60du1aIiJKTk4mPT09evz4MXd9ly5dSkREJ0+epAYN\nGpS6VkREISEhNH78eCIi8vHxoa1bt5a6TmVR0obKIyQkhHr06MEd+/v7k5mZGTVp0oS6du1KV69e\n5c5du3aNjI2N6cyZM2RmZkaXLl2SKFMsFtP+/fu542XLllG7du0oLy+PcnNzydPTk5YvX16mTp07\nd6Z69epRw4YN6dGjR+Xq7+/vT0uWLOGl6ejo0KxZs6hly5bUrFkz3j0hbRuK9z8RUVZWFvf9yJEj\n1Lx5cyKS/h7Ky8ujpk2b0rVr14iI6OXLl2RtbU0pKSlEVHjN5syZU2Y7i9sXUaFtGxoaSrTtZ8+e\ncffHgwcPyNDQkNNLLBaTr68vERU+w/T09Lj7rjiSngGmpqZcn2RkZJCGhgZlZ2dLbJuNjQ13XJyQ\nkBAaN24cERG9evWK3r17x31v3LgxV6boHiy6x0QiEQ0ePJiIiO7evcvVXaRXQEAAERE9fPiQDAwM\n6PLly/T27VsyMDCg2NhYIiLasWMHCQQCrh+L7kkiotWrV3P9UpLIyEjq2bMnd1zSvosfl9e/YrGY\n5s2bx5Urqt/f3597JhDxn9eXL18mAwMD7pkzdepU6t27NxERrVu3jnR0dOjOnTtERDR06FDuGSaJ\nomcLo4q4d49oxQqi4OAPn9BQoiNHiHJzJRYpKCC6eJEoPJxfbM4coqSkwvOF+Qro0v1LFH4inIKj\ng7nP3JNzKelR2f5JWcjLJfvihmLV1NRw4MABjB8//pNuZyIQCNC1a1coKSkBAJo2bYobN24AAPbu\n3Ytz585x8z5ycnJgbGwstWxHR0dcvXoVI0eOhFgshpeXFwDJEZvi7N+/H0FBQVxEREdHB0DhkEWf\nPn2g8d+8gB9++AGjRo3iynXq1ImLyBRF+xQVFaGoqAgbGxukp6fDwsICADBw4EAIhUKoq6vD19cX\nx44d4/Tr27cvhP/NDN27dy8SEhLQtGlTAIXDs9ra2rw6tbS0AABubm5IT0/nyr169Qo7d+4EALx7\n9w6Ojo5l9nl6ejo8PT3Rtm1bDBgwAN9++y28vLxgZmbG65tXr14hMTERgwYNAgBuLtvp06c5/Yui\nOm5ubrh37x5yc3O5uorPgym6Dm3btsXMmTORnp6O9u3bw9XVtdzrIy3btm3D1q1bceLECS4tLCwM\nBgYGAICNGzeiU6dOuHHjBoRCIWxsbDB9+nS0bNkSERERcHBwKFN2cRuKiorCoEGDuGHzQYMGYffu\n3Rg+fLjEsgcOHAAR4bfffkP//v1x8OBBmdqVn5+PO3fu4OTJk8jKyoK7uztsbGzQqlWrSrfh/Pnz\nmDVrFp49ewahUMhFaqW9h65fv45r165x1x4A8vLycPXqVVhbWwMALzJcEQKBAN26dZNo248ePcKg\nQYOQlpYGBQUFPH36FCkpKZzdfP/99wAALS0tNGrUCGlpadx9J6ntxSnS38TEBDo6Orhz5w7ev39f\nqm25ubm4du0abMp4ITkAZGdnY/jw4fj3338hFApx7949JCYmwsbGhrsHiz8fivrWwMCAq7uo7wIC\nAgAAIpEIXl5eiI6ORkFBAdTV1eHh4cG1+4cffuDqP3DgAJYtW4bXr1/jffGNv0og6/O9ZP+mp6dD\nX18f8fHxiIqK4vLp/p+9O4+L6r73x/+ahX3fZd8ZRRRUEEURFMUVbNI2zaJttl/SNk3T7Tb3mhg1\nbZPc3jZNm+Y2eSTNftOkzaP5CgiCbIqKu6IiDpvsIPsOAzNzfn98nDlzcEYHGYbF9/Px6KPMmzkz\n55DD+OHz+bzfbze3u75HUVERtm/frt1e8eyzzwru2TVr1sDX1xcAm30/cuSIwfOiPXYzZHycLzR8\np9TVCXp6gKws4NavtVZMDEuUtbFhj/tG+5BZmYnqbuF+5JgFMdgcuhk2FjYmvJjJmXMDO6lUql3S\nnG5WVlbaryUSieBD6ODBgwjSVB6cpODgYFy7dg35+fnIycnBnj17cOXKFaOO1fdBNHGvy8TnTLyO\nO13XxNfRnT62n7ChdO/evXj88cf1ns/E9xgdHdU+/tvf/mbww27icZqEln//+984e/YsCgsLsX79\nerz77rvYsmXLbcdPPGdd1rem2jXLoUqlUjuw0+eFF15Aeno6jhw5gueffx6pqan4zW9+Y/D5xvjm\nm2/w8ssvo7CwULB8qxnUAcDu3bvx85//HM3NzfD39wfANnZ7eXmhsbHxjq8/8drvdF8YOv7JJ580\nKnlj4nsFBATgkUceAcCWYTdt2oQzZ84gMTFxUtegMTY2hu985zs4fvw4YmJi0NLSAj8/PwDG/w5x\nHAd3d3dcnFifSsfE+/puDN3bP/rRj/Ctb30L33zzDQBAJpMJ7ntrnaUeiUQClW7vybuYeKxSqTTq\n2nRp/nvt2bMHPj4++PTTTyEWi7F582bBeU68Pn3vraHv80Lf758mVl9fj1/84hc4d+4cAgMDcfLk\nSTz22GNGnb9UKhX8zHTPGdD/M9J3nvrOS1/8Tr87uu8lFovvOEAlM6CujhWY0y3MLpUCycmsyJye\nQsNqNXD6NFBYyMaEGi4ubKU2JIQ95jgOZ1vOIr82H2OqMe3znK2dkRaRhlDXUMw02mN3i4ODA3p7\njevZlp6ejtdff127X6mzsxN1Ezdj3kFzczNEIhF27tyJN998Ex0dHejp6YGjo6N2f58+O3bswN/+\n9jcM3qq303UrNWfjxo346quvMDg4CI7j8MEHHyA1NdXo89HgOA6ff/45VCoVhoaG8K9//UuQyav7\n4Zaeno533nlH+zNTKBS4fPnybc/TPNbE0tPT8cc//lH7oTwwMIDrE9PNJ1CpVKipqUFcXBxefPFF\npKam4tKlS4LnODg4ICYmBp988gkAoKKiAmVlZVi1apVR167ZB6N77pWVlQgODsYzzzyDn/70p4IM\nuok0x91p8JSVlYVf/vKXyMvLu22Gt7m5Wft1bm4upFKpdkbgm2++wYkTJ3D16lVkZWUZnElzdHQU\n3MMbN27EJ598AqVSifHxcXzyySd674vOzk50dnZqH//rX/+66+yk7n9TjUcffRQ5OTkA2KxQSUmJ\ndjbW2GvQfc3R0VGoVCrtYO5///d/td8z9ndIJpPB1tZWsCn++vXrGNDdMHMHE3+mHMcJBqa6P4e+\nvj4E3moKfuTIkdsyi40ZWN/tM0DXwoULjb62iefp5+cHsViMq1evCmaO9TF03hzHaff0dXR0ICcn\nB+vXr0dERARGRkZw/PhxAMDXX3+t/Rn29/fD0tISXl5eUKvVePfdd426VgAICwvT/g5eu3btts8A\nfedpb2+PhIQE/OlPf9LGNJ+bE//b6tLs39PsnXv//ffv6TMVoD12ZjU6ygZ0H38sHNRpUlfXrtU7\nqLt5E/j734HcXH5QJxKx8iU/+hE/qOsc7sRHlz5CdlW2dlAnggjxvvH4cdyPZ8WgDpjBgd1f//pX\nxMbGwtraWrt8ptHd3Y0HHngA9vb2CAoKMlj2wJSbEX/5y19iw4YNguQJQ6//1ltvQSKRIDo6GkuX\nLsXWrVvR0tKi9/x0X0Pz9eXLl5GQkICYmBjEx8djz549WLBgAaKjoyGTybBkyRK9yROapchVq1Zh\n2bJleOCBBwCwZc9du3Zh9erVWLp0KcRiMV5++WW953Cn6xKJRFi4cKH23Hbs2IFt27bpPW7Xrl14\n7LHHkJSUhOjoaMTGxuLkyZMGr1vz+D//8z8RHR2NuLg4REdHIzExUTCw03euKpUKTzzxhHYZua2t\nTW/26v/93//h888/R3R0NHbt2oXPP/9cu+wymZ+B5ntvv/02oqKisHz5crzzzjv43e9+p/eYBx98\nEAEBARCJRJDJZNi6dav2e8uWLUNbWxsA4Mknn8T4+Di+/e1va8uaaFrbPf7449rre+2115CRkQGx\nWIy6ujq88MIL+Oqrr+Di4oKvvvoKzz77rN777ZlnnsGrr76qTZ545plntGVHli9fjpiYGL3Zom1t\nbdiyZYt2c/nRo0e1A+SJ1/CPf/wD/v7++Prrr7F37174+/tr//v9/Oc/R2NjI6KiohAfH4/du3cj\nJSVlUteg+/N3dHTEq6++iri4OMTGxsLe3n7Sv0NSqRSZmZn48ssvER0djaioKPzkJz/RzgTf7TPk\npz/9KZ544gksX74cFRUVeu8jTeyNN97Ar371Kyxbtgz/+te/bltuNubz6m6fAbokEoneaxsbG7vt\nubrn+fLLL+P9999HdHQ0Dhw4gKSkpDue551+Vzw8PBAbG4uEhATs2bMHixcvhpWVFf7xj3/gxz/+\nsfZ+0gx4lyxZgu9+97uIjIzEqlWrEBISYtTvIgD8+te/RnZ2NpYuXYrf//732m0gdzvPzz//HCdO\nnMCSJUsQExODDz/8EACbGf/iiy+0yRO67xcVFYU33ngDmzZtQnR0NK5cuaJNMrnT5xuZQRUVwDvv\nABcu8DFrazbd9oMfsB5fEyiVQFER8N57gM7f1vDyAp5+mi29WlqydmAl9SV499y7aOjjk33cbd3x\n5LInsTV8Kywlhld/zG3GWop98803EIvFyM3NxcjICD766CPt9zTLOX//+99x8eJFbN++HSdPnkRk\nZKT2OU888QR+9atfYfHixXpfn1qKTd769evxH//xH4LBHCGEEDJrDQ6yzhET6yQuXAhs387q0+nR\n2Mgm93SrMkkkrBXYmjXsawBoHWjFQflBtA22aZ8nFomRGJCIxMBESMWm29E251uKaWabzp07Jyj7\nMDQ0hH//+98oLy+Hra0t1qxZg507d+Kzzz7T7vvZtm0bysrKIJfL8eyzz05q8zMhhBBC5jiOY71d\n8/LYEqyGvT3rHLFokd52YAoFUFAAnD3LXkLD35/lVGi2Po+rxnG0/ihONp6EmuOTL3wcfLBTthNe\n9neuWzqTZjx5Ql8Wm1QqRVhYmDYWHR0t2KegqV10N48//rg2wcHZ2RkxMTHaTfua16PH/ON9+/bN\nqvMx5+O33nqL7g96bNRjzdez5Xzo8ex+TPfLNDzOyABOnkTyrSSW4lt73JMfeABITUXx6dNAe/tt\nx/v6JiMrCygrY4+DgpJhaQm4uBQjJATw8GDP/zLrS5xsOAnXSFcAQN2lOkhEEjz14FNY5bcKx44e\nQwUqpnw9mq8ns0ffGDO2FKuxd+9eNDU1aZdiS0pK8NBDD6G1tVX7nPfffx9ffPEFioqKjH5dWool\nk1FcXKz9pSPkTuheIZNB94sJqdXAqVNsY9ydUlcnGB4GDh8GbuX3aYWHAzt2AE5O7LFCqUB+bT7O\ntggT5YKcg5AuS4erjaspr+Y2c34pVmPiRdjb26O/v18Q6+vrE1TGJ8TU6IOXGIvuFTIZdL+YSFsb\n2xSnm3SlSV1dvx641Q5UF8cBV68COTlscKdhawts3QpERfGrtVVdVciszES/gh9/WEmskBqaiuXe\ny+dUgsyMD+wm/rAiIiKgVCpRXV2tXY4tKytDVFTUTJweIYQQQmaKUgkcPQqcOCEsNOzlBezcCejU\nANXV18cKDVdVCeNLl7Js11t1/jE8PozD1Ydx+aZwOi/CLQI7InbA0crRlFdjFjM2sFOpVBgfH4dS\nqYRKpYJCoYBUKoWdnR0efPBBvPLKK/jggw9w4cIFZGZmorS0dNLvsX//fiQnJ9NfTOSuaLmEGIvu\nFTIZdL9MQX09m6W7VXsQACs0nJTECg1rUld1cBxLjMjPB3Qr/zg5sWXX8HDN8ziUd5Qjuyobw+P8\ndJ6thS22hW/DYo/FZpulKy4uFuy7m6oZ22O3f//+25pu79+/H6+88gp6enrw5JNP4siRI3B3d8cb\nb7whaJtjDNpjRyaDPnyJseheIZNB98s9UCiAI0eAc+eE8cBAtpfOQPepzk42Dmxo4GMiERAXB6Sk\nAJqmKv2KfhyqPAR5l1xw/FKvpdgStgW2FramvBqjmWrcMuPJE9OFBnaEEELIHCOXA4cOAbp77a2s\ngE2bgBUr9JYwUanYSu3Ro+xrDXd3VsJE0+iH4zhcaL2AvJo8KFQK7fMcrRyRFpGGcLfw6boqo8yb\n5AlCCCGE3OcGB1mWQ3m5MC6TsULDjvr3ujU3s1m6W93fALCuYYmJ7H/SW6Oc7pFuZMgzUNdbJzg+\nzicOG0M2wkpqhfnC4MBu9+7dRr2AlZUVPvjgA5OdkCnRHjtiLFouIcaie4VMBt0vd8FxQFkZa9Q6\nMsLH7exY6urixXpn6cbGWNWTU6eEhYZ9fdksndet+sFqTo3SxlIU1RVBqVZqn+dm44Z0WToCnQOn\n68qMZrY9dlZWVtizZ4/BaUHNlOEf//hHoxtqmxMtxZLJoA9fYiy6V8hk0P1yBz09LHW1pkYYj4kB\nUlNZXRI9amuBzEx2uIaFBbBhAxAfz2bsAKBtsA0Z8gy0DPAlUsQiMRL8E5AUmAQLye0lUmbStO+x\nCw0NRc3EH7YeMpkMcrn8rs8zNxrYEUIIIbOQWg2cPg0UFgoLDTs7s+SI0FC9h42MsA5iFy8K4yEh\n7DAXF/ZYqVbiWP0xHG84LmgHtsB+AXbKdsLbwdvUV2QSlDxxFzSwI4QQQmaZmzfZprjmZj4mErGp\ntg0bAEvL2w7hOKCiAsjOZlvxNKytWU26mBh+tbaxrxEZ8gx0DHdonycVS5EUmIQE/wRIxLeXSJkt\nZjR5ora2FmKxWNuHlZC5jpZLiLHoXiGTQffLLUolUFLC/qdbaNjTk22K8/PTe9jAAEuSvX5dGI+M\nBLZtA+zt2eMx1RgKagtwpvkMOPCDowCnAKTL0uFuq79Eynxk1MDu4Ycfxk9/+lMkJCTgo48+wo9/\n/GOIRCL85S9/wdNPPz3d50gIIYSQuaqhgc3SdXbyMYkEWLcOWLvWYKHhCxdYObvRUT7u4MCSZBcu\n5GM13TXIrMxE72ivNmYpscSmkE2I9YmdU+3ATMGopVgPDw80NzfD0tISUVFReO+99+Ds7IydO3ei\nurraHOc5aSKRCPv27aOsWEIIIWQmKBRAQQFrBaE71PD3Z7N0Hh56D+vuZuPAujphfMUKVs7O2po9\nHhkfQW5NLi61XRI8L8w1DGkRaXCydjLhxUwfTVbsgQMHzLfHztnZGb29vWhubsbKlSvRfGtt3MHB\nYVZmxAK0x44QQgiZMVVVLOO1r4+PWVoCGzeyVhB6ZtHUaqC0lJUxUfKVSeDqysaBuru/rnVcQ3ZV\nNgbH+E13NlIbbA3fiiWeS+bkLJ1Z99hFR0fj9ddfR11dHbZv3w4AaGpqgpPT3BgNE3I3tA+GGIvu\nFTIZ9939MjQEHD4MXLkijIeHs2atBsYNbW3AwYNAaysfE4uB1auB5GRWzgQABhQDyK7KRkVnheD4\nKM8obA3bCjtLOxNezNxk1MDu73//O/bu3QtLS0v8/ve/BwCUlpbisccem9aTI4QQQsgcwHFsMHf4\nMDA8zMdtbVmh4agovbN0SiVrBXbihDCnYsECYOdOwNtb8/IcLrVdQm5NLkaV/KY7B0sHbI/YjoXu\nC0EYKndCCCGEkHvX28tSV6uqhPGlS4EtWwwWGq6vZ3vpurr4mFTKZuhWr+ZzKnpGepBZmYnanlrB\n8Su8V2BT6CZYS61NeDEzx+zlTkpKSnDx4kUMDAxo31wkEmHPnj1TPonpQi3FCCGEkGmiVrPEiIIC\n1uNLw8mJLbuGh+s9bHQUyM8Hzp0TxgMD2V46N7dbL8+pcab5DApqCzCu5gsZu1i7IF2WjmCXYFNf\n0YwwW0sxXc8//zz++c9/IjExETY2NoLvffbZZyY7GVOiGTsyGffdPhhyz+heIZMxb++Xjg423dbY\nyMdEImDlSlZo2MpK72FyOZvc6+/nY1ZWrIPY8uX8am37UDsy5Blo6m/iXx4irPZfjfVB62ddOzBT\nMOuM3eeff47y8nL4+PhM+Q0JIYQQMkepVHyhYZWKj3t4sOk2f3+9hw0OAjk5QHm5MC6Tsbp0jo63\nXl6twvGG4zhWfwwqjn99LzsvpMvS4evoa+ormneMmrFbunQpCgsL4e4+dyo304wdIYQQYkJNTWyW\nrr2dj0kkQGIiKzQsvX2uiOOAy5dZTsXICB+3s2OdIyIj+Vm65v5mHJQfRPsQ//oSkQTrAtdhbcDa\nWd0OzBTM2iv27NmzeO211/Doo4/Cy8tL8L1169ZN+SSmAw3sCCGEEBMYGwMKC4HTp4WFhv382Cyd\np6few3p7gcxMoKZGGI+JYUuvmpyKcdU4Cm8U4lTTKUE7MD9HP6TL0uFpp//15xuzLsWeP38e2dnZ\nKCkpuW2PXaPu+johc9S83QdDTI7uFTIZc/5+qa5mhYZ7+XZdsLAAUlLYfjqx+LZD1GrgzBk2FtTN\nqXB2BtLSgNBQPnaj5wYy5BnoGe3hX15sgZSQFKz0XQmx6PbXJ3dm1MDupZdeQlZWFjZt2jTd50MI\nIYSQmTY8DOTmAmVlwnhoKBudOTvrPay9na3WNvE5DxCJgPh4llNhaclio8pR5NXk4ULrBcHxIS4h\nSItIg4uNiymv5r5i1FJsQEAAqqurYan5LzIH0FIsIYQQMkkcxzIccnJYFwkNGxtWk27pUoOFhktK\ngOPHhTkVnp5stdbPj49d77yOQ5WHMDDGtyS1llpjc+hmxCyImZPtwEzBrEuxr776Kn72s59h7969\nt+2xE+uZhp0tqI4dIYQQYqS+PlaLpLJSGI+KYt0j7PS362psZLN0HR18TCIB1q1jORWaQsODY4PI\nqcpBeYcwNTbSIxJbw7bCwcrBlFczZ8xIHTtDgzeRSASV7tB8FqEZOzIZc34fDDEbulfIZMyJ+4Xj\nWLXg/HxAoeDjjo6s0HBEhN7DxsZYbeIzZ4Q5Ff7+bJbOw0Pz8hwu37yMw9WHMaLkU2PtLe2xLXwb\nIj0ip+Oq5hyzztjV1tbe/UmEEEIImVs6O9l0W0ODMB4XB2zcaLDQsL6cCktLllMRF8fnVPSO9iKr\nMgvV3dWC45ctWIbU0FTYWAgTMsnUUa9YQggh5H6jUgEnTgBHjwo3xbm7s+m2gAC9hxnKqQgLY5N7\nmpwKjuNwtuUs8mvzMabiU2OdrZ2RFpGGUNdQEKFpn7Hbu3cvfvOb39z1Bfbt24cDBw5M+UQIIYQQ\nYgbNzWyW7uZNPiYWsw1x69YZLDSsL6fC1pblVCxZwudUdA53IkOegYY+fhZQBBHi/eKxIXgDLCVz\nJxFzLjI4Y2dvb4/Lly/f8WCO47BixQr06s7FzhI0Y0cmY07sgyGzAt0rZDJm1f0yNgYUFQGnTgk3\nxfn6slm6CcmRGv39bNl1Yk7FkiVsUKfJqVCpVTjZeBLFdcWCdmAeth5Il6XD30l/uzHCTPuM3fDw\nMMLCwu76AlYG1t8JIYQQMkvU1rI2ED18IWBYWLDicvHxegsNTyanonWgFQflB9E22KaNiUViJAYk\nIjEwEVKxUVv6iQnQHjtCCCFkvhoZAfLygIsXhfGQEFZo2EV/IeDOTjYOrK8XxifmVIyrxnG0/ihO\nNp6EmlNrn+fj4IOdsp3wstc/C0huZ9as2LmK6tgRQgi5L3EccO0a2xQ3OMjHbWyAzZuB6Gi9hYZV\nKuDkSZZToVTycX05FfW99ciQZ6BrpEsbsxBbYH3weqzyW0XtwIw0I3Xs5iKasSOTMav2wZBZje4V\nMhkzcr/09wPZ2cD168L44sWs0LC9vd7DWlpYTkUbv5qqN6dCoVQgvzYfZ1vOCo4Pcg5Cuiwdrjau\nprya+wbN2BFCCCGEx3HAhQts6VV3U5yDA7B9O7Bwod7DxsdZTkVpqTCnwseHzdItWMDHKrsqkVWZ\nhX5FvzZmJbFCamgqlnsvv2/bgc0mNGNHCCGEzHVdXWxTXF2dMB4byzbFWVvrPezGDXZYdzcfs7AA\n1q8HVq3icyqGxoZwuPowrrRfERwvc5Nhe8R2OFo5mvBi7k9mnbFrb2+HjY0NHBwcoFQq8emnn0Ii\nkWD37t2zulcsIYQQMq+pVGyqrbhYuCnOzY0lRwQF6T1sZAQ4coRN8OkKDmaHud5aTeU4DlfbryKn\nOgfD48Pa59lZ2GnbgdEs3exi1IzdypUr8d5772HZsmV48cUXkZWVBQsLCyQnJ+Ott94yx3lOGs3Y\nkcmgfVPEWHSvkMmY1vultRU4ePD2TXEJCUBSEpt606OiAjh0SJhTYW3NcipiYvicin5FP7Iqs1DZ\nJSxgF+0Vjc1hm2FrYWvqK7qvmXXGrqqqCjExMQCAzz//HCdPnoSDgwMiIyNn7cCOEEIImZfGx9kM\nXWkpoOZLjMDbm22K8/bWe9jAAMupqKgQxiMjWU6FgwN7zHEczreex5GaI1Co+L16TlZO2BGxA+Fu\n4Sa+IGJKRs3Yubu7o6mpCVVVVXj44YdRXl4OlUoFJycnDOoO+WcRmrEjhBAy7+jbFCeVsk1xq1cb\nLDR88SLLqRgd5eP29iynYtEiPtY13IXMykzU9dYJXmOl70qkBKfASkpNCaaLWWfstmzZgoceeghd\nXV343ve+BwC4du0a/Pz8pnwChBBCCLmL0VE2MrvbprgJurvZOPDGDWF8+XIgNZXPqVBzapQ2lqKo\nrghKNb9Xz83GDemydAQ6B5ryasg0MmrGbnR0FJ988gksLS2xe/duSKVSFBcXo62tDQ8//LA5znPS\naMaOTAbtmyLGonuFTIZJ7hdDm+JSU4Fly/QWGlarWUvYoiK2cqvh6srGgcHBfKxtsA0Z8gy0DLRo\nY2KRGGv81yApKInagZmJWWfsrK2t8eyzzwpi9MFGCCGETCNDm+IWLQK2beM3xU3Q1sYKDbfw4zSI\nRCynIjmZz6lQqpU4Vn8MxxuOC9qBedt7I12WDm8H/Xv1yOxmcMZu9+7dwife+ouA4zhBavOnn346\njad370QiEfbt20ctxQghhMwtk9kUp0OpZK3ATpwQ5lQsWMByKnx8+FhjXyMOyg+ic7hTG5OKwmdn\nggAAIABJREFUpUgOSsZqv9WQiCWmvipigKal2IEDB0wyY2dwYLd//37tAK6zsxOffPIJ0tLSEBgY\niPr6emRlZeEHP/gB/vKXv0z5JKYDLcUSQgiZc+60KW7TJtbrVY/6enZYJz9Og1TKqp4kJACSW+O0\nMdUYCmoLcKb5DDjw/0YGOgUiTZYGd1t3U18RMZKpxi1G7bFLTU3F3r17kZiYqI0dP34cr776KvLy\n8qZ8EtOBBnZkMmjfFDEW3StkMoy+X9RqVr6kqEhYaNjFhU236W6K06FQAPn5wFlh21YEBrK9dO46\n47Tq7mpkVWahd7RXG7OUWGJTyCbE+sRSoeEZZtY9dqdOncKqVasEsfj4eJSWlk75BAghhJD7mrGb\n4iaorASysoB+vm0rrKzYxN6KFXxOxcj4CHJrcnGp7ZLg+HDXcOyI2AEnaycTXxCZSUbN2CUlJSEu\nLg6/+c1vYGNjg+HhYezbtw+nT5/GsWPHzHGek0YzdoQQQma1yWyK0zE0BOTkAFevCuMyGduC53ir\nbSvHcajorMChykMYGh/SPs/WwhZbwrZgiecSmqWbRcw6Y/fxxx/j0UcfhaOjI1xcXNDT04PY2Fh8\n8cUXUz4BQggh5L5TX89m6bq6+Ji+TXE6OA64fBnIzQWG+batsLNjnSMWL+Zn6QYUA8iuykZFpzCj\nNsozClvDtsLO0m46rorMAkbN2Gk0NDSgpaUF3t7eCAyc3cUKacaOTAbtmyLGonuFTMZt98voKNsU\nd+6c8ImBgWyWzs1N7+v09rJl1+pqYTw6mvV4tb3VtpXjOFxqu4TcmlyMKvmMWgdLB+yI2AGZu8wE\nV0Wmg1ln7DSsra3h6ekJlUqF2tpaAEBISMiUT4IQQgiZ9+RyNjobGOBj+jbF6VCrWWJEQQEwNsbH\nnZ2BHTuAsDA+1jPSg8zKTNT21ApeI9YnFhtDNsJaam3qKyKzkFEzdocPH8ZTTz2F1tZW4cEiEVQq\n1bSd3FTQjB0hhJBZYXCQbYorLxfGJ26Km6CjAzh4EGhq4mMiERAfD2zYAFhaspiaU+N002kU3ijE\nuJpvM+Fq44p0WTqCnINMfEFkOpi13ElISAh+/etf4/vf/z5sNfO9sxwN7AghhMyUerkcNUeOQFxf\nD3VlJUL9/RGoqT1iZ8c6R0RG6p2lU6mAkhL2P925E09Ptlqr26a9fagdGfIMNPXzoz8RREjwT0By\nUDIsJPozasnsY9aBnaurK7q6uuZU9gwN7Mhk0L4pYiy6V8jd1MvlqH73XaTU1aH4xg0kOzujQKlE\nWEwMAjduZJviDBQabmpiORXt7XxMIgESE9n/NDkVKrUKJQ0lKKkvgYrjR39edl7YuXAnfBz0Z9SS\n2cuse+yeeuopfPjhh3jqqaem/IaEEELIvKVWo+a995BSViYoYZJib49CDw8Efutbeg8bGwMKC4HT\np1n2q4afH5ul8/TkY839zTgoP4j2IX70JxFJkBSUhDX+a6gd2H3OqBm7tWvX4syZMwgMDMSCBQv4\ng0UiqmNHCCGEAKzAcEYGiv/f/0Oybo9XPz8gOBjFbm5I/tnPbjuspoa1A+vlG0LA0hJISQHi4gCx\nmMXGVGMoulGEU02nBO3A/B39kS5Lh4edx3RdGTEDs87YPf3003j66af1ngQhhBByXxsbY63ATp0C\nOA5qzUjMzo4lSNxKjlBrsh1uGR4G8vKAS8KGEAgLYxmvzs587EbPDWTIM9Az2qONWYgtsDFkI+J8\n4yAWiafl0sjcM6k6dnMJzdiRyaB9U8RYdK8Qgaoq4NAhwXRbfU8Pqru7kRIcjOKGBiQHBaFAoUDY\n448jUCYDx7EE2Zwc1kVCw8YG2LIFWLqUz6kYVY4iryYPF1ovCN421CUUabI0OFs7g8wPZp2x4zgO\nH330ET777DM0NzfDz88Pu3btwhNPPEGzdoQQQu4/Q0PA4cPAlSvCeEgIAnfsADo6UFhQgMvd3VB7\neiIsJQWBMhn6+9k4UC4XHhYVxbpH2Ok0hLjeeR2HKg9hYIyve2cjtcHmsM2I9oqmf3+JXkbN2P3u\nd7/Dp59+il/+8pcICAhAQ0MD/vSnP+Gxxx7Dyy+/bI7znDSRSIR9+/YhOTmZ/romhBBiGhzH1k7z\n8oCRET6ub7ptwmHnzwNHjgAKBR93dGTLrhERfGxwbBA5VTko7xDWvYv0iMS28G2wt7Q39VWRGVRc\nXIzi4mIcOHDAfOVOgoKCcPToUUEbsfr6eiQmJqKhoWHKJzEdaCmWEEKISXV1sc4RN24I40uXshIm\ndvr7r3Z1sRIm9fXCeFwcsHEjaz4BsNWxyzcv43D1YYwo+UGjvaU9todvxyKPRaa8GjLLmHUpdnh4\nGO6awoq3uLm5YVQ364eQOYz2TRFj0b1yH1KpgJMngaNHAaWSj+vr63WLXF6PvLwaFBVdxtjYUgQF\nhcLdnU2OuLmxEia6Ldd7R3uRVZmF6m5hM9hlC5YhNTQVNhb6694RMpFRA7stW7Zg165deP311xEY\nGIi6ujq89NJL2Lx583SfHyGEEDJzmppYLZKbN/mYSASsXg0kJ/N9vXTI5fV4551q1NamoKlJDGfn\nZFy6VIBly4BvfSsQSUmA9Na/vhzH4WzLWeTX5mNMxTeDdbZ2RrosHSEu1I+dTI5RS7F9fX14/vnn\n8dVXX2F8fBwWFhZ46KGH8Pbbb8PZeXZm5NBSLCGEkHumULCKwWfOCCsGe3uz6TZvb72HjY8DP/95\nIa5e3SCI29sDa9YU4r/+i493DnciQ56Bhj5+S5MIIsT7xWND8AZYSm4fNJL5y6wtxTRUKhU6Ozvh\n7u4OiWR2V7amgR0hhJB7Ipez1NX+fj5mYQFs2ADEx/MVgyeoqWFb8HJyijE6mgyAPTU4mNUodnEp\nxs9+lgyVWoWTjSdRXFcsaAfmYeuBnQt3ws/RT+/rk/nNrHvsPvnkE8TExCA6OhpeXl4AgLKyMly+\nfBm7d++e8kkQMtNo3xQxFt0r89jAACthUi7MRkVYGLB9O+Diovew4WEgNxcoK2OPxWLWSszZGbCx\nKYa/fzIAwNJSjZaBFmTIM9A22KY9XiwSIzEgEYmBiZCKjfpnmRCDjLqD9u7di0sTSmP7+fkhLS2N\nBnaEEELmNo4DLlxgtUh0kwLt7FgJk6gogyVMrlxhY8HhYT4eGRmKnp4C+PmlaDNhhxW5sF7cgQ8u\nHIea43vI+jr4Il2WDi97r+m6OnKfMWop1sXFBZ2dnYLlV6VSCTc3N/T19U3rCd4rWoolhBByV52d\nLDliYi2SmBggNRWwtdV7WG8vW3atFiaxYskSNhZsaqpHQUENxsbEGJC0QBFUBakD/2+ShdgCG4I3\nIN4vntqBEQBmXopdtGgRvv76a3zve9/Txr755hssWkQ1dQghhMxBSiVw/DhQUsLKmWi4ugJpaWxj\nnB5qNXD6NMurGB/n405OrPJJePitgGQUYy6XUd5Rjua+ZoQMhMDdgZUNC3YORposDa42rtN0ceR+\nZtSM3fHjx7Ft2zZs2rQJISEhqKmpQX5+PrKzs7F27VpznOek0YwdmQzaN0WMRffKPNDQwGbpOjr4\nmFgMrFkDrFvHEiX0aGtjhYZbWviYSASsXMnyKjSFhuXVcryZ/SbqXOpws/wmnBc6Q1mtxMqoldiV\ntAvLFiyjdmDkNmadsVu7di2uXLmCL774Ak1NTVi5ciX+/Oc/w9/ff8onQAghhJjF6CiQnw+cOyeM\n+/qyEiZe+ve5jY+z2sQnT7IZOw0vLza556eTxDo0NoT/yfofVDtXAzoTgV5LvOCr9MVy7+UmvCBC\nbjfpcic3b96Ej4/PdJ6TSdCMHSGEEK2KCiA7m2W+alhaAikprLeXgRImN26wyb3ubj4mlQJJSUBC\nAqDZes5xHMpuliG3OhdFxUUY9WNJGBZiC4S7hcPD1gMuN13ws4d/Nl1XSOY4s87Y9fT04LnnnsPX\nX38NqVSK4eFhZGRk4MyZM/jtb3875ZMghBBCpkV/PxvQXb8ujEdEsBImTk56DxsZAfLygIsXhfGg\nIDZL5+bGx3pGepBVmYWanhoAgBhskLjAfgFCXUJhIWFLu5ZiKjhMpp9RqTg//OEP4ejoiPr6eljd\n2kSwevVqfPnll9N6coSYS3Fx8UyfApkj6F6ZIzgOOHsWeOcd4aDO3h747neBRx7RO6jjOODqVeCv\nfxUO6qyt2WrtD37AD+rUnBqljaX437P/qx3UAUD0omgsHFiIhe4L0XylGQCgqFIgZXnKtFwqIbqM\nmrErKChAa2srLHQ2lHp4eKC9vX3aTowQQgi5J+3tbP20sVEYX7EC2LgRsLHRe1hfH2s4UVkpjEdG\nAlu3Ag4OfOzm4E0clB9EywCfSSGCCKv8VmF94nrcuHEDBRcK0NndCc92T6SsT4EsTGaqKyTEIKP2\n2IWFheHYsWPw8fGBi4sLenp60NDQgNTUVFyfOL09S9AeO0IIuc8olcCxY8CJE8ISJu7ubP00MFDv\nYWo1m9wrKADGxvi4oyNbrZXpjMeUaiWO1h3FicYTgkLDXnZeSJelw9fR19RXRe4TZt1j9/TTT+M7\n3/kOfvvb30KtVqO0tBR79uzBs88+O+UTIIQQQqasro7N0nV18TGJBFi7FkhMZBkPerS3sxImTU18\nTCQCYmPZ5J6mhAkA1PfWI0Oega4R/j2kYimSApOQ4J8AiXh291An9wejZuw4jsNf/vIXvPfee6ir\nq0NAQAB++MMf4oUXXpi1tXhoxo5MBtUmI8aie2WWGRlhrcAuXBDG/f3ZLJ2np97DNJN7x48LS5h4\neLC9dLrVvEaVo8ivzce5FmGZlECnQKTJ0uBu627w9Oh+IcYy64ydSCTCCy+8gBdeeGHKb0gIIYRM\nGccB5eWsUevgIB+3smJTbbGxevu7Aqx7WEbG7ZN7iYlsgk93cu9653UcqjyEgTG+TIqVxAqbQjdh\nhfeKWTu5Qe5fRs3YFRYWIigoCCEhIWhtbcWLL74IiUSC119/HQsWLDDHeQq8+OKLKC0tRVBQED78\n8ENI9Uyx04wdIYTMU4ayHBYtYlkOjo56DxsdZZN7588L4wEBbHLPw4OPDY4NIrsqG9c6rgmeu9B9\nIbaFb4Ojlf73IORemWrcYtTAbuHChcjLy0NAQAAeeeQRiEQiWFtbo7OzExkZGVM+ickoKyvDH/7w\nB3z22Wd47bXXEBISgocffvi259HAjhBC5hm1GjhzhjVq1c1ycHBgWQ4LF+o9jONYfeKcHGF9Yisr\nYNMmliyrmXjjOA6X2i4htyYXo8pR7XPtLe2xLXwbFrkvolk6Mi3MuhTb0tKCgIAAjI+PIzc3V1vP\nztvbe8onMFmlpaXYvHkzAGDLli346KOP9A7sCJkM2gdDjEX3ygwx1Kg1NpZ1j7C21nuYofrECxcC\n27YJJ/e6R7qRKc/Ejd4bgucuW7AMqaGpsLHQXyblTuh+IeZm1MDO0dERbW1tKC8vx+LFi+Hg4ACF\nQoHx8fHpPr/b9PT0aAeUjo6O6Nbt80IIIWR+MdSo1dOTrZ8a6FnOcawlbH4+oFDwcXt7Nrm3aBEf\n0xQaLqorglKt1MZdrF2QJktDiEuIqa+KkGlj1MDu+eefx8qVK6FQKPDWW28BAE6cOIFFur8Zk/TX\nv/4VH3/8Ma5evYpHHnkEH330kfZ73d3deOqpp3DkyBG4u7vj9ddfxyOPPAIAcHZ2Rn9/PwCgr68P\nrq6u93wOhGjQX9TEWHSvmFFtLZCVJWzUKpGwRq1r1vCNWifo6GCVTxoahPEVK9jSq+7kXutAKzLk\nGWgdbNXGRBAhwT8ByUHJ2nZg94ruF2JuRg3sXnzxRXzrW9+CRCJBWFgYAMDPzw8ffPDBPb+xr68v\n9u7di9zcXIyMjAi+99xzz8Ha2hrt7e24ePEitm/fjujoaERGRiIhIQFvvvkmdu/ejdzcXKxdu/ae\nz4EQQsgsNDwM5OYCZWXCeGAgm6Vz119eRKlk5UtKSoT1id3cWAkT3frE46pxFNcVo7SpVFBoeIH9\nAuyU7YS3g/m3GhFiCkYlT0ynvXv3oqmpSTtjNzQ0BFdXV5SXl2sHkT/4wQ/g4+OD119/HQDw61//\nGqdOnUJgYCA++ugjyoolU0b7YIix6F6ZRhwHXLnCSpgMD/Nxa2sgNRVYtsxgCZPGRrYFr6ODj4nF\nrHzJunXCEiY3em4gszIT3SP8TKBULEVyUDJW+602aaFhul+IsaY9eWLhwoXadmH+BvYwiEQiNEyc\n656kiRdRWVkJqVSqHdQBQHR0tKDx9u9//3ujXvvxxx9HUFAQALaEGxMTo/0F07wePabHAHDp0qVZ\ndT70mB7fd48HBpDc2wvU1KC4ro59PygIWLwYxfb2QH8/km8N6nSPVyiAN98shlwOBAWx16urK4aH\nB/Af/5EMT0/++fFr4nGk9gj+nfNvAEBQTBAAYLR6FKv9VmNtwNrZ8/Ogx/P+sebrulv3u6kYnLEr\nKSlBYmLibScxkeZE79XEGbuSkhI89NBDaG3l9zu8//77+OKLL1BUVGT069KMHSGEzAFqNXDqFFBU\nxBIlNJycWJZDRITBQ69fZxmvt7ZdAwAsLfn6xGIxH7/WcQ3ZVdkYHOOLGVtLrZEamoplC5ZRCRMy\n46Z9xk4zqAOmPni7k4kXYW9vr02O0Ojr64ODg8O0nQMhhJAZ0NLCshx0/pCHSATExwMbNrBRmh4D\nA6wm3TVh7WBERLCxoJOTznMVA8iuykZFZ4XguZEekdgathUOVvRvC5lfDA7s9u7da3D0qImLRCK8\n+uqrUzqBiX8lRUREQKlUorq6WrscW1ZWhqioqCm9DyF3UlxcPK1/wJD5g+4VExgbYzN0p06xfXUa\nXl4sy8HXV+9hHMdawh45wrpIaNjZsYYTixcLCw1faL2AvJo8KFR8vRMHSwdWaNjj3qs6TAbdL8Tc\nDA7sGhsb7zg1rRnY3SuVSoXx8XEolUqoVCooFApIpVLY2dnhwQcfxCuvvIIPPvgAFy5cQGZmJkpL\nSyf9Hvv370dycjL9UhFCyGxRXc1KmPT28jGpFEhOBlavNljCpLOTTe7V1wvjy5axvAobndrBXcNd\nyKzMRF1vneC5K7xXYFPoJlhL9RczJmQmFBcX33HL22TNWFbs/v37b5vt279/P1555RX09PTgySef\n1Naxe+ONNybdXYL22BFCyCwyNMSyXa9cEcZDQoAdOwADNUlVKuDECeDYMVbORMPVlVU+CQ7Wea5a\nhZONJ3G0/qig0LCbjRvSZGkIcg4y4QURYlrT3iu2trbWqBcICQmZ8klMBxrYEULILMBxwKVLQF4e\noFuz1MYG2LwZiI42WMKkqYmVMGlv52NiMZCQwGoUW+jUDm4ZaEGGPANtg238c0VirPFfg3WB66Zc\naJiQ6TbtAzuxbjrRHU5CpVsFchahgR2ZDNoHQ4xF98okdHWxZdcbwt6rWLqUDers7PQeplAAhYXA\nmTPCLXg+PmwL3oIFfGxMNcYKDTeWggP/ZB8HH6TL0rHAXufJM4DuF2Ksac+KVev25COEEEKMpVKx\n3q5HjwrXT52d2bKrTp3SiSorgUOHgL4+PmZhwZJk4+OFJUxqe2qRKc9Ez2gP/1yxBdYHr8cqv1UQ\ni+4+QUHIfDPjnSemi0gkwr59+yh5ghBCzKmpiWU53LzJx0QilhiRnGywhMngINuCd/WqMB4WxsaC\nzs58bGR8BLk1ubjUdknw3BCXEKRFpMHFxsVEF0PI9NMkTxw4cGB6l2I3b96M3NxcAMKadoKDRSIc\nO3ZsyicxHWgplhBCzMjQ+qm3N1s/9dbfe9XQFjxbW1bCJCpKWMJEU2h4aHxI+1wbqQ02h21GtFc0\nFRomc9a0L8V+//vf13791FNPGTwJQuYD2gdDjEX3ih5yOVs/1S0ub2EBrF8PrFolXD/V0d3NtuBN\nzNWLjmZb8Gxt+Vi/oh+HKg9B3iUXPHexx2JsDd8Ke0t7U12NSdH9QszN4MDuscce0379+OOPm+Nc\nCCGEzCUDA2z9tLxcGA8LYy0gXPQviapUQGkpUFws3ILn4sKWXUND+RjHcTjXcg75tfmCQsOOVo7Y\nHr4dMneZCS+IkLnP6D12x44dw8WLFzE0xKa/NQWK9+zZM60neK9oKZYQQqbJnVpAbNkiXD+doKWF\nlTBp46uSGNyC1znciQx5Bhr6GgSvEecTh5SQFCo0TOaVaV+K1fX888/jn//8JxITE2GjW957lqPO\nE4QQYmKGWkDExLAWELrrpzoMdRHz9maFhn18+JhKrcKJxhM4WncUKo4vqeVu6450WToCnAJMeUWE\nzKgZ6Tzh4uKC8vJy+Oj+5s1yNGNHJoP2wRBj3bf3ilLJt4DQrV+qrwXEBPq6iFlY8F3EdLfgNfU3\nIUOegfYhviqxWCTG2oC1WBe4DlKxUfMRs8Z9e7+QSTPrjJ2/vz8sDaSoE0IImecaGtgsXUcHHxOL\ngTVrgHXrhC0gdAwPsy14ly8L4/q6iI2pxlB4oxCnm04LCg37OvgiXZYOL3svU14RIfOWUTN2Z8+e\nxWuvvYZHH30UXl7CX65169ZN28lNBc3YEULIFI2OAvn5wLlzwrivLyth4qV/sMVxbDCXm8sGdxqG\nuohVd1cjqzILvaP8lJ6F2AIpISlY6buSCg2T+4JZZ+zOnz+P7OxslJSU3LbHrrGxcconQQghZJap\nqACys1nmq4alJZCSAsTFGSxh0tPDll1raoTxJUtYXoVuF7Hh8WHkVuei7GaZ4LmhLqFIk6XB2doZ\nhJDJMWrGzs3NDV9++SU2bdpkjnMyCZqxI5NB+2CIseb9vdLfzwZ0168L4xERrISJk5Pew9RqlhhR\nVASMj/NxJye27Boezsc4jsPV9qvIqc7B8Dg/pWcjtcGWsC1Y6rV03tRJnff3CzEZs87Y2dnZISkp\nacpvZm6UFUsIIUbiOLbkmp/Pukho2NuzFhCRkQZLmLS2si14LS18TCRivV03bBCWMOkb7UNWZRaq\nuqsEr7HEcwm2hG2BnaUdCLmfzEhW7Mcff4wzZ85g7969t+2xExuYjp9pNGNHCCFGam9nI7OJW2tW\nrAA2bmSb4/QYH2dFhktL2YydhpcX24Ln68vH1JwaZ5vPouBGAcZUY9q4k5UTtkdsR4RbhAkviJC5\nx1TjFqMGdoYGbyKRCCrdtPdZhAZ2hBByF0olK19y4oSwhIm7OythEhho8NDaWraXrrubj0mlQFIS\nkJAASCR8vH2oHZnyTDT28wNHEUSI841DSnAKrKRWprwqQuYksy7F1k5s5EfIPEP7YIix5s29UlfH\nZum6uviYRAKsXQskJrJRmh7Dw0BeHnDpkjAeFMTGgm5ufEypVuJ4w3GU1JcICg172HogXZYOfyd/\n013PLDVv7hcyZxg1sAsKCprm0yCEEGIWIyOsFdiFC8K4vz8bmXl66j2M41hL2Jwc4FZnSQCAtTVr\nOLFsmXALXmNfIzLkGegY5mvfSUQSJAYmYm3A2jlXaJiQucLoXrFzDS3FEkKIDs3I7PBhYHCQj1tZ\nsX10sbEGkyP6+tiya5Uw3wGLF7O8Cnt7PqZQKlBwowBnm88KCg37OfohXZYOTzv9A0dC7ndmXYol\nhBAyh/X1AYcOAZWVwviiRWxk5uio9zC1GjhzBigsZL1eNRwdWeUTmUz4/MquShyqPIQ+RZ82Zimx\nREpwCuJ846jQMCFmMK8HdlTuhBiL9sEQY82pe8XQyMzBAdi2jQ3sDLh5E8jIAJqb+ZhIxGoTp6Sw\niT6NobEhHK4+jCvtVwSvEe4ajh0RO+Bkrb/23f1gTt0vZEaYutzJvB/YEULIfamtjSVHTByZxcay\nkZm1td7DlErg6FGWKKtbwsTDg5Uw8dfJd+A4DpdvXkZuTa6g0LCthS22hm1FlGfUvCk0TMh00UxA\nHThwwCSvZ9Qeu9raWrz00ku4dOkSBnX2ZohEIjQ0NJjkREyN9tgRQu5L4+NsZHby5N1HZhMYSpRd\nt44ly+qWMOkd7UWmPBM1PcLeYdFe0dgcthm2FrYmuiBC7g9m3WP36KOPIiwsDG+++eZtvWIJIYTM\nEvqKy0kkrLjcmjXCkZkOQ4myAQEsUdbDg4+pOTXONJ9BQW0BxtV87zBna2fsiNiBMNcwU14RIWSS\njJqxc3R0RE9PDyQGPhRmI5qxI5NB+2CIsWblvWKouFxgIBuZubvrPYzjgIoK1hp2YqLspk2s8YTu\nSurNwZvIkGegeYBf3hVBhHi/eGwI3gBLiU7vMAJglt4vZFYy64zdunXrcPHiRcTGxk75DQkhhJgI\nxwFXrrASJsP8HjeDxeV09PezRFm5XBjXlyirVCtxrP4Yjjcch5rjl3c97TyRLkuHn6OfKa+KEDIF\nRs3YPffcc/jqq6/w4IMPCnrFikQivPrqq9N6gveKZuwIIfNaTw9bdq0R7nHTW1xOB8cBZ88CBQWA\nQsHHDSXKNvQ1IEOegc7hTm1MIpIgKSgJa/zXQCKeOys5hMxmZp2xGxoawo4dOzA+Po6mpiYALBuK\nsp0IIcTM1Grg1CmgqIglSmg4ObHichERBg/t6GAlTBobhfHYWFajWDdRVqFUIL82H2dbzgqeG+AU\ngHRZOtxt9S/vEkJmllEDu48//niaT2N6UB07YizaB0OMNaP3SksLS1ttbeVjIhEQHw+sXy8sLqdD\nqQSOHwdKSgAV37IV7u5sC15goPD58k45DlUdQr+iXxuzklhhY8hGxPrE0h/1k0CfLeRuzFbHrq6u\nTtsjtra21uALhISEmOxkTI3q2BFC5oWxMTZDd+oUW0vV8PJiJUx8fQ0e2tDAxoIdfMtWSCSsfEli\nIiDV+VdgcGwQOVU5KO8oF7yGzE2G7RHb4Wilv0MFIeTema2OnYODAwYGBgAAYrH+NjAikQgq3T//\nZhHaY0cImReqq9leut5ePiaVAsnJwOrVBkuYjI4C+fnAuXPCuJ8fGwt66rRs5TgOl9r6vQ2BAAAg\nAElEQVQuIa8mDyPKEW3czsIO28K3IdIjkmbpCJlmphq3GJU8MRfRwI4QMqcNDbFs1yvCNl0ICQF2\n7ABcXQ0eev06y3i99bc5AMDSku2ji40FdP9W7xnpQWZlJmp7hCszMQtisDl0M2wsqHYpIeZg1uQJ\nQuY72gdDjDXt9wrHAWVlQG4uqxysYWMDbN4MREcbLGEyMMBq0lVUCOMyGct4ddJp2arm1DjVdApF\nN4oEhYZdrF2QJktDiMvs3WYzl9BnCzE3GtgRQshs0d3NNsTduCGML13KBnV2dnoP4zjWNeLIEbYE\nq2FvzyqfREYKx4Jtg23IkGegZaBFGxNBhNX+q5EclEyFhgmZw2gplhBCZppKxXq7Hj3KUlg1nJ3Z\nsmuY4TZdnZ1sLFhfL4wvX866R+h2gRxXjeNo/VGcbDwpKDS8wH4B0mXp8HHwMdUVEUImiZZiCSFk\nPmhqYiOzmzf5mEjEEiOSk9nmOD1UKlbC5NgxYQkTV1dWwiQ4WPj8ut46ZMoz0TXSpY1JxVIkBSYh\nwT+BCg0TMk9MemCnVqsFjw1lzBIyl9A+GGIsk90rCgVQWAicOSMsYeLtzdJWvb0NHtrUxAoNt7fz\nMbEYWLMGWLcOsLDg46PKURypOYLzrecFrxHoFIh0WTrcbN2mfi3EIPpsIeZm1MDu/Pnz+MlPfoKy\nsjKM6mzgmM3lTgghZNaSy1naaj9fABgWFqzI8KpVwrRVHQoFawV29qxwLOjry2bpFiwQPr+iowLZ\nVdkYGOPTY60kVkgNTcVy7+VUwoSQecioPXZRUVFIT0/Hrl27YGtrK/iepojxbEN77Aghs87AACth\nUi4sAIywMNYOzMXF4KGVlaycne5Y0NIS2LABWLlSOBYcUAwguyobFZ3C9NiF7guxPXw7HKwcTHE1\nhBATMmsdO0dHR/T19c2pv+5oYEcImTUMpa3a2QFbtgBRUQZLmAwOAjk5t48Fw8PZWNDZWfdtOFxs\nu4i8mjyMKvn3sbe0x/bw7VjksciUV0UIMSGzJk888MADyM3NxZYtW6b8huZEvWKJsWgfDDHWpO8V\nQ2mrMTFAaiowYRVEg+OAS5eAvDxhOTtDY8Gu4S5kVmairrdO8DrLvZdjU8gmKjQ8Q+izhdyN2XrF\n6hoZGcEDDzyAxMREeHl5aeMikQiffvqpyU7G1KhXLCFkxtwpbXXHDtZBwoCuLrbsOrGcnb6xoEqt\nQmlTKYrriqFU86VSXG1ckRaRhmCXCemxhJBZxWy9YnUZGiCJRCLs27fPJCdiarQUSwiZMQ0NbJau\no4OPGUpb1aFSAaWlQHGxsJydiwsbC4aGCp/fOtCKg/KDaBts499GJEaCfwKSApNgIdH/PoSQ2Yd6\nxd4FDewIIWY3Ogrk5wPnzgnjvr6shInOisdEzc1sLNjGj9EgFvPl7HTHguOqcRTXFaO0qVRQaNjb\n3hvpsnR4OxgulUIImZ3MXqC4qKgIn376KZqbm+Hn54ddu3Zhw4YNUz4BQmYD2gdDjGXwXqmoYI1a\nB/jSIrC0BFJSgLg4gyVMxsZYObvTp40rZ3ej5wYyKzPRPdKtjUnFUqwPWo/V/qshFlFt0dmEPluI\nuRk1sPvggw+wZ88ePP3004iPj0dDQwMeffRRvPrqq3jmmWem+xwJIWT26u9nA7rr14XxiAiWturk\nZPDQ6mq2l663l48ZKmc3Mj6CvJo8XGy7KHiNYOdgpMnS4GrjaoqrIYTMcUYtxYaHh+Prr79GdHS0\nNnb58mU8+OCDqK6untYTvFe0FEsImVYcx5Zc8/NZ5WANe3tg61YgMtJgCZOhIVbO7soVYTwkhBUa\n1i1nx3EcrnVcQ051DgbHBrVxa6k1NoduRsyCmDlViooQop9Z99i5ubmhtbUVljo9CxUKBXx8fNDV\n1XWHI2cODewIIaZUL5ejJj8f4vFxqEdGEKpSIVA3wwEAVqwANm4EbPSXFuE44PJlIDcXGB7m4zY2\nrITJ0qXCsWC/oh/ZVdm43imcDYz0iMTWsK1UaJiQecSse+zWrFmDX/ziF/jv//5v2NnZYXBwEP/1\nX/+FhISEKZ8AIbMB7YMhd1Ivl6P644+RYmGB4tOnsWFsDAXj40BMDALd3QE3NzbVdodOPD09bNm1\npkYYX7oU2LyZ1afT4DgO51vP40jNEShU/Gygg6UDtkdsx0L3hSa+QjJd6LOFmJtRA7t3330XDz/8\nMJycnODq6oru7m4kJCTgH//4x3SfHyGEzLia/HykDA+zvl43bwLOzkiRSlFYV4fAb38bSEwEpPo/\nTtVq4NQpoKgIGB/n487ObAteeLjw+Z3DnciUZ6K+T1jQONYnFhtDNsJaam3qyyOEzCOTKnfS2NiI\nlpYW+Pj4wN/ffzrPa8poKZYQYhJDQyj+yU+QPLFasKMjiuPikLx3r8FDW1uBjAz2/xoiEUuMWL+e\nJc1qqNQqnGg8gWP1xwSFht1s3JAuS0egc6CprogQMgtN+1Isx3HaDblqNauT5OvrC19fX0FMbCCF\nnxBC5jSd/q7qmzf5uETCshx8fKB2c9N76Pg4m6E7dYrN2Gl4ebESJrc+RrWa+5uRIc/AzSH+fcQi\nMdb4r0FSUBKkYqMrUxFC7nMGPy0cHR0xcKsek9TAEoNIJIJKt1UOIXMU7YMhAjdvsg1xjY0AgNCQ\nEBRcuoQUHx8US6VI9vVFgUKBsJSU2w6trWWFhnt6+JhUyooMr17NxoUaY6oxFN0owqmmU+DA/6Xu\n4+CDdFk6FtgvmK4rJGZCny3E3AwO7MrLy7Vf19bWmuVkCCFkRo2NAUePsr5eOlNtgeHhQHo6Cqur\ncfnaNag9PRGWkoJAmUz7nOFhIC8PuHRJ+JLBwawd2MTJvZruGmRWZqJ3lC9iZyG2wIbgDYj3i6dC\nw4SQe2LUHrs//OEP+NWvfnVb/M0338QvfvGLaTmxqaI9doSQSZHLWaHhvj4+JpGw/q6JiQb7u3Ic\ncPUqq0s3NMTHra1ZtmtMjLCEyfD4MPJq8nCpTTgCDHUJxY6IHXCxcQEh5P5j1jp2Dg4O2mVZXS4u\nLujRXW+YRUQiEfbt24fk5GSaBieEGNbXx0ZlFRXCeGAgm2rz8DB4aG8vW7GdWKd98WJWo9jeno9x\nHIfyjnLkVOVgaJwfAdpIbbA5bDOivaKp0DAh96Hi4mIUFxfjwIED0z+wKywsBMdxSEtLQ1ZWluB7\nNTU1+O1vf4v6+noDR88smrEjk0H7YO5DajVr0FpUxJZgNWxtgdRUIDpab+eI4uJirFuXjDNnWI9X\n3UMdHdlYMCJCeEzfaB8OVR1CZVelIB7lGYUtYVtgb2kPMj/RZwsxllkKFD/55JMQiURQKBR46qmn\nBG/u5eWFt99+e8onQAghZtfczDIc2tqE8WXLgE2b2OBuArm8Hvn5NTh37jLee08NN7dQuLuzEiQi\nERAXB6SkAFZW/DEcx+Fsy1nk1+ZjTMWPAB2tHLE9fDtk7rKJb0MIIVNi1FLs7t278dlnn5njfEyG\nZuwIIbcZHQUKCliPV93PBw8PNtUWqL9WnFxejw8/rEZrawoaG9mhSmUBYmLCEBkZiLQ0YGJpz46h\nDmTIM9DY3yiIr/RdiZTgFFhJrUAIIRpm3WM3F9HAjhCipclwyM0FBgf5uIUFkJR0ex2SCQ4cKMSp\nUxswMsLHRCIgJqYQf/jDBsGhKrUKxxuO41j9Mag4vhyUu6070mXpCHAKMOWVEULmCbP2iu3r68P+\n/ftx9OhRdHV1aYsTi0QiNDQ0TPkkCJlptA9mHuvuBg4dur1Ja3g4sG0b4GI4C3VoiI0FS0vFGB1l\nsd7eYgQGJkMmA3x8xIJBXVN/EzLkGWgfatfGJCIJ1gasRWJgIhUavg/RZwsxN6M+ZZ577jk0Njbi\nlVde0S7L/s///A++/e1vT/f5EULIvVEqgRMngJIS9rWGgwNLWV20SG9yBMAm+MrK2KBuZAQQi9kf\ns1Ip4OfHlzCxtGTxMdUYCmoLcKb5jKDQsJ+jH9Jl6fC085y+6ySEEB1GLcV6eHigoqIC7u7ucHJy\nQl9fH5qbm5GWloYLFy6Y4zwnjZZiCbmP3bjBZuk6O/mYSASsXAls2CDMcJigq4uVMNFtDdvZWY+W\nlmosXJii7e+qUBTg8cfDIHYfQ1ZlFvoUfP07S4klUoJTEOcbR4WGCSFGMetSLMdxcHJyAsBq2vX2\n9sLb2xtVVVVTPgFCCDGZoSHW/qGsTBj38WHJET4+Bg9VqdgE37Fjwgk+Z2fgsccCoVYDBQWFGBsT\nw9JSjdVJPihXn8flK5cFrxPmGoYdETvgbO1syisjhBCjGDWwW7p0KY4dO4aUlBSsXbsWzz33HOzs\n7CCTUao+mR9oH8wcx3HAhQtAfj4EGQ5WVqwGSWwsIDY8c9bQwKqfdHTwMbEYWLWK9Xi1tATk1aPg\n3CpQdfUaHAMcUVs+DntPvv6crYUttoRtwRLPJVRomGjRZwsxN6MGdu+//7726z//+c/Ys2cP+vr6\n8Omnn07biRFCiFFu3mRrp43CsiJYvBjYsoXtqTNgZISNBc+fF8Z9fIC0NMDbmz2WV8vxcdHH4II5\nXBZdhsRCAuVlJWIiY+Du446lXkuxOXQz7CztTHxxhBAyOUbtsTt9+jTi4+Nvi585cwYrV66clhOb\nKtpjR8g8NzYGHD0KlJayLhIaLi4s2zU83OChHAeUl7NOYrrVTywt2Ra8lSuFE3xvf/k2LtlcQl1v\nnaCEiVubG974/95AuJvh9yKEEGOYdY/dxo0b9faK3bJlC7q7u6d8EoQQMimVlUB2NmvWqiGRAAkJ\nwLp1rD6dAb29LK9i4hZhmYyNB29tJ9ZqHWjFsYZj6PDsEMR9HXwRYxFDgzpCyKxyx4GdWq3Wjh7V\nun8Rg/WKlUqpJhOZH2gfzBzR3w/k5AAVFcJ4YCBLjvDwMHioWg2cOsVaw46P83EHBzagW7hQWP1k\nTDWG4rpilDaWYmhsSBsfrRrF6sTVcLJ2gl07Lb2SO6PPFmJudxyZ6Q7cJg7ixGIxXnrppek5K0II\n0aVWA2fOAIWFbAlWw9aW9XbVFJYzoKWFJUe0tvIxkYjlVKSkANbWwudXdVXhUNUh9I6yGcGQkBBc\nrriM0BWhULmr4GTtBEWVAinrU0x5lYQQMmV33GNXV1cHAFi3bh1KSkq0s3cikQgeHh6w1dMoe7ag\nPXaEzBPNzSw5QndUBgDLlrFB3R0+hxQKNkN3+rSwNaynJ/T2dx0cG8Th6sO42n5VEA92DoZMIsOF\nigsYU4/BUmyJlOUpkIVRZQBCiGlQr9i7oIEdIXPc6CiboTt7Vjgq8/Bgy66BgXc8XC5n2/D6+LrB\nkEpZ+ZKJrWE5jsPFtovIq8nDqHJUG7e1sP3/27vv6Kiuc23gzxSNehcS6kIICWSK6AhQAQw2zYlJ\n4hjH2Bhf28uxfRPflC+JTQvO8nLidt3iGydxI+Cybu6yBZiqSpUBgakqgAqSUO91NHO+P7ZVzghJ\nM2g0Mxo9v7W84tlzzpw9ZCO/2uV9cc/EezA9YDpTmBDRiLLo4YkNGzbctgMAmPKE7AL3wdiQgY6s\nqtVAUpI4INE3KjPQ1CS24V2+LG+PjBTxoI+PvL2qpQp78vagqKFI1j4jYAbuiboHLg7yGUGOFTIF\nxwtZmlGB3cSJE2WR5K1bt/C///u/+NnPfjainSOiMaa2VhxZvXZN3h4VBaxeLVKZDECSgNOnRV66\njo7edhcXkc5u2jT5NrwufReOFh9FVlGWLIWJj7MP1kSvQaR3pLm+FRGRxdzxUuzp06exbds27Nmz\nx9x9MgsuxRKNIl1dwPHj/et5ubuLqCw2dtDDEZWV4nCEYY7igbbhFdYXYk/eHlS39taSVSqUWBS6\nCInhiXBQDZwuhYhoJFh9j11XVxe8vb1vm9/OFjCwIxolCgvF4Yjq3iALCoXIErx0qSgLNgCtVsSC\nx47JcxT7+opl1wkT5Ne3adtw6PohnC0/K2sP8QjB2ui1CHALMMMXIiIynUX32B05ckS2cbilpQWf\nffYZ7rrrrmF3wFSNjY24++67ceXKFZw6dQqxsbEW7wPZH+6DsYKWFuDQIeDcOXl7YKA4shoUNOjt\n16+LeLBvjnSVCli8GEhIEFvyukmShEtVl/BN/jdo0fbmpHNUOWJZ5DLMCZoDpWLgWrJ9cayQKThe\nyNKMCuwef/xxWWDn6uqKuLg47N69e8Q6NhAXFxfs27cPv/nNbzgjRzQaSRKQkyOCura23nZHRzFD\nN3euvJ6XgZYW4MAB4Lvv5O1hYSIeNMxRXNdWh735e1FQWyBrn+I3BSsnrYSHo8dwvxERkc0wKrDr\nzmdnC9RqNfz8/KzdDbIz/I3aQiorxTRbcbG8PTZW7KXzGDjIkiTg/HkR1PWNB52cxD66WbPk2/D0\nkh4nb55E2o00aPW9pSY8HD2watIqTPabfEdfgWOFTMHxQpZmdE2w+vp67N27F2VlZQgKCsKqVavg\nPcgJNSKiHlotkJEhDkj03Qzn5SVOu04avN5qTY2IB2/ckLdPnSriQTc3eXtZUxlSclNQ3tyb1FgB\nBeYFz8PSCUvhqB543x4R0Whm1KaS1NRURERE4K233sK3336Lt956CxERETh8+LBJD3vnnXcwZ84c\nODk54bHHHpO9V1tbi/vvvx9ubm6IiIiQLfO+8cYbWLJkCV577TXZPUwYSuaSnp5u7S7Yr7w84N13\ngaNHe4M6pVJshHvmmUGDOp1OxIN//as8qPPyAn72M+DHP5YHdZ26Tuwv2I8PznwgC+oCXAPw+KzH\nsXLSymEHdRwrZAqOF7I0o2bsnnnmGfztb3/DAw880NP25Zdf4tlnn8XVq1eNflhwcDA2b96MAwcO\noK3vWsr3z3ByckJlZSVycnKwevVqzJgxA7GxsXj++efx/PPP9/s87rEjsmGNjSLJsGGm4LAwcWTV\n33/Q24uLRQqTqqreNqUSWLBAVI/QaOTX51bnYl/+PjR09JaaUCvVSI5IRnxIPFTKgZMaExHZC6PS\nnXh5eaGmpgaqPtnetVotxo0bh/r6epMfunnzZty8eRMffvghAHHK1sfHB5cuXUJUVBQA4NFHH0VQ\nUBBefvnlfvevWrUK58+fR3h4OJ566ik8+uij/b+YQoFHH30UERERPd8hLi6uZ79D929RfM3XfG3m\n13o90t9/Hzh7FskhIeL9wkJAo0Hy008DcXFIz8gY8P62NuCNN9KRlwdERIj3CwvT4esL/PrXyQgM\nlF/f1NGEV3e9iqKGIkTERYjrzxUiyC0Iv/7Zr+Hj7GNbfz58zdd8zdffS09P7znH8PHHH1suj91z\nzz2HqKgo/OIXv+hpe+utt5Cfn4+3337b5Ie++OKLKC0t7QnscnJysHjxYrS09KYheP3115Geno6v\nv/7a5M8HmMeOyCpKS8VmuPJyeXtcHLBiRf9MwX0MVElMoxGHZefNkx+WlSQJZ8rP4PD1w/3qu94b\ndS+m+U/jdg0iGjUsmsfu7NmzeP/99/HnP/8ZwcHBKC0tRWVlJebPn4+EhISeDmVmZhr1UMMfts3N\nzfAwOA3n7u5us8mPyf6kp6f3/DZFd6C9HUhNBb79VkRo3fz8xLLr9zPnA6mvF5XE8vPl7TExwKpV\ngKenvL2ypRIpuSkoaZSXmogbH4cVE1f0q+9qThwrZAqOF7I0owK7J554Ak888cSg15jym7FhROrm\n5obGxkZZW0NDA9zd3Y3+TCKyAkkSe+j27wf6/iKmVgOJicCiRSJr8AD0euDkSSAtTRyc7ebuLgK6\nyZP713fNLMrEseJjsvquvs6+WBO9BhO8DUpNEBGNMUYFdhs3bjTrQw2DwOjoaHR1daGgoKBnj935\n8+cxderUYT1n27ZtSE5O5m9LNCSOkTtQWwvs2wcUyBP/IipKRGU+PoPeXlYmDkf0XbVVKIA5c4Bl\ny0R+ur5u1N3Anrw9qGmr6WlTKpRYHLYYieGJUCuNzt40LBwrZAqOFxpKenq6bN/dcBldKzYzMxM5\nOTk9++AkSYJCocAf/vAHox+m0+mg1Wqxfft2lJaW4oMPPoBarYZKpcL69euhUCjw97//HWfPnsWa\nNWtw4sQJTJky5c6+GPfYEY0MnU4UZ83MBLq6etvd3ICVK0Wy4UFm8Ds6xAzdqVPyVVt/f1E5IjRU\nfn2rthWHrh1Czq0cWXuoRyjWxqyFv+vgp2uJiEYDi+6xe+655/DFF18gISEBzs7Od/ywHTt24I9/\n/GPP6507d2Lbtm3YsmUL3nvvPWzatAn+/v7w8/PD+++/f8dBHZGpuA/GSIWFYjNc3xwkCoUoA7Z0\naf9pNgO5uWKSr6E3IwnUaiA5GYiPl6/aSpKEC5UXsL9gP1q1rT3tjipHLJ+4HLMDZ1vlcATHCpmC\n44UszagZO29vb1y6dAlBQxTltiWcsSNT8IfvEFpbgYMHgXPn5O2BgeJwRHDwoLc3NQHffNM/pd3E\niaLwhOGqbV1bHfbk7cG1umuy9thxsVgZtRLujtbbf8uxQqbgeCFjmStuMSqwmz59OlJTU0dVjVYG\ndkRmIEkimDt4UF6gdaAcJLe5/fRp4PBhsQTbzcVFlAKbNk2+aqvT63Di5glkFGbI6rt6Onpi1aRV\niPGLMee3IyKyGRZdiv3HP/6BJ554Ag899BACAgJk7yUmJg67EyOFhyeIhqGyUiy7FhXJ22NjRVRm\nkKLIUEWFOBxx86a8feZMYPny/intShtL8XXu16hoqehpU0CB+SHzsSRiCeu7EpFdssrhiffffx+/\n+MUv4O7u3m+PXUlJyQB3WRdn7MgUXC7pQ6sVBVqPH++t7QqIAq2rVgHR0UPenpkpzlf0vd3XV6za\nTjDISNLR1YHUG6nILs2GhN6/s+PdxuO+mPsQ5G5bW0A4VsgUHC9kLIvO2L3wwgvYs2cPli9fPuwH\nEpENy88Xpxvq6nrblEpg4UIgKQlwcBj09uvXReGJ2treNpUKWLwYSEgQByX6ulp9Ffvy96GxozeP\npYPSAUsmLMGCkAVQKgZe5iUiov6MmrELCwtDQUEBNBqNJfpkFpyxIzJBY6NIMmx4uiEsTEyz+Q+e\nUqSlBThwAPjuu/63r10LjBtn8LiORnyT/w2uVF+RtUf5RGH1pNXwdva+029CRDQqWfTwxEcffYTs\n7Gxs3ry53x475SAbp62JgR2REfR6UQYsNVV+usHZWWyEmzlz0Jx0A52tcHISt8+aJb9dkiScLjuN\nw9cPo0PX+zxXB1fcG3UvpvpPZX1XIhqTLBrYDRS8KRQK6HS6275nbQqFAlu3buXhCTLKmNwHU1Ym\n1k3LyuTtcXEiKnN1HfT2mhpx+40b8vapU8XZCjc3eXtFcwVS8lJws1F+mmJW4Cwsj1wOZ4c7z5Fp\nSWNyrNAd43ihoXQfnti+fbvl9thdv3592A+yhm3btlm7C0S2p71dzNB9+6289IOfn1h2jYgY9Had\nDjh6FMjKkhee8PISOekmTZJfr9VpRX3XkmPQS72nKfxc/LAmeg0ivAZ/HhGRPeuegNq+fbtZPs/o\nkmIAoNfrUVFRgYCAAJtdgu3GpVgiA5Ik9tDt3y8yBndTq4HERHFAwvB0g4HiYpHCpG/hCaUSWLBA\nVI8w3IZ7ve469uTtQW1b72kKlUKFhPAELA5bbLH6rkREts6ip2IbGxvx7LPP4rPPPkNXVxfUajUe\nfPBBvP322/D09Bx2J4hohNXVidOu+fny9oFKPxhoaxNJhs+ckbcHBQH33QeMHy9vb9W24kDBAZyv\nOC9rD/cMx5roNRjnanCagoiIzMKoabfnnnsOLS0tuHjxIlpbW3v+97nnnhvp/hFZhDmTQ9oUnU6s\nmb77rjyoc3MDfvxj4OGHBw3qJAm4eFHc3jeo02jEPrr/+A95UCdJEs7fOo93st+RBXVOaiesjV6L\njXEbR31QZ7djhUYExwtZmlEzdvv378f169fh+v1m6ujoaHz00UeIjIwc0c4R0TAUFYnTDX3XTRUK\nYM4cYNkycXR1EPX1ovCE4SRfTIzIU2w4WV/bVos9eXtwvU6+J3eq/1TcG3Uv3DQGpymIiMjsjArs\nnJ2dUVVV1RPYAUB1dTWchvgPg7WxpBgZy67GSGsrcOgQkJMjbx8/XiSVCw4e9Ha9Hjh5EkhLE1Uk\nurm7i4Bu8uT+9V2PlxxHRlEGuvS9pym8nLywetJqTPI1OE0xytnVWKERx/FCQ7FKSbGXXnoJH3/8\nMX71q18hPDwchYWFeOONN7BhwwZs3rzZbJ0xJx6eoDGnO6ncoUMiuOum0QBLlwLz5omTDoMoKxOH\nI8rLe9sGm+QraShBSl4KKlsqe6+HAvGh8UiOSIZGNXqSmhMRWZNF89jp9Xp89NFH+Ne//oXy8nIE\nBQVh/fr12LRpk80mE2VgR6YY9bmmqqrEsmtRkbx9yhRg5UrAw2PQ2zs6xAzdqVPyDCj+/mKSLzRU\nfn17VzuOXD+C02WnZfVdA90CcV/MfQh0DxzuN7JZo36skEVxvJCxLHoqVqlUYtOmTdi0adOwH0hE\nZqTVApmZwPHj4qBENy8vsW4aHT3kR+TmigOzDQ29bWq1SF8SHy9qvXaTJKmnvmtTZ2/KFAelA5ZO\nWIr5IfNZ35WIyIqMmrF77rnnsH79eixcuLCn7fjx4/jiiy/w5ptvjmgH7xRn7MjuFRSI0w11db1t\nSqWIxpKS+ieVM9DUBHzzTf/ysANlQGnsaMS+/H24Wn1V1h7tG41Vk1bBy8lrON+GiGhMs+hSrJ+f\nH0pLS+Ho6NjT1t7ejtDQUFT1PXFnQxjYkd1qahJJhi9dkreHhorKEQb1nA1JEnD6tMhL17c8rIuL\nSGEybZr8cIRe0uPb0m9x5MYRdOo6e9rdNG5YGbUSseNibXZLBhHRaGHxpVi9Xi9r0+v1DJzIboyK\nfTB6vSgDlpoqj8icnUVt15kz5RHZbVRUiMMRN+XlWjFzpvgIFxd5+63mW0jJTYe2KRsAACAASURB\nVEFpU6msfXbgbNwdefeoqe9qTqNirJDN4HghSzMqsFu8eDFefPFF/OUvf4FSqYROp8PWrVuRkJAw\n0v0bFqY7IbtRViYOR5SVydtnzABWrAD6pCK6ne6teMeOifiwm6+vmOSbMMHgep0WGUUZOF5yXFbf\ndZzLOKyNWYswz7DhfiMiIoKV0p2UlJRgzZo1KC8vR3h4OIqLixEYGIiUlBSEGh6XsxFciiW70NEh\nZuiys+XHVQeKyG7j+nURE9b2lmuFSgUsXgwkJPQvD3ut9hr25O1BXXvv3j2VQoXE8EQsClvE+q5E\nRCPAonvsAECn0yE7OxslJSUIDQ3F/PnzoRwiJ5Y1MbCjUU2SgCtXxOmGpt7Tp1CrRTS2aFH/iMxA\nSwtw4ADw3Xfy9rAwkcJknEFlr5bOFhy4dgDfVchviPCKwJroNfBz8RvONyIiokFYPLAbbRjYkSls\nah9MXZ3IP2JYyysyUhxX9fUd9PbuPMUHDwJtbb3tTk5iH92sWfKteJIk4dytczh47SDaunpvcFY7\nY8XEFYgbH8fDEX3Y1Fghm8fxQsay6OEJIrIAnQ44cQLIyJDX8nJzA+65B5g6dcjDETU14nBEYaG8\nfepUceLVzaBca01rDVLyUlBYL79hmv803Bt1L1w1g+/dIyIi28IZOyJbUFQkctJV9pbmGrSWlwGd\nDjh6FMjKArp6y7XCy0tM8k0yKNeq0+twrOQYMosyZfVdvZ28sTp6NaJ8oszxrYiIyEgWm7GTJAk3\nbtxAWFgY1EPs6SEiE7W2itquOTny9vHjxeGIkJAhP6K4WMzS9U0pqVQCCxaI6hGGeYqLG4qRkpuC\nqtbeG5QKJeJDRH1XB5XDML4QERFZ05AzdpIkwdXVFc3NzTZ9WMIQZ+zIFBbfByNJwPnzYiNca2tv\nu0YDLFkCzJ8vorNBtLWJJMNnzsjbg4KA++4TsWFf7V3tOHz9ME6XnZZf7x6E+2Luw3g3gxvotrhn\nikzB8ULGstiMnUKhwMyZM5Gbm4spU6YM+4FEY15VlVh2NdwIN3kysHIl4Ok56O2SJIpO7N8PNDf3\ntms0wNKlwLx58phQkiRcrrqMbwq+QXNn7w0alQbLJizD3OC5rO9KRGQnjFpbXbJkCVauXImNGzci\nNDS0J6pUKBTYtGnTSPfxjjFBMRnLImNEqxWb4I4dE5viunl6AqtWATExQ35Efb2ICQ0PzMbEiI8w\njAkb2huwN38v8mry5Nf7xmDVpFXwdBo8iKT++POETMHxQkOxSoLi7oF5u5QHaWlpZuuMOXEplmxK\nQYGIyOp6k/5CqQTi44GkpP4b4Qzo9cDJk0BamvzArLu7COgmT+5f3zW7NBupN1Jl9V3dNe5YOWkl\npvhNYQoTIiIbwjx2Q2BgR6YYsX0wTU0iS/DFi/L20FBxOCIgYMiPKCsThyPKy3vbFApg7lyx9Gp4\nYLa8qRwpeSkoa+otP6aAAnOC5mBZ5DI4qQc/YUuD454pMgXHCxnL4nnsampqsHfvXty6dQu//e1v\nUVpaCkmSEGLEqT2iMUevB06fBo4cEWXBug2UJfg2OjrEDN2pU/JqYv7+4nCE4V+9Tl0n0gvTcfLm\nSVl9V39Xf6yNXotQT9ss/0dEROZj1IxdRkYGfvSjH2HOnDk4duwYmpqakJ6ejtdeew0pKSmW6KfJ\nOGNHVlNeLoqzlpbK26dPF4mGXYdO+pubK4pPNDT0tqnVIn1JfLyo9dpXfk0+9ubvRX17fe/1SjWS\nwpOwMHQhVEqDG4iIyKZYdCk2Li4Or776Ku6++254e3ujrq4O7e3tCAsLQ2XfhKo2hIEdWdxAU2y+\nviJLcGTkkB/R2CjKw165Im+fOFF8hI+PvL25sxn7C/bjYqV8qXeC1wSsiV4DX5fBy48REZFtsOhS\nbFFREe6++25Zm4ODA3R9T/YRjWLD2gcjSSIS279fRGbd1GogIQFYtEj8+xAfcfq0yEvXd+XWxUWU\nAps2rX9915xbOTh47SDau9p7r3dwwYqJKzAjYAYPR4wQ7pkiU3C8kKUZFdhNmTIF+/fvx7333tvT\nduTIEUybNm3EOkY0KtTXizXTPHk6EURGiik236FnzCoqxOGImzfl7TNniu14Li7y9urWaqTkpqCo\noUjWPiNgBlZMXMH6rkREY5hRS7EnT57EmjVrsGrVKnz55ZfYsGEDUlJS8NVXX2HevHmW6KfJuBRL\nI0qnA06cADIy5PlHXF3FFNvUqUMejtBqgcxMkdZO33vWAb6+wNq1QESE/PoufReOFh9FVlEWdFLv\nbLm3kzfWRK/BRJ+JZvhiRERkDRZPd1JaWoqdO3eiqKgIYWFhePjhh236RCwDOxoxxcXicETf/aUK\nBTB7NrBsGeDsPORHXL8uPqK2trdNpQIWLxart4Yrt0X1RUjJS0F1a3VPm1KhxMLQhUgKT2J9VyKi\nUc4qeez0ej2qq6sxbtw4m9+/w8COTGHUPpjWVrEJ7uxZeXtAgJhiM+IXnZYWkdbuu+/k7WFh4iPG\njZO3t2nbcOj6IZwtlz8zxCMEa6PXIsBt6Dx4ZF7cM0Wm4HghY1n08ERdXR3+8z//E1988QW0Wi0c\nHBzwk5/8BG+99RZ8DI/p2RCWFCOzkCQRiR04IIK7bhqNyD+yYIG8OOsAH3HuHHDwINDW1ts+UFo7\nSZJwqeoS9hfsl9V3dVQ5YlnkMswJmsP6rkREdsAqJcV++MMfQq1WY8eOHQgLC0NxcTG2bNmCzs5O\nfPXVV2brjDlxxo7MorparJkWFsrbJ08GVq7sX5z1NmpqxOEIw4+YOlVsx3Nzk7fXt9djb95e5NfK\nC8JO8ZuClZNWwsPRw/TvQURENs2iS7Genp4oLy+HS5/jea2trQgMDERD3wyqNoSBHQ2LVgtkZYmT\nDX3T+nh6ioBu8uQhP0KnA44eFR/T1dXb7uUlDsxOmiS/Xi/pcfLmSaTdSINW33sgw8PRA6smrcJk\nv6GfSUREo5NFl2InT56MwsJCxMbG9rQVFRVhshH/cSMaDWT7YK5dA/bulZ9sUCrFkmtysliCHUJx\nsZilq6oy7iPKmsqQkpuC8ubegrAKKDA3eC6WTVgGR7XjHX83Mi/umSJTcLyQpRkV2C1duhQrVqzA\nI488gtDQUBQXF2Pnzp3YsGED/vnPf0KSJCgUCmzatGmk+0s0cpqaxD66i/IqDggJAdasAcaPH/Ij\n2trE+YozZ+TtQUGivqvhR3TqOpF2Iw0nb56EhN7f1AJcA7A2Zi1CPGz35DkREdkeo5Ziu3/b6HsS\ntjuY6ystLc28vRsGLsWSMYpyc3Ht4EEob9yAvqAAE8PCEO7nJ950cgLuvlukMRniFLgkAZcuieIT\nzb1nHaDRAEuXAvPm9T9fkVeTh715e9HQ0budQa1UIzkiGfEh8azvSkQ0hlgl3clowsCOhlJ07hwK\n3n4byyorxWwdgCNdXYiKi0P40qXAihX9TzbcRn29WLnNl591QEwMsGpV//MVTR1N2F+wH5eqLsna\nJ3pPxOro1fBxtt2T5kRENDIsuseOyC5IkqjflZcH5Ofj2r//jWUtLQCA9Pp6JHt5YZm7O1L9/RG+\nbt2QH6fXAydPAmlp8uIT7u4ioJs8uX8KkzPlZ3D4+uF+9V3vjboX0/yn2Xx+SOKeKTINxwtZGgM7\nsm+dnaLMQ36++KexsectZd/TroCo4RUWBqXH0OlEysrE4Yjy3rMOUCiAuXPF0quTk/z6qpYqpOSl\noLihWNYeNz4OKyaugIuDQUFYIiKiO8DAjuxPba0I4vLyRPI4wwDue3qVCvDwAHx9kezv31MKTD/I\nqdeODjFDd+qUmADs5u8vDkcYFp/o0nchqygLR4uPyuq7+jj7YG30WkzwnnDHX5Osg7MvZAqOF7I0\n7rGj0U+nA4qKeoO5mpqBr3VyAqKigOhoFOn1KPj8cyxz7E0lcqSjA1EbNyI8Jqbfrbm5Yi9dn0k/\nqNUifUl8vKj12ldhfSFSclNQ09bbH6VCicVhi5EQlsD6rkRE1MPie+yuXLmCL7/8EhUVFXj33Xdx\n9epVdHZ2Yvr06cPuBJHJmpt7A7nr18VU2kD8/YHoaJERODS053hqOAA4OyP1yBF8d/kypsfGImrZ\nsn5BXWMj8M03wJUr8o+dOFEkGjasqtembcPBaweRcytH1h7qEYq1MWvh7+p/h1+abAH3TJEpOF7I\n0owK7L788kv8/Oc/x7p167Br1y68++67aGpqwu9//3scPnx4pPtIJNY9y8p6Dj6grGzgax0cgAkT\neoO5Qcp+hcfEIDwmBsrb/PCVJOD0aZGXrm/c6OIiSoFNm9b/cMTFyovYX7AfLdqWnnZHlSOWT1yO\n2YGzeTiCiIhGlFFLsZMnT8Znn32GuLg4eHt7o66uDlqtFoGBgaiurrZEP03GpVg70N4uqkB0H3xo\naRn4Wi+v3kAuIkIEd8NQUSEOR9y8KW+fORNYvlwEd33VtdVhT94eXKu7JmuPHReLlVEr4e7oPqz+\nEBGRfbPoUmxVVdVtl1yVhhlXiYZDkoDq6t5ZueJikVPkdpRKICxMBHLR0YCf35BJhI2h1QIZGcDx\n4/JH+/oCa9eKmLEvnV6HkzdPIr0wvV9919WTViPGr/9ePSIiopFiVGA3a9YsfPrpp3j00Ud72j7/\n/HPMmzdvxDpGY4RWK06udu+Xq68f+FpX156DD5g4sX9OkWFIT09HaGgy9uwB6up621UqYPFiICFB\nHJToq7SxFCl5KbjVfKunTQEF5ofMx5KIJazvaqe4Z4pMwfFClmZUYPf2229j+fLl+Mc//oHW1las\nWLECeXl5OHjw4Ej3b1i2bduG5ORk/qWyNQ0NvYHcjRvy7L6GgoJ6Z+WCgswyK9dXbm4R9u69hoMH\nv4NSqUdk5ET4+YUDEBOCa9cC48bJ7+no6kDqjVRkl2bL6ruOdxuPtdFrEewRbNY+EhGR/UpPT0d6\nerrZPs/odCctLS3Ys2cPioqKEBYWhtWrV8Pd3Xb3DXGPnQ3R64GSkt69chUVA1/r6AhERopALipK\nlHEYIZcvF+HPfy7AzZvL0NUl2rq6jmD+/Cg89FA4Zs3qH0fmVudib/5eNHb05jxxUDpgyYQlWBCy\nAEoFtycQEZHpWCt2CAzsrKy1FSgoELNy164BbW0DX+vnJ2blJk0CwsP7J4QzM0kCLl4Etm9PRXX1\nUtl7/v7AggWp+K//krc3dTRhX/4+XKmW5zyJ8onC6kmr4e3sPaJ9JiIi+2bRwxNFRUXYvn07cnJy\n0NzcLOtEXl7esDtBdsCgDitu3pSXZuhLpRKnELpPsRomghvBLl67Bhw5IkqBNTf3zq61tqZj/vxk\n+PrKDwVJkoTTZadx+PphdOh6c564Orji3qh7MdV/KlOYjDHcM0Wm4HghSzMqsPvJT36CKVOmYMeO\nHXAy44Z1GuUGqcPaj7t7byAXGQkMUrZrJJSWinx0N270timVeqjVYpKws1OcfAUAjUYch61sqURK\nbgpKGktknzUrcBaWRy6Hs4OzpbpPRERkFKOWYj09PVFbWwvVCC+RmROXYkdIbW3vrNwgdVihUIjC\nqd0HHwICzH7wwRjV1UBqKnD5srzdwQEIDi7CpUsFcHVd1tPe0XEEP3skHBWORThWcgx6qTfniZ+L\nH9ZEr0GEV4SFek9ERGOFRZdi16xZg4yMDCxdunToi8m+mFKH1dlZHHiYNEn8r2EWXwtqbBT56HJy\n5PnolEpg1iwgKQlwdw9Hbi5w5EgqOjuV0Gj0iJ6vQWrjPtS21fbco1KoRH3X8ASolUZX4SMiIrI4\no2bsqqurER8fj+joaPj799a5VCgU+Oc//zmiHbxTnLEbBlPqsAYE9M7KhYT01GG1lrY24Ngx4ORJ\n9Jx07XbXXcDSpb1LrgCQW5CLw2cO47vvvoMUIEHjp4FfkF/P+2GeYVgbvRbjXA1yntCYxT1TZAqO\nFzKWRWfsNm3aBI1GgylTpsDJyann4dw0bidGqA6rJWm1QHY2kJUlKpH1FRkJLFsGBBukl8styMVH\naR+hLrAOp/Wn4e7hjq7LXYhDHELCQrA8cjlmBc7iOCciolHDqBk7d3d3lJaWwsPDwxJ9MgvO2A2h\nuw5rXp5IS2LBOqzmpNeL5db0dKCpSf5eYCBw992iSIUhSZKw9Z9bkeOcg6ZO+Y1RDVF48+k34aZx\nG7mOExER9WHRGbvp06ejpqZmVAV2ZOBO6rB2B3NmqsNqTpIEXLkiDkZUV8vf8/ERS6533dW/25Ik\n4Wr1VWQUZSC7PBvtIb3Te44qR0T7RmOi00QGdURENCoZFdgtXboU99xzDx577DEEBAQAQM9S7KZN\nm0a0gzQMptZh7U4SbOY6rOZ244ZIXVJaKm93cxOHImbN6p/jWC/pcbnqMjKLMlHZUgkAUELsB1Qq\nlFAVqTBv8TyolCpomi2bioVGF+6ZIlNwvJClGRXYZWVlISgo6La1YRnY2ZiGht5ZOSvXYTW38nKR\nXLigQN7u6AgsWgQsWNA/PZ5e0uNi5UVkFmWiulU+tRc9MRq3Sm4hcnYkyurKoFKq0JHfgWVLloGI\niGg0Ykmx0a5vHda8PKCycuBrHR3FbFz3zJzb6FhurK0F0tKACxfk7SoVMG8ekJDQP7OKTq/DhcoL\nyCzKlKUuAQCNSoN5wfMQHxKPm8U3ceTsEXTqO6FRarBs1jLERMWM8DciIiKSG/FasX1PveoH2osF\nefklW2LXgd2d1GGNjhb75kZRkunmZiAzEzh9Wr4dUKEAZswAlizpfyhXp9fhfMV5ZBVloa69Tvae\no8oR80PmY0HIArg4WC/HHhERkaERPzzh4eGBpu+PGarVt79MoVBAN1DlATKfUVCH1Zw6OoDjx4ET\nJ0Spr75iYkTqkj7pFAEAXfou5JTn4GjxUTR0NMjec1I7IT4kHvND5sNJffu9g9wHQ8biWCFTcLyQ\npQ0Y2F26dKnn369fv26RzlAf3XVYu4M5w1wefXl49C6vWqEOq7l0dYnZucxMMSnZV1iYSF0SFiZv\n1+q0OFt+FsdKjqGxQ16r1sXBBfEh8ZgXPA+OascR7j0REZH1GbXH7tVXX8Wvf/3rfu2vv/46/uu/\n/mtEOjZco3Ip1tQ6rN2zclaqw2ouer3YP5eW1v/gbkCAmKGbNEn+FTt1nThTdgbHSo6hubNZdo+r\ngysWhi7E3OC50KhGZ5BLRERjy4jvsevL3d29Z1m2L29vb9TV1d3mDusbFYHdKK3Dai6SJL764cP9\nz3x4eYk9dNOmyauUdXR14Nuyb3Gi5ARatPKkym4aNywKXYQ5QXPgoLKdJMpERERDsUiC4tTUVEiS\nBJ1Oh9TUVNl7165dY8LiO9HU1HvwYZTVYTWnkhLg0CGRJ7kvFxcgMRGYMwfou7Wzvasd2aXZOFFy\nAm1d8sMiHo4eWBy2GDPHz7zjgI77YMhYHCtkCo4XsrRBA7tNmzZBoVCgo6MDjz/+eE+7QqFAQEAA\n3n777RHvoKHs7Gz88pe/hIODA4KDg/HJJ58MeLjDJkiSyKTbPStXXj7wtQ4OYo9c9345G6nDak6V\nlSIXXW6uvF2jAeLjgYULRVaWbm3aNpwqPYWTN0+ivUteBNbT0RMJ4QmIGx8HtdKGxwAREZGFGLUU\nu2HDBnz66aeW6M+Qbt26BW9vbzg6OuIPf/gDZs+ejR/96Ef9rrPqUqwpdVi9vXtn5SIi5NNUdqSh\nQeyhO39efqBXpQJmzxazdH3T6rVqW3Gi5ASyS7PRoZPPano7eSMhPAEzAmZApRw96VuIiIgGYtFa\nsbYS1AHA+PHje/7dwcEBKlvIy2ZndVjNqbUVyMoCvv1WnHrta9o0UdPV27u3raWzBcdLjuPbsm/R\nqZPnOvF19kVieCKmBUyDUmE/y9JERETmMmorTxQVFWH9+vXIysq6bXA34jN2dlqH1Vw6O4GTJ4Fj\nx/pvI4yKEqlL+sToaOpowvGS4zhddhpavbwM2jiXcUgMT8Rd/neNWEDHfTBkLI4VMgXHCxnLojN2\n5vLOO+/go48+wsWLF7F+/Xp8+OGHPe/V1tbi8ccfx6FDh+Dn54eXX34Z69evBwC88cYb+Prrr7Fm\nzRr86le/QmNjIx555BF8/PHHlp2xM7UOa/es3Ciow2ouOh1w9iyQkSEqR/QVHAwsXy5WnLs1djTi\nWPExnCk/gy69fEovwDUAieGJiB0X21MFhYiIiAZm0Rm7//u//4NSqcSBAwfQ1tYmC+y6g7h//OMf\nyMnJwerVq3H8+HHExsbKPqOrqwv33Xcffv3rX2Pp0qUDPssske8YqMNqLpIEXLoEpKaKdHx9+fmJ\nXHSTJ/fGt/Xt9ThafBQ55TnQSfJ8fYFugUgMT8Rkv8kM6IiIaEywaB47c9u8eTNu3rzZE9i1tLTA\nx8cHly5dQlRUFADg0UcfRVBQEF5++WXZvZ9++imef/55TJs2DQDw9NNP44EHHuj3jDv+AxojdVjN\nRZJE1pbDh/sf+PXwAJKTgbi43kwtdW11yCrOwrlb56CX5PsQg92DkRSRhEk+kxjQERHRmDIql2K7\nGXY8Ly8ParW6J6gDgBkzZiA9Pb3fvRs2bMCGDRuMes7GjRsR8f26n5eXF+Li4nr2OnR/dnJSEnDr\nFtK/+AK4eRPJbm6AJCG9sFC8//396YWFgFKJ5KVLgUmTkF5RAXh49P+8MfS6qgpobU3GjRtAYaF4\nPyIiGc7OgItLOqZMAWbNEtd/tf8rXKi8AH24HnpJj8JzheL6uAiEeoTC6aYTgqQgRPtGW+X7vPnm\nm7cfH3zN1wav+/5csoX+8LVtv+Z44euBXnf/e+H38Ya52MSMXVZWFh544AGU95ny+eCDD7Br1y6k\npaXd0TMGjXzvpA5rdDQwYcKorcNqTjU1Ihfd5cvydrUaWLAAWLRIFMoAgKqWKmQWZeJi5UVIkP//\nEe4ZjqSIJEzwmmD1Gbr09PSev3REg+FYIVNwvJCx7GrGzs3NDY2N8gLuDQ0NcHd3N99Dx2gdVnNq\nagLS04GcHHk2F6USmDkTSEoScTAAVDRXILMoE5erLvcL6CK9I5EYnogIrwiL9X0o/MFLxuJYIVNw\nvJClWSWwM5ydiY6ORldXFwoKCnqWY8+fP4+pU6cO6zmp27djYnAwwtvajKvDGh0tDkDYQR1Wc2pv\nB44eBU6d6n8QODZW5KLz8xOvy5vKkVmUiSvVV/p9TpRPFJLCkxDqGWqBXhMREY09Fg3sdDodtFot\nurq6oNPp0NHRAbVaDVdXV6xbtw5btmzB3//+d5w9exYpKSk4ceLEsJ6X+eGHyHF3x4+TkhDeHXl0\nCwjonZWzszqs5qLVAtnZIqgzPEMyYYLIRRccLF6XNpYisygTuTW5/T4n2jcaSeFJCPYItkCv7wyX\nS8hYHCtkCo4XGkp6erps391wWTSw27FjB/74xz/2vN65cye2bduGLVu24L333sOmTZvg7+8PPz8/\nvP/++5gyZcqwnrft+4MPqTduIDww0O7rsJqLXg+cOyeWXQ1WyBEYKAK6yEixQl3SUIKMogwU1Bb0\n+5zJfpORFJ6EQPdAy3SciIholElOTkZycjK2b99uls8btZUnhqJQKCDdcw/g64v0iAgkb99ut3VY\nzUWSgKtXxcGI6mr5ez4+Ysn1rrtEQFdUX4SMogxcr7suu04BBWLHxSIxPBEBbgEW7D0REdHoNaoP\nT1jM/PmAQgG9vz+DuiEUFopcdDdvytvd3MShiFmzAKVSQmF9ITKKMlBYXyi7TgEFpvpPRWJ4Isa5\njrNYv4mIiKiXXUc72zIy4B4UhB9/X9WC+rt1SwR0BQYrqY6OIm3JggWAg4OEa3XXkFGYgZLGEtl1\nSoUS0/ynISE8AX4uBvsYRxHugyFjcayQKTheaCijeo+dpSU+8AAmLluG8JgYa3fF5tTVifJfFy7I\n21UqYN48ICEBcHaWkF+bj4zCDJQ2lcquUyqUiBsfh8Vhi+Hj7GPBnhMREdkP7rEzkrnWqu1NczOQ\nmQmcOSNP5adQADNmAMnJgKenhNyaXGQUZqC8WV4nTKVQYWbgTCwOWwwvJy/Ldp6IiMhOjepasZbA\nwE6uowM4fhw4cUIU3ugrJgZYtgwYN07C5arLyCzKREVLhewatVKNWYGzsCh0ETydeKKYiIjInBjY\nDYGBndDVBZw+LWbpWlvl74WFidQlIaF6XKq8hMyiTFS1VsmucVA6YHbQbCwKXQR3RzNWArEx3AdD\nxuJYIVNwvJCxeCrWCNu2betZux5r9Hqxfy4tDaivl7/n7y8CuolRelysvICvsjNR0yavzKFRaTA3\naC7iQ+PhpnGzYM+JiIjGDnMfnuCMnZ2RJFEO98gRoEK+mgpPT5GLLvYuHS5WfYes4izUttXKrnFU\nOWJe8DzEh8bDxYGl1YiIiCyBS7FDGIuBXUmJSF1SVCRvd3EBEhOBuFlduFh9DkeLj6K+XT6N56R2\nwvzg+VgQsgDODs4W7DURERExsBvCWArsqqrEDN3Vq/J2jQaIjwfmLejCpdqzOFp8FI0d8hphzmpn\nxIfGY17wPDipnSzYa9vCfTBkLI4VMgXHCxmLe+wIDQ2inuu5c2IJtptSCcyZA8Qv0iK36QzezzmG\nps4m2b0uDi5YGLoQc4PmwlHtaNmOExER0YjgjN0o1NoKHD0KZGeLU699TZsGLErsxLW2b3G85Dha\ntC2y9900blgYuhBzguZAo9JYsNdEREQ0EM7YGcHeTsV2dgInTwLHjom8dH1FRQGLkztQosvGJ3kn\n0KqV5zZx17hjUdgizA6cDQeVgwV7TURERAPhqVgj2dOMnU4HnD0LZGSIyhF9BQcDCUvaUaE+hZM3\nT6Ktq032vqejJxaHLcbMwJlQK+06jh8W7oMhY3GskCk4XshYnLEbAyQJuHRJ1HStlWclgZ8fsDCp\nDfWuJ/F/pSfRoZNP4Xk5eSEhLAFx4+OgUqos2GsiIiKyFs7Y2ahr18RJAHCaxwAAFSdJREFU17Iy\nebu7OzB/cQtafU/gdHk2OnXy+mA+zj5ICEvA9IDpDOiIiIhGCaY7GcJoDezKykQuuuvX5e1OTsCs\nBc3oCjyOnIpvodVrZe/7ufghMTwRU/2nQqlQWrDHRERENFwM7IYw2gK7mhqx5HrpkrxdrQamzWkC\nQo/hQs1pdOnlx2DHuYxDUkQSYsfFMqAbBu6DIWNxrJApOF7IWNxjZyeamsShiLNnRX3XbkolED29\nAaqIo7jQmIOuKnlAF+AagKSIJEzxmwKFQmHhXhMREZEtsusZu61bt9psupP2dpG25ORJQCtfVUV4\nTD00UVm43noOOkkney/QLRBJEUmI8Y1hQEdERDTKdac72b59O5diB2OrS7FdXSKxcFYW0CbPTIJx\nYbVwnpyFm9rz0Et62XshHiFICk9ClE8UAzoiIiI7w6XYUUavB86fB9LSgEZ5uVa4+VfDZUoWKhXf\nAZ3y/1PDPMOQFJ6ESO9IBnQjiPtgyFgcK2QKjheyNAZ2I0ySgNxckbqkqkr+ntqzEs5TMtHkdAnN\nkAd0EV4RSApPQoRXBAM6IiIiMgqXYkdQUZFIXVJSIm/Xu9yCU0wm2t0vQ2lwkHWi90Qkhici3Cvc\nch0lIiIiq+JSrA2rqBABXX6+vL1DUwaHqExIvlfRqQL6xnSTfCYhMTwRoZ6hFu0rERER2Q8GdmZU\nVyf20F24IJZguzUrb0IVmQmVfx6UGvk9Mb4xSIpIQpB7kGU7SzLcB0PG4lghU3C8kKUxsDODlhYg\nMxM4fRrQ9clO0qAohj40A85B1+DkJL9nit8UJIYnItA90LKdJSIiIrvFPXbD0NEBnDgBHD8OdH5f\nslWChAYUoX18BtxCbsDNrU+foMBd/nchISwBAW4BI9o3IiIiGj24x84I27ZtG5EExV1dwJkzYpau\npUW0SZBQjxto9MmAV3gRxnv2Xq+AAtMCpiEhLAHjXMeZtS9EREQ0enUnKDYXztiZQJLE/rnUVKC+\n/vs2SKhFAWrdM+AdfhO+vkB3dhKlQonpAdOREJYAXxdfs/aFzIv7YMhYHCtkCo4XMhZn7CxIkoCC\nAnHStaLi+zZIqEEeKp0z4BNehqgAeUA3c/xMLA5bDG9nb+t1nIiIiMYUztgN4eZN4NAhkZMOEAFd\nNa6iXJMB79BbCA5GTy46lUKFWYGzsChsEbycvIb9bCIiIhobOGM3wqqqRLWIq1fFawl6VOEybqoy\n4R1SidhQQP39n55aqcbswNlYFLYIHo4e1us0ERERjWkM7Aw0NADp6cC5c2IJVoIelbiIYkUmPAOr\nMTUC0Hyfi85B6YA5QXOwMHQh3B3drdltGibugyFjcayQKTheyNIY2H2vrQ3IygKys8WpVz10qMQF\nFCET7v61mDoBcHYW12pUGswLnof4kHi4alyt23EiIiKi7435PXZaLXDyJHDsGNDeLgK6CpxHEbLg\n7F2HyEjA/fvJOEeVI+aHzMeCkAVwcXAZ4W9AREREYwX32A2TTgfk5AAZGUBTE6BHF8qRg2Ichca9\nATGRgPf3B1qd1E5YELIA84Pnw9nB2bodJyIiIhrAmAvsJAm4fFnkoqupAXTQohxnUYJjUDo3IioS\n8PMTqUuc1c5YGLoQc4PnwkntNPSH06jFfTBkLI4VMgXHC1namArsrl8XuejKygAdOlGGMyjBMUDT\njIgIIDBQBHSuDq5YGLoQc4LmwFHtaO1uExERERnFrvfYbd26FcnJyYiOTsaRI8C1a0AXOlCGb1GC\nE5DULQgLA4KDAZUKcNO4YVHoIswOmg2NSmPtr0BERER2rruk2Pbt282yx86uA7tXXjkCR8eJqKsL\nRxfaUYpslOAE9Mo2BAcDYWGAgwPg4eiBRaGLMCtwFhxUDtbuOhEREY0x5jo8YdeBXXKyhHbdPoyf\n3Ygmz0J0KdoROB6IiAAcHQFPR08khCcgbnwc1MoxtSpNBrgPhozFsUKm4HghY/FUrBHyFP8JxTg1\nqltqMGPiBERGAi4ugLeTNxLCEzAjYAZUSpW1u0lERERkFnY9Y+f5iySoirsQGeaH1T+Mg6+zLxLC\nEzDNfxoDOiIiIrIZnLEzgqsboJ6uhq6sHuumrMNU/6lQKpTW7hYRERHRiLDrKMdF7QpfnQdW3JWA\n6QHTGdTRgNLT063dBRolOFbIFBwvZGl2PWM3ycUXkZE+CNMHWrsrRERERCPOrvfYbU3bio78Dmxc\nshExUTHW7hIRERHRbXGPnRH8K/2xbMkyBnVEREQ0Jtj1prOfP/BzBnVkFO6DIWNxrJApOF7I0uw6\nsCMiIiIaS+x6j52dfjUiIiKyM+aKWzhjR0RERGQnGNgRgftgyHgcK2QKjheyNAZ2RERERHbCrvfY\nbd26FcnJyUhOTrZ2d4iIiIj6SU9PR3p6OrZv326WPXZ2HdjZ6VcjIiIiO8PDE0RmxH0wZCyOFTIF\nxwtZGgM7IiIiIjvBpVgiIiIiK+NSLBERERHJMLAjAvfBkPE4VsgUHC9kaQzsiIiIiOwE99gRERER\nWRn32BERERGRDAM7InAfDBmPY4VMwfFClsbAjoiIiMhOcI8dERERkZVxjx0RERERyTCwIwL3wZDx\nOFbIFBwvZGkM7IiIiIjsBPfYEREREVkZ99gRERERkQwDOyJwHwwZj2OFTMHxQpbGwI6IiIjIToy6\nPXYVFRVYt24dNBoNNBoNdu3aBV9f337XcY8dERERjRbmiltGXWCn1+uhVIqJxo8//hjl5eX43e9+\n1+86BnZEREQ0WozZwxPdQR0ANDY2wtvb24q9IXvBfTBkLI4VMgXHC1naqAvsAOD8+fOYP38+3nnn\nHaxfv97a3SE7cO7cOWt3gUYJjhUyBccLWZpFA7t33nkHc+bMgZOTEx577DHZe7W1tbj//vvh5uaG\niIgI7N69u+e9N954A0uWLMFrr70GAJgxYwZOnTqFl156CTt27LDkVyA7VV9fb+0u0CjBsUKm4Hgh\nS7NoYBccHIzNmzdj06ZN/d575pln4OTkhMrKSvzrX//C008/jcuXLwMAnn/+eaSlpeFXv/oVtFpt\nzz0eHh7o6OiwWP9HgiWm6c3xjDv9DFPuM+baoa4Z7H17WBIZ6e9grs+/k88x91gx5jp7Hi/82WLa\ntWN5rAD82WLqtbY8Xiwa2N1///34wQ9+0O8Ua0tLC/79739jx44dcHFxwaJFi/CDH/wAn376ab/P\nOHfuHJKSkrB06VK8/vrr+O1vf2up7o8I/vA17dqR+stUWFg45LNtAX/4mnbtSIwXjhXzPoM/W2wD\nf7aYdq0tB3ZWORX74osvorS0FB9++CEAICcnB4sXL0ZLS0vPNa+//jrS09Px9ddf39EzoqKicO3a\nNbP0l4iIiGgkTZw4EQUFBcP+HLUZ+mIyhUIhe93c3AwPDw9Zm7u7O5qamu74Geb4wyEiIiIaTaxy\nKtZwktDNzQ2NjY2ytoaGBri7u1uyW0RERESjmlUCO8MZu+joaHR1dclm2c6fP4+pU6daumtERERE\no5ZFAzudTof29nZ0dXVBp9Oho6MDOp0Orq6uWLduHbZs2YLW1lYcPXoUKSkp2LBhgyW7R0RERDSq\nWTSw6z71+sorr2Dnzp1wdnbGn/70JwDAe++9h7a2Nvj7++Phhx/G+++/jylTpliye0RERESj2qir\nFTscjY2NuPvuu3HlyhWcOnUKsbGx1u4S2bDs7Gz88pe/hIODA4KDg/HJJ59ArbbKeSOycRUVFVi3\nbh00Gg00Gg127drVL60TkaHdu3fjF7/4BSorK63dFbJRhYWFmDt3LqZOnQqFQoEvvvgCfn5+g94z\nKkuK3SkXFxfs27cPP/7xj81SaJfsW1hYGNLS0pCRkYGIiAh89dVX1u4S2ahx48bh2LFjSEtLw0MP\nPYQPPvjA2l0iG6fT6fDll18iLCzM2l0hG5ecnIy0tDSkpqYOGdQBYyywU6vVRv2hEAHA+PHj4ejo\nCABwcHCASqWyco/IVimVvT9KGxsb4e3tbcXe0Giwe/duPPDAA/0OExIZOnbsGBITE/HCCy8Ydf2Y\nCuyI7kRRUREOHTqEtWvXWrsrZMPOnz+P+fPn45133sH69eut3R2yYd2zdT/96U+t3RWycUFBQbh2\n7RoyMzNRWVmJf//730PeMyoDu3feeQdz5syBk5MTHnvsMdl7tbW1uP/+++Hm5oaIiAjs3r37tp/B\n35LGjuGMl8bGRjzyyCP4+OOPOWM3BgxnrMyYMQOnTp3CSy+9hB07dliy22Qldzpedu7cydm6MeZO\nx4pGo4GzszMAYN26dTh//vyQzxqVO8GDg4OxefNmHDhwAG1tbbL3nnnmGTg5OaGyshI5OTlYvXo1\nZsyY0e+gBPfYjR13Ol66urrw4IMPYuvWrZg0aZKVek+WdKdjRavVwsHBAQDg4eGBjo4Oa3SfLOxO\nx8uVK1eQk5ODnTt3Ij8/H7/85S/x5ptvWulbkCXc6Vhpbm6Gm5sbACAzMxN33XXX0A+TRrEXX3xR\n2rhxY8/r5uZmSaPRSPn5+T1tjzzyiPS73/2u5/XKlSuloKAgKT4+Xvroo48s2l+yLlPHyyeffCL5\n+vpKycnJUnJysvT5559bvM9kHaaOlVOnTkmJiYnSkiVLpBUrVkglJSUW7zNZz538t6jb3LlzLdJH\nsg2mjpV9+/ZJs2fPlhISEqRHH31U0ul0Qz5jVM7YdZMMZt3y8vKgVqsRFRXV0zZjxgykp6f3vN63\nb5+lukc2xtTxsmHDBibJHqNMHSvz5s1DRkaGJbtINuRO/lvULTs7e6S7RzbE1LGycuVKrFy50qRn\njMo9dt0M9yc0NzfDw8ND1ubu7o6mpiZLdotsFMcLGYtjhUzB8ULGssRYGdWBnWHk6+bmhsbGRllb\nQ0MD3N3dLdktslEcL2QsjhUyBccLGcsSY2VUB3aGkW90dDS6urpQUFDQ03b+/HlMnTrV0l0jG8Tx\nQsbiWCFTcLyQsSwxVkZlYKfT6dDe3o6uri7odDp0dHRAp9PB1dUV69atw5YtW9Da2oqjR48iJSWF\n+6TGOI4XMhbHCpmC44WMZdGxYq6THpa0detWSaFQyP7Zvn27JEmSVFtbK/3whz+UXF1dpfDwcGn3\n7t1W7i1ZG8cLGYtjhUzB8ULGsuRYUUgSE7oRERER2YNRuRRLRERERP0xsCMiIiKyEwzsiIiIiOwE\nAzsiIiIiO8HAjoiIiMhOMLAjIiIishMM7IiIiIjsBAM7IiIiIjvBwI6IyMDGjRuxefNms37m008/\njZdeesmsn0lEZEht7Q4QEdkahULRr1j3cP31r3816+cREd0OZ+yIiG6D1RaJaDRiYEdENuWVV15B\nSEgIPDw8MHnyZKSmpgIAsrOzER8fD29vbwQFBeG5556DVqvtuU+pVOKvf/0rJk2aBA8PD2zZsgXX\nrl1DfHw8vLy88OCDD/Zcn56ejpCQELz88ssYN24cJkyYgF27dg3Ypz179iAuLg7e3t5YtGgRLly4\nMOC1zz//PAICAuDp6Ynp06fj8uXLAOTLu2vXroW7u3vPPyqVCp988gkA4OrVq1i+fDl8fX0xefJk\nfPnllwM+Kzk5GVu2bMHixYvh4eGBe+65BzU1NUb+SRORPWJgR0Q2Izc3F++++y5Onz6NxsZGHDx4\nEBEREQAAtVqN//7v/0ZNTQ1OnDiBI0eO4L333pPdf/DgQeTk5ODkyZN45ZVX8MQTT2D37t0oLi7G\nhQsXsHv37p5rKyoqUFNTg7KyMnz88cd48sknkZ+f369POTk5ePzxx/HBBx+gtrYWTz31FO677z50\ndnb2u/bAgQPIyspCfn4+Ghoa8OWXX8LHxweAfHk3JSUFTU1NaGpqwhdffIHAwEAsW7YMLS0tWL58\nOR5++GFUVVXhs88+w89//nNcuXJlwD+z3bt346OPPkJlZSU6Ozvx6quvmvznTkT2g4EdEdkMlUqF\njo4OXLp0CVqtFmFhYYiMjAQAzJo1C/PmzYNSqUR4eDiefPJJZGRkyO7/7W9/Czc3N8TGxmLatGlY\nuXIlIiIi4OHhgZUrVyInJ0d2/Y4dO+Dg4IDExESsXr0an3/+ec973UHY3/72Nzz11FOYO3cuFAoF\nHnnkETg6OuLkyZP9+q/RaNDU1IQrV65Ar9cjJiYG48eP73nfcHk3Ly8PGzduxBdffIHg4GDs2bMH\nEyZMwKOPPgqlUom4uDisW7duwFk7hUKBxx57DFFRUXBycsIDDzyAc+fOmfAnTkT2hoEdEdmMqKgo\nvPnmm9i2bRsCAgKwfv16lJeXAxBB0Jo1axAYGAhPT0+88MIL/ZYdAwICev7d2dlZ9trJyQnNzc09\nr729veHs7NzzOjw8vOdZfRUVFeG1116Dt7d3zz83b9687bVLlizBs88+i2eeeQYBAQF46qmn0NTU\ndNvv2tDQgB/84Af405/+hIULF/Y869SpU7Jn7dq1CxUVFQP+mfUNHJ2dnWXfkYjGHgZ2RGRT1q9f\nj6ysLBQVFUGhUOD//b//B0CkC4mNjUVBQQEaGhrwpz/9CXq93ujPNTzlWldXh9bW1p7XRUVFCAoK\n6ndfWFgYXnjhBdTV1fX809zcjJ/+9Ke3fc5zzz2H06dP4/Lly8jLy8Nf/vKXftfo9Xo89NBDWLZs\nGf7jP/5D9qykpCTZs5qamvDuu+8a/T2JaGxjYEdENiMvLw+pqano6OiAo6MjnJycoFKpAADNzc1w\nd3eHi4sLrl69alT6kL5Ln7c75bp161ZotVpkZWVh7969+MlPftJzbff1TzzxBN5//31kZ2dDkiS0\ntLRg7969t50ZO336NE6dOgWtVgsXFxdZ//s+/4UXXkBrayvefPNN2f1r1qxBXl4edu7cCa1WC61W\ni2+//RZXr1416jsSETGwIyKb0dHRgd///vcYN24cAgMDUV1djZdffhkA8Oqrr2LXrl3w8PDAk08+\niQcffFA2C3e7vHOG7/d9PX78+J4Tths2bMD//M//IDo6ut+1s2fPxgcffIBnn30WPj4+mDRpUs8J\nVkONjY148skn4ePjg4iICPj5+eE3v/lNv8/87LPPepZcu0/G7t69G25ubjh48CA+++wzBAcHIzAw\nEL///e9ve1DDmO9IRGOPQuKve0Q0xqSnp2PDhg0oKSmxdleIiMyKM3ZEREREdoKBHRGNSVyyJCJ7\nxKVYIiIiIjvBGTsiIiIiO8HAjoiIiMhOMLAjIiIishMM7IiIiIjsBAM7IiIiIjvx/wGpwoOsPsdJ\n4gAAAABJRU5ErkJggg==\n", + "text": [ + "
\n", + "
\n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "# Speed of numpy functions vs Python built-ins and std. lib." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## `sum()` vs. `numpy.sum()`" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "from numpy import sum as np_sum\n", + "import timeit\n", + "\n", + "samples = list(range(1000000))\n", + "\n", + "%timeit(sum(samples))\n", + "%timeit(np_sum(samples))" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "10 loops, best of 3: 18.2 ms per loop\n", + "10 loops, best of 3: 138 ms per loop" + ] + }, + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "\n" + ] + } + ], + "prompt_number": 20 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "funcs = ['sum', 'np_sum']\n", + "orders_n = [10**n for n in range(1, 6)]\n", + "times_n = {f:[] for f in funcs}\n", + "\n", + "for n in orders_n:\n", + " samples = list(range(n))\n", + " times_n['sum'].append(min(timeit.Timer('sum(samples)', \n", + " 'from __main__ import samples')\n", + " .repeat(repeat=3, number=1000)))\n", + " times_n['np_sum'].append(min(timeit.Timer('np_sum(samples)', \n", + " 'from __main__ import np_sum, samples')\n", + " .repeat(repeat=3, number=1000)))" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 26 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%pylab inline" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 24 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "import matplotlib.pyplot as plt\n", + "\n", + "labels = [('sum', 'in-built sum() function'), \n", + " ('np_sum', 'numpy.sum() function')]\n", + "\n", + "matplotlib.rcParams.update({'font.size': 12})\n", + "\n", + "fig = plt.figure(figsize=(10,8))\n", + "for lb in labels:\n", + " plt.plot(orders_n, times_n[lb[0]], \n", + " alpha=0.5, label=lb[1], marker='o', lw=3)\n", + "plt.xlabel('sample size n')\n", + "plt.ylabel('time per computation in milliseconds [ms]')\n", + "plt.legend(loc=2)\n", + "plt.grid()\n", + "plt.xscale('log')\n", + "plt.yscale('log')\n", + "plt.title('Performance of explicit for-loops vs. list comprehensions')\n", + "\n", + "max_perf = max( n/i for i,n in zip(times_n['sum'],\n", + " times_n['np_sum']) )\n", + "min_perf = min( n/i for i,n in zip(times_n['sum'],\n", + " times_n['np_sum']) )\n", + "\n", + "ftext = 'the in-built sum() is {:.2f}x to '\\\n", + " '{:.2f}x faster than the numpy.sum()'\\\n", + " .format(min_perf, max_perf)\n", + "plt.figtext(.14,.75, ftext, fontsize=11, ha='left')\n", + "\n", + "plt.show()" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "display_data", + "png": "iVBORw0KGgoAAAANSUhEUgAAAnYAAAIECAYAAACUvmMzAAAABHNCSVQICAgIfAhkiAAAAAlwSFlz\nAAALEgAACxIB0t1+/AAAIABJREFUeJzs3XdYFMf/B/D3HUXKnYLoAYJyCorSi2CnCGKLJhqjWKLY\n4tcWG6JfY4EYTewFMSY2NMbY8k00xoJKURQlIpKAiiCiWCgWRBCp8/vDHxsWDrjDExA/r+fhedjZ\n2dnP7M4ew+zsnoAxxkAIIYQQQt57wvoOgBBCCCGEKAd17AghhBBCGgnq2BFCCCGENBLUsSOEEEII\naSSoY0cIIYQQ0khQx44QQgghpJGgjh2pE8XFxZgwYQJatGgBoVCI8+fP13dI76XDhw/D1NQUqqqq\nmDBhQr3G4u/vj/bt23PLwcHBUFNTk3t7RfOXl5aWBg8PD4hEIqioqNSqjOpIpVKsWLFC6eW+r9zc\n3DB58mRu2cfHB3369KnHiN4vDak9CYVC7N+/v77DIO8QdewIx8fHB0KhEEKhEGpqapBKpZg6dSqe\nPXv21mX/+uuv+OWXX3D8+HGkp6ejW7duSoj4w1JSUoIJEybA29sbaWlp2LRpU32HBIFAwP3u7e2N\nR48eyb1txfz79u2DUCjfR9LKlSvx5MkTxMXF4fHjx/IHLCeBQMCr24eu4vEIDAzEkSNH5N5eVVUV\ne/fufRehvRcaUntKT0/Hp59+Wt9hkHdItb4DIA2Li4sLDh06hOLiYly9ehWTJ09GWloajh8/Xqvy\nCgsLoa6ujqSkJBgZGaFr165vFV9ZeR+iR48eIS8vD/3794ehoWF9hwMAKP9+cw0NDWhoaMi9raL5\ny0tKSoKTkxNMTU1rtX2Z4uJiqKrSx6CixGKxQvkFAgEa47vw38f2I5FI6jsE8o7RiB3hUVNTg0Qi\nQatWrTB48GDMmjULp06dQkFBAQDgwIEDsLOzg6amJtq2bYt58+bh1atX3PZubm6YNGkSlixZglat\nWsHExATu7u5YunQpUlJSIBQK0a5dOwBAUVERFi5cCGNjYzRp0gSWlpb45ZdfePEIhUIEBgZi1KhR\n0NHRwdixY7lbeOHh4bC2toaWlhZ69+6N9PR0hIWFwc7ODiKRCH369OGNCN29exdDhw6FkZERtLW1\nYWNjg3379vH2V3bLafny5TA0NISenh7GjRuHvLw8Xr6DBw/C0dERmpqaaNGiBQYMGIDs7GxufWBg\nIDp27AhNTU106NABK1euRElJSbXH/vLly3BxcYGWlhaaN2+O0aNHIysrC8Cb25YmJiYA3nS+a7qd\nXd3+Dx06hCZNmuCvv/7i8u/duxdaWlqIj48H8O+ttg0bNnDHa/jw4Xj+/HmV+5R1azUmJgb9+vVD\ns2bNIBaL0aVLF0RHR1fKHx4ejrFjxwIAN2pc1a1moVCI0NBQ7Nq1i5fv8ePH8Pb2hq6uLrS0tODu\n7o6YmBhuu/DwcAiFQpw4cQI9e/aEpqYmdu7cWWV9ynv58iWmTJkCiUQCDQ0NODk54cyZM7w8iYmJ\nGDhwIMRiMcRiMQYPHow7d+5UOj7nzp2DpaUlNDU10bVrV8TFxXF5cnJyMH78eBgaGkJDQwNt2rTB\nvHnzqoyrR48emDJlSqX0Tp06YenSpQCAhIQE9O3bF7q6uhCJRLCwsKjU7hVV8VZsdfuQSqUoKSnB\n+PHjIRQKa7x1HhQUBAsLC2hoaEBfXx/Dhg3j1tV0HlJTUyEUCvHLL7+gb9++0NbWhoWFBSIjI3H/\n/n3069cPIpEIlpaWiIyM5LYraxvHjx+Hs7MzNDU1YW1tjbCwsEp5ZLUfea73goICzJo1C3p6ejAw\nMMDcuXMr5ampHKlUimXLllVbTmRkJHr06IGmTZuiadOmsLOzQ0hICLe+4q1Yea+bs2fPwsXFBdra\n2rC0tMSpU6d4sa9cuRKmpqbQ0NCARCJBv3798Pr162rPNXlHGCH/b9y4caxPnz68tHXr1jGBQMBy\nc3PZ7t27ma6uLtu3bx+7e/cuO3/+PLOxsWGff/45l9/V1ZWJxWI2depUdvPmTRYfH8+ePXvGfH19\nWdu2bVlGRgZ78uQJY4wxX19fpqenx44cOcKSkpLYypUrmVAoZOfOnePKEwgETE9PjwUFBbGUlBSW\nlJTEdu/ezYRCIXN3d2fR0dHs2rVrrH379qxnz57MxcWFXblyhV2/fp117NiRjRgxgivrn3/+YUFB\nQezvv/9mKSkpLDAwkKmqqrKwsDBe/Do6Omzu3LksMTGRhYSEsObNm7MlS5ZweXbt2sXU1NTYN998\nw9Vxy5YtXL2WLVvGTExM2O+//85SU1PZiRMnWJs2bXhlVPT48WMmFovZ6NGjWXx8PIuMjGQ2NjbM\nxcWFMcZYfn4+++uvv5hAIGB//PEHy8jIYIWFhTLLkmf/kydPZqampiwnJ4clJiYysVjMvv/+e15b\naNq0Kfv4449ZfHw8Cw8PZ+3bt2dDhgzh7cfMzIxb3r17N1NVVeWW4+PjmZaWFhs1ahSLiYlhd+7c\nYYcOHWJRUVGV8hcWFrKgoCAmEAhYRkYGy8jIYDk5OTLrl56ezrp3787GjBnD5SstLWXOzs7M3t6e\nXbx4kf3zzz9sxIgRTFdXlzsvYWFhTCAQsI4dO7Ljx4+z1NRU9vDhQ5n7kEqlbMWKFdzysGHDWNu2\nbVlISAi7desWmzVrFlNXV2e3bt1ijDH26tUr1qZNG+bp6cmuXbvGYmJimLu7OzMzM+POU1m7dXR0\nZOfPn2d///03++ijj5iRkRHLz89njDE2c+ZMZmtry6Kjo1laWhq7dOkS27Fjh8wYGWPsxx9/ZLq6\nuqygoIBLu3LlChMIBCwpKYkxxpi1tTUbPXo0u3nzJrt79y47efIkO378eJVlyuLm5sYmT57MLfv4\n+PA+K6rbR1ZWFlNVVWWbN2/mzm1Vli5dykQiEQsKCmJJSUns+vXr7Ntvv+XW13Qe7t69ywQCATM1\nNWVHjx5lt2/fZkOGDGFGRkbMzc2N/f777+z27dts2LBhrHXr1qyoqIgx9m/baN++Pfvzzz/ZrVu3\n2MSJE5m2tjZ7/PgxL0/59vPgwQO5rjcTExOmq6vLVq1axZKTk9mhQ4eYmpoa27lzJ5dHGeUUFRUx\nXV1dNm/ePJacnMySk5PZ77//zi5cuMCVIRAI2M8//8wYYwpdN7a2tuz06dMsOTmZjR8/njVt2pQ9\nf/6cMcbYr7/+ypo2bcqOHz/O0tLS2PXr19mmTZu4dk3qFnXsCGfcuHHM09OTW05ISGDt2rVj3bp1\nY4y9+VD54YcfeNtEREQwgUDAsrOzGWNvOkbm5uaVyq7YCcjLy2NNmjThdSYYY2zIkCGsd+/e3LJA\nIGCTJk3i5dm9ezcTCAQsLi6OS1uzZg0TCATs2rVrXNqGDRtYixYtqq3zxx9/zPuD5erqyuzs7Hh5\npk6dyh0Dxhhr3bo1mzlzpszy8vLymJaWFjt9+jQvfc+ePUxHR6fKOBYvXsz7Q8MYY3FxcUwgELDz\n588zxv79o3Xx4sUqy5F3/69evWKWlpZs+PDhzM7Ojg0dOpSXf9y4cUwsFvM6VyEhIUwgELA7d+4w\nxmru2I0ZM6bSsSyvYv6ffvqJCQSCKvOXV7GjcfbsWSYQCNjNmze5tIKCAmZoaMi+/vprxti/f6D2\n7dtXY/nlO3ZJSUlMIBCwkydP8vI4ODiwCRMmMMYY27FjB9PS0mJPnz7l1mdkZDBNTU22d+9err4C\ngYCFhoZyeZ4/f85EIhHbtWsXY+xNe/Tx8ZHrGJRtr6mpyQ4fPsylTZ8+nXXv3p1bbtasGQsODpa7\nTFkqHu+KnxU17UNVVZXt2bOn2n3k5uYyDQ0Ntm7dOpnr5TkPZdfIpk2buPVl/xCtX7+eS4uNjWUC\ngYAlJCQwxv5tG2XngTHGiouLmYmJCdexktV+5L3eTExM2Mcff8zL079/fzZy5EillvPs2TMmEAhY\neHh45QP4/8p37BS5bn777TcuT0ZGBhMIBCwkJIQxxtj69etZhw4deJ9fpP7QrVjCEx4eDrFYDC0t\nLVhbW8PMzAw///wzsrKycP/+fcyZM4e71SQWizFgwAAIBAIkJydzZTg6Ota4n+TkZBQWFsLFxYWX\n7uLigoSEBF6as7Nzpe0FAgGsra25ZX19fQCAjY0NL+3p06fc3J5Xr15h4cKFsLKygp6eHsRiMU6c\nOIH79+/zyrW1teXty9DQEBkZGQCAzMxMPHjwAF5eXjLrlZCQgPz8fAwdOpR3nP7zn/8gJycHT58+\nrXK7rl278ubr2NjYoFmzZrhx44bMbd5m/5qamjh48CB+/fVXPHnyROYtSQsLC95cqu7duwOA3PHE\nxMTAw8ND7tjfRkJCAvT09NCxY0cuTV1dHV26dKm2PfXv3593nGQpq291bTUhIQGWlpZo3rw5t14i\nkcDc3LzS8Sr/4JCOjg46derElTNt2jQcOXIE1tbWmD17Nk6dOlXt3DQdHR0MHjwYP/30E4A30xsO\nHDjA3dYGAF9fX0yaNAnu7u4ICAhAbGxsleXVljL2kZCQgIKCgiqvLXnOQ5ny13BVnw3Am+u5vPLn\nRkVFBc7OztW2H3mvN4FAADs7O1455T9XlFWOrq4uJk2ahL59+2LAgAFYtWoVbt++jaooct2U369E\nIoGKigq33xEjRqCoqAgmJiYYP3489u3bh9zc3Cr3S96t92vWJ3nnunbtij179kBVVRWtWrXiOhpl\nF/DmzZvh7u5eaTsjIyMAbz54tLW1lRqTrPKEQiHvKbOy38vP3ylLY4xBIBBg/vz5OHbsGDZs2ABz\nc3NoaWlh3rx5ePHiBa/sig9nCAQClJaWyhVrWb4jR46gQ4cOldbr6urK3E5Zk8sV2f+FCxcgEAjw\n4sULZGZmQkdHh5f3beNpCBPmy859eeXb086dO2s9D6hi2bLqKk/9y+fx8vLC/fv3cfr0aYSHh2PM\nmDGwtrbGuXPnqnxieOzYsRgyZAiePHmCyMhI5OXlwdvbm1u/ePFijB49GqdOnUJoaChWrlwJPz8/\nLF++XJHqVqsu9lEVWce4/FzPsnMkK62m67qm9qPI9Vbd54qyygGAH3/8EbNmzUJISAjOnDmDJUuW\nYMuWLfjiiy+qrWt5suot66G1sv22atUKt27dQlhYGEJDQ7F8+XIsWLAAV65cgbGxsdz7JcpBI3aE\nR0NDA+3atUObNm14o0f6+vpo3bo1bt26hXbt2lX6adKkiUL7MTMzQ5MmTRAREcFLj4iI4I3EKdOF\nCxcwZswYDBs2DNbW1mjbti0SExMVeg2BRCKBsbExTp8+LXO9paUlNDQ0cOfOHZnHqao/zpaWlrh8\n+TKKioq4tLi4OLx48QJWVlZyxyfv/uPj4zFv3jzs3LkTHh4e8Pb2RmFhIa+smzdv4uXLl9zypUuX\nALwZyZOHo6Mjzp07J3fnruwPR206g5aWlnj69Clu3rzJpRUUFODKlSvVHr9WrVrxjk9VZQOo1FbP\nnz/PlW1paYkbN27wRmQzMjJw+/btSvuPiorifs/OzsatW7d4x1RXVxfe3t7Ytm0b/vzzT0RERPDq\nVZGXlxeaN2+OAwcOYO/evRg0aBCaNWvGy9O2bVtMnToVhw8fRkBAAL7//vsqy5NXxeumun2oq6vX\n+PBQ2QMT1V1bgOzzoKzPjPLnpri4GNHR0dW299pe7++qnPLlzZkzBydOnMDEiRPx448/VpmvNteN\nLOrq6ujbty9WrVqFf/75B69evcLRo0cVKoMox3s5YhcdHY3Zs2dDTU0NRkZG2Lt373v3yPn7aMWK\nFZg4cSJ0dXUxePBgqKmp4ebNmzh16hS2bdsG4M0fZXn+MGtpaeHLL7/EkiVL0LJlS9jY2ODIkSM4\nduwYzp49+07iNzc3x++//46hQ4dCW1sb69evx+PHj2FgYMDlkSf+ZcuWYerUqdDX18enn36K0tJS\nhIWFYeTIkdDT08OiRYuwaNEiCAQCeHh4oLi4GP/88w+uX7+O7777TmaZM2bMwKZNm+Dj44NFixbh\n+fPnmDZtGlxcXNCjRw+56ygSiWrc/+vXrzFy5EgMGTIEY8eOxaBBg2Braws/Pz9s3LiRK0sgEGDs\n2LH45ptv8PTpU0yfPh0ff/xxlR2givz8/NClSxeMHj0a8+bNg46ODq5du4bWrVvLfO1N27ZtAQBH\njx5Fjx49oKWlVeXob8Xz5OHhAWdnZ4waNQpBQUFo2rQpli9fjsLCQkydOlXu41e+/DKmpqb47LPP\nMG3aNPzwww9o06YNvv/+e9y4cQMHDhwAAIwePRrLly/HiBEjsGbNGpSWlsLX1xfGxsYYMWIEV5ZA\nIMCCBQuwbt066Ojo4KuvvkLTpk0xatQoAMBXX32Fzp07w8LCAkKhEPv27YNYLEabNm2qjFVVVRWj\nRo3C1q1bkZKSgl9//ZVbl5eXBz8/PwwbNgxSqRTZ2dk4deoU10kC3oz4CQQC7Nmzp9rjUfG6KFvO\nzc3FggULqt1H27ZtERoain79+kFNTQ0tWrSotA+RSIR58+bB398fmpqa8PT0RH5+Pk6ePImFCxfK\ndR7e1qpVq2BgYACpVIr169fj6dOnmDZtWpX55bneyh+rd11OcnIytm/fjsGDB8PY2BiPHj3ChQsX\nqpweo6zrZufOnWCMwcnJCTo6Ojh37hxevnwp9z+BRLneyxG7Nm3aICwsDBEREZBKpfRfgZLU9BLN\nMWPG4NChQzh+/Di6dOkCZ2dnBAQE8IbaqypDVvqKFSswefJkzJ49G9bW1ti/fz9+/vlnmbd6ZZWn\naNqGDRu41694enqidevWGDZsWKVbuhXLqZg2ceJEBAcH48iRI7C3t4erqytOnz7N/XOxePFirF+/\nHtu3b4ednR169eqFTZs2cR0XWSQSCUJCQvDgwQM4OTlh0KBBXGe3pjpWVNP+58yZg/z8fK4zrqur\ni/3792Pr1q04efIkV46zszN69uyJPn36oH///rC1tcWuXbtqPFZlrKysEB4ejqysLLi6usLe3h4b\nNmzg/RNWPr+TkxNmzZqFKVOmQF9fHzNnzqyyjrL2/fvvv6Njx44YOHAgnJ2dkZmZiTNnzvDmvck7\nOlsx344dO9C3b1+MGTMGdnZ2iIqKwvHjx7nbZhoaGggJCUGTJk3g4uICNzc3iMVinDp1ildfoVCI\nlStXYsqUKXByckJmZib+/PNP7n1+mpqaWLp0KTp37gwnJyfEx8fj5MmTNb43bty4cbh16xZ0dHTQ\nv39/Ll1VVRXZ2dmYOHEiLCws0K9fPxgaGvJed5GWloa0tLQaj0dV14mamlqN+1i3bh1iYmIglUq5\n+W2yLF++HCtWrMDmzZthbW2Nvn378ubr1XQeymKTFb88aWvXrsWSJUtgb2+PqKgoHD16lPePn6xt\n5Lne5flMVEY5IpEIycnJ8Pb2hrm5OYYNG4YePXpgy5YtlbYro4zrpnnz5ti9ezfc3d1hYWGBjRs3\nYvv27XJ9lhPlE7D6ngTzlpYtWwZ7e3t88skn9R0KIY2Gj48PHj58WOldbaT2goODMXnyZN7tdtIw\nhIeHo3fv3njw4AFatWpV3+EQ8lbeyxG7Mvfu3cOZM2cwaNCg+g6FEEIIIaTe1WvHbsuWLejcuTM0\nNDQwfvx43rpnz55hyJAhEIlEkEqllb6RICcnB2PHjsWePXveyZeAE/Iha0jfbdmY0DFtuOjckMai\nXm/F/vbbbxAKhTh9+jTy8/Oxe/dubt3IkSMBvJmUGRsbi4EDB+LSpUuwsLBAcXExBg8eDF9fX/Tu\n3bu+wieEEEIIaVje+SuQ5bB48WLe29Zzc3OZuro695U4jDE2duxYtnDhQsYYY3v37mV6enrMzc2N\nubm5sYMHD1Yqs1WrVgwA/dAP/dAP/dAP/dBPg/8xNTVVSp+qQcyxYxUGDW/fvg1VVVWYmZlxaba2\nttybsD///HM8efIEYWFhCAsLw/DhwyuV+ejRI+4R/Yb8s2zZsvdiH7UtQ5Ht5MlbU57q1td2XUP6\neddxKqv82pSj7LYiT77atAlqK8rdB322NIwf+mxRLO+7aC937txRSp+qQXTsKs5tyM3NRdOmTXlp\nYrGY97LUxsLNze292Edty1BkO3ny1pSnuvXVrUtNTa1x3w3Bu24vyiq/NuUou63Ik6827YXainL3\nQZ8tDQN9tiiW9121F2VoEK87Wbx4MR4+fMjNsYuNjUXPnj2Rl5fH5Vm7di3Onz+PY8eOyVVmQ/g6\nI/L+8PHxQXBwcH2HQd4D1FaIIqi9EHkpq9/SIEfsOnTogOLiYt4Xy8fFxSn8FSf+/v4IDw9XRoik\nkfPx8anvEMh7gtoKUQS1F1KT8PBw+Pv7K628eh2xKykpQVFREQICAvDw4UNs374dqqqqUFFRwciR\nIyEQCLBjxw5cu3YNH330EaKiotCpUye5yqYRO0IIIYS8LxrFiN3y5cuhpaWFVatWYd++fdDU1MSK\nFSsAAFu3bkV+fj4kEgnGjBmDbdu2yd2pI0RRNLJL5EVthSiC2gupa6o1Z3l3/P39qxx+1NXVxW+/\n/Va3ARFCCCGEvMfqtWP3rvn7+8PNza3SEyjNmzfH8+fP6ycoQhoZXV1dPHv2rL7DqDN18bQpaTyo\nvZCahIeHK3Vkt0E8FfsuVHevmubfEaI8dD0RQsjbaxRz7Agh5H1Dc6aIIqi9kLpGHTtCCCGEkEai\nUXfs6D12hBBlozlTRBHUXkhNGtV77N4lmmNHSN2g64kQQt4ezbH7gPn4+KBPnz5vXU54eDiEQiEe\nPXr01mUJhULs37+fW5ZKpdw7Cd8nsbGxMDAwwKtXrwAAkZGRkEqlKCgoqHHbtLQ0eHh4QCQSQUVF\n5V2HWqPU1FQIhUJcunSpvkNpVOguAFEEtRdS16hj9x4KDAzEkSNH6jsMnvT0dHz66afcskAg4H1V\nnJmZGQICAuojNIX4+flh7ty50NLSAgD07NkTZmZm2LJlS43brly5Ek+ePEFcXBweP378rkPlkXV8\n27Rpg/T0dDg7O9dpLIQQQupPo36P3dtITLyHs2fvoKhICDW1Unh6msLc3KRBlCcWi2sdx7sikUiq\nXV/x+4AbooSEBERERPBGHgFgwoQJ+OqrrzB37txq65GUlAQnJyeYmpq+61ArkRWXUCis8bwQxdGc\nKaIIai+krjXqEbvaPjyRmHgPwcHJyMrqjexsN2Rl9UZwcDISE+/VKg5ll1fxVmzZ8o8//ggTExM0\na9YMH3/8MTIzM+Uq79q1a3B2doampiasra0RFhbGravqdq2qqir27NnDLQuFQvz8888yy3dzc8Od\nO3cQEBAAoVAIoVCI+/fvy8ybkJCAvn37QldXFyKRCBYWFti3bx9vPxU7Xp6enhg/fjy3LJVKsXTp\nUkydOhU6OjowMDDA999/j9evX2P69Olo3rw5jI2NERQUxCtn37596N69O1q2bMlLHzRoENLS0hAZ\nGSkz5rK4QkNDsWvXLgiFQkyYMEGheJctW4ZZs2ZBT08PBgYGmDt3LkpKSnjbBQUFwcLCAhoaGtDX\n18ewYcOqPb6ybsUmJiZi4MCBEIvFEIvFGDx4MO7cucOtDw4OhpqaGi5dugQHBwdoa2ujc+fOuHr1\napV1J4QQUnvKfniiUY/Y1fZAnT17B02aeIDfJ/TA33+HwslJ8VG26Og7ePXKg5fm5uaBc+dCazVq\nV/E2JwD89ddfkEgkOHnyJHJycjBq1Cj4+vpi7969NZY3d+5cbNy4EaamplizZg0GDRqE5ORkGBgY\nKBRDVaNZv/32GxwdHTFs2DD4+voCAFq0aCEz78iRI2FjY4OoqChoaGjg1q1blTo48sQSGBiIZcuW\n4dq1a/jll18wY8YMHD16FP369cPVq1dx6NAhfPnll+jduzf3HcQRERHo1atXpfLFYjEsLS0RGhoq\ncz0APH78GEOHDkW7du2wbt06aGpqKhzvwoULER0djWvXrmH06NGwsrLiOojLli3D+vXrsWrVKnh5\neSEvLw8nT54EUPXxrdh5zs/Ph5eXFzp06IDz58+DMQZfX1/069cPN27cgJqaGgCgtLQUixYtQmBg\nIFq0aIE5c+Zg+PDhSEpKahBzB+tbeHg4jcIQuVF7ITUp+4YsZU1XatQjdrVVVCT7sJSU1O5wlZbK\n3q6wsHblMcYqPTmjoaGB4OBgWFhYoGvXrvjPf/6Ds2fPylXef//7XwwYMADm5ub44Ycf0KJFC2zd\nurVWscmiq6sLFRUViEQiSCQSSCQSCIWy637//n306dMHHTt2hFQqRb9+/TBw4ECF9+nu7o7Zs2ej\nXbt2WLRoEUQiEZo0acKlLViwAM2aNUNoaCi3TVJSEtq0aSOzPKlUitu3b1e5P319fairq0NTUxMS\niUTh2+UuLi7w8/ODqakpPvvsM3h6enLnLy8vD6tXr0ZAQACmTZsGMzMz2NraYuHChQDkP7779+/H\nkydPcPDgQdjb28PBwQEHDhzAw4cPceDAAS4fYwwbN25Ejx49YG5uDn9/f6SmpiIlJUWhOhFCCKl7\n1LGTQU2tVGa6iors9JoIhbK3U1evXXmydOzYkRtxAQBDQ0NkZGRwyyKRiLv9VrGj1K1bN+53FRUV\nODs7IyEhQWmxKcLX1xeTJk2Cu7s7AgICEBsbq3AZAoEAtra2vOWWLVvCxsaGlyaRSJCVlcWlvXjx\nosoOmVgsRnZ2tsKxyBuvnZ0dL638+UtISEBBQQG8vLzeaj8JCQmwtLRE8+bNuTSJRAJzc3PcuHGD\nF0/542doaAgAvPb0IaPRF6IIai+krjXqW7G15elpiuDgc3Bz+/f2aUHBOfj4mMHcXPHyEhPflNek\nCb88Dw8zZYQLALxOHVD5fTh///0393t1twmBNyM2ZbcKy0Z+ypdVUlKC0lLldUrLW7x4MUaPHo1T\np04hNDQUK1euhJ+fH5YvXw5A9nt+CgsLK5Uj63jISitfDx0dHbx8+VJmXC9evICurq7C9ZE3XnV1\n9WpjUxZZ70iqmCYUCnm3ist+f1fnnBBCiPLQiJ0M5uYm8PExg0QSCh2dcEgkof/fqavdU6zKLk+W\nmp46bdeg43pVAAAgAElEQVSuHfdTNgJTJioqivu9uLgY0dHRsLCwAPDv064PHz7k8ly/fl3hlyiq\nq6vXOFeuTNu2bTF16lQcPnwYAQEB+P7777l1EomEF0tBQQFvtOlttG/fHqmpqTLX3bt3Dx06dFC4\nTGXEW/bAxOnTp6vMI8/xtbKywo0bN/D06VMuLSMjA7dv34aVlZVCMX3I6L1kRBHUXkhda9Qjdv7+\n/tykREWZm5soteOl7PIqepu3Va9atQoGBgaQSqVYv349nj59imnTpgF409kxMTGBv78/NmzYgKys\nLCxatKjGjmTFeNq2bYvIyEikpaVBU1MTenp6lcrIy8uDn58fhg0bBqlUiuzsbJw6dQqWlpZcHk9P\nT2zbtg0uLi4QiURYsWIFioqKePuTZ1RKVpqrqysuXrxYKd/Lly9x48aNGtuRrLmPtY23PJFIhHnz\n5sHf3x+amprw9PREfn4+Tp48yc2zk3V8Kxo1ahS+/vprjBgxAmvWrEFpaSl8fX1hbGyMESNGVBsD\nIYSQdyM8PFyp/wA06hG7so5dY1PxqUpZT1mWpctT1tq1a7FkyRLY29sjKioKR48e5Z6IVVFRwcGD\nB5GZmQl7e3vMnDkTK1eurPLhh6r2HRAQgOzsbJibm0NfXx9paWmVtlFVVUV2djYmTpwICwsL9OvX\nD4aGhrzXhaxduxZWVlbo27cvBg4cCDc3Nzg5Ocm8dVjTsaiYNmbMGERFRfHm3QHAsWPH0Lp1a7i4\nuNRY54plvk285dOXL1+OFStWYPPmzbC2tkbfvn158w+rOr7ly9DQ0EBISAiaNGkCFxcXuLm5QSwW\n49SpU1BVVeXtu6Zj9SFrjJ8p5N2h9kJq4ubmRt8VKw/6rlhSG15eXvDw8MCCBQu4NA8PD/Tv3597\nlQjho+uJEELeHn1XLCHvwOrVq7Fx40bed8WmpKTgyy+/rOfISENBc6aIIqi9kLrWqOfYEaIoOzs7\n3ve89uzZE3fv3q3HiAghhBD50a1YQshboeuJEELeHt2KJYQQQgghPI26Y+fv70/zGwghSkWfKUQR\n1F5ITcLDw+mpWHnQrVhC6saHdj3Rl7oTRVB7IfJS1mcpdewIIW+FridCCHl7NMeOEEIIIYTwUMeO\nEEIUQHOmiCKovZC6Rh078sGLjY2FgYEB76XEUqkUBQUFNW6blpYGDw8PiEQiqKiovOtQa5Samgqh\nUIhLly7VdyiEEELqAXXsyAfPz88Pc+fOhZaWFoA3LyU2MzPDli1batx25cqVePLkCeLi4ngvNq4L\nZmZmCAgI4KW1adMG6enpcHZ2rtNYPiQ0EZ4ogtoLqWvUsSMftISEBERERGD8+PG89AkTJmDLli01\nTmRNSkqCk5MTTE1NIZFI3mWolQgEgkppQqEQEokEqqr0pTKEEPIhoo5dFRKTExF0MAgbD2xE0MEg\nJCYnNpjy3NzcMHnyZCxfvhyGhobQ09PDuHHjkJeXx+Xx8fFBnz59eNvt27cPQuG/p9zf3x/t27fH\n4cOHYWZmBm1tbXz66afIzc3F4cOHYW5ujqZNm+Kzzz5DTk5OpbI3bNgAIyMjaGtrY/jw4Xj+/DmA\nN3NKVFVV8eDBA97+9+7dCx0dHeTn5/PSExIS0LdvX+jq6kIkEsHCwgL79u3j1guFQuzfv5+3jaen\nJ68zJpVKsXTpUkydOhU6OjowMDDA999/j9evX2P69Olo3rw5jI2NERQUVOmYdO/eHS1btuSlDxo0\nCGlpaYiMjKzyPAiFQoSGhmLXrl0QCoWYMGGCQvEuW7YMs2bNgp6eHgwMDDB37lyUlJTwtgsKCoKF\nhQU0NDSgr6+PYcOGAXjTBu7cuYOAgAAIhUIIhULcv39f5q3YxMREDBw4EGKxGGKxGIMHD8adO3e4\n9cHBwVBTU8OlS5fg4OAAbW1tdO7cGVevXq2y7h8ymjNFFEHthdS1Rt2xq+0LihOTExEcFows/Sxk\nG2QjSz8LwWHBte6MKbs8ADhy5Aiys7MRERGBAwcO4Pjx41i1ahW3XiAQyBzRqejx48fYu3cvfv/9\nd5w8eRIXLlzA0KFDERwcjCNHjnBpK1eu5G0XHR2NiIgIhISE4MSJE7h+/TomTpwI4E2no3379ti1\naxdvm+3bt2P06NHQ1NTkpY8cORItW7ZEVFQU4uPjsX79eujq6lYbt6z6BQYGwtzcHNeuXcPMmTMx\nY8YMfPLJJ2jfvj2uXr2KGTNm4Msvv8TNmze5bSIiItClS5dK5YvFYlhaWiI0NLTaY9etWzeMHj0a\n6enp2LRpk8LxGhkZITo6GoGBgdiyZQv27NnDrV+2bBkWLlyIGTNmID4+HiEhIejcuTMA4LfffoNU\nKoWvry/S09ORnp4OY2PjSvvNz8+Hl5cXCgsLcf78eURERCA3Nxf9+vVDUVERl6+0tBSLFi1CYGAg\nrl27BolEguHDh1fqaBJCCFEuZb+guFHfr6ntgTobcxZN2jdBeGr4v4lqwN8H/oZTTyeFy4uOjMYr\n41dA6r9pbu3dcO7aOZibmdcqRqlUinXr1gEAOnTogBEjRuDs2bP4+uuvAQCMMbneh1NQUIA9e/ag\nefPmAIDhw4dj27ZtyMjIgJ6eHgDA29sb586d423HGMNPP/0EsVgM4M3IUt++fZGSkoJ27drhiy++\nwKZNm7BkyRIIBALcunULFy9elDlv7f79+5g3bx46duzI1a023N3dMXv2bADAokWLsHr1ajRp0oRL\nW7BgAVavXo3Q0FB06tQJwJtbqaNHj5ZZnlQqxe3bt6vcn76+PtTV1aGpqVmr27AuLi7w8/MDAJia\nmmL37t04e/YsJkyYgLy8PKxevRorVqzAtGnTuG1sbW0BALq6ulBRUYFIJKp23/v378eTJ08QGxvL\nneMDBw5AKpXiwIED+PzzzwG8OZ8bN26EnZ0dgDfXTteuXZGSkoL27dsrXLfGjOZMEUVQeyE1cXNz\ng5ubW6U507XVqEfsaquIFclML0HtRi9KUSozvbC0sFblCQQC7g98GUNDQ2RkZChclpGREfcHH3jT\nWTEwMOA6dWVpmZmZvO0sLCy4Th0AdO/eHQBw48YNAMDYsWORmZmJ06dPAwB27NiBzp07V4obAHx9\nfTFp0iS4u7sjICAAsbGxCtej4jERCARo2bIlbGxseGkSiQRZWVlc2osXL3j1KE8sFiM7O1vhWOSN\nt6wTVab8OUxISEBBQQG8vLzeaj8JCQmwtLTknWOJRAJzc3PuXJXFU/74GRoaAkCt2hQhhJD6Qx07\nGdQEajLTVVC711kIqzjM6kL1WpUHAOrq/G0FAgFKS//tQAqFwkojduVvvZVRU+PXVSAQyEwrXzaA\nGkcD9fT0MGzYMGzfvh1FRUXYu3cvvvjiC5l5Fy9ejNu3b2P48OGIj49H165dsWTJEt7+K+6vsLBy\np7g2ddHR0cHLly9lxvXixYsabwnLIm+8NZ1DZZF1riqmCYVC3q3ist/fRTzvO5ozRRRB7YXUtUZ9\nK7a2PB09ERwWDLf2blxaQVIBfLx9anXrNNH4zRy7Ju2b8MrzcPdQRrgy6evr4/Lly7y0a9euKa38\nmzdv4uXLl9xoV9lkfQsLCy7PlClT4O7ujm3btuH169cYOXJkleW1bdsWU6dOxdSpU/Hdd99h7dq1\nWL58OYA3I0wPHz7k8hYUFODGjRswNTV963q0b98eqampMtfdu3ePe1hBEcqIt+yBidOnT8PKykpm\nHnV19RrnwFlZWeGHH37A06dPuVHYjIwM3L59G/Pnz5c7HkIIIe8HGrGTwdzMHD7uPpBkSqCTrgNJ\npgQ+7rXr1L2L8uSZP+fp6Ylbt25h69atuHPnDrZv347Dhw/Xan+yCAQCjB07FgkJCTh//jymT5+O\njz/+GO3atePy9OjRA+bm5pg/fz5GjhwJbW1tAICHhwcWLVoEAMjNzcX06dMRFhaGu3fvIjY2FqdO\nnYKlpSWvLtu2bcPly5cRHx8PHx8fFBUV8Y6BPKNSstJcXV0RHR1dKd/Lly9x48aNGufHyDoXtY23\nPJFIhHnz5sHf3x9bt27F7du3ERcXh++++47L07ZtW0RGRiItLQ1PnjyRWeaoUaPQsmVLjBgxArGx\nsYiJiYG3tzeMjY0xYsSIamMgstGcKaIIai+krtGIXRXMzcxr3fF61+XJesKyYpqHhwe++eYbrFy5\nEgsWLMDgwYOxdOlSzJw5U6FyqkpzdnZGz5490adPH7x48QIDBgzAjz/+WCnWSZMmYc6cObzbsCkp\nKTAxMQHw5vZpdnY2Jk6ciMePH6Np06bo3bs31q5dy+Vfu3YtJk+ejL59+0JHRweLFi3CkydPZN46\nrBh3TWljxozB2rVrkZWVxXvlybFjx9C6dWu4uLhUKqOmY/M28ZZPX758OVq2bInNmzdjzpw50NXV\nhaurK7c+ICAAX3zxBczNzVFQUIC7d+9WKltDQwMhISGYM2cOVxd3d3ecOnWK9647eY8fIYSQhk3A\n5Hl08j0ka56TPOtIzXx8fPDw4UOcOXOmxrx+fn44d+4cYmJi6iCy2vHy8oKHhwcWLFjApXl4eKB/\n//7w9fWtx8jeDx/a9RQeHk6jMERu1F6IvJT1WUq3Ysk78eLFC/z111/Yvn075syZU9/hVGv16tXY\nuHEj77tiU1JS8OWXX9ZzZIQQQohiaMSOKGz8+PF4+PAhQkJCqszj5uaG6OhojBw5Ejt37qzD6Ehd\no+uJEELenrI+S6ljRwh5K3Q9EULI26NbsYQQUg/ovWREEdReSF2jjh0hhBBCSCPRqDt2/v7+9N8S\nIUSp6AlHoghqL6Qm4eHhtf5ue1lojh0h5K3Q9UQIIW9PWZ+lH+QLinV1denlq4QoSW2+T/d9Ru8l\nI4qg9kLq2gfZsXv27Fl9h0AaGPrwJYQQ0hh8kLdiCSGEEEIaEnrdCSGEEEII4aGOHSGgd00R+VFb\nIYqg9kLqGnXsCCGEEEIaCZpjRwghhBBSz2iOHSGEEEII4aGOHSGgeTBEftRWiCKovZCaJCYnIuhg\nkNLK+yDfY0cIIYQQUt8SkxOx9fRWPGz5UGll0hw7QgghhJA6llOQg/k/zkeiOBEAEDE+gr5SjBBC\nCCHkffK6+DUu3r+Iyw8u496Le4BYueXTHDtCQPNgiPyorRBFUHshZYpLixGVFoVNlzfhwv0LKCot\ngvD/u2G6Gsr7zm0asSOEEEIIeUdKWSn+yfgHYalhyH6dzVvXxaoL0tPSoS/Vx+/4XSn7ey/n2OXk\n5MDT0xM3b97ElStXYGFhUSkPzbEjhBBCSH1hjOHO8zs4c+cMMvIyeOt0NXTRu21vWEmscPvObZy7\ndg7TR0xXSr/lvezYFRcXIzs7G/Pnz4evry8sLS0r5aGOHSGEEELqw8OchzibchZ3s+/y0rXUtOBq\n4orOrTpDRajCW/dBv6BYVVUVLVq0qO8wSCNC82CIvKitEEVQe/mwPMt/hsMJh7H92nZep05NqAZX\nE1fM6jILXYy7VOrUKRPNsSOEEEIIeQu5hbmISI1AzOMYlLJSLl0oEMLB0AFuUjeI1EV1Eku9jtht\n2bIFnTt3hoaGBsaPH89b9+zZMwwZMgQikQhSqRS//PKLzDIEAkFdhEoaOTc3t/oOgbwnqK0QRVB7\nadwKigsQnhqOzVc2469Hf/E6dRYtLTDdaTo+6vBRnXXqgHoesTMyMsKSJUtw+vRp5Ofn89ZNnz4d\nGhoayMzMRGxsLAYOHAhbW9tKD0rQPDpCCCGE1KWS0hLEPI5BRGoE8oryeOtMmpmgj2kfGDc1rpfY\nGsTDE0uWLMGDBw+we/duAEBeXh6aN2+OhIQEmJmZAQDGjRuHVq1a4dtvvwUADBgwAHFxcTAxMcGU\nKVMwbtw4Xpn08ARRRHh4OP1nTeRCbYUogtpL48IYQ0JWAkLvhuJZ/jPeOom2BH3a9YFZc7Na3U1U\nVr+lQcyxq1iR27dvQ1VVlevUAYCtrS1vEuqJEydqLNfHxwdSqRQAoKOjAzs7O+4CKyuLlmkZAK5f\nv96g4qFlWqZlWqblhrVsYmuCMylncOnCJQCA1E4KAMhKyIK9oT0muE6AUCCUu7yy31NTU6FMDXLE\n7sKFCxg+fDgeP37M5dm+fTv279+PsLAwucqkETtCCCGEvK303HScTTmL5GfJvHRNVU30MukFZyNn\nqArffpysUY/YiUQi5OTk8NJevHgBsVjJX6hGCCGEECJD9utshN4NxT8Z/4Dh336KqlAVXYy6oGeb\nntBU06zHCGUT1ncAQOUnWzt06IDi4mIkJ//bO46Li4OVlZVC5fr7+/OGPAmpCrUTIi9qK0QR1F7e\nP6+KXuF08mkEXgnE3xl/c506AQSwN7DHTOeZ6GPaR2mduvDwcPj7+yulLKCeR+xKSkpQVFSE4uJi\nlJSUoKCgAKqqqtDW1sbQoUOxdOlS7NixA9euXcMff/yBqKgohcpX5oEihBBCSONVVFKEyw8uI/J+\nJApKCnjrzPXM4dHOAxJtidL36+bmBjc3NwQEBCilvHodsVu+fDm0tLSwatUq7Nu3D5qamlixYgUA\nYOvWrcjPz4dEIsGYMWOwbds2dOrU6Z3F4u/vj6KiIm7Zx8cHQUFBb1VmTEwMxowZo/B2qampaNmy\n5Vvtr2IZFetX14KCgrBq1SoAwPHjxzF9+vQq8y5btgyHDh1SeB+MMXh6elZ77CIiIuDk5AR7e3tY\nWVnhhx9+AAC4urri008/RceOHWFnZwcvLy+kpKQoHMPGjRuRlZWl8HYAsHTpUlhYWMDOzg69evXC\njRs3AACXLl1Cjx49YGlpCUtLS/j5+VVZRkZGBry8vGBubg47OztER0dz69auXYuOHTtCRUUFf/75\nZ61ivHTpEqysrODo6IiIiAiFt4+Li8Phw4drtW9Z3sV1W524uLhK51coFOLVq1fvbJ8fgoyMDHTr\n1g0AUFBQgM6dOyMvL6+Grd4PZRPmScNVykoR8ygGm69sxrm753idOuOmxhhvNx4jrUe+k07dO8Ea\nKUWrJhAIWG5uLrfs4+PDtmzZouyw5HL37l3WokULpZZRsX51qaCggJmamvL2b2Njw9LS0pS6n82b\nN7OJEyeyli1bVpmnY8eO7M8//2SMMZaens5EIhHLzMxkpaWl7NixY1y+LVu2MA8PD4VjkEqlLD4+\nXuHtLl++zExMTNirV68YY2/qMmDAAMYYY/Hx8Sw5OZkx9uZY9uzZk/30008yyxk/fjxbsWIFY4yx\nyMhI1r59e27dX3/9xe7cucPc3Ny4Y6Co//znP2zNmjW12pYxxnbv3s2GDRtWq22Li4srpdX1dSsr\n/vq8thqLOXPmsODgYG551apV7LvvvqvHiMiHoLS0lN3MuskCrwSyZWHLeD+BVwLZjcwbrLS0tM7i\nUVaXrEHMsXtX5J1jVzZ61L17dzg4OODFixcAgPj4eHh4eKBDhw689+Tl5ORg0qRJ6NKlC2xtbTF7\n9myUlpZWKjc8PBxOTk4A3oygtWjRAosXL4aDgwM6duyIixcvVhuXr68vbG1tYWNjg8jIyEplVlyu\nuE5W/ezt7bn6lcnMzISnpydsbGxgY2ODefPmAXhz/ObPn8/lK7/s7+8Pb29vDBw4EO3bt8fw4cNx\n9epVuLu7w8zMjDeq9Mcff8DZ2Rna2tpc2tChQ7F3716Z9S4/6nL06FHY2NjA3t4e1tbWVY4SJSUl\n4eDBg1i4cGG1TxUZGxsjOzsbwJsHcpo1awZtbW1ERERg0KBBXL6uXbvi3r173PFp27YtYmJiAAB7\n9uxBr169Kp3zFStW4NGjRxg2bBjs7e1x69Yt5ObmYvz48bC2toa1tTXWrFkjMy6JRAKBQMCNUmRn\nZ6N169YAAEtLS5iamgIA1NXVYWdnh/v378ss5/Dhw/jPf/4DAOjRoweaNGmCq1evAgA6d+6Mdu3a\nVdpG3vqtWbMGhw4dwqZNm+Dg4IDXr1/D19cXzs7OsLOzg6enJxeXrDb17NkzLF26FGfPnoW9vT1m\nz54NALhy5Qp69+6Nzp07o3PnztyrjMqumfnz58PR0RE7d+7kxaPodbt//3507doVDg4OcHBwQGho\nKLdOKpVi2bJl6N69O9q2bStz1O/p06dYtmwZTp06xYsfADZv3gxnZ2eYmprif//7H5deVd0q8vHx\nwdSpU2XG7ebmxhthdXNz48pxc3ODr68vevXqhTZt2mDt2rXYt28fV48jR45w2wmFQvj7+8Pe3h4d\nO3bk4lyzZg1mzJjB5cvIyICBgQFev37NpQUEBKBTp06wt7eHg4MDcnJyKt0VKL9cdu4WLVoEBwcH\ndOrUCVevXsXEiRNhY2ODrl27IiMjAwBQXFyM/fv3Y9iwYVxZI0aMqHS+31c0x65huv/iPnbF7sKB\n+AN48uoJly5WF2NQh0GY5jQNnVp2qpNvt1L2HDsasft/AoGA5eXlccvjxo1jvXr1YgUFBaywsJBZ\nWlqyM2fOMMYYmzhxIjdiUlJSwry9vdn27dsrlRkWFsY6d+7MGHszgiYQCLiRkp9//pn16NFDZixl\necv2ER4ezoyNjVlBQQGvzIr7qLi/iiN25etX3vr169mUKVO45ezsbMYYY/7+/szX15dL9/f3Z/Pn\nz2eMMbZs2TLWvn17lpOTw0pKSpitrS3z8vJihYWFLC8vj0kkEm6Uadq0aWzz5s28fYaEhLDevXvL\njMfHx4cFBQUxxhiztbVlly9fZoy9+e8qJyenUv6SkhLm6urK4uLiahztvH37NjM2NmZt2rRhIpGI\nHT16lDH25thVjGHevHnccnh4OOvQoQOLiopiJiYm7MGDBzLLl0qlLCEhgVv28/NjPj4+jDHGcnJy\nmKWlJTt58qTMbQMDA5mWlhYzMjJilpaW7OnTp5XyZGRksFatWrHr169XWvfkyROmra3NSxswYAD7\n3//+x0uTNWInb/3Kn5uyfZbZvn078/b2ZoxV3aaCg4N5I17Pnz9n9vb27PHjx4wxxh49esSMjY3Z\nixcvuOvg0KFDMmNhTLHrtvzxvHXrFjM2NuaWpVIp17ZTU1OZSCSSeb0EBwczV1fXSjGUHZOLFy8y\nIyOjautWdizKkxX32bNnGWOVz1f5ZTc3N+6YP3r0iGlqarKvvvqKMcZYdHQ0r44CgYAtX76cMcZY\nYmIi09PTY1lZWezZs2fMwMCAq+/XX3/N5s6dy2339OlTpqOjw16/fs0YYyw3N5cVFxdXutbKL5ed\nuxMnTjDGGFuzZg1r1qwZi4uLY4y9+UxYvHgxF6e9vX2lY9KqVSulj+rXh4qfLaR+ZeZmsv1/7680\nQrfy/Ep2PvU8KyguqLfYlNUla9Qjdm9DIBDgk08+gbq6OtTU1ODg4MDNuTp27BjWrFkDe3t7ODo6\nIjY2FklJSTWWKRKJMGDAAABAly5dcOfOnSrzqqurc/PlXF1doampicTERCXUrLJu3brh5MmT8PPz\nw59//skbWauIlRsN69evH8RiMYRCIWxsbNC3b1+oqalBS0sL5ubmXP1SU1NhZGTEK8fIyEiuOWy9\ne/fG7NmzsXbtWty4cUPmK2/Wrl0LV1dX2NjYVFsWYwxDhw7Fhg0bcO/ePcTExGD69OlIS0vjzYNZ\nvXo1EhMT8c0333Bprq6uGDlyJHr16oWgoKBK9anKuXPnMHnyZACAWCzGyJEjcfbs2Ur5Ll++jC1b\ntiAlJQUPHjyAj49PpW9TefnyJQYPHsyN5MpLnv84Falf+TZw4sQJdOvWDdbW1li3bh33oueq2hSr\nMJp66dIl3L17F/3794e9vT0GDBgAoVDIPRGvoaGBzz77TKG6Vrxuy9phcnIyvLy8YGVlBW9vb6Sn\npyMzM5Pb1tvbGwBgYmICXV1dPHjwQGbdZc3hLNu2S5cuePToEQoLC6usm6zrvrq4a1J2fAwNDdGi\nRQsMHToUAODg4ICHDx+isLCQyztx4kQAb9484ODggKioKOjq6mLw4MHYu3cviouLsWPHDkybNo3b\nRkdHB2ZmZvj888+xY8cOvHz5EioqKjXGJRKJ0L9/fwCAvb09WrduzV2jjo6O3Dm+e/euzPZmbGxc\nq3muDQ3NsWsYcgpycCzxGLb+tRWJT//9W6oiUEFX466Y1XUWepn0grqKej1GqRwN4j12DVWTJk24\n31VUVFBcXMwtHz16lPtWi7ctb8WKFdwtk40bN8LExATAmz8i5f8oCwQCqKqq8m6Rlb9dUltdu3bF\n9evXERISgp9++gnfffcdLly4UGlf+fn5XDwCgaBSfao7XhX/oNf0IsaydevXr0dCQgLOnTuHzz77\nDHPnzsWkSZN4eS9cuIC///6b+8P0/PlztGvXDn///TdEon+/eDkrKwspKSncLZ8OHTrA2toaV65c\n4W57BgYG4sCBAwgNDYWGhgZvP7GxsZBIJEhLS6vmaFZdl7LfZXW0zp8/Dw8PD+jr6wMAPv/8c97Q\n/KtXr/DRRx+hX79+mDNnjsz96OnpAXhzy7Ds9/v373N1q4mi9bt37x7mzp2Lq1evwsTEBJcuXcLo\n0aMBVN2mZLGxsZF5iz01NbXafzKqUrEdlpSUAABGjhyJDRs2YPDgwWCMQUtLi3f9lD/fFdtvTcq2\nLevwFBcXgzFWZd3kibts/6qqqlwdgMrXfMW4ZcWirv7mj5Ws6xAAZs6cidGjR6Nly5awsLDgbv0D\nb27hXr58GRcvXkRoaCgcHR1x6tQp6OrqVvtZVLE+5eMUCoU1Hl96yTxRhtfFrxF5PxKXH1xGcSm/\nzdno28Bd6g5dTd16iu7daNQjdoq8x04sFnNzr2oyePBgfPvtt9yH2pMnT97qK0G++uorxMbGIjY2\nFq6urgCAwsJC7N+/H8Cbjsvr16/RsWNHtGvXDikpKcjOzgZjDL/88otc+6iufqmpqRCJRBgxYgTW\nrVvHzbUyMzNDTEwMGGN4+fIljh8/zm2jyAeuVCrFw4cPeWkPHjxA27Zta9w2MTERlpaW+PLLLzFm\nzLKSvDcAACAASURBVBhuvlh5f/zxB+7du4e7d+8iMjISurq6SElJ4XXqAKBly5YQi8VcByM9PR3X\nr1+HpaUlwsPD8cMPP2D79u0ICQmBjo4Ob9sNGzagpKQEMTExWLVqFeLi4mTG27RpU95x9vT05OYK\nvXz5EgcPHkSfPn0qbWdlZYXIyEju6coTJ07A2toawJs/mIMGDUK3bt1qnIfx2WefYdu2bQCAyMhI\nvH79Go6Ojrw8jLFK50/e+pWXk5MDdXV16Ovro7S0lNsvUHWbatq0KW+OZ/fu3ZGUlMS7Tv/6668a\n911Gkev2xYsX3D9jO3fuREFBQfUbyNCsWTO5r/W3rVsZMzMzbrsbN25wo6JlFLkWy77dJykpCbGx\nsejatSuAN+1PT08Pc+bMqfTEem5uLjIzM+Hi4gJ/f39YWVkhISEBBgYGKCoq4kYWyz6vFCXr8wGQ\n/zOioaM5dvWjuLQYl9IuYdPlTYi8H8nr1Jk1N8MUxykY2mlog+jUKXuOXaPv2Mk7DD5v3jz07t2b\nNwm7qltYGzduhIqKCvdgQ//+/fHo0aNK+QQCQaURt4rrq6Knp4fr16/D1tYWM2bMwC+//AJVVVW0\natUK8+bNg6OjI3r06IFWrVpVuY/yv8uqX5nw8HA4Ojpyt4vKXgEydOhQNG/eHJ06dcKnn37KezCj\nYt2qq4+7uzsuX77MS7t06RI8PT2rrH9ZWf/9739hbW0Ne3t7nD17FgsWLKhyG6DyiNijR49gb2/P\nlXn48GH4+vrC3t4effr0wddff41OnTrh1atXmDZtGvLy8tCnTx/Y29tzr1+Ijo5GYGAg9uzZAwMD\nA2zfvh3e3t4yX8fw5ZdfYvz48dzDE0uWLAFjDNbW1ujevTvGjh0LLy+vStsNGDAAQ4YMgZOTE+zs\n7LB3717uj/DOnTsRERGBkJAQ2Nvbw97eHt9++22l+gHAd999h/DwcHTo0AEzZszATz/9xK1bs2YN\nWrdujStXrsDHxwdt2rRBbm6uQvUrf26sra3x2WefwcLCAl27dkW7du24dWFhYTLblKenJ/Ly8mBn\nZ4fZs2dDR0cHx44dQ0BAAOzs7GBhYYGvv/660r6qouh1+8knn8DR0RF3795FixYtqi1bFg8PD+Tn\n53Pxy9pf2bKurq7MulXVEasqbj8/P5w4cQI2NjZYvXo1HBwc5NpO1rqSkhI4ODhg0KBB+PHHH3nH\nYOLEiVBRUcFHH30E4M3t0/T0dLx48QJDhgyBra0trK2tYWhoiKFDh0JVVRWbNm1Cnz590KVLF6iq\nqlb7WVTVsr29PR4+fMh7Zcy9e/egoaGBNm3aVFk3QmQpZaWIS49D4JVAhNwJQX5xPrfOUGSIsbZj\nMcZmDAzFhvUYJZ+bm5tSO3YN4rti3wUaxm84CgoKYGlpibi4OO7Wmp2dHY4fPw5jY+N6jo6QD4NQ\nKERubi60tLRkrp80aRI6derEPRVfl2bPng17e3tuXunq1atRWlqKhQsX1nks5P3EGEPys2ScTTmL\njLwM3jpdDV14tPOAZUvLOnnKtbaU1W+hjh2pE0FBQcjLy4Ofnx+OHz+OkydPvtMXyRJC+FRUVPDy\n5ctKHbtHjx6hd+/eMDQ0xMmTJyvNLa0LGRkZ+OSTTxAVFYWCggL06NEDERERtZpjST48D3Me4kzK\nGaRmp/LStdW04Sp1haOhI1SENT/wU9+oY1cD6tgRRYSHh9PTa0Qu1FaIIqi9vDtPXz1F6N1QJGQl\n8NLVVdTRzbgburfujiaqTarYuuFRVr+lUT8VWzbHji4qQgghpHHILcxFRGoEYh7HoJT9+2S2UCCE\no6EjXKWuEKmLqimhYQkPD1fqQzY0YkcIIYSQBq+guACX0i4h6kEUCksKeessW1qid9ve0NPSq6fo\n3h6N2BFCCCGk0SspLUHM4xhEpEYgr4j/tL5UR4o+7frAqKl8L43/EDTq150QIi961xSRF7UVoghq\nL7XHGEN8Zjy2RG/BiaQTvE6dvrY+RluPxjjbcdSpq4BG7AghhBDSoKQ8T8HZlLN49JL/jthmTZqh\nd9ve+D/27js6qvPMH/j3zox676jOqJiOEKbZgAoIsI+Nk9jZxcYxLtiOs3GI0zY5i40ROGtvNnFL\nnLa4YMy6pW1sx7/YRjCSaBYGgYwAgZBGDRUk1IXKzNzfH9cacZEEI2bmTvt+zuEs874zcx/2vJEf\nveV558TNgUrg3NR4uMeOiIiIXEJzbzN2V+9G1cUqWXuAJgDZ2mwsSlwEjcoz56S4x84KPBVLRETk\n+joHOrGnZg/KW8pl7RqVBjcl3YRlKcvgr1G+xqISFDsVu379equ+wM/PD6+++qrdArIXztjRZLDW\nFFmLY4Umg+Pl6vqH+1FcW4zDjYdhEk2WdgEC5sXPQ54uD6F+oU6MUDkOn7F7//33sWnTpqveayiK\nIp5//nmXTOyIiIjINQ2ZhvB5w+fYV7cPg6ZBWd+0qGnIT8tHbFCsk6JzbxPO2KWnp+PcuXPX/IJp\n06ahsrLS7oHZijN2RERErsUsmlHWVAa9QY+eoR5ZX3JoMlalr0JKWIqTonMuXil2DUzsiIiIXIMo\nijjddhqFNYVo62+T9UUHRmNl2kpMi5oGQRCcFKHz2Stvua6zwtXV1TAYDDY/nMhVsNYUWYtjhSaD\n4wWo66rD62Wv472K92RJXYhvCO6Yege+u/C7mB493auTOnuyKrG75557cODAAQDAG2+8gVmzZmHm\nzJncW0dERETjau1rxTtfvoPXy15HfXe9pd1P7Yf81Hx8f/H3MT9hPuvR2ZlVS7ExMTFobGyEr68v\nZs+ejT/+8Y8IDw/H17/+dVRVVV3r404hCAK2bNnCcidEREQK6h7sxt6avTjWfAwiRlMMtaDGosRF\nyNZmI9An0IkRupaRcidbt25Vbo9deHg4Ojs70djYiEWLFqGxsREAEBISgp6enmt82jm4x46IiEg5\nl4YvYX/9fhxqOASj2WhpFyBgTtwcrEhdgXD/cCdG6NoULVA8d+5cPPfcczAYDLj99tsBAA0NDQgL\nC7M5ACJXwFpTZC2OFZoMbxgvRrMRpY2lKKktwSXjJVlfRmQGVqatxJTgKU6KzvtYldi99tpr2Lx5\nM3x9ffHf//3fAICDBw/iW9/6lkODIyIiItdkFs0obynH3pq96BrskvUlhCRgVdoqpEakOik678Vy\nJ0RERGQ1URRRdbEKu6t3o6WvRdYXGRCJFakrMCtmFk+5TpLid8WWlJSgrKwMPT09locLgoBNmzbZ\nHAQRERG5vobuBuyu3g1Dp0HWHuQThFxdLubHz4dapXZOcATAysRu48aNeP/995GdnY2AgABHx0Sk\nOG/YB0P2wbFCk+Ep46W9vx2FNYU4eeGkrN1X7YslyUtwc9LN8NP4OSk6upxVid2uXbtQUVGBhIQE\nR8dDRERELqJ3qBd6gx5Hm47CLJot7SpBhfnx85Gry0Wwb7ATI6QrWbXHLjMzE3v27EF0dLQSMdkF\n99gRERFdn0HjIA7UH8CB+gMYNg/L+mbFzMKK1BWICoxyUnSeSdG7Yg8fPoxnn30W9957L+Li4mR9\nOTk5NgfhCEzsiIiIJsdkNuGL81+guLYYfcN9sr7U8FSsTFuJxNBEJ0Xn2RQ9PHHkyBF8/PHHKCkp\nGbPHrr6+foJPOV9BQQFvniCreMo+GHI8jhWaDHcZL6IoouJCBQqrC9Ex0CHriwuKw6r0VUiPSOdJ\nVwcYuXnCXqyasYuKisK7776LVatW2e3BjsYZO5oMd/nhS87HsUKT4Q7jpbqjGp+d+wxNvU2y9jC/\nMKxIXYE5cXN4n6sCFF2KTUlJQVVVFXx9fW1+oFKY2BEREU2sqacJu6t341zHOVl7gCYAOdocLExc\nCI3K6qpoZCNFE7sdO3agtLQUmzdvHrPHTqVyzSyeiR0REdFYHZc6sKdmD75s/VLWrlFpcFPSTViW\nsgz+Gn8nRee9FE3sJkreBEGAyWSyOQhHYGJHk+EOyyXkGjhWaDJcabz0D/ejuLYYhxsPwySO/rdb\ngIB58fOQp8tDqF+oEyP0booenqiurrb5QURERKS8IdMQDjUcwv66/Rg0Dcr6pkdPR35qPmKCYpwU\nHdkb74olIiLyQGbRjKNNR6E36NE71CvrSw5Nxqr0VUgJS3FSdHQle+UtE26Q27x5s1VfsGXLFpuD\nICIiIvsQRRGnLpzCb0t/i4/OfCRL6qIDo3HP7HuwYd4GJnUeasIZu+DgYJSXl1/1w6IoYv78+ejs\n7HRIcLbgjB1NhivtgyHXxrFCk6H0eKntrMVn1Z+hobtB1h7iG4LlqcuRNSWLpUtclMP32PX39yMj\nI+OaX+Dnx0t/iYiInKm1rxW7q3fjTPsZWbu/xh/LUpZhceJi+Kh9nBQdKYl77IiIiNxU10AX9AY9\njjUfg4jR/+apBTUWJS5CtjYbgT6BToyQrKXoqVgiIiJyHZeGL2Ff3T583vg5jGajpV2AgMy4TCxP\nXY5w/3AnRkjOwoV2IsCu9/SRZ+NYocmw93gxmo3YX7cfL3/+MvbX75cldTdE3oDHFjyGO2fcyaTO\ni3n0jF1BQQHy8vK40ZmIiNyaWTSjvKUce2v2omuwS9aXEJKAVWmrkBqR6qToyBZ6vd6uvwBwjx0R\nEZGLEkURZy+exe7q3Wjta5X1RQZEIj81HzNjZkIQBCdFSPai6B671tZWBAQEICQkBEajETt37oRa\nrcb69etd9q5YIiIid9bQ3YDPzn2G2q5aWXuQTxDydHm4Mf5GqFVqJ0VH9lJZWYvdu8/Z7fusysrW\nrFmDqqoqAMCTTz6J559/Hi+++CJ+9KMf2S0QImfivimyFscKTcb1jJe2/ja8X/E+Xj36qiyp81X7\nIk+XhyduegILExcyqfMAlZW1+MMfqrB37wq7fadVM3Znz55FVlYWAGDXrl04cOAAQkJCMHPmTLz0\n0kt2C4aIiMhb9Qz2oKi2CEebjsIsmi3tKkGFBQkLkKPNQbBvsBMjJHu6dAn49a/P4eTJfNhz55hV\niZ1arcbg4CDOnj2L8PBwaLVamEwm9Pb2XvvDRG6AB2zIWhwrNBnWjJdB4yD21+/HwfqDGDYPy/pm\nx87GitQViAyIdFCEpDSTCTh8GCgqAs6dU9k1qQOsTOxuvfVWrF27Fu3t7bj77rsBACdPnkRSUpJ9\noyEiIvISRrMRR84fQVFtEfqH+2V9qeGpWJW+CgkhCU6KjuxNFIHTp4HPPgMuXpTaVCppZjYszH7P\nsepU7MDAAN588034+vpi/fr10Gg00Ov1aG5uxj333GO/aOyIp2JpMnj/J1mLY4UmY7zxIooiTrSe\nwJ6aPegY6JD1TQmegpVpK5Eekc6Trh7k/Hngk0+AWvk5GAwP1+LChSrEx+dj2zYFT8X6+/vjscce\nk7XxBxsREdHknLt4Drurd6Opt0nWHu4fjhWpKzAndg4TOg/S1QUUFgLl5fJ2f38gNxdYtEiLqiqg\nsHCP3Z454Yzd+vXr5W/8aqCJoigbdDt37rRbMPbEGTsiInIVTT1N2F29G+c65GUtAjQByNHmYGHi\nQmhUHn1ngFcZHAT27QMOHgSMo5eDQKUCFi0CcnKAwCuu8HV4Hbv09NFp4La2Nrz55pu44447oNVq\nUVtbi48++ggPPPCAzQEQERF5msqqSuw+shtdg12ovlgN/1h/RCdEW/p9VD64KekmLE1ZCn+NvxMj\nJXsym4GyMmDPHqCvT943fTqwahUQFeXYGCZM7AoKCix/X716Nf7xj38gOzvb0rZv3z5s27bNocER\nKYX7pshaHCt0LZVVldi+ezuaYppQUVaB8OnhMJ40IgtZiEmIwY3xNyJXl4tQv1Bnh0p2VFUFfPop\n0Cq/IAQJCcDq1YBOp0wcVs37Hjp0CDfddJOsbfHixTh48KBDgiIiInJHPYM9+PU/f41TIadg7hmt\nRafJ0KC/tR/f/fp3ERMU48QIyd5aWqSE7twVl0eEhgIrVwJz5gBKbpu06lRsbm4uFi5ciGeeeQYB\nAQHo7+/Hli1b8Pnnn6O4uFiJOCeNe+yIiEgp3YPd2F+3H0eajmBf8T4MJA1Y+sL8wpAWkQZtpxY/\nuOcHToyS7Km3F9i7Fzh6FLJadL6+wLJlwM03Az4+1n+fonfF7tixA/feey9CQ0MRERGBjo4OLFiw\nAG+//bbNARAREbmrroEu7Kvbh6NNR2ESTQAA1Ve3dQb7BkMXrkNUQBQEQYCvyteZoZKdDA9LhyL2\n7QOGhkbbBQG48UZg+XIg2IkXhFg1Yzeirq4O58+fR3x8PLRarSPjshln7GgyuG+KrMWxQgDQOdCJ\nfXX7UNZUZknoRqi71GiobcCUOVNQe7wWuiwdBs8O4sHlD2JaxjQnRUy2EkWpbElhIdDdLe/LyJAO\nRsTFXf/3KzpjN8Lf3x+xsbEwmUyorq4GAKSlpdkcxGT97Gc/w8GDB6HT6fD6669Do+ERcSIicrzO\ngU6U1JbgWPOxMQldUmgScrW5yIjMwJlzZ1B4tBDtF9sR2xqL/OX5TOrcWG2tVGD4/Hl5e2ysdDAi\nI8M5cY3Hqhm7f/7zn3j44YfR1CQvqCgIAkwm0wSfcozjx4/jV7/6Fd566y08++yzSEtLG/f2C87Y\nERGRvXRc6kBxbTGOtxyHWTTL+pJDk5Gny0NaRBqLC3uY9nZg927g1Cl5e1AQsGIFMG+eVJvOHhSd\nsfvud7+LzZs34/7770fglRX1FHbw4EHccsstAKQ7bN944w2XvdaMiIjcW3t/O0rqSlDeUj4modOG\naZGry0VqeCoTOg9z6RJQVASUlkq16UZoNNKhiGXLAD8/58V3NVYldp2dnXjsscdcYuB2dHQgPj4e\nABAaGoqLIzfpEtmA+6bIWhwr3qGtvw0ltVJCJ0I+i6IL1yFXmwtduO6a/13keHEvJpOUzBUXS8nd\n5TIzgfx8ICzMObFZy6oJxIcffhivv/66XR/8yiuvYMGCBfD398dDDz0k67t48SLuvPNOBAcHQ6fT\n4Z133rH0hYeHo/urXYtdXV2IjIy0a1xEROS9LvRdwF9O/gW/Lf0tjrcclyV1qeGpeCjrITyY9SBS\nIzhL50lEETh5Evjtb6W9dJcndVot8OijwF13uX5SB1i5x27ZsmUoLS2FVqvFlClTRj8sCNddx+5v\nf/sbVCoVPvnkE1y6dAlvvPGGpW/dunUAgNdeew1lZWW4/fbbceDAAcycORPHjx/HCy+8gDfffBPP\nPvss0tPTcffdd4/9h3GPHRERWam1rxXFtcWoaK0YM0OXHpGOXF0uUsJSnBQdOVJjo5TM1dXJ2yMj\npZOu06crU2BY0T12jzzyCB555JFxg7hed955JwDgiy++QENDg6W9r68Pf/3rX1FRUYHAwEAsXboU\nX//61/HWW2/hueeew9y5cxEXF4ecnBxotVr89Kc/nfAZDz74IHRf3eERHh6OrKwsy5S4Xq8HAL7m\na77ma7724tctvS34/Z9+D0OXAbosHQDAcMwAAFi5YiVytbk4V3YO1R3VSMlLcXq8fG2/1/Pm5WH3\nbuDDD6XXOp3Uf/68HllZwGOP5UGtdtzzR/5uMBhgT5OqY+cITz31FBobGy0zdmVlZVi2bBn6Lrs9\n94UXXoBer8cHH3xg9fdyxo4mQ6/XW/5HR3Q1HCueobm3GUWGIpxqOzWm74bIG5Cry0VSaJLNz+F4\ncT2Dg1Jx4YMHAaNxtF2tBhYuBHJzgYAA5eNSdMZOFEW88cYbeOutt9DY2IikpCTcd999eOihh2ze\nY3Dl53t7exEaKr8YOSQkBD09PTY9h4iIqKmnCUW1RTjddnpM37SoacjV5SIhJMEJkZGjmc3S9V97\n9wKXzR0BAGbMkJZdPWHbvlWJ3bPPPoudO3fixz/+MVJSUlBXV4df/vKXOH/+PJ566imbArgyOw0O\nDrYcjhjR1dWFkJAQm55DdDX8jZqsxbHins73nEeRoQiV7ZVj+qZHT0euNhfxIfF2fy7Hi2uoqpL2\n0V24IG9PSABuuUU6IOEprErstm/fjqKiItk1Yrfccguys7NtTuyunLGbOnUqjEYjqqqqkPFVKefj\nx49j9uzZk/7ugoIC5OXl8X9YREReqrG7EXqDHmcvnh3TNzNmJnK0OZgSPGWcT5InaGkBPv0UOHdO\n3h4aCqxcCcyZo8zBiKvR6/WyfXe2smqPXWxsLGpqahAUFGRp6+3tRVpaGlpbW6/rwSaTCcPDw9i6\ndSsaGxuxfft2aDQaqNVqrFu3DoIg4NVXX8XRo0exZs0aHDx4EDNmzLD+H8Y9djQJ3AdD1uJYcQ/1\nXfUoqi1C1cUqWbsAwZLQxQXbcLGnlThenKO3F9izBygrk0qZjPD1BbKzgZtuAnx8nBffeBTdY3fr\nrbfivvvuw3PPPQetVguDwYAnn3zScgPE9XjmmWewbds2y+tdu3ahoKAATz/9NH73u99hw4YNiI2N\nRXR0NP7whz9MKqkjIiLvVNdVB71Bj+qOalm7AAGzYmchR5uD2KBYJ0VHjjY8LB2K2LcPGBoabRcE\n4MYbgeXLgeBg58WnBKtm7Lq6urBx40a89957GB4eho+PD9auXYvf/OY3CA8PVyLOSRMEAVu2bOFS\nLBGRFzB0GlBkKEJNZ42sXYCAOXFzkJ2SjZigGCdFR44mikB5OVBYCFyxTR8ZGcDq1UCsi+bzI0ux\nW7dutcuM3aTKnZhMJrS1tSE6OhpqtdrmhzsSl2KJiDybKIpSQldbBEOnQdYnQEBmXCaytdmIDox2\nToCkCINB2kd3/ry8PTZWSui+2q7v8uyVt1iV2L355pvIysrC3LlzLW3Hjx9HeXk51q9fb3MQjsDE\njiaD+2DIWhwrzieKImo6a1BkKEJtV62sTyWokBmXiRxtDiIDnF+7guPFcdrbgc8+A05fUbkmKAhY\nsQKYNw9QqZwT2/VQdI/d5s2bcezYMVlbUlIS7rjjDpdN7IiIyLOIoohzHedQZChCfXe9rE8lqJA1\nJQvZKdmICIhwUoSkhP5+oKgIOHxYqk03QqMBliwBli4F/PycF5+zWTVjFxERgba2Ntnyq9FoRFRU\nFLq6uhwa4PXijB0RkWcQRRFVF6tQVFuEhu4GWZ9aUEsJnTYb4f6uueeb7MNolJK5oiJgYEDel5kJ\n5OcDYWHOic0eFJ2xmzFjBv785z/j7rvvtrT97W9/c/mTqqxjR0TkvkRRxNmLZ1FkKEJjT6OsTy2o\nMS9+HpalLGNC5+FEETh1Slp27eiQ92m1UoHhBDe+LMQpdez27duH2267DatWrUJaWhrOnTuH3bt3\n4+OPP8ayZcvsFow9ccaOJoP7YMhaHCuOJ4oiKtsrUWQoQlNvk6xPLagxP2E+liYvRZi/60/PcLzY\nprFRujGirk7eHhkpXQE2fbrzCwzbi6IzdsuWLcOXX36Jt99+Gw0NDVi0aBFefvllJCcn2xwAERER\nICV0p9tOo6i2CM29zbI+jUqD+fHzsTRlKUL9Qif4BvIUnZ1S6ZIvv5S3BwQAubnAwoWAixfncJpJ\nlztpaWlBghvMeXLGjojIPYiiiFNtp1BkKEJLX4usT6PSYEHCAixNXooQP94Z7ukGB4GSEuDQIWlP\n3Qi1Gli0CMjJkZI7T6TojF1HRwcef/xx/PnPf4ZGo0F/fz8++OADlJaW4uc//7nNQRARkfcxi2ac\nvHASxbXFaO2TX0/po/LBwsSFWJK8BMG+Hn5VAMFsBo4eBfbuBfr65H0zZkjLrpHOr17jFqyq8PKd\n73wHoaGhqK2thd9XZ4hvvvlmvPvuuw4NzlYFBQV23ZBInovjhKzFsWI7s2jGly1f4veHf48/n/yz\nLKnzVftiafJS/OCmH2B1+mq3T+o4Xq5OFIGzZ4Hf/x746CN5UpeYCDz0EHD33Z6d1On1ehQUFNjt\n+6xaio2OjkZTUxN8fHwQERGBjq+OpYSGhqL7yrs7XASXYmkyuMGZrMWxcv3MohknWk+guLYYbf1t\nsj5ftS8WJS7CkuQlCPQJdFKE9sfxMrGWFunGiHPn5O1hYVLpkjlzPOdghDUUvXkiIyMDxcXFSEhI\nsCR2dXV1WL16NU5fWfLZRTCxIyJyDWbRjPKWcpTUlqD9Urusz0/th8VJi3FT0k0eldDRxHp6pCXX\nsjJpxm6Ery+QnQ3cdBPg4+O8+JxF0T12jzzyCP7lX/4FP//5z2E2m3Hw4EFs2rQJjz32mM0BEBGR\nZzKZTShvKUdxbTE6BuQFyPw1/licKCV0AT4euhueZIaHgQMHgP37gaGh0XZBAObPB/LygGD3Xnl3\nCVbN2ImiiF//+tf44x//CIPBgJSUFHznO9/BE088AcFF50k5Y0eTweUSshbHyrWZzCYcbzmOktqS\ncRO6m5NuxuKkxfDX+DspQuVwvEizcuXlUvmSK3dvZWQAq1cDsbHOic2VKDpjJwgCnnjiCTzxxBM2\nP5CIiDyTyWxCWXMZ9tXtQ+dAp6wvQBOAm5NvxqLERV6R0JHEYJAKDDfJ60wjNla6MSI93SlheTSr\nZuz27NkDnU6HtLQ0NDU14Wc/+xnUajWee+45TJkyRYk4J00QBGzZsoVXihEROZjRbERZk5TQdQ3K\n7w8P9AnEzUlSQuen8eKb2b1MW5t0BVhlpbw9OBhYvhyYNw9QWVWXw/ONXCm2detW5Q5PTJ8+HZ9+\n+ilSUlKwbt06CIIAf39/tLW14YMPPrA5CEfgUiwRkWMZzUYcbTqKfXX70D0oX2ML9AnE0uSlWJi4\nEL5qXydFSErr7weKioDDh6XadCM0GmDJEmDpUsCP+f24FD0VO1LWZHh4GHFxcZZ6dvHx8Whvb7/W\nx52CiR1NBvfBkLU4VoBh0zCONB3B/rr96BnqkfUF+QRhacpSLEhYwIQO3jNejEagtBQoLgYGBuR9\nc+cCK1ZIZUxoYorusQsNDUVzczMqKiowa9YshISEYHBwEMPDwzYHQERE7mHYNIwvzn+B/fX7RorE\nBAAAIABJREFU0TvUK+sL9g3G0mQpofNRe2GtCi8lisDJk8Du3UCH/JwMdDrpYIQb3ELqUaxK7DZu\n3IhFixZhcHAQL730EgBg//79mDFjhkODI1KKN/xGTfbhjWNlyDQkJXR1+9E3LL/vKcQ3BMtSluHG\n+BuZ0I3Dk8dLQ4N0MKK+Xt4eFSVdATZtmncVGHYVVi3FAkBlZSXUajUyMjIAAGfOnMHg4CDmzJnj\n0ACvF5diiYhsM2QaQmljKQ7UH0D/cL+sL9Qv1JLQaVRWzRGQh+jslGboTpyQtwcEALm5wMKFgFrt\nnNjcmaJ77NwREzuaDG/ZB0O284axMmgcRGljKQ42HByT0IX5hWFZyjLMi5/HhM4KnjReBgaAffuA\nQ4ekPXUj1Gpg0SIgJ0dK7uj6OHyP3fTp0y3XhSUnJ08YRF1dnc1BOEpBQQHLnRARWWnAOIDPGz7H\noYZDuGS8JOsL9w9Hdko25k6Zy4TOy5jNwJEj0jVg/fI8HzNnAitXApGRzonNE4yUO7GXCWfsSkpK\nkJ2dbXnoRFw1aeKMHRGRdQaMAzjUcAiHGg5hwCg/0hjhH4FsbTbmxs2FWsX1NW8iisDZs1I9ugsX\n5H2JiVKB4ZQU58TmibgUew1M7IiIru7S8CVLQjdoGpT1RQZEIjslG5lxmUzovFBzM/Dpp0B1tbw9\nLEyaoZs9mwcj7M3hS7GbN2+e8CEj7YIgYNu2bTYHQeRsnrQPhhzLE8ZK/3A/DtYfRGlj6ZiELiog\nCjnaHMyJmwOVwKsBbOVu46WnB9izBzh2TJqxG+HnB2RnA4sXAz48/OzSJkzs6uvrIVwlHR9J7IiI\nyD30DfXhYIOU0A2ZhmR90YHRyNHmYHbsbCZ0XmhoCDh4UDoccXmJWkEA5s+XrgELCnJefGQ9LsUS\nEXm4vqE+HKg/gMPnD49J6GICY5CjzcGs2FlM6LyQKALHjwOFhdJs3eVuuEGqRxcb65zYvI3Dl2Kr\nr1xYn0BaWprNQRARkf31DvVif91+fHH+Cwyb5TcFxQbFIlebi5kxM7n64qVqaqR9dE1N8va4OOnG\niPR058RFtplwxk6luvZvboIgwGQy2T0oe+CMHU2Gu+2DIedxh7HSM9iD/fVSQmc0G2V9cUFxyNXl\nYkb0DCZ0CnDF8dLWJp10rayUtwcHS3e6ZmUBVqQAZGcOn7Ezm802fzkRESmne7Ab++v240jTkTEJ\n3ZTgKcjV5mJ69HQmdF6qvx/Q64EvvpBq043w8QGWLAGWLgV8fZ0WHtmJR++x27JlCwsUE5HH6xro\nwr66fTjadBQmUb6KkhCSgFxtLqZGTWVC56WMRqC0FCgulm6PuNzcuUB+PhAa6pzYaLRA8datWx1b\nx+6WW27BJ598AgCWQsVjPiwIKC4utjkIR+BSLBF5us6BTuyr24eyprIxCV1iSCJydbm4IfIGJnRe\nShSBkyele107OuR9Op20jy4hwSmh0TgcvhR7//33W/7+8MMPTxgEkSdwxX0w5JpcYax0XOrAvrp9\nONZ8bExClxSahDxdHtIj0vkz2gU4a7w0NACffALU18vbo6Kkk67TprHAsKeaMLH71re+Zfn7gw8+\nqEQsRER0FRcvXURJbQmOtxyHWZTvg04OTUaeLg9pEWlM6LxYZ6c0Q3fihLw9IADIywMWLADUvEjE\no1m9x664uBhlZWXo6+sDMFqgeNOmTQ4N8HpxKZaIPEV7fztK6kpQ3lI+JqHThmmRq8tFangqEzov\nNjAAlJQAn38u7akboVZLt0VkZ0vJHbkuhy/FXm7jxo14//33kZ2djQCODCIiRbT1t6GkVkroRMh/\n4OvCdcjT5UEXrnNOcOQSTCbgyBHptGt/v7xv5kzpXtfISKeERk5i1YxdREQEKioqkOBGuyw5Y0eT\n4Qr7psg9KDFWLvRdQHFtMU60nhiT0KVFpCFXmwttuNahMZB9OGq8iCJw9qxUYLitTd6XmAjccguQ\nkmL3x5IDKTpjl5ycDF8WtyEicqjWvlYU1xajorViTEKXHpGOXF0uUsL4X2tv19wsJXRXXhAVFibN\n0M2ezYMR3syqGbvDhw/j2Wefxb333ou4uDhZX05OjsOCswVn7IjIXbT0tqC4thgnL5wck9BlRGYg\nV5uL5LBkJ0VHrqKnB9izBzh2TJqxG+HnJ+2hW7xYKjZM7knRGbsjR47g448/RklJyZg9dvVXnqUm\nIiKrNPc2o8hQhFNtp8b0TY2ailxtLhJDE50QGbmSoSHgwAFg/35g+LIrf1UqYP586bRrUJDTwiMX\nY9WMXVRUFN59912sWrVKiZjsgjN2NBncY0fWssdYaeppQlFtEU63nR7TNy1qGnJ1uUgIcZ89zTQx\nW8aL2QwcPy7N0vX0yPtuuEEqMBwTY3uM5BoUnbELCgpCbm6uzQ8jIvJm53vOQ2/Q40z7mTF906On\nI1ebi/iQeCdERq6mpkYqMNzcLG+Pi5MSuvR058RFrs+qGbsdO3agtLQUmzdvHrPHTqVSOSw4W3DG\njohcRUN3A4oMRTh78eyYvpkxM5GjzcGU4ClOiIxcTVubdDDizBW5f3AwsGIFkJUlLcGS57FX3mJV\nYjdR8iYIAkwm07h9ziYIArZs2YK8vDwusRGRU9R31aOotghVF6tk7QIES0IXFxw3wafJm/T1AUVF\nwBdfSEuwI3x8gCVLgKVLARan8Ex6vR56vR5bt25VLrEzGAwT9ul0OpuDcATO2NFkcI8dWcuasVLX\nVQe9QY/qDnk9CgECZsXOQo42B7FBsQ6MklzFtcaL0SjdFlFcDAwOjrYLAjB3rjRLFxrq+DjJ+RTd\nY+eqyRsRkSsxdBpQZChCTWeNrF2AgDlxc5CjzUF0YLSToiNXIopARYV0r2tnp7xPp5MKDMdzuyVd\nB6vvinU3nLEjIiWIoigldLVFMHQaZH0CBGTGZSJHm4OowCjnBEgup75e2kd3ZbWwqCjpYMTUqSww\n7I0UnbEjIiI5URRR01mDIkMRartqZX0qQYW5cXORrc1GZAAv6iRJR4c0Q1dRIW8PDARyc4EFCwC1\n2jmxkedgYkcE7rGja6usqsTuI7tx8sRJRKdGwyfaB8ZQo+w9KkGFrClZyE7JRkRAhJMiJVei1+tx\n0015KCkBDh0CLj9vqFZLt0Xk5AD+/s6LkTwLEzsiomuorKrE/+z+H/Qn9eM4jsNX5QtjqRFZM7MQ\nnRANtaCWEjptNsL9w50dLrmAyspafPrpORQVleOll8xISkpHdLTW0j9rlnSvawTzf7Izq/bYVVdX\n48knn8SxY8fQ29s7+mFBQF1dnUMDvF7cY0dEtugb6oOh0wBDpwFv/t+baI1tHfOe4IZg/Nvaf8Oy\nlGUI8w9zQpTkio4ercXLL1fhwoV89PdLbUZjIbKyMpCVpcXq1UBKinNjJNej6B67e++9FxkZGXjh\nhRfG3BVLROQJBowDlkSupqMGLX0tlr7uoW7ZewUISAhJwGztbNw+9XalQyUXNDQEnD4NlJcD77xz\nDn19+bL+4OB8hIfvwcMPa3kwghzKqsTu5MmT2L9/P9Tc1UkeinvsvM+gcRB1XXWo6ayBodOApp4m\niBj/t2UVVFAJKoT6hWKgagDzbpoHP40fwkycpfNmZjNQXS0lc6dPS8kdAJhMo0X9u7v1yMrKQ1IS\nEBmpYlJHDmdVYpeTk4OysjIsWLDA0fEQETnEsGkY9d31qOmoQU1nDc73nIdZNE/4fpWgQmJIIlIj\nUrEsdBk+O/wZArQBMDQb4Kfxw+DZQeQvz5/w8+SZRBFoapKSuRMngMt2J1moVGaEhUn3uvb3jy67\n+vpOPN6I7MWqPXaPP/443nvvPdx1112yu2IFQcC2bdscGuD14h47Iu9mNBvR0N1gWVpt6G6ASZz4\nCsSR5VVduA6pEalICUuBr3r0DqfKqkoUHi3EkHkIvipf5N+Yj2kZ05T4p5AL6OgAvvxSSuja2sZ/\nT0wMkJkJ+PvX4q9/rYKf32jiPzhYiAcfzMC0adrxP0xeT9E9dn19fVizZg2Gh4fR0NAAQKrhJHBO\nmYhchMlswvme81Ii11mDuq46GM3Gq35mSvAUpIanQheugzZcC3/NxDUnpmVMYyLnZfr7gZMnpWRu\nonOCwcHAnDlSQjdlykhhYS1CQ4HCwj0YGlLB19eM/HwmdaQM3jxBBO6xc0dm0Yzm3mbUdEh75Gq7\najFkGrrqZ2ICY5AakYrU8FRow7UI9Amc9HM5Vjyb0QicOSMlc2fPyuvOjfD1BWbMkJK51FRApRr7\nnhEcL2Qth8/YGQwGyx2x1dXVE70NaWlpNgdBRHQtoiiita8VNZ01qOmoQW1XLQaMA1f9TGRAJFLD\nU5EaIc3KBfsGKxQtuRNRBGprpWTu5ElgYJxhpVIB6elSMjdtmpTcEbmiCWfsQkJC0NPTAwBQTfDr\niCAIMI3364wL4IwdkXsTRRHtl9othx0MnQb0D/df9TNhfmGWGbnUiFSE+oUqFC25o9ZWKZn78kug\nq2v89yQmSsnc7NlAUJCy8ZF3sVfe4nZLsd3d3Vi5ciVOnTqFzz//HDNnzhz3fUzsiNyLKIroGOiw\nHHao6axB79A4Rw4vE+IbYpmNSw1PRbh/OPf+0lV1d0unWcvLgebm8d8TESElc5mZQFSUsvGR91L0\n8IQrCQwMxMcff4x///d/Z+JGdsN9MM7RNdBlmY2r6ahB1+AE0yZfCfIJspxa1YXrEBUQpXgix7Hi\nfgYHgVOnpGSupkZaer1SYKB0zVdmJpCUBLvVm+N4IaW5XWKn0WgQHR3t7DCI6Dr0DvXKllYvXrp4\n1ff7a/wts3GpEamICYzhjBxZxWQCzp0bLR5sHOeAtEYDTJ8uJXPp6QBr8JMncLvEjsgR+Bu1Y/QP\n98uWVtv6JygA9hVftS+0YVrLPrm44DiohKscOXQCjhXXJYpAQ4O0Z+7ECVjuab2cIEgnWTMzpZOt\nfn6OjYnjhZSmaGL3yiuvYMeOHThx4gTWrVuHN954w9J38eJFPPzww/jss88QHR2N5557DuvWrQMA\nvPjii/jggw+wZs0a/PjHP7Z8hr+5E7mWAeMAajtrLSdXL79vdTw+Kh+khKVYllcTQhJcLpEj19fe\nPnoI4uIEk8BTpoweggjlmRryYJNO7Mxm+ZUoE52YHU9iYiI2b96MTz75BJcuXZL1Pf744/D390dr\nayvKyspw++23Y+7cuZg5cyZ++MMf4oc//OGY7+MeO7IX7oO5PiP3rY4UBb7afasAoBbUSA5LthQF\nTgxNhEblXgsHHCuuoa9v9BBEY+P47wkNlZK5OXOk672cgeOFlGbVT9QjR47ge9/7Ho4fP46Bywr8\nTLbcyZ133gkA+OKLLyw3WADSzRZ//etfUVFRgcDAQCxduhRf//rX8dZbb+G5554b8z233XYbjh8/\njsrKSjz22GN44IEHrI6BiK7f5fetGjoNaOxptPq+1dTwVCSFJsFH7aNgxORJhoel/XLl5dL+OfM4\nQ8/Pb/QQhFZrv0MQRO7CqsTugQcewNe+9jW89tprCAycfKX2K10503bmzBloNBpkZGRY2ubOnQu9\nXj/u5z/++GOrnvPggw9aiiyHh4cjKyvL8pvTyHfzNV+PuPw3a2fH4yqvl+UsQ2N3I/72z7+hqacJ\nwVODYRJNMBwzAAB0WToAsLxOzUpFfEg8uk93Iz4kHv96+7/CV+0LvV6P2tpapOalutS/73pe5+Xl\nuVQ8nv7abAbeeUeP6mpArc7D0BBgMEj9Op30/ro6PZKSgHvuycPUqcC+fXoYDKP9HC987YqvR/5u\nMBhgT1bVsQsNDUVXV5fd9rRt3rwZDQ0Nlj12JSUlWLt2LZqamizv2b59O95++23s3bv3up7BOnZE\nk2cWzTjfc95y2KG+qx7D5uGrfmZK8BTLydVr3bdKZA1RlGrMlZdLy61f1cofIyVFmpmbOVMqV0Lk\nzhStY3fnnXfik08+wa233mrzA4GxM3bBwcHo7u6WtXV1dSEkJMQuzyO6Fv1ls3XeZOS+1ZGTq5O5\nb1UXroMuXHdd9626M28dK0ro7JQOQJSXAxcujP+e6OjRfXMREcrGdz04XkhpViV2ly5dwp133ons\n7GzEXbYDVRAE7Ny5c9IPvXLmb+rUqTAajaiqqrIsxx4/fhyzZ8+e9HdfrqCgwDIVTkTy+1YNnQYY\nOg28b5Wc6tIl6X7W8nLpvtbxBAdLp1kzM4H4eO6bI8+i1+tly7O2smoptqCgYPwPCwK2bNli9cNM\nJhOGh4exdetWNDY2Yvv27dBoNFCr1Vi3bh0EQcCrr76Ko0ePYs2aNTh48CBmzJhh9fdfGRuXYsnb\n2Xrfqi5chzD/MIWiJW9hNAJnz0rJ3JkzUjHhK/n4SHXmMjOBtDRApVI+TiIlueVdsQUFBdi2bduY\ntqeffhodHR3YsGGDpY7df/3Xf+Gee+657mcxsSNvJIoiOgc6LXXkDJ0G9AxNsEHpKyG+IZY6crxv\nlRxFFIG6OimZq6gABsaZKBYE6QaIzEzpRghfX+XjJHIWxRO7vXv3YufOnWhsbERSUhLuu+8+rFix\nwuYAHIWJHU2GO++D6RrostSRs+a+1UCfQMtsXGpEqlPuW3Vn7jxWnOHCBSmZKy8HuiYYmgkJo8WD\ngz1spZ/jhayl6OGJV199FZs2bcIjjzyCxYsXo66uDvfeey+2bduGb3/72zYH4SjcY0eeaOS+1ZFk\nztr7VkdOrsYGxTKRI4fq6RktHnxZsQOZ8HApmcvMlA5EEHkrp+yxu+GGG/DnP/8Zc+fOtbSVl5fj\nrrvuQlVVld2CsSfO2JGnGLlvdeTk6oX+CY4LfuXy+1Z14TpMCZ7Ca7rI4QYHR4sHV1dLS69XCggY\nLR6cnMxDEESXU3QpNioqCk1NTfC9bMPD4OAgEhIS0N7ebnMQjsDEjtzV5fetGjoNaO5tvur7fVQ+\nlmu6UiNSER8cD7VKrVC05M1MJimJKy+XkrrhcUoeajTA1KlSMnfDDYCaQ5NoXIomdl/72teQkpKC\nX/ziFwgKCkJvby/+4z/+AwaDAR9++KHNQTgCEzuaDGfugxkyDaGuq85yctXa+1ZHllbd8b5Vd+bt\ne6ZEETh/frR4cF/f+O/T6UaLB/t7cc1qbx8vZD1F99j94Q9/wD333IOwsDBERkbi4sWLWLJkCd55\n5x2bA3Ak7rEjVzRy3+rI0qq1962OHHZIDk3mfaukuIsXR4sHT7RQExs7Wjw4jFVyiKzilD12I+rr\n63H+/HkkJCQgOTnZbkE4AmfsyFWYzCY0dDdYDjvUd9XDJI5TuOsrAgTEh8RbTq6mhKXAT+OnYMRE\nkv5+qTRJeTlQXz/+e0JCRpO5uDjumyO6Xg5fihVF0XJyzmy+ymyCi1aNZGJHznL5fauGTgPquuqu\ned9qXFCcpY4c71slZxoelooGl5dLRYTH+/Hv5yctsWZmAlotiwcT2YPDl2JDQ0PR89XNyxrN+G8T\nBAGm8UqGE7kZW/bBmEUzWnpbLHXk6rrqMGgavOpnYgJjLEur3njfqjvzxD1TZjNgMEjJ3KlT0gnX\nK6lU0uGHzEzpMIQPdwNYxRPHC7m2CRO7iooKy9+rq6sVCYbIHYiiiAv9F2TXdFlz3+rIYQdduA4h\nfiEKRUs0PlEEWlqkZO7LL6Xac+NJTpaSuVmzgED+/kHk8qzaY/erX/0KP/nJT8a0v/DCC/jRj37k\nkMBsNXKPLQ9PkK1G7lsdOexg6DSgb3iCo4BfGblvdSSZ432r5Cq6ukYPQbS2jv+eqKjRfXORkcrG\nR+RtRg5PbN26VblyJyEhIZZl2ctFRESgo6PD5iAcgXvsyBYdlzoss3E1HTXXvG812DfYUkdOF65D\nhH8Eb3cglzEwAJw8KSVztbXjFw8OCpKu9MrMlK744vAlUpYi5U727NkDURRhMpmwZ88eWd+5c+cQ\nGhpqcwBEzlRZVYndR3bj2PFjiE6NRqI2EcZQIzoHOq/6uUCfQMtsHO9b9S7usmfKZJIOP5SXS4ch\njMax7/HxAaZPl5K5tDQWD3YEdxkv5Dmumtht2LABgiBgcHAQDz/8sKVdEATExcXhN7/5jcMDJLIH\nURTRM9SDzoFOy58TZ07g49KPYdQa0WxuRrhPOIzFRmTNzEJ0gvzySn+Nv+WaLt63Sq5KFKWyJOXl\nUpmSS5fGvkcQpCQuM1NK6vxYSYfIo1i1FLt+/Xq89dZbSsRjN1yK9S5m0YzeoV5Z4nb5n66BrjG1\n40r3laI/qX/MdwU1BGFp7lJow7SWk6u8b5VcWVublMyVlwOdE0w2x8dLydzs2VLtOSJyLYrePOFu\nSR15nutJ3K75nRgt0KUSVAjzC0O4fzhSxBT8bOnPeN8qubTeXulKr/Jy6Yqv8YSFSclcZiYQE6Ns\nfETkHFYldl1dXSgoKEBRURHa29stBYsFQUBdXZ1DA7QFrxRzH2bRjJ7B0aXSrsEumxO3KwX6BCLc\nP9zypy+mDwOxA/DX+KOlogVp89IAALFDsUzqaEJOvVd4CDh9Wkrmzp0b/xCEv79UmiQzE0hJ4SEI\nZ+MeO7oWe18pZlVi9/jjj6O+vh5PP/20ZVn2l7/8Jb75zW/aLRBHKCgocHYI9JUrE7cxM26DXVe9\nL9UaQT5BssTt8j9h/mHwVfvK3q8Vtdixdwf8bvCzLLMOnh1E/vJ8m+IgsiezGaiuHi0ePDzOJSZq\ntVQ0ODNTKiI8QU15InJBIxNQW7dutcv3WbXHLiYmBqdOnUJ0dDTCwsLQ1dWFxsZG3HHHHTh69Khd\nArE37rFTlismbtaorKpE4dFCDJmH4KvyRf6N+ZiWMc2mOIlsJYpAU9No8eC+CcomarVSMjdzJhAQ\noGyMRGRfDr8r9nLR0dFoamqCj48PkpKScOLECYSGhiIsLGzc+naugImdfblr4kbkTjo6RosHt7WN\n/56YmNHiweHhysZHRI6j6OGJzMxMFBcXIz8/H8uWLcPjjz+OoKAgTJvGmQ1P4e2JG/fBkLXsPVb6\n+0eLB0+0ZTk4WErkMjOBKVO4b86d8GcLKc2qxG779u2Wv7/88svYtGkTurq6sHPnTocFRvZlFs3o\nHuye8ESpPRK3YN9gy8nS8f74qHlrOBEgFQs+c0ZK5s6elYoJX8nXF5gxQ0rmUlMBFavtEJEVrFqK\n/fzzz7F48eIx7aWlpVi0aJFDArOVty3FXi1x6xzoRPdgt10Stwln3PzCmLgRXYUoStd5jRQPHhwc\n+x6VCkhPl5K5adOk5I6IvIOiS7ErV64cdy/drbfeiosXL9ochKN4UrkTJm5E7qmlZfQQRHf3+O9J\nTBwtHhwUpGx8RORc9i53ctUZO7PZDFEUER4ejq6uLlnfuXPnsHTpUrS2ttotGHtytxk7Jm7OxX0w\nZC1rxkp39+ghiJaW8d8TGTl6CCIqyv5xkmvgzxayliIzdprLiiFpriiMpFKp8OSTT9ocgLcwmU2W\nxO3K4rtM3Ijc3+Dg6CEIg2H84sGBgdKsXGamNEvHQxBEZG9XnbEzGAwAgJycHJSUlFgySUEQEBMT\ng8DAQEWCvB5Kz9hdnrhNNOMmwrZ4QnxDZKdImbgROZfJBFRVSclcZaV0KOJKGg0wfbqUzKWnS8WE\niYiupGgdO3dk78RO6cRtvHIgGhXLyRM5S2VlLXbvPoehIRX6+syIjk5Hd7cW/f1j3ysI0knWzEzp\nZKufn/LxEpF7UfTwxPr168cNAIDHlDxh4ubduA+Grqayshavv16FCxfyceKEHoGBK2A0FiIrC4iO\n1lreN2XK6CGI0FAnBkwugz9bSGlWZRLp6emyTLK5uRl/+ctf8K1vfcuhwdnqt+/9Fivnr8S0jGlM\n3IjouphMwGuvncOxY/kYGgKGhqS9chpNPmpq9iAtTWs5BBEX5+xoicjbXfdS7BdffIGCggJ89NFH\n9o7JLgRBwNr316K3shdzps9BQHQAEzcispooSidb9+4F/t//02NgIM/Sp1YDsbHADTfosXVrHg9B\nEJHNFF2KHU9WVhaKiopsDsCRWvpagCTgy9NfYuGyhVd9rwABIX4TJ26hfqFM3Ii8gChKt0EUFo6W\nKlGppBPrvr6ATictuapUQGysmUkdEbkUqzKVwsJCy546AOjr68O7776LWbNmOSwwezLBxMSNror7\nYAiQboYoLBx7Z+vMmeloaSmETpeP+no9VKo8DA4WIj8/wylxkvvgzxZSmlWZzMMPPyxL7IKCgpCV\nlYV33nnHYYHZQ88nPdBl6bBYtxg/yvkREzciGldzs5TQnT0rb/fxAW6+GViyRPtV0rcHHR3liI01\nIz8/A9Omacf/QiIiKyl684Q7EwQBW/ZuweDZQTy4/EFMy5jm7JCIyMVcvAjs2QOcOCFvV6uBBQuA\n7GwgONg5sRGRd1F8j11nZyf+8Y9/4Pz580hISMBtt92GiIgImwNwpNjWWOQvz2dSR0QyPT1AURFw\n9ChgvuzCF0GQypXk5QEu/uONiGhcVs3Y7dmzB3fddRemTZsGrVaL2tpanD59Gn/5y1+wcuVKJeKc\nNHe7K5aci/tgvMOlS8C+fcDnn4+9JWL6dGDFCum069VwrNBkcLyQtRSdsXv88cfxP//zP1i7dq2l\n7U9/+hO+973v4fTp0zYHQUTkSENDUjK3fz8wMCDv0+mAlSuBpCSnhEZEZFdWzdiFh4ejvb0d6ssu\nORweHkZMTAw6OzsdGuD14owdEZlMwJEjQHEx0Nsr74uPlxK6tDSwZAkROZ3iV4q98soreOKJJyxt\nv//978e9aoyIyNnMZulAxN69QEeHvC8qSlpynTmTCR0ReR6rZuyWLl2K0tJSxMbGIjExEY2NjWht\nbcXixYstZVAEQUBxcbHDA7YWZ+xoMrgPxjOIInDmjFS6pLVV3hcaKh2KyMqSigtfL44VmgyOF7KW\nojN2jz76KB599NFrBkRE5CwGg5TQ1dfL2wMCpLIlCxdKdemIiDyZR9ex89B/GhFdpqmXEOuHAAAg\nAElEQVRJSuiqquTtvr5SceGbbwb8/Z0TGxGRtRSvY1dcXIyysjL09fUBAERRhCAI2LRpk81BEBFN\nVnu7tIduouLCOTlAUJBzYiMicharEruNGzfi/fffR3Z2NgICAhwdE5HiuA/GfXR3S8WFy8rGFhee\nO1faRxce7rjnc6zQZHC8kNKsSux27dqFiooKJCQkODoeIqJx9fdLxYVLS8cWF54xQzrpGhPjnNiI\niFyFVYldcnIyfH19HR2L3RUUFCAvL4+/LdE1cYy4rqEh4NAhqbjw4KC8LzUVyM9XtrgwxwpNBscL\nXYter4der7fb91l1eOLw4cN49tlnce+99yIuLk7Wl5OTY7dg7ImHJ4jcm9E4Wlz4q629FgkJUkLH\n4sJE5CkUPTxx5MgRfPzxxygpKRmzx67+ytoCRG6I+2Bch9kMfPmldDDiyottoqOlJdcZM5yX0HGs\n0GRwvJDSrErsnnzySXz00UdYtWqVo+MhIi8likBlJbBnj+OKCxMReTqrlmJTUlJQVVXlVvvsuBRL\n5D4MBmD3bqChQd4eGDhaXFhjdXEmIiL3Y6+8xarEbseOHSgtLcXmzZvH7LFTueivz0zsiFzf+fNS\nceFz5+Ttvr7AkiVScWE/P+fERkSkJEUTu4mSN0EQYDKZbA7CEZjY0WRwH4yy2tqkPXQVFfJ2tVqa\nncvOdt3iwhwrNBkcL2QtRQ9PVFdX2/wgIqLubkCvB44dG1tcOCsLyM11bHFhIiJPN6m7Ys1mM1pa\nWhAXF+eyS7AjOGNH5DpYXJiI6OrslbdYlZ11d3fj/vvvh7+/PxITE+Hv74/7778fXV1dNgdARJ5r\naEi6/uvll4EDB+RJXVoa8OijwN13M6kjIrIXqxK7jRs3oq+vDydOnEB/f7/l/27cuNHR8REpwp5V\nv0lK4D7/XEro9u6V3xiRkADcf7/0JzHReTFeL44VmgyOF1KaVXvs/vnPf6K6uhpBX+1mnjp1Knbs\n2IG0tDSHBkdE7sVsBsrLpX104xUXzs8Hpk/nbRFERI5i1R47nU4HvV4PnU5naTMYDMjJyUFdXZ0j\n47tu3GNHpBxRBE6flooLX7gg7wsLk4oLz53L4sJERBNR9FTsI488glWrVuHHP/4xtFotDAYDXnzx\nRTz66KM2B0BE7q2mRiou3Ngobw8MBHJygAULWFyYiEgpVs3Ymc1m7NixA//7v/+LpqYmJCQkYN26\nddiwYQMEF11T4YwdTQZrTU3eRMWF/fykwsKeWlyYY4Umg+OFrKXojJ1KpcKGDRuwYcMGmx9oD6Wl\npfjBD34AHx8fJCYmYufOndBwSoBIEW1t0pLryZPydo1GKi68bJnrFhcmIvJ0Vs3Ybdy4EevWrcOS\nJUssbQcOHMD777+Pl156yaEBjqe5uRkRERHw8/PDpk2bMH/+fHzzm9+UvYczdkT21dUllS4pK5P2\n1I0YKS6clyftpyMioslT9Eqx6OhoNDY2wu+ydZWBgQEkJyfjwpU7pRW2ZcsWzJs3D9/4xjdk7Uzs\niOyjvx8oKQEOHx5bXHjmTGD5ctahIyKylaIFilUqFcyX3/8Dad+dsxOn2tpafPbZZ7jjjjucGge5\nP9aaGmtwUCpb8vLLwMGD4xcXXrvW+5I6jhWaDI4XUppVid2yZcvw1FNPWZI7k8mELVu2IDs7e9IP\nfOWVV7BgwQL4+/vjoYcekvVdvHgRd955J4KDg6HT6fDOO+9Y+l588UUsX74czz//PIDR2zDefPNN\nqNXqScdBROMzGoFDh6SETq+XFxdOTAQeeMB9iwsTEXk6q5Zi6+vrsWbNGjQ1NUGr1aKurg7x8fH4\n8MMPkZycPKkH/u1vf4NKpcInn3yCS5cu4Y033rD0rVu3DgDw2muvoaysDLfffjsOHDiAmTNnyr7D\naDTia1/7Gn7yk59gxYoV4//DuBRLNClmM3D8uJTMXXlbYEyMdJ8riwsTETmGonvsAGmWrrS0FPX1\n9UhOTsbixYuhsqHa6ObNm9HQ0GBJ7Pr6+hAZGYmKigpkZGQAAB544AEkJCTgueeek332rbfewg9/\n+EPMmTMHAPBv//ZvWLt2rfwfxsSOyCojxYULC6UTr5cLC5P20GVmsrgwEZEjKVruBADUajVuvvlm\n3HzzzTY/FMCY4M+cOQONRmNJ6gBg7ty54+5PWL9+PdavX3/NZzz44IOW2zLCw8ORlZVlqSc08r18\nzdcA8NJLL3nl+EhJyUNhIbB/v/Rap5P6m5v1yMwEvv3tPGg0rhOvK7we+burxMPXrv2a44WvJ3o9\n8neDwQB7snrGzt6unLErKSnB2rVr0dTUZHnP9u3b8fbbb2Pv3r2T/n7O2NFk6PV6y//ovEFjozRD\nV10tb/fzA5YsAW66yTOLC9uDt40Vsg3HC1lL8Rk7e7sy+ODgYHR3d8vaurq6EBISomRY5KW85Qfv\nhQtSceFTp+TtGg2waJFUXDgw0DmxuQtvGStkHxwvpLRrJnaiKKKmpgYpKSl2vd3hyqvIpk6dCqPR\niKqqKsty7PHjxzF79uzrfkZBQQHy8vL4Pyzyel1dgF4PHDs2trjwvHlAbi6LCxMROYNer5ctz9rq\nmkuxoigiKCgIvb29Nh2WGGEymTA8PIytW7eisbER27dvh0ajgVqtxrp16yAIAl599VUcPXoUa9as\nwcGDBzFjxoxJP4dLsTQZnrpc0tc3WlzYZJL3zZolHYyIjnZObO7KU8cKOQbHC1lLsQLFgiBg3rx5\nqKystPlhAPDMM88gMDAQv/jFL7Br1y4EBATgP//zPwEAv/vd73Dp0iXExsbivvvuwx/+8IfrSuqI\nvN3lxYUPHZIndenpwLe/DfzrvzKpIyLyNFYdnnjqqaewa9cuPPjgg0hOTrZklYIgYMOGDUrEOWmc\nsSNvZDRKs3MlJdJVYJdLSgLy84HUVOfERkREE1P08MS+ffug0+lQVFQ0ps9VEzuAe+zIe5jN0v65\noqLxiwvn5wPTprG4MBGRq1F8j5274owdTYa77oMRRemE6549Y4sLh4dLe+jmzGFxYXty17FCzsHx\nQtZSvNxJe3s7/vGPf6C5uRk//elP0djYCFEUkZSUZHMQRDQ5oijVoCssBM6fl/cFBQE5OcD8+VIZ\nEyIi8h5WzdgVFRXhm9/8JhYsWID9+/ejp6cHer0ezz//PD788EMl4pw0ztiRp2psBHbvBmpq5O1+\nfsDSpVJxYV9f58RGRETXR9G7YrOysvCrX/0KK1euREREBDo6OjAwMICUlBS0trbaHIQjMLEjT8Pi\nwkREnkuxcicAUFtbi5UrV8rafHx8YLqyMJaLKSgosOuGRPJcrjxOOjuB//s/4He/kyd1KpW03Pr9\n7wOrVzOpU4orjxVyPRwvdC16vR4FBQV2+z6rduDMmDED//znP3Hrrbda2goLCzFnzhy7BeII9vx/\nFJHS+vqA4mLgiy/GLy68YgUQFeWc2IiIyD5Gqnds3brVLt9n1VLsoUOHsGbNGtx2223405/+hPXr\n1+PDDz/E3//+dyxatMgugdgbl2LJXQ0MAAcPSn+GhuR9GRlS6ZL4eOfERkREjqHoHjsAaGxsxK5d\nu1BbW4uUlBTcd999Ln0ilokduZvhYam48L59Y4sLJydLCZ1O55TQiIjIwRRP7ADAbDajra0NMTEx\nEFy80ikTO5oMZ9aaGikurNcD3d3yvthYKaGbOpXFhV0F65LRZHC8kLUUPTzR0dGB9evXIyAgAFOm\nTIG/vz/uu+8+XLx40eYAHImHJ8iViSJQUQH89rfABx/Ik7rwcOCuu4DvfIc3RhAReTJ7H56wasbu\nG9/4BjQaDZ555hmkpKSgrq4OTz/9NIaGhvD3v//dbsHYE2fsyFVdrbhwcPBocWG12jnxERGR8hRd\nig0LC0NTUxMCL6un0N/fj/j4eHRdeTGli2BiR66ooUEqLmwwyNtZXJiIyLspuhQ7ffp0GK74L1Ft\nbS2mT59ucwBErsDRS/atrcC77wKvvipP6jQaKaH7wQ+kmTomda6P2ztoMjheSGlW1bFbsWIFVq9e\njfvvvx/Jycmoq6vDrl27sH79erz++usQRRGCIGDDhg2OjpfIrXR2Anv3AuXl0hLsCJUKuPFGKZkL\nDXVefERE5FmsWoodOdFz+UnYkWTucnv37rVvdDYQBAFbtmyxFP4jUlJvL1BSMn5x4dmzgeXLWVyY\niIikWV29Xo+tW7cqX+7EnXCPHTnDwABw4ABw6NDY4sI33CDdFsHiwkREdCVF99gReTpb98EMDwP7\n9wMvvyxdA3Z5UpecDDz0EPCtbzGp8wTcM0WTwfFCSrNqjx0Rjc9sBsrKgKKiscWF4+Kk4sI33MA6\ndEREpAwuxRJdB1EETp4E9uwB2tvlfRER0h662bOlQxJERETXYq+8hTN2RJMgisC5c1Jx4aYmeV9w\nMJCbK512ZXFhIiJyBqvnE06dOoVt27bh8ccfBwCcPn0a5eXlDguMSEnW7IOprwfefBPYtUue1Pn7\nS0uu3/8+sHAhkzpPxz1TNBkcL6Q0qxK7P/3pT8jJyUFjYyN27twJAOjp6cGPfvQjhwZH5ApaW4F3\n3gFee01eXNjHB1i2DHjiCSA7m8WFiYjI+azaYzd9+nS8++67yMrKQkREBDo6OjA8PIz4+Hi0tbUp\nEeeksY4d2aqjQyou/OWX4xcXzs0FQkKcFx8REbk/p9Sxi4qKwoULF6BSqWSJXWJiIlpbW20OwhF4\neIKuV2+vVLLkyJGxxYXnzJEORkRGOic2IiLyTIrWsbvxxhvx1ltvydree+89LFq0yOYAiFyBXq/H\nwIB0KOLll4HSUnlSN3Uq8J3vAN/8JpM6b8c9UzQZHC+kNKtOxf7mN7/BqlWr8Nprr6G/vx+rV6/G\nmTNn8Omnnzo6PiKHqqysxSefnINeX44XXzQjOTkd0dFaS39KinQwQqu9ypcQERG5CKvr2PX19eGj\njz5CbW0tUlJScPvttyPEhTcYcSmWrqWioha//GUVzp/Pt9wUYTQWIisrA7NmaVlcmIiIFGOvvIUF\nisnrmEzAsWPAs8/uQXv7Clmfvz8wf/4e/PznK5jQERGRYhQtUFxbW4utW7eirKwMvb29siDOnDlj\ncxBESjCZgPJy6fqvzk6gr290i2lvrx7z5uUhPh6IjFQxqaMJ6fV6nrQnq3G80P9v7+6Do6rvPY5/\nNk8kkAQChkBCQ9AQBJFQAZUHIYiWyYhSGBWxIKAVxqIXdGyrg0AYZChTpHhFpFKrIhLFOzo+0cFe\nw4ZAIUANuZQHCSiPWiIE8ggh2ez945SVk4DshuWc3c37NZORPb+ze37JfIkffud7zrGaV8HugQce\nUM+ePTV//nxFR0df6zkBftXQYNyyJD9fKiv7cXtYWIMiI40+uvPnpZQUY3tUVIM9EwUA4Cp5dSq2\nbdu2KisrU3gQ3VKfU7FoaJB275aczqbPc23dWvrZzw5r584DiokZ4dleW/ulJk9OV48eXC0BALCO\npadiR40apfz8fN15551X3jmA5OTkcIPiFsjtNgJdfr70ww/msZgYadAg6dZbpVatuuqWW6Qvv8zT\n+fNhiopq0IgRhDoAgHUu3KDYX7xasTt58qQGDhyojIwMdezY8cc3Oxz661//6rfJ+BMrdi2P2y3t\n3Wus0DW+b3Z0tDRwoHTbbcafG6MPBt6iVuAL6gXesnTF7tFHH1VUVJR69uyp6Ohoz8EddJgjALjd\n0tdfG4Hu3/82j7VqJd1+uxHqaA8FAIQ6r1bs4uLidPz4ccXHx1sxJ79gxS70ud1SSYnxPNfvvzeP\nRUUZq3ODBhmnXwEACGSWrtj16dNHp06dCqpgh9DldksHDxqB7vhx81hkpNE/N2iQ1KaNPfMDAMAu\nXgW7O++8UyNHjtSUKVOUlJQkSZ5TsY8++ug1nSBwgdstffutEeiOHjWPRURIAwZIgwdLsbG+fzZ9\nMPAWtQJfUC+wmlfBrqCgQMnJyZd8NizBDlY4dMgIdIcPm7dHREj9+klDhkgB/IQ7AAAswSPFENCO\nHDEC3bffmreHh/8Y6OgQAAAEu2veY3fxVa8NDZe/E39YWNhlx4DmOnrUuMr14EHz9rAw6ZZbpDvu\nkNq2tWVqAAAErMumsosvlIiIiLjkV2RkpCWTRMtx/Lj07rvSG2+YQ92FQPfUU9KoUf4Pdf68OSRC\nG7UCX1AvsNplV+x2797t+fM333xjyWTQcn3/vbFC9/XX5u0Oh5SZKQ0dKrVvb8vUAAAIGl712C1e\nvFjPPvtsk+1LlizRM888c00mdrXosQsOJ04YgW7vXvN2h0O6+WZp2DCpQwdbpgYAgGX8lVu8vkFx\nZWVlk+0JCQk6ffr0VU/iWiDYBbbSUuNZrhctDEsyAt1NNxmBLjHRnrkBAGA1S25QnJeXJ7fbLZfL\npby8PNPYwYMHA/6GxTk5OcrKyuIeQgHk5Ekj0P3rX8Z96S7Wq5cR6P5zq0RLca8peItagS+oF1yJ\n0+n0ay/mT67YpaWlyeFw6MiRI0pNTf3xTQ6HkpKS9Pzzz+u+++7z22T8iRW7wHLqlBHodu1qGuhu\nvFHKypI6dbJlapL45QvvUSvwBfUCb1l6KnbixIl65513rvpgViLYBYbTp6WNG6XiYqnxXXMyMoxA\nl5xsy9QAAAgYlga7YESws9eZM0ag27mzaaBLT5eGD5dSUuyZGwAAgcZfuYW7C8Ovysulzz6TXnlF\n+uorc6i7/nrpscekCRMCL9Rxryl4i1qBL6gXWM2rZ8UCV1JZKRUUSP/8p+RymcfS0owVuq5dbZka\nAAAtBqdicVWqqqRNm6QdO6T6evNYaqoR6Lp1s2duAAAEC0tudwJcTnW1tHmztH27VFdnHuvSxQh0\n119v3JcOAABYgx47+KSmRvrf/5Vefln6xz/MoS45WfrVr4w+uhtuCK5QRx8MvEWtwBfUC6zGih28\ncvastGWLtHWrdP68eaxzZ+O2JRkZwRXmAAAINfTY4SedO2eEuS1bpNpa81hSknHKtUcPAh0AAFeD\nHjtcU7W1UmGhcbr13DnzWGKiEeh69iTQAQAQSOixg8n588ZVrkuXSnl55lB33XXS/fdLTzxhPNc1\nlEIdfTDwFrUCX1AvsBordpBkXASxfbsR6mpqzGPt2xs9dL17S2H8UwAAgIBFj10LV1dn3FR40ybj\nnnQXS0iQhg2T+vQh0AEAcC3RY4erUl9vPPKroMB4asTF2rY1Al1mphQebs/8AACA74JuHebEiRMa\nPHiwhg8frpEjR+rUqVN2TymouFzGUyL++7+ldevMoS4+Xho1Svqv/5JuuaVlhTr6YOAtagW+oF5g\ntaBbsUtMTNTmzZslSW+//bZWrlyp5557zuZZBT6XSyoulvLzpfJy81hcnHTHHUaYiwi6igAAABcE\ndY/dK6+8oqioKE2bNq3JGD12hoYGI9Bt3CidPm0ei42VhgyR+vWTIiPtmR8AAGjhPXbFxcWaOnWq\nzpw5o+3bt9s9nYDU0CDt2mWs0JWVmcfatJEGD5YGDCDQAQAQSiztsVu2bJn69++v6OhoTZkyxTRW\nVlamMWPGKDY2VmlpacrNzfWM/elPf9Lw4cP10ksvSZIyMzNVWFioF198UfPnz7fyWwh4FwLd8uXS\nRx+ZQ11MjHTXXdKMGdKgQYS6i9EHA29RK/AF9QKrWbpil5KSotmzZ2v9+vU6e/asaWz69OmKjo5W\naWmpioqKdM899ygzM1O9evXS008/raefflqSVFdXp8j/JJL4+HjVNn7OVQvldkt79khOp/TDD+ax\n6GgjyN12m9SqlS3TAwAAFrClx2727Nk6duyY3nzzTUlSdXW12rdvr927dys9PV2SNGnSJCUnJ2vh\nwoWm927fvl3PPvuswsPDFRkZqTfeeENdunRpcgyHw6FJkyYpLS1NktSuXTv17dtXWVlZkn78V1Sw\nvx42LEv79kmvv+7U6dNSWpoxfuiQU5GR0q9+laXbb5e2bg2M+fKa17zmNa95zWt5/nzo0CFJxgWh\n/ohktgS7F154QcePH/cEu6KiIg0ZMkTV1dWefZYsWSKn06lPPvmkWccI9Ysn3G5p/37J6ZS+/948\nFhUl3X67NHCgcfoVAAAENn/lljA/zMVnjkYPGa2qqlJ8fLxpW1xcnCob3zkXcrulkhJp5UopN9cc\n6qKijKtcZ86U7ryTUOeLi/8FBfwUagW+oF5gNVuuim2cSGNjY1VRUWHaVl5erri4OCunFdDcbumb\nb6QNG6Rjx8xjkZHGFa6DBxtXvAIAgJbJlmDXeMUuIyND9fX1OnDggKfHrri4WL17976q4+Tk5Cgr\nK8tzXjtYffutEeiOHDFvj4j4MdDFxtozt1AR7DUC61Ar8AX1gitxOp1+Xdm1tMfO5XKprq5O8+bN\n0/Hjx7Vy5UpFREQoPDxc48ePl8Ph0F/+8hd99dVXGjVqlLZs2aKePXs261ih0GN3+LAR6P7TV+kR\nHi7172+cdmVREwCA4BeUPXbz589X69attWjRIq1evVoxMTFasGCBJGn58uU6e/asOnbsqAkTJmjF\nihXNDnXB7uhRadUq6c03zaEuPNxYoZsxQ8rOJtT5E30w8Ba1Al9QL7Capadic3JylJOTc8mxhIQE\nffTRR1ZOJ+AcO2Zc5XrggHl7WJj0858bz3Nt186WqQEAgCAQ1M+K/SkOh0Nz584Nih67774zAt3+\n/ebtYWFSZqY0dKiUkGDL1AAAwDV0ocdu3rx5wXsfOysEQ4/dv/9tBLp9+8zbHQ6pTx8j0HXoYMvU\nAACAhYKyxw6G0lJp7VppxQpzqHM4pJtvlqZPl8aMIdRZiT4YeItagS+oF1jNltudtFQ//CDl50u7\ndxv3pbvYTTdJWVlSYqItUwMAACEgpE/FBkqP3cmTRqD717+aBrqePY1Al5Rky9QAAICN6LHzUiD0\n2JWVGYHu//6vaaDr0cMIdJ072zI1AAAQQOixC2CnT0sffywtWyYVF5tDXffu0tSp0vjxhLpAQh8M\nvEWtwBfUC6xGj50flZdLGzdKRUVSQ4N57IYbpOHDpS5d7JkbAAAIfZyK9YOKCqmgQPrqK8nlMo91\n62YEutRUS6YCAACCkL9yS0iv2OXk5FzTiycqK6VNm6R//lOqrzePde1qBLq0tGtyaAAAEAIuXDzh\nL6zYNUNVlbR5s7R9e9NA97OfGYGuWzfjvnQIDk6n0/arpxEcqBX4gnqBt1ixs0FNjRHotm2T6urM\nY126GIHu+usJdAAAwB6s2HmhpkbaskUqLJTOnzePJScbgS49nUAHAACahxU7C5w7ZwS6rVul2lrz\nWKdORqDLyCDQAQCAwMB97C7h3DnjxsJLlxr/vTjUdewojRsnTZtm3GSYUBcauNcUvEWtwBfUC6wW\n0it2vl4VW1tr9M/94x/S2bPmscRE40kRvXoR5gAAgH9wVayXfDlXff68cYXr5s1GP93FOnQwAt1N\nN0lhrG8CAIBrgB47P6irk3bsMO5FV11tHmvfXho2TLr5ZgIdAAAIDi0ystTXG1e4vvyytH69OdS1\nayeNHi1Nny5lZhLqWgr6YOAtagW+oF5gtRa1Yldfbzz2q6DAeGrExdq2lYYOlfr2lcLD7ZkfAADA\n1WgRPXYul1RUZAS68nLzfvHx0h13SD//uRTRomIuAAAIFPTYeeGVV/KUknKDjhzpqjNnzGOxsUag\n69ePQAcAAEJDSHeQvfzyRs2d+z86cOCwZ1ubNtLIkdKMGdJttxHqYKAPBt6iVuAL6gVX4nQ6lZOT\n47fPC+lY06VLjiTp22/zlJraVYMHSwMGSFFR9s4LAABAkud+u/PmzfPL54V0j92wYW5FREi9ejm1\neHEWgQ4AAAQkeuy80K2blJIiJSc3EOoAAEDIC+keu65dJZfrS40YcYPdU0GAow8G3qJW4AvqBVYL\n6RW7jh3zNGJEunr06Gr3VAAAAK65kO6xC9FvDQAAhBh/5ZaQPhULAADQkhDsANEHA+9RK/AF9QKr\nhXSwy8nJ4S8VAAAIWP6+QTE9dgAAADajxw4AAAAmBDtA9MHAe9QKfEG9wGoEOwAAgBBBjx0AAIDN\n6LEDAACACcEOEH0w8B61Al9QL7AawQ4AACBE0GMHAABgM3rsAAAAYEKwA0QfDLxHrcAX1AusRrAD\nAAAIESHdYzd37lxlZWUpKyvL7ukAAAA04XQ65XQ6NW/ePL/02IV0sAvRbw0AAIQYLp4A/Ig+GHiL\nWoEvqBdYjWAHAAAQIjgVCwAAYDNOxQIAAMCEYAeIPhh4j1qBL6gXWI1gBwAAECLosQMAALAZPXYA\nAAAwIdgBog8G3qNW4AvqBVYj2AEAAIQIeuwAAABsRo8dAAAATAh2gOiDgfeoFfiCeoHVCHYAAAAh\nImh77HJzczVjxgyVlpZecpweOwAAECxadI+dy+XSBx98oNTUVLunAgAAEDCCMtjl5ubqwQcflMPh\nsHsqCBH0wcBb1Ap8Qb3AakEX7C6s1o0bN87uqSCE7Ny50+4pIEhQK/AF9QKrWRrsli1bpv79+ys6\nOlpTpkwxjZWVlWnMmDGKjY1VWlqacnNzPWNLlizR8OHDtXjxYr377rus1sHvzpw5Y/cUECSoFfiC\neoHVLA12KSkpmj17th599NEmY9OnT1d0dLRKS0v17rvv6oknntCePXskSc8884w2bNigZ599Vnv2\n7NGqVauUnZ2tkpISzZw508pvwe+sWKb3xzGa+xm+vM+bfa+0z0+Nh8IpkWv9Pfjr85vzOf6uFW/2\nC+V64XeLb/u25FqR+N3i676BXC+WBrsxY8Zo9OjR6tChg2l7dXW1PvzwQ82fP1+tW7fW4MGDNXr0\naL3zzjtNPuMPf/iD1q9fr7/97W/KyMjQ0qVLrZr+NcEvX9/2vVZ/mQ4dOnTFYwcCfvn6tu+1qBdq\nxb/H4HdLYOB3i2/7BnKws+V2Jy+88IKOHz+uN998U5JUVFSkIUOGqLq62rPPkram5CcAAAs5SURB\nVCVL5HQ69cknnzTrGOnp6Tp48KBf5gsAAHAt3XDDDTpw4MBVf06EH+bis8b9cVVVVYqPjzdti4uL\nU2VlZbOP4Y8fDgAAQDCx5arYxouEsbGxqqioMG0rLy9XXFycldMCAAAIarYEu8YrdhkZGaqvrzet\nshUXF6t3795WTw0AACBoWRrsXC6Xzp07p/r6erlcLtXW1srlcqlNmzYaO3as5syZo5qaGm3atEmf\nfvqpJk6caOX0AAAAgpqlwe7CVa+LFi3S6tWrFRMTowULFkiSli9frrNnz6pjx46aMGGCVqxYoZ49\ne1o5PQAAgKBmy1WxdqmoqNBdd92lvXv3qrCwUL169bJ7Sghg27Zt08yZMxUZGamUlBStWrVKERG2\nXG+EAHfixAmNHTtWUVFRioqK0po1a5rc1gloLDc3VzNmzFBpaandU0GAOnTokAYMGKDevXvL4XBo\n7dq1uu66637yPUH3SLGr0bp1a61bt073339/kws4gMZSU1O1YcMG5efnKy0tTR9//LHdU0KASkxM\n1ObNm7VhwwY9/PDDWrlypd1TQoC78HjM1NRUu6eCAJeVlaUNGzYoLy/viqFOamHBLiIiwqsfCiBJ\nnTp1UqtWrSRJkZGRCg8Pt3lGCFRhYT/+Kq2oqFBCQoKNs0EwyM3N5fGY8MrmzZs1dOhQzZo1y6v9\nW1SwA5rj8OHD+vvf/657773X7qkggBUXF+u2227TsmXLNH78eLungwB2YbVu3Lhxdk8FAS45OVkH\nDx7Uxo0bVVpaqg8//PCK7wnKYLds2TL1799f0dHRmjJlimmsrKxMY8aMUWxsrNLS0pSbm3vJz+Bf\nSS3H1dRLRUWFHnnkEb399tus2LUAV1MrmZmZKiws1Isvvqj58+dbOW3YpLn1snr1albrWpjm1kpU\nVJRiYmIkSWPHjlVxcfEVjxWUneApKSmaPXu21q9fr7Nnz5rGpk+frujoaJWWlqqoqEj33HOPMjMz\nm1woQY9dy9Hceqmvr9dDDz2kuXPnqnv37jbNHlZqbq3U1dUpMjJSkhQfH6/a2lo7pg+LNbde9u7d\nq6KiIq1evVolJSWaOXNm0D/3HD+tubVSVVWl2NhYSdLGjRt10003Xflg7iD2wgsvuCdPnux5XVVV\n5Y6KinKXlJR4tj3yyCPu5557zvM6OzvbnZyc7B44cKD7rbfesnS+sJev9bJq1Sp3hw4d3FlZWe6s\nrCz3+++/b/mcYQ9fa6WwsNA9dOhQ9/Dhw92/+MUv3EePHrV8zrBPc/5fdMGAAQMsmSMCg6+1sm7d\nOne/fv3cd9xxh3vSpElul8t1xWME5YrdBe5Gq2779+9XRESE0tPTPdsyMzPldDo9r9etW2fV9BBg\nfK2XiRMncpPsFsrXWrn11luVn59v5RQRQJrz/6ILtm3bdq2nhwDia61kZ2crOzvbp2MEZY/dBY37\nE6qqqhQfH2/aFhcXp8rKSiunhQBFvcBb1Ap8Qb3AW1bUSlAHu8bJNzY2VhUVFaZt5eXliouLs3Ja\nCFDUC7xFrcAX1Au8ZUWtBHWwa5x8MzIyVF9frwMHDni2FRcXq3fv3lZPDQGIeoG3qBX4gnqBt6yo\nlaAMdi6XS+fOnVN9fb1cLpdqa2vlcrnUpk0bjR07VnPmzFFNTY02bdqkTz/9lD6pFo56gbeoFfiC\neoG3LK0Vf13pYaW5c+e6HQ6H6WvevHlut9vtLisrc//yl790t2nTxt21a1d3bm6uzbOF3agXeIta\ngS+oF3jLylpxuN3c0A0AACAUBOWpWAAAADRFsAMAAAgRBDsAAIAQQbADAAAIEQQ7AACAEEGwAwAA\nCBEEOwAAgBBBsAMAAAgRBDsAaGTy5MmaPXu2Xz/ziSee0IsvvujXzwSAxiLsngAABBqHw9HkYd1X\n67XXXvPr5wHApbBiBwCXwNMWAQQjgh2AgLJo0SJ16dJF8fHxuvHGG5WXlydJ2rZtmwYOHKiEhAQl\nJyfrqaeeUl1dned9YWFheu2119S9e3fFx8drzpw5OnjwoAYOHKh27drpoYce8uzvdDrVpUsXLVy4\nUImJierWrZvWrFlz2Tl99tln6tu3rxISEjR48GDt2rXrsvs+/fTTSkpKUtu2bdWnTx/t2bNHkvn0\n7r333qu4uDjPV3h4uFatWiVJ2rdvn+6++2516NBBN954oz744IPLHisrK0tz5szRkCFDFB8fr5Ej\nR+rUqVNe/qQBhCKCHYCA8fXXX+vVV1/Vjh07VFFRoS+++EJpaWmSpIiICL388ss6deqUtmzZoi+/\n/FLLly83vf+LL75QUVGRtm7dqkWLFunxxx9Xbm6ujhw5ol27dik3N9ez74kTJ3Tq1Cl99913evvt\ntzV16lSVlJQ0mVNRUZEee+wxrVy5UmVlZZo2bZruu+8+nT9/vsm+69evV0FBgUpKSlReXq4PPvhA\n7du3l2Q+vfvpp5+qsrJSlZWVWrt2rTp37qwRI0aourpad999tyZMmKAffvhB7733nn7zm99o7969\nl/2Z5ebm6q233lJpaanOnz+vxYsX+/xzBxA6CHYAAkZ4eLhqa2u1e/du1dXVKTU1Vddff70k6ZZb\nbtGtt96qsLAwde3aVVOnTlV+fr7p/b/73e8UGxurXr166eabb1Z2drbS0tIUHx+v7OxsFRUVmfaf\nP3++IiMjNXToUN1zzz16//33PWMXQtjrr7+uadOmacCAAXI4HHrkkUfUqlUrbd26tcn8o6KiVFlZ\nqb1796qhoUE9evRQp06dPOONT+/u379fkydP1tq1a5WSkqLPPvtM3bp106RJkxQWFqa+fftq7Nix\nl121czgcmjJlitLT0xUdHa0HH3xQO3fu9OEnDiDUEOwABIz09HQtXbpUOTk5SkpK0vjx4/X9999L\nMkLQqFGj1LlzZ7Vt21azZs1qctoxKSnJ8+eYmBjT6+joaFVVVXleJyQkKCYmxvO6a9eunmNd7PDh\nw3rppZeUkJDg+Tp27Ngl9x0+fLiefPJJTZ8+XUlJSZo2bZoqKysv+b2Wl5dr9OjRWrBggQYNGuQ5\nVmFhoelYa9as0YkTJy77M7s4OMbExJi+RwAtD8EOQEAZP368CgoKdPjwYTkcDv3+97+XZNwupFev\nXjpw4IDKy8u1YMECNTQ0eP25ja9yPX36tGpqajyvDx8+rOTk5CbvS01N1axZs3T69GnPV1VVlcaN\nG3fJ4zz11FPasWOH9uzZo/379+uPf/xjk30aGhr08MMPa8SIEfr1r39tOtawYcNMx6qsrNSrr77q\n9fcJoGUj2AEIGPv371deXp5qa2vVqlUrRUdHKzw8XJJUVVWluLg4tW7dWvv27fPq9iEXn/q81FWu\nc+fOVV1dnQoKCvT555/rgQce8Ox7Yf/HH39cK1as0LZt2+R2u1VdXa3PP//8kitjO3bsUGFhoerq\n6tS6dWvT/C8+/qxZs1RTU6OlS5ea3j9q1Cjt379fq1evVl1dnerq6rR9+3bt27fPq+8RAAh2AAJG\nbW2tnn/+eSUmJqpz5846efKkFi5cKElavHix1qxZo/j4eE2dOlUPPfSQaRXuUvedazx+8etOnTp5\nrrCdOHGi/vznPysjI6PJvv369dPKlSv15JNPqn379urevbvnCtbGKioqNHXqVLVv315paWm67rrr\n9Nvf/rbJZ7733nueU64XrozNzc1VbGysvvjiC7333ntKSUlR586d9fzzz1/yQg1vvkcALY/DzT/3\nALQwTqdTEydO1NGjR+2eCgD4FSt2AAAAIYJgB6BF4pQlgFDEqVgAAIAQwYodAABAiCDYAQAAhAiC\nHQAAQIgg2AEAAIQIgh0AAECI+H/3Q2KUDc+3ZgAAAABJRU5ErkJggg==\n", + "text": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## `range()` vs. `numpy.arange()`" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "from numpy import arange as np_arange\n", + "\n", + "n = 1000000\n", + "\n", + "def loop_range(n):\n", + " for i in range(n):\n", + " pass\n", + " return\n", + "\n", + "def loop_arange(n):\n", + " for i in np_arange(n):\n", + " pass\n", + " return\n", + "\n", + "%timeit(loop_range(n))\n", + "%timeit(loop_arange(n))" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "10 loops, best of 3: 50.9 ms per loop\n", + "10 loops, best of 3: 183 ms per loop" + ] + }, + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "\n" + ] + } + ], + "prompt_number": 36 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "funcs = ['loop_range', 'loop_arange']\n", + "orders_n = [10**n for n in range(1, 6)]\n", + "times_n = {f:[] for f in funcs}\n", + "\n", + "for n in orders_n:\n", + " for f in funcs:\n", + " times_n[f].append(min(timeit.Timer('%s(n)' %f, \n", + " 'from __main__ import %s, n' %f)\n", + " .repeat(repeat=3, number=1000)))" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 38 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "import matplotlib.pyplot as plt\n", + "\n", + "labels = [('loop_range', 'in-built range()'), \n", + " ('loop_arange', 'numpy.arange()')]\n", + "\n", + "matplotlib.rcParams.update({'font.size': 12})\n", + "\n", + "fig = plt.figure(figsize=(10,8))\n", + "for lb in labels:\n", + " plt.plot(orders_n, times_n[lb[0]], \n", + " alpha=0.5, label=lb[1], marker='o', lw=3)\n", + "plt.xlabel('sample size n')\n", + "plt.ylabel('time per computation in milliseconds [ms]')\n", + "plt.legend(loc=2)\n", + "plt.grid()\n", + "plt.xscale('log')\n", + "plt.yscale('log')\n", + "plt.title('Performance of explicit for-loops vs. list comprehensions')\n", + "\n", + "max_perf = max( a/r for r,a in zip(times_n['loop_range'],\n", + " times_n['loop_arange']) )\n", + "min_perf = min( a/r for r,a in zip(times_n['loop_range'],\n", + " times_n['loop_arange']) )\n", + "\n", + "ftext = 'the in-built range() is {:.2f}x to '\\\n", + " '{:.2f}x faster than numpy.arange()'\\\n", + " .format(min_perf, max_perf)\n", + "plt.figtext(.14,.75, ftext, fontsize=11, ha='left')\n", + "\n", + "plt.show()" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "display_data", + "png": "iVBORw0KGgoAAAANSUhEUgAAAnYAAAIECAYAAACUvmMzAAAABHNCSVQICAgIfAhkiAAAAAlwSFlz\nAAALEgAACxIB0t1+/AAAIABJREFUeJzs3XdYFMf/B/D3Hr0KgihFinSQJs0uCPaORkENosRY0kws\n8WssqJFoDPYWKyoaoyb2XjiU2LBGUUEQFQvFAkgXmN8f/Fg5OOAODxHyeT3PPc/t7OyUvdm7udnZ\nXY4xxkAIIYQQQho8QX0XgBBCCCGEyAZ17AghhBBCGgnq2BFCCCGENBLUsSOEEEIIaSSoY0cIIYQQ\n0khQx44QQgghpJGgjh35KIqKijBmzBjo6upCIBDg3Llz9V2kBmnPnj0wNzeHvLw8xowZU69lCQkJ\ngaWlJb8cHh4OBQUFibeXNn55ycnJ8PHxgbq6OuTk5GqVRnVMTU2xYMECmafbUHl5eWHs2LH8clBQ\nELp161aPJWpYPqX2JBAIsHPnzvouBqlD1LEjvKCgIAgEAggEAigoKMDU1BQTJkzA69evPzjtv/76\nC3/88QcOHz6MlJQUtGvXTgYl/m8pLi7GmDFj4O/vj+TkZCxfvry+iwSO4/j3/v7+eP78ucTbVowf\nEREBgUCyr6TQ0FC8fPkSt27dwosXLyQvsIQ4jhOp239dxf2xcuVK7N27V+Lt5eXlsW3btrooWoPw\nKbWnlJQUDB48uL6LQeqQfH0XgHxaOnfujN27d6OoqAhXr17F2LFjkZycjMOHD9cqvcLCQigqKuLB\ngwcwNDRE27ZtP6h8Zen9Fz1//hw5OTno1asX9PX167s4AIDy9zdXVlaGsrKyxNtKG7+8Bw8ewN3d\nHebm5rXavkxRURHk5elrUFoaGhpSxec4Do3xXvgNsf3o6enVdxFIHaMROyJCQUEBenp6MDAwQP/+\n/fHdd9/h+PHjKCgoAADs2rULzs7OUFFRgZmZGSZPnozc3Fx+ey8vL3zxxReYNWsWDAwMYGJiAm9v\nb8yePRsPHz6EQCBAq1atAADv3r3D9OnTYWRkBCUlJdjb2+OPP/4QKY9AIMDKlSsxfPhwaGlpITAw\nkD+FJxQK4eDgAFVVVXTt2hUpKSmIjIyEs7Mz1NXV0a1bN5ERoaSkJPj5+cHQ0BBqampwdHRERESE\nSH5lp5zmz58PfX196OjoYNSoUcjJyRGJ9+eff8LV1RUqKirQ1dVF7969kZGRwa9fuXIlbGxsoKKi\nAisrK4SGhqK4uLjafX/p0iV07twZqqqqaNq0KUaMGIH09HQApactTUxMAJR2vms6nV1d/rt374aS\nkhJiYmL4+Nu2bYOqqiru3LkD4P2ptqVLl/L7a+jQoXjz5k2VeYo7tXrt2jX07NkTTZo0gYaGBjw9\nPXHlypVK8YVCIQIDAwGAHzWu6lSzQCDA2bNnsXnzZpF4L168gL+/P7S1taGqqgpvb29cu3aN304o\nFEIgEODo0aPo2LEjVFRUsGnTpirrU97bt28xbtw46OnpQVlZGe7u7jh16pRInLi4OPTp0wcaGhrQ\n0NBA//79kZiYWGn/nDlzBvb29lBRUUHbtm1x69YtPk5WVhZGjx4NfX19KCsrw9jYGJMnT66yXB06\ndMC4ceMqhdva2mL27NkAgNjYWPTo0QPa2tpQV1eHnZ1dpXYvrYqnYqvLw9TUFMXFxRg9ejQEAkGN\np85Xr14NOzs7KCsro3nz5hgyZAi/rqbP4dGjRxAIBPjjjz/Qo0cPqKmpwc7ODtHR0Xjy5Al69uwJ\ndXV12NvbIzo6mt+urG0cPnwYHh4eUFFRgYODAyIjIyvFEdd+JDneCwoK8N1330FHRwctWrTADz/8\nUClOTemYmppizpw51aYTHR2NDh06QFNTE5qamnB2dsbJkyf59RVPxUp63Jw+fRqdO3eGmpoa7O3t\ncfz4cZGyh4aGwtzcHMrKytDT00PPnj2Rn59f7WdN6ggj5P+NGjWKdevWTSQsLCyMcRzHsrOz2ZYt\nW5i2tjaLiIhgSUlJ7Ny5c8zR0ZF9/vnnfPwuXbowDQ0NNmHCBHbv3j12584d9vr1azZlyhRmZmbG\nUlNT2cuXLxljjE2ZMoXp6OiwvXv3sgcPHrDQ0FAmEAjYmTNn+PQ4jmM6Ojps9erV7OHDh+zBgwds\ny5YtTCAQMG9vb3blyhV2/fp1ZmlpyTp27Mg6d+7MLl++zG7evMlsbGzYsGHD+LRu377NVq9ezf79\n91/28OFDtnLlSiYvL88iIyNFyq+lpcV++OEHFhcXx06ePMmaNm3KZs2axcfZvHkzU1BQYD///DNf\nx1WrVvH1mjNnDjMxMWH79+9njx49YkePHmXGxsYiaVT04sULpqGhwUaMGMHu3LnDoqOjmaOjI+vc\nuTNjjLG8vDwWExPDOI5jhw4dYqmpqaywsFBsWpLkP3bsWGZubs6ysrJYXFwc09DQYGvXrhVpC5qa\nmmzAgAHszp07TCgUMktLSzZo0CCRfCwsLPjlLVu2MHl5eX75zp07TFVVlQ0fPpxdu3aNJSYmst27\nd7OLFy9Wil9YWMhWr17NOI5jqampLDU1lWVlZYmtX0pKCmvfvj0bOXIkH6+kpIR5eHgwFxcX9s8/\n/7Dbt2+zYcOGMW1tbf5ziYyMZBzHMRsbG3b48GH26NEj9uzZM7F5mJqasgULFvDLQ4YMYWZmZuzk\nyZPs/v377LvvvmOKiors/v37jDHGcnNzmbGxMfP19WXXr19n165dY97e3szCwoL/nMraraurKzt3\n7hz7999/Wd++fZmhoSHLy8tjjDH2zTffMCcnJ3blyhWWnJzMLly4wDZu3Ci2jIwxtn79eqatrc0K\nCgr4sMuXLzOO49iDBw8YY4w5ODiwESNGsHv37rGkpCR27Ngxdvjw4SrTFMfLy4uNHTuWXw4KChL5\nrqguj/T0dCYvL89WrFjBf7ZVmT17NlNXV2erV69mDx48YDdv3mS//PILv76mzyEpKYlxHMfMzc3Z\ngQMHWHx8PBs0aBAzNDRkXl5ebP/+/Sw+Pp4NGTKEtWzZkr17944x9r5tWFpasiNHjrD79++z4OBg\npqamxl68eCESp3z7efr0qUTHm4mJCdPW1maLFi1iCQkJbPfu3UxBQYFt2rSJjyOLdN69e8e0tbXZ\n5MmTWUJCAktISGD79+9n58+f59PgOI7t2LGDMcakOm6cnJzYiRMnWEJCAhs9ejTT1NRkb968YYwx\n9tdffzFNTU12+PBhlpyczG7evMmWL1/Ot2vycVHHjvBGjRrFfH19+eXY2FjWqlUr1q5dO8ZY6ZfK\n77//LrJNVFQU4ziOZWRkMMZKO0bW1taV0q7YCcjJyWFKSkoinQnGGBs0aBDr2rUrv8xxHPviiy9E\n4mzZsoVxHMdu3brFhy1evJhxHMeuX7/Ohy1dupTp6upWW+cBAwaI/GB16dKFOTs7i8SZMGECvw8Y\nY6xly5bsm2++EZteTk4OU1VVZSdOnBAJ37p1K9PS0qqyHDNnzhT5oWGMsVu3bjGO49i5c+cYY+9/\ntP75558q05E0/9zcXGZvb8+GDh3KnJ2dmZ+fn0j8UaNGMQ0NDZHO1cmTJxnHcSwxMZExVnPHbuTI\nkZX2ZXkV42/fvp1xHFdl/PIqdjROnz7NOI5j9+7d48MKCgqYvr4+mzdvHmPs/Q9UREREjemX79g9\nePCAcRzHjh07JhKnTZs2bMyYMYwxxjZu3MhUVVXZq1ev+PWpqalMRUWFbdu2ja8vx3Hs7NmzfJw3\nb94wdXV1tnnzZsZYaXsMCgqSaB+Uba+iosL27NnDh3311Vesffv2/HKTJk1YeHi4xGmKU3F/V/yu\nqCkPeXl5tnXr1mrzyM7OZsrKyiwsLEzsekk+h7JjZPny5fz6sj9ES5Ys4cNu3LjBOI5jsbGxjLH3\nbaPsc2CMsaKiImZiYsJ3rMS1H0mPNxMTEzZgwACROL169WIBAQEyTef169eM4zgmFAor78D/V75j\nJ81xs2/fPj5Oamoq4ziOnTx5kjHG2JIlS5iVlZXI9xepP3QqlogQCoXQ0NCAqqoqHBwcYGFhgR07\ndiA9PR1PnjzB999/z59q0tDQQO/evcFxHBISEvg0XF1da8wnISEBhYWF6Ny5s0h4586dERsbKxLm\n4eFRaXuO4+Dg4MAvN2/eHADg6OgoEvbq1St+bk9ubi6mT5+O1q1bQ0dHBxoaGjh69CiePHkikq6T\nk5NIXvr6+khNTQUApKWl4enTp+jevbvYesXGxiIvLw9+fn4i+2n8+PHIysrCq1evqtyubdu2IvN1\nHB0d0aRJE9y9e1fsNh+Sv4qKCv7880/89ddfePnypdhTknZ2diJzqdq3bw8AEpfn2rVr8PHxkbjs\nHyI2NhY6OjqwsbHhwxQVFeHp6Vlte+rVq5fIfhKnrL7VtdXY2FjY29ujadOm/Ho9PT1YW1tX2l/l\nLxzS0tKCra0tn87EiROxd+9eODg4YNKkSTh+/Hi1c9O0tLTQv39/bN++HUDp9IZdu3bxp7UBYMqU\nKfjiiy/g7e2NuXPn4saNG1WmV1uyyCM2NhYFBQVVHluSfA5lyh/DVX03AKXHc3nlPxs5OTl4eHhU\n234kPd44joOzs7NIOuW/V2SVjra2Nr744gv06NEDvXv3xqJFixAfH4+qSHPclM9XT08PcnJyfL7D\nhg3Du3fvYGJigtGjRyMiIgLZ2dlV5kvqVsOa9UnqXNu2bbF161bIy8vDwMCA72iUHcArVqyAt7d3\npe0MDQ0BlH7xqKmpybRM4tITCAQiV5mVvS8/f6csjDEGjuMwdepUHDx4EEuXLoW1tTVUVVUxefJk\nZGZmiqRd8eIMjuNQUlIiUVnL4u3duxdWVlaV1mtra4vdTlaTy6XJ//z58+A4DpmZmUhLS4OWlpZI\n3A8tz6cwYb7ssy+vfHvatGlTrecBVUxbXF0lqX/5ON27d8eTJ09w4sQJCIVCjBw5Eg4ODjhz5kyV\nVwwHBgZi0KBBePnyJaKjo5GTkwN/f39+/cyZMzFixAgcP34cZ8+eRWhoKKZNm4b58+dLU91qfYw8\nqiJuH5ef61n2GYkLq+m4rqn9SHO8Vfe9Iqt0AGD9+vX47rvvcPLkSZw6dQqzZs3CqlWr8OWXX1Zb\n1/LE1VvcRWtl+RoYGOD+/fuIjIzE2bNnMX/+fPz444+4fPkyjIyMJM6XyAaN2BERysrKaNWqFYyN\njUVGj5o3b46WLVvi/v37aNWqVaWXkpKSVPlYWFhASUkJUVFRIuFRUVEiI3GydP78eYwcORJDhgyB\ng4MDzMzMEBcXJ9VtCPT09GBkZIQTJ06IXW9vbw9lZWUkJiaK3U9V/Tjb29vj0qVLePfuHR9269Yt\nZGZmonXr1hKXT9L879y5g8mTJ2PTpk3w8fGBv78/CgsLRdK6d+8e3r59yy9fuHABQOlIniRcXV1x\n5swZiTt3ZT8ctekM2tvb49WrV7h37x4fVlBQgMuXL1e7/wwMDET2T1VpA6jUVs+dO8enbW9vj7t3\n74qMyKampiI+Pr5S/hcvXuTfZ2Rk4P79+yL7VFtbG/7+/li3bh2OHDmCqKgokXpV1L17dzRt2hS7\ndu3Ctm3b0K9fPzRp0kQkjpmZGSZMmIA9e/Zg7ty5WLt2bZXpSaricVNdHoqKijVePFR2wUR1xxYg\n/nOQ1XdG+c+mqKgIV65cqba91/Z4r6t0yqf3/fff4+jRowgODsb69eurjFeb40YcRUVF9OjRA4sW\nLcLt27eRm5uLAwcOSJUGkY0GN2J35coVTJo0CQoKCjA0NMS2bdsa3OXmDdWCBQsQHBwMbW1t9O/f\nHwoKCrh37x6OHz+OdevWASj9UZbkh1lVVRXffvstZs2ahWbNmsHR0RF79+7FwYMHcfr06Topv7W1\nNfbv3w8/Pz+oqalhyZIlePHiBVq0aMHHkaT8c+bMwYQJE9C8eXMMHjwYJSUliIyMREBAAHR0dDBj\nxgzMmDEDHMfBx8cHRUVFuH37Nm7evImFCxeKTfPrr7/G8uXLERQUhBkzZuDNmzeYOHEiOnfujA4d\nOkhcR3V19Rrzz8/PR0BAAAYNGoTAwED069cPTk5OmDZtGpYtW8anxXEcAgMD8fPPP+PVq1f46quv\nMGDAgCo7QBVNmzYNnp6eGDFiBCZPngwtLS1cv34dLVu2FHvbGzMzMwDAgQMH0KFDB6iqqlY5+lvx\nc/Lx8YGHhweGDx+O1atXQ1NTE/Pnz0dhYSEmTJgg8f4rn34Zc3NzfPbZZ5g4cSJ+//13GBsbY+3a\ntbh79y527doFABgxYgTmz5+PYcOGYfHixSgpKcGUKVNgZGSEYcOG8WlxHIcff/wRYWFh0NLSwk8/\n/QRNTU0MHz4cAPDTTz/Bzc0NdnZ2EAgEiIiIgIaGBoyNjassq7y8PIYPH441a9bg4cOH+Ouvv/h1\nOTk5mDZtGoYMGQJTU1NkZGTg+PHjfCcJKB3x4zgOW7durXZ/VDwuypazs7Px448/VpuHmZkZzp49\ni549e0JBQQG6urqV8lBXV8fkyZMREhICFRUV+Pr6Ii8vD8eOHcP06dMl+hw+1KJFi9CiRQuYmppi\nyZIlePXqFSZOnFhlfEmOt/L7qq7TSUhIwIYNG9C/f38YGRnh+fPnOH/+fJXTY2R13GzatAmMMbi7\nu0NLSwtnzpzB27dvJf4TSGSrwY3YGRsbIzIyElFRUTA1NaV/BDJU0000R44cid27d+Pw4cPw9PSE\nh4cH5s6dKzLUXlUa4sIXLFiAsWPHYtKkSXBwcMDOnTuxY8cOsad6xaUnbdjSpUv526/4+vqiZcuW\nGDJkSKVTuhXTqRgWHByM8PBw7N27Fy4uLujSpQtOnDjB/8GYOXMmlixZgg0bNsDZ2RmdOnXC8uXL\n+Y6LOHp6ejh58iSePn0Kd3d39OvXj+/s1lTHimrK//vvv0deXh7fGdfW1sbOnTuxZs0aHDt2jE/H\nw8MDHTt2RLdu3dCrVy84OTlh8+bNNe6rMq1bt4ZQKER6ejq6dOkCFxcXLF26VOSPWPn47u7u+O67\n7zBu3Dg0b94c33zzTZV1FJf3/v37YWNjgz59+sDDwwNpaWk4deqUyLw3SUdnK8bbuHEjevTogZEj\nR8LZ2RkXL17E4cOH+dNmysrKOHnyJJSUlNC5c2d4eXlBQ0MDx48fF6mvQCBAaGgoxo0bB3d3d6Sl\npeHIkSP8/fxUVFQwe/ZsuLm5wd3dHXfu3MGxY8dqvG/cqFGjcP/+fWhpaaFXr158uLy8PDIyMhAc\nHAw7Ozv07NkT+vr6Ire7SE5ORnJyco37o6rjREFBocY8wsLCcO3aNZiamvLz28SZP38+FixYgBUr\nVsDBwQE9evQQma9X0+dQVjZx5Zck7LfffsOsWbPg4uKCixcv4sCBAyJ//MRtI8nxLsl3oizSUVdX\nR0JCAvz9/WFtbY0hQ4agQ4cOWLVqVaXtysjiuGnatCm2bNkCb29v2NnZYdmyZdiwYYNE3+VE9jhW\n35NgPsCcOXPg4uKCgQMH1ndRCGlUgoKC8OzZs0r3aiO1Fx4ejrFjx4qcbiefBqFQiK5du+Lp06cw\nMDCo7+IQ8kEa3IhdmcePH+PUqVPo169ffReFEEIIIeSTUG8du1WrVsHNzQ3KysoYPXq0yLrXr19j\n0KBBUFdXh6mpaaWnEWRlZSEwMBBbt26tkweAE/Jf9yk927IxoX366aLPhjQW9XYqdt++fRAIBDhx\n4gTy8vKwZcsWfl1AQACA0gmZN27cQJ8+fXDhwgXY2dmhqKgI/fv3x5QpU9C1a9f6KDohhBBCyKep\nzm+BXIOZM2eK3Gk9OzubKSoq8o/DYYyxwMBANn36dMYYY9u2bWM6OjrMy8uLeXl5sT///FNsugYG\nBgwAvehFL3rRi170otcn/zI3N5dJv6re59ixCgOG8fHxkJeXh4WFBR/m5OTE3wX7888/x8uXLxEZ\nGYnIyEgMHTpUbLrPnz/nL9H/lF9z5sxpEHnUNg1ptpMkbk1xqltf23Wf0quuyymr9GuTjqzbiiTx\natMmqK3INg/6bvk0XvTdIl3cumgviYmJMulX1XvHruK8huzsbGhqaoqEaWhoiNwotTHx8vJqEHnU\nNg1ptpMkbk1xqltf3bpHjx7VmPenoK7bi6zSr006sm4rksSrTXuhtiLbPOi75dNA3y3Sxa2r9iIL\n9X67k5kzZ+LZs2f8HLsbN26gY8eOyMnJ4eP89ttvOHfuHA4ePChxup/C44xIwxEUFITw8PD6LgZp\nAKitEGlQeyGSklW/5ZMbsbOyskJRUZHIQ+Vv3bol9eNNACAkJARCofBDi0j+A4KCguq7CKSBoLZC\npEHthdREKBQiJCREZunV24hdcXEx3r17h7lz5+LZs2fYsGED5OXlIScnh4CAAHAch40bN+L69evo\n27cvLl68CFtbW4nTpxE7QgghhDQUDX7Ebv78+VBVVcWiRYsQEREBFRUVLFiwAACwZs0a5OXlQU9P\nDyNHjsS6deuk6tQRIi0a2SWSorZCpEHthXxs8jVHqRshISFVDj1qa2tj3759H7dAhBBCCCENXL1f\nPFFXqhvSbNq0Kd68efORS0SIeNra2nj9+nV9F4MQQkg9ktWp2HobsfsYQkJC4OXlVenS4jdv3tD8\nO/LJoEcZEULIf5dQKJTpKfv/5IgdXVhBPiXUHhsWoVD4Ue4RRxoHai9EUg3+4glCCCGEECJbNGJH\nSD2j9kgIIYRG7AghhBBCiIhG3bFrjE+eCAoKQrdu3T44HaFQCIFAgOfPn39wWgKBADt37uSXTU1N\n+XsSNlbPnz+Hjo4Onj17BgBISkqCjo4O0tPT67lkpK41tu8UUreovZCayPrJE42+Y9fYJq2uXLkS\ne/fure9iiEhJScHgwYP5ZY7jRK70tLCwwNy5c+ujaHVmzpw5GDZsGAwNDQEAZmZmGDRoEObNm1fP\nJSOEENKQeHl5ybRj16hvd1JbcXGPcfp0It69E0BBoQS+vuawtjb5JNLU0ND4oHLUBT09vWrX1/Z2\nHu/evYOCgkKttq1Lr1+/RkREBC5cuCASPmbMGPTo0QO//PIL1NXV66l0pK41tj+LpG5ReyEfW6Me\nsauNuLjHCA9PQHp6V2RkeCE9vSvCwxMQF/f4k0iz4qnYsuX169fDxMQETZo0wYABA5CWliZRetev\nX4eHhwdUVFTg4OCAyMhIfl1Vp2vl5eWxdetWflkgEGDHjh1i0/fy8kJiYiLmzp0LgUAAgUCAJ0+e\nVFu3lStXwtTUFCoqKigoKMCpU6fg5eUFHR0daGlpwcvLCzExMSLbCgQCrF27Fp9//jk0NTXRsmVL\nLFy4UCTOq1ev8Nlnn0FdXR36+vqYN2+e2FPbK1euhI2NDVRUVGBlZYXQ0FAUFxfz6/fs2YPmzZvD\nxcVFZLt27dpBTU2NnppCCCGk3tCIXQWnTydCSckHotMifPDvv2fh7l67UbsrVxKRm+sjEubl5YMz\nZ85KPWpX8TQnAMTExEBPTw/Hjh1DVlYWhg8fjilTpmDbtm01pvfDDz9g2bJlMDc3x+LFi9GvXz8k\nJCSgRYsWUpWhqlG5ffv2wdXVFUOGDMGUKVMAALq6ulWmfeXKFWhqauLQoUMQCARQUFBATk4Ovv76\nazg5OaGoqAhLlixBz5498eDBAzRt2pTfdu7cuViwYAHmzZuHY8eO4euvv4aHhwe6du0KABg9ejTi\n4+Nx5MgRNGvWDL/99hsOHDgAd3d3Po2QkBCEh4dj+fLlcHZ2xt27dzF+/Hjk5+fzp1mjoqLg6ekp\ndr94enri7Nmz+Pzzz6usI2nY6L5kRBrUXsjHRiN2Fbx7J36XFBfXfleVlIjftrBQ+jQZY5Uuh1ZW\nVkZ4eDjs7OzQtm1bjB8/HqdPn5Yovf/973/o3bs3rK2t8fvvv0NXVxdr1qyRulxV0dbWhpycHNTV\n1aGnpwc9PT0IBFXXW05ODtu3b4eDgwPs7e0hEAgwcOBADBkyBJaWlrC1tcXvv/8OxhiOHz8usq2/\nvz+Cg4NhZmaGiRMnwsbGht8PDx48wOHDh7F27Vp06dIFdnZ2WL9+vcip7dzcXCxevBjr16/HgAED\nYGJigl69emH+/PlYuXIlHy8+Ph7GxsZiy29iYoL4+PgP2WWEEEJIrTXqEbuqHilWHQWFErHhcnLi\nwyUhEIjfVlGx9mmWZ2NjIzIXTV9fH6mpqfyyuro6P6LWuXNnHDlyhF/Xrl07/r2cnBw8PDwQGxsr\nk3LVhq2tLVRVVUXCkpKSMHv2bFy6dAlpaWkoKSlBbm5upVO6zs7OIssGBgb8Kem7d+8CANq2bcuv\nl5eXh5ubG7KzswEAsbGxyMvLg5+fn8gIZHFxMQoKCvDq1Svo6OggKyuryrmOmpqayMjIqGXtSUNA\noy9EGtReSE1k/UixRt+xk5avrznCw8/Ay+v9qdOCgjMICrKAtXXtyhEXV5qmkpJomj4+FrVLsIKK\nFxhUvMnhv//+y79XUVGpNi3GGN+pKRtZK59WcXExSkpk0yEVp2KnDgD69u0LPT09rFmzBi1btoSC\nggI6duyIwsJCkXiKioqVtq1Y1oqnjMvXrSzu3r17YWVlVSktbW1tAICWlhbevn0rtvyZmZl8PEII\nIaQmZQNQsrp7BJ2KrcDa2gRBQRbQ0zsLLS0h9PTO/n+nrvZXxdZFmuXVdNVpq1at+Je+vr7IuosX\nL/Lvi4qKcOXKFdjZ2QF4f7Vr2b3aAODmzZtS3xlbUVFR5OIDabx69Qr37t3D9OnT0a1bN9jY2EBJ\nSUmii0PK75eyOpW/krWoqAjXrl3jl+3t7aGsrIzExESRfVb2KuvoWlpa4tGjR2LzfPz4sdhOIWk8\n6L5kRBrUXsjH1qhH7GrL2tpEZp2uukyzzIc8gmTRokVo0aIFTE1NsWTJErx69QoTJ04EUNqBMTEx\nQUhICJYuXYr09HTMmDGjxo5kxfKYmZkhOjoaycnJUFFRgY6OjsS3QNHW1kazZs2wfv16tGrVCi9f\nvsS0adNqHHksK0dZWSwtLdGvXz989dVX/FzCsLAwZGVl8WVRV1fHjBkz+Dr6+PigqKgIt2/fxs2b\nN/mrbLvUs/HVAAAgAElEQVR06SL2BsyMMVy5cgW//vqrRHUjhBBCZI1G7BqYilekirtCtSxckrR+\n++03zJo1Cy4uLrh48SIOHDjAXxErJyeHP//8E2lpaXBxccE333yD0NDQai9+EJf33LlzkZGRAWtr\nazRv3hzJyckS1Q0oPR28Z88eJCYmwtHREWPGjMH3339faeRRkvS2bNmC1q1bo1evXujatSuMjIzQ\nvXt3KCsr83FmzpyJJUuWYMOGDXB2dkanTp2wfPlymJmZ8XEGDx6MtLQ0XL9+XSS/CxcuIDs7G35+\nfjWWjTRcNGeKSIPaC/nYONZInz5e3cN06aHrBCidL2hjY4OBAwdi8eLFUm375ZdfQk5ODmvXruXD\ngoODoaKiglWrVkmVFrVHQgghsvotoI4d+c84f/48UlNT4eLigrdv32Lp0qXYtWsXrl+/Dnt7e6nS\nevHiBVq3bo1///0XhoaGSEpKgru7O+7du4dmzZpJlRa1x4aF7ktGpEHthUhKVr8FjXqOXW1ud0Ia\nr+LiYixYsAAJCQlQUFDgn7QhbacOKL2lzKtXr/hlMzMzvHz5UpbFJYQQ8h8g69ud0IgdIfWM2iMh\nhBBZ/RbQxROEEEIIIY0EdewIIUQKdF8yIg1qL+Rjo44dIYQQQkgjQXPsCKln1B4JIYTQHDtCCCGE\nECKCOnaEECIFmjNFpEHthXxs1LEjRAYOHDgABwcHfjkiIgJt27atxxIRQgj5L2rUHbuQkBD6t0Tq\nXElJCaZNm4ZZs2bxYcOHD8ebN2/w119/1WPJSF2gG54TaVB7ITURCoUICQmRWXp08QRpMAoLC6Go\nqFjfxajkyJEjGDVqFFJSUiAv//5hLqGhoThx4gSioqKq3Z7aIyGEELp4og7FJcRh9Z+rsWzXMqz+\nczXiEuI+iTS9vLwwduxYzJ8/H/r6+tDR0cGoUaOQk5MDAAgKCkK3bt1EtomIiIBA8P5jDgkJgaWl\nJfbs2QMLCwuoqalh8ODByM7Oxp49e2BtbQ1NTU189tlnyMrK4rcrS3vp0qUwNDSEmpoahg4dijdv\n3gAo/cchLy+Pp0+fiuS/bds2aGlpIS8vTyT8+vXr6NWrF5o3bw4NDQ14eHjgxIkTInFMTU0xa9Ys\nTJw4Ebq6uujSpQsAYPny5XBxcYGGhgb09fUREBCAlJQUfjuhUAiBQIDTp0+jc+fOUFNTg729PY4f\nPy6S/o0bN9C2bVuoqKjAxsYGf//9N0xNTbFgwQI+TnZ2Nr777jsYGRlBTU0Nbdq0wb59+yrt4759\n+4p06gBg4MCBOH/+PJKTkyt+lKQBo7MARBrUXsjHRh27CuIS4hAeGY705unIaJGB9ObpCI8M/6DO\nnSzT3Lt3LzIyMhAVFYVdu3bh8OHDWLRoEb+e47ga03jx4gW2bduG/fv349ixYzh//jz8/PwQHh6O\nvXv38mGhoaEi2125cgVRUVE4efIkjh49ips3byI4OBhAaafT0tISmzdvFtlmw4YNGDFiBFRUVETC\n3759i4CAAAiFQty4cQM9evRA//798eDBA5F4K1asQIsWLXDp0iVs2bKFr2NYWBju3LmDffv24cmT\nJ/D3969UzylTpmDmzJn4999/4enpiWHDhiEjIwMAkJubi969e6N58+aIiYnB1q1bERYWhvT0dH4f\nMsbQr18/3L59G7t370ZsbCwmTJgAf39/nD17ls/n3Llz8PT0rJS/ra0tmjRpIhKXEEIIqUvyNUf5\nbzl97TSULJUgfCR8H6gA/LvrX7h3dK9VmleiryDXKBd49D7My9ILZ66fgbWFtVRpmZqaIiwsDABg\nZWWFYcOG4fTp05g3bx4ASDSMW1BQgK1bt6Jp06YAgKFDh2LdunVITU2Fjo4OAMDf3x9nzpwR2Y4x\nhu3bt0NDQwMAsHr1avTo0QMPHz5Eq1at8OWXX2L58uWYNWsWOI7D/fv38c8//2DVqlWVylA2+lZm\n/vz5OHToEPbs2YMZM2bw4R4eHpg9e7ZI3G+//ZZ/b2JiglWrVsHV1RUvXryAvr4+vy4kJATdu3cH\nACxcuBDh4eGIiYlBt27dsGPHDmRnZyMiIoKvz+bNm2Fra8tvHxUVhUuXLiE1NRWampoAgLFjx+Li\nxYtYuXIlunbtiuzsbLx48QLGxsaV6shxHIyNjSt1VknDRnOmiDSovZCPjUbsKnjH3okNL0ZxrdMs\nQYnY8MKSQqnS4TgOTk5OImH6+vpITU2VKh1DQ0O+UwcAzZs3R4sWLfhOXVlYWlqayHZ2dnZ8JwgA\n2rdvDwC4e/cuACAwMBBpaWn8KdWNGzfCzc2tUpkBID09HRMnToStrS20tbWhoaGB2NhYPHnyRKS+\nHh4elbYVCoXo0aMHjI2NoampiU6dOgEAHj9+LBLP2dmZf6+npwc5OTl+X929e7dSfaytraGlpcUv\nx8TEoLCwEIaGhtDQ0OBfO3bsQEJCAgAgMzMTAETSKU9TU5MfJSSEEELqGo3YVaDAKYgNl4NcrdMU\nVNF/VhRIfyFAxYsHOI5DSUlpx1EgEFQasXv3rnJHVUFBtI4cx4kNK0u3TE2jgTo6OhgyZAg2bNgA\nHx8fbNu2rdLp3DJBQUF4+vQpFi9eDDMzMygrK8Pf3x+FhaKdXTU1NZHlJ0+eoHfv3hg1ahRCQkKg\nq6uL5ORk+Pr6VtpW3IUWFetUnZKSEjRp0gRXr16ttK4s7bKO4Nu3b8WmkZmZKdJZJA2fUCikURgi\nMWov5GOjjl0Fvq6+CI8Mh5elFx9W8KAAQf5BUp82LRNnVDrHTslSSSRNH2+fDy2uCD09PVy6dEkk\n7Pr16zJL/969e3j79i0/OnXhwgUApSN5ZcaNGwdvb2+sW7cO+fn5CAgIEJvW+fPnsXjxYvTt2xcA\nkJOTg8TERJF7wYkTExOD/Px8LFu2DEpKSnyYtOzt7bFp0yZkZWXxp1nj4uJERtfc3NyQkZGBvLw8\n2Nvbi01HTU0N+vr6lUYLgdKOcHJyMqysrKQuHyGEEFIbdCq2AmsLawR5B0EvTQ9aKVrQS9NDkHft\nO3WyTJMxVu2oma+vL+7fv481a9YgMTERGzZswJ49e2pd7oo4jkNgYCBiY2Nx7tw5fPXVVxgwYABa\ntWrFx+nQoQOsra0xdepUBAQE8CNuPj4+InPnrK2tERERgTt37uDmzZsICAhASUmJSP3E1dXKygoc\nx+G3335DUlIS9u/fj/nz50tdlxEjRkBdXR2BgYG4ffs2Ll++jODgYKioqPAXT/j4+MDX1xd+fn44\ncOAAHj58iGvXrmHlypXYuHEjn1aXLl1w+fLlSnncu3cPmZmZ9G+9kaHPk0iD2gv52KhjJ4a1hTUm\nDp2ISf6TMHHoxA/q1MkyTY7jKl31Wj7M19cXP//8M0JDQ+Hs7AyhUIjZs2eLbFNTGtWFeXh4oGPH\njujWrRt69eoFJyenSlfBAsAXX3yBwsJCfPnll3zYw4cPRW5JsmXLFpSUlMDDwwN+fn7o3bs33N3d\nK5W1IgcHB6xcuRK///477O3tsWTJEixbtkxs+aujoqKCo0ePIjU1Fe7u7ggMDMSkSZOgrq4OZWVl\nPt7Bgwfh5+eH77//Hra2tujbty+OHTsGCwsLPs7IkSNx5MgRFBUVieSxb98+dOzYUeyFFYQQQkhd\noBsUE4kEBQXh2bNnOHXqVI1xp02bhjNnzuDatWsfoWSy8/jxY5iZmeHQoUPo06ePxNsxxmBnZ4eQ\nkBAMGzYMQOn8PBsbG4SGhmLIkCHVbk/tsWGhOVNEGtReiKToBsUSoEeKfVyZmZmIiYnBhg0b8P33\n39d3cWoUERGByMhIPHr0CFFRURg6dChMTU35W6RIiuM4LFq0SOTGxjt37uQvJiGEEEKqQo8UkxCN\n2MnW6NGj8ezZM5w8ebLKOF5eXrhy5QoCAgKwadOmj1i62lmxYgVWrFiBZ8+eoWnTpujYsSPCwsJg\nZGT0UctB7ZEQQoisfguoY0dIPaP2SAghhE7FEkJIPaDpHUQa1F7Ix0YdO0IIIYSQRoJOxRJSz6g9\nEkIIoVOxhBBCCCFExH/ykWLa2to13sCWkI9FW1u7votApED3JSPSoPZCPrb/ZMfu9evX9V0E8omh\nL19CCCGNwX9yjh0hhBBCyKeE5tgRQgghhBAR1LEjBHSvKSI5aitEGtReyMdGHTtCCCGEkEaC5tgR\nQgghhNQzmmNHCCGEEEJEUMeOENA8GCI5aitEGtReyMfWqDt2ISEhdFARQggh5JMlFAoREhIis/Ro\njh0hhBBCSD2JS4jD6Wun8bX/1zLpt/wnnzxBCCGEEFLf4hLiEB4ZDkEr2Z1AbdSnYgmRFJ2yJ5Ki\ntkKkQe2FVOfU1VPINMjElWdXZJYmjdgRQgghhHxkmfmZuPz8MpKbJss0XZpjRwghhBDykTDGEPM8\nBqcfnkZ0VDRyjXIBAFGjo2iOHSGEEEJIQ5Gek46DcQeRnFU6SteqVSvcvHsTxi7GMsuD5tgRApoH\nQyRHbYVIg9oLAYCikiIIHwmx7uo6vlMHADYWNpg3eB46FneUWV40YkcIIYQQUkeSM5NxMO4g0nPT\n+TABJ0An407oZNIJ8gJ5dHXpiq+GfSWT/GiOHSGEEEKIjBUUFeBM0hnEPIsBw/v+iJGmEfpZ9UNz\n9eYi8WXVb6ERO0IIIYQQGYp/FY/D8YeRVZDFhynKKcLHzAfuhu4QcHU3E47m2BECmgdDJEdthUiD\n2st/S05hDvbe3Yudt3eKdOosm1piovtEeBp51mmnDqARO0IIIYSQD8IYw63UWziRcAJ5RXl8uKqC\nKnpZ9EJrvdbgOO6jlIXm2BFCCCGE1NKbvDc4FH8ID988FAl3au6EHhY9oKqgKlE6NMeOEEIIIaSe\nlLASXHp6CZFJkXhX8o4P11LWQl+rvrBoalEv5aI5doSA5sEQyVFbIdKg9tI4pWSnYOP1jTiZeJLv\n1HHg0M6oHSa6T6y3Th1AI3aEEEIIIRJ5V/wOUY+jcCH5AkpYCR/eXK05+lv3h6GmYT2WrhSN2P2/\nkJAQvHv3fig1KCgIq1ev/qA0r127hpEjR0q93aNHj9CsWbMPyq9iGhXr9ymaOnUq/vzzTwDA6tWr\nsXjx4irjjh07Fv/884/Ead+8eROurq5wcXGBvb09xowZg7y89xNcvby8+PempqawtbWFi4sLXFxc\ncPLkSX7dwIED4ezsDBcXF3To0AExMTFS1LBUbT8LoVAIVVVVvlzt2rUTG+/Ro0d8HBcXF5iamkJH\nR4dfn5+fjwkTJsDKygqOjo4YN26c1GXZv38/7Ozs4Orqivj4eKm3j4qKwqlTp6TeTpzMzEz8+uuv\nImFeXl44cuSITNKvqHxbIbVz5MgRTJgwAQBw+/Zt9OvXr55LVHeovTQeSW+SsPbqWkQ/ieY7dfIC\nefiY+eBL1y8/iU4dAIA1UtJWjeM4lp2dzS8HBQWxVatWybpYEklKSmK6uroyTaNi/apTVFT0QXnX\nRmpqKrO1teWX8/PzWatWrVheXp5M0s/Ly2Pv3r1jjDFWUlLCBg8ezMLCwsTGNTU1ZbGxsWLXZWZm\n8u8PHDjAHBwcpC6LNJ9FeZGRkczNzU3q7SZNmsS++eYbfvmbb75hP/zwA7+cmpoqdZo9e/Zke/fu\nlXq7MnPmzGFTpkyp1bbFxcUiy+KOFy8vL3b48OFal++/oj6OdcYYc3V1ZUlJSfxyr1692KVLl+ql\nLITUJLcwlx24f4DNiZwj8tp8fTNLz0mXWT6y6pLRiB2Ar74qfYxH+/bt0aZNG2RmZgIA7ty5Ax8f\nH1hZWWHUqFF8/KysLHzxxRfw9PSEk5MTJk2ahJKSkkrpCoVCuLu7AygdRdHV1cXMmTPRpk0b2NjY\n1DjiNGXKFDg5OcHR0RHR0dGV0qy4XHGduPq5uLjw9StTVrapU6fC1dUVGzduxNmzZ/n94ejoyI+k\nAaX/QKdNm4ZOnTrB3Nwc//vf//h1d+/ehaenJxwcHPD555+jXbt2/MjJixcv8Nlnn8HT0xOOjo74\n5Zdf+O22b9+OgQMH8stKSkro3Lkz/v77b7H7pvyIzPr162FnZwcXFxc4OTkhLi6uUnxlZWXIy5fO\nPCgsLEReXh709PRE9mN5rIorkzQ1Nfn3GRkZfBr379+HsbExnjx5AgCYO3cuAgICKm1f8bPIyspC\namoqBg0axH/W27dvF5t3bRQWFmLHjh0YM2YMACA7Oxvbt2/H/Pnz+TjS1uH7779HdHQ0pk2bBh8f\nHwDAiBEj4O7uDkdHR/j5+SEjIwMAEBcXh3bt2sHZ2RkODg4ICwvDnTt38Pvvv2Pbtm1wcXHhR9uO\nHj2Kjh07ws3NDe3bt8fly5cBlH42jo6OGDNmDFxcXHD8+PFK+zQjIwMuLi7o2PH98xajoqLEttGw\nsDB4eHigTZs2aN++PW7dusWvEwgE+OWXX+Dh4QFzc3Ox7U8oFCIkJAQBAQHo06cPbG1t0bdvX34E\nuOJof/nloKAgjB8/Hj4+PjA1NcWkSZNw6tQpdOrUCWZmZlixYgW/nampKf73v//Bzc0NlpaWfBp7\n9uxB3759+XgFBQXQ19fH06dP+bCdO3eibdu2aNOmDdq0aYOzZ89WStfT0xPjx49HamoqunbtCjc3\nN7Ru3Ro//vgjH7e6emZmZmLw4MGwtbWFr68vAgMDMXXqVACl7W7q1Knw9PSEs7MzAgMDkZOTA6D0\nzIKCggJMTU35fIYNG4ZNmzZV2teNAc2xa7gYY7ibfherY1bj+ovrfLiSnBL6WfVDkHMQdFV167GE\nVZBJ9/AjyszMZO7u7kxdXb3KURXGajdil5OTwy+PGjWKderUiRUUFLDCwkJmb2/PTp06xRhjLDg4\nmG3fvp0xVjp64O/vzzZs2FApzfIjLElJSYzjOHbkyBHGGGM7duxgHTp0EFuWsrhleQiFQmZkZMQK\nCgoqjdqUX66YX8URu/L1E5ff7t27+bA3b97wIyMpKSnMyMiIZWRkMMZKR0P8/f0ZY6Wfh66uLktI\nSGCMMdamTRu2Y8cOxhhjV69eZXJycnydfX192blz5xhjjBUUFLCOHTvy+7RPnz7s4MGDIuVav349\nGzNmjNgye3l58ek2adKEpaSkMMYYKywsZLm5uWK3ef78OXNycmIaGhps8ODBIusiIyP596ampszB\nwYE5ODiwiRMn8vUuExwczIyNjZmBgQG7d+8eH759+3bWtm1bduLECWZtbc3evn0rthwVP4uhQ4ey\n2bNnM8YYe/HiBTMwMGB37typtF1kZCTT0NBgzs7OzNPTk23dulVs+uXt2bOHubi48Ms3b95k5ubm\nbOrUqczNzY15eXmx6OhoqetQfv8zxtjLly/59z/99BObPn06Y4yxb7/9lv3yyy/8urJ9GRISwqZO\nncqHJyQksHbt2rGsrCzGGGN37txhxsbGfL3l5OSqHNF59OhRpRG7Ll26VNlG09Pf/8M+deoUa9u2\nLb/McRxbvXo1Y4yxf/75hxkaGlbKLzIyks2ZM4dZWlryI7jdu3fnvwOCgoL4NCoul32vlLVTPT09\nvo0/e/aMqaur823D1NSUBQcHM8ZKR1UNDAzY7du3WVFRETMxMeFHvLZt28b8/PxEyvjq1Sv+/f37\n95mRkRG/bGpqyr766it+OT8/nx9BLiwsZF27dmXHjx9njLFq6/nDDz+wsWPHMsYYe/36NTMzM+M/\n0/nz57Off/6Zz2PatGnsp59+Yowx9uuvv4qMGDPGWFxcHGvVqlWlfd0YlP9uIQ1HZn4m++P2H5VG\n6Xbd3sWy8rPqJE9Zdcka3MUTqqqqOHr0KKZOnVqn96njOA4DBw6EoqIiAKBNmzZ4+LD0HjUHDx5E\nTEwMwsLCAAB5eXkwNjauMU11dXX07t0bAODp6YnJkydXGVdRUZGfL9elSxeoqKiIHYmSFWVlZXz2\n2Wf8clpaGkaPHo2EhATIy8vj9evXiIuLg4eHBwDwcTU1NWFra4vExEQ0a9YMsbGxGD58OADA1dUV\njo6OAICcnBwIhUK8fPmSzyM7Oxv379+Hr68vkpKSYGgoOj/B0NCQ3+fV6dq1KwIDA9GvXz/06dMH\nZmZmYuPp6+vj5s2byM3NxfDhw7Fw4UJMnz4dgOg8mOjoaBgaGqKwsBCTJk3C119/LTKKtnHjRgBA\nREQE/Pz8EBsbC47jMHLkSJw+fRqDBg1CdHQ01NXVayw7AJw5cwZLly4FALRo0QK9e/dGZGQk7O3t\nReK5urri2bNn0NDQwKNHj+Dr6wtDQ0N+1EyczZs386N1AFBcXIyHDx+iTZs2+PXXX3HlyhX069cP\nCQkJ0NDQkKoO5Y+/rVu3YufOnSgsLEROTg6sra0BlLbdadOmITc3F97e3vD29ha7/YkTJ5CYmIjO\nnTuLlDU9vfSh2ZaWlvD09KyxHGU4jhPbRs3NzXH16lWEhobizZs3EAgEleYI+vv7Ayg9Rp8/f47C\nwkL+ewAobStRUVHo2bMnP4Lr6emJxMTEastUVq6BAwdCQUEBCgoKsLa2Rp8+fQAABgYG0NbWxtOn\nT2FlZQUACA4OBlA6qtqnTx9ERkaidevWGDduHNatW4eFCxdi9erVCA0NFcknISEBM2fOxPPnz6Gg\noICUlBSkpaXxo7OBgYF83KKiIkyZMgUXL14EYwwpKSm4desWevToAQBV1lMoFGLVqlUAAG1tbZER\n94MHD+Lt27fYu3cvgNJRRWdnZwClZwhatWolUl4jIyM8fvxY7D5r6GiOXcPCGMO1F9dwKvEUCooL\n+HB1RXX0sewD22a29Vg6yTS4jp28vDx0dT/O0KeSkhL/Xk5ODkVFRfzygQMHRE4lfEh6CxYs4L8A\nly1bBhMTEwClDaz8nao5joO8vLzIad/8/HypylAVNTU1keUJEyZg4MCB2LdvHwDA2tpaJC9lZWWx\ndalKSUkJBAIBrl69Cjk5OYnKJOnNGv/++2/ExMTg7Nmz8Pb2xrp169CzZ88q46uqqsLf3x87duwQ\nu76sg6moqIgJEyZgwIABYuONHDkSX375Jd68eYOmTZuisLAQsbGx0NbWRkpKigQ1fK98PSt+7mU0\nNDT496amphg4cCD++eefKjt2z549w7lz50TqaWxsDHl5eb7z4uHhAV1dXTx48ABt2rSRqg5lZTx/\n/jzWrVuHixcvQkdHBzt37sSGDRsAAH5+fmjfvj1OnDiBhQsXYvPmzdi+fbvYz7Vnz57YunWr2Lwk\n7SSXJ66NFhYWYsiQIYiOjoazszOeP38OIyMjsduVtdOioiKRjl2Zisdz2fFR8Rgtf5GOuO2qO5aq\nahdjx45FmzZt0K9fP2RmZqJr164ieQQEBGDp0qXo378/GGNQVVUVOX7L788lS5YgIyMDV65cgaKi\nIsaNG8fH5TiuynqKK195a9eulbhTU3asV9X2CfkYXua+xKG4Q3icKfonw1XfFd3Mu0FZXrmKLT8t\nNMfu/2loaPDzgmrSv39//PLLL/yX98uXL/Ho0aNa5/3TTz/hxo0buHHjBrp06QKgdI7Kzp07AZT+\ncObn58PGxgatWrXCw4cPkZGRAcYY/vjjD4nykKZ+QOn8mbIO5qlTp5CQkCCyXtwPs6amJuzt7fky\nXb9+Hbdv3+bz79Spk8i8uuTkZKSmpgIo7aiUnyMEAE+fPq30z76i4uJiJCYmwt3dHT/++CO6d++O\nmzdvVoqXlJSEgoLSf1+FhYU4cOAAP/oIvJ8Hk5uby89BZIxh165dcHFxAVA66picnMxvc+jQIRgY\nGKBp06YASq/qdXd3x8mTJzF+/Hg8e/ZMbJkrfha+vr58RyglJQXHjh2r9ENdtq5sv79+/RonT57k\nyybO1q1b0bdvX2hra/Nhurq68Pb25q9IjY+PR1paGiwsLKSqQ3kZGRlo0qQJmjZtioKCAmzevJlf\nl5CQAD09PYwaNQqzZ8/mryJu0qSJyFzP7t274/jx47h79y4fJukVx5qamsjNzUVxcbFIuLg2mp+f\nj+LiYr4zt2bNGonyKE8oFFZKu6xTAgAWFhZ82V+8eFHjHKvq/ryEh4cDANLT03Hs2DF+xFNXVxe+\nvr4ICAjg522Wl5mZyf/x3LRpE9/2xcnMzIS+vj4UFRXx7NkzHDhwoMqyla+nl5cXtm3bBqC0DRw8\neJCP179/f4SFhfGdwLdv3+L+/fsASo/1iu3q6dOnMDY2bpSdOppj9+krLinGucfnsDZmrUinTkdF\nB0HOQehn3a/BdOqAeuzYrVq1Cm5ublBWVsbo0aNF1r1+/RqDBg2Curo6TE1Nq+y8yPJLYPLkyeja\ntavIxRNVpb9s2TLIycnxk9179eqF58+fiy1fxRE3Scuvo6ODmzdvwsnJCV9//TX++OMPyMvLw8DA\nAJMnT4arqys6dOgAAwODKvMo/15c/aory8KFCzFlyhS4uLhgz549cHJykqjs27Ztw7Jly+Do6Iiw\nsDA4ODigSZMmAIAdO3bg7t27cHR0hKOjI/z9/fmyeHt749KlSyJpXbhwodrTjEBpx2706NFwdHSE\ns7MzUlJSxN6+48KFC3B3d4ezszPc3NxgYGDAT6i/evUqf0o2JSUF3t7ecHJygoODAxISEvgf/5yc\nHAwdOhSOjo5wcXHB6tWr+R/B/fv349y5c1i2bBns7OwwZ84cBAQEiL2opvxnkZWVhRUrVuDWrVtw\ncnJC9+7dsWjRItjaVh7u/+uvv+Dg4AAXFxd06dIFo0aN4m8TcfXqVf6UXpmtW7eKnIYts27dOoSG\nhsLR0REBAQGIiIiApqamVHUor1evXjA3N4eVlRW8vLzg6urKt489e/bA0dERbdq0wbfffovly5cD\nAAYNGoSYmBj+4gkLCwtEREQgODgYzs7OsLOz4zu7QPXHStOmTTFixAg4ODiIXDwhbhtNTU3MmzcP\n7u7ucHNzg7q6eq2OUXHHdvnRtKdPn8Le3h4TJ05E27Ztq02zurrp6uryF5PMmDFD5PR8cHAw3rx5\nw55QpxsAACAASURBVF/Y1adPH1y/XjrBe9myZRg4cCBcXV2RlJRU7VmOb7/9Fv/88w8cHBzwxRdf\nwNfXV6J6zp49G2lpabC1tYWfnx/c3Nz4Y3369OlwcnKCu7s7nJyc0KlTJ75jV9WxXj5fQj6WZ1nP\nsP7aepxNOotiVvrnUMAJ0Mm4E8a7jYeplmn9FrAW6u1Zsfv27YNAIMCJEyeQl5eHLVu28OvKrsTb\ntGkTbty4gT59+uDChQuws7Pj44wePRpTpkypNA+pDD0rtn7k5OTwp3Xv3r0Lb29vxMfH81/4VUlL\nS4OXlxc/YlNQUAA7OzvExsaKnKoi5L/CzMwMR44cEfneK+/nn39GamoqVq5c+ZFLVqqoqAjFxcVQ\nUlJCVlYWOnXqhKVLl4odba6oTZs2+Pvvv/lRxT59+mDWrFmVOsGE1JXC4kKcTTqLy08vg+F9X8FA\nwwD9rfujhXqLj16mBv+s2EGDBgEoHWkofwouJycHf//9N2JjY6GqqooOHTpgwIAB2L59O38ar3fv\n3rh16xbi4uIwbtw4kVuRkPp14cIFkQtbNm7cWGOnDng/OXz37t0YOnQoNm7ciPHjx1OnjhAx7O3t\noaioiBMnTtRbGV6/fo3evXujuLgY+fn5GDFihESdOgCYP38+fv31V6xZswa3b9+GQCCgTh35aBJe\nJ+Bw/GFk5L+fEqMgUEBXs67wNPKEgGvYs9Tq/eKJir3T+Ph4yMvL83N+AMDJyUlknsLRo0clSjso\nKIj/R6ilpQVnZ2d+Mm9ZerQs2+Vu3brh5s2btdq+T58+/HLFkdi6Lv+yZcuofdCyRMtl7+s6vy1b\ntvCjdRXXl93Truwq1/raH1evXhVZLlPT9mpqahg6dCgAwMHBAZMnT4ZQKPwkPl9ZL3+s9kLLNS97\ndPDA8YTjOHiidD6oqbMpAKAwoRCuLV3RrmW7j1qesvcfMkdfnHo7FVtm1qxZePr0KX8q9vz58xg6\ndChevHjBx9mwYQN27tyJyMhIidOlU7FEGuV/VAipDrUVIg1qL/WPMYbbabdxPOE4ct/l8uEq8iro\nadETjs0dP4kLdxr8qdgyFSuhrq6OrKwskbDMzEyRWz0QImv0xUskRW2FSIPaS/3KyM/A4fjDSHgt\nemcHBz0H9LToCTVFtSq2bLjqvWNXsZdsZWWFoqIiJCQk8Kdjb926hdatW9dH8QghhBDSwJSwElx5\ndgVnk86isLiQD2+i1AR9rPrASseqHktXtwT1lXHZhNuyK6sKCgpQXFwMNTU1+Pn5Yfbs2cjNzUV0\ndDQOHTqEzz//XOo8QkJCKs39IEQcaidEUtRWiDSovXx8qdmp2HR9E44nHOc7dRw4eBp6YqL7xE+u\nUycUlj5/WlbqbY5dSEgI5s2bVyls9uzZePPmDcaMGYNTp05BV1cXCxcu5O+ULymaY0ekQfNgiKSo\nrRBpUHv5eIpKinDu8TlEP4lGCXt//81mqs3Q37o/WjZpWY+lq5ms+i31fvFEXaGOHSGEEPLf8Djj\nMQ7FH8LL3PfPI5fj5NDZpDM6GneEnECyR1nWp0Zz8QQhhBBCSG3kF+Xj9MPTuPr8qki4cRNj9LPq\nh2ZqzeqpZPWnyo6dpHPalJSUsHHjRpkVSJZCQkLg5eVFw+CkRnS6hEiK2gqRBrWXunP/5X0ciT+C\nt4Vv+TAlOSX4tvKFm4HbJ3ELE0kIhUKZzsWs8lSskpISZsyYUeWwYNmQYVhYGN6+fSs2Tn2iU7FE\nGvTlSyRFbYVIg9qL7L0teItjCcdwN/2uSLi1jjV6W/ZGE+Wan3b0KarzOXbm5uZITEysMQFra2vE\nxcV9cEFkjTp2hBBCSOPBGMONlBs4mXgS+UX5fLiaghp6W/aGXTO7BjNKJw5dPFED6tgRQgghjcOr\n3Fc4FH8IjzIeiYS7tHBBd/PuUFFQqZ+CyZCs+i21uo/dw4cPZf5sM0LqE91rikiK2gqRBrWXD1Nc\nUozoJ9FYe3WtSKdOW1kbgU6BGGAz4P/Yu++ouK5rf+DfKQy9N9ERVaCGegMJCSGKAJe8OHKRixzH\n+dnW84uTX7yWbVnI8Xt+vxQnsfMSO26yrJc4tl+xQCBE0QgkoS4hq1E19CJ6hyn398c1M1wkpBmY\nmXtn2J+1smw2gtl2jofNPufuYxVFnTHpVdjt2LEDp06dAgB89tlnWLhwIWJjYwX70AQhhBBCLFvL\nQAs+uvgRiuqKoNKoALCDhjcEbcALq15AmHsYzxkKk15bsd7e3mhuboZMJsOiRYvw4Ycfws3NDQ88\n8ABqamru9+W8EIlE2Lt3Lz0VSwghhFgQpVqJY4pjKG8sBwNdieLn5Ies6Cz4OfvxmJ3xTTwVu2/f\nPvOdsXNzc0Nvby+am5uxevVqNDc3AwCcnZ0F+UQsQGfsCCGEEEtT212L3Kpc9Iz2aGNSsRSbQzdj\nXdA6iEW83YRqcmYdULx06VK88847UCgU2L59OwCgqakJrq6W+UgxIVPRSAKiL1orxBC0XvQzrBzG\n0dqjuNx2mROf7zYfmdGZ8LD34Ckzy6NXYffJJ59gz549kMlk+PWvfw0AKC8vx+OPP27S5AghhBBi\nvRiGwbXb15BfnY8h5ZA2bie1Q0p4CuLmxVn0CBM+0LgTQgghhJhd32gfDlcfRlVXFSe+0Hsh0iLT\n4CRz4ikzfpj9rtiysjJcunQJAwMD2hcXiUR47bXXZp2EqdCVYoQQQoiwMAyDcy3nUFRXhHH1uDbu\nYuuC7ZHbEe0VzWN25me2K8Um2717N7766iskJCTA3p47L+aLL74wWjLGRB07Ygg6B0P0RWuFGILW\nC1fHUAdyKnPQ2N/Iia/yX4WtYVthK7XlKTP+mbVjd/DgQVy7dg3+/v6zfkFCCCGEzC0qjQonGk6g\nrL4MakatjXs5eCErOgvBrsE8Zmdd9OrYLVmyBCUlJfDy8jJHTkZBHTtCCCGEf419jThUeQi3h29r\nYxKRBPHB8UgISYBUrPepMKtm1o7dJ598gueeew6PPfYYfH19OZ/buHHjrJMghBBCiHUZU42h+FYx\nzjWf4wwaDnQJRFZ0FnwcfXjMTjgqK+tRVFRrtO+nV2F34cIF5OXloays7I4zdo2NjdN8FSGWg87B\nEH3RWiGGmKvrpaqrCrlVuegf69fGZBIZkuYnYVXAKqseNGyIysp67N9fA5EoyWjfU6/C7vXXX0du\nbi6Sk5ON9sKEEEIIsS6D44M4UnMEVzuucuKRHpHIiMqAqx1dbDBZYWEturuTUGu8hp1+Z+yCg4NR\nU1MDmUxmvFc2MTpjRwghhJgHwzCoaK9AQU0BRlQj2riDjQPSItKwyGcRDRqeorsbeOklOVpaEgEA\nx48bp27Rqxf61ltv4V/+5V/Q2toKjUbD+Z+QZWdnG3U2DCGEEEK4ekZ68MWVL/C/N/+XU9Qt9V2K\nl1a/hMW+i6mom0SjAU6dAv7yF6C3V4PeXjkUimyjfX+9OnZi8d3rP5FIBLVafdfP8Y06dsQQc/Uc\nDDEcrRViCGteLxpGg9NNp3Hs1jEoNUpt3M3ODZlRmQj3COcxO2FqawMOHQJaWtiPOzvrUVFRg9DQ\nJBw8aManYuvq6mb9QoQQQgixDq0DrThUeQitg63amAgirA1ci83zN0MmsZyjW+agVALHj7Odusmb\nnQsXhuCRR4CrV0tw8KBxXovuiiWEEEKIXpRqJY7XH8epxlPQMLoKxdfRF1nRWQhwCeAxO2FSKICc\nHKCrSxeTSoFNm4D16wGJhI0Zq26Z9ozdnj179PoGe/funXUShBBCCBG2Wz238Jfzf8GJhhPaok4q\nliJpfhJ+suInVNRNMTrKFnT793OLupAQ4Kc/BRISdEWdMU3bsXNycsKVK1fu+cUMw2DFihXo7e01\nfmazRB07YghrPgdDjIvWCjGENayXEeUICusKcbH1Iice6haKzKhMeDp48pSZcN28CRw+DAwM6GK2\ntkByMrBiBXC3Z0lMfvPE8PAwIiIi7vsNbG3n7oW9hBBCiLViGAY3Om8grzoPg+OD2rid1A7JYclY\n7recnnadYnAQyMsDrl/nxhcsANLTARcX0+dAZ+wIIYQQwtE/1o+86jzc7LzJicd4xSA9Mh3Ots48\nZSZMDANcugQcPcpuwU5wcmILupiYu3fpJjPrXbGWKjs7G4mJiRbfBieEEELMgWEYXGi9gMLaQoyp\nx7RxZ5kz0iPTEeMdw2N2wtTdzZ6lu3WLG1+2DNi2DZhyE+sd5HK5UWfuUseOEFjHORhiHrRWiCEs\nab10DnfiUOUhNPQ1cOIr/FYgOTwZdlI7njITJo0GKC8Hjh0DVCpd3N0dyMwEwsIM+37UsSOEEELI\nrKk1apxsPInjiuNQM7pLBzztPZEZnYlQt1D+khOotjbg22+BVt0YP4hEwLp1wObNgI0Nf7lRx44Q\nQgiZo5r6m3Co8hA6hjq0MbFIjA1BG7ApdBOkYur/TDbdoOF584CsLMDff+bf26wdu46ODtjb28PZ\n2RkqlQoHDhyARCLBzp07p71ujBBCCCHCNK4eR8mtEpxpOgMGumLC39kfWdFZmOc0j8fshEmhYK8D\n6+7WxaRSIDGR7dSZYibdTOjVsVu9ejU+/PBDLFu2DK+++ipyc3NhY2ODxMRE/OEPfzBHngajjh0x\nhCWdgyH8orVCDCHE9VLdVY3cqlz0jfVpYzZiG2yZvwVrAtdALKKGzWSjo0BhIXDhAjceGsqepfM0\n0hg/s3bsqqurERcXBwA4ePAgTp06BWdnZ8TGxgq2sCOEEEKIztD4EApqC3ClnXv5QLh7ODKiMuBu\n785TZsJ14wY7l27qoOFt24Dly+8/woQPenXsvLy80NTUhOrqauzYsQPXrl2DWq2Gq6srBgcH7/fl\nvKCOHSGEEMKOMPmu4zscqTmCYeWwNm4vtUdqRCqW+C6hQcNTDAywBd2NG9z4ggXA9u2AswnG+Jm1\nY5eamopHHnkEXV1d+NGPfgQAuH79OgIDA2edACGEEEJMo3e0F7lVuajpruHEF/ssRmpEKhxljjxl\nJkz3GzQcG8tfbvrSq2M3OjqKzz//HDKZDDt37oRUKoVcLkdbWxt27NhhjjwNRh07YgghnoMhwkRr\nhRiCr/WiYTQ423wWxXXFUGqU2rirrSsyojIQ6Rlp9pyEbrpBw8uXs3e83m/Q8GyZtWNnZ2eH559/\nnhOjNzZCCCFEeNoH23Go8hCaB5q1MRFEWB2wGklhSZBJZDxmJzzTDRr28GAfjpg/n7/cZmLajt3O\nnTu5f/D7/XeGYTh78QcOHDBhejMnEomwd+9eulKMEELInKDSqFBaX4oTDSegYXRD1nwcfZAVnYVA\nFzo+NVVrKzvCZPKgYbGYHV+SmGieQcMTV4rt27fPKB27aQu77OxsbQHX2dmJzz//HJmZmQgJCUF9\nfT1yc3Px1FNP4b333pt1EqZAW7GEEELmivreehyqPISukS5tTCKSYFPoJmwI2gCJWCBD1gRCqQTk\ncrZTN3XQ8AMPAH5+5s/JWHWLXmfstm3bhj179iAhIUEbO3HiBN566y0cPXp01kmYAhV2xBB0boro\ni9YKMYSp18uoahRFdUU433KeEw92DUZWdBa8HLxM9tqW6tYt9iyd0AYNm/WM3enTp7F27VpObM2a\nNSgvL591AoQQQggx3I3bN5BXnYeBcd2QNVuJLbaGbcVK/5U0wmSK0VH2adeLF7lxYw8a5pteHbtN\nmzZh1apV+NWvfgV7e3sMDw9j7969OHPmDEpLS82Rp8GoY0cIIcQaDYwNIK86Dzc6uUPWoj2jsT1q\nO1xsXXjKTLhu3AAOHwYmj961s2MHDS9bJoxBw2bt2O3fvx+PPfYYXFxc4O7ujp6eHqxcuRJ/+9vf\nZp0AIYQQQu6PYRhcaruEo7VHMarSDVlzkjkhLSINsd6x1KWbYrpBwzEx7Fw6Uwwa5pteHbsJDQ0N\naGlpgZ+fH0JCQkyZ16xRx44Ygs5NEX3RWiGGMNZ66RruQk5VDhS9Ck58ud9yJIclw97GxEPWLAzD\nsFuuhYV3Dhrevp0t7ITGrB27CXZ2dvDx8YFarUZdXR0AICwsbNZJEEIIIeROao0a5U3lkCvkUGl0\nQ9Y87D2QGZWJ+e4WNmTNDLq62IcjFApufPlyduvVzo6XtMxGr47dkSNH8Oyzz6J18qAXsNWlWq02\nWXKzQR07QgghlqxloAWHKg+hbbBNGxOLxFgXuA6JoYmwkZhhyJoFUavZ8SVyuWUOGjbruJOwsDD8\n8pe/xJNPPgkHB4dZv6g5UGFHCCHEEo2rxyFXyFHeWA4Gup9jfk5+yIrOgp8zD0PWBG66QcPr1wOb\nNpln0PBsmbWw8/DwQFdXl0UdyqTCjhiCzk0RfdFaIYYwdL3UdtcityoXPaM92piN2Aab52/G2sC1\nEIvEJsjSck03aNjPD8jK4mfQ8EyZ9Yzds88+i08//RTPPvvsrF+QEEIIIVzDymEU1BSgor2CEw9z\nD0NGVAY87D14yky4phs0vHkzO2hYPEdrYL06dvHx8Th79ixCQkIwb9483ReLRDTHjhBCCJkhhmFw\nteMqjtQcwZBySBu3l9ojJSIFS32XWtRumTmMjLBPu04dNDx/PnuWzsNCa2CzbsXu379/2iSeeuqp\nWSdhClTYEUIIEbK+0T4crj6Mqq4qTnyRzyKkRqTCSebEU2bCxDDsPLq8PGEPGp4psxZ2logKO2II\nOjdF9EVrhRjibutFw2hwrvkcim8VY1w9ro272Lpge+R2RHtFmzlL4RsYYG+OuHmTG4+NBdLSrGPQ\nsFnP2DEMg88++wxffPEFmpubERgYiCeeeALPPPMMtYgJIYQQPXUMdeBQ5SE09TdpYyKIsCpgFZLm\nJ8FWastjdsIzMWj46FFgbEwXd3Zmb44Q4qBhvunVsfvXf/1XHDhwAD//+c8RHByMhoYG/P73v8fj\njz+ON954wxx5GkwkEmHv3r1ITEyk364JIYTwSqVRoay+DCcaTkDN6Oa/ejt4IzM6E8GuwTxmJ0zT\nDRpesQJITraeQcNyuRxyuRz79u0z31ZsaGgojh8/zrlGrL6+HgkJCWhoaJh1EqZAW7GEEEKEoKGv\nATmVObg9fFsbk4gkSAhJQHxwPKRigy6Bsnr3GjSclQWEhvKVmWmZdSt2eHgYXl5enJinpydGJ1/A\nRogFo3NTRF+0Vog+KmsqkX8uH4WnCsH4MggLC4OXP/tzNNAlEFnRWfBx9OE5S+FpaWEHDbfpLtuw\nuEHDfNOrsEtNTcUTTzyBd955ByEhIVAoFHj99deRkpJi6vwIIYQQi1JZU4l3D78LhYcC7Y7tcAt0\nw+Xrl7FKvAqPJjyKlf4radDwFEolcOwY26mb3LSyxEHDfNNrK7avrw+7d+/GP/7xDyiVStjY2OCR\nRx7B+++/Dzc3N3PkaTDaiiWEEGJuA2MDePkvL6POrY4T97D3wAblBvz8iZ/zlJlw1dWxZ+l6dJdt\nwMaGHTS8du3cGTTMy7gTtVqNzs5OeHl5QSKRzPrFTYkKO0IIIebCMAwutl5EYV0h5HI5RgPZo0o2\nYhtEekbC28Eb7u3u+Jcd/8JzpsIxMsI+7XrpEjdu6YOGZ8pYdYtedfDnn3+OiooKSCQS+Pr6QiKR\noKKiAl988cWsEyBECORyOd8pEAtBa4VMdXvoNj67/BlyqnIwqhqF+PsfrfOc5sH3ti98HH0gEokg\nE8t4zlQYGAa4fh34j//gFnV2dsADDwBPPjn3ijpj0uuM3Z49e3D58mVOLDAwEJmZmdi5c6dJEiOE\nEEKETKVR4UTDCZTVl3FGmCyLXYbu5m74hvpC0aQAAIxVjyFpcxJPmQpHfz97c8TdBg2npwNOdNnG\nrOm1Fevu7o7Ozk7O9qtKpYKnpyf6+vpMmuBM0VYsIYQQU7nbCBOxSIwNQRuwMWQj6m7VofhiMcY1\n45CJZUhanoToiLl7owTDABcusHe8Th00vH07sGABf7kJhVnHncTExOCbb77Bj370I23sf/7nfxBD\nI58JIYTMIaOqURTVFeF8y3lOPMA5AFnRWfB18gUAREdEz+lCbrLOTvbhiPp6bnzlSmDrVusZNCwU\nenXsTpw4gfT0dCQnJyMsLAy1tbUoKipCXl4e4uPjzZGnwahjRwxBs8mIvmitzE0Mw+BG5w3kV+dj\nYHxAG5dJZEian4RVAavuOsJkLq8XtRo4dQo4fpw7aNjTk304wloHDc+UWTt28fHx+O677/C3v/0N\nTU1NWL16Nf74xz8iKCho1gkQQgghQtY32oe86jxUdlVy4tGe0UiPTIernStPmQnXdIOGN2wANm6k\nQcOmZPC4k/b2dvj7+5syJ6Ogjh0hhJDZ0DAanG85j6K6Ioyrx7VxJ5kT0iPTEeMVA5FIxGOGwjM+\nzl4FNnXQsL8/O2h43jzeUhM8s3bsenp68OKLL+Kbb76BVCrF8PAwDh06hLNnz+Ltt9+edRKEEEKI\nkLQPtiOnKgdN/U2c+Aq/FUgOT4adlA6GTUWDhoVBr3/NP/3pT+Hi4oL6+nrY2toCANatW4cvv/zS\npMkRYi40m4zoi9aKdVOqlSiuK8aHFz7kFHVeDl54Ju4ZZEZnGlTUzYX1MjICfPstcOAAt6gLCwP+\nz/9h73mlos589OrYFRcXo7W1FTaTNsW9vb3R0dFhssQIIYQQc7rVcws5VTnoHunWxiQiCRJCEhAf\nHA+pWK8fmXPGxKDhvDxgaEgXt7MDUlKAuDiAdqrNT68zdhERESgtLYW/vz/c3d3R09ODhoYGbNu2\nDTenThkUCDpjRwghRB/DymEU1hbiUhv3bqtg12BkRmXC29Gbp8yEq78fOHwYqOQ+T4KFC4G0NBo0\nPBNmPWP34x//GP/0T/+Et99+GxqNBuXl5Xjttdfw/PPPzzoBQgghhA8Mw+Bqx1UcqTmCIaWu5WQr\nsUVyeDJW+K2ghyOmoEHDwqdXx45hGLz33nv48MMPoVAoEBwcjJ/+9Kd4+eWXBbvoqWNHDDGXZ00R\nw9BasQ69o704XHUY1d3VnHisdyzSItLgbOtslNexpvVCg4ZNy6wdO5FIhJdffhkvv/zyrF+QEEII\n4YuG0eBM0xmU3CqBUqPUxl1sXZAemY4FXtRymkqtBk6eBEpL7xw0nJUFhITwlxu5k14du5KSEoSG\nhiIsLAytra149dVXIZFI8M4772AeD0NpXn31VZSXlyM0NBSffvoppNI761Pq2BFCCJmsdaAVOVU5\naBlo0cZEEGFVwCokzU+CrdSWx+yEqbmZHTTc3q6LTQwa3rQJuMuPXzJDxqpb9HoA+YUXXtAWT6+8\n8gpUKhVEIhF+8pOfzDoBQ1VUVKClpQWlpaVYsGABvvnmG7PnQAghxHKMq8dxtPYoPrr4Eaeo83H0\nwbPLn0V6ZDoVdVOMjwMFBcDHH3OLOn9/4Cc/AZKSqKgTKr3+b2lpaUFwcDCUSiUKCgq08+z8/PxM\nnd8dysvLkZKSAgBITU3FZ599hh07dpg9D2JdrOkcDDEtWiuWpba7FrlVuegZ1Q1Yk4ql2BSyCeuD\n1kMilpj09S1xvdTWArm5dw4a3rIFWLOGZtIJnV6FnYuLC9ra2nDt2jUsXLgQzs7OGBsbg1KpvP8X\nG1lPT4+2oHRxcUF3d/d9voIQQshcMzQ+hILaAlxpv8KJz3ebj4yoDHg6ePKUmXCNjLBdusuXufGw\nMCAzE3B35ycvYhi96u7du3dj9erVeOyxx/DCCy8AAE6ePImYmJgZv/Cf/vQnrFy5EnZ2dnjmmWc4\nn+vu7sZDDz0EJycnhIaG4u9//7v2c25ubujv7wcA9PX1wcPDY8Y5EDLB0n6jJvyhtSJsDMPgcttl\n/OnsnzhFnb3UHg9EP4Anlz5p1qLOEtYLwwBXrwJ/+hO3qLO3Bx58ENi5k4o6S6JXx+7VV1/Fgw8+\nCIlEgoiICABAYGAgPv744xm/cEBAAPbs2YOCggKMjIxwPvfiiy/Czs4OHR0duHTpErZv346lS5ci\nNjYW69evx7vvvoudO3eioKAA8fHxM86BEEKI9ege6UZuVS7qeuo48cU+i5EakQpHmSNPmQnXdIOG\nFy0CUlNp0LAl0uupWFPas2cPmpqa8NlnnwEAhoaG4OHhgWvXrmmLyKeeegr+/v545513AAC//OUv\ncfr0aYSEhOCzzz6jp2LJrFniORjCD1orwqPWqFHeVA65Qg6VRjePw83ODdsjtyPSM5K33IS6XhgG\nOH8eKCriDhp2cWEHDUdH85fbXGXyOXYLFizQXhcWFBQ0bRINDQ2zSmDqP0RVVRWkUqm2qAOApUuX\nci5S/vWvf63X93766acRGhoKgN3CjYuL0/4HNvH96GP6GAAuf7//IJR86GP6mD7W7+Pm/mb89m+/\nRc9oD0LjQgEAissKxHrH4oUdL0AmkQkqXyF8/O23cpw8CTg4sB8rFOznf/jDRCQlAadPy9HaKpx8\nrfXjib9XKBQwpmk7dmVlZUhISLgjiakmEp2pqR27srIyPPLII2htbdX+mY8++gh/+9vfcOzYMb2/\nL3XsCCHEeo2pxnBMcQxnms6Age693s/JD5nRmfB39ucxO2GaGDR8/Dj79xO8vNiHI2jQML9M3rGb\nKOqA2Rdv9zL1H8LJyUn7cMSEvr4+ODsb53oXQgghlq2qqwqHqw6jb6xPG7MR22Dz/M1YG7gWYpGY\nx+yEabpBw/HxwMaNNJPOmkz7f+WePXumrR4n4iKRCG+99dasEph612xUVBRUKhVqamq027EVFRVY\ntGjRrF6HkHuRy+Um/QWGWA9aK/wZHB9EfnU+rt2+xomHu4cjIyoD7vbCe3ST7/UyPg4cOwacPs2e\nq5sQEMBeB+bry1tqxESmLewaGxvvKLommyjsZkqtVkOpVEKlUkGtVmNsbAxSqRSOjo54+OGH8eab\nb+Ljjz/GxYsXkZOTg/LycoNfIzs7G4mJifQmTAghFoxhGFxsvYjCukKMqka1cQcbB6RGpGKx20js\nJwAAIABJREFUz+JZ/TyyVrW1QE4O0Nuri9GgYeGRy+X3PPJmKN6eis3Ozr6j25ednY0333wTPT09\n2LVrFwoLC+Hl5YV///d/N/h2CTpjRwghlq9zuBM5lTmo76vnxOPmxWFb+DY42DjwlJlwDQ+zg4Yr\nKrjx8HAgI4Nm0gmVseqWaQu7urq6u4XvEBYWNuskTIEKO0IIsVxqjRonGk6gtL4UakZ30t/D3gMZ\nURkIcxfmzx4+MQxw7RqQnw8MDeni9vbsTLolSwBqbAqXyQs7sR49WpFIBPXkR2sEhAo7Ygi+z8EQ\ny0FrxfQa+hqQU5mD28O3tTGxSIz1QeuxKWQTbCQ2PGZnGHOtl74+dtBwVRU3ToOGLYfJn4rVaDSz\n/uaEEEKIvkZVoyiqK8L5lvOceIBzADKjMzHPaR5PmQkXDRomU/F+84SpiEQi7N27lx6eIIQQC3Dj\n9g3kVedhYHxAG5NJZEian4RVAatohMld3L7NjjBpbOTGV60Ctm4FbG35yYsYZuLhiX379pl2KzYl\nJQUFBQUAuDPtOF8sEqG0tHTWSZgCbcUSQojw9Y/1I686Dzc7b3LiUZ5R2B65Ha52rjxlJlxqNXDi\nBFBaeueg4awsIDiYv9zIzJl8K/bJJ5/U/v2zzz47bRKEWAM6N0X0RWvFOBiGwfmW8yiqK8KYWreH\n6CRzQlpEGmK9Y63iZ4yx10tTE9ul6+jQxcRiICGB/R8NGibTLoHHH39c+/dPP/20OXIhhBAyB3QM\ndSCnMgeN/dw9xBV+K7A1bCvsbex5yky4xseBkhLgzBkaNEzuTe8zdqWlpbh06RKGvn+GemJA8Wuv\nvWbSBGeKtmIJIURYVBoVSutLcaLhBDSM7gE9LwcvZEZlIsSNLiu9m5oaIDf3zkHDSUnA6tU0aNha\nmHwrdrLdu3fjq6++QkJCAuztLec3Kbp5ghBChEHRq0BOZQ66Rrq0MYlIgvjgeCSEJEAqpj3Eqe41\naDgzE3Bz4ycvYly83Dzh7u6Oa9euwd/f32gvbGrUsSOGoHNTRF+0VgwzohxBYV0hLrZe5MSDXIKQ\nGZ0JH0cfnjIzj5msF4YBrl4FjhyhQcNziVk7dkFBQZDJZLN+MUIIIXMDwzC4dvsa8qvzMaTUVSe2\nElskhydjhd8Kq3g4wtimGzS8eDFb1Dk68pMXsRx6dezOnTuHf/u3f8Njjz0G3yknNDdu3Giy5GaD\nOnaEEMKP3tFeHK46jOruak48xisGaZFpcLF14Skz4WIY4Nw5dtDw+Lgu7uLC3u8aFcVfbsQ8zNqx\nu3DhAvLy8lBWVnbHGbvGqZMRCSGEzEkaRoMzTWdQcqsESo1SG3exdUF6ZDoWeC3gMTvhutugYZGI\nHTSclESDholh9OrYeXp64ssvv0RycrI5cjIK6tgRQ9C5KaIvWit31zbYhkOVh9Ay0KKNiSDCqoBV\nSJqfBFvp3KxO7rVeaNAwmcysHTtHR0ds2rRp1i9mbvRULCGEmJZSrYRcIUd5UzlnhImPow8yozIR\n5BrEY3bCdbdBwxIJEB9Pg4bnGl6eit2/fz/Onj2LPXv23HHGTizQATrUsSOEENOq7a5FblUuekZ7\ntDGpWIqNIRuxIWgDJGIJj9kJ0/g4UFwMnD3LHTQcGMh26Xys+yFhcg/Gqlv0KuymK95EIhHUk/vH\nAkKFHSGEmMbQ+BCO1h5FRTt3wFqoWygyozLh6eDJU2bCVlMD5OSwT75OkMmALVto0DAx81ZsXV3d\nrF+IECGjc1NEX3N5rTAMgyvtV1BQW4Bh5bA2bi+1x7bwbYibF0cjTKaQy+VYvToRR44AV65wPxcR\nwT7xSoOGiTHpVdiFhoaaOA1CCCFC1j3SjdyqXNT1cH/RX+SzCKkRqXCSOfGUmTBVVtajsLAWJ05c\nwR/+oEFgYDi8vNgr0xwc2Jl0ixfToGFifHrfFWtpaCuWEEJmT61R43TTacgVcs4IE1dbV2REZSDS\nM5LH7ISpsrIeH35YA4UiCd3dbEylKkZcXAQ2bw6hQcPkrsy6FUsIIWTuae5vRk5VDtoG27QxEURY\nG7gWm+dvhkxCNxJNxTDAp5/WoqIiiTPCxNExCZ6eJfjBD0L4S47MCVZ9VDM7O9uojxAT60XrhOhr\nLqyVcfU4jtQcwccXP+YUdfOc5uHHy3+MlIgUKuruorcXOHgQqKgQa4u63l45AgLYYcPu7lb9I5fM\nkFwuR3Z2ttG+n1V37Iz5L4oQQuaC6q5q5Fblom9M9+imjdgGiaGJWBu4lkaY3AXDABcuAEePsuNM\nxGJ2np+9PTtsOPL73WqZTHOP70Lmqol5u/v27TPK99PrjF1dXR1ef/11XL58GYODg7ovFonQ0NBg\nlESMjc7YEUKI/gbHB3Gk5giudlzlxMPdw7E9ajs87D14ykzYenvZQcOTh0d0ddXj9u0aREYmQfJ9\nHTw2Voynn45AdDRtxZK7M+scu7Vr1yIiIgKPP/74HXfFCvWxfyrsCCHk/hiGwaW2SzhaexSjqlFt\n3MHGASnhKVjiu4RGmNzF1C7dBC8v4IEHgOHhehQX12J8XAyZTIOkpHAq6sg9mbWwc3FxQU9PDyQS\ny2nBU2FHDDGXZ5MRw1jTWuka7kJOVQ4UvQpOfKnvUqREpMDBxoGfxASup4ft0t26pYuJRMD69UBi\nImBjo4tb03ohpmXWp2I3btyIS5cuYeXKlbN+QUIIIfxSa9Q42XgSpfWlUGlU2ri7nTsyojIQ7hHO\nY3bCxTDA+fNAYeGdXboHH2SvBSOEb3p17F588UX84x//wMMPP8y5K1YkEuGtt94yaYIzRR07Qgi5\nU2NfI3KqctAxpLt9XiwSY33QemwK2QQbic09vnru6ukBvv0WUCh0MZEI2LCB7dJJrfpRRGIOZu3Y\nDQ0NISMjA0qlEk1NTQDYcxl07oIQQizDqGoUxXXFON9yHgx0Pzz8nf2RFZ2FeU7zeMxOuBgGOHeO\n7dIpdfOZ4e3NdukCAvjLjZC70auw279/v4nTMI3s7GztY8SE3AudgyH6ssS1crPzJg5XHcbA+IA2\nJpPIsGX+FqwOWA2xiOar3U13N3uWbmqXLj4e2LRJvy6dJa4XYl5yudyo8zGnXZYKhUJ7R2xdXd10\nfwxhYWFGS8bYaI4dIWQu6x/rR351Pm503uDEIz0isT1qO9zs6Pb5u2EY4OxZoKiI26Xz8WGfeKUu\nHTEms82xc3Z2xsAA+9udWHz33+ZEIhHUk+9MERA6Y0cImasYhsH5lvMoqivCmHpMG3eSOSEtIg2x\n3rF0lGYa3d3sWbr6el1MLGa7dBs30lk6YjpmHXdiiaiwI4TMRR1DHcipzEFjfyMnvtxvOZLDkmFv\nYz/NV85tDAOcOQMUF9/ZpXvwQcDfn7/cyNxg1ocnCLF2dA6G6Euoa0WlUaGsvgwnGk5Azeh2Ujzt\nPZEZnYlQt1D+khO4ri62Szf5IiVjdemEul6I9aLCjhBCLFx9bz1yqnLQOdypjYlFYsQHx2NjyEZI\nxfRWfzcaja5Lp9KN84OvL9ul8/PjLzdCZoq2YgkhxEKNKEdQWFeIi60XOfEglyBkRmfCx9GHp8yE\nr6sL+N//BRon7ViLxUBCAtuls6CLloiVoK1YQgiZoxiGwfXb15Ffk4/B8UFt3FZii61hW7HSfyU9\nHDENjQY4fRooKeF26ebNY594pS4dsXQGF3YajYbz8XRPzBJiSegcDNEX32ulb7QPh6sPo6qrihNf\n4LUA6ZHpcLF14Skz4evsZLt038/ZB8B26TZuZDt1pujS8b1eyNyjV2F34cIFvPTSS6ioqMDo6Kg2\nLuRxJ4QQYk00jAZnm8+i5FYJxtW6i0qdZc5Ij0xHjHcMj9kJ2726dA8+yP6VEGuh1xm7RYsWISsr\nC0888QQcHBw4n5sYYiw0dMaOEGIt2gbbkFOZg+aBZk58lf8qJIUlwU5qx1Nmwne3Lp1Ewnbp4uPp\nLB0RDrPOsXNxcUFfX59Fndmgwo4QYumUaiWO1x/HqcZT0DC6YzDeDt7IjM5EsGswj9kJm0YDlJcD\nx45xu3R+fmyXzteXv9wIuRtj1S16HZB76KGHUFBQMOsXM7fs7Gyj3r9GrBetE6Ivc62Vup46/OX8\nX3Ci4YS2qJOIJNgyfwt+uvKnVNTdw+3bwCefAIWFuqJOIgG2bAF+/GPzFnX03kLuRy6XG/UKVL3O\n2I2MjOChhx5CQkICfCf9FyESiXDgwAGjJWNsdFcsIcTSDCuHUVBTgIr2Ck48xDUEmdGZ8HLw4ikz\n4dNogFOn2C7d5OPf1KUjQma2u2Inm65AEolE2Lt3r1ESMTbaiiWEWBKGYfBdx3c4UnMEw8phbdxO\naodt4duwbN4yizoOY24dHeztEc2TjiFKJEBiIrB+PZ2lI8JHd8XeBxV2hBBL0TPSg9yqXNT21HLi\nC70XIi0yDU4yJ54yEz6NBjh5EpDLuV06f3+2S+dDM5qJhTD7gOJjx47hwIEDaG5uRmBgIJ544gls\n2bJl1gkQIgQ0a4roy5hrRcNoUN5YDrlCDqVGd/O8q60rtkdtR5RnlFFex1p1dLBPvLa06GITXboN\nG9gZdXyj9xZibnot+48//hg/+tGP4Ofnh4cffhjz5s3DY489hr/+9a+mzo8QQqxSy0AL/nrhryis\nK9QWdSKIsDZwLV5c/SIVdfegVgOlpcCHH3KLuoAA4Pnn2WHDQijqCOGDXluxkZGR+Oabb7B06VJt\n7MqVK3j44YdRU1Nj0gRnirZiCSFCNK4ex7Fbx3C66TQY6N6jfB19kRWdhQCXAB6zE772drZL19qq\ni0kkwObN7Fk6KuiIpTLrGTtPT0+0trZCJpNpY2NjY/D390dXV9eskzAFKuwIIUJT3VWNw9WH0Tva\nq41JxVIkhiZiXeA6SMR0wn86ajVw4gTbqZt8li4ggD1L5+3NX26EGINZ59ht2LABr7zyCoaGhgAA\ng4OD+MUvfoH169fPOgFChIBmTRF9zWStDI4P4pvr3+A/v/tPTlEX5h6GF1a9gPjgeCrq7qGtDfj4\nY+4YE6kUSE4Gnn1W2EUdvbcQc9Pr4YkPPvgAO3bsgKurKzw8PNDd3Y3169fj73//u6nzI4QQi8Uw\nDC63XcbR2qMYUY1o4w42DkgJT8ES3yU0wuQeJrp0x4+zT79OCAxku3ReNNKPkDsYNO6ksbERLS0t\n8Pf3R1BQkCnzmjXaiiWE8KlruAs5VTlQ9Co48aW+S7EtfBscZY78JGYh2trYs3RtbbqYVMreHrF2\nLZ2lI9bH5GfsGIbR/iapmfyr0hRigf7XRYUdIYQPao0aJxtPorS+FCqN7pJSdzt3ZERlINwjnMfs\nhE+tBsrK2LN0k3/0BAUBDzxAXTpivUw+x87FxQUDAwPsH5Le/Y+JRCKoJ59iJcRC0awpoq97rZWm\n/iYcqjyEjqEObUwsEmNd4DpsCt0EmUR2168jrNZWtkvX3q6LSaVAUhKwZo1ldunovYWY27SF3bVr\n17R/X1dXZ5ZkCCHEEo2pxlB8qxjnms9xRpj4O/sjMyoTfs5+PGYnfBNz6crKuF264GC2S+fpyV9u\nhFgavc7Y/fa3v8UvfvGLO+LvvvsuXnnlFZMkNlu0FUsIMYebnTeRV52H/rF+bcxGbIMt87dgTeAa\niEUW2GYyo5YW9o7XyV06Gxu2S7d6tWV26QiZCbPOsXN2dtZuy07m7u6Onp6eWSdhCiKRCHv37kVi\nYiK1wQkhRjcwNoD8mnxcv32dE4/0iMT2qO1ws3PjKTPLoFKxXboTJ6hLR+Y2uVwOuVyOffv2mb6w\nKykpAcMwyMzMRG5uLudztbW1ePvtt1FfXz/rJEyBOnbEEHQOhtxPZU0lii4U4frV63AMcoTKRQWX\neS7azzvaOCItMg0LvRfSCJP7aGlhz9J16I4iwsYG2LqV7dJZ078+em8h+jL5wxMAsGvXLohEIoyN\njeHZZ5/lvLivry/ef//9WSdACCFCV1lTif3H9kMVosJlXIaNzAaqqyrEaeLg5e+FZfOWYVv4Ntjb\n2POdqqCpVOxMupMnuV26kBC2S+fhwV9uhFgLvbZid+7ciS+++MIc+RgNdewIIcbyh7//ARdsL6Cp\nv4nzcIRPuw/+30/+H+a7z+cxO8vQ3Mx26W7f1sWstUtHyEyYpWM3wdKKOkIIMQaGYfBdx3coUZSg\n30/3cIQIIgS7BmOJbAkVdfehUgFyOdulm/wzKzQUyMqiLh0hxqZXYdfX14fs7GwcP34cXV1d2oHF\nIpEIDQ0NJk2QEHOgczBkqrbBNuRV56GhrwEqtW7QsLJWiXUJ6+Akc4K9krZe76WpiX3idXKXTiZj\nu3SrVs2NLh29txBz06uwe/HFF9HY2Ig333xTuy37m9/8Bj/4wQ9MnR8hhJjViHIEJbdKcL7lvHbb\nNSwsDDcqbyBqVRSGe4bhJHPCWPUYkjYn8ZytMKlUwLFjwKlTd3bpHngAcHfnLTVCrJ5eZ+y8vb1x\n48YNeHl5wdXVFX19fWhubkZmZiYuXrxojjwNRmfsCCGG0DAaXGq9hOJbxRhWDmvjEzdH+Kp8caLi\nBMY145CJZUhanoToiGgeMxampib2LF1npy4mkwHJycDKlXOjS0fITJj1jB3DMHB1dQXAzrTr7e2F\nn58fqqurZ50AIYTwrbGvEfk1+WgZaOHEw93DkRaZBi8H9oLSJdFL+EjPIiiV7Fm6qV26+fPZs3TU\npSPEPPQq7JYsWYLS0lIkJSUhPj4eL774IhwdHREdTb+tEutA52DmpsHxQRTVFeFy22VO3M3ODakR\nqYj2jL5jJh2tlTs1NrJn6aZ26bZtA1asmNtdOlovxNz0Kuw++ugj7d//8Y9/xGuvvYa+vj4cOHDA\nZIkRQoipqDVqnG0+C7lCjjH1mDYuFUsRHxyPDUEbYCOx4TFDy6BUsmfpysu5XbqwMLZL50aXbxBi\ndnqdsTtz5gzWrFlzR/zs2bNYvXq1SRKbLTpjRwi5m7qeOuRX5+P28G1OPMYrBikRKXQVmJ4aGtgu\nXVeXLmZry3bpli+f2106QmZCEHfFenh4oLu7e9ZJmAIVdoSQyfpG+1BQW3DH3a5eDl5Ii0hDuEc4\nT5lZFqUSKCkBTp/mdunCw9ku3ffHsQkhBjLLwxMajUb7IprJ97+AvStWKtVrJ5cQwaNzMNZLpVHh\nVOMplNWXQalRauMyiQyJoYlYE7AGErFE7+83l9dKQwP7xOvk3+dtbYGUFGDZMurS3c1cXi+EH/es\nzCYXblOLOLFYjNdff900WRFCyCwxDIOqriocqTmCntEezueW+C5BclgynG2decrOsiiVQHExcOYM\nt0sXEQFkZlKXjhAhuedWrEKhAABs3LgRZWVl2u6dSCSCt7c3HBwczJLkTNBWLCFzV9dwF47UHEF1\nN3ck0zyneUiPTEewazBPmVme+nr2LN3ULl1qKhAXR106QozFrGfsLBEVdoTMPePqcZTVl+FU4ymo\nGbU2bi+1x5b5W7DCfwXEIjGPGVqO8XG2S3f27J1duqwswMWFv9wIsUZmHVC8c+fOuyYAgEaeEKtA\n52AsG8MwuHb7Go7WHkX/WL82LoIIK/xXYMv8LXCwMc4Ow1xYKwoF26XrmbSDbWfHnqWjLp1h5sJ6\nIcKiV2EXHh7OqSTb2trwX//1X3j88cdNmhwhhNxP+2A78mvyoehVcOKBLoFIj0yHv7M/P4lZoPFx\noKiI7dJNFhnJnqWjLh0hwjfjrdjz588jOzsbubm5xs7JKGgrlhDrNqoaxbFbx3Cu5Rw0jO6pfSeZ\nE7aGbcVS36V33BpBpjddly41FVi6lLp0hJga72fsVCoV3N3d7zrfTgiosCPEOjEMg8ttl1FUV4Qh\n5ZA2LhaJsSZgDTaFboKd1I7HDC3L+DhQWAicO8eNR0UBGRnUpSPEXMx6xq64uJjzm+/Q0BC+/PJL\nLFy4cNYJGKq/vx9bt27FjRs3cObMGcTGxpo9B2J96ByMZWjub0ZedR6aB5o58TD3MKRFpMHb0dvk\nOVjTWrl1i+3S9fbqYnZ2QFoasGQJdemMwZrWC7EMehV2zz77LKewc3R0RFxcHP7+97+bLLHpODg4\nIC8vD//3//5f6sgRMkcMjQ+h+FYxLrZe5MRdbV2REpGCGK8Y2nY1wNgYe5ZuapcuOprt0jnTeD9C\nLJZehd3EPDshkEql8PLy4jsNYmXoN2ph0jAanGs+h2OKYxhVjWrjUrEU64PWIz44HjKJzKw5Wfpa\nqasDDh3iduns7dku3eLF1KUzNktfL8Ty6H0nWG9vLw4fPoyWlhb4+/sjPT0d7u7upsyNEDKHKXoV\nyK/OR/tQOyce7RmNlIgUeNh78JSZZRobY8/SnT/PjVOXjhDrotekzpKSEoSGhuK9997DuXPn8N57\n7yE0NBRFRUUGvdif/vQnrFy5EnZ2dnjmmWc4n+vu7sZDDz0EJycnhIaGcrZ5f//732Pz5s343e9+\nx/ka2nohxiKXy/lOgXyvf6wf31z/Bvsv7+cUdZ72nnh88eN4dPGjvBZ1lrhWamuBP/+ZW9TZ2wM/\n+AGwYwcVdaZkieuFWDa9OnYvvvgi/vrXv+KRRx7Rxr7++mu89NJLuHnzpt4vFhAQgD179qCgoAAj\nIyN3vIadnR06Ojpw6dIlbN++HUuXLkVsbCx+9rOf4Wc/+9kd34/O2BFiPVQaFU43nUZpfSnG1ePa\nuI3YBptCN2Ft4FpIxXpvMhCwXbqjR4ELF7jxBQvYLp2TEz95EUJMR69xJ25ubujq6oJEItHGlEol\nvL290Tv5oIae9uzZg6amJnz22WcA2KdsPTw8cO3aNURERAAAnnrqKfj7++Odd9654+vT09NRUVGB\nkJAQPP/883jqqafu/AcTifDUU08hNDRU+88QFxenPe8w8VsUfUwf08f8f3zw24M423wWHrFsJ05x\nWQEAyNiWgW3h23Cx/KKg8rWEj5ubgY6ORPT1AQoF+/nY2ESkpwO3b8shEgkrX/qYPp5rH0/8/cRz\nDJ9//rn55tjt3r0bERERePnll7Wx9957D9XV1Xj//fcNftE33ngDzc3N2sLu0qVLiI+Px9CQbibV\nu+++C7lcjkOHDhn8/QGaY0eIJege6UZBTQEquyo5cR9HH6RHpiPULZSfxCzY6CjbpbvIfYAYMTHA\n9u3UpSNEqMw6x+7ixYv44IMP8Otf/xoBAQFobm5GR0cH1qxZg4SEBG1CpaWler3o1LNxg4ODcJky\nBdPZ2Vmww4+J9ZHL5drfpojpKdVKlDWU4VTjKag0Km3cTmqHzaGbsSpgFcQiMY8ZTk/Ia6Wmhn3i\ntV93XS4cHNiCLjaWnnjlg5DXC7FOehV2zz33HJ577rl7/hlDHmSYWpE6OTmhf/I7EYC+vj4404le\nQqwKwzC40XkDBTUF6Bvr43xuud9yJM1PgqPMkafsLNfoKFBQAFy6xI3HxrJFnSP9KyVkztCrsHv6\n6aeN+qJTi8CoqCioVCrU1NRoz9hVVFRg0aJFs3qd7OxsJCYm0m9L5L5ojZje7aHbyK/JR11PHSce\n4ByA9Mh0BLgE8JSZYYS2VqqrgZycu3fpeLgciEwhtPVChEcul3PO3c2W3nfFlpaW4tKlS9pzcAzD\nQCQS4bXXXtP7xdRqNZRKJfbt24fm5mZ89NFHkEqlkEgkePTRRyESifDxxx/j4sWLyMjIQHl5OWJi\nYmb2D0Zn7AgRhFHVKI4rjuNM8xloGI027mjjiKSwJCybt4xGF83A6Chw5Ahw+TI3vnAhkJ5OXTpC\nLI1Zz9jt3r0bX331FRISEmBvbz/jF/vVr36Ft956S/vxwYMHkZ2djTfffBN//vOfsWvXLvj4+MDL\nywsffPDBjIs6QgxF52CMj2EYVLRXoKiuCIPjg9q4WCTGKv9V2Dx/M+ykdjxmODNCWCtVVWyXbvIx\nZEdH3Vk6IhxCWC9kbtGrY+fu7o5r167B39/fHDkZBXXsiCHozde4WgdakVedh8b+Rk481C0UaRFp\n8HXy5Smz2eNzrYyMsGfppnbpFi1iu3QODrykRe6B3luIvoxVt+hV2C1ZsgQlJSUWdUcrFXaEmN+w\nchglt0pwoeUCGOj++3OxdcG28G1Y6L2Qtl1nqLISyM29s0uXkcGOMiGEWDazbsV+8skneO655/DY\nY4/B15f7m/bGjRtnnYSp0MMThJiHhtHgQssFlNwqwYhKd6uMRCTBuqB12BiyETKJjMcMLdfICHuW\nrqKCG1+8GEhLoy4dIZaOl4cnPvjgA7z88stwdna+44xdY2PjNF/FL+rYEUPQdsnMNfQ1IK86D22D\nbZx4pEckUiNS4engyVNmpmHOtVJZyZ6lG9QdUYSTE3uWjrp0loHeW4i+zNqxe/3115Gbm4vk5ORZ\nvyAhxDoMjA2gsK4QV9qvcOLudu5IjUhFlGcUbbvO0PAw26W7wv1XiyVLgNRU6tIRQqanV8cuODgY\nNTU1kMksZyuFOnaEmIZao8aZ5jOQK+QYV49r4zZiGySEJGB90HpIxXr9zkju4uZN9izd1C5dRgaw\nYAF/eRFCTMusD0/s378fZ8+exZ49e+44YycWC/PaHyrsCDG+2u5a5Nfko3O4kxOP9Y5FSngKXO1c\necrM8g0PA/n5wHffceNLl7JdullMmiKEWACzFnbTFW8ikQhqtXrWSZiCSCTC3r176eEJohc6B3Nv\nvaO9KKgpwI3OG5y4t4M30iLTEOYexlNm5meKtXLjBtul+37+OwDA2Znt0kVHG/WliJnRewu5n4mH\nJ/bt22e+M3Z1dXX3/0MClJ2dzXcKhFg0pVqJk40ncaLhBFQalTZuK7FFYmgiVgeshkQs4TFDyzY8\nDOTlAVevcuPUpSNk7phoQO3bt88o30/vK8UAQKPRoL29Hb6+voLdgp1AW7GEzBzDMLjZeRMFtQXo\nHe3lfC5uXhy2hm2Fk8yJp+ysw/XrwOHDd3bpMjOBqCj+8iKE8MOsT8X29/fjpZdewpf7OtnjAAAg\nAElEQVRffgmVSgWpVIodO3bg/fffh6srnakhxJp0DncivzoftT21nLifkx/SI9MR5BrEU2bWYWiI\n7dJdu8aNx8UBKSnUpSOEzI5ebbfdu3djaGgIV69exfDwsPavu3fvNnV+hJiFMYdDWqox1RgKawvx\nl3N/4RR1DjYOyIzKxHMrnqOiDrNbK9euAX/+M7eoc3EBHn8cePBBKuqsEb23EHPTq2N35MgR1NXV\nwdHREQAQFRWF/fv3Iyxs7hyYJsRaMQyD7zq+Q2FtIQbGdfdViSDCSv+V2DJ/C+xtqOKYjaEhdtv1\n+nVufNkytktnZ8dPXoQQ66NXYWdvb4/bt29rCzsA6OzshJ3A343oSjGir7m6RtoG25BXnYeGvgZO\nPNg1GOmR6ZjnNI+nzITLkLXCMLqzdMPDuriLC5CVBUREGD8/Iixz9b2F6I+XK8XefvttfP755/j5\nz3+OkJAQKBQK/P73v8fOnTuxZ88eoyVjTPTwBCHTG1GOoORWCc63nAcD3X8nzjJnJIcnY7HPYro1\nYpYGB9mzdFO7dMuXA9u2UZeOEMJl1jl2Go0G+/fvx3/+53+itbUV/v7+ePTRR7Fr1y7BvvlTYUcM\nMVdmTWkYDS61XkLxrWIMK3UtJLFIjHWB67AxZCNspbY8Zih891srDMOeocvL43bpXF3ZJ16pSze3\nzJX3FjJ7Zn0qViwWY9euXdi1a9esX5AQwo/Gvkbk1+SjZaCFEw93D0daZBq8HLx4ysx6DA6y2643\nuHOcsWIFkJxMXTpCiOnp1bHbvXs3Hn30Uaxfv14bO3XqFL766iv84Q9/MGmCM0UdO0JYg+ODKKor\nwuW2y5y4m50bUsJTsMBrgWA775aCYdghw3l5wMiILu7qyp6lCw/nLzdCiGUw61asl5cXmpubYWur\n26IZHR1FUFAQbt++PeskTIEKOzLXqTVqnG0+C7lCjjH1mDYuFUsRHxyPDUEbYCOx4TFD6zAwwHbp\nbt7kxleuZLt0trSzTQjRg9m3YjUaDSem0WiocCJWw9rOwdzquYW86jzcHub+4hXjFYOUiBS42bnx\nlJnlm1grDAN89x2Qn8/t0rm5sV06mgZFAOt7byHCp1dhFx8fjzfeeAO/+c1vIBaLoVarsXfvXiQk\nJJg6v1mhcSdkrukb7UNBbQGu3+Y+iunl4IW0iDSEe9CeoDEMDAC5uUBlJTe+ahWwdSt16Qgh+uNl\n3EljYyMyMjLQ2tqKkJAQNDQ0wM/PDzk5OQgKEuYketqKJXOJSqPCqcZTKKsvg1Kj1MZlEhk2hWzC\n2sC1kIglPGZo+Sor61FYWIv6ejEqKzUIDg6Hl1cIALZL98ADwPz5PCdJCLFYZj1jBwBqtRpnz55F\nY2MjgoKCsGbNGojFet1Ixgsq7MhcUdVVhfzqfPSM9nDiS3yXIDksGc62zjxlZj0qK+vx0Uc1uHUr\nCV1dbEylKkZcXATS0kKQnAzIZPzmSAixbGYv7CwNFXbEEJZ4DqZruAtHao6guruaE5/nNA/pkekI\ndg3mKTPr88YbJThzZguUSqC3Vw43t0TY2QHr15fgzTe38J0eETBLfG8h/DDrwxOEEOEYV4+jrL4M\npxpPQc2otXF7qT22zN+CFf4rIBYJt5tuScbGgCNHgPPnxVDqdrgREMA+HOHiQv+eCSHCQoUdIbCM\n+xwZhsG129dwtPYo+sf6tXERRFjutxxJYUlwsHHgMUPrUl8P/M//AL29gFjMTgWwtQU2bUqEuzv7\nZ2QyzT2+AyGW8d5CrMt9CzuGYXDr1i0EBwdDKqU6kBA+tA+2I78mH4peBSce6BKI9Mh0+Dv785OY\nFVKpgGPHgFOn2MHDABAWFo6WlmLExCTB5vvRf2NjxUhKovvBCCHCct8zdgzDwNHREYODg4J+WGIq\nOmNHDCHUczCjqlEcu3UM51rOQcPoukNOMidsDduKpb5L6dYII2pvB/77v9m/TrCzAzIyABubehQX\n1+L69SuIjV2CpKRwREeH8JcssQhCfW8hwmO2M3YikQjLli1DZWUlYmJiZv2ChJD7YxgGl9suo6iu\nCEPKIW1cLBJjTcAabArdBDspXTxqLAwDlJcDxcWAWndsEeHh7BgTFxcACEF0dAjkcjH9oCaECJZe\ne6ubN29GWloann76aQQFBWmrSpFIhF27dpk6xxmjAcVEX0JaI839zcirzkPzQDMnPt9tPtIi0+Dj\n6MNTZtapr489S6dQ6GJSKXsd2OrVwNSGqJDWChE+Wi/kfngZUDyxMO+25XPs2DGjJWNMtBVLLM3Q\n+BCKbxXjYutFTtzV1hUpESmI8YqhbVcjYhjgyhUgL499+nWCnx/w8MOAtzd/uRFC5h6aY3cfVNgR\nQ/B5DkbDaHCu+RyOKY5hVDWqjUvFUqwPWo/44HjIJDT91piGh9krwa5PunlNJAISEoBNmwDJPS7p\noDNTxBC0Xoi+zD7HrqurC4cPH0ZbWxt++ctform5GQzDIDAwcNZJEDJXKXoVyK/OR/tQOyce7RmN\nlIgUeNh78JSZ9aqpAb79lr3vdYKHB/DQQ4BAb0gkhBC96dWxO378OH7wgx9g5cqVOHnyJAYGBiCX\ny/G73/0OOTk55sjTYNSxI0LWP9aPo7VHcbXjKifuYe+BtIg0RHpG8pSZ9VIqgaNHgXPnuPEVK4CU\nFLoSjBDCL7NuxcbFxeG3v/0ttm7dCnd3d/T09GB0dBTBwcHo6OiYdRKmQIUdESKVRoXTTadRWl+K\ncfW4Nm4jtsGm0E1YG7gWUjHNizS25mZ2jMnEPa8A4OjIPvEaFcVfXoQQMsGsW7H19fXYunUrJ2Zj\nYwP15LkAhFgwc5yDqe6qxpGaI+ga6eLEF/kswrbwbXCxdTHp689FajVQVgaUlgKaSZdELFgAZGay\nxZ2h6MwUMQStF2JuehV2MTExOHLkCFJTU7Wx4uJiLF682GSJEWItuke6UVBTgMquSk7cx9EH6ZHp\nCHUL5ScxK9fVxXbpmidNjbG1BVJTgbi4O8eYEEKINdBrK/b06dPIyMhAeno6vv76a+zcuRM5OTn4\n9ttvsXr1anPkaTDaiiV8U6qVKGsow6nGU1BpVNq4ndQOm0M3Y1XAKohFlnObi6VgGOD8efY8nVKp\niwcHsw9ITNzzSgghQmL2cSfNzc04ePAg6uvrERwcjCeeeELQT8RSYUf4wjAMbnTeQEFNAfrG+jif\nW+63HEnzk+Aom8EeILmvgQHg0CGguloXk0iAzZuB9esBC7oVkRAyx/Ayx06j0aCzsxPe3t6CH5RK\nhR0xhLHOwdweuo38mnzU9dRx4gHOAUiPTEeAS8CsX4Pc3fXr7Gy64WFdzMeHHTY8b57xXofOTBFD\n0Hoh+jLrwxM9PT3453/+Z3z11VdQKpWwsbHBD3/4Q7z33nvw8BDunC26UoyYy6hqFMcVx3Gm+Qw0\njO6UvoONA7aGbcWyecsE/8uQpRodBfLzgYoKbnzdOiApib0ejBBChIqXK8UefPBBSKVS/OpXv0Jw\ncDAaGhrw5ptvYnx8HN9++63RkjEm6tgRc2AYBhXtFSiqK8Lg+KA2LoIIqwNWIzE0EfY29jxmaN0U\nCvae175JO96ursCDDwLz5/OWFiGEGMysW7Gurq5obW2Fg4ODNjY8PAw/Pz/09fXd4yv5Q4UdMbXW\ngVbkVeehsb+REw9xDUF6ZDp8nXx5ysz6qVRASQlQXs4+LDFh6VIgLQ2ws+MvN0IImQlj1S16HSVe\nsGABFAoFJ1ZfX48FCxbMOgFChMCQNviwchi5Vbn464W/coo6Z5kz/in2n/B03NNU1JlQezvw0UfA\nqVO6os7eHvjhD9mnXk1d1Blzy4RYP1ovxNz0On2yZcsWbNu2DU8++SSCgoLQ0NCAgwcPYufOnfj0\n00/BMAxEIhF27dpl6nwJ4Y2G0eBCywWU3CrBiGpEG5eIJFgXtA4bQzZCJqF7qUxFo2E7dCUl7ODh\nCeHh7NarszN/uRFCiFDotRU78fDB5MPfE8XcZMeOHTNudrNAW7HEmBr6GpBXnYe2wTZOPNIjEqkR\nqfB08OQps7mht5c9S1dfr4vZ2ADJycCqVTRsmBBi+XgZd2JJqLAjxjAwNoDCukJcab/CibvbuSM1\nIhVRnlH0tKsJMQz7tGt+PjA2posHBLDbrl5e/OVGCCHGZNZxJ4RYu6mzptQaNc40n4FcIce4elwb\ntxHbICEkAeuD1kMqpv98TGl4GMjJAW7c0MXEYmDjRiAhgR08zAeaS0YMQeuFmBv9ZCJkitruWuTX\n5KNzuJMTj/WOxbbwbXCzc+Mps7mjuhr49ltgUDdBBp6ebJdOwBfeEEII72grlpDv9Y72oqCmADc6\nb3Di3g7eSItMQ5h7GE+ZzR3j4+wdr+fPc+MrVwLbtgEyejaFEGKlaCuWECOorKlEwbkCVPVUQdGt\nQGhYKLz82YNbthJbJIYmYnXAakjEPO37zSFNTcB//zfQ3a2LOTkBDzwAREbylxchhFgSvTt2N27c\nwNdff4329nb8x3/8B27evInx8XEsWbLE1DnOCHXsyP1cqbyCdw+/izafNrRdbYPbAjeoalSIi43D\n1uVbsTVsK5xkTnynafXUaqC0FCgrY0eaTIiJATIzgUlz0QWBzkwRQ9B6Ifoy64Dir7/+Ghs3bkRz\nczMOHDgAABgYGMArr7wy6wQIMbeWgRYcqjyEN755AwoPBUZVo9rPucW6YZ5yHh5c8CAVdWbQ2Ql8\n8glw/LiuqLO1Zc/SPfKI8Io6QggROr06dgsWLMCXX36JuLg4uLu7o6enB0qlEn5+fujs7Lzfl/OC\nOnZksjHVGK52XMX5lvNoHWwFAJw+cRqjgWxRJxVLEeYeBj8nP7j///buPDrK8t4D+Hcmk8lkm+zb\nBEOAkJCwBJiIUhUCESkXFEkrBY8o0BYPLrfaVYuyFD3WU7X2lqqttQpY4nKOWkF6wQsEcQMzhKAJ\nEAgQloTsZLJOZnnvH28zk2EmMBMm72zfzzmck3neJ/P+Jn1Mf/m9z9IQh8eWPObNcAOeIADffCPO\npzOZbO0jR4pJXSzXpxBRkJF0jl1TU5PTR65yuUsFPyKvqe+oh65eh6MNR+22LQEAOeSIDI1EWnQa\nUqNSrduXKOWcoT+cOjqAjz4CampsbSEhQFERcPPN4pYmREQ0NC79Cp06dSq2bt1q1/buu+9i2rRp\nwxIU0fXoM/fhcP1h/E33N/xV91eU1ZXZJXUKuQL5Kfn41YJfYVL3JIxQj8CFoxcAAIaTBhRNLfJW\n6AGvshJ45RX7pC4lBVi1Cvje9/wjqePZn+QOjheSmksVuz//+c+YM2cO3njjDXR3d+OOO+5AdXU1\ndu/ePdzxXZf169ejsLCQE1eDxKXOS9DVidU5g9ngcD0xIhEFmgLkp+QjPDQcAKCJ0mDP4T1obm1G\ncmMyimYVIScrR+rQA15vL7BzJ3B0wAEeMpmYzM2aBSi4Pp+IglRpaalH/wBweVVsV1cXduzYgdra\nWmRkZGD+/PmI9uFTtznHLjj0mftQ2VgJXb0OF/QXHK4r5ArkJeVBm6ZFRkwGj//ygjNnxEev7e22\ntpgYcS5dZqbXwiIi8ik8K/YamNgFtsauRpTVleFow1G7Va39EsITxOpcaj4iQrm00htMJmDPHuCr\nr+zbJ08Gvv99QKXyTlxERL5I0sUTtbW12LBhA8rLy9E54IwfmUyG6urq6w6CyBVGsxGVTZXQ1elw\nXn/e4XqILAS5Sbko0BRgZMxIt6pz3GvKsy5dEjcbbmy0tUVEAAsWAHl53ovLEzhWyB0cLyQ1lxK7\ne+65B7m5udi4cSNU/DObJNbU1YSyujJUNFQMWp3TarTIT8lHpDLSCxFSP4sF+PJLYN8+cePhfmPH\nAnfdBfjw7A0iooDg0qPYmJgYtLa2IiTEf45V4qNY/2aymFDVVIWyujKcaz/ncD1EFoJxieNQoClA\nZmwm5875gLY24MMPgXMD/ucKDQXmzgW0WnGxBBEROSfpo9gFCxZg//79mD179nXfkOhqmruboavT\n4cilI+gx9Thcj1PFQavRYkrqFFbnfIQgAEeOAP/+N9A3YKvA9HSguBhISPBebEREwcalil1zczOm\nT5+O7OxsJCcn275ZJsM//vGPYQ1wqFix8x8miwnHmo6hrK4Mte21DtflMjnGJY6DNk2L0XGjh6U6\nx3kwQ9PVBWzfDhw/bmuTy4GZM4HbbvOPfencxbFC7uB4IVdJWrFbuXIllEolcnNzoVKprDfn4y+6\nHi3dLdDVi9W5bmO3w/VYVSy0aVpMSZvCc1t90IkTwMcfi8ldv4QEsUqXnu69uIiIgplLFbvo6Ghc\nvHgRarVaipg8ghU732S2mHGs+Rh0dTqcuXzG4bpcJkdOQg60Gi3GxI3hHw8+qK8P2LUL0Ons26dN\nA+bMEefVERGReySt2E2aNAktLS1+ldiRb2ntabXOnesydjlcjwmLsc6diw7j0klfdf68uI1JW5ut\nLToaWLgQyMryXlxERCRyKbGbPXs25s6dixUrViAlJQUArI9iV65cOawBkv8yW8w43nwcunodTred\ndrgugwzZCdko0BRgTPwYyGXem5DFeTBXZzYD+/cDBw6IiyX65eWJe9NFBNEe0Bwr5A6OF5KaS4nd\ngQMHoNFonJ4Ny8SOrtTW02adO9fZ1+lwXR2mxtS0qZiaNhXqMFaBfV1Tk1ilq6+3tYWFAfPnAxMn\nchsTIiJfwiPFyCPMFjOqW6pRVleGmrYah+syyDA2YSy0aVqMTRjr1eocuUYQgEOHgE8/FY8H6zdq\nFHD33eJ5r0RE5BnDPsdu4KpXi8Uy6BvIA3E/A3LZ5d7LOFx/GIfrDzutzkUro63VuRgVMwF/odcD\nH30EnB7wBD0kBLj9duDmm1mlIyLyVYMmdmq1Gh0dHWInhfNuMpkM5oHnBlFQsAgWW3WutQYC7P/C\nkEGGrPgsaDVaZCdk+0V1jvNgbL77DtixA+gdcHpbaqq4jcmAbSyDFscKuYPjhaQ2aGJXWVlp/fr0\naceJ7xR82nvbrdW5jr4Oh+tRyihrdS5WFeuFCOl69PQAO3cC335ra5PJgFtuAQoLgUH+viMiIh/i\n0hy7F154Ab/85S8d2l966SX8/Oc/H5bArhfn2HmGRbDgZMtJ6Op1ONly0ml1bkz8GGjTxOpciNx/\nzhMmm9OnxUever2tLTYWWLQIGDnSe3EREQULT+UtLm9Q3P9YdqC4uDi0DdzQyocwsbs+eoPeWp3T\nG/QO16OUUZiSOgVT06YiLjzOCxGSJxiNwJ49wNdf27dPmQJ8//vi6lciIhp+kmxQvHfvXgiCALPZ\njL1799pdq6mp4YbFAcYiWFDTWoOyujJUt1Q7VOcAYHTcaBRoCpCTkBNQ1blgnAdTXy9uY9LUZGuL\niADuvBPIzfVeXL4uGMcKDR3HC0ntqondypUrIZPJYDAY8OMf/9jaLpPJkJKSgj//+c/DHuCVDh06\nhMceewyhoaFIT0/Hli1bBl3cQa7pMHRYq3PthnaH65GhkZicOhlajRbx4fFeiJA8yWIBvvgC2LdP\n/LpfdjZw111AFI/lJSLyWy49il22bBm2bt0qRTzXdOnSJcTFxSEsLAy//e1vodVq8YMf/MChHx/F\nXp0gCKhps1XnLILjljajYkdBq9FiXOI4KORMngNBayvw4Yfi0WD9lEpg7lxg6lRuY0JE5C2SnhXr\nK0kdAKSmplq/Dg0NRUhI4DwOlEJnXyfK68uhq9fhcu9lh+sRoRFidS5Ni4SIBC9ESMNBEIDycuB/\n/xfo67O1jxghbmMSz0IsEVFA8NuTJ2pra7F06VIcOHDAaXLHip2NIAg43XYaunodjjcfd1qdy4zN\nhDZNi9yk3KCszgXyPJjOTmD7duDECVubXC5uYXLrreLX5LpAHivkeRwv5CpP5S2S/krftGkTCgoK\noFKpsGLFCrtrra2tWLRoEaKiopCZmYmSkhLrtT/+8Y+YNWsWXnzxRQCAXq/H/fffj82bN7NidxWd\nfZ34/Nzn+J+D/4OtR7eiqqnKLqkLV4Rj+ojpeGTaI1g+eTkmpkwMyqQukB0/Drz6qn1Sl5gI/OQn\nwIwZTOqIiAKNpBW7Dz/8EHK5HLt27UJPTw/efPNN67WlS5cCAN544w2Ul5dj/vz5+PLLL5GXl2f3\nHiaTCXfddRd++ctfYvbs2YPeK1grdoIg4MzlM9DVidU5s+B4MkhGTAYKNAXIS8pjIhegDAZg1y7g\n8GH79ptuEo8FCw31TlxEROScpPvYedrTTz+NCxcuWBO7rq4uxMfHo7KyEllZWQCABx54ABqNBs89\n95zd927duhWPP/44Jk6cCABYvXo1Fi9e7HCPYEvsuvq6cOTSEejqdWjtaXW4rlKokJ+SD61Gi+RI\nngsVyM6dExdIDNxiMjoauPtuYMwY78VFRESDk3TxhKddGXh1dTUUCoU1qQOA/Px8lJaWOnzvsmXL\nsGzZMpfus3z5cmRmZgIAYmNjMXnyZOtch/739ufXgiBg1JRRKKsrw87dO2GBBZmTxc979shZAMBt\nM25DgaYATZVNUFxUIHlsss/E70uvX375Zb8fH2YzIAiF+OIL4MwZ8XpmZiEmTACio0tx/jwwZozv\nxOuvrwf+XvKFePjat19zvPD1YK/7vz579iw8yScqdgcOHMDixYtRX19v7fP6669j27Zt2Ldv35Du\nEcgVu25jNyouVUBXr0Nzd7PDdZVChUkpk6BN0yIlKsULEfqf0tJS6390/qixUdxs+NIlW5tKBcyf\nD/ynuE0e4u9jhaTF8UKuCqiKXVRUFPR6+2Or2tvbER0dLWVYPk0QBJxrP4eyujJUNVU5nTs3Qj0C\n2jQtJiRPQGgIJ1G5w19/8QqCeBzYnj2AyWRrHz0aWLgQiInxXmyByl/HCnkHxwtJzSuJneyKXVCz\ns7NhMplw6tQp6+PYiooKTJgwwRvh+ZQeYw8qGiqgq9OhqbvJ4XpYSJhYndNokRqV6uQdKFC1twMf\nfQScOWNrUyjExRE33cTNhomIgpGkiZ3ZbIbRaITJZILZbIbBYIBCoUBkZCSKi4uxdu1a/P3vf8fh\nw4exfft2fPXVV9d1v/Xr16OwsNDv/mISBAHn9eehq9OhsqkSJovJoU96dDq0GrE6pwxReiHKwOJP\nj0sEAfjuO+CTT4DeXlt7Wpq42XBSkvdiCwb+NFbI+zhe6FpKS0vt5t1dL0nn2K1fvx6/+93vHNrW\nrl2LtrY2rFy5Ep9++ikSExPx+9//HkuWLBnyvfxxjl2PsQdHG45CV69DY1ejw3VliNI6dy4tOs0L\nEQYuf/nl29MD7NgBVFba2mQycaPhwkKA2zoOP38ZK+QbOF7IVX693YkU/CWxEwQBF/QXoKvXobKx\nEkaL0aFPWlQaCjQFmJA8AWGKMC9ESb6gpkZ89NrRYWuLiwMWLQIyMrwXFxERXT+/XjxBQK+pV6zO\n1enQ0NXgcF0ZosSE5Ako0BRAE63xQoTkK4xG4NNPgUOH7NunTgXmzgXCmOsTEdF/BHRi52tz7ARB\nQF1HHcrqyvBd43dOq3OpUako0BRgYvJEVuck5KuPS+rqxG1MmgfsahMZCdx1F5CT4724gpmvjhXy\nTRwvdC2enmMX8ImdLzCYDPi28VuU1ZXhUuclh+uh8lC76tyVq4Yp+FgswIEDwP794tf9cnLEpC4y\n0nuxERGR5/QXoDZs2OCR9+Mcu2FU11EHXZ0O3zZ+iz5zn8P1lMgUaDVaTEqZBJVC5YUIyRe1tIhH\ngl24YGtTKoHvfx+YMoXbmBARBSLOsfNRBpMB3zV+h7K6MtR31jtcV8gVmJA8Ado0LUaoR7A6R1aC\nAOh0wK5d4ry6fjfcIC6QiI/3XmxEROQfmNh5SH1HPXT1OhxtOOq0OpcUkYQCTQEmpUxCeGi4FyKk\nq/H2PJjOTuBf/wJOnrS1yeXArFnALbeIX5Nv8PZYIf/C8UJSC+jEbrgXT/SZ+/Bd43fQ1elwseOi\nw3WFXIHxSeOh1Whxg/oGVufIqWPHgO3bge5uW1tSkrjZcBq3KyQiCmh+vUGxlIZzjl1DZwPK6spw\ntOEoDGaDw/XEiEQUaAqQn5LP6hwNymAA/v1v4MgR+/abbwaKioBQHvdLRBQ0OMdOYkazUazO1etw\nQX/B4XqILAR5SXko0BQgIyaD1Tm6qtpacYHE5cu2NrUauPtuYPRo78VFRET+jYndNTR2NVqrc72m\nXofrCeEJ0Gq0mJw6GRGhEV6IkDxBqnkwJhOwbx/w5ZfiYol+EycC//VfQDgLvD6Pc6bIHRwvJDUm\ndk4YzUZUNVWhrK4M5/XnHa6HyEKQm5QLbZoWmbGZrM6RSxoaxM2GGwYcNKJSAQsWABMmeC8uIiIK\nHJxjN0BTVxN09TpUXKpAj6nH4Xp8eDy0aWJ1LlLJHWLJNYIAfPUVsGcPYDbb2seMARYuFB/BEhFR\ncOMcOxe4sirWZDFZq3Pn2s85XJfL5MhNzIVWo8Wo2FGszpFbLl8GPvoIOHvW1qZQAHPmANOmcbNh\nIqJgx1WxLrpW5tvc3QxdnQ5HLh1xWp2LU8VZ585FKaOGM1TyAZ6eByMIwNGjwM6d4urXfhqNuNlw\nUpLHbkUS45wpcgfHC7mKFbshMFlMONZ0DLp6Hc5ePutwXS6TIychBwWaAoyOG83qHA1JdzewYwdQ\nVWVrk8mAGTPEfyEh3ouNiIgCW1BU7Fq6W6CrF6tz3cZuh76xqlhMTZuKKalTEB0WLXWoFEBOnRJP\nkOjosLXFx4tVuhtu8F5cRETk21ixc8Gav69BdGo0eqMctymRy+TITsi2VufkMp7ZREPX1wd8+inw\nzTf27VotMHcuoFR6Jy4iIgouAZ3YfaH4AqavTZicNxmJmkQAQExYjFidS5sCdRiXI5LoeubBXLwo\nbmPS0mJri4wUV7xmZ3smPvIdnDNF7uB4IakFdGIHAIosBc6cPoNbJt4CrUaLrJFrcWIAABYGSURB\nVPgsVufII8xm4MAB4LPPAIvF1j5uHHDnnWJyR0REJKWAnmM3+u7RGKsdi5kjZ+LJZU96OyQKIC0t\nYpXu4kVbW1gYMG8ekJ/PbUyIiMg1/dudbNiwwSNz7AI6sVu7dy1kMhmSG5Px0OKHvB0SBQBBAMrK\ngN27AaPR1p6RIS6QiIvzXmxEROS/PLV4IqCfScpkMhhOGlA0tcjboZCPc2VzyI4O4J//BD75xJbU\nhYQAt98OLF/OpC5YeHIjUQp8HC8ktYCeY5fcmIyiWUXIycrxdijk56qqgO3bgZ4Be1knJwPFxUBq\nqvfiIiIiGiigH8UG6EcjCfX2Av/+N1BRYWuTyYDp04HZs8XjwYiIiK4X97EjGmZnzwIffgi0t9va\nYmKAu+8GRo3yWlhERESDCug5dkSuGjgPxmQSF0ds3myf1OXnA6tXM6kLdpwzRe7geCGpsWJHNEBD\ng7iNSUODrS08HFiwABg/3ntxERERuYJz7IggbjD81VfA3r3ixsP9srLEEySieYQwERENI86xc8H6\n9etRWFjI41xoUCdO1OLjj2tw+LAc7e0WjB49BomJIxEaCtxxB1BQwM2GiYho+PRvUOwprNhRUBIE\nYP/+Wrz22ik0NRWhpaUUsbGFMJn2YO7cLKxePRKJid6OknwRz/4kd3C8kKtYsSNyk8UCnD8v7kl3\n7Bjwf/9Xg+5u2+bVMhmQlVWE2Ni9SEwc6cVIiYiIhoaJHQU0iwWorbUlc52dA6/ZFoWnpRUiNxdQ\nqwGzmYvFaXCsvpA7OF5IakzsKOBYLOIedP3JXFeX834qlQVxcUBSEhAbC8j/k88plRbJYiUiIvIk\nzrGjgGA2A2fOiMnc8eNAd7fzfpGRQG4ukJcHGAy12LLlFMLCinD2bCkyMwthMOzB8uVZyMnho1hy\njnOmyB0cL+QqzrGjoGcyAadP25K53l7n/aKjbclcRoatMgeMxPLlwJ49e9HcfBTJyRYUFTGpIyIi\n/8WKHfkVkwk4dUpM5k6cAAwG5/3UajGRy8sDbriBW5YQEZFvY8WOgobRKCZzlZVAdTXQ1+e8X0yM\nmMiNHw+kpzOZIyKi4MPEjnxSXx9w8qRYmauuFpM7Z+LibJU5jWboyRznwZCrOFbIHRwvJDUmduQz\nDAYxiauqEit0gyVz8fFiVS4vD0hNZWWOiIioX0DPsVu3bh2PFPNxvb3iXLmqKqCmRpxD50xioi2Z\nS05mMkdERIGh/0ixDRs2eGSOXUAndgH60fxeT499Mmc2O++XnGybM5eUJG2MREREUuLiCfIr3d3i\nliRVVeIWJZZB9gBOTbXNmZPyrFbOgyFXcayQOzheSGpM7GjYdHWJJz9UVYknQQyWzGk0tmQuPl7S\nEImIiAIKH8WSR3V02JK52lpgsP8JRowQE7ncXHFlKxERUTDjo1jyGXq9LZk7d27wZO6GG2yVuZgY\naWMkIiIKBkzsaEguX7Ylc+fPO+8jk4lHePVX5tRqaWN0B+fBkKs4VsgdHC8kNSZ25LK2NjGRq6oC\nLl503kcmAzIzxWRu3DjxnFYiIiKSBufY0VW1tNiSufp6533kcjGZGz9eTOYiIyUNkYiIyO9xjh0N\nm+ZmMZGrrAQaGpz3kcuB0aNtlbmICGljJCIiIkdM7AiCADQ12SpzjY3O+4WEAGPGiMlcTg4QHi5t\nnMOJ82DIVRwr5A6OF5IaE7sgJQhiNa4/mWtudt5PoQCyssRkLjsbUKmkjZOIiIhcxzl2QUQQxHly\n/clca6vzfqGhwNixYjI3diwQFiZtnERERMGGc+zIJYIA1NXZkrm2Nuf9QkPFilx/MqdUShsnERER\nXT8mdgFIEIALF2zJXHu7835KpThXLi9PfNwaGiptnL6E82DIVRwr5A6OF5IaE7sAYbGIGwVXVYkb\nB+v1zvuFhYmrWPPyxIUQCo4AIiKigBHQc+zWrVuHwsLCgP1ryWIRz2PtT+Y6O533U6nEZG78eGDU\nKCZzREREvqK0tBSlpaXYsGGDR+bYBXRiF4gfzWIBzp61JXNdXc77RUTYKnOjRolblRAREZFv4uKJ\nIGI2A2fOiMnc8eNAd7fzfpGR4pmseXniSRByuaRh+jXOgyFXcayQOzheSGpM7HyUyQScPi0mcydO\nAD09zvtFRYmJXF4ekJHBZI6IiCiY8VGsDzGZgFOnbMmcweC8n1otVubGjwdGjGAyR0RE5O/4KDZA\nGI32yVxfn/N+MTG2ytyIEYBMJm2cRERE5PuY2HlBXx9w8qSYzFVXi8mdM3FxtmROo2EyN5w4D4Zc\nxbFC7uB4IakxsZOIwSAmcVVVYoVusGQuPl58xJqXB6SmMpkjIiIi13GO3TDq7RUfr1ZVATU14hw6\nZxITbZW5lBQmc0RERMGGc+x8VE+PfTJnNjvvl5xsS+aSkpjMERER0fVjYucB3d3i/nJVVeIWJRaL\n836pqWIil5srJnPkOzgPhlzFsULu4HghqTGxG6KuLvHkh6oq8SSIwZK5tDRxzlxuLpCQIGmIRERE\nFGQ4x84NHR22ZK62Fhjs7dPTbY9Z4+I8GgIREREFIM6xk4heb0vmzp0bPJm74QbbY9bYWGljJCIi\nIgKY2DnV3i4mclVVwPnzzvvIZOIRXv3JnFotbYzkWZwHQ67iWCF3cLyQ1JjY/Udbmy2Zu3jReR+Z\nDMjMFJO5ceOA6GhJQyQiIiK6qqCeY9fSYkvm6uud95HL7ZO5qCjPx0pERETBjXPshqi52ZbMXbrk\nvI9cDowebUvmIiKkjZGIiIhoKAI+sRMEoKnJlsw1NjrvFxICjBkjJnM5OUB4uLRxkndxHgy5imOF\n3MHxQlIL6MTuv/97L9TqMVAoRjq9rlCIydz48UB2NqBSSRwgERERkQf53Ry7hoYGFBcXQ6lUQqlU\nYtu2bUhwsvOvTCbDzJkCTKY9mDw5C4mJYnKnUABjx4qVuexsICxM6k9AREREZM9Tc+z8LrGzWCyQ\ny+UAgM2bN6O+vh5PPPGEQ7/+xA4A1Oq9WLZsNvLyxKROqZQ0ZCIiIqKr8lRiJ/dALJLqT+oAQK/X\nI+4qRzskJ4uPWWfOlOOee8SvmdSRM6Wlpd4OgfwExwq5g+OFpOZ3iR0AVFRU4KabbsKmTZuwdOnS\nQfvl5QFJSUB4+CAHuRL9x5EjR7wdAvkJjhVyB8cLSU3SxG7Tpk0oKCiASqXCihUr7K61trZi0aJF\niIqKQmZmJkpKSqzX/vjHP2LWrFl48cUXAQD5+fk4ePAgnnnmGWzcuPGq9zQY9qCoaIznPwwFlMuX\nL3s7BPITHCvkDo4XkpqkiV16ejqefvpprFy50uHaww8/DJVKhcbGRvzzn//E6tWrUVVVBQB4/PHH\nsW/fPvziF7+A0Wi0fo9arYbBYBj0fsnJe7F8eRZycpyvivUFUpTpPXGPob6HO9/nSt9r9bna9UB4\nJDLcn8FT7z+U9/H0WHGlXyCPF/5uca9vMI8VgL9b3O3ry+NF0sRu0aJFWLhwocMq1q6uLnzwwQfY\nuHEjIiIicMstt2DhwoXYunWrw3scOXIEM2fOxOzZs/HSSy/h17/+9aD3e+ih2T6d1AH85etu3+H6\nj+ns2bPXvLcv4C9f9/oOx3jhWPHsPfi7xTfwd4t7fX05sfPKqtinnnoKFy9exJtvvgkAKC8vx623\n3oquri5rn5deegmlpaX4+OOPh3SPrKws1NTUeCReIiIiouE0ZswYnDp16rrfxysbFMtkMrvXnZ2d\nUKvVdm3R0dHo6OgY8j088cMhIiIi8ideWRV7ZZEwKioKer3erq29vR3R0dFShkVERETk17yS2F1Z\nscvOzobJZLKrslVUVGDChAlSh0ZERETktyRN7MxmM3p7e2EymWA2m2EwGGA2mxEZGYni4mKsXbsW\n3d3d+Pzzz7F9+3YsW7ZMyvCIiIiI/JqkiV3/qtfnn38eb7/9NsLDw/Hss88CAF555RX09PQgOTkZ\n9913H1577TXk5uZKGR4RERGRX/O7s2Kvh16vx+23345jx47h4MGDyMvL83ZI5MMOHTqExx57DKGh\noUhPT8eWLVugUHhlvRH5uIaGBhQXF0OpVEKpVGLbtm0O2zoRXamkpAQ/+9nP0NjY6O1QyEedPXsW\nN954IyZMmACZTIb33nsPiYmJV/0evzxSbKgiIiKwc+dO/PCHP/TIQbsU2DIyMrBv3z7s378fmZmZ\n+Ne//uXtkMhHJSUl4YsvvsC+fftw77334vXXX/d2SOTjzGYz3n//fWRkZHg7FPJxhYWF2LdvH/bu\n3XvNpA4IssROoVC49EMhAoDU1FSEhYUBAEJDQxESEuLliMhXyeW2X6V6vR5xcXFejIb8QUlJCRYv\nXuywmJDoSl988QVmzJiBNWvWuNQ/qBI7oqGora3Fp59+ijvvvNPboZAPq6iowE033YRNmzZh6dKl\n3g6HfFh/te5HP/qRt0MhH6fRaFBTU4PPPvsMjY2N+OCDD675PX6Z2G3atAkFBQVQqVRYsWKF3bXW\n1lYsWrQIUVFRyMzMRElJidP34F9JweN6xoter8f999+PzZs3s2IXBK5nrOTn5+PgwYN45plnsHHj\nRinDJi8Z6nh5++23Wa0LMkMdK0qlEuHh4QCA4uJiVFRUXPNefjkTPD09HU8//TR27dqFnp4eu2sP\nP/wwVCoVGhsbUV5ejvnz5yM/P99hoQTn2AWPoY4Xk8mEJUuWYN26dRg7dqyXoicpDXWsGI1GhIaG\nAgDUajUMBoM3wieJDXW8HDt2DOXl5Xj77bdx8uRJPPbYY3j55Ze99ClICkMdK52dnYiKigIAfPbZ\nZxg/fvy1byb4saeeekpYvny59XVnZ6egVCqFkydPWtvuv/9+4YknnrC+njdvnqDRaITp06cLb731\nlqTxkne5O162bNkiJCQkCIWFhUJhYaHw7rvvSh4zeYe7Y+XgwYPCjBkzhFmzZgl33HGHcP78eclj\nJu8Zyv8X9bvxxhsliZF8g7tjZefOnYJWqxVuu+024YEHHhDMZvM17+GXFbt+whVVt+rqaigUCmRl\nZVnb8vPzUVpaan29c+dOqcIjH+PueFm2bBk3yQ5S7o6VadOmYf/+/VKGSD5kKP9f1O/QoUPDHR75\nEHfHyrx58zBv3jy37uGXc+z6XTk/obOzE2q12q4tOjoaHR0dUoZFPorjhVzFsULu4HghV0kxVvw6\nsbsy842KioJer7dra29vR3R0tJRhkY/ieCFXcayQOzheyFVSjBW/TuyuzHyzs7NhMplw6tQpa1tF\nRQUmTJggdWjkgzheyFUcK+QOjhdylRRjxS8TO7PZjN7eXphMJpjNZhgMBpjNZkRGRqK4uBhr165F\nd3c3Pv/8c2zfvp3zpIIcxwu5imOF3MHxQq6SdKx4aqWHlNatWyfIZDK7fxs2bBAEQRBaW1uFu+++\nW4iMjBRGjhwplJSUeDla8jaOF3IVxwq5g+OFXCXlWJEJAjd0IyIiIgoEfvkoloiIiIgcMbEjIiIi\nChBM7IiIiIgCBBM7IiIiogDBxI6IiIgoQDCxIyIiIgoQTOyIiIiIAgQTOyIiIqIAwcSOiOgKy5cv\nx9NPP+3R91y9ejWeeeYZj74nEdGVFN4OgIjI18hkMofDuq/Xq6++6tH3IyJyhhU7IiIneNoiEfkj\nJnZE5FOef/55jBgxAmq1GuPGjcPevXsBAIcOHcL06dMRFxcHjUaDRx99FEaj0fp9crkcr776KsaO\nHQu1Wo21a9eipqYG06dPR2xsLJYsWWLtX1paihEjRuC5555DUlISRo0ahW3btg0a044dOzB58mTE\nxcXhlltuwbfffjto38cffxwpKSmIiYnBpEmTUFVVBcD+8e6dd96J6Oho67+QkBBs2bIFAHD8+HHM\nmTMHCQkJGDduHN5///1B71VYWIi1a9fi1ltvhVqtxty5c9HS0uLiT5qIAhETOyLyGSdOnMBf/vIX\nlJWVQa/XY/fu3cjMzAQAKBQK/OlPf0JLSwu++uor7NmzB6+88ord9+/evRvl5eX4+uuv8fzzz+On\nP/0pSkpKcO7cOXz77bcoKSmx9m1oaEBLSwvq6uqwefNmrFq1CidPnnSIqby8HD/+8Y/x+uuvo7W1\nFQ8++CDuuusu9PX1OfTdtWsXDhw4gJMnT6K9vR3vv/8+4uPjAdg/3t2+fTs6OjrQ0dGB9957D2lp\naSgqKkJXVxfmzJmD++67D01NTXjnnXfw0EMP4dixY4P+zEpKSvDWW2+hsbERfX19eOGFF9z+uRNR\n4GBiR0Q+IyQkBAaDAZWVlTAajcjIyMDo0aMBAFOnTsW0adMgl8sxcuRIrFq1Cvv377f7/l//+teI\niopCXl4eJk6ciHnz5iEzMxNqtRrz5s1DeXm5Xf+NGzciNDQUM2bMwPz58/Huu+9ar/UnYX/729/w\n4IMP4sYbb4RMJsP999+PsLAwfP311w7xK5VKdHR04NixY7BYLMjJyUFqaqr1+pWPd6urq7F8+XK8\n9957SE9Px44dOzBq1Cg88MADkMvlmDx5MoqLiwet2slkMqxYsQJZWVlQqVRYvHgxjhw54sZPnIgC\nDRM7IvIZWVlZePnll7F+/XqkpKRg6dKlqK+vByAmQQsWLEBaWhpiYmKwZs0ah8eOKSkp1q/Dw8Pt\nXqtUKnR2dlpfx8XFITw83Pp65MiR1nsNVFtbixdffBFxcXHWfxcuXHDad9asWXjkkUfw8MMPIyUl\nBQ8++CA6Ojqcftb29nYsXLgQzz77LL73ve9Z73Xw4EG7e23btg0NDQ2D/swGJo7h4eF2n5GIgg8T\nOyLyKUuXLsWBAwdQW1sLmUyG3/zmNwDE7ULy8vJw6tQptLe349lnn4XFYnH5fa9c5drW1obu7m7r\n69raWmg0Gofvy8jIwJo1a9DW1mb919nZiR/96EdO7/Poo4+irKwMVVVVqK6uxh/+8AeHPhaLBffe\ney+Kiorwk5/8xO5eM2fOtLtXR0cH/vKXv7j8OYkouDGxIyKfUV1djb1798JgMCAsLAwqlQohISEA\ngM7OTkRHRyMiIgLHjx93afuQgY8+na1yXbduHYxGIw4cOIBPPvkE99xzj7Vvf/+f/vSneO2113Do\n0CEIgoCuri588sknTitjZWVlOHjwIIxGIyIiIuziH3j/NWvWoLu7Gy+//LLd9y9YsADV1dV4++23\nYTQaYTQa8c033+D48eMufUYiIiZ2ROQzDAYDnnzySSQlJSEtLQ3Nzc147rnnAAAvvPACtm3bBrVa\njVWrVmHJkiV2VThn+85deX3g69TUVOsK22XLluGvf/0rsrOzHfpqtVq8/vrreOSRRxAfH4+xY8da\nV7BeSa/XY9WqVYiPj0dmZiYSExPxq1/9yuE933nnHesj1/6VsSUlJYiKisLu3bvxzjvvID09HWlp\naXjyySedLtRw5TMSUfCRCfxzj4iCTGlpKZYtW4bz5897OxQiIo9ixY6IiIgoQDCxI6KgxEeWRBSI\n+CiWiIiIKECwYkdEREQUIJjYEREREQUIJnZEREREAYKJHREREVGAYGJHREREFCD+HxiurmiM2mt9\nAAAAAElFTkSuQmCC\n", + "text": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## `statistics.mean()` vs. `numpy.mean()`" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "# The statistics module has been added to\n", + "# the standard library in Python 3.4\n", + "\n", + "import timeit\n", + "import statistics as stats\n", + "import numpy as np\n", + "\n", + "def calc_mean(samples):\n", + " return sum(samples)/len(samples)\n", + "\n", + "def np_mean(samples):\n", + " return np.mean(samples)\n", + "\n", + "def np_mean_ary(np_array):\n", + " return np.mean(np_array)\n", + "\n", + "def st_mean(samples):\n", + " return stats.mean(samples)\n", + "\n", + "n = 1000000\n", + "samples = list(range(n))\n", + "samples_array = np.arange(n)\n", + "\n", + "assert(st_mean(samples) == np_mean(samples)\n", + " == calc_mean(samples) == np_mean_ary(samples_array))\n", + "\n", + "%timeit(calc_mean(samples))\n", + "%timeit(np_mean(samples))\n", + "%timeit(np_mean_ary(samples_array))\n", + "%timeit(st_mean(samples))" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "100 loops, best of 3: 26.2 ms per loop\n", + "1 loops, best of 3: 144 ms per loop" + ] + }, + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "\n", + "100 loops, best of 3: 3.21 ms per loop" + ] + }, + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "\n", + "1 loops, best of 3: 1.12 s per loop" + ] + }, + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "\n" + ] + } + ], + "prompt_number": 2 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "funcs = ['st_mean', 'np_mean', 'calc_mean', 'np_mean_ary']\n", + "orders_n = [10**n for n in range(1, 6)]\n", + "times_n = {f:[] for f in funcs}\n", + "\n", + "for n in orders_n:\n", + " samples = list(range(n))\n", + " for f in funcs:\n", + " if f == 'np_mean_ary':\n", + " samples = np.arange(n)\n", + " times_n[f].append(min(timeit.Timer('%s(samples)' %f, \n", + " 'from __main__ import %s, samples' %f)\n", + " .repeat(repeat=3, number=1000)))" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 3 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%pylab inline" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 6 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "import matplotlib.pyplot as plt\n", + "\n", + "labels = [('st_mean', 'statistics.mean()'), \n", + " ('np_mean', 'numpy.mean() on list'),\n", + " ('np_mean_ary', 'numpy.mean() on array'),\n", + " ('calc_mean', 'sum(samples)/len(samples)')\n", + " ]\n", + "\n", + "matplotlib.rcParams.update({'font.size': 12})\n", + "\n", + "fig = plt.figure(figsize=(10,8))\n", + "for lb in labels:\n", + " plt.plot(orders_n, times_n[lb[0]], \n", + " alpha=0.5, label=lb[1], marker='o', lw=3)\n", + "plt.xlabel('sample size n')\n", + "plt.ylabel('time per computation in milliseconds [ms]')\n", + "plt.legend(loc=2)\n", + "plt.grid()\n", + "plt.xscale('log')\n", + "plt.yscale('log')\n", + "plt.title('Performance of different approaches for calculating sample means')\n", + "\n", + "max_perf = max( s/c for s,c in zip(times_n['st_mean'],\n", + " times_n['np_mean_ary']) )\n", + "min_perf = min( s/c for s,c in zip(times_n['st_mean'],\n", + " times_n['np_mean_ary']) )\n", + "\n", + "ftext = 'using numpy.mean() on np.arrays is {:.2f}x to '\\\n", + " '{:.2f}x faster than statistics.mean() on lists'\\\n", + " .format(min_perf, max_perf)\n", + "plt.figtext(.14,.15, ftext, fontsize=11, ha='left')\n", + "\n", + "plt.show()" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "display_data", + "png": "iVBORw0KGgoAAAANSUhEUgAAAnYAAAIECAYAAACUvmMzAAAABHNCSVQICAgIfAhkiAAAAAlwSFlz\nAAALEgAACxIB0t1+/AAAIABJREFUeJzsnXlcjen//1/30X46aU+UdocKWSNUlOzMpBmhUXYxGHuT\n1omGMdbiYyamIs2iGcPMd6ytwky0GELSggzJ0oJou35/9Oue7s6pzmkOkev5eJxH3e/7ut7X+762\n8z7XdjOEEAIKhUKhUCgUyjsPr70NoFAoFAqFQqHIBurYUSgUCoVCoXQQqGNHoVAoFAqF0kGgjh2F\nQqFQKBRKB4E6dhQKhUKhUCgdBOrYUSgUCoVCoXQQqGPXAampqcGcOXOgra0NHo+HlJSU9jbpneTw\n4cMwMzODnJwc5syZI3G8oKAgWFhYNHvdnO6kpCRYW1tDQUEBo0aNks1DUGQKj8dDbGxsu6T9+eef\nQ09PDzweDwcOHGgXG1rCy8sLo0ePlqlOcW2nrbRn2b1NODo6Yv78+e1tBuU1Qh27dsLLyws8Hg88\nHg/y8vIwNjaGt7c3njx58p91//zzz/j+++/x+++/48GDBxg6dKgMLH6/qK2txZw5c+Du7o67d+9i\n586dbda1Zs0a/PXXX63q9vb2xsCBA1FQUIBffvnlPz+DLDA3N0dwcHB7m/He89dff2Hz5s3Yv38/\nHjx4gI8//ri9TRKBYRgwDPNa9EqDs7MzZs+eLSJ/8OABpk6dKiuz3lleVzlR3h7k2tuA9xl7e3v8\n9NNPqKmpwaVLlzB//nzcvXsXv//+e5v0VVVVQUFBAbm5uejWrRuGDBnyn+xr0Pc+8s8//+D58+cY\nN24c9PX1/5MuPp8PPp/fom5CCG7duoX169ejW7dubU6LEILa2lrIycmmab8LXwDvQz3Nzc0Fj8fD\nxIkT/5Oe15lXhBC8jvPuZaVTV1dXJnoolLcdOmLXjsjLy0NXVxddu3bF5MmTsXz5cpw4cQKvXr0C\nAPzwww+wsbGBsrIyTExMsGrVKrx48YKN7+joiHnz5sHf3x9du3aFkZERRo4ciYCAAOTn54PH48HU\n1BQAUF1dDR8fHxgYGEBRURFWVlb4/vvvOfbweDyEhYVhxowZUFdXx6xZsxAVFQV5eXkkJSWhd+/e\nUFFRwahRo/DgwQMkJibCxsYGqqqqGD16NP755x9WV0FBAVxdXdGtWzfw+Xz06dMHMTExnPQapgRC\nQkKgr68PLS0teHp64vnz55xwP/74IwYMGABlZWVoa2tj/PjxKC0tZe+HhYWhZ8+eUFZWRo8ePRAa\nGora2toW8/7PP/+Evb09VFRUoKmpiZkzZ6KkpAQAEBUVBSMjIwD1zndL09kvX76Et7c31NXVoamp\nicWLF7Pl10Dj6aSmujt16oTk5GR06tQJtbW1mDVrFmeq7datW5g6dSo0NDSgqamJMWPG4OrVq6zu\nxuXTr18/KCkpIT4+HtXV1QgKCoKpqSmUlZVhbW2Nb7/9VqS8//e//+GTTz6BmpoaDA0NsWnTJk75\n5OXlITg4mB1dvnPnjth8yMjIwLhx46CnpweBQIDBgwfj5MmTnDDGxsbw8/PDvHnz0LlzZ+jo6GD9\n+vWcL25Jw/j7+2Px4sXQ1taGg4MDAOCPP/7AgAEDoKSkBD09PSxZsoTTXiSxsaamBsHBwTAzM4OS\nkhIMDAywbNkyTpiysrJm8wyARHm/b98+9OrVC8rKytDS0oKDgwPu3bsnNm+9vLwwa9Ys1NXVgcfj\noVOnTgDqHZ6vv/4apqamUFRUhLm5ucjIcnN5JY709HSMHTsWnTt3hkAggK2tLdLS0gBI1p7F0VLb\nFTcluGHDBpiYmDSrrzU7vLy8kJCQgOjoaLbONrRdHo+HQ4cOsWFbq/8A8PjxY3z00UdQVVWFvr4+\nvvjiC4mmnENDQ9k6pKuri7Fjx+Lly5cS52VD3+7n5wddXV1oaGggICAAhBAEBgaiS5cu0NXVhZ+f\nHyeeJO1HHNL2oUlJSeDxeDh+/DiGDh0KFRUVDBo0CNevX8fff/+NYcOGgc/nw9bWFtevX+fETU9P\nh4uLCwQCAXR1dTF16lROvyKr747s7GyMGTMGGhoaUFVVhaWlpUR1tkNAKO2Cp6cnGT16NEe2detW\nwjAMefbsGYmMjCQaGhokJiaGFBQUkJSUFNKnTx/yySefsOEdHByIQCAg3t7e5Pr16+Tq1avkyZMn\nZPXq1cTExIQUFxeTR48eEUIIWb16NdHS0iJxcXEkNzeXhIaGEh6PR+Lj41l9DMMQLS0tsnv3bpKf\nn09yc3NJZGQk4fF4ZOTIkSQtLY1kZGQQCwsLMnz4cGJvb0/++usvkpWVRXr27EmmTZvG6rpy5QrZ\nvXs3+fvvv0l+fj4JCwsjcnJyJDExkWO/uro6WblyJcnJySGnTp0impqaxN/fnw3z3XffEXl5ebJh\nwwb2GcPDw9nnCgwMJEZGRuTXX38lhYWF5I8//iDdu3fn6GjK/fv3iUAgIDNnziRXr14lqamppE+f\nPsTe3p4QQkhlZSW5ePEiYRiG/Pbbb6S4uJhUVVWJ1fXZZ58RXV1dcuzYMZKTk0NWr15N1NTUiIWF\nBRsmMDCQvW5O94MHDwjDMGTPnj2kuLiYVFZWkgcPHhA9PT2yePFicvXqVXLz5k2ydOlSoqWlRUpK\nSgghhC0fW1tbkpSURAoKCkhJSQnx9PQkffv2JadPnyaFhYXkxx9/JOrq6mT//v2c8tbT0yP79u0j\n+fn5ZPfu3YRhGLZOPHnyhJiYmJA1a9aQ4uJiUlxcTGpra8XmQ1JSEomOjibXrl0jubm5xM/Pjygo\nKJCbN2+yYYyMjIiamhoJDAwkN2/eJAcPHiR8Pp/s3LmzTWGCg4NJbm4uuX79Orl8+TLp1KkTW5eO\nHz9Ounfvzmkvktg4a9YsoqurS2JiYkh+fj65ePEiJ+3W8owQ0mreX7p0icjJyZGDBw+SO3fukCtX\nrpD9+/eToqIisXlbVlZGdu7cSeTk5NhyIISQ8PBwoqysTCIiIsitW7fI3r17iZKSEqeMxeWVOK5e\nvUpUVFTIjBkzSHp6OsnLyyM//fQTuXDhAiFEsvbs6elJnJ2d2evW2q6joyOZP38+x46QkBBibGzM\nXgcGBhJzc3P2ujU7ysrKiL29PXF3d2fzqqHtMgxDDh06JFVZTpo0iQiFQpKUlESys7PJ7Nmzibq6\nukjf3Ziff/6ZqKmpkd9//53cvXuXZGVlkZ07d5LKykqJ89LBwYF07tyZ+Pj4kNzcXPLdd98RhmHI\nmDFjyLp160hubi6Jjo4mDMOQ48ePi5R3S+2nab63pQ9NTEwkDMOQ/v37k8TERHLt2jUydOhQ0qdP\nHzJs2DCSkJBArl+/ToYPH05sbW3ZeNnZ2URVVZUEBQWRnJwccvXqVfLRRx+RHj16kJcvX0qVP619\nd/Tu3ZvMnDmTXL9+nRQUFJDjx4+T33//vdln6khQx66daNoJZmdnE1NTUzJ06FBCSH0D/eabbzhx\nkpOTCcMwpLS0lBBSX7mFQqGI7qad4fPnz4mioiL53//+xwn34YcfklGjRrHXDMOQefPmccJERkYS\nhmHI5cuXWdmWLVsIwzAkIyODlW3fvp1oa2u3+MxTpkzhdCgODg7ExsaGE8bb25vNA0IIMTQ0JEuX\nLhWr7/nz50RFRYWcPHmSI4+Ojibq6urN2uHn50cMDQ1JdXU1K7t8+TJhGIakpKQQQggpKCggDMOQ\nc+fONavn2bNnRElJiezbt48jHzhwoIhj17g8mtPd9IsnMDCQDBkyhBOmrq6OmJmZkR07dhBC/i2f\n1NRUNkx+fj7h8XgkJyeHEzc4OJiT3wzDkOXLl3PC9OrVi3z++efstbm5OQkODm42D1qib9++ZOPG\njey1kZER6zw34OvrSwwNDaUO07jtEEKIh4cH5wuEEEKOHj1KeDweuXPnjkQ25ubmEoZhyM8//9xs\n+NbyTJK8/+WXX0jnzp1JeXl5s+k0JTIyksjJyXFkBgYGZN26dRzZihUriKmpKXstLq/E4eHhIdIW\nW6Npe27ap7XUdglpm2MniR3Ozs5k9uzZIuHEOXYtleXNmzcJwzAkISGBvV9dXU0MDQ1bdOy2bdtG\nevTowelfWkNc39ivXz9OGCsrK9KnTx+OrG/fvmT16tXstSTtp3G+t7UPbXDsjh49ysoOHz5MGIYh\nv/zyCys7cuQIYRiGPH/+nBBSX0fc3d05ul6+fElUVFTIr7/+2mx6bfnu6Ny5M4mKimpWZ0eGTsW2\nI0lJSRAIBFBRUUHv3r1hbm6OQ4cOoaSkBHfu3MGKFSsgEAjYz/jx48EwDG7dusXqGDBgQKvp3Lp1\nC1VVVbC3t+fI7e3tkZ2dzZENHjxYJD7DMOjduzd7raenBwDo06cPR/b48WN2yP/Fixfw8fGBtbU1\ntLS0IBAI8Mcff3CG3BmGQd++fTlp6evro7i4GADw8OFDFBUVwcXFRexzZWdno7KyEq6urpx8WrRo\nEcrLy/H48eNm4w0ZMoSzDq1Pnz7o3Lkzrl27JjaOOPLy8vDq1SvY2dlx5MOGDZPJuqCLFy8iPT2d\n82xqamq4ffs2pw4AwKBBg9j/L126BEIIBgwYwIn75ZdfisSzsbHhXHft2hUPHz6U2taSkhIsXrwY\nvXr1goaGBgQCAbKzs0XKu+lGHjs7OxQVFeHZs2dShWlaT69duya2fhNC2DJtzcaMjAwAaLa+NdBS\nnkmS9y4uLjA1NYWJiQmmT5+OiIiIZutqc5SXl+PevXtin7mwsJCd9hOXV+JIT0+Hk5NTs/clac+N\naa3tthVp7WiNlsqyod40XqssJyeHgQMHtqhz2rRpqK6uhpGREWbPno2YmBi27kr6DOL6xi5dunD6\n3AZZ4/YqSftpTFv70AYa29jc9wIA1saLFy/iyJEjnLS0tbXx6tUrtn3I4rsDAFavXo158+Zh5MiR\nCA4ORmZmZovP0pGgmyfakSFDhiA6OhpycnLo2rUr62g0VM5du3Zh5MiRIvEaFtczDMNZlC8LxOnj\n8XicRfQN/zes9WksI4SAYRisWbMGx44dw/bt2yEUCqGiooJVq1ahrKyMo7vpQm6GYVBXVyeRrQ3h\n4uLi0KNHD5H7GhoaYuMxDPNaFnnLGkIInJ2dER4eLnKvc+fO7P+dOnXi5GNDvly4cAEqKiqceE03\nQ/yX/G+Ml5cXioqKsGXLFpiYmEBJSQnu7u6oqqqSWpckiKunrZWprGwUt/mgIc8kyXs+n49Lly7h\n3LlzOHPmDPbu3Yu1a9ciPj4e/fv3l8oWSZCkj2itTUjanqWBx+OJpFldXd1iHFnb0VJZNtC0zbRW\nz7p27YobN24gMTERCQkJCAkJwbp16/DXX3/BwMBA4meQl5cXsaOpTBJ7WqKtfag4GxvySZysIR1C\nCGbNmgUfHx8RXVpaWgAkL+PW+i4/Pz/MnDkTJ06cQEJCAkJDQ7F27VqEhIS0+EwdAerYtSNKSkrs\n5obG6OnpwdDQEDdu3MDcuXP/czrm5uZQVFREcnIyLC0tWXlycjJnJE6WnD17Fh4eHnBzcwNQ37Bz\ncnKk2mGqq6sLAwMDnDx5UuxuQCsrKygpKSEvLw9jx46VWK+VlRUiIyNRXV3NdkKXL19GWVkZrK2t\nJdZjZmYGBQUFnDt3Dr169WLl586dk8lu0oEDByIqKgrdunWDoqKixPEaRnFv376NCRMm/CcbFBQU\nWt2IAtSX95YtW9hyev78OfLy8jj1ixCCCxcucOKdP38eBgYGUFVVlTiMOKysrEQ2uCQnJ4NhGFhZ\nWUlkY4NTdfLkSamOxWhc1pLmPY/Hw4gRIzBixAgEBwfD0tISsbGxEjt2ampqMDAwQHJyMsaPH895\nZlNTUygpKUlsf4Pd8fHx7A+zpkjbnltruw1hmm4YycjIaLHtSGKHgoICampqWn7gZmicdkNfef78\nefZcyZqaGqSnp6Nnz54t6lFQUMCYMWMwZswYhISEQE9PD0ePHsWSJUuQkpLyn/tGcfYC0reftvah\nbWXgwIG4fPmy2O+9BmTx3dGAiYkJvL294e3tjU2bNuHrr7+mjt3bSHFxMVxdXaGgoAAFBQXExsay\nnn5HYuPGjZg7dy40NDQwefJkyMvL4/r16zhx4gT27t0LQPLjBVRUVLBs2TL4+/tDR0cHffr0QVxc\nHI4dO4YzZ868FvuFQiF+/fVXuLq6gs/nY9u2bbh//z66dOnChpHE/sDAQHh7e0NPTw9Tp05FXV0d\nEhMTMX36dGhpacHX1xe+vr5gGAZOTk6oqanBlStXkJWVJbLDrYFPP/0UO3fuhJeXF3x9ffH06VMs\nXrwY9vb2GDZsmMTPyOfzsWjRIvj5+UFPTw89evTA/v37cfPmTZkcrfDpp59i//79mDJlCvz8/GBg\nYICioiIcP34cEydObPZ8QnNzc8yZMwfz58/HV199hSFDhuD58+dIT0/Ho0ePsHbt2mbTbFomJiYm\nSE1Nxd27d9kdnOK+eIVCIWJiYjBs2DDU1NQgICAAdXV1IuWblZWF4OBgTJ8+HZcuXcKuXbuwYcMG\nqcKIqzNr1qxB//79sXLlSixYsACFhYVYunQpPDw8YGBgIJGN5ubmmDlzJhYvXoyXL19iyJAhePLk\nCS5cuCCyM7a5PJMk748ePYqCggKMGDECOjo6SE9Px927d1kHVFI+//xzrFq1ChYWFnBwcEBCQgL2\n7t2LPXv2tJhX4li7di1sbW0xc+ZMrFq1Curq6sjIyIChoSGGDBkiUXtuSmtt19nZGd7e3oiLi4ON\njQ3i4uKQmpoKdXX1ZnVKYoeJiQkSExORn58PNTU1qKurS3z8T+OytLCwwKRJk7BkyRJ888030NbW\nxtatW1FeXt6i87l//34QQjBo0CCoq6sjPj4eFRUVrKPYs2fPNvWNksokaT8NcVRVVdvUh7YVX19f\nDB48GB4eHli+fDm0tbVRWFiIo0ePYvny5TAxMZHJd8ezZ8+wbt06uLm5wdjYGKWlpThx4oTUbexd\n5Z1bY6ejo4Nz584hMTERM2bMQERERHub1CZaOyTSw8MDP/30E37//XfY2tpi8ODBCA4OZr+kWtIh\nTr5x40bMnz8fn332GXr37o3Y2FgcOnRI7FSvOH3SyrZv384ev+Ls7AxDQ0O4ubmJTOk21dNUNnfu\nXERFRSEuLg79+vWDg4MDTp48yXbUfn5+2LZtGyIiImBjY4MRI0Zg586dLR6ZoKuri1OnTqGoqAiD\nBg3CpEmTWGe3tWdsyqZNm/DBBx/gk08+ga2tLcrLy7FkyRKJnrM1dHV1ceHCBWhra8PV1RU9e/aE\nh4cH7t69i65du7ao69tvv8WKFSuwceNGWFlZwdnZGQcPHoSZmVmLaTa1NTg4GKWlpRAKhdDT08Pd\nu3fFxouMjERdXR0GDx4MV1dXjB8/HoMGDRLJh2XLluH27dsYNGgQli9fjqVLl3KcJknDNKV37944\nduwYUlJSYGNjg1mzZmHSpEnsjyBJbYyMjMTChQvh5+cHS0tLuLq6orCwUKo8ay3vNTU18dtvv2Hc\nuHEQCoXw8fGBv7+/2EN1m6bTGG9vb3zxxRcIDQ2FlZUVtmzZgs2bN3P0SDpybG1tjaSkJJSUlMDB\nwQH9+vXD9u3b2XbWlvbcWtv19PTEkiVLsGTJEgwaNAj37t3DsmXLWtQpiR2rVq2CtrY2+vbtC11d\nXZw/f16iPBCXXmRkJKytrTFu3DiMGjUKBgYGcHFxaXFEVFNTE5GRkRg5ciQsLS2xY8cOREREsH2t\nrPpGcTJJ20/jOG3pQxv0SCvr2bMnzp8/j2fPnmHMmDGwsrLCggUL8PLlS9ahl0X+yMvLo7S0FHPn\nzoWlpSXGjh0LfX399+bNIwx5FxYbNUNYWBgUFBSwcOHC9jaFQqG0gomJCebPnw9fX9//FIZCaS9q\na2vRs2dPfPDBB9iyZUt7myMCbT8U4B2cigXq10MtWLAApaWluHjxYnubQ6FQJECS35Dv8O9MSgfk\n7NmzKC4uRr9+/VBRUYHt27fjzp078PLyam/TxELbDwVox6nY8PBwDBw4EEpKSiJTEE+ePMGHH34I\nVVVVGBsbi7whoW/fvvjrr7+wYcOG92IhJIXSEZBkWlAWm04oFFlRW1uLjRs3wsbGBqNGjUJhYSES\nExPf2rVatP1QgHYcsevWrRv8/f1x8uRJVFZWcu4tWbIESkpKePjwITIzMzFhwgT07dsXlpaWnJ2M\nampqIq9volAobycFBQUyCUOhvCkcHR3fqfPPaPuhAG/BGjt/f38UFRUhMjISQP0RBJqamsjOzoa5\nuTmA+kW2Xbt2xZdffom0tDSsWbMGnTp1gry8PPbv38/ZUNBAt27dOO8upVAoFAqFQnlbMTMzEzlE\nvi20+67Ypn7lzZs3IScnxzp1QP3Ua8MbEgYPHozk5GQkJCTg5MmTYp06APjnn3/YLdFv8ycwMPCd\nSKOtOqSJJ0nY1sK0dL+t996mz+u2U1b626JH1nVFknBtqRO0rsg2Ddq3vB0f2rdIF/Z11Je8vDyZ\n+FXt7tg1XRPw7NkzqKmpcWQCgQAVFRVv0qw3hqOj4zuRRlt1SBNPkrCthWnpfkv3WjvS4m3hddcX\nWelvix5Z1xVJwrWlvtC6Its0aN/ydkD7FunCvq76IgvafSrWz88P9+7dY6diMzMzMXz4cDx//pwN\n8/XXXyMlJQXHjh2TWC/DMAgMDISjo+Mb6eAo7zZeXl6IiopqbzMo7wC0rlCkgdYXSmskJSUhKSkJ\nwcHBkIVL9taN2PXo0QM1NTWceebLly9L9aqnBoKCgqhTR5GIt/X4AsrbB60rFGmg9YXSGo6OjggK\nCpKZvnZz7Gpra/Hy5UvU1NSgtrYWr169Qm1tLfh8PlxdXREQEIAXL14gNTUVv/32Gz755JP2MpXy\nHkB/AFAkhdYVijTQ+kJ507SbYxcSEgIVFRVs3rwZMTExUFZWxsaNGwEAe/bsQWVlJXR1deHh4YG9\ne/dyXrJOociapKSk9jaB8o5A6wpFGmh9obxp2u0cu6CgoGaHHjU0NHDkyBGZpEHX2FEoFAqFQnlb\naVhjJyvaffPE64JhmGYXIWpqauLp06dv2CIKhfIm0dDQwJMnT9rbDAqFQpGIlvwWqfS8j46drDKP\nQqG8vdB2TqFQ3iVk1We1+67Y10lQUBBd30ChUGQK7VMo0kDrC6U1kpKSZLorlo7YUSiUDsnraudJ\nSUl03S5FYmh9oUgKnYptBerYUSjvN7SdUyiUdwk6FUuhUCgUCoVC4UAdO0qL8Hg8xMbG/icdXl5e\nGD16tIwsenfJzMxEly5d8OLFCwBAamoqjI2N8erVq3a2jCINdM0URRpofaG8aTq0Y/e+bZ4oKioC\nj8dDSkqK1HGdnZ0xe/ZsEfmDBw8wdepUiXTExMSAxxOtUmFhYYiLi5Papo7G2rVrsXLlSqioqAAA\nhg8fDnNzc4SHh7ezZRQKhUJpL+jmCQn5r2vscnJu48yZPFRX8yAvXwdnZzMIhUZttkfW+sRRVFSE\n7t27IzExEQ4ODlLFdXZ2hqGhISIjI9ucfkxMDGbNmoW6uro26+ioZGdno1+/frh37x50dHRYeWxs\nLNavX4/8/HyR9yZT/ht0jR2FQnmXoGvsXiM5ObcRFXULJSWjUFrqiJKSUYiKuoWcnNtvhb7U1FQM\nGzYMampqUFNTg42NDU6dOoXu3bsDAEaOHAkejwdTU1MAQEFBAVxdXdGtWzfw+Xz06dMHMTExrD4v\nLy8kJCQgOjoaPB6PM+rXdCp237596NWrF5SVlaGlpQUHBwfcu3cPSUlJmDVrFhuHx+Nhzpw5rP6m\nU7E//vgjBgwYAGVlZWhra2P8+PEoLS1t8fmao0F/WFgYDAwMIBAIsGjRItTW1iI8PBxGRkbQ1NTE\nwoULUV1dzYkbFhaGnj17QllZGT169EBoaChqa2vZ+7GxsbC1tYW6ujp0dHQwceJE5ObmsvcLCwvB\n4/Fw+PBhTJw4EXw+H2ZmZoiOjuakExMTAzs7O45TBwCTJk3C3bt3kZqa2uzzUSgUCoUiKe32SrG3\nmTNn8qCo6ATuLK4T/v47AYMGST/KlpaWhxcvnDgyR0cnxMcnSD1qV1NTg8mTJ2POnDk4cOAAAODq\n1atQUVFBRkYG+vfvj19++QV2dnbo1KkTAOD58+dwdnZGcHAwVFVV8X//93+YPXs2DAwM4OjoiF27\ndqGgoABdu3bFzp07AdSf2t+U9PR0eHt7IzIyEg4ODigrK0NaWhoAYNiwYQgPD8enn36KBw8eAACU\nlZXZuI1HoyIjI7Fw4UIEBgbi0KFDqK2tRVJSEurq6lp8vpbzOA0GBgaIj49Hbm4uPvroIxQWFqJL\nly44deoU8vLy4Obmhn79+mHRokUA6qfqo6KisHPnTtjY2ODatWtYtGgRXr58iS+++AIAUFVVhYCA\nAFhaWqK8vBwBAQGYMGECsrOzIS8vz6bv4+ODzZs3Y9euXdi/fz/mzZsHOzs7WFhYAACSk5MxYsQI\nEbsFAgGsrKyQkJAg9j7l7YMeX0GRBlpfKG8a6tiJobpa/EBmbW3bBjjr6sTHq6qSXl9FRQVKS0sx\nadIkmJmZAQD7t6ioCED9K9N0dXXZONbW1rC2tmavP/30U5w5cwaxsbFwdHSEmpoaFBQUoKyszInX\nlDt37oDP52PKlCkQCAQwNDTk6FVTUwMAsToaDy8HBgZi0aJFWL9+PSuzsrICADx9+rTZ52sJZWVl\nREREQE5ODkKhEE5OTkhLS8O9e/cgLy8PoVAIFxcXxMfHY9GiRXjx4gW2bNmCI0eOwMXFBQBgZGSE\nkJAQLF++nHXsvLy8OOlERkZCW1sbly5dwtChQ1n50qVL4ebmBgAICQlBWFgYEhMTWccuNzcXM2fO\nFGu7sbExbt682eozUigUCoXSGh16Kratmyfk5cWvEevUqW1rx3g88fEUFKTXp6GhgXnz5mHMmDEY\nP348Nm/e3KpT8OLFC/j4+MDa2hpaWloQCAT4448/cOfOHanSdnFxgampKUxMTDB9+nRERETg8ePH\nUul4+PCh91h8AAAgAElEQVQhioqKWGeqKW15PgDo1asX5OT+/Z2ip6cHoVDIGVXT09PDw4cPAdSv\neausrISrqysEAgH7WbRoEcrLy9nnysrKwocffghTU1OoqanByKh+hPX2be40uo2NDfs/j8eDrq4u\nmxYAlJWVQSAQiLVdIBCw09CUtx86+kKRBlpfKK0h680THXrErq0Z5exshqioeDg6/jt9+upVPLy8\nzCEUSq8vJ6den6IiV5+Tk3mb7Pv222+xfPlynDp1CqdPn4a/vz/Cw8Mxfvx4seHXrFmDY8eOYfv2\n7RAKhVBRUcGqVatQVlYmVbp8Ph+XLl3CuXPncObMGezduxdr165FfHw8+vfv36ZnEUdzz7dgwYJm\n4zR26oD6qV9xsoaNHQ1/4+Li0KNHDxF9GhoaePHiBVxcXGBvb4+oqCjo6emBEAIrKytUVVVxwiso\nKDSbFgCoq6ujoqJCrO1lZWVip74pFAqF0vFxdHSEo6MjgoODZaKvQ4/YtRWh0AheXubQ1U2AunoS\ndHUT/r9T17ZdrLLWB9RPXa5YsQJ//PEH5s6di2+//RaKiooAwFn8DwBnz56Fh4cH3Nzc0Lt3b5iY\nmCAnJ4ez7k1BQQE1NTWtpsvj8TBixAgEBwcjPT0d+vr6+P7771kdAFrc1aOrqwsDAwOcPHlS6udr\nCWl3lFpZWUFJSQl5eXkwNTUV+fB4PFy/fh2PHj3Cxo0bYW9vD6FQiCdPnrRp15KFhQUKCwvF3rt9\n+7ZY55LydvI+HaFE+e/Q+kJ503ToEbv/glBoJNPjSGSlLy8vD99++y0mT54MAwMD/PPPP0hJScHA\ngQOhra0NVVVVnDx5Er169YKioiI0NDQgFArx66+/wtXVFXw+H9u2bcP9+/fRpUsXVq+JiQkSExOR\nn58PNTU1qKuri4x4HT16FAUFBRgxYgR0dHSQnp6Ou3fvwtLSktXREG7YsGFQUVEBn88XeYbAwEB4\ne3tDT08PU6dORV1dHZKSkuDu7o7S0lKR5zt79iwGDBjAxndycoKtrS1CQ0NZmbTOlqqqKnx9feHr\n6wuGYeDk5ISamhpcuXIFWVlZ2LRpE4yMjKCoqIhdu3Zh5cqVKCwshI+Pj0ROZFN7HBwccO7cOZFw\nFRUVuHbtGp2uoVAoFIpMoCN27xh8Ph+3bt2Cu7s7hEIh3NzcMHz4cISHh4NhGOzevRs//fQTDA0N\nWWdo+/btMDIywsiRI9nz6tzc3DgOyqpVq6CtrY2+fftCV1cX58+fF0lbU1MTv/32G8aNGwehUAgf\nHx/4+/uzBxsPGjQIy5cvx8KFC6Gnp4elS5cCqB9Na5zW3LlzERUVhbi4OPTr1w8ODg44ceIE5OTk\nxD5fw47bBvLz89mdt+L0Syrz8/PDtm3bEBERARsbG4wYMQI7d+5kHVRtbW3ExMTg9OnTsLa2xtq1\na7F161aRQ5jFOXpNZR4eHrhw4QJKSko48mPHjsHQ0BD29vYiOihvJ9QJp0gDrS+UNw09oJhCeUO4\nuLjAyckJ69atY2VOTk4YN24cVq9e3Y6WdUxoO6dQKO8S9IBiCXjfXilGebv56quvsGPHDs67YvPz\n87Fs2bJ2towiDbRPoUgDrS+U1qCvFJMQOmJHobzfvK52Tg+cpUgDrS8USZFVn0UdOwqF0iGh7ZxC\nobxL0KlYCoVCoVAoFAoH6thRKBSKFNA1UxRpoPWF8qahjh2FQqFQKBRKB4GusaNQKB0S2s4pFMq7\nBF1jR6FQKBQKhULh0KEdO3qOHYVCkTW0T6FIA60vlNaQ9Tl2Hd6xo+cHUWRFZmYmunTpwjlg2NjY\nGK9evWo3m4KCgmBhYcFeR0VFQV5evt3soVAoFIp0ODo6UseOQmkP1q5di5UrV0JFRQUAMHz4cJib\nm3PeY9seNH4vrbu7O/755x+J4zo7O7Pv+qVIBv2xSJEGWl8obxrq2FEoEpCdnY3k5GQRJ2jOnDkI\nDw9v10X6jdNWUlKCjo5Ou9lCoVAolPaFOnbNkHMrB7t/3I0dP+zA7h93I+dWzlujz9HREfPnz0dI\nSAj09fWhpaUFT09PPH/+nA3j5eWF0aNHc+LFxMSAx/u3yBum8Q4fPgxzc3Pw+XxMnToVz549w+HD\nhyEUCqGmpoaPPvoI5eXlIrq3b9+Obt26gc/n4+OPP8bTp08B1K8XkJOTQ1FRESf9AwcOQF1dHZWV\nlRx5g76wsDAYGBhAIBBg0aJFqK2tRXh4OIyMjKCpqYmFCxeiurqaEzcsLAw9e/aEsrIyevTogdDQ\nUNTW1rL3Y2NjYWtrC3V1dejo6GDixInIzc1l7xcWFoLH4+Hw4cOYOHEi+Hw+zMzMEB0dLZJ3dnZ2\nIk7TpEmTcPfuXaSmpjZfYABycnIwYcIECAQCCAQCTJ48GXl5eez9hinU8+fPo3///uDz+Rg4cCAu\nXbrUot6mNJ2KLS8vx+zZs6Gvrw8lJSV0794dq1atAlCf7wkJCYiOjgaPxwOPx0NKSopU6b2P0DVT\nFGmg9YXypqGOnRhybuUgKjEKJXolKO1SihK9EkQlRrXZGZO1PgCIi4tDaWkpkpOT8cMPP+D333/H\n5s2b2fsMw3Cm6Jrj/v37OHDgAH799VccP34cZ8+ehaurK6KiohAXF8fKQkNDOfHS0tKQnJyMU6dO\n4Y8//kBWVhbmzp0LoN7xtLCwwHfffceJExERgZkzZ0JZWVnEjrS0NGRkZCA+Ph7ff/89oqOjMWHC\nBFy6dAmnTp1CTEwMDh48iP3797NxgoKCsHXrVmzevBk3btzAzp078c033yA4OJgNU1VVhYCAAGRm\nZuLMmTPo1KkTJkyYIOIg+vj4wMvLC1euXIG7uzvmzZvHcQCTk5Nha2srYrdAIICVlRUSEhKazePK\nykq4uLigqqoKKSkpSE5OxrNnzzB27FiOHXV1dfD19UVYWBgyMjKgq6uLjz/+mOOoSoufnx8yMzNx\n7Ngx3Lp1Cz/++CMsLS0BALt27cKIESMwbdo0PHjwAA8ePMDQoUPbnBaFQqFQ2h+59jbgbeRM+hko\nWigiqTDpX6E88PcPf2PQ8EFS60tLTcMLgxdA4b8yRwtHxGfEQ2gubJONxsbG2Lp1KwCgR48emDZt\nGs6cOYMvvvgCQP30nCTTg69evUJ0dDQ0NTUBAB9//DH27t2L4uJiaGlpAahftxUfH8+JRwjBwYMH\nIRAIAAC7d+/GmDFjkJ+fD1NTUyxYsAA7d+6Ev78/GIbBjRs3cO7cuWbXoykrKyMiIgJycnIQCoVw\ncnJCWloa7t27B3l5eQiFQri4uCA+Ph6LFi3CixcvsGXLFhw5cgQuLi4AACMjI4SEhGD58uVsPnh5\neXHSiYyMhLa2Ni5dusRxYpYuXQo3NzcAQEhICMLCwpCYmMhuTMjNzcXMmTObLYubN282m8exsbF4\n9OgRMjMz2Xz+4YcfYGxsjB9++AGffPIJm6c7duyAjY0NgHrHdciQIcjPz+dskJCGO3fuoF+/fhg0\nqL7eGhgYsM+tpqYGBQUFKCsrQ1dXt03630fomimKNND6QnnT0BE7MVSTarHyWrRt5KQOdWLlVXVV\nbdLHMAz69u3Lkenr66O4uFhqXd26dWOdDQDQ09NDly5dWKeuQfbw4UNOPEtLS9apAwA7OzsAwLVr\n1wAAs2bNwsOHD3Hy5EkAwL59+zBw4EARuxvo1asX5OT+/Z2hp6cHoVDImVZsbEd2djYqKyvh6urK\nTm82TOGWl5fj8ePHAICsrCx8+OGHMDU1hZqaGoyMjAAAt2/f5qTf4EwBAI/Hg66uLueZy8rKOM/b\nGIFAgNLSUrH3Gmy1srLi5LOuri6EQiGbX4Bouerr6wNAm8q1gcWLFyMuLg69e/fGZ599hhMnTtBD\neykUCqUDQx07Mcgz4o+L6IRObdLHayabFXgKbdIHAAoK3LgMw6Cu7l8HksfjiXyBN51+BCByNAbD\nMGJljXUDaNU50NLSgpubGyIiIlBdXY0DBw5gwYIFzYZv7NQ1pClO1mBHw9+4uDhcvnyZ/Vy9ehW5\nubnQ0NDAixcv4OLigk6dOiEqKgoXL17ExYsXwTAMqqq4TnVr+amuro6KigqxtpeVlUFDQ6PF/BCX\nX01lPB6PM33e8H/TvJcGFxcX3LlzB+vXr8fLly/h4eGBUaNG/Sed7zt0zRRFGmh9obxp6FSsGJwH\nOCMqMQqOFo6s7FXuK3i5e7Vp6jTHoH6NnaKFIkef00gnWZgrFj09Pfz5558cWUZGhsz0X79+HRUV\nFewo1vnz5wGAXb8FAAsXLsTIkSOxd+9evHz5EtOnT29WnyTrARtjZWUFJSUl5OXlYezYsc3a+OjR\nI2zcuBFCoZC1sy0jVhYWFigsLBR77/bt2+w0rjisra3xzTff4PHjx+xIaHFxMW7evIk1a9ZIbYu0\naGhowN3dHe7u7pg9ezaGDh2K69evw8rKCgoKCqipqXntNlAoFArlzUBH7MQgNBfCa6QXdB/qQv2B\nOnQf6sJrZNucutehT5L1c87Ozrhx4wb27NmDvLw8RERE4PDhw21KTxwMw2DWrFnIzs5GSkoKlixZ\ngilTpsDU1JQNM2zYMAiFQqxZswbTp08Hn88HADg5OcHX11fkmaRBVVUVvr6+8PX1xZ49e5CTk4Ps\n7Gz88MMP8PHxAVC/5k5RURG7du1CXl4e4uPjsXz5comcyKb2ODg4IC0tTSRcRUUFrl271uI6mhkz\nZkBHRwfTpk1DZmYm0tPT4e7uDgMDA0ybNk2q55aW9evX48iRI8jJyUFubi5iYmIgEAjQvXt3AICJ\niQnS09ORn5+PR48eUSdPAuiaKYo00PpCedN06BG7hjdPtKVhCc2FbXa8Xrc+cTtem8qcnJywYcMG\nhIaGYt26dZg8eTICAgKwdOlSqfQ0Jxs8eDCGDx+O0aNHo6ysDOPHj8e3334rYuu8efOwYsUKzjRs\nfn4+u9btv9jh5+cHfX19hIeHY9WqVVBWVoZQKGQ3TGhrayMmJgaff/45vvvuO1haWmL79u1wcnIS\n0duUpjIPDw98/fXXKCkp4Rx5cuzYMRgaGsLe3l5ERwNKSko4deoUVqxYwYYbOXIkTpw4wZlulsQO\ncffF5VMDysrKCAgIQGFhITp16oR+/frh+PHj7EjrqlWrcOXKFfTt2xcvXrxAYmJii89CoVAoFNmS\nlJQk0yl7hnTQldQMwzQ7CtTSPUrreHl54d69ezh9+nSrYdeuXYv4+Hikp6e/ActeLy4uLnBycsK6\ndetYmZOTE8aNG4fVq1e3o2UUcbyudp6UlERHYSgSQ+sLRVJk1WfRqVjKa6GsrAwXL15EREQEVqxY\n0d7myISvvvoKO3bs4LwrNj8/H8uWLWtnyygUCoVCqYeO2FGkZvbs2bh37x5OnTrVbBhHR0ekpaVh\n+vTpnEOFKZQ3BW3nFArlXUJWfRZ17CgUSoeEtnMKhfIuQadiKRQKpR2g55JRpIHWF8qbhjp2FAqF\nQqFQKB0EOhVLoVA6JLSdUyiUdwk6FUuhUCgUCoVC4UAdOwqFQpECumaKIg20vlDeNNSxo1AoFAqF\nQukg0DV2FAqlQ0LbOYVCeZega+wolDdMZmYmunTpwnnzhLGxMV69etXOllEoFAqFUs876dilpaXB\nzs4ODg4OmDFjBmpqatrbJMp7wNq1a7Fy5UqoqKgAAIYPHw5zc3OEh4e3s2WUNwldM0WRBlpfKG8a\nufY2oC10794diYmJUFRUhK+vL44ePYqpU6fKNI3bOTnIO3MGvOpq1MnLw8zZGUZC4Vujj/Jmyc7O\nRnJyMmJjYznyOXPmYP369Vi5ciUYhmkn66SjqqoKCgoKIvKamhrIyb2TXQKFQqG8s+Tk3MaZM3ky\n0/dOjth16dIFioqKAAB5eXl06tRJpvpv5+TgVlQURpWUwLG0FKNKSnArKgq3c3LeCn2Ojo6YP38+\nQkJCoK+vDy0tLXh6euL58+dsGC8vL4wePZoTLyYmBjzev0UeFBQECwsLHD58GObm5uDz+Zg6dSqe\nPXuGw4cPQygUQk1NDR999BHKy8tFdG/fvh3dunUDn8/Hxx9/jKdPnwKo/4UqJyeHoqIiTvoHDhyA\nuro6KisrOfIGfWFhYTAwMIBAIMCiRYtQW1uL8PBwGBkZQVNTEwsXLkR1dTUnblhYGHr27AllZWX0\n6NEDoaGhqK2tZe/HxsbC1tYW6urq0NHRwcSJE5Gbm8veLywsBI/Hw+HDhzFx4kTw+XyYmZkhOjpa\nJO/s7Oygo6PDkU+aNAl3795Fampq8wUGICcnBxMmTIBAIIBAIMDkyZORl/dvQ46KioK8vDzOnz+P\n/v37g8/nY+DAgbh06VKLejMyMjBu3Djo6elBIBBg8ODBOHnyJCeMsbEx/P39sXjxYmhra8PBwQHJ\nycng8Xj4448/MHz4cCgrK2P//v0oLS2Fh4cHjIyMoKKigp49e2Lbtm2sLmnLtiPi6OjY3iZQ3iFo\nfaG0RE7ObURE3MLFi6NkpvOd/nl++/ZtnD59GgEBATLVm3fmDJwUFYFGQ+hOABL+/htGgwZJry8t\nDU7/f10Wq8/REQnx8W0etYuLi8OcOXOQnJyM27dvw93dHUZGRvjiiy8A1C/ClGQE6f79+zhw4AB+\n/fVXPHnyBG5ubnB1dYW8vDzi4uJQXl6OqVOnIjQ0FJs2bWLjpaWlgc/n49SpU3j06BHmz5+PuXPn\n4pdffoGjoyMsLCzw3XffccomIiICM2fOhLKysogdaWlpMDAwQHx8PHJzc/HRRx+hsLAQXbp0walT\np5CXlwc3Nzf069cPixYtAlDvmEZFRWHnzp2wsbHBtWvXsGjRIrx8+ZLNh6qqKgQEBMDS0hLl5eUI\nCAjAhAkTkJ2dDXl5eTZ9Hx8fbN68Gbt27cL+/fsxb9482NnZwcLCAgCQnJyMESNGiNgtEAhgZWWF\nhIQEsfcBoLKyEi4uLujRowdSUlJACMHq1asxduxYXLt2jbWjrq4Ovr6+CAsLg7a2NlasWIGPP/4Y\nubm5zf54qaiowPTp07Ft2zbIy8sjOjoakydPxtWrV1nbAWDXrl1YtWoV/vzzT9TU1ODBgwcAgFWr\nVuHrr7+GtbU15OTk8OrVK/Tu3RurV6+GhoYGUlNTsWjRImhqasLLy6tNZUuhUCgUUZ48AXbsyENO\njhPq6mSomLQjYWFhZMCAAURRUZF4eXlx7j1+/Jh88MEHhM/nEyMjIxIbG8u5X1ZWRuzt7cnNmzfF\n6m7p0Vp77MTt2wkJDCTEwYHzSRwzpl4u5SdxzBgRXSQwsD6dNuDg4EBsbGw4Mm9vbzJ06FD22tPT\nkzg7O3PCHDx4kDAMw14HBgYSOTk58vjxY1a2ZMkS0qlTJ/Lo0SNWtnz5cjJw4ECOboFAQMrLy1nZ\nqVOnCMMwJC8vjxBCyLZt24iRkRGpq6sjhBBy/fp1wjAMycrKEnkeT09PoqenR6qrq1nZhAkTiI6O\nDqmqqmJlU6ZMIW5uboQQQp4/f05UVFTIyZMnObqio6OJurq6SBoNPH78mDAMQ86fP08IIaSgoIAw\nDEO2NyqL2tpaIhAIyDfffMPKtLW1SXh4uFidkydPJjNmzGg2zX379hEVFRVOPhcXFxNlZWVy4MAB\nQgghkZGRhGEYkpmZyYb566+/CMMwzdbx5ujbty/ZuHEje21kZCRSFxITEwnDMCQmJqZVfcuWLSOj\nR49mr6Up2/bkdXVviYmJr0UvpWNC6wulKXfuEPLDD4QEBREyZkwi6xbIqs9q16nYbt26wd/fH3Pm\nzBG5t2TJEigpKeHhw4c4dOgQvL29ce3aNQD1a4Hc3d0RGBjIGZWQFXWNRnI48jZO+dbxxGdznZh1\nTpLAMAz69u3Lkenr66O4uFhqXd26dYOmpiZ7raenhy5dukBLS4sje/jwISeepaUlBAIBe21nZwcA\nbBnNmjULDx8+ZKcF9+3bh4EDB4rY3UCvXr0467v09PQgFAo5o2qN7cjOzkZlZSVcXV3Z6c2GKdzy\n8nI8fvwYAJCVlYUPP/wQpqamUFNTg5GREYD60d7G2NjYsP/zeDzo6upynrmsrIzzvI0RCAQoLS0V\ne6/BVisrK04+6+rqQigUsvkFiJarvr4+ALRYriUlJVi8eDF69eoFDQ0NCAQCZGdn486dOxy9gwcP\nFhu/qbyurg6bNm2CjY0NdHR0IBAI8M0333D0eXp6SlW2FAqF8r5TVwdcvw7s31//uX4dIATg8eqH\n6lRVZZdWu07FfvjhhwCAS5cucdbsPH/+HL/88guys7OhoqKCYcOGYcqUKTh48CC+/PJLfP/990hL\nS0NISAhCQkLg7e2Njz/+WES/l5cXjI2NAQDq6uqwsbGRaL2DmbMz4qOi4NQobPyrVzD38gLaMHVq\nlpNTr+//rwtk9Tk5Sa2rgaaL3xmGQV2jsVwejydyHk7T9WkAOI5Tgx5xsrom48RNdTdFS0sLbm5u\niIiIgJOTEw4cOIDQ0NBmwzddtM8wjFhZgx0Nf+Pi4tCjRw8RfRoaGnjx4gVcXFxgb2+PqKgo6Onp\ngRACKysrVFVVccK3lp/q6uqoqKgQa3tZWRk0NDSafTZAfH41lfF4PM70ecP/TfO+MV5eXigqKsKW\nLVtgYmICJSUluLu7izwfn88XG7+pfOvWrdi0aRN27NiBfv36QSAQYNu2bfi///s/NoympqZUZfs2\n0LAzsaH9/5drR0dHmeqj1x37mtaX9/u6uhrYvz8J2dmAllb9/cLC+vsAUF19DPfvb4KSUlfIDJmM\n+/1H1q9fz5mKzcjIICoqKpwwW7duJZMmTZJYZ0uPJsljF964QeJ37yaJ27eT+N27SeGNGxKn/br1\nOTo6kvnz53NkISEhxNjYmL328fEhvXr14oT59NNPRaZizc3NW9RDCCFffvklMTAwYK8lmYolhJDU\n1FQiLy9Pdu3aRQQCAXn27JnY5/H09ORM9RFCyNy5c4mjoyNHtnDhQjJ8+HBCCCEVFRVEWVm52elR\nQgi5dOkSYRiG3GiU1+fOnSMMw5Do6GhCyL9TsefOnePENTc3J8HBwey1nZ0dWb16tdh0evfuzQnb\nlP379xMVFRXO9PaDBw+IsrIyOxUaGRlJ5OTkOPHu3r1LGIYhycnJzeoWCARk79697PWzZ8+IhoYG\nmT17NiszNjbmTM0S8u9U7L179zjyiRMnEnd3d45s9OjRxMTEhCOTtGzbk7eke6NQKO8hFRWEJCQQ\nsnmz6AqtL74g5NdfCSkurg9740Yh2b07XmZ91luxeaLpIv9nz55BTU2NIxMIBM2OmLwOjIRCmR5H\nIkt9hJBWR8ycnZ2xefNm7NmzB2PGjEFCQgIOHz4sk/SB+jKbNWsWNmzYgMePH2PJkiWYMmUKTE1N\n2TDDhg2DUCjEmjVr4OnpyY4OOTk5wdbWljPK09rzNEVVVRW+vr7w9fUFwzBwcnJCTU0Nrly5gqys\nLGzatAlGRkZQVFTErl27sHLlShQWFsLHx0eiTSVN7XFwcMC5c+dEwlVUVODatWvsLzVxzJgxA198\n8QWmTZuGLVu2oK6uDqtXr4aBgQGmTZsm1XM3RSgUIiYmBsOGDUNNTQ0CAgJQV1fHsV+avO3ZsycO\nHjyIpKQkdO3aFQcOHEBaWhpnGhlovmzfB5KSklosbwqlMbS+vF88egRcuABcvgw0PWJXWRkYOBAY\nPBhovLJHKDSCUGiEJUtkY8NbcdxJ0y8eVVVVzvEaQMtrnJojKCiIHRLtSIjb8dpU5uTkhA0bNiA0\nNBQ2NjZISkpCQECAyFRfa3qakw0ePBjDhw/H6NGjMW7cOPTt2xffffediK3z5s1DVVUVFixYwMry\n8/PZXZn/xQ4/Pz9s27YNERERsLGxwYgRI7Bz506YmJgAALS1tRETE4PTp0/D2toaa9euxdatWzlH\nvjTobUpTmYeHBy5cuICSkhKO/NixYzA0NIS9vb2IjgaUlJRw6tQpKCoqwt7eHo6OjhAIBDhx4gRn\nulkSO5oSGRmJuro6DB48GK6urhg/fjwGDRokdkpXEt3+/v5wcHDAlClTYGdnh7KyMixbtkxsfHFl\nS6FQKO8bhACFhUBsLBAeDqSnc506dXVg3DhgxQrAyYnr1AH1zn9QUJDM7Hkr3hXr7++PoqIiREZG\nAqhfY6epqYns7GyYm5sDAD755BMYGhpKvJaHviv29eHl5YV79+7h9OnTrYZdu3Yt4uPjkZ6e/gYs\ne724uLjAyckJ69atY2VOTk4YN24cVq9e3Y6WtQ9ve9nSdk6hUF4ndXXAtWvA+fPAP/+I3u/WDbCz\nA3r1AngSDKPJqs9q16nY2tpaVFdXo6amBrW1tXj16hXk5OTA5/Ph6uqKgIAA7Nu3DxkZGfjtt99w\n4cKF9jSXIgVlZWW4efMmIiIiEBYW1t7myISvvvoK48aNw9KlS6GiooLU1FTk5+c3O6LVUemIZUuh\nUCiS8uoVkJkJ/PknIO5ABKGw3qHr3h1ojxcStetUbEhICFRUVLB582bExMRAWVkZGzduBADs2bMH\nlZWV0NXVhYeHB/bu3YtevXpJpb+jTsW2N5IcfjxlyhQ4ODjA1dUVHh4eb8iy14uNjQ3u37/PeVds\nQUGB2NdzdWQ6YtlKA+1TKNJA60vHoaICOHMG2L4dOHGC69TJydWvn/v0U2D6dMDISHKnrkNOxb4O\n6FQshfJ+87raOV0MT5EGWl/efYqL6zdEXLkCNHpjJQBARaV+M8SgQcB/3UMmqz6LOnYUCqVDQts5\nhUJpK4QABQX16+du3RK9r6UFDB0K9O0LNPNOA6npEGvsKBQKhUKhUN4WamuB7Ox6h67R4Q0s3bvX\nr5/r0UOyDRHtQYd27IKCgtiTvykUCkUW0Kk1ijTQ+vJu8PIlkJFRvyGiyWlrYJj6na12doCBgezT\nTkpKkulaTDoVS6FQOiR0jR3lbYDWl7ebsrJ6Zy4jo363a2Pk5YF+/YAhQ4AmZ7S/Fugau1agjh2F\n8la9kTUAACAASURBVH5D2zmFQmmO+/frp1uzs+vPo2uMqmr9hoiBA+s3R7wp6Bo7CoVCoVAoFAkh\npH4jxPnz9RsjmqKjU78hok+f+uNL3lXe0qV/soGeYyd7jh49it69e7e3GRJRWFgIHo+H8+fPy0zn\nhg0bJH6/K4/HQ2xsrMzSbkxmZia6dOmCFy9evBb9skbWeRETE4MhQ4bITJ800D6FIg20vrQ/NTX1\nBwrv2QMcOiTq1JmYADNnAosXA/37v3mnTtbn2HV4x46ubZAddXV1WLt2Lfz9/dvblHbjs88+w+nT\np3Hp0iWx9x0cHN7I2xjWrl2LlStXsoclv2/MmDEDT58+xc8//9zeplAolLeUykrg7Flgxw7g6FGg\n8au+eTygd29gwQLA0xOwsGift0QAgKOjo0wdu3d4sPH1kpOfjzPZ2agGIA/A2coKQlPTt0Zfe3D8\n+HE8fvwYrq6u7W1Ku6Gqqgo3NzeEhYUhOjqac6+kpATnz5/HoUOHXqsN2dnZSE5Ofm2jge8CPB4P\nnp6e2LVrF6ZOnfpG06Y/FinSQOvLm+fp0383RFRXc+8pKAADBgC2toC6evvY97rp0CN2bSUnPx9R\nGRkosbZGqbU1SqytEZWRgZz8/LdCX2pqKoYNGwY1NTWoqanBxsYGp06danbq0dzcHMHBwew1j8dD\neHg4pk2bBlVVVRgbG+PIkSN4+vQppk+fDjU1NZiZmeGXX37h6ImJicHEiRMh12icuqioCFOnToWO\njg6UlZVhZmaGr7/+mr0fGxsLW1tbqKurQ0dHBxMnTkRubi57v8Hm77//HmPGjAGfz4elpSVSU1Nx\n584djB07FqqqqrCyskJqaiobLykpCTweD7///jsGDx4MZWVl9O7dG4mJiS3mXXFxMby8vKCrqws1\nNTUMHz4cZ8+eZe9XV1dj5cqVMDQ0hJKSErp27Yrp06dzdHz44YeIi4tDVVUVR3706FH069cPBs3s\nh3/27BmWL18OAwMD8Pl89O/fH0eOHBHJi8OHD2PixIng8/kwMzMTcSBjYmJgZ2cHHR0dVlZeXo7Z\ns2dDX18fSkpK6N69O1atWsXeP336NBwdHaGlpQV1dXU4Ojri4sWLHL1tqRcNNh86dAhOTk5QUVGB\nmZkZfvzxxxbLobW8AIDQ0FCYmZlBSUkJurq6GDt2LF6+fMne/+CDD3D27FncvXu3xbQoFMr7QVER\ncPgwsGsX8NdfXKdOTQ0YPRpYuRIYM6bjOnUAdezEciY7G4oDBiCptJT9XDAzw8qUFAQVFEj9WZGS\nggtmZhx9igMGID47W2rbampqMHnyZAwdOhSZmZnIzMxEcHAw+C28y0Tcu103btyIiRMn4u+//8aE\nCRPwySefwN3dHePGjUNWVhYmTJiAWbNm4cmTJ2yclJQU2NracvQsXrwYFRUViI+PR05ODvbv389x\nbKqqqhAQEIDMzEycOXMGnTp1woQJE1Dd5GeUv78/lixZgqysLPTs2RPu7u7w9PSEt7c3MjMzYWlp\niRkzZqCmpoYTb+XKlQgKCkJWVhZsbW0xadIkPBB3qiSAyspKjBw5Es+fP8eJEyeQlZWF8ePHY/To\n0bhx4wYAICwsDIcPH8ahQ4dw69YtHDt2DEOHDuXosbW1RWVlJf7880+O/MiRI82OZhJCMGnSJFy5\ncgU//fQTsrOz4e3tDXd3dyQkJHDC+vj4wMvLC1euXIG7uzvmzZvHcYaTk5NFysHPzw+ZmZk4duwY\nbt26hR9//PH/sXfncXFX5/7AP9+ZYVhnY2dYQ0hYskBgskE2gzFpo9W0/WlcYpPqbXuNGvX26q02\nJtGq1Xqjtd5ea2xMjbW3rVdvfdlaNSSYhCXJANkIkABhh7DNDMM2zPL9/fENAzMDyQCz87xfL19m\n5gzfOcBh5plzzvMcZGRkmNsHBgbw6KOPorS0FCUlJZg3bx42bdpk8fsFpjcuAG5p+OGHH8a5c+dw\n33334f7778fZs2en/bP45JNP8Oqrr+Ktt95CbW0tvv76a3z729+2uE56ejokEonNz8/ZaM8UmQoa\nL87FskB1NXDwIPDee1yW6/jE0qgoYMsWYNcuIC8PCAhwX19dhvVRANg9e/awx44dm7DtRt747DN2\nT309u7a83OK/jYcOsXvq66f838ZDh2yutae+nn3js8+m/H319vayDMOwhYWFNm1Xr15lGYZhi4qK\nLO5PSUlh9+3bZ77NMAz75JNPmm93dXWxDMOwjz/+uPk+lUrFMgzD/v3vf2dZlmW1Wi3LMAz7+eef\nW1w7MzOT3bt3r9397+npYRmGYYuLiy36/Otf/9r8mDNnzrAMw7D79+8331dRUcEyDMNWVlayLMuy\nx44dYxmGYQ8ePGh+jMFgYBMTE9ndu3dP+PN4//332bi4ONZgMFj06ZZbbmGfeOIJlmVZdteuXez6\n9etv+n2IxWL2wIED5tt9fX1sQEAAW11dbb6PYRj2j3/8o7m/AQEBrEajsbjOjh072Lvuusuiv2+8\n8Ya53Wg0siKRiP3d735nvi88PJx9++23La5z5513stu3b79pv8dfVyaTmfs32t+pjovRPj///PMW\n18/NzWW3bds27Z/F/v372fnz57N6vf6G38fixYvZ5557bsI2Z728TfSaQshkaLw4x8gIy545w7Jv\nvcWye/bY/vfBByxbW8uyJpNbu2mXY8eOsXv27HHYa5ZP77Gb7mbEyY5940+zvgxvkq8TTuNaMpkM\nDz/8MDZu3Ij169dj7dq12LJlC+bPnz+l62RmZpr/HR4eDj6fj8WLF5vvk0qlEAqF6OzsBABoNBoA\ngEgksrjOE088gR//+Mf44osvsG7dOmzevBmrV682t589exb79u3DuXPn0N3dba7R09jYaDETNr4/\nUVFRAGDRn9H7Ojs7LWaixl+Dz+dj2bJlqJxkJvTMmTPo6OiA1GoOXqfTmWc8d+zYgQ0bNiAlJQUb\nNmzAhg0bcMcdd8DP6jBAsVgMtVptvv33v/8dc+bMQWpq6qTPPTIygtjYWIv7R0ZGbH53WVlZ5n/z\neDxERkaafw8A97uw/j088sgj+N73vgelUon8/Hxs2rQJGzduNM/UXr16Fc8//zxKS0vR2dkJk8mE\nwcFBNDU1WVxnquNilPWsZl5eHgoKCqb9s7jnnnvwm9/8BomJibjtttuQn5+Pu+66CyEhIRZfY/17\ncAXaM0WmgsaLYw0MAGfOAKdPA9ZFAfh8LiFi5Upups5bjJ6QNX7L1Ez4dGA3XbcuWIBDZWVYl5Nj\nvk9XVobta9Ygdc6cKV+vhmVxqLwc/lbXy8/Onlb/3n33XezatQtfffUVvv76a+zevRtvv/02Nm3a\nBAA2BQ6tlz0B2AQqE93HMAxM1ys3jgZDWq3W4jHbt2/Hpk2b8M9//hPHjh3Dt771LWzZsgWHDx/G\n4OAgbrvtNqxZswaHDh1CVFQUWJbFggULbPanjX/u0WBkovtM1pUkrbAsa7PsPMpkMiE9PR3/93//\nZ9M2ml2amZmJq1ev4uuvv8axY8ewa9cu7N69G6WlpRbBlEajsQgQb7QMO/rcEolkwmxaoVB4w9vj\nfw8A97uw/j3cdtttaGpqwpdffonCwkI88MADWLRoEQoKCsDj8XD77bcjMjISv/3tbxEfHw8/Pz+s\nWrXqhr+Hye6z7s9ErMfgePb8LORyOaqrq3Hs2DEcPXoUL774Ip555hmcOnXKYqnf+vdACPFNPT1A\nSQlw9ixXvmS8gACumPCyZdxeutmOArsJpCYnYzuAgosXMQJuZi0/O3vaWayOvh4ALFiwAAsWLMCT\nTz6Jf/3Xf8W7776LBx98EADQ2tpqflxnZ6fF7ekKDg5GTEwMGhsbbdqio6Oxfft2bN++Hd/61rdw\n33334b//+79RU1OD7u5uvPTSS+aZrOLiYoeeBlBSUoK0tDQA3P7D06dP4wc/+MGEj126dCkOHz4M\nkUhkkXhgLTg4GHfddRfuuusuPPvss4iJicHx48exefNmAEBPTw/6+/vNs0s6nQ5ffPEFnn766Umv\nqVAooFarMTQ0hAULFkz32wUAzJs3Dw0NDTb3y2QybN26FVu3bsWOHTuwcuVKVFVVITo6GlVVVdi/\nfz82bNgAgEt6sZ51m4mSkhLzBwuA+z1P9n3a+7MQCoXYuHEjNm7ciBdffBFRUVH429/+hp07dwLg\ngsfm5uYpz1bPFB0RRaaCxsv0sSzQ3MwVFK6psdw7BwASCTc7t2QJ4O/vnj56IgrsJpGanOzQciSO\nul5dXR3effddfOc730FcXBza2tpw/PhxKBQKBAQEIC8vD6+99hrS0tKg1+vx3HPPwd9BI37t2rU4\ndeoUHnnkEfN9jz76KDZv3oz58+djeHgYn3zyCRISEhASEoLExET4+/vjrbfewlNPPYWGhgb8x3/8\nx6QzatPx6quvIjo6GklJSdi/fz96enos+jfe/fffjzfeeAObN2/GSy+9hHnz5uHatWs4evQoMjIy\ncOedd+JXv/oVYmNjkZmZiaCgIPzpT3+CQCCwCB5OnTqFgIAAc4Hcr7/+GjKZDDnjZmSt5efn49Zb\nb8V3v/tdvPbaa1i0aBFUKhWKi4sRGBiIhx9+eNKvtQ6E165di6KiIov7nnvuOSgUCmRkZIDH4+HD\nDz+ESCRCQkICgoODERERgXfffRfJycno7u7G008/jcDAwJv+fO118OBBpKWlIScnBx9++CFKS0vx\nX//1XxM+1p6fxe9//3uwLIulS5dCKpWioKAAWq3WYhm+qqoKGo2G3jQJ8TEmE5cQUVzMZbpak8uB\n3FwgI4OrR0csUWDnZYKDg1FbW4utW7eiq6sLYWFhuP32280lRg4ePIh/+Zd/QW5uLmJjY/HLX/4S\ndXV1DnnuBx54AA8++CAMBoNFyZMnnngCzc3NCAoKwsqVK/HFF18A4PZoffjhh/jZz36GgwcPIiMj\nA2+88Qby8/MtrjtRoGfvfa+//jp2796NixcvIiUlBX/7298QHR094df4+/vjm2++wc9//nPs2LED\nXV1diIiIwPLly80ZlxKJBPv378eVK1dgMpmQkZGB//3f/8W8efPM1/n000/x/e9/37xs+Omnn2LL\nli03/fl99tln2LdvH5588km0trYiNDQUS5YssZjps+f7fuCBB/D666+js7MTkZGRAIDAwEA8//zz\naGhoAJ/Px5IlS/DFF1+Yl4//+te/4vHHH8fixYuRlJSEl156Cc8888xN+2yvX/7yl3j33XdRWloK\nuVyOP/7xjxZ7Ba3d7GcRGhqK119/HU8//TR0Oh3mzp2LAwcO4JZbbjFf49NPP8WqVauQkJDgsO/D\nHhRIkqmg8WK/kRFuqbWkhKtFZ23+fC6gS0x0XzFhb8CwjlwX8yA3OkyXDgefHpZlkZGRgb1799p9\nrJazFBYWYv369WhpaYFcLnfZ82q1WiQmJuKrr76CQqGA0WhETEwM/vrXv2Lt2rUu68doQoEjg7Pp\naGhoQHJyMk6ePInc3FyXPa/JZEJaWhpefvllfP/735/wMfR3Toh30Gq5ZAilkjstYjw+H8jM5JZc\nb7CDxic46jXLpycx6axYx2IYBq+++ipeeukld3fFbd566y3cdtttUCgUAIDe3l48/vjjFpnArvDa\na6/hzTff9JqzYh3to48+QlhY2KRBnTPRawqZChovk+vq4o76evNN7uiv8UFdYCCwZg3w5JPAd77j\n20Gdo8+KpRk74pUKCwuRn5+P5uZml87YEUsNDQ2YO3cuTpw44dIZO3s46++cNsOTqaDxYollgYYG\nbv/cuLrrZqGh3OxcZiZ3/Nds4qjXLArsCCE+if7OCfEcRiNw6RIX0LW327bHxXEnQ6Smzt6ECEe9\nZlHyBCGEEEKcQqcDysuB0lLgep17M4YB0tK4hIj4ePf0zxdRYEcIIVNAS2tkKmbreOnrA06d4hIi\ndDrLNoGAqz23YgUQFuae/vkyCuwIIYQQ4hAdHVy5kgsXuHp04wUHc6dDLF0KXD/shzgB7bEjhPgk\n+jsnxDVYFqir4wK6icqmhodzCRGLFwMTnFpIrqM9djMgk8kcevoBIcTzyGQyd3eBEJ9mNHIzcyUl\nwLVrtu2Jidz+ufnzqaCwK00a2G3bts2uC/j7++O9995zWIccae/evVi3bp3N/obe3l73dIh4rNm6\nD4ZMHY0VMhW+OF6Gh7m9c6dOccWFx2MY7qiv3FwgNtY9/fM2hYWFDq13OOlSrL+/P5599tmbLmf+\n53/+J7TWv1kPQMswZCp88cWXOAeNFTIVvjRe1Gouu7W8nDv+azyhEMjOBpYvB2iyfHqcXsdu7ty5\ndp0xmpqaipqamhl3xNEosCOEEEJmrq2Nqz9XWcntpxtPJOKCuZwc7rQIMn1UoPgmKLAjhBBCpodl\nuZMhiou5kyKsRUZyy60LF3LlS8jMuTV5or6+HjweD0lJSTPuACGewJeWS4hz0VghU+Ft48VgAM6f\n5wK67m7b9uRkLqCbO5cSIjyVXQd3bN26FcXFxQCA999/HwsWLEBGRobHJk0QQgghxH6Dg8A33wBv\nvAF89pllUMfjcaVKfvIT4MEHgZQUCuo8mV1LsREREWhtbYVQKMTChQvxu9/9DlKpFHfeeSdqa2td\n0c8po6VYQggh5MZ6e7lyJWfPAnq9ZZu/P7d3bvlyQCJxT/9mE5cuxer1egiFQrS2tkKlUiEvLw8A\ncG2iwjWEEEII8WjNzdxya3W1bUKEWMwd95WdDQQEuKd/ZPrsCuwyMzPxyiuvoKGhAZs3bwYAtLS0\nQEIhPPER3rYPhrgPjRUyFZ40XkwmoKaGC+iam23bo6O5/XMLFgB8vuv7RxzDrsDu97//PXbv3g2h\nUIjXXnsNAFBSUoL777/fqZ0jhBBCyMzo9dxSa0kJt/Rqbd48LqBLSqK9c76Ayp0QQgghPqi/Hzhz\nhvtvcNCyjc/nEiJWruRKlxD3c3m5kxMnTqCiogJardb85AzD4Nlnn51xJ5xlsiPFCCGEEF/V3c3N\nzp07x5UvGS8wEFAogGXLuOLCxP1cdqTYeI899hj+8pe/YPXq1Qi0Ki19+PBhh3XGkWjGjkyFJ+2D\nIZ6NxgqZCleNF5YFGhu5gG6iw6CkUm52bskS7vgv4nlcOmP34YcforKyEnK5fMZPSAghhBDHMJmA\nS5e4hIi2Ntv22Fhu/1x6OlePjvg+u2bsFi9ejKNHjyI8PNwVfXIImrEjhBDiq3Q6oKICKC0F1Grb\n9tRULqBLSKCECG/h0rNiz5w5g5dffhn33XcfoqKiLNrWrFkz4044AwV2hBBCfI1WC5w6BSiVwPCw\nZZtAAGRlcTXovGgehlzn0qXYsrIy/OMf/8CJEyds9tg1T1QMhxAvQ/umiL1orJCpcNR46ezkllsv\nXACMRsu2oCAuGWLpUiA4eMZPRbycXYHdc889h88//xwbNmxwdn8IIYQQAi4h4upVLqCb6PTOsDAu\nISIzE/Dzc33/iGeyayk2ISEBtbW1EHpRKg0txRJCCPFGRiNQWckFdB0dtu0JCdz+ufnzKSHCF9TU\n1uBI2RE8uvVR1y3FvvDCC3jiiSewe/dumz12PBpVhBBCyIwNDwPl5VxCRF+fZRvDcJmtublAXJx7\n+kccr6a2BgePHoQ6ZoIMmGmya8ZusuCNYRgYrRf7PQTN2JGpoH1TxF40VshU2DNeNBoumCsv57Jd\nx/Pz42rPrVgBhIY6r5/E9YYNw3j2vWdxMfgiRowj+GbHN66bsauvr5/xExFCCCFkTHs7t9xaWcnV\noxsvJIRLiFAouOQI4ju0Oi1KW0qhbFPiUvcljASMOPT6dFYsIYQQ4iIsyyVCFBdziRHWIiK4hIjF\ni7nyJcR3dA92o7i5GOc6zsHIcqudp0+exmDcIIR8Ib5+8GuHxC2TbpDbvXu3XRfYs2fPjDsxVX19\nfVi2bBlEIhEuXbrk8ucnhBBCpsJg4AoK//d/A3/8o21QN2cOcP/9wCOPANnZFNT5kmZNM/7n4v/g\n7dNvo7y93BzUAUB2RjaSVclYEbfCYc836YxdSEgIzp8/f8MvZlkWOTk5UE9U9tqJDAYD1Go1/v3f\n/x0//elPsWDBApvH0IwdmQraN0XsRWOF2KOmphFHjtTh/PnzCApaDD+/uQgKSrR4DI8HLFjAzdDR\niZ2+hWVZXOm9gpNNJ9GkabJpjxfHIy8hD6lhqbhcdxkF5QXYec9O5+6xGxwcREpKyk0v4O/vP+NO\nTJVAIPCq480IIYTMHjU1jXjnnVp0dubj0iUeJJJ1MBgKkJUFhIcnQigEcnKA5csBqdTdvSWOZDQZ\ncaHzAoqaitA12GXTnhqWiryEPCRIEsbuS0lFakoqdt6z0yF9mDSwM1nv5CTEh9EMDLEXjRVyI62t\nwGuv1aGuLh8AIJGsAwAIBPlobz+Ke+9NRE4OEBDgxk4Sh9MZdChrL0NpSyn6dJa1avgMH4uiFiEv\nPg8RwRFO74tbi9C9/fbbUCgUCAgIwI4dOyzaent7sWXLFoSEhCApKQl/+tOfJrwGQ6cbE0IIcSOW\nBS5fBt5/HzhwAGhttXxrDQ4G0tKA3Fwe8vIoqPMl/SP9KKgvwBulb+Cruq8sgjohX4jc+FzsWrEL\nd6Xd5ZKgDrCz3ImzxMbGYvfu3fjyyy8xNDRk0bZz504EBASgs7MTFRUV2Lx5MzIzM5GRkWHxONpH\nRxyB9k0Re9FYIaMMBuD8eaCkBOgat+rG43ErXjIZwDCFWLRoHRgGCAyklTBf0TPYg+LmYpztOGuR\nDAEAIcIQrIhbAYVcgQCB66N4twZ2W7ZsAQAolUq0tLSY7x8YGMAnn3yCyspKBAUFIS8vD3feeScO\nHz6MV155BQDw7W9/G+fOnUNNTQ1+/OMf4wc/+IFbvgdCCCGzy9AQcOYMcPo00N9v2cbjAd/61lxc\nuVKA0NB8NDRwp0bodAXIz7/5vnXi2Vr7WnGy6SSqu6vBwnJiKSwwDLnxuciMzoSA577wyiMSqq1n\n3S5fvgyBQGCRvJGZmYnCwkLz7X/84x83ve727duRlJQEAJBKpcjKyjJ/0h69Ft2m26PGz8S4uz90\n23Nvr1u3zqP6Q7dddzszcx1KS4GPPy6E0QgkJXHtDQ2F8PMD/t//W4cVK4Dy8quQSLrQ338UUikP\nDQ37kZ0tR2pqvkd9P3TbvtvHjh1Da18rDIkGNKgb0HC2AQCQlJUEAOiv6ceiqEXYtnYbeAzP7uuP\n/ruhoQGOZFeB4s7OTgQGBkIkEsFgMOCDDz4An8/Htm3bHHJW7O7du9HS0oL3338fAHDixAncfffd\naG9vNz/mwIED+Oijj3Ds2DG7rknlTgghhDhCaytXUPjSJW4/3XhiMXfcV3Y27Z3zNUaTERc7L6Ko\nuQidA5027fNC52FVwiokSBIcst/fUXGLXTN2t99+O373u99hyZIleO655/D555/Dz88PFRUVePPN\nN2fcCetvJCQkBH1WJyBrNBqIRKIZPxchEykcN1tHyI3QWJkdWBa4cgUoKgIaG23bo6KAvDyuDh2f\nP/l1aLx4nxHjCMrby1HSXAKNTmPRxmN4WBS5CLnxuYgKiXJTD2/MrsDuypUryMrKAgB8+OGHKC4u\nhkgkQkZGhkMCO+tId/78+TAYDKitrTUvx547dw4LFy6c0nX37t2LddeXTgghhJCbmSwhYtTcuUBu\nLpCczO2dI75jYGQAp1pP4UzrGQwZLBM6hXwhcmJysCJuBSQBEoc+b2FhocXy7EzZtRQbHh6OlpYW\nXLlyBVu3bkVlZSWMRiMkEgn6rXeOToHRaIRer8e+ffvQ2tqKAwcOQCAQgM/n49577wXDMHjvvfdQ\nXl6O22+/HSUlJUhPT7fvG6OlWEIIIXa6WULEokXcCRHR0e7pH3Ge3qFelDSXoKKjAgaTwaIt2C8Y\ny+OWY6l8KQL9Ap3aD5cuxW7atAl33303enp6cM899wAALl26hLi4uBk9+YsvvogXXnjBfPvDDz/E\n3r178fzzz+O3v/0tfvjDHyIyMhLh4eF455137A7qCCGEEHuoVEBpKVBeDuj1lm3+/twJEStWcHvp\niG9p07ahqKkIl7ou2WS4hgaGchmuUZnw4/u5qYfTY9eM3fDwMP7whz9AKBRi27ZtEAgEKCwsREdH\nB7Zu3eqKfk4ZwzDYs2cPLcUSu9A+GGIvGiu+wVUJETRePAvLsqhT1aGoqQhX1Vdt2uUiOfLi85Ae\nkQ4eM/PkUHuMLsXu27fPITN2dgV23oiWYslU0IsvsReNFe/lqISIqaDx4hlMrAmVnZUoai5CR3+H\nTXtKaAry4vOQJE1y24lWjopbJg3stm3bZvOEABftjv+mP/jggxl3whkosCOEEAJQQsRsNmIcQUV7\nBUpaSqAeVlu08RgeFkQsQF5CHqJD3L950ul77ObOnWsO4Lq7u/GHP/wBd9xxBxITE9HY2IjPP/+c\nTnsghBDisSghYvYa1A/idOtpnG49jUH9oEWbH88P2THZWBm/EtIAqZt66DyTBnZ79+41//u2227D\n3//+d6xevdp838mTJy0SHzwRlTsh9qLlEmIvGiuez56EiOXLAYljq1ZMiMaLa6mGVChpKUFFewX0\nJstffpBfEJbHLsfS2KUI8gtyUw9tuaXciVgsRk9PD/z8xjJD9Ho9QkNDodVqHdYZR6KlWDIV9OJL\n7EVjxXN54gkRNF5co13bjqLmIlR2VtpkuEoDpMiNz8WS6CUeneHq9D12461duxZLly7Fiy++iMDA\nQAwODmLPnj04deoUjh8/PuNOOAMFdoQQ4vtGEyKKi4GJjtyMiuL2zy1c6LiECOIZWJbFVfVVFDUV\noU5VZ9MeExKDvIQ8ZERkuCzDdSZcWsfu0KFDuO+++yAWiyGTyaBSqaBQKPDRRx/NuAOEEELIVFFC\nxOxlYk241HUJRU1FaO9vt2lPliUjLz4PybJkt2W4upNdgd2cOXNQUlKCpqYmtLW1ISYmBomJEUuv\nEwAAIABJREFUic7u24zRHjtiL1ouIfaiseJeQ0OAUgmcOuUdCRE0XhxHb9TjbMdZFDcXQzWssmhj\nwGBB5ALkxudCLpK7qYfT45Y9dqM6OzttjhBLTk52WGcciZZiyVTQiy+xF40V9/CkhIipoPEyc4P6\nQZxpPYNTradsMlwFPAGX4Rq3ErJAmZt66Bgu3WP3z3/+Ew899BDa2y2nPBmGgdFonHEnnIECO0II\n8X6emBBBXEM9rEZpSynK2spsMlwDBYFYFrsMy2KXIVgY7KYeOpZLA7vk5GQ8/fTTePDBBxEU5Dkp\nwjdCgR0hhHgnSoiY3a71X0NRcxEudl6EiTVZtEn8JVyGa8wSCPlCN/XQOVwa2IWGhqKnp8erNiFS\nYEemgpZLiL1orDiPLyZE0HixD8uyaFA3oKi5CLW9tTbtUcFRyEvIw4KIBeDzfDOad2lW7EMPPYSD\nBw/ioYcemvETuhIlTxBCiOe7WULEwoVcQOcpCRHEcUysCdXd1ShqKkKrttWmfY50DvIS8jBXNter\nJpemwi3JE6tWrcLp06eRmJiI6HF/WQzDUB07Qggh0zKaEFFRAYyMWLZ5ckIEmTmDyWDOcO0d6rVo\nY8AgPSIdefF5iBXHuqmHrufSpdhDhw5N2glPPS+WAjtCCPFMbW1AURElRMxGQ/ohKNuUKG0pxYB+\nwKJNwBMgKzoLK+NWIiwozE09dB+XBnbeiAI7MhW0D4bYi8bK9MzWhAgaLxzNsIbLcG0vw4jRcno2\nQBBgznANEYa4qYfu59I9dizL4v3338fhw4fR2tqKuLg4PPDAA9ixY4fPrnkTQgiZOV9MiCD26xzo\nRFFTES50XrDJcBX7i7EybiWyY7LhL/B3Uw99j10zdi+99BI++OAD/Nu//RsSEhLQ1NSEN954A/ff\nfz9+/vOfu6KfU0YzdoQQ4j6UEDF7sSyLJk0TipqLcLnnsk17ZHAk8uLzsDByoc9muE6HS5dik5KS\n8M0331gcI9bY2IjVq1ejqalpxp1wBoZhsGfPHsqKJYQQF6KEiNmLZVkuw7W5CC19LTbtiZJE5CXk\nYV7oPFrtG2c0K3bfvn2uC+wiIyNx9epVBAePVXfu7+9HcnIyOjs7Z9wJZ6AZOzIVtA+G2IvGysTa\n2rj9c5WVEydELF/OBXWzLSFiNowXg8mA89fOo6ipCD1DPRZtDBikhachLyEPceI4N/XQO7h0j92m\nTZvwwAMP4JVXXkFiYiIaGhrw3HPPYePGjTPuACGEEO80WxMiCGfYMGzOcO0fsVxv5zN8LsM1fiXC\ng8Ld1MPZya4ZO41Gg8ceewx//vOfodfr4efnh7vvvhu/+c1vIJVKXdHPKaMZO0IIcQ6DAbhwgQvo\nJkqISE4G8vIoIcJX9en6zGe46ow6izZ/vj+Wxi7F8tjlEPmL3NRD7+SWcidGoxHd3d0IDw8H38M/\nflFgRwghjkUJEbNb10AXipuLcf7aeRhZo0WbSCjCyviVyInJoQzXaXJpYPeHP/wBWVlZyMzMNN93\n7tw5nD9/Htu2bZtxJ5yBAjsyFbNhHwxxjNk4VighYvp8Ybw0aZpQ1FSEmp4am7bwoHDkxedhcdRi\nynCdIZfusdu9ezfOnj1rcV9cXBzuuOMOjw3sCCGEzAwlRMxeLMvics9lFDUXoUljW/0iQZKAvPg8\nzA+bTxmuHsauGTuZTIbu7m6L5VeDwYCwsDBoNBqndnC6aMaOEEKmjhIiZjeDyYAL1y6guLkYXYO2\nGyhTw1KRl5CHBEmCG3rn21w6Y5eeno6PP/4Y99xzj/m+Tz/9FOnp6TPugDPt3buX6tgRQogdKCFi\ndtMZdChrL0NJcwm0I1qLNj7Dx+KoxciNz0VEcISbeui7RuvYOYpdM3YnT57Et7/9bWzYsAHJycmo\nq6vDkSNH8I9//AOrVq1yWGcciWbsyFT4wj4Y4hq+NlYoIcK5PH28aHVanGo9hTOtZybMcFXIFVge\ntxxif7Gbejh7uHTGbtWqVbhw4QI++ugjtLS0YNmyZfj1r3+N+Pj4GXeAEEKI66nV3PmtlBAxO3UP\ndqO4uRjnOs7ZZLiGCEOwIm4FFHIFAgS0gdLbTLncybVr1yCXy53ZJ4egGTtCCLFFCRGzW0tfC4qa\nilDdXQ0WlgMgLDAMeQlchquAZ9e8D3Egl87YqVQq7Ny5Ex9//DEEAgEGBwfx2Wef4fTp0/jFL34x\n404QQghxHkqImN1YlsWV3isoaipCo6bRpj1OHIe8+DykhadRhqsPsCuw+8lPfgKZTIbGxkZkZGQA\nAFauXImnnnqKAjviEzx9HwzxHN40Vighwv3cOV6MJiMudl5EUXMROgdsz3WfHzYfefFchisFdO5T\nU1+PI5WVDrueXYFdQUEB2tvb4efnZ74vIiICnZ22A4UQQoh7UULE7KYz6FDeXo6SlhL06fos2ngM\nD4siFyEvIQ+RwZFu6iEZVVNfj0Pl5RjJynLYNe0K7KRSKbq6uiz21jU1NXnFXjtC7OEtMzDE/Tx5\nrFBChOdx5XjpH+nHqZZTONN2BsOGYYs2IV+InJgcrIhbAUkADQBP8bfz51GbloYurfbmD7aTXYHd\nww8/jO9///v4xS9+AZPJhJKSEjz77LP48Y9/7LCOEEIImR5KiJjdeod6UdxcjLMdZ2EwGSzagv2C\nzRmugX6BbuohsaY1GFCoVuOoRoNhvd6h17YrsHvmmWcQGBiIRx99FHq9Hjt27MBPfvIT7Nq1y6Gd\nIcRdvGnfFHEvTxkrlBDhHZw5Xlr7WlHUXISqriqbDNfQwFDkxuciKzqLMlw9yJDRiCKNBqe0WuhN\nJvCcUL3Drt82wzDYtWsXBXKEEOJm9iRE5OYCc+dSQoQvYlkWdao6nGw6iQZ1g027XCTHqoRVSAtP\nA4/hub6DZEJ6kwmntVqcUKsxbDKZ709OTkbDuXOYv2oVvnHQc9lVx+7o0aNISkpCcnIy2tvb8cwz\nz4DP5+OVV15BtIfuvqU6doQQX0IJEbOb0WREZVclipqKcG3gmk17SmgK8uLzkCRNogxXD2JiWVT0\n96NQrYbWYLlMHuPvj1tlMhja21FQWYmd3/mO6+rYPfLII/jqq68AAE899RQYhoFAIMCPfvQjfPbZ\nZzPuhLPQWbGEEG9HCRGz24hxBBXtFShuLoZGp7Fo4zE8LIxciNz4XESHUETvSViWRdXgII6qVOi2\n2kMX6ueHfJkMGUFBYBgGhU1N6Cwrc9hz2zVjJxaL0dfXB71ej6ioKDQ2NsLf3x8xMTHo6elxWGcc\niWbsyFR4yr4p4vlcNVYoIcI3THe8DIwM4HTraZxuPY0hw5BFmx/PD9kx2VgZvxLSAKmDekoc5erQ\nEI6oVGjVWZ69G8LnY51UiiUiEfgTzKq69OQJsViMjo4OVFZWYsGCBRCJRNDpdNA7OJODEEJmM0qI\nIKohFYqbi1HRUWGT4RrkF4TlscuxNHYpgvyC3NRDMpl2nQ5HVCrUDVkG4gE8HvIkEiwXiyHkOX/f\no12B3WOPPYZly5ZBp9PhzTffBAAUFRUhPT3dqZ0jxFVoto7YyxljhRIifJe946Vd246i5iJUdlba\nZLjKAmTmDFc/vt8kVyDu0qvX46hKhYsDAxb3CxgGy8RirJJIEOTCT2J2LcUCQE1NDfh8PlJSUgAA\nly9fhk6nw6JFi5zawemipVhCiKejhIjZjWVZ1KvqUdRchHpVvU17TEgM8hLykBGRQRmuHqjfYMA3\nGg3KtFqYxsUbDMMgKyQE66RSSAT2l5pxVNxid2DnbSiwI1NBe+yIvRwxVtRqoLQUKC+3TYgQCrm9\ncytWUEKEL5hovJhYEy51XUJRUxHa+9ttvmaubC7yEvIwRzqHMlw90LDRiOK+PpT09UE/rnQJAKQH\nB2O9VIoIoXDK13X6Hru0tDRUV1cDAOLj4yftRFNT04w7QQghs8GNEiJEIi6Yo4QI36U36lHRUYGS\n5hKohlUWbQwYLIhcgLz4PMSIYtzUQ3IjhtFadBoNhoxGi7akgADcKpMhzgP+eCedsTtx4gRWr14N\ngPvEMRlPneWgGTtCiCeghIjZqaa2BkfKjkDP6sGaWITFhqFD0IFB/aDF4/x4flgSswQr41ZCFihz\nU2/JjZhYFueu16LTWNWiixYKkS+TISUwcMazq7QUexMU2BFC3IkSImavmtoaHDp2COwcFs2aZrT3\nt2PkygiyMrIQLg8HAAQKArE8bjmWypciWBjs5h6TibAsi5rBQRSo1eiy2jMh8/PDeqkUC4ODHbZc\n7vSl2N27d0/6JKP3MwyDF154YcadIMTdaI8dsdfNxgolRMxuLMvio+Mf4YrkCnpaeqCqVkGaJoUg\nRYCr9VeRkpyClXErsSRmCYT8qe/DIq7RODyMIyoVmoeHLe4P5vOxVipFziS16DzBpIFdc3PzDaPQ\n0cCOEEIIJUTMdgMjA6joqEBZWxlOtZ3CcJxlQBAiDEFGZAYeX/44Zbh6sGsjIziiUuHKoOWSuT+P\nh1yJBCtdVItuJmgplhBCZoASImYvlmXRqGmEsk2Jqq4qGFluQ/3pk6cxGMcFBtIAKRIkCZAFyBDV\nFYVH7n7EnV0mk1Dp9TimVuPCwIBF7MBnGCwVibBaKkWwkzfBOn0ptr7etqbORJKTk2fcial65pln\nUFJSgqSkJBw8eBCCKdSJIYSQ6aipacSRI3XQ63kQCEyYN28u2tsTKSFiFhrSD+HctXNQtinRPdht\n054xLwMtjS1IWJJgPiFCd0WH/FvyXd1VchMDRiOOq9VQarUwWtWiywwOxjqpFFI/7yoKPWlENFqI\n+EYYhoHRKuXX2c6dO4e2tjYcP34cL7/8Mj7++GNs3brVpX0gvof22JEbqalpxKFDtfDzy0dFRSGM\nxvXo6ytAVhYQHp5ofhwlRPgulmXRqm2Fsk2Ji50XbY77AoB4cTwUcgUyVmeg/mo9CsoLcOniJWQs\nzED+LflITUl1Q8/JRHQmE0o0GhT39WHEqhZdalAQ8mUyRE6jFp0nmDSwM1l9o56ipKQEGzduBABs\n2rQJ77//PgV2hBCn0euBw4frUF+fj54eoLsbkEoBgSAfV68eRWRkIhYuBFauBGKo/JjP0Rl0uNB5\nAco2JTr6O2zahXwhMqMykSPPQXTIWEZMakoqUlNSURhJHxo9icFkglKrxXGNBoNWE1MJ12vRJXj5\nvgmvW8NUqVSIuf7qKRaL0dvb6+YeEV9AL7xkPJ2Oqz136RL3f6WSh9HkOKl0HQBuiTUujodduygh\nwhd19HdA2abE+WvnMWIcsWmPCYmBQq7AwsiF8Bf4T3odem3xDCaWxYWBARxTqaC2qkUXKRTiVpkM\n8xxQi84TTBrYbdy4EV9++SUAmAsVW2MYBsePH5/WE7/99ts4dOgQLl68iHvvvRfvv/++ua23txcP\nPfQQvv76a4SHh+OVV17BvffeCwCQSqXo6+sDAGg0GoSGhk7r+QkhZLyhIaCmhgvm6uqA8R/mebyx\nFYyAAEAuH/3PREGdD9Eb9ajsqoSyTYmWvhabdj+eHxZGLoRCroBcJPeJIMDXsSyLK0NDKFCpcM0q\nXV0qEOAWmQyLgoPB86Hf5aSB3YMPPmj+90MPPTThY2YyqGNjY7F79258+eWXGBoasmjbuXMnAgIC\n0NnZiYqKCmzevBmZmZnIyMhAbm4u9u/fj23btuHLL7/EqlWrpt0HQkbRHrvZqb8fqK7mgrmGBmCy\nHSjZ2XNRX1+AmJh8dHcXIiFhHXS6AuTn33wvMvF8XQNdKGsvw9mOsxg2DNu0RwRFQCFXYHHUYgT6\nBU7p2vTa4j7Nw8P4WqVCk1UtuiA+H2skEihEIgg8vHTJdEwa2N1///3mf2/fvt3hT7xlyxYAgFKp\nREvL2CejgYEBfPLJJ6isrERQUBDy8vJw55134vDhw3jllVeQmZmJqKgorFmzBomJiXj66acd3jdC\niO/SaICqKu6/pibbEiWjoqOB9HQgIwOIiEhETQ1QUHAUOt15REaakJ+fgtTUxIm/mHg8g8mAqq4q\nlLWXoUHdYNPOZ/jIiMiAQq5AgiSBZue8SOfICApUKtRY1aIT8nhYKRYjVyKBvw8GdKPs3mN3/Phx\nVFRUYGBgAMBYgeJnn312Rh2wrtly+fJlCAQCi6zczMxMi/NqX3vtNbuuvX37diQlJQHglnCzsrLM\nn5xGr0e36fao8Z+s3d0fuu3Y23/7WyGamoCAgHVobQUaGrj2pCSuffT2qlXrkJ4O9PQUQiwG1q61\nvN4jj6wHsB6FhYVob79qDuzc/f3Rbftv9w714uAnB3Gl9wpiFnH7tRvONgAAkrKSEBoYCqaBQUpo\nCr6V8a0ZP9+6des86vv35dtZeXkoVKvxfwUFAMsiacUKAEDTqVNIDQzEzs2bESIQeEx/R//dMFHN\npBmwq0DxY489hr/85S9YvXo1AgMtp6EPHz48ow7s3r0bLS0t5j12J06cwN1334329nbzYw4cOICP\nPvoIx44ds/u6VKCYkNmLZYHOzrGZuWvXJn4cwwCJidzMXHo6IBa7tp/ENUysCTXdNVC2KVGnqrNp\n5zE8pIalQiFXIFmWTLNzXmbQaMQJjQan+/psatEtCg7GLVIpZF5Qi87pBYrH+/DDD1FZWQm5XD7j\nJ7Rm/U2EhISYkyNGaTQaiEQihz83IaMKx83WEe/EskB7O7dfrqoK6OmZ+HE8HldvLj0dSEsDgqd4\n/jqNFe+hGdagvL0c5e3l0I5obdrF/mLkxOQgOyYbIn/nvMfQeHGeEZMJpX19KNJooLPaIDsvKAj5\nUimi/SfPWPZVdgV28fHxEAqFTumA9Sej+fPnw2AwoLa21rwce+7cOSxcuHDK1967d695KpwQ4ntY\nFmhuHgvmNJqJHycQACkpXDA3fz4QOLX978SLmFgT6nrroGxT4nLPZbCwnDxgwC2zKuQKzAubR+e2\neiEjy6JMq8VxtRr9VrXo4vz9catMhiQv+iMvLCy0WJ6dKbuWYs+cOYOXX34Z9913H6Kioiza1qxZ\nM60nNhqN0Ov12LdvH1pbW3HgwAEIBALw+Xzce++9YBgG7733HsrLy3H77bejpKQE6enpdl+flmIJ\n8U1GI9DYyAVz1dVcZutEhEJg3jwu+WHePO428V39I/2oaK9AWXsZ1MNqm/YQYQiyY7KRHZMNaYDU\nDT0kM8WyLC4ODOCoWg2VXm/RFiEUIl8qRWpQkNcupTsqbrErsHvnnXewa9cuiEQimz12zc3N03ri\nvXv34oUXXrC57/nnn4dKpcIPf/hDcx27X/7yl1M+XYICO0J8h8EA1Ndzs3LV1VzNuYkEBACpqVww\nl5wMeMG2GjIDLMuiQd0AZZsSVd1VMLG29WqSZclQyBVIDUsFn0cH93ojlmVRNzSEIyoVOqxq0YkF\nAtwilSIzJMTra9G5NLALCwvD//zP/2DDhg0zfkJXYRgGe/bsoaVYYhfaB+N5RkaA2loumLt8mTsN\nYiLBwdxeuYwMICmJOxHCmWisuN+gfhDnOs5B2aZEz5DtZspAQSCWxCxBTkwOwoLC3NDDMTReZqZl\neBhHVCo0WNWiC+TzsVoiwVKRCH48715OH12K3bdvn+uSJ4KDg7F27doZP5mr7d27191dIIRMwfAw\nF8RVVXFBndVqi5lYPFZjLj6eS4ggvo1lWbT0tUDZpkRlVyUMJoPNYxIkCVDIFciIyICA53UnZpJx\nukdGUKBWo+p6ibVRfjweVojFyBOLEeDsT3EuMjoBtW/fPodcz64Zu0OHDuH06dPYvXu3zR47noe+\notJSLCHeYXCQW16tquKWW632QpuFho6VJYmN5UqVEN+nM+hw/tp5KNuUuDZgW7fGn++PzOhM5MTk\nICokaoIrEG/SZzCgUK1GRX+/xXs4j2GQIxJhjUQCkcA3g3aXLsVOFrwxDAPjZK/CbkaBHSGeS6sd\nqzHX0DD56Q+RkWPBXFQUBXOzSbu2Hco2JS50XsCIccSmXS6SQyFXYGHkQgj5lBnj7YaMRpzUaHCq\nrw8GqxeEhcHBWC+TIdTHN826tI5dfX39jJ/IHajcCbEX7YNxPrV6rCzJjXKu5PKxYC483HX9sxeN\nFecZMY6gsrMSyjYlWrWtNu1+PD8siloEhVwBucjxdVWdgcbLjenH1aIbtqpFNzcwEPkyGeQ+XovO\nLeVOvBHN2JGpoBdf5+ju5gK5S5e44sGTiY/n9sulpwNSD69EQWPF8ToHOlHWVoZz185h2DBs0x4Z\nHAmFXIHFUYsRIAhwQw+nj8bLxIwsiwqtFt9oNNAaLPdLxl6vRTfHi2rROYJLl2K9EQV2hLgey3LH\nd40Gc11dEz+Ox+OO8srI4DJa6WCZ2cdgMuBS1yWUtZWhUdNo0y7gCZARkQGFXIF4cbzX1iYjlliW\nxaXBQRxVqdBjlR0V5ueHfJkM6V5ci24mXLoUSwghk2FZoLV1bM9cb+/Ej+PzudpyGRlcrbmgINf2\nk3iGnsEelLWX4WzHWQzqB23aQwNDoZArkBWdhSA/GiS+pP56Lbo2q9pFIoEA66RSLPGBWnSewKcD\nO9pjR+xFyyVTYzIBTU1jwZzV8c5mfn6WR3kFeNcq2oRorEyd0WRETU8NlG1K1Kts92zzGB7SwtOg\nkCswRzrHp2ZraLwAbTodjqhUqLeqLB7A42GVRILlYrHX16KbCUfvsfP5wI4Q4hhGI3D16tjpD1bl\npcz8/bkgLj2dC+roKK/ZSzOsQVl7Gcrby9E/Ynv2m8Rfghx5DpZEL4HIn9bjfU2PXo+jKhUqrV4s\nBAyD5WIxVkkkCPSRWnQz4ZY6dvX19Xjuuedw9uxZ9I87mJFhGDQ1NTmkI45Ge+wImTm9nqstd+kS\nUFPDFRCeSGAgt1cuPZ1bbvXRMlPEDibWhNreWijblLjScwUsLF+HGTCYHzYfOfIcpISmgMfM3pka\nX6U1GPCNWo3y/n6YrGrRLQkJwVqpFGJ6kbDh0j129913H1JSUrB//36bs2IJIb5lZAS4coUL5q5c\n4W5PJCRkrCxJUhKd/jDbaXVaVHRUoKytDBqdxqZdJBQhOyYb2THZkARI3NBD4mzDo7XotFrorUqX\nZAQHY71UinCawnc6u2bsxGIxVCoV+F40ZUozdmQqZvs+mKEh7iivS5eAujrAYHtaEwBAIhkrSxIf\nPzsLBs/2sTIey7K4qr4KZZsS1d3VMLEmm8fMlc2FQq7A/LD54PO85z3EUWbDeNGbTDit1eKkRoMh\nq0ML5gQG4laZDLE+XovOEVw6Y7dmzRpUVFRAoVDM+AldiZInCJncwAC3V+7SJW7vnMn2PRkAEBY2\nFszFxMzOYI5YGtQP4mzHWSjblOgdsk2DDvILwpLoJciR5yA0MNQNPSSuYGJZnO3vR6FajT6rT4Mx\n12vRJQcE+FQyjDO4pUDxzp078ec//xnf/e53Lc6KZRgGL7zwgsM640g0Y0eIrb6+sRpzTU2TH+UV\nFcUFchkZQEQEBXOEm51r7muGsk2Jys5KGFnb4yQTJYlQyBVIj0iHgEd7qHwVy7KoHhxEgUqFbqta\ndKF+flgvlWJBcDAFdFPk0hm7gYEB3H777dDr9WhpaQHA/WLpl0aI5+vtHStLcv3Pd0KxsWN75sLC\nXNc/4tmGDcM4f+08lG1KdA502rQHCAKQGZWJHHkOIoMj3dBD4kpXr9eia7WqRRfC52OtVIpskQh8\nig3cik6eIAS+tQ+GZbkTH0aDuY6OiR/HMEBCwlgwJ6H97HbxpbFyI23aNijblLhw7QL0Jr1Ne6wo\nFgq5AgsjF8KP79uHs8+Er4yXdp0OBSoVaq1q0fnzeMiTSLBCLIaQMqhmxOkzdg0NDUhKSgLAlTuZ\nTHJy8ow7QQiZGZblArhLl7hgrrt74sfxeMCcOVwgl5bGZbYSMmrEOIKLnRehbFOiTdtm0y7kC7Eo\nchEUcgViRDFu6CFxtV69HsfUalzot6xDKGAYLLteiy7IixIrZ4NJZ+xEIhG0Wi0AgDdJFM4wDIxG\n230WnoBm7IivY1luaXU0mFOrJ36cQADMncsFc6mpXM05Qsa71n8NZe1lONdxDjqjzqY9KjgKCrkC\ni6MWw19A2Y2zQb/BgOMaDZRarUUtOoZhkBUSgnVSKSRUi86hnD5jNxrUAYBpsnQ5D0dZscTXmExA\nYyMXzFVXA+P+TC34+QHz5nHJD/PmcadBEDKewWRAZWcllG1KNPc127QLeAIsiFgAhVyBOHEc7ame\nJXQmE4o0GpT29WHE6r0/LSgI62UyRFItOodyS1asN6IZOzIVnrwPxmCwPMpr0PbcdADcOazz53PB\n3Ny5XHBHHM+Tx4o9egZ7oGxT4mzHWQwZhmzawwLDoJArkBWdhUA/mt6dKW8ZLwaTCWe0WpzQaDBo\ntRKXGBCAW2UyxPvCYc8ezKVZsYQQ19LrgdpaLpirqQF0tqtjAICgIG6vXEYGt3eOtrqQiRhNRlR3\nV0PZpsRV9VWbdh7DQ3p4OhRyBZKkSTQ7N4uYWBbn+/txTK2GxqoWXZRQiFtlMqQEBtKY8CI0Y0eI\nh9DpuNMfqqq4o7z0tomIAACRaKzGXEICHeVFJqceVqOsrQwVHRXoH+m3aZcGSM2zcyFCyqSZTViW\nxeWhIRSoVOi0OjdQ5ueHW6RSLKJadC5FM3aE+IDBQW5GrqqKO8prslwkmWwsmIuNpYLBZHIm1oQr\nPVegbFOitrcWLCzfKBgwSA1PhUKuwFzZXHrjnoUah4dxRKVC8/Cwxf3BfD7WSKVQUC06rzblwM46\nkWKyjFlCvIkr98H094/VmGtomPwor4iIsRpz0dEUzHkKT90z1afrQ0V7Bcrby6HRaWzaRUIRcuQ5\nWBK9BJIAKlroKp40Xq6NjKBApcJlq426Qh4PuWIxVkok8Kf3dK9nV2BXVlaGRx99FOfOncPwuAjf\nk8udEOJJ1OqxYK65efKjvGJixoK5iAjX9pF4H5ZlUa+qh7JNiZqeGphY208JKaEpUMgmFdWjAAAg\nAElEQVQVmB82HzyG3rRnI/X1WnTnBwYslvr4DIOlIhFWS6UIpg26btNYU4O6I0ccdj279tgtXLgQ\n3/nOd/DAAw8gKCjIom20iLGnoT12xN16esbOZW2zrfVqFhfHLbGmp3NLroTczMDIAM52nIWyTQnV\nsMqmPdgvGEtiliA7JhuhgaFu6CHxBANGI46r1VBqtTBa1aJbHByMW6RSSCl93q0aa2pQe/Ag8gcH\nwbz9tkPiFrsCO7FYDI1G41V7MRiGwZ49e6iOHXEZlgU6O8eCuU7bYzUBcEuqiYlcMJeWBojFru0n\n8U4sy6JJ0wRlmxKXui7ByNquliRJk6CQK5AWngYBj7ZQz1Y6kwklGg2KJ6hFNz8oCPkyGaKoFp17\n6XTAlSs4+sYb4FVUoLC3F/saG10X2P3gBz/Avffei02bNs34CV2FZuzIVEx3HwzLcrNxo8Fcb+/E\nj+PzuXIkGRnc6Q/BwTPrL3EfV++ZGtIP4fy181C2KdE12GXTHiAIQFZ0FnJichARTOv3nsaV48Vg\nMqGsvx/H1WoMWG2Tir9eiy6RatG5T38/ly1XXQ3U1wNGIwpLS7Hu+hY35ptvXJcVOzQ0hC1btmD1\n6tWIiooy388wDD744IMZd4IQb2IycfvkRvfMaWz3qQPgjvJKSRk7yoteT4m9WJZFm7YNyjYlLnZe\nhN5kW/smThwHhVyBBREL4Men5bTZjGVZXBgYwFGVCmqrWnSRQiHyZTLMp1p07qFSjVWXn2CDtWk0\nWcWBZz3aNWO3d+/eib/4+nKnJ6IZO+JIRiOXwTr699lvWxIMACAUcqc/pKdzR3nRageZCp1Bh4ud\nF6FsU6K9v92mXcgXYnHUYijkCkSHRLuhh8STsCyLK9dr0V2zqkUnEQhwi1SKxSEh4FFA5zosC1y7\nNvZmce3a5I+NiUFjUBBqz5xBvkwG5oUXXLcU640osCMzZTBwteVGT38Ysj19CQD3QSs1lQvm5s7l\nZuoImYqO/g6UtZXh/LXz0BltjxmJDomGQq7AoshF8BfQwb8EaL5ei67RqhZdEJ+P1RIJlopEEFDp\nEtcYXcaprubeMNTqiR83usE6LY37TyoFcD0rtqAA+Tt3ujawO3bsGD744AO0trYiLi4ODzzwANav\nXz/jDjgLBXbEHjU1jThypA5VVeeRnr4Ya9bMBZ+fiEuXuNMfJjvKKzh4rCxJUhId5TWbOGrPlN6o\nR2VXJZRtSrT0tdi0C3gCLIxcCIVcgVhRLC2jeSlH77Hrul6LrnqCWnQrxWLkUi061zAYuH1y1dXc\nJ/+BgYkfJxAAycncm8X8+TfcYO3Skyfee+89PPvss3j44YexfPlyNDU14b777sMLL7yAH/3oRzPu\nBCHuUFPTiEOHasHn56OpiYeOjnX4858LsHgxEB6eaPN4iWQsmIuPp6O8yPR0D3ZD2abEuY5zGDLY\nTgOHB4VDIVcgMyoTgX6O23dDvJvGYMAxlQrnrGrR8RgGCpEIayQShNBygXNdz2Q1n/totfxt5u8/\nticnJcXle3LsmrGbN28ePv74Y2RmZprvO3/+PL773e+itrbWqR2cLpqxI5PR6YCmJuCtt46isXE9\ntFrL/azBwUexdCk3Gx0aOlZjTi6n0x/I9BhNRlR1V0HZpkSDusGmnc/wkR6RjqXypUiQJNDsHDEb\nNBpxQqPBmb4+GKze0xaFhOAWqRShVIvOeUYzWauqgKtXJz/3MSRkbIl1zpxpLeO4dMaut7cX6enp\nFvelpqZCpbItjEmIpxke5gK5hgagsRFob+e2RFy5woPV9hQAQGAgD2vXcgFdZCQFc2T6VEMqlLWX\noaK9AgN626UaWYAMCrkCWdFZCBZSDRwyZsRkQmlfH4o0GuisatGlBAbiVpkM0f6039Ipenu5JdZJ\nMlnNQkO5T/1paVyleQ95s7ArsMvLy8NTTz2FV199FcHBwejv78fPfvYz5ObmOrt/hEzZ0NBYINfQ\nAHR0TPx3yeONvVjqdIVIS1uHiAggKcmEW25xWXeJl7nZnikTa8LlnstQtilR11sHFpaDj8fwkBqW\nCoVcgWRZMs3O+bip7rEzsizKtVp8o1aj32p2KNbfHxtkMiQ5sDQGwZQzWZGWNnbuowf+/doV2L3z\nzjvYunUrJBIJQkND0dvbi9zcXPzpT39ydv8IuanBQctA7tq1yT9gAdzfYVQUsGXLXJw5U4CIiHy0\ntnLJSjpdAfLzU1zVdeJD+nR9KG8vR3l7Ofp0fTbtYn8xcmJysCRmCcT+dNwIscSyLCoHBnBUrUav\n3rJuYbifH/JlMqQFBdEHAUcZX5C0unrKmayebErlTpqbm9HW1ga5XI74+Hhn9mvG6Egx3zUwwC2p\nNjaOBXI3wjDch6zERC6DNSFhrBZkTU0jCgrqMDLCg1BoQn7+XKSm2iZOEDIRlmVRp6qDsk2Jmu4a\nm9k5BgxSQlOgkCswL2weeAxl3BBLLMuibmgIBWo12q3S8MUCAdZJpciiWnSO4YRMVkcoLCxEYWEh\n9u3b59xyJyzLmj8ZmKzW98fjeWhqICVP+I7+/rEgrrFx8jNYR/F4XCCXlMQFcwkJdOoDmbma2hoc\nKTsCPauH0WhETEIMuvy6oBq23Wsc7BeM7JhsZMdkQxYoc0NviTdo1elwRKXCVasimYF8PlZJJFgm\nEsHPQ99jvcbwMJfBWl3t0ZmsgOPilkkDO5FIBK1WC2Dy4I1hGBgnyxBxMwrsvJdWaxnIddkej2mB\nx+MyVscHclPdU+zq8z+Jd6mprcGBIwcwnDCMi6cvgklioK/VIysjC+HycPPj5kjnQCFXIC08DXwe\nFTckE7+2dI+M4KhajUtWM0Z+PB5WiMXIE4sRQMUxp8+FmayO5PSs2MrKSvO/6+vrZ/xEhEymr28s\nkGtoAHp6bvx4Pn8skEtK4mrK0dFdxJFYlkX3YDea+5rRpGnCH//2R3RFdQGdgHpYDSmkEKQIcLX+\nKuIT4pEVnYUceQ7Cg8JvfnEya/UZDChUq3G2vx8mq1p02SEhWCuVQkS16KZnNJO1qgpoafG6TFZH\nsmuP3euvv46f/vSnNvfv378fTz31lFM6NlM0Y+e5NJqx2biGBu7v8Ub4fO7vb3SPXFwcBXLEsfRG\nPdq0bWjSNKG5rxnNmmaL4sGlJ0sxHGdZG0fsL0aqNhUvP/Qy/PhUR4xMbshoxEmNBqcmqEW3IDgY\n62UyhFEtuqnxsUxWwAVLseONX5YdTyaTeWwtOwrsPIdaPTYb19gI3GzICAS2gRy95hFH6h/pR7Om\n2RzItWvbYWQn31Zy+uRpDMUNQeQvgsRfgqiQKIQIQxDZGYlH7n7EhT0n3qKmvh5fXryI2uFhNAwN\nIWHOHISPSzpMvl6LTk616Oznw5msgIsKFB89ehQsy8JoNOLo0aMWbXV1dRCLKWWfWGJZ20Busr+9\nUQIBt5w6ukcuLo67z5Voj53vYlkWXYNdXBCnaUZzXzN6h24yTQwgyC8ICZIExIvjsU66Dl+c/gKB\nSYFoONuAkKwQ6K7okH9Lvgu+A+Jtzl+5gl+dOoVrixah8/RpSBUKnFUqkQVgcUoKbpXJkEy16Ozj\noZmsnuyGb58//OEPwTAMdDodHnroIfP9DMMgKioKv/nNb5zeQeLZWJZbSh2/R67PtoSXBT+/sUAu\nKYnbL0fbSoij6I16tGpbLQK5YcMER4xYCQ8KNwdyCZIEhAaGjtUMSwBiQmJQUF6A7t5uRHZGIv+W\nfKSmpDr5uyHeRKXX41RfH35bXIy+xYu5GabrRMuWQVpbi39ZvZpq0d2MF2WyeiK7lmK3bduGw4cP\nu6I/DkNLsc7Bslxyw/g9chOs0lsQCm0DOUr4Io6i1WnNSQ7Nmma097fDxE5eogkABDwBYkWxiJfE\nI14cj3hJPIL8glzUY+JLWJZFi06Hkr4+VA0OgmVZlB4/juHFiwEAQh4PSf7+iPb3R+jFi3jijjvc\n3GMP5aWZrI7k0rNivS2oI47DskB3t2Ug199/468RCrkl1dE9cjExPvW3R9zIxJrQNdBl3hvXpGmC\nevgma/3g6solSBIQL+Fm42JCYqgcCZkRE8uianAQJRoNWqwKC/NYFsF8PuL8/RHl52cuLkzzSVYo\nk9Up7ArsNBoN9u7di2+++QY9PT3mgsUMw6CpqcmpHSSuxbJc3bjxe+Qm29Iwyt/fNpDztpqatMfO\nM40YR9Da12qRraoz6m76dZHBkeaZuARJAmQBMoctf9FYmd2GjUaU9/fjVF8fNAaDTXtKYCDyli/H\n0YsX4a9QoKG0FEkrVkBXVob87Gw39NiDsCx3eHd1tc9ksnoiuwK7nTt3orm5Gc8//7x5WfZXv/oV\nvve97zm7f8TJWJY7yWF8IDc4eOOvCQgYC+ISE4HoaO8L5Ihn6tP1mZdUmzRNuDZw7abLqn48P8SK\nY8174+LEcQj0o43pxLFG98+V9/djxOo0JgHDYHFICFaIxYgUCoHoaCT4+6Pg4kV0X72KyJAQ5Gdn\nIzU52U29dyMfz2T1RHbtsYuIiEBVVRXCw8MhkUig0WjQ2tqKO+64A+Xl5a7o55TRHruJmUzch6Tx\nJztYnWZjIzBwLJBLSgIiIymQIzNnYk3oHOi0COQ0Os1Nvy5EGGKR5BAdEk3LqsRp/j97dx7fVJX+\nD/xz0yRN2yRtoRToRoVCKXQHhA4IlU0EFEFROoKU4o6iiPqanwsUmPmiA4K4+xUFpQwOiKMgKl8E\nOiqCLG3ZdyhLKZRCaZO0zXp+f6S5JGnS3rRpmqbP+/XiRW+We0/Sk9sn5zznuZdqa23y56wF+flh\ngEKB/goF5LQC7DbLStbjx815c85GC8RioEcPcyDXzleyAh7OsWOMITg4GIC5pt2tW7fQtWtXnD59\nutkNcFVVVRVGjhyJ48eP488//0SfPn083oa2xGQyj3xbgrgLF8wLjhoSGFg/kKNRcNJcWoMWl6su\n87lxl6suQ2d0stqtDgfOPK1aN6UarYxGiCyEVhWSFtVQ/hwAdJJKkaFUIikoiK7lakErWb2GoMAu\nOTkZv/76K0aMGIEhQ4Zg1qxZCAoKQny855f6BwYG4scff8Qrr7xCI3IOmExAaentqdWLFwEH5yUb\nQUG3p1VjY9tnOgPlTblfZW2lzSKHa+prYGj4MysRSRCljOIDuShlFGRimYdaLAz1Fd+lNZlQoFLh\nz6oq3HKQP9cjIAAZSiV6BAQI/nLh0/1Frb6dL9dOV7J6I0GB3Weffcb/vGLFCrz22muorKzEV199\n1WINc0YsFiMsjK7HaGE01g/knH1RspDLb4/GdesGhIW1v0COuJeJmXBVfZWvG3ex8iKqtI0UNIT5\nslyWKdXo4Gh0DupM06rE427p9fhTpUKBSgWtXf6cn1X+XGcaXaKVrG2AoMCuvLwcAwcOBAB07twZ\nn3/+OQBg7969Ldcy4pDRCJSU3M6Ru3Sp8UBOobgdyMXGmj9v9Bmz5bPfqFtIraHWPK1alxtXoioR\nNK3aWd7ZJpAL9g9uc9Oq1Fd8x+W6/LljDvLnAuvy5wY0M3+uzfcX65Wsx4+bV9s507Xr7WCuPU79\neAlBvXXkyJEOrxU7ZswY3GzsCu5WPvjgA6xevRpHjhxBVlYWVq1axd938+ZNzJw5E9u2bUNYWBgW\nL16MrKwsAMDy5cuxadMmjB8/HnPnzuWf09b+IDSFwWAO5Cw5cpcuAXp9w89RKm0DudBQ+nyRpmOM\n4VbtLb7cyMXKiyjTlDU6rSr1k5qnVa1Wq/qL6bqYpHWZGMOJ6mrsrqrCJQcJx2ESCTKCg5HcnvPn\naCVrm9ZgYGcymfhvMSa74emzZ89C7OK3mMjISLz55pvYunUrauyWYs6aNQsymQxlZWUoLCzEuHHj\nkJKSgj59+mDOnDmYM2dOvf35Yo6dwWAe3bYO5ByketgICbHNkQsJoUDOVT6dB+Mio8lonla1upqD\nStfI5UUABPsH2yxy6CzvDBHne38Yqa+0TVqTCYUqFfY4yZ/rXpc/F+dC/pwQbaa/0EpWn9FgZGYd\nuNkHcSKRCK+//rpLB5s4cSIAYP/+/bh8+TJ/u0ajwbfffoujR48iMDAQgwcPxoQJE7BmzRosXry4\n3n7Gjh2LgwcP4uTJk3jqqacwffp0h8fLzs5GbGwsACAkJASpqan8Byw/Px8AWn178OBMXL4MfPdd\nPq5eBRSKTBgMQHGx+f7YWPPjrbdDQwGVKh9dugAPPZSJkBDz/iorgdBQ73p9bWW7qKjIq9rjye0a\nfQ02/rgRZdVl6JDQASVVJThdYF7xHpsaCwAoLiqut90hoANGDh+JmOAYFBcVQw45Mvvc3v9JnPSK\n10fb7Xu70mDA//74I05VVyOyLqWoeM8eAECPjAwkBQXBVFSEDhIJenpBez26PWgQcPo08jduBC5f\nRmZUlPn+4mLz/XV/P/OvXAEiI5E5eTIQF4f8P/4AKiuRWRfUec3raWPblp+L695vd2mwjp3lYEOH\nDsVvv/3Gj5BxHIdOnTohMLBp11Z84403UFJSwk/FFhYWYsiQIdBYXeJg2bJlyM/Px6ZNm5p0DG+t\nY6fTmUfhLDlyJSXOFxJZdOhgu9ihrvIMIS5jjKGitoKfUr1UdQnXNdcbnVb19/NHlDKKz42LUkZB\n6if1UKsJcV2JVovdlZU4Vl0Nk4P8uf51+XOK9lZ/ztWVrAkJ5j8+tJK1xXmkjp1ltMvdlw2zH+ZW\nq9VQKpU2tykUCod5fW2NTmdeqWqZWi0pMacvNKRjR9tAzu6tIUQwo8mIUnWpTSCn1jVysV8AIbIQ\nm0UO4UHhPjmtSnyLiTGcrMufu+gkf26QUokUubx95c/RStZ2RdBXlWnTptW7zRKcNaXkiX1EKpfL\nUVVlWxqhsrISCoXC5X1by83NRWZmJj/86QlarW0gd+VK44Fcp062l+hq5ssmTZCf30byYBpRo6+x\nyY0rUZXAYGo4SVPEidBF3sUmkFP607cJZ3ylr/gSrcmEIrUae6qqUOFgddkddflzPd2cPydEq/QX\nWsnapuTn59tMzzaXoMCuR48eNkOEV69excaNG/Hoo4826aD2H6xevXrBYDDgzJkziIuLAwAcPHgQ\niYmJTdq/RW5ubrOeL0Rt7e1ArrjYXFOusZHU8PDbQVy3buYRb0JcxRjDzZqbNoHc9errjT5PJpbd\nnlZVRiNSGUnTqqRNqjQYzNdvValQ66D+XGJQEDKUSnTxbwersU0m8x8jyzQrrWRtMywDUAsWLHDL\n/gRdK9aR/fv3Izc3Fz/88IPg5xiNRuj1eixYsAAlJSX47LPPIBaL4efnh6ysLHAch5UrV6KgoADj\nx4/H7t27kZCQ0JTmtViOXU3N7UtzFRebvxQ1dpjOnW0DOVpERJrCYDKgVFXKT6leqrwEjV7T6PNC\nZaE2q1XDg8LbRakg4rsayp8LsKo/5/P5c01ZyRofb75uJPE67opbmhzYGQwGhIaGupQHl5ubi4UL\nF9a7bd68eaioqEBOTg5fx+6tt97ClClTmtI0AO57g6qrbQO5a9caDuQ47nYgFxsLxMTQZ4g0TbW+\n2iY37orqiqBp1a7yrvyUarQyGgp/mtsnbV9j+XMdJRJktIf8OaHXZJXJgJ496ZqsbYhHA7vt27fb\nfMPXaDT4+uuvcfbsWeypWzbubTiOw/z5813OsdNobgdxFy6YA7mGj2NOUbDkyMXEAAEBzWk5aQ2t\nnTfFGMONmhs2gVx5dXmjz5OJZTa5cZGKSEj8JB5ocfvV2n2lvdGZTCj00vw5IdzSX2glq0+z5Ngt\nWLCg5VfFWsycOdPmAxMUFITU1FSsW7eu2Q1oSUJy7NTq24FccTFwvZEUJY4DIiJsAzmZd12jnLQB\nBpMBV1RX+Ny4S1WXUK13Mo1ipUNAB35KNTo4Gp0CO3nlHzNCmquqLn/ugIP8ORHHISkoCIOUSnT1\n1fy5mzdvX/mBVrL6NK/JsfN2zoY0VSrbQK68kUERkcgcyFly5GJiAF89j5CWo9FpbBY5XFFdgZE1\nXMDQj/NDV0VXm0BOLqWVNsS3XdFqsbuqCkc1Gof5c5b6c0pfy5+jlaztnkfq2Fm7desWtmzZgitX\nriAiIgJjx45FaGhosxvQksaOnY4JE+5FRsYUPpi7caPh54hEQGTk7Ry56GhKTSCuYYyhvLqcn1K9\nWHkRN2sav6ZyoCSQD+CildGIUETQtCppF0yM4VRd/twFJ/lzlvpzUl/Kn6OVrATuL3ciaMRux44d\nmDRpEuLj49GtWzdcuHABJ06cwMaNGzFy5Ei3NcadOI7DmDEMKtV2pKbGISysm8PH+fnZBnJRURTI\ntUfNyYPRG/W3p1XrVqvWGGoafV7HgI58blxMcAw6BnSkadU2gHLs3EdnVX/upoP8uViZDBnBwejl\npflzQtTrL7SSlTjh0RG7WbNm4X//93/x8MMP87dt2LABzz33HE6cONHsRrSUmhpALB6B8+d38IGd\nWGwO3iw5clFRgIQGRYgL1Do1P6V6sfIiStWlMLGGq1D7cX6IUETYrFYNklLdG9I+VRkM2FtVhQNq\nNWrsFgKIrOrPteX8uQsnT+LsL7/g0PHjMBUVoUf37uhWWwucOUMrWUmLEjRiFxISghs3bsDPaoWN\nXq9Hp06dcMvZ0HEr4zgOw4YxiERAly75eOqpTHTrZg7kfC01g7QcxhiuV1+3CeQqaisafV6gJJDP\njYsJjkFXRVeIRdTxSPtWWpc/d8RB/pxMJEJ/hQJ3KpVtN3/OaATUalwoKsKZf/0LI4xG8yKIigps\n1+sRl5qKbmFhts+hlaykjkdH7KZNm4YPPvgAL7zwAn/bxx9/7PBSY94kNdV8ndUuXUwYNqy1W0O8\n0ckzJ/HLgV+gZ3pIOAmGpg6FPFzOT6leqrqEWkP9nB97YYFhNoFch4AObXbqiBB3YozhVE0NdldW\nothB/lyHuvy5VG/Nn2PMXDtOrTavvlOrbf9Z31ZjTsE4u3cvRthNsY4Qi7Hj/HlzYEcrWUkLEhTY\nFRQU4JNPPsE///lPREZGoqSkBGVlZRg4cCDuuusuAOZI89dff23RxrqqqCgXEREKPProQ63dFNIK\nGGPQm/QwmAzQG83/G0wG/rZTZ0/hm9+/gaiHCGcLzkIWJ8O/8/6N5IRkhEWEOd2vWCRGpCKSn1KN\nDo5GoITyX9oLyrETRmcy4WBd/twNB/lz3WQyZCiV6BUYCFFrBDYGQ/0gzVGwplY7rxvnhMiqPEv+\nrVvIDAkB5HKIYmOBZ5+llazERqtcK/aJJ57AE0880eBjvHF04uGHh2LEiB6Ij3e8cIJ4jomZHAZX\n1kFXc26zvt1yW2PlRPb+vhfVUdVAOXCr+hZCdCEQ9RDh/LnzNoFdkCTIZpFDV3lX+IlouoQQR1QG\nA/aqVNivUjnMn+tblz8X0RL5c4yZR80aC9SsRtfciuMAuRym0FBAqzXnyAUFAUlJgEwGU3i4+WLh\nhFihOnYCtdS1Yts6xhhMzNQigVRDtzW2uKA17Pl9D2qj6k8NhZWFYfqE6XwgFyoL9covLoR4k1Kt\nFnvq8ueMDvLn+tXlzwU3JX/OenTNUZBmuV2jcXl0TRB/f3MunOWfQmG7bfkXGAiIRLhw8iTOrF6N\nEVbB63atFnHZ2egWH+/+9hGf4PE6dr/++isKCwuh0ZgvOs4YA8dxeO2115rdiPaKMQYjM3osuLLc\nxtB+Al6JSAKxSAyJn/l/sUjM33Y+8Dw0gRqIOBFkYhmC/YOh9FciMjAS98Xf19pNJ8TrNZY/F1qX\nP5fmKH/OMrrWUKBm+dnBvptNJDKPpjUUqCkU5se4uDq1W3w8kJ2NHdu3Q6TTwSSVIm7ECArqiEcI\nCuyef/55rF+/HnfddRcC2tCFUD/894cY2W8k4uMa/zBZgixPBVeW29pLkMWBswmwLMFVU26zvr2h\n2/w4vwZH2v6i+AtW71wN/57+KC4qRsfUjtCe1mLE3SM8+M6QtoZy7AC9Vf25evlzJhNiGEMGxyG+\nqgqi0lLnI22mFhjJ9/dvOFCzHl1rwZH4bvHx6BYfj/z8fAxv5/2FeJagwC4vLw9Hjx5FRERES7fH\nrT7e+DG+/uVrPDDxAYRHhjcasLUXIk7U7KCpsdvsgzMRJ/K66cz4uHhkIxvbC7aj/GY5wsvCMeLu\nEYK+CBDSrjAGVFdDVVmJvRUV2K9Wo6a2FtDrzTXZdDqItFr0razEoPJyRDqr09ZUIlHjgZrlHxUm\nJW1Mq1x5Ijk5GTt27ECYff0dL8ZxHIatMtc4CbochAFDBrRyixwTcaJmB02u3kaJ/4QQAObATEDu\n2lWdDrsVChwJCoLR7guazGRCP5UKd1ZVIdjV/DaZTHjumpd9MSTE3TyaY/f555/jiSeewF//+ld0\n7tzZ5r6hQ4c2uxEtzQhhJxs/zs9t04VCR8NEnBfWbSKEtF11o2uCcte0Wue7AXA6IAC7lUqc79ix\n3v2hej0GVVUhVa2Gv/UfI8voWmPToUFBNLpGSAsQFNgdOHAAP/74I3777bd6OXaXLl1qkYa5Q3zH\neIg4EcKMYZiaPLXB0TAKsto3ypsiQrVaX9HphNVc02ialbum5zgclMuxR6lEuXXgJRYDUiliRCJk\niMWIDwiAyFEAFxBAo2tW6NxCPE1QYPf666/jhx9+wKhRo1q6PW7VVdEV2tNaTL17KuI6xLV2cwgh\nxJbJZB5da6zmWiOja03m58cHZGqFAnuVSuwPDES1RGJeCVr3T+Tvjz5yOQYplYiSydzfDkKI2wjK\nsYuJicGZM2cgbUMXJOY4Dh/++0OMSKdkeEJI81ku6i7S62GSSNBj5Ejn5SusR9caq7vWEvU2AwIa\nz1tTKACZDNf0euyurMRhB/Xn/C315xQKhNC0KSEtyqM5dgsXLsSLL76IN998s16Oncgbr+1Xp+xY\nGUrDSymwI4Q0jjHzCJqDfxdOnDBf1F0iMRfA1emw/cABYMQIdOvQoX4A5+5VoRY44VcAACAASURB\nVMDt0TUhuWuNFAFmjOFMTQ12X7uGcw6uwBAiFpvrzykU8PficzwhvqBVVsU6C944joOxJap8uwFd\neYK4gvJgrDgLcIxGp4GPoH/e/vwGzhc79u7F8LqLuvPX/gSwIygIwwc0c8V9YKCwUh4yWbNz1/Qm\nEw5pNNhdWYlyB9dvja67fmvv1rp+qw+icwsRyqMjdufOnWv2gVrDjg8/bHi6hLR7lum1Q8ePw3T0\nqG1/sQ5wmhtUtIXARkCA015ZX9Td5nZnX2zFYmFlPORy80hcC1MbDNinUmGfSoVquzZzHIc+gYHI\noPw5QnyCS9eKNZlMuHbtGjp37uzVU7BAXeQ7bRq26/WIGz0a3aKjbf9gWX52dJuzn9vzY9tCG118\n7IVr13Bm/36MEIvNtzNm7i+pqejWsSMFOO2Rn5+5XIfdvx1//IHhGo15xEwk4hcV7AgPx/C//rV+\nAOfv7xUrQ6/pdNhTVYVDarXD/Ll0hQIDKX+OEK/grhE7QYFdVVUVnnvuOXz99dcwGAwQi8WYMmUK\n3n//fQQHBze7ES2B4ziwYeYCxW6ZLiE+x3p6zeZ26i9OA5wG/zXlOe56vjuO3UAg1pYu6s4Yw9ma\nGuyuqsJZJ/lzA5VKpFP+HCFexaNTsc8//zw0Gg2OHDmCmJgYXLx4Ea+99hqef/55fPXVV81uREtz\nOl1C2jXr6TXrvCmb/uJqwNDWgxuRyCtGmryN9UXdDx07huQ+fbzuou56kwmHNRrsrqrCdQeLN6L8\n/ZERHIwEyp/zKMqxI54mKLD7+eefce7cOQQFBQEAevXqhdWrV6N79+4t2rhmu+MOAIApJAQYMeL2\nHyzrk1pjP7fWY9tCG73hsc3Yl+nTT4EbN8zbFy8CsbHm28PDgeeeowCH2LBc1F3kZX+o1QYD9tfl\nz2kc5M8l1OXPRVP+HCHtgqDALiAgANevX+cDOwAoLy+HzNtPFN268dMl8KJv1sQ79Bg/Htvrptcy\n48wFrLdrtYgbPZqCOuKUtwR1ZToddldV4bBaDYOT/Lk7FQqEUv5cq/KW/kLaD0GB3eOPP45Ro0Zh\n7ty56NatG4qLi7F8+XI88cQTLd2+Zpm+bx/ufeQRjKCgjjhgPb0m0ulgkkq9bnqNEGuN5c8FW+rP\nyeWQeWC1LSGk+Vqljp3JZMLq1auxdu1alJaWIiIiAllZWcjJyQHnpSMbVMeOuILyYIhQrdFXDHX1\n5/ZUVaHMQf5cpL8//kL5c16Jzi1EKI8unhCJRMjJyUFOTk6zD0gIIUQYjdGIfVVVjebPRfn7e+2X\nbEKIZwkasXv++eeRlZWFv/zlL/xtf/zxB9avX4933323RRvYVDRiRwhpq67X5c8dcpA/JxWJkC6X\nY6BSSflzhPgQj9axCwsLQ0lJCfytajjV1tYiOjoa169fb3YjWgIFdoSQtoQxhnO1tdhdWYkzTvLn\nBiqVSKf8OUJ8krviFkHVKUUiEUx2l9QxmUwUOBGf4c7EVeLb3N1XDCYTClUqfHzlCtZcvVovqIvw\n98dDnTphdlQU/hIcTEFdG0PnFuJpgnLshgwZgjfeeANLliyBSCSC0WjE/Pnzcdddd7V0+wghxCdp\njEbsV6mwt6rKYf5cb0v9OcqfI4S4QNBU7KVLlzB+/HiUlpaiW7duuHjxIrp27YrNmzcjOjraE+10\nGU3FEkK80fW667cedJI/l1aXP9eB8ucIaVc8mmMHAEajEXv37sWlS5cQHR2NgQMHQuTF1xmkwI4Q\n4i0YYzhfW4vdVVU47eD6xEqr/LkAmmolpF3yeGDX1lBgR1xBtaaIUK70FYPJhCN112+95qD+XIS/\nPzKUSvQJCoIfTbf6JDq3EKE8WseOEEKIcNVW+XNqB/lz8QEByAgORgzlzxFC3MynR+zmz5+PzMxM\n+rZECPGI8rr8uSIH+XOSuvy5QZQ/RwixYrmk2IIFCzwzFcsYw/nz5xETEwOxuO0M8NFULCHEExhj\nKK7LnzvlIH9OIRZjoEKBfgoF5c8RQpzyaB27xMREr14oQUhzUa0pIpSlrxgZw0G1Gp9euYIvr16t\nF9R19ffHpE6d8GJUFIaEhFBQ107RuYV4WqNDcBzHIS0tDSdPnkRCQoIn2kQIIV7n5Llz+OXoURw6\neBDflZVBGhGBwMhIm8dwHIdeAQHIUCrRTSaj/DlCiMcJyrF74403kJeXh+zsbERHR/PDhRzHIScn\nxxPtdBlNxRJC3OXE2bP4YN8+3EhKwjW9HibGYNi/H6nx8QiLjubz5wYqlehI+XOEkCbwaLkTy+ID\nR98+d+7c2exGtAQK7AghzVVtNOKQWo3lmzfjWt++9e7vcPgwXrr/fsqfI4Q0m0fLnVCOAPF1VGuK\nWFiKCReoVDheXQ0jY6i0Kllya/9+RA0ciGh/f8SFhmJISEgrtpZ4Ozq3EE8TvMz1xo0b2LJlC65e\nvYpXX30VJSUlYIwhKiqqJdtHCCEeUWUwoFCtRqFKhVsGg819IsYg4jiESyQICwhAX4UCHICA1mkq\nIYQ4JWgq9r///S8efPBB9O/fH7t27YJKpUJ+fj7eeecdbN682RPtdBlNxRJCGmNkDKeqq1GgVuNM\nTY3Dc0akvz863LiBP48fR+CAAfzt2gMHkJ2ejvju3T3ZZEKIj/Jojl1qaiqWLl2KkSNHIjQ0FBUV\nFaitrUVMTAzKysqa3YiWQIEdIcSZcp0OhWo1itRqaOyuDAEAAX5+SA4KQrpCgc5SKQDzqtjtR49C\nB0AKYETfvhTUEULcxqOBnSWYs/7ZaDQiPDwcN27caHYjWgIFdsQVlAfj+3QmE45pNChQq3Gxttbh\nY7oHBCBdLkfvwECIndTupL5CXEH9hQjl0cUTCQkJ+PnnnzFmzBj+tu3btyMpKanZDSCEkJbCGEOp\nTocClQqHNRpoTaZ6j1GKxUiVy5EmlyOUSpUQQto4QSN2e/bswfjx4zF27Fhs2LAB06ZNw+bNm/H9\n99/jzjvv9EQ7XUYjdoS0XzVGIw5pNChQqXBNp6t3v4jjEB8YiHS5HD0CAiCiQsKEkFbm0alYACgp\nKUFeXh4uXLiAmJgYTJ06tdVWxO7duxcvvvgiJBIJIiMj8dVXX9W7ji0FdoS0L5Zrthao1Tiu0cDg\n4PPfUSJBukKBlKAgyNvQta8JIb7P44EdAJhMJpSXl6NTp06teqmcq1evIjQ0FP7+/njttdfQr18/\nPPjggzaPocCOuILyYNquKoMBRWo1CtVqVOj19e6XiEToGxiIdIUC0f7+zT53UV8hrqD+QoTyaI5d\nRUUFZs+ejfXr10Ov10MikWDy5Ml477330KFDh2Y3wlVdunThf5ZIJPCjiu+EtCtGxnC6rkzJaSdl\nSiL8/ZEulyMxKAgyOkcQQtoJQSN2DzzwAMRiMRYtWoSYmBhcvHgR8+bNg06nw/fff++Jdjp04cIF\nZGVl4bfffqsX3NGIHSG+54ZejwKVCgfVaqgdlCmRiURIlsuRLpeji79/K7SQEEKaxqNTscHBwSgt\nLUVgYCB/W3V1Nbp27YrKykqXDvjBBx9g9erVOHLkCLKysrBq1Sr+vps3b2LmzJnYtm0bwsLCsHjx\nYmRlZQEAli9fjk2bNmH8+PGYO3cuqqqqcN9992HlypXo2bNn/RdGgR0hPkFvMuFYdTUKVCpccFKm\n5A6rMiUSJ2VKCCHEm3l0KrZ3794oLi5Gnz59+NsuXLiA3r17u3zAyMhIvPnmm9i6dStqamps7ps1\naxZkMhnKyspQWFiIcePGISUlBX369MGcOXMwZ84cAIDBYMCUKVMwf/58h0EdIa6iPBjvU6rVokCt\nxiG12mGZEoVVmZIOHixTQn2FuIL6C/E0QYHd8OHDMXr0aDz22GOIjo7GxYsXkZeXh2nTpuGLL74A\nYwwcxyEnJ6fRfU2cOBEAsH//fly+fJm/XaPR4Ntvv8XRo0cRGBiIwYMHY8KECVizZg0WL15ss491\n69Zh7969WLRoERYtWoRnnnkGDz/8cL1jZWdnIzY2FgAQEhKC1NRU/gOWn58PALRN2wCAoqIir2pP\ne90eeNddOKzR4OutW3HTYEDsoEEAgOI9ewAA3TMy0CsgAPrCQkT6+2P43Xd7Vftpm7Zpm7aFblt+\nLi4uhjsJmoq1NMZ6NZklmLO2c+dOwQd+4403UFJSwk/FFhYWYsiQIdBoNPxjli1bhvz8fGzatEnw\nfi1oKpaQtsFSpqRQrcYxJ2VKOkgkSJfLkSKXQ0FlSgghPsijU7HW0aW72AeFarUaSqXS5jaFQgGV\nSuX2YxNCWp/KqkzJTQdlSsQch75112uNcUOZEkIIaQ9a7auvfVQql8tRVVVlc1tlZSUUCkWTj5Gb\nm4vMzEx+xJEQZ/Lz86mfeICJMZyuqUGBSoXTNTUwOfh22rWuTEmSl5Ypob5CXEH9hTQmPz/frQNo\nrRbY2X/77tWrFwwGA86cOYO4uDgAwMGDB5GYmNjkY+Tm5janiYQQN7mh16NQpUJRI2VK0uRydKUy\nJYSQdsQyALVgwQK37M+lK0+4g9FohF6vx4IFC1BSUoLPPvsMYrEYfn5+yMrKAsdxWLlyJQoKCjB+\n/Hjs3r0bCQkJLh+HcuwIaV16kwnH68qUFDspUxIrkyFdoUAClSkhhLRzHs2xc6dFixZh4cKF/HZe\nXh5yc3Mxb948fPTRR8jJyUF4eDjCwsLwySefNCmos6CpWEI8z1Km5LBajVoHZUrkfn5IUyg8XqaE\nEEK8kbunYgWP2B0/fhwbNmzAtWvX8OGHH+LEiRPQ6XRITk52W2PciUbsiCsoD6Z5ao1GHNZoUKBW\no1SrrXe/iOPQMyAA6QoFegYEQNSGF0JQXyGuoP5ChHJX3CJo7mPDhg0YOnQoSkpK8NVXXwEAVCoV\nXnrppWY3gBDSNjHGUFxTg/9cv46lly5hy40b9YK6DhIJRoSGYk5UFLI6d0Z8YGCbDuoIIcTbCRqx\n6927N77++mukpqYiNDQUFRUV0Ov16Nq1K8rLyz3RTpfRiB0hLUNtVabkhpMyJX2CgpAul6ObTEZl\nSgghRACP5thdv37d4ZSryMuTnSnHjhD3MDGGM3VlSk45KVPSRSpFukKBpKAgBHhhmRJCCPFGrZJj\nN2rUKEydOhXTp0/nR+zy8vLw9ddf44cffnBbY9yJRuyIKygPxrGbej0K1WoUqdVQGQz17peJREiS\ny5HejsqUUF8hrqD+QoTy6Ijd+++/j1GjRuHzzz9HdXU1Ro8ejVOnTuH//u//mt0AQoh3MVjKlKjV\nOF9T4/Ax3erKlPShMiWEEOJVBK+K1Wg0+OGHH3DhwgXExMRg3LhxzboqREujETtCXHPVUqZEo0GN\ngyLCcj8/pMrlSFMo0JHKlBBCiFt5vI5dUFAQHnnkkWYf0JMox46QhtUajThSV6bkioMyJZylTIlc\njp6BgfCjhRCEEOJWrZJjd+HCBSxYsACFhYVQq9W3n8xxOHXqlNsa4040Ykdc0Z7yYBhjuKjVolCl\nwtHqaugdFBEOlUiQJpcjVS6HUtxqVx70Su2pr5Dmo/5ChPLoiN3kyZORkJCARYsWQSaTNfughBDP\nUxsMOKjRoEClclqmJKGuTEkslSkhhJA2SdCIXXBwMG7evAm/NlTCgEbsCDGXKTlbU4MCtRonq6sd\nlinpXFemJJnKlBBCSKvx6Ijd+PHj8d///hfDhw9v9gEJIS2vwqpMSZWDMiX+IhGSgoKQrlCgq1RK\no3OEEOIjBAV2K1asQEZGBnr16oXw8HD+do7j8MUXX7RY45qLFk8QoXwhD8ZgMuFEXZmSc07KlMTI\nZEiXy9EnKAhSKlPSJL7QV4jnUH8hjXH34glBgV1OTg6kUikSEhIgq8u9YYx5/bf83Nzc1m4CIS3u\nmk6HApUKh5yUKQmylCmRyxEmlbZCCwkhhDhjGYBasGCBW/YnKMdOoVCgpKQESqXSLQf1BMqxI75M\nazKZy5SoVChxUqYkrq5MSS8qU0IIIV7Pozl2ycnJuHHjRpsK7AjxNYwxXNJqUahW44hG47BMSYhY\njHSFgsqUEEJIOyXozD98+HDcc889mDFjBjp37gwA/FRsTk5OizaQEE/w5jwYjdGIg2o1ClQqlDso\nU+LHcUgIDES6QoE7qExJi/PmvkK8D/UX4mmCArvffvsNERERDq8NS4EdIe5nKVNSqFbjhJMyJeFS\nKdLlciTL5QikMiWEEELgwrVi2xrKsSNt0a26MiWFTsqUSK3KlERQmRJCCPEZLZ5jZ73q1eQgl8dC\n5MUlE6jcCWkLDCYTTtbUoEClwrnaWocf7BiZDGlyOfpSmRJCCPEpHrtWrEKhgEqlAuA8eOM4DkYH\n5RW8AY3YEVe0Rh5MmVWZkmonZUpS6sqUdKIyJV6DcqaIK6i/EKFafMTu6NGj/M/nzp1r9oEIIeYy\nJUfrypRcdlKmpIdMhnSFAvFUpoQQQoiLBOXYLV26FC+//HK925ctW4aXXnqpRRrWXDRiR7wFYwyX\ntVoUqNU4qtFA56RMSVpdmZJgKlNCCCHtjrviFsEFii3TstZCQ0NRUVHR7Ea0BArsSGvTGI04pFaj\nQK3GdZ2u3v1+HIfedWVKulOZEkIIadc8UqB4x44dYIzBaDRix44dNvedPXuWChYTn+GuPBgTYzhn\nVabESGVKfA7lTBFXUH8hntZgYJeTkwOO46DVajFz5kz+do7j0LlzZ7z//vst3kBC2oJKgwGFKhUK\n1WpUOilTkhgUhHS5HJH+/jQ6RwghpEUImoqdNm0a1qxZ44n2uA1NxZKWZmQMJ6urUaBS4ayTMiXR\nVmVK/KlMCSGEECc8eq3YthbUWVAdO9ISrut0KFCrcVCtdlimJLCuTEk6lSkhhBDSCI/VsWvraMSO\nuKKxPBidpUyJWo1LtbX17uc4Dt0tZUoCAiCm0TmfRTlTxBXUX4hQHh2xI6Q9YoyhpK5MyREnZUqC\nxWKkyeVIlcsRIpG0QisJIYSQ22jEjhA71VZlSsqclCmJDwxEulyO7gEBENFCCEIIIc1EI3aEuMHJ\nc+fwy9Gj0AGo1OsRGh2Nqk6dHJYp6WRVpiSIypQQQgjxQpQIRNqtw2fOYMWff2JfXBzWXbuG/Dvu\nwIaiIly7eJF/jFQkQppCgZldu+LZiAhkBAdTUNfOuTPJmfg+6i/E02jEjrQLjDHc0OtxWavl/23e\ntQua5GSgthY6xhAIQNy/P84fPIjUuDikKxRUpoQQQkibQjl2xCfVGo0o0elwWavFpdpalOh0qLEr\nTbLn119Rm5zMb4s5Dl2kUsSfOoU3H3jA000mhBDSjlGOHSF1TIzhut1onKNrs9rzAxDk5welnx9C\nJRKEicUQcRw60lQrIYSQNooCO9LmaIxGmyCuRKt1WIrEXqCfH6L8/fl/kwYPxrqiIvj364fiPXsg\nGjQI2gMHMCI93QOvgrRVVJeMuIL6C/E0CuyIVzMyhqt1U6qWfxV6faPPE9VNq1oHcqFise01WuPi\n4C8SYfuRIyg/fx7hcjlGpKcjvnv3FnxFhBBCSMvx6Ry7+fPn0yXF2phKg8EmiCvVamEQ0EUVYjGi\nrYK4rlIpJLTogRBCiJezXFJswYIFbsmx8+nAzkdfms/Qm0wotRuNqzIYGn2emOPQ1SqIi/L3h9LP\nz3Y0jhBCCGlDaPEEaVMYY6iwG427qtPBJKATh0okNkFcF6kUfm4O4igPhghFfYW4gvoL8TQK7EiL\n0JpMKLEK4i5rtai2KzfiiFQkQqTdaBwVBCaEEEKEoalY0mzMUbkRvV7Q+9/JboFDJ4mErr1KCCGk\n3aGpWNJqqh2UG9EKKDcS4OeHSKkU0TIZovz9ESmVQkajcYQQQojbUGBHGmRkDGV2CxxuCCg3wnEc\nOtvlxnWUSLx2gQPlwRChqK8QV1B/IZ5GgR2xobJb4HBFp4NewGic3K74b4S/P6RUboQQQgjxKMqx\na8cMDsqNVAooN+LHcehqlxsXbF/8lxBCCCGCUY4dcQljDLcclBsxCuhEIWJxvXIjYhqNI4QQQrwO\nBXY+Sueg3IhGQLkRiUiECLvROIXY97sJ5cEQoaivEFdQfyGe5vt/sdsBxhhu2JUbuSaw3EhHuwUO\n4S1Q/JcQQgghnkE5dm1QjdFYbzSuVsACB3+RyCaIi/T3RyCVGyGEEEJaXbvNsbt27RomTZoEqVQK\nqVSKf/3rX+jYsWNrN6vFmByUGykXWG4k3G40LsyLy40QQgghpPna3IidyWSCqC5x/8svv0RpaSn+\n9re/1XtcWx2xUzsoN6ITMBoXaFVuJLqu3Ig/LXAQjPJgiFDUV4grqL8QodrtiJ3IKlipqqpCaGho\nK7ameYyM4apOh0u1tXwgd0tAuRERx6GL3QKHUCo3QgghhLR7bW7EDgAOHjyIJ598Erdu3cK+ffug\nVCrrPcbbRuwYY6iyuxRXqVYLg4A2Ku3KjXSVSiGh0ThCCCHEZ7grbvFoYPfBBx9g9erVOHLkCLKy\nsrBq1Sr+vps3b2LmzJnYtm0bwsLCsHjxYmRlZQEAli9fjk2bNmH8+PGYO3cu/5wNGzZg7969WLJk\nSb1jtXZgp3NQ/FclYDROzHGIsAriovz9oWwH5UYIIYSQ9qxNTsVGRkbizTffxNatW1FTU2Nz36xZ\nsyCTyVBWVobCwkKMGzcOKSkp6NOnD+bMmYM5c+YAAPR6PSQSCQBAqVRCq9V68iU4xBjDTbvcuGs6\nHUwCfkEd7BY4dKZyI62C8mCIUNRXiCuovxBP82hgN3HiRADA/v37cfnyZf52jUaDb7/9FkePHkVg\nYCAGDx6MCRMmYM2aNVi8eLHNPoqKivDyyy/Dz88PEokEn3/+udPjZWdnIzY2FgAQEhKC1NRU/gOW\nn58PAE3arjUasfGXX3Bdr0enAQNwWavF8V27AACxgwYBAIr37Km3LeY43JWZiSh/f1z580+ESSS4\nd8QIfv+nAES4oX207fp2UVGRV7WHtmmbtmmbtn172/JzcXEx3KlVcuzeeOMNlJSU8FOxhYWFGDJk\nCDQaDf+YZcuWIT8/H5s2bWrSMdw1pGliDOV1xX8vWZUbEbLvTlLbBQ6dJBKIaDSOEEIIIXba5FSs\nhf3qTbVaXW8BhEKhgEql8mSzAAAau+K/JVottALKjQRYlRuJ8vdHpFQKGRX/JYQQQogHtUpgZx+R\nyuVyVFVV2dxWWVkJhULRrOPk5uYiMzOTH/60Z2QM1+wWONwUUPxXxHHobDca14HKjbRp+fn5TvsJ\nIdaorxBXUH8hjcnPz7eZnm0urxix69WrFwwGA86cOYO4uDgA5pImiYmJzTpObm6uzXaVffFfgeVG\n5H5+iJbJbMqNSEWiZrWNEEIIIcQyALVgwQK37M+jOXZGoxF6vR4LFixASUkJPvvsM4jFYvj5+SEr\nKwscx2HlypUoKCjA+PHjsXv3biQkJDTpWBzHYeHGjYiLi4Nfly64rNWiSkC5ET+OQ1e70bhgGo0j\nhBBCSAtqkzl2ixYtwsKFC/ntvLw85ObmYt68efjoo4+Qk5OD8PBwhIWF4ZNPPmlyUGfx+datUPzx\nB4bdfz/CoqMdPibErvhvF6kUYhqNI4QQQogHuHsqtk1eeUIIjuMwrKAAABB08CAGDBsGiUiESLvR\nODkV/yWgPBgiHPUV4grqL0SoNjli52mBIhGUYjEigoLwdEQEwqVSKjdCCCGEEJ/l0yN288+dAwCE\nHzmCZ++7r5VbRAghhBDiGI3YCZD/7ruI6NABWdOmtXZTCCGEEELqoRw7gTiOw4ebNmFE376I7969\ntZtDvBzlwRChqK8QV1B/IULRiJ0ANP1KCCGEkPbEp0fsfPSlEUIIIcTHuCtu8emCbbm5uW6dtyaE\nEEIIcaf8/Px6V8pqDhqxIwSUB0OEo75CXEH9hQhFI3aEEEIIIcQGjdgRQgghhLQyGrEjhBBCCCE2\nfDqwo8UTRCjqJ0Qo6ivEFdRfSGPcvXjCp+vYufONIoQQQghxt8zMTGRmZmLBggVu2R/l2BFCCCGE\ntDLKsSOEEEIIITYosCMElAdDhKO+QlxB/YV4GgV2hBBCCCE+wqdz7ObPn88nJRJCCCGEeJv8/Hzk\n5+djwYIFbsmx8+nAzkdfGiGEEEJ8DC2eIMSNKA+GCEV9hbiC+gvxNArsCCGEEEJ8BE3FEkIIIYS0\nMpqKJYQQQgghNiiwIwSUB0OEo75CXEH9hXgaBXaEEEIIIT7Cp3PsqI4dIYQQQrwZ1bETiBZPEEII\nIaStoMUThLgR5cEQoaivEFdQfyGeRoEdIYQQQoiPoKlYQgghhJBWRlOxhBBCCCHEBgV2hIDyYIhw\n1FeIK6i/EE+jwI4QQgghxEdQjh0hhBBCSCujHLtWtnnzZrz66qut3Yx25fDhwxg/fjwAoKysDBkZ\nGRS8N8Onn36Kd99916XnPProo4iMjIRIJEJ1dbXTx508eRLDhw9HSkoKUlJS8Msvv/D35eXlITk5\nGRKJBB9++GGT2r569WqcPn3a5ecxxpCRkYHU1FSkpKRg5MiROHPmDADgjz/+QFpaGv8vMjIS/fr1\n45978+ZNZGVlIT4+HomJiVi0aJHDY1RXV+ORRx5Bz549kZCQgC1btvD3NfS+CHX69GmkpaWhX79+\nWLduncvPv3DhAj777DOXn+fMu+++i+vXr/Pbubm5eOWVV9y2f6H++9//Ytu2bYIee/DgQWzYsMHm\ntrS0NGi1WqfPqaysxD//+U+b25544gns2rXL9cZ6oYcffhh79+4FALzyyiv13h93y87O5j//Qs5F\n33//Pfbt29eibfIZzEf58EtrtyZOnMjy8/P57WeeeYZ9/fXXbtn3zp073bKf5jAYDDbber2+lVri\n3M6dO1lZWRnjOI5pNBqnj8vIyGB5eXmMMcZOnz7NoqKiWHV1NWOMsSNHjrBjx46xxx57jH344YdN\nasewYcPYDz/80KTnVlVV8T+vWLGC3XfffQ4f98ADD7B33nmH377vvvvY+9UDZAAAIABJREFUihUr\n+L5y9epVh89bsGABe/LJJxlj5tfepUsX/r1q6H0R6q233mKzZs1y6TnWdu7cyfr379+k59r3UcYY\ni42NZUeOHOG3c3Nz2csvv9zk9jXV/PnzBR931apV7KGHHnJp/+fPn2dhYWEut8sbzi2NKSoqYpmZ\nmfz2lStXWGJiYoseMzs726XP//Tp09kHH3zQgi1qfe6KW3w2+nHlDbL/wFpvX7t2jY0YMYIlJSWx\npKQk9tJLLzHGbE8MO3fuZCkpKeypp55iycnJLCUlhR0/fpzf32uvvcbi4uLYwIED2auvvur0pDps\n2DD2yiuvsCFDhrDu3buzv/3tb/x93bp1Y0ePHnW43a1bN/bGG2+wjIwMFh0dzfLy8tjSpUvZgAED\nWFxcHPv111/519WxY0c2d+5clpyczJKSkthvv/3GGGNs1qxZbMmSJfz+CwoKWHx8vE37hB6HMca2\nbNnCBg8ezPr168cyMjLYnj17GGOMlZaWsrvvvpv169eP9e3bl7366qv8c+bPn8+mTJnCxo4dy3r3\n7s3GjRvH/9G7du0ai46OtmlPfn4+GzVqlMP30mAwsLlz57LExESWmJjIXn75ZWY0Ghlj5hPE008/\nzYYPH8569uzJHnvsMYcnX8v79frrr7O0tDQWHx/Pfv/990bfS3uW9ygtLY1lZGSwoqIi/j6O41hu\nbi4bMGAAe/PNN1l2djabOXMmu+uuu1haWhpjjLG//vWvrH///iwpKYlNnDiRVVRUMMYYGzduHNuw\nYQO/r40bN7LRo0czxsx/XHv37s1SU1NZWloau3XrVr12Wf8h3LVrF0tPT2epqamsb9++bN26dQ5f\ni3W7GwrsgoKCWHl5Ob+dnJzMNm7caPOY7OxsmxN1dXU1S05OZt9//z1jjLHt27ez3r17M7VabfO8\nL774gsnlcta9e3eWmprKtm/fzoxGo9Pfd0MWLlzIZs6cWe/2a9euscDAQFZWVsYYY+zUqVMsNjaW\nMdb4H+q+ffuyAwcO8Nvjx4/nf0/O3peamhpBrz0vL4916dKFhYeHs9TUVHb27Fmn/Uuj0bCHHnqI\n9enTh6WkpLBHHnmEMcZYnz59WGBgIEtNTWWTJ09mjDF24sQJdu+997IBAwawlJQUtmrVKv6Y1n10\n3rx5Nu35+9//zqRSKd/Xjh07xnJzc1lWVpbDz/Evv/zCMjIyWFpaGktKSrL5YtbQ+c/aiRMn2KBB\ng1hKSgpLTExkS5cuZYcPH7Z5X95++21mMBjYPffcw/r378/69u3LZsyYwXQ6HSsvL2cxMTEsJCSE\npaamshdeeIF/nRqNhhmNRvbMM8+w3r17s5SUFDZkyBDGGGNjx45lYrGYpaamssGDB/NttnzBuHXr\nFpsxYwZLSkpiKSkp7Pnnn2eMMbZo0SKWlJTEUlNTWWJios2XU4tVq1axUaNGscmTJ7PevXuz4cOH\ns0OHDrExY8awXr16sUcffZR/bGVlJZs5cya78847WXJyMnvhhRf4vt7YueZ//ud/2IABA1j37t1t\nPo/PPfcc+/TTT23alJmZyXbt2uXwd7B37142aNAglpyczDIyMti+ffsYYw2fM+1ZB3aNnYu2bt3K\nOnTowKKiolhqaipbs2aNw37Q1lFg1wgAbP78+YK+LTUU2C1btow99dRT/H2WP5D2gZ1EIuE/RP/4\nxz/4D+KmTZtYSkoKq66uZiaTiU2aNIkNGDDAYTsyMzPZlClTGGPmD29YWBg7c+YMY8z8rdg6sLPe\njo2N5QOkffv2sYCAAPbRRx8xxhhbv349f2I6f/484ziOrVmzhjFmDoyioqKYTqdjx48fZ3Fxcfz+\nc3Jy2HvvvWfTPqHHOXPmDMvIyOBHRo4cOcJiYmIYY4zV1tbyf6x0Oh0bPnw4+/nnnxlj5g93z549\nWWVlJWOMsdGjR7PPPvuM3//EiRNt2lNdXc3kcrnDka2PPvqIjRw5kun1eqbT6diIESPYxx9/zBgz\nB3Z33XUX02q1TKfTsb59+7Jt27bV24fl/dqyZQtjjLG1a9fyJ/SG3kt7169f53/etm0bGzRoEL/N\ncRz75z//yW9Pnz6dDRgwwGYUxzoQeP311/k/eD///DO7++67+fuGDx/ONm3axG7cuMFCQkJYbW0t\nY4wxtVrtcKQlNzeXvfLKK4wxxu6//36bYM5RIGitscBu6NChbMWKFYwxc1/x9/dny5cvt3mMfWDH\nmPmPdkxMDPvzzz/ZHXfcYfOHyVpmZib/e2Gs4d+3I/feey/r0qUL6927Nx+8WVuyZIlNf/vuu+/Y\nkCFD2MyZM1l6ejobO3aszefRmkKhsPmdPfvss/xrb+h9EfrarX9vjDnvX99++y275557+Pssv9P8\n/HybL5d6vZ6lp6ezEydOMMbMI5q9evViJ0+eZIzV76P27M9NDX2OKyoq+CDk6tWrLCoqim9XQ+c/\na7Nnz2aLFy+u97rs3xfGGLtx4wZjjDGTycQee+wx9sknnzDGGFu9enW9ETtLny4oKGAJCQn19l9c\nXFxvxM66H2ZnZ7PZs2fXO3ZKSgr/xdZkMtmMGFusWrWKhYaGspKSEsaY+ctAcnIyq6ysZAaDgSUn\nJ7NffvmFMcbYzJkz+fOO0WhkU6ZM4d/fxs41lkBq165dLDIykr+vb9++7NChQzZteu2119jChQvr\ntVWr1bLo6Gi2Y8cOxpg5WI+JiWF6vb7Bc6Y968BOyLnIfoTPvh9YvvC2RTt37mTz5893W2Dn0zl2\nubm5yMzMbNY+MjIy8NNPP+HVV1/Fli1bEBQU5PBx8fHxSElJAQAMHDgQZ8+eBQDs3LkTjzzyCAIC\nAsBxHKZPn95gXtjkyZMBAEqlEgkJCfx+GvPII48AMOeJ1NbW8tvp6el8DhEASKVSTJ06FQAwbNgw\nBAQE4OTJk+jduze6d++On3/+GRUVFdi8eTOys7ObdJytW7fi7NmzGDp0KNLS0jB16lQYjUZcv34d\nBoMBL7/8MlJTU9G/f38cOXIEBw8e5Pc/ZswYKJXKeu/j+fPnERkZadOWgIAABAYG4sqVK/XauX37\ndsyYMQNisRgSiQQzZszg85k4jsMDDzwAqVQKiUSC9PR0p++zXC7H2LFj67WnoffS3v79+zF06FAk\nJSVh7ty5KCoqsrl/+vTp/M8cx+Ghhx5CQEAAf9uXX36J/v37Izk5GevWreOfP3r0aJSWluLEiRM4\nfvw4zp07h/HjxyM4OBhxcXGYNm0aVq5cCZVKBT8/P4evz9IXhw8fjr///e/4xz/+gb179yI4ONjh\n44VavXo1duzYgbS0NCxfvhxDhgyBWCxu9Hnx8fFYuHAh/vKXv2Du3Ln8Z6qhtgMN/74d+fHHH3Hl\nyhVMmzYN06ZNq3f/qlWrkJOTw28bjUbs2bMHM2bMwIEDB/D444/j/vvvb/T12Le1ofelqa/dWf9K\nTU3F8ePH8dxzz+Gbb76BVCqt91wAOHXqFE6cOIEpU6YgLS0NQ4cOhV6vx/Hjx/nHWPfRxnAc5/Rz\nXFZWhgcffBBJSUkYM2YMbt68afOZsT//WZ+7LIYNG4aVK1di3rx52Llzp01ftX5tJpMJS5YsQVpa\nGlJSUrBjxw7+XNPQObh79+7Q6/XIyclBXl4e/9iGngMAW7Zssckt7NChAwDzZ+vFF1/E0qVLcezY\nMSgUCofPHzx4MCIiIgCYz6/Dhg2DUqmEn58fUlJS+Pdw06ZN/Ovq168fCgoK+HzTxs41U6ZMAWD+\nnVy5cgU6nQ6A4/NrVFQUzp07V6+dJ0+ehL+/P+6++24AwIgRIyCVSvnfY0PnzIYIORdZ/w7s+0FI\nSIig43ijzMxM5Obmum1/Ph3YCSUWi2Eymfjt2tpa/udBgwahqKgI/fr1w5o1a/jObE8mk/E/+/n5\nwWAwAKi/yqWxk4Oz/TTURuvnWf6AW29b9tFYG2bPno2PPvoIX3zxBR588EGHJyChxxkzZgwKCwv5\nf5cvX0anTp2wbNky3Lp1C3v37sXBgwfxwAMP8K+F4zj4+/s7fP0cxzlsc0OriOzfd+t92B/n2LFj\nDvfhrD2OjuGITqfDQw89hPfeew+HDx/GTz/9VC9BWy6X22xbf3n47bff8Mknn2Dr1q04dOgQFi1a\nZPN+Pffcc/jwww/x8ccf4+mnnwbHcfDz88OePXvw3HPP4fLly+jXrx8OHz7cYDtfeOEFbN68GZ06\ndcLzzz+PN998s8HHN+aOO+7Ad999h8LCQqxduxalpaXo06dPvcc5+r0eOHAAnTt3xqVLlxo8hv1z\nXfmcWZ6fk5NTL/l9z549qKio4P84AUC3bt0QExODwYMHIz8/HxMnTkRpaSlu3rxZb78xMTEoLi7m\nty9cuICYmBgAjb8vQl+7RUP964477sCxY8cwatQo/PLLL0hJSXG4OIAxhrCwMJvP67lz5zBhwgT+\nMfZ9tDH2nxuj0QgAeOaZZzB8+HAcPnwYhYWFiIqKsjmX2Z//LM+zNmnSJPz+++/o0aMH3nrrLT4w\nt/+dr127Frt27cLvv/+OQ4cO4dlnn0VNTU2jbQ8ODsbRo0cxZcoUHDp0CH379kVZWZmg1+2o391/\n//1YuXIlpFIpJk+ejJUrVzp8rv1rb+jc8/333/O/q5MnT+Ltt98WdK6xP3/bn8+EvJ7GNHbObExD\n5yLrz7yzfkAosAMAdOnSBXq9nv9m8a9//Yu/r7i4GHK5HI888gjeeecdHDhwwKV9Z2Zm4ptvvkFN\nTQ1MJhPWrFnjNEgBnH+Q4uLi+BVL27dvx7Vr11xqh4VOp+Nf32+//Yba2lr07t0bADB27FicPHkS\ny5cvx6xZs5q0fwAYNWoUfv75Z5tgybKaqbKyEl27doVUKkVJSQm+//57/jH2r916OzY2FiUlJTb3\n19TUQK1W899yrY0cORJffvklDAYD9Ho9vvzyS4waNarJr8mRht5Li9raWhiNRkRFRQEAPvroI5eO\nUVlZieDgYHTo0AFarRZffPGFzf3Tp0/Hd999h/Xr1+Pxxx8HAKjVapSVlWHo0KHIzc1FYmIijh49\nWm/f1u/vqVOncMcdd+DJJ5/E7NmzG1x9JmQE4/r16zajVDKZDMOHD6+3H/t9/Oc//8GuXbtw5MgR\n/PDDD/j5558d7l+pVOLWrVv8tqPf9+jRo+s9r7y8HOXl5fz2hg0bcOedd9o85osvvsBjjz0Gkej2\n6TE9PR1BQUF8n/7111/RsWNHflTG2uTJk/Hpp58CMK9g3b9/P8aMGdPo+yL0tVu/Zw31r5KSEnAc\nhwkTJmDZsmW4fv06KioqoFQqUVlZyT8uPj4egYGByMvL4287ceIEVCqVw+Pbs/9dOPocW26rrKxE\nt27dAADbtm2rNyInJJA4e/YswsPDMX36dMybN4/vq8HBwTavq7KyEmFhYQgKCkJlZSXWrl3Ln3vt\nH2utvLwcGo0Go0ePxuLFixEcHIxz585BqVSiurraYbAJAOPHj8eSJUv47Rs3bgAALl68iL59+2L2\n7NmYOnUq9u/f3+hrbOh9uP/++7F48WL+i355eTmKi4ubda6JjY3F5cuXbW67fPkyunfvXu+x8fHx\n0Ol0fOHlHTt2wGAwID4+XvDxLBydS5ydi+z7mX0/sPx9JEDjcyPtgFgsxooVKzBq1Ch06tQJ48aN\n408AO3fuxPLly+Hn5weTycSfsDmOswnQ7H+2bN933334448/kJycjA4dOmDQoEE2ndOes6Bv0aJF\nmD59Ot5//30MHz6cPzkK2Yf1dseOHVFUVMQv21+3bh0/FcRxHB577DFs3boViYmJuHLlCsaNG4fC\nwkKXjtOzZ0/k5eVh5syZqKmpgU6nw5AhQzBgwADMnj0bkydPRlJSEqKiojBy5Eib5zt7H++66y68\n/PLLNsfbu3cvMjIyIJFI6rXtySefxJkzZ5CWlgbAPIL4xBNPOG27s5NSU9/LtLQ0/PTTT+jSpQsW\nLlyIAQMGoGPHjnjooYec9htHt40ZMwZ5eXno1asXwsLCMHToUJugSy6X495770VtbS06duwIwPwH\n7cEHH+S/TPTr1w+TJk1yeBzLsd5//33s3LkTUqkUMpkM77//vsP3Y9KkSdi3bx84jkN8fDySkpLw\n008/1XvNmzZtwttvvw2O4xAXF4f//Oc//D7WrVuHV199FRUVFdi0aRPeeustbNu2DTKZDC+88AJ2\n7NiB0NBQ/Pvf/8bYsWOxe/fuesH7k08+iblz52LJkiV45513Gv19W1y9ehXZ2dnQ6/UAzH31yy+/\n5O+vqanB+vXr6/2R4DgOq1atwowZM6DVahEUFIRvv/2Wv9/6tb/yyivIzs5Gz5494efnh88++4wf\nhXX2vhQXFwt+7da/N6VS6bR/HTp0CP/v//0/AOap5Ndeew1dunRBp06d+N9dQkIC1q9fj82bN+PF\nF1/EkiVLYDQa0aVLF6xfv54/XkNmz56NGTNmICgoiA+enH2O33rrLTz77LOYP38+BgwYUG+6ubFj\nAcD69euxdu1aSKVScByHFStWAAAmTpyIr776CmlpacjKysLTTz+N77//HgkJCQgPD8ewYcP4EbsR\nI0Zg6dKlSE1NRWZmJt59913+2BcvXsSTTz4Jg8EAg8GAsWPHYtCgQQDM5X6SkpLQoUMH/P777zbt\nWr58OV588UUkJiZCLBbz+/3uu++wZMkSiMVihIaG4vPPPwcAzJ8/HxEREXjqqacafM/svfvuu3j1\n1VeRkpLCz3KsWLECsbGxLp1rrLfvvvtu7NmzB8nJyfxtu3fvdljSRyqVYuPGjZg9ezY0Gg3kcjm+\n+eYbm78jzo5jz3KfkHPRtGnTkJ2djQ0bNuCll17C5cuXbfrBe++95/Q47Q0VKPYAtVoNuVwOk8mE\nxx9/HFFRUVi4cKHH21FcXIwBAwbY1JyyN2rUKDz99NN48MEHPdgyYR544AHMmTMHw4YNAwA8++yz\nGDp0KJ834klC3ktPMBgMSElJwVdffWVTc40QQoQqKirCSy+9hB07dgAASktLMXLkSIcj/aTlUIHi\nNuSxxx5Deno6+vbtC71e36qFjZ19e9q/fz/i4uIQGhrqlUEdYB61XLp0KQBzEnZhYSG/eKO5mnI9\nRyGjCy1p06ZNiIuLwz333ENBnQfRtT+JK9pCf0lNTUVYWBg/G7Bs2TK3JvMTz6IRO0JgPvk2dwU1\naR+orxBXUH8hQrkrbqHAjhBCCCGkldFULCGEEEIIsUGBHSFoG3kwxDtQXyGuoP5CPI0CO0IIIYQQ\nH9Fmc+zWrVuHF154wWlFcMqxI4QQQkhb0a5z7IxGIzZs2MBfpocQQgghhLTRwG7dunV4+OGHW72O\nGPEdlAdDhKK+QlxB/YV4WpsL7Cyjde4qTEsIYK68TogQ1FeIK6i/EE/zaGD3wQcfoH///pDJZJgx\nY4bNfTdv3sTEiRMhl8sRGxuLdevW8fctW7YMd999N5YuXYq1a9fSaB1xu4au30uINeorxBXUX4in\neTSwi4yMxJtvvomcnJx6982aNQsymQxlZWVYu3YtnnnmGRw7dgwA8NJLL2Hnzp14+eWXcezYMXz1\n1Ve49957cfr0abz44ouefAlu54lhencco6n7cOV5Qh7b2GMaut8XpkRa+jW4a/9N2Y+7+4qQx/ly\nf6Fzi2uPbc99BaBzi6uP9eb+4tHAbuLEiZgwYQI6duxoc7tGo8G3336LRYsWITAwEIMHD8aECROw\nZs2aevt46623sHXrVvz000/o1asX3n33XU81v0XQyde1x7bUh6m4uLjRY3sDOvm69tiW6C/UV9x7\nDDq3eAc6t7j2WG8O7Fql3Mkbb7yBkpISrFq1CgBQWFiIIUOGQKPR8I9ZtmwZ8vPzsWnTpiYdIy4u\nDmfPnnVLewkhhBBCWlKPHj1w5syZZu9H7Ia2uMw+P06tVkOpVNrcplAooFKpmnwMd7w5hBBCCCFt\nSausirUfJJTL5aiqqrK5rbKyEgqFwpPNIoQQQghp01olsLMfsevVqxcMBoPNKNvBgweRmJjo6aYR\nQgghhLRZHg3sjEYjamtrYTAYYDQaodVqYTQaERQUhEmTJmHevHmorq7G77//js2bN2PatGmebB4h\nhBBCSJvm0cDOsur17bffRl5eHgICAvCPf/wDAPDRRx+hpqYG4eHhmDp1Kj755BMkJCR4snmEEEII\nIW1aq6yKbS1VVf+/vfuPaeL84wD+Lj9KwVItSvhloC7ADDoxc7igEzHqlkadSrYJRhDjhGxqpn+o\nMyi4qDFGTVyi041/kBCKmCxZ+JGAkZ8zs0hSiZlggUTmMgMODG1Fa4HbHwv9WoqzRb/ttX2/kibc\n9bl7Pnf5BD48d8+dAWvWrEFnZye0Wi2SkpLcHRKJWFtbG/bt24fAwEDExMSgtLQUAQFumW9EItff\n34+MjAxIpVJIpVKUl5fbPdaJaDKNRoNvvvkGAwMD7g6FROrBgwdISUnBwoULIZFIUFlZiTlz5vzn\nNh73SrE3ERISgtraWnz22Wd2EziIJouNjUVjYyOam5uhUqnwyy+/uDskEqnw8HDcvHkTjY2N2Lp1\nK4qLi90dEoncxOsxY2Nj3R0KiVx6ejoaGxvR0NDw2qIO8LHCLiAgwKGTQgQAkZGRCAoKAgAEBgbC\n39/fzRGRWPn5/e9XqcFggFKpdGM05Ak0Gg1fj0kOuXnzJtLS0lBQUOBQe58q7Iimo6+vD9evX8eG\nDRvcHQqJWEdHBz788ENcuHABWVlZ7g6HRGxitG7Lli3uDoVELjo6Gr29vWhpacHAwAB+/vnn127j\nkYXdhQsX8MEHH0Amk2HHjh023w0NDWHz5s2Qy+VQqVTQaDRT7oP/JfmON8kXg8GAnJwcXLlyhSN2\nPuBNciU5ORlarRYnTpzA8ePHXRk2ucl086WsrIyjdT5murkilUoRHBwMAMjIyEBHR8dr+/LIO8Fj\nYmJw9OhR1NXV4dmzZzbf7d69GzKZDAMDA9DpdFi3bh2Sk5PtJkrwHjvfMd18GR0dRWZmJoqKipCQ\nkOCm6MmVppsrFosFgYGBAACFQgGz2eyO8MnFppsvnZ2d0Ol0KCsrQ3d3N/bt2+fx7z2n/zbdXDGZ\nTJDL5QCAlpYWLFiw4PWdCR7syJEjQm5urnXZZDIJUqlU6O7utq7LyckRvv32W+uyWq0WoqOjhdTU\nVKGkpMSl8ZJ7OZsvpaWlwuzZs4X09HQhPT1duHr1qstjJvdwNle0Wq2QlpYmrFq1Svj444+Fhw8f\nujxmcp/p/C2akJKS4pIYSRyczZXa2lphyZIlwooVK4Tt27cLY2Njr+3DI0fsJgiTRt30ej0CAgIQ\nHx9vXZecnIympibrcm1travCI5FxNl+ys7P5kGwf5WyuLF26FM3Nza4MkURkOn+LJrS1tf2/wyMR\ncTZX1Go11Gq1U3145D12Eybfn2AymaBQKGzWhYaGwmg0ujIsEinmCzmKuULOYL6Qo1yRKx5d2E2u\nfOVyOQwGg8264eFhhIaGujIsEinmCzmKuULOYL6Qo1yRKx5d2E2ufBMTEzE6Ooqenh7ruo6ODixc\nuNDVoZEIMV/IUcwVcgbzhRzlilzxyMJubGwMz58/x+joKMbGxmA2mzE2NoYZM2YgIyMDhYWFGBkZ\nwa+//oqqqireJ+XjmC/kKOYKOYP5Qo5yaa68rZkerlRUVCRIJBKbz3fffScIgiAMDQ0JmzZtEmbM\nmCHExcUJGo3GzdGSuzFfyFHMFXIG84Uc5cpckQgCH+hGRERE5A088lIsEREREdljYUdERETkJVjY\nEREREXkJFnZEREREXoKFHREREZGXYGFHRERE5CVY2BERERF5CRZ2RERERF6ChR0R0SS5ubk4evTo\nW93nV199hRMnTrzVfRIRTRbg7gCIiMRGIpHYvaz7TV26dOmt7o+IaCocsSMimgLftkhEnoiFHRGJ\nyunTpzF37lwoFArMnz8fDQ0NAIC2tjakpqZCqVQiOjoae/fuhcVisW7n5+eHS5cuISEhAQqFAoWF\nhejt7UVqaipmzZqFzMxMa/umpibMnTsXp06dQnh4OObNm4fy8vJXxlRdXY3FixdDqVRi+fLluHv3\n7ivb7t+/HxEREZg5cyYWLVqEe/fuAbC9vLthwwaEhoZaP/7+/igtLQUAdHV1Ye3atZg9ezbmz5+P\na9euvbKv9PR0FBYW4qOPPoJCocAnn3yCwcFBB880EXkjFnZEJBr379/HxYsX0d7eDoPBgPr6eqhU\nKgBAQEAAvv/+ewwODuK3337DjRs38MMPP9hsX19fD51Oh1u3buH06dPYtWsXNBoN/vjjD9y9exca\njcbatr+/H4ODg/jrr79w5coV5OXlobu72y4mnU6HnTt3ori4GENDQ8jPz8enn36KFy9e2LWtq6tD\na2sruru7MTw8jGvXriEsLAyA7eXdqqoqGI1GGI1GVFZWIioqCqtXr8bTp0+xdu1abNu2DY8fP0ZF\nRQW+/vprdHZ2vvKcaTQalJSUYGBgAC9evMDZs2edPu9E5D1Y2BGRaPj7+8NsNuP333+HxWJBbGws\n3nnnHQDA+++/j6VLl8LPzw9xcXHIy8tDc3OzzfYHDx6EXC5HUlIS3nvvPajVaqhUKigUCqjVauh0\nOpv2x48fR2BgINLS0rBu3TpcvXrV+t1EEfbTTz8hPz8fKSkpkEgkyMnJQVBQEG7dumUXv1QqhdFo\nRGdnJ8bHx/Huu+8iMjLS+v3ky7t6vR65ubmorKxETEwMqqurMW/ePGzfvh1+fn5YvHgxMjIyXjlq\nJ5FIsGPHDsTHx0Mmk+GLL77AnTt3nDjjRORtWNgRkWjEx8fj/PnzOHbsGCIiIpCVlYVHjx4B+LcI\nWr9+PaKiojBz5kwUFBTYXXaMiIiw/hwcHGyzLJPJYDKZrMtKpRLBwcHW5bi4OGtfL+vr68O5c+eg\nVCqtnz///HPKtqtWrcKePXuwe/duREREID8/H0ajccpjHR4exsbM9OUyAAACSElEQVSNG3Hy5Eks\nW7bM2pdWq7Xpq7y8HP39/a88Zy8XjsHBwTbHSES+h4UdEYlKVlYWWltb0dfXB4lEgkOHDgH493Eh\nSUlJ6OnpwfDwME6ePInx8XGH9zt5luuTJ08wMjJiXe7r60N0dLTddrGxsSgoKMCTJ0+sH5PJhC1b\ntkzZz969e9He3o579+5Br9fjzJkzdm3Gx8exdetWrF69Gl9++aVNXytXrrTpy2g04uLFiw4fJxH5\nNhZ2RCQaer0eDQ0NMJvNCAoKgkwmg7+/PwDAZDIhNDQUISEh6OrqcujxIS9f+pxqlmtRUREsFgta\nW1tRU1ODzz//3Np2ov2uXbtw+fJltLW1QRAEPH36FDU1NVOOjLW3t0Or1cJisSAkJMQm/pf7Lygo\nwMjICM6fP2+z/fr166HX61FWVgaLxQKLxYLbt2+jq6vLoWMkImJhR0SiYTabcfjwYYSHhyMqKgp/\n//03Tp06BQA4e/YsysvLoVAokJeXh8zMTJtRuKmeOzf5+5eXIyMjrTNss7Oz8eOPPyIxMdGu7ZIl\nS1BcXIw9e/YgLCwMCQkJ1hmskxkMBuTl5SEsLAwqlQpz5szBgQMH7PZZUVFhveQ6MTNWo9FALpej\nvr4eFRUViImJQVRUFA4fPjzlRA1HjpGIfI9E4L97RORjmpqakJ2djYcPH7o7FCKit4ojdkRERERe\ngoUdEfkkXrIkIm/ES7FEREREXoIjdkRERERegoUdERERkZdgYUdERETkJVjYEREREXkJFnZERERE\nXuIfaui5S+SyaUwAAAAASUVORK5CYII=\n", + "text": [ + "
\n", + "
\n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "# Cython vs regular (C)Python" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Here, we implement a linear regression via least squares fitting (with vertical offsets) by solving to fit *n* points $(x_i, y_i)$ with $i=1,2,...n,$ via linear equation of the form \n", + "$f(x) = a\\cdot x + b$. \n", + "\n", + "Therefore we calculate the following parameters as follows:\n", + "\n", + "$a = \\frac{S_{x,y}}{\\sigma_{x}^{2}}\\quad$ (slope)\n", + "\n", + "$b = \\bar{y} - a\\bar{x}\\quad$ (y-axis intercept)\n", + "\n", + "where \n", + "\n", + "\n", + "$S_{xy} = \\sum_{i=1}^{n} (x_i - \\bar{x})(y_i - \\bar{y})\\quad$ (covariance)\n", + "\n", + "\n", + "$\\sigma{_x}^{2} = \\sum_{i=1}^{n} (x_i - \\bar{x})^2\\quad$ (variance)\n", + "\n", + "I have described the approach in more detail in this [IPython notebook](http://sebastianraschka.com/IPython_htmls/cython_least_squares.html)." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
\n", + "**First, the implementation in Python (CPython)**:" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "def py_lstsqr(x, y):\n", + " \"\"\" Computes the least-squares solution to a linear matrix equation. \"\"\"\n", + "\n", + " x_avg = sum(x)/len(x)\n", + " y_avg = sum(y)/len(y)\n", + " var_x = 0\n", + " cov_xy = 0\n", + " for x_i, y_i in zip(x,y):\n", + " temp = (x_i - x_avg)\n", + " var_x += temp**2\n", + " cov_xy += temp*(y_i - y_avg)\n", + " slope = cov_xy / var_x\n", + " y_interc = y_avg - slope*x_avg\n", + " return (slope, y_interc)" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 1 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
\n", + "**And now, adding type definitions and compiling the code via Cython**:" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%load_ext cythonmagic" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 2 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%%cython\n", + "\n", + "def cy_lstsqr(x, y):\n", + " \"\"\" Computes the least-squares solution to a linear matrix equation. \"\"\"\n", + " cdef double x_avg, y_avg, temp, var_x, cov_xy, slope, y_interc, x_i, y_i\n", + " x_avg = sum(x)/len(x)\n", + " y_avg = sum(y)/len(y)\n", + " var_x = 0\n", + " cov_xy = 0\n", + " for x_i, y_i in zip(x,y):\n", + " temp = (x_i - x_avg)\n", + " var_x += temp**2\n", + " cov_xy += temp*(y_i - y_avg)\n", + " slope = cov_xy / var_x\n", + " y_interc = y_avg - slope*x_avg\n", + " return (slope, y_interc)" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 3 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
\n", + "**A small visual proof of concept that our least squares fit works as intended:**" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%pylab inline" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 5 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "from matplotlib import pyplot as plt\n", + "\n", + "import timeit\n", + "import random\n", + "random.seed(12345)\n", + "\n", + "n = 500\n", + "x = [x_i*random.randrange(8,12)/10 for x_i in range(n)]\n", + "y = [y_i*random.randrange(10,14)/10 for y_i in range(n)]\n", + "\n", + "slope, intercept = cy_lstsqr(x, y)\n", + "\n", + "line_x = [round(min(x)) - 1, round(max(x)) + 1]\n", + "line_y = [slope*x_i + intercept for x_i in line_x]\n", + "\n", + "plt.figure(figsize=(8,8))\n", + "plt.scatter(x,y)\n", + "plt.plot(line_x, line_y, color='red', lw='2')\n", + "\n", + "plt.ylabel('y')\n", + "plt.xlabel('x')\n", + "plt.title('Linear regression via least squares fit')\n", + "\n", + "ftext = 'y = ax + b = {:.3f} + {:.3f}x'\\\n", + " .format(slope, intercept)\n", + "plt.figtext(.15,.8, ftext, fontsize=11, ha='left')\n", + "\n", + "plt.show()" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "display_data", + "png": "iVBORw0KGgoAAAANSUhEUgAAAgIAAAH4CAYAAAA4pIUuAAAABHNCSVQICAgIfAhkiAAAAAlwSFlz\nAAALEgAACxIB0t1+/AAAIABJREFUeJzs3Xd8jdcfwPHPvZn3ZkkkEpIQYsaMhColVmLP2qMoSukw\na+8WtYqqllbt1qoaLapmVa0aNWtTIoIMIzcy7v3+/gj3Z8SIJG7Ieb9eeVXu8zznfM9zb3O+93nO\nc45GRARFURRFUbIlraUDUBRFURTFclQioCiKoijZmEoEFEVRFCUbU4mAoiiKomRjKhFQFEVRlGxM\nJQKKoiiKko2pREDJ8nbs2EHRokUtHcYrq0SJEvzxxx8vtc7333+fTz/99IWO9fPzY/PmzRkcUfb2\n888/4+vri7OzM4cOHbLIZ0LJujRqHgElq/Dz82POnDnUqFHD0qEoFpQ/f37mzJlD9erVM6X8bdu2\n0b59ey5dupQp5WdF/v7+TJ06lQYNGjy2beTIkZw9e5aFCxdaIDIlK1BXBJQsQ6PRoNFoLB2GmdFo\nzJB9npeIoPJy5b6M+myJCP/99x8BAQEZUp7y+lGJgJLlbdu2DV9fX/Pvfn5+TJ48mdKlS5MjRw5a\ntWpFQkKCefsvv/xCmTJlcHV1pVKlShw5csS8bfz48RQsWBBnZ2eKFy/OqlWrzNvmzZtHpUqV6NOn\nD+7u7owaNeqxWEaOHEmzZs1o3749Li4uzJ8/n5s3b9K5c2fy5MmDj48Pw4YNw2QyAWAymejbty8e\nHh4UKFCAGTNmoNVqzdurVq3K0KFDqVSpEg4ODpw/f55///2X0NBQcubMSdGiRVm+fLm5/nXr1lG8\neHGcnZ3x8fFh8uTJANy4cYP69evj6upKzpw5qVKlykPn6/6l9oSEBHr16oW3tzfe3t707t2bxMRE\n83n28fFhypQpeHp6kidPHubNm5fqe7J06VLKlSv30GtffPEFjRo1AqBjx44MGzYMgJiYGOrXr0+u\nXLlwc3OjQYMGhIeHp1ruo0TE/J65u7vTsmVLYmJizNubN29O7ty5yZEjByEhIRw/fvyJ52rKlCkY\nDAbq1KnDlStXcHJywtnZmatXrz5W75POM8DEiRPN7/X333+PVqvl3LlzQMr7OWfOHPO+8+bNo3Ll\nyubfP/74Y/LmzYuLiwvBwcH8+eef5m1p/WydOXOGkJAQcuTIgYeHB61atXqsHQkJCTg5OWE0Gild\nujSFChUC/v+Z2LBhA+PGjWPp0qU4OTkRGBj4XO+L8poRRcki/Pz8ZPPmzY+9vnXrVvHx8Xlovzfe\neEMiIiIkOjpaihUrJt98842IiBw4cEBy5cole/fuFZPJJPPnzxc/Pz9JTEwUEZHly5dLRESEiIgs\nXbpUHBwc5OrVqyIiMnfuXLG2tpYZM2aI0WiU+Pj4x2IZMWKE2NjYyOrVq0VEJD4+Xho3bizdu3cX\ng8Eg165dk/Lly8usWbNEROTrr7+WgIAACQ8Pl5iYGKlRo4ZotVoxGo0iIhISEiL58uWT48ePi9Fo\nlNjYWPHx8ZF58+aJ0WiUgwcPiru7u5w4cUJERLy8vOTPP/8UEZHY2Fg5cOCAiIgMHDhQunfvLsnJ\nyZKcnGze59HzOmzYMHnzzTfl+vXrcv36dalYsaIMGzbMfJ6tra1lxIgRkpycLOvWrRO9Xi+xsbGP\nnQeDwSBOTk5y+vRp82vBwcGydOlSERHp2LGjudyoqChZuXKlxMfHy+3bt6V58+bSuHHj1D8Ej8Q7\ndepUefPNNyU8PFwSExOlW7du0rp1a/O+c+fOlTt37khiYqL06tVLypQpY972pHO1bdu2hz5PqXnS\nsevXrxdPT085duyYxMXFSevWrUWj0cjZs2dFRKRq1aoyZ86ch+J76623zL8vWrRIoqOjxWg0yuTJ\nk8XLy0sSEhJEJO2frVatWsnYsWNFRCQhIUF27tz5xPY8GOOj53jkyJHSvn37p54P5fWmrggor6SP\nPvoILy8vXF1dadCgAYcOHQJg9uzZdOvWjXLlyqHRaHjnnXews7Nj165dADRr1gwvLy8AWrRoQaFC\nhdizZ4+53Dx58tCzZ0+0Wi329vap1l2xYkUaNmwIwM2bN1m/fj1ffPEFOp0ODw8PevXqxZIlSwBY\ntmwZvXr1Ik+ePOTIkYNBgwY9dPlfo9HQsWNHihUrhlarZcOGDeTPn58OHTqg1WopU6YMTZs2Zdmy\nZQDY2tpy7Ngxbt26hYuLi/kbnK2tLREREVy4cAErKysqVaqUauw//PADw4cPx93dHXd3d0aMGPHQ\nvWEbGxuGDx+OlZUVderUwdHRkZMnTz5Wjk6no1GjRvz4448AnD59mpMnT5rPC2Bup5ubG02aNMHe\n3h5HR0cGDx7M9u3bU39jHzFr1iw+/fRT8uTJg42NDSNGjGDFihXmb8UdO3bEwcHBvO2ff/7h9u3b\nTz1X8hy3X5507LJly3j33XcJCAhAr9enetXoadq2bYurqytarZY+ffqQkJDw0PlNy2fL1taWCxcu\nEB4ejq2tLRUrVkxTLPeJuiWV7alEQHkl3e/MIaVTunPnDgAXL15k8uTJuLq6mn8uX75MREQEAAsW\nLCAwMNC87ejRo0RFRZnLevAWxJP4+PiY/33x4kWSkpLInTu3uczu3btz/fp1ACIiIh4q88FjU6vz\n4sWL7Nmz56H4f/jhByIjIwH46aefWLduHX5+flStWpXdu3cD0L9/fwoWLEhYWBj+/v58/vnnqcZ+\n5coV8uXLZ/49b968XLlyxfx7zpw50Wr//2dBr9ebz+2j2rRpY04EfvjhB3Nn/yiDwUC3bt3w8/PD\nxcWFkJAQbt68+Vydz4ULF2jSpIn5XAQEBGBtbU1kZCRGo5GBAwdSsGBBXFxcyJ8/PxqNhhs3bjz1\nXD2PJx376PuZN2/e5y4TYNKkSQQEBJAjRw5cXV25efOmOV5I22drwoQJiAjly5enRIkSzJ07N02x\nKMp91pYOQFEywv1Bhnnz5mXIkCEMHjz4sX0uXrzIe++9x5YtW3jzzTfRaDQEBgY+9g39WfU8uI+v\nry92dnZERUU91IHelzt37odGp6c2Uv3B8vLmzUtISAgbN25Mtf7g4GBWrVqF0Wjkyy+/pEWLFvz3\n3384OjoyadIkJk2axLFjx6hevTrly5enWrVqDx2fJ08eLly4QLFixQD477//yJMnz1Pb/CQ1a9bk\n+vXr/PPPPyxZsoSpU6em2q7Jkydz6tQp9u7dS65cuTh06BBly5ZFRJ55vvPmzcvcuXN58803H9u2\ncOFC1qxZw+bNm8mXLx+xsbG4ubmZ388nnavnGZD6pGNz587Nf//9Z97vwX8DODg4EBcXZ/79wfEH\nO3bsYOLEiWzZsoXixYsDPBTvg+cMnv3Z8vT0ZPbs2QDs3LmTmjVrEhISQoECBZ7ZvgdlpQG6imWo\nKwJKlpKYmMjdu3fNP887cvr+H9OuXbvyzTffsHfvXkSEuLg4fv31V+7cuUNcXBwajQZ3d3dMJhNz\n587l6NGjaYrv0W+xuXPnJiwsjD59+nD79m1MJhNnz541P6PdokULpk2bxpUrV4iNjeXzzz9/7A/v\ng2XWr1+fU6dOsWjRIpKSkkhKSmLfvn38+++/JCUlsXjxYm7evImVlRVOTk5YWVkBKQMkz5w5g4jg\n7OyMlZVVqp1H69at+fTTT7lx4wY3btxg9OjRtG/fPk3n4D4bGxuaN29Ov379iImJITQ09KE23W/X\nnTt30Ol0uLi4EB0dnabL6d27d2fw4MHmDvf69eusWbPGXK6dnR1ubm7ExcU9lPw97Vx5enoSFRXF\nrVu3Uq3zace2aNGCefPmceLECQwGw2NtKVOmDCtXriQ+Pp4zZ84wZ84c8/t9+/ZtrK2tcXd3JzEx\nkdGjRz8xBnj2Z2v58uVcvnwZgBw5cqDRaFJ9z5/Fy8uLCxcuqNsD2ZhKBJQspW7duuj1evPPqFGj\nnvlY4YPbg4KC+Pbbb/nggw9wc3OjUKFCLFiwAICAgAD69u3Lm2++iZeXF0ePHuWtt95KtZznqeu+\nBQsWkJiYSEBAAG5ubjRv3tz8TbBr166EhYVRqlQpgoKCqFev3mOd9IPlOTo6snHjRpYsWYK3tze5\nc+dm0KBB5pH9ixYtIn/+/Li4uDB79mwWL14MpIwgDw0NxcnJiYoVK9KzZ09CQkIei3/o0KEEBwdT\nqlQpSpUqRXBwMEOHDk01lufRpk0bNm/eTPPmzR9r0/2yevXqRXx8PO7u7lSsWJE6deo8dz0ff/wx\nDRs2JCwsDGdnZ95880327t0LwDvvvEO+fPnw9vamRIkS5qs89z3pXBUtWpTWrVtToEAB3NzcUn1q\n4EnH1q5dm169elG9enUKFy782JwXvXv3xtbWFk9PTzp16kS7du3M22rXrk3t2rUpXLgwfn5+6HS6\nh24tpPWz9ffff1OhQgWcnJxo1KgR06dPx8/PL9Xz+LTz3bx5cyDltlBwcPAT91NeXxabUOjkyZMP\nPe5y7tw5xowZQ7t27WjZsiUXL17Ez8+PZcuWkSNHDgDGjRvH999/j5WVFdOnTycsLMwSoSvKC1u/\nfj3vv/8+Fy5csHQoSgbRarWcOXMmzZfkFSWrsNgVgSJFinDw4EEOHjzI/v370ev1NGnShPHjxxMa\nGsqpU6eoUaMG48ePB+D48eMsXbqU48ePs2HDBnr06GEeOawoWdXdu3dZt24dycnJhIeHM2rUKJo2\nbWrpsBRFUcyyxK2BTZs2UbBgQXx9fVmzZg0dOnQAoEOHDuYJX1avXk3r1q2xsbHBz8+PggULmi8R\nKkpWJSKMHDkSNzc3ypYtS/HixRk9erSlw1IykBpsp7zqssRTA0uWLKF169YAREZG4unpCaQM6rn/\n2NSVK1eoUKGC+RgfH5/nnp1MUSxFp9OphPU1l5HTTCuKJVg8EUhMTGTt2rWpPvf8PIPEHlWwYEHO\nnj2boTEqiqIoSlbl7+/PmTNnXvh4i98aWL9+PUFBQXh4eAApVwHuj4qNiIggV65cAHh7ez/0DPbl\ny5fx9vZ+rLyzZ8+aH13Kjj8jRoyweAyq7ar9qv2q/artL+8nvV9+LZ4I/Pjjj+bbAgANGzZk/vz5\nAMyfP5/GjRubX1+yZAmJiYmcP3+e06dPU758eYvErCiKoiivC4veGoiLi2PTpk18++235tcGDhxI\nixYtmDNnjvnxQUh5BrxFixbmKUZnzpypBukoiqIoSjpZNBFwcHB4aJ5tSJlyc9OmTanuP3jw4FSn\njlX+r2rVqpYOwWKyc9tBtV+1v6qlQ7CY7Nz2jGCxCYUyi0aj4TVrkqIoiqI8UXr7PYuPEVAURVEU\nxXJUIqAoiqIo2ZhKBBRFURQlG1OJgKIoiqJkYyoRUBRFUZRsTCUCiqIoipKNqURAURRFUbIxlQgo\niqIoSjamEgFFURRFycZUIqAoiqIo2ZhKBBRFURQlG1OJgKIoiqJkYyoRUBRFUZRsTCUCiqIoipKN\nqURAURRFUbIxlQgoiqIoSjamEgFFURRFycZUIqAoiqIo2ZhKBBRFURQlG1OJgKIoiqJkYyoRUBRF\nUZRsTCUCiqIor6DIyEgqV66DnZ0juXMXZOPGjS+1/vj4ePbs2cORI0cQkZdad5YSHw/Ll1s6inRR\niYCiKMorqF69luzeXYLExHCuXp1FkyZtOXPmzEup++LFixQqVJqwsO5UqNCAOnXeJjk5+aXUnaXc\nvQtNmkCLFvDll5aO5oWpREBRFOUVk5CQwMGDf5GcPB5wAWqg0dRmx44dmV53YmIilSvXIzy8Hbdu\nHcRgOMWOHbf45ptZmV43gIhw69YtTCbTS6nviRISoGlT+O03cHeHatUsG086qERAURTlFWNra4ut\nrT1w/wqAEY3mJDlz5sz0ut99tyeXLkUCze5Hg8FQjyNHTmZ63SdPniR//hLkzOmFo6Mby5evyPQ6\nU5WQAG+/DevXQ86csGULlChhmVgygEoEFEVRXjEajYapU6eg19fAxqY3Dg7VKV3albp162ZqvSaT\niaVLFwFvAosBAQzY2i4nKChzO0IRISysCf/914Pk5Dji47fSsWMPTp06lan1PiYxEZo3h19/BTc3\n2LwZSpZ8uTFkMI28ZqM8NBpN9h64oihKtvHXX3/x119/4eXlRcuWLbGxscnU+kQEnc6ZhIStwLtA\nPHCD4OCS7N69FSsrq0yrOzY2lly5fElKum1+zcmpBbNmNaF169aZVi9ATEwM169fJ1/u3Ni1bw+r\nV4Ora8qVgDJlMrXu55Hefk8lAoqiKMpzGzp0FF98sRKDoSfW1n/g6rqDkycP4erqmqn1Go1GHB3d\nuHt3B1AKMODgEMivv84mJCQk0+qdOnUGAwcOQWedk4VJEdRPvAs5cqRcCShbNtPqTYv09nvWGRiL\noiiK8poxGo1MmPAF69dvx9fXk7Fjh1GgQD5+/XUrPj65GTJkX6YnAQBWVlbMmTOLLl1qYm1dHZPp\nEI0bV6FKlSqZVufBgwcZMmQcxoT9fJswkPr8RKxGi8vvv6PJIklARlBXBBRFUZQn6tbtYxYtOoDB\n0BcrqwO4us7j338PvpSBian5999/2b9/P97e3oSEhKDRaDKlnkWLfuC993qQGF+ZH9DRguXE4kId\nq3h+j43C0dExU+p9EerWwCNUIqAoyuvkwoULLFiwkORkI61atSAgIOCl1W0ymbCzcyA5ORxwA8DB\n4W2++qohHTp0eGlxvGx79uyhevUmJBhGs4hetCKOmzgTxjhOuowmJiYi0xKQF6FuDSiKorymTp06\nRblylYmLa4WIPVOmhLB16zrKlSv3UuqPjY0l5cmABzu9rNMBZpatW7eSnNCK+WylFXHcQkMtcnHc\ncSSrVy7JUklARlCPDyqKomRRY8ZM4s6dDzEap2EyfU5c3FgGDvws0+u9ceMG5cpVxcsrH0ajFdbW\n9YA1WFmNwt5+D/Xq1cv0GCzJw82NeSyhLT9wG0dqM5hTrgbOnj1K9erVLR1ehlOJgKIoShYVE3Mb\nkynvA6/k5ebN20/cP6N06NCDf/4pRVLSTUROo9FcoGjRsTRrdpH9+//E3d0902OwGKORjn/8QWtj\nBHeworFtTf7Rz2bBglnkypXL0tFlCnVrQFEUJYtq06YRW7cOw2AoBtjj4DCY1q3fyfR6d+/eTVLS\nNlK+K+YhKakn9evfZuLE8Zlet0WZTNClC1aLFyN6PX/17k2jXLmYWm00JV/xSYOeRiUCiqIoWVSb\nNq2Iiopm3Lh2GI3JvP/+u/Tp81Gm1+vt7Ut09J9AAcCETvcX+fLVyfR6Lcpkgq5dYd480OvRrFtH\nWEgIYZaO6yVQTw0oiqIoDzlw4ADVqtUF3sBkCicgwJk//liPnZ2dpUPLHCYTdOsG330HOl3K9MGv\n0CJC6vHBR6hEQFEUJf2uXr3Kzp07cXR0pEaNGlhbv6YXkE0m6NEDZs0Ce3v45ReoUcPSUaWJSgQe\noRIBRVEU5bmIQM+e8PXXYGcHa9dCaKilo0qz9PZ76qkBRVEUJfsRgQ8//H8SsHr1K5kEZASVCCiK\noijZiwj06gVffQW2trBqFdSqZemoLEYlAoqiKEr2IQJ9+sD06SlJwM8/Q+3alo7KolQioCiKomQP\nItC/P0ydCjY28NNPULeupaOyOJUIKIqiKK8/ERgwACZPTkkCVqyA+vUtHVWW8Jo+D6IoiqJkd7Gx\nsfTpM4RDB48xxniLekcOgrU1LFsGDRtaOrwsQ10ReI35+flx/PjxTCl748aNBAcHY29vT//+/Z+4\nX2JiIrVr18bDwwMPD4+Htl24cAFra2sCAwPNP9HR0Q/tIyLUrFnzsWPTq23btnh7e6PVajEYDKnu\n8+677z51+6JFiyhVqhQ2NjZ89dVXD207evQob731FoGBgQQEBDBq1Cjzto4dO+Lr62tu87hx4zKu\nYYqiAJCcnEyVKnVYvCiRtw/lod6RgyQDxsWLoXFjS4eXpVg0EYiNjaVZs2YUK1aMgIAA9uzZQ3R0\nNKGhoRQuXJiwsLB7y2CmGDduHIUKFaJo0aJs3LjRgpFb1rx58x7qWJ4kM+dU8Pf3Z86cOU9NAgCs\nrKz45JNP2LRpU6rbXV1dOXjwoPnHzc3toe0zZszAz8/vuZf97NixI9u3b3/mfl27duXQoUNP3L52\n7Vq0Wu1T6w0MDGTp0qW0adPmsf369+9PmzZtOHjwIPv27WPu3Ln8/fffQMr7MmjQIHObBw0a9Fxt\nUxTl+Z04cYJz564zJMmLIfxIMlZ0svfiaJEilg4ty7FoIvDxxx9Tt25dTpw4weHDhylatCjjx48n\nNDSUU6dOUaNGDcaPT1nk4vjx4yxdupTjx4+zYcMGevTogclkyrBYJk6cyAcffGD+PTIyEi8vL+7e\nvZvusiMjI6levTrBwcGUKFGCAQMGmLd17dqVPn36mPcrUKAAhw8ffmp5aVkLe9GiRQQHB1OoUKHH\nvrWmh7+/P6VLl37mbGNWVlZUr14dFxeXNNdx+vRpli5dysCBA587odFoNM91fqpWrfrEqwxRUVGM\nHj2aKVOmPLXe4sWLU6xYMbRa7WP7+fj4mJPYO3fuoNFoHlq5LLVy4+PjKV26NGvWrAFgy5YtFCtW\njLi4uGe2R1GUh1lZWTEo8QbD+RQjWtqykJ+tdFhZWVk6tCzHYmMEbt68yY4dO5g/f35KINbWuLi4\nsGbNGvM3ug4dOlC1alXGjx/P6tWrad26NTY2Nvj5+VGwYEH27t1LhQoVMiSeLl26EBAQwIQJE9Dr\n9cyePZu2bdtib2//2L7NmzfnzJkzj72u0WjYtWvXY/Nx58iRg7Vr1+Lg4EBSUhK1a9fmt99+o1at\nWnz55Ze88cYbrF69mi+//JJPPvmEUqVKPTXWtHzLv379On///TfXrl0jMDCQKlWqPLaK1okTJ2jT\npk2qx4eFhfH5558/d31pdevWLYKCgtBoNLRq1Yp+/foBYDKZ6Nq1KzNnzkzz1KbpvQrSs2dPRo8e\njbOz8wuXMX78eKpUqcLMmTOJiYlh0qRJ5M37/+VkJ0+ezKxZs/D392fcuHEULVoUnU7HsmXLCAsL\nw8vLiy5duvDzzz/j4OCQrvYoSnZUdMUKhiTdxAi0oydr7FdRpmQBAgICLB1almOxROD8+fN4eHjQ\nqVMn/vnnH4KCgpg6dSqRkZF4enoC4OnpSWRkJABXrlx5qNP38fEhPDw8w+JxdXWlYcOGLFiwgC5d\nuvDdd9+xZcuWVPddvnx5mspOTk6mX79+7Nq1CxHh6tWrHDp0iFq1amFvb8+yZcsICgqibt26dO/e\nPdUy1q1bx5AhQwCIjo4mMTGRVatWAfDhhx/y7rvvpnpc586dAciVKxf16tVj27ZtjyUCxYoV4+DB\ng2lqU0bIkycP4eHhuLu7c/36dRo2bIirqyudO3dm0qRJhISEUKpUKS5cuPDUcsaMGcPKlSsB+O+/\n//jzzz9xdHQEYP78+c9MrB60bNky7OzsqFOnjjmheJHEokOHDrz77rv07duXq1evUrVqVYKCgihf\nvjyfffYZefLkAWDhwoXUrl2bc+fOodVqKVKkCKNHj6ZixYpMmzaN0qVLp7luRcn2PvsM7YgRiFbL\n8roNiUq4wkeBxRgxYhBarRoa9yiLJQLJyckcOHCAGTNmUK5cOXr16mW+DXDfsy7zPmnbyJEjzf+u\nWrUqVatWfa6YPvzwQ9q2bYuHhwcBAQH4+/unul+zZs04e/Zsqtt27dr12FWEKVOmEBsby969e7G1\ntaVbt24P3XI4duwYLi4uXL16FaPRmOqlq7p161L33vOu8+fP5+LFiwwfPvyZbXqwExORVM/Z8ePH\nadu2barHh4aGMmHChGfW8yJsbW1xd3cHwMPDg7Zt27Jz5046d+7Mjh07OHz4MAsWLCA5OZmYmBjz\nbZP7nfx9w4YNY9iwYQB06tSJTp06UaVKlReKafv27WzZsoX8+fObXytRogTr16+naNGiTzzu0fO6\ndetW5s2bB4CXlxfVq1fnjz/+oHz58uYkAKB9+/b07t2b8PBwfH19Adi/fz+enp5cunTphdqgZE1x\ncXEcP34cV1dXChYsaOlw0uXvv/+mR48BREZeIyysGtOnf45Op7N0WCnGj4ehQ0GjQTNvHq3at6eV\npWPKYNu2bWPbtm0ZV6BYSEREhPj5+Zl/37Fjh9StW1eKFi0qERERIiJy5coVKVKkiIiIjBs3TsaN\nG2fev1atWrJ79+7Hyk1vk6pVqya+vr6ydu3adJXzoL59+0rv3r1FROTy5cvi6ekpo0aNEhGRc+fO\nSd68eeXMmTPSoUMHGThw4DPLmzt3rowcOfKZ++XLl0+6du0qIiLXrl0Tb29vOXr0aDpa8rgRI0ZI\nv379nrnf+fPnxd3d/aHXrl27JomJiSIiEhcXJzVr1pTp06c/duyFCxceO/ZJOnbsKNu2bXuufU0m\nk2g0Grlz584T99FoNBIXF/fUcjp06CAzZsx46LVy5crJggULRETk1q1bUqJECdmwYYOIpHwG7tuw\nYYN4enqK0WgUEZGVK1dK2bJlJTo6WooXLy7r169/rrYoWdvRo0clZ05fcXYuIzpdLunS5QMxmUyW\nDuuFXLhwQRwdPQTmCRwUe/sm0rRpO4vFYzKZZNmyZTJq1Cg53K6dCIhoNCLz5lksppctvf2exRIB\nEZHKlSvLyZMnRSSlQ+nfv7/0799fxo8fLyIpnf+AAQNEROTYsWNSunRpSUhIkHPnzkmBAgVS/R8p\nvSdk0aJFDyUoGeHixYtSvnx5KVGihNSuXVvatm0ro0aNksTERClfvrz8+OOPIpLSGQYEBJg7jCeZ\nN2+eOZF4Gj8/Pxk8eLAEBQVJwYIF5auvvsqQ9oikJG4+Pj7i7OwsTk5O4uPjIxs3bhQRkW+++UaG\nDx9u3jc4OFhy584t1tbW4uPjY05OfvrpJylRooSULl1aAgICZMCAAam+p+fPnxcPD4/niqtjx46y\nffv2Z+7XpEkT8fHxEa1WK97e3lK7du1U99NqtQ8lAmXKlDEnqj/88IP4+PiIg4ODuLq6io+Pj5w4\ncUJEUv4BlgPUAAAgAElEQVTwV6lSxdy2MWPGmMuoWbOmlCxZUkqXLi1VqlSRPXv2mNvp6+srp0+f\nNpeRN29eCQ8Pf662K1lXsWLlBL6VlFltboqDQ0lZtWqVpcN6Id98843odB3vtUUEbom1tZ05mX2Z\nTCaTtG//njg4lJW+1BABMYLI99+/9Fgs6ZVOBA4dOiTBwcFSqlQpadKkicTGxkpUVJTUqFFDChUq\nJKGhoRITE2Pe/7PPPhN/f38pUqTIEzvL9J6Qzp07y6RJk9JVhqIoyoPs7BwFYsydp7V1v4eucL5K\n5s+fLw4O9R5IBM6Jvb2zRa5wnDx5UnQ6L+nFuPvBSDdrRzl//vxLj8WS0tvvae4V8tp40Wfnr1y5\nQvXq1cmdOzfr169P9WkBRVGUF1G8+BucOPEuIt2Amzg4VOKHH8bSMI2z20VERJCYmIivr+8TB70d\nPXqUWbPmYjQa6dy5PUFBQRnQgv+7ffs2JUu+QUREBRITS6PXz2To0C4MGvT0OUUyw969e1lRpSkT\nElIGjr/HLJY4TeePPxZRpkyZlx6PpaR3zhiVCCiKomSyEydOEBJSh4QEJxITr9KpU3u++mryc88J\nkpycTKtW7/LLL7+g1dpRrFghNm9eQ44cOR7a759//qFSpZrExfUEbNHrv+C3337mrbfeytD2xMTE\n8MUX07ly5Tp16lTj7bffztDyn1fC5MnY3XvkuDsTmK2xw9NzGufPH8tWX+ZUIvAIlQgoipIVxcfH\n8++//+Lm5ka+fPnSdOyECZMYMeJX7t79FbDH1rY7zZoJixd/+9B+LVt2YtmyEkDfe698T/Xqa9i8\neVWGtCFLmTkTevYEYGye/HwWe5PChQNYtux7ChUqZOHgXq709nvqgUpFUZSXQKfTERgYmOYkYPv2\n7QwZMo67d9sCekBLYuK77Nv3+BTZd+7EA+4PvOJBXFx8esJ+ovDwcDp06E6NGk2YNGlqhs70+kyz\nZpmTAKZPZ3D4OeLiojh4cEe2SwIyglp9UFEUJYsyGAw0bNiC5OQGwG/Au4AWK6t1FCny+DwnnTu3\nZNu23hgM3qTcGuhPly6fZHhc0dHRlC1biaioNhiNNdm9+wvOnbvIzJlfZHhdj/n2W7g/8drUqfDh\nh5lf52tO3RpQFEXJok6ePElwcD3u3DkM1AVuAFrc3W9y8OBOfHx8Hjtm/vyFjB37JUajkV69utCz\nZ/c0rU/yPBYuXMj7768kLu7ne69EYW3tTUKCIXNn7pszB7p0Sfn3lCnQu3eGFv/333+zfft23N3d\nadWq1WPTxWdVaozAI1QioCjK6+LWrVt4eubl7t2dQBHgV2xtO/D33zsemyr8ZVqwYAE9eqwhLm7F\nvVdisbb24u7duMxb1GfuXOjcOeUhwYkT4d4gwYzy449L6Ny5F8nJrbC1PUqRIons2rUJW1vbDK0n\nM6gxAoqiKK8pZ2dnvv12JjpdVVxc6qDTvcfo0cMtmgQA1KlTB3v7PVhZfQqsQ69vSrt2nTIvCZg/\n//9JwOefZ3gSAPD++72Jj/+FpKSpxMX9zsmT1mleV+ZVpcYIKIqiZGHt2rWhcuVKnDhxggIFClC4\ncOFMq+vu3btERUXh5eX11E7dw8ODffv+oF+/EYSH76BOnZoMGZLxYxEAWLQIOnVKSQLGjoVPMq4e\nEWH27DnMnLmAmzdvAPdXJtSQnFyUqKioDKsrK1O3BhRFURQWLfqBrl3fR6PR4ehox2+/rSIwMNCy\nQf3wA7RvDyYTfPop3FuBNaN888239O07GYNhGjAcKAlMBo6g0zVl9+5NaVq91FLUGIFHqERAURQl\nbU6fPk2ZMpUwGLYAJYAl5Mo1kIiIc5ZbtnfJEmjbNiUJGDUKnmO11bQqVaoyR44MB0KBaKAmWu1x\ncub0YvbsqTRu3DjD68wMaoyAoiiKki6HDx/G2roiKUkAQCtu3rzNjRs3XnosIgLLlkG7dilJwPDh\nmZIEANjY2AB37v3mhkbTnI4dO3Pt2oVXJgnICCoRUBRFyeby5ctHcvJBIObeKwfRao24urq+tBj2\n7dtHvnwBtLSyJrllSzAaYehQGDky0+ocObI3Ol0PYCYwHr1+Ch9/3C3T6suq1K0BRVEUhd69BzF7\n9iJsbEqTlLSH+fO/oVmzl7OGQExMDH5+xahxqx1LmYYNyUzTu9A96ip2mbxmwObNm5k9ezH29rb0\n7dvjlRgT8Cg1RuARKhFQFOV1s2HDBgYNGo/BYKBjxxYMGNAnXffuTSYTkyZNZdmyX3Fzc+Hzz4cR\nGBjIwYMH+e+//yhVqhT58+fPwBY83R9//MGsOu8xz3AWG5IZzwDGOv7Enr1rKFas2EuL41WV3n5P\nPT6oKIqShe3atYumTTsQH/814MGnn/bCaDQydOiAFy5zyJBRTJ++HoNhDHCOv/4K4+DBvwgMDLTI\nkwL5jxxhruEkNsBE+jGI3tglfUPOnDlfeizZkRojoCiKkkXdvn2bDz7oR3x8b6ApUBmDYSZz5vyY\nrnJnzZqDwbAIqAW8T0JCG1asWPGswzLHL7/g27s3tsB0azeG2RpwcKhEv359yJUrl2ViymbUFQFF\nUZQsKDExkUqVwjh27A7w4MQ2sdjbp28OfK3WCkgw/67RJGTerIBPs24dvP02JCUhH31EwbAwPj9z\nhpIl36Z69erpKvr27dusXbuWu3fvUqtWLby9vTMo6NePSgQURVGyoNmzZ3Pq1B1MplVAJcAG8ESn\nm8CYMdNeqMxbt26h0+kYMKAXI0e2wGAYjFZ7Dr1+NW3a7MvI8J9twwZo0gQSE+HDD9FMnUrdDFoc\nKWV1xLeIivLDZHLF2nowO3ZsfCUHAr4M6taAoijKc1i+fAWVK9enevXGbN68OVPr+v77efTtO4KE\nBD3gD+wEDGi1I5g/fyrNmjVLU3mRkZGUKVOJnDlz4+Dggkaj4euvh1Cv3jratYvgwIHUVzLMNBs3\nQuPGKUlAz54wbRpk4AqJEyd+QUREJe7cWYfBsJhbt0bRo8eLj6l47clr5jVskqIoFvbjj0tEr88r\nsExgvuj1uWT79u2ZUpfJZBJ7e2eBvQIFBEYK/Ck2Nm2lYsVQMZlMaS6zSpW6Ym39iYBJ4KLo9X6y\nefPmTIj+/1auXClVqjSQ6tUby++///7/Db//LmJvLwIi778v8gLteZa2bbsKzJSUBQpEYK/4+wdm\neD1ZRXr7PXVFQFEU5RmmTPkOg2E60Bx4B4NhODNmzM2UupKSkkhMNACBwFbgOBpNa8qXv8qGDT+h\neYFvzvv2/UVycn9AA+QlIaEFu3fvztjAH/DTTytp1+4j/vijLVu2NKFhw7Zs3boVtmyBBg3g7l14\n7z2YMSNDrwTcV6tWFfT6mUA4EIe9/VhCQ0MyvJ7XhUoEFEVRniGl8zU+8EoyWm3Gd2AAtra2lC37\nFtbWA4EcQHd0uni+/34mTk5OL1Smp6cPKbcXAIzY2+/J1MFzkyd/i8EwFWgJvEN8/Gg2DxsL9eun\nJAFdusDXX0MmrWPQrl1b+vRpjq1tYays3KhTx4EpU8ZmSl2vAzWhkKIoyjOsWrWKNm16Eh8/FjCg\n0w1ny5a1VKhQIVPqi4yMpFmzjuzZ8wdubl7MnTuDOnXqvHB5O3bsoE6dpmi1IYico2xZLzZvXoO1\ndeaMF69UqS5//dURaAFAFXrzm9WX2BuNKUsKf/ddpiUBDxIRjEZjprUzq1AzCz5CJQKKomSGdevW\nMWPGfGxsrBkwoCcVK1a0dEhpcunSJf766y9y5MhBzZo1M/xxQRHh7t276HQ61q5dS8uW3YiPH8tb\nHGU9k3EE6NABvv/+pSQB2YlKBB6hEgFFUTLClStXiIiIoFChQjg7O1s6nCxt165dNGrUmqioK7i7\n52H16iXExMSwadREPv17BzpjMrRvD3PngiXmK3jNqWWIFUVRMti4cZPw9y9B9eqd8fUtzM6dO599\n0EtiMBiIjY1N83GnTp1i0qRJzJgxI0OXF7516xa1azfh+vXpmEwJXLs2jdq1m1DFxobJx/alJAFt\n277SSYDJZGLq1BlUq9aIVq3e5dy5c5YOKUOpKwKKoigPOHDgAJUrN8Rg2AvkAX7F1fU9oqIuv9CI\n/YwiIvTo0YfvvpuFRmNFhQpv8csvS5/rasXu3bupWbMBiYmtsLKKwdn5T/75ZzdeXl7piun48ePU\nqFGfq1c1wFnz69X1hdioicAqLg5at4YFC+AVvk//ySdD+eqr3zAYBqLVHsfZ+WuOH99P7ty5LR0a\noK4IKIqiZKgTJ05gZfUWKUkAQD3u3LnFzZs3LRkWc+Z8z4IFO0lOvkJSUgx793rywQf9n3mcwWCg\nU6ePiIubTFLSl9y9u4jo6MZMnDg1XfEkJydTvXp9rl7tDtwErgFQjvWsNJxNSQJatnzlkwCAr76a\nicGwEngbk2kYd+/WZOXKlZYOK8OoREBRFOUBRYoUwWj8E7h675XfcHBwxMXFxZJhsW3bHgyGTqQ8\nUmhNQsIH/Pnn3qcec+3aNYoXL8fJkxeBIubXk5MLc+1adLriCQ8P5/btROAToBdQjiBqsJF6uCDQ\nvDksWvTKJwHAvW/bD14Ner26zterNYqiKOkUHBzMgAEfYG9fHGfnIJyc3mH16qUWvS0AULCgL3Z2\nO4CUS8Ba7Q78/HzZvn07jRq1pWHDNg9NfWwwGHjzzTAuXKiOSFdgCHAZOIpeP4UmTWqnKx43NzeS\nk28CF4ChBDKQ39lKDiRlIaHFi1+LJACgW7du6PXNgLVotZ9jZ/cbTZo0sXRYGUaNEVAURUnFpUuX\niIiIoHDhwuTIkcMiMURHR/PBB5+wf/9hChfOz5kzZwkPtwZcsbU9yrRp4+natTfx8WMAK3S6oaxa\ntYCwsDAaN27DmjW7EfkSCAP6AguxthYmTfqUjz/+IM3xJCQksHnzZuLj4wkJCeGHH5bxySdjKGUs\nw4bkjbhhSllDYNkysLHJ2JNhQSaTicmTp7Fq1e94eubk88+HU6hQIUuHZaYeH3yESgQURUkLEeG7\n775n5crf8PBwZfToQfj5+Vk6LIxGI2XKVOLUqSASE9thbb0ab+81TJ8+HhGhcuXKvPNOT379tTrQ\n9d5RC6lW7Sc2bVqJra0Oo/EjUgbxLQFM2Ng04ZNPKvDppyPSHE9cXBwVKtTgwgXQaNyxtj5Iq1bN\n+HvOYtYn3iQnyRz1L0SJ40fB1jbDzoPybOnt916P6zaKoigvaPTocUyYsOTeiPCT/PJLJY4d+9vi\nI8LPnj3LuXMRJCZ+CWhJTq5AdPQGcuXKZZ7R0GQykbI88X02GI0mNBoNNjb2GI2dSLkl4AkkEhBQ\nmuHDB71QPF98MY3Tp/ORkLAE0KDRfMnOr4ewGRtyksxawmgXfoCDly9ToECBdLVdebnUGAFFUbK1\nKVOmYzD8BLTBZBqFwRDG8uXLLR0Wtra2iCQASfdeMSFiQKPRMGbMOOrUaYFOBzrdYOBHYBn29n3x\n9XXhiy++oH//vuj1jYBK2NjUJF++vPz550ZsX/Db+pkzl0hIeIv7g+YCJDebuIM70fxKXZqxBo1d\nfiIjI9PfeOWlUlcEFEXJ1kwmI/D/zlHEFqPR+OQDHvHvv/9y7NgxChQoQGBgYIbFlS9fPqpWfYtt\n2xoRH98Ce/tfKFkyH2PGTGLLljvEx7+Dnd1veHo6kTfvIm7evMXJk7dZssSDFSsO4+Kyja++Gsne\nvYfIkyeQDz/8DkdHxxeOp1q1N1m+fCoGQxsCuMwW2uOBsJ7SvM1yElmPvfxHQEBAhp0D5SVJ1yLG\nWdBr2CRFUTJRr14DRK9/U2CDaDTTxMnJQy5cuPBcx86a9Z3odB7i7NxI9HpvGTZsTIbGlpiYKOPH\nT5SmTd+RkSM/lfPnz4udnatAvIAImMTJqYwsWLBAbGw8BJbce13E2vpD6d9/UIbFYjKZ5KOP+ktJ\nK1u5eq+SqHLlpEAef9ForMTTM7/s2rUr3XW8LBcvXpQqVepKzpx5pWLFMDl79uxLqzujpbffe+16\nTZUIKIqSFkajUT77bIIEBVWX2rWbyZEjR57ruJiYGLGzcxY4da/zjRSdzlP+/fffTIv10qVLYm/v\nIZBk7vCdnd+QnDnzCBQQ2Gt+HabKu+/2yNgATpwQk6dnSgWhoSIGg4iIJCQkvFBxRqNR4uPjJSoq\nSqpVqy9WVrbi7OwpCxcuzsioH5OQkCB58xYVK6tPBc6IVjtRcuf2l7i4uEytN7Okt99TYwQURcnW\ntFotgwf35++/N7N+/XJKlCjxXMdFRkZiY+MB3H+MLBe2tkW5dOlSpsXq7e1NUFAgdnYdgS3Y2HxC\nUtI5oqJuAa2AgaQ8178fW9tJNG1aN+MqP3kSqlVDExkJNWrAqlWg0wG80LiDAQMGY2fniIODM35+\npfjzTzeMxhhu3VpPt2592bv36ZMlpcepU6eIiRGMxiGAPyZTP+LinDh69Gim1ZmVqURAURQlDUwm\nEx06vEeJEkHcuXMFWHNvy26Sk49RvHjxTKtbo9GwYcNPdOzoTunSo3BzW0VyciVSJhlqDZQC3gSq\n07t3O+rVq5cxFZ8+DdWqwdWrKf9dswb0+hcurnfvfkyYMIvk5EOYTHe5fbsFSUnhgB4IJDGxNdu3\nb8+Y2FPh6OhIUlI0EHfvlbskJ99I1xiKV1oGXZnIMl7DJimKkkH++ecfadq0vdSs2fSFLz83atRM\noITAZYE1Ao5ibZ1D9HpX+eWXXzI44idLSkoSKysbAYPAXAFPgaZiZ1dAOnXqkXH320+fFvH2Trkd\nEBIicudOuoqLj48XKytbgY8fuI0RK2BvHvfg4BAq33//fcbEnwqTySRt2nQWvf4NgbGi178ljRu3\nealjFDJSevs9NaGQoijZwr///ktwcGXi4gYDudHrhzFxYj969Oj23GVcuXIFH5+SiHwH3J9idiVl\nykxl797N2GTgbHoiwt69e4mKiiIoKAhPT8/Htut0ziQkHAbyA4exs2vNJ5+8zahRo546JfLJkyfZ\nv38/Pj4+VK5c+cn7nj0LVavC5ctQuTKsXw8ODulqV2RkJD4+/iQnlwW2AlbAJqAJ9vbtsLI6SeHC\nSezatQk7O7t01fU0JpOJhQsX8s8/xyhevAgdO3bE6hVdJjnd/V66U5Es5jVskqIoGeCTTwaJRjPo\ngW+hOyVv3hJpKmPPnj1ibZ1HYPQD5YyWGjUapDkeo9Eoly5dkqioqFS3NW3aThwc/MXFJUwcHT1k\n586dj+03YcIU0ev9BcaJvX1zKVKkrBjuDeB7kh9/XCp6vYc4ObUQB4dC0r79e6l/Ez57VsTXN6WR\nlSqJ3L6d5jamxmQySd68xe5dVQkWaCHgIOPGjZMvv/xSFi9e/MKDD7Or9PZ7r12vqRIBRVFS06/f\nQIGhD3Tgu8XXt3iayoiOjhadzlUgl0BbgTYCejlw4ECayomMjJTixcuLTucptrZO8v77vR/qjFes\nWCEODmUfeEzwZ/H1LZpqWWvXrpXevfvLhAkT5fYzOuvk5GTR6VwEDt0r9444OBSS7du3P7zj+fMi\nefOmnKiKFUVu3UpT+57l7NmzEhBQXjQarTg5ucuCBQsytPzsRiUCj1CJgKIoqTl69Kg4OLgLfCWw\nUvT6YjJlyrQ0l7Nx40bR693ExsZZ7OwcZOnSpWkuo1att8Xauq+ASSBaHBwCZdGiRebtkyZNElvb\nB++h3xZra7s01/Og3bt3S8GCZQTsHihXxMamgYSE1JS9e/em7HjhgoifX8rGChVEbt5MV71PYzQa\nM63s7CS9/Z56akBRlGyhePHibNu2njp1tlGp0ndMm9aPXr0+THM5oaGhxMREcO7cMW7diqZFixZp\nLuPAgQMkJ3cjZbpeV+LiWrJnzwFiY2P5/fff0el0WFuvBq4AoNV+Q0BAUJrrue/y5cvUrNmAM2cG\nAHmBr+5t+YekpD/Zvr0MISF1+HvlypSnAi5cgPLlYcMGcHZ+4XqfRavN+C5o+/btNGnSniZN2vPH\nH39kePmvIzXFsKIo2UZwcDDr1i1Ldzm2trb4+Pi88PF+fvm5cWMjIoWAZHS6rTg6lsXfvwTJyQUx\nGq+QJ48zFy8WxtraiVy5XPn5519eqK4bN24wcuRIkpIqkDLXQBlSBjr2IWXBorlAc9zic+Ld7h2I\nj4PgYPjtN3BxeeE2WsKWLVto0KA1BsMoADZubM6vvy6latWqlg0si1NPDSiKorxkJ06c4K23wkhO\nLojJFEmJEnk4fPgEBsMwoDuQiF5fi88/f5tGjRqRJ0+eFxrRHh4eTmBgRW7d8iMhIRY4SMr0MZdI\nedJgLtCePISznSAKEglBQfD77+DqmoEtfjlq1WrGxo31gE73XplDrVob2LDB8otIZab09nsWvTXg\n5+dHqVKlCAwMpHz58gBER0cTGhpK4cKFCQsLIzY21rz/uHHjKFSoEEWLFmXjxo2WCltRFCVdihUr\nxtmzR1i+fBAbNnxLUlISBsNdoM69PWwxGKpz+fIVfH19X/ixtlGjxhMd3ZqEhE2AG1ALGIG9fQhW\nVjmAYeRmMVt5g4JEEu3nBxs3vpJJAEBy8sMLSIEdRqPJUuG8MiyaCGg0GrZt28bBgwfN00mOHz+e\n0NBQTp06RY0aNRg/fjwAx48fZ+nSpRw/fpwNGzbQo0ePe2txK4qivHpy5MhBWFgYZcuW5dCh3aTM\nCPgdKbMExmBnt4SgoPStZnjlyg2MxuKk3ALYABQjV675LF8+nWrVKpHXxoqtdKIw4VzI4Yrb/v3g\n5pbepllMr17votcPAJYDy9HpBvDxx52eddgLS0hI4PTp0w99YX0VWXyw4KOXM9asWUOHDh0A6NCh\nA6tWrQJg9erVtG7dGhsbG/z8/ChYsGCmzkWtKIqS2davX0/Pnn1J+VM8GPgF8AHyULNmMZo1a5au\n8hs1qomDw2RS1h+IRa8/QrduXahfvz4b5s3ksMddipDEHX9//M6cfqWTAIAGDRqwePEMKlb8nooV\n5/LjjzOpX79+ptR14MABvL0LUrZsLby88jFt2lfPPiirSvdzC+mQP39+KVOmjAQFBcns2bNFRCRH\njhzm7SaTyfz7Bx988NDjNZ07d5YVK1Y8VqaFm6QoyissPj5e2rXrKk5OucTT018WL/4h0+r69ts5\notfnFZgqGk01AXfRavuITldBypULkaSkpHTXYTKZZOjQUaLXu4qdnZN07fphSrlXr4oEBKQ8Iliy\npMj16xnQouzDZDJJrlx+AkvvPYZ5XvT63HLo0CGLxJPefs+iTw3s3LmT3Llzc/36dUJDQylatOhD\n2zUazVOnyXzStpEjR5r/XbVqVTViVFGU59KjR19WrIjk7t393L59iS5d3sbHx5sqVapkeF1Dh47F\nYFgBlEPkY6yt6xIaeoIWLbrRtm1brK3T9udZRNi3bx+RkZGULVsWb29vNBoNY8YMZ8yY4f/f8dq1\nlNUDjx+H4sVh82Zwd8/Yxr3mbt++TXT0NeD+o6N+aLUhHDlyhNKlS2d6/du2bWPbtm0ZVp5FE4Hc\nuXMD4OHhQZMmTdi7dy+enp5cvXoVLy8vIiIiyJUrF5Cy/OaDy3tevnwZb2/vVMt9MBFQFEV5XmvW\n/Mrdu5tIuTzvw9273Vi37jeqVKnCyZMnmTjxS27fNtChQzPq1n3yEr8iQnR0NC4uLk/s0BMT7wI5\nzb+bTCWoUMGJjh07pjluEaFjx/f56aeNWFkVxWjcx6pVP1KzZs2Hd7x+HWrWhGPHoFixlCTAwyPN\n9T1a97Fjx4iNjaVUqVI4Z+K8AwsWLGLixG8QEfr160bHju9kWl1P4+TkhF7vyK1b24EQIBqR3fj7\nf/RS6n/0C+6oUaPSV2D6L0q8mLi4OLl1b9rKO3fuSMWKFeW3336T/v37y/jx40VEZNy4cTJgwAAR\nETl27JiULl1aEhIS5Ny5c1KgQIFU58e2YJMURXnFFShQWmCDedY9W9v2MmHCBDl9+rQ4OXmIRjNK\n4GvR631l0aLUVy/ct2+fuLp6i5WVo9jaOsqCBYtS3a9nzz6i11cT2CewVPR6dzl8+PALxb1x40Zx\ncCgmcOde7FvE1TWPefudO3ekS+PWclhrLQJyy8dHJCLihep6kNFolJYtO4pe7yMuLm+Im5uPHDly\nJN3lpmbZsuWi1/vde39+E70+vyxZkvZZHTPK77//Lg4O7uLiEiI6nZf07TvYYrGkt9+zWK957tw5\nKV26tJQuXVqKFy8uY8eOFRGRqKgoqVGjhhQqVEhCQ0MlJibGfMxnn30m/v7+UqRIEdmwYUOq5apE\nQFGUF/Xrr7+KTuchVlb9xd6+lfj4FJYbN25IgwZNRKPp/8DUvJukUKGgx46PiooSKysXgRn39jsq\ndnbucvTo0cf2TUpKkn79hkj+/KWlTJkqsnXr1heOe/bs2aLXd3ogPqNoNFaSmJgoIiKdGraUg5oc\nIiD/4iMFdDll//79L1zffUuWLBEHh2CBuHv1ficBAW+ku9zUVK/eWOCHB9q4VKpWbZgpdT2vq1ev\nyqZNm+T48eMWjeOVTQQyi0oEFEVJj/3798unn34m06ZNk+joaGnSpK1YW3sLjHmgE9oj+fKVfOzY\nIUOGCtjeW0MgZV+ttlGmL6qzb98+0elyC5wTENFoZoq/f6mUjVFRclBrJQJykkKSm3Cxtu4r48aN\nS3e9o0ePfmRFx2ui17umu9zU1KvXUuDrB+qaJXXqNM+Uul416e331BTDiqIoDyhbtixly5YFYM+e\nPWzcuJvk5CVAU8Af8ESv70337v+/Py0ijBz5GWPHTiJlQpv9QDAQh8gBfHw+ztSYg4OD+fzzYfTr\nVworKwfc3Jz55Zc1EBMDYWGUMRk5jQ/V2EoEedDZnMfFxT/d9ZYsWRK9fhhxcf0BV7TahRQtWjL9\nDQKuX7/O+vXrsbKyol69ev9j776joyreBo5/t2Sze3eTkELvhkBo0kFEeu+9iaA08QUVFRuKSlFA\nBcknKDwAACAASURBVBRBQX4goiBdCSJKD72DSFM6hBpKICGbZLO7z/tHYgwKpGxCCfM5x3Nk773P\nzKxlnp07hWHDXmHdutbY7TcAHZr2KcOGhWVJWY+8rMlHHhw5sEmKotwny5cvFz+/xinv3aGJ6HSB\n8t57w2+ZozRnzg9itZYV+E4gSCBAoL1AISlRouJt5zN5Ijw8XJ54oomUK1dLPv30s5T4sbGxEhER\nIU6nUyQqSqRq1aQ5AXnzSglzbtHp3hGzubM89li5lDlannC73TJo0BDx9vYXH58QKVgwRI4dO+Zx\n3OPHj0tAQEGxWjuIzdZa8uV7TM6fPy+7du2Sfv0GSZ8+A2Xnzp0el5NTeNrvqbMGFEVR7iAyMpIS\nJcoTE/MV0Bi9fiqFC8/i+PE/btn2t1evAXz//ePAIOB/wFAgmkaNGrN06SIsFkuW1WnRokV0794P\np/NLID9W6+s88UR+Dhw4gl6v4803X+KV3r2gSRPYsQOKF4f169kSEcHKlavw989Fnz598PHxuWMZ\nLpeLS5cu4e/vn666nz9/nhs3bhAcHIzJZErz/rS0a9eDn38ui9v9DgBG45s8+6yd6dMnexw7J/K4\n38uCZOSBkgObpCjKfbRt2zYpWrSseHlpUqlSbTlx4sR/7hk69D0xmfqmen/9P6lZs8lt4126dElm\nz54t8+fPl5iYmNve43Q6Zdq0afLyy6/JjBkzxOVyiYjInj17xGi0CXyQqqxdotMFCOwT2CV5LSUk\nMrhE0sVixUROncpQew8fPiwFCpQQiyWPmEw2mTJlWoaezwpVqjS4ZfUGzJPGjTve83o8LDzt93Jc\nr6kSAUVR7ubGjRuyc+dOiYiIyLKY165dk6JFS4vV2lw07Rnx8ckje/fu/c99f/75p+TKlV9stg5i\nszWRIkVC5cqVK7fck5iYKMWKlReoLDBWjMZq0rHjM+J2u6V16+4CTQReT9VJhguUFBCxES2bCEm6\nUKSIyMmTGW5L8eLlRKebmhz7qGhaftmzZ09mv5pMeeed4aJpjQVuCFwRTaspEyZMvKd1eJioROBf\nVCKgKMqdbN68WXx984qvb0UxmwNk+PDRWRY7OjpaZs+eLdOnT79jktG4cXvR6candOJeXv8nr776\n5i33DB48RCBQwJ58X6x4eQXJ0aNHpU6d1slLE/MkjwpMTZ6T8KzYiJaN1BIBuaxZRW4zcpGWuLg4\n0euNt6x6sFp7yYwZMzL1nWSWw+GQHj36icFgEqPRWwYOfDVlVET5L5UI/ItKBBRFuR232y2BgYUE\nfk7u5C6IphWW7du337M6lC79hMDGVL/mv5EOHXqmXL906ZIEB1cUKJ3qHhGDoZjs3btXvvnmW9G0\nUgILBDqLwZBXunXrLrmMvrKegiIgZ9DJT+PGZap+brdb/PzyCmxILvumWK2hsnr16qz6Cm4RFRUl\nzz77gjz+eG15+um+cvlfZx4kJiYmTXxU7srTfu++nz6oKIpyL8TGxnLjxlXg79Po8qHX1+bPP/+8\nZ3Vo1Kg2Fss4wA5cQdO+onHjpwCYNm0GRYuGcurUReA6MAY4CozE29tO6dKlee65Xowe/SKFCw+n\ncOFDjB8/jDcHDSTMFUcdznEWX+rzEX0+Gp+pyWM6nY55875F0zrg69sSq7U8nTrVo0GDBln3JSRz\nuVzUr9+KuXNd/PHHCBYutPLkk41xOBwp9xiNxlsmZSrZJGvykQdHDmySoihZILtHBG7cuCF79uyR\nS5cu3fGeuLg4adfu6ZQh75dffkPcbrecOHFCLJYggaMCx5JfDRQVCBSjMUC2bNkiIiJr1qyR0NDq\nki9fiPTr95LEXb0qF5JPETxHfinBkeRXDrZbdmXNqDNnzkhYWJhs3749y5c+iiTNlRg4cJB4eRUQ\ncCX/83CLj09Z2bFjR5aXl9N52u/luF5TJQKKotzJli1bbpkjMGKE57vriSR10DZbbvH1LS9mcy6Z\nPHnqXe9PSEi45ZjhFStWiJ9fg1SvAy6KyRQoH330kURGRoqIyIEDB0TTggR+EjgoAebmciBfARGQ\nC+ilZMpw/q/i758/zQ587dq1kj9/CTEavaVKlbpy5swZz7+IdNizZ49YrUGi0/UVyC3gSK63U2y2\nkCzZ+vhRoxKBf1GJgKLkLE6nU06fPi3Xr1/PkniZWTWQkJAgEyZ8Jv37vyjTp0+/ZeKaw+EQH5/c\nAmuSO7TjYrHkkT///DPd8Y8fP548InA8OcZWsVoDJDY2NuWeTz75RIzGwQIiZuyykjpJWUPevPLV\ny0PEbA4QX99K4uOTRzZs2HDX8k6fPi1Wa5DArwIxYjAMl9DQKtny6z+1b7/9Lnkr5MnJExJbCbQQ\nmC1mc1epVq2emhOQCSoR+BeVCChKznHy5EkpVqysWCz5xWSyybBhI+95HZxOpzz1VFOxWJoLfCaa\n9oT06jUg5XpERIRYLPlumdzn69tCwsLCMlTOl19+LWazv/j6VhFNC5Sff172r+tfisXSRbyJk99o\nIgJySacXST7wJiIiQnbs2JGuhGn+/Pni49M+VZ3dYjL5yLVr1zJU54wICwsTTSsi8ESq1zNxAl0k\nKChEhg59/5bER0k/lQj8i0oEFCXnqFKlruj1Y5J/PV4Uq7XEHU8ezS5bt24Vmy1UIDG584oWk8kv\nZS5AQkKC2GyBqVYDnBGLJW+mTqQ7f/68bNu27T97C4gkzbAPLlhCftMlrQ6IRC+LRozKVJtWr16d\nvCXy38PyJ8TLy5JyWmF26NChl8A0ga8EKkjSBki7RdNKybffZu+hTDmdp/2eWjWgKMoD68CBPbjd\nAwAdkJf4+Hbs2bPnntbBbrej1wcCf5/RZsNotGK32wEwmUwsXDgbq7U9fn7VMZsrMnLkUEqXLp3h\nsvLnz0+NGjUIDAz8z7VcFguHyhSnqZzjptnC0a+n0PH9YZlqU/369alduxRWa21MpsFoWh3GjfsU\nLy+vTMVLD19fDZ3uEvAC0A1ogZdXEz78cCC9ej2TbeUqaVNnDSiK8sAKDq7AiRPvAF2BeKzWusyY\n8Rpdu3a9Z3WIiYmhRInHuXJlEG53M7y8vqFkyc388cdW9Pp/fktdvXqVI0eOUKhQIQoXLpy1lUhI\ngE6dYNkyCAyEtWvh8cc9CulyuVi8eDFnz56levXqPPXUU1lU2dv766+/qFatNrGxz+F2W9G0yfz6\n62Lq1KmTreU+Cjzt91QioCjKA2vnzp00atQana4cTucpGjWqzo8/zr6lA86omzdvomlahmIcP36c\nPn1e5tix41SuXJFvvplE7ty5M12HDHE4oHNnWLoUAgKSkoAKFe5N2ZkUGRnJvn37yJcvH+XL/3Ms\n8bFjx5g+fSaJiU569Oiactyz4hmVCPyLSgQUJWe5fPkyu3fvJiAggGrVqqHT6TIV58iRIzRr1pGI\niGOYTBa++246HTt2yOLaesbtdnP69GmMRiOFChVC53RCly6wZAn4+8OaNVCp0v2u5l2Fh4fTunUX\nDIZyJCYeoVevznz11YRM/3NT0qYSgX9RiYCiKP8mIhQvXpYzZwYhMhDYg6Y15/ffNxMSEnK/qwdA\ndHQ0DRu24dCho7jdiTSq+xRhFkG/ZAnkygWrV0OVKumO53K5SExMxGw2Z2OtbxUWFkaXLs/hcMwH\nmgDRWK1VWbp0arbsTqgk8bTfU5MFFUXJ8W7cuMH58xGIDCJp4mEVDIa67N6926O4YWFhBAdXIl++\nErz00uu3bI97N3/99RcvvTSEAQNeZuvWrQC88spQ9u8Pxm4/gzP+BL1XbU1KAvz8YNWqDCUBY8aM\nw2LxwWbzo3btZkRFRWWqfRkxefIUund/BYcjBmic/KkvIrU4fvx4tpeveMCjNQcPoBzYJEVRPOR0\nOsVs9hXYn3Kin9VaUtavX5/pmFu3bhVNyyuwQuCQaFpjGTjwtTSfO3TokNhsuUWnGybwsWhaHlmx\nYoWUK1dLIFwMJMo8uoiA3DR6iWRwC+RffvlFNC1Y4IxAophMA6R1626ZbWaaEhMTZdmyZWK15kle\nElhaYGbKUkpNK3hPD3Z6FHna76kRAUVRcjyDwcD06VPRtIbYbE9jtVahbds61K5dO9Mxlyz5Gbv9\nBZKGwEtjt09i4cKf0nxu3LjJxMa+hMgo4E3s9km0afMs58+fw9sQxvf0pCsLiMbIt92fgerVM1Sv\n9es3Ybc/CxQGjDgcb7Np06ZMtDBtiYmJ1K3bgm7dhhMbawcCgAXAcKAABkMoI0e+QfUMtkG5t4xp\n36IoipJ9zpw5Q/v2Pdm3bxt58xZlzpxp1KtXL8vL6dGjO5UqVWD37t0UKtSfevXqodPpiI6OZvr0\n6Vy5co0mTRqlu2xfXxteXidITPz7k3NYrbY0n4uNjUck9T4BgSQkFCAxoSff8QbdcRKDnpdLluHL\nKZMy2kwKFcqP2byS+Hgh6TXIDvLmzZ/hOOnx/fff8/vvLuz2bcCrQC/gY2AkFssrhIevU0nAwyCL\nRiYeGDmwSYqSY7ndbilRooLo9R8KxAosF6s1KEPnAHgiJiZGgoPLi9ncRXS64aJpBWXGjJnpejYy\nMlLy5CkmXl79BEaKpuWTxYsXp/ncb7/9JhZLAYHlApsEyomeiTKLniIg8V5esn/q1FsOJcqIuLg4\nqVixlthsT4rN1lVsttyydevWTMVKy6hRo0Svfzv5NYBDYLDodP5SpUp92bx5c7aUqfyXp/2eWjWg\nKMp9c/nyZQoXLklCwjWSfr2Cr29bZs58lg4dsn9p3/Tp0xk8eCl2+9LkT/bi79+K33/fypIlS9Dr\n9XTq1Il8+fLd9vnIyEimTfsfN27cpF27VpQtW5Z58+Zht9tp2bIlpUqVuu1zCxcu4oMPxvPXX0fA\n3Z8ZXOQ5ZnETI6tfe5l248dnuC0nT57kvfdGc+nSNVq3bkCRIgWJiYmhbt26FClSJMPx0mPt2rW0\nbt0Huz0cKILR+Aa1ax9l7dqlaT2qZCGP+70sSEYeKDmwSYqSI9ntdlm8eLEYDBaBU8m/KhPEZist\n4eHh96QO48ePF5PpxVSH71wVk8kqPj55xNu7r5jNvcTfv4CcPHkyzViXL1+WAgVKiMXSWUymgWK1\nBsmmTZvu+sy4TybILINP0sRAjNLWP99tzxlIy4ULF8Tfv4Do9cMF5ovVWknefXd4huNkxvjxE8XL\nyyIGg7dUqVIn5QwG5d7xtN/Lcb2mSgQU5cF37do1KVGigvj41BaTqZxAkJhML4rVWk1ateqS7cfh\n/u3AgQOiaUECywSOi9ncRfLmDRGdbnxKcqDXvy89ez6fZqxhwz4QL6/+qZKKuRIYGCwFC5aW8uVr\n/WeFws3oaNlVqYoISJxeL1927iYXL17MVDsmTZokZvOzqco+IVZrYKZiZYbT6ZSbN2/es/KUW3na\n76lVA4qi3HOjR3/KmTNViYlZj8OxH52uI489tpFvv32TsLC592wXurJly7JkyQ8EBw8jIKAe7dv7\nULBgIUT+OTDI7S7NxYtX04wVGXmNxMTQVJ+Ecu1aDOfOzWP//sE0b96Rw4cPA3D18mUWBOWnyt7d\nxOFFO0NpNhk18ubNm6l2uN1uRFLP/TYC9+4VqcFgwGq13rPylKylEgFFUe65Y8cicDie4u95ASI9\n0Os1OnXq5NE5ApnRuHFjjh3by9WrZ/jhh+l07NgcTRsFRAAn0LSxtG/fJM04rVo1RtMmAweASGAI\nIu2Bx4HOJCR0YeTIkSTExxNe9nF6O2KJw0xrlrMicRuLFi3kypUrmWpD+/bt8fZehk43HliOpnWh\nX79+mYqlPHpUIqAoyj1Xv/4TaNr/gBuAA7N5EnXrPnHH+91uNx999AmPP16bOnVasn379myr29tv\nD6F//9poWkVsthq89loHXnihf5rPtW7dmrFjX8fXtxHe3iF4ef0O/NMZu1wX+HHxbsIKP0bHyxeJ\nR0dbwlhDI8CCiDHdOxP+zel0cv78efLmzcu2beto2XIXNWpM5L33OjFhwpgMtlx5ZGXRK4oHRg5s\nkqLkOC6XS/r2HSQGg7cYjRZp2rS9xMbG3vH+t956TzSthsBagW/Eag2SQ4cOZUldDh8+LJMmTZJZ\ns2aJ3W73KNbYsePF29tXbLYSYrUGitlcWGCcQF+BYJlI/6QlgiBNyS0wSmCbQE957LHHMzQ3YtOm\nTZIrVz6xWPKIpvnL0qU/e1T3++mrr76WQoVKS758IfLBBx/eszkiOYWn/V6O6zVVIqAoDw+73S4x\nMTFp3hcYWETgz5TJcAbDEBk5cpTH5a9evVo0LUjM5gFitTaV0NAqmZ70tn37dtG0QgJnk+v5vQQG\nFhCdziLwtnzG88lJgF4+fLKWmM0FBCqKTldAfH0LyYULF9Jdlt1uF1/fvAK/JJe1TazWoAzFeFAs\nXLhINO0xga0C+0TTKsunn352v6v1UPG031OvBhRFuW8sFgs2W9q78RmNXkBsyp/1+li8vNK3MarL\n5eLSpUsk/rMFYIr+/V/Dbv+W+PipxMb+yunTRfjmm2/uGs/pdHLp0iVcLtctn//xxx9AQ6Bg8ic9\niIqKpE7tunymX84rTMOBkadNPrSYPImff57F0KEt+eyzNzl//s877lVwO6dOnULEF2iR/EkNjMbS\nKZMRM2v+/AU0bNie1q27s2PHDo9ipdecOUuw298FngAex24fy5w5S+5J2UoSlQgoivLAe/fd19C0\nrsAM9Pr3sFrDeOaZZ9J8bsWKFfj55aNQoVB8fXOzZEnYLdevXbsMlE/+k474+HJERt55wt6qVavw\n989PsWJlCQoqxMaNG1OuhYSEoNNtAq7/XTr+ufLya/lgXnH/gQN4zuZLt9nTqVSpEo0aNWL06A8Z\nPHhwhmfc58+fn8TESOBo8icXcTj+onDhwhmKk9rMmbPo0+dt1q7tyrJldahfvyV79+7NdLz0ypXL\nhl4fkeqTCHx9004OlSyURSMTD4wc2CRFUURk/vwF0r59T+nf/8V0bfBz7Ngx0emsAouSh893iNkc\ncMv2xe3bPyPe3j0FYgT2i6YVknXr1t023pUrV8RqDRJYnxzvV7HZcsvy5cvl4MGDIiLSo0cfgVwC\njwuYZUPNmknvMoxGkSVLsuJrSDFt2gyxWHKLr28r0bT8MnLkWI/ihYbWEFiVai+Cj2TAgJezqLZ3\ndvToUfH1zSsGw2DR6YaKpgXJli1bsr3cnMTTfk8dOqQoSrbZv38/3bs/z+nTxylXrgLz58/I9Ha3\nXbp0pkuXzum6NzExkY4duyNiAzomf1qNxMRQ3n9/BH/8cYLAwFy8996rxMdPZNWqICwWX8aPH3PH\nQ4cOHz6M0VgCqJP8STNu3rTQpctQXK7LdO/ejrVr1wOvAzX5iJnU3jobMRrRLVgAbdtmqt130r9/\nH+rWfYpDhw4RHDya8uXLp/3QXST1J6n3b7g327WXKFGCffu2MWvWdyQmOunWbR3lypXL9nKVVLIk\nHXmA5MAmKcpDKSoqSvz9CwjMEDgvBsMoKVasbKYP00kvt9stzZt3FJ2umIBV4HDyL9wrAv5iNpcT\n+FXgS7HZcsvRo0fTNUv91KlTYjYHCpxPjndSwFcgUiBaNK2MGI0WAbeMZJgISCI62fjKK9na3qwy\nffo3yZP25gtMEU0Lkt27d9/vainp4Gm/p+YIKIqSLfbs2YPbHQz0AfLjcr3L5cvRnDlzJlvLPX36\nNOvWbURkBOAH1AbaAKEYDG7i438EmgEDiY/vxqJFi9K1k2HRokV599030bQq+Pq2AyoAI4HcgA9x\ncbVwuZx8wAu8x4c4MdDHOze6Tp2yra2ZdfDgQfr3f5EePfqzdu1aAPr27c3//vch9er9QIsW61iz\n5mcqV658n2uq3Avq1YCiKNnCz88Pp/MckAB4A1EkJt7A19c3W8t1OBzo9d7AM8AV4GNgJQ0b1mfP\nngNERSWk3KvXJ2AwGNIde9iwN2nTphlHjhxhyJC/iIgwkzR6fhmRVbyv82c403AB/bwDMPfqSq1a\ntdKMe+HCBb777jsSEhx06NA+W4fGDx06RI0a9bDbByPix08/9WDevK9p06YNTz/dnaef7p5tZSsP\nqCwamXhg5MAmKcpDye12S+vWXcVqfVJgmFitZWXw4DfveP+OHTukadNOUrNmM5k69X+Z3lTG6XTK\n44/XFJPpBYFNYjS+KcWKlZW4uDgZO3acaFppge9Frx8ufn75bpk8mBGHDh0Sf/+CAgUF/ORd6ouA\nOEHCunWTrVu3pivO2rVrxWDwEXhW4DUxmwNk48aNmapTejz//Eui041MNSnwJ6lYsW62ladkP0/7\nvRzXa6pEQFEeHE6nU7777jt5//0P5Keffrpj575///7kGflTBJaIppWRceM+z3S5165dkx49+kto\naA3p0KFnykY7brdbZs6cJS1adJWePZ+XY8eOZboMEZH169eLppWQt3lHBMSFTnqgSe7cRdJ1nHBc\nXJxYLAECr6XqmGdL0aLlZf/+/R7V7U569Rog8Hmq8tZImTI1s6Us5d5QicC/qERAUR4+b7wxVHS6\nd1N1TtulcOGyWV6O2+2W1atXy/Tp07NkIpzD4ZAJ+QqnJAE9qSTQWkymp+Wzz9LeHW/fvn1iNOYT\n+DJV2zeLTldALJbcMnfuPI/r+G/h4eFiseRNXla5WjStjHzxxZdZXo5y73ja76nJgoqi3Hd6vQ5I\nvVOfM8uPIhYR+vZ9kbZtBzF48EZq127FV1997VFMr4kTefViBG6gD+X4nrbAQlyuAsTGxqb1OAEB\nAeh0N4FPgJ0kbRD0EiKdiYtbSd++L3i0hC8+Pp4NGzawefPmlAON6taty8KFM6hS5UvKlfuAjz9+\nkRdf/L9Ml6E8/HTiyb9lDyCd7t6sfVUUJev89ddfVKnyFHb724gUQNPe59NPX2fgwAFZVsauXbuo\nV68zsbH7ARtwApOpAlFRl9A0LeMBJ0yAIUMAmPlUfQbtNhEXNw44jsXSj61bV1OhQoU0w7z00htM\nmzYHh8MFxAGhwFZAh15vJjY2GrPZnOHqXb58mRo1GnDlijeQSJEiZrZsWZXtkzWVe8/Tfk+NCCiK\nct+VKlWKrVvX0rHjHzRpsphp00ZmaRIASTPzDYbSJCUBAI9hMNi4du1axoN9/nlKEsC0afRY8xu9\ne5elQIFOhIaOZunSuelKAgC++OITFi+exquv9sBk0gFfkZQEjCU0tGKmkoATJ07QsmVnzpxpSEzM\nTmJifufYsbK8996HGY6l5HxqREBRlEfC2bNnCQ2tRGzsYpL2FphO/vwfExHxV4aWEDJpErz8ctLf\nT50KA7IuYVm0aDHPPfc8cXHRhIZWYvnyhRQtWjRDMbZt20ajRq2x230RmQo0Tr6ygEaN5rNq1eIs\nqevx48c5ffo0pUuXJn/+/FkSU8kcNSKgKIqSDoUKFeLHH+fg59cFvd6bYsUmsmbNzxlLAr788p8k\n4KuvsjQJAOjUqSMxMVew229y8OCODCcBAP/3f28SGzsRkbbAt4ATSMBimU3NmhWzpJ5jxoyjfPma\ndOgwghIlyvPTT+q0wIeZGhFQFOWRIiIkJCTcdsh93rz5DBz4GjEx16hbtwndu7flhx9+xsfHysTS\nhSk6dmzSjZMnw6BB97jm6VOoUGnOnVsAFAc6AHswGFw0alSfsLC5eHt7exT/8OHDVKlSn7i4PUAB\nYBea1oSrV89n6jWG4rmHfkTA5XJRqVIlWrduDcC1a9do3LgxJUuWpEmTJly/fj3l3jFjxhASEkJo\naCgrV668X1VWFOUhptPpbtth7dq1iz59BhMVtQSn8zLr1uWmX7/XWbu2M3nCnP8kARMnPrBJAECT\nJg0wm0cAbuBLzGYr06aN59dfF3ucBEDS/AOTqSJJSQBAVUDj0qVLHsdW7o/7nghMnDiRMmXKpCwV\nGjt2LI0bN+bIkSM0bNiQscn/8R06dIj58+dz6NAhfvvtNwYOHIjb7b6fVVcUJQNiY2N57bW3adCg\nHa+//k66ltfdS+Hh4Tid3YFqgA23exzgoC92pjEfgMW16v7zauABYLfbCQsLY/HixURFRQEwefKn\ntGhhwWjMi8VSnVGjXqVPnz5ZthyzdOnSJCbuAf5M/mQlRmOimifwMPNoFwIPRURESMOGDWXt2rXS\nqlUrEREpVaqUXLx4UURELly4IKVKlRIRkdGjR8vYsf+ct920adPbbuF5n5ukKMptJCQkSFBQcYH2\nAovEYOgo1arVE5fLJcePH5cePfpJ48YdPdpa2FMzZ84Uq7WxgDt5Y59N0ptc4kInAvIazWTgwAfn\nJMGrV69K8eLlxGarKz4+zSUoqIicOHEi5Xp2fo/ffDNLzGY/8fEJEV/fPBIeHp5tZSlp87Tfu68j\nAq+++iqffvopev0/1bh06RJ58+YFIG/evCnDTefPn6dQoUIp9xUqVIhz587d2woripIpr732Jleu\nxAMLgY64XPPZt+8Ya9eupWLFmsydW5BVqzoxZMhkhg//6L7UsVu3bpQoEYvV2hij8f/oRSOmcx09\nwht0YAIb6devV5pxlixZQsuW3ejc+Tn27t0LJL0CzWojR47l3Lla3Ly5jpiY5URFDeCll95OuZ7V\nGzKl1rt3Ly5cOMX27WFcuHCSunXrZltZSva7b6cPLlu2jDx58lCpUiXCw8Nve49Op7vrv8x3ujZ8\n+PCUv69Xrx716tXzoKaKonhq48ZdJJ1A+Pd/szqcTqF37/7ExDQChgMQG1uVzz9/ihEjhmVp+SLC\nvHnz2LRpB8HBRRg48P/+M0/AbDazfftaFixYQNCvv9J0bjx6YKgukAm6dUwYN4pKlSrdtZzZs39g\nwICh2O0jgSiWLWuIt7eJ6OjLlChRgWXL5lGyZEmP2rJ3717Cw8NZv34bDkd//v5OXa5anD693KPY\nGZErVy5y5cp1z8pT/hEeHn7HfjMz7lsisGXLFpYuXcry5cuJj48nOjqanj17kjdvXi5evEi+2XUg\nJwAAIABJREFUfPm4cOECefLkAaBgwYJERESkPH/27FkKFix429ipEwFFUbJOYmIiDocDq9WaoedC\nQh7jjz8uAs8DnYB5mEwOLl70A1LH8sqWuT9DhrzD118vx27vidkczty5YWzZsgovL6+Ue9xuN97e\n3vTU62HePAAO9ehBSIMG/Fm7NiEhIWmWM3r0JOz2/wFNAIiPjyY+fh+wgGPHvqJhw9acOnUoY0sW\nU1m0aDG9eg3E5eqCyGV0uomItAHMmM1fUK9ezUzFVR4u//6BO2LECM8CZs0bCs+Eh4enzBF44403\nUuYCjBkzRt566y0RETl48KBUqFBBEhIS5MSJE/LYY4/d9h3YA9IkRclxRowYLUajWYxGs9Ss2Uiu\nXbuW7mdPnjwp/v4FxGgMFb2+qJjNgdK0aTuBMQJ5BCYI/CI6XQUZMmRoltb75s2bYjRaBK4kv/t3\nic1WWVauXCkiImvWrJHcuYuJTqeXQQH5xa3XJ53+M3JkhssqVaq6wNpUBwiNFXgx5c8WSx45e/Zs\nptsSGFhYYEtyPKcYjcVFrzeJ0WiWFi06id1uz3Rs5eHlab/3QPSa4eHh0rp1axFJmgDTsGFDCQkJ\nkcaNG0tUVFTKfR999JEEBwdLqVKl5LfffrttLJUIKErWCwsLE00rKXBWwCkm0wvSpk33DMW4fPmy\nzJo1SyZOnCiDBg2WypVrisnUSGCvQFeBklKuXHVxuVxZWvcrV66IyeQr4EzpkH19m8tPP/0ku3bt\nEp3OKjBHujBbnMkTA+OHpj8ZOXXqlOzYsUOio6NlypRpomkhAksEvhGwCWxILvekmExWiY2NzXRb\nvLw0gesp7TCZXpKxY8fKzZs3Mx1Tefh52u+pDYUURUnTG2+8zbhxPsC7yZ8cJzCwAVeunM5QnMuX\nL1O2bFWiotrhdD6GXj8Kg0GHyeRLkSIBbNjwK0FBQVladxGhRo0G7NtXGofjJXS6Dfj5jeDo0T9o\n0KA1+/dfpxMjmUt3jLgYbQii0ZZfqF69epqx33zzPSZNmoLJVAS9/iJTpkxg48ZNbN68n4AAfwwG\nJ9u2ncblegKd7ldGjx7K4MGZ34OgUaO2bNyYH4fjE+AQFktbNm36lcqVK2c6pvLw87jf8zwXebDk\nwCYpyn03ceJEsVhaC7iSf43OlnLlamY4zmeffSbe3r1SDZ3vlly58svBgwclMTHxrs96shzu6tWr\n0rbt05IvXwmpVq2BHDhwQERE8uYNlg5YJRGDCMgoXhOD3ldOnjyZZsx169aJ1Rqc6pXDAtHpfMTX\nt41YLHnls88midvtluXLl8uUKVNk27ZtGarz+fPnZeTIUfLmm0Nlx44dKe1o2LCteHlZJCCgkCxY\nsDDD30Vqbrf7vi3XVLKOp/1ejus1VSKgKFkvLi5OKlV6Smy2J8THp5PYbLlTOqeMGDNmjBiNr6RK\nBE6Jr2/euz6zcOEi8fcvIAaDSZ56qplERkZmthn/8UnNeuJIfh0wmooCBaRp0zZ3fcbtdssvv/wi\nXbt2FW/v3qna4hTQCyQKnBKzOZdcuHAhU/U6d+6cBAYWEqPxBYH3xWLJLb/++mumYt2O0+mUQYNe\nE29vm3h7+8hrr72d5a9klHtHJQL/ohIBRckeCQkJsnTpUvnhhx8yPeHt0KFDomlBArMFtoumNZAB\nAwbf8f69e/eKxZJHYJtArHh5vSy1azfPbBNutWSJuI1GEZBPdUbR64zSrl2Xu/5Cdrvd8uyzL4jV\nWlZMpjYC+QQuJycC8wRKpZqH8Ljs3r07w9VauXKlBAQUFBiYKskIkzJlnvCktSmioqKkV6++4u39\nhMBFgXOiadXk888nZUl85d5TicC/qERAUR5smzZtkqpVG0hwcGV5/fV3xeFw3PHeiRMnirf3/6Xq\nEGPFYDB5Ppy9dKmIl1dS0CFD5OqVK+maxLd3717RtMICMcn1eVXAKjZbeQFNYGry56vEZsst169f\nz1C1fv/9d7FYggTaCXxyyyuUIkXKZba1KY4cOSJBQYVFry8ssCxV/IXSoEE7j+Mr94en/d5920dA\nUZRHU61atdi5c0267s2dOzdG42ISEtwkHY1yAF/fIM92zfvlF+jUCRIT4ZVX4NNPCUhnvMjISLy8\nSgC25E8mYLHMZ86cD9HpdPTo0YfExGGYTHrCwhbg5+eXoaotX74ch+NZoAXQk6RzD/Kgaa/StWu7\nDMX6N7vdTu/eL3Pt2qu43buBP4CWAOj1+8mfP2snaSoPkSxKSB4YObBJivLISkhIkOrV64vVWke8\nvQeKxZLHswlyy5eLmExJP4NfflkkgyMLkZGR4uOTR2CpgEN0uimSP39wyqhGYmKinD9/Ps2Jj3cy\nceJEMZu7p3rVUFJ0Oj959dW3Mx3T5XJJ376DxGg0C/gJ/C5wRCC/QDcxGDpKQEBBOXXqVKbiK/ef\np/2eWj6oKMoDQ0T45ptvWbFiA0WK5OOdd97Ax8eHhQsXcuXKFerUqUPFihUzF3zFCmjbFhIS4MUX\n4YsvIBMjC5s3b6Zz52e5ePEkJUpUYOnSHwgNDc1cnf4lKiqKcuWqc+VKbRyOEmjaFL788iOeey7t\nMw7u5IsvvmTo0B+w25cDLwEmYBpwFC+vZnTrVptx48al7OKqPHw87fdUIqAoygPjjTfeZcqUX4mN\nHYjJtIv8+Tewf/92fHx8/nOv2+1m9OhPmT59DiaTiQ8/fIsuXTrfNq57xQpcLVvi5XLxrcWXmx+P\n4cWXBnpUV7fbfcuBaVnl6tWrTJ36NVevXqdVq2Y0aNAgU3FOnz7NoUOHGD/+a9asaQv0BqKAxuh0\nf2E0Cs8914evv56YrQcUKdlP7SPwLzmwSYrySHA6ncnD1xdTJrHZbE1l7ty5t71/9OhPRdOqCOwQ\nWCkWS4GUbYNvsXq1JBiS9gmYQhfRsUM0LVjmz1+QzS26f2bP/kEslkDx82ssBoOfGAy95O/jlXW6\n0VK/fqsMT2RUHlye9nv39RhiRVGUv7ndbkTcwD+//kX8cDgct71/1qyF2O2fkTShrjFxcW8ze/bi\nW29auxZHs2aYXC6m0ZKBzEWoht3+DvPm/Zxtbbkbt9vNzZs3sy1+TEwM/fr9H3Fx4dy4sRKXaxtu\n909YrU/h49OcwMApTJ8+McMTGZWcSyUCiqJw4sQJVq1axenTGdsyOCt5eXnRunUnLJYewFZ0ukkY\njRto0qTJbe+3WjUgMuXPev0lfHy0f25Yvx5p2RKT08l0CvIC7ZDk/+XpdCcJCPDNxtbc3jfffIum\n+eHvn5syZapx5syZLC/j4sWLGI0BQLnkT0Lx8anMm282Zdas5zlyZB+PPfZYlperPMSyaGTigZED\nm6Qo2WrSpClisQSJn199sViCZPr0mfetLnFxcTJo0BApWbKa1K/fWg4dOnTHe1esWCGalkfgI9Hr\n3xBf37xy7NixpIvr14tomgjI90Zf0bFFIEjgdYHnxWrNLcePH79HrUqya9cusVjyCRwWcIvBMEoq\nVKiVJbEPHDggM2fOlBUrVojdbhdf3zwCvyW/YtkrFkugnDlzJkvKUh48nvZ7arKgojzCzp49S0hI\nBeLjdwHFgb8wm58gIuJolh/+kx22bdvGnDkLMZtNlC9fmvXrt1MhJooXfwlDb7fj6tGDQis3cenK\n24hUBT7G23sFu3dvpWzZsvekjgkJCQwe/BZz5swmNrYlIrOSrzjQ6zUSEx0eTTqcM2cuzz//Cjpd\nE2AvzZpV5uWX+9GmTRecThMuVzSzZk2nS5dOWdIe5cGjJgv+Sw5skqJkm82bN4ufX/VUO8yJ+PqW\nk717997vqmXInDk/iKYVlJq8LNEk7Rh4s0MHEadTDh8+LKGhVcVo9JbixctnattfT/Tt+6JYLM0F\nJgtUFEhI/q43iL9/AY9iO51OMZt9Bf5IjhknVmtpWb16tcTHx8vx48c9OvZYeTh42u+pnQUV5REW\nEhJCYuJxYCdJk+424XJdoHjx4ve5Zhnz7rtjKW9/h994Gx8SmUMIGwOCmGowEBoayuHDO+95nc6f\nP8/EiV/y/fc/4HBMIGmnwHDgcUym0hiNm5g9e9bdg6TBbrfjdCbyz3wAMzrd41y4cAFvb281F0BJ\nFzVZUFEeYblz5+aHH75B05pisz2G1dqOxYvnPDQzyi9fvsz69espdf0KK3gLX2KYSzeepSszvv2B\nffv23Zd6Xbx4kQoVnmD8+FgcjmHAu8BsYD5GYyE6drSwb99WWrRo4VE5Pj4+FC0agk73GSDAblyu\nNVSrVs3zRiiPDDVHQFEeMW63mwULFnDq1CmqVq1Ko0aNiI2N5fz58xQsWBBN09IO8gBYu3Ytbdp0\npRoFWBL7B37AAhrwNC/gYhDQmeeeczNz5pR7Wq+VK1fStWtPrl9vBvz9i38L0BmTqSmBgRs4cGAH\nAQEBWVLe8ePHad68EydOHMLb28p3302nY8cOWRJbeTh42u+pVwOK8ggREdq06UZ4+BkSEp7CZBrA\n0KHPM2zYW4SEhNzv6qWbiNC+fXdCY0fyE0PxAxbhRQ9O4mISMAf4E4fjj3tar7FjP+addz5EpA1Q\nINWVIMxmB++/X5Lnn/80y5IAgODgYI4c2UtcXBxms1ntEqhkmBoRUJRHyObNm2natC+xsX+QtOf8\neby8QoiKisRqtd5yr4jw/fffs3nzLkJCivLii4Mwm824XC62bdvGzZs3qVGjBrly5brn7bhx4waN\ng/KxwmnGn+v8SHt6mSCRrTgcM4FoLJaXWb58HvXq1bsndVqwYCHPPDOAxMSOwCCgCTAFKIqmvU7/\n/jX4/POPMxz3xIkTvP/+GC5dukbHjs0YMKCf6uyVW6gRAUVR0u3atWsYDMVJSgIA8mMwaERHR/8n\nERg0aAizZq3Hbn8GszmcBQuWER7+C02bduD33yPQ6/Pg5XWMTZtWpRy643A4mDlzJhERZ3nyyZoe\nvwO/E9/jx1nhSsCfeJbQlm58gpexDm8NGcDPP4/F29vEBx/M/E8S4HK5MBgM2VKnqVPnkJjYHTgK\nVADmAW+h0x1n0KBBjB79QYZjnj9/nipVniI6+gXc7kZs3TqGCxcuMWLEsCyuvfJI82jNwQMoBzZJ\nUbLMhQsXko/RXSRwTQyGURIc/Li4XK5b7rtx44Z4eVkFriUvS3OJzVZBXnzxRbFYmgk4k/et/0Jq\n1GgkIklH8D7xREPRtCYC74umlZAPP/w46xvx++8Sb7OJgIThJV4UETDLK6+8fsdHdu7cKQUKhIhO\np5dChUplyxLCFi26CHwhUE2gjcBQ0etzy/Tp32Q65meffSbe3n1SLe88Ir6+ebOw1kpO4Gm/p1YN\nKMojJF++fKxcGUaxYsPx9i5K5cprWbv25/9saBMXF4de7w38vXpAj16fmxMnIoiLawgk/aoWacyp\nUycBWLVqFQcOXMdu/xUYgd0ezvvvv0dgYDFy5y7Oe++NxO12p7uuq1evpnXrbnTs2IvNmzcnfbh/\nP6769fG+eZNllKEzkSTyKzCSv/66/fbIf/75J3XrNuf8+Y8QcXD27HAaNWpNbGxsuuuSHsOGvYKm\njQTaAHqMxolMmjScvn17ZzqmiHDrKm+v5PMYFCULZU0+8uDIgU1SlHvO7XZL5cq1xctroMAB0em+\nkICAgvLVV1+J1VpFIErALV5er0qLFp1FRGTevHni49Mh1a9Xl4BJIFzggHh7V5RPPpmQrvJHjRol\n4CswVeBLMZuDZPesWSJBQSIgy9HExHepyvpRatVq8Z84mzZtEosll0DovzZNejxbRgV27twpffoM\nlN69/0927tzpcbzTp0+Lj08e0ekmCPwimlZDhgwZmgU1VXIST/u9HNdrqkRAUbLG1atXpV27HlKg\nQCmpWLG2LFq0SOLi4mTAgMHi5WUTiyWPlCtXQyIjI0VE5OzZs2Kz5RaYLxAhMFCgbKoOeKWUK/dU\nmuWePn1a9PoAge9Tni3N+xJl8hYBOV+hgviaygiUENgosEWg8H+G4Ldv3y6ali85mQgSuJIcL1K8\nvQMkIiIiW763zLp586ZcuHBB3G73LZ8fPHhQWrXqKtWrN5aPPx7/n9c4iuJpv6dWDSiKckdOp5N2\n7Z5m3bqtGAx++Pu72bx5JZqmYbfb8fb2ZsKELzh//jKtWjWkSJEi9Oo1iHPnzmC3uxDpC3ySHG0G\nVarMYdeutSnxz5w5w4wZM4mPT6Bbt85UqlSJn3/+mfbtX8Ll+gToQiiHWccT5CMaZ/36OBYtouIT\nDThxwozLdQmdLoZmzZ7il19+SplNv2PHDurXb4ndHgecBiYA84FawErq1CnP+vUr7+VXeVfDh3/E\n6NGjMRgsFC1alDVrllKwYMH7XS3lIaFWDSiKckdxcXF4e3tn+lCbqVO/Zt26q9jtxwBv7Pbh9O79\nEqtW/YTBYKBs2WpERjYgMbE8ixa9x6uvdiU4uDhXrlwiNvYq8C0QB1iALxk7Niwl9smTJ6lU6Ulu\n3uyCy+XH5MlN+OWXhRQoUACDIRaXawgluchaPiAf0azW+bO8VHkmBASwa9cGvvhiMhERF2ncuA6d\nOt16oM748VOw24cBy4HPgA+BisD/ATUpX/7B2DMhJiaGhg1bsnPnMeAYiYn5OHZsOF269GHz5hX3\nu3rKoyILRiUeKDmwSYqSYZcuXZKqVeuJwWASk8kqkydPyVSc/v1fFPgs1fD+filYMFRERGbMmCGa\n1i7VtWOi01nEy2uwwDGB5wW8xcsrjxiNmnz++cRbYg8c+Iro9UNTPT9XqlVrKCIiAwYMlhD85BwG\nEZDVFBILC247D+B2qlWrn/xK4FzyLH6bgJfAm6JpxWXZsmWZ+j6yWufOz4rB8LjA26m+h0uiaQH3\nu2rKQ8TTfk+tGlCUHKhBgzbs3l0Bl8uOw/E7b745mg0bNmQ4TsWKZdC0MCAeEAyG+ZQtWwaA+Ph4\n3O7UO+QFIpJIYuJ4IBj4GputFh9//DaRkWcZPPjlW2LfuBGL2516+Lsg0dExAEwdMoitZgcFcLGO\nqrThIGL+mcqVy6RZZxFh374dwPvARuAdkkYkBB+fmYwZM4SWLVtm+LvISmFhYdSs2Ywff1yCy/Us\nsAlITL4aToECRe5j7ZRHTtbkIw+OHNgkRcmQzz+fJGBONTlOxGAYIqNHj85wrMTERGnTpptYLPnF\nxydUihUrK2fPnhURkZMnTyZPDvxGYKeYzS1Fp7MJXE4u1yk2WwVZs2bNLTFjYmIkOjpafvnlF9G0\nwgLrBfaJplWXUaPGihw9KlKwoAjILpuf5NaKitVaXKpWrSsxMTFp1vny5csCRoFlAs0EGglUkbFj\nx2a4/dlh2bJlYrEUEFgoECKwWKCdQGmBuuLtnUu2b99+v6upPEQ87ffUHAFFyWHee28UUAjYATQH\nXBgM2ylQoF+GYxmNRpYs+YGjR48SFxdH6dKlMZmSdiUsVqwY4eG/MmjQ20RGXqF58waYTGX53//q\nExv7NBbLRsqXz02dOnUASExM5Omn+7JkySIAWrVqz6RJIxg5chAOh4M+fZ7mnW4doX59OHcOnnqK\nisuWsTYiAp1OR2hoaLp2Bfz4488Ab+AwSXME9gH1iImJyXD7s8OkSd8SFzcG6AQEAR2AVpjNPgQG\nRrBhw251fLByT6lEQFFyGIcjjqQ97nuRtN/9AQoUcPH0009nKp5Op6NkyZK3vValShW2bVuV8mcR\n4cknq7Jlyw6Cg1vw/PPPYzQm/W/mww8/ZvnySJzOK4COFSs6Ubr0aU6d2p/08KlTULcunD0LtWrB\n8uUYfHwol8EjkffuPUzSCevfAsNIei1QmBIlSmSs4dnEaDSQ9KoFoB7wBqVKzefdd4fQsWPHh+b0\nRyUHyZqBiQdHDmySomRIjx79xGJpKbBU4Hnx9vaRQ4cO3e9qSe3arQR+SjUp7mepWbNZ0sVTp0SK\nFk26ULOmyI0bacb76quvpXjxClKs2OMyceLklPX3Q4YMFaOxukAxgRECncVmy5eu1wr3woYNG0TT\ncgtMEvhKNC2PrF69+n5XS3mIedrvqcmCipLDzJgxmd69S1G06LtUrnyEdetWULp06ftdLUqUKIzR\nuDHlz0bjRoKDC8OZM1CvHpw+DTVqwG+/ga/vXWPNnv0Dr7/+KSdPTubUqSkMHTqJmTNnATBixLtU\nrGjB2zseL69J5MmzlQMHtmGz2e4a0+12Z2gL5LQ4nU5GjBhN9epN6NChJydOnACgdu3arFz5E506\n7aJjx20sX76Ahg0bZlm5ipJRakMhRXnExcfH88Yb77F69UYKFcrP5MljKVWqVMr1vXv3smvXLooU\nKUKTJk0yfQRuZGQkVavW4fr1/IAeX98I9oTNJU+XLnDiBFSrBqtWQTpeBTRq1IE1a7oA3ZI/+ZGA\ngLc4cmQbgYGBuFwuDh48iNvtply5cimvJ24nMTGRvn1fZO7cWeh0el56aTDjxo32+KjfPn0GMn/+\nn9jtb6DX78XP70v+/HMvefLk8Siuovybx/1eFoxKPFByYJMUJVu1bdtdzOa2AptEp5sguXLll0uX\nLomIyNdfTxdNyyea1lus1rLSrVvv/2yBmxExMTGydOlSCQsLk5jDh0WCg5NeB1StKhIVla4YDodD\nQkIq/2t/g69EpysrFSo8meH6DR36gVgsjZPPT7gomlY10/suiIjs379fRowYKXq9Sf45vVHEau0s\nM2fOzHRcRbkTT/u9HNdrqkRAUdIvISFBDAaTgD2lw7LZOsjs2bMlISFBTCarwF/J1+xitYbIxo0b\nPS/47FmRkJCkAitXFrl2Ld2P9u07SMzm6gL+Au8IvCsQKLBNzObcKcsb06tixboCq1MlFd9LixZd\nM9oiEREZNuw9AavAy5J04NLlVIlAe/n2228zFVdR7sbTfk+tGlCUR5her08eAo8laXY9QAwmk4kb\nN26g05mAv1cMWDAYynDx4kXPCr1wARo0gKNHoWLFpNcB/v7pfvz772fhcBwHrgDfAb8BzwOhuN3x\nWCyWuz4PcO7cOXbs2EFgYCD58+fhjz/24nYnvac3Gn+ncOG8GW7Wxo0b+eijL4HpJL2y8AKaAm9j\nMOzBat1D69bTMxxXUbKbmiyoKI8wo9HIwIEvo2nNgW8wmV4gKOgczZs3JygoiLx586LTfQG4gU04\nnZupWrVqpsqKi4vj2KZNuOrWhSNHoEIFWL0aAgLSfjjZ1q1bcTicwE2gDDAWKA4cQ9Oa0LNnLwLS\niLdx40ZCQyvx3HMzadnyBez2GHx9J6Bp3bBa2xEY+CMffPB2htu3YsUqRHyAv/cA+AR4jHz5PqBn\nz+vs2bM5zbopyv2gRgQU5RGTmJjI6NGfsnr1FooVK8DYsR9QpkwIK1eup1ix/Lz77saUGfarVy+l\nZcsuHDv2Gr6+Qcyd+x3FihXLcJlbtmzh2WbtWHbzOgZJ5FrBQgSsXg2BgRmK8+qrHwBtkv96DfgD\nL691dO3aiQYNXuC555676/NLly6lW7f+xMXVB74A/Nm1qzYTJgzHaDRiMBho02Ym/hkYofhbYKA/\nBoMfLtdbwAzgGjrdJqZNm0br1q0zHE9R7hW1akBRHjFPP92XsLDT2O0vYTRuJShoIX/+uQe/u8zW\ndzgceHl5ZWomvd1up3zeYiy9aaYsEeynJA04R+32rZk//zu8vLzSHatkyWocPfo5cAJYDZynffs8\n/PjjnDSfnTx5Cm+99Sl2+yvAX8CvwE5MplF89FEhXn/99Qy3LbXo6GgqVKhJRIQbl+sS4GTQoOeY\nPPkLj+IqSlo87fdUIqAoj5D4+HhsNj9crqtA0q9+m60ZM2f2+89Rvlnh4MGDdKjdmMVRlyiHm4OU\noT7ruEwfTKazDB7cmk8+GXXLMy6Xi1WrVnH16lWefPJJihcvnnLtnXeGM3FiOHb7TOAGmtaR2bPH\n0b59+zTrEhBQiKio5cDjyZ/0AEqhaTNYvvw76tat63F7Y2JimDNnDtHR0TRt2pQKFSp4HFNR0uJp\nv6deDSjKIy47k+eezTuyMEpPOdwcohgNWMtlBNiDwzGaX36Zdksi4HQ6adq0Azt2RAAlcbtfISxs\nLo0aNQJg5MhhREfH8P33T+LlZeKDD95KVxIA4HDEA6lfRfig13/IiBGfZEkSAODj48MLL7yQJbEU\n5V5RIwKK8ojp3r0PYWERxMW9hNG4jaCg+Wm+GrgdEWHq1P+xePFv5Mnjz6hR7xAcHJxyPfHiRQ7m\nz09F4DCFqI+dSwQDx/l/9u47vsbrD+D4567k5mYRkYTYEiFW7BFUkVhFW9Vp/FClKKpV1WHWqFbR\n6jCqRltUbC1qj5qJ1YgRM0RsSWTc3Jt7v78/bprSmE3UOu/Xy4v7POec55zzkjzfe57znAPvoNHk\no2HDlaxbtzQ7z5w5c+jWbRKpqRtxfE9ZjZ/fmyQkHM11u19/vTc//xxLevpI4AgmUx+2bl2rvrUr\njzz1aOAfVCCgPOkSExNJTk7G39//prv1Wa1WRo4cmz1ZcOzYYRQuXPierzN48HDGjVtEWtogtNoj\neHh8Q3T0Lvz9/eHKFWjcGPbu5TD+NGQX57gG1MNgKI5eXwG9/je2bFlNpUqVSE9PZ9Kkr1m4cDG7\ndlXBZvsq6yrXMBh8sVjSctcpOOY5DBw4mCVLVuLllZ+JEz8hNDQ01+UqyoOmAoF/UIGA8iQbNGgI\nX3zxBTqdG4UL+7B+/XKKFi2a59eZOnU6b7zxFo4tfh27+jk7d2XMmIr069QJmjSB3btJ8/cn5Go6\nFwyVsVpjeeGFFtSrV53MzEyeeeYZihYtitVqpXbtxsTEFMBsLgr8DOwASqHTfUz16tvZvn3NXdft\n8OHD7Nmzh6JFi6obvfJEUHMEFEUBYPny5Xz11TwsluOANydPjuCll7qydevveXqdP/74g379BgMm\n4O8RBxEdTmlpEBYGu3dD6dKYNmxgi5MT+/btw9fXl0qVKuUob+PGjRw5korZvAHH0iYv+C9+AAAg\nAElEQVQlgLLo9QbKlavMwoUL7rpu3bu/yZQpPwM69Ho9HTu+yPffT8pVexXlcffAFhQym83UqlWL\nkJAQgoODGTRoEABXrlwhLCyMMmXKEB4eTmJiYnae0aNHExgYSNmyZfn997z95aYoj7rdu3eTltYW\nKAhosNm6s3//7rvKazabmTVrFuPHj2ffvn23Tbt582YslleA7jhW0FsOfIZr5hy6zJ0LUVFQqhSs\nXw9FiuDj40NYWNhNgwBwvF6o1Rbk719HfTEYTBw9epD9+7fd9WOLZcuWMWVKBLAK2EFmZllmzfqF\n7du331X+6509e5Zt27Zx4cKFe86rKI+aBxYIGI1G1q9fz969e9m/fz/r169ny5YtjBkzhrCwMI4c\nOULjxo0ZM2YMADExMcybN4+YmBhWrlxJz54983TLUEV51JUsWRKTaSNgyTqymqJFS+ZIl5KSwrp1\n69i6dSs2my0rKG9Ez56zef/949StG87ixYtveR1fX1+cnfcAQ4FXgJG4M5YVosP4559QsqQjCLjL\nRxKhoaHo9dFoNF8D0Tg59aJKlWoUK1bsrvKLCBcuXGDmzHnAR0BtIBAYh82m48yZM3dVzl+mTPme\ngICKNGvWlxIlyhERsfCe8ivKIydXOxXkkdTUVKlevbpER0dLUFCQnDt3TkREEhISJCgoSERERo0a\nJWPGjMnO07RpU9m2bVuOsh6SJinKfy4zM1NatHhBXF0DxcOjkXh6+klUVNQNaU6cOCF+fqXEw6Ou\nuLkFS+3ajaVv377i5PS0gD1rg5zNUrBgiVteJyMjQ2rUaCg6XVWB9uKOl/xBORGQOJ1ePu3ZW44e\nPXpPdY+JiZE6dcKlcOEgef75DnLlLjchSk5Olnr1moqzcz7RaIwCva7bPOgX0WjyS2xs7F3XIy4u\nTlxcCgjEZpWxW1xc8ktSUtI9tUdR/ku5ve/dMffEiRPv+ofyXtlsNqlcubK4ubnJgAEDREQkX758\n2eftdnv25969e8uPP/6Yfa5r164SERGRo0wVCChPMrvdLtu3b5eVK1fKpUuXcpxv3LiN6HQjs25y\nmaLXtxCt1k2g93U30Cvi7Ox+2+tYLBYJC2shntq6soUqIiAncZcSVBKt9l1xdy8oBw8evF/NzNal\nSy9xdm4vYM3aJdFdoJPAAAFXGTJk6D2Vt2HDBvH0DL2uL0Tc3YMkOjr6PrVAUXIvt/e9O04WPH/+\nPDVq1KBq1ap06dKFpk2b/qtlRm9Gq9Wyd+9ekpKSaNq0KevXr7/hvEajue218qoeivK40Gg01KpV\n65bnjx49js02JOuTjszMFjgm/M0HOgBlMRjeo2HDsNtex2AwMHfatxwLCqaGOZU4DDyNGydZDXYf\nUlLyMWrUeGbNmpwj76VLlzhx4gTFixfHx8fn3zYVgK1bI8nIGIdj3nMZYBjly/9IixZNePnlTVSt\nWvWeygsICMBiOQTE4NjUaAc228W7fkyhKI+iOwYCI0eOZMSIEfz+++/MmDGD3r178+KLL9K1a9cb\nFg/JDU9PT1q2bElUVBS+vr6cO3cOPz8/EhISsn9R+Pv7c/r06ew8Z86ccbyvfBNDhw7N/nfDhg1p\n2LBhntRTUR5mFy9eZPjwMcTFnSM8vD49e3bPESzXqFGFs2e/x2r9EkgDfgBeB9oDzwMXCAmpx5w5\nN5+pLyKOMlNT8erQAS9zKmZvbzq7FuTEqVGAT1Y6f5KSYrLzpaenc/DgQbZt286AAR9jMBTHaj3J\nlCmTaN/+1X/d5tKlixMbuw6bLRQQnJ330KpVS0aPHv6vyvP392fy5Il0714Pg6EINttZ5syZibu7\n+7+uo6LktQ0bNrBhw4a8K/Buhw727Nkjffr0kTJlykiPHj0kJCRE3n333X89FHHx4kW5evWqiIik\npaVJ/fr1Zc2aNTJgwIDsuQCjR4+WgQMHiojIgQMHpHLlypKRkSHHjx+XUqVKid1uz1HuPTRJUR4b\nycnJUqRIGTEY3hKYKSZTTXnrrZw/n5cuXZLKleuKi0shcXbOJ05OBQQmC6QJ/Cz58xe+6fPwiIgF\nkj+/v+h0TtK4TphkhIY6xs0LFxaJjZXx478Sk6mSQKTAFjGZSskvv8wXEZFDhw6Jr29JcXMrnzV0\n/3LWfIRocXHxkoSEhLtuZ0TEAnn++Q7SpUtPOXr0qJw6dUp8fUuKh0dDcXevLuXL15Tk5OR/35HX\n9dPu3buzf0cpysMst/e9O+aeMGGCVK1aVcLCwmTevHlisVhExPF8v1SpUv/6wvv375cqVapI5cqV\npWLFijJ27FgREbl8+bI0btxYAgMDJSws7IYfxJEjR0rp0qUlKChIVq5cefMGqUBAecz9/vvvUq9e\nC6lZM0xmzJglIiJz5swRN7fm1z3bviR6vbNkZmbmyG+z2eTkyZNy7tw5iY6OlsDAKqLTGaREiQo5\nJheKiOzcuVOMxoIC28SFi7JWU9RxkUKFRA4fFhHH3IRRo8aKv39ZKVasvEyePDU7f6VKoaLRfJVV\nrySBygILBEQ8PWvKH3/8cVft/u67KWIylRSYJhrNEPHw8JWTJ09KUlKS/Pbbb7JmzRoxm83/pksV\n5ZGW2/veHVcWHDJkCF26dKF48eI5zsXExBAcHJx3wxN5QK0sqDzONm/eTNOmL5CePh5ww2Tqz6RJ\nH2Mw6OjRYxGpqX8N6aei03mRnp5yT9v8/nWNP//8kzJlyqDX62nevBVmc1uMfMtSWhPGGhIAv5gY\nNOXK3bE8V1cv0tIO41jfAOB9HDsfPo+LS32OHz+An5/fHcspUqQc8fEzAMccCJ2uD4MHF2Tw4I/v\nqX2K8ri57ysLDhs27JbnHrYgQFEed999N4v09A8Bx3P1tDQnJk4cze+/z8fJaSDp6Z9jt1fHaPyM\nOnUasXr1aqpVq4avr+9dlT9s2CjGjp2CSFO02i+xWM5jtb6FkY0syQoCzuFFGzctO+8iCAAoXbos\nf/75C9ALuAYsBK7h7DyOyZMn3VUQAJCZacWxmqGD3e6K1Zp5V3kVRbm1B7agkKIo906v1wEZ1x0x\no9Pp8PHxYceODYSH76B8+Q/x949nx44jvPLKBAIDK7Ft27Y7ln3hwgVGjx5LWtp20tMnk5q6A6tV\ngzPPsYhDhLOGCxhpZrDz7vff3HWdZ8yYBLwHhOBY6Kcezs6VGTt2GB06vHbX5bzxRidMpi7AWmAG\nLi7TeOmldnedX1GUW8iDxxMPlcewSYoidrtdrl69KpGRkWIyeQt8ITBVTKbCsmjRohvSduzYWSBI\nID3rufxCKVas3B2vceDAAXFzC7zhHXonysmvVBQBuYCbhOhdZPHixfdcd1dXL4EfshbqSRVX17Ky\nevXqW+aJjo6Wbt16S4cOb8j69etFxDG3YcyYcVKpUn0JDW1+13MLFOVxl9v73mN311SBgPK42bx5\ns+TPX0icnNwlf/5CMnXqVHnppc7Sps1rsmLFihvSrlu3TgyGfAJ9r7uhXxO93njH66Snp0uBAkUF\nvhewihPzZRlaEZBLGoNUd/aS0aM//1dtWLp0qZhM3uLu/qy4ugbIa6+9ftO3fkQci/q4uHiJRjNc\nYKK4uPjKsmXL/tV1FeVJkNv7ntqGWFEeYklJSRQrFkRy8g9Ac2Al7u6dOH36CJ6enjnSjx49mo8+\n2ovdvgvYAhQGxuDjM40zZw7mmDh44cIFVq1ahU6no2XLlvTv/x7Tpy/DwDkicKE1aSTq9MR89SXe\njRtTpkyZf92WEydOEBkZSeHChalbt+5NFwSbMWM2r7/eG5vtHWBw1tFFVKnyJbt3r8+RXlGU3N/3\n1BwBRXkAUlJSeP759phM+fHxKcncufNumm7u3LlYLAVwBAEAzdBo/Dly5MhN0xcpUgQXlwSgG46V\n9nyA0SQlFaJDhzduSHv06FHKlq3Cm28uoXv3nwkOrk5aWgYG3uUXWtGaNK7gwas+Jan75pu5CgLA\nsSlSu3btCA0NvWkQcP78eXr27IfN1gzId90ZTywWS470iqLkjTu+NaAoSt5r3fplNm0yYbMdIT39\nGF26PEfx4sWoU6dOdpq3336f776bg9l8BTiL49t9AhbLKQoVKnTTcl955RV++OEXtm37CbO5DhAF\nLCEjozoREd5kZHyHs7MzAH37fkhSUl/s9vcAyMjox7Urx5ivG0wbWypX8eQZ5/JUa1v3vvaF3W5n\n69atHDhwAIOhJOnpb+LY1bAo4InR2JMePfre1zooypNMBQKK8h+bN++XrH01juB4t74gGRmdWLNm\nTXYg8NVXXzFhwrfASWAaUAOojIvLPj788H2KFCly07L1ej2rVy9m6NChfP75MszmfThuqMnAjUOH\n8fHnsNurZ38WawiD9i8g1JZKIhpaOmUS8GIFPv/8kzzugb9duHCBgIAqXLuWllU/O45XBKcCI9Bo\njjJ06Pv06tXjvtVBUZ506tGAovzH+vR5H/DFEQgACDrdQby8vADYunUrffoMAMoC+YEBwDKcnfcw\na9ZEPvxwwG3L1+l0DBgwAC+vVPT6L4EFmExtaN++c/ZoAEBYWH1cXD4HUtBxjrm6dwg9d4YUvYFm\nWh277Jnky+eBk5NTXnfBdXVow7VrdYCLWX8aoNE0wsNjECbTUSIipjFw4AC1wZii3EdqsqCi/Mcc\nK+1NwHGDfw04SIECMZw8eQA3NzeqV69PVFQ6cBqYATQD5uLp+Q7nzh3HaDTe1XXOnTvHBx8M5+TJ\nszRpEsrAgf3R6XTZ5y0WCx07dmfR/J+YKTZeFjvpTk40pQ6bLSuANEymZnzxxRt0794tj3vBwc2t\nGKmpk/l7DsRCPDzeYePGRRQvXpz8+fPfl+sqyuNETRZUlEdM06YtcXZeCcwGUjEY/mDJkp9xc3MD\nIDHxGnAGx86AbwBOQDeWLp1710EAgJ+fH9Onf8O6dYv54IMBNwQBAE5OTsz9aRrpL7blZbGDuztd\nCgWw2TIScAEKkJb2Jr//viVH2Xa7nQkTJvH00214+eUuHD9+/F/1ReHC3sBiHI8FBFhMsWI+hISE\nqCBAUf4jKhBQlP/YrFnf8swzetzc2lO48Abmz59NaGho9vkWLcLR6QoAvYH6gBsajZ3Zs+fn6WiX\n1WzmTFgY2rlzsbu6wsqVXAosg0azIzuNk9MOSpTIOTHx/fcH8+GHM9mwoSPz55eiWrV6JCQk3HMd\nVqyYj8EQgeMNh9I4O6/g11/n56JViqLcK/VoQFEeMmazmZdf7sKSJb9kHXkF+ApX1zC++64v7du3\nz/01UlNZU7w0z1w+Two6nndx55MNq8iXLx+1az+N1VoTuIa3dwJRUZuz5y/8xfF4Yy9QDACjsROf\nf16TXr163fKaNpsNIMfIRHJyMnPmzEGj0dC+fXtMJtPNsiuKcgv3fdMhRVH+W0ajkcWLf8bHZzsX\nLy4HHJt7paa2Y+fOPbkPBOx2TjRuzDOXz5OKiRasYHP6Wc787y1iYnZw6NAe1qxZg5OTE82aNct+\nZGGz2Vi7di1XrlzJ+qVz/QQ+7S1/EaWnp9Os2XNs2bIOjUZL9+49+eqrz9FqHQOSHh4edO/ePXdt\nUhTlX1OBgKI8pEqVKs2lS2sQCQasmExrKFv22dwVardDt26U27GDVAy04Dc20wA4yblz8QD4+Pjw\n6quv3pAtMzOTsLBniYw8g0YTgNVqx2h8HrP5Y7TaGJydV/LcczlfM7RarZQpU4UzZwoClwAr33/f\nnICASbz9dp/ctUVRlDyh5ggoykNq1qyvKVBgHB4e9XF1rUCtWk5065aL2ft2O3TvDtOnY3Ny4gVn\nHzYRANgwGD6nTp2cCwfFxcUxaNBHtGz5LNu3nyElJZJr1yLIzJyNs3M8det+R5s2B9i1axP+/v45\n8kdERHD2bArwEeABFCAj4x1WrNj079uhKEqeUiMCivKQKlOmDMeO/UlkZCSurq7UqFEjezj9ntnt\n0LMnTJsGRiO65csJ3RbJmmEBAFStWpdZs365IcvJkycJCalDSsor2Gx1gHHAahyv+tUjIyOVP/74\n7baXPX/+PI61EKKApllHt1OsmN+/a4eiKHlOTRZUlMedCPTqBd9+C87OsGwZhIUBjiF/s9mcPQ/g\nem+//R5ffqnBbv8068hiYDSwHb3+Q2rWjOKPP1bd9tKRkZHUr98cs1kPhALXMBh2cupUzC2XSVYU\n5d6odQQU5TGQkpLCwYMHSU5OztuCRaBPn7+DgCVLsoMAcCxJ/FcQYLfbWbhwIePHj2fz5s1cu5aG\n3e57XWF+wBH0ejfKlVtPRMSMO16+evXqTJkyAZPJjEaziFKlzhAdvUMFAYryEFEjAorygK1cuZIX\nXmiPVutFZuYlZs6cSrt2bXNfsAj06wdffglOTo4goFmzmybNyMigbNnqnDyZCTyFs/NyOnVqw48/\nLiYtbQZQAJPpTd555xneeafPTbdAvn1VBJvNhl6vnkYqSl7L7X1PBQKKksdE5K7Xxk9OTqZw4VKk\npi7BMXS+F5OpCceORePn9/dz9L/+T9/1mvsi0L8/TJgABgMsWgQtW94y+bPPvsiSJTtw7H/gDMSh\n15dl+vQpDB8+HrPZTOfOLzN06If/fp6Coij3hXo0oCgPicOHD1O2bHX0egP+/mXYtm3bHfOcOnUK\nrdYHRxAAEILBUIbY2FjA8e7+W2+9i9HojouLB/36DcRut9++UBESu3WDCROw6/WwYMFtgwCAdes2\nAOVxBAHg2LHQmbCwJsTGRnH69AGGD/84z4MAm83GuXPnsFqteVquoih3TwUCipIHrFYrTz/dkiNH\numC3p3L27Gc0bfosFy9evG2+IkWKYLUmAAeyjhzHYjlCiRIlAPjsswlMn74Ni+UYGRmxTJ26kfHj\nv7p1gSIcevZZ8n3/PVY0vKzNR981OfcKAMfjgK+//ppnn32B1FQrsAP4DUgDPsXd3Q0fH59764h7\nsGvXLnx9S1CyZEXy5fNh8eIl9+1aiqLchjxmHsMmKY+A2NhYcXUtLo4xeccfT8+Gsnr1ahERmTZt\nunh4+Iheb5TmzV+Q5OTk7LyzZ/8kLi4FxNOzvri4FJCvv56cfa5u3eYCS64rd4GUKBEiNpstZyXs\ndskcMEAExIJenmWhwFUxmYrJzp07b0iakpIiAQGVRadrJPCJQCEBdwFfAb3o9V4SFRV1y/bu2bNH\nSpasKDqdQUqXriz79++/p/7KyMgQLy9/gYisdu0Sk8lb4uLi7qkcRVFyf99TIwKKkge8vLywWq8A\nf228k4rVepyCBQuyYcMG+vQZTHLyajIzL7BunQudO/+9Jn/79q9y5MheFi0axsGDUfTs+Ub2ucKF\nC6LV7r/uSvs5ffoiI0aMubECIvDRR+g++4xM4GXmspjngHzo9VWIi4vLTmo2m6lcuRZHj5qx2X4H\nPgT2AhlAI7y9C5GQcJiqVavetK3Xrl2jUaOWnDjxHjZbEseO9ePpp1uSlpZ21/0VHx9PRoYW+GtS\nZHUMhqr8+eefd12Goih5QwUCipIHvLy8+PDDDzCZ6uLs3AtX19q88EJzKlWqxNq160hP7wxUAtzJ\nyBjJmjVrbshfpEgRnn76aYoXL37D8TFjBmMwfAG8iGPzoWnYbN8yffqcGyswZAiMGoXodPTw8GYh\nGVkn/iQzcyuVKlXKTvrjjz9y+rQBCAT+2gDIGzACzSlSpCTe3t63bGtMTAw2mx/QHsd2xf/Das3H\n4cOH77q/fHx8sNmSgINZRy5jtUZTrFixuy5DUZS8oQIBRckjgwe/z2+/zWDs2CDmzRvDjBnfotFo\nKFjQG2fnaOCvWb3R5MtX4K7KLF26NG+80RGNJhFoBOwGjDg7O/+daNgwGDECtFo0P/1E742r8fH5\nAKPRB6OxHlOnfklgYGB28k2bNmGx1AZ2ATOA40AvwA+dri8jRgy4bZ28vb2xWE4DiVlHrpCREU+B\nAnfXJgBXV1e+/fYrXFyewsPjWUymEHr37kqFChXuugxFUfJIHj2ieGg8hk1SHnEpKSkSFFRV9Pqn\nBf4n4CZeXoUlNjZWNm3aJAEBVcTTs5C0bPmirF69WlasWCEXLlzIzn/ixAnx8PAVjaanQEvR693l\niy/GO04OH+6YPKDVivz8c3aezMxMiY+PF7PZfENdhg4dKUZjUYGCAvMF6ggUEPAWrdZdpkyZcldt\n6tmzv7i6lhUnp97i6lpG3n77/X/VN4cPH5aIiIjbzkdQFOX2cnvfU+sIKMp/YM6cOfzvf+9hsfQG\nWqPVrqRcuTmcPHmC1NQpQHW02qHAEtzdq2C372f16qXUqlULgLVr19KiRVus1lfQaJxxcfmJI51f\npvCkSaDVYp8xA22HDje9dnx8PG3bdmLPnu1YLBpgFpAEvAskYjDkJyCgFBMnfkLYdasOAiQlJfHx\nx59w8OBx6tQJ4aOPBuLk5ISIsHLlSg4dOkT58uUJDw+/b32nKMrt5fq+l/tY5OHyGDZJeQwMGzZM\nNJoPrpv9f16cnFzF1fW1645ZBAwCGQIRUrx4eRERSU9Pl8aNnxGN5pPstANpLQJiA+mkNQlopGDB\n4rJhw4Ybrmu32yU4uIZotR8KJAosF/AWiBMQcXNrLfPnz79pnTMyMiQ4uIY4OXUWmCcuLi2lefO2\nYrfb73t/KYpy93J731NzBBTlPxAUFITJ9DuOd/RBo1mMr68/Gs0p/p47cAZwAgxAOGfPHufChQsE\nBVVlw4Y9iJQE4F0+YwxLsQOd+YaZ9h+Bgly8OJpnnmnHuXPnsq+bmJjIoUPR2O0jAE+gJVADWAb8\njEazndDQUG5mx44dnD5twWL5HniR9PQFrFu3noSEhJumVxTl0aQCAUX5D7z44ou0bl0Jk6kMHh41\n8PL6hKVL5xEQILi4tAIGA3WAQYAGrXYyQUGVKF++NnFx9bHZRgEj6E9/PuM9AF7nFWbxJvAcUA1w\nRaerzJ49e7Kve/z4cez2TOCv1wetwCFcXT+mXLmJrF27/KYbAEnW3gCOoOQvOjQaXdZxRVEeF2oH\nEEX5D2g0Gn76aRqHDh0iMTGRChUq4O7uzrZta5g+fTpnz54jLq4F8+aNQa//Bm9vTypVqsOBA5eA\nVkBL+rGQcYwH4A2NgR/ks6zSLThm/ruQmRmLr+/fOwYmJyfj7FyCjIwGON7Z34FOl0xU1FaCgoJy\n1HP69Bn07TuA9PQk6tULI3/+VMzmd7Bam2E0zqB69aoUKVLk/naWoij/KTVZUFHug8jISL744jus\n1kzefLMjjRo1uqt8SUlJJCUl4e/vT61a4URF+QBW3iKUL+kPwMKmLTlYvwGjRn2H2dwGu30NWq0V\no1FLu3ZP8cMP32RvTnTlyhVKlgwmObkHjm/3sRQqtIVTpw5iMBhuuPYff/xBePiLpKWtBAJwcupL\nvXoJ+Pn5cejQMWrXrsLYscNxdXW9IV98fDzr16/H1dWV5s2bYzQac9l7iqLcC7X74D+oQEB50Hbt\n2kXDhi1IS/sAMGIyDSciYjrNmze/p3I6derBzz9r6J65mUlZexF8kN+PwWdPYDQaWbt2LVFRUWRk\nZODl5UVAQADh4eE5dijctWsX7dp1Jj7+KGXKVGbRotmUKVMmx/VGjhzJkCHJ2GyfZh05j6treVJS\nLt2yjlFRUTz9dAtEngISKF7cwo4d63IEC4qi3D8qEPgHFQgoD9orr3Rl7tyKQL+sI79Qu/Z0tm1b\neU/lXL58ma/KhTD04hkARvgWpe+RaDw8PPK2wlmmTJnC228vJi3tV0ADrKZIkT6cPn3wlnmqVGnA\n3r2vAx0BwWhsx/DhtRkw4N37UkdFUXJS2xArykPGYsnEsfQuQCawnOjoGDp16s7Zs2dvmU9EyMjI\nyP5cICIiOwg42b8/g84cv29BAEDHjh0JCEjE1bUxLi5vYDK9yrRpE26bx/EGQc2sTxrM5lqcOnXr\nNiqK8vBRgYCi5LE33+yAyTQU+AWoCBwmJWUCs2a5U7VqKImJiTnyLFu2DE9PX0wmN4KCqnJh5Ejo\n0cNxcsIESowbh15/f+f2Go1Gdu5cz9Sp3fj88xAiIzfRtGnTG9IcP36cL774gq+++orz58/ToEEo\nzs6f4ngb4Swm03QaNap3X+upKEoey9UqBA+hx7BJyiNo+fLlEhBQUUAvkJS9EJBO97TMnTv3hrRH\njx4Vk8lbYJuATbry4t97GX/xxQNqQU579uwRN7eC4uTUQ4zGDlKgQBE5cOCANGzYUnQ6J9HrjTJk\nyCcPupqK8sTJ7X1PvT6oKPdBy5Yt+e231Rw9eojrB95sNnI8y9u1axc6XUOgNp2YwRTmA3Bp4EC8\n3377ltc4e/YsY8Z8wfnzV3juuaa8/PJLd6zXiRMniI+Pp1y5cve0SRBA//6DSUkZDjhGKqzW9/ny\ny8msX78cs9mMwWBAp9PdvhBFUR466tGAouSC3W5n6dKlTJ06lejo6BvOBQaWQKsthmPBn+XAR+j1\nUTmG2/38/LDbo+nA90ynC1qEgego/tU0VqxYcdPrXrx4kZCQOnz7rfDLL7Xp2nUIY8d+cdu6Dh78\nCcHBNXnmmQGUKFGO9evX31NbL126Cvy99oDNVpbz568AjscKKghQlEdU3gxMPDwewyYpDymbzSbh\n4c+Jm1s1MZk6i8nkI/Pm/ZJ9PjExUZydCwoUEigm4CYvvdQ+RznTpk2Xbi5eYst6HPA+rgIzBf4Q\nV1cvycjIyJFn0qRJYjRev0/BQfH09LtlPRcuXChGo7/A+az0a8XDw0dsNttdt/fjj0eIyVQ/a5+C\nQ2IylZOZM2ffdX5FUe6P3N731KMBRfmXfv31V7ZuPUVKynYci/XspmvXcNq1ewGNRsPGjRsxGALI\nyBiCY4+BQBYsqI7VOj17MZ+ZM2ezpef7TLMkogU+wsAYPsXxOh7Y7QYuXLiQYzU/i8WC3X79GwTu\nWK2WHHVMTU0lPPw5IiMjsVjqAD5ZZxphNqeTlJRE/vz5c+TbsmULO3fupGjRorRRYooAACAASURB\nVLRt2xatVsvgwe9z5cpVZsyogk6nZ+DA/nTo8FruOlFRlAdOPRpQlH/p/Pnz2O2V+Hs9fgOpqVfZ\ntWsX586dY9++fYAv0BTH44GygAar1ZpdRuyIz5hmuYQOO4MZxki+AdZknV2NwSA3LBn8l9atW+Pk\nFAF8D2zBZOpA+/btc6T76KMR7N5dAItlNbAHOJ11Zilubh7ky5cvR54JEybRtOmrDBp0is6dP6dV\nq5ew2+3o9XomTRpHSsolkpLO8cEH7+VYvEhRlEdQHo1MPDQewyYpD6no6GhxcSkoECnwjYCXaLXh\n4uTkKzqdh7i7VxYwCUwViBYnp87SoEHzvwuYN0+saERAhjI4a8h+nICHQKBotW45thW+XmRkpNSr\n11zKlq0lAwcOFqvVmn3OYrFIfHy81KnTTGBpVtkTBdxFpysunp5+sm3bthxlms1mMRhMAiey8mSI\nm1uwrF+/Pi+7TlGUPJTb+556NKAoOCb9bdq0icTERGrXro2fn98d85QvX56ZM7+hS5dwUlJSgRjs\n9lJYLBeA8ly7FgHsRqPpjo+PNw0b1mfy5DmOzBER8Oqr6BFGaowMlfzAp8AooAWwhHHjRvHUU08R\nHR3NsmXLMJlMtG/fPnu2f7Vq1di8+bcc9Vq9ejXPP/8qNpuWzEwLBkNhrNZngLdwctpJmzYafvjh\nu5suA3zt2jU0GgNQPOuIE1ptIJcvX77XLlUU5RGhlhhWnniZmZk0b96W7duPodWWQGQXa9Yso2bN\nmnfMe+rUKZ57rj179pwAzlx3pj4wAmiIu3sFNm36kZCQEMephQuRl15Ck5nJxrqhtP7zMMnXAFoC\nHoAXev081qz5DpvNRqtWL2GxdESnO0/+/NvYv387BQsWvGl9rly5QrFiQaSmRgBPAUvRaDrg6uqY\nY1C8uBt//PE7np6e2XlWrlzJmjXrKVTIh27dulG9+lMcP94Wm60fsAVX1w4cPLibokWL3mPPKory\nX8j1fS8PRiX+lbi4OGnYsKEEBwdL+fLlZeLEiSIicvnyZWnSpIkEBgZKWFiYXL16NTvPqFGjJCAg\nQIKCgmTVqlU3LfcBNkl5RM2YMUNcXesLWLKGw+dJQECVO+Yzm83i719GtNohAj7XDcFvFvDKmqEf\nKS4u+eXKlSuOTIsWiV2vFwEZQ03R8J5oNK4C/gKjBRIEpohO5y4uLvkFDAKNBdIFRAyG7jJ48NBb\n1mnbtm3i4VHturcJRNzdK8qMGTNkx44dYrFYbkg/ceIkMZlKCHwizs4vSmBgiBw6dEiqVm0ger1R\nChcOVI8FFOUhl9v73gO7ayYkJMiePXtEROTatWtSpkwZiYmJkQEDBsinn34qIiJjxoyRgQMHiojI\ngQMHpHLlymKxWOTEiRNSunTpm776pAIB5V4NHz5ctNpB1908z4nJ5HVDmgsXLsi0adPk+++/lwsX\nLojFYpHmzZ8VKJqVZ6tAYQE3cXJyFycnN/HwqCRGo6cMHPi+rFu3Tlb37SuZOp0IyGc0ELBn5Z0k\nGk0BgYoCHqLRmMTZuZRArMBVgVYCfbPSfiZvvtn3lm05ffq0GI1eWa/4icAJMRrzS0JCQo60drtd\nTKb8Aoez0trF1bWpzJ6tXglUlEdJbu97D+ytAT8/v+yhUjc3N8qVK0d8fDxLly6lU6dOAHTq1InF\nixcDsGTJEl555RUMBgMlSpQgICCAnTt3PqjqK4+RmjVr4uIyH0gABJ3uK6pW/fuxwMmTJylbtgp9\n+vzOW2+toly5qrz99nusW3cJxxr7aUAd4AhGo4k9e7YTH3+CoUP/h0Zj4Jtv9jOu0bM0mDgRnc3G\neI2BAbTAscMfQFHKlClN27ZV6Nz5NZ5//jkyMvoAAUA+YDjwOxCJyfQVbdr8vZ1xYmIiXbr0okqV\nhnTs2B2TycSIEYNxcamBh0crXFxq8+mnn9x0zoOIkJGRCvhnHdFgt/uTkpKStx2sKMrDLW/ikdw5\nceKEFCtWTJKTkyVfvnzZx+12e/bn3r17y48//ph9rmvXrhIREZGjrIekScoj4OLFi7Jx40aJjY2V\noUNHicHgIs7O+aVcueoSHx8vIiJpaWlSpEiwwMfX7RcwVLy8Sgv8JtBRoK7AaNFqq8nzz7cXu90u\nNpst69v2DmnOr2LGSQRkPO0EJohW6yOwQyBKTKYK8vXX32XX66OPhojB0PW6EYpZotV6ScGCJWXq\n1O+z02VmZkrlynXFyambwBpxcuolZctWE4vFIgcOHJBFixZJTEzMbfugRYt24uz8qsARgQhxdfWW\n2NjY+9PhiqLcF7m97z3wtwZSUlJo27YtEydOxN3d/YZzGo3mtu8p3+rc0KFDs//dsGFDGjZsmBdV\nVR4j69ato02bl9HpArFYYunXrxdJSZdJSUnB29s7+/9Wr17vEh9vBqpl57XZKqPRzEKni8Rm+wGY\nhUbzLaGhnsyfPxONRkNSUhJWq4WmXGEhz+OMhS8J4G3aAEVxczPi7t4JEaFPn268+eYb2eW//XYf\nZs0K5fLlNthsBdHplrBu3YockxcPHz7M0aMJWCybAS0WSyPOnCnP/v37qVatGsHBwXfsh3nzptOt\nW1/Wrm2Kt3dBJk9eSEBAQB70sKIo98uGDRvYsGFD3hWYN/HIv2OxWCQ8PFzGjx+ffSwoKCj7eebZ\ns2clKChIRERGjx4to0ePzk7XtGlT2b59e44yH3CTlEeA3W4XT09fgbVZ37gviMlUTHbs2JEjbaFC\nZQQGCIQK/CnwmkAhqVYtVAoUKCKurq1Fr28iOp2HNGjQPPvbtNlsluY6V0nHMTFwEq8IeGfNJWgi\n4CEXL168ZR2TkpJk+vTp8vXXX8uxY8dumubQoUNiMhURsGa1I1NcXUtnz71RFOXJkNv73gObIyAi\ndO3aleDgYPr165d9vHXr1sycOROAmTNn8uyzz2Yfnzt3LhaLhRMnThAbG3tXr3cpyj8lJyeTlpYC\nNMo6UhCdrg6xsbE50jre2a8GVAFq41ii9weio30ICAigWLE4NJo0bLYFbNnyNHXqNOLKlSusef99\nFtjSMJLJt7jQm4VAUtY1vQEN3303mZ07d7Jq1SouXbp0w3U9PDzo3LkzPXv2pFSpUjdtR2BgINWq\nVcDF5SVgDkbja5QvX4yKFSvmST8pivKEyJt45N5t3rxZNBqNVK5cWUJCQiQkJERWrFghly9flsaN\nG9/09cGRI0dK6dKlJSgoSFauXHnTch9gk5RHhN1uF2/vogKLsr5JnxKTqdBNv0lv2rRJTCZv0eka\nCFS57rl9hoCLgGv2q32OV/Way6Zhw8SS9Yrgd3QTDXFZz+ALZP39jEBtCQioLK6upcXTs5F4ePjK\nrl277rkt6enp8uGHQyU8/AV5772PJDU1NS+6SFGUR0hu73tqQSHlibRz506aNn0Wm82DjIwEQkKq\ncvbsBXx9ffj227HUqFEjO22XLm8ya9ZcbLaiwD4cs/3TcewjUBL4H/A2IDR3qcgy+1F0GRn8oHOn\nqy0aoQgaTR9EfgZMQC0MhrXodAGYzVsAIzCXgIBPiY3d8992hKIoj7zc3vfUpkPKE6lmzZrExx9l\n27YFNG4cxv79BThz5meiov5Ho0YtOXnyJAC//PILP/20DJvtAI4fl+7ALzg2EWoFdEanGw9Mo4mh\nORHmg+gyMqBzZ84O+QCdPgi93p2QkP106NAWLy8tRYocpG3b5litjXAEAQDhnDlz/D/vB0VRFDUi\noDzR7HY7Tk4u2GyXATcATKb/MX58XcqUKUPTpq2xWBoCS4GrOJYN/gHoBQxCp6uKRnOOULHxm5gx\n2W3QqRPy/fdodDqsVivp6el4eDi2DBYRDh06xNq1axk48EvS0jYDPmi1n1K16ip27Vr/AHpBUZRH\nmRoRUJRc0Gg0GAzOwMXrjl3AxcWFHj0GYLF8AuwEjgL5gVpotZm4u/+GwVASEQM1M6ey3CaY7DZ2\nl69I4KYo9E7OFC8ezL59+7KDAKvVSrNmz1O9elM++OAbDIZUDIYATKYiFC06m/nzf3gAPaAoypNO\nBQLKE02j0TB48MeYTM2ACTg5/Q8fnziee+45Ll++hGMnwE+AqkAhjMaebNq0klWrvqZ8+WBq2juw\nkq64kcaP1KfOoQSOnngHuz2NuLghhIW1JikpCYAvv5zE5s3ppKUd5dq1A6SmdqJRozBiYrZy7Nh+\nSpQo8cD6QVGUJ5cKBJQn3qBBA5g1axSvv36UQYNKsXv3Ftzc3AgPb4LR+BHQDliL0ahlwYLZhIaG\nUqdOHcI9jKxiCO6kMIeX6aIJw67R45g86AS8hEgRYmJiANi37xDp6a2zzmnIzGzHoUNHKV68ODqd\n7gG1XlGUJ90DX1lQUR4Gbdu2pW3btjccmzJlAikp3fjtNz/0eiMDBvSjRYsWjpM7dzJq9x/oyGC+\nthRddSaMxkmYzWbgEo61AhKxWE7h7e0NQJUqwURELCY9vSvghF4/j0qVyv+XzVQURclBBQLKEys6\nOppx474mPT2Drl1fISws7IbzJpMJo9EZo7EkUJlx476mQoVgXixVEsLD0aWkkNaiBWcaNmSkXk+7\ndsOYNGkqkybVxmYLR6dbR7t2z7FixQp+/fVXWrV6htWrN7NxY2m0Wjd8fZ2ZMmXVg2m8oihKFvXW\ngPJEOnDgALVqNSQtrT8inri4fMLPP3+TvZIlwNq1a2nT5i1SU6MAF2APdY0N2GLUo0lMJLVpU/QR\nETi7ud1Q9rp16zhw4AAeHh706zeI9PRmiBhwdl7Mtm3r0Ov1mM1mypUrh5OT013X2WKxsGfPHnQ6\nHSEhIej1Ko5XFCX39z31m0R5In355WTS0vogMghIIT09hp4936FUqVJUqlQJgLNnz6LRhOAIAiAE\nYZk5BY0ZlmoNdNx8APEvzbJl82nQoEF22Y0aNaJEiRK8+GInkpK6ITIMAKu1LO+9N5xff50HQGZm\nJqdOnaJAgQK4/SOY+KfLly8TGhrO2bMWRDIJCCjA5s0r75hPURTlTtRkQeWxcunSJTp27E6NGk3o\n1esdUlJSbpouI8OKiBuQAJQBDpOQEE6tWo1YuXIlANWrV8dmWwPsoxJ7WUN9vICl6HjBvoWktNMk\nJ8+mVat2pKenZ5e9e/duKleuze7dlxEpl31cpCwXL14BYP/+/fj7BxAcHIq3d2G++WbybdvVv/+H\nHD9eh2vX9pOScoCDB0syZMjIXPSUoihKllwtUPwQegybpNwls9kspUtXEoOhj8BKMRpflbp1w8Ru\nt+dI69hDwFegpECr6/YQWCVFipTLTjdnzjyp4ewmF7MSLEMnBmpfl17Eza2UHDp0SE6ePCm9er0t\nBQsGCPQSmCpQSSBW4ISYTLVk9OjPxW63i79/oMDMrDKOisnkJ3v37r2hjna7XRYvXiyfffaZBAZW\nF1h13XXnSZMmz9/3PlUU5eGX2/ueGhFQHhuzZ8/m9OlMrNYJQFPM5pns2fMncXFxOdLWr1+f+fOn\nAyeB63frK8vlyxeyP71cIZgd7ka8gd80OtrSBSvHgLNZKQ6QmXkZu91OhQo1+PprPRcv9sexEqEG\nx6uHtdDrK9Gjx9OEhTVk9uzZnDt3GuiQVUZptNpG7Nu374Y6du7ck9deG8wHH8Rz4kQcOt0swAZY\nMRrnUrOm2mVQUZTcU4GA8lhYt24dvXsPwGKxXXfUjogNrVZLfHw8zZq9QNGi5WnWrG3W52bo9U44\nlgyOBC4D/alcubIj+4ED0KgRmkuXWKkx8Lw8gwUX4D0cCww1QqOpzdSp3zBhwgRSUtoAY4E3gXnA\nGKAdJpMPM2Z8h6enJw0atKFXr8XYbBpgS1Y9kxDZQcmSJbNrHhMTw/z5y0hN/QOrdTyZmTuw25fi\n4lIcF5fi1KqVwccfv38/u1RRlCeECgSUx8LAgSPJyJgE5AO6AvPR6dpQv35dfHx8qFevKWvWlOPM\nmTmsWVOe0NBwrFYrgwcPxWAAaA4UwdV1K0uWzIGYGGjUCC5eZL3ByLNSjAxex7Hh0FWgBxrNHj74\noB/t2rUlMnIvUOC6GuUDzuHu3oD33+/Irl1RDBs2mrS0SFJSFgLfAc1wdw/DZKpAp05tqF+/fnbu\ny5cvYzAU46/9D6AErq6+LFgwlT//3Mz69csxGo0oiqLklnp9UHksBAfX4eDBT4EQHEsCb6B8eYiM\n3MRnn41j2LDvsdmO4RiuF9zdK7Bx449UqVKFhQsX8vvvGylWzI+33uqNe3w8NGwI58+T8dRT+G3/\nk8SMrkA0MBz4Co1mKUajFjAAZqpWDeGPP/YC3wDFgHfw8jrPpUsnaNmyHWvXxmOxWHGMPDi4ugYy\nduzbhIaG/j0KkSUxMZGSJYNJTPwcaIVGMxsfn885deogzs7O97s7FUV5hOT6vpcH8xQeKo9hk5S7\nMGrUZ2IyVROIElgnJlMR+e2332TMmM/FaCwl4CtgzppoZxaTyV8OHjyYs6BDh0T8/Bwz8ho1Ekti\nori4eApsE3hDwF3ARZycPAUissrbI0ZjAXF2dhcoJ1BG9HovmTNnrpw+fVqMRm+BMwLeAuuz8iwX\nDw9fSUlJuWWboqKipFSpSqLXGyU4uKYcOnToPvagoiiPqtze99SIgPJYsNvtjBgxhmnTfsLJyYnh\nwwfQuHEjSpeuSFraehzbB18C2mAwLCAszIfly39Bo9H8XUhsLDz1FCQkwNNPw/LlYDKxYMFCOnbs\njl5fg8zM/YSF1WHJkrXAleysLi5NGDeuLVFRB0hJSed//2tHs2bNOHXqFOXK1SY9/QywHngVSMPN\nzcTKlYsIDQ39L7tJUZTHUG7veyoQUB5LcXFxVKlSlytXUoAooDjwHRrNdFq1KsaCBRE3rsx39Kjj\ncUB8PNa6dfmsYRgRKzbi5ZWPsWM/xsvLi/3791OkSBFef/0t9uzZB2wHKgCX0WiC2LdvPRUr3jiT\nX0SoU6cJe/cWISOjEzrdcvz9f+Pgwd2YTKb/qjsURXmMqUDgH1QgoAC88cZbTJ/ugc0mOL6JDweO\n4OY2jL17t1G6dOm/Ex875ggCzpxhr3s+GqRkcE1sQBvgaVxdB7Nnz1YCAwP58MNhfPrpVGy2hsDv\nQC0gkoIFXblw4ehN63Lt2jXeffcjduzYS9mypZk4cTS+vr73tf2Kojw5cnvfU28NKI+lCxeuYrMF\n4pg4+DwwCFfXT9m8+fcbg4Djxx2PAc6c4WCBgjQyt+KaXMOx4uAxwITZ/Crz50cAMGXKD9hs44BV\nQF8gEL0+kwkTRtyyLu7u7kyePJG9ezcyd+50FQQoivJQUYGA8lhq27Y5JtOnwCEc7/IbGTDgTUJC\nQv5OdPKkIwg4fZp9rh7UvGzmqvVdQAd4AZ2AHWi1ZgwGx2MErVaHY0nixcB+NJqVtGsXzquvvvKf\ntk9RFCWvqEBAeWxcvXqV2bNnM2PGDMLCGjN48Ovkzx+Ou3stOnaszkcfvQc4NvvZ8csvpNeuDXFx\n7HZ2oX5qDVKoA2zIKs2e9e8juLou49VXXwVg0KB+mEwvA8fQaoPx8LjKp5+O+c/bqiiKklfUHAHl\nsRAbG0tISF0yMqqh0RjRaNaj0Qh2O9hsVnQ6LRUqVGXx4h/p3fplvvwzipJiJVJnoInNnSTmAkWB\nJkBZtNoE3NySaN26GcOGfUCpUqWyr/XTT3OYO3cZBQp48PHHA2581PAPNpuN0aM/Z+nSNfj5efP5\n58MoU6bMfe8PRVGeHGqy4D+oQODJk5mZia9vaa5caQt8AbTAsRfAMhxr87cCeqLXH6Vm4eXMPn2G\nUpLGLqoTTksS+QbogWNC4SXgVZo3d2Hx4vk4OTnlqm69evVnxoxdpKV9gEYTjYfHF8TERFG4cOFc\nlasoivIXFQj8gwoEnjyRkZHUrt0qaxJfraw/s3AEBOBY9/8nCjOEDdQkEDuRVCOM1SRyCmgIuAPl\ngGQMhiMkJMRSoECBm1zt3hiNHmRkxAKOCYIuLh354ou69OjRI9dlK4qigHprQFGw2+1ZawJMxLEM\nsAtw8LoUByiEsJ5WBGJnr8ZEOBEk4glMBmoA6cAaihW7SnT09jwJAoCsBYus1x3JRKtVP3aKojw8\n1IiA8lCz2Wxs3LiR5ORk6tSpc9NX7ywWCxUr1uboUR12+34c+wm4A62BDPxYwAaNliBJIzUggIHV\n6zE5IoLMTC1QCVgCeOHuXp7Nm3/Ose5/bgwY8CHffLOKtLT30On+JF++GcTEROHj45Nn11AU5cmm\nHg38gwoEHh9Wq5UmTdqwe3c8Wm1RRHaxbt2vVK9ePUfaK1eu0L//h/z552H8/DzYuHEzZnMaRQ16\ntrkY8Lt6FWu5chg2bQJvb2JiYqhWLRSzeSOOYGAXJlMz4uOPkS9fvjxrg4jw9dffsWTJGgoXLsiI\nER9QrFixPCtfURRFBQL/oAKBx8e0adPo23cOaWmrAD3wM2XLTuDgwZ13zGu321k8eTJle71FsNg4\noNESN2MGzTt2yE4zZ848unZ9E4PBn8zMs/z003SefbbN/WuQoijKfaACgX9QgcDj4+OPB/PJJxpg\nWNaReDw8qpGUdO6Oea8ePkxCufIEi41oytOIz0k1tScu7vANz/+vXLlCXFwcJUqUyNORAEVRlP+K\nmiyoPLZq1aqJq+tc4Bwg6PUTqVat5p0zXryI8zPPECw2YihHY9ZykWZkZBT8f3t3HhBVuf9x/D0z\ngDJgaqlYoGKIIoi466+ycCFy19zNpUxt1ex2y7LfvWn3XtR2tcwWLa+ZW2maW26hlqWppRYtdsVE\nRJMUlXVg5vn9Mcgvs7qZygjn8/qLOWfmzPMdjp4P8zzneXjhhWln/YO58soradKkyTkh4ODBg8yb\nN4+VK1dSVFR0cQsTEbmMKAjIZatr16489NAQ/P2vpUKFK4mK2sz8+a8B8N1339GxYw8aN27LuHF/\nx+VyeV+UmQkdO+L8/nu+sdlpz2v8SAjwPW73YZ59dh7Tpr30u+/78ccfEx3dnHvuWcaAARNp2/aW\n/z++iEg5o64Buezl5eWRk5PDVVddhc1mY+/evTRpch0ez6NAaxyOJOrXP8noQb0YtXgxjj17KKhT\nh3qHMjnk9gNigN1AP+BuwsJuJy0t5TffLyKiCfv3PwH0Ajw4nYlMndqfESNGlEa5IiLn5UKve37/\n/SkivhUYGIifnx9ZWVlUqVKFfv2G4PHcBDwOgNvdhiNfV6HN3/fjMFl4IiJ4sm0HDr15NXAH8D3w\nI/AsYP/VfzDGGHbs2MGxY8c4ejQdaFO8x05eXisOHUovjVJFREqdugbksvfyy68SHFyVmjXr0KBB\nM3744QDeqYO9qvAja3HT1GTxvc3Jm8OGsevIEbyndx2gAxAKZOPvP5CxY+866/jGGG67bQTt2vVn\n4MCp5OcX4XBMxrvw0EECA+dzww3Xl1K1IiKlS10Dclnbvn077dr1Ijd3ExCB3T4Zh+NZCgsDgAFU\npiFrGUsrcvkPtUnwj+dYhffxeBqSm7sbeAaoDvwFOMXo0bczdepzxTP+eb3//vsMGvS/ZGd/AjiB\n+fj53Q/kYrPZSEpK4q9/HVv6xYuI/AHqGpAyzxjDgQMHKCoqIiIigqKiIjZu3Ehubi7ffvstHk93\noB4AHs9DeDyPA06u4F0+4DCtKGI/dtpxP4c9/8TkDsHjeRGYC9yP9zQvYvjwQUyb9vw57+997+vx\nhgCA3ng8Qzh5MovAwEAcDkdpfAwiIj6hICA+5XK56NatP1u2fILNFkBkZBgejyE11Y3NVgOPZxs2\nW3XABQQAH3PVVWGEVgrmlQNHaE0RqTjpaPfDr85b1LHVYf/+W4uP/i5wMzAWu30Ly5fP4MSJE1St\nWvWsNjRr1gy7/RngEBCGzTaT+vWbEhwcXIqfhIiIb2iMgPjUlCnPsmVLIXl5B8nNPcDevTl8+eVV\nZGd/yunTK8jN/QcVKuQSHNyUSpV643T2Y9Gs6eyoUYk2nCAjoCJvDh3E3lOH2b9/N927dyYwcDpw\nDFgHzAOux+N5lIKCGJKTkwHvzINnXH/99TzxxFgCAhridF5DaOhLLF/+tg8+DRGR0qcgID61ffte\n8vL64/1rfyEezwGMac+ZU9OYtjidFVm+/EVef70/Kds+pP3TT+O/fTvUqsXV36Qwcc5rBAUFATBp\n0gQ6dAjA4aiN91uEvOJ3MsBpkpM3U6lSNQICKtK+fTdOnDgBwCOPPMiXX+7i8cfvZ/z4sVSuXLl0\nPwgRER/RYEHxqfHjn+D5578lP/8NoCYwHngb2ABUxWYbwa23enjnnTmQnQ2dO8OWLZyuXJlpt/al\nSe+edOnS5ZzjZmdnM3r0wyxatJvc3BEEBHxESMhWMjNPkZf3ARBJQMADtG//E6tXv8O+ffto1eom\nCgo6AB4CAzexc+dHhIeHl+KnISJy/rTWwC8oCJQtubm53HRTZ1JSDpGbewzIwhsGXgDAZqvI0qVv\n0qNjR+jSBTZt4oifPx3sHUlxtcXpfJ2JE0czZsy9fP3111SsWJH69etjs9nweDy89NJMkpO3ERER\nRoUKdiZNcuF2Tyl+9x9xOhuSk/MTt946hGXLYoonKQK7/QkGDjzMW2+95ouPRUTkD9NaA1KmOZ1O\nPv10Axs3zqNKlWBgPjAJSAacVKjQkcyDB6FbN9i0ibwqVbgloAUprpXAY+TmruPxxx+nQYNm3HBD\nf5o160Dnzn0oLCzEbrczevS9vPvuHJ566l/Url2bChX24O0mANhN1arVAcjIOIbH06ikXR5PLBkZ\nmaX5UYiI+ISCgPicw+Hg0KF0mjdvhs02CqgMJAKTCLJtpc+cOfDhh1CzJiseeoh9RAJn5gEowuXy\n48CBRLKzvyY39z9s2pTN9OnnricwZMgQIiNPERTUnsDAUTidg5g1ayoA3bp1wOmcjHeBo8M4nU/R\nvXuHUqlfRMSX1DUgPjdz5ms89NAkcnPHAzuAuQQGVsO/6Di761xN+Pf7jO+KBQAAHd5JREFUICQE\nkpNJrVCB2NhW5OS8BDTFZuuI99e9FGhWfMSX6dx5A9988x0//PAt114bw5Il/6ZRo0YUFBSwZMkS\nsrKyiI+Pp2HDhgC43W4eeOARXn/9VcDGfffdx9NP/wu7XVlZRC5vGiPwCwoCZU9YWEPS098EWgIe\n7PYHGHu3jclfp+D/4YdQowYkJ0PxRXvr1q3ceecDZGQc4tSp4xjTDYgEkoBC/P074XDsJD//ReBW\nYAHVqk3k4MFvCAwM/N22nDl3fj7zoIjI5UxjBKRM+PHHH7nttpE0b96B0aP/Sk5OTsm+wkIXMBsI\nAoLw93zEqDWrvSGgenXYuLEkBADUqFGDzMwfMSYK72yAPYAVeFcZDKN27Uz8/SOAwcX7h1NQUIlV\nq1Zx4MCB3/0HY7PZFAJExFJ8GgSGDx9OSEgIsbGxJduOHz9OQkIC9evX5+abbyYrK6tk36RJk4iM\njCQqKoq1a9f6osnyJ+Tl5dG6dTsWLarMrl3jeP31dDp16sOePXtYtWoVUVF1gc3AAQLI4F3SabB/\nP1SrBhs2QEzMWce7884HOH78AU6d+hBjlgD3ERBwNYGB+dx4YzNWrlxEUdEh4GTxK46Tnf0DQ4c+\nRMOGrenbdxhutxsREQGMD23evNns2rXLNGrUqGTbww8/bKZMmWKMMWby5Mlm3LhxxhhjvvrqKxMX\nF2dcLpdJTU01ERERxu12n3NMH5ckv2Ljxo2mUqVWBjwGjIFC43BUMRUrhhins5mBSgZeNwHkm+V0\nNQbMcYefMbt3n3OsEydOmGrVrjWwu/hYxsBE0759oklOTi45J+6+e6wJCmpoAgJGG4ejtrHZbih+\n/xzjdLY1M2fOLO2PQUTkkrjQ655PvxFo27btOfO+L1++nGHDhgEwbNgw3nvvPQCWLVvGwIED8ff3\nJzw8nHr16rF9+/ZSb7OcP++iPYU/21KE211Ifv775Ob+AHTHn+0soh/dWMFPOHm0+Q3QuHHJK4wx\njBp1H1WrhpKZeQyYhneZ4JM4nSsZNKgfN910U8ngvhkznmPRomeYNCmcypU9GDMT750GTnJze7Fz\n55elVb6IyGXtshsjcPToUUJCQgAICQnh6NGjABw+fJiwsLCS54WFhZGenu6TNsr5adOmDWFhDgIC\nRgCLCAjogcNRAwgGrsSPp1nI2/RgOcfxJ9Fu+KzQw5w5c0uOMX36DGbNSga+xNuNsByogsNxDQMH\ntuSOO24/6z1tNhudO3fmL3/5C3FxTXE43ive4yIwcBWxsQ0ufeEiImXAZb364H8buPVb+yZMmFDy\nc3x8PPHx8Re5ZXI+AgIC+OST9fztb//k668X0qBBQ2bN2oXbnYMfx1jAYHqRzQkCSaCQXZ6x8HkL\n7r77EU6ePElCQgcWLlyBx/MPoG7xUV8FxjNyZAdefnn6777/G29M5/rrEzh9+l2Kio7Ttm0z7rnn\n7ktdtojIJZGcnFyygNrFcNkFgZCQEI4cOULNmjXJyMigRo0aAISGhpKWllbyvEOHDhEaGvqrx/h5\nEJDLQ+XKlZk27emSxy1aNGP03e1505VPb7ORLBwk4GEXnfHeBgj5+T8wduw4goNDyck5DFTCezsg\nwDc4HCe58cbr/ut716lTh337drNnzx6cTieNGjXSnQEiUmb98g/ciRMnXtDxLruuge7duzNnzhwA\n5syZQ8+ePUu2L1iwAJfLRWpqavEiMa182VQ5Dzt27CApKYmXXnqJo0ePcvvgQZzokkBvU4inUiUe\njG7MTpoDzYtfsR14AmNWcPr0d3g824CVQE9gCPAk/frdzIABA/7Q+wcGBtK6dWtiY2MVAkREfsan\nEwoNHDiQTZs2kZmZSUhICE8++SQ9evSgX79+HDx4kPDwcBYtWkSVKlUASEpKYvbs2fj5+TF16lQS\nExPPOaYmFLr8vPfeewwadBcFBfF4PGtwkMOiioHcmp+NqVSJlQ88wP1zl/PDD6OACUBfYB7eMQT/\nPw6kUqV4Bg+OpkaNGvTu3fus205FRKxKMwv+goLA5WPjxo3Mm/cOb7+9gPz8ycD/Ymcuc5jLYOaR\njY1B1a5hY14LCgqOUlR0CngQeByYDtwFfAg0AdIIDGzO3r2fEBER4buiREQuMxd63bvsxghI+TB3\n7luMHPkQBQWPAMOAR7ETzRvMKw4BQXRxONmadQNFRQvw3grYGngACAAaA68BHYEIHI5v+cc/nlQI\nEBG5yBQE5KIoKChgzJhHWL58NU5nIAcOpOHxLAQSALBzhFksZSgfk00QnXiTrWYYHneL4iOcBr4D\nPsUbAP4KvALMAQYCLlJTf8AYoz5+EZGL6LIbLChl08iRY5g7dz9HjrzH/v0GjycIqAaADQ+v8R23\nU0AONnr5t2aX8wEGDuyH0zkbOAB8g3f54VjgKSAMaID324TpuN0ZvPnmBt555x1flCciUm4pCMgF\n279/P/Pnzycv7xXgTbwX9t7Avdj4lFe4heHsIhc7Sde15bZXh7B583Lmzp3No48OpWLFOByOm3A4\nTuCdLKgicC9g8N4pMAyoSk5Obz77bJdPahQRKa80WFAuSEZGBtHRzcnKKgKeBJ4D6gHNseHgZaZz\nF8fJAx6LbcU/t24gODj4rGOc+X2tW7eO3r1vw+UKwuXKBGoBY/EOGnThdCYydeptjBgxohQrFBG5\nvGkZYvGpxYsXk5eXCEwBxgPX4e3jf4+XeK4kBPQJqMZXITWpUKHCOcc4M4Nk8+bN2b17G2PG9MXf\nvzewEO8dBC2AurRqFcTtt99eWqWJiFiCgoBcEGNMcRL9GO/I/1WAnRe5gXs4TT7Qg9WscqXz8cf5\nzJjx8jnHcLvdDBp0J9dcU5fo6JYkJ3+G0/kBMALvOIFbsNlak5q6H5fLVYrViYiUfwoCckF69eqF\n270cOAF8DnRmKrW4j5kUAD2pzDq6ADeSl9eGPXu+PecY06a9xLJl/8HlyqCg4Ch794YTHx+PzbYH\nWAv8E2Pe5fjxamzcuLE0yxMRKfcUBOSC1K5dG4ejEO8iQOE8TxXG4KYAG72owAcsA/KBnthsM2jW\nLOas1xtj+OijneTmDgWCAD8KCkby/vtri28TPHOHqw0IwO12l15xIiIWoCAgF+THH3/E43EAB3iG\nvzKWqbiw0ZtrWU0H4CbAHxgH5NCvX18ANm/ezDXXXIvd7mD9+g8ICFiH9y4BgPV4PP8DXIWf363A\nWhyOvxEUlEq7du1Kv0gRkXJMEwrJn2aMISGhJ+6iVjzFjTxEbnEIuJKVHMN7ehUAFYDv8fe3U6VK\nFVauXEnXrn3x3l1QhVOn2mO3r8LPL4aioiuAY8CHeDwHCQ4eQGTkFOrWDeP55zdxxRVX+K5gEZFy\nSEFA/rRjx47xdcpXJGHjYXIpBPrixwoKgLeB+UALbLZmBAau47nnXsBms9G//zBgBdAeSAOaYrP5\n0aZNDbZurYrHsw7vksM7qFMnnB07NviqRBGRck9BQP6UEydOsOOzz5hQlM+juCjCQX9eZTmPAxFA\nXxyOAKpWvYJ77w2na9dlrFy5lmrVapGTU4A3BIB3roAWuN0evvjic664wp/s7Edwu6sRGPgKzz33\nts9qFBGxAgUBOW87duygY4euPF7gz/jiEDCABSylD7ATWILN1onatVNp2DCKDh06sH37Tp555h1y\nctYB8cAGoAPebwR2A+vJzX2LoUN/ol69WuTm5tG79xqaNWvmszpFRKxAMwvKeatdO5rhaTFM4B2K\ngEH8hcU8C+TgXUHwIN6M+TzeGQEfp169KPbsGQvcindp4d7AlUAm3hkJxwAzuO22L3jrrVd9UJWI\nSNmkZYil1BQWFjJ48EhuT/uGCXyNGzuDGcFiZuJdNfAg4MB7Ws0E+gGQm1tIZuZ0bLb9eM/VdsBY\nwsPnc+TIteTntwc+wumcwsCBM3xSm4iIVSkIyB+WlPQ0Dd9dxwQMbmAIQSykKgEBwQQF/UBRURF+\nfnayswMoLPz5nal2GjaM5PTpp8nP3w9AxYrvsmLFRubNW8zs2b0ICAhg4sQn6dKli09qExGxKnUN\nyB/2St2G3HXgGzzYGMq/mUciEM2wYT35+98fo3Hj1uTkTAV+BJKAaUABgYGPsGrVQurWrcvChQsB\n6N+/P3Xq1PFdMSIi5cSFXvcUBOSPmTIFHn0UD3A7c5jLUAD8/Hryxht9OH36NA89tIO8vFnFL1gA\n3E379u0YP/5+OnTo4KuWi4iUaxojIJfeM8/Ao49ibDaGG3/mUql4RxoOxzauuuou3G43dvuRn72o\nBYGBNtavX1I8VbCIiFyOFATk9z3/PDz8MAB3mgrMIRAYClQGjuPxBHPrrUNJTEygevUfKCwcisvV\nGKdzJhMnPqEQICJymVPXgPy2qVNh7FgA7rI14FXzEzAKqAL8E+gLvAbk4XR25sknu5Ofn8/hw8dI\nTGxH9+7dfdZ0ERGrUNeAXBrTp5eEgHvtfrzquQUIBP5V/IRXgPvxrgroJDe3D8nJm3n//UU+aa6I\niPw5Wn1QzvXSSzBmDAAPB1VlpvEHTgJVf/akOsDS4p+LgPdZu3Y1mZmZpdpUERG5MAoCcpYttw2G\n++8HYDQJPJMzAGNqAu/i/TZgKfAR3qmBnwVaAg0BBy5XFJMmTfFNw0VE5E/RGAEpsfWO4Vz35hsA\njOFKptMaCAIGAy8DXwPhQB7QDfgHsAS4Bm8YiCUg4CiHDx/gqquuKv0CREQsSPMI/IKCwJ/0+usw\nciQAD/I3XuBFIADvX/7+wCfAAGA/3mmEc4CaeAcO3gZsAmpTqdKXbNnyNnFxcaVfg4iIBV3odU9d\nAwKzZ8OoUQA8RD9e4EngZsCFdzAgeLsAsrHbuwAzgHY4HA78/bOAU8Bo4H48nqOEh4eXdgUiIvIn\nKQhY3Zw5MGIEGEPqPfcw07kRmIz3L/1CoA/wPnAfEEydOj/QqdN6xoy5joyMfSxY8G+czgUEBz9B\nUFAvFi+eS+XKlX1YkIiInA91DVjZ3LkwbBgYw6nHHmNNkyZMmzaLQ4fSSUs7jMcTDlyLtxsgEmiB\nwzGe6tUr8NFHa4mIiADg9OnTpKenU6tWLYKCgnxXj4iIBWmMwC8oCPxB8+bB0KHg8fBM1RDGnTyN\nxxOA906AAmAcMAb4AFgEGKAHcA92ex5t2qzl448/8FnzRUTES0HgFxQE/oD582HwYPB4mOSswvjc\nLsBPwB1Av+InXQfUB8LwTh6UB3TBu5jQf6hWLYFjx1J90HgREfk5DRaU87NwYUkIWBLbhPG5Drz9\n/wV47w44oy/wHvAMkIV3zMDrADgcb9GoUaPSbbeIiFwSmmLYShYvhttuA4+Hb/v1o987H+C9wK/H\ne1fAKMAN5ANTgO7Ap9jtPzBgwC0sWXItfn5VqFatAnPnrvFZGSIicvGoa8Aq3n0X+vcHt5uPb7qJ\nm7Zsx+1pBaTgzYPVgP8AVwAN8IaAZ4C7qVr1FY4fT+fgwYNkZ2cTGRmJv7//b76ViIiUHo0R+AUF\ngV+xdCn06wdFRcyrVZfBaSeB/wUm4e3/TwK+KX7yFXgHBuYBtXA6jzN9+j8ZPvx2X7RcRET+C60+\nKL9v2bKSELD1hrYM/uiL4h03A1cBtwMV8Y4PWA0cB7YBkwkJOcWCBW8RHx/vg4aLiEhpUBAoz95/\nH/r2haIitrRuQ/tPvgJuwXuHwCPAPCAGbyioVfxzMLAXu93J6tVLadq0qa9aLyIipUBBoLxauRL6\n9IHCQl4LqsK9O2Iocn+Fd9bAOKAj3sWCAGKBCkAofn7VCA4uYPXqVQoBIiIWoDEC5dHq1dCzJ7hc\n7G3XjqabKuL2rMI7J8ALeFcTzMO7jsBxbLa2tG0bRv/+PbjuuuuIioqiYsWKvqxARET+II0RkLN9\n8AH06gUuF9x/P+O+34/bUw3YjffiPwqoDvwd74DBr/D3z2DWrGXUq1fPhw0XERFfUBAoT9at834T\nUFCAuftu9tx5Jx/+TyLeyYKCgRvxdg0sxjtDYBIVKlTh3XfnKgSIiFiUugbKiw0boGtXyM/HjBrF\ngON5LF/xAfn5LryTBX2F9xuBr4CrgQ0EBw/k5Mkj2O2aYFJEpKzSFMMCu3dDt26Qnw8jRvD2DTew\nYtW35Ocvw3tbYA5wZfHPkfj5RRMcPJDlyxcqBIiIWJyuAuVBTAx07w7Dh8Mrr7AxOZnc3AS8F/88\nvHcH9AUaAoH07duEgwe/pV27dr5stYiIXAY0RqA88PODt94Cu51Va9Ywe/Y8oDbexYTyi3/eA9wA\nZJCYmEjVqlV92GAREblcaIxAOZKSkkJMTBugEAjCu4CQH1AVuBtYT0TEIVJSdhAQEODDloqIyMVi\nuTECa9asISoqisjISKZMmeLr5lw2Tp06RfPm7QAH4AGewjtPQBPgBE7nZCZPjmfv3m0KASIiUqJM\nfSPgdrtp0KAB69evJzQ0lJYtWzJ//nwaNmxY8hwrfiNw7NgxwsKicLk8eC/8R4EjQHvgE2y2k3zz\nzS7q16/v03aKiMjFZ6lvBLZv3069evUIDw/H39+fAQMGsGzZMl83y+fatu2Iy2XHO02wE+8KgrnA\nSipVKmTv3m0KASIi8qvKVBBIT0+nVq1aJY/DwsJIT0/3YYt8Lysri+++2w9Uwzs2IBvYBRhsNgcH\nDnxDTEyMT9soIiKXrzJ114DNZvtDz5swYULJz/Hx8eV+GV27vRC3uwDv4MAUvGMEAli2bD5XXnml\nbxsnIiIXVXJyMsnJyRfteGUqCISGhpKWllbyOC0tjbCwsHOe9/MgUN5VqVKFzp17smrVbtzuXMCN\nw1GRPXs+Izo62tfNExGRi+yXf+BOnDjxgo5XpgYLFhUV0aBBAzZs2MA111xDq1atNFgQcLlcTJiQ\nxKZN26lXrxZPP/0PatSo4etmiYhIKbjQ616ZCgIAq1evZuzYsbjdbu68804ee+yxs/ZbMQiIiIh1\nWS4I/DcKAiIiYiWWun1QRERELi4FAREREQtTEBAREbEwBQERERELUxAQERGxMAUBERERC1MQEBER\nsTAFAREREQtTEBAREbEwBQERERELUxAQERGxMAUBERERC1MQEBERsTAFAREREQtTEBAREbEwBQER\nERELUxAQERGxMAUBERERC1MQEBERsTAFAREREQtTEBAREbEwBQERERELUxAQERGxMAUBERERC1MQ\nEBERsTAFAREREQtTEBAREbEwBQERERELUxAQERGxMAUBERERC1MQEBERsTAFAREREQtTEBAREbEw\nBQERERELUxAQERGxMAUBERERC1MQEBERsTAFAREREQtTEBAREbEwBQERERELUxAQERGxMAUBERER\nC1MQEBERsTAFAREREQtTEBAREbEwBQERERELUxAQERGxMJ8EgcWLFxMTE4PD4WDXrl1n7Zs0aRKR\nkZFERUWxdu3aku07d+4kNjaWyMhIHnjggdJucpmRnJzs6yb4jJVrB9Wv+pN93QSfsXLtF4NPgkBs\nbCxLly7lxhtvPGt7SkoKCxcuJCUlhTVr1nDvvfdijAHgnnvuYdasWezbt499+/axZs0aXzT9smfl\nfxBWrh1Uv+pP9nUTfMbKtV8MPgkCUVFR1K9f/5zty5YtY+DAgfj7+xMeHk69evXYtm0bGRkZnD59\nmlatWgEwdOhQ3nvvvdJutoiISLlzWY0ROHz4MGFhYSWPw8LCSE9PP2d7aGgo6enpvmiiiIhIueJ3\nqQ6ckJDAkSNHztmelJREt27dLtXbEhERgc1mu2THLwsmTpzo6yb4jJVrB9Wv+q1bv5Vrj4iIuKDX\nX7IgsG7duvN+TWhoKGlpaSWPDx06RFhYGKGhoRw6dOis7aGhob96jO+///78GysiImJRPu8aODMY\nEKB79+4sWLAAl8tFamoq+/bto1WrVtSsWZMrrriCbdu2YYxh7ty59OzZ04etFhERKR98EgSWLl1K\nrVq1+PTTT+nSpQudOnUCIDo6mn79+hEdHU2nTp2YMWNGydf8M2bMYMSIEURGRlKvXj1uueUWXzRd\nRESkXLGZn/9JLiIiIpbi866BP0uTEp1tzZo1REVFERkZyZQpU3zdnEti+PDhhISEEBsbW7Lt+PHj\nJCQkUL9+fW6++WaysrJK9v3WeVAWpaWl0a5dO2JiYmjUqBHTpk0DrFN/fn4+rVu3pkmTJkRHR/PY\nY48B1qn/DLfbTdOmTUsGXFup/vDwcBo3bkzTpk1LbiW3Sv1ZWVn06dOHhg0bEh0dzbZt2y5u7aaM\n+vrrr823335r4uPjzc6dO0u2f/XVVyYuLs64XC6TmppqIiIijMfjMcYY07JlS7Nt2zZjjDGdOnUy\nq1ev9knbL7aioiITERFhUlNTjcvlMnFxcSYlJcXXzbroNm/ebHbt2mUaNWpUsu3hhx82U6ZMMcYY\nM3nyZDNu3DhjzK+fB2632yftvhgyMjLM559/bowx5vTp06Z+/fomJSXFMvUbY0xOTo4xxpjCwkLT\nunVrs2XLFkvVb4wxzz77rBk0aJDp1q2bMcY6578xxoSHh5uffvrprG1WqX/o0KFm1qxZxhjv+Z+V\nlXVRay+zQeCMXwaBpKQkM3ny5JLHiYmJ5pNPPjGHDx82UVFRJdvnz59v7rrrrlJt66WydetWk5iY\nWPJ40qRJZtKkST5s0aWTmpp6VhBo0KCBOXLkiDHGe7Fs0KCBMea3z4PyokePHmbdunWWrD8nJ8e0\naNHCfPnll5aqPy0tzXTo0MFs3LjRdO3a1RhjrfM/PDzcZGZmnrXNCvVnZWWZunXrnrP9YtZeZrsG\nfosVJyVKT0+nVq1aJY/P1GwFR48eJSQkBICQkBCOHj0K/PZ5UB4cOHCAzz//nNatW1uqfo/HQ5Mm\nTQgJCSnpJrFS/Q8++CBPP/00dvv//7dtpfptNhsdO3akRYsWvPbaa4A16k9NTaV69erccccdNGvW\njJEjR5KTk3NRa79k8whcDL6alKissfoESmfYbLbf/SzKw+eUnZ1N7969mTp1KpUqVTprX3mv3263\n88UXX3Dy5EkSExP58MMPz9pfnutfsWIFNWrUoGnTpr85r355rh/g448/5uqrr+bYsWMkJCQQFRV1\n1v7yWn9RURG7du3ixRdfpGXLlowdO5bJkyef9ZwLrf2yDgK+mpSorPllzWlpaWclwvIsJCSEI0eO\nULNmTTIyMqhRowbw6+dBWf99FxYW0rt3b4YMGVIyj4aV6j+jcuXKdOnShZ07d1qm/q1bt7J8+XJW\nrVpFfn4+p06dYsiQIZapH+Dqq68GoHr16vTq1Yvt27dbov6wsDDCwsJo2bIlAH369GHSpEnUrFnz\notVeLroGjMUnJWrRogX79u3jwIEDuFwuFi5cSPfu3X3drFLRvXt35syZA8CcOXNKfqe/dR6UVcYY\n7rzzTqKjoxk7dmzJdqvUn5mZWTIqOi8vj3Xr1tG0aVPL1J+UlERaWhqpqaksWLCA9u3bM3fuXMvU\nn5uby+nTpwHIyclh7dq1xMbGWqL+mjVrUqtWLb777jsA1q9fT0xMDN26dbt4tV+sAQ2lbcmSJSYs\nLMxUrFjRhISEmFtuuaVk37/+9S8TERFhGjRoYNasWVOyfceOHaZRo0YmIiLCjB492hfNvmRWrVpl\n6tevbyIiIkxSUpKvm3NJDBgwwFx99dXG39/fhIWFmdmzZ5uffvrJdOjQwURGRpqEhARz4sSJkuf/\n1nlQFm3ZssXYbDYTFxdnmjRpYpo0aWJWr15tmfr37NljmjZtauLi4kxsbKx56qmnjDHGMvX/XHJy\ncsldA1apf//+/SYuLs7ExcWZmJiYkv/jrFL/F198YVq0aGEaN25sevXqZbKysi5q7ZpQSERExMLK\nRdeAiIiI/DkKAiIiIhamICAiImJhCgIiIiIWpiAgIiJiYQoCIiIiFqYgICIiYmEKAiIiIhamICAi\nf9pnn31GXFwcBQUF5OTk0KhRI1JSUnzdLBE5D5pZUEQuyN/+9jfy8/PJy8ujVq1ajBs3ztdNEpHz\noCAgIheksLCQFi1aEBgYyCeffFJml3sVsSp1DYjIBcnMzCQnJ4fs7Gzy8vJ83RwROU/6RkBELkj3\n7t0ZNGgQ+/fvJyMjg+nTp/u6SSJyHvx83QARKbv+/e9/U6FCBQYMGIDH4+G6664jOTmZ+Ph4XzdN\nRP4gfSMgIiJiYRojICIiYmEKAiIiIhamICAiImJhCgIiIiIWpiAgIiJiYQoCIiIiFqYgICIiYmH/\nB2H+zOtjC0adAAAAAElFTkSuQmCC\n", + "text": [ + "
\n", + "
\n", + "**Eventually, the benchmark:**:" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "import timeit\n", + "import random\n", + "random.seed(12345)\n", + "\n", + "funcs = ['py_lstsqr', 'cy_lstsqr']\n", + "\n", + "orders_n = [10**n for n in range(1, 6)]\n", + "times_n = {f:[] for f in funcs}\n", + "\n", + "for n in orders_n:\n", + " x = [x_i*random.randrange(8,12)/10 for x_i in range(n)]\n", + " y = [y_i*random.randrange(10,14)/10 for y_i in range(n)]\n", + " for f in funcs:\n", + " times_n[f].append(min(timeit.Timer('%s(x,y)' %f, \n", + " 'from __main__ import %s, x, y' %f)\n", + " .repeat(repeat=3, number=1000)))" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 13 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "import matplotlib.pyplot as plt\n", + "\n", + "labels = [('py_lstsqr', 'regular Python (CPython)'), \n", + " ('cy_lstsqr', 'Cython implementation')]\n", + "\n", + "\n", + "matplotlib.rcParams.update({'font.size': 12})\n", + "\n", + "fig = plt.figure(figsize=(10,8))\n", + "for lb in labels:\n", + " plt.plot(orders_n, times_n[lb[0]], alpha=0.5, label=lb[1], marker='o', lw=3)\n", + "plt.xlabel('sample size n')\n", + "plt.ylabel('time per computation in milliseconds [ms]')\n", + "plt.xlim([1,max(orders_n) + max(orders_n) * 10])\n", + "plt.legend(loc=2)\n", + "plt.grid()\n", + "plt.xscale('log')\n", + "plt.yscale('log')\n", + "max_perf = max( py/nu for py,nu in zip(times_n['py_lstsqr'],\n", + " times_n['cy_lstsqr']) )\n", + "min_perf = min( py/nu for py,nu in zip(times_n['py_lstsqr'],\n", + " times_n['cy_lstsqr']) )\n", + "ftext = 'Using Cython is {:.2f}x to '\\\n", + " '{:.2f}x faster than regular (C)Python'\\\n", + " .format(min_perf, max_perf)\n", + "plt.figtext(.15,.8, ftext, fontsize=11, ha='left')\n", + "plt.title('Performance of least square fit implementations in Cython and (C)Python')\n", + "plt.show()" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "display_data", + "png": "iVBORw0KGgoAAAANSUhEUgAAAnIAAAIECAYAAACdVcNJAAAABHNCSVQICAgIfAhkiAAAAAlwSFlz\nAAALEgAACxIB0t1+/AAAIABJREFUeJzs3XdYFFf3B/DvLB2k4yIg0gVBRWONhaIYY0mssRdsMZpo\nNOaNvkYFVGKLNWLeiL3GjgajWLE3xIqKIkJEBY1dRGnn98f+GFhYYEHYBTyf5/FJ9u7MnTNnyl7u\n3JkRiIjAGGOMMcYqHYm6A2CMMcYYY6XDDTnGGGOMsUqKG3KMMcYYY5UUN+QYY4wxxiopbsgxxhhj\njFVS3JBjjDHGGKukuCGnBpmZmRg6dCgsLCwgkUhw/PhxdYdUKW3btg1OTk7Q1NTE0KFDFU7j7++P\ndu3aqTgyln/bHDt2DBKJBA8fPixxXQ4ODvjll1/KIcqC7O3tERwcrJJlVVQSiQSbNm1Sdxjw8fHB\n119/re4wilVV95mEhARIJBKcPn26yOmICI0aNcK2bdtKVH9WVhbq1KmDffv2fUiYJVJVfw+4IVcI\nf39/SCQSSCQSaGlpwd7eHqNGjcKzZ88+uO4dO3Zg8+bNCA8PR3JyMj799NMyiPjjkpWVhaFDh6JP\nnz64f/8+Fi9erHA6QRAgCIJKY9uwYQMkko/30FK0bVq0aIHk5GRYWVkBAE6ePAmJRIJ//vmn2Pqi\noqIwfvz48g4bgHr2lw+RlJRU6j8G/fz8MGTIkALlycnJ6NGjR1mE90HCwsKwYMGCMqkrJiYGAwcO\nRM2aNaGrqwt7e3t0794dkZGRStcxc+ZMODg4FCivbPtMWdu0aRPev3+Pr776Sq68uJxraGjg559/\nxsSJE+Xmi4yMFH97JRIJLCws0KZNG5w8eVLpmAo7v1TVbfXx/toowcvLC8nJyUhMTMSSJUuwc+dO\nDBo0qNT1paenAwDu3LkDGxsbNG/eHFKpFFpaWh9U38fo4cOHSE1NRYcOHWBlZQVDQ0OF0xER+JnX\nJZednY3s7OxSzato22hpaUEqlRY4iSqzbczNzaGnp1eqWD4WZbmPS6VS6OjolFl9pWViYoJq1ap9\ncD0RERFo3LgxkpOTsXLlSty8eRPh4eFo3rw5vvnmmzKI9OO2aNEiDBs2TK5M2Zz36NEDiYmJOHr0\naIF6L126hOTkZBw+fBh6enro0KEDEhMTSxRb/uOiyv4WEFNo8ODB5OfnJ1cWHBxMGhoa9O7dOyIi\n2rx5M3l6epKuri7Z29vTDz/8QKmpqeL03t7eNGzYMJoyZQpZWVlRjRo1yMfHhwRBEP85ODgQEVF6\nejpNnDiRbGxsSFtbm9zd3WnTpk1yyxcEgZYsWUJ9+/YlY2Nj6t27N61evZo0NTXp6NGjVLduXdLT\n0yNfX1969OgRHTlyhDw9PcnAwID8/PzowYMHYl3x8fHUrVs3sra2Jn19fapXrx6tX79ebnne3t40\nfPhwmj59OtWoUYPMzMxo0KBB9ObNG7np/vzzT/rkk09IV1eXzM3NqUOHDvT8+XPx+yVLlpCrqyvp\n6uqSi4sLBQcHU2ZmZpH5P3PmDLVu3Zr09PTI1NSU+vXrR48fPyYiotWrV8vlUBAEOnbsmNLbsbjt\nduDAAfL29iYzMzMyNjYmb29vOn/+vFwdoaGh5ObmRrq6umRmZkZeXl6UlJRER48eLRDbkCFDCl3P\n4OBgcnR0JB0dHapevTq1b9+e0tLS5HJnY2ND+vr61L59e1q7di0JgiBuy5ztn9f9+/cL5GT48OHk\n5OREenp65OjoSJMnT6b379+L3wcEBJCzszNt2bKFXF1dSVNTk27dukWvX7+msWPHijE0bNiQdu7c\nWej6FLZtcvLy4MEDunfvXoFpfH19C63Tzs6OZs6cKfd56tSp9M0335CxsTFZWlrSsmXLKC0tjUaP\nHk2mpqZkY2NDS5culatHEARavHgxde/enQwMDMjGxoYWL14sN429vT0FBweLn9PT0ykgIIAcHBxI\nV1eXPDw86I8//ihQ72+//Ua9evUiAwMDsrOzo507d9KzZ8+oT58+ZGhoSI6OjrRjxw65+ZKTk2nw\n4MFUvXp1MjQ0pJYtW9Lx48fF73NydvDgQWrdujXp6+uTu7s77du3T27Zis4nxR3fgwcPLvQYEgSB\nNm7cKE778OFD6t27N5mYmJCenh75+PhQVFRUieIkKn5fzy/n/JP/c3Hno7xSU1NJKpVSx44dFX7/\n4sULMR+fffZZge99fX1p2LBhtGbNmgL5CgoKIiLZPjNt2jQaO3YsmZmZkaWlJY0fP17uHKfs+X3Z\nsmU0YMAAMjQ0pJo1a9KsWbMKXbccyh7bu3fvJldXVzIwMCAfHx+6c+eOXD1btmwhJycn0tXVpRYt\nWtDu3btJEAQ6depUocuOjY0lQRDo3r17YpmyOc/Rq1cvGjp0qPg577kix4MHD0gQBFq+fDmtXr2a\nTExM6O3bt3L1BAUFkYuLS5Hnl5zfgz/++INq1apFRkZG9OWXX1JKSopcXWvWrKE6deqQtrY21axZ\nk6ZMmSK3PUuzL5YnbsgVYvDgwdSuXTu5svnz55MgCPTmzRtavXo1mZqa0oYNG+jevXt0/Phxql+/\nPg0cOFCc3tvbmwwNDWnUqFF08+ZNun79Oj179ox+/PFHcnBwoJSUFPr333+JiOjHH38kc3Nz2r59\nO925c4d++eUXkkgkdPjwYbE+QRDI3NycQkJCKD4+nu7cuUOrV68miURCvr6+dP78eYqOjiYXFxdq\n1aoVeXl50blz5+jy5cvk5uZGvXv3Fuu6du0ahYSE0NWrVyk+Pp5+++03sUGYN34TExP64YcfKDY2\nlg4cOEBmZmY0depUcZpVq1aRlpYWzZw5U1zHpUuXiusVEBBAdnZ2FBYWRgkJCfT3339TrVq15OrI\n79GjR2RoaEj9+/en69ev08mTJ6l+/frk5eVFRERpaWl04cIFEgSB/vrrL0pJSaH09PRCt2Pehpwy\n223Xrl20bds2un37Nt24cYOGDx9OZmZm9PTpUyIiioqKIk1NTVq/fj39888/dO3aNVq5ciUlJSVR\neno6hYSEkCAIlJKSQikpKfTq1SuFse3YsYOMjIwoPDyc7t+/T5cvX6bFixeLP25hYWGkqalJCxcu\npDt37tDKlStJKpWSRCIpUUMuOzubfv75Zzp//jwlJibSnj17yMrKigICAsR5AgICSF9fn3x8fOj8\n+fN0584dev36Nfn4+JCvry+dOnWK7t27R8uXLydtbW25/TKvwrZN3pNzVlYW7dmzhwRBoKioKEpJ\nSZFr+OeXv3FlZ2dHJiYmtHDhQrp79y7NnDmTJBIJtW/fXiybNWsWSSQSunHjhjifIAhkZmZGS5cu\npTt37tDixYtJU1OTdu/eXeiyBg8eTJ6ennTw4EFKSEigLVu2kImJCa1cuVKu3ho1atC6devo7t27\nNHr0aDIwMKDPPvuM1q5dS3fv3qUxY8aQgYGBuA+9ffuW6tSpQz179qSLFy/S3bt3KTg4mHR0dOjm\nzZtElPuD5unpSRERERQXF0dDhgwhIyMjMV+XLl0iQRBo165dcueT4o7vly9fkpeXF/Xp00fcT3OO\nobwNuezsbGratCk1bNiQTp06RdeuXaPevXuTqampuCxl4ixuX1fEx8eHRowYIX5W5nyU365du4pt\njBDJ/nCUSCRyDZI7d+6QRCKh8+fPU1paGk2aNIlsbW3FfOX88WdnZ0empqY0Z84ciouLo61bt5KW\nlpbcPqLs+d3S0pJWrFhB8fHx4nmksGONSPlj28DAgDp06EDR0dF05coVatSoEbVu3VqcJjo6mjQ0\nNGjy5Ml0+/Zt2rlzJ9nb2xebuz/++IOqV69eqpznmD9/Ptnb24ufFTXknj59SoIgUEhICKWlpZGp\nqSmtXbtW/D4rK4vs7Oxo7ty5RZ5fBg8eTMbGxtSvXz+KiYmhM2fOkIODg9z5Pzw8nDQ0NGj27Nl0\n584d2rJlC5mamsrtZ6XZF8sTN+QKkb8BEBMTQ46OjvTpp58Skezgzf+X+bFjx0gQBPEvDm9vb3J1\ndS1Qd85fSDlSU1NJR0eHfv/9d7npunXrRm3atBE/C4Ig9xcqUW4PyJUrV8SyefPmkSAIFB0dLZYt\nXLiQLCwsilznLl26FDhxNmjQQG6aUaNGiTkgIrK1taUxY8YorC81NZX09fUpIiJCrnzt2rVkYmJS\naBxTpkwhW1tbysjIEMuuXLlCgiCIPRY5f3UVd7LIvx2V2W75ZWVlkampqfjjtnPnTjI2Ni60gbZ+\n/XoSBKHIuIiIFixYQLVr15Zbz7xatmxJAwYMkCv78ccfS9Ujp2jZLi4u4ueAgACSSCR0//59sezo\n0aOkq6tLL1++lJt3yJAh1LVr10LrVrRt8p+cT5w4QYIgUGJiYqH15FDUkOvWrZv4OTs7W/zLOm+Z\nqampXK+cIAg0aNAgubr79esn94OWd1nx8fEkkUgoNjZWbp6goCC540IQBBo/frz4+cmTJyQIAo0d\nO1Yse/78OQmCQHv37iUi2XarWbNmgZ5pX19fGjduHBHl5mzXrl3i9ykpKSQIAh04cICIlNvWOfIf\n335+fgp7i/M25A4dOkSCIIiNSyKi9+/fk5WVFU2fPl3pOIvb1xVR1JAr7nyU35w5c0gQhCL/UMhR\nv359mjJlivh50qRJcsubMWOGXIMjh52dHXXp0kWurEOHDtS3b18iKtn5/fvvv5ebpk6dOvTf//63\n2NjzUnRsa2pqig1vIlnvm0QiEXvu+vfvT61atZKrZ+nSpcWeYydMmECNGzeWKytJzolkjXxBEMR9\nI2d/SkpKIiKiV69e0fDhw0lbW5tiYmKIiGjs2LFy8e7fv5+0tbXpyZMnRFT4+WXw4MFkaWkp94f/\nnDlzyMrKSvzcqlUruU4PIqLFixeTnp6eGGNp9sXyxGPkihAZGQlDQ0Po6+ujXr16cHZ2xsaNG/Hk\nyRP8888/GD9+PAwNDcV/HTt2hCAIiIuLE+to1KhRscuJi4tDeno6vLy85Mq9vLwQExMjV9a0adMC\n8wuCgHr16omfLS0tAQD169eXK3v69Kk4RuDt27eYNGkS6tatC3NzcxgaGuLvv/+WGxwqCAI8PT3l\nlmVlZYWUlBQAwOPHj5GUlITPPvtM4XrFxMQgLS0N3bt3l8vTN998g1evXuHp06eFzte8eXNoamqK\nZfXr14exsTFu3LihcB5lKLvd7t27h4EDB8LFxQXGxsYwNjbGy5cvxdx89tlncHR0hIODA/r27YvQ\n0NBC16UovXv3RkZGBuzs7DBkyBBs2LABb968Eb+/efMmWrRoITdPy5YtS7XuoaGhaNasGWrUqAFD\nQ0NMnjy5wEBgS0tL1KxZU/x84cIFpKenw8bGRi5fGzdulNvHVS3/fikIAqpXry63vwuCAKlUiidP\nnsjNm//GohYtWhQ4xnJERUWJd+TlXf9Zs2YVWP+88VhYWEBDQ0MuHhMTE2hra+Px48cAZLlNTk6G\niYmJXN0nT54sUHeDBg3E/5dKpdDQ0BCPwcIoc3wrIyYmBubm5nBzcxPLtLW10axZswJ5KyrO4vZ1\nZRR3PlKESjAmauTIkVi9ejWICJmZmVizZg1GjBihVFx51z1/XCU5v+evx9raWtxnCqPMsW1tbQ1z\nc3O5+IhIrLu055qXL18WGMdYkpwDgJGREQDgxYsXcuWurq4wNDSEiYkJDh48iLVr18Ld3R2AbFud\nOnUKsbGxAGQ56NKlCywsLIpdnpubm9y49Pz70I0bNxRuq3fv3uHu3btiWUn3xfKkWfwkH6/mzZtj\n7dq10NTUhLW1tdiwyNlYS5Ysga+vb4H5bGxsAMgOcAMDgzKNSVF9EolEbhB5zv9raGgUKCMiCIKA\n//znP9izZw8WLlwIV1dX6OvrY8KECXj58qVc3dra2nKfBUFQehB8znTbt29H7dq1C3xvamqqcD5B\nEMplUGpOPMVtt86dO0MqlWLZsmWwtbWFlpYWWrVqJd5cYmBggKioKJw6dQqHDh3C//73P/z00084\nfPgwPvnkE6Xjsba2xq1bt3D06FEcOXIEM2bMwMSJE3Hu3Dm5BlVRFN0dm5GRIfd527Zt+O677zBn\nzhx4e3vDyMgIW7duxc8//yw3Xf59Kzs7G8bGxoiKiiqwjPz7harlv0FIEASFZaW9YQPI3V/OnDkD\nfX39AnUXFU9hMebUmZ2djTp16iAsLKzAfPmXpSjXxa2Xssd3aeWcR5SNsyz2dUXLKG4b5zRAY2Ji\nim2YDBgwABMnTkR4eDiysrLw6tUrDBgwoFziKqt6lD22FdULyO9HpTnnmpiY4PXr13JlJck5AHGf\nNDExkSs/cOAArKysYGZmBmNjY7nv3N3d0apVKyxfvhwTJ07EX3/9hb179yoVs6LjsqTrLghCmW3z\nssA9ckXQ1dWFo6MjatWqJdc7ZGlpCVtbW9y6dQuOjo4F/pX0ji9nZ2fo6Ojg2LFjcuXHjh2T62kr\nSydOnMCAAQPQs2dP1KtXDw4ODoiNjS3RrdlSqRQ1a9ZERESEwu89PDygq6uLu3fvKsxTYY/o8PDw\nwNmzZ+UaJFeuXMHLly9Rt27dkq1oHspst6dPn+LmzZuYNGkS2rVrBzc3N+jo6BT4q1gikaB169YI\nCgrCxYsXYWVlhc2bNwPIPWkqc3LQ1tZG+/btMWfOHFy7dg1v377F7t27AchOVqdOnZKbPv9nqVSK\nrKwsufiio6Plpjl+/DgaNmyIcePGoWHDhnBycsK9e/eKja1JkyZ48eIF0tLSCuSqJD++iuTkKCsr\n64PqKakzZ87IfT59+jQ8PDwUTpvTm56YmFhg/RU9hqIkmjRpgvj4eBgaGhaou0aNGkrXU1gelTm+\ntbW1kZmZWWT9Hh4e4jGR4/379zh37lyJj8Wi9vXy8tlnn0EqlRb6nLfnz5+L/29kZIQ+ffogNDQU\nK1asQK9evcTeopz4S7O/luf5vbTHdn7u7u4FnheX/1yjiIuLS4E7SUuSc0B2fNnb28v9xgKy5/M5\nODgUaMTlGDlyJNatW4fly5ejZs2a8PPzE78r6vxS3G+ch4eHwm2lr68PJyenIudVl0rXI5eSkoLu\n3btDW1sb2tra2LRpk1yXsaoEBwdj2LBhMDU1xZdffgktLS3cvHkT+/fvx//+9z8Ayj/6Ql9fH2PH\njsXUqVPFS0Tbt2/Hnj17cOjQoXKJ39XVFWFhYejevTsMDAywYMECPHr0SO5HRJn4AwICMGrUKFha\nWqJHjx7Izs7G0aNH0bdvX5ibm2Py5MmYPHkyBEFA27ZtkZmZiWvXruHy5cuYPXu2wjq/++47LF68\nGP7+/pg8eTKeP3+O0aNHw8vLq9SXFnMUt91MTU1RvXp1LF++HI6Ojvj333/x008/yT3+Yvfu3bh3\n7x5at26N6tWr4+LFi7h//77Y7Z/zI7979260bNkS+vr6CntSV65cCSJCkyZNYGJigsOHD+P169di\nPRMmTMBXX32Fpk2bokOHDjh58iQ2bNggdyJq1qwZDA0NMWnSJPz3v//F3bt3MX36dLnluLm5YdWq\nVdizZw88PDwQHh6OXbt2FZurNm3awM/PD927d8fcuXNRr149PH/+HKdPn4aenh6GDx9e8g3w/+zs\n7CCRSLB371706tULOjo6hZ6w8++DivZJZcv27t2LkJAQfPbZZ9i/fz+2bt2K7du3K5zH2dkZQ4cO\nxYgRIzB37lw0b94cqampuHjxorhflFb//v2xcOFCdOrUCcHBwXBxcUFKSgqOHDkCd3d3dOnSRal6\nLCwsUK1aNURERKBOnTrQ0dGBqampUse3g4MDjh49ivj4eBgZGcHExKTAj2nbtm3RtGlT9OvXDyEh\nITAyMsKMGTOQnp6OUaNGKb2+xe3riuQ//yh7Ps1LT08Pa9asQbdu3dCuXTtMmDABtWvXRmpqKiIi\nIrBixQrcunVLnH7kyJFo3rw5BEEo8Gw+R0dHJCcn4+zZs3B2doaBgQH09PSKjelDzu/FrXNpj+38\nxo8fjyZNmmDKlCkYNGgQYmJilHqGn7e3N54+fYqEhATY29sDKHnOz549Cx8fnxLH3LNnT4wbNw4z\nZ85EQECA3Hf5zy+6urpio7y47fXf//4XX3zxBebMmYNu3brh8uXLCAoKwoQJE8TjozT7YnmqdD1y\n1atXx6lTp3D06FH069cPoaGh5bIcoZgHBw4YMABbt25FeHg4mjVrhqZNmyIoKEiup6KwOhSVBwcH\nY8SIERg3bhzq1auHTZs2YePGjQovASqqr6RlCxcuhJ2dHXx9feHn5wdbW1v07NmzwCXa/PXkLxs2\nbBjWrFmD7du3o2HDhvD29kZERIS4w0+ZMgULFixAaGgoGjRogNatW2Px4sVF9mhIpVIcOHAASUlJ\naNKkCb744gvx5FfcOipa57zTFbfdJBIJtm3bhrt376J+/foYOnQoxo8fLz7IFgDMzMzw119/oUOH\nDnB1dcWkSZMwdepU8eGqTZo0wffff4+RI0fC0tISY8aMURibmZkZVq9eDV9fX7i7u2PRokUIDQ0V\nt3nXrl0xf/58zJ07F56enti8eTPmzJkjdwIxNTXF5s2bcfbsWXh6eiI4OBjz5s2TW+eRI0di4MCB\nGDJkCD755BNcuHABgYGBxW5rANizZw+6d++O8ePHo06dOujcuTP27dsHZ2fnYvNeVJmlpSVmzZqF\n2bNnw9raGt26dVO6rtLs7zmmTZuGQ4cOoUGDBpg9ezbmzZsn12jKP8/y5csxfvx4BAcHw8PDA35+\nfli/fv0H/2We00PTuHFjDBkyBK6urujRoweioqLEH8TC1iEviUSCkJAQbN26Fba2tmIvojLH94QJ\nE2BhYQFPT09IpdJCn+AfFhYGNzc3dOrUCU2bNsXjx49x8OBBmJmZKR1ncfu6Ivn3SWXOR4p8/vnn\nuHDhAiwtLTFs2DBxPz5x4gSWLFkiN23jxo1Rr149uLm5FRhP2bVrV3z11Vfo1KkTpFIp5s2bV+i6\n54+rtOf34tbvQ47tvGWffPIJNm3ahD///BP169fH3LlzsXDhwmJz6+rqisaNG2Pnzp1y5crmPC0t\nDREREQUuYStzbtfR0cGAAQNARAXe7JP//NK1a1elc9GhQwesWrUKa9euRb169fDDDz/g22+/lWss\nlnZfLC8CVaRmZQn99ttv0NbWxsiRI9UdCmMqERkZiTZt2iApKQnW1tbqDqdSkUgk2LBhA/r166fu\nUFgFlZGRAXt7e0yaNKnQP8CYvE2bNiE4OLjQm4aKsn79esybNw9Xr14t1bJ79eqFrKws7Nixo1Tz\nVxWVrkcOkI2XatasGZYuXYq+ffuqOxzGGGOVWM4dnLNnz0ZaWprCV5cxxfr16wc9Pb1SvWv1l19+\nwdy5c0u8zOfPnyMiIgJhYWEqe31fRaa2htzSpUvRuHFj6OrqFjhonj17hm7duqFatWqwt7cXB5Hn\n8PT0xLlz5zBz5kzMmDFDlWEzpnbq6r5nrKpKTExEjRo18Mcff2DVqlVl8mqwj0lUVFSBd60WR0ND\nAzdv3sTnn39e4uU1bNgQX331FSZOnIhWrVqVeP6qRm03O9jY2GDq1KmIiIhAWlqa3HfffvstdHV1\n8fjxY1y6dAmdOnWCp6cn3N3dkZGRId4+bGRkhPfv36sjfMbUwsfHR+V3elYV6no0AKv47O3tef+o\nRBISEtQdQoWi9jFyU6dORVJSElavXg0ASE1NhZmZGWJiYsQB1YMHD4a1tTVmzZqF8+fP4z//+Q80\nNDSgpaWFlStXKnwUgo2NDR4+fKjSdWGMMcYYKw1PT09cvny5xPOpfYxc/nbk7du3oampKXdXnKen\npziQsmnTpjh27BiOHDmCiIiIQp9n9fDhQ/EWYVX8CwgIUOn8ykxf1DSFfadsuaLpPjQHqsx3Seso\nr3yXJJfKbIOKnPOy3sdL+z3nu/TT8zml7Orgc0rV3sdLk+8rV66Uqh2l9oZc/vE+b968kXsIIwAY\nGhoWeHp0RVOa5+B8yPzKTF/UNIV9p2y5oulU2d39ofkuaR3lle/CvlOmTNWXFyraPl7a7znfpZ+e\nzyllVwefU6r2Pq7KfKv90uqUKVPw4MED8dLqpUuX0KpVK6SmporT/Prrrzh+/Dj27NmjdL3l9Zon\nVjh/f3+sWbNG3WF8NDjfqsX5Vj3OuWpxvlUrf75L226pcD1ytWvXRmZmptyLo69cuVKqVzMFBgYi\nMjLyQ0NkSvL391d3CB8Vzrdqcb5Vj3OuWpxv1crJd2RkJAIDA0tdj9p65LKyspCRkYGgoCA8ePAA\noaGh0NTUhIaGBvr27QtBELBixQpER0ejc+fOOHPmDOrUqaN0/dwjxxhjjLHKotL1yM2YMQP6+vqY\nM2cONmzYAD09PfElu8uWLUNaWhqkUikGDBiA//3vfyVqxDH14N5P1eJ8qxbnW/U456rF+Vatssq3\n2p4jFxgYWGhXoqmpaale/MsYY4wx9jFR+80O5UUQBAQEBMDHx6fAnSJmZmZ4/vy5egJjjBXJ1NQU\nz549U3cYjDGmEpGRkYiMjERQUFCpLq1W6YZcYavG4+cYq7j4+GSMfYwq3Rg5xhirTHj8kOpxzlWL\n861aZZVvbsgxxhhjjFVSVfrSamFj5PjSDWMVFx+fjLGPCY+RKwSPkWOscuLjkzH2MeIxckxlJBIJ\nNm3apO4wSqwixD1+/HiMHj1arTHkp6q8ZGVloU6dOti3b1+5L6s88Pgh1eOcqxbnW7V4jByrkvz9\n/SGRSCCRSKClpQV7e3uMGjWqRI+jGD58OHx9fcsxytK5d+8eQkNDMWXKFLnyp0+f4qeffoKbmxv0\n9PRgaWkJb29vrF+/HllZWQCqRl40NDTw888/Y+LEiWqLgTHGqhq1PRC4IouNTcShQ3eRkSGBllY2\n/Pyc4OpqV+HqBID09HRoa2t/cD2qlpmZCU1Nxbufl5cXtm7diszMTERFRWHEiBG4f/8+wsPDVRxl\n2Vq2bBnatm0La2trsez+/fto1aoVtLW1MX36dDRs2BBaWlo4deoUfv31V3h6eqJ+/foAqkZeevTo\ngW+//RbKJLnuAAAgAElEQVRHjx6tkI3touQfa8vKH+dctTjfqlVW+a7SPXKBgYEl7rqMjU3EmjVx\nePKkDV688MGTJ22wZk0cYmMTSx1HWdbp4+OD4cOHY+rUqbCysoK9vT0AIC4uDj169ICpqSnMzMzQ\nvn17XL9+XW7ezZs3w8nJCXp6emjdujX27t0LiUSC06dPA5B180okEjx8+FBuPk1NTaxdu7bQmBYv\nXoyGDRvC0NAQVlZW6Nu3L5KTk8Xvc+r9+++/0apVK+jp6WHlypWF1qelpQWpVApra2t8+eWX+P77\n77F//368e/cOPj4+GDlypNz0RAQnJyfMnDkTQUFBWLVqFY4dOyb2YK1bt06c9uXLlxg4cCCMjIxg\na2uL2bNny9X1+vVrjBw5ElKpFLq6umjSpAkOHjwofp+QkACJRIJt27ahc+fOMDAwgJOTU5H5ybFx\n40Z069ZNrmz06NHIyMhAdHQ0+vbtCzc3Nzg5OWHQoEGIjo6Gs7NzlcqLnp4ePv/8c2zYsKHYfDHG\n2McgMjKy0DddKYWqqKJWrajvli49TAEBRN7e8v86dpSVl+Zfhw6HC9QXEEAUEnK4xOvl7e1NhoaG\nNGrUKLp58yZdv36dkpOTydLSkkaPHk3Xr1+n27dv05gxY8jc3JyePHlCRERRUVEkkUho6tSpdPv2\nbQoLCyNnZ2eSSCR06tQpIiI6evQoCYJADx48kFumpqYmrV27VvwsCAJt3LhR/Lx48WI6fPgwJSQk\n0JkzZ6hFixbk7e0tfp9Tr5ubG4WHh1NCQgIlJSUpXL/BgwdTu3bt5Mrmz59PgiDQmzdvaPPmzWRo\naEhv3rwRvz906BBpamrSo0eP6M2bN9S/f39q2bIlpaSkUEpKCr17906M29LSklasWEHx8fEUEhJC\ngiDQ4cO526Fnz57k4OBABw4coFu3btH3339P2tradOvWLSIiunfvHgmCQI6OjrRt2za6e/cuTZ48\nmTQ1Nen27duFbrfY2FgSBIFu3Lghlj19+pQ0NDQoODi40PmqYl7mz59P9vb2ha5rRT0tHT16VN0h\nfHQ456rF+Vat/Pku7bmvSvfIlUZGhuKUZGWVPlXZ2YrnTU8vXZ3W1tZYtmwZ3Nzc4OHhgd9//x0O\nDg4ICQmBh4cHXFxcsHjxYpiYmGDjxo0AgAULFqBVq1aYPn06XFxc0KVLF/z4449lcnfg2LFj0aZN\nG9jZ2aF58+ZYunQpjh8/jkePHslNN2XKFHTq1Al2dnawsbEptL68Md24cQMhISFo3rw5DAwM0K1b\nN+jq6uLPP/8Up1mxYgU6d+6MGjVqwMDAALq6umLvlVQqhY6Ojjhtnz59MGzYMDg4OGD06NFwc3PD\noUOHAMh6NXfs2IFly5ahXbt2cHV1xaJFi1C3bl3MnTtXLsYxY8agZ8+ecHR0xIwZM6Cnp1dk7+/t\n27cBALVq1RLL4uLikJ2dDXd39yKyW/XyYm9vj8TERGRmZiq13owxxgrHDbl8tLSyFZZraCguV4ZE\nonhebe3S1dmoUSO5zxcuXMDFixdhaGgo/jMyMkJiYiLi4uIAyH74mzdvLjdf/s+lFRkZifbt26NW\nrVowMjJC69atAQCJifKXjps2bap0fYaGhtDX10e9evXg7OwsNkh1dHTg7++P0NBQALIbBcLCwjBi\nxAil6m7QoIHcZ2trazx+/BiALEeAbCxaXl5eXoiJiSm0HolEAqlUipSUlEKX+/LlSwCAgYGBWFbS\nRnRVyYuRkREA4MWLF0rFVlHw+CHV45yrFudbtcoq33yzQz5+fk5Ys+YwfHzaimXv3x+Gv78zXF1L\nV2dsrKxOHR35Otu2dS5iLsUEQZBrDACyBoGfnx+WLl1aYHpjY2NxPkEQiqxbIpGI9eXIyspCdnbh\nDc5//vkHHTt2xODBgxEYGAgLCwvcv38ffn5+SE9Pl5s2f9yFad68OdauXQtNTU1YW1sXuCli5MiR\nmD9/Pq5du4bDhw9DKpWiQ4cOStWt6MaQotYPUNzgyl+PIAhF1mNiYgIASE1NFfPg4uICiUSCmJgY\ndO3atdjYq0pechq1OTlhjDFWetwjl4+rqx38/Z0hlR6BiUkkpNIj/9+IK/0dpuVRZ16NGzfG9evX\nYWNjA0dHR7l/5ubmAAB3d3fxpoYcZ8+elfsslUoBAA8ePBDLLl++XGTP0YULF/Du3TssWrQIn376\nKVxcXORudCgNXV1dODo6olatWgrvbHVyckKbNm0QGhqKlStXYujQoXKNVG1tbfGxHcXJO5+HhwcA\n4NixY3LTHD9+HPXq1SvNqohcXFwAyPdSmpmZoUOHDli6dClevXpVYJ6MjAy8fftW/FxV8pKYmAh7\ne/tC71quqPgZW6rHOVctzrdqlVW+K9eZtIQCAwMVvqKrOK6udmXWyCrrOomoQMPqu+++w8qVK9Gl\nSxdMmTIFNWvWRFJSEvbt24fOnTvj008/xQ8//IAmTZogICAA/fv3x61bt7BgwQIAuT/azs7OsLOz\nQ2BgIBYuXIgnT55g8uTJRfbkubi4QBAE/Prrr+jXrx+uXLmCGTNmfPB6FmfkyJHo378/srOzMXz4\ncLnvHB0dsX37dty4cQNSqRRGRkaFPqIlbz6dnJzw1VdfYfTo0fjjjz9Qq1Yt/P7777hx44bc2LPC\n6ilK7dq1UaNGDZw7d05uTNyyZcvQsmVLNGrUCNOnT4enpye0tbVx9uxZ/Prrr1i3bp34+BFlVIa8\nnD17li/hMMbY/8t5RVdpVekeuZyGXFWi6BKpVCrFmTNnYGFhge7du8PNzQ0DBgzA/fv3xWeWffLJ\nJ9i4cSM2btyI+vXrY86cOWKDS1dXF4DsMSNbtmzB48eP0bBhQ4wZMwa//PKLeMlVkfr16+O3337D\nH3/8AQ8PDyxYsACLFi0qEGNxl3WLWj9FunbtChMTE3z++ecFbpwYNmwYmjRpghYtWkAqlRbZ2Mi/\nvBUrVqB9+/YYMGAAGjRogDNnziA8PBy1a9cucl2UiXnAgAHYtWuXXJmtrS2io6PRtWtXBAYGolGj\nRmjZsiVCQ0MxatQosTesquQlLS0NERERGDBgQLHrUtFUtXNJZcA5Vy3Ot2rl5NvHx+eDHj/C71r9\niK1btw5Dhw7Fs2fPxAHolcXTp09ha2uLLVu24IsvvlB3OEpJSEhA3bp1ERsbW+Rdux+ioudl/fr1\nmDdvHq5evVroNHx8MsY+RvyuVVasX3/9FRcvXsS9e/ewdetWTJo0Cb169apUjbjMzEwkJyfj559/\nRs2aNStkY6Uw9vb2+Prrr/HLL7+Ued2VIS9ZWVn45ZdfCjyypLLg8UOqxzlXLc63avEYOVZi165d\nw4IFC/Ds2TPY2tpi4MCBCAoKUndYJXLy5Em0adMGjo6OWL9+vbrDKbGccYllrTLkRUNDAzdv3lR3\nGIwxVqXwpVXGWIXCxydj7GPEl1YZY4wxxj4yVbohFxgYyNf8GWNlgs8lqsc5Vy3Ot2rl5DsyMvKD\n7lqt0mPkPiQxjDHGGGPlLed5t6Uds85j5BhjFQofn4yxjxGPkWOMMcYY+8hwQ44xxpTA44dUj3Ou\nWpxv1SqrfHNDjsnx9/dHu3bt1LZ8BweHcnlgriL29vYIDg5WybIqKolEgk2bNqk7DMYYY6XEDblK\n6OnTp/jpp5/g5uYGPT09WFpawtvbG+vXr0dWVpZSdZw8eRISiQT//POPXLmy7/QsL1FRURg/frxK\nlqXudS2ppKQkSCQSHD9+vMTz+vn5YciQIQXKk5OT0aNHj7IIr8rj91CqHudctTjfqlVW+a7Sd61W\nRffv30erVq2gra2N6dOno2HDhtDS0sKpU6fw66+/wtPTE/Xr11e6vvwDK9U9yNzc3Fyty68MynIb\nSaXSMquLMcaY6nGPnAKxcbEI2RKCRX8uQsiWEMTGxVaYOkePHo2MjAxER0ejb9++cHNzg5OTEwYN\nGoTo6Gg4OztjzZo1MDU1RVpamty806dPR+3atZGQkAAvLy8AskuZEokEbdq0EacjIixfvhx2dnYw\nNjZGly5d8PjxY7m61q5dC3d3d+jo6MDW1hZTp06V6w308fHBiBEjMGPGDFhZWcHc3ByDBw9Gampq\nkeuX/3Knvb09pk2bhlGjRsHExAQ1atTA77//jnfv3uHbb7+FmZkZatasiZCQELl6JBIJlixZgh49\neqBatWqoWbMmlixZUuSyMzIyEBgYCEdHR+jp6aFu3bpYvnx5gXqXLl2K3r17o1q1arC3t8euXbvw\n/Plz9O3bF0ZGRnBycsLOnTvl5ktJSYG/vz+kUimMjIzQqlUrnDhxQvw+MjISEokEhw4dgpeXFwwM\nDODh4YH9+/eL09SqVQsA4OvrC4lEAkdHRwDAvXv30L17d9jY2MDAwAD169fHhg0bxPn8/f1x5MgR\nrF27FhKJRK5XL/+l1UePHqFPnz4wNTWFvr4+fH19cfHixRLFWVXx+CHV45yrRmxsIhYtOoJhwxYh\nJOQIYmMT1R3SR4HHyCmhNA8Ejo2LxZqja/DE8gle1HiBJ5ZPsObomg9qzJVVnc+ePcO+ffvw3Xff\nwdDQsMD3Ghoa0NfXR58+fSAIArZt2yZ+l52djVWrVmHEiBGoVasWdu/eDQC4cOECkpOT5RoeFy5c\nwLFjx7Bv3z5ERETg2rVr+PHHH8Xv9+7di2HDhmHw4MGIiYnB/PnzERISUuAZONu3b8eLFy9w7Ngx\n/PnnnwgPD8ecOXOKXEdFlzt/++03uLq6Ijo6GmPGjMF3332Hrl27wsXFBVFRUfjuu+8wduzYAu/x\nDAoKQps2bXD58mX89NNPmDBhAvbs2VPoskeMGIGwsDAsX74ct27dwrRp0zBx4kSsWrVKbrrg4GB0\n7twZV69eRadOnTBw4ED06dMHHTp0wOXLl9GpUycMGjQIz549AwCkpaXB19cXqamp2L9/Py5fvoyO\nHTuiXbt2uHXrllzdP/74I6ZMmYKrV6+iWbNm6N27N168eAEAiI6OBgDs3LkTycnJuHDhAgAgNTUV\nfn5+2L9/P65fv46vv/4aQ4YMEff9JUuWoHXr1ujduzeSk5ORnJyMTz/9tMD6ExG6du2K27dvY+/e\nvTh//jwsLS3Rrl07PH36VOk4GWOVR2xsIn7/PQ4HDrTB1asN8OBBG6xZE8eNORX60AcC83Pk8gnZ\nEoInlk8QmRApV26QZIAmrZqUKpbzJ8/jbc23cmU+9j6QPpZidK/Rytdz/jyaN2+OnTt3omvXrkVO\n+/333yM6Olrs9YmIiMCXX36JBw8ewMLCAidPnoSXlxcSEhLEnh5A1nuzf/9+3L9/H1paWgCAuXPn\nYtGiRXj48CEAoHXr1rCxscGff/4pzrdkyRJMmjQJr169gqamJnx8fPDy5UtcunRJnGb06NG4fPky\nTp8+XWjcDg4OGDFiBCZPngxA1iP3ySefiA1NIoKJiQl8fHzExigRwdzcHDNmzMC3334LQNbTNHDg\nQKxdu1asu3///rh//77YG5V3Wffu3YOzszNu3ryJ2rVri/NMnz4du3btEtdDIpFg3LhxWLBgAQDg\n33//hVQqxZgxY7B48WIAwIsXL2BmZobw8HB07NgRa9aswdSpU5GQkAANDQ2x7jZt2sDT0xMLFy5E\nZGQk2rRpI7dtHz9+jBo1aiAiIgLt2rVDUlISatWqhcjISLFHtTBdu3aFVCoVexTbtWsHW1vbAo1S\niUSCDRs2oF+/fjh8+DDatWuHGzduwM3NDQCQnp4Oe3t7jBo1ClOnTlUqzg/Fz5FjTHUCAo7g1Kk2\nyMyUfTYyAho2BCwtj2D06DZFz8zKVGnPfTxGLp8MylBYngXlbiJQJBvZCsvTs9NLVE9JNvDIkSNR\nt25dxMbGwtXVFaGhoejSpQssLCyKndfNzU1sxAGAlZUVUlJSxM83btxA37595ebx8vLCu3fvcPfu\nXbi6ugIAPD095aaxsrJCRESE0usAyHbsvPUIgoDq1avLjQMUBAFSqRRPnjyRmzd/r1OLFi0wbdo0\nhcuJiooCEaFRo0Zy5ZmZmdDUlD9M8sZjYWEBDQ0NuXhMTEygra0tXo7O6fU0MTGRq+f9+/cwMDCQ\nK2vQoIH4/1KpFBoaGnK5V+Tt27eYPn06wsPD8ejRI6Snp+P9+/dyl8uVERMTA3Nzc7ERBwDa2tpo\n1qwZYmJiPjhOxljFQQScOgWcOycRG3ESCWBtDQgCkJ5epS/YVSnckMtHS9BSWK4BDYXlypAUcgVb\nW6JdonpcXFwgkUgQExNTbI+cu7s7WrVqheXLl2PixIn466+/sHfvXqWWk7cRB5TurwRBEKCtrV2g\nLDtbcaO2pPEoKitN3Tly5j1z5gz09fUL1F1UPIXFmFNndnY26tSpg7CwsALz5V9W/pzlja0w//nP\nf7Bnzx4sXLgQrq6u0NfXx4QJE/Dy5csi51MWERXIQWnirOwiIyP5rj4V45yXj/R0YPduICYGkEhk\nx62ODmBsHIkaNXwAANraVft4rgjKav/mhlw+fo38sOboGvi4+Ihl7++8h38ff7g6u5aqztiasjFy\nOi46cnW29W1bonrMzMzQoUMHLF26FGPGjIGRkZHc9xkZGcjIyBAbByNHjsS4ceNgamqKmjVrws/P\nT5w254dY0eNKinskh4eHB44dO4bRo3MvCx87dgz6+vpwcnIq0TqVpzNnzuCbb74RP58+fRoeHh4K\np83piUtMTESnTp3KNI4mTZpg/fr1MDQ0RPXq1UtdT2Hb7MSJExgwYAB69uwJQNagio2NhZWVldy8\nmTl/dhfCw8MDT58+xc2bN1GnTh0Asl7Dc+fO4bvvvit13IyxiuPFC+DPP4HkZNlnR0cn3Lt3GJ6e\nbfH/o2fw/v1htG3rrL4gWYlw32k+rs6u8Pf1h/SxFCbJJpA+lsLft/SNuLKuc9myZdDS0kKjRo2w\nefNm3LhxA3FxcdiwYQOaNGmCuLg4cdqcH/aZM2di+PDhcvXY2dlBIpFg7969ePz4MV69eiV+V1zv\n23//+1/s2LEDc+bMwe3bt7F161YEBQVhwoQJ4mVIIirVtX5lHoeibNnevXsREhKCO3fu4LfffsPW\nrVsxYcIEhfM4Oztj6NChGDFiBDZs2IC4uDhcuXIFq1atwty5c0u8Hnn1798fDg4O6NSpEw4ePIiE\nhAScO3cOs2bNEsf5KcPCwgLVqlVDREQEkpOT8fz5cwCAq6srwsLCcOHCBdy4cQNff/01Hj16JLd+\nDg4OuHjxIuLj4/Hvv/8qbNS1bdsWTZs2Rb9+/XD69Glcv34dgwYNQnp6OkaNGvVBOagKuGdI9Tjn\nZevePWD58txGHAB06GCHefOcUbPmETRoAEilR+Dv7wxXVzv1BfqR4OfIlSNXZ9cPariVZ522traI\njo7GnDlzEBgYiH/++QdGRkZwc3PDqFGj5HqcdHR0MGDAACxbtgxDhw6Vq8fS0hKzZs3C7NmzMW7c\nOHh5eeHIkSOFPiQ3b1mHDh2watUqzJ49G9OmTUP16tXx7bffIiAgQG76/PUo8wBeRfMUN01hZdOm\nTcOhQ4fw008/wcTEBPPmzUOXLl0KnWf58uWYP38+goODER8fDyMjI9StW/eDe6N0dHRw7NgxTJky\nBUOGDMGTJ09QvXp1NGvWDB07dixyHfKSSCQICQlBQEAA5s+fD1tbW8THx2PhwoUYPnw4fH19YWRk\nhJEjR6Jnz56Ij48X550wYQKuXbsGT09PpKamFnrDRFhYGMaPH49OnTrh/fv3aNasGQ4ePAgzMzOl\n42SMVSxEwLlzwIEDQM4ICA0NoGNHQHYxwg7u7txwq6z4rtUqrlevXsjKysKOHTvUHYpK5b0bk1Uu\nFfX45PFaqsc5/3CZmUB4OHD5cm5ZtWpA796Ara38tJxv1cqfb75rlcl5/vw5zp8/j7CwMBw5ckTd\n4TDGGFOxV6+ALVuABw9yy2xsZI24fEOsWSXGPXJVlL29PZ49e4bvv/8eM2bMUHc4Ksc9cpXXx3B8\nMlbe/vkH2LoVePMmt6xBA6BzZ0CTu3AqpNKe+7ghxxirUPj4ZOzDREUB+/YBOTe4SyRA+/ZA06ay\nZ8Sxiqm05z6+a5UxxpTA7/1UPc55yWRlycbDhYfnNuL09YGBA4FmzYpvxHG+Vaus8s0drIwxxlgl\n9+aN7FLqP//kltWoAfTpA+R7qQyrYqr0pdWAgAD4+PgUuAuHL90wVnHx8clYyTx4ILupIc/jQFG3\nLtClC6DgRTSsgomMjERkZCSCgoJ4jFxePEaOscqJj0/GlHflCvDXXxDflyoIgJ8f0KIFj4erbPjx\nIyVgamrKDzVlrIIyNTVVdwgK8TO2VI9zXrjsbNkDfs+ezS3T1QV69gScS/l2Lc63avG7Vj/As2fP\n1B1ClcQnAdXifDP2cXr7Fti2TfbKrRzVq8vGw5mbqy8uph4f5aVVxhhjrDJKTpa99P7Fi9wyNzeg\nWzdAR0d9cbEPx5dWGWOMsSosJgYICwMyMnLLfHwAb28eD/cx4+fIsTLDzyBSLc63anG+VY9zLpOd\nDRw+LLucmtOI09aWXUr18Sm7RhznW7X4OXKMMcZYFffuHbBjB3DnTm6ZubmsEVe9uvriYhUHj5Fj\njDHGKqAnT2Tj4Z4+zS1zdpbdmaqrq764WPngMXKMMcZYFREbC+zcCbx/n1vWqhXQpo3s3amM5eDd\ngZUZHl+hWpxv1eJ8q97HmHMi4NgxYPPm3Eaclhbw1VeyB/2WZyPuY8y3OvEYOcYYY6wKef9edlfq\nzZu5ZSYmsvFwNWqoLy5WsfEYOcYYY0zNnj2TjYd7/Di3zMFB1hOnr6++uJjq8Bg5xhhjrBKKiwO2\nb5fdoZqjeXPgs894PBwrHu8irMzw+ArV4nyrFudb9ap6zomAU6eAjRtzG3GamkDXrsDnn6u+EVfV\n813R8Bg5xhhjrJLKyAD27AGuXcstMzICevcGbGzUFxerfCrdGLnz589j3Lhx0NLSgo2NDdatWwdN\nzYLtUR4jxxhjrCJ68QLYsgV49Ci3zNZW1oirVk19cTH1Km27pdI15JKTk2FqagodHR1MnjwZjRo1\nQo8ePQpMxw05xhhjFU1CArB1K/D2bW5Zo0ZAx46AhobawmIVQGnbLZVujFyNGjWgo6MDANDS0oIG\n7/kVBo+vUC3Ot2pxvlWvKuWcCDh/Hli3LrcRJ5EAnTsDX3xRMRpxVSnflUFZ5bvSNeRyJCYm4uDB\ng/jiiy/UHQpjjDFWqMxM2Xi4v/8GsrNlZdWqAf7+QOPGag2NVQFqu7S6dOlSrFmzBtevX0ffvn2x\nevVq8btnz55h2LBhOHjwICwsLDBr1iz07dtX/P7Vq1f44osvsGLFCri4uCisny+tMsYYU7fXr2Xj\n4ZKScsusrWUP+TUyUl9crOKpdM+Rs7GxwdSpUxEREYG0tDS577799lvo6uri8ePHuHTpEjp16gRP\nT0+4u7sjMzMTffr0QUBAQKGNOMYYY0zd7t+XNeLevMkt8/SUXU7V0lJfXKxqUdul1W7duqFLly4w\nNzeXK09NTcXOnTsxY8YM6Ovro2XLlujSpQvWr18PANi8eTPOnz+PGTNmwNfXF1u3blVH+EwBHl+h\nWpxv1eJ8q15lznl0NLBmTW4jTiKRPRuua9eK24irzPmujKrMc+TydyPevn0bmpqacHZ2Fss8PT3F\nFR44cCAGDhyoVN3+/v6wt7cHAJiYmKBBgwbw8fEBkJtA/lx2ny9fvlyh4qnqnznfnO+q/jlHRYlH\nmc9ZWcDcuZGIjQXs7WXfP3wYCR8foHlz9cdX1OccFSWeqv758uXLiIyMREJCAj6E2h8/MnXqVCQl\nJYlj5E6cOIFevXrhUZ4H7ISGhmLTpk04evSo0vXyGDnGGGOqlJoqe7RIYmJumaWlbDycqan64mKV\nQ6UbI5cjf9DVqlXDq1ev5MpevnwJQ0NDVYbFGGOMKe3hQ9l4uJcvc8s8PIAuXQBtbfXFxao+iboD\nEARB7nPt2rWRmZmJuLg4sezKlSuoW7duiesODAws0GXMyg/nWrU436rF+Va9ypLzq1eBVatyG3GC\nAPj5AT17Vq5GXGXJd1WRk+/IyEgEBgaWuh619chlZWUhIyMDmZmZyMrKwvv376GpqQkDAwN0794d\n06ZNw4oVKxAdHY2//voLZ86cKfEyPiQxjDHGWFGys4GDB4G8P0+6ukCPHgA/VIEpy8fHBz4+PggK\nCirV/GobIxcYGIjp06cXKJs2bRqeP3+OoUOHis+Rmz17Nvr06VOi+nmMHGOMsfKSlgZs3w7cvZtb\nVr26bDxcvocxMKaUj+Zdq8rihhxjjLHykJIC/Pkn8Px5bpmrK9C9O/D/b5BkrMQ+mnetlgSPkVMt\nzrVqcb5Vi/OtehUx5zduACtXyjfivL1lPXGVvRFXEfNdlVX6MXKqwGPkGGOMlQUi4OhR4Pjx3DJt\nbaBbN6BOHfXFxSq/SjtGrrzxpVXGGGNl4d07YOdO4Pbt3DIzM1kvnFSqvrhY1VJpnyPHGGOMVVT/\n/isbD/fvv7llTk6yR4vo6akvLsZy8Bg5VmY416rF+VYtzrfqqTvnt28DoaHyjbiWLYH+/atmI07d\n+f7Y8Bg5JfAYOcYYYyVFBJw4IRsTl3OlS0sL+PJLoF499cbGqh4eI1cIHiPHGGOspNLTgbAw2d2p\nOYyNZePhrKzUFxer+niMHGOMMfYBnj+XjYdLSckts7cHvvoKMDBQW1iMFalKj5FjqsXjK1SL861a\nnG/VU2XO4+OB5cvlG3HNmgEDB348jTjex1WrrPJdpXvkAgMDxWvPjDHGWH5EwNmzwIEDuePhNDSA\nzp2Bhg3VGxv7OERGRn5Qo47HyDHGGPsoZWQAf/0FXL2aW2ZoCPTuDdSsqb642MeJx8gxxhhjSnr5\nUjYe7tGj3DJbW6BXL1ljjrHKgsfIsTLD4ytUi/OtWpxv1SuvnCcmysbD5W3EffIJMHjwx92I431c\nte6wy6gAACAASURBVHiMHGOMMVYCREBUFLBvH5CdLSuTSIAOHYDGjQFBUG98jJUGj5FjjDFW5WVm\nAn//DURH55YZGMgupdrZqS8uxnLwGDkF+K5Vxhhjr18DW7cC9+/nlllZyR7ya2ysvrgYA8rxrtWB\nAwcqVYGOjg5WrFhR6gDKC/fIqV5kZCQ3mlWI861anG/VK4ucJyUBW7bIGnM56tcHvvhC9totlov3\ncdXKn+8y75HbunUrJk+eXGilOQucP39+hWzIMcYY+7hdugSEhwNZWbLPggB89hnQvDmPh2NVR6E9\nck5OTrh7926xFbi6uiI2NrbMA/tQ3CPHGGMfp6ws2QN+z53LLdPTk71qy9FRfXExVpTStlv4ZgfG\nGGNVRmoqsG0bkJCQWyaVAn37AqamaguLsWKVtt1SqufIxcfHIyHvUcIY+BlEqsb5Vi3Ot+qVNOeP\nHsmeD5f358ndHRg+nBtxyuB9XLXKKt9KNeT69OmD06dPAwBWr14NDw8PuLu789g4xhhjFcK1a8Cq\nVbI3NgCyMXBt2sgup2prqzc2xsqTUpdWq1evjgcPHkBbWxt169bFH3/8ARMTE3Tp0gVxcXGqiLPE\nBEFAQEAAP36EMcaqsOxs4PBh4NSp3DIdHaBHD6B2bfXFxZiych4/EhQUVH5j5ExMTPDixQs8ePAA\nTZs2xYMHDwAAhoaGeJ33nu4KhMfIMcZY1ZaWBmzfDuS9L8/CQvZ8OAsL9cXFWGmU6xg5T09PzJo1\nC9OnT0enTp0AAElJSTDmJymyPHh8hWpxvlWL8616ReX88WMgNFS+EVe7tmw8HDfiSof3cdVS6Ri5\nlStX4urVq3j37h1mzJgBADhz5gz69+9fJkEwxhhjyrp5E1ixAnj2LLfMy0t2Z6qurvriYkwd+PEj\njDHGKgUiIDISOHYst0xbG+jaVXZ3KmOVWbm/a/XEiRO4dOkSXr9+LS5MEARMnjy5xAtljDHGSuL9\ne2DnTiDv8+dNTWXj4Swt1RcXY+qm1KXVMWPGoGfPnjh+/Dhu3bqFmzdviv9lLAePr1Atzrdqcb5V\nLyfnT5/KLqXmbcQ5OgJff82NuLLE+7hqlVW+leqR27BhA2JiYmBtbV0mC2WMMcaUcecOsGMH8O5d\nblmLFoCfHyAp1SPtGatalBojV79+fRw5cgQWlehWIB4jxxhjlVNsbCIOHryL27cluHs3Gw4OTrCw\nsIOmJvDll0D9+uqOkLGyV67vWr1w4QJ++eUX9OvXD5b5+rG9vLxKvFBV4IYcY4xVPrGxiVi1Kg7x\n8W3x5ImsLDPzMFq1csbYsXbgC0OsqirXmx0uXryIv//+GydOnICenp7cd/fv3y/xQlUlMDCQ3+yg\nQpGRkZxrFeJ8qxbnWzV2776L69fbIjUVePEiEiYmPjA3bwtLyyOwtrZTd3hVGu/jqpWT75w3O5SW\nUg25n3/+GeHh4WjXrl2pF6QOgYGB6g6BMcaYkuLigOPHJUhNzS2zsQGcnABB4AFxrGrK6XAKCgoq\n1fxKXVqtVasW4uLioF2J3jzMl1YZY6xyIJK9K/XwYeDcuSN4+7YNBEH2pgYrK9k0UukRjB7dRr2B\nMlaOyvUVXdOnT8e4cePw6NEjZGdny/1jjDHGSis9Hdi2DTh0SNagc3R0gobGYTRsmNuIe//+MNq2\ndVJvoIxVUEr1yEkKucdbEARkZWWVeVBlgXvkVI/HV6gW51u1ON9l7+lTYMsW2XtTc9SqBTRokIiz\nZ+/ixo2rcHevj7ZtneDqyuPjyhvv46qVP9/lerNDfHx8iStmjDHGCnP7tuxNDXmfD9e0KdC+PaCh\nYYdPPrFDZKSEGxaMFYPftcoYY0xliIATJ4CjR2X/DwCamkDnzkCDBuqNjTF1KvMxclOnTlWqgoCA\ngBIvlDHG2Mfn/XvZpdQjR3IbccbGwNCh3IhjrLQK7ZGrVq0arl69WuTMRIRGjRrhxYsX5RLch+Ae\nOdXj8RWqxflWLc73h/n3X+DPP2X/zWFvD3z1FWBgoHgezrlqcb5Vq9zHyL19+xbOzs7FVqCjo1Pi\nhTLGGPt4xMbKxsO9f59b9umnQLt2/L5Uxj4Uj5FjjDFWLoiAY8eAvA+t19ICvviC35fKWH7letcq\nY4wxVhLv3sl64W7fzi0zMQF69859Phxj7MNxpzYrMx/yrjhWcpxv1eJ8K+/JEyA0VL4R5+gIfP11\nyRpxnHPV4nyrVlnlu0r3yAUGBorvMGOMMVb+bt4Edu2SvbEhR8uWQNu2PB6OMUUiIyM/qFHHY+QY\nY4x9sOxs2bPhTpzILdPSArp0AerWVV9cjFUW5TpG7vHjx9DT04OhoSEyMzOxbt06aGhoYODAgYW+\nvosxxtjHIS0N2LEDiIvLLTM1Bfr0ASwt1RcXYx8DpVphnTt3Rtz/H6E///wz5s+fj4ULF+KHH34o\n1+BY5cLjK1SL861anG/FHj+WjYfL24hzdpaNh/vQRhznXLU436ql0jFyd+7cQYP/f+z2hg0bcPr0\naRgaGsLd3R2LFi0qk0AYY4xVLjExQFgYkJGRW9a6NeDry+PhGFMVpcbIWVhYICkpCXfu3EGfPn0Q\nExODrKwsGBsb482bN6qIs8R4jBxjjJWP7Gzg8GHg1KncMm1toOv/sXfnUVFe9//A38MmIKsiCAoi\nIsjiGqNxx92qcRfFxrgmpk1y0qRpe35JjBjPt/n22yZdkraJGmPU1jXuSdQojkvc9wgIArLjwr4v\ns/z+eMIzjFtmhpnnmRner3M8Ze4Iz6efM7Uf7v3ce2cA0dHyxUVkyyzaIzdp0iTEx8ejpKQE8+bN\nAwCkpKSga9euRj+QiIhsV22t0A+Xmakb69hROB/O31++uIjaKoMmv9evX48pU6Zg+fLleOeddwAA\nJSUlSExMtGRsZGPYXyEt5ltazDdw9y6wdq1+ERcRAbz0kmWKOOZcWsy3tCTtkXN1dcWKFSv0xng2\nGxFR2/Hjj8D+/fr9cKNGAXFxgEIhW1hEbd4Te+QWLlyo/xd/+l+qVqsVvwaATZs2WTA807FHjoio\n9TQa4PvvgbNndWPt2gEzZwK9eskXF5G9MbVueeLSao8ePRAeHo7w8HD4+Phg7969UKvVCA4Ohlqt\nxr59++Dj49OqoImIyHrV1ACbN+sXcX5+wlIqizgi62DQrtUJEyZg5cqVGDFihDh2+vRpfPDBBzhy\n5IhFAzQVZ+Skp1QqueQuIeZbWm0t30VFwLZtQEWFbqxXL2Emrl07aWJoazmXG/MtrYfzbdFdq+fO\nncNzzz2nNzZ48GCcbflrGhER2YXr14EDBwCVSnitUAi9cCNHsh+OyNoYNCM3atQoPPvss1izZg3c\n3NxQW1uLVatW4fz58zh58qQUcRqNM3JERMZRq4EjR4Dz53Vj7doBs2cLu1OJyHIsOiO3ceNGLFiw\nAF5eXvD19UVZWRkGDhyI//73v0Y/kIiIrE91NbBzJ5CToxvr1Em4L7VjR/niIqKnM+gcue7du+Ps\n2bPIzMzE/v37kZGRgbNnz6J79+6Wjo9sCM8gkhbzLS17zndBgXA+XMsiLjoaWL5c3iLOnnNujZhv\naZkr30bdhufq6gp/f3+o1WpkZWUhKyvLLEEYo7KyEoMGDYKnpydSUlIkfz4RkT25ehX48kugslJ4\nrVAAY8cCc+dKt6mBiExnUI/coUOHsGzZMhQVFel/s0IBtVptseAeR6VSoby8HL/73e/w9ttvIyYm\n5rF/jz1yRERPplYDhw4BFy/qxlxdgTlzgPBw+eIiaqvMfo5cS7/+9a+xcuVKVFdXQ6PRiH+kLuIA\nwMnJCX5+fpI/l4jIXlRXA199pV/EBQQAL7/MIo7I1hhUyJWXl2PFihVwd3e3dDxkw9hfIS3mW1r2\nku+8PODzz4HcXN1YTAywbBnQoYN8cT2OveTcVjDf0pK0R27ZsmXYsGGDWR7Y7NNPP8XAgQPh6uqK\nJUuW6L1XWlqKmTNnwsPDA6Ghodi6detjf4aCBxoRERns8mVg40agqkp4rVAA48cLy6kuLrKGRkQm\nMqhHbvjw4bhw4QK6deuGzp07675ZoTD5HLk9e/bAwcEBhw8fRl1dHb788kvxvYSEBADAF198gatX\nr2LKlCk4c+YMoqOjxb+zZMkS9sgRERlApQK++04o5Jq5uQkbGsLC5IuLiHQseo7c8uXLsXz58sc+\n1FQzZ84EAFy6dAn5+fnieE1NDXbv3o3k5GS4u7tj2LBhmD59OjZv3owPP/wQADB58mRcv34daWlp\nWLFiBRYtWvTYZyxevBihoaEAAB8fH/Tr10+8DqN5SpOv+Zqv+dqeX1dWAmvWKPHgARAaKrxfWanE\nM88AYWHyx8fXfN1WXzd/nZ2djdYwaEbOkt577z0UFBSIM3JXr17F8OHDUVNTI/6djz/+GEqlEvv3\n7zf453JGTnpKpVL8oJLlMd/SssV85+YCO3YImxua9ekDPP884OwsX1yGssWc2zLmW1oP59uiu1a1\nWi02bNiA0aNHIyIiAmPGjMGGDRvMUig9PKtXXV0NLy8vvTFPT09UNTd1EBHRU2m1wo7UjRt1RZyD\nAzBpknDpvS0UcURkGIOWVv/4xz9i06ZN+O1vf4uQkBDk5ubiz3/+MwoLC/Hee++1KoCHi0EPDw9U\nNp9M+ZOKigp4enq26jlkefxNTlrMt7RsJd8qFfDNN8JBv83c3YV+OFu7jMdWcm4vmG9pmSvfBhVy\n69atw4kTJ9CtWzdxbOLEiRgxYkSrC7mHZ+QiIiKgUqmQkZGB8J8ONLp+/TpiY2ON/tmJiYmIi4vj\nh5OI2oSKCmD7dqCwUDcWFATMmwd4e8sXFxE9mVKp1OubM5ZBS6u1tbWPHMLbsWNH1NfXm/xgtVqN\n+vp6qFQqqNVqNDQ0QK1Wo3379pg1axbef/991NbW4vTp0zhw4AAWLlxo9DOaCzmSRms+iGQ85lta\n1p7v7GzhvtSWRVzfvsCSJbZbxFl7zu0N8y2tlpsgEhMTTf45BhVykyZNwgsvvIBbt26hrq4Oqamp\nePHFFzFx4kSTH7xmzRq4u7vjT3/6E7Zs2QI3Nzf8z//8DwDgX//6F+rq6uDv748XXngBn332GaKi\nokx+FhGRvdJqgXPngE2bgOY9Yg4OwOTJwIwZ7IcjsncG7VqtqKjA66+/ju3bt6OpqQnOzs6Ij4/H\nJ598Ah8fHyniNJpCocCqVau4tEpEdqupCTh4ELh+XTfWvj0QHw+06IQhIivWvLS6evVqkzaRGnX8\niFqtRnFxMfz8/ODo6Gj0w6TE40eIyJ6Vlwv9cEVFurEuXYR+uIc2/hORDbDo8SNfffUVrl+/DkdH\nRwQEBMDR0RHXr1/H5s2bjX4g2S/2V0iL+ZaWNeU7K0voh2tZxA0YIPTD2VMRZ005bwuYb2mZK98G\nFXIrV65EcHCw3ljXrl3x7rvvmiUIIiL6eVotcOYMsHkzUFsrjDk6AlOnCof8Ohl0DgER2RODllZ9\nfX1RXFyst5yqUqnQsWNHVFRUWDRAU3FplYjsSWMjsH8/cPOmbszDQ1hKfej3bCKyQRZdWo2KisKu\nXbv0xvbs2WP1O0kTExM5VUxENq+sDPjiC/0iLjgYWLGCRRyRrVMqla06fsSgGbnTp09j8uTJGD9+\nPMLCwpCZmYmjR4/i22+/xfDhw01+uCVxRk56vKdPWsy3tOTKd2YmsGsXUFenGxs4EPjFL4RlVXvG\nz7i0mG9pSXrX6vDhw/Hjjz9i4MCBqK2txaBBg5CcnGy1RRwRka3TaoHTp4EtW3RFnKMjMG2a0BNn\n70UcERnG6ONH7t27h6CgIEvGZBackSMiW9XYCOzdC6Sk6Ma8vITz4bp2lS8uIrIci87IlZWVYcGC\nBXBzcxPvP92/f3+r71klIiJ9paXA+vX6RVxICPDyyyziiOhRBhVyr7zyCry8vJCTk4N27doBAIYM\nGYJt27ZZNLjW4mYHaTHX0mK+pSVFvm/fFs6Hu39fNzZoELBokbBDta3hZ1xazLe0mvPd2s0OBp06\ndOzYMRQVFcG5xaV9nTp1wv2W/9pYodYkhohIKlotcOoUcPy48DUgnAk3dSrQr5+8sRGRZTVfJbp6\n9WqTvt+gHrnw8HCcPHkSQUFB8PX1RVlZGXJzczFhwgTcunXLpAdbGnvkiMgWNDQI/XCpqboxb2/h\nfDgbaEcmIjOxaI/c8uXLMWfOHCQlJUGj0eDs2bNYtGgRVqxYYfQDiYhIUFws9MO1LOJCQ4V+OBZx\nRGQIgwq5P/zhD5g3bx5ee+01NDU1YcmSJZg+fTp+85vfWDo+siHsr5AW8y0tc+c7LQ1Ytw548EA3\nNmQI8OKLQPv2Zn2UzeJnXFrMt7TMlW+DeuQUCgXeeOMNvPHGG2Z5KBFRW6XVAidOAC3/DXdyEs6H\n69NHtrCIyEYZ1COXlJSE0NBQhIWFoaioCH/4wx/g6OiIDz/8EJ07d5YiTqMpFAqsWrVKbCIkIpJb\nfT2wZ48wG9fMx0fohwsMlC8uIpKPUqmEUqnE6tWrTeqRM6iQ69WrF44cOYKQkBAkJCRAoVDA1dUV\nxcXF2L9/v0mBWxo3OxCRNXnwANi2DSgp0Y2FhQFz5gDu7vLFRUTWwaKbHQoLCxESEoKmpiYcPnwY\nn3/+OT777DP88MMPRj+Q7Bf7K6TFfEurNflOTRX64VoWccOGAS+8wCLuafgZlxbzLS1Je+S8vLxw\n9+5dJCcnIyYmBp6enmhoaEBTU5NZgiAiskcajdALd/KkbszZGZg+HYiNlS0sIrIjBi2t/ulPf8I/\n//lPNDQ04G9/+xsSEhKQlJSE//f//h/Onz8vRZxG49IqEcmprg7YvVu4raGZry8wfz4QECBfXERk\nnUytWwwq5AAgLS0Njo6O4l2r6enpaGhoQO/evY1+qBRYyBGRXO7fF/rhSkt1Y+HhwOzZgJubfHER\nPU5aRhqOXj6KRk0jXBxcMO6ZcYgMj5Q7rDbHoj1yABAZGSkWcQAQERFhtUUcyYP9FdJivqVlaL6T\nk4VDflsWcSNGAAsWsIgzFj/jlpeWkYaNxzfisutlHEg/gPv+97Hx+EakZaT9/DdTq1i8R65Xr17i\n9VvBwcGP/TsKhQK5ublmCcQSEhMTefwIEUlCowGSkoDTp3VjLi7AjBlAdLR8cRE9zeGLh5Hpk4n7\nZfdRXleO/Mp8BPcMxrErxzgrJ5Hm40dM9cSl1VOnTmHEiBHiQ57EWoskLq0SkVTq6oBdu4DMTN1Y\nx47C+XD+/vLFRfQ0lQ2VWP735bjvf18c83H1Qd+AvvC954vfzOftTVIytW554oxccxEHWG+xRkQk\nt7t3ge3bgbIy3VhEBDBrFuDqKl9cRE+TX5mPbTe3obqxWhwL9AhEz449oVAo4OLgImN0ZIwnFnIr\nV658YnXYPK5QKPDBBx9YNECyHUqlkkW/hJhvaT0u3z/+COzfD7Q8iWnUKCAuDlAoJA3PLvEzbhnX\n7l7DgbQDUGvVCAsLw43UG4h8NhKNmY1w8HNAw+0GjB09Vu4w7Z65Pt9PLOTy8vKgeMq/RM2FHBFR\nW6PRAEePAmfO6MbatQNmzgR69ZIvLqKn0Wg1+D7ze5zNPyuOhXQLwYxeM5CekY6UshT43/fH2NFj\n2R9nQww+fsTWsEeOiCyhthbYuRO4c0c35ucnnA/n5ydfXERPU9dUh10pu5BZpmvk9G/vj4TYBPi6\n+coYGTUze49cVlaWQT8gLCzM6IcSEdmioiLhfLiKCt1YZKTQD9eunXxxET1NcW0xtv64FSV1ujvi\nevn1wsxeM9HOiR9cW/fEGTkHh58/Yk6hUECtVps9KHPgjJz02M8iLeZbGmlpOTh6NBMnT95ATU0f\nhIb2gJ9fNygUQi/cyJHsh7MUfsZb73bJbexK2YUGdYM4NqrbKMSFxj3SHsV8S+vhfJt9Rk6j0ZgU\nGBGRvUhLy8GXX2YgL28s0tIc4OMTh2vXjmHQIOCVV7ohIkLuCIkeT6vV4kzeGRzNOgothOLA2cEZ\nM3rNQIx/jMzRkTnZdY/cqlWreCAwEZnsL39JwsmTY1BZqRtzdwfi4pLw+9+PkS8woqdoUjfhQPoB\n3Lh3QxzzbueN+bHzEegZKGNk9DjNBwKvXr3avHetTpw4EYcPHwagf6ac3jcrFDh58qTRD5UCl1aJ\nqDVu3wbefFOJ6uo4cczPT9iV6uenxG9+E/fE7yWSS2VDJbbf3I6CqgJxLMQ7BPNi5qG9S3sZI6Of\nY/al1RdffFH8etmyZU98KFEz9ldIi/m2jJZXbTW3mCgUgJubEjExcVAoABcXtp5IgZ9x4+RX5mP7\nze2oaqwSxwYEDsCUnlPg6OD4s9/PfEvL4ufI/fKXvxS/Xrx4casfRERk7aqqhKu2cnKE12FhPZCa\negx9+oxFWZlQ0DU0HMPYseHyBkr0kOt3r+NA+gGoNCoAgIPCAZPCJ+HZoGc56WLnDO6RO3nyJK5e\nvYqamhoAugOB33nnHYsGaCourRKRMbKygK+/Bn76Jw4AEB4OxMbm4MyZTDQ2OsDFRYOxY3sgMrKb\nfIEStaDRanA06yjO5OlOp3ZzckN8TDy6+3aXMTIyltmXVlt6/fXXsWPHDowYMQJubm5GP4SIyFpp\nNMCpU4BSCTT/G6pQAKNHAyNGAApFN/Trx8KNrE+9qh67UnYhozRDHPNv74/5sfPRwa2DjJGRlAya\nkfP19UVycjKCgoKkiMksOCMnPfZXSIv5br2aGmD3biBTd9g9PDyA2bOB7g9NZjDf0mPOn+xxh/xG\ndozErKhZJh/yy3xLy+LnyLUUHBwMFxcXo384EZG1yskR+uGqdH3hCA0F5swRijkia5VRmoFdKbtQ\nr6oXx0Z2G4nRoaPZD9cGGTQjd/HiRfzxj3/EggULEBAQoPfeyJEjLRZca3BGjogeR6sVLrs/dkxY\nVm02cqRwU4MBl9oQyUKr1eJs/ll8n/m93iG/03tNR6x/rMzRUWtZdEbu8uXL+Pbbb3Hq1KlHeuTy\n8vKMfigRkRzq6oC9e4G0NN2Yu7twV2o4N6KSFVNpVDiQdgDX710Xx7zaeSEhNoGH/LZxBv3u+e67\n7+LgwYMoLi5GXl6e3h+iZkqlUu4Q2hTm2zgFBcDnn+sXccHBwIoVhhVxzLf0mHNBVUMVNl7bqFfE\nBXsF4+VnXjZrEcd8S8tc+TZoRq59+/YYNWqUWR5IRCQlrRa4cAE4cgRQq3XjQ4cCY8cCjj9/TiqR\nbAoqC7Dt5rZHDvmd3HMynBwM+r9wsnMG9cht3LgRFy5cwMqVKx/pkXOw0oYS9sgRUX09sH8/kJKi\nG3N1BWbMEK7aIrJmN+7dwP60/XqH/E7sMRGDugzipgY7ZGrdYlAh96RiTaFQQN3yV1wrolAosGrV\nKsTFxXE7NVEbdPcusGMHUFqqGwsKAubOBXx95YuL6OdotBocyzqGH/J+EMfcnNwwN2YuwnzDZIyM\nLEGpVEKpVGL16tWWK+Sys7Of+F5oaKjRD5UCZ+SkxzOIpMV8P55WC1y5Anz3HaBS6cYHDQImTACc\nTFyNYr6l1xZzXq+qx9cpX+N26W1xrJN7JyT0TrD4Ib9tMd9ykvQcOWst1oiIWmpsBA4eBG7c0I25\nuADTpgGxPJ2BrFxJbQm23tyK4tpicSyiYwRmR802+ZBfsn8G37VqazgjR9S2PHggLKU+eKAbCwgQ\nllL9/OSLi8gQmaWZ2JmyU++Q3xEhIzC6+2g4KKyzF53My6IzckRE1uzGDeDAAaCpSTc2YADwi18A\nzs7yxUX0c7RaLc7ln8ORzCPiIb9ODk6YHjkdvQN6yxwd2QKW+WQ2PINIWsy3ULgdOCDcl9pcxDk7\nC7tSp00zbxHHfEvP3nOu0qiwL20fDmceFos4r3ZeWNp/qSxFnL3n29pIeo4cEZG1KSkBdu4Udqc2\n8/MD4uMBf3/54iIyRFVDFbYnb0d+Zb44FuwVjHmx8+Dhwst+yXAG9chlZWXh3XffxbVr11BdXa37\nZoUCubm5Fg3QVOyRI7JfycnC+XANDbqx3r2B558XNjcQWbPCqkJsu7kNlQ2V4li/zv0wNWIqD/lt\nwyzaI7dgwQKEh4fj448/fuSuVSIiqajVwg0N58/rxhwdhV64Z54BeEYqWbsf7/2IfWn7xEN+FVBg\nYvhEDO4ymIf8kkkMmpHz8vJCWVkZHG3oLhvOyEmPZxBJq63lu7xcWEotKNCN+foKS6mBEtwZ3tby\nbQ3sKecarQZJd5JwOve0OObq5Iq50XPRo0MPGSPTsad82wJJz5EbOXIkrl69ioEDBxr9ACKi1kpP\nB/bsAerqdGNRUcD06cKVW0TWrEHVgK9Tv0Z6Sbo45ufuh4TYBHR07yhjZGQPDJqRe/XVV7F9+3bM\nmjVL765VhUKBDz74wKIBmoozckS2T60GkpKAH3Q3FcHBQbihYfBgLqWS9SutK8XWH7fiQa3ugMOI\njhGYFTULrk78LYR0LDojV1NTg6lTp6KpqQn5+cIOG61Wy/V8IrKYykpg1y6g5X4qb2/hgN+uXeWL\ni8hQWWVZ2Jm8E3Uq3VTy8JDhGNN9DA/5JbPhzQ5kNuyvkJY95zszE/j6a6C2VjfWsycwcybg7i5P\nTPacb2tlqznXarU4X3AehzMO6x3yOy1yGvoE9JE5uiez1XzbKov3yGVnZ4t3rGZlZT3xB4SFhRn9\nUCKix9FogBMngJMngeZ/zxQKYOxYYNgwLqWS9VNpVPgm/RtcvXtVHPN08cT82Pno4tVFxsjIXj1x\nRs7T0xNVVVUAAAeHx08BKxQKqNVqy0XXCpyRI7It1dXCDQ0tf2/09ARmzwZ++p2SyKpVN1Zj+83t\nyKvME8e6enXFvJh58GznKWNkZAtMrVtscmn1D3/4A86ePYvQ0FBs2LABTk6PTiyykCOyHTk5Qj/c\nT787AgDCwoBZswAPHnJPNoCH/FJrmVq32Fy35fXr11FYWIiTJ0+iV69e2LVrl9wh0U94T5+0Thpw\nnwAAIABJREFU7CHfWi1w+jSwcaOuiFMogFGjgBdesK4izh7ybWtsJec379/EhqsbxCJOAQUm9piI\n6ZHTbaqIs5V82wtz5dvmCrmzZ89i4sSJAIBJkybhh5bnEhCRzaitBf77X+DoUV0/nLu7UMCNHi0c\nM0JkzbRaLY5lHcOulF3iTQ2uTq74ZZ9fYkjwEJ7sQJKwnV8VflJWVobAn45x9/LyQmlpqcwRUTPu\ndpKWLec7P1+4paGiQjcWEgLMmQN4eckX19PYcr5tlTXnvEHVgN2pu5FWkiaO2fohv9acb3tkrnzL\n9jvvp59+ioEDB8LV1RVLlizRe6+0tBQzZ86Eh4cHQkNDsXXrVvE9Hx8fVFYK09cVFRXo0KGDpHET\nkem0WuDcOWDDBv0ibtgwYNEi6y3iiFoqrSvF+ivr9Yq4nh16YvmA5TZbxJHtMrqQ02g0en9M1aVL\nF6xcuRJLly595L1XX30Vrq6uuH//Pv7zn//gV7/6FVJSUgAAQ4cOxdGjRwEAhw8fxvDhw02OgcyL\n/RXSsrV819cDO3YAhw4Jx4wAgJsbkJAAjB8PWPtVzraWb3tgjTnPKsvCusvr9G5qGBY8DAm9E2z+\npgZrzLc9k7RH7vLlyxgyZAjc3d3h5OQk/nF2djb5wTNnzsT06dPRsaP+by81NTXYvXs31qxZA3d3\ndwwbNgzTp0/H5s2bAQB9+/ZFQEAARo4cidTUVMyePdvkGIhIGkVFwOefA6mpurEuXYAVK4DISPni\nIjKUVqvF+fzz2HJji3hTg5ODE2ZFzcL4HuN5UwPJxqAeuUWLFmHatGn44osv4G7mY9Uf3mqbnp4O\nJycnhIeHi2N9+/bVq1z/7//+z6CfvXjxYvFQYx8fH/Tr109ck27+eXxt3tfNrCUee3/dzFriefj1\nqFFxuHwZ+Pe/lVCrgdBQ4X03NyXCwgAfH+uK19bzzdeWeX0s6RjO5Z9DU0gTACD7WjbcnNzw3ovv\noYtXF9nj42vbfA0AiYmJyM7ORmsYdI6cl5cXKioqLLIDZ+XKlcjPz8eXX34JADh16hTi4+NRVFQk\n/p1169bhv//9L44fP27wz+U5ckTyamwEDhwAfvxRN9auHTBtGhATI19cRMaoaazB9uTtyK3IFce6\neHbB/Nj5POSXzMqi58jNnDkThw8fNvqHG+LhoD08PMTNDM0qKirg6cn/wVi7lr9lkOVZc77v3wfW\nrtUv4jp3Bl5+2XaLOGvOt72SO+dFVUVYe3mtXhHXN6AvlvRfYpdFnNz5bmvMlW+Dllbr6uowc+ZM\njBgxAgEBAeK4QqHApk2bWhXAw7N8ERERUKlUyMjIEJdXr1+/jtjYWKN/dmJiIuLi4sTpTCKyvGvX\ngG++AZqadGPPPANMmgS0oq2WSFLJ95Ox99ZeNGmED7ICCozvMR5DuvJ8ODIvpVLZqqLOoKXVxMTE\nx3+zQoFVq1aZ9GC1Wo2mpiasXr0aBQUFWLduHZycnODo6IiEhAQoFAqsX78eV65cwdSpU3H27FlE\nRUUZ/PO5tEokraYm4Ntvgau6u8Lh7AxMnQr07StfXETG0Gq1OJ59HCdzTopjrk6umB01Gz079pQx\nMrJ3NnfXamJiIj744INHxt5//32UlZVh6dKl+P777+Hn54f//d//xfz58436+SzkiKRTUiIcLXLv\nnm6sUydg7lzA31++uIiM0aBqwJ5be3Cr+JY41tGtIxJ6J8DP3U/GyKgtsHghd/z4cWzatAkFBQXo\n2rUrXnjhBYwZM8boB0qFhZz0lEoll7ElZC35vnkT2L9f2NzQrE8fYSbOxUW+uMzNWvLdlkiZ87K6\nMmy9uRX3a+6LY+EdwjEneo7Nnw9nKH7GpfVwvi262WH9+vWYN28eAgMDMWvWLHTu3BkLFizA2rVr\njX6glBITE9m8SWQhKpWwlLprl66Ic3ICnn8emDnTvoo4sm93yu5g7eW1ekXc0OChWNB7QZsp4kg+\nSqXyiS1shjBoRq5nz57YtWsX+rZodLlx4wZmzZqFjIwMkx9uSZyRI7KcsjLhrtTCQt1Yhw5AfLyw\nO5XIFmi1WlwsvIhDGYeg0QrXjTgqHDEtchr6dmZjJ0nLokurHTt2RFFREVxa/Ird0NCAoKAglJSU\nGP1QKbCQI7KMW7eAvXuFK7eaRUcL58O5cvKCbIRao8a3t7/F5aLL4piHiwfmx85HV6+uMkZGbZVF\nl1aHDRuGt956CzU1NQCA6upqvP322xg6dKjRDyT7xWVsaUmdb7UaOHIE2LZNV8Q5OgK/+IWwqcHe\nizh+vqVnqZzXNNZg0/VNekVcF88uePmZl9t0EcfPuLQkPUfus88+w/z58+Ht7Y0OHTqgtLQUQ4cO\nxdatW80ShKXwHDki86isFJZS8/J0Yz4+QgHXpYt8cREZ6271XWz9cSsqGirEsT4BffB8xPNwduRB\nhyQ9Sc6Ra5aXl4fCwkIEBQUhODjY5IdKgUurROaRkQHs3g3U1urGIiKEDQ1ubvLFRWSslAcp2JO6\nR++Q33Fh4zA0eCgP+SXZmb1HTqvVih9sjUbzxB/g4GDQ6qzkWMgRtY5GAyiVwKlTQPP/lBwcgLFj\ngaFDAf7/HtkKrVYLZbYSJ3JOiGPtHNthTvQcHvJLVsPsPXJeXl7i105OTo/948z7dqgF9ldIy5L5\nrq4GNm8GTp7UFXGensCiRcCwYW2ziOPnW3rmyHmjuhE7knfoFXEd3Dpg+YDlLOIews+4tCzeI5ec\nnCx+nZWVZZaHEZH1y84WzoarrtaNhYUBs2cD7dvLFhaR0crqyrDt5jbcq9FdOdLDtwfmRM+BmzP7\nAsg+GNQj95e//AVvv/32I+Mff/wx3nrrLYsE1lrN98ByswORYbRa4PRpIClJNwunUABxccCIEcKy\nKpGtyC7Pxo7kHaht0jV3Duk6BON7jIeDgh9msh7Nmx1Wr15tuXPkPD09UVVV9ci4r68vysrKjH6o\nFNgjR2S42lphQ0PL873btxdm4cLC5IuLyBQXCy7iu4zv9A75fT7yefTr3E/myIiezNS65anHjyQl\nJUGr1UKtViMpKUnvvczMTL0+OiLe0yctc+U7L084WqSyUjfWrRswZ47QF0cCfr6lZ2zO1Ro1vsv4\nDpcKL4ljHi4emBczD8He1n3SgjXgZ1xa5sr3Uwu5pUuXQqFQoKGhAcuWLRPHFQoFAgIC8Mknn7Q6\nACKSh1YLnDsHfP+9sEO12fDhwJgxXEol21LTWIOdKTuRXZ4tjgV5BmF+7Hx4teOkA9kvg5ZWFy5c\niM2bN0sRj9lwaZXoyerrhWu2bt3Sjbm5CWfDRUTIFxeRKe5V38PWm1tRXl8ujvX2741pkdN4yC/Z\nDIvetWqLWMgRPV5hobCU2rK9tWtXYSnVx0e+uIhMkfogFXtu7UGjuhGAcMjv2LCxGBY8jIf8kk2x\n6F2rFRUVePPNNzFgwAB069YNwcHBCA4ORkhIiNEPlFJiYiLPxZEQcy0tY/Ot1QIXLwJffKFfxD33\nHLBkCYu4n8PPt/SelvPmQ363J28Xi7h2ju2Q0DsBw0OGs4gzAT/j0mrOt1KpRGJiosk/x6C7Vl99\n9VXk5eXh/fffF5dZ//znP2P27NkmP1gKrUkMkT1paAAOHABu3tSNtWsHTJ8OREfLFxeRKRrVjdh7\nay9SHqSIYx3cOiAhNgGd2neSMTIi4zUfk7Z69WqTvt+gpdVOnTohNTUVfn5+8Pb2RkVFBQoKCvD8\n88/jypUrJj3Y0ri0SiS4dw/YsQMoKdGNBQYKF9536CBfXESmKK8vx9Yft+od8hvmG4a50XN5yC/Z\nNIscP9JMq9XC29sbgHCmXHl5OQIDA3H79m2jH0hE0rl6FfjmG0Cl0o0NHAhMmgQ4GfS/fiLrkVOe\ng+3J2/UO+X2u63OY0GMCD/mlNsugT36fPn1w8uRJAMDw4cPx6quv4pVXXkFkZKRFgyPbwv4KaT0t\n301Nwq7Ufft0RZyLCzBrFjB1Kos4U/DzLb2WOb9UeAlfXf9KLOIcFY6YHjkdk8InsYgzE37GpWXx\nu1ZbWrdunfj13//+d7zzzjuoqKjApk2bzBIEEZlPcbGwlHr/vm7M319YSu3E9iGyMWqNGocyDuFi\n4UVxjIf8EukY1CN3/vx5DB48+JHxCxcuYNCgQRYJrLXYI0dt0Y8/CpsaGht1Y337AlOmCDNyRLak\ntqkWO5J36B3yG+gRiPmx8+Ht6i1fYEQWYNEeuXHjxj32rtVJkyahtLTU6IdKJTExUdwNQmTPVCrg\n0CHgku5mIjg5CQVcv34AT2IgW5KWkYbdZ3bjQuEFNKgaEBYWBr8gP8T6x2J65HQe8kt2RalUtmqZ\n9amNBRqNBmq1Wvy65Z/bt2/DycobbZoLOZIG+yuk1Zzv0lLhbLiWRVzHjsDy5UD//izizIWfb2mk\nZaTho28+wlHtUdwpv4ParrW4nnIdPR16YnbUbBZxFsTPuLSa8x0XF2e5c+RaFmoPF20ODg549913\nTX4wEbVeaqqwoaG+XjcWEwNMmyacE0dkS7RaLf515F9I904HflphclQ4InZoLCruVvCQX6LHeGqP\nXHZ2NgBg5MiROHXqlLh2q1Ao0KlTJ7i7u0sSpCnYI0f2Ki0tB0eOZOLmTQfk5WkQFtYDfn7d4OgI\nTJwIPPssZ+HI9tSr6rE7dTc27duE+q7CbyauTq7o7d8b7V3aw+euD34z/zcyR0lkORbpkQsNDQUA\n5ObmmhQUEZlXWloO/vnPDGRnj0VlpTB27doxjBgBvPZaN3TpIm98RKZ4UPMA225uQ0ldCRx+6vjx\ndfVFdKdocSnVxYG7dYgex6Amt4ULFz4y1jzFzSNIqJlSqWRPogVVVAAffZSJ9PSxAIDyciV8fOIQ\nEDAWAQFJ6NKlm8wR2jd+vi0jrTgNu1N3o0HdAAAICwtDcX4xIp+NRM71HIT2C0XD7QaMHT1W5kjt\nHz/j0jJXvg0q5Hr06KE35Xf37l18/fXX+OUvf9nqAIjo6errgdOngXPngNxc/f1JPXoAXbsCBp7t\nTWQ1tFotTuScgDJbKY45OzjjlXGvwLnSGceuHENJaQn87/tj7OixiAznAfREj2PQOXKPc+nSJSQm\nJuLgwYPmjsks2CNHtk6lAi5eBE6eBOrqhLELF5JQWzsGHToAYWGAh4cw7u+fhF//eox8wRIZoUHV\ngN2pu5FWkiaO+bj6YH7sfHT26CxjZETyMbVuMbmQU6lU8PX1fez5ctaAhRzZKq1WONg3KQkoL9d/\nz8EhB0VFGQgI0C0zNTQcw+LF4YiM5NIqWb/i2mJsu7kNxbXF4liYbxjmRM+Bu7P1bqAjsjSLHgh8\n7NgxvW3fNTU12LZtG2JiYox+INkv9le0XlYW8P33QFGR/rivLzB2LBAT0w3p6cCxY0lISbmB6Og+\nGDuWRZwU+PluvYf74QBgaPBQjAsb99j7UplzaTHf0pK0R27ZsmV6hVz79u3Rr18/bN26tdUBWBJv\ndiBbcfeuUMBlZuqPu7sDo0YBAwcCjo7CWGRkN0RGdoNS6cDPNtkErVaLkzkncTz7uDjm7OCMaZHT\n0Dugt4yREcmvtTc7mLy0au24tEq2oLxcWEL98UdhSbWZszPw3HPAsGGAq6t88RG1VoOqAXtu7cGt\n4lviGPvhiB5l0aVVACgvL8c333yDwsJCBAUFYfLkyfD19TX6gUQkbF44dQo4fx746RY8AMJBvv37\nA3FxgJeXbOERmUVJbQm23tyq1w/X3ac75sbMZT8ckZkYdGZBUlISQkND8Y9//AMXL17EP/7xD4SG\nhuLo0aOWjo9sCO/p+3lNTcAPPwB//ztw5ox+ERcZCfz618L1WoYUccy3tJhv46SXpGPt5bV6RdyQ\nrkOwsO9Cg4s45lxazLe0zJVvg2bkXn31Vaxduxbx8fHi2M6dO/Haa6/h1q1bT/lOIgIAjQa4cQM4\nflw42Lelrl2B8eOBbtyvQHZAq9XiVO4pHL9zHNqfLkx1cnDCtMhp6BPQR+boiOyPQT1yPj4+KCkp\ngWNztzWApqYmdOrUCeUPn49gJdgjR9ZAqwUyMoCjR4F79/Tf69hR2IkaFcW7Uck+NKgasPfWXqQW\np4pj3u28MT92PgI9A2WMjMj6WbRHbuHChfj000/xxhtviGP//ve/H3t1FxEJCguFnah37uiPt28v\n9MANGKDbiUpk60pqS7Dt5jY8qH0gjnX36Y450XPQ3qW9jJER2TeDZuSGDRuGCxcuwN/fH126dEFB\nQQHu37+PwYMHi8eSKBQKnDx50uIBG4ozctLjGUSCsjLg2DHg5k39cRcXYMgQYOhQoF271j+H+ZYW\n8/1kt0tu4+vUr1GvqhfHnuv6HCb0mPDY8+EMxZxLi/mW1sP5tuiM3EsvvYSXXnrpqX9HwbUhauNq\na4XrtC5e1N/E4OAgzL7Fxemu1CKyB1qtFqdzTyPpTpJeP9zzEc+jb+e+MkdH1DbwHDmiVmpqEi60\nP30aaGjQfy8qSuiD8/OTJzYiS2lUN2Lvrb1IeZAijnm388a82HkI8gySMTIi22Txc+ROnjyJq1ev\noqamBoDwm5hCocA777xj9EOJ7IFGA1y7JuxEffjK4ZAQYSdqcLA8sRFZUmldKbbd3Ib7NffFsVCf\nUMyNnst+OCKJGdS88Prrr2Pu3Lk4deoUUlNTkZqailu3biE1NfXnv5najLZyBpFWC6SnA//+N7B/\nv34R5+cHzJ8PLFli+SKureTbWjDfgozSDKy9vFaviBvcZTAW9llo9iKOOZcW8y0tSc+R27JlC5KT\nkxEUxOlyatvy84WdqDk5+uMeHsDo0cKtDA6m93YTWS2tVosf8n7Asaxjev1wUyOmol/nfjJHR9R2\nGdQj16dPHyQlJcHPhhp9FAoFVq1ahbi4OO7CoVYrKRF2oqak6I+7uAj3oQ4ZInxNZI8a1Y3Yd2sf\nkh8ki2Ne7bwwL2Yeunh1kTEyItunVCqhVCqxevVqk3rkDCrkLl68iD/+8Y9YsGABAgIC9N4bOXKk\n0Q+VAjc7kDnU1AAnTgCXLgk9cc0cHICBA4FRo4Rz4YjsVVldGbbd3IZ7NboTrbt5d8PcmLnwcOE2\nbCJzsehmh8uXL+Pbb7/FqVOn4ObmpvdeXl6e0Q8l+2RPZxA1NgJnzwr3ojY26r8XEyPsRO3QQZ7Y\nmtlTvm1BW8x3ZmkmdqXsQp2qThwb1GUQJvaYCEcHy59m3RZzLifmW1rmyrdBhdy7776LgwcPYvz4\n8a1+IJE102iAK1cApRKortZ/LzRU2InahStJZOe0Wi3O5J3B0ayjYj+co8IRUyOmon9gf5mjI6KW\nDFpaDQkJQUZGBlxsqAmIS6tkDK0WSEsT7kQtLtZ/z98fGDcO6NmTd6KS/WtUN2J/2n7cvK+7moT9\ncESWZ2rdYlAht3HjRly4cAErV658pEfOwUq36LGQI0Pl5QFHjgj/2ZKXl7ATtW9f7kSltuFx/XAh\n3iGIj4lnPxyRhVm0kHtSsaZQKKBueReRFWEhJz1b668oLhZm4G7d0h9v1w4YPhx47jnA2Vme2Axh\na/m2dfae76yyLOxM3qnXD/ds0LOYFD5Jkn64x7H3nFsb5ltakt61mpWVZfQPJrJWVVXCTtQrV/R3\nojo6As8+C4wcCbi7yxcfkZS0Wi3O5p/F95nf6/XDTYmYggGBA2SOjoh+jlF3rWo0Gty7dw8BAQFW\nu6TajDNy9LCGBuDMGeFPU5P+e717A2PGAL6+8sRGJIcmdRP2p+3Hj/d/FMc8XTwxL3Yeunp1lTEy\norbHojNylZWVeO2117Bt2zaoVCo4OTlh/vz5+OSTT+Dt7W30Q4mkpFYDly8Ls3A/XRUs6t5d2InK\nS0uorSmvL8e2m9twt/quOBbsFYz4mHh4tvOUMTIiMobBd63W1NTg5s2bqK2tFf/z9ddft3R8ZEOs\n7Z4+rVa4ieGf/wS+/Va/iAsIAF54AXjxRdst4qwt3/bOnvKdVZaFtZfX6hVxA4MGYnG/xVZVxNlT\nzm0B8y0tSe9aPXToELKystD+pyPsIyIisHHjRoSFhZklCCJzy8kR7kTNz9cf9/YWllB79+ZOVGp7\ntFotzuWfw5HMI3r9cJN7TsYzQc/IHB0RmcKgHrnQ0FAolUqEhoaKY9nZ2Rg5ciRyc3MtGZ/J2CPX\nNj14IOxETUvTH3d1FTYxDBoEOBn06wuRfWlSN+FA+gHcuHdDHPN08UR8TDyCvYNljIyIAAv3yC1f\nvhzjx4/Hb3/7W3Tr1g3Z2dn461//ipdeesnoBxJZQmWlcBvD1avCkmozJyeheBsxAnjodjmiNqO8\nvhzbb25HUXWROMZ+OCL7YNCMnEajwcaNG/Gf//wHRUVFCAoKQkJCApYuXQqFlR51zxk56clxBlF9\nvXAf6rlz+jtRFQqgTx/hQF8fH0lDkgzPfJKWreb7Ttkd7EzZidqmWnHsmcBn8Iuev4CTg3VPT9tq\nzm0V8y0tSc+Rc3BwwNKlS7F06VKjH2BulZWVGDduHFJTU3H+/HlER0fLHRLJQK0GLl4ETp4Eamv1\n3wsPF67U6txZntiIrIFWq8X5gvM4knkEGq1wYKKjwhG/6PkLDAwaKHN0RGQuBs3Ivf7660hISMDQ\noUPFsTNnzmDHjh3429/+ZtEAH6ZSqVBeXo7f/e53ePvttxETE/PYv8cZOfuk1QLJycCxY0BZmf57\ngYHCUSLcg0NtXZO6CQfTD+L6vevimIeLB+Jj4hHiHSJjZET0JBa9osvPzw8FBQVo166dOFZfX4/g\n4GA8ePDA6Ieaw5IlS1jItTF37gg7UQsL9cd9fICxY4HYWF5qT1RRX4FtN7fp9cN19eqK+Jh4eLXz\nkjEyInoaU+sWgw5gcHBwgKblXUYQ+uZYKFFLljqD6N494D//Ab76Sr+Ic3MDJk4EXntNOE6krRVx\nPPNJWraQ7+zybKy9vFaviBsQOACL+y22ySLOFnJuT5hvaZkr3wYVcsOHD8d7770nFnNqtRqrVq3C\niBEjjHrYp59+ioEDB8LV1RVLlizRe6+0tBQzZ86Eh4cHQkNDsXXrVvG9v/71rxg9ejQ++ugjve+x\n1o0WZB4VFcDevcBnnwG3b+vGnZyES+3feAMYMoTHiRBptVqczz+PTdc3oaZJOPnaQeGAKT2n4PmI\n561+UwMRmc6gpdW8vDxMnToVRUVF6NatG3JzcxEYGIgDBw4gONjw84f27NkDBwcHHD58GHV1dfjy\nyy/F9xISEgAAX3zxBa5evYopU6bgzJkzT9zMwKVV+1VXB5w+DZw/D6hUunGFAujXT9iJ6mV7kwtE\nFqHSqHAw/SCu3b0mjrV3bo/4mHh08+kmY2REZAyL9sgBwizchQsXkJeXh+DgYAwePBgOJh6Nv3Ll\nSuTn54uFXE1NDTp06IDk5GSEh4cDABYtWoSgoCB8+OGHj3z/5MmTcf36dXTr1g0rVqzAokWLHv0v\nxkLO5qhUwIULwKlTQjHXUkSEsBPV31+e2IisUUV9BbYnb0dhla7noItnF8yLnWeTS6lEbZlFjx8B\nAEdHRwwZMgRDhgwx+iEPezjQ9PR0ODk5iUUcAPTt2/eJ68fffvutQc9ZvHixeBuFj48P+vXrJ57Z\n0vyz+dp8r69du4bf/OY3Rn+/Vgts2KDE1auAn5/wfna28P6wYXEYP154nZIC+Ptbz39fuV+bmm++\nto98362+izzfPNQ01SD7WjYAYMakGZgaMRWnT56WPT5zvG4es5Z47P1185i1xGPvr69du4by8nJk\nZ2ejNQyekTOnh2fkTp06hfj4eBQV6Rp0161bh//+9784fvy4Sc/gjJz0lEql+EE1VGamsBP17l39\n8Q4dhJ2o0dFtbxODoUzJN5nOWvKt1WpxsfAiDmUcEs+Hc1A4YFL4JDwb9Kxd9Q5bS87bCuZbWg/n\n2+Izcub0cKAeHh6orKzUG6uoqICnJ6+OsSXG/ANQVCTciZqZqT/u7g6MGgUMHAg4Opo3PnvDf3Cl\nZQ35VmlU+Cb9G1y9e1Ucs+d+OGvIeVvCfEvLXPn+2UJOq9Xizp07CAkJgZOZtgc+/BtjREQEVCoV\nMjIyxOXV69evIzY2tlXPSUxMRFxcHD+cVqS8HEhKAm7c0B93dhZ2oA4bBrQ4rpCIflLZUIntN7ej\noKpAHAvyDMK8mHnwdvWWMTIiag2lUqm3vG2sn11a1Wq1aN++Paqrq03e3NBMrVajqakJq1evRkFB\nAdatWwcnJyc4OjoiISEBCoUC69evx5UrVzB16lScPXsWUVFRJj2LS6vSe9q0fG2tsInhwgXheq1m\nCgUwYAAQFwdwAtY4XAaRlpz5zq3Ixfab28WjRQCgX+d+mNJzCpwdnWWJSQr8jEuL+ZaWZEurCoUC\n/fv3R1pamslFVbM1a9bggw8+EF9v2bIFiYmJeP/99/Gvf/0LS5cuhb+/P/z8/PDZZ5+1+nkkv6Ym\n3U7U+nr993r1EvrgOnWSJzYia6fVanGp8BK+y/hOrx9uYo+JGNRlkF31wxGRaQza7PDee+9hy5Yt\nWLx4MYKDg8WqUaFQYOnSpVLEaTTOyMlLoxGWT5OSgIfaH9G1KzBhAhDCKx+JnkilUeHb29/iStEV\ncczd2R3xMfEI9QmVLzAisgiLbnY4ffo0QkNDceLEiUfes9ZCDmCPnBy0WiAjQ9jIcO+e/nsdOwpn\nwfXqxZ2oRE9T2VCJHck7kF+ZL44FegRifux89sMR2RmL98jZKs7ISSctLQdHj2bi4sUbUKv7wNu7\nB/z8dDvoPDyEHrj+/bkT1ZzYzyItqfKdW5GLHck7UN1YLY71CeiD5yOet+t+uMfhZ1xazLe0JD9+\npKSkBN988w3u3r2L3//+9ygoKIBWq0XXrl2NfijZj7S0HHz2WQYKCsYiPd0BPj5xuHPnGPr1A4KC\numHoUGDoUMDFRe5IiazfpcJL+O72d1BrhR1BDgoHTOgxAYO7DGY/HBE9lkEzcidOnMDVCttnAAAg\nAElEQVTs2bMxcOBA/PDDD6iqqoJSqcRHH32EAwcOSBGn0TgjJ43Vq5OgVI5By1QrFECvXkn485/H\nwMNDvtiIbIVKo8J3t7/D5aLL4pi7szvmRs9Fd9/uMkZGRFKx6IzcG2+8gW3btmHcuHHw9fUFADz3\n3HM4f/680Q8k++Lh4QAXF6ChQXjdqRPQvTsQFOTAIo7IAFUNVdiRvAN5lXniWGePzpgfOx8+rj4y\nRkZEtsCgg+FycnIwbtw4vTFnZ2eoWx4IZoUSExNb1UBIP8/VVYPQUMDbG/DzUyImRridwcVFI3do\ndo+fbWlZIt95FXn4/PLnekVcb//eWNZ/GYs48DMuNeZbWs35ViqVSExMNPnnGFTIRUVF4dChQ3pj\nx44dQ+/evU1+sBSad62S5Ywb1wMdOgg9cc0zcA0NxzB2bA95AyOycpcLL2PjtY3ipgYFFJjYYyJm\nRc1qc5saiNqyuLi4VhVyBvXInTt3DlOnTsXkyZOxc+dOLFy4EAcOHMC+ffswaNAgkx9uSeyRk05a\nWg6OHctEY6MDXFw0GDu2ByIj7e/eRyJzUGvU+C7jO1wqvCSOuTm5YW7MXIT5hskYGRHJydS6xeDj\nRwoKCrBlyxbk5OQgJCQEL7zwglXvWGUhR0TW5kn9cPNi5sHXzVfGyIhIbhYv5ABAo9GguLgYnTp1\nsvqt8CzkpMcziKTFfEurtfnOr8zH9pvbUdVYJY7F+sdieuR0LqU+AT/j0mK+pWWuc+QM6pErKyvD\nwoUL4ebmhs6dO8PV1RUvvPACSktLjX6glLjZgYiswZWiK/jy6pdiEaeAAhN6TMDsqNks4ojauNZu\ndjBoRm7GjBlwcnLCmjVrEBISgtzcXLz//vtobGzEvn37TH64JXFGjojkptaocSjjEC4WXhTH3Jzc\nMCd6Dnp04IYgItKx6NKqt7c3ioqK4O7uLo7V1tYiMDAQFRUVRj9UCizkiEhO1Y3V2JG8A7kVueJY\nQPsAzI+dz344InqERZdWe/XqhezsbL2xnJwc9OrVy+gHkv3iMra0mG9pGZPvgsoCrL28Vq+Ii+kU\ng2UDlrGIMwI/49JivqVlrnwbdLPDmDFjMGHCBLz44osIDg5Gbm4utmzZgoULF2LDhg3QarVQKBRY\nunSpWYIiIrJVV4uu4mD6QfG+VAUUGBc2DkODh1r9JjEisj0GLa0276po+Y9Qc/HW0vHjx80bXSso\nFAqsWrUKcXFx3IVDRBan1qhxOPMwLhRcEMdcnVwxJ3oOwjuEyxgZEVkzpVIJpVKJ1atXW/74EVvC\nHjkikkp1YzV2Ju9ETkWOOObf3h/zY+ejg1sHGSMjIlth0R45IkOwv0JazLe0npTv5n64lkVcdKdo\nLB+wnEVcK/EzLi3mW1qS9sgREdGjrt29hoPpB6HSqAAI/XBjw8ZiWPAw9sMRkSS4tEpEZCS1Ro0j\nmUdwvuC8OObq5IrZUbPRs2NPGSMjIltlat3CGTkiIiPUNNZgZ8pOZJdni2PshyMiuRjcI5eamooP\nPvgAr776KgDg1q1buHHjhsUCI9vD/gppMd/SUiqVKKwqxNrLa/WKuCi/KCzrv4xFnAXwMy4t5lta\n5sq3QYXczp07MXLkSBQUFGDTpk0AgKqqKrz11ltmCYKIyFqlZaThn9v/iX8f+Dde/vvLyLyTCUDo\nhxvTfQziY+LRzqmdzFESUVtlUI9cr169sG3bNvTr1w++vr4oKytDU1MTAgMDUVxcLEWcRuM5ckTU\nWmkZafgy6Uvk++UjvzIfAKDKUGFw78FYMW4FIjpGyBwhEdk6Sc6R69ixIx48eAAHBwe9Qq5Lly64\nf/++SYFbGjc7EFFr/XnLn3HK8RQqGyrFMXdnd8Rp4/D7F34vY2REZG8seo7cgAEDsHnzZr2x7du3\nY9CgQUY/kOwX+yukxXxbVnpJOk7knhCLuPJb5fBz98OAwAFwcXKRObq2gZ9xaTHf0pL0HLlPPvkE\n48ePxxdffIHa2lpMmDAB6enpOHLkiFmCICKyFmqNGkl3kvBD3g/QaDQAhH64QI9AxHSKgUKhgIsD\nCzkisg4GnyNXU1ODgwcPIicnByEhIZgyZQo8PT0tHZ/JuLRKRMaqbKjEzuSdyKvMAwAUFxYjNT0V\nfZ7rA29XbwBAw+0GLB69GJHhkXKGSkR2xtS6hQcCExEBuF1yG3tu7UFtU604Ft4hHLEusThz4wwa\nNY1wcXDB2AFjWcQRkdlZtEcuJycHS5cuRf/+/dGzZ0/xT0QEd2yRDvsrpMV8m4dGq8GxrGP4z4//\nEYs4BRQY230sftn7l+jXqx9+Hf9r9Oss/CeLOOnwMy4t5ltakvbIzZ07F1FRUVizZg1cXV3N8mAi\nIrlVNVRhV8ouvQvvPV08MSd6Drr5dJMxMiIiwxi0tOrt7Y3S0lI4OjpKEZNZcGmViJ4mszQTu1N3\no6apRhzr4dsDs6Jmob1LexkjI6K2yKJ3rU6dOhUnTpzAmDFjjH6AnBITE3kgMBHp0Wg1OJF9Aidz\nTkIL4R9NBRSIC43DiG4j4KAw+OZCIqJWaz4Q2FQGzcgVFxdjyJAhiIiIgL+/v+6bFQps2LDB5Idb\nEmfkpKdUKlk0S4j5Nl51YzW+Tvkad8rviGMeLh6YHTUb3X27P/V7mW/pMefSYr6l9XC+LTojt3Tp\nUri4uCAqKgqurq7iwxQKhdEPJCKSw52yO/g69WtUN1aLY919umN29Gx4uHjIGBkRkekMmpHz9PRE\nQUEBvLy8pIjJLDgjR0SAsJR6KucUlNlKvaXUkd1GYlToKC6lEpFVsOiMXJ8+fVBSUmJThRwRUU1j\nDXan7kZmWaY41t65PWZFzUKPDj1kjIyIyDwM+lV0zJgxmDhxIj788ENs2LABGzZswBdffGG1/XEk\nD55BJC3m++lyynPw2aXP9Iq4bt7d8MrAV0wq4phv6THn0mK+pSXpOXKnTp1CUFDQY+9WXbp0qVkC\nISIyB61Wi9O5p5F0J0lcSgWAESEjMLr7aC6lEpFd4RVdRGQ3aptqsTt1NzJKM8Qxd2d3zIqahfAO\n4TJGRkT0dGbvkWu5K1Wj0TzxBzg48LdbIpJfbkUudqXsQmVDpTgW4h2COdFz4NWO/b1EZJ+eWIW1\n3Njg5OT02D/Ozs6SBEm2gf0V0mK+BVqtFj/k/oCN1zbqFXHDgodhUd9FZivimG/pMefSYr6lZfEe\nueTkZPHrrKwsszyMiMic6prqsOfWHqSXpItjbk5umBk1ExEdI2SMjIhIGgb1yP3lL3/B22+//cj4\nxx9/jLfeessigbUWe+SI7Ft+ZT52Ju9ERUOFONbVqyvmRs+Ft6u3jJERERnP1LrF4AOBq6qqHhn3\n9fVFWVmZ0Q+VAgs5Ivuk1WpxLv8cvs/6Hhqtrn93SNchGBc2Do4OjjJGR0RkGoscCJyUlAStVgu1\nWo2kpCS99zIzM63+gODExETExcXx7jiJ8J4+abXFfNc11WFf2j7cKr4ljrk6uWJGrxno5dfLos9u\ni/mWG3MuLeZbWs35ViqVreqXe2oht3TpUigUCjQ0NGDZsmXiuEKhQEBAAD755BOTHyyFxMREuUMg\nIjMpqCzAzpSdKK8vF8e6eHbBnOg58HXzlTEyIiLTNU84rV692qTvN2hpdeHChdi8ebNJD5ALl1aJ\n7INWq8WFggs4knkEaq1aHB/cZTDG9xgPJweDzjUnIrJqFu2Rs0Us5IhsX72qHvvT9iPlQYo41s6x\nHab3mo7oTtEyRkZEZF6m1i08zZfMhmcQScve811UVYS1l9fqFXGBHoFYMXCFLEWcvefbGjHn0mK+\npSXpXatERFLRarW4VHgJhzIO6S2lPhv0LCaGT+RSKhFRC1xaJSKr0aBqwIH0A7h5/6Y41s6xHZ6P\nfB6x/rEyRkZEZFkWOX6EiEgqd6vvYmfyTpTUlYhjAe0DEB8Tj47uHWWMjIjIerFHjsyG/RXSspd8\na7VaXC68jPVX1usVcc8EPoPlA5ZbTRFnL/m2Jcy5tJhvabFHjohsXqO6EQfTD+LGvRvimIujC6ZG\nTEWfgD4yRkZEZBvYI0dEsrhfcx87kneguLZYHPNv74/4mHj4ufvJGBkRkfTYI0dENuPa3Wv4Jv0b\nNGmaxLH+nftjcs/JcHZ0ljEyIiLbwh45Mhv2V0jLFvPdpG7C3lt7sffWXrGIc3ZwxoxeMzC913Sr\nLuJsMd+2jjmXFvMtLfbIEZFNeVDzADtTduJ+zX1xrJN7J8yNmQv/9v4yRkZEZLvYI0dEFnfj3g0c\nSDugt5TaN6AvpkRMgYuji4yRERFZB/bIEZHVaVI34buM73Cl6Io45uTghMk9J6N/5/5QKBQyRkdE\nZPtsrkfuwoULGDp0KEaNGoUFCxZApVLJHRL9hP0V0rL2fJfUlmD9lfV6RVxHt454acBLGBA4wOaK\nOGvPtz1izqXFfEvLXPm2uUIuJCQEx48fx4kTJxAaGop9+/bJHRIRPeTm/Zv4/PLnuFdzTxzr7d8b\nLz/zMgI8AmSMjIjIvth0j9yqVavQv39/zJgx45H32CNHJD2VRoVDGYdwqfCSOObk4IRJ4ZPwTOAz\nNjcLR0QkFVPrFpst5HJycpCQkIBTp07B0dHxkfdZyBFJq7SuFDuTd6Koukgc6+DWAXOj5yLQM1DG\nyIiIrJ+pdYukS6uffvopBg4cCFdXVyxZskTvvdLSUsycORMeHh4IDQ3F1q1bxff++te/YvTo0fjo\no48AAJWVlXjxxRfx1VdfPbaII3mwv0Ja1pTvlAcp+PzS53pFXEynGKx4ZoXdFHHWlO+2gjmXFvMt\nLZs8R65Lly5YuXIlDh8+jLq6Or33Xn31Vbi6uuL+/fu4evUqpkyZgr59+yI6Ohpvvvkm3nzzTQCA\nSqXC/PnzsWrVKvTs2VPK8InoISqNCkcyj+BCwQVxzFHhiInhE/Fs0LNcSiUisjBZllZXrlyJ/Px8\nfPnllwCAmpoadOjQAcnJyQgPDwcALFq0CEFBQfjwww/1vnfz5s1488030bt3bwDAr371K8THxz/y\nDIVCgUWLFiE0NBQA4OPjg379+iEuLg6ArhLma77ma9NeVzVUocivCIVVhci+lg0A6P9cf8yNmYv0\ny+myx8fXfM3XfG3Nr5u/zs7OBgB89dVXttMj995776GgoEAs5K5evYrhw4ejpqZG/Dsff/wxlEol\n9u/fb9Iz2CNHZDmpD1KxL20f6lX14liUXxSm95oOVydXGSMjIrJNNtEj1+zh5Zbq6mp4eXnpjXl6\neqKqqkrKsKiVWv6WQZYnR77VGjUOZRzC9uTtYhHnqHDEpPBJiI+Jt+sijp9v6THn0mK+pWWufMty\ns8PDFaeHhwcqKyv1xioqKuDp6SllWET0FOX15diVsgv5lfnimI+rD+ZGz0UXry4yRkZE1HbJUsg9\nPCMXEREBlUqFjIwMsUfu+vXriI2NbdVzEhMTERcXJ65Lk2Uxz9KSMt9pxWnYe2sv6lS6TUqRHSMx\no9cMuDm7SRaHnPj5lh5zLi3mW1ote+ZaMzsnaY+cWq1GU1MTVq9ejYKCAqxbtw5OTk5wdHREQkIC\nFAoF1q9fjytXrmDq1Kk4e/YsoqKiTHoWe+SIWk+tUePYnWM4k3dGHHNQOGBc2DgM6TqEu1KJiMzE\nJnrk1qxZA3d3d/zpT3/Cli1b4Ob2/9u796Cm7rQP4N9wCfcIKMhNLhUUeW2xVbwBKdR2u7xbq2Iv\n2lettquul27t7PQ2VsVRp+Nu69p91dr1nV11rVitdrdau7UVEaQK2irLCgqoBK+ggoRrCMl5/+hL\nXiNaIcLvcJLvZ8aZ5vxOyMN30szDOc858cCqVasAABs2bEBzczMCAwMxbdo0bNy40eYmjuTB+Qqx\nejrvupY6bD612aqJ07hpMGvYLIwdMNbhmji+v8Vj5mIxb7EUOSOXkZGBjIyMu675+fnhiy++EFkO\nEd1D2c0yfHHmCzQZmyzbYvxjMGnIJHi6espYGRER3U6xX9F1PyqVCsuWLeOMHFEXmCUzsi5k4Ujl\nEcs2J5UTnoh6AokDEh3uKBwRUU9rn5Fbvny5cu4jJwJn5Ii6Rm/QY3fxbujqdJZtPmofPBf3HCJ8\nI2SsjIjI/iliRo7sG+crxOrOvM/VnMMnJz6xauIG+g3Eb0b8hk3c/+H7WzxmLhbzFkuRM3JE1LuY\nJTMOVxxGji4HEn76S1AFFVKjUpEcnsxTqUREvZxdn1rljBzRvTW0NmB38W5cuHXBss1b7Y3JQyYj\nyi9KxsqIiBwHZ+TugTNyRPd2ofYCdpfsRkNrg2VblG8UJsdNhrfaW8bKiIgcE2fkSHacrxDLlrzb\nT6VuLdxqaeJUUCElMgXT46ezifsZfH+Lx8zFYt5icUaOiLqksbURu0t243ztecs2L1cvTI6bjIf8\nHpKxMiIishVPrRI5gIpbFdhdvBv1rfWWbZG+kZg8ZDJ83HxkrIyIiADb+xa7PiKXkZHBix3IoUmS\nhCOVR5B1IctyVSoAaCO0SIlMgZOK0xVERHJqv9jBVjwiR90mOzubTbNA98u7ydiEPSV7UF5Tbtnm\n6eqJ9CHpiPaPFlChfeH7WzxmLhbzFuvOvHlEjogsKusq8Xnx59Ab9JZt4X3C8Vzcc9C4aWSsjIiI\nuhOPyBHZEUmS8P3F73HwwkGYJbNle1J4ElIjU+Hs5CxjdUREdC88Ikfk4JqMTfj7mb+j9GapZZuH\niwcmDZmEQX0HyVgZERH1FE46U7fhPYjEuj3vS/pL+OTEJ1ZN3ADNAPxmxG/YxHUTvr/FY+ZiMW+x\neB+5TuBVq2TvJEnCsUvH8O35b61OpY4dMBbjosbxVCoRUS/Hq1bvgTNyZK/Olp/Fdz98h8a2Rpy5\nfgaegZ7oF9IPAODu4o6JsRMR2y9W5iqJiKgrOCNH5ADOlp/F5kObYQg3oPh6MVp8W9BW3IZhGIb4\nwfF4/j+eh6+7r9xlEhGRIJyRo27D+Yqe9+WxL6Hz1+Hk1ZO49u9rAACXaBdINRJeefQVNnE9iO9v\n8Zi5WMxbLM7IETmQmuYa5OpycbDiIFrCWizbnVXOiO0Xixj3GM7DERE5IM7IEfVi1xuvI7cyF0VV\nRZAgoeBIAZrCmgAAfdz6ILZfLDxcPRBYHYj5L8yXuVoiIrIVZ+SI7Mi1hmvI0eWg5HqJ1XekPvTQ\nQ6g4X4GBwwfC190XKpUKhjIDxqWOk7FaIiKSi13PyGVkZPCcv0DM+sFd1l9GZlEmNp7YiOLrxVZN\n3EC/gXjzP9/Eh1M+xOD6wbh56CYCqwMxM3UmBkcPlrFqx8D3t3jMXCzmLVZ73tnZ2cjIyLD559j1\nEbkHCYZIpMq6ShyuOIxztec6rA3uOxjJEckI04T9tMEXGBw9GNmB/IJrIiKla7/f7fLly216Pmfk\niGQiSRIu3LqAHF0OKm5VdFiPC4iDNkKLIO8g8cUREZFQnJEjUghJklBeU44cXQ4u6i9aramgwtDA\noUiOSEagV6BMFRIRkVLY9YwcicX5ip8nSRLO3DiDTT9uwqdFn1o1cU4qJzwa9CgWjlyIyXGTO9XE\nMW+xmLd4zFws5i0W7yNHpBBmyYzi68XI1eWiqrHKas1Z5YxHgx9FUngSb+ZLRERdxhk5oh5ilswo\nqipCbmUubjTdsFpzcXLB8ODhSAxPhMZNI1OFRETUW3BGjqiXMJlNKKwqRK4uF7UttVZramc1EkIS\nMGbAGHirvWWqkIiI7AVn5KjbOPp8RZu5DQWXC/Cn/D/hy7NfWjVxbs5u0EZosWj0Ijw18KluaeIc\nPW/RmLd4zFws5i0WZ+Q6ISMjw3J/FqKe0mpqxQ9XfkDexTw0tDZYrXm4eGB02GiMChsFdxd3mSok\nIqLeKjs7+4GaOs7IEdnI0GZAweUCHL10FE3GJqs1L1cvjB0wFiNCRsDNxU2mComISCk4I0ckSLOx\nGfmX85F/KR/Nbc1Waz5qHySGJ2J48HC4OrvKVCERETkKzshRt7H3+YrG1kYcPH8Qa4+tRXZFtlUT\n5+vui2cGPYPXR7+O0WGjhTRx9p53b8O8xWPmYjFvsTgjRyRIvaEeRy8dxfHLx2E0G63W/D38kRye\njEf6PwJnJ2eZKiQiIkfFGTmie6hrqUPexTz8ePVHtJnbrNYCPAOQHJGMoYFD4aTigW0iInownJEj\n6ia1zbU4UnkEp66dgkkyWa0FeQdBG6HFkH5DoFKpZKqQiIjoJzyUQN1G6fMVN5pu4IuSL/DfBf+N\nH67+YNXEhfiEYOrQqZg7fC7iAuJ6RROn9LyVhnmLx8zFYt5icUaOqJtUN1YjR5eD09WnIcH6sHZ4\nn3BoI7QY6DewVzRvREREt+OMHDmsq/VXkaPLQcmNkg5rUb5R0EZoEekbyQaOiIh6HGfkiDrpkv4S\nDlccRllNWYe1GP8YaCO0GNBngAyVERERdQ1n5Kjb9Pb5iopbFdhauBX/8+P/dGjiYvvFYs7wOfiv\nR/5LMU1cb8/b3jBv8Zi5WMxbLM7IEXWCJEk4X3seOboc6Op0VmsqqBAXEAdthBb9vfvLVCEREZHt\n7HpGbtmyZUhJSUFKSorc5ZBgkiShrKYMObocXNJfslpzUjnh4cCHkRSehACvAJkqJCIi+unIXHZ2\nNpYvX27TjJxdN3J2+qvRz5AkCSU3SpCjy8G1hmtWa04qJwwLGoak8CT4e/jLVCEREVFHtvYtnJGj\nbiPnfIVZMqOoqggbjm/AztM7rZo4FycXJIQk4PVRr+PZwc/aTRPHeRaxmLd4zFws5i0WZ+SIAJjM\nJhRVFyFXl4ubzTet1lydXDEiZATGDhgLHzcfmSokIiLqOTy1SorUZm7DqWuncKTyCG613LJaUzur\nMTJ0JMaEjYGX2kumComIiDqP95Ejh2A0GfHj1R+RdzEPeoPeas3dxR2jQkdhdNhoeLh6yFQhERGR\nOJyRo27Tk/MVraZW5FXm4aP8j/B1+ddWTZynqyfGRY3DotGLkBqV6jBNHOdZxGLe4jFzsZi3WJyR\nI4fQ0taCgssFOHbpGJqMTVZr3mpvjB0wFiNCRkDtrJapQiIiIvlwRo56pWZjM45dOob8y/loaWux\nWtO4aZA4IBGPBT8GV2dXmSokIiLqPpyRI7vQ2NqI7y9+j+NXjqPV1Gq15uvui+TwZMQHxcPFiW9d\nIiIizshRt3mQ8/16gx7/LP8n1h5bi7yLeVZNXF+PvpgYOxGvjXwNw0OGs4n7P5xnEYt5i8fMxWLe\nYnFGjuzCrZZbyKvMw49Xf4RJMlmtBXoFQhuhRVxAHJxU/JuDiIjoTpyRI1nUNNcgV5eLwqpCmCWz\n1VqwdzC0EVrE9ouFSqWSqUIiIiJxOCNHinC98TpyK3NRVFUECdZv2DBNGLQRWsT4x7CBIyIi6gSe\nr6Ju83Pn+681XMOu07uw4fgG/KvqX1ZNXESfCMyIn4FXH30Vg/oOYhPXSZxnEYt5i8fMxWLeYnFG\njhThsv4ycnQ5OHvzbIe1gX4DoY3QIsI3QobKiIiIlE9xM3JVVVVIT0+HWq2GWq3G9u3b0bdv3w77\ncUZOXpV1lcjR5aC8przD2qC+g6CN0CJMEyZDZURERL2PrX2L4ho5s9kMJ6efzghv2bIFV69exTvv\nvNNhPzZy4pwtP4vvfvgOreZW1LfUwzXAFQZvQ4f9hvQbAm2EFsE+wTJUSURE1HvZ2rcobkauvYkD\nAL1eDz8/PxmrobPlZ7H50Gac1ZzFruJdyFJl4atjX+HGlRsAABVUeDjwYcxPmI8Xh77IJq4bcZ5F\nLOYtHjMXi3mL1V15K66RA4DCwkKMGjUK69atw9SpU+Uux6Ht+X4P/u31b/yr6l+oPlcNAHCJdsGF\n8xcwLGgYFo5ciMlxkxHoFShzpfbn1KlTcpfgUJi3eMxcLOYtVnflLbSRW7duHUaMGAF3d3fMmjXL\naq2mpgaTJk2Ct7c3IiMjkZmZaVn74x//iNTUVHz44YcAgPj4eOTn52PlypVYsWKFyF+B7qQCGlob\nAABtTW1QQYVg72AkhidiYuxE9PXsOL9I3ePWrVtyl+BQmLd4zFws5i1Wd+UttJELDQ3FkiVL8Mor\nr3RYW7BgAdzd3VFdXY1PP/0U8+bNQ3FxMQDgjTfewKFDh/C73/0ORqPR8hyNRgODoeMslhwe9BBp\nV5/fmf1/bp97rXV2e/tjjVqDYJ9gOKmc4KP2weiw0RjcbzB83X3vW9+D6I5D0l35GT2V973WOrtN\npN72Hrd1nXnbvr+IzxS58DNFPHt+j4vMW2gjN2nSJEyYMKHDVaaNjY3Ys2cPVqxYAU9PTyQmJmLC\nhAn429/+1uFnnDp1Co8//jieeOIJrFmzBm+99Zao8n+WPb8h77a9/fGTw59EyPUQjA4bDWe9M9xc\n3GAoM2DcY+PuW9+D4IcuUFFRcd+aulNve4+LbuQcPe/77dMTjZzIzPmZwvf4/fbprZ8psly1+t57\n7+Hy5cv461//CgA4efIkkpKS0NjYaNlnzZo1yM7OxpdffmnTa0RHR+PcuXPdUi8RERFRT4qPj7dp\nbk6WGwLfeef+hoYGaDQaq20+Pj6or6+3+TXKyzvev4yIiIjInshy1eqdBwG9vb2h1+utttXV1cHH\nx0dkWURERESKIksjd+cRuUGDBqGtrc3qKFphYSGGDh0qujQiIiIixRDayJlMJrS0tKCtrQ0mkwkG\ngwEmkwleXl5IT0/H0qVL0dTUhCNHjmDv3r2YPn26yPKIiIiIFEVoI9d+Verq1auxbds2eHh4YNWq\nVQCADRs2oLm5GYGBgZg2bRo2btyIIUOGiCyPiIiISFEU912rD+rtt9/G0aNHERkZib/85S9wcZHl\neg+Hodfr8eSTT6KkpAT5+fmIi4uTuyS7VlBQgEWLFsHV1RWhoaHYunUr3+M9qKqqCunp6VCr1VCr\n1di+fXuH2ytRz8jMzMTrr7+O6upquUuxaxUVFUhISMDQoUOhUqmwc+dO9OvXT0U9SNUAAAriSURB\nVO6y7Fp2djZWrlwJs9mM3/72t5g4ceLP7q/Ir+iyVWFhIa5cuYKcnBzExsbi888/l7sku+fp6Yn9\n+/fjueees+nLgKlrwsPDcejQIRw+fBiRkZH4xz/+IXdJdi0gIAB5eXk4dOgQXnrpJWzatEnukhyC\nyWTCrl27EB4eLncpDiElJQWHDh1CVlYWm7ge1tzcjDVr1uDrr79GVlbWfZs4wMEauaNHj+Lpp58G\nAPzyl79EXl6ezBXZPxcXF/6PL1BQUBDc3NwAAK6urnB2dpa5Ivvm5PT/H6F6vR5+fn4yVuM4MjMz\n8cILL3S4cI56Rl5eHrRaLRYvXix3KXbv6NGj8PDwwPjx45Geno6qqqr7PsehGrna2lrLLU00Gg1q\nampkroioZ+h0Onz77bcYP3683KXYvcLCQowaNQrr1q3D1KlT5S7H7rUfjXvxxRflLsUhhISE4Ny5\nc8jJyUF1dTX27Nkjd0l2raqqCuXl5di3bx9mz56NjIyM+z5HkY3cunXrMGLECLi7u2PWrFlWazU1\nNZg0aRK8vb0RGRmJzMxMy5qvr6/lfnV1dXXw9/cXWreS2Zr57fjXc+c9SN56vR4zZszAli1beESu\nkx4k7/j4eOTn52PlypVYsWKFyLIVzdbMt23bxqNxNrA1b7VaDQ8PDwBAeno6CgsLhdatVLbm7efn\nh8TERLi4uOCJJ57A6dOn7/taimzkQkNDsWTJErzyyisd1hYsWAB3d3dUV1fj008/xbx581BcXAwA\nGDt2LL777jsAwDfffIOkpCShdSuZrZnfjjNynWdr3m1tbZgyZQqWLVuGmJgY0WUrlq15G41Gy34a\njQYGg0FYzUpna+YlJSXYunUr0tLSUFZWhkWLFokuXZFszbuhocGyX05ODj9XOsnWvBMSElBSUgLg\np++WHzhw4P1fTFKw9957T5o5c6blcUNDg6RWq6WysjLLthkzZkjvvPOO5fGbb74pJScnS9OmTZOM\nRqPQeu2BLZmnpaVJISEh0pgxY6TNmzcLrVfpupr31q1bpb59+0opKSlSSkqK9NlnnwmvWcm6mnd+\nfr6k1Wql1NRU6Re/+IV08eJF4TUrnS2fKe0SEhKE1GhPupr3/v37peHDh0vJycnSyy+/LJlMJuE1\nK5kt7+/169dLWq1WSklJkc6fP3/f11D0fQmkO47wlJaWwsXFBdHR0ZZt8fHxyM7Otjz+/e9/L6o8\nu2RL5vv37xdVnt3pat7Tp0/njbQfQFfzHjlyJA4fPiyyRLtjy2dKu4KCgp4uz+50Ne+0tDSkpaWJ\nLNGu2PL+nj9/PubPn9/p11DkqdV2d85INDQ0QKPRWG3z8fFBfX29yLLsGjMXi3mLxbzFY+ZiMW+x\nROSt6Ebuzk7X29vbcjFDu7q6OsuVqvTgmLlYzFss5i0eMxeLeYslIm9FN3J3drqDBg1CW1sbysvL\nLdsKCwsxdOhQ0aXZLWYuFvMWi3mLx8zFYt5iichbkY2cyWRCS0sL2traYDKZYDAYYDKZ4OXlhfT0\ndCxduhRNTU04cuQI9u7dy5mhbsDMxWLeYjFv8Zi5WMxbLKF5P+gVGXJYtmyZpFKprP4tX75ckiRJ\nqqmpkSZOnCh5eXlJERERUmZmpszV2gdmLhbzFot5i8fMxWLeYonMWyVJvLkXERERkRIp8tQqERER\nEbGRIyIiIlIsNnJERERECsVGjoiIiEih2MgRERERKRQbOSIiIiKFYiNHREREpFBs5IiIiIgUio0c\nEdEdZs6ciSVLlnTrz5w3bx5WrlzZrT+TiMhF7gKIiHoblUrV4cuuH9THH3/crT+PiAjgETkiorvi\ntxcSkRKwkSOiXmX16tUICwuDRqNBbGwssrKyAAAFBQUYM2YM/Pz8EBISgtdeew1Go9HyPCcnJ3z8\n8ceIiYmBRqPB0qVLce7cOYwZMwa+vr6YMmWKZf/s7GyEhYXh/fffR0BAAKKiorB9+/Z71rRv3z4M\nGzYMfn5+SExMRFFR0T33feONN9C/f3/06dMHjzzyCIqLiwFYn64dP348fHx8LP+cnZ2xdetWAMCZ\nM2fw1FNPoW/fvoiNjcWuXbvu+VopKSlYunQpkpKSoNFo8PTTT+PmzZudTJqI7AEbOSLqNc6ePYv1\n69fjxIkT0Ov1OHDgACIjIwEALi4u+Oijj3Dz5k0cPXoUBw8exIYNG6yef+DAAZw8eRLHjh3D6tWr\nMXv2bGRmZqKyshJFRUXIzMy07FtVVYWbN2/iypUr2LJlC+bMmYOysrIONZ08eRKvvvoqNm3ahJqa\nGsydOxfPPvssWltbO+z7zTffIDc3F2VlZairq8OuXbvg7+8PwPp07d69e1FfX4/6+nrs3LkTwcHB\nGDduHBobG/HUU09h2rRpuH79Onbs2IH58+ejpKTknpllZmZi8+bNqK6uRmtrKz744IMu505EysVG\njoh6DWdnZxgMBpw+fRpGoxHh4eF46KGHAACPPfYYRo4cCScnJ0RERGDOnDk4fPiw1fPfeusteHt7\nIy4uDg8//DDS0tIQGRkJjUaDtLQ0nDx50mr/FStWwNXVFVqtFr/61a/w2WefWdbam64///nPmDt3\nLhISEqBSqTBjxgy4ubnh2LFjHepXq9Wor69HSUkJzGYzBg8ejKCgIMv6nadrS0tLMXPmTOzcuROh\noaHYt28foqKi8PLLL8PJyQnDhg1Denr6PY/KqVQqzJo1C9HR0XB3d8cLL7yAU6dOdSFxIlI6NnJE\n1GtER0dj7dq1yMjIQP/+/TF16lRcvXoVwE9NzzPPPIPg4GD06dMHixcv7nAasX///pb/9vDwsHrs\n7u6OhoYGy2M/Pz94eHhYHkdERFhe63Y6nQ4ffvgh/Pz8LP8uXbp0131TU1OxcOFCLFiwAP3798fc\nuXNRX19/19+1rq4OEyZMwKpVqzB27FjLa+Xn51u91vbt21FVVXXPzG5vFD08PKx+RyKyf2zkiKhX\nmTp1KnJzc6HT6aBSqfD2228D+On2HXFxcSgvL0ddXR1WrVoFs9nc6Z9751WotbW1aGpqsjzW6XQI\nCQnp8Lzw8HAsXrwYtbW1ln8NDQ148cUX7/o6r732Gk6cOIHi4mKUlpbiD3/4Q4d9zGYzXnrpJYwb\nNw6//vWvrV7r8ccft3qt+vp6rF+/vtO/JxE5FjZyRNRrlJaWIisrCwaDAW5ubnB3d4ezszMAoKGh\nAT4+PvD09MSZM2c6dTuP209l3u0q1GXLlsFoNCI3NxdfffUVnn/+ecu+7fvPnj0bGzduREFBASRJ\nQmNjI7766qu7Hvk6ceIE8vPzYTQa4enpaVX/7a+/ePFiNDU1Ye3atVbPf+aZZ1BaWopt27bBaDTC\naDTi+PHjOHPmTKd+RyJyPGzkiKjXMBgMePfddxEQEIDg4GDcuHED77//PgDggw8+wPbt26HRaDBn\nzhxMmTLF6ijb3e77duf67Y+DgoIsV8BOnz4dn3zyCQYNGtRh3+HDh2PTpk1YuHAh/P39ERMTY7nC\n9E56vR5z5syBv78/IiMj0a9fP7z55psdfuaOHTssp1Dbr1zNzMyEt7c3Dhw4gB07diA0NBTBwcF4\n991373phRWd+RyKyfyqJf84RkYPJzs7G9OnTcfHiRblLISJ6IDwiR0RERKRQbOSIyCHxFCQR2QOe\nWiUiIiJSKB6RIyIiIlIoNnJERERECsVGjoiIiEih2MgRERERKRQbOSIiIiKF+l+7eRzDyLZFUAAA\nAABJRU5ErkJggg==\n", + "text": [ + "
\n", + "
\n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "# Numba vs. Cython vs. regular (C)Python & Numpy" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Numba is using the [LLVM compiler infrastructure](http://llvm.org) for compiling Python code to machine code. Its strength is to work with NumPy arrays to speed-up the code. If you want to read more about Numba, please see refer to the original [website and documentation](http://numba.pydata.org/numba-doc/0.13/index.html)." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Here, we implement a linear regression via least squares fitting (with vertical offsets) by solving to fit *n* points $(x_i, y_i)$ with $i=1,2,...n,$ via linear equation of the form \n", + "$f(x) = a\\cdot x + b$. \n", + "\n", + "\n", + "\n", + "$\\Rightarrow \\pmb a = (\\pmb X^T \\; \\pmb X)^{-1} \\pmb X^T \\; \\pmb y$" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "I have described the approach in more detail in this [IPython notebook](http://sebastianraschka.com/IPython_htmls/cython_least_squares.html)." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### Matrix equation " + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "In order to obtain the parameters for the linear regression line for a set of multiple points, we can re-write the problem as matrix equation \n", + "$\\pmb X \\; \\pmb a = \\pmb y$ \n", + "\n", + "$\\Rightarrow\\Bigg[ \\begin{array}{cc}\n", + "x_1 & 1 \\\\\n", + "... & 1 \\\\\n", + "x_n & 1 \\end{array} \\Bigg]$\n", + "$\\bigg[ \\begin{array}{c}\n", + "a \\\\\n", + "b \\end{array} \\bigg]$\n", + "$=\\Bigg[ \\begin{array}{c}\n", + "y_1 \\\\\n", + "... \\\\\n", + "y_n \\end{array} \\Bigg]$ \n", + "\n", + "With a little bit of calculus, we can rearrange the term in order to obtain the parameter vector \n", + "$\\pmb a = [a\\;b]^T$ \n", + "\n", + "We will implement this matrix equation in \n", + "- Python/CPython: `py_mat_lstsqr()` \n", + "- Numba: `numba_mat_lstsqrs()` \n", + "- Cython: `cy_mat_lstsqr()`" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### \"Classic\" approach " + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "In the more \"classic\" approach that is often found in statistics textbooks, we calculate the following parameters as follows:\n", + "\n", + "$a = \\frac{S_{x,y}}{\\sigma_{x}^{2}}\\quad$ (slope)\n", + "\n", + "$b = \\bar{y} - a\\bar{x}\\quad$ (y-axis intercept)\n", + "\n", + "where \n", + "\n", + "\n", + "$S_{xy} = \\sum_{i=1}^{n} (x_i - \\bar{x})(y_i - \\bar{y})\\quad$ (covariance)\n", + "\n", + "\n", + "$\\sigma{_x}^{2} = \\sum_{i=1}^{n} (x_i - \\bar{x})^2\\quad$ (variance)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "We will implement this \"classic\" approach in\n", + "- Python/CPython: `py_lstsqr()` \n", + "- Numba: `numba_lstsqrs()` \n", + "- Cython: `cy_lstsqrs()` " + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
\n", + "
" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "import numpy as np\n", + "import scipy.stats\n", + "from numba import jit" + ], + "language": "python", + "metadata": {}, + "outputs": [] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%load_ext cythonmagic" + ], + "language": "python", + "metadata": {}, + "outputs": [] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### Matrix equation:" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "def py_mat_lstsqr(x, y):\n", + " \"\"\" Computes the least-squares solution to a linear matrix equation. \"\"\"\n", + " X = np.vstack([x, np.ones(len(x))]).T\n", + " return (np.linalg.inv(X.T.dot(X)).dot(X.T)).dot(y)" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 59 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "@jit\n", + "def numba_mat_lstsqr(x, y):\n", + " \"\"\" Computes the least-squares solution to a linear matrix equation. \"\"\"\n", + " X = np.vstack([x, np.ones(len(x))]).T\n", + " return (np.linalg.inv(X.T.dot(X)).dot(X.T)).dot(y)" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 60 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%%cython\n", + "def cy_mat_lstsqr(x, y):\n", + " \"\"\" Computes the least-squares solution to a linear matrix equation. \"\"\"\n", + " X = np.vstack([x, np.ones(len(x))]).T\n", + " return (np.linalg.inv(X.T.dot(X)).dot(X.T)).dot(y)" + ], + "language": "python", + "metadata": {}, + "outputs": [] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### \"Classic\" approach:" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "def py_lstsqr(x, y):\n", + " \"\"\" Computes the least-squares solution to a linear matrix equation. \"\"\"\n", + " x_avg = sum(x)/len(x)\n", + " y_avg = sum(y)/len(y)\n", + " var_x = 0\n", + " cov_xy = 0\n", + " for x_i, y_i in zip(x,y):\n", + " temp = (x_i - x_avg)\n", + " var_x += temp**2\n", + " cov_xy += temp*(y_i - y_avg)\n", + " slope = cov_xy / var_x\n", + " y_interc = y_avg - slope*x_avg\n", + " return (slope, y_interc)" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 61 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "@jit\n", + "def numba_lstsqr(x, y):\n", + " \"\"\" Computes the least-squares solution to a linear matrix equation. \"\"\"\n", + " x_avg = sum(x)/len(x)\n", + " y_avg = sum(y)/len(y)\n", + " var_x = 0\n", + " cov_xy = 0\n", + " for x_i, y_i in zip(x,y):\n", + " temp = (x_i - x_avg)\n", + " var_x += temp**2\n", + " cov_xy += temp*(y_i - y_avg)\n", + " slope = cov_xy / var_x\n", + " y_interc = y_avg - slope*x_avg\n", + " return (slope, y_interc)" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 62 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%%cython\n", + "def cy_lstsqr(x, y):\n", + " \"\"\" Computes the least-squares solution to a linear matrix equation. \"\"\"\n", + " cdef double x_avg, y_avg, temp, var_x, cov_xy, slope, y_interc, x_i, y_i\n", + " x_avg = sum(x)/len(x)\n", + " y_avg = sum(y)/len(y)\n", + " var_x = 0\n", + " cov_xy = 0\n", + " for x_i, y_i in zip(x,y):\n", + " temp = (x_i - x_avg)\n", + " var_x += temp**2\n", + " cov_xy += temp*(y_i - y_avg)\n", + " slope = cov_xy / var_x\n", + " y_interc = y_avg - slope*x_avg\n", + " return (slope, y_interc)" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 63 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### NumPy and SciPy libraries:" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "def numpy_lstsqr(x, y):\n", + " \"\"\" Computes the least-squares solution to a linear matrix equation. \"\"\"\n", + " X = np.vstack([x, np.ones(len(x))]).T\n", + " return np.linalg.lstsq(X,y)[0]" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 70 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "def scipy_lstsqr(x,y):\n", + " \"\"\" Computes the least-squares solution to a linear matrix equation. \"\"\"\n", + " return scipy.stats.linregress(x, y)[0:2]" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 71 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### Verifying that the different approaches yield the same results" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "import random\n", + "random.seed(12345)\n", + "\n", + "n = 500\n", + "x = [x_i*random.randrange(8,12)/10 for x_i in range(n)]\n", + "y = [y_i*random.randrange(10,14)/10 for y_i in range(n)]\n", + "\n", + "np.testing.assert_array_almost_equal(\n", + " py_lstsqr(x, y), py_mat_lstsqr(x, y), decimal=6)\n", + "np.testing.assert_array_almost_equal(\n", + " numpy_lstsqr(x,y), py_lstsqr(x, y), decimal=6)\n", + "np.testing.assert_array_almost_equal(\n", + " scipy_lstsqr(x,y), py_lstsqr(x, y), decimal=6)\n", + "\n", + "print('ok')" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "ok\n" + ] + } + ], + "prompt_number": 80 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### Visual checking the least square fit" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%pylab inline" + ], + "language": "python", + "metadata": {}, + "outputs": [] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "from matplotlib import pyplot as plt\n", + "\n", + "slope, intercept = py_mat_lstsqr(x, y)\n", + "\n", + "line_x = [round(min(x)) - 1, round(max(x)) + 1]\n", + "line_y = [slope*x_i + intercept for x_i in line_x]\n", + "\n", + "plt.figure(figsize=(7,6))\n", + "plt.scatter(x,y)\n", + "plt.plot(line_x, line_y, color='red', lw='2')\n", + "\n", + "plt.ylabel('y')\n", + "plt.xlabel('x')\n", + "plt.title('Linear regression via least squares fit')\n", + "\n", + "ftext = 'y = ax + b = {:.3f} + {:.3f}x'\\\n", + " .format(slope, intercept)\n", + "plt.figtext(.15,.8, ftext, fontsize=11, ha='left')\n", + "\n", + "plt.show()" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "display_data", + "png": "iVBORw0KGgoAAAANSUhEUgAAAdQAAAGQCAYAAAAX7pEWAAAABHNCSVQICAgIfAhkiAAAAAlwSFlz\nAAALEgAACxIB0t1+/AAAIABJREFUeJzs3Xl4TNf/wPH3zGSbySabLbGEiCzErpYiKEVbpaVqD6qr\nVrXWtvZqS1v90mpRFKW0iKVaamnsRWyx7xHEFktC9mTm8/sjMT+RRCIZBOf1PHnq3nPuWe7czmfu\nveeeqxERQVEURVGUQtE+6gYoiqIoypNABVRFURRFsQAVUBVFURTFAlRAVRRFURQLUAFVURRFUSxA\nBVRFURRFsQAVUBWCg4N58803H3UznjobNmxAq9Vy4cKFR90Uzpw5g1arZdu2bYUqJyQkhBYtWlio\nVYolpKen07t3b9zd3dFqtWzcuFF9Tg+I1aNugPLghYSEEB0dzdq1a3NMX7ZsGVZW6lB42Bo2bMil\nS5fw8PB41E2hbNmyXLp0CVdX10KVo9Fo0Gg0FmpV/sybN48ePXpgMpkear2PiyVLlrBgwQLCwsKo\nUKECLi4u1KxZM8v+euONNzh16hRhYWGPsKWPP/Ut+hTI60uuWLFiD7E195aamoqNjU2e+W5/GWi1\nD+YiS37bURjW1tYUL178gdaRX1qt1iJtERHUXDH370EezydOnMDT05N69eqZ11lbW1u8HkVd8n0q\n5PUlFxwcTN++fbMtjx07llKlSuHm5kbPnj1JSEjIst3ChQupXr06er0eb29vPv74YxITE83pa9eu\nJTg4GDc3N4oVK0ZwcDDh4eFZytBqtXz//fd06dKFYsWK0bNnzxzbOGrUKCpVqsQff/yBn58ftra2\nnDhxgvj4ePr374+Xlxf29vbUrFmTpUuXZtl279691KtXD71ej5+fH6GhoZQvX55x48bl2Y61a9fS\nsGFDDAYDXl5e9O7dm+vXr5u3O3ToEM8//zwuLi44ODgQEBDAvHnzzOkzZszA398fvV6Pm5sbTZo0\nITo6Gsj5ku/27dtp3LgxBoMBV1dXunbtSkxMTLb9sGLFCvz8/HBwcKBp06acPHkyl08Xfv75Z4oV\nK0ZKSkqW9ePHj6dcuXJAzpd8P/30UwICArC3t6ds2bK888473Lx5M9d6cmOJ4yS3/bhhwwZ69OgB\nZHyGWq2W3r1759qWL774gooVK2JnZ0fx4sVp1aoVycnJ5vTvv//efCy1atWKuXPnZvmMZs+enS0Y\nnT9/Hq1Wy6ZNm8zr+vbti4+PDwaDgYoVK/Lpp5+SmppqTi/M8ZxXH+4UHBzMiBEjOH36NFqtlgoV\nKgBZL82PGjWKWbNmsXHjRvM+nDt3bq77ULkHUZ54PXv2lOeeey7X9ODgYOnbt695uUmTJlKsWDH5\n6KOP5NixY7JmzRpxdXWV4cOHm/P88ssv4uLiIvPmzZPIyEjZtGmTBAUFSffu3c15li5dKosWLZLj\nx4/L4cOH5Y033hBXV1e5du2aOY9GoxE3NzeZMmWKnD59Wk6ePJljG0eOHCkGg0GCg4Nl586dcuLE\nCbl165YEBwdL06ZNZevWrRIZGSnTp08XGxsbWb9+vYiIJCQkSMmSJaVt27Zy4MAB2b59uzRo0EAM\nBoOMGzfunu1Yv369GAwG+eGHH+TkyZMSHh4uTZs2lSZNmpi3q1q1qnTt2lWOHDkikZGRsmrVKlm5\ncqWIiOzatUusrKzk119/lbNnz8qBAwdk5syZcv78eRERCQsLE41GI9HR0SIicvHiRXF0dJSuXbvK\nwYMHZcuWLRIUFCSNGzfOsh/s7e2ldevWsmfPHomIiJBatWpJo0aNcv184+LiRK/Xy++//55lfUBA\ngHz66aciIhIZGSkajUa2bt1qTv/8889ly5YtEhUVJevXrxc/Pz/p2bNnrvWIZD/WLHGc3Gs/pqam\nypQpU0Sj0cjly5fl8uXLcvPmzRzbtmTJEnFycpKVK1fKuXPnZN++fTJp0iRJSkoSEZFly5aJlZWV\nfPfdd3LixAmZOXOmFC9eXLRarfkz+uWXX8TKyipLuefOnRONRiMbN24UERGTySSffvqp7Ny5U6Ki\nomTFihVSqlQpGTlypHmbgh7PefXhbtevX5eBAweKt7e3XL58Wa5evWr+nFq0aCEiIvHx8dK1a1dp\n2LCheR/mVp5ybyqgPgUKElCrV6+eJc8777wj9evXNy+XK1dOpk2bliXPxo0bRaPRSGxsbI71GI1G\ncXFxkfnz55vXaTQaeeONN/Lsw8iRI0Wr1cq5c+fM68LCwsTOzk7i4uKy5O3Vq5e0a9dORESmT58u\nDg4OWb5kjx49KhqNJltAvbsdTZo0kWHDhmVZFxUVJRqNRiIiIkRExNnZWWbPnp1jm0NDQ8XZ2TnX\nL/i7A+pnn30mZcqUkbS0NHOeiIgI0Wg0snnzZvN+sLKyMn8xioj8/vvvotVqJSUlJcd6RERef/11\neeGFF8zL4eHhotFo5Pjx4yKSc0DNqT+2tra5potkP9YscZzktR9//fVX0Wg092yXiMjEiRPF19c3\ny/69U8OGDaVbt25Z1g0cODDLZ5SfgJpb3ZUqVTIvF/R4zqsPORk5cqT4+PhkWXf359SnTx8JDg7O\nd5lKztQlXyUbjUZDtWrVsqwrVaoUly9fBiAmJoazZ88yYMAAHB0dzX9t2rRBo9GYLz9GRkbSvXt3\nKlWqhLOzM87OzsTFxXH27NksZdetWzdf7SpRogReXl7m5fDwcFJTU/H09MzSjvnz55vbcPjwYQIC\nAnB0dDRvV7ly5RzvG9/djvDwcL777rssZQcGBqLRaDhx4gQAAwcO5I033qBp06aMHj2avXv3mrdv\n2bIlFSpUwNvbm86dO/Pzzz9z7dq1XPt36NAh6tWrl2WAWFBQEM7Ozhw6dMi8rnTp0ri5uZmXS5Uq\nhYhw5cqVXMvu2bMna9as4erVqwDMnTuXZ555hkqVKuW6TWhoKI0bNzbv327dupGWlsalS5dy3eZO\nljpO7nc/5qZTp06kpaVRrlw5evXqxbx584iPjzenHzlyhAYNGmTZpmHDhvddD2RcZn/mmWcoWbIk\njo6OfPLJJ9mO+4Icz3n1QXm0VEBVcnT3gByNRmMeOHH7v5MnTyYiIsL8t3//fk6cOEGVKlUAePHF\nFzl//jw//vgjO3bsYN++fRQvXjzLvSQAe3v7fLXp7nwmkwlnZ+csbYiIiODIkSOsWrXqvvt8d/ki\nwtChQ7OVf+LECVq1agXAZ599xvHjx3nttdc4ePAg9erVY/jw4ebydu3axdKlS/H19WXq1Kn4+Piw\nZ8+eHOvXaDT5GtCT02dze3/kpkWLFri7uzN//nzS0tJYuHBhrverAXbs2MFrr71GcHAwy5YtY+/e\nvUydOhURyfb55cZSx8n97sfclC5dmqNHjzJr1iyKFy/O2LFjqVy5MufPn893GTkNGkpLS8uyvGjR\nIvr160fnzp1ZtWoV+/btY8SIEXke9/k5ni3RB+XBUQH1KWHJRxlKlChBmTJlOHr0KBUqVMj2Z2tr\ny7Vr1zhy5AhDhw6lRYsW5oEX9zqLul916tQhNjaWpKSkbG24/cs/MDCQI0eOZBlMc+zYMWJjY/Ms\nv3bt2hw8eDDHPt75Zejt7c0777zDokWLGD16ND/99JM5TavV0qhRI0aPHs3u3bspVaoUCxYsyLG+\nwMBAtm/fnuULOiIigri4OHPwKSidTkfXrl359ddfWbVqFTdv3uT111/PNf+WLVtwd3dnzJgx1KlT\nBx8fH86dO3dfdVryOLnXfrz9AyO/P0aef/55xo8fz4EDB0hMTGT58uUABAQEsHXr1iz5714uXrw4\nRqMxS/vuDuybNm2iRo0afPjhh9SoUYOKFSsSGRmZZ9vyczzn1YeCsrGxwWg0FqoMRT0289S4desW\nERERWb509Ho9lStXzjYK+O7lnIwbN44+ffrg4uJC27Ztsba25siRI6xevZqpU6fi4uKCh4cH06dP\np0KFCly9epXBgwej1+st1qdmzZrx3HPP8corrzBhwgSqVq3KjRs32LZtG3q9njfeeIOuXbsyYsQI\nevTowdixY0lMTOTjjz9Gr9fn+SNjzJgxtGzZko8//pju3bvj6OjIiRMnWLx4MT/88ANGo5HBgwfT\noUMHypcvT2xsLKtXryYwMBCA5cuXExkZSaNGjfDw8GD37t2cO3eOgICAHOvr168fkyZNIiQkhE8+\n+YQbN27w7rvv0rhx4wJferxTjx49+Pbbbxk1ahQvvfTSPR+X8vPzIyYmhlmzZhEcHMyWLVuy/FDI\nL0scJ3ntR29vb3O+2yOyc7rqMXPmTESEOnXqUKxYMdavX8+tW7fM5Xz88cd07NiRunXr0rp1a7Zs\n2cK8efOyHCfPPPMMjo6ODB06lGHDhnHq1CnGjBmTbd/NmjWLFStWEBgYyMqVK7ON1M1Jfo7nvPpQ\nUBUqVGDx4sUcPnyY4sWL4+Tk9MAfG3siPZpbt8rDFBISIhqNJtufv7+/iGQflHT3skjGiE9vb+8s\n65YtWyb169cXg8EgTk5OUr16dRk7dqw5fePGjVKtWjWxs7MTPz8/WbJkifj4+Mjo0aPNeTQaTZZB\nSrkZNWpUlkEdtyUlJcnQoUPF29tbbGxspGTJktK6dWsJCwsz59m7d6/Uq1dPbG1txdfXVxYtWiTF\nixeXiRMn5tmOzZs3y3PPPSeOjo5ib28v/v7+MmDAAElPT5fk5GTp0qWLeHt7i52dnRQvXlxef/11\n8yjeTZs2SbNmzcTDw0Ps7OzE19dXxo8fby47LCwsywhSEZHt27dL48aNRa/XS7FixaRr164SExNz\nz/2wefNm0Wq1EhUVled+rFGjhmi1WlmxYkWW9ZGRkaLVarMMSho+fLiUKFFC7O3t5YUXXpAFCxbk\nWU9ISIh59OhthT1O8tqPIiIffvihFC9eXDQajfTq1SvHtoWGhkqDBg3ExcVFDAaDVK1aVWbNmpUl\nz6RJk8TT01P0er20aNFC5syZk2VQkojIX3/9Jf7+/qLX6+XZZ5+Vf/75R7RarXlQUlpamrz11lvi\n6uoqTk5O0rVrV/nhhx9Eq9Wayyjo8ZyfPtwtp7ru/pyuX78ubdq0EWdnZ9FoNDJnzpx7lqnkrEgF\nVHt7e3FwcDD/6XQ6ef/9983p69atk8qVK4vBYJCmTZtm+x978ODB4ubmJm5ubjJkyJCH3XzlMXHm\nzBnRaDTmx1sUJTd3j8RWlHspUvdQ4+PjuXXrFrdu3eLSpUvo9Xpee+01AK5evcqrr77KuHHjuHHj\nBrVr16ZTp07mbadNm8by5cvZv38/+/fv588//2TatGmPqitKETJv3jzCwsI4c+YMGzdu5LXXXqN8\n+fK0bNnyUTdNUZQnSJEKqHdavHgxJUqU4NlnnwUyhvBXqVKFV199FRsbG0aNGkVERATHjx8HYM6c\nOQwcOJDSpUtTunRpBg4cyOzZsx9hD5Si4vr16/Tt2xd/f3+6dOlC+fLl2bRpk5p+TcmXhz03sfL4\nKrKDkubMmWOeUgwyntG789lIg8GAj48Phw4dwtfXl8OHD2dJDwoKyvLsnvL0+uCDD/jggw8edTOU\nx1BwcLAa/arkW5E8Q42KimLTpk1ZnpNLSEjAyckpSz4nJydu3boFZFwudnZ2zpKmHnhWFEVRHpYi\neYb666+/0qhRI/PE3QAODg7ZJuaOi4szz4Bzd3pcXBwODg7Zyvbx8eHUqVMPqOWKoijK46hixYr3\nfMlEfhTJM9S5c+dmm8UlMDCQiIgI83JCQgKnTp0yP/MXGBjIvn37zOkRERE5Pgx/6tQp83OWT+Lf\nyJEjH3kbVP9U31T/nry/J71/ljjRKnIBddu2bVy4cIGOHTtmWd++fXsOHjxIaGgoycnJjB49murV\nq+Pr6wtkPLQ+ceJELly4QHR0NBMnTiQkJOQR9EBRFEV5GhW5S75z587l1VdfzTbTibu7O0uWLKFf\nv35069aNevXqsXDhQnP6W2+9xenTp6latSqQ8T7CN99886G2XVEURXl6aUQk7wkwnyD5nYD8cbVh\nwwaCg4MfdTMemCe5f09y30D173H3pPfPErFBBVRFURTlqWeJ2FDk7qEqiqIoyuNIBVRFURRFsQAV\nUBVFURTFAlRAVRRFURQLUAFVURRFUSxABVRFURRFsQAVUBVFURTFAlRAVRRFURQLUAFVURRFUSxA\nBVRFURRFsQAVUBVFURTFAlRAVRRFURQLUAFVURRFUSxABVRFUZQC2rZtGy+91JmWLTuwfPnyB15f\nbGwsx44dIzk5+YHX9UgZjY+6BQWiAqqiKEoB7Ny5kxYtXmblyiasXduOLl368ccfix5YfZMn/0jJ\nkuWoXbsNnp4+7Nq164HV9UidOwe1a8Offz7qltw39T5URVGUAuje/U3mzfMDPspc8yc1akxkz54w\ni9c1YcI3DBnyObAX8AYW4+HxMZcvn0Gj0Vi0rvT0dGbPns2pU5HUrl2TV155xeJ15GrPHnjxRbh4\nEWrVgp07QftwzvssERusLNQWRVGUp0rGd++dX/baB/Jj/dixYwwfPgZoQUYwBehAXFxvYmNjcXFx\nsVhdJpOJ1q1fZdu2myQmNsXefhRbt+5i4sQvLVZHrv76Czp1goQEaNIEQkMfWjC1lMertYqiKEXE\ne+/1wmD4ApgF/IHB8B4DB75p8Xp2796NlVVdYBdwLXPtZmxsbHF2drZoXTt27OC//46RmLgGGEFC\nwgamTPmB2NhYi9YDsH//fhYuXMiePXvgxx+hbduMYNqtG/zzD7i6WrzOB02doSqKohRA/fr1+fvv\nRYwbN5mUlDTee+9rXnuto8XKj4+PZ+/evSQmJqLRnAM6A1UAH2APf/yxGK2Fz+Bu3ryJTlcasM5c\n44pO50B8fDzFihWzWD3fffc9n376Bda6hoxJ/oea6fEZCcOHw+jR8LAuMVuYuoeqKIpSxJw8eZKG\nDVuQnFyC9PQrODtbc/MmiHhjNG5n8uSvefPNvhav9/r16/j4VOXGjYxLzDrdNHx8VnP4cLhFgnd6\nejrDh49m/PiJ2MkefuUTXiWUNCB2wgQ8Bg0qdB0FZYnYoAKqoihKHi5fvsxXX33LhQtXadv2Obp0\n6fxAB+rUr9+SnTtbYzINAFLR65/n3XdrUatWLWrWrEnlypUfWN0HDhygW7d3iIo6TY0atZg/fxql\nS5e2SNl9+rzHb7/twDH5BivwoB47iMWZHvalGPLPDBo2bGiRegpCDUpSFEV5wG7cuEH16g24evUF\n0tPrsXLl55w+fZbhw4davK6kpCSmT59OREQEJtOUzLU2JCW1Ii0ths6dO1u8zrtVrVqViIgtFi/X\naDQyd+5MKqb/w980pwKniaIsbRjJWc1QfvHzs3idD5salKQoinIPS5Ys4ebNGqSnTwbeJDFxJV99\nNcHi9aSkpFCvXnOGDg0jKakkGYOdBLiFwbCEWrWCLF7nw9ZITGyjHRUwEo41zxDDabuPWbZsAW5u\nbo+6eYWmzlAVRVHuISUlBZPpztG0zqSnp1q8nr///pvTp3UkJy8FLpLxmMwMbG2NvPZaJ7p3727x\nOh8m3cKFrBEjVsSynDp00wRj6/ob547uw93d/VE3zyLUGaqiKMo9vPjii1hZ/QlMB7ah13ehU6eu\nFq/n1q1bgBegAUoDu9Bq4zh6dA+//PLTw5tcwdJE4PPPoVs3rEwmdtZvyMSGHrTtfJ29e/97YoIp\nqEFJiqIoedq3bx8ffPApV65c5YUXmvPll6OwsbGxaB1nz54lMLA28fGTgTrY2HxFvXrRbNz4t0Xr\neahSU+Gtt2D27IxHYb77Dvr3f9StypEa5VsAKqAqilJU7dixg969+3P58kWeffZZZs+eYtHnPx+q\n2Fh49VX491/Q62HBAnj55UfdqlypgFoAKqAqiqI8YFFR0KYNHD4MJUpkTHRfp86jbtU9WSI2qHuo\niqIoiuXs2gX16mUEU39/2L69yAdTS1EB9SlVvnx5Dh8+/EDKXrNmDbVr18bOzo5B95j5JDU1lVat\nWuHh4YGHh0eWtDNnzmBlZUWNGjXMf9evX8+SR0R47rnnsm1bWF27dsXT0xOtVktiYmKOeXr37n3P\n9Hnz5hEUFIS1tTVTpkzJknbw4EGeffZZatSoQUBAAKNHjzanhYSEUKZMGXOfv/zyIUxKriiWsmJF\nxsT2ly5Bs2awbRuUL/+oW/XQqID6hJk9e3aWL+jcPMhL3xUrVmTmzJn3DKYAOp2OwYMHs27duhzT\nXVxc2Lt3r/nP9a7Jsn/44QfKly+f79GPISEhbNy4Mc98ffv2Zd++fbmm//nnn2i12nvWW6NGDX7/\n/Xe6dOmSLd+gQYPo0qULe/fuJTw8nF9++cX8bkuNRsOwYcPMfR42bFi++qYoj8L169dZs2YN27dv\nRyZPhnbtIDERevaEVavgcb3/W0BFMqAuXLgQf39/HBwc8PHxYcuWjFk71q9fj5+fH/b29jRr1oyz\nZ89m2W7IkCG4u7vj7u7O0KGWm8Xk66+/pl+/fubly5cvU7JkSZKTkwtd9uXLl2nWrBm1a9emSpUq\nDBkyxJzWt29fPvroI3O+ChUqsH///nuWdz9D6+fNm0ft2rWpVKlStrOowqhYsSLVqlXDyurejznr\ndDqaNWtWoDdmnDhxgt9//52hQ4fm+4eBRqPJ1/4JDg7O9az32rVrjBkzhokTJ96z3sDAQPz9/dFq\ns7/Sy8vLy/z2jvj4eDQaDcWLFzen51RuUlIS1apVY8WKFQD8+++/+Pv7k5CQkGd/FOVBiIiIoGLF\nKrze8Ut2NWqNpn//jEdkRo+GX34BC4+CfixIEbNmzRopV66c7NixQ0RELly4INHR0RITEyNOTk6y\nePFiSUlJkUGDBkm9evXM202dOlUqV64s0dHREh0dLQEBATJ16tRs5Reky9evX5eSJUtKQkKCiIiM\nGTNGPvrooxzzdujQQapXr57tr0aNGpKcnJwtf3JyssTHx4uISGpqqjRr1kxWr14tIiJJSUkSFBQk\ny5Ytk+bNm8tPP/2UZ1t/+eUXGTVqVJ75ypcvL3369BERkcuXL0vp0qVl//792fIdPnw4x/5Ur15d\nBg8efM86Ro0aJQMHDsyzLZGRkeLu7p5tnY2NjdSsWVNq1aolX3/9tTnNaDRKkyZNJCIiIsdtcxMS\nEiIbNmzIV14REY1GY/7Mb+vUqZP8/fffuabnVOcPP/yQZd3Vq1clICBAPD09xWAwyI8//pglv7e3\nt1StWlXatWsnR44cMacdPXpUypYtKzt27BBvb2/Zt29fvvuiKJYWGFhPDEyVpbwsApKCRjb17fuo\nm1VglgiHRS6g1q9fX2bNmpVt/bRp06Rhw4bm5YSEBNHr9XLs2DHzdj///LM5fdasWVkC7m0F3Wlv\nvvmm/PTTT5KWliZly5aVkydPFqicu8XHx8vbb78t1apVk6CgIClevLh89dVX5vSjR4+Kvb29dOzY\nMdcy/vrrL3OQK1u2rJQsWdK8PHPmzBy3KV++vGzbts283LdvX5k8ebJF+nRbYQJqSkqKxMTEiIjI\nlStXpF69ejJjxgwRERk/fryMGDEi123vNGbMGPO+cHV1FR8fH/NyRETEPdt1d8D8/fffpUePHiIi\nYjKZRKPRmH8M5SangPrCCy/IN998IyIiFy9elMqVK5t/QEZHR5vzzZ07V8qVKydGo9G8bvbs2aLT\n6bKVqSgPWwV7d9lBdRGQ6xSTJnSX4cNHPOpmFZglAmqRmnrQaDSye/duXn75ZSpVqkRycjLt2rXj\n66+/5tChQ1SrVs2c12Aw4OPjw6FDh/D19eXw4cNZ0oOCgjh06JDF2vb+++/TtWtXPDw8CAgIoGLF\nijnm69ChA6dOncox7b///sPOzi7LuokTJxIbG8vOnTuxsbHhrbfeynIp+dChQzg7O3Pp0iWMRiM6\nnS5buW3atKFNmzYAzJkzh6ioKEaMGJFnn+SOS4sikuPl0MOHD9O1a86zwrRo0YIJEyw/pymAjY2N\neQYVDw8PunbtytatW+nTpw+bN29m//79zJ07l/T0dG7cuGG+HO7g4JClnOHDhzN8+HAAevXqRa9e\nvWjcuHGB2rRx40b+/fdfvL29zeuqVKnCqlWr8LvHxN5379ewsDBmz54NQMmSJWnWrBmbNm2ibt26\nWd7q0b17dwYMGEB0dDRlypQBMl42XaJECc6dO1egPiiKRRw+zOb0BEqzj0jK04b5nLPvy4Barz7q\nlj1SReoe6uXLl0lLS2PJkiVs2bKFffv2sXfvXj7//HMSEhJwcnLKkt/JySlzuq6Me1F33otzcnIi\nPj7eYm2rUqUKbm5uDBgwgPfeey/XfIsXL84ykObOv7uDKUBcXBylSpXCxsaG6Oholi9fbv4CjoyM\nZMCAAWzatIkKFSrw2Wef5dlOybjqkK98t7/UY2JiWLVqFU2bNs2WLyAgINf+5BVM89OO3MTExJCW\nlgZAYmIiy5cvp0aNGkDGoKCoqCgiIyPZsmULLi4unD59OlswLUybbue7M/+UKVM4d+4ckZGRREZG\nAhk/eO4VTHP6PAIDA1m1ahWQMd3c5s2bqVq1KgDR0dHmfP/88w9WVlZ4enoCsHTpUrZu3crBgwdZ\nuXIlq1evzldflOxOnTrFrFmzWLZsmfk4K8qMRiPDho2kTJlAKleuw/Llyx9dY/79Fxo0oHRKEhG2\nBprpEzlt8zxvv92Otm3bPrp2FQWFPse1oOvXr4tGo5G5c+ea1y1ZskRq1Kgh/fv3l3fffTdL/ipV\nqkhoaKiIiDg7O0t4eLg5LTw8XBwdHbPVAcjIkSPNf2FhYflu37x586R8+fL32at7i4qKkrp160qV\nKlWkVatW0rVrVxk9erSkpqZK3bp1ZcGCBSKScYk7ICDAfH81N7Nnz5bRo0fnWW/58uXlk08+kVq1\naomPj49MmTLFIv0REdm8ebN4eXmJk5OTODo6ipeXl6xZs0ZEMu51375cKyJSu3ZtKVWqlFhZWYmX\nl5f0zbwHs2TJEqlSpYpUq1ZNAgICZMiQIWIymbLVFRkZKR4eHvlqV0hIiGzcuDHPfO3btxcvLy/R\narXi6emXpMTjAAAgAElEQVQprVq1yjGfVqvNckm4evXqcvHiRRER+e2338TLy0vs7e3FxcVFvLy8\nzPdDDx48KI0bNzb3bezYseYynnvuOalatapUq1ZNGjdubL4UHBkZKWXKlJETJ06YyyhbtmyWS8RK\n/qxbt04MBnext+8uDg715JlnmklKSsqjbtY9DRkyXAyGhgJ7BFaJXl9CNm/e/FDqjouLk9df7y2e\nnn4y0ttfjFZWIiDSvr2k37wpZ86ckevXrz+UtlhSWFhYllhgiXBYpAKqiEiZMmVyDKjTp0/Pcg81\nPj4+yz3UBg0aZLmHOmPGDKlfv3628guz0/r06WO+96UoyuPJy8tPYJVkDEk1isHQPMdxG0WJp6e/\nwN7MNovAF/LBBx8/lLqbNn1RbG16yEjeuV253HrzTZH09IdS/8NiiYBapC75QsZ9ru+//56YmBhu\n3LjBd999x0svvUT79u05ePAgoaGhJCcnM3r0aKpXr46vry8APXr0YOLEiVy4cIHo6GgmTpxISEiI\nRdp04cIF/Pz8OHXq1D0v9yqKUvRdu3YJqJ25pCUlpSYXL17M17ZJSUn8/fffrFixgri4uGzpIsKM\nGbNo06YTISFvc+bMGYu02WAwAJfNyzrdJRwcDBYp+14SExP5b+MapqcaGcVPGNHykU01/gwOhhzG\nczz1Ch/XLSstLU3effddKVasmJQsWVL69+9vvhyzbt068fPzE71eL02bNpWoqKgs2w4ePFhcXV3F\n1dVVhgwZkmP5RbDLiqI8RE2bviTW1v0F0gSOi8HgJZs2bcpzu+vXr0vFikHi6PisODq2kOLFy8uZ\nM2ey5Bk1apwYDFUF5olWO1xcXEqbbwMUxrJly8RgKCnwheh0H4irq6ecP3++0OXmJfXyZQnTaERA\n4jHIC6wQB4eGsmzZsgde98NmidigJsdXFOWpcvXqVV588XXCwzdhbW3Ld999yzvvvHnPbZKSkqhb\ntwkHDwYCswANOt1YXnzxKMuWzTfnc3Iqwa1bW4BKANjahjBhQk0++OCDQrd7y5YtTJ78E5cvX6Rl\ny+Z8+OGH2NvbF7rcXEVGZkxwf/Qol7DiBfpxyPYCPj5n2LVrY46DLB9nlogNReqxGUVRlAfN3d2d\n7dvXkZqairW1db5mz+re/S0OH74BNCfjBeBgND7L6dNrs+QzmYyArXlZxBaj0WiRdp85c5a//goj\nObkn4eHhzJvXhN27N2VeDrawnTvhpZfgyhUkMJBd775Lnf3H6Fi+Ju+/P+uJC6YWU+hz3MfMU9hl\nRVEKwWQyiZWVrcCXAo0Fbgoki5VVO+nXL+vAoP79B4vB0EBgtWg0k8TR0SPbZeGCcnX1EtiZOS7I\nJAZDm1wnbimU0FARvT5jANJzz4nExhaquNjYWOnS5Q0pV66qNGnyonkgaVFjidigzlAVRVHyYGVl\nS3p6ByASKAEIfn5BTJjwW5Z83377BR4e37J06QQ8PFz5+ut/KVeunEXakJAQC9yeUEZDenpF85zQ\nFiEC//sffPxxxr9794apU8HauhBFCm3adGTXLi9SU+dw7twGGjRozvHjEdledvFEKHxcf7w8hV1W\nFKWQvvhighgMvgKTxNq6u3h5VZKbN29apGyTySQ7duyQ5cuXy7lz53LN9+KLncTWtrtAtMB60es9\n8pw+M9/S00X69TM/FiOffy6Sw3Pf92PdunVStWoDAX3mALCMop2cWsry5cst024LskRsUGeoiqIo\neRg2bBCVKnmzalUYpUtXYMCA/+Ho6FjockWEkJB3WLJkDTqdP0bjTkJD59OyZctseefPn05IyLus\nXVudYsXc+OmnWQQFBRW6DcTHQ+fOsHJlxhtiZs/OWC6E3bt307ZtZxITvwXeBOKBYoAgcuOJvQer\nRvkqiqI8ImvXrqV9+/4kJIQD9sAGXFy6cP36hYfTgIsX4cUXYc8ecHGB5cuhUaNCFzto0DC++cYG\nGA18CGwBemNntxFf33OEh2/Apoi93s0SsaHITeygKIrytDhz5gwiz5ARTAEaExt7hdTU1AdW56ZN\nm/Dyqkx1K1sulvfOCKYVKsB//1kkmAIYDHbodDcyl74DmmFv/zmffBLEtm1ri1wwtRR1hqooipIP\nRqORDRs2EBsbS/369bO8Geh+iAh79uzhypUraLVaXnmlF4mJW4AKaDRT8fb+kVOn9lu28Zmio6Op\nXLk69RI+ZAkTcOYme+0MVI+KRHPHS+4L69y5cwQFPcPNm90xmTwxGL5m2rSv6NYt5zdXFQXqOVRF\nUZSHID09nRYt2rFr11m0Wm9E3mXNmuXUq1fvvsoREXr1epdFi1Zhbe2L0biPN9/swY8/BqHT2ePi\n4sRff/35gHoBO3fupKfRk/8xCmvSWUQH+prWcUKjwcOC9ZQpU4a9e7fxv/9NITb2CJ07z+D555+3\nYA1FkzpDVRRFuQeTyUTnzl1ZvDgSk2kLGechi/HxGceJE3vvq6zVq1fTocPHJCTsJOMy7z94eLzF\nmTOHuXHjBiVLlszxnccWIUJUSAjl5s4FYAKDGMq7WFkHcvPmtSd2oFB+qXuoiqIoD1i/fh+zdOkW\nTKam/P9FvUZcvHj/L3mPjIzEZGrI/98zbc7Vq+ewtbXF09PzwQXTlBTo1o1yc+diBPrblGe4TSp6\nQxO+/PKL+w6mcXFxhIS8Q0BAfdq165rlPb5PM3XJV1GUx9ahQ4f4/fdF2NhY07NnD8qUKWPR8j//\nfDw//TQFmAd8AvQDSqPVfkfNmnXzVYbRaOTzz8ezePHfWFkJImeAc0AZNJqf8fEJenCBFOD6dWjX\nDjZvBgcHNAsX0jAhgTJnz1KnzlyaNGlyX8WJCC1atCMiogKpqV9z4sQ/7N7djKNH9zzYuYUfB4V+\nkvUx8xR2WVGeSNu2bRODwV202iFiZfWeODuXlFOnTlms/FWrVoleXyFzYoIYgfGZ/7aX0qV95cKF\nC/kq58MPh2S+HDxM4GextnYSa2t7MRhKS+nSPuYXzxfGxYsX5dVXu0tgYAMJCXlHYm9PF3jypEjl\nyhkzKpQuLbJ3b6HrioqKEr2+hED6HZM11JN///230GU/SpaIDeoMVVGUx9KgQWNJTPwaCMFkglu3\nXPjqq++YPv17i5S/bdt/JCV1AW4B7YFBwEcUKzaTPXs2UaJEiXyV88svc0hM3ApUAIIR2c+IER70\n6NEDT09PrKwK9zWclJREvXrNiY5+kfT0NzhxYg4HD77Mjv+NQ9uuHVy9CkFB8Ndf4OVVqLoArK2t\nMZlSgVRAD5gwmRKe2Edh7ocKqIqiPJbi4m4C/z9PrslUjhs37v++Zm7KlPHCYPidxMS/gUnAOBwc\nzrNnz7Z8B1MAnc4aSDAva7UJ2NuXt9gcv7t27eL6dT3p6eMBSE1tSKX97tC8eca90+efhz/+ACcn\ni9RXqlQpWrduzZo1bUlM7Iqd3RoqVXKmbt38XQJ/kqlBSYqiPJY6d34Zg2EocBjYicHwJZ06vWSx\n8nv27En16locHOrj6PgfDg6RrF27BG9v7/sq55NPPsZg6ADMRKcbhoPDP3Tp0qXQ7YuOjiYsLIyr\nV68ikgSYAOFjJvBbaizalBTo2xf+/NNiwfS2RYvmMGpUa9q1W89HH/myefNqrAsxif6TQj02oyjK\nYyMsLIwZM37Dzs6GDz98mwULljBjxlysrKwZPvzjPF8Ufr/S09NZv349cXFxNGzYEE9PzwKVs2DB\nQhYsWE7x4i6MGDGUsmXLFqpdv/22kDfe6IeNTQApKYdxd/fg+pUgvk69yrv8C4B8+SWaIUMgH+97\nfVRiY2PZv38/rq6uBAYG5uvdtA+KRWJDoe/CPmaewi4ryhNhxYoVoteXFJgsGs3nYm/vLgcPHnzo\n7Zg+fYZUrfqs1KgRLKGhoXnmj4uLk0aNWom1tYNYWenl7bc/FFMB3+RiMpnkm2++E43GILA/c1BQ\npLjbusiBcuVFQNKsrCRt3rwClf8w7dmzR4oVKyXOzvXFYPCSbt36Fni/WIIlYsNTF11UQFWUx1ON\nGsECS80jSzWasdK3b7+H2oaZM38Rg6GSwD8Cy8VgKC2rVq3KNX9iYqLUr99crKy6Zb7C7LoYDHVk\nxoyCvRj8228niZ1dBYGK5v1QmvMSobXPWHBzE9mypaDde6h8fKoL/JrZj3ixt6+erx8oD4olYoO6\nh6ooymMhY8L4/78XKOJIcvKDm0Q+Jz/++CuJid8BLYG2JCaOZPr03zhy5Ahbt27l5s2b5rzXrl3D\n17cG//13gPT0D8kYA+pCYmIIGzfuKFD9338/k+TkaUAssIWq7Gc7tQgyJZDu7Z0xwX3DhoXv6ENw\n9uwJ4PY9b3tSUppx/PjxR9mkQlMBVVGUx8Lbb3fDYOgHrAdC0eu/pHfvwr23M7/CwsLo1+8jLl++\nBNy8IyWOvXv3UatWc9q0+Qhv7wAiIiKIi4vD378658+XBeoCmzPzC7a2W6hYMf8TUKSmprJr1y72\n7t2LVqsjIzDPpyUvsIXqlOEyMZUrY7VzJ1SqZKkuP3C+vlXRaOZmLl3D1vYvqlWr9kjbVFhqUJKi\nKEXWnDm/Mn36AgwGO0aMGMD+/YeYOnUeNjY2jBnzMS+88MIDb8PChb/Tp89HJCZ+gFa7DZMpDBgD\nJGFr+zUajTvJyXsAB2AOlSpN4rXX2vLVV8sxGlsCfYBmQBXgPIGB9mzfvh4HB4c867527RoNG7Yk\nOjoZkTScnY1cuJDAG7TmJ+ZihYmrLVrgvmIFPGZz8R4/fpzg4DbcugVpaVfp1+89vvlm3CNrjyVi\ngwqoiqIUST//PJMPP/yKxMQJwA0MhiFs2PA3derUeajtKFeuCmfPTgEypujT6V6iatVrVKxYAZ0u\njdBQL9LTv83MHYeNjSft27/G77+7AfOBFYArEEK9elo2bPgHW1vbfNXdo8dbLFxoQ1raZMCElo58\nzn8M4xIAE3Ql8Zj2Ob369LFspx+S1NRUTp8+jYuLy3092/sgqNe3KYryxJo48WcSE6eRcXYHiYkx\n/Pzz3DwDalJSEocOHcLe3h4/P79CP4qRmJgAlDQvm0zVMRpX8fff6wF70tO3A5+SETTnkpZmxaJF\n87G29iUtbSTQDriKp2cF/vlne76DKcCBA8dISxsOaLAljdlc4nUukY6Od/iJGcYLDD15ulD9e5Rs\nbGzw8/N71M2wGHUPVVGUIikjEKbfsSY9z+B45swZKlWqRvPmfahduwXt23fFaDQWqh2dO3fEYHgH\niAD+xNr6J06cuEVS0gmSkk4AVdBoygJewFhEtmMynUTkEjrdB+h0V2nZsjXHjoXjdJ8TLNSsWQVb\n2wW4cYV1NOd1/uMmOl5gJTN4FXv7UGrVqnFfZSYkJBAeHs7Jkyfva7v8MplMjBgxlpIlffDy8mfa\ntJ8fSD1FUqHHCT9mnsIuK8pjae7ceWIwlBWYKzBJ7O3dZd++fffcplGjNqLVfpn5KEaSGAyNZfr0\n6YVqR2pqqgwYMFS8vALEz6+u9OjRQ3S6webHVuCqgE3m32XzeiurAfLFF19IQkJCgeuOjY2Vtv61\n5ITGSgTkko2ttPb0EYPBS2xsHKVfv4H5enZzzZo18sknn8rw4cPFzc1LnJyqi15fQnr3fs/iz35+\n+eU3YjDUFjggsF0MBm9ZvHiJRet4ECwRG5666KICqqI8PpYsCZVWrTrKK690l127duWaLykpSd56\nq5/odC4Cx+8IduPl/fc/smibli1bJvb2VQRiM+uYJOAkEHDHc7IpYmtbSxYsWFC4yrZsEZObmwhI\nkr+/mM6dk/T0dDl9+rTExMTkubnJZJLWrdsKuAt8ItBMoJJAssBNsbevZvFnP6tWfVZg/R2fwc/y\nyis9LFrHg2CJ2KAu+SqKUiSEhobSoEFrGjZsw59//gnAK6+0Z9WqP1iyZC61atXKcTuj0UiFCtWY\nNu0IRmNlMgYCASRiMKygWrUAi7azbdu29OjRAju7ilhZVQA+J+PS9GfAm0AboBLVqzvRsWPHglf0\n++/QvDmaa9egTRvsduxA4+WFTqfD29sbd3f3PItYtGgRq1b9A2wFxgHryLjXuxxwJDn5eY4cOVLw\nNubA2dmRjPe9ZtBqz+Hq6mjROoosCwT2x8pT2GVFKfKWLl0qBoOXwCKB30WvLyV//fVXvradPHmy\ngJ1AkkCUgJ9AWbGx8ZCOHXuI0WgscLuMRqNMmDBRGjd+Sbp06SNRUVHmtKioKPH3ryPwp8AsgZIC\nHUWrLS2NGrXIsd7jx4/LSy+9LrVqNZPhw8dKWlpa9kpNJpGvvhLzKd7bb4vklC8f3nrrfQFd5hnp\n7SJfE5guECf29lVl6dKlBSo7N1u3bhWDwV00mqGi070vTk4l5OTJkxat40GwRGx46qKLCqiKUvQ0\nafKSwII7vvRny/PPd8jXtm+/3U/AkBlQJTN4+MnMmTPzdX/w4MGD0rBhKylfPkh69nxbbt26ZU57\n772PxGBoIBAqOt0IcXcvk+VS69ChI0SvbyVwSyBMrK0rSu/eOc9Je/HiRSlWrJRotV8JrBaDoan0\n6vVO1kypqSJ9+/5/MP3664wAW0BffvmVaDTlBd4QOJ8Z/A3i4FBJ7Ozc5a23+j+Q+XMPHDggn302\nQkaPHpPlR0hRpgJqAaiAqihFT7Nm7QRm3xFQp8kLL3TK17YzZswQrdZToI3AMoH3xdbWTeLj4/Pc\n9tKlS+LsXFI0mikCu8XWtrO0aNFORDLuP1pb6wWumNtlb99BZs2aZd4+JSVFOnUKEZ3ORnQ6G+nb\n9/0cz0zXr18v7u5eAm3v6OM10WisZdSocRlnqnFxIs8/n5FoZyeyaFE+917ubt26JZUr1xCdroyA\nk2g0zvL999/Lnj17JDIyssDlPspJ7B+UJzKgNmnSROzs7MTBwUEcHBzEz8/PnLZu3TqpXLmyGAwG\nadq0abZfPoMHDxY3Nzdxc3OTIUOG5Fi+CqiKUvSsXbtW9PriAlMFpojB4CEbNmzI17bp6enSrl1n\n0emKiVZbSgyG4hIeHp6vbX/77TdxcGh/R5BLEZ3OVr74Yrw0bfqyaDQ2AjHmdIPhNZk5M/vE9ikp\nKZKamppjHYcPHxaDwV2gv0C7O+q6JGAnWu2z8t7Lr4lUrZqR4O4u8t9/+Wp/fiQlJcnSpUtl/vz5\nEh0dXaiyYmJipFGj1qLT2YiLS2lZtGixhVr56D2RATU4ODjHAzYmJkacnZ1l8eLFkpKSIoMGDZJ6\n9eqZ06dOnSqVK1eW6OhoiY6OloCAAJk6dWq2clRAVZSi6d9//5V27brJK690l02bNt3XtiaTSY4d\nOya7d++WpKSkfG8XGhoqDg6NBUyZQe6KgJVoNDUF/hB4RqC2wArR6caIq6unXLlyJd/lT536s+h0\nNgJdBa4JlBMYnHl5u47AYKnGVjl/O8r6+ooU4fuNjRq1FmvrDwQSBLaLXl9c9u7d+6ibZRFPbECd\nMWNGtvXTpk2Thg0bmpcTEhJEr9fLsWPHRESkfv368vPPP5vTZ82alSXg3qYCqqIotyUmJoqvbw2x\nte0m8L1YWflnDnC6mhlg08Xauqr4+NSSjh173tdl0u3bt4vBUFrgG4EWmUE7WqCLQDGBKdKalXIT\nBxGQ9IYNRa5de3CdLSSTyZT54yDBfJZta/uuTJo06VE3zSIsERuK5GMzw4YNw8PDg2effZaNGzcC\ncOjQoSxvIjAYDPj4+HDo0CEADh8+nCU9KCjInKYoipITvV5PePgGhg715dVXd2A0HgdMgC4zhw6t\ntiKfffY+f/wxm/Lly+e77O3bt2M0tgPeBW4ArwA/odGsBeAt/uRP2uJIPKtdPdCtXw+urpbsnkVp\nNBocHd2Ag5lrTFhZHcTDw8NidRw5coT+/Qfy7rsfsnPnTouV+7AUuYA6fvx4IiMjuXDhAm+++SYv\nvfQSp0+fJiEhIdu0XU5OTty6dQuA+Ph4nJ2ds6TFx8c/1LYrivJgpKamMmbMFzz/fEcGDBhCXFyc\nxcp2dHTk7NmL/PnnX2TMjd6NjOD3NzAaK6sttGnT5r7L9fT0xMoqnIwp0zcAXuj1P/DLzPFMsktn\nKqvRYeInt5IEn4+C+5jj91GZPn0yev1L2Nq+h719UwIDdXTo0MEiZR88eJA6dRozebI9P/1UgqZN\nX2TDhg0WKfthKXKT49etW9f87x49erBgwQL+/vtvHBwcsry8FyAuLg5Hx4wHhu9Oj4uLy9frkRRF\nKfpefbU769ffIimpBxs3rmHNmhbs3bsFGxubQpe9YMEC/vhjJ6mpUcBA4AhQARiMtfVFNmz4J19n\nYcnJySxcuJBr167RtGlT2rdvz8yZC9iypRYajT9G478sWzCHFr/+CsnxmHQ6Lo8ezduffFLoCfwt\nLTY2FltbW/R6fZb1HTt2oFIlHzZt2oS7e0M6duyItbW1Rer86qtJJCYOBIYAkJjoxfDhX7N5c7BF\nyn8YilxAzU1gYCBz5swxLyckJHDq1CkCAwPN6fv27aN27doAREREUKVKlRzLGjVqlPnfwcHBBAcH\nP7B2K4pSOJcuXWLt2rWkpFwA7EhJ6cS5c7X477//2LVrH2PHfkVqajKdO3dh6tT/ZfuCT0hIYNKk\nScTEXKdly+a0bt06S/revftJSGgPOAJTgKHodHNo27YV3377J97e3nm2MTk5mWeeacbJkw6kp/uj\n07Vm7twf+euvRaxfv579+/ejvVaRGgMHwokT4OSEdskSSj333H3vj5s3bzJ69JccOXKaBg1qMGTI\nxxYLarGxsbRu3YHdu3cgks4HHwzgm2/GZQn41atXp3r16hap704JCcmI3Dn7kzuJiUkWr+e2DRs2\nWP4M2AL3ci0mNjZWVq9eLUlJSZKWlibz5s0Te3t7OXHihHmU75IlSyQpKUkGDRok9evXN287depU\n8ff3l+joaDl//rwEBATItGnTstVRxLqsKEoezp8/L3Z27gJp5sEwjo7PSL9+/cTWtoLAYYGLote3\nkI8+GpZl24sXL2Y+jtNG4HOxsioj33zzXZY8M2fOFIPhWfNsQhrNJKlTp9l9tfGXX34Re/vbA49E\nYJu4u5cVkYxHgqrZFZNTGkcRkAvWtpJ8j3mJ7yUlJUUCA+uKrW1PgfliMDwvbdu+XqCyctKhQw+x\nsekrkC4QIwZDkPz2228WK/9eVqxYIQZDGYE1ApvF3j5Afvwx+3f4g2KJ2FCkoktMTIzUqVNHHB0d\npVixYlK/fn1Zt26dOX3dunXi5+cner0+1+dQXV1dxdXVVT2HqihPCJPJJM8++7zY2XUWWCvW1gPF\n3b2caLUZI2X//7nO7VKxYk3zdkajUapUqSVQ/45Ad0qsrAxZJiZIT0+Xl17qJAZDOXFyqiMeHuXM\nTw/k17fffis2Nh/c0ZZYsbGxFxGRDsXLybXMkby7qCkV9MEFfgPOhg0bxMGh+h39SRJb22Jy8eLF\nApV3t1KlfAUO3dGPb+Sdd/pbpOz8mD//N6lcua5UrFhT/ve/7x/qBBKWiA1F6pKvu7v7PUd2NW/e\n/J4TOY8fP57x48c/iKYpivKIaDQaVq9ewqBBw9m+fRw+PmVZseIaJlMn4OgdOY/i4eEGZFyCbd68\nLQcPHiZjgNHtS5alMRpTMRqNWFllfP3pdDqWL1/AgQMHuHXrFtWqVcvX+IsLFy4QHh6Ou7s7TZo0\nQadrA7wOBGBjM4jg4Jbw22/MuxKFLbCCl+jCbySljOPy5csF2hdGoxGNxu6O/lih0VgV+J2vly5d\nol+/wRw5cpLatYMoXbo0ly5tRiQAMGFntxVv7wYFKrsgunTpTJcunR9afZamyYzMTw2NRsNT1mVF\neaKcPXsWf//6JCbuAepl/tljZxfKxo2rqVu3LqNGfc748XtITr5ExiCj6UBNYASVKh3i+PF9hWrD\nvHnzeOOND9DpaqPRRPHMM76kpSWzY8ceTKZkmjVtybK6gejHjQPgR20V3jftxEQUBkMLVq2aT+PG\njbOVazQac3yi4bbExET8/Wtx4cLLpKe3xNZ2FrVrx7B58+r7HtiUnJyMv38tzp9/kfT0F7G1nUuF\nCnu4ePEiRmNN4AoVK9qwbdvabIOTnkQWiQ2FPsd9zDyFXVaUJ0paWpq4u5fJnMnoqsBQsba2zzJV\nYfv23SXjDTAHBNwESgk4ibOzV5bLo6tXr5bPPhsuP/74oyQnJ2era+3atdKqVXt5+eXO5tmbVq5c\nKeAosNJ82TXjDTfdBH4UJ31JOfPcc5J5Q1bix42TJo1bi1ZrJQZDMZk69eds9YhkvFDdzs5JrKwM\nUrFikJw6dSrHfBcvXpTXXguRoKDG8tZb/bNM5n8/Nm/eLI6ONe+4fGwUg8FTdu7cKaGhobJ69epc\np1N8ElkiNjx10UUFVEUpmuLj4+XTT0dK+/bdZcKEb3N+tVmmXbt2ibt7GbGzcxO9vpgsXrwkS/q4\ncV+JXt9aIEXgslhZtZImTVpKSkqKOc+ECRPFYPAWGCF6fWupXbtJlgDyv//9TzLeYvO1wGSxtnaV\ntWvXiptbGYGsMwbBWwLfiROxsoZqGSv1epE7Xt6dnp6e6z3BAwcOZA6eOiBgEo3mG/H1rZljXkvZ\nvn27ODj4CxjNPwzs7Nwfm7fDWJoKqAWgAqqiFD1paWlSo8azYmvbSWCWGAzN5ZVXut5zm/T0dLl4\n8WKOZ1EpKSnSsmU70etLib29t1Sp8oxcu2Nav/T0dLGxMQicMZ+dOTjUl2XLlomIyNWrVzMHPU26\nI2jOk9q1m4lOZy1QT2B85tndWYESUoZFcoBAEZAYnbXIjh357v+sWbPE3r77HXUZRau1vq95ifOy\nadMm+eijwTJ27Ody5coVSUtLk1q1GoudXSeB2aLXt5I2bTo8kW+SyQ9LxAZ1D1VRlEdu69attGr1\nNvHxEWRM4JaEra0nZ84cpmTJkgUqU0Q4ffo0qamp+Pr6otPpzGnJyck4ODhjNCZw+3F8B4fO/Phj\nG8OxohoAACAASURBVJKSUnn//f6kptoAXwN9Mrdajq/vOFxcnNi1ywejcStwEbhJTbSsRE8pYjmC\nNT3di7P98lm02vxNRvfPP//QocNA4uPDATsgHAeH1ty8GWORSR/++GMRvXr1JzHxXayto3B1Xc+B\nAzswGAyMGzeBQ4dO8swzQQwa9JHFnml93Kh7qAXwFHZZUYq8sLAwcXJ65o4ztHTR60sU+PKj0WiU\nKVN+ko4dQ2TYsOESFxeXLU+DBi3E2vrdzDPMxeLg4CFr164Vg6GEwEEBJ4GSAqECfwmUlK++Gi+t\nW7+SmaYRjcZWprRqLfFoRED+xVmK8Z3Y2RWXc+fO5dq+/fv3y8SJE2XGjBkSHx8vJpNJXnmlmzg4\nBIijYycxGDwkNHRpgfp+p0uXLknLlq9knm1vMO9fG5teMn78+EKX/ySxRGwoUo/NKIry+EpMTERE\n+D/2zjM+qqIL48+92+9uNhVIJYSahI5AhEgX6QJSBJHyCgIivQlIE1QEFWkKCCJgAaVJEUFCBxOQ\n0Lt0CKFDQrJpu/u8H3ZZExNIW5SQ+/8iuXfmzJzrjzzMzJlz9Hp9rvvWrFkTRuN9JCaOg8XSFGr1\nIlSoEIKAgIAn9tu+fTvOnDmD0NDQDFGzffsOxg8//AmTqSc0mt1Ys6YhDh3aA61W62izbt0ydOv2\nDvbuDUPRoj5YvPgXXL58GQpFXQDlAayC7cpNHwiCBV26vApSxM6d8QBuACAGiPXQZ9NvUABYii7o\nhUVIQwLU1nGP/Q6bNm1Cu3bdYDa/DqXyIqZN+xLR0buwcuVSbN++HbGxsahRYxLKli2b6++Ynm3b\ntuG117ohPv51kAYAvo53aWl+iI+Xc507nfzresGiELosI/NUSUtLY+fOb1Gp1FKp1LJ1684Zgn9y\nyrVr19i69RsMDg5j9+59+eDBgye2Hzp0NPX60tTpelGvD+L7739A0hbcpFRqCTywr8isdHGpxV9/\n/TXbOURGRlKvL0Hgnr3v79RqjY7I4JYtOxNYShFmfo4hj5bTXFSiDCVdIwIfUa+vzAEDhj92jMDA\n8gQ2O+am073GWbNm5eJLZc/69eup1RaxRzhbaStu/gptWaV+o05XlPtyccZbGHCGNhQ6dZEFVUbG\nuUye/AklqRGBBAIm6nQtOHLkuKc65vnz5+2CcZePCoNrNO68du0aHzx4QJVKTyDVscXp4tKUa9bk\nbAt10KD3KEl+dHVtTEnyyiDEw4ePoZv6Da5CWxJgCkTOrFaLqampnDt3LocNG8nly5c/MbDHaPS2\nbzM/ulkzluPHT8jvJ8lAlSr1CCy1X++5Z492HkxBcKOfXwjXr1/v1PGeB2RBzQOyoMrIOJeGDdsQ\nWJHu/HMja9Zs/FTHjIyMpKtr9XRjkkZjBR46dIgk2aBBS2o0bxCIpChOo5dXQIYo3+w4cuQIN27c\nmOkc9OG5czyi05MA70PBDl5+vHbtWq7m3q5dV2o0XexCF01J8nXccXUWISEvEthGYCiBKgSmUKGo\nz7p1m9FsNjt1rOcFZ2jDM1cPVUZGpmBRqlQAVKo9jp8Vir0ICvLPtZ2rV68iIiICFy5cyLZtSEgI\ngGuwnXNaACyDUnkfZcqUAWA7H+3SxR1lygxAo0aRiIraDo9cFO+uVKkSmjVrBn//dH6cOgXDyy+j\nUlIikosVw6mF87Ho4mn4+fnlys9vv/0Sr7xigVodAHf3Vvjqq09Qp06dXNlIj8ViwcqVKzF79mwc\nOHAAANCnTxfo9QMBtADQEArFFPTvXxkREWszRDvLOBknCHuBohC6LCPzVLl16xaLFw+mi0sDurg0\nprd3yVyv2r777gfqdJ50da1Pnc6LM2bMybbPvn376OtbhoIgMiAgmNHR0Xl1IRMbN27kW2/14/Dh\noxgTE0Nu3066udmWwjVqkE5KRp9f4uPjGRbWgHp9TWo071Cn8+aiRYtptVo5Y8ZshoS8yCpV6nLd\nunX/9VSfeZyhDfI9VBkZmXyTkJCArVu3giQaNmz42Fy0WfHgwQP4+AQhOXkPbNG1l6HVvoDTp6MR\nGBiYbX+LxeJYdaWkpOCzz75AdPRJVKkSjPfeGwaNRvPE/gcPHkRkZCR8fHzQunVrLFnyHQYMmACT\naSgUikvorVuKL1MeQkhLA9q0AX74AZCkJ87n448/xerVm+Dl5Y7PPpuIypUr5/h75JT9+/ejYcOm\nSEwsCuA4bPdpT0GrDYPJFPfMFS1/1pHvoeaBQuiyjMwzzfHjx+niUi7Deaira3iG3Lw5wWq1skGD\nltTpWhBYRJ3uVdat24wWi+WxfRYvXkpJ8qZW24cGQ002aNCSRYuWIhBFwMpx+ODvSQ0eTObg/HHg\nwBGUpHACWwjMocFQhBcuXMiVL0/CbDZz8eLFlCQvAv0JdMmQYUmhUNNkMjltvMKCM7Sh0KmLLKgy\nMnnn8OHD/Oyzz/jNN9847Zf2w4cPaTB4EdhpF4VDlCRPXr9+nUlJSbx06VKOruGcOnWKkuSfLro3\nlXp9II8fP55le6vVSq3WaE/iQHu/MgQ0VOEEv0V3EqAZAn9t2izH/th8uZwuiUJffv755znu/ySs\nViubNWtHSXqRgIbAGQKPvl0SRXEMK1Wq7ZSxChvO0AY5KElGRiZHrFu3DrVqNcaYMZcxYMAKvPBC\nXZhMpnzbNRgMWL36RxgM7WAwlIZO1wCLF89HVNQ+eHr6IjQ0HEWKBGDXrl1PtJOWlgZR1OFRKkFb\nrVAdUlNTs2yfkpKCtLRkACH2JyoANeCK2tiEWuiBJUiEBp3UBvhP/STH/igUSgDJjp8FIdlpgUD7\n9+/Hrl1HYDLtAOAO4AqAJQC6ADCgXLnf8dtvK5wylkwecIKwFygKocsyMk7Bx6c0ge2OhASS1ILz\n5893mv3ExESePn2a8fHxjImJoSR5EvjTPt5murgU5dy5c9m79wB+/vn0TOXW0tLSGBz8AlWqgQT2\nUqUawuLFy7J//yEcNuw9njp1KtOYFSvWokIxnkAygT0MhDtPoCQJ8DpEdi5bldu2bcvR/A8ePMgm\nTdrRz68cVaqyBJZQoXifHh5+GUrG5YfNmzfT1bWB/ZtsJ1CEQCg1Gi+OGjXBKWMUVpyhDYVOXWRB\nlZHJHfHx8ezbdzBFUSJw07GVqVCM4Mcff/xUxty6dStdXetmOFdVKgOo01UkMJ06XQvWrt04053K\n27dvs2PHHixXriYbNGhGnc6TwEQKwvvU6704c+ZMLlu2jNevXydpy85UrVpdAiKrw4WxsEXyHoMX\nB7Z+PcfzPXv2LPV6LwJzCGygSlWCZcq8wLfe6ufUcmh3796lm5sPgW8JXKcojqSvbymeOXPGaWMU\nVmRBzQOyoMrI5ByLxcKwsIbUaLoSaEDgDQL3CeyjJHkzKirqqYz7119/UacrQiDGLqin7GeGjzIM\nmWkwhHLPnj2PtdGwYWsCC9OJ8lQqFAF0cWlLo7GYIwlEZGQkO6iNTLQ3jBCMDPYOcohuTpg0aTIV\nisHpxjpJT8/i+f4OWXH48GGGhobRYCjC8PAmT0zCL5NznKENcnJ8GRmZx3Lx4kUcO3YGKSm/A0gE\n0AtAMRiNHpg37wuEhYVl6pOQkACtVgulMu+/XkqXLo1x40Zi8uRqUKmqIDX1ACwWHdLSHiVRUEAU\nvZ54hpuQYAJQNN0TH1gstfHw4XIAC9Gs2evYsOFH/BzeCMutJogAFqEB3hWu4rOxg+Hj45Pj+SoU\nCghCWronqRDFp5NAoXLlyjhxIuqp2JbJH3JQkoyMzGMRRRGkBYAVgBHAchgMpbFly1p07twpQ9u7\nd++iZs2GcHcvCkky4qOPpuVr7NGjhyMycjO+/bY3jh6NRHBwMFSqoQBOQBS/gFp9CTVr1nxs/7fe\n6gidbiSASAA7AYwB0MH+Nhy3bjxAVNhLmG4X07GYjJ7YimTr2zhzJvtsTQBw9OhRjBgxGteuXYdW\n+zNE8SMAP0KSOmHEiAH5cf+pkJycjJUrV2LJkiW4evXqfz2d5w8nrJQLFIXQZRmZPGO729mCOl1b\nAj9To+nGSpVsyeD/SdOm7ahS9SdgJnCNklQ6RxVessJisbB374FUKjVUKNRs1ep1Xr16la++2pm+\nvuVYp04znj179ok2Lly4QK3WjUAAAT/79ZIbBNIooRt/gT8JMBngG+ho36pNIlA9R8FWkZGR9rug\nYymKIylJ7mzRogObNu3AxYuXPjFB/n9BQkICQ0Kq02CoR72+Mw2GIty/f/9/Pa1nBmdoQ6FTF1lQ\nZWRyR1JSEkePHs9Gjdpy8OCRjI+Pz7Kdm5tvhvuXwASOGTP2ibbv37/PTZs2cdeuXRkCjGbOnENJ\nqmU/r02iTteGAweOzNKG1Wrlvn37+Ouvv/LGjRuO5yNGjKYoDnNEJQNdCShZDBr+CSMJ8C50bCDq\nCPgSKE/Ai0FBFXOUQN52RrvA4a8gTOEbb/TMtt8/MZvNnDRpCqtWrc/GjdvyyJEjubaRE6ZOnUaN\npr39W5DA96xUKfypjFUQcYY2yGeoMjIyT0Sr1eLjjz/Itl2xYr548CAKQHEAVuh0+xAQ0Pqx7bds\n2YLmzTvAYikNhSIOVav6YffuzdBoNNi69Q+YTH0BuAEAkpIGYdu2sZlskESnTv/Dr7/uhkJRClbr\nYfz222q89NJLMJmSYbUWsbcUAAxEDel3rEi6j0DG4wK80VZjwbuzPoT5+7W4fj0GTZp0wcyZn+Xo\n3ujDh4lIX7Sb9EVc3JFs+/2ToUNHY+HCP2AyTYAgnEF4+Ms4dmw/SpQokWtbT+LatRtISakB27cA\ngOq4efOGU8co9ORf1wsWhdBlGZkcYbVa+fXXC9m8+evs2fPdXEePRkVF0WAoQheXdjQYarBmzQaZ\n7oo+4vz58xRFdwJfOKJ2BeFldu78BocOHcmGDZtSperrWP2J4hQ2b94xk51ffvmFBkMVAiZ723UU\nRSPd3HzZpk0HiqIrgV8IRLIhSjBeoSABRqu0rFmiIlesWJGnb0WSs2d/RUmqaL8ru4eSVJI//fRz\nru1kzqzUh9OnT8/zvB7HmjVrKEll7ZHSKdRourNjxx5OH6eg4gxtKHTqIguqjEzWjB37ASWpMoHv\nqFCMoqenP2/dupUrG1evXuWPP/7IDRs2ZHnOSpLXr1+nq2tRAh4ETqTbIp5OhcKbwEfU6WpTpytC\nF5dGdHFpRU9Pf54/fz6TrS+++IIaTf90NkwElATOU6OpS4XCnUA4uyOQqRBtRcFffZV0QtpEq9XK\nqVM/p79/KIsXr8D58xfkyY7RWIy2FII2H7Ta7pw5c2a+55cVkyd/QrVaoiiq2KjRq4yLi3sq4xRE\nZEHNA7KgyshkjSS5E7jo+MWu03Xm3LlznT5O//5DqVB0JOBNYAABC4EHBEIIfGYfP5mSVJyffvop\nf/rpJ965cydLWzt37qQkBRK4Zu/3OYEw+5//JODFDzDOkR1iulLitStXnji/+Ph4Xr9+/akEFd26\ndYv167ekVmukr29Z/v7775w0aQolqTyBpU7PrJQVFoslR7mRCxvO0Ab5DFVGRgYAYLVaAGgdP5Na\nWCyWTO1iY2Px7beLkZSUjHbt2qJKlSq5GufGjbuwWBoCuA5gJYDvAZggCFqQQ+ytNFAqfRAWFvbE\n4tt169bF2LH9MXFiOVitOpjNIoA/AABqnMFCxKMrJsMCEYPFBthe7iEGPaYguNlsRqNGLbB7906I\nohYlSwZh167f4O3tnSv/nkSrVp0QHV0JZvMSXL8ejTZt3sChQ3vh5+eN1as3olgxD0yY8IdTx/wn\noihCrVY/NfuFGicIe4GiELosI5Mj+vYdTEmqS2ALBWEGjcaivPKP1dzVq1fp4eFHlaoPBWEMJakI\nt27dmqtxli79npIUQuAYgXEUxQA2bPgKS5Qob8+re56CMINFigQ+NqL4n8THx/PPP/+kh4cfgc50\nQ39uh5IE+BACWwo6vvxymwxRwP+kcePmBIII3LZHwg5lQECo0wqXp6SkUBSVBNIcuwB6fVd+8803\nTrEvkz+coQ2FTl1kQZWRyRqz2cwJEz5k1ar12aRJO544cSJTm+HDR1GhGJLuzPJnVqlSN1fjWK1W\nTp78CQ0GL2q1ruzVqz9TU1P566+/0tU1kAqFG/39Q7NMZp8dV69eZWlRxVNwJwHGwJtVEECN5sUn\nbl+bzWYKgpLAB+l8u0TAlZJUhJs3b871XP6JrVycC4HTfFS71GB4kWvWrMm3bZn84wxtkDMlycjI\nALClz5s48X0cPLgdmzatRGhoaKY2d+/Gw2Ipnu5JIOLjH+ZqHEEQMHbse3j48DaSkh5gwYLZuHjx\nIjp27I64uPdhsazEvXtGfPPN97n2wT8mBkf1WgTjPo7CDWEw4DBeQUpKI8TGPv6KiP33KYAdAMz2\npxEAfGEyLcTgweNzPZcdO3YgMLA89HpPvPxya9y9exczZkyHJDWCUjkcev3LqFBBj5YtW+batswz\nSv51vWBRCF2WkXEav/32GyUpgMAeAqcoSXU4evSEfNv96KOPqVAMSrc6/Ivu7n65M7JyJanVkgD3\nuXnSTQwn8CuBM5Sk4tluTXft2oui6EegHIG6BCQCOwgcpr9/aI6nYTab+fHHH1OpNBJYT+AmVaqB\nDAtrSJLcs2cPP/nkEy5evPixkdAy/z7O0IZnVl3Onj1LjUbDN9980/EsIiKC5cqVoyRJbNCgQaay\nSCNHjqSnpyc9PT353nvvZWlXFlQZmfzx7bdL6OcXTC+vQA4ZMoppaWn5tjlt2jSq1b3TCeoRenkF\n5qyz1Up+/jkpCLbOvXrx7o0brFOnGUVRRa3WhbNnf5WtmbS0NI4fP5nFi4dSEFwIfEPgLHW6hhw8\nOOvfJ5mnYuWrr3aiWl2WQPt0/pipUKiZlJSUM5+yICIigq1bd2G7dt24d+/ePNuRyZrnWlAbN27M\nOnXqsGvXriRtdQ5dXV25cuVKpqSkcMSIEXzxxRcd7efNm8dy5coxJiaGMTExDA0N5bx58zLZlQVV\nRubZIyYmhm5uPhTFsQQWUZLK8vPPZzje7969m82adWDDhm34/fc/cM6cOfzgg0ncu3Mn2a/f30VT\np0yxCayd1NTUPF1/+e67H+jnF0xPz+Ls129ojleSJ06coCT5E/jZfn3HYp/aOarVelosllzPhbTt\nDOh0xQjMJzCHOp2XLKpO5rkV1GXLlrFjx46cOHGiY4U6f/58hof/nXcyMTGROp3OUVi3Vq1aXLDg\n74vVixYtyiC4j5AFVUYm5/ybCd4vXrzInj3fZZs2b/LHH5c5nkdFRVGn8yLwNYEfaUtyX5F6DOav\ngvpReiFy2bInWP932L9/P43GygRSaasf+zKB4dRq/Tlr1pd5tlu3bksCP6Rb8c5h27ZvZt9RJsc4\nQxueuaCk+Ph4TJgwAV988YU9UMDGiRMnULlyZcfPkiShdOnSOHHiBADg5MmTGd5XqlTJ8U5GRiZ3\n/PTTTzAafaBQqFCmTFX89ddfAIArV65g9+7duHHD+TlgS5QogYUL52DNmu8ylIabPfsbJCWNAfA2\ngM4AvoUPDNiFnWjOVNwTBGDrVqBTp8eZfmrcuHEDnTv3RPXqjTBo0EiULFkSBkMiRHE6gNkQBD1c\nXZdg3bpFGDCgX57HMZstADTpnmjsz2SeJZ65xA7jxo1Dr1694OvrC0EQIAi2RM6JiYkoUqRIhrZG\noxEPH9oiDBMSEuDq6prhXUJCwr83cRmZ54QdO3agU6e3APwMoDHOnfsKdeo0weDB/TBx4hRoNGWR\nlnYW33+/EK+91vZfn18FXMKvOIjiSME5BKGVGItTL7302PYRERGIiopCQEAAunTpYi8GLjy2fU5J\nTExEWFgDXL/+Kszm13HixEIcP/4/7N69GT169Mfp0/NRvnx5LFlyAMWLF8/e4BMYPPgtHD48FCaT\nACAVOt04DBiwNN8+yDiZ/C+UncehQ4dYvnx5x3nFhAkTHFu+gwYNYr9+/TK0r1ChAlevXk2SdHV1\n5Z9//ul49+eff9LFxSXTGM+YyzIyzxzVq79kj3L9+2hSEDzsUa+PUhMeoCS5MyEhId/jWSwWzpgx\nm61adeagQSN49+7dDO8jIyMdW76NMYpxsAUf7UF1euFVVq9e77G2p079nJIURFEcRUmqS6PRj6Ko\npIeHX57vf5rNZs6fP58tW7ahVlsj3XdKoUbjxps3b+bJbnb89NPPfPHFJnzppeZ5rjMr83icoQ3P\n1Ap1586duHTpkuNfcwkJCbBYLDh58iT69u2LJUuWONomJibi/PnzKF++PACgfPnyOHz4MKpXrw4A\nOHLkCCpUqJDlOBMnTnT8uX79+qhfv/7TcUhG5j/m1q1bWLx4MRISTGjbtjWqVq2abR+TKRVALAAT\nAAnANZAJAKoDKGFv9QIEwQ0xMTEoW7ZsvubYp88g/PjjQZhMfaBW/4F16+ri2LF90Ov12LVrF/bu\n3YvBg99GkXXzMODEQSgB/AQtuuMUDJ73sGVLdJZ2U1NTMXbsWKSlnQEQAJPJDKAigHm4d88Tb7zR\nGvv3l37s74msIIn27bvh99+vwWQKAXAOtvurAgALSCtE8emcpHXs2AEdO3Z4KrYLIzt27MCOHTuc\nazT/uu48TCYTb968yZs3b/LGjRscPnw427dvzzt37jiifFetWsWkpCSOGDGCtWrVcvSdN28eQ0JC\nGBMTw2vXrjE0NJTz58/PNMYz5rKMzFMjNjaWRYoUp1rdk4IwmpJUhFu2bMm239Spn1Oh8KOt4HYf\nAkUoCEUIeBI4aV+N7aQkeTIxMTFfc0xOTqZCobEnx7cVAndxqc81a9awTZsOBDwpYDA/Fvwcy+Ub\nPXpwzqxZXLFiRZaFwJOTk3nu3DleuXKFKpWBfxfUJoE29ghcUqfrlevk/2fOnKFO50NbVZskAlUI\ndCewnDpdU7Zu3Tlf30Pmv8MZ2vBMq8vEiRMd12ZI2z2s4OBg6nS6x95D9fDwoIeHh3wPVabQM2bM\nOCqV76YTkzUsX75Wtv0sFgtHjhxLrdaVgJqiWJKAK4EZBNzsiQ+0XLt2bb7nmJiYSKVSaxcn2zxd\nXFpw6NChBNTU4Dh/RCcSYBrAw/849vkn+/fvp7u7L/X6QGo0Rrq7B1AUhxO4RWA1bSXjrhIw02Co\nlet6qIcOHaKLS0i6b3qPKpUPa9Z8mR988JGcqKEA89wL6tNAFlSZwkLfvoMIfJrul/9BBgSUz3H/\nevVaEviKj2qVAm5UqV6gRuPKRYsWZ2hrtVq5Zs0aTp06lb/++muurtu0aNGBWm1bAjsoilPo5RXA\nDh3epAcU3IVwEmA8DGyKWvz2228fayc5OZkeHv4EVtnnfJqAgUApAlpKkjfVandqtf1oMLzE8PBX\nsk1KcePGDTZu3JaensVZtWpdHjhwgIGBoVQoJhA4SYViEgMCgnOVsMFqtXL16tWcOnUqN27cmON+\nMk8XWVDzgCyoMoWFLVu22JMM7CHwF3W6Bhw6dHSO+wcHhxHYnU6QP2KdOo159erVTG27d+9Lvb4y\nlcqh1OuDOWTIqByPYzKZ2L//cFaoEM4WLTry/Pnz/LRPP56xV4u5Ch9WwpcURUOWRcatViuHDx9D\nhUJNQJ8hmAp4lcAKArFUqfTctWsXZ86cyWXLlmUrptu2baOrqz8FYQhtFXC+pru7L48ePcpGjVrT\n27sMGzRolWmn7Encu3ePzZq9RkmqZP9W5XL1/0Tm6SELah6QBVWmMLFkyXf08ytHT8/i7N9/WK62\nJIcOHU2drgmBuwQuUZIq8Ntvl2Rqd/LkSUqSL4GHdhG7S43GnTExMXmb9N69tHh6kgAPQUVfaCkI\nRn75ZdaJEZYsWUpJqkxbkXE3Avvs87hNwJ9ANAErdbpiWf5jICs2bNhAna4IAfcMZ7BG48t5jrDd\nvn27vYi7R7pvdYcajdtTLSgukzOcoQ3PVJSvjIyMc+nW7U106/ZmnvpOmTIRt28PwPLl/lAoVBg2\nbDi6d++aqd29e/egVPoDMNifeECtLor79+/D19c3d4OuWAF07QoxJQVpjRrhaNu2GE2iWbNmKFWq\nVJZdIiL2wGTqDcAPwFIAzSAIpUGeAdAUQEmI4ifw9i6S4/lMmDAdSUmfA3gHwD0AngDMsFqvw2Aw\nPLnzP7h9+za+/noBJk2ahtTUiQCW4+9v5QmVqgju3bv3VIuKy/w7yIIqIyOTJWq1GkuXzseSJfMA\nIEMyBJJYv349zp07h1KlSkGhiIFNzFpDEL6HJKWidOnSOR+MBD79FHjvPdvPfftCNXs2uimz/xVV\nooQvNJp9SEl5B0ArAMMQGroGY8fOw+jRHyE2NgAVKryAVas25PhKS2pqKgAfAP0BNADwOhSKLahW\nLRDh4eE5duv27duoUKEm7t2rDbOZAP4H4FMA3wF4FYKwFAaD5bH/WJApYOR/oVywKIQuy8g4FavV\nym7d+lCvr0i1eiD1+lLs2bMvy5SpSrVaz/Llw3j69GmStqT2FSrUYrFipdm9e98sr9lYUlIYWbma\n4+Bzb5vXMiS4z44HDx6wTJnKNBjq02B4ja6u3jx+/Hi+fJwzZy4lKYTAFgLDqVQaOWjQoBxvmd+/\nf58LFixgy5YtqVT2JGAmUNRu7xCBigRULF26iiMfucx/izO0odCpiyyoMjL549ixY5Qkv3TngDep\nVht569atDO3Onj1Lvd6LwE8ETlKrbc+2bbtkaGO6eZN7je4kwCSo2R5TKEkluXLlqlzNKTExkatW\nreIPP/zAGzdu5NtHq9XKL7+cx/Llw1m1an2uX78+x31v3bpFH59SlKR2VCiqEJho/07bbfdqhdLU\nat05der0fM9TxnnIgpoHZEGVeZ5ISEjgkSNHnlq6u6zYvn07XV3DM0TTGgylHKvSR8yZM4da7dvp\n2j2gUqn9+0rN1au84GoT01twZS3stbebzw4dejx1P7Zu3cratZuyatX6/Oqr+U6rrDN8+CiqO0Yf\nPAAAIABJREFUVP3svuyhrTpOBIHT1Grrs127N3jt2jWnjCXjPJyhDfIZqoxMAWXfvn1o0qQNrFZ3\npKZex+TJEzFixOCnPq6tqtN52JLnt4AgLIHBQAQFBWVop9froVDEpnsSC41GbzuLPXIEbNECQXH3\ncRYSmuF9XEBtAIAgXISHh/GpzZ/289/XX++J5OSZADwwfPhQpKWlYeDAd/Nkc9++fdi48Te4uhpx\n+fJ1pKW9aH8TDmAElMrX4e5uRIcObfHFF1OgVqud5Y7Ms0T+db1gUQhdlnkOsVqt9PT0J7DGvhK6\nQknyZXR09L8y/v79++0JDtQMDq6eaXVKkg8fPmRQUHmq1W8SmEqFwo9qtSs7uRZjqlZLAtwtiPTA\nUvsqbgSBXpQkT168ePGpzDsuLo5hYQ2pULgSmJpu9byTZcvWyJPNVatWUacrRkEYQ43mDXp4+FCS\nyhE4R+AOdbpmHDhwpJM9kXE2ztCGZ64eqoyMTPbEx8cjPv4+gDb2JwFQKOrg5MmT/8r4NWrUwKVL\nJ2A2p+DUqT9Rrly5TG0MBgMOHdqLCRNCUb78z1AofNE9dTS+i7sDVXIyrterh22j3keyNBVAH4ji\nNri5rcXBg3tRokQJp8/5wIEDaNKkJaKjCYulB4DEdG9NUOYgojgrBg16H0lJy0F+hJSUH5CYWB/1\n65eDXl8TanUgXnvND9OmTXKGCzLPOPKWr4xMAcRoNEKSDIiLiwDwMoDbsFj+QNmyQ//rqWXAarXC\n19cH165cx6TU5ngPowAAH6Ieph8+gysbNqB0hRBs2rQD/v7NMWLEELi7uzt9Dp988ikmT56BlJQm\nIA8DOA0gGoAWgCckaTLGj/88T/YTEuLxdxUeIDU1CNWrq/Drr2vzP3mZgoUTVsoFikLossxzxE8/\n/czg4DCWLFmVvXu/Q4OhCF1dw6jVenHs2En/9fQycP36dRYrFkRP6VX+BK09wb2Cb2EhgR5UKkMf\nm/3IWVy8eJFlylQhoCJwho/qltoq6cynKIawbNkX8lVftGvX3tTpWtNWK3YbdbpijIqKcqIXMv8G\nztAGeYUqI1NA2Lx5M/73vyEwmb4B4ILvv38Ho0cPRYMGdeDj44OSJUsCAO7evYsNGzaAJFq2bAkv\nLy+kpqbi9OnTkCQJpUqVypCk4WkxceInwJ2mWGs5gnAkIw5Ae7yKCGwFcAAWS3PcuXP3qY1/4sQJ\nVKtWF6mpdQH8BaCM/Y0aQCA0mg8RHByAPXs2Z5v9yGq1Yvny5Th//jyqVKmCli1bOr7h/PkzIAhD\nsG5dOAwGV8yaNRdhYWFPzS+ZZ5j863rBohC6LPOc0KnTWwS+TBdIs5Xly9fO0Oby5cv08gqgXv8a\n9fp29PIKYFRUFEuUKE+DoRx1Om+2afOGo47opUuXOGDAMHbr1sfplU/+91ITnkUxEuBlBLACxttz\n7X5MIIKS5MO9e/eStNVu3b17d45z7eaEmjUb2RMobCZQlcBHBFIJ7KRG48Hly5fnKFGD1WplmzZv\nUK9/kYIwmnp9CIcNG+O0eco8GzhDGwqdusiCKlNQ6dXrXQrCB+kE9SfWqNEoQ5s33uhFhWKco41C\nMZHFipWxP7MSMFGS6nLevHm8cuUK3dx8KIrvEZhFSQrgiBEj+fXXX/PPP//M8byuXbvGPXv2ZEio\n8GDDBt6BQAI8gIr0wUmKYh2WKlWJGo0LPTz8+d13P5Akly//mTqdB11dX6RW68H58xc65Xt5e5ch\n8DqB/vbt2NoERGq1Hrn6x0N0dDT1+iD+XbP1NtVqF965c8cp85R5NpAFNQ/IgipTUDl16hQNhiIU\nhDEEPqEkFeXvv/+eoU3duq1oK6T9SHTXUaUqRuBEumfT2bv3AE6cOIlKZf90z3fZq6u8TrW6KGfN\n+irbOfXo0YuCYKBCUYEajRt//nkl+eOPTBFEEuB6lKIeOgIaiqI7jx496uhrtVo5YMAwAloCh+1z\n+Is6nSevXLmS7+/Vpk0XqlQ9CFQhUIlAEAMCghkfH58rOxEREXR1rZshkYVeH5BlKTmZgosztEG+\nNiMjU0AIDg5GdPQeDBqUhr59YxER8QsaN26coU3z5vUgSdMB3AVwH2r1Jyha1BUKxSp7ixRI0gZU\nrhyMpKRkmM3pI2rdYKuqshypqVEYOnQ4TCbTY+czc+ZsLF68DOQhWCzHkJKyFUc6vQG88QbUtGIO\nXNEGp5CIRACJEEW949zx4cOHqFu3MebM+QVAIIDKdquloVKVxcWLF/P9vb75ZhaqVLkEleo8FIrT\neP31l3Du3BG4uLg8tg9J7N+/H+vXr8f169cBANWqVYMg/AVb8v9bEMVP4OXlguLFi+d7jjLPGfnX\n9YJFIXRZphBhNpvZu/cACoKKgJJKZQWq1R708Aigi0tFarXedHMrTknyYPHiwdRqPe25dvfYV3Lj\n020XezjONE+ePMlq1erR3d2fDRq0YkxMDIODXyBQkwCpRCoXoCcJ0CoInFkimEAJAgMI7CDQi0WL\nlmRaWhotFgtr1KhPQShLYBFt9UEj7eMeo0JhzHst1X9gtVp5584dJiQkZNv25s2brF69DtVqfxoM\nr1Cv9+LWrVtJkocOHWJISA1Kkgdr1mzIS5cuOWV+Ms8OztCGQqcusqDKFAROnjzJpk3bs0qVehw7\ndhLT0tJy3DciIsJeKSXRIVIajYFRUVH27EbjCdwk8CMVCj0FoQgBTwIutFVDsRCYQ72+KM1mM+/f\nv09PT38KwlcELlKheJ9lylRh9eoNCLjRiEhuRmMSoAlgB6WGp06doru7L5XKIApCMXp4BDry1546\ndYpqtS+BtwmMJbDBPn4ZAhJDQ/OWsSg/XLt2jUajF4FS6b7b7/TyCvjX5yLz3+AMbZCvzcjIPGMc\nP34cL7zwElJTxwGojLNnP8aNG7ewYMHsHPWPjY2FKFYGINmflIfFYoGbmxtu3rwNi2UiAAFAZ1gs\nswEMA+AFWy3RjgDiIIouWL9+DRQKBaKjo2E2B4F8BwBgsUxGTMxivP/+YEx460+ss4ajEqy4BaAV\nBuIgFmKBjw/Onz+OyMhI6HQ6vPTSS1CpVACAyMhIe73RUQDqwZYXuA6AnVCrK6FDh5ZO+Io5IyEh\nAQMHvodVq9YiPr4ygCD8/d0a4O7dGFgsFigUin9tTjIFGCcIe4GiELosU4AwmUz08vIl0CldEMxt\nqtVSjquhnDlzhjqdF4H9BCwUxU9ZunRlxsfHU6XSE7iRLsFBoH27lwQOU6GQOHLkSMbExHD//v18\n770x7NOnLyWptP3KCQncp1pt5L2ICCa6uZEAT0HHICylKH7OkiUrPnGuderUJ1DOHoG7iEB1Ahoq\nlRJbt+6c45qjzqBu3WbUaN4k0ILAGAJ+9ohgEpjBMmWq/mtzkflvcYY2FDp1kQVV5llm48aN1GhK\nEeiYTlCvUqNxyVV5sVWrVtNg8KQoKhkc/IIjInXs2EnU68tQFEdRkmpTFF3t0b1WCsJc+vmVocVi\n4caNGylJRQmMp0LRhyqVO3W6+gQ+pF5fhXOatSb1ehLg7fLlWdzgQVFUsly5ajx37txj55WamkpB\nUBB4hcAoAq0ItCWg/9evody9e5dqtQuBNAI/EAi2b0HrCbjRxcVbLv5diHCGNshbvjIyzxAWiwUq\nlS9SUvYBeA9ARQCT0bdv31xlN3rttbZo27YNUlNTodFoHM8nTx6H2rWrIzo6GoGBfeHu7o4uXdoj\nIeE+AgPLYePGdRBFEcOGfQCTaSGAVrBYAECB+vUvo3z5h+h0rxqqLV4MWK3Am2/Ca+FCXFKrM411\n6dIldO/+Lk6fPo3Q0FAsXfoVJEmCIChBHgVgBlAcwDrodCp4eno64Qs+GYvFAqvVCpVKBZVKBdIC\nwASgM4AYAJOgVivQsWM7LFz4ZQZ/ZGSyxQnCXqAohC7LFCDi4uLo41OKojiQQFuKYhmGhlZ77Or0\n1q1bXLt2Lbdv3+7IfvSI1NRUWiyWbMe0Wq00mUwZngUElCdwKN0qeSr79xtMDhv292XM8ePJx8wr\nKSmJ/v5lqVB8TOAMRXEiixYNYmJiIv39yxDwJxBO4FUCbpwyZWqWdmJjY/nFF19w2rRp/Ouvv7L1\n5Uk+jhw5lkqllqKoYuvWnWkymfi//71DSapNYAE1mjcZElKdycnJeR5HpuDiDG0odOoiC6rMs87V\nq1fZrl1XVqlSjwMGjMgkdo84dOgQXV29aTQ2pcFQgXXqNGVqaipNJhNbt+5MhUJNlUrHUaPG52q7\nmCSHDx9DSapH4CSB7fTQevNWnTo2IVUqycWLn9h/z5491GjKZUiGABRneHgjnjp1ioGBIRQEBZVK\nPSdNyjqp/+XLl+nh4UeN5n9Uqd6lXu+Vp3qvp0+fZu/efajVVrCfHydSq23Ld94ZQovFwtmzv2T7\n9t05Zsz4XCd9kHl+kAU1D8iCKvO8ULFibQLf2sUqjZL0MufNm8c+fQZRq33Nfv3jOiWpIpcu/S5X\nttPS0jhkyCgWK1aKLwSE8E7p0jZVdHUlIyKe2Dc1NZXly1enrWj4o3R9iQS8KUkVuHr1apJkYmLi\nE4W+d+8BVChGpRPk+axXr2WOfbBarWzX7nUCBvt1mPnpbEWydOkXcmxL5vnHGdogZ0qSkSmgXLt2\nGUAD+09KmEx1ceHCZWzZsgvJyaNgu/7hA5PpHWzatDNXtpVKJaZPn4IbOzbggDIZnufOAYGBwB9/\nAI0aPbHv+vXrcfq0CbY6rU0AfAogHEBDWCzhiImJAQD7eWrGc+HY2Fh07dobtWo1xfbtf8BiKZPu\nbRncu/cgxz7Mnfs1Vq36BcBeAK8D+NPxThAOwM/PO8e2ZGRyghyUJCNTgEhKSsLixYsRG3sDQUGl\nkJAwB2lp0wDcgV6/HGFhk/HHH4dx8WI0yBoAALU6GsWL++R4jLi4OPTuPRiWbduw6P51GC1moHp1\nYP16wDt7ERo37kNYLN4AvgewCMAJAKcAzIdC0RZhYT2z7Ldr1y60bt0B8fG1YLW+C5VqAhSKybBY\nwgDoIUnj0K5d8xz78fPPGwCoAFQC4A/bXde60GiKQKv9A19+GZFjWzIyOcIJK+UCRSF0WeY5ITk5\nmZUq1aJO15KCMJY6nR99fMpQo/GgSiVx5MixtFqtPHr0KI3GYtTrO9FgeIUlSoTy3r17ORojLi6O\nISHV2VWsyxQoSYC/QMHxw97LUf+EhAQqFBoCPgTm2c9g3yJgoFqtf2wlma+//oYaTVECPez3UtsT\nSKQoaunm5kujsRgHDhyRKfDqSXTu/Fa6eZDAXgqCjlOnTuX169dzbEemcOAMbSh06iILqkxBZfny\n5TQY6tNWho0EzlCjcWFsbCwfPnyYoW1MTAwXL17MZcuWZXr3OK5evcqiRQL5PhSOSKIvMIgimlOt\n9uMPP/yYof2yZcsZFvYKw8ObOcqhpaSkUKnU0Ja/tyGBMlQo/Llw4cLHiqHZbKZGYyBwJl3CiYq0\nVcqRGBcXl4evRf711180GLwIFKWtoo2Gn38+PU+2ZJ5/nKENgt1QoUEQBBQyl2WeExYsWIDBg/+A\nyfSt/UkKFAoXJCeboFTm7PTm3r17+O6775CYmIgWLVqgcuXKjnedXuuCpmuOoQeOwQoBQ/AFZmEg\ngFoAaqJDh4f4+Wfb2MuWLUevXqNgMs0AkAydbjDWr/8BjRo1wqhR4zF79hqYTP+DVhuF0qWv4MCB\nnY+90/nw4UN4eBSD2ZwIW0pEAGgHleovtGpVGatWfZeXzwUAiImJwfLly2EymdClSxeULFkyz7Zk\nnm+cog35luQCRiF0WaYAYTKZ+Ntvv3HDhg2ZrnD89ddflCQvAusIXKNa/TYbNMh51OvFixfp6RlA\ntbozFYphlKQi3LJli+3l/fvcZ7ClEUyEkq+iBIHp9nuitahQDODAgcP58OFDDh8+hq6uQQTWpIua\nnce2bbuStEXXLlu2jH36DOQnn0zNUaWX4OAXqFBMtq9Od1IUXdi//5B/NQ2hTOHGGdrwzKlLly5d\n6O3tTRcXFwYFBfHDDz90vIuIiGC5cuUoSRIbNGjAy5cvZ+g7cuRIenp60tPTk++9l/WZjyyoMs8q\n9+7dY+nSleniUptGYwP6+JRyVGh5xPbt21myZCUajd5s2fJ13r9/P0e2T58+TUlyI9A9nQj+wuDg\nmuSlS2RoKAnwBjSsjt0EFhLwpyAEUqttx6JFS/DKlSusXLk2NZouBF6grezbI1sz2aFDj2zncezY\nMYaF1WFQUCj79h3kuGN7+fJlVqnyEkVRQU/PAMcWsozMv8VzKajHjx9nUlISSdsvgWLFinHTpk28\nffs2jUYjV65cyZSUFI4YMYIvvviio9+8efNYrlw5xsTEMCYmhqGhoZw3b14m+7Kgyjyr9O8/jGp1\nb8cZqUIxlu3adXtiH6vVyqioKP7yyy+8cuXKY9u98EI9Ao0ITEkngsf5ikcA6e1NArQEB7NH/WZU\nKiUqlTq2atWR06dP59y5c3nnzh3+8ccfNBgq0FbebQMBbwJfE5hFSfJiVFTUE+d6+PBhCoKeQGva\nktG7s27dJhnuomaXgOLGjRu8cOFCjjJAycjkhudSUNNz+vRp+vv7Mzo6mvPnz2d4eLjjXWJiInU6\nnSN5da1atbhgwQLH+0WLFmUQ3EfIgirzrNK0aQcCP6YTvAhWqVIvQ5s//viDM2bM4KpVq2g2m9mt\nWx/q9SVpNLagJHlx06ZNWdp2dfWhLQG8P4EoApfZChWZKAi2wRo2JO2r3bi4uCwDmXbt2kUXlxfS\nBUX9RlH0YNOm7RgZGflYv8xmM3fs2MGyZav9Q9CHUhQNvH37drbfxmKxsGvX3tRo3ChJvgwNrcFb\nt25l209GJqf8a4I6aNAgHjx4MN+D5ZR33nmHkiRRoVBw7ty5JMmBAweyX79+GdpVrFjRkXXF1dWV\n+/fvd7w7cOAAXVxcMtmWBVXmWWXKlE8pSQ0JJBBIoVbbjgMGjHC8nzNnLiXJjxrNu9Trq9PXtyxt\n5dcS7AK1k25u3lmu8mrVakxR/JDAEgJB7A81LXZlW6514dyZc7Kdn8lkYokS5alSDSewjRpNd9as\n2eCJq8qUlBSGh79Cg6EiBcGPQEQ6Qf2BguCWoyozCxcupCS9SCCegJUq1RC2atUp234yMjnlXxPU\nAQMGsGjRoixfvjw/+eQTXr16Nd8DZ4fVauX27dvp6enJffv2sWfPnhw1alSGNuHh4VyyZAlJUqFQ\nZCi1dPbsWQqCkMmuLKgyzyppaWl8/fUeVKkkqlQGNm7c2nHGmJqaSrVaInA+3dWSQAIvpxMoK0VR\nmWVy94sXL7JIkRIU4cvp0DiuxYyFksAOSlJJ/vzzikz94uLieOHCBUdw0I0bN9ixYw9WqlSHPXv2\nz/ZKy+zZs6nTNSVgJjDefpUmjsAtApVYsWKNHH2bt9/uT+CLdL4eo59fcI76ysjkBGdoQ45i7WfN\nmoXp06dj06ZN+P777/Hhhx8iLCwMXbt2Rbt27WAwGPIXapwFgiCgfv366NChA5YtWwaDwYD4+PgM\nbeLi4uDi4gIAmd7HxcU9dl4TJ050/Ll+/fqoX7++0+cvI5NbLl26BIvFimrVwtGsWT2MGzcaomjL\nDvrw4UMACgBB9tZq2Eq77QFwHkApAHNRokRIhuspV69eRWxsLMqVK4fzRyMRVaosGptSkAoF3oI/\nfsBrAOrBZBqDZcvWoUOH9rh58ya2bNmCLVu2YtmyFVCrPWAwKLFt2waEhobip5++RU45f/4ykpLq\n2ec+FkBXAB4QBBE1a9bGzp2bc2QnJKQUdLotSErqD0AJUdyE0qVL5XgeMjL/ZMeOHdixY4dzjeZF\nhY8dO8aKFStSEARKksSePXtmikZ0Fj179uT777/Pr7/+OsMZakJCQoYz1Nq1a2c4Q124cCFr1aqV\nyV4eXZaRcSp37tzhpk2bGBkZSYvFwuvXr9PNzYei+BGBXyhJL3LQoJGO9larlWXKVLFv2yYR2ELA\nw34maSDgRqPRm6dOnXL0+eCDKdRqPejqWo2lDF6MDwkhAZq0WrZx8yXwP8d5qCiO51tv9ePJkyfp\n5uZDna4NgRdpK7p9j8DXDAqqkK1ft27dYt26zalSSSxatARHjhxJvb4SgTsELFSphrBJk3a5DipK\nSUlhnTpNaTCUo9FYm0WLlnhiIXMZmdziDG3IsYUHDx5wwYIFrFevHt3d3dmrVy/u3r2bV65c4aBB\ng1ihQvZ/2bLj1q1bXLZsGRMSEmg2m7lp0yYajUbu37+ft2/fpqurK1etWsWkpCSOGDEig2DOmzeP\nISEhjImJ4bVr1xgaGsr58+dndlgWVJn/mL/LrjWkXl+WTZu+Zt8a7ZZuS/MadTrXDP3+vlqipNHo\nbU/VN5uiOIhubt7cvHkzf/vtN169epVvvdWHguBJIJYhOMELKEYCtAYFkSdPMjIykpLkRVEcSaWy\nP11di/HcuXNs0KAVBWGGYwsZ6EngfQIWCoIi23uhtWo1pko1iMADAjuo03mxW7e3qVJJ1Go9WalS\nrTwHE5nNZkZFRXH79u1ymTUZp/OvCWq7du2o1+vZtGlTLlu2LFN9RovFQr1en+/J3L59m/Xq1aOb\nmxtdXV1Zo0YNrl271vE+IiKCwcHB1Ol0j72H6uHhQQ8PD/keqswzS0hITXtwkO0sVK9/id26daNO\n1zWdoG6jSqXlwoULM0XcPgoCWr9+Pbt168OBA4fxzTd7UZL86er6MlUqVyqVfgTaswG28j5cSYD7\nBJFx6Yp0Hz9+nOPGTeCkSZMdf5fKlKlOIDLdPObTlov3d3p5BTzRL7PZTFFU2s93bf0l6S3OmzeP\ncXFxjI2NzXVdVhmZf4t/TVCnTZvG2NjYJ7bJSTaUZwFZUGX+awyGIgSupxOtsRwyZCjd3X0pipMJ\n9COgpyD0ok7XgiVLVnhi8M/27dup15exB/uQwHYCnuwKN6ZARQJchVr0fUwEcHr69BlIoLl9W/kW\ngXIESlGjceO2bdsytU9ISODBgwcZExNDq9VKvd6DwFH7PCw0GMK5YkXmYCcZmWeNf3XL93lBFlSZ\n/4KkpCQOHDiSoaG16eYWSFEcb99SvUW9PoRr167l+fPn2bBhcwKuBNY6BFet7sRp06Y91vaiRYso\nSW+mE2gLJ0B0RPJ+Bnfq1K7cs2dPtvNMSEigIBgIqAloCLxFjaY6v/zyy0xtt2zZQqOxGF1cKlKj\nceeECR9x8eKl1Om8qVYPol5fn2FhDeX0gTIFAmdog1wPVUbGCcTGxuL27dsoXbo0JEnK9L5z557Y\nvDkRSUlTIAhbIAgzoNXOh9WaiH79BuPVV18FAKSlAYAOQKijb2pqKI4ePZHBXlJSEhYuXIjY2Jvw\n8/NBSspvAC5CBT8sRH10gxUWCBgk1MJSw1lE79uLkJCQbP3Q6/UYPnwQ5sxZj6SkflCpjqBo0Yd4\n8803M7SbPHkqxo+fDGAxgPYAbuLTT2siImI5tm//Bbt370axYi+gU6dOUKlUufmUMjIFFycIe4Gi\nELos85QZNWo81Wo3uriE0tPTn4cPH3a8s1qtnDFjNgElAZNjFanXt+GsWbMy5eKtVKkOgZYEOtqj\na48QKEKt1si/7OefycnJrFjxRep0rxKYSEkqQZXKSDdI3Ga/Y5oABd8NLMMRI0Y/MSVhVlitVi5c\nuIjt23fn0KHvZUq8sGvXLup0xe0+WdP51J0LF2Zd71RG5lnHGdpQ6NRFFlQZZzJx4kQCxQjctAvL\nEgYGhjret2/fiUCQfQv1jkN8DIbG/OmnnzLZmzLlM+p0VQm0IiAR0BN4h0plf06dOpUkuWLFChoM\nddOJ2XkGQc2TsOXkjYEPa2nqcObMmU719bPPPqNGU4yC4EagKm2JJR5tTd+mJJXI0bayjMyziDO0\nQfxv18cyMgWXS5cu4aOPPgXQCkBR+9M3cOXKaVgsFgwePBQrV24GsBTAAADNACyGKPaGp+c1NG/e\nPJPNkSOHYNiwNhCEXQB8AEwH8BVE0eTYOo2Li4PVGohHtUNrIBaRSEUIbuAY9HgRetwOiEfv3r2d\n5uuqVaswfPgEpKR8BXIHABcAVQD0tv+3JAYM6Ibw8HCnjSkjU+BwgrAXKAqhyzJPibVr11KSahIo\nY9+eJYGVLFasJB88eEBR9CAQQmCzfTU5j0A1VqoUlm3+2mnTplOSyhFYRIViND08/ByR9hcuXKBe\n70VgFdtgHhOhIAH+jkAa8Q2Bz6nTeWVIxUnatnKPHz/OnTt35rjs2yMaN25MYEi6wKfLBCQajeGU\nJHc5klemwOMMbZCDkmRk8kjx4sUBxABoByAYgD+A01ixYhNiYmIgCCoAdQH0BDAewF0Iwml8910k\nPD09n2h7xIgh8PX1xqpVv6FIETe8//4f8Pb2BgAEBQVh2Y/f4GC3tzEu7hZEAN9ARF+chRlqAIBC\nEY2oqCiULVsWAEASPXq8g5UrN0ClCoQgXMTWrRtQrVq1HPmq1+sBXE/35CYEQYnVqz9A5cqV4eXl\nlcOvJiPzHJN/XS9YFEKXZZ4iw4aNoST502BoSLXalV9/bQvKefjwIbVaIwFPAn0I1CZgoFJZlm+/\nPSBfY0Zs3sy5Sq3jWsymeg2p1bgQOO5IFmEwVMpQyu2XX36xpwB8aG/zI4OCKmZp32w288CBA4yM\njHTUJj579ixF0YW2zEnTCHixR4+e+fJDRuZZwhnaUOjURRZUGWcTHR3N1atXO6JwH/H7779TqZQI\neNnvlk4mcJlGY7E8j2WJi+NGhZoEmAw1O2EeJSmAEyZMpE5XjDpdTxoM1diiRQdaLBZGR0ezSZN2\n9PcvQ4ViQLot24dUKrWZ7F+4cIEeHsUpCP5UKssxMDDUsdV88uRJ1q1bnxUqVOfHH3+cZx9kZJ5F\nnKENgt1QoUEQBBQyl2X+Q6ZNm4axY/cgLe0H2AJ5olGkSHvcunUx98ZiY2Fu1gzKI0cSidk+AAAg\nAElEQVRwFx5og1+wB3Xg4tIR8+a1Qej/2bvv8KiqrYHDvymZzEwaCQkldAi9FyWhKAhSpaPAVRBR\nQcQu7QoqCAiKggUUpBcFpUsRERQpUlSQEqRICRBaaAkpk0xZ3x8z5iOCXpVAQljv88wDc+reh8Di\nnLP3WpUqsW3bNiIjI7HZbGzevJkxY8bjcIwGzgOfADuBcAyGiVSsOJvY2G2Zh3c4HBQqVIrExLuA\nJYARGED79mdYsmTujV8MpXKx7IgN+g5VqZuoV69ejBv3MRcuDMLlisJu/4CRI4f+8wPt3QutW2M+\nfpwjRhMtPCM4REMgDrd7E1WqDKVatWrUqFGDZ57pz8yZX5KREY3TaQOuAK/ifQdanMDAIgQGeli8\neFWWU+zcuZOUFBPQHm+5NYCO7NrV7waugFJ3Dg2oSt1E4eHh7N69jfffn8D588do3/5jWrZs+c8O\nsnYtdOoESUkQE8Ol114joVsvgtzvk5FxhlGjRlG5cmWSkpKIj49n+vTPSEv7FQjBG0Qr4h0Y9SQR\nEavZsGE5pUuXxmKxZDmNxWLBYHACC4CHAT9gOlWrVrjxC6HUHUAf+SqVTZKTk3n66ZdZt+57ChYs\nyOTJ73DXXXfd2EGnT4c+fcDlgs6dYfZssNlIS0vj6NGjFCxYkDfeGMmECZMRESIiIklLC+PKlR+v\nOkgJ4Bns9mmMHPk0L7743HVP5Xa7iYlpys8/x+HxeAuaBwYaOXZsz/8clazU7S5bYsMNv4W9zdyB\nXVY34PTp07J169a/VcOzVavO4u/f1TfadrYEBkZkKTF46NAh2b59+9+rzOTxiAwdmjmSVwYOFLlO\nUe5XX31dIFhgj2+u69tiMAQLrBBwC8wUf/9Qadask8yaNed/njY1NVVefXW4NGvWXl56aUDmKF+l\n8rrsiA13XHTRgKr+runTZ4nVGirBwbXFZguTL75Y+KfbulwuMZn8/pCv9xGZNm2aeDwe6dmzr9hs\nBSU4uIZERJSQffv2/emxfouNlQ3FSomAuA0GcV+n0ovb7ZZjx45JQEB+gf9kqTRjMJglIqK4GAxG\nKV68ouzateuGr8W+fftk8OAh8t//Dr0mYYRSeYEG1H9BA6r6O+Lj48VmCxP41ReodojNFioXL16U\nw4cPy0MP9ZQGDVrL22+PE7fbLR6PRyyWAF8GIRHwSGBgU5k3b54sWLBAAgJqXDUHdIKUKFHluuc9\nExsrm33TYpKwSnv/CvLss/2zbLN3716JjCwrBkOogFWgknjrl4rAj+LnFyQej+easmnr1q2TZs06\nSZMmHWT58uV/+1r8/PPPEhAQLgbDYDEYBkpgYES2BGmlchMNqP+CBlT1d2zcuFFCQqKvuvMTCQ6u\nLOvWrfMVAn9DYKnY7THyzDMvi4jIyJFvid1eTuAd8ff/j5QtW11SUlJkxIgRYjQOuupY5wSs8tFH\nk7Oe9Lff5FKBAiIgJ4mU6uwUOCX+/oGZhcF/+eUXMZnCBAb7HvEOEG9y/vICHQUCZfz4967pz3ff\nfSc2WwGB6QJzxWaLlKVLl/6ta9Gq1UMCEzLbbzC8Kx07dr+xC6xULqMB9V/QgKr+jlOnTvnuUGN9\ngeQnsdvDZPz48WK3d7sqOJ7OEvAWL14sffs+L6NGvSmJiYkiIrJo0SKxWCoJJGbeoUItyZ+/+P+f\n8IcfRMLDRUB2GfJJEU74to0Xf/+gzONXrhzty750PPMRLzQTiBSwyZw5139P2rbtf8SbS/j3dn8u\n9eu3/FvXokGD1gKLr9p3vjRp0uEGrq5SuU92xAatNqPUdRQuXJjJkz/AZmtAcHAN7PZmzJ07zZfT\nNiuXy43D4WDcuPeZNu0LLBYLrVq15OjRozgcDjp06ED16vnxjratBLwBtMPhSPYeYOFCuO8+OH+e\n9MaNaRNi5YxpOrAEu70jTz3VF4PBwPz5XxAbuxsoDazxnd2Nd55pA8qWrXBNIfDfGQyG6y39W9fi\nkUfaY7cPBX4CtmG3v0b37h3+1r5K3Ul02oxSf+H8+fPExcVRqlQpwsLCOHv2LOXKVScp6Sm8ZcvG\nYDIlULZsKMeP20lN7Y3RuB6RLwgIiCQ42M2GDatJT0+nVq16pKdbgapABjbrXvY93o2SEycCcKFz\nZ8I+/ZRj8fH8978jOH36PM2bN+DHH3exevVKHA43UAqoC6wCygG/AZexWq3s2bOFqKgo3G43Y8a8\ny7Jl31CoUDjvvDOc06dP06rVQ6SmjgYs2O2DmDfvY9q2bfs/r4GI8O677/Pee5MxGAwMGNCP557T\nZA8qb9FpM//CHdhllc169+4rcJfAAwLjBH4S8BNIyhyQBA0FvhSj8R2pU6exHDx4UAoVipLfS6CZ\ncMpEama+oB2ITaz+kdKiRUdxOp2Z52rbtqsYjZ3FW8D8B4EI8ZaECxMwC1ikQYPmWcqx9ev3ktjt\nDQRWicHwtoSEFJL4+HhZt26dNG/e+R8PSlLqTpAdsUHvUJX6h0aOHMUbbxzH6ZzsW7ICbwm3JMDf\nt6wF0Ae4G6u1Mk6nB7e7FDCMQJoyn660ZhUOjPTgMxawG/gVuz2Jt97qwDPP9ENEMJuD8XgOAJG+\n4w4C7EAMFktnLl48fc1jaKs1mPT0g4C33JvN9ijvvhtN3759b95FUeo2lx2xQd+hKvUPPfnkE+TL\ntxqz+SlgFDbbk9SseRdW63+A74BhwK9AY2A66elpuN0PAd2I5F020IDWrOI8fjShFwvoAnQD9pGa\n2po9ew4AcOnSJTweA3DAd2YBduPnNw+b7WFmzpxCQEAAIsKrr76B3R6K1RqMy+UCXFe12Pkn71CV\nUtlJA6pS/0N8fDx16zbBz89G4cJR7Nmzhz17tjN0aBFefDGRNWsWsHnzNzzxRGmqVn2dIkU+x2p1\nEBx8H0FBH2A2lwcsVKUZW/mFmuzmIFDfGMEPvOc7y2KgLHb7EmrXrgJ4c+sajelAV+AloD0Gwzb6\n9+9IbOx2unXrAsDUqdMZP34xaWm/kJ5+CIOhMH5+rYHPMZlew2bbQMeOHTP78+2339KgQSvq1GnC\nlCnT9ImNUtnlhh8a32buwC6rG1SlSl0xmYb6EjOsEbs9XI4cOfKn23s8Htm/f79s27ZN5s6dK3Z7\nXWlGiCTiLwKyEZsMeepZadGio9hsxcRgKCMGQ7D4++eXTp26i8vlyjxW797PidVaWaC9mM11pFKl\nOuJwOLKcr3XrrgKzr5rW8o0UKVJOmjbtKN2795Zjx45lbvvDDz+IzRYhMFdgpdjt5a+dD6vUHSg7\nYoNWm1HqL1y5coX9+3fjdm/BO81kD2lp/jRp0oFJk96mWbNm1+wTFxfHmTNnKF++PNWqVeNA/yG8\nlpqEGWEeVuY0vpcVE9/DYDCwb98+kpKSsFqthIaGUqJEiSyPZydNeo/ataexYcN2ypatTf/+L+Lv\n75/lfIULh2MyxeJ2e78bDPuoXLkKX3+96Jq2TZ06l7S0gXiryUBqqpUPPxxC3769s+uSKXXH0kFJ\nSv0Fl8tFQEAIGRm7gRHA18AkIAU/v2dZs2YRjRo1ytx+7Nj3eP31UVgs5XFl/MqOVo0pt8gb2FbX\nrEPy4AF0fuihbG3jyZMnqVmzHikpDRCx4ue3ki1bvqVy5cq4XC4GD36dRYtWEBwcTPHiBVm5sgYi\nv9dkXUWVKm+yZ8+mbG2TUreb7IgNGlCV+h8++mgyL788HIfDAcwGHvCtmUjbtptZtuwzAA4cOEDN\nmveQlvYz/oQzk7Z05RvEZMIwaRI88cQ1x96+fTubN2+mYMGCPPjgg/j5+f1pOy5cuMCFCxcoWbLk\nNbVMExISWLRoEW63m7Zt21KsWDEAnn22P9On7yA1dSxwFKv1SQwGI2lpg4FQ7PZhzJgxnoceevDG\nL5RSt7HsiA36yFepP1i2bBmbN2+lZMliPPHEEzz9dB/MZgNPPfUaIhlXbZlORoYj89vhw4fx86uO\nPc3KUprSgM0kYeCpkAKMvv9+SvzhPDNnzqZfv8G4XJ3x81vCRx/NYv36lZjN1/61HDHiLUaNehM/\nv3Dsdg/ffruCypUrZ66PiIjgqaeeuma/Tz+dT2rq90AZoDYZGT/Su/cVkpIOkprq4MknP6FVq1Y3\ndsGUUl43/Bb2NnMHdln9A0OGDJeAgAoCb4jN1lyio5uI0+mU8+fPi9Fo8yWinyLwnoBdVqxYkbnv\nCy8MkChscpASIiDHiZCqhIrROEyio5tmOY/H4xG7PZ9465iKgEsCA6Nl8eLFWbY7deqUdOnysJhM\nhQVO+badKqVLV7tu+51OZ2YOYRHxJZPYmjlgyWLpJW+//Xb2XTCl8ojsiA06bUYpH4fDwVtvjSYl\nZT3wKmlps9i16zhffPGF73GvE0gFXgGGYbEU9S2HlStXsmfSPLZgpCxx7MBCXVLYwyI8nt7s3bsr\ny7lcLpcvl28F3xITHk8FLly4kLnN+fPnqVEjhgUL4nG7WwKFfWse5ejRvXg8nizHnDx5KgEB+cif\nvzAVK9bhxIkTjBo1BLv9QWA8ZvNzBAd/Q48ePbL5yimlQB/5KpUpLS0No9EPiABWAo+SllaI7t37\nAILHI0AccBkojMXSxZdEAZKmTGOF4zRW3KzkfrrwLCk8iTe5wyeULBmV5Vx+fn7Urt2QnTsH43IN\nA3YgsoIGDQbhdrvZtGkT8+bNIzGxAR5PD+B5vJmYgoGvKFSoFEbj//9/eNu2bbz00utkZOwEojh0\naCTt2j3Mjh0biIwsxKJFKwkLC+aFF7ZRsGDBm3odlbpjZcOdcrZJT0+XXr16SYkSJSQoKEhq1Kgh\nX331Veb6tWvXSvny5cVut0vjxo0lLi4uy/4DBw6U/PnzS/78+WXQoEHXPUcu67K6yY4cOSKdO/eQ\nmJgWMmLEmCxzPP/I4/FIrVoNxWR6RCDIlztXBI4JFBBoLdBS4DsxGkdL/vxF5XxCgjhHjcrMyTuR\np8SEU2CeL99ubTGbg2XcuHFSrlwdKV68irz++khxu91y9uxZqV+/uZjNVomIKCkrV66U9PR0qV+/\nmQQGVhF//yiBp3y5gZ/zlWirLkFBBWTz5s3i8XgkPj5eTp06Je+9955YrU9fNRc1TYxGc2bZN6XU\nX8uO2JCroktKSooMGzYsM1CuWLFCgoKCJC4uThISEiQ4OFgWLlwo6enpMmDAAImOjs7cd9KkSVK+\nfHmJj4+X+Ph4qVSpkkyaNOmac2hAvXOcO3dOwsKK+IqBfyl2e0Pp3fvZv9znyy+/FKMxwJeEXq76\nNBNYIvCkmM0R0qrVQ3J4/3451769CIgb5CUsAuUEmgsECJQUsEvhwhXEaAwTmCXeuqp3yRtvjL7u\n+T/88EOx2VoIuAT2C+TzJW3YKlZrXWnVqr0kJCRISkqKNGrUWqzW/OLvHyY1atSVgIC7BNJ97V0n\nERElbsJVVSpvynMB9XqqVasmixYtksmTJ0v9+vUzl6ekpIjNZpMDBw6IiEhMTIxMmTIlc/306dOz\nBNzfaUC9c0yfPl0CAh68KiieF7PZKm63W0RETpw4Ia+/PlwGDvyv/Pzzz/LDDz+IyRQqsEAgXGC9\nb79DvgC7VQyGTlKsWJS88syzsr9kKRGQVPykIwvFW4zcJtBN4EuBpgK1BJYJDBMo4qsas0XKlKmV\npa1paWnicrnkuedeFhhzVZvni9kcLmXK1JL+/YdIRkaGiIg8//xAsVofFMgQcIjV+oCUKVNNAgOr\nSFDQg2K3h8vXX399y6+5Urer7IgNufod6tmzZzl48CBVqlRh4sSJVK9ePXOd3W4nKiqK2NhYypUr\nx759+7Ksr1atGrGxsTnRbJVLeDMOXT2vzJOZhWj37t3ExDTG4eiKx5OfDz9shsEAbrcBqAd8BnQG\nwoAT+PkF4XTej0hzPCeq0mXCRMrj4RwG2jKCbXTynaM+3vevS/EOYjqF951sW7wJ85cD4QQE2AG4\nePEibdp0Zdu27zGZzHTq1ImAgLWkpDwJ5MPPbyv339+YlSu/yNK3LVt24nC8CHjnrTocPSlWbDaT\nJj1LQkIC0dFvUapUqWy+okqpv5JrA6rT6eThhx+mZ8+elCtXjpSUFCIiIrJsExwczJUrVwBITk4m\nJCQky7rk5ORb2maVO8yf/zkvv/waycmJuFwZmExDcbtrYbe/y6OP9uH8+fPExNxHampP4F0A0tJM\nGI1LgChgFPA+sBaLpQVz5sxk7tyFrFxZhqqeLqykNUXwsJ9AWjGRo4zGW1btAlbrrzRr1ox8+fLx\n2Wef4rq66AvpwCpstk2MHj0DgO7dn+LHH6Nwu1fhdp9i2bLGNG1ai6++Ko7JZKNcubLMnLnsmj5W\nqFCanTu/xulsAYDFsoZKlcrQtGnTm3RVlVL/S64MqB6Ph+7du2O1WpkwYQIAgYGBJCUlZdkuMTGR\noKCg665PTEwkMDDwuscfNmxY5u8bNWqUJXWcur1t3LiRxx9/kdTUL4Bi+Pv3omTJlRQtupvWrTvz\n8svPM3DgENLSigMlgIPAf4FYPJ6zeO8s++CtOQodOz7EQw89xLhxU2jucfI5DQkime+pSgeCucRd\nQDzBwU1xOvfxzDOP8/bbIwAICgpmxoz2pKb2x2j8BYtlAw8//BCPP76UmJgYADZv3oTTuQ3vX8Xi\npKZ2p0oVN7NnTyU1NZVChQpdt/TaO++MYOPG+zh/vi7gJjISRo5ce3MvrlJ5yPr161m/fn22HjPX\nBVQR4fHHHychIYFVq1ZhMpkAqFy5MrNmzcrcLiUlhcOHD2dmi6lcuTK//PILderUAWDXrl1UqVLl\nuue4OqCqvGXFiq9ITX0KaABAevrHJCW1YP36nZnbnDlzAZF6wBi8d6MDgBeBx4B78E51KQg05csv\nv2bt2rXcvWMz41mLCfiU9vTiNBncDTwLpFOhQhoilTh7NoH4+HiKFCnCBx+MpUSJ91mxYgaRkRGM\nGbODEiWy5ksqWDCSxMTtQDHAg832I0WKtCYkJCTLE5c/ioiIIDZ2O1u3bsVgMBATE3NN0nyl1J/7\n483U8OHDb/ygN/4qN3v16dNHoqOjJTk5OcvyhIQECQkJkUWLFklaWpoMGDBAYmJiMtdPmjRJKlas\nKPHx8XLy5EmpVKmSTJ58bVmqXNhllY3efHO0WCw9rxrUM0qCgiLluedelqNHj4qIyOeffyFWa0mB\nQr6BQ79vmyJg9C1fKyBiYLwsq1Alc7jvcGy+KTWhvmksnQTyidl8t8BCMZn+KwUKlJSLFy/+rfZu\n3rxZAgLCJTCwiwQGxkitWg0lLS3tJl4hpdT1ZEdsyFXR5dixY2IwGMRms0lgYGDm57PPPhMR7zzU\nChUqiM1m+9N5qGFhYRIWFqbzUO9Q58+fl8jIKPH3f1igrW8u6HtiNA6SkJBCcuzYMTl16pT4+4cJ\nPPKHgHpewF+gssDHYiVBvqCiCEgGSE/6+bY74Quo0QL5BcwCFzOPExjYTubMmfO32xwXFyezZ8+W\npUuXZo7iVUrdWtkRG7TajMpzLl68yMyZMxk58gMuXZoO3AeAyfQizz1nYs6czzl/XoD9wN3Avb5f\nJwD5gC2EE8GXxBODG4e/P1OaN+eVb7dgMOQnI+M0YWH5OXMmDhEr3sFG5wHvI9qAgE58/HF7unfv\nfsv7rpT6d7IjNmguX5XnhIWF8dJLL2G32/FOe/Fyu8PYsWMHSUnRQDKQAWwEbHjfoxYD9lCOr9iK\nmRjcxGGgoaEaL60qi9tt5uWXu3HixCFmzvwYqzU/MBqoC7QAVmIyvYHN9iMtW7a8xb1WSuU0vUNV\nedarr45g3LgVpKaOB05js/Wle/eOTJsW4ptv+iXe2qbLKVHCTGpqEpUvFGORZx9hXOInatOGPZzh\nDBAK7MdqrUtq6mUaNGjFDz/0ALoBLqAD+fPv57776jN27PBrBh8ppXI3rYeq1J/wJkxoTnq6gyVL\nniMwMIC33ppLgQIFmDOnGWlpU4FiGAzvEhNTnE2bvuPIyJEUfe01/IEvaUM3niCVR/A+BgaIIj09\nhTNnzpCR4QR+n5ZlBtrSsGE+vvhi5q3vrFIqV9A7VJXnrFy5ii5demAyFSMjI4733x9L796PZ66v\nX/8+tmzZj4gFKE54/qMcf+pRbKNGAfCRycrQgGqkuw7jdjtJT18I1MZo7I7It1gsVvz9LWRkBOBw\nTAQc2Gz9WLJkBs2bN//Ltnk8Ho4dO4bZbKZYsWLXnWOqlLr1siM2aEBVtx0RISMj47rzLlNSUihY\nsDgpKSuAGsAgjMZZvP32a7z00kukp6cTGJgPt/sSYMOMk2nm0vRwncQD9Dda2FgrmlGjhlCnTh1+\n+uknHnvsGc6ejcPttuB951oDmEVw8ABKlqyI2WyiQ4f7qFq1KjExMRQoUOC67U5KSqJp03bs3XsQ\nERf33lufL7+cj8ViuWnXSin19+igJHXHmTx5KnZ7CHZ7EHfd1Zhz585lWX/q1CkMhhCgMt40gvvx\neIYxcOBM+vR5/qo7wgyCSWQVLenhOkkqBjoym/GeFPbsKc2sWQsIDQ2lTJkyVK1aCaOxJNAQbzAF\neJT0dDcrV36K3W7hrbcW0KPHZMqVq84vv/xyTbuTk5N5/vnB7N5dkrS04zgcJ9iwIYPRo8ferEul\nlLrVbnjizW3mDuxynrF06VKxWAr6ypq5xGx+Se65p1WWbZKTk8VuD/PNLy0n4PbND70kRqNNLl26\nJL16PS3lrHVkD5EiIGcwyl28fNV81B1SvHgVadasvZhMYb4EDlECRQUSfdvsFn//QHnxxRfFZmvq\nK7cmAjOlatV6me1JSkqSe+9tJWazzVeKbd1V5/lUWrZ86FZfRqXUdWRHbNA7VHVb2Lt3L126PEJG\nRhegPGDC5RrKtm0bOHPmDC1adKZw4XK0bPkg77//NrAVKMz/P4QJBvxwOBx80udRfjLvpwqn2Icf\n0bTlR07y/5VpNmIwZLBmzQnc7pPASaCL7xg1gGaYTA3weMxMnPgFaWn3ACbfvvdy8mRcZrv79n2Z\nrVvDcbkSgdZ4cwUL4MHffxWVK0fdxKumlLqV9B2qui3cc09rNm4MB+KAdXgD2DcULvw0fn5+xMc/\ngNv9GEbjcgoUmEzVqlX45ptNwOt4EztMICLiG85OHY+hWzdITeU7/OlIcy5TCm8AzgASMRrPULBg\nQU6ffhZvjl+AfUAbDIaGGAxL8HgMQCywHRiM991qOGbzABo3PsqaNYsBKFGiKsePz8EbiM8Bd+Hn\nF4C/v4GoqFA2blz9p0UclFK3jr5DVXeMU6fOAL0BC956pV0xmTrj72/m+PHzuN1vARXxeAaSmhpO\n3749KVgwGJPpLQyGRgQELGdfvx4YOnSA1FQ+NfnRnPpcpi7e+qUmvJVnmuPxDOLMmQvAdLxZkAAW\nABcJCvoWb+L9hkARoD3euahFsVjyU6nSFubOnZTZ7mLFimIwbPZ9i8BiiaZHjwZ88800fvxxvQZT\npfIQvUNVt4Wnn36JGTOO4nDMAhZhsbxG8eKBHDlSEo9nO94710DASUBABTZsWEC5cuXYvHkzJqDR\nypWYP/wQgIxXXqHsnCUcP9EPb7WZGsAmvNVm3vedcZnvuw0IxWCIZ+DAfowd+xYez09AM7x3taWA\nbwgKepgDB3ZdU25t37591K/fFLe7DiIXKF7cybZt32ogVSqX0Wkz/4IG1NtTWloa//nPEyxfvhCj\n0USpUuU5dOgiIm/hfQS8F+iMwbCUJk3C+PrrJRiNRkhNhUcegSVLcBqM9DGamSUQFlaAlJRUjMZ6\nZGTswulMAEYCL/vO+CPBwZ0ICQkgPT2DPn16MmzYEMLCIklMXAn8iLeOaj6s1iRWr17Mvffee922\nJyQksH79emw2G/fff7+WWVMqF9KA+i9oQL29uVwujh8/TuXK0TgcT+B9TPsZMBuD4UPq1s3Hhg1r\n8fPzg7NnoW1b2L6dyxjowGOsJw3YBTxH/vwjGTv2DZ57bgDJyZeAQsAsIAJ4nDZtSvLllwuznH/+\n/M/p1et5PJ7OmEw7KF3axfffryYsLAyl1O1LUw+qO47ZbCY9PR2TKRAYCnTAO9/URZkyBVm9+ktv\nMP31V6RVKwzHjnEUA61YzH7a+47SHjCSlgYlS5bEO3q3tm/dS8AVTKYLjB//+TXn79q1C+XKlWXj\nxo1ERNSjc+fOmphBKQXooCR1G/nkkymUL383nTs/jsWSjsHQCggHIggOzuDHH79jyZJlvFy7IcnV\na2A4dozt2IjGzn4aXnWkokAcbnciZcuWxem8ALyNt/zaOQyGy8yYMZEyZcpc0waHw8GVK1eoWbMm\nnTp10mCqlMqkd6jqtvDSSy8zfvwsYAbekmkuoALwHRBERkZ56tdvSsxv8XyUkYAFN0sw8TCDSOMw\n0Ad4E/gVmIPVauGdd8ZStGhRxowZxZAhrTAaGyMSwBNP9LxuLdMLFy4QHd2Es2fNgFC4sLB16zpC\nQ0Nv1WVQSuVi+g5V5XoXLlygQIGyeDzvA35AT+AIEAmkAWWB+bzK/byBA4DxvEB/BA+bge/xPspd\nisXi4YUXHqNbt27UqFEj8xw7duxgz549lClThgYNGly3Hb169WPuXANOp3e0sMXyND17Wpg8+f3r\nbq+Uun3oO1R1R4iLi8NotOLxJONNpBCCN5gC2PAjkk8YTk8cuDHyPO8zkWeA/sABoB1gxWBIZtu2\nTVkC6e9q1apFrVq1/rId+/cfwensB3inxWRkNGffvqnZ1U2l1G1O36GqHLdv3z4qVKiD2exPqVJV\n2bFjR5b1pUqVwmRKxjsI6QzgAcYByYQwidX8TE/WkgJ0sRRlIkWAD4GZQBVgO4GB29i69dvrBtO/\nKyamJlbrTMAJZGCzzaZevZr/+nhKqbxFH/mqHOVwOChRoiIJCf9F5BFgKfny9efYsV8JCQnJ3G7p\n0mV07doDl8uK250MBFKCZFaRRiWEswYTqV/MZ82FywwePJrLl/MBU4FqWK2dedxbBHkAAB/xSURB\nVPXVurzyyuAbamtaWhqtWz/Eli1bAaFBg/osX/45Vqv1ho6rlMp5Og/1X9CAmrvs3buXevU6c+XK\n/sxlISHRrFz5LvXr18+ybWJiInFxcYSGhjKy3YMM37mLQjjYS0U6Wqz0fP0hXnllMMePHycmpgnJ\nyfnxeBKpUqUI3323IjPweTweb9KHf0FEOHXqFACRkZFaIFypPEID6r+gATV3OX36NKVKVSI9/Tcg\nP3AFm608O3Z8S1JSEp9+Oo+wsFCeeOJxihQp4t1p2TLSOnTEJh7W0oROLCKJRZhM/Vm5ch7Nmzcn\nJSWFn376CX9/f+666y5MJhMbNmzgwQcfJSHhOFFR1Vm+fB7ly5fPye4rpXIJDaj/ggbU3GfAgKF8\n/PEXOJ0t8fP7lm7dGlOyZEGGDh0DvAAk4Oc3j/bt2/FOsQiKjx8PIkynFk+xBSd+wH8AfwIDV3D5\n8llMJlOWc5w9e5aoqKokJ88C7sdgmELhwuOIi/sVs1nH5il1p9OA+i9oQM2dvvrqK2bNmkVGhotG\nje7hhReGITIV6AiAkWcZxxKeJx6AoQYTo6Qw3rmoV/CO5v0Cf/+HOHHiIBEREQCcOXOGQYOGsWPH\nTg4d8ic9fUPmOQMCirF370ZftiSl1J1Mp82oPEFE+OSTuaxZc5LU1JZ89dVHiJjxZjQCOyl8xne0\nI550zPT1D2JGuhvYjbdKjAVvpqNN2O128ufPD0BSUhK1azfk3Ln2uFwPAu8AKUAAcAKn87Lm4FVK\nZRsNqOqW8Xg8rFu3joSEBKKjo0lNTWXlypUkJiby9dfrSUs7DFhxOFoAjYAXKcgoVtCHOhzkIiba\nM4SN6QuBZKAXcBLvHWocBsNWVq1alzngaO3atVy5UhqXaywgeLMkVcFqvR+jcTWvv/4GwcHBOXAl\nlFJ5kQZUdUt4PB4eeOAhNm48gMFQkYyMvoAJt7snRmM8Tmcy3rtHK95E9SYqsYeVNKEkHg5joRV3\nc5CPgffwBtGhwFy8d6jdeeONp4iOjs48p/fxze+jcA3A+xiNYYweXYm6dR8jJibmlvVfKZX36TtU\ndUssXryYHj1Gk5KyGW/JtcbAFMisANMDSAA+wmSaR6d8E5h84Qz5ELYaQmhvcBESVYGjR/fjdP6A\nN59vC+BR3/5fUrHiaIoUKUxGhovnnutJ06ZNqFChFgkJHXC7g7FYltOmTQUWLpxzi3uvlMrtsiM2\naKYkdUvEx8fjdt+FN6l9K7x3omWv2qIyBQseJDT0Ht4oPY95l8+RD+HbsPysevlJdp8+woEDPzFt\n2mRstmYYjbHAhav2v8iBAwdZu7YdGzY8TI8eL/D112tYuXIB/v6zMZmWAU52797DpUuXbl3HlVJ3\nDL1DVbfE9u3badSoPWlpH+N9VNsE+A34BDgNNGfpkqm027EDRowAYCyNGERrbPa3eP/90Wzc+BMi\nQpMm9YiNjeWDD6aSnv4C4I/J9CZudy9gvO+MS7n77o8oVaooixYVwOUaAwgWy1M8+WQgEya8e8uv\ngVIq99JpM/+CBtScM3nyVJ555jlcLiNwCBgOLAIcBPpZOdE8mnwrVuAGnuFhJjHXt2cbvGXahgFG\n7PYxfPXVQkJDQ/n442m4XG6OHDnGunXNgGd9+yyibt1PcDqd7NjxX+B+3/LPuf/+BaxZs/BWdVsp\ndRvQaTPqtlKvXl2s1kBSUgogchfQAYgglKoscy8m34oVEBDAkNJVmLTnAd9ei4EtwFigLwCpqSE8\n9tjznDt3Eo/HzRNPPMGwYYPYsqUjqakWwIbN9l8GD57It9/+wL59U3A47gXc2O0zaNjw3hzovVIq\nr8tV71AnTJhAnTp1sFqtPPbYY1nWrVu3jgoVKhAQEMB9993H8ePHs6wfNGgQ4eHhhIeHM3jwjSVB\nVzfut99+Y/Xq1Rw5ciRz2csvv05ycn9EegPFgU+IMl5hi2EhDT0uKFwYNm6kzmsDsNkGAJ3xZkoq\nDwRddfQgjh07R3LyNlJTdzF16iY2bNjC118v5oEH1tOs2XIWLPiE9u3bM2bMMOrXd+DvXxCLpSDN\nm+dn8OD+t/JSKKXuFJKLLF68WJYuXSp9+/aVnj17Zi5PSEiQkJAQWbhwoaSnp8uAAQMkOjo6c/2k\nSZOkfPnyEh8fL/Hx8VKpUiWZNGnSdc+Ry7qcJ7333gSx2SIkJKSp2GzhMmnSFNmzZ49ERdUWKCbw\nkEB/uZsgOW8yi4BI1aoix4+LiIjL5ZJSpaqIwdBFoJFAH99+ywVWChQQeF5AfJ9V4udXUIoVqywf\nfDDxmvZ4PB45e/asnD9//lZfCqXUbSI7YkOujC5Dhw7NElAnT54s9evXz/yekpIiNptNDhw4ICIi\nMTExMmXKlMz106dPzxJwr6YB9eY6fvy4WK1hAsd8we6AGAx2sduLCfj7gqlIRxZKKhZvRGzWTCQx\nUURE3G63vPPOO+LnV0zALfCDQLjAwwIVxWAIkypVaonR+NpVAXWsQHOBLWK3l5VZs+bk8FVQSt1u\nsiM25KpHvr+TP7wYjo2NpXr16pnf7XY7UVFRxMbGAt4C1Vevr1atWuY6dWsdP34cf/+yQAnfknKI\nFCY1tQJQHajIi4xjAQ9iI4NZfnacS5ZAcDBXrlwhKqoG/fuPwul0+vaPAVZiNn9N27bV2Lr1K5Yv\nX0RIyFSs1kcxGHrgnZM6HogmNXUEs2cvvuX9VkqpXBlQ/1hjMiUl5ZoUccG+f4ABkpOTsxSjDg4O\nJjk5+eY3VF2jXLlyOJ2/Adt9SzYAF4EDmOjHBMYyjpcxIgwmil6uQGre3Zi4uDheeGEQR49WBM4A\nZfAme1iB2fwh1apVZPHiT7n77rspWbIk+/b9zNixd1Ghwh7gRaAiAAbDKUJCAm95v5VSKleO8v3j\nHWpgYCBJSUlZliUmJhIUFHTd9YmJiQQG/vk/qsOGDcv8faNGjWjUqNGNN1oBEBERwbx5M+jWrQUG\nQxDp6RcRaYrVfZ75vMMDpOLAwKNY+IJIkO3ExtahdOlKiHiABnjTBK7CG1CfoF69aqxYsTJLSbZC\nhQrxzDPPEBMTw733tiA1NRXwYLdP4/XX1+VI35VSt4/169ezfv36bD1mrgyof7xDrVy5MrNmzcr8\nnpKSwuHDh6lcuXLm+l9++YU6deoAsGvXLqpUqfKnx786oKrsJSLs3buf/PkjSU29TIMGDTj3yw6m\nnztDDcngAmbaEcFmigNTgXpAeTyefYAbbxald4BBQCAmk4OJE8dl/ufpj2rXrs327d8ze/anGAxm\nevbcpEXDlVL/0x9vpoYPH37jB73ht7DZyOVySVpamgwePFi6d+8uDodDXC5X5ijfRYsWSVpamgwY\nMEBiYmIy95s0aZJUrFhR4uPj5eTJk1KpUiWZPHnydc+Ry7qcZ7jdblmyZIm0aNFK/PzKCMQINJIq\nPCnHMYmAHDIYJQqbwD0CDwpECVQSWH3VAKPPBSIECovJlE/mzZuX011TSt0BsiM25Kro8vrrr4vB\nYMjyGT58uIiIrF27VipUqCA2m00aN24scXFxWfYdOHCghIWFSVhYmAwaNOhPz6EBNfu43W45fvy4\nXLhwQerWbSxmc2WBHgL5BCrL/aySRIJEQDZhkPx8LjBIoLqAXaCsQC+Bwb5g6hGzuY907vyw7Nix\nQ5xOZ053USl1h8iO2KCpB9W/cuLECRo3foBTp87hcFxGpDCwH28ptQ/pxUQm8xtm3HzOgzzKctI5\nhXeAUk28j3sHAevxpgUsDFzCYDjGN98soUmTJjnTMaXUHUmrzagc4fF4uO++thw50oG0tGcRMQF1\nAAsGPIzkENM4gBk3Y3iaR4xFyDD4+/Y+AVTFmwWpOtAbeBxv6baCiHzAs8++khPdUkqpG6IBVf0j\nIkLnzj347beDiFQA3geigO/w5xs+pRtD+BAX8CRWRgYs4J5GR3jyyR4EBNTGbn8P2AXEAQvw3q2O\nAPoAq4EmnDp1Imc6p5RSNyBXjvJVudfOnTtZs2Yz3iA6C291lwmE0YOlPEBDMrgCdMaPe0YOJXnI\nkMx9e/Tows8//8z8+ZfYvr0GbncJvHesdqAL3uky71Knzl23vF9KKXWj9A5V/SOXL1/GYAjFGwi/\nB05Rhg/Zwoc0JIOTGGhADdYwlgkTZmTZt0yZMowY8Q4//lgFt/spDIZjQFHABpQGwjAYPqN27T+f\n8qSUUrmV3qGqf6RWrVqkpR0HxgD1iOFeljGdCJzsxJ8HmMQpkoARXLyYCnjfue7du5dJkyaTmNgC\nl2siACItgE7Ax0A7IAWR7Xz33bgc6ZtSSt0IDajqH3G73b7EG7XpzK/MIQUr6azCRBc2kszvj2sP\nERn5DYcPH6Z+/WacP5+MiAOP57mrjlYY70OSnXgf+VoxGvdSuHDELe6VUkrdOA2o6h/p3ftFXK5A\nBvAIb7MfgI8J5VlcuLFetaWTjh1bU61aNKmp3fCO6B0JfADUxVsP9XngQWAGRuNB/P0DsFjWMnbs\n97e2U0oplQ10Hqr6WzweDytWrODhLo8x1pHOU6QAMAAj72AG7gIuA68Ch7FY3sZo9OBw3AOs8B3l\nKN4k9jYgGPgPMAKD4VXuvnszjz32MG3atCEyMvJWd08pdYfLjtigAVX9TyJCp07d2bJmN9NT9tMS\nJ2lY6c5MFiFAX8ADXAFCKV26KKdOuXA4OuENonN8R0oASgHjgBcxGvthMKQTEDCfH3/cSLly5XKi\ne0oppYkd1M0nIvTr9xzbl6zjq5QMWuIkgUDu41sW0QU4hzeYjgYygNkcOxaPwzEGb8KGNcBEYCPe\nO9IeQA+MxgyGDLHx2mvh7Nq1TYOpUuq2p3eo6i+NGfMOXwx5ky89lygKHMBIKywcYSAgeN+Jgvdx\nr5fZXBK3+wVEXgB2Aw/hvTvtCozDZBpJzZqb+PHH725tZ5RS6k/oHaq66WLfeZ/vfcF0Aw2pxwKO\nYADexjt1xgw4gXjfHsmYzenYbCMxm5/Fz+9jgoMv8+abA7Fa52AyBVO58rcsW/ZpDvVIKaVuDr1D\nVX/uk09w9emDGfiMbjzGDDLwByJ4/PF2zJxpx+3+AHgXbwrCxgQE/Eznzg157bUBLFiwAIPBQNeu\nXSlevDgigsPhwGaz5Wi3lFLqj3RQ0r+gAfVv8HjwDBqE8Z13ABiJhdf4DaEYsAOoT0BAEE5nTTIy\nvvbtNI2AgIEsWDCXFi1aXFMkXimlcrPsiA06D1VllZYGjz6KccECnEAfAphBC6Aa8Hvu3U9JSXFg\nNPbBz+8xnM5y2O0f8cEH79KyZcscbb5SSuUUDajq/50/T0rTpgTs2kUiBjrTkLW0wlu39EPgM2AP\n4J0n6vE8SZ8+dqzWS7RqNYOmTZvmXNuVUiqHaUBVuFwuxj/9PJ2mT6G028lx7LTGyl6O8P8/InXx\nDkSy+77vBGDt2i0cOPCzPuJVSt3xdJSvYnL3XvSa8gml3U5+JpRoQtnLNKARMByYAZwEUoEKQBug\nGfA0hw+fYfny5TnVdKWUyjU0oN7p5s3jiflzyI+L5RTjXopwmg+B9r5PeWAZMBVvEM2Ht0LMSmAT\nHk9l9uzZk1OtV0qpXEMD6p1KBEaPhv/8B39gAu1oz1RSOAU4fBvdgzd1YGNgFN4C4PHA08B9QDR2\n+0XNcqSUUmhAvTM5nfDkk/DKK3iA14NCeZb1eFgOpODNzTsZmAe48Sa8b0yRIj/zySfjCA0NJTi4\nMnb7Qtq1q0Hnzp1zri9KKZVL6DzUO01iIjz4IHzzDanAw5RlKYWAl4D38ObjPY737tSC9+70BWAh\nBQv25syZwyQlJbF7925CQ0OpVKmSDkhSSt32NLHDv3BHB9QTJ6B1a9izh3NAGzqyndN4H+c2xjty\ntznQEtiMt9TaemAW0AR//6I4HFdypu1KKXUTaS5f9fft2AF168KePRwwmonGxHaq+1b+4vu1JlAZ\n+AJIxlspZiTQBj+/14mOvvfWt1sppW4Teod6J1i5Erp0gZQUvsNIR4K5jAVv2bV2eN+VtsOb5H47\n4AKCKF/enxMnTuBwXKFu3cYsW/YpEREROdcPpZS6SfSR779wxwXUjz6CZ58Fj4c5mHiComRQBu98\n0rmADUjE+7CiAJAOmLFaM/jyyzk0bdoUt9uN2aw5QJRSeZc+8lV/zuOB/v2hXz/weBiOhR50JIN0\nvGkEvwIKAWlANNAN8AcuExxsZMGCqdx///0YDAYNpkop9Tfov5R5UWoqdO8OixfjBJ6gO7P5FGiL\nd7BRf+BF4BO8j3c7ADWA/eTLl8pvv/1M/vz5c6r1Sil1W9JHvnnNuXPQti1s20aG3U6L1HC+4zug\nEjAE2AfUAr4BtgAODIZACheOpE2bJrz//lj8/f1zsANKKXXrafk2ldX+/dCqFRw9SnJ4OE0cHrbj\nAFYDtYHxeO9I2wITgKHExFxi48bVmEymHGy4Ukrd/vQdal7x/fdQrx4cPcrposWociWY7cmpeKvD\njAZMeN+bPgYMA2pz330OVq9epMFUKaWygQbUvCAlxZv96NIlaNeOSmevEJe+DO8DCDMQgTdpw0ig\nEP7+6YwdO4J1674kODg4J1uulFJ5Rp4KqBcvXqRDhw4EBgZSsmRJ5s2bl9NNujUCAuCzz/A8/zzV\nfzvJZWcG3ukwJrypBIsC5YA1BAW9x6JFE+nf/4WcbLFSSuU5eeodar9+/bBarZw7d46dO3fSunVr\nqlevTqVKlXK6aTedNGlClef68+uve4EiwIN4sx45gVAgEkhiwICetG7dOgdbqpRSeVOeGeWbkpJC\nWFgYsbGxREVFAfDoo48SGRnJ6NGjM7fLi6N8nU4nRYtGce7cWcAKFANO461dmghcBuDhh7szZ840\nTWavlFJ/oKN8r3Lw4EHMZnNmMAWoXr0669evz7lG3SIPPNCBc+cS8VaHCcUbTIvirV96gSJFinLk\nyAEsFktONlMppfK0PPMONTk5+ZoBNkFBQVy5kvero3z33XdAAN7/HxnxDkLaBxygZcsGHD16UIOp\nUkrdZHnmDjUwMJCkpKQsyxITEwkKCrpm22HDhmX+vlGjRjRq1Ogmt+7m8vOz4HSC9w41BW8KQRs9\nez7IjBlTc7RtSimVG61fvz7bn2Dm6Xeo3bt3p1ixYrz55puZ2+XFd6gjRoxh+PD3cLt/f+zrZPDg\n57O8O1ZKKfXntNrMH3Tr1g2DwcDUqVPZsWMHDzzwAFu2bKFixYqZ2+TFgCoizJo1h88+W0JISABj\nxgynTJkyOd0spZS6bWhA/YNLly7Rq1cvvvnmG8LDwxkzZgxdu3bNsk1eDKhKKaVujAbUf0EDqlJK\nqT/SeqhKKaVULqEBVSmllMoGGlCVUkqpbKABVSmllMoGGlCVUkqpbKABVSmllMoGGlCVUkqpbKAB\nVSmllMoGGlCVUkqpbKABVSmllMoGGlCVUkqpbKABVSmllMoGGlCVUkqpbKABVSmllMoGGlCVUkqp\nbKABVSmllMoGGlCVUkqpbKABVSmllMoGGlCVUkqpbKABVSmllMoGGlCVUkqpbKABVSmllMoGGlCV\nUkqpbKABVSmllMoGGlCVUkqpbKABVSmllMoGGlCVUkqpbKABVSmllMoGGlCVUkqpbKABVSmllMoG\nGlCVUkqpbJBrAuqECROoU6cOVquVxx577Jr169ato0KFCgQEBHDfffdx/PjxLOsHDRpEeHg44eHh\nDB48+FY1WymllAJyUUAtUqQIr776Kr169bpm3fnz5+nUqROjRo3i0qVL1KlThy5dumSunzx5MsuW\nLWP37t3s3r2b5cuXM3ny5FvZfKWUUne4XBNQO3ToQLt27cifP/816xYvXkyVKlXo1KkTFouFYcOG\nsWvXLg4ePAjArFmz6N+/P5GRkURGRtK/f39mzpx5i3uQO6xfvz6nm3BT5eX+5eW+gfbvdpfX+5cd\nck1A/Z2IXLMsNjaW6tWrZ3632+1ERUURGxsLwL59+7Ksr1atWua6O01e/6HPy/3Ly30D7d/tLq/3\nLzvkuoBqMBiuWZaSkkJwcHCWZcHBwVy5cgWA5ORkQkJCsqxLTk6+uQ1VSimlrnJLAmqjRo0wGo3X\n/dxzzz1Ztr3eHWpgYCBJSUlZliUmJhIUFHTd9YmJiQQGBt6EniillFJ/QnKZoUOHSs+ePbMs++ST\nT6R+/fqZ35OTk8Vms8mBAwdERKRevXoyZcqUzPVTp06VmJiY6x6/TJkyAuhHP/rRj370k/kpU6bM\nDccvM7mE2+3G6XTicrlwu92kp6djNpsxmUx06NCBAQMGsHjxYlq1asXw4cOpUaMG5cqVA6BHjx6M\nGzeOVq1aISKMGzeO559//rrn+e23325lt5RSSt0hcs071BEjRmC323nrrbeYO3cuNpuNUaNGARAe\nHs6iRYsYMmQIYWFh/PTTT8yfPz9z3z59+tCmTRuqVq1KtWrVaNOmDb17986priillLoDGUSu89JS\nKaWUUv9IrrlDVUoppW5neTag3mmpDC9evEiHDh0IDAykZMmSzJs3L6eb9Lf91Z9VXvhzysjI4PHH\nH6dkyZIEBwdTs2ZNVq9enbk+L/TxkUceoXDhwgQHB1O6dOnM1zWQN/oHcOjQIaxWK927d89clhf6\n1qhRI2w2G0FBQQQFBVGxYsXMdXmhfwDz58+nYsWKBAYGEhUVxaZNm4Cb0L8bHtaUSy1evFiWLl0q\nffv2vWbUcEJCgoSEhMjChQslPT1dBgwYINHR0ZnrJ02aJOXLl5f4+HiJj4+XSpUqyaRJk251F/6R\nrl27SteuXSUlJUU2bdokISEhEhsbm9PN+lv+7M8qr/w5paSkyLBhwyQuLk5ERFasWCFBQUESFxcn\nCQkJEhwcfNv3ce/evZKWliYiIvv375eCBQvK6tWr80z/RETuv/9+adiwoXTv3l1E8s7PZ6NGjWTa\ntGnXLM8r/VuzZo2UKFFCtm3bJiIip06dkvj4+Jvys5lnA+rvrjcNZ/LkyVmm4aSkpGSZhhMTE5Nl\nGs706dOzXOjcJjk5WSwWixw6dChzWY8ePWTw4MH/1969hES1x3EA/w4mNEzmqEgPrBaVVCMJxUSS\nkNE6alFRpJEoCcJQRE8qNWjjotpED4hW0aHFWbgRiQmyWhnmSBRh+FiUNDMhEkfHmMnvXVycm497\nSzre8Zz5fuAsPGcW/y+/A1/OjPOfDK5q/mbOym1z+tnWrVtpmqYrM3748IElJSXs7u52TT7DMHj4\n8GG2tLSwurqapHvuz6qqKj548GDWebfkq6io4MOHD2edX4h8rn3LdwqzYCvDvr4+LFmyBBs2bEif\nKy8vX9RrnsvMWbltTlOi0Sj6+vpQVlbmqoyNjY3w+XwIBAK4fPkytm3b5op83759Q3NzM27dujXt\nHnVDtimXLl1CcXExKisr0dnZCcAd+X78+IHu7m7EYjFs3LgRa9asQSgUwsTExILkc32hZsNWhpZl\nzcqTl5eXzuMUM2fltjkBQDKZxLFjx3DixAmUlpa6KuOdO3dgWRbC4TCuXLmCrq4uV+S7evUq6uvr\nsXr1ang8nvR96oZsANDa2orBwUEMDw/j5MmT2LdvHwYGBlyRLxqNIplMwjRNvHr1CpFIBD09Pbh+\n/fqC5HNkoWorw+l+lccpZs7KbXOanJxETU0Nli5ditu3bwNwX0aPx4OqqiocOnQIhmE4Pl8kEsGz\nZ89w+vRpAH/fo1P3qdOzTdmxYwd8Ph9yc3Nx/Phx7Nq1C+3t7a7I5/V6AQChUAgrVqxAUVERzpw5\ns2D5HFmoz58/x+Tk5JzHixcvpr12rifUQCCA3t7e9N9jY2Po7+9HIBBIX49EIunrvb29KCsrW6A0\nf660tBSpVGraLlCLfc1zmTkrN82JJOrq6hCPx2GaJnJycgC4K+PPkslk+u1fJ+fr7OzE0NAQ1q5d\ni1WrVuHGjRswTRPbt293fLZfcUO+goIClJSUzHltQfL94ee9i1YqlWIikeDFixdZU1PDiYkJplIp\nkv/895ppmkwkEjx37ty0vX/v3bvHzZs38/Pnz/z06RO3bNnC+/fvZyrKbzly5AiPHj3KsbExvnz5\nkvn5+Xz//n2ml/Vb/m1WbppTQ0MDd+7cScuypp13Q8ZYLEbDMGhZFlOpFDs6Orh8+XJ2dXU5Pt/4\n+Dij0Sij0Si/fPnCs2fP8uDBg/z69avjs5Hk6OgoOzo6mEgkmEwm+ejRI/p8Pn78+NEV+UiyqamJ\nwWCQsViMIyMjrKysZFNT04Lkc22hNjc30+PxTDuuXbuWvh4Oh7lp0yZ6vV7u2bMn/ZWGKefPn2dh\nYSELCwt54cKF/3v58zYyMsIDBw7Q5/Nx3bp1NAwj00v6bf81KzfMaWhoiB6Ph16vl8uWLUsfjx8/\nJun8jPF4nLt376bf72d+fj6DwSDb2trS152e72ctLS3pr82Qzs8Wj8cZDAaZl5dHv9/PiooKhsPh\n9HWn5yPJZDLJxsZG+v1+rly5kqdOneL3799J2p9PWw+KiIjYwJGfoYqIiCw2KlQREREbqFBFRERs\noEIVERGxgQpVRETEBipUERERG6hQRUREbKBCFRERsYEKVURExAYqVBERERuoUEWySH9/P4qKitDT\n0wMAGB4eRnFx8axfaRKR+VOhimSR9evXo7W1FdXV1UgkEqitrUVtbe2s3xEWkfnT5vgiWWj//v0Y\nGBhATk4OXr9+jdzc3EwvScTx9IQqkoXq6+vx7t07hEIhlamITfSEKpJlLMtCeXk59u7di/b2drx9\n+xYFBQWZXpaI46lQRbJMXV0dxsfHYRgGGhoaMDo6iidPnmR6WSKOp7d8RbJIW1sbnj59irt37wIA\nbt68iTdv3sAwjAyvTMT59IQqIiJiAz2hioiI2ECFKiIiYgMVqoiIiA1UqCIiIjZQoYqIiNhAhSoi\nImIDFaqIiIgNVKgiIiI2UKGKiIjY4C99KgLU7MAmZwAAAABJRU5ErkJggg==\n", + "text": [ + "
\n", + "I would be happy to hear your comments and suggestions. \n", + "Please feel free to drop me a note via\n", + "[twitter](https://twitter.com/rasbt), [email](mailto:bluewoodtree@gmail.com), or [google+](https://plus.google.com/+SebastianRaschka).\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### watermark is now located and maintained in a separate GitHub repository: [https://github.com/rasbt/watermark](https://github.com/rasbt/watermark)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "# IPython magic function documentation - `%watermark`" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "I wrote this simple `watermark` IPython magic function to conveniently add date- and time-stamps to my IPython notebooks. Also, I often want to document various system information, e.g., for my [Python benchmarks](https://github.com/rasbt/One-Python-benchmark-per-day) series.\n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Installation" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "The `watermark` line magic can be directly installed from my GitHub repository via" + ] + }, + { + "cell_type": "code", + "execution_count": 1, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Installed watermark.py. To use it, type:\n", + " %load_ext watermark\n" + ] + } + ], + "source": [ + "install_ext https://raw.githubusercontent.com/rasbt/watermark/master/watermark.py" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Loading the `%watermark` magic" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "To load the `date` magic, execute the following line in your IPython notebook or current IPython shell" + ] + }, + { + "cell_type": "code", + "execution_count": 2, + "metadata": { + "collapsed": false + }, + "outputs": [], + "source": [ + "%load_ext watermark" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Usage" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "In order to display the optional `watermark` arguments, type" + ] + }, + { + "cell_type": "code", + "execution_count": 12, + "metadata": { + "collapsed": false + }, + "outputs": [], + "source": [ + "%watermark?" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
%watermark [-a AUTHOR] [-d] [-e] [-n] [-t] [-z] [-u] [-c CUSTOM_TIME]\n", + " [-v] [-p PACKAGES] [-h] [-m] [-g] [-w]\n", + "\n", + " \n", + "IPython magic function to print date/time stamps \n", + "and various system information.\n", + "\n", + "watermark version 1.2.1\n", + "\n", + "optional arguments:\n", + " -a AUTHOR, --author AUTHOR\n", + " prints author name\n", + " -d, --date prints current date as MM/DD/YYYY\n", + " -e, --eurodate prints current date as DD/MM/YYYY\n", + " -n, --datename prints date with abbrv. day and month names\n", + " -t, --time prints current time\n", + " -z, --timezone appends the local time zone\n", + " -u, --updated appends a string \"Last updated: \"\n", + " -c CUSTOM_TIME, --custom_time CUSTOM_TIME\n", + " prints a valid strftime() string\n", + " -v, --python prints Python and IPython version\n", + " -p PACKAGES, --packages PACKAGES\n", + " prints versions of specified Python modules and\n", + " packages\n", + " -h, --hostname prints the host name\n", + " -m, --machine prints system and machine info\n", + " -g, --githash prints current Git commit hash\n", + " -w, --watermark prints the current version of watermark\n", + "File: ~/.ipython/extensions/watermark.py\n", + "" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Examples" + ] + }, + { + "cell_type": "code", + "execution_count": 4, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "06/29/2015 15:34:42\n", + "\n", + "CPython 3.4.3\n", + "IPython 3.2.0\n", + "\n", + "compiler : GCC 4.2.1 (Apple Inc. build 5577)\n", + "system : Darwin\n", + "release : 14.3.0\n", + "machine : x86_64\n", + "processor : i386\n", + "CPU cores : 4\n", + "interpreter: 64bit\n" + ] + } + ], + "source": [ + "%watermark" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "code", + "execution_count": 5, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "06/29/2015 15:34:43 \n" + ] + } + ], + "source": [ + "%watermark -d -t" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "code", + "execution_count": 6, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Last updated: Mon Jun 29 2015 15:34:44 EDT\n" + ] + } + ], + "source": [ + "%watermark -u -n -t -z" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "code", + "execution_count": 7, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "CPython 3.4.3\n", + "IPython 3.2.0\n" + ] + } + ], + "source": [ + "%watermark -v" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "code", + "execution_count": 8, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "compiler : GCC 4.2.1 (Apple Inc. build 5577)\n", + "system : Darwin\n", + "release : 14.3.0\n", + "machine : x86_64\n", + "processor : i386\n", + "CPU cores : 4\n", + "interpreter: 64bit\n" + ] + } + ], + "source": [ + "%watermark -m" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "code", + "execution_count": 9, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "CPython 3.4.3\n", + "IPython 3.2.0\n", + "\n", + "numpy 1.9.2\n", + "scipy 0.15.1\n", + "\n", + "compiler : GCC 4.2.1 (Apple Inc. build 5577)\n", + "system : Darwin\n", + "release : 14.3.0\n", + "machine : x86_64\n", + "processor : i386\n", + "CPU cores : 4\n", + "interpreter: 64bit\n" + ] + } + ], + "source": [ + "%watermark -v -m -p numpy,scipy" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "code", + "execution_count": 10, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "John Doe 06/29/2015 \n", + "\n", + "CPython 3.4.3\n", + "IPython 3.2.0\n", + "\n", + "compiler : GCC 4.2.1 (Apple Inc. build 5577)\n", + "system : Darwin\n", + "release : 14.3.0\n", + "machine : x86_64\n", + "processor : i386\n", + "CPU cores : 4\n", + "interpreter: 64bit\n", + "Git hash : 06830b939358f35f2daf50af42509e603c93d9b4\n" + ] + } + ], + "source": [ + "%watermark -a \"John Doe\" -d -v -m -g" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": { + "collapsed": true + }, + "outputs": [], + "source": [] + } + ], + "metadata": { + "kernelspec": { + "display_name": "Python 3", + "language": "python", + "name": "python3" + }, + "language_info": { + "codemirror_mode": { + "name": "ipython", + "version": 3 + }, + "file_extension": ".py", + "mimetype": "text/x-python", + "name": "python", + "nbconvert_exporter": "python", + "pygments_lexer": "ipython3", + "version": "3.4.3" + } + }, + "nbformat": 4, + "nbformat_minor": 0 +} diff --git a/ipython_magic/watermark.py b/ipython_magic/watermark.py new file mode 100644 index 0000000..35349a9 --- /dev/null +++ b/ipython_magic/watermark.py @@ -0,0 +1,172 @@ +""" +Sebastian Raschka 2014 + +watermark.py +version 1.1.0 + + +IPython magic function to print date/time stamps and various system information. + +Installation: + + %install_ext https://raw.githubusercontent.com/rasbt/python_reference/master/ipython_magic/watermark.py + +Usage: + + %load_ext watermark + + %watermark + +optional arguments: + + -a AUTHOR, --author AUTHOR + prints author name + -d, --date prints current date + -n, --datename prints date with abbrv. day and month names + -t, --time prints current time + -z, --timezone appends the local time zone + -u, --updated appends a string "Last updated: " + -c CUSTOM_TIME, --custom_time CUSTOM_TIME + prints a valid strftime() string + -v, --python prints Python and IPython version + -p PACKAGES, --packages PACKAGES + prints versions of specified Python modules and + packages + -h, --hostname prints the host name + -m, --machine prints system and machine info + -g, --githash prints current Git commit hash + + +Examples: + + %watermark -d -t + +""" +import platform +import subprocess +from time import strftime +from socket import gethostname +from pkg_resources import get_distribution +from multiprocessing import cpu_count + +import IPython +from IPython.core.magic import Magics, magics_class, line_magic +from IPython.core.magic_arguments import argument, magic_arguments, parse_argstring + +@magics_class +class WaterMark(Magics): + """ + IPython magic function to print date/time stamps + and various system information. + + """ + @magic_arguments() + @argument('-a', '--author', type=str, help='prints author name') + @argument('-d', '--date', action='store_true', help='prints current date') + @argument('-n', '--datename', action='store_true', help='prints date with abbrv. day and month names') + @argument('-t', '--time', action='store_true', help='prints current time') + @argument('-z', '--timezone', action='store_true', help='appends the local time zone') + @argument('-u', '--updated', action='store_true', help='appends a string "Last updated: "') + @argument('-c', '--custom_time', type=str, help='prints a valid strftime() string') + @argument('-v', '--python', action='store_true', help='prints Python and IPython version') + @argument('-p', '--packages', type=str, help='prints versions of specified Python modules and packages') + @argument('-h', '--hostname', action='store_true', help='prints the host name') + @argument('-m', '--machine', action='store_true', help='prints system and machine info') + @argument('-g', '--githash', action='store_true', help='prints current Git commit hash') + @line_magic + def watermark(self, line): + """ + IPython magic function to print date/time stamps + and various system information. + + watermark version 1.1.0 + + """ + self.out = '' + args = parse_argstring(self.watermark, line) + + if not any(vars(args).values()): + self.out += strftime('%d/%m/%Y %H:%M:%S') + self._get_pyversions() + self._get_sysinfo() + + else: + if args.author: + self.out += '% s ' %args.author.strip('\'"') + if args.updated: + self.out += 'Last updated: ' + if args.custom_time: + self.out += '%s ' %strfime(args.custom_time) + if args.date: + self.out += '%s ' %strftime('%d/%m/%Y') + elif args.datename: + self.out += '%s ' %strftime('%a %b %M %Y') + if args.time: + self.out += '%s ' %strftime('%H:%M:%S') + if args.timezone: + self.out += strftime('%Z') + if args.python: + self._get_pyversions() + if args.packages: + self._get_packages(args.packages) + if args.machine: + self._get_sysinfo() + if args.hostname: + space = '' + if args.machine: + space = ' ' + self.out += '\nhost name%s: %s' %(space, gethostname()) + if args.githash: + self._get_commit_hash(bool(args.machine)) + + + + + print(self.out) + + + def _get_packages(self, pkgs): + if self.out: + self.out += '\n' + packages = pkgs.split(',') + for p in packages: + self.out += '\n%s %s' %(p, get_distribution(p).version) + + + def _get_pyversions(self): + if self.out: + self.out += '\n\n' + self.out += '%s %s\nIPython %s' %( + platform.python_implementation(), + platform.python_version(), + IPython.__version__ + ) + + + def _get_sysinfo(self): + if self.out: + self.out += '\n\n' + self.out += 'compiler : %s\nsystem : %s\n'\ + 'release : %s\nmachine : %s\n'\ + 'processor : %s\nCPU cores : %s\ninterpreter: %s'%( + platform.python_compiler(), + platform.system(), + platform.release(), + platform.machine(), + platform.processor(), + cpu_count(), + platform.architecture()[0] + ) + + + def _get_commit_hash(self, machine): + process = subprocess.Popen(['git', 'rev-parse', 'HEAD'], shell=False, stdout=subprocess.PIPE) + git_head_hash = process.communicate()[0].strip() + space = '' + if machine: + space = ' ' + self.out += '\nGit hash%s: %s' %(space, git_head_hash.decode("utf-8")) + + +def load_ipython_extension(ipython): + ipython.register_magics(WaterMark) diff --git a/normalize_data.py b/normalize_data.py deleted file mode 100644 index 117d2fb..0000000 --- a/normalize_data.py +++ /dev/null @@ -1,15 +0,0 @@ -# Sebastian Raschka, 03/2014 - -def normalize_val(x, data_list): - """ - Normalizes a value to a data list returning a float - between 0.0 and 1.0. - Returns the original object if value is not a integer or float. - - """ - if isinstance(x, float) or isinstance(x, int): - numerator = x - min(data_list) - denominator = max(data_list) - min(data_list) - return numerator/denominator - else: - return x diff --git a/not_so_obvious_python_stuff.ipynb b/not_so_obvious_python_stuff.ipynb deleted file mode 100644 index c48d0de..0000000 --- a/not_so_obvious_python_stuff.ipynb +++ /dev/null @@ -1,3037 +0,0 @@ -{ - "metadata": { - "name": "", - "signature": "sha256:9654ab25e9d989ee8f196ec7cd5c2b7222157dbe195ddf70e6ec34329c9db545" - }, - "nbformat": 3, - "nbformat_minor": 0, - "worksheets": [ - { - "cells": [ - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Sebastian Raschka \n", - "last updated: 04/27/2014 ([Changelog](#changelog))\n", - "\n", - "[Link to this IPython Notebook on GitHub](https://github.com/rasbt/python_reference/blob/master/not_so_obvious_python_stuff.ipynb)\n", - "\n", - "\n" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "#### All code was executed in Python 3.4" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "# A collection of not-so-obvious Python stuff you should know!" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "I am really looking forward to your comments and suggestions to improve and \n", - "extend this little collection! Just send me a quick note \n", - "via Twitter: [@rasbt](https://twitter.com/rasbt) \n", - "or Email: [bluewoodtree@gmail.com](mailto:bluewoodtree@gmail.com)\n", - "
" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "# Sections\n", - "- [The C3 class resolution algorithm for multiple class inheritance](#c3_class_res)\n", - "- [Assignment operators and lists - simple-add vs. add-AND operators](#pm_in_lists)\n", - "- [`True` and `False` in the datetime module](#datetime_module)\n", - "- [Python reuses objects for small integers - always use \"==\" for equality, \"is\" for identity](#python_small_int)\n", - "- [Shallow vs. deep copies if list contains other structures and objects](#shallow_vs_deep)\n", - "- [Picking `True` values from logical `and`s and `or`s](#false_true_expressions)\n", - "- [Don't use mutable objects as default arguments for functions!](#def_mutable_func)\n", - "- [Be aware of the consuming generator](#consuming_generator)\n", - "- [`bool` is a subclass of `int`](#bool_int)\n", - "- [About lambda-in-closures and-a-loop pitfall](#lambda_closure)\n", - "- [Python's LEGB scope resolution and the keywords `global` and `nonlocal`](#python_legb)\n", - "- [When mutable contents of immutable tuples aren't so mutable](#immutable_tuple)\n", - "- [List comprehensions are fast, but generators are faster!?](#list_generator)\n", - "- [Public vs. private class methods and name mangling](#private_class)\n", - "- [The consequences of modifying a list when looping through it](#looping_pitfall)\n", - "- [Dynamic binding and typos in variable names](#dynamic_binding)\n", - "- [List slicing using indexes that are \"out of range](#out_of_range_slicing)\n", - "- [Reusing global variable names and UnboundLocalErrors](#unboundlocalerror)\n", - "- [Creating copies of mutable objects](#copy_mutable)\n", - "- [Key differences between Python 2 and 3](#python_differences)\n", - "- [Function annotations - What are those `->`'s in my Python code?](#function_annotation)\n", - "- [Abortive statements in `finally` blocks](#finally_blocks)\n", - "- [Assigning types to variables as values](#variable_types)" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "
\n", - "
" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "## The C3 class resolution algorithm for multiple class inheritance" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "If we are dealing with multiple inheritance, according to the newer C3 class resolution algorithm, the following applies: \n", - "Assuming that child class C inherits from two parent classes A and B, \"class A should be checked before class B\".\n", - "\n", - "If you want to learn more, please read the [original blog](http://python-history.blogspot.ru/2010/06/method-resolution-order.html) post by Guido van Rossum.\n", - "\n", - "(Original source: [http://gistroll.com/rolls/21/horizontal_assessments/new](http://gistroll.com/rolls/21/horizontal_assessments/new))" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "class A(object):\n", - " def foo(self):\n", - " print(\"class A\")\n", - "\n", - "class B(object):\n", - " def foo(self):\n", - " print(\"class B\")\n", - "\n", - "class C(A, B):\n", - " pass\n", - "\n", - "C().foo()" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "class A\n" - ] - } - ], - "prompt_number": 2 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "So what actually happened above was that class `C` looked in the scope of the parent class `A` for the method `.foo()` first (and found it)!" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "I received an email containing a suggestion which uses a more nested example to illustrate Guido van Rossum's point a little bit better:" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "class A(object):\n", - " def foo(self):\n", - " print(\"class A\")\n", - "\n", - "class B(A):\n", - " pass\n", - "\n", - "class C(A):\n", - " def foo(self):\n", - " print(\"class C\")\n", - "\n", - "class D(B,C):\n", - " pass\n", - "\n", - "D().foo()" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "class C\n" - ] - } - ], - "prompt_number": 3 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Here, class `D` searches in `B` first, which in turn inherits from `A` (note that class `C` also inherits from `A`, but has its own `.foo()` method) so that we come up with the search order: `D, B, C, A`. " - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "
\n", - "
" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "## Assignment operators and lists - simple-add vs. add-AND operators" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Python `list`s are mutable objects as we all know. So, if we are using the `+=` operator on `list`s, we extend the `list` by directly modifying the object directly. \n", - "\n", - "However, if we use the assigment via `my_list = my_list + ...`, we create a new list object, which can be demonstrated by the following code:" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "a_list = []\n", - "print('ID:', id(a_list))\n", - "\n", - "a_list += [1]\n", - "print('ID (+=):', id(a_list))\n", - "\n", - "a_list = a_list + [2]\n", - "print('ID (list = list + ...):', id(a_list))" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "ID: 4366496544\n", - "ID (+=): 4366496544\n", - "ID (list = list + ...): 4366495472\n" - ] - } - ], - "prompt_number": 6 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Just for reference, the `.append()` and `.extends()` methods are modifying the `list` object in place, just as expected." - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "a_list = []\n", - "print('ID:',id(a_list))\n", - "\n", - "a_list.append(1)\n", - "print('ID (append):',id(a_list))\n", - "\n", - "a_list.append(2)\n", - "print('ID (extend):',id(a_list))" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "ID: 4366495544\n", - "ID (append): 4366495544\n", - "ID (extend): 4366495544\n" - ] - } - ], - "prompt_number": 7 - }, - { - "cell_type": "code", - "collapsed": false, - "input": [], - "language": "python", - "metadata": {}, - "outputs": [] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "
\n", - "" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "## `True` and `False` in the datetime module\n", - "\n" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\"It often comes as a big surprise for programmers to find (sometimes by way of a hard-to-reproduce bug) that, unlike any other time value, midnight (i.e. `datetime.time(0,0,0)`) is False. A long discussion on the python-ideas mailing list shows that, while surprising, that behavior is desirable\u2014at least in some quarters.\" \n", - "\n", - "(Original source: [http://lwn.net/SubscriberLink/590299/bf73fe823974acea/](http://lwn.net/SubscriberLink/590299/bf73fe823974acea/))" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "import datetime\n", - "\n", - "print('\"datetime.time(0,0,0)\" (Midnight) ->', bool(datetime.time(0,0,0)))\n", - "\n", - "print('\"datetime.time(1,0,0)\" (1 am) ->', bool(datetime.time(1,0,0)))" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\"datetime.time(0,0,0)\" (Midnight) -> False\n", - "\"datetime.time(1,0,0)\" (1 am) -> True\n" - ] - } - ], - "prompt_number": 8 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "
\n", - "" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "## Python reuses objects for small integers - use \"==\" for equality, \"is\" for identity\n", - "\n" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "This oddity occurs, because Python keeps an array of small integer objects (i.e., integers between -5 and 256, [see the doc](https://docs.python.org/2/c-api/int.html#PyInt_FromLong))." - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "a = 1\n", - "b = 1\n", - "print('a is b', bool(a is b))\n", - "True\n", - "\n", - "c = 999\n", - "d = 999\n", - "print('c is d', bool(c is d))" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "a is b True\n", - "c is d False\n" - ] - } - ], - "prompt_number": 9 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "(*I received a comment that this is in fact a CPython artefact and **must not necessarily be true** in all implementations of Python!*)\n", - "\n", - "So the take home message is: always use \"==\" for equality, \"is\" for identity!\n", - "\n", - "Here is a [nice article](http://python.net/%7Egoodger/projects/pycon/2007/idiomatic/handout.html#other-languages-have-variables) explaining it by comparing \"boxes\" (C language) with \"name tags\" (Python)." - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "This example demonstrates that this applies indeed for integers in the range in -5 to 256:" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "print('256 is 257-1', 256 is 257-1)\n", - "print('257 is 258-1', 257 is 258 - 1)\n", - "print('-5 is -6+1', -5 is -6+1)\n", - "print('-7 is -6-1', -7 is -6-1)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "256 is 257-1 True\n", - "257 is 258-1 False\n", - "-5 is -6+1 True\n", - "-7 is -6-1 False\n" - ] - } - ], - "prompt_number": 11 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "#### And to illustrate the test for equality (`==`) vs. identity (`is`):" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "a = 'hello world!'\n", - "b = 'hello world!'\n", - "print('a is b,', a is b)\n", - "print('a == b,', a == b)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "a is b, False\n", - "a == b, True\n" - ] - } - ], - "prompt_number": 6 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "We would think that identity would always imply equality, but this is not always true, as we can see in the next example:" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "a = float('nan')\n", - "print('a is a,', a is a)\n", - "print('a == a,', a == a)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "a is a, True\n", - "a == a, False\n" - ] - } - ], - "prompt_number": 12 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "
\n", - "" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "## Shallow vs. deep copies if list contains other structures and objects\n", - "\n" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "**Shallow copy**: \n", - "If we use the assignment operator to assign one list to another list, we just create a new name reference to the original list. If we want to create a new list object, we have to make a copy of the original list. This can be done via `a_list[:]` or `a_list.copy()`." - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "list1 = [1,2]\n", - "list2 = list1 # reference\n", - "list3 = list1[:] # shallow copy\n", - "list4 = list1.copy() # shallow copy\n", - "\n", - "print('IDs:\\nlist1: {}\\nlist2: {}\\nlist3: {}\\nlist4: {}\\n'\n", - " .format(id(list1), id(list2), id(list3), id(list4)))\n", - "\n", - "list2[0] = 3\n", - "print('list1:', list1)\n", - "\n", - "list3[0] = 4\n", - "list4[1] = 4\n", - "print('list1:', list1)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "IDs:\n", - "list1: 4346366472\n", - "list2: 4346366472\n", - "list3: 4346366408\n", - "list4: 4346366536\n", - "\n", - "list1: [3, 2]\n", - "list1: [3, 2]\n" - ] - } - ], - "prompt_number": 1 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "**Deep copy** \n", - "As we have seen above, a shallow copy works fine if we want to create a new list with contents of the original list which we want to modify independently. \n", - "\n", - "However, if we are dealing with compound objects (e.g., lists that contain other lists, [read here](https://docs.python.org/2/library/copy.html) for more information) it becomes a little trickier.\n", - "\n", - "In the case of compound objects, a shallow copy would create a new compound object, but it would just insert the references to the contained objects into the new compound object. In contrast, a deep copy would go \"deeper\" and create also new objects \n", - "for the objects found in the original compound object. \n", - "If you follow the code, the concept should become more clear:" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "from copy import deepcopy\n", - "\n", - "list1 = [[1],[2]]\n", - "list2 = list1.copy() # shallow copy\n", - "list3 = deepcopy(list1) # deep copy\n", - "\n", - "print('IDs:\\nlist1: {}\\nlist2: {}\\nlist3: {}\\n'\n", - " .format(id(list1), id(list2), id(list3)))\n", - "\n", - "list2[0][0] = 3\n", - "print('list1:', list1)\n", - "\n", - "list3[0][0] = 5\n", - "print('list1:', list1)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "IDs:\n", - "list1: 4377956296\n", - "list2: 4377961752\n", - "list3: 4377954928\n", - "\n", - "list1: [[3], [2]]\n", - "list1: [[3], [2]]\n" - ] - } - ], - "prompt_number": 25 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "
\n", - "" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "## Picking `True` values from logical `and`s and `or`s" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "** Logical `or`: ** \n", - "\n", - "`a or b == a if a else b` \n", - "- If both values in `or` expressions are `True`, Python will select the first value (e.g., select `\"a\"` in `\"a\" or \"b\"`), and the second one in `and` expressions. \n", - "This is also called **short-circuiting** - we already know that the logical `or` must be `True` if the first value is `True` and therefore can omit the evaluation of the second value.\n", - "\n", - "** Logical `and`: ** \n", - "\n", - "`a and b == b if a else a` \n", - "- If both values in `and` expressions are `True`, Python will select the second value, since for a logical `and`, both values must be true.\n" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "result = (2 or 3) * (5 and 7)\n", - "print('2 * 7 =', result)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "2 * 7 = 14\n" - ] - } - ], - "prompt_number": 9 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "
\n", - "" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "## Don't use mutable objects as default arguments for functions!" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Don't use mutable objects (e.g., dictionaries, lists, sets, etc.) as default arguments for functions! You might expect that a new list is created every time when we call the function without providing an argument for the default parameter, but this is not the case: **Python will create the mutable object (default parameter) the first time the function is defined - not when it is called**, see the following code:\n", - "\n", - "(Original source: [http://docs.python-guide.org/en/latest/writing/gotchas/](http://docs.python-guide.org/en/latest/writing/gotchas/)" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "def append_to_list(value, def_list=[]):\n", - " def_list.append(value)\n", - " return def_list\n", - "\n", - "my_list = append_to_list(1)\n", - "print(my_list)\n", - "\n", - "my_other_list = append_to_list(2)\n", - "print(my_other_list)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "[1]\n", - "[1, 2]\n" - ] - } - ], - "prompt_number": 1 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Another good example showing that demonstrates that default arguments are created when the function is created (**and not when it is called!**):" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "import time\n", - "def report_arg(my_default=time.time()):\n", - " print(my_default)\n", - "\n", - "report_arg()\n", - "\n", - "time.sleep(5)\n", - "\n", - "report_arg()" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "1397764090.456688\n", - "1397764090.456688" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n" - ] - } - ], - "prompt_number": 10 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "
\n", - "\n" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "## Be aware of the consuming generator" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Be aware of what is happening when combining \"`in`\" checks with generators, since they won't evaluate from the beginning once a position is \"consumed\"." - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "gen = (i for i in range(5))\n", - "print('2 in gen,', 2 in gen)\n", - "print('3 in gen,', 3 in gen)\n", - "print('1 in gen,', 1 in gen) " - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "2 in gen, True\n", - "3 in gen, True\n", - "1 in gen, False\n" - ] - } - ], - "prompt_number": 9 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Although this defeats the purpose of an generator (in most cases), we can convert a generator into a list to circumvent the problem. " - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "gen = (i for i in range(5))\n", - "a_list = list(gen)\n", - "print('2 in l,', 2 in a_list)\n", - "print('3 in l,', 3 in a_list)\n", - "print('1 in l,', 1 in a_list) " - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "2 in l, True\n", - "3 in l, True\n", - "1 in l, True\n" - ] - } - ], - "prompt_number": 27 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "
\n", - "\n" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "## `bool` is a subclass of `int`\n", - "\n" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Chicken or egg? In the history of Python (Python 2.2 to be specific) truth values were implemented via 1 and 0 (similar to the old C). In order to avoid syntax errors in old (but perfectly working) Python code, `bool` was added as a subclass of `int` in Python 2.3.\n", - "\n", - "Original source: [http://www.peterbe.com/plog/bool-is-int](http://www.peterbe.com/plog/bool-is-int)" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "print('isinstance(True, int):', isinstance(True, int))\n", - "print('True + True:', True + True)\n", - "print('3*True + True:', 3*True + True)\n", - "print('3*True - False:', 3*True - False)\n" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "isinstance(True, int): True\n", - "True + True: 2\n", - "3*True + True: 4\n", - "3*True - False: 3\n" - ] - } - ], - "prompt_number": 28 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "
\n", - "" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "## About lambda-in-closures-and-a-loop pitfall" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Remember the section about the [\"consuming generators\"](consuming_generators)? This example is somewhat related, but the result might still come unexpected. \n", - "\n", - "(Original source: [http://openhome.cc/eGossip/Blog/UnderstandingLambdaClosure3.html](http://openhome.cc/eGossip/Blog/UnderstandingLambdaClosure3.html))\n", - "\n", - "In the first example below, we call a `lambda` function in a list comprehension, and the value `i` will be dereferenced every time we call `lambda` within the scope of the list comprehension. Since the list is already constructed when we `for-loop` through the list, it will be set to the last value 4." - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "my_list = [lambda: i for i in range(5)]\n", - "for l in my_list:\n", - " print(l())" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "4\n", - "4\n", - "4\n", - "4\n", - "4\n" - ] - } - ], - "prompt_number": 7 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "This, however, does not apply to generators:" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "my_gen = (lambda: n for n in range(5))\n", - "for l in my_gen:\n", - " print(l())" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "0\n", - "1\n", - "2\n", - "3\n", - "4\n" - ] - } - ], - "prompt_number": 9 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "And if you are really keen on using lists, there is a nifty trick that circumvents this problem as a reader nicely pointed out in the comments: We can simply pass the loop variable `i` as a default argument to the lambdas." - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "my_list = [lambda x=i: x for i in range(5)]\n", - "for l in my_list:\n", - " print(l())" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "0\n", - "1\n", - "2\n", - "3\n", - "4\n" - ] - } - ], - "prompt_number": 10 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "
\n", - "" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "## Python's LEGB scope resolution and the keywords `global` and `nonlocal`" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "There is nothing particularly surprising about Python's LEGB scope resolution (Local -> Enclosed -> Global -> Built-in), but it is still useful to take a look at some examples!" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "### `global` vs. `local`\n", - "\n", - "According to the LEGB rule, Python will first look for a variable in the local scope. So if we set the variable `x = 1` `local`ly in the function's scope, it won't have an effect on the `global` `x`." - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "x = 0\n", - "def in_func():\n", - " x = 1\n", - " print('in_func:', x)\n", - " \n", - "in_func()\n", - "print('global:', x)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "in_func: 1\n", - "global: 0\n" - ] - } - ], - "prompt_number": 31 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "If we want to modify the `global` x via a function, we can simply use the `global` keyword to import the variable into the function's scope:" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "x = 0\n", - "def in_func():\n", - " global x\n", - " x = 1\n", - " print('in_func:', x)\n", - " \n", - "in_func()\n", - "print('global:', x)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "in_func: 1\n", - "global: 1\n" - ] - } - ], - "prompt_number": 34 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "### `local` vs. `enclosed`\n", - "\n", - "Now, let us take a look at `local` vs. `enclosed`. Here, we set the variable `x = 1` in the `outer` function and set `x = 1` in the enclosed function `inner`. Since `inner` looks in the local scope first, it won't modify `outer`'s `x`." - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "def outer():\n", - " x = 1\n", - " print('outer before:', x)\n", - " def inner():\n", - " x = 2\n", - " print(\"inner:\", x)\n", - " inner()\n", - " print(\"outer after:\", x)\n", - "outer()" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "outer before: 1\n", - "inner: 2\n", - "outer after: 1\n" - ] - } - ], - "prompt_number": 36 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Here is where the `nonlocal` keyword comes in handy - it allows us to modify the `x` variable in the `enclosed` scope:" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "def outer():\n", - " x = 1\n", - " print('outer before:', x)\n", - " def inner():\n", - " nonlocal x\n", - " x = 2\n", - " print(\"inner:\", x)\n", - " inner()\n", - " print(\"outer after:\", x)\n", - "outer()" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "outer before: 1\n", - "inner: 2\n", - "outer after: 2\n" - ] - } - ], - "prompt_number": 35 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "
\n", - "" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "## When mutable contents of immutable tuples aren't so mutable" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "As we all know, tuples are immutable objects in Python, right!? But what happens if they contain mutable objects? \n", - "\n", - "First, let us have a look at the expected behavior: a `TypeError` is raised if we try to modify immutable types in a tuple: " - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "tup = (1,)\n", - "tup[0] += 1" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "ename": "TypeError", - "evalue": "'tuple' object does not support item assignment", - "output_type": "pyerr", - "traceback": [ - "\u001b[0;31m---------------------------------------------------------------------------\u001b[0m\n\u001b[0;31mTypeError\u001b[0m Traceback (most recent call last)", - "\u001b[0;32m
\n", - "
\n", - "However, **there are ways** to modify the mutable contents of the tuple without raising the `TypeError`, the solution is the `.extend()` method, or alternatively `.append()` (for lists):" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "tup = ([],)\n", - "print('tup before: ', tup)\n", - "tup[0].extend([1])\n", - "print('tup after: ', tup)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "tup before: ([],)\n", - "tup after: ([1],)\n" - ] - } - ], - "prompt_number": 44 - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "tup = ([],)\n", - "print('tup before: ', tup)\n", - "tup[0].append(1)\n", - "print('tup after: ', tup)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "tup before: ([],)\n", - "tup after: ([1],)\n" - ] - } - ], - "prompt_number": 5 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "### Explanation\n", - "\n", - "**A. Jesse Jiryu Davis** has a nice explanation for this phenomenon (Original source: [http://emptysqua.re/blog/python-increment-is-weird-part-ii/](http://emptysqua.re/blog/python-increment-is-weird-part-ii/))\n", - "\n", - "If we try to extend the list via `+=` *\"then the statement executes `STORE_SUBSCR`, which calls the C function `PyObject_SetItem`, which checks if the object supports item assignment. In our case the object is a tuple, so `PyObject_SetItem` throws the `TypeError`. Mystery solved.\"*" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "#### One more note about the `immutable` status of tuples. Tuples are famous for being immutable. However, how comes that this code works?" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "my_tup = (1,)\n", - "my_tup += (4,)\n", - "my_tup = my_tup + (5,)\n", - "print(my_tup)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "(1, 4, 5)\n" - ] - } - ], - "prompt_number": 6 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "What happens \"behind\" the curtains is that the tuple is not modified, but every time a new object is generated, which will inherit the old \"name tag\":" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "my_tup = (1,)\n", - "print(id(my_tup))\n", - "my_tup += (4,)\n", - "print(id(my_tup))\n", - "my_tup = my_tup + (5,)\n", - "print(id(my_tup))" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "4337381840\n", - "4357415496\n", - "4357289952\n" - ] - } - ], - "prompt_number": 8 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "
\n", - "\n" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "## List comprehensions are fast, but generators are faster!?" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\"List comprehensions are fast, but generators are faster!?\" - No, not really (or significantly, see the benchmarks below). So what's the reason to prefer one over the other?\n", - "- use lists if you want to use the plethora of list methods \n", - "- use generators when you are dealing with huge collections to avoid memory issues" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "import timeit\n", - "\n", - "def plainlist(n=100000):\n", - " my_list = []\n", - " for i in range(n):\n", - " if i % 5 == 0:\n", - " my_list.append(i)\n", - " return my_list\n", - "\n", - "def listcompr(n=100000):\n", - " my_list = [i for i in range(n) if i % 5 == 0]\n", - " return my_list\n", - "\n", - "def generator(n=100000):\n", - " my_gen = (i for i in range(n) if i % 5 == 0)\n", - " return my_gen\n", - "\n", - "def generator_yield(n=100000):\n", - " for i in range(n):\n", - " if i % 5 == 0:\n", - " yield i" - ], - "language": "python", - "metadata": {}, - "outputs": [], - "prompt_number": 11 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "#### To be fair to the list, let us exhaust the generators:" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "def test_plainlist(plain_list):\n", - " for i in plain_list():\n", - " pass\n", - "\n", - "def test_listcompr(listcompr):\n", - " for i in listcompr():\n", - " pass\n", - "\n", - "def test_generator(generator):\n", - " for i in generator():\n", - " pass\n", - "\n", - "def test_generator_yield(generator_yield):\n", - " for i in generator_yield():\n", - " pass\n", - "\n", - "print('plain_list: ', end = '')\n", - "%timeit test_plainlist(plainlist)\n", - "print('\\nlistcompr: ', end = '')\n", - "%timeit test_listcompr(listcompr)\n", - "print('\\ngenerator: ', end = '')\n", - "%timeit test_generator(generator)\n", - "print('\\ngenerator_yield: ', end = '')\n", - "%timeit test_generator_yield(generator_yield)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "plain_list: 10 loops, best of 3: 22.4 ms per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "\n", - "listcompr: 10 loops, best of 3: 20.8 ms per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "\n", - "generator: 10 loops, best of 3: 22 ms per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n", - "\n", - "generator_yield: 10 loops, best of 3: 21.9 ms per loop" - ] - }, - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "\n" - ] - } - ], - "prompt_number": 13 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "
\n", - "" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "## Public vs. private class methods and name mangling\n", - "\n" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Who has not stumbled across this quote \"we are all consenting adults here\" in the Python community, yet? Unlike in other languages like C++ (sorry, there are many more, but that's one I am most familiar with), we can't really protect class methods from being used outside the class (i.e., by the API user). \n", - "All we can do is to indicate methods as private to make clear that they are better not used outside the class, but it is really up to the class user, since \"we are all consenting adults here\"! \n", - "So, when we want to mark a class method as private, we can put a single underscore in front of it. \n", - "If we additionally want to avoid name clashes with other classes that might use the same method names, we can prefix the name with a double-underscore to invoke the name mangling.\n", - "\n", - "This doesn't prevent the class user to access this class member though, but he has to know the trick and also knows that it his own risk...\n", - "\n", - "Let the following example illustrate what I mean:" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "class my_class():\n", - " def public_method(self):\n", - " print('Hello public world!')\n", - " def __private_method(self):\n", - " print('Hello private world!')\n", - " def call_private_method_in_class(self):\n", - " self.__private_method()\n", - " \n", - "my_instance = my_class()\n", - "\n", - "my_instance.public_method()\n", - "my_instance._my_class__private_method()\n", - "my_instance.call_private_method_in_class()" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "Hello public world!\n", - "Hello private world!\n", - "Hello private world!\n" - ] - } - ], - "prompt_number": 11 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "
\n", - "" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "## The consequences of modifying a list when looping through it" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "It can be really dangerous to modify a list when iterating through it - this is a very common pitfall that can cause unintended behavior! \n", - "Look at the following examples, and for a fun exercise: try to figure out what is going on before you skip to the solution!" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "a = [1, 2, 3, 4, 5]\n", - "for i in a:\n", - " if not i % 2:\n", - " a.remove(i)\n", - "print(a)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "[1, 3, 5]\n" - ] - } - ], - "prompt_number": 3 - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "b = [2, 4, 5, 6]\n", - "for i in b:\n", - " if not i % 2:\n", - " b.remove(i)\n", - "print(b)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "[4, 5]\n" - ] - } - ], - "prompt_number": 4 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "
\n", - "**The solution** is that we are iterating through the list index by index, and if we remove one of the items in-between, we inevitably mess around with the indexing, look at the following example, and it will become clear:" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "b = [2, 4, 5, 6]\n", - "for index, item in enumerate(b):\n", - " print(index, item)\n", - " if not item % 2:\n", - " b.remove(item)\n", - "print(b)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "0 2\n", - "1 5\n", - "2 6\n", - "[4, 5]\n" - ] - } - ], - "prompt_number": 7 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "
\n", - "" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "## Dynamic binding and typos in variable names\n" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Be careful, dynamic binding is convenient, but can also quickly become dangerous!" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "print('first list:')\n", - "for i in range(3):\n", - " print(i)\n", - " \n", - "print('\\nsecond list:')\n", - "for j in range(3):\n", - " print(i) # I (intentionally) made typo here!" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "first list:\n", - "0\n", - "1\n", - "2\n", - "\n", - "second list:\n", - "2\n", - "2\n", - "2\n" - ] - } - ], - "prompt_number": 14 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "
\n", - "" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "## List slicing using indexes that are \"out of range\"" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "As we have all encountered it 1 (x10000) time(s) in our live, the infamous `IndexError`:" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "my_list = [1, 2, 3, 4, 5]\n", - "print(my_list[5])" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "ename": "IndexError", - "evalue": "list index out of range", - "output_type": "pyerr", - "traceback": [ - "\u001b[0;31m---------------------------------------------------------------------------\u001b[0m\n\u001b[0;31mIndexError\u001b[0m Traceback (most recent call last)", - "\u001b[0;32m
\n", - "
\n", - "" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "## Reusing global variable names and `UnboundLocalErrors`" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Usually, it is no problem to access global variables in the local scope of a function:" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "def my_func():\n", - " print(var)\n", - "\n", - "var = 'global'\n", - "my_func()" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "global\n" - ] - } - ], - "prompt_number": 37 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "And is also no problem to use the same variable name in the local scope without affecting the local counterpart: " - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "def my_func():\n", - " var = 'locally changed'\n", - "\n", - "var = 'global'\n", - "my_func()\n", - "print(var)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "global\n" - ] - } - ], - "prompt_number": 38 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "But we have to be careful if we use a variable name that occurs in the global scope, and we want to access it in the local function scope if we want to reuse this name:" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "def my_func():\n", - " print(var) # want to access global variable\n", - " var = 'locally changed' # but Python thinks we forgot to define the local variable!\n", - " \n", - "var = 'global'\n", - "my_func()" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "ename": "UnboundLocalError", - "evalue": "local variable 'var' referenced before assignment", - "output_type": "pyerr", - "traceback": [ - "\u001b[0;31m---------------------------------------------------------------------------\u001b[0m\n\u001b[0;31mUnboundLocalError\u001b[0m Traceback (most recent call last)", - "\u001b[0;32m
\n", - "
\n", - "" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "## Creating copies of mutable objects\n", - "\n" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Let's assume a scenario where we want to duplicate sub`list`s of values stored in another list. If we want to create independent sub`list` object, using the arithmetic multiplication operator could lead to rather unexpected (or undesired) results:" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "my_list1 = [[1, 2, 3]] * 2\n", - "\n", - "print('initially ---> ', my_list1)\n", - "\n", - "# modify the 1st element of the 2nd sublist\n", - "my_list1[1][0] = 'a'\n", - "print(\"after my_list1[1][0] = 'a' ---> \", my_list1)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "initially ---> [[1, 2, 3], [1, 2, 3]]\n", - "after my_list1[1][0] = 'a' ---> [['a', 2, 3], ['a', 2, 3]]\n" - ] - } - ], - "prompt_number": 24 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "
\n", - "In this case, we should better create \"new\" objects:" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "my_list2 = [[1, 2, 3] for i in range(2)]\n", - "\n", - "print('initially: ---> ', my_list2)\n", - "\n", - "# modify the 1st element of the 2nd sublist\n", - "my_list2[1][0] = 'a'\n", - "print(\"after my_list2[1][0] = 'a': ---> \", my_list2)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "initially: ---> [[1, 2, 3], [1, 2, 3]]\n", - "after my_list2[1][0] = 'a': ---> [[1, 2, 3], ['a', 2, 3]]\n" - ] - } - ], - "prompt_number": 25 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "And here is the proof:" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "for a,b in zip(my_list1, my_list2):\n", - " print('id my_list1: {}, id my_list2: {}'.format(id(a), id(b)))" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "id my_list1: 4350764680, id my_list2: 4350766472\n", - "id my_list1: 4350764680, id my_list2: 4350766664\n" - ] - } - ], - "prompt_number": 26 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "
\n", - "" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "## Key differences between Python 2 and 3\n", - "\n" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "There are some good articles already that are summarizing the differences between Python 2 and 3, e.g., \n", - "- [https://wiki.python.org/moin/Python2orPython3](https://wiki.python.org/moin/Python2orPython3)\n", - "- [https://docs.python.org/3.0/whatsnew/3.0.html](https://docs.python.org/3.0/whatsnew/3.0.html)\n", - "- [http://python3porting.com/differences.html](http://python3porting.com/differences.html)\n", - "- [https://docs.python.org/3/howto/pyporting.html](https://docs.python.org/3/howto/pyporting.html) \n", - "etc.\n", - "\n", - "But it might be still worthwhile, especially for Python newcomers, to take a look at some of those!\n", - "(Note: the the code was executed in Python 3.4.0 and Python 2.7.5 and copied from interactive shell sessions.)" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "### Unicode...\n", - "####- Python 2: \n", - "We have ASCII `str()` types, separate `unicode()`, but no `byte` type\n", - "####- Python 3: \n", - "Now, we finally have Unicode (utf-8) `str`ings, and 2 byte classes: `byte` and `bytearray`s" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "#############\n", - "# Python 2\n", - "#############\n", - "\n", - ">>> type(unicode('is like a python3 str()'))\n", - "
\n", - "
\n", - "" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "\n", - "## Function annotations - What are those `->`'s in my Python code?\n" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Have you ever seen any Python code that used colons inside the parantheses of a function definition?" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "def foo1(x: 'insert x here', y: 'insert x^2 here'):\n", - " print('Hello, World')\n", - " return" - ], - "language": "python", - "metadata": {}, - "outputs": [], - "prompt_number": 8 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "And what about the fancy arrow here?" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "def foo2(x, y) -> 'Hi!':\n", - " print('Hello, World')\n", - " return" - ], - "language": "python", - "metadata": {}, - "outputs": [], - "prompt_number": 10 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Q: Is this valid Python syntax? \n", - "A: Yes!\n", - " \n", - " \n", - "Q: So, what happens if I *just call* the function? \n", - "A: Nothing!\n", - " \n", - "Here is the proof!" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "foo1(1,2)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "Hello, World\n" - ] - } - ], - "prompt_number": 9 - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "foo2(1,2) " - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "Hello, World\n" - ] - } - ], - "prompt_number": 11 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "**So, those are function annotations ... ** \n", - "- the colon for the function parameters \n", - "- the arrow for the return value \n", - "\n", - "You probably will never make use of them (or at least very rarely). Usually, we write good function documentations below the function as a docstring - or at least this is how I would do it (okay this case is a little bit extreme, I have to admit):" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "def is_palindrome(a):\n", - " \"\"\"\n", - " Case-and punctuation insensitive check if a string is a palindrom.\n", - " \n", - " Keyword arguments:\n", - " a (str): The string to be checked if it is a palindrome.\n", - " \n", - " Returns `True` if input string is a palindrome, else False.\n", - " \n", - " \"\"\"\n", - " stripped_str = [l for l in my_str.lower() if l.isalpha()]\n", - " return stripped_str == stripped_str[::-1]\n", - " " - ], - "language": "python", - "metadata": {}, - "outputs": [] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "However, function annotations can be useful to indicate that work is still in progress in some cases. But they are optional and I see them very very rarely.\n", - "\n", - "As it is stated in [PEP3107](http://legacy.python.org/dev/peps/pep-3107/#fundamentals-of-function-annotations):\n", - "\n", - "1. Function annotations, both for parameters and return values, are completely optional.\n", - "\n", - "2. Function annotations are nothing more than a way of associating arbitrary Python expressions with various parts of a function at compile-time.\n" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "The nice thing about function annotations is their `__annotations__` attribute, which is dictionary of all the parameters and/or the `return` value you annotated." - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "foo1.__annotations__" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "metadata": {}, - "output_type": "pyout", - "prompt_number": 17, - "text": [ - "{'y': 'insert x^2 here', 'x': 'insert x here'}" - ] - } - ], - "prompt_number": 17 - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "foo2.__annotations__" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "metadata": {}, - "output_type": "pyout", - "prompt_number": 18, - "text": [ - "{'return': 'Hi!'}" - ] - } - ], - "prompt_number": 18 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "**When are they useful?**" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Function annotations can be useful for a couple of things \n", - "- Documentation in general\n", - "- pre-condition testing\n", - "- [type checking](http://legacy.python.org/dev/peps/pep-0362/#annotation-checker)\n", - " \n", - "..." - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "
\n", - "" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "## Abortive statements in `finally` blocks" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "Python's `try-except-finally` blocks are very handy for catching and handling errors. The `finally` block is always executed whether an `exception` has been raised or not as illustrated in the following example." - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "def try_finally1():\n", - " try:\n", - " print('in try:')\n", - " print('do some stuff')\n", - " float('abc')\n", - " except ValueError:\n", - " print('an error occurred')\n", - " else:\n", - " print('no error occurred')\n", - " finally:\n", - " print('always execute finally')\n", - " \n", - "try_finally1()" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "in try:\n", - "do some stuff\n", - "an error occurred\n", - "always execute finally\n" - ] - } - ], - "prompt_number": 24 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "
\n", - "But can you also guess what will be printed in the next code cell?" - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "def try_finally2():\n", - " try:\n", - " print(\"do some stuff in try block\")\n", - " return \"return from try block\"\n", - " finally:\n", - " print(\"do some stuff in finally block\")\n", - " return \"always execute finally\"\n", - " \n", - "print(try_finally2())" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "do some stuff in try block\n", - "do some stuff in finally block\n", - "always execute finally\n" - ] - } - ], - "prompt_number": 21 - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "Here, the abortive `return` statement in the `finally` block simply overrules the `return` in the `try` block, since **`finally` is guaranteed to always be executed.** So, be careful using abortive statements in `finally` blocks!" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "
\n", - "
\n", - "" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "#Assigning types to variables as values" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "I am not yet sure in which context this can be useful, but it is a nice fun fact to know that we can assign types as values to variables." - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "a_var = str\n", - "a_var(123)" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "metadata": {}, - "output_type": "pyout", - "prompt_number": 1, - "text": [ - "'123'" - ] - } - ], - "prompt_number": 1 - }, - { - "cell_type": "code", - "collapsed": false, - "input": [ - "from random import choice\n", - "\n", - "a, b, c = float, int, str\n", - "for i in range(5):\n", - " j = choice([a,b,c])(i)\n", - " print(j, type(j))" - ], - "language": "python", - "metadata": {}, - "outputs": [ - { - "output_type": "stream", - "stream": "stdout", - "text": [ - "0
\n", - "
\n", - "
\n", - "
\n" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "# Changelog" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "[[back to top](#sections)]" - ] - }, - { - "cell_type": "markdown", - "metadata": {}, - "source": [ - "#### 04/27/2014\n", - "- minor fixes of typos\n", - "- single- vs. double-underscore clarification in the private class section." - ] - }, - { - "cell_type": "code", - "collapsed": false, - "input": [], - "language": "python", - "metadata": {}, - "outputs": [], - "prompt_number": 1 - }, - { - "cell_type": "code", - "collapsed": false, - "input": [], - "language": "python", - "metadata": {}, - "outputs": [] - } - ], - "metadata": {} - } - ] -} \ No newline at end of file diff --git a/numpy_matrix.py b/numpy_matrix.py deleted file mode 100644 index 06d5eb2..0000000 --- a/numpy_matrix.py +++ /dev/null @@ -1,36 +0,0 @@ -# numpy matrix operations -# sr 12/01/2013 - -import numpy - -ary1 = numpy.array([1,2,3,4,5]) # must be same type -ary2 = numpy.zeros((3,4)) # 3x4 matrix consisiting of 0s -ary3 = numpy.ones((3,4)) # 3x4 matrix consisiting of 1s -ary4 = numpy.identity(3) # 3x3 identity matrix -ary5 = ary1.copy() # make a copy of ary1 - -item1 = ary3[0, 0] # item in row1, column1 - -ary2.shape # tuple of dimensions. Here: (3,4) -ary2.size # number of elements. Here: 12 - - -ary2_t = ary2.transpose() # transposes matrix - -ary2.ravel() # makes an array linear (1-dimensional) - # by concatenating rows -ary2.reshape(2,6) # reshapes array (must have same dimensions) - -ary3[0:2, 0:3] # submatrix of first 2 rows and first 3 columns - -ary3 = ary3[[2,0,1]] # re-arrange rows - - -# element-wise operations - -ary1 + ary1 -ary1 * ary1 -numpy.dot(ary1, ary1) # matrix/vector (dot) product - -numpy.sum(ary1) # sums up all elements in the array -numpy.mean(ary1) # average of all elements in the array diff --git a/os_shutil_fileops b/os_shutil_fileops deleted file mode 100644 index d517f22..0000000 --- a/os_shutil_fileops +++ /dev/null @@ -1,22 +0,0 @@ -# sr 11/19/2013 -# common file operations in os and shutil modules - -import shutil -import os - -# Getting files of particular type from directory -files = [f for f in os.listdir(s_pdb_dir) if f.endswith(".txt")] - -# Copy and move -shutil.copyfile("/path/to/file", "/path/to/new/file") -shutil.copy("/path/to/file", "/path/to/directory") -shutil.move("/path/to/file","/path/to/directory") - -# Check if file or directory exists -os.path.exists("file or directory") -os.path.isfile("file") -os.path.isdir("directory") - -# Working directory and absolute path to files -os.getcwd() -os.path.abspath("file") diff --git a/other/happy_mothers_day.ipynb b/other/happy_mothers_day.ipynb new file mode 100644 index 0000000..de8f0db --- /dev/null +++ b/other/happy_mothers_day.ipynb @@ -0,0 +1,128 @@ +{ + "metadata": { + "name": "", + "signature": "sha256:c47e61a194d5b7fe9015e4a6494048a41f535f22a9a874def78f1a7fc13b1243" + }, + "nbformat": 3, + "nbformat_minor": 0, + "worksheets": [ + { + "cells": [ + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[Sebastian Raschka](http://sebastianraschka.com) \n", + "last updated: 05/11/2014\n", + "\n", + "- Link to [this IPython Notebook](https://github.com/rasbt/python_reference/blob/master/funstuff/happy_mothers_day.ipynb) on GitHub\n", + "- Link to the [GitHub repository](https://github.com/rasbt/python_reference)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "# Happy Mother's Day!" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "import numpy as np" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 1 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "x = np.arange(-1,1.001,0.001)" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 2 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "y1 = (x**2)**(1.0/3) + np.sqrt(1-x**2)\n", + "y2 = (x**2)**(1.0/3) - np.sqrt(1-x**2)" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 4 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%matplotlib inline" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 5 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "import matplotlib.pyplot as plt\n", + "\n", + "plt.rcParams.update({'font.size': 12})\n", + "fig = plt.figure(figsize=(8,8))\n", + "plt.plot(x, y1, lw=15, color='r')\n", + "plt.plot(x, y2, lw=15, color='r')\n", + "plt.xlim(-1.2, 1.2)\n", + "plt.title(\"Happy Mother's Day\")\n", + "\n", + "ftext = 'x = np.arange(-1,1.001,0.001)'\\\n", + " '\\ny1 = (x**2)**(1.0/3) + np.sqrt(1-x**2) '\\\n", + " '\\ny2 = (x**2)**(1.0/3) - np.sqrt(1-x**2)'\\\n", + " '\\n\\n\\n[inspired by a MATLAB impl. by TU Delft]'\n", + "plt.figtext(.32,.45, ftext, fontsize=11, ha='left')\n", + "\n", + "plt.show()" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "display_data", + "png": "iVBORw0KGgoAAAANSUhEUgAAAesAAAHtCAYAAAA5qla8AAAABHNCSVQICAgIfAhkiAAAAAlwSFlz\nAAALEgAACxIB0t1+/AAAIABJREFUeJzs3XdYFFf3B/DvAmKhWFFQpAjYaKuigImKxqgx8afGHhv2\nGvVNNLFE0ajJa4lJXktMbJhYorEkJtFYohixYEMU7GIFsaCooPTz++PKhqUssDPb4HyeZx/dnZk7\nZwt7dmbuPVdBRATGGGOMGS0zQwfAGGOMMc04WTPGGGNGjpM1Y4wxZuQ4WTPGGGNGjpM1Y4wxZuQ4\nWTPGGGNGjpM1Y6xYQkNDUa5cOUOHwViZxMmalUnBwcF4++23C1xmZmaGTZs26Tki7ZmZmcHMzAy/\n//57vmXdu3eHmZkZRowYUaI2LSws8OOPP8oVYrGEhobC1dVV6+1v3bqlei3MzMxgbW2N+vXrY9Cg\nQTh+/LiMkTKmf5ysWZmkUCigUCgMHYZsnJ2dsXr1arXH4uPjsXv3bjg5OZX4uSoUCuirXhIRITMz\nU7b2du3ahYSEBFy8eBHfffcdiAhvvvkmvv76a9n2wZi+cbJmZVJJEtG3336LJk2awMbGBg4ODujX\nrx8SEhJUy8PCwmBmZoY//vgDLVq0QMWKFeHt7Y1Dhw4Vex0iQr169fDll1+q7TslJQW2trbYuHGj\nxhiHDBmCv/76C/Hx8arH1q5di1atWsHV1VXt+WZkZGDq1KlwdHRE+fLl4enpic2bN6uWu7i4ICsr\nC0OGDIGZmRnMzc3V9nXs2DE0bdoUVlZW8PPzw+nTp9WWX79+HT169EDVqlVRrVo1dOzYEdHR0arl\nOafTw8LC0KRJE1SoUAEHDhzI95zu3buHHj16wM7ODhUrVoSbmxsWL16s8XUAgGrVqqFmzZpwcnLC\nW2+9hZ9++gmTJ0/Gp59+itjYWNV6I0aMgLu7OypVqgQ3NzfMmDED6enpAIDY2FiYmZnlOyL/559/\nYGFhgbt37xYZB2Ny4mTNyqziJmyFQoGvvvoK0dHR2LlzJ+7cuYO+ffvmW++jjz7C7Nmzce7cOfj7\n+6NLly5qSV3TOgqFAiNHjsSaNWvU1v/5559haWmJXr16aYzRzc0NrVu3xrp16wAA2dnZWLt2LUaO\nHKl6DjmmT5+O1atX49tvv0VMTAwGDBiAAQMG4ODBgwCA06dPw9zcHN9++y0SEhJw//591bbZ2dmY\nPn06li5dirNnz6JmzZro3bs3srKyAAAPHjzAm2++CXt7e4SHhyMiIgINGjRAUFAQHj9+rNbO1KlT\n8c033+DKlSvw8/PLd7Zj7NixePHiBf7++29cuXIFa9asQd26dTW+DoX55JNPkJWVhZ07dwIQ732t\nWrWwefNmXL58Gd988w3WrVuHL774AgBQr149dOjQAatWrVJrZ9WqVejYsaPWcTCmNWKsDBo8eDBZ\nWFiQtbV1vptCoaCNGzcWuu3Zs2dJoVBQfHw8EREdOnSIFAoFrV27VrVOZmYmOTs708yZM4u9TkJC\nAllaWtKBAwdU6wQEBNCkSZM0PpeceLdu3Uqurq5ERLRnzx6qWbMmpaenU5s2bWjEiBFERJSSkkLl\ny5en7777Tq2N7t27U7t27VT3LSwsaP369WrrrFu3jhQKBUVGRqoei4iIIIVCQVevXiUiopCQEAoI\nCFDbLjs7m9zc3Oibb75Rayc8PFzj8/L19aXZs2drXCe3mzdvkkKhoKNHjxa43N7ensaNG1fo9kuW\nLCEPDw/V/R07dpCVlRU9f/6ciIiePn1KlSpVol9//bXYMTEmFz6yZmVWQEAAoqKi1G7nzp3Lt15Y\nWBg6duwIJycn2NraolWrVgCA27dvq60XGBio+r+5uTlatGiBmJiYYq9Tq1YtdO3aVXU0Fx0djYiI\niGJ3DuvWrRtSUlKwf/9+/PDDDxg0aFC+3tvXr19Heno6WrdurfZ469at88VaEIVCAV9fX9V9BwcH\nAOKIGgBOnTqFM2fOwMbGRnWztbXF7du3cf36dbW2mjdvrnFfkyZNwhdffIGAgABMnToVR44cKTI+\nTYhI7ch91apV8Pf3h729PWxsbDB9+nTcuXNHtbxLly6oXLmy6hLEhg0bUKVKFXTp0kVSHIxpw8LQ\nATBmKBUqVEC9evU0rnPnzh107twZgwcPxuzZs1GjRg3cvXsX7du3V13fLEze5FCcdUaPHo3OnTsj\nMTERq1evRsuWLdG4ceNiPZ9y5cohODgY8+bNw4kTJ3DhwgUAkLUjnZmZmVp7Of/Pzs4GIJ5P+/bt\nsWzZsnzbVq5cWfV/c3NzWFpaatxXcHAwOnXqhL/++guHDh3CO++8g+7du+Onn34qcdyPHj3Co0eP\nVO/3L7/8gvHjx2PBggVo06YNbG1tsXXrVsyYMUO1jYWFBYYNG4ZVq1Zh9OjRWL16teo6PmP6xp86\nVmYVJ4mdOnUKqamp+OabbxAYGAgPD49816Fz5O6MlJmZiZMnT+ZLtEWt07ZtWzg5OWHlypXYsGFD\niYdcjRw5EuHh4QgMDET9+vXzLXd3d0f58uVx+PBhtccPHz4Mb29v1X1LS0vVdeiS8PPzQ3R0NOrU\nqYN69eqp3apXr17i9uzt7REcHIz169dj9erV2LhxI5KTk0vczqJFi2BhYYHu3bsDEB3FmjRpgkmT\nJqFJkyZwc3PDzZs3830mhg8fjqioKKxcuRIXLlzA8OHDS7xvxuTAR9aszKJidDDz8PCAQqHA4sWL\n8cEHHyAqKgpz584tcN0FCxbA3t4eLi4uWLJkCRITEzF27NgSrZPT0WzGjBmwsrJCnz59SvSc3Nzc\nkJiYiPLly6s9z5znWqlSJUyYMAEzZ86EnZ0dfHx8sG3bNuzatUutR7arqysOHjyITp06oVy5cqhR\no0ax9j9+/HisWbMGXbt2xWeffQZHR0fcu3cPe/bswXvvvad2GaA4bb377ruoX78+UlNTsWPHDjg5\nOcHa2lrjdomJiUhISEBaWhquX7+O0NBQbNq0CUuWLIGLiwsAoGHDhli7di127doFT09P/PHHH9i5\nc2e+z4STkxM6deqESZMmoX379qrtGdM7w10uZ8xwgoOD6e233y5wWd4OZsuXL6e6detSxYoVqVWr\nVvTXX3+RmZkZHT58mIj+7Tz2+++/U7Nmzah8+fLk6emp1lGsOOvkePz4MVlaWtL48eOL9VyK6hAX\nFBSk6mBGRJSRkUFTp06lOnXqkKWlJXl6etLmzZvVtvnrr7+oUaNGZGlpSWZmZkQkOoaVK1dObb27\nd++qvRZERLdv36b+/fuTnZ0dlS9fnpydnWngwIF069atQtspyLhx46h+/fpUsWJFql69Or333nt0\n8eLFQtfP6WCWc6tUqRK5u7vToEGD6Pjx42rrZmRk0KhRo6hatWpka2tL/fv3p2XLlqmea26//vor\nKRQK2rZtW5ExM6YrsiTrtLQ0Gjp0KDk7O5ONjQ0plUras2dPoesvWbKE7O3tydbWloYOHUppaWly\nhMGYQeQk4ri4OEnr5IiOjiaFQkHnz5+XM0ympeXLl5O9vT1lZGQYOhRWhslyzTozMxNOTk74559/\n8Pz5c8ybNw+9e/fO11sWAPbu3YsFCxbg4MGDuH37NmJjYxESEiJHGIyZtPT0dMTFxWHatGlo166d\n2jVkpn8pKSm4fPkyFi5ciHHjxsHCgq8aMsORJVlXqlQJISEhcHJyAgC8++67cHV1xdmzZ/Otu379\negwfPhyNGjVClSpVMGvWLISGhsoRBmMGU5zOakWts2nTJjg5OeH27dv47rvv5AqNaWncuHHw9fWF\nt7c3pkyZYuhwWBmnIJK/APCDBw/g4uKCqKiofD1SlUolZsyYoarIlJiYCDs7OyQmJqJq1apyh8IY\nY4yZPNnP62RkZKB///4IDg4ucOhIcnKy2nhLW1tbAMCLFy/UkrVSqURUVJTc4THGGGNGydfXt8DC\nTIDM46yzs7MxcOBAVKhQocCiCABgbW2N58+fq+4/e/YMAGBjY6O2XlRUlGrIibHcQkJCDB5DWbvx\na86veVm48WvOrzkRaTxAlS1ZExGGDRuGR48eYfv27flm6snh6emp9sshKioKtWrV4lPgjDHGWCFk\nS9ZjxozB5cuXsWvXLrWCDHkNGjQIa9aswaVLl/D06VPMnTsXQ4YMkSsMxhhjrNSRJVnfvn0bP/zw\nA6KiolRF8W1sbLB582bcuXMHNjY2uHfvHgCgY8eO+OSTT9C2bVu4uLjAzc0Nc+bMkSMMnQsKCjJ0\nCGUOv+b6x6+5/vFrrn+m9prrpDe4HBQKBYw0NMYYY0x2mvIeT+TBGGOMGTlO1owxxpiR42TNGGOM\nGTlO1owxxpiR42TNGGOMGTlO1owxxpiR42TNGGOMGTlO1owxxpiR42TNGGOMGTlO1owxxpiR42TN\nGGOMGTlO1owxxpiR42TNGGOMGTlO1owxxpiR42TNGGOMGTlO1owxxpiR42TNGGOMGTlO1owxxpiR\n42TNGGOMGTlO1owxxpiR42TNGGOMGTlO1owxxpiR42TNGGOMGTlO1owxxpiR42TNGGOMGTlO1owx\nxpiR42TNGGOMGTlO1owxxpiR42TNGGOMGTlO1owxxpiR42TNGGOMGTlO1owxxpiR42TNGGOMGTlO\n1owxxpiR42TNGGOMGTlO1owxxpiR42TNGGOMGTlO1owxxpiR42TNGGOMGTlO1owxxpiR42TNGGOM\nGTlO1owxxpiR42TNGGOMGTlO1owxxpiR42TNGGOMGTlO1owxxpiR42TNGGOMGTlO1owxxpiR42TN\nGGOMGTlO1owxxpiR42TNGGOMGTlO1owxxpiR42TNGGOMGTlO1owxxpiR42TNGGOMGTlO1owxxpiR\n42TNGGOMGTlO1owxxpiR42TNGGOMGTkLQwfATEBGBvDgAfDkCfD8+b+3Fy+ArCwgO1vciACFAqhY\nEbCyAipVEv/a2AA1a4pbpUqGfjaMmb7sbCAxEUhIEH+LKSlAcrK4paSI5TkUCnGrVEn8Ldrain8r\nVwbs7cW/CoXhngsrFk7WTEhMBC5dAi5fFv/GxgL37gFxceILgUie/Vhbi6Rtbw+4ugL16gFubv/+\n6+DAXxyMpacDt24B168DN26If2Njgfh48ff44IH4oSyHSpWA2rWBOnUAJyegYcN/b+7ugKWlPPth\nkiiI5PoWlpdCoYCRhmb6kpKAU6eAiAhxO3VK/PEbg6pVAR+ff2++voC3N1ChgqEjY0w3HjwAzp0D\noqL+/ffKFfmSsRTm5oCnJ9Cixb83T0/Ago/zdEFT3uNkXRakpgLh4cD+/cC+feILwZSUKwc0bQq0\nbPnvrXZtQ0fFWMllZgLnzwNHj4rbsWPA3buGjqpkbGyAtm2B9u2Bt98GGjTgs2Ey4WRdFj19Cvz2\nG7BtG/D33yJhlyaurv9+Wbz1FlCtmqEjYiw/IiA6WvxI3rdPJOfkZENHJS8nJ6B7d6BnT/FD2oz7\nLWtLL8l62bJlCA0NRXR0NPr164d169YVuF5oaCiGDRuGSrk6Gv35559o3bp1sYNmhXj5Eti+Hdi8\nGThwQHQMKwsUCsDPD+jQAfi//xP/5y8MZihJScBffwF794oEHR9v6Ij0x8EB6NEDCA4GmjUzdDQm\nRy/JeufOnTAzM8PevXvx6tUrjcl67dq1+OeffzS2x8m6BKKigFWrgA0bgGfPDB2N4dWpA3TrJm5t\n2ojT6Izp0sOH4kzW9u3iTFZmpqEjMrwmTYARI4APPhA9zlmRNOU92Q4/unfvjq5du6J69epFrstJ\nWAbZ2eLL4Y03AKUSWL6cE3WOuDjxerz9NlCrFjB0KHDwoPpwFsakevxYfM6CgsQR5ciR4miaE7UQ\nGQmMHStem3HjRG92pjXZzxUWlYgVCgUiIyNhZ2eHBg0aYN68ecgyhl6PpiItDVi7VvTI7NZNXANj\nhXv6FFi3TlzXdnYGPv1UXENkTBtpacCOHeJvz8EBGD8eOHyYfwhq8uoVsGIF4OEB9OsnkjgrMdmT\ntaKIXoGtW7dGTEwMHj16hO3bt2Pz5s1YtGiR3GGUPllZwI8/ip6Xw4aJ8dCsZO7dAxYuFEPBlErx\nBfL8uaGjYqbg3Ll/jxJ79BBntfgIumSys4GffxYjO3r0EMPTWLHJ3hv8s88+Q1xcXKHXrPPasmUL\nFi1ahNOnT6sHplAgJCREdT8oKAhBQUFyhmoaiIA9e4CpU4ELFwwdTeljZQX07w+MHi2usTGWIzVV\njKZYsQI4ftzQ0ZQ+5ubiEtXs2WV2KGZYWBjCwsJU9+fMmaO/oVszZ87EvXv3SpSsFy5ciDNnzqgH\nxh3MxDWe8eNFsma65+8vXu8+fbhTWll26xawciWwZo24Ls10y8oKmDMHmDChzP/d6aWDWVZWFlJT\nU5GZmYmsrCykpaUVeC16z549ePC6Wtbly5cxb948dOvWTa4wSof0dOCLL8R1aU7U+hMRAQwcKEqf\nfvUVnyIvayIjRc9ld3dgwQJO1PqSkgJMniyGXPIZjMKRTEJCQkihUKjd5syZQ7dv3yZra2u6e/cu\nERFNnjyZatWqRVZWVlSvXj0KCQmhzMzMfO3JGJppiYwk8vIiEifA+WbIm60t0eTJRK8/u6wUys4m\n2r+f6O23Df9545u4TZhA9PKloT8ZBqEp73EFM2ORlSWO5j77rOwUMzEV5cqJTn3TpolqTcz0EQG7\ndgGffw6cPWvoaFhejRuLuhFlrB8Jlxs1dvHx4vTb4cOGjqRwlpaiE0iNGqLAga2tuFWoIKqFmZmJ\nSmKZmaKS2suX4vTWixfAo0disoKkJEM/C2nKlQOGDxdJu25dQ0fDtEEE7N4NzJpl+knawkLUEahV\nS/xNWluLm5XVvzNl5XyHZmSIMqcvXojLO0lJwP37xn2qv1w54MsvgY8+KjO1xzlZG7OjR0VN3YQE\nQ0ci/ji8vMSv2kaNxBR59eoBjo4iSUv9g0lPF5We7twRnedu3BD/XrsmerqbSs1kS0uRtGfMKLO9\nWE0OkZjIZtYs0TfBVLi4iOGa7u5iCll3d/GYg4Oohy+1rG5amvjuuXNHDKW6fFnczp83nglGevcW\nnf2srQ0dic5xsjZGRKLH6cSJhjvtbWcnioUEBoqe0EolUL68YWLJzha9cM+fF7fTp0VnE2P+5V+p\nEvDxx8CUKWImImacTp8W71MRJY4NSqEQP5SbNxd/h0qlmCLWkGU6ExKAkyfFj5vDh4ETJww3baeX\nF7Bzp/ixUopxsjY2WVlimMKKFfrfd2CgmOyiY0cxV7QxT3hBBFy/Lqq0HT0qSobeuGHoqPKrVUsM\nPRk2jOf5NSZ37gDTpwMbNxo6kvwsLcUMVW++KUoGBwQAVaoYOirNnj8HwsLECJWdO8WlLX2qUkX0\nM2jVSr/71SNO1sYkNVUU4dixQ3/7bN5cXBPv0cP0r7XGxorTmfv3iwkTjOk6eMOGwOLFwLvvGjqS\nsu3FC+C//wWWLDGuqWEbNhQ/kjt2FBPM5Jp50ORkZYkf0Fu3Aps2ibK++lC+vJhVsHt3/exPzzhZ\nG4ukJKBrV/2cjqtcGRgwQMx64+ur+/0ZQkaGeC1//VXc7t0zdERCly7At9+KObeZ/hCJxPHxx/o/\n6iuIQiGOAt9/X9QSd3Y2dES6kZoqDj5WrRJH3rpmZiYmUBk9Wvf70jNO1sYgKQlo3x7IU6lNdo6O\novfk8OFl6zoqkbg2uX27+MI2dOeYChVEr/FPPhH/Z7p16ZKo3a2PZKGJmZnoB9Kzp/hhXquWYePR\nt/PnRf39n3/W/fXtr78GJk3S7T70jJO1oT17BnToIDpr6IqTk6ix27//v8M2yqrsbHHEvWED8Msv\nhq1E5uYGLF0KvPOO4WIozVJSgHnzxOUHQ06s4eUFDB4sLjfxCAHg9m1RhXHNGt0m7aVLRYngUoKT\ntSG9eCGuUemqjF61aqKQypgxfARXkFevxAxJK1cadhx7//7AN9+IIXBMHnv3ijmk79wxzP4rVwYG\nDQKGDBG9t8vIWOASuXJFfD9t26a7fXz3Xak5Jc7J2lAyMkRno/37ddP+yJGiaEC1arppv7S5dAn4\n/nsgNFSc7dA3Oztxra1nT/5ilyIpSVyXXrvWMPtXKoFx48TczFZWhonB1Bw+LA4oLl3STfu//CL+\nrkwcJ2tDIBJDeYo5+1iJeHqKzhyBgfK3XRa8fCmG83z1lWHm1O3eXSRtBwf979vU7d4tfqTGxel3\nv+bmYja2Dz8UNQn4x1bJpaeLyxVz5oj/y6l8eTG0s2VLedvVM07WhjB3rqiWJLePPgLmz+dT3nLI\nzgb+/FN8gei7YEbVqsAPP5SKowG9ePZMFBBav16/+61USYyo+M9/Sm9vbn07f16MVLlwQd52q1cX\nlxs9PORtV484Wevbjh1iTLOcatYUvZzfekvedpkQESHG5v76q373O2SIGOZVlnrul9SxY+Ka/61b\n+ttnjRqicNHYsSIJMHmlpgJTp4rPvpwaNABOnTLZvydO1vp09aqYl/XFC/na9PcXQ5Lq1JGvTVaw\nyEhxmu633/S3Tzc3cVre319/+zQFmZmiR/Hnn+uvzGW1aqJ87PjxZaIWtcFt2iSGmb56JV+bvXoB\nW7aY5KUKTtb6kpIiygZGR8vX5tChoiypoWp2l1Vnz4qkvWuXfvZnbg6EhIjymObm+tmnMbtzRxxN\nh4frZ3+VK4tOaxMnitnkmP5ERYkSyHL26jfRMdicrPVl2DB5e6jOni2ue5vgL8RS48QJYPJkUVpR\nH95+Wxxl29npZ3/G6LffxJhlffTYr1BBXI/+5BPjr81dmsXHA506yXcd28JC/M22aCFPe3rCyVof\nfv9d/DqUg0Ihxg6OGiVPe0waInEt+9NPxXSeuuboKIaiBATofl/GJDMTmDlT9B3QhwEDxGl2U6+X\nX1okJYmyrHLVQ2jQQJwhM6Ea7JrynhFPuWRCHj8WPUbloFCI4V6cqI2HQiGGW8XEAMuW6X5c+717\nQOvWojqTKf1gleLhQ1E8SB+JunVr0Qnpp584URuTKlXE0Lw2beRp78oV0YmtlOAjazn06iVfhZ7V\nq8XpdGa8Hj8W15ZXr9Z9Mu3bV5RsNKGjgxKLiBBD2HQ9EUvt2mImrt69+dKSMUtOFuV55eqv8Pff\nQLt28rSlY3waXJfkPP29ZIm4fsZMQ0SEqGSl68lZmjUT13FL42iA9evFWamMDN3tw9xcDMOaPZs7\nj5mK58/FGZCoKOlt1a8vxnabQCddPg2uK69eid6jcpgwgRO1qfH3Fwl7xQrdjus8c0bMSX7qlO72\noW/Z2WJWsuBg3Sbqli3F67dkCSdqU2JrKwoWOTpKb+vqVVGt0MTxkbUUc+aIX+tSdekC7NzJQ3ZM\n2b17op/B7t2620eFCqKueZ8+utuHPqSkAAMHis+8rlhZAQsWiHrUZnxMYrIuXBBllVNSpLVTsaKo\nS27kVej4NLgu3LoFNGokKvFI4e4u5mGuXFmWsJgBEYlhVxMnAk+e6G4/n38uZjIyxeuucXHistHZ\ns7rbR9u2oj9BvXq62wfTny1bRN8NqXr2FKMsjBifBteFkBDpibpiRVGZjBN16aBQiOFAFy+KISi6\nMmuWKIOpr6pecrl8WRwl6SpRW1mJSxIHDnCiLk369JHncuO2beLAyETxkbU2Ll4EvL3FdTcpStE8\nrCwPItGLe+JEMcuXLrz/vjiSN4VJXSIixHSxiYm6ad/fX7wWbm66aZ8ZVnq6qDsQGSmtnbffBvbt\nkycmHeAja7nNmiU9UXfsyGOpSzOFQtQ8jowUvbl1YccO8TlKStJN+3LZu1cMndFFolYogBkzgCNH\nOFGXZpaWYly81B7d+/eLqTRNECfrkoqMFKeupahSRRx1meI1R1Yy9euLWaOmTdPN+/3PP2KIy4MH\n8rcth82bgffe083ZBUdH4NAhYN48oFw5+dtnxsXTU7zXUuli6mI94GRdUgsXSm9jwYLSOWaWFczS\nUpS13L1bN9XPLlwAgoKA+/flb1uKdevEZByZmfK3/d57YgyuXNWumGmYNAlQKqW1cfSomPfaxPA1\n65K4c0d0XJHSscfPT0wOwcO0yqY7d0TFu5Mn5W/bw0Oc4pNjbKpUq1fLV4I3N4UCmDtXnKngIVll\n0/HjYvy8FN27i8tIRoavWcvl22+lJWqFAli+nBN1WebkJE5djxsnf9vXrokjzdu35W+7JL7/XjeJ\nunp14K+/xDVqTtRlV2AgMGSItDZ+/VUUSzEh/IkvrufPgVWrpLXxwQcmN2Ub04Hy5cWEIHJ0mMkr\nNtawCXvFCt2McGjWTFQi69BB/raZ6Zk7V9ooCCJx8GVCOFkX1+bNwIsX2m9vYSFPtTNWegwYIDpI\n1awpb7u3b4shKvrudLZ2rW7OGPTqJc5GGHn1KaZHdeoA48dLa2PjRt0Nq9QBTtbFtXattO2HDhXV\nyhjLLTBQXL/28ZG33WvXxFHo06fytluYHTt0c+p75kzg559L96xjTDtTp0qr9/7smfSRPXrEybo4\noqOldQgyNxfX2RgriLOz6KHatau87Z4/LwqRJCfL225eBw4A/fpJrz2Qm6UlsGGDKK3K16dZQapX\nF7XfpZB6aVOP+K+gOKQeVffuLToWMVYYa2vxK1/uQjnHj4uer2lp8rab48QJUVo1PV2+NqtUET8A\n+veXr01WOk2YIG2M/ZEj4iyUCeBkXZTsbHG9WoqPP5YnFla6mZuLErRy9204cECcopZ7KOT162K8\ns9QZkXKrXVtcn27VSr42WelVu7bo+yHF1q3yxKJjnKyLcuwYkJCg/fatWumu3CQrfRQKMUnMypXy\nnv796SfRg1YuT57IX+u7fn1xOcDbW742WekndZKPbdvkiUPHOFkXRerAeV10umGl36hR4he/nGU0\nQ0LEdWCp0tPFJCJyjlNt1gwIDwdcXORrk5UNvr5A8+bab3/unDhLZOQ4WWtCJC1Z29oCPXrIFw8r\nW3r0ENexLS3la3PoUODwYe23JxI/QKW0kZe/vzhVb2cnX5usbJF6UGTk81wDnKw1i4yUVlyiXz8e\ncsKk6dJokezzAAAgAElEQVRFVFuSq3hKRoYYt3z3rnbbL1kC/PijPLEAYujavn2iUxlj2urbF6hY\nUfvtd++WLxYd4WStyf790rbn3qxMDu+8A/zxh7Qvo9wePQJ69ix5D/HDh4FPP5UnBgB44w0xfaaU\nsbKMAYCNjehDoa3jx0WVSiPGyVqTAwe037ZWLenF5hnL0b69OMKW6xr2yZNiBqPiiosTQxCl1MbP\nLTAQ2LNHfMkyJoeePbXfNivL6Oe55mRdmFevxBg8bXXrxhN2MHl16CCGEcrVS3zlSiA0tOj10tNF\non74UJ79+vgAf/7JiZrJ6913pdUL37dPvlh0gJN1YY4dk1ZI4v335YuFsRw9egBr1sjX3tixwOXL\nmteZMUP8PcjB3V2c+q5aVZ72GMthbS3OQGnr77/li0UHOFkXRspRdYUKQOvW8sXCWG7BwcA338jT\n1qtXom9FYRXIDh4EvvpKnn3VqSP6gdjby9MeY3l17Kj9tlevyls3QGacrAsTEaH9tq1aSTsdw1hR\nJk4EPvpInrbOngVmzcr/+NOnwODB8lQ+s7YWPW55HDXTJalTqJ44IU8cOsDJuiBE0ibukHIqhrHi\nWrhQ9I2Qq61Dh/69TyQmSbh3T3rbZmaiwIvcM4sxlpeHh7QfhMePyxaK3DhZFyQ2VpRT1Fa7dvLF\nwlhhzM1FRTI/P+ltEQHDhv1b53vbNmDLFuntAsDSpWL4GWO6plBIqyvPydrESDmqrlBBlL9jTB+s\nrIBdu8SEBlLdvAnMmQMkJYnZjOQwYYLoxMaYvgQEaL9tVJT8E97IhJN1QS5c0H7bpk3lrefMWFEc\nHES5RDk+d0uWiE46UiavydGqFbB4sfR2GCuJwEDtt01MlG+Iosw4WRfk0iXtt23RQr44GCuuli2B\nr7+W3k5WlrQzSzns7cVpdP7hyvTN21tamefoaPlikREn64JISdZyXD9kTBtjxwIDBxo6CnEtfetW\nccTPmL5ZWABeXtpvz8naRKSnS5suzdNTvlgYKwmFAlixQvSINaR586R18mFMKinJOiZGvjhkxMk6\nr+vXta9/rFAADRrIGw9jJWFtDWzcKI4uDKF1a2DKFMPsm7EcUpL1rVuyhSEnTtZ5SXmjXFzkmxmJ\nMW01bw58/rn+92trK6bP5Jr4zNCknOHkZG0ipBSBaNhQvjgYk+KTT6QNYdHG0qWAs7N+98lYQdzc\ntN/29m0gO1u+WGTCyTqvuDjtt3V1lS8OxqQwNwdWrdJfb+wOHYyjcxtjAFC3rrgsqY30dOD+fXnj\nkQEn67ykHFk7OsoXB2NSeXkBn36q+/1UqCA6tmn75ciY3CwtxcQx2rp7V75YZMLJOi8pyVrKh4Mx\nXZgxQ9opweKYNUv3+2CspKRcknn8WL44ZMLJOi8ppz/4yJoZmwoVgAULdNe+i4t8s38xJicpU7Fy\nsjYBUuYz5SNrZozefx94803dtP3ll0D58rppmzEpatTQfttHj+SLQyacrHNLSZGWrKV8OBjTFYUC\nWLRI/nb9/IA+feRvlzE5SPk+5iNrIxcaCqSlab99lSqyhcKYrAIC5J9nfeZM7lTGjJednfbbcrI2\ncmYSXg5zcy4GwYzbZ5/J15anJ/Dee/K1x5jc+DR4KSY1WTNmzFq3FlO4ymHyZGl/L4zpGh9Z57ds\n2TL4+fmhQoUKGDJkiMZ1v/76azg4OKBy5coYNmwY0tPT5QhBHlK+fPh0IDN2CgUwapT0dmxtgd69\npbfDmC7xkXV+derUwcyZMzF06FCN6+3duxcLFizAwYMHcfv2bcTGxiIkJESOEOTByZqVdv36Se+9\n3bevtPmCGdMH7mCWX/fu3dG1a1dUr15d43rr16/H8OHD0ahRI1SpUgWzZs1CaGioHCEYHp8SZKbA\nxgZ44w1pbXTvLk8sjOmSlGSdlARkZMgXiwxkzTBEpHH5xYsX4evrq7rv4+ODBw8e4OnTp3KGwRjT\n5P33pW0fFCRLGIzplNSzP0Z2dC3rpLeKIk4FJycno3Llyqr7tra2AIAXL16gatWq+dafPXu26v9B\nQUEI4i8JxqTL9YO5xBo1ElXRGDN2UmfOSk2VJw4NwsLCEBYWVqx1ZU3WRR1ZW1tb4/nz56r7z549\nAwDY2NgUuH7uZM0Yk4m3N2BhAWRmlnxbuXqTM6ZrL15ov225cqKUro7lPQidM2dOoevKehq8qCNr\nT09PnDt3TnU/KioKtWrVKvCo2iCK+LGhs20Z06fKlYG2bbXblnuBM1Mh5fJq1apG12lYlmSdlZWF\n1NRUZGZmIisrC2lpacjKysq33qBBg7BmzRpcunQJT58+xdy5c4sc6qVXUk6bcLJmpkSbpFu1KtCx\no/yxMKYLSUnab2uE1ShlSdZz585FpUqVsGDBAmzYsAEVK1bE/PnzcefOHdjY2ODe62knO3bsiE8+\n+QRt27aFi4sL3NzcNB726x0fWbOyYuhQ4J13SrbNmjU8aQczHVKPrI2Mgoq60GwgCoWiyGvgsvvu\nO2DsWO22LVcOMKYCL4wV5ckTUTP82rWi1503T8yNzZip2LED6NFDu207dQL27JE3nmLQlPd4cHBu\nBZy6LzapPQ8Z07dq1YDISODTT0WHs4LUrw/88Qcwfbp+Y2NMqlJ2ZC1rb3CT17y59ttmZYkZu/g0\nITMlVlbAf/8rypAeOQKcOSOmiW3UCPDyEqfKLS0NHSVjJVfKrllzss7N31/UPc41vKxEnj4F7O3l\njYkxfXB1FbdBgwwdCWPyKGVH1nwaPC8pb5KRVbxhjLEyKzFR+205WZuAatW03zYhQb44GGOMae/+\nfe23lZIHdISTdV5STmNL+XAwxhiTj5TvYyO8nMnJOi8HB+235WTNGGPGQcr3sZQ8oCOcrPOS8ibx\naXDGGDM8Imnfx5ysTYCU0x/x8fLFwRhjTDtPnmg/H7WZGWBnJ288MuBknZeUX1S3bskWBmOMMS3d\nvKn9tjVrAubm8sUiE07WedWpo/22N27IFwdjjDHtSPkurl1bvjhkxMk6r3r1tN/28WPtC6owxhiT\nh5Rk7eYmXxwy4mSdl50dYG2t/fZ8dM0YY4Z1/br223KyNhEKhbQ3i5M1Y4wZFh9ZlxFS3qxLl+SL\ngzHGWMlJ+R7mZG1CpLxZFy7IFwdjjLGSefAAePRI++05WZuQ+vW135aTNWOMGY6U7+CKFaWNCNIh\nTtYF8fbWfttr14DUVPliYYwxVnxSkrWnp1GOsQY4WRfM01P7bbOy+Lo1Y4wZipRkLeVATcc4WRfE\n2lraeGs+Fc4YY4bBybqMkfKmnT0rXxyMMcaKJz2dk3WZ4+Wl/bYREfLFwRhjrHjOnwfS0rTfXsr3\nvo5xsi6Mj4/22549K+0DwxhjrOSkHCjVqAHUqiVfLDLjZF2YZs203zY9HYiKki8WxhhjRZOSrP38\nRAVLI8XJujD16olfWtriU+GMMaZfUr53/f3li0MHOFkXRqEAWrTQfntO1owxpj9PnwJXr2q/PSdr\nEyblzfvnH4BIvlgYY4wV7sgRadtLOTjTA07WmkhJ1nfvArGx8sXCGGOscIcOab+tuztQvbp8segA\nJ2tNpP7SkvLhYYwxVnxSvm+N/BQ4wMlas6pVgcaNtd8+LEy2UBhjjBXiyRMxxlpbLVvKF4uOcLIu\nSlCQ9tseOsTXrRljTNcOH5b2Xdu2rXyx6Agn66JIeRPj48UsXIwxxnRHyilwe3ugYUP5YtERTtZF\nadNG2vb798sTB2OMsYIdOKD9tkFBRl0MJQcn66LY2UmrF7tnj3yxMMYYU3fzprRpiaVc6tQjTtbF\nIeVU+MGDwKtX8sXCGGPsX1IPiEzgejXAybp4pLyZr15xr3DGGNOVP//UftvatQEPD/li0SFO1sXR\nti1gbq799rt3yxcLY4wx4dUrcfZSW+3bm8T1aoCTdfFUqSJtHN7u3TyEizHG5BYWBqSmar/9u+/K\nFoqucbIuLilvamwscPGifLEwxhgDfv9d+23NzYEOHeSLRcc4WRdX587Stt++XZ44WJkxZcoUbNmy\npcBl586dwxtvvAErKyv06tVLYzuTJ09GvXr1YGZmhosafjRmZWVh3LhxcHd3h4eHB9asWVOsZfv2\n7YOfnx8qVKiAKVOmaIzlxIkT8PX1RYMGDdCxY0c8evSowPVevnyJPn36wMPDA40aNcKfua5Lalq2\nYcMG+Pj4oFy5cli+fLlam71798apU6c0xsdMSHY2sHOn9tu/8YY4a2oqyEgZXWjZ2UR16xKJE9ol\nv/n4GPoZMC1lZmbqfZ8PHjygRo0aFbo8Pj6eIiIi6Pvvv6eePXtqbCs8PJzu3r1LLi4uFBMTU+h6\n69evp44dOxIR0aNHj8jR0ZFu3bpV5LLr16/TuXPn6LPPPqPJkycX2n5WVha5ubnR0aNHiYho3rx5\nNHTo0ALXnTNnDo0cOZKIiK5du0b29vaUkpJS6LLk5GQiIoqOjqaLFy/SoEGDaPny5WptRkREUKdO\nnTS8UsykHDmi/fcxQPTf/xr6GeSjKe/xkXVxKRTSjq7Pny9T1cwuX74MJycn3LlzBwAwZ84c9OvX\nT5a2w8LCoFQqMXr0aPj6+kKpVOLy5cuqZb6+vhg8eDC8vLzg7++PS4WMwZwyZQpatGgBpVKJ9u3b\nq2K9desWatSogSlTpqBZs2ZYvXo1Dh48iJYtW6Jp06bw8fFRO+INCgrCJ598glatWsHNzQ3Tpk1T\nLbt48SL8/f3h7e2NgQMHIjAwUHUkeP/+ffTq1Qv+/v7w8fHBl19+qdrup59+Qrdu3Qp9DRwcHNCi\nRQtYWloW+Xq98cYbcHR0LHK9rVu3YuTIkQCAGjVqoFu3bvjll18AAFu2bCl0mZubG3x9fWFhYaGx\n/TNnzqBixYpo+br/x6hRo7B169ZCYxk1ahQAwN3dHX5+ftj9uqNmQcv2vB6+4+npiUaNGsHMzAyU\np59IixYtcPXqVcTFxRX5WjATsG2btO2lni3VM07WJSG1M0IZOhXesGFDfPHFF+jTpw/27duHzZs3\nY9WqVQWu26tXLzRp0iTfrWnTpkhLSytwm4sXL2LMmDGIiopC7969MW/ePNWyCxcuYPjw4YiOjsa4\nceMwaNCgAtuYOnUqTp48iXPnzqFv37749NNPVcuePHmCFi1a4MyZMxg1ahSaNm2K8PBwnD17Fvv3\n78fkyZPx7NkzAIBCocDdu3dx5MgRREZGYvXq1bhx4wYAYODAgZg4cSIuXLiASZMm4dSpU1C87n06\naNAgTJgwARERETh9+jR2796NA68rMR06dAiBgYElfNWluXPnDpydnVX3nZyccPfuXQDA3bt3C12m\nbfs1atRAdnY2kpKSihXLvXv3ioyzKAEBATgopfcwMw5EwI4d2m9ft660YlcGoPmnMFPXvj1gZQWk\npGi3/bZtwNSp8sZkxAYMGIADBw6ge/fuCA8Ph7W1dYHr5RyhlUSDBg3g6+sLAPD398fvuTqauLu7\no1WrVqoYRo4cieTk5Hz73717N1asWIHk5GRkZmaqLatQoYLateCHDx9iyJAhuH79OiwsLPDkyRNc\nuXIFLV5Po5qzrq2tLRo1aoQbN27Azs4OMTEx+OCDDwAAzZo1g4+PDwAgJSUFYWFhePz4sWofycnJ\nuHz5Mtq3b4+bN2+iTp06JX5d5KYwkWEtxeXo6IhYnmfe9J06BZTwx6Ka7t1NZshWDk7WJVGxoji6\nLuTUXZHOnBGl8Vxd5Y3LSKWnpyMmJgZVq1ZFQkJCoev17NlTdSSa1/Hjx1GhQoV8j+d+zNzcPF+y\nLcrt27fx0Ucf4fTp03B2dsaxY8fQv39/1XIrKyu19ceMGYNu3bph5+sOLQ0aNEBqriEjJY0nOzsb\nZmZmOH36NMyLMYY/ICAAaWlpsLW1xeHDh1WPy5lMnZyccOvWLTRr1gyAeI1cX39WNS0rLmdnZ9y+\nfVt1//HjxzAzM0OVAjr55OyvevXqqv21a9euyGW5FfTaKBSKfKfHmQnS9js4R48e8sShR3wavKR6\n9pS2/c8/yxOHCZgyZQqaN2+Offv2YfTo0YVeK9y2bRsiIyMLvBWUqIty48YNhIeHAwA2bdoEHx+f\nfEfVz58/h6WlJWrVqoXs7GysXLlSY5vPnj1TnXrdv38/rl+/rra8oARga2sLT09PbN68GQBw9uxZ\nXLhwAQBgY2ODVq1aqV2nvnv3Lh48eAAAcHFxUZ32BUQv6sjISLVEXdh+T548ifbt2xf4PDQlql69\nemHVqlUgIjx69Ai//fYber7+vGtapqntnTt3YvDgwQCApk2b4tWrVzh69CgAYOXKlejdu3ehsXz/\n/fcAgGvXruH06dPo1KlTkctyx1JQPPfu3UO9evUKfQ2YCcjKAl7/TWmlVi3RE9zU6Lx7m5aMNrQX\nL4gqVNC+B2KjRqJneSm3c+dOUiqVlJaWRkREa9asoVatWlFWVpbktsPCwqh58+YF3j906BD5+vrS\n4MGDycvLi/z9/enSpUtERBQXF0dKpVK13cSJE8nV1ZWaN29OISEh5OrqSkREN2/eJDs7O7V97t+/\nnzw8PEipVNKIESOoSZMmdPjwYSIiCgoKoj///FO1bu770dHR1KJFC/L29qYPPviAmjRpQuHh4URE\nlJCQQP369SNvb2/y9vamli1b0pUrV4iIaNGiRTRt2rRCX4ObN2+So6MjVatWjSpVqkSOjo60du1a\nIiLaunUrdenSRbXuhx9+SI6OjlSuXDmyt7cnLy8v1bLOnTvTmTNniEj01h4zZgy5ubmRm5sbrVq1\nSrWepmVHjhwhR0dHsrW1JRsbG3J0dKR9+/YREdHChQvpww8/VK177Ngx8vb2Jg8PD+rQoQM9fPhQ\ntUypVNL9+/eJiCglJYV69epF7u7u1KBBA9q1a5dqPU3LNm3aRI6OjmRlZUVVq1YlR0dH1ftPROTm\n5kZ37twp9HVlJuDAAWm9wEePNvQzKJSmvGekGdGIkzURUffu0j4sr78cmfwOHTpEfn5+hg5DJWdI\nERFRTEwM1axZk5KSkorcrqihW5pMmDBBNTzK0Hr06EG3b982dBhEJIZu5Qw/YyYsOFja9+/+/YZ+\nBoXSlPf4NLg2pF7v2LBBnjhYPgqFwqg6RR07dgxKpRK+vr7o168fVq9ejcqVKxe5Xc2aNfHuu+8W\nOrRJk2+//VY1PMrQtm3bBicnJ0OHAQBYvHgxPv/8c0OHwaR4+VLaqJrq1YE2beSLR48Ur7O50THq\njiDPngE1awLp6dptX6sWcO8eUMS4VMYYY7ls2QL07av99sOHA4UMITUGmvIeH1lro3Jl4L33tN/+\nwQPg77/li4cxxsoCqWclBw6UJw4D4GStLalv+vr18sRhgkpS11obqamp8PPzw8uXL4u1/pAhQ/I9\ndvjwYazP9R5dvXoVbdu2RaNGjeDt7Y2hQ4eqhm49fPgQgYGB+X4Rr1+/Pl/v7dz7y73+n3/+iTFj\nxgAA0tLS0KlTJ9jZ2cHOzk5j7A8ePECHDh3QoEEDKJVKnDx5Um15eHg4evXqBSJCYGCg6pR8+/bt\nVT3aC4u/tJg9ezYyMjLUHtP29S5K3s9Njrzv+apVq+Dr6wsfHx/4+vpi48aNqnWXL1+ORYsWSYqj\nVLp/H3hdqU4rTk7Am2/KF4+e8WlwbaWnAw4OwJMn2m1vaQnEx4trKGXM/fv3cffuXZw7dw779+/X\nqiiKJv/73//w9OlThISEaFzvo48+UpWqDAgIwIsXL1TV0AICAvDo0SPcv38fAwYMgIODA5KSkuDr\n6wsiQr9+/eDl5YXPPvsMADB27Fi0adMGffr0wYkTJ/DTTz+hTp06qF69Ok6ePInp06dj69atsLGx\nQUREBDp37ozTp0/jq6++AgD4+flh27ZtcHFxQVZWFg4fPozq1aujffv2hU52AQBDhw6Fu7s7pk+f\njqNHj2LIkCG4evWqavnkyZPRtGlTfPDBB3jx4gVsbGxUr9GBAwewa9eufPFrEhYWhvXr12PdunVF\nvxEGlpWVBXNzc5iZmeHFixdqY+e1fb0LExsbW+DnJjo6GgkJCYiOjkZwcDB++eUXrFq1CkeOHIGv\nry+qVKmCuLg4KJVK1Zj/tLQ0NG7cGDExMVoNXSy1vvgCmDFD++2nTRNtGDGNeU+nXdskMOLQ/jV6\ntLReiUuWGPoZ6MzChQtp3LhxqvsJCQlUq1YtevXqleqxdevWFTkJhTa8vb1VQ6AePHhALi4udPr0\naSIiCg0NpTfffFM1hGzatGmkUCjUhiIlJiaSp6cnOTo6FtqTefHixTR8+HDV/bCwMHr77bdV92/e\nvEm1a9cmb29vevr0qerxlStXkkKhoBkzZqgeO336NAUEBOTbx82bN6lGjRoan6u1tTUlJiaq7nt5\nedGpU6dU9xs1alRg7/PPP/+chg0bVmj8hQkLC6Pg4OAi12vTpg1NmTKF3nzzTapXrx5NnTpVbdmk\nSZOoRYsW5O7uTtOnTy+wjaNHj1LTpk1JqVSSp6cnbd68mYiI7t27R+3ataPGjRtT586d6d1336Vl\ny5YREdHgwYNp2LBh1KpVK1IqlTRu3DhSKBTk4+NDSqWSkpKSJL3eP/30E/n7+1NGRgZlZWXRW2+9\nRd9//z0RFf65+eOPP6hcuXI0cODAQtv19vZW68EfHBxMGzdu1BhLmZKVReTqKu37VsMkNsZCU97j\n0+BSSD0V/sMP4mNUCg0fPhzbt29XnYr+4Ycf0L9//xIfKZS0bvijR48QFxeH+vXrAxC9qkNDQ/HB\nBx/gxIkTCAkJwc8//wwzMzNMmTIFXl5e6N+/P1JTU7Fw4ULcvHkTU6ZMwYQJEzBmzBgsWrQIERER\navt49eoV1q1bh65du6oea9GiBY4fP47MzEycOHECixYtwvjx4zFu3DhMnjwZsbGxWLhwITIyMtC/\nf380btxYNZ1kziQhJZWYmAgiQrVq1VSP5a6THRMTAwcHB7Xe5507d4aDgwM2bdqkVpAld/yaUDE/\nr5rqpSsUCly6dAnHjx/HuXPn8Pvvv6tNc5lj4cKFmDJlCiIjIxEdHY133nkHADBhwgQEBQUhJiYG\ny5Ytwz///KM2AuD8+fPYu3cvIiMjsWzZMgCiEl5kZCQqV66s9esNiPK1Xl5emDp1KubNm4caNWpg\n5MiRiI2NLfBzs27dOkRGRuL9999H//79MWLECGRnZ6u1GRYWhmfPnqmqwwFAy5Yt8Tf3a/nXwYOi\n+qO2mjQBGjeWLx5D0NtPhhIy4tD+lZ1N5OYm7dfeP/8Y+lnozMiRI+m7776jjIwMcnJyouvXr6st\n18WRdUREhFrRjxwhISFkYWFBf/zxR75lBR0phoWFUWhoaL7HMzIy6P/+7/9owoQJ+ZbVrFlT7Ygq\nNDSUwsLCitzf2LFjafHixfnWK+pI7/Hjx2RlZaX2WOfOnWnnzp1ERDR//nz63//+l2+77Oxsmj9/\nfr4xx3njzxEZGUlKpZKUSiW5u7tTtWrVVPfnzp1bYGxBQUG0fft21f1WrVrR3r17Vcs2bdqkWjZ/\n/nz66KOP8rXxzTffkKenJ82bN48iIiJUj1erVo3i4+NV97t166aaDjM4OJgWLFig1o5CoVBNr0mk\n/eud49WrV9SoUSPy8PCgFy9eqC0r7HNT2NmImJgYcnJyyjcu/s8//6SgoKAiYykzeveW9j371VeG\nfgbFoinv8dghKRQKcXQ9e7b2baxaBbyedKK0+fDDD9G/f3/Y2dmhcePGcHNzU1tenPHQ2tQNL0hk\nZCRq1qxZ4OxMBV1/bdOmDdrkGY+ZlZWF/v37o3r16vj222/zbZP3elNOmc3i7E8bObWxExMTVf+/\nc+cO6tatCwDYtWtXgeO0FQoFhg4dqnZkXVD8OZRKJSIjIwGIDlShoaHFeg6a6qXn3g8RFfhZmDhx\nIv7v//4P+/fvx4cffogOHTpg7ty5BW6fW9667lKMHz9eVR5169at8PDwwP3795GSkgIzMzM8e/ZM\nrZRtQZ8boOD3/Nq1a3j33Xfxww8/5DvSN/o+O/r06BHwuia/ViwsgNeT6ZgyPg0uVXCwtNlbtm7V\nvpOakfPy8kL16tXxn//8B+PGjcu3vDhfRiWtG+7i4oL4+Hi1x77++mtkZWXhzJkzWLBgAaKiokr8\nXLKzsxEcHAwLCwusXr063/JXr14hOTkZtWvXLnHbLi4uWs+x3KtXL1Vd8/DwcKSmpqJZs2aIj49H\nWlqaqiDJ48eP1Wb4+uWXX1QzhpUk/pIkkMLWJSJs2LABWVlZSElJwS+//FLgRBxXr16Fq6srRo4c\niQkTJuDUqVMAgHbt2qmS382bN4uc8tLGxkZtGs6SvN7Lli1Tfd48PDyQnp6OPn36YNGiRQgJCUHf\nvn2RlZVVrLZyi42NRceOHbF06VJ07Ngx33KuYZ7Ljz8CeXrzl0iXLoC9vXzxGAgna6mcnYE8kwiU\nSFoaEBoqWzjGZtiwYTA3N8d7ucal37p1C3Xr1sXHH3+M3bt3o27durIdbdasWRMODg6qHtEnT57E\n0qVLsX79etjb22PVqlXo27cvUko4zemePXuwceNGREdHo1mzZmjSpAk+/PBD1fKTJ08iMDAQ5cqV\nK3HMbdu2xYkTJ9Qea968OVq2bImkpCTUrVsXI0eOBADEx8ejSZMmqvX++9//IiwsDPXr18f48ePx\n008/AQB+++03tWvqCQkJ6NSpE3x9feHr65tviFFx4y9JhbjC1lMoFGjYsCFatmwJpVKJ9957D507\ndwYAhISEqCbpWLp0Kby8vNC0aVMsX74c8+fPByAqtB06dAienp748MMPERQUpHG/H3/8Mdq1a4em\nTZvi2bNnJXq98/r000/RtGlT9O7dG8HBwXB1dcXMmTOL9XrkNnXqVDx9+hQzZ85U9cPYt2+favmx\nY8fw1ltvlbjdUicrC1ixQlobI0bIE4uh6focvLaMOLT8duyQdj2lXj2izExDPwudGDZsWIHXB3Xp\n63SZ3IEAACAASURBVK+/ptmzZ+t1n2PGjFH1VtZGkyZN6ObNm7LF06lTJzp79myx15caf0nknfhE\nquDgYFVv8OKS+/WWU2pqKtWrV09t5ESZ9fvv0r5b69Y1qe9WTXmPj6zl8N570k6zxMYCBfSGNWXx\n8fFo2LAhbty4UeApcF0aPXo0fv/992IXRZHq4cOHiIyMLHKMsiZz587FwoULZYtpz549akfgmsgR\nv6GVtB683K+3nFavXo3Ro0fzGGsA+N//pG0/dChQjPniTQEXRZHL9OlAng47JfLWW8CBA/LFwxhj\npuzSJWnDrRQK4NYtUbnMRHBtcH0YPlza9n//DcTEyBMLY4yZutdj5LXWqZNJJeqicLKWS716QPv2\n0tqQesrHRGiqiywHrg3+r7JUG/zZs2cFntpevnw5FixYAACIi4tD27ZtUaVKFTRv3lzS/op6f3PM\nnTsXXl5e8PX1hZ+fn1pHsilTpshebrdUSEqSPn9CIZ0ETZZcF8YTExOpW7duZGVlRc7OzmpFD3Jb\nt24dmZmZkbW1tep2+PDhfOvJGJr+7NwprTNExYpEuUpHllZhYWGqEpz37t2jGjVq0K1bt2Rr/9tv\nvy1WB7P//Oc/tHHjRhowYAAtW7aMvvzyS7px4wYNGTKEvv/+e5o3bx6NGzeOjh8/Trdu3aJz584R\nkSgq0qdPH7WCIGPGjKGff/6ZiIiOHz9OY8eOpfnz59PKlStp6NChdP36dfriiy9o6dKlNGDAANq0\naZNaIZBmzZqpOjxlZmbS33//TefOnSuySMeQIUNo/vz5REQUHh5OHh4eass//vhjVdnK58+fq71G\nXbp0KTB+U5SZmVlgUZO0tDRyc3Oj5ORkIiJ69uwZhYeH059//kl+fn5a7auk7+/evXtVncWioqKo\nSpUqlJqaSkRE8fHxBRbxKfO++krad6mzs0l1LMuhKe/JlhH79u1Lffv2pZSUFAoPD6fKlStTTAG1\nWNetW0etWrUqsj2TTNaZmUQuLtI+ZPPmGfpZyKI4tcFz5K2LLBXXBtddbfCCODs706xZsygwMJBc\nXFzUemY7OzvT1KlTqVmzZuTu7l5or+1ff/2VvL29SalUkpeXl6ryW0xMDLVo0YI8PT2pb9++5O/v\nr6pCl1NjPCAgQFUj3MLCgpRKJb3xxhtERLRt2zbq169fvv0dOnSoyGQ9d+5cev/994mIKCUlhby8\nvGjPnj1EVLL3N7fs7GyqXLkyxcXFqR4LCgqS9fNv8tLSiBwdpX2PLlpk6GehFZ0n6+TkZLK0tKRr\n166pHhs0aJBa8f4c69atozfffLPINk0yWRMRLV4s7UNmZ0f08qWhn4VkT548IXt7e1WZx88//7zA\nkpKHDh0iJycn1ZFGXj179lSVtsx9a9KkSYHbPHz4kKpVq6b2WFhYGNWvX5+OHz9Ozs7OdO/ePSIi\nmjx5surIeunSpbRgwQKKjY2loUOH0vfff0/z58+n8ePH04kTJ9Tae/nyJXl6etLvv/+u9pi1tTVl\nZGSojry++OILWrlyJQ0bNoxu3LhBCxYsUB15bdy4kSZPnkxE4odNQa+NtuVGd+zYQURE0dHR1K5d\nO7Xl77zzDtnb21PDhg3p4cOHBcZfUi4uLjRlyhQiIrp16xZZW1ur3ncXFxfVj4IHDx5Q7dq16fz5\n8/na8PX1Vb3O2dnZqrMATZs2pR9//JGIiE6cOEHm5uaqYV9BQUHUtWtX1Q+vW7du5Xu9xo4dW2DJ\n1eIk6+zsbOrQoQMtXbqUhgwZQp9++qkqjpK8v7mFhoZSs2bN1B6bPn06ff755xpjKVPWr5f2HVqp\nEtGTJ4Z+FlrRebI+e/YsVapUSe2xr776Su00W47Q0FCysrKiGjVqUP369Wnu3LmUWcDpCpNN1k+f\nig+LlA/bihWGfhayKKo2eGF1kaXg2uDy1wYviouLC505c0Z1v27duqozGy4uLnTs2DHVshEjRhQY\n03/+8x8KCAigRYsWUXR0NBGJU9bly5dXW0+pVKol6y1btqiWFfR6de7cWa1GeY7iJGsi8ePPwcGB\nAgMDVT8KchT3/c0RFhZGTk5OdPXqVbXHV6xYUayZzMqErCwiT09p35+jRhn6WWhNU96TpTZ4cnIy\nbG1t1R6zsbHBixcv8q3bunVrxMTEwNnZGdHR0ejTpw8sLCwwdepUOUIxvCpVgMGDge++076NxYtF\n1R0L0y7drqk2uKa6yLlxbfDC6as2+Pvvv4+bN29CoVDgn3/+UauFnUNqHfAlS5YgJiYGf//9N3r1\n6oWPPvoIvXv31vj8ARQYS14FPaeCYsj9PI8cOQIrKyvExsbC3NwcSUlJePnypdr+SvL+Hj9+HAMH\nDsSuXbvg4eFRrBjLpD17pI+KyVVZsDSRJRtYW1vj+fPnao89e/ZMNdF9bq6urqr/e3l5YdasWVi0\naFGByXp2rgkygoKC8pUVNFoffigtWcfGAtu3AyZcpAJQrw2+IlfJwKLqIue2bdu2Eu2zqNrggYGB\nCAwMhK+vb4naNfba4DNmzCiyNjgA1KhRA0DJaoPv2LFDq9hyhIaGomXLlnj06BH27NmDSZMm5Vvn\nypUr8PT0hKenJ5KTk3H69GkMHz4c3t7e2LhxI/r374+TJ0/iwoULhe7H1tYWL1++RFZWFsxfF8Io\n7LUtKDnmfZ5Pnz7FgAEDsGXLFuzbtw8jRozA5s2bS/r0cerUKfTp0wfbt2+HUqnMt5zrgOfyute+\n1t56C/D0lCcWPQgLC0NYWFjxVpbj0L2ga9YDBgygadOmFbntzz//TE2bNs33uEyhGU6HDtJO5TRp\nIqbgNHEbNmwgFxcXtcd69epFVapUUbsGvW/fPtn26enpqToNGxERQa6urvT48WMiItq/fz81bNhQ\n1Tu4uP744w9SKBTk4+Ojinn8+PGq5WFhYdS+fXut4j116hQFBgaqPebn50cODg5kYWFBjo6ONGLE\nCCIiiouLI6VSqVovISGB2rdvTx4eHuTr60vHjx8nInFqNSQkRLXehQsXqFmzZuTj40M+Pj7Uo0cP\nunv3rizxu7i4qHUmzX3fxcWFpk2bpupgljOVJRHR8OHDVdf9u3fvTl5eXqRUKqlt27YUGxtLREQX\nL14kf39/8vLyon79+lFAQIDaafC8ZUtHjBhBjRo1UnUw++WXX9Q6mGVmZlKdOnXIzs6OLC0tydHR\nkebMmVPg8+revTt9+eWXRESUlZVFQUFB9P3335f49WnevDnVrFlT7fN+4cIF1fK2bdtSeHh4idst\ndY4dk/adCRD99puhn4UkmvKebBmxb9++1K9fP0pJSaEjR45Q5cqV6eLFi/nW2717NyUkJBAR0aVL\nl8jLy6vAzhUmn6z37pX+wZMxgRkK1wYvntJaGzxvIpeqpHXFU1NT1YZuGZv4+Hhq3LixocMwDl27\nSvu+bNhQXPM2YXpJ1k+ePFEbZ53zh3/79m2ytrZW/YqfPHky1apVi6ysrKhevXoUEhJSujqY5cjO\nFkfHUj58rVub7NF1XFwcNWjQgIKCgvQ+IcGrV6+oWbNmqh7JuvbgwQMKCAigbAnv1R9//EFjxoyR\nMarikyP+whg6WRMRLVu2jBYsWCBbDHKaPHkybd261dBhGF5kpPSDmzVrDP0sJNOU97g2uC79/DPQ\nr5+0Ng4eBNq2lScexhgzRu+/D+zcqf32tWuLvj7ly8sXkwFwbXBD6dkTyNWhTitz5sgTC2OMGaPz\n56UlagCYNMnkE3VROFnrkoUF8PHH0to4fBgobm9BxhgzNZ9/Lm17W1tg1Ch5YjFinKx1bcgQ4PVw\nGa3lGsLGGGOlxvnzYpiqFGPGiIRdynGy1rVKlYAJE6S1wUfXjLHSaO5cadtbWgITJ8oTi5HjDmb6\n8OQJ4OwMJCdr30ZQEHDokGwhMcaYQUVHA97e0toYMwbIVXDJ1HEHM0OrVk360XVYGPD337KEwxhj\nBjdzprTty5UDSkuZ6mLgI2t9SUwEXFykHV03bw5ERAAF1DVmjDGTceIEEBgorY1Ro4CVK+WJx0jw\nkbUxqF5d+tH1qVPShzgwxpghEUk/Ii5XDpg2TZ54TAQfWeuTHEfXDRqIaz0mPiMXY6yM2rsX6NRJ\nWhsjRgA//CBPPEaEj6yNRfXq0qdvu3IFWL9enngYY0yfsrOlHxFbWADTp8sTjwnhI2t9e/xYVDWT\ncnTt6AhcvQpUrChfXIwxpmtbtgB9+0prY/hwYNUqeeIxMnxkbUxq1JB+dH3vXqkarsAYKwMyMoDP\nPpPWRhk9qgY4WRvG5MlA5crS2pg/H3j6VJ54GGNM1374Abh+XVobI0dKn2/BRHGyNoRq1YBPPpHW\nxtOn0qv/MMaYPiQlASEh0tqoWFH6kbkJ42RtKBMnArVqSWtj2TLg2jV54mGMMV2ZP1+MhpFi0iTA\nwUGeeEwQdzAzpBUrgHHjpLXRtSvw66/yxMMYY3K7cQNo3BhIT9e+japVxXzVVarIF9f/t3ffYVWW\n/x/A3wcIF7gR9ZuKW1wM90JU/GqpuDdOUDO10rRyomWlpQ3NvZA0UtNKvmqWA0dqSgIqrlyYluRE\nQBSB8/vjTn6ywefD4Yz367q4rqCH93k8Hs/nPPdz35/bCHGCmbEaNQqoXl1bxo8/smc4ERmvd9/V\nVqgB1UTFzAt1TnhlXdCCgoBBg7RluLoCoaGAtbXMORERSTh0CPDw0JZRsaK63Ve0qMw5GTFeWRuz\n/v0BFxdtGeHhbJRCRMYlJQWYNEl7jr+/RRTqnPDK2hj89BPwyivaMsqXV41S7O1lzomISIvAQGDY\nMG0ZtWsDp0+rXuAWgFfWxq5TJ+C//9WWcesWl3IRkXGIidG+PBUAFiywmEKdE15ZG4szZ9RweErK\ni2fY2ACnTgHOznLnRUSUVxMnAl98oS3Dywv4+WeL2hKYV9amoH591Z1Hi6Qk1crUkj7kEJFxOX0a\nWLxYW4ZOByxcaFGFOics1sZkzhygeHFtGXv3At99J3M+GllZWcHV1RX79u0DAPj7+2Pz5s358lij\nRo3Cr7/+qjknICAAffv2zfT/OTk54ezZs5ofIzdmz54NKysr7Ny5M/VncXFxsLOzQ5MmTdIce+7c\nOVhZWeGLf69k1q1bBzc3N7i5uaFMmTKoVKlS6vfHjx/H8OHDsWTJkiwfe9myZbCyskJ4eHian3t6\neqJ69epwc3NDvXr10KdPHzx8+DDTjC5duuDq1asv+sfPlJWVFR49eiR2XFaaN2+e+me0sbFJfe5G\njhyJAwcOZHj+z5w5g6pZtMCcPXs2HB0d4e7ujtq1a6Np06ZYtGgRUnIxghYSEpLmsZYtWwZnZ2c0\natQIhw8fxpYtW9Ic7+XlhTJlymT7d5vv9Hpg/HggOVlbjq8v0LChzDmZCRZrY1KuHDB9uvacSZO0\n7eol6MiRI2jfvj0AYM6cOejXr1++PM6qVavQqlWrDD9PSkoSewxD3prR6XRwd3fH+udm+W/ZsgV1\n6tSBLt3Vxtq1a9GrVy+sW7cOADBixAiEhYUhLCwM3t7emDp1aur3TZs2hU6ny5CRPq93795Yu3Zt\nhnNavHgxwsLCEBkZCRsbGyxfvjzTjB07dmRZwAxBy9/TsWPHEBYWhp07d6JUqVKpz93atWvznKvT\n6TBs2DCcPHkSFy5cwKZNm7Bp0yZMnDgxz+e1ePFibNiwAb///jsuXbqU4YPvnj174O3tne3fbb4L\nCgIOHtSWYWfH+TeZYLE2Nm++qb1R/Y0bqr2fkXn+im727NkYOHAgunTpAmdnZ3Tt2hUJCQkAgB9/\n/BENGzaEm5sbGjRogIP//uP39PTExIkT0axZM9SsWRPTn/tg4+npiR07dqQ+jp+fHzw8PNC0aVMA\nwPr169G8eXM0btwYHTp0wMWLFwEAiYmJGDNmDGrVqoWWLVvixIkT2f4ZNmzYgMaNG6NmzZqpf5Yt\nW7aga9euqcc8efIEFSpUwI0bNzL8/pQpU9C0aVO4urrCy8sL169fz/KxPD09cerUKTx48AAAEBgY\niOHDh6cpGElJSfjmm2/w+eefIyUlBaGhoRlyMiswWRWdM2fOICYmBosXL8amTZuQmK6ZxbPfe/r0\nKR49eoTSpUtnmvP8KISnpycmT56MNm3aoHLlyliwYAE2bNiAli1bomrVqvjuuZEgKysrzJ49G25u\nbqhTpw62bduW5fOTnU8//TRDxqefforx48enHhMdHY3y5cvj8ePHmWZIfTB7Pqdq1apYu3Ytli1b\nhtjYWADAzp070bp1azRu3BgtW7bEb7/9liGjf//+uHz5Mnx8fPDqq6/C398fe/bsgZubG9588818\nOe88e/hQbVKk1dSpanULpcFibWwKFQI++UR7zsKFwIUL2nMEpb+i+/333xEUFIRz587h6dOn2Lhx\nIwA1XL5q1SqEhYXh1KlTcHNzS/39c+fO4ejRowgPD0dwcHBqgU6fferUKezevRsnT57EoUOHsGXL\nFhw8eBChoaGYPHkyRo4cCQBYsWIFoqKicO7cOezduxfHjx/P9srk9u3bCA0Nxa+//oqPPvoIZ86c\nQa9evXDmzBlcu3YNALB582a0bNkSL7/8cobff++993D8+HGEh4djwIABePfdd7N9vvr3749vv/0W\nV65cQXx8PBo0aJDmmB07dsDZ2RmVKlXCkCFDMlwN59WaNWswdOhQlC9fHo0aNcIPz7Wy1ev1eOON\nN+Dm5oby5cvj7t27GDp0aJbn/vx/37x5E4cOHcJvv/2GWbNm4fz58zhy5Ag2b96c4SrTxsYGYWFh\n2L59O0aPHo07d+7k+c+RWYafnx+2bt2aOkS+cuVKDB48GIULF85zvha1a9dG0aJFceHCBVy+fBlz\n587Frl27EBoailWrVmU6+rRp0yZUrFgRW7duxc6dO/H+++/Dy8sLYWFh+PLLLw16/ll6/33g77+1\nZVSqpCanUQYs1saod2/tXX+ePlX3jox4slnnzp1R/N979M2aNcPly5cBAO3bt8dbb72FBQsW4OzZ\ns7B/bu34sGHDYGVlhWLFimHAgAGp98Ofp9Pp0KdPHxQpUgQAEBwcjIiICDRr1gxubm6YOnVq6lXv\n/v37MWzYMFhbW6NIkSLw8fHJ9srE19cXAFCuXDl06dIF+/fvh7W1NcaMGZM6JLxkyRKMy6Ln+86d\nO9GiRQs0aNAACxcuzHBfOL1hw4Zh/fr1CAwMxLBM1qyuXbsWQ4YMAQAMHjwYmzdvxpMnT7LNzMrT\np08RFBQEHx8fAMhQ/J8fBr99+zbq1auX7YeN5z2bB1ChQgWULVsWvXr1AgC4u7vj5s2baa7gnz3H\ntWrVgru7O44dO5bnP0v6jKNHj6JUqVLw9vZGYGAgkpKSsHr1arz++ut5ys3qg1xeh56fvcZ2796N\ny5cvw8PDA25ubvDx8UFycjJu376dq983GqdPAxIfGhYuVLtrUQY2BX0ClAmdTu2o5eambaLGnj3A\nxo3Av2++xkSn06FQoUKp31tbW6cOg3/22WeIjIzE3r170bdvX0yaNAl+fn4A0r5J6fX6LN8kixUr\nlub7kSNHYs6cOZmeR/rM7GT1+KNGjYK7uzu6deuGmJiY1Pv0z4uKisKkSZMQGhqKKlWq4MiRIxg8\neHCWj6XT6VC1alUULlwYq1evxunTpxEREZH6/6Ojo/Hzzz8jPDwcs2fPBgAkJCRg69atGJRDC9vM\nnrft27cjJiYG7dq1AwCkpKQgOjoaN2/exH/+8580x1pZWaFnz56YMmVKto/zzPNXr9bW1qnfW//b\nIjcpKQm2trYAZApR+oxnf94JEyZg8ODBcHBwQN26dVE9j735HRwccDfd7lF37txBuXLlcp1x4cIF\nJCQkoE6dOjh+/Dg6d+6cZm6CyUlOVvscaJ0f0qED0KePzDmZIV5ZG6sGDdSVsVYTJ2rfmk7QszfR\n9G+mz39/4cIF1KtXD2+88QZ8fHxS78Pq9Xps2LABycnJiI+Px5YtWzItiul169YNgYGBuHnzJgAg\nOTkZJ0+eBKCu4r/++mskJycjISEB33zzTbbnHhAQAEANh+/atSu1sJUtWxZeXl4YOHBgllfVDx8+\nhK2tLRwdHZGSkpLl5Kxnj/XsOfn4448xf/58lCpVKs0xgYGB6Nu3L6KionD16lVcvXoVa9asydVQ\neGYFce3atViyZElqVlRUFIYPH546cS397+3fvx+1a9fO8bGyerysPHu8P/74A2FhYWjevHmufzen\njPr166NMmTKYOHFiln9P2alZsyYA4OeffwagXksrV65Ep06dMj0+/Z/72rVr8PX1xeuvvw47Ozt0\n7NgRP/30U5pVBjnNmwCAEiVKICYmJs/nny+WLwcyuc+eJzY2arkXl2plicXamM2Zo33P6zt3ZCZ9\nCHl2hZP+HvPz30+dOhUNGjSAm5sb9uzZkzrUqtPpUKdOHbRs2RKurq7o2rUrXn311WwfBwDatGmD\nDz/8EN7e3nB1dUWDBg2wfft2AMDo0aNRuXJlODs7o0OHDqmzpbPKdHBwSJ0ING3aNNSrVy/1//v6\n+uL+/fuZDlcDQIMGDdC3b1/UrVsXzZs3R7Vq1bJ9rGf/r3nz5mmuwJ/9PCAgIMOVube3N0JDQ9NM\nXMvsMWbOnIlKlSqhUqVKqFy5MoKCgnDw4EH0SXdlM3jw4DRXfc/uWdevXx+RkZGpy8Vykt0wcfr/\nl5ycnDpKsXLlSpQtWzbDcaNGjUJwcHCWmVllAOrvydraOs2kwNye20svvYRt27Zh3rx5cHNzg7u7\nO8qWLYtp06Zl+fuBgYFwd3dHnTp10K9fP/Tr1w+ff/45AFX8N2zYAF9fX7i6uqJu3bpYtWpVlo//\nTIcOHRAfHw9XV1e89dZbOf458s3Nm2pCmFYTJ7KZUw7YwczYSfTXBdT661xchUqysrJCbGxshiHp\nF9WuXTtMmTIlywJd0ObOnYvo6Ggs1toQwoJZWVkhLi4ORfNx4wY/Pz84Ozvj7bffzrfHKCjDhw9H\nkyZNXmjU4IX07g284Iz9VBUrAufPc18DsIOZaRsyBMhk/XCevfYakMUSlfzi6OiI1q1bZzoJzNzU\nq1cPW7duxcyZMwv6VExafq4R/uuvv1CnTh1cvnzZcMXMgLy8vHDo0CHY2dkZ5gG3b9deqAHV/5uF\nOke8sjYFERGAu7u2vuEAMGMGmw0QkXaxsUDduqqngxZt2wL79/Ne9b94ZW3qXFyAPC4xydS8eWrD\nECIiLWbO1F6ora3VqhcW6lzhlbWpePBATcC4dUtbTvPmwOHD6h8KEVFeHTkCtG6tvYfDO+8A8+fL\nnJOZ4JW1OShZEli0SHvOsWPat64jIsuUkACMHKm9UFetCvj7y5yThWCxNiV9+gBdumjPmTHD6FqR\nEpEJ8PeXee9YtgzIxxn/5ojD4KYmKgqoVw+Ij9eW07Kl2h2Hw+FElBvHjqmVKVonug4cCGTTfMiS\ncRjcnFSpAsydqz3nyBGZYXUiMn+PHwMjRmgv1KVKAf82hKG8YbE2RRMmAI0aac+ZNg344w/tOURk\n3ubMUY1LtPr0U+1dGS0Uh8FNVVgY0KSJto0+ADWsdeAAh8OJKHMnTqhVJFqvqj08gJAQLtXKBofB\nzZGbm8y+r7/+qtY6EhGl9+SJzPC3rS2wYgULtQYs1qZs9mygWjXtOVOncnY4EWU0cyYQGSmTU6eO\n9hwLxmFwUxcSAvy7TaMmTZqoq+yXXtKeRUSm78AB9d6i9X3Y3V3NJOd7S444DG7OPD0BiU0JTpyQ\nmWVORKbv4UO125/WQv3SS8C6dSzUAliszcG8eYCTk/acDz9Un4CJyLK9+abq6aDVjBlAw4bac4jD\n4GZj3z6gQwftOTVqAOHhgNAe1ERkYrZtU/tUa+XqChw/zqvqPOAwuCVo317tWa3VpUvA229rzyEi\n03PrFjB6tPYcGxsOfwtjsTYnn3wCVK6sPWfFCmDHDu05RGQ69HrA1xe4e1d71rRp6sqaxHAY3Nzs\n2QN07Kg9x9EROH0acHDQnkVExm/FCpnRuQYNgNBQtbaa8oTD4JbEywsYM0Z7TnQ04OenfTYoERm/\nc+dkmizZ2ADr17NQ5wMWa3O0YAFQvbr2nO3bgSVLtOcQkfF6/BgYMEDtVa3V+++r7ookjsPg5uro\nUaB1a+1tAgsVAn77DXBxkTkvIjIuEybItBzmtruacRjcErVoAUyfrj3nyROgf3/t+2cTkfH58UeZ\nQm1nB3z9NQt1PmKxNmczZ8pspXnhgmqSQETm48YNYORImawvvpDZp4CyxGFwc3f+vOrNK3E/6ttv\n1VU2EZm25GTVROnAAe1Z3t7ADz9wRy0BHAa3ZHXqqA3fJYweDVy9KpNFRAXno49kCrWDA7BqFQu1\nAfDK2hLo9cArrwC7d2vPatYMOHSInYmITNXhw0DbttonnwLqnre3t/YcAsAra9LpgLVrgdKltWf9\n9ptqzk9EpufuXWDQIJlC7efHQm1AvLK2JD/8APTsKZMVHAx07SqTRUT5LyVF/ZvdtUt7Vu3awO+/\nc8MfYbyyJqVHD+D112Wyhg4Frl2TySKi/DdvnkyhtrVVk01ZqA2KV9aWJiFB3Xc+fVp7VpMm6v51\noULas4go/+zfr1oRSwx/f/kl8MYb2nMoA15Z0/8rUkR9Ki5SRHvWiRPA5Mnac4go/9y6BQwcKFOo\nu3ZVHc/I4MSK9b1799CzZ0/Y2dnByckJQUFBWR77+eefo0KFCihRogR8fX2RmJgodRqUG3Xrqk/H\nEr76Cti0SSaLiGQlJalCHR2tPatiRbVHNZdpFQixYj1u3DgULlwY//zzDzZu3IixY8fi7NmzGY7b\nvXs35s+fj3379iEqKgpXrlyBv7+/1GlQbvn5AX37ymVduCCTRURy/P2BkBDtOTodsGEDULas9ix6\nISL3rOPj41G6dGlERkaiRo0aAIBhw4ahYsWK+Pjjj9McO2jQIFSrVg1z584FAOzfvx+DBg3C+G16\nPgAAHkpJREFU33//nfbEeM86/z14oDaIj4rSnlW/vlrWVbSo9iwi0m7nTqBLF5ms6dOBf9+zKf/k\n+z3rixcvwsbGJrVQA4CLiwsiIyMzHHv27Fm4PLeDU8OGDREdHY379+9LnArlRcmSQFCQTPP9M2eA\nceO05xCRdtevA0OGyGS1aAHMni2TRS9MpFjHxcWhePHiaX5mb2+P2NjYTI8tUaJE6vfPfi+zY8kA\nWrQAPvhAJisgAFi5UiaLiF7M48dA797AvXvas0qVAr75BrCx0Z5Fmoj8DdjZ2eHhw4dpfhYTEwN7\ne/scj42JiQGATI+d/dynOU9PT3h6ekqcLqX37rtqCZbEGszx44GGDYHmzbVnEVHe6PVqhCs0VCYv\nMBBwcpLJogxCQkIQkss5Bfl2z3rIkCGoVKkSPvroozTHDh48GFWrVk29Z7137174+PjwnnVBu3sX\ncHMD/vxTe1bFiqq7Ufny2rOIKPeWLwfGjpXJevdd1UiFDCa7uifWFGXgwIHQ6XRYvXo1Tp48ia5d\nu+Lo0aNwdnZOc9zu3bsxfPhw7Nu3D+XLl0fPnj3RsmXLDEWdxboAHDsGeHgAT59qz2rTBti7lxt+\nEBnK0aNqgw6Jf78eHurfL4e/DcogTVGWLl2KhIQElCtXDj4+Pli+fDmcnZ1x/fp12Nvb48aNGwCA\nTp064Z133kG7du3g5OSE6tWrY86cOVKnQVo0by63neahQ8Dbb8tkEVH2/v5b3aeWKNTlyqnGSSzU\nRoXtRiktvV6tv966VSYvMFBuVioRZZSYCLRvD/z6q/YsKyvgl19UHhkc241S7ul0wJo1wHPL8DQZ\nPRo4eVImi4gymjRJplADwJw5LNRGilfWlLnwcDUs/uSJ9qwqVdTsVHY/IpK1fj0wfLhMVqdOqpGK\nFa/hCgqvrCnvXF1V328JUVGqP3FSkkweEakVF2PGyGS9/LJqJ8pCbbT4N0NZ8/UFRoyQydqzB5gy\nRSaLyNLduqX2p5cY+XrpJWDLFo58GTkWa8qaTgcsXQo0biyT98UXwNq1MllElurxY6BnT+DfFTaa\nffUVmxiZAN6zppxdvw40agTcuaM966WXgH37gNattWcRWRq9Xt2jDgyUyfPzA1atkskizQzSFEUa\ni7WR2b8f6NgRSE7WnuXgAJw4oSaeEVHuLVggdzupaVPg4EGgUCGZPNKME8xIu3btgE8+kcm6fRvo\n3h2Ii5PJI7IEO3cC77wjk1WunOqlwEJtMlisKfcmTgQGDJDJiogAhg0DUlJk8ojM2blzakWFxGij\ntTWwebOaAU4mg8Wack+nA1avVrtqSdi2TTVhIKKs3b0LdOsGpNvZ8IUtXKh6iJNJ4T1ryrsrV9QM\n8fv3ZfI2bQL69ZPJIjInT58CnTurSZkSBg8Gvv5affAmo8N71iSrWjW1Ib3UP/jhw1WDByJKa+JE\nuULt6gqsXMlCbaJYrOnFdO4MfPyxTFZCAuDtLbdulMgcfPUVsGSJTFaZMuq2U9GiMnlkcBwGpxen\n16sdtTZulMlzdVVba9rZyeQRmaodO9QHWIkJmDY2am9qDw/tWZSvOAxO+ePZhLOmTWXywsPVjFeJ\ntdxEpioiQq26kFopsXQpC7UZYLEmbQoXBn74AahYUSbvf/9TW/4RWaK//gK6dpXrQTBhAjBqlEwW\nFSgOg5OM0FCgTRvVt1jC4sXA+PEyWUSmIC5OXQGHhcnkeXkBu3apYXAyCRwGp/zXuLHsJh1vvqnu\n2xFZguRkYNAguUJdo4ZaEslCbTZYrEnOwIHAtGkyWSkp6r5dRIRMHpExmzwZCA6WySpeXGWVLi2T\nR0aBw+AkKyUF6NUL+PFHmbyXXwZ++03unjiRsVm6FBg3TibLykrN+3jlFZk8MijuukWGFRcHtGwJ\nnD4tk+fmpnYH4pIuMje7dqkJZVIzvxcu5ARNE8ZiTYZ37Zpa0nX7tkxet26qqQPvwZG5iIhQ+7pL\nzfweMQJYs4YdykwYJ5iR4Tk5qaFwqS34goPVMhR+gCNzcP068OqrcoW6XTtg+XIWajPGYk35p0UL\nIDBQLm/5crkWp0QF5f59dU/5r79k8mrXVntT29rK5JFRYrGm/NWvH/Dhh3J506fLfgAgMqTHj4Ee\nPYCzZ2XyypZVSxxLlZLJI6PFe9aU//R6YORIICBAJs/GBti5E+jYUSaPyBBSUtTyxs2bZfJsbdWO\nXK1ayeRRgeM9aypYOh2wYoW6ryYhKQno3Vv1EicyFZMnyxVqQH34ZaG2GCzWZBi2tuq+Wu3aMnmx\nsWqCTlSUTB5Rfvr8c/Ul5f331VU6WQwOg5NhXb4MNG8O3Lkjk+fsDBw+zG5NZLw2bwb695fLGzpU\nXVVz5rfZ4TprMi6//gq0bw8kJsrktWkD/Pyz2gGMyJgcOAD8979yr/W2bdVrnTO/zRLvWZNxadVK\nbrIZABw6BAwZItcFikhCZKSa+S1VqGvVUo2BWKgtEos1FYyBA2WXdH33HTBxIpumkHG4eVOtpX7w\nQCbv2RIt3u6xWCzWVHCmTgXGjJHLW7SITVOo4N27B3TqBPz5p0xekSJqc44aNWTyyCTxnjUVrKQk\ntUuX1PaAALBqFeDnJ5dHlFuPHqn1/0eOyORZWQE//KB645PZ4z1rMl42NkBQkNr0Q8qYMcD338vl\nEeXG06eqY59UoQbU9pks1AQWazIGxYrJDvM96xR14IBMHlFOUlIAX191X1nK9Omyt4nIpHEYnIzH\npUtqH2ypbTWLF1cF29VVJo8oM3o98Pbbsk1PuJbaInGdNZmO48dVW9JHj2TyHB3Vuu7q1WXyiNKb\nPx947z25vI4d1UgTl2hZHBZrMi3/+x/Qvbvcuulq1VTBLl9eJo/omTVrZCczuroCBw8C9vZymWQy\nOMGMTEvXrsCyZXJ5V66oNa8xMXKZRD/8AIweLZdXpYraTY6FmjLBYk3GafRoYMYMubzwcHW1/vix\nXCZZroMHgQED5EZ/SpUCdu0CKlSQySOzw2FwMl7S+2ADQM+ewJYtgLW1XCZZlvBw1aP74UOZvEKF\ngD17gNatZfLIZHEYnEyTTqcanHTtKpf5/fdqOQw/CNKLuHwZ6NxZrlBbWQGbNrFQU45YrMm42djI\nv5mtWQNMmcKCTXnz119qB63oaLnMVavU7RmiHLBYk/ErWhTYvh1o0EAuc+FC4KOP5PLIvN25o5ZU\nXbkilzlvnrrNQ5QLvGdNpuOvv9T2mteuyWUuXgyMHy+XR+YnJgbo0AH4/Xe5zEmTgAUL2PSE0uA6\nazIfFy+qIXGpLmcAsH696hhFlF58vLpHffiwXObQocC6dep+NdFzWKzJvPz+O+DpCcTFyeRZW6v9\nsHv0kMkj8/DkCeDtDfz8s1xmly5qkuNLL8llktngbHAyL40aAT/+KNeOMTkZ6N8f2LtXJo9MX1KS\n2gxGslC3bAls3sxCTS+ExZpMU/v2wMaNcvf8EhPVrNxjx2TyyHSlpKiJX5LbrNavr9roFi0ql0kW\nhcWaTFefPmq/Xynx8aot6alTcplkWvR6NeHw66/lMqtUAXbvVl3KiF4QizWZttdeA+bMkct78ECt\npb10SS6TTINer3bPkuxL7+CghtIrVpTLJIvEYk2mb+ZM2eVX0dGAlxdw44ZcJhm/jz4CPvlELs/O\nTvX7rlVLLpMsFmeDk3lISQGGDQM2bJDLrFNHbdjg4CCXScZp0SLgzTfl8goVUjtotW8vl0lmj0u3\nyDIkJan72D/+KJfp7g7s2weUKCGXScZl3TrZTmLW1mpyWrducplkEbh0iyyDjQ3w7beyVzMnT6qN\nRB49kssk47FlC+DnJ5en06nJaSzUJIzFmsxL4cLADz8AzZrJZR4+DPTqpZpkkPnYuRMYNEhuT2oA\nWLFCrc8mEsZiTebH3l69EdevL5e5ezcweLAaaifTFxIC9O4t+/f52WfAqFFyeUTPYbEm81S6tFoy\nU62aXObWrerNWPJKjAzv+HE1TP34sVzm7NnAxIlyeUTpsFiT+apQAdizR3aNa0CAelPm5EfTdOqU\n2phDqq88oHbQmjVLLo8oEyzWZN6qVgV++QUoU0Yuc9EivjmboosXVcOb+/flMkeN4laXZBBcukWW\nITRUzRKPjZXL/OQTYMoUuTzKP9evq61V//xTLnPgQDXz29paLpMsGtdZEwHAgQNqCFTyXuXy5cCY\nMXJ5JO/WLaBNG9kWst26qTkM3EGLBLFYEz2zY4fat1pqFrBOp3b/4nId43TvHtC2LXDmjFxm+/bq\ndVS4sFwmEdgUhej/demihi6l7jHq9cCQIcD27TJ5JCc2Vo2kSBbqFi1UhzwWajIwzcX63r176Nmz\nJ+zs7ODk5ISgoKAsjw0ICIC1tTXs7e1Tvw4ePKj1FIjyZsAANXwtJTkZ6NcP2LtXLpO0efRIDVWf\nOCGX6eqq1u/b2cllEuWSjdaAcePGoXDhwvjnn38QFhaGLl26wMXFBXXr1s30+FatWrFAU8EbPRp4\n+FBugtiTJ0D37mqpWPPmMpn0YhITVY/4AwfkMmvXVo1xSpaUyyTKA01X1vHx8di2bRs++OADFC1a\nFK1atUL37t3xdTYbt/M+NBmNyZOBGTPk8uLjgVdeASIi5DIpb5KSVKe5XbvkMp2c1IewcuXkMony\nSFOxvnjxImxsbFCjRo3Un7m4uCAyMjLT43U6HcLCwuDg4IDatWtj7ty5SE5O1nIKRNq8/z4wYYJc\n3oMHai3vxYtymZQ7KSlq3fN338llPmus8/LLcplEL0DTMHhcXByKFy+e5mf29vaIzWItq4eHByIj\nI1GlShWcOXMG/fv3h42NDd57771Mj589e3bqf3t6esLT01PL6RJlpNMBX3yhJiMFBMhk/vMP4OWl\nNgCpXFkmk7Kn1wNvvSX3dwioRjq//AJUry6XSfSckJAQhISE5OrYbJdueXp6Znl/uXXr1li0aBFa\ntWqF+Pj41J8vWLAABw8exPZczI7dtGkTPv30U4SGhmY8MS7dIkNKSlITz7ZulcusWRM4dAhwdJTL\npMzNmAF8+KFcnr09sH8/0KiRXCZRDrKre9leWedU8ePj45GUlIRLly6lDoVHRESgfh52O2JBJqNg\nY6PWS8fHAz/9JJP5xx9Ax45qh6fSpWUyKaP582ULdZEiah01CzUZEU33rIsVK4ZevXph1qxZePTo\nEQ4fPozg4GAMGTIk0+N37dqF6OhoAMD58+cxd+5c9OjRQ8spEMkpVEhdWbdpI5d5+jTw6quybU7p\n/y1bBmRxG+2F2Nqq/dAlXwNEAjSvs166dCkSEhJQrlw5+Pj4YPny5XB2dgYAXL9+Hfb29rhx4wYA\nYN++fXBxcYGdnR26dOmC3r17Y9q0aVpPgUhO0aJAcDDg7i6X+dtvqmuaZJtTUs1tXn9dLs/aGvj2\nWzVBkMjIsN0oUWbu3FFtKs+elctkP2k5338P9O2rGtJICQxU3eiICgjbjRLlVdmyaiZw1apymcHB\nwLBhsgXGEv3yi5oMKPk8Ll3KQk1GjcWaKCsVK6o1thUrymUGBamhW44avZgjR9QthcREucz584Gx\nY+XyiPIBizVRdqpVU1dyZcrIZa5cCbzzDgt2XoWFqcl6jx7JZU6frv4uiIwcizVRTurWVX2h0zUA\n0mTBAtnlRubu/HmgUycgJkYu8403gA8+kMsjykecYEaUW4cOqYKRkCCX+eWXqmhQ1q5dA1q3Bm7e\nlMscMQJYvRqw4vUKGY/s6h6LNVFe7N6tZnU/fSqXuXatKh6U0d9/qzXPly/LZfbtq+YOWFvLZRIJ\n4GxwIimdOqk3eskrMj8/2c0nzMXdu2rNs2ShfvVVYMMGFmoyOSzWRHnVuzewZo1cXkoKMGiQXJtT\ncxAbq7YbPXNGLrNtW/WhyNZWLpPIQFisiV7E8OHAokVyeU+fAr16qfvili4hQd1qOHFCLrNJE2D7\ndtX3m8gEsVgTvagJE4C5c+XyEhKALl2ATHahsxhPn6p7ygcOyGXWrw/s2iU7m5/IwFisibSYNg2Y\nMkUuLzYW6NxZts2pqUhJURPtduyQy6xeHfj5Z9l18kQFgMWaSAudTnXAGjNGLvPuXcDLC7hyRS7T\n2On1wKRJaptSKS+/rDrQVaggl0lUQFisibTS6YAlS9QkMSl//w106CC7ttiYffyxWnMuxcFBFWon\nJ7lMogLEddZEUp4+Bfr0UROZpDg7q/u3Dg5ymcZm1Spg9Gi5vBIlgJAQwNVVLpPIANgUhchQHj9W\nk8T27ZPLdHdXeSVKyGUai23b1ISylBSZvKJFVS/3li1l8ogMiMWayJDi4oCOHYFjx+QyW7dW3dOK\nFpXLLGj796vJdFI7aNnaqslpXl4yeUQGxg5mRIZkZwfs3Ak0bCiXefiwWoctuTVkQTp5EujeXe7P\nY20NbN7MQk1mi8WaKD+UKqWWDNWsKZe5ezcwapTpb635xx/qijo2ViZPpwMCAlTxJzJTLNZE+cXR\nUc1IrlRJLjMwEJg5Uy7P0G7dUv3Vb9+Wy1y6FPDxkcsjMkIs1kT5qXJlYO9eVbilfPghsGKFXJ6h\nxMWpyXdXr8plzp8PvPaaXB6RkWKxJspvNWuqIfGSJeUyX39ddolYfktKAvr3V/eqpbzzjvoisgAs\n1kSG0LCh6k9drJhMXkoKMGCA7Izz/KLXqw8XO3fKZY4cCcybJ5dHZORYrIkMpXlzdTVcqJBM3rPd\nqf74QyYvv3z8sWp8IsXbW90G0OnkMomMHNdZExlacLBahpWUJJNXrRpw5IjsfXEpGzYAQ4bI5bVp\no2bFc6tLMkNsikJkbIKCgMGD5ZZhNW6s2pIaU9OUffvUEq2nT2XyGjZUf0bJe/9ERoRNUYiMzcCB\nwPLlcnmhocCwYXJtO7WKjAR69pQr1FWrAj/9xEJNFovFmqigjB4NfPKJXN533wH+/nJ5L+r2baBr\nV+DhQ5m8cuXUbHpudUkWjMPgRAVt4kTgiy/k8r7+uuCahCQmqpafhw7J5Nnbq6FvNzeZPCIjxnvW\nRMbs2TKsLVtk8mxt1SYZht55Sq8HfH2Bdetk8mxt1dB3u3YyeURGjvesiYyZlZVqI+rhIZOXmAj0\n6AFcuyaTl1uffSZXqAHV75uFmggAr6yJjMf9+2orzLNnZfLq1wd+/RUoXlwmLzv/+59a/yz1b3be\nPODdd2WyiEwEh8GJTMX160CLFsBff8nkeXsD33+vrt7zy5kz6pzj4mTyxo4Flixh0xOyOBwGJzIV\nlSurtpz29jJ527erDmL55cEDtURLqlB36wYsWsRCTZQOr6yJjNHevcArr8isU9bp1AeAzp21Zz0v\nJUXdGw8Olslr0kRNjJPqn05kYnhlTWRqOnQAVq+WydLrgUGDgCtXZPKe+egjuUJdrZq6781CTZQp\nXlkTGbNp0+SGsRs2BI4elWlJ+tNPwKuvykwoK1lS7R5Wu7b2LCITxglmRKYqJQXo2xfYtk0mb/Bg\n1TRFyz3hq1eBRo3U7HWtrK1V4ffy0p5FZOI4DE5kqp6twXZ3l8nbuFHbWujERKBfP5lCDQCLF7NQ\nE+UCizWRsStWTM3qrlhRJm/8eODcuRf73alT1aYhEsaNU8u0iChHHAYnMhW//672c05I0J5Vvz5w\n/Hje9oXetUvdp5bg5aXybGxk8ojMAIfBicxBo0ZqSFzCmTPApEm5P/7vv9UWnBJq1QI2b2ahJsoD\nFmsiU9KnD/DeezJZy5cDW7fmfFxKCjBkiNr6Uit7ezWkX6qU9iwiC8JhcCJTk5ysGpzs2aM9q0wZ\nIDIScHTM+pgvvlDbeEr4/nvVSIWIMuAwOJE5sbYGvv0WqFJFe9bdu8Brr2W9XvriRTWpTMLUqSzU\nRC+IV9ZEpurkSaBVK+DxY+1ZX38N+Pik/VlysprQdvSo9vyOHdWEMmtr7VlEZopX1kTmyN1d3XeW\nMGFCxp2+PvtMplBXrgx88w0LNZEGvLImMnUjRgABAdpzunVTk78ANfzdsCHw5Im2TBsbtad206ba\nz4/IzPHKmsicffWVTF/t4GBVrPV61bBEa6EG1GYfLNREmvHKmsgcREQAzZppL7BVqgD+/sDIkdrP\nqVMntTWnFa8JiHKDG3kQWYKlS9UVsTFwdFQfILJbEkZEaXAYnMgSjB0LeHsX9FkogYEs1ESCWKyJ\nzIVOB6xYAZQuXbDnMXYs8N//Fuw5EJkZDoMTmZtNm4ABAwrmsZ2cgNOnATu7gnl8IhPGYXAiS9K/\nP9C3b8E89rp1LNRE+YDFmsgcLV2q+n4b0rhxgKenYR+TyEJwGJzIXK1dC/j6GuaxHB2BCxeAEiUM\n83hEZojD4ESWaPhwoGVLwzzWZ5+xUBPlI15ZE5mzU6dUD/Hk5Px7jHbtgL171Wx0InphvLImslQN\nGwJjxuRfvk4HfPklCzVRPuOVNZG5u3ULqFEDiI+Xzx4+XM0AJyLNeGVNZMnKlwfefls+t3Bh4P33\n5XOJKAMWayJL8PbbQMmSspljxgCVKslmElGmWKyJLEHx4sDrr8vl2djkz9U6EWWKxZrIUrzxhhq6\nljBoEK+qiQxIc7H+6quv0LhxYxQuXBgjRozI8fjPP/8cFSpUQIkSJeDr64vExEStp0BEueHoKNeG\ndPx4mRwiyhXNxfo///kPZs6ciZG52Kx+9+7dmD9/Pvbt24eoqChcuXIF/v7+Wk/BYEJCQgr6FCwO\nn3NhuehoFpLTAQ0aAI0bS5wN/Yuvc8Mztedcc7Hu2bMnunfvjjK56EO8fv16+Pn5wdnZGSVLlsSs\nWbMQEBCg9RQMxtT+cs0Bn3NhHh5A5crZHhKSU4aPD9dVC+Pr3PBM7TkXu2edmzXRZ8+ehYuLS+r3\nDRs2RHR0NO7fvy91GkSUHZ0OaNtWW0bXrjLnQkS5Jlasdbn4pB0XF4cSz/UPLl68OAAgNjZW6jSI\nKCfdu7/475YsCTg7y50LEeWOPhtt27bV63S6TL/atGmT5tjp06frhw8fnl2c3sXFRb9ly5bU72/f\nvq3X6XT6e/fuZXosAH7xi1/84he/LOLLxcUly/ppg2zkZUw/N1fW9erVQ3h4OPr06QMAiIiIgKOj\nI0qVKpXh2PDw8Fw/NhERkTnTPAyenJyMx48fIykpCcnJyXjy5AmSs9jhZ+jQoVizZg3OnTuH+/fv\n44MPPsjVci8iIiJLprlYf/DBByhatCjmz5+PDRs2oEiRIvjwww8BANevX4e9vT1u3LgBAOjUqRPe\neecdtGvXDk5OTqhevTrmzJmj9RSIiIjMmtHuukVEREQK240SEREZORbrbLCVquHdu3cPPXv2hJ2d\nHZycnBAUFJTlsQEBAbC2toa9vX3q18GDBw14tqYrL88zX9cycvuc83UtIy/v36bwGmexzoYltVI1\nFuPGjUPhwoXxzz//YOPGjRg7dizOnj2b5fGtWrVCbGxs6peHh4cBz9Z05fZ55utaTl5e23xda5fb\n92+TeY1nuzCa9Hq9Xj9jxowc15APHDhQP3369NTv9+3bpy9fvnx+n5pZiYuL09va2ur/+OOP1J8N\nHTpU/95772V6/Lp16/StW7c21OmZjbw8z3xdy8jLc87Xtayc3r9N5TXOK+tc0LOVqkFcvHgRNjY2\nqFGjRurPXFxcEBkZmenxOp0OYWFhcHBwQO3atTF37twslw3S/8vL88zXtYy8POd8XcvK6f3bVF7j\nLNa5wFaqhhEXF5f6vD1jb2+f5XPo4eGByMhI3L59G1u3bkVQUBA+/fRTQ5yqScvL88zXtYy8POd8\nXcvK6f3bVF7jFlusPT09YWVllelX+vtDubmytrOzw8OHD1O/j4mJAaD+QZKS03Nub2+f5jkE1POY\n1XNYtWpVVKlSBQBQv359zJo1C999912+/zlMXfrXKpD188zXtYy8POd8XcvK6f3bVF7jFlusQ0JC\nkJKSkulX+pmXeWml+kx2rVQtVU7Pec2aNZGUlIRLly6l/k5ERATq16+f68fIzQcrS1erVq1cP898\nXcvIy3OeGb6uX1xO79+m8hq32GKdG2ylaljFihVDr169MGvWLDx69AiHDx9GcHAwhgwZkunxu3bt\nQnR0NADg/PnzmDt3Lnr06GHIUzZJeXme+bqWkZfnnK9rGbl9/zaZ13hBzm4zdv7+/hl2G5szZ45e\nr9fro6Ki9HZ2dvo///wz9fjPPvtM7+joqC9evLh+5MiR+sTExII6dZN17949fY8ePfTFihXTV6lS\nRR8UFJT6/9I/55MnT9Y7OjrqixUrpq9WrZre399fn5SUVFCnblKyep75us4/uX3O+bqWkdX7t6m+\nxtlulIiIyMhxGJyIiMjIsVgTEREZORZrIiIiI8diTUREZORYrImIiIwcizUREZGRY7EmIiIycizW\nRERERu7/AP8FKy4MQovVAAAAAElFTkSuQmCC\n", + "text": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Table of Contents" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "- [Bitstrings from positive and negative elements in a list](#Bitstrings-from-positive-and-negative-elements-in-a-list)\n", + "- [Command line arguments 1 - sys.argv](#Command-line-arguments-1---sys.argv)\n", + "- [Data and time basics](#Data-and-time-basics)\n", + "- [Differences between 2 files](#Differences-between-2-files)\n", + "- [Differences between successive elements in a list](#Differences-between-successive-elements-in-a-list)\n", + "- [Doctest example](#Doctest-example)\n", + "- [English language detection](#English-language-detection)\n", + "- [File browsing basics](#File-browsing-basics)\n", + "- [File reading basics](#File-reading-basics)\n", + "- [Indices of min and max elements from a list](#Indices-of-min-and-max-elements-from-a-list)\n", + "- [Lambda functions](#Lambda-functions)\n", + "- [Private functions](#Private-functions)\n", + "- [Namedtuples](#Namedtuples)\n", + "- [Normalizing data](#Normalizing-data)\n", + "- [NumPy essentials](#NumPy-essentials)\n", + "- [Pickling Python objects to bitstreams](#Pickling-Python-objects-to-bitstreams)\n", + "- [Python version check](#Python-version-check)\n", + "- [Runtime within a script](#Runtime-within-a-script)\n", + "- [Sorting lists of tuples by elements](#Sorting-lists-of-tuples-by-elements)\n", + "- [Sorting multiple lists relative to each other](#Sorting-multiple-lists-relative-to-each-other)\n", + "- [Using namedtuples](#Using-namedtuples)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "code", + "execution_count": 1, + "metadata": { + "collapsed": false + }, + "outputs": [], + "source": [ + "%load_ext watermark" + ] + }, + { + "cell_type": "code", + "execution_count": 2, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Sebastian Raschka 26/09/2014 \n", + "\n", + "CPython 3.4.1\n", + "IPython 2.0.0\n" + ] + } + ], + "source": [ + "%watermark -d -a \"Sebastian Raschka\" -v" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[More information](https://github.com/rasbt/watermark) about the `watermark` magic command extension." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Bitstrings from positive and negative elements in a list" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[back to top](#Table-of-Contents)" + ] + }, + { + "cell_type": "code", + "execution_count": 3, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "input values [ 1. 2. 0.3 -1. -2. ]\n", + "bitstring [1 1 1 0 0]\n" + ] + } + ], + "source": [ + "# Generating a bitstring from a Python list or numpy array\n", + "# where all postive values -> 1\n", + "# all negative values -> 0\n", + "\n", + "import numpy as np\n", + "\n", + "def make_bitstring(ary):\n", + " return np.where(ary > 0, 1, 0)\n", + "\n", + "\n", + "def faster_bitstring(ary):\n", + " return np.where(ary > 0).astype('i1')\n", + "\n", + "### Example:\n", + "\n", + "ary1 = np.array([1, 2, 0.3, -1, -2])\n", + "print('input values %s' %ary1)\n", + "print('bitstring %s' %make_bitstring(ary1))" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Command line arguments 1 - sys.argv" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[back to top](#Table-of-Contents)" + ] + }, + { + "cell_type": "code", + "execution_count": 5, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Overwriting cmd_line_args_1_sysarg.py\n" + ] + } + ], + "source": [ + "%%file cmd_line_args_1_sysarg.py\n", + "import sys\n", + "\n", + "def error(msg):\n", + " \"\"\"Prints error message, sends it to stderr, and quites the program.\"\"\"\n", + " sys.exit(msg)\n", + "\n", + "args = sys.argv[1:] # sys.argv[0] is the name of the python script itself\n", + "\n", + "try:\n", + " arg1 = int(args[0])\n", + " arg2 = args[1]\n", + " arg3 = args[2]\n", + " print(\"Everything okay!\")\n", + "\n", + "except ValueError:\n", + " error(\"First argument must be integer type!\")\n", + "\n", + "except IndexError:\n", + " error(\"Requires 3 arguments!\")" + ] + }, + { + "cell_type": "code", + "execution_count": 6, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Everything okay!\n" + ] + } + ], + "source": [ + "% run cmd_line_args_1_sysarg.py 1 2 3" + ] + }, + { + "cell_type": "code", + "execution_count": 7, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "ename": "SystemExit", + "evalue": "First argument must be integer type!", + "output_type": "error", + "traceback": [ + "An exception has occurred, use %tb to see the full traceback.\n", + "\u001b[0;31mSystemExit\u001b[0m\u001b[0;31m:\u001b[0m First argument must be integer type!\n" + ] + } + ], + "source": [ + "% run cmd_line_args_1_sysarg.py a 2 3" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Data and time basics" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[back to top](#Table-of-Contents)" + ] + }, + { + "cell_type": "code", + "execution_count": 7, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "13:28:05\n", + "26/09/2014\n" + ] + } + ], + "source": [ + "import time\n", + "\n", + "# print time HOURS:MINUTES:SECONDS\n", + "# e.g., '10:50:58'\n", + "print(time.strftime(\"%H:%M:%S\"))\n", + "\n", + "# print current date DAY:MONTH:YEAR\n", + "# e.g., '06/03/2014'\n", + "print(time.strftime(\"%d/%m/%Y\"))" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Differences between 2 files" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[back to top](#Table-of-Contents)" + ] + }, + { + "cell_type": "code", + "execution_count": 9, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Writing id_file1.txt\n" + ] + } + ], + "source": [ + "%%file id_file1.txt\n", + "1234\n", + "2342\n", + "2341" + ] + }, + { + "cell_type": "code", + "execution_count": 10, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Writing id_file2.txt\n" + ] + } + ], + "source": [ + "%%file id_file2.txt\n", + "5234\n", + "3344\n", + "2341" + ] + }, + { + "cell_type": "code", + "execution_count": 11, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "5234\n", + "3344\n", + "Total differences: 2\n" + ] + } + ], + "source": [ + "# Print lines that are different between 2 files. Insensitive\n", + "# to the order of the file contents.\n", + "\n", + "id_set1 = set()\n", + "id_set2 = set()\n", + "\n", + "with open('id_file1.txt', 'r') as id_file:\n", + " for line in id_file:\n", + " id_set1.add(line.strip())\n", + "\n", + "with open('id_file2.txt', 'r') as id_file:\n", + " for line in id_file:\n", + " id_set2.add(line.strip()) \n", + "\n", + "diffs = id_set2.difference(id_set1)\n", + "\n", + "for d in diffs:\n", + " print(d)\n", + "print(\"Total differences:\",len(diffs))" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Differences between successive elements in a list" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[back to top](#Table-of-Contents)" + ] + }, + { + "cell_type": "code", + "execution_count": 12, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "[1, 1, 2, 3]\n" + ] + } + ], + "source": [ + "from itertools import islice\n", + "\n", + "lst = [1,2,3,5,8]\n", + "diff = [j - i for i, j in zip(lst, islice(lst, 1, None))]\n", + "print(diff)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Doctest example" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[back to top](#Table-of-Contents)" + ] + }, + { + "cell_type": "code", + "execution_count": 17, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "ok\n" + ] + } + ], + "source": [ + "def subtract(a, b):\n", + " \"\"\"\n", + " Subtracts second from first number and returns result.\n", + " >>> subtract(10, 5)\n", + " 5\n", + " >>> subtract(11, 0.7)\n", + " 10.3\n", + " \"\"\"\n", + " return a-b\n", + "\n", + "if __name__ == \"__main__\": # is 'false' if imported\n", + " import doctest\n", + " doctest.testmod()\n", + " print('ok')" + ] + }, + { + "cell_type": "code", + "execution_count": 18, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "**********************************************************************\n", + "File \"__main__\", line 4, in __main__.hello_world\n", + "Failed example:\n", + " hello_world()\n", + "Expected:\n", + " 'Hello, World'\n", + "Got:\n", + " 'hello world'\n", + "**********************************************************************\n", + "1 items had failures:\n", + " 1 of 1 in __main__.hello_world\n", + "***Test Failed*** 1 failures.\n" + ] + } + ], + "source": [ + "def hello_world():\n", + " \"\"\"\n", + " Returns 'Hello, World'\n", + " >>> hello_world()\n", + " 'Hello, World'\n", + " \"\"\"\n", + " return 'hello world'\n", + "\n", + "if __name__ == \"__main__\": # is 'false' if imported\n", + " import doctest\n", + " doctest.testmod()" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## English language detection" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[back to top](#Table-of-Contents)" + ] + }, + { + "cell_type": "code", + "execution_count": 1, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "0.2\n" + ] + } + ], + "source": [ + "import nltk\n", + "\n", + "def eng_ratio(text):\n", + " ''' Returns the ratio of non-English to English words from a text '''\n", + "\n", + " english_vocab = set(w.lower() for w in nltk.corpus.words.words()) \n", + " text_vocab = set(w.lower() for w in text.split() if w.lower().isalpha()) \n", + " unusual = text_vocab.difference(english_vocab)\n", + " diff = len(unusual)/len(text_vocab)\n", + " return diff\n", + " \n", + "text = 'This is a test fahrrad'\n", + "\n", + "print(eng_ratio(text))" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## File browsing basics" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[back to top](#Table-of-Contents)" + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": { + "collapsed": false + }, + "outputs": [], + "source": [ + "import os\n", + "import shutil\n", + "import glob\n", + "\n", + "# working directory\n", + "c_dir = os.getcwd() # show current working directory\n", + "os.listdir(c_dir) # shows all files in the working directory\n", + "os.chdir('~/Data') # change working directory\n", + "\n", + "\n", + "# get all files in a directory\n", + "glob.glob('/Users/sebastian/Desktop/*')\n", + "\n", + "# e.g., ['/Users/sebastian/Desktop/untitled folder', '/Users/sebastian/Desktop/Untitled.txt']\n", + "\n", + "# walk\n", + "tree = os.walk(c_dir) \n", + "# moves through sub directories and creates a 'generator' object of tuples\n", + "# ('dir', [file1, file2, ...] [subdirectory1, subdirectory2, ...]), \n", + "# (...), ...\n", + "\n", + "#check files: returns either True or False\n", + "os.exists('../rel_path')\n", + "os.exists('/home/abs_path')\n", + "os.isfile('./file.txt')\n", + "os.isdir('./subdir')\n", + "\n", + "\n", + "# file permission (True or False\n", + "os.access('./some_file', os.F_OK) # File exists? Python 2.7\n", + "os.access('./some_file', os.R_OK) # Ok to read? Python 2.7\n", + "os.access('./some_file', os.W_OK) # Ok to write? Python 2.7\n", + "os.access('./some_file', os.X_OK) # Ok to execute? Python 2.7\n", + "os.access('./some_file', os.X_OK | os.W_OK) # Ok to execute or write? Python 2.7\n", + "\n", + "# join (creates operating system dependent paths)\n", + "os.path.join('a', 'b', 'c')\n", + "# 'a/b/c' on Unix/Linux\n", + "# 'a\\\\b\\\\c' on Windows\n", + "os.path.normpath('a/b/c') # converts file separators\n", + "\n", + "\n", + "# os.path: direcory and file names\n", + "os.path.samefile('./some_file', '/home/some_file') # True if those are the same\n", + "os.path.dirname('./some_file') # returns '.' (everythin but last component)\n", + "os.path.basename('./some_file') # returns 'some_file' (only last component\n", + "os.path.split('./some_file') # returns (dirname, basename) or ('.', 'some_file)\n", + "os.path.splitext('./some_file.txt') # returns ('./some_file', '.txt')\n", + "os.path.splitdrive('./some_file.txt') # returns ('', './some_file.txt')\n", + "os.path.isabs('./some_file.txt') # returns False (not an absolute path)\n", + "os.path.abspath('./some_file.txt')\n", + "\n", + "\n", + "# create and delete files and directories\n", + "os.mkdir('./test') # create a new direcotory\n", + "os.rmdir('./test') # removes an empty direcotory\n", + "os.removedirs('./test') # removes nested empty directories\n", + "os.remove('file.txt') # removes an individual file\n", + "shutil.rmtree('./test') # removes directory (empty or not empty)\n", + "\n", + "os.rename('./dir_before', './renamed') # renames directory if destination doesn't exist\n", + "shutil.move('./dir_before', './renamed') # renames directory always\n", + "\n", + "shutil.copytree('./orig', './copy') # copies a directory recursively\n", + "shutil.copyfile('file', 'copy') # copies a file\n", + "\n", + " \n", + "# Getting files of particular type from directory\n", + "files = [f for f in os.listdir(s_pdb_dir) if f.endswith(\".txt\")]\n", + " \n", + "# Copy and move\n", + "shutil.copyfile(\"/path/to/file\", \"/path/to/new/file\") \n", + "shutil.copy(\"/path/to/file\", \"/path/to/directory\")\n", + "shutil.move(\"/path/to/file\",\"/path/to/directory\")\n", + " \n", + "# Check if file or directory exists\n", + "os.path.exists(\"file or directory\")\n", + "os.path.isfile(\"file\")\n", + "os.path.isdir(\"directory\")\n", + " \n", + "# Working directory and absolute path to files\n", + "os.getcwd()\n", + "os.path.abspath(\"file\")" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## File reading basics" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[back to top](#Table-of-Contents)" + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": { + "collapsed": false + }, + "outputs": [], + "source": [ + "# Note: rb opens file in binary mode to avoid issues with Windows systems\n", + "# where '\\r\\n' is used instead of '\\n' as newline character(s).\n", + "\n", + "\n", + "# A) Reading in Byte chunks\n", + "reader_a = open(\"file.txt\", \"rb\")\n", + "chunks = []\n", + "data = reader_a.read(64) # reads first 64 bytes\n", + "while data != \"\":\n", + " chunks.append(data)\n", + " data = reader_a.read(64)\n", + "if data:\n", + " chunks.append(data)\n", + "print(len(chunks))\n", + "reader_a.close()\n", + "\n", + "\n", + "# B) Reading whole file at once into a list of lines\n", + "with open(\"file.txt\", \"rb\") as reader_b: # recommended syntax, auto closes\n", + " data = reader_b.readlines() # data is assigned a list of lines\n", + "print(len(data))\n", + "\n", + "\n", + "# C) Reading whole file at once into a string\n", + "with open(\"file.txt\", \"rb\") as reader_c:\n", + " data = reader_c.read() # data is assigned a list of lines\n", + "print(len(data))\n", + "\n", + "\n", + "# D) Reading line by line into a list\n", + "data = []\n", + "with open(\"file.txt\", \"rb\") as reader_d:\n", + " for line in reader_d:\n", + " data.append(line)\n", + "print(len(data))\n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Indices of min and max elements from a list" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[back to top](#Table-of-Contents)" + ] + }, + { + "cell_type": "code", + "execution_count": 19, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "min_index: 0 min_value: 1\n", + "max_index: 4 max_value: 5\n" + ] + } + ], + "source": [ + "import operator\n", + "\n", + "values = [1, 2, 3, 4, 5]\n", + "\n", + "min_index, min_value = min(enumerate(values), key=operator.itemgetter(1))\n", + "max_index, max_value = max(enumerate(values), key=operator.itemgetter(1))\n", + "\n", + "print('min_index:', min_index, 'min_value:', min_value)\n", + "print('max_index:', max_index, 'max_value:', max_value)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Lambda functions" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[back to top](#Table-of-Contents)" + ] + }, + { + "cell_type": "code", + "execution_count": 20, + "metadata": { + "collapsed": false + }, + "outputs": [], + "source": [ + "# Lambda functions are just a short-hand way or writing\n", + "# short function definitions\n", + "\n", + "def square_root1(x):\n", + " return x**0.5\n", + " \n", + "square_root2 = lambda x: x**0.5\n", + "\n", + "assert(square_root1(9) == square_root2(9))" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Private functions" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[back to top](#Table-of-Contents)" + ] + }, + { + "cell_type": "code", + "execution_count": 2, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "My message: Hello, World\n" + ] + } + ], + "source": [ + "def create_message(msg_txt):\n", + " def _priv_msg(message): # private, no access from outside\n", + " print(\"{}: {}\".format(msg_txt, message))\n", + " return _priv_msg # returns a function\n", + "\n", + "new_msg = create_message(\"My message\")\n", + "# note, new_msg is a function\n", + "\n", + "new_msg(\"Hello, World\")" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Namedtuples" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[back to top](#Table-of-Contents)" + ] + }, + { + "cell_type": "code", + "execution_count": 25, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "1 2 3\n" + ] + } + ], + "source": [ + "from collections import namedtuple\n", + "\n", + "my_namedtuple = namedtuple('field_name', ['x', 'y', 'z', 'bla', 'blub'])\n", + "p = my_namedtuple(1, 2, 3, 4, 5)\n", + "print(p.x, p.y, p.z)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Normalizing data" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[back to top](#Table-of-Contents)" + ] + }, + { + "cell_type": "code", + "execution_count": 28, + "metadata": { + "collapsed": false + }, + "outputs": [], + "source": [ + "def normalize(data, min_val=0, max_val=1):\n", + " \"\"\"\n", + " Normalizes values in a list of data points to a range, e.g.,\n", + " between 0.0 and 1.0. \n", + " Returns the original object if value is not a integer or float.\n", + " \n", + " \"\"\"\n", + " norm_data = []\n", + " data_min = min(data)\n", + " data_max = max(data)\n", + " for x in data:\n", + " numerator = x - data_min\n", + " denominator = data_max - data_min\n", + " x_norm = (max_val-min_val) * numerator/denominator + min_val\n", + " norm_data.append(x_norm)\n", + " return norm_data" + ] + }, + { + "cell_type": "code", + "execution_count": 31, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "data": { + "text/plain": [ + "[0.0, 0.25, 0.5, 0.75, 1.0]" + ] + }, + "execution_count": 31, + "metadata": {}, + "output_type": "execute_result" + } + ], + "source": [ + "normalize([1,2,3,4,5])" + ] + }, + { + "cell_type": "code", + "execution_count": 30, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "data": { + "text/plain": [ + "[-10.0, -5.0, 0.0, 5.0, 10.0]" + ] + }, + "execution_count": 30, + "metadata": {}, + "output_type": "execute_result" + } + ], + "source": [ + "normalize([1,2,3,4,5], min_val=-10, max_val=10)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## NumPy essentials" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[back to top](#Table-of-Contents)" + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": { + "collapsed": false + }, + "outputs": [], + "source": [ + "import numpy as np\n", + "\n", + "ary1 = np.array([1,2,3,4,5]) # must be same type\n", + "ary2 = np.zeros((3,4)) # 3x4 matrix consisiting of 0s \n", + "ary3 = np.ones((3,4)) # 3x4 matrix consisiting of 1s \n", + "ary4 = np.identity(3) # 3x3 identity matrix\n", + "ary5 = ary1.copy() # make a copy of ary1\n", + "\n", + "item1 = ary3[0, 0] # item in row1, column1\n", + "\n", + "ary2.shape # tuple of dimensions. Here: (3,4)\n", + "ary2.size # number of elements. Here: 12\n", + "\n", + "\n", + "ary2_t = ary2.transpose() # transposes matrix\n", + "\n", + "ary2.ravel() # makes an array linear (1-dimensional)\n", + " # by concatenating rows\n", + "ary2.reshape(2,6) # reshapes array (must have same dimensions)\n", + "\n", + "ary3[0:2, 0:3] # submatrix of first 2 rows and first 3 columns \n", + "\n", + "ary3 = ary3[[2,0,1]] # re-arrange rows\n", + "\n", + "\n", + "# element-wise operations\n", + "\n", + "ary1 + ary1\n", + "ary1 * ary1\n", + "numpy.dot(ary1, ary1) # matrix/vector (dot) product\n", + "\n", + "numpy.sum(ary1, axis=1) # sum of a 1D array, column sums of a 2D array\n", + "numpy.mean(ary1, axis=1) # mean of a 1D array, column means of a 2D array" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Pickling Python objects to bitstreams" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[back to top](#Table-of-Contents)" + ] + }, + { + "cell_type": "code", + "execution_count": 35, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "{1: 'some text', 2: 'some text', 3: 'some text', 4: 'some text', 5: 'some text', 6: 'some text', 7: 'some text', 8: 'some text', 9: 'some text'}\n" + ] + } + ], + "source": [ + "import pickle\n", + "\n", + "#### Generate some object\n", + "my_dict = dict()\n", + "for i in range(1,10):\n", + " my_dict[i] = \"some text\"\n", + "\n", + "#### Save object to file\n", + "pickle_out = open('my_file.pkl', 'wb')\n", + "pickle.dump(my_dict, pickle_out)\n", + "pickle_out.close()\n", + "\n", + "#### Load object from file\n", + "my_object_file = open('my_file.pkl', 'rb')\n", + "my_dict = pickle.load(my_object_file)\n", + "my_object_file.close()\n", + "\n", + "print(my_dict)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Python version check" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[back to top](#Table-of-Contents)" + ] + }, + { + "cell_type": "code", + "execution_count": 36, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "executed in Python 3.x\n", + "H\n", + "in for-loop:\n", + "e\n", + "l\n", + "l\n", + "o\n" + ] + } + ], + "source": [ + "import sys\n", + "\n", + "def give_letter(word):\n", + " for letter in word:\n", + " yield letter\n", + "\n", + "if sys.version_info[0] == 3:\n", + " print('executed in Python 3.x')\n", + " test = give_letter('Hello')\n", + " print(next(test))\n", + " print('in for-loop:')\n", + " for l in test:\n", + " print(l)\n", + "\n", + "# if Python 2.x\n", + "if sys.version_info[0] == 2:\n", + " print('executed in Python 2.x')\n", + " test = give_letter('Hello')\n", + " print(test.next())\n", + " print('in for-loop:') \n", + " for l in test:\n", + " print(l)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Runtime within a script" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[back to top](#Table-of-Contents)" + ] + }, + { + "cell_type": "code", + "execution_count": 4, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Time elapsed: 0.49176900000000057 seconds\n" + ] + } + ], + "source": [ + "import time\n", + "\n", + "start_time = time.clock()\n", + "\n", + "for i in range(10000000):\n", + " pass\n", + "\n", + "elapsed_time = time.clock() - start_time\n", + "print(\"Time elapsed: {} seconds\".format(elapsed_time))" + ] + }, + { + "cell_type": "code", + "execution_count": 6, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Time elapsed: 0.3550995970144868 seconds\n" + ] + } + ], + "source": [ + "import timeit\n", + "elapsed_time = timeit.timeit('for i in range(10000000): pass', number=1)\n", + "print(\"Time elapsed: {} seconds\".format(elapsed_time))" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Sorting lists of tuples by elements" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[back to top](#Table-of-Contents)" + ] + }, + { + "cell_type": "code", + "execution_count": 37, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "[(2, 3, 'a'), (2, 2, 'b'), (3, 2, 'b'), (1, 3, 'c')]\n" + ] + } + ], + "source": [ + "# Here, we make use of the \"key\" parameter of the in-built \"sorted()\" function \n", + "# (also available for the \".sort()\" method), which let's us define a function \n", + "# that is called on every element that is to be sorted. In this case, our \n", + "# \"key\"-function is a simple lambda function that returns the last item \n", + "# from every tuple.\n", + "\n", + "a_list = [(1,3,'c'), (2,3,'a'), (3,2,'b'), (2,2,'b')]\n", + "\n", + "sorted_list = sorted(a_list, key=lambda e: e[::-1])\n", + "\n", + "print(sorted_list)" + ] + }, + { + "cell_type": "code", + "execution_count": 38, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "[(2, 3, 'a'), (3, 2, 'b'), (2, 2, 'b'), (1, 3, 'c')]\n" + ] + } + ], + "source": [ + "# prints [(2, 3, 'a'), (2, 2, 'b'), (3, 2, 'b'), (1, 3, 'c')]\n", + "\n", + "# If we are only interesting in sorting the list by the last element\n", + "# of the tuple and don't care about a \"tie\" situation, we can also use\n", + "# the index of the tuple item directly instead of reversing the tuple \n", + "# for efficiency.\n", + "\n", + "a_list = [(1,3,'c'), (2,3,'a'), (3,2,'b'), (2,2,'b')]\n", + "\n", + "sorted_list = sorted(a_list, key=lambda e: e[-1])\n", + "\n", + "print(sorted_list)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Sorting multiple lists relative to each other" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[back to top](#Table-of-Contents)" + ] + }, + { + "cell_type": "code", + "execution_count": 49, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "input values:\n", + " ['c', 'b', 'a'] [6, 5, 4] ['some-val-associated-with-c', 'another_val-b', 'z_another_third_val-a']\n", + "\n", + "\n", + "sorted output:\n", + " ['a', 'b', 'c'] [4, 5, 6] ['z_another_third_val-a', 'another_val-b', 'some-val-associated-with-c']\n" + ] + } + ], + "source": [ + "\"\"\"\n", + "You have 3 lists that you want to sort \"relative\" to each other,\n", + "for example, picturing each list as a row in a 3x3 matrix: sort it by columns\n", + "\n", + "########################\n", + "If the input lists are\n", + "########################\n", + "\n", + " list1 = ['c','b','a']\n", + " list2 = [6,5,4]\n", + " list3 = ['some-val-associated-with-c','another_val-b','z_another_third_val-a']\n", + "\n", + "########################\n", + "the desired outcome is:\n", + "########################\n", + "\n", + " ['a', 'b', 'c'] \n", + " [4, 5, 6] \n", + " ['z_another_third_val-a', 'another_val-b', 'some-val-associated-with-c']\n", + "\n", + "########################\n", + "and NOT:\n", + "########################\n", + "\n", + " ['a', 'b', 'c'] \n", + " [4, 5, 6] \n", + " ['another_val-b', 'some-val-associated-with-c', 'z_another_third_val-a']\n", + "\n", + "\n", + "\"\"\"\n", + "\n", + "list1 = ['c','b','a']\n", + "list2 = [6,5,4]\n", + "list3 = ['some-val-associated-with-c','another_val-b','z_another_third_val-a']\n", + "\n", + "print('input values:\\n', list1, list2, list3)\n", + "\n", + "list1, list2, list3 = [list(t) for t in zip(*sorted(zip(list1, list2, list3)))]\n", + "\n", + "print('\\n\\nsorted output:\\n', list1, list2, list3 )" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Using namedtuples" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[back to top](#Table-of-Contents)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "`namedtuples` are high-performance container datatypes in the [`collection`](https://docs.python.org/2/library/collections.html) module (part of Python's stdlib since 2.6).\n", + "`namedtuple()` is factory function for creating tuple subclasses with named fields." + ] + }, + { + "cell_type": "code", + "execution_count": 1, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "X-coordinate: 1\n" + ] + } + ], + "source": [ + "from collections import namedtuple\n", + "\n", + "Coordinates = namedtuple('Coordinates', ['x', 'y', 'z'])\n", + "point1 = Coordinates(1, 2, 3)\n", + "print('X-coordinate: %d' % point1.x)" + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": { + "collapsed": true + }, + "outputs": [], + "source": [] + } + ], + "metadata": { + "kernelspec": { + "display_name": "Python 3", + "language": "python", + "name": "python3" + }, + "language_info": { + "codemirror_mode": { + "name": "ipython", + "version": 3 + }, + "file_extension": ".py", + "mimetype": "text/x-python", + "name": "python", + "nbconvert_exporter": "python", + "pygments_lexer": "ipython3", + "version": "3.4.3" + } + }, + "nbformat": 4, + "nbformat_minor": 0 +} diff --git a/python_true_false.html b/python_true_false.html deleted file mode 100644 index 079d7cb..0000000 --- a/python_true_false.html +++ /dev/null @@ -1,2297 +0,0 @@ - - - - -
Sebastian Raschka, 03/2014
Code was executed in Python 3.4.0
True and False in the datetime module
-Pointed out in a nice article "A false midnight" at http://lwn.net/SubscriberLink/590299/bf73fe823974acea/:
-"it often comes as a big surprise for programmers to find (sometimes by way of a hard-to-reproduce bug) that,
unlike any other time value, midnight (i.e. datetime.time(0,0,0)) is False.
A long discussion on the python-ideas mailing list shows that, while surprising,
that behavior is desirable—at least in some quarters."
import datetime
-
-print('"datetime.time(0,0,0)" (Midnight) evaluates to', bool(datetime.time(0,0,0)))
-
-print('"datetime.time(1,0,0)" (1 am) evaluates to', bool(datetime.time(1,0,0)))
-Boolean True
-my_true_val = True
-
-
-print('my_true_val == True:', my_true_val == True)
-print('my_true_val is True:', my_true_val is True)
-
-print('my_true_val == None:', my_true_val == None)
-print('my_true_val is None:', my_true_val is None)
-
-print('my_true_val == False:', my_true_val == False)
-print('my_true_val is False:', my_true_val is False)
-
-print(my_true_val
-if my_true_val:
- print('"if my_true_val:" is True')
-else:
- print('"if my_true_val:" is False')
-
-if not my_true_val:
- print('"if not my_true_val:" is True')
-else:
- print('"if not my_true_val:" is False')
-Boolean False
-my_false_val = False
-
-
-print('my_false_val == True:', my_false_val == True)
-print('my_false_val is True:', my_false_val is True)
-
-print('my_false_val == None:', my_false_val == None)
-print('my_false_val is None:', my_false_val is None)
-
-print('my_false_val == False:', my_false_val == False)
-print('my_false_val is False:', my_false_val is False)
-
-
-if my_false_val:
- print('"if my_false_val:" is True')
-else:
- print('"if my_false_val:" is False')
-
-if not my_false_val:
- print('"if not my_false_val:" is True')
-else:
- print('"if not my_false_val:" is False')
-None 'value'
-my_none_var = None
-
-print('my_none_var == True:', my_none_var == True)
-print('my_none_var is True:', my_none_var is True)
-
-print('my_none_var == None:', my_none_var == None)
-print('my_none_var is None:', my_none_var is None)
-
-print('my_none_var == False:', my_none_var == False)
-print('my_none_var is False:', my_none_var is False)
-
-
-if my_none_var:
- print('"if my_none_var:" is True')
-else:
- print('"if my_none_var:" is False')
-
-if not my_none_var:
- print('"if not my_none_var:" is True')
-else:
- print('"if not my_none_var:" is False')
-Empty String
-my_empty_string = ""
-
-print('my_empty_string == True:', my_empty_string == True)
-print('my_empty_string is True:', my_empty_string is True)
-
-print('my_empty_string == None:', my_empty_string == None)
-print('my_empty_string is None:', my_empty_string is None)
-
-print('my_empty_string == False:', my_empty_string == False)
-print('my_empty_string is False:', my_empty_string is False)
-
-
-if my_empty_string:
- print('"if my_empty_string:" is True')
-else:
- print('"if my_empty_string:" is False')
-
-if not my_empty_string:
- print('"if not my_empty_string:" is True')
-else:
- print('"if not my_empty_string:" is False')
-Empty List
-It is generally not a good idea to use the == to check for empty lists...
my_empty_list = []
-
-
-print('my_empty_list == True:', my_empty_list == True)
-print('my_empty_list is True:', my_empty_list is True)
-
-print('my_empty_list == None:', my_empty_list == None)
-print('my_empty_list is None:', my_empty_list is None)
-
-print('my_empty_list == False:', my_empty_list == False)
-print('my_empty_list is False:', my_empty_list is False)
-
-
-if my_empty_list:
- print('"if my_empty_list:" is True')
-else:
- print('"if my_empty_list:" is False')
-
-if not my_empty_list:
- print('"if not my_empty_list:" is True')
-else:
- print('"if not my_empty_list:" is False')
-
-
-
-[0]-List
-my_zero_list = [0]
-
-
-print('my_zero_list == True:', my_zero_list == True)
-print('my_zero_list is True:', my_zero_list is True)
-
-print('my_zero_list == None:', my_zero_list == None)
-print('my_zero_list is None:', my_zero_list is None)
-
-print('my_zero_list == False:', my_zero_list == False)
-print('my_zero_list is False:', my_zero_list is False)
-
-
-if my_zero_list:
- print('"if my_zero_list:" is True')
-else:
- print('"if my_zero_list:" is False')
-
-if not my_zero_list:
- print('"if not my_zero_list:" is True')
-else:
- print('"if not my_zero_list:" is False')
-List comparison
-List comparisons are a handy way to show the difference between == and is.
While == is rather evaluating the equality of the value, is is checking if two objects are equal. The examples below show that we can assign a pointer to the same list object by using =, e.g., list1 = list2.
a) If we want to make a shallow copy of the list values, we have to make a little tweak: list1 = list2[:], or
b) a deepcopy via list1 = copy.deepcopy(list2)
Possibly the best explanation of shallow vs. deep copies I've read so far:
-*** "Shallow copies duplicate as little as possible. A shallow copy of a collection is a copy of the collection structure, not the elements. With a shallow copy, two collections now share the individual elements. Deep copies duplicate everything. A deep copy of a collection is two collections with all of the elements in the original collection duplicated."***
-(via S.Lott on StackOverflow)
-a) Shallow vs. deep copies for simple elements
-List modification of the original list doesn't affect
shallow copies or deep copies if the list contains literals.
from copy import deepcopy
-
-my_first_list = [1]
-my_second_list = [1]
-print('my_first_list == my_second_list:', my_first_list == my_second_list)
-print('my_first_list is my_second_list:', my_first_list is my_second_list)
-
-my_third_list = my_first_list
-print('my_first_list == my_third_list:', my_first_list == my_third_list)
-print('my_first_list is my_third_list:', my_first_list is my_third_list)
-
-my_shallow_copy = my_first_list[:]
-print('my_first_list == my_shallow_copy:', my_first_list == my_shallow_copy)
-print('my_first_list is my_shallow_copy:', my_first_list is my_shallow_copy)
-
-my_deep_copy = deepcopy(my_first_list)
-print('my_first_list == my_deep_copy:', my_first_list == my_deep_copy)
-print('my_first_list is my_deep_copy:', my_first_list is my_deep_copy)
-
-print('\nmy_third_list:', my_third_list)
-print('my_shallow_copy:', my_shallow_copy)
-print('my_deep_copy:', my_deep_copy)
-
-my_first_list[0] = 2
-print('after setting "my_first_list[0] = 2"')
-print('my_third_list:', my_third_list)
-print('my_shallow_copy:', my_shallow_copy)
-print('my_deep_copy:', my_deep_copy)
-b) Shallow vs. deep copies if list contains other structures and objects
-List modification of the original list does affect
shallow copies, but not deep copies if the list contains compound objects.
my_first_list = [[1],[2]]
-my_second_list = [[1],[2]]
-print('my_first_list == my_second_list:', my_first_list == my_second_list)
-print('my_first_list is my_second_list:', my_first_list is my_second_list)
-
-my_third_list = my_first_list
-print('my_first_list == my_third_list:', my_first_list == my_third_list)
-print('my_first_list is my_third_list:', my_first_list is my_third_list)
-
-my_shallow_copy = my_first_list[:]
-print('my_first_list == my_shallow_copy:', my_first_list == my_shallow_copy)
-print('my_first_list is my_shallow_copy:', my_first_list is my_shallow_copy)
-
-my_deep_copy = deepcopy(my_first_list)
-print('my_first_list == my_deep_copy:', my_first_list == my_deep_copy)
-print('my_first_list is my_deep_copy:', my_first_list is my_deep_copy)
-
-print('\nmy_third_list:', my_third_list)
-print('my_shallow_copy:', my_shallow_copy)
-print('my_deep_copy:', my_deep_copy)
-
-my_first_list[0][0] = 2
-print('after setting "my_first_list[0][0] = 2"')
-print('my_third_list:', my_third_list)
-print('my_shallow_copy:', my_shallow_copy)
-print('my_deep_copy:', my_deep_copy)
-Some Python oddity:¶
-a = 1
-b = 1
-print('a is b', bool(a is b))
-True
-
-a = 999
-b = 999
-print('a is b', bool(a is b))
--# else inserts new record an sets purchasable to 0 -c.execute("""INSERT OR REPLACE INTO zinc_db1 (zinc_id, rotatable_bonds, purchasable) - VALUES ( 'ZINC123456798', - 3, - COALESCE((SELECT purchasable from zinc_db1 WHERE zinc_id = 'ZINC123456798'), 0) - )""") - - - -# delete rows -t = ('NO', ) -c.execute("DELETE FROM zinc_db1 WHERE purchasable=?", t) -print "Total number of rows deleted: ", conn.total_changes - -# add column -c.execute("ALTER TABLE zinc_db1 ADD COLUMN 'keto_oxy' TEXT") - -# save changes -conn.commit() - -# print column names -c.execute("SELECT * FROM zinc_db1") -col_name_list = [tup[0] for tup in c.description] -print col_name_list - - - -# close connection -conn.close() diff --git a/templates/webapp_ex1/README.md b/templates/webapp_ex1/README.md new file mode 100644 index 0000000..d07794d --- /dev/null +++ b/templates/webapp_ex1/README.md @@ -0,0 +1,13 @@ +Sebastian Raschka, 2015 + +# Flask Example App 1 + +A simple Flask app that calculates the sum of two numbers entered in the respective input fields. + +A more detailed description is going to follow some time in future. + +You can run the app locally by executing `python app.py` within this directory. + +
+ + diff --git a/templates/webapp_ex1/app.py b/templates/webapp_ex1/app.py new file mode 100644 index 0000000..5314e37 --- /dev/null +++ b/templates/webapp_ex1/app.py @@ -0,0 +1,31 @@ +from flask import Flask, render_template, request +from wtforms import Form, DecimalField, validators + +app = Flask(__name__) + + +class EntryForm(Form): + x_entry = DecimalField('x:', + places=10, + validators=[validators.NumberRange(-1e10, 1e10)]) + y_entry = DecimalField('y:', + places=10, + validators=[validators.NumberRange(-1e10, 1e10)]) + +@app.route('/') +def index(): + form = EntryForm(request.form) + return render_template('entry.html', form=form, z='') + +@app.route('/results', methods=['POST']) +def results(): + form = EntryForm(request.form) + z = '' + if request.method == 'POST' and form.validate(): + x = request.form['x_entry'] + y = request.form['y_entry'] + z = float(x) + float(y) + return render_template('entry.html', form=form, z=z) + +if __name__ == '__main__': + app.run(debug=True) \ No newline at end of file diff --git a/templates/webapp_ex1/img/img_1.png b/templates/webapp_ex1/img/img_1.png new file mode 100644 index 0000000..bfc3510 Binary files /dev/null and b/templates/webapp_ex1/img/img_1.png differ diff --git a/templates/webapp_ex1/static/style.css b/templates/webapp_ex1/static/style.css new file mode 100644 index 0000000..8abda7b --- /dev/null +++ b/templates/webapp_ex1/static/style.css @@ -0,0 +1,7 @@ +body{ + width:600px; +} + +#button{ + padding-top: 20px; +} \ No newline at end of file diff --git a/templates/webapp_ex1/templates/_formhelpers.html b/templates/webapp_ex1/templates/_formhelpers.html new file mode 100644 index 0000000..5790894 --- /dev/null +++ b/templates/webapp_ex1/templates/_formhelpers.html @@ -0,0 +1,12 @@ +{% macro render_field(field) %} +
-
+ {% for error in field.errors %}
+
- {{ error }} + {% endfor %} +
\n", + "I would be happy to hear your comments and suggestions. \n", + "Please feel free to drop me a note via\n", + "[twitter](https://twitter.com/rasbt), [email](mailto:bluewoodtree@gmail.com), or [google+](https://plus.google.com/+SebastianRaschka).\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 1, + "metadata": {}, + "source": [ + "Awesome things that you can do in IPython Notebooks (in progress)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 2, + "metadata": {}, + "source": [ + "Writing local files" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%%file hello.py\n", + "def func_inside_script(x, y):\n", + " return x + y\n", + "print('Hello World')" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "Writing hello.py\n" + ] + } + ], + "prompt_number": 13 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 2, + "metadata": {}, + "source": [ + "Running Python scripts" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "We can run Python scripts in IPython via the %run magic command. For example, the Python script that we created in the [Writing local files](#Writing-local-files) section." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%run hello.py" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "Hello World\n" + ] + } + ], + "prompt_number": 14 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "func_inside_script(1, 2)" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "pyout", + "prompt_number": 16, + "text": [ + "3" + ] + } + ], + "prompt_number": 16 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 2, + "metadata": {}, + "source": [ + "Benchmarking" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%timeit [x**2 for x in range(100)] " + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "10000 loops, best of 3: 38.8 \u00b5s per loop\n" + ] + } + ], + "prompt_number": 4 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%timeit -r 5 -n 100 [x**2 for x in range(100)] " + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "100 loops, best of 5: 39 \u00b5s per loop\n" + ] + } + ], + "prompt_number": 3 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 2, + "metadata": {}, + "source": [ + "Using system shell commands" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "By prepending a \"`!`\" we can conveniently execute most of the system shell commands, below are just a few examples." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "my_dir = 'new_dir'\n", + "!mkdir $my_dir\n", + "!pwd\n", + "!touch $my_dir'/some.txt'\n", + "!ls -l './new_dir'\n", + "!ls -l $my_dir | wc -l" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "/Users/sebastian/Desktop\r\n" + ] + }, + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "total 0\r\n", + "-rw-r--r-- 1 sebastian staff 0 Jun 27 10:11 some.txt\r\n" + ] + }, + { + "output_type": "stream", + "stream": "stdout", + "text": [ + " 2\r\n" + ] + } + ], + "prompt_number": 12 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 2, + "metadata": {}, + "source": [ + "Debugging" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "def some_func():\n", + " var = 'hello world'\n", + " for i in range(5):\n", + " print(i)\n", + " i / 0\n", + " return 'finished'" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 1 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%debug\n", + "some_func()" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "> \u001b[0;32m
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 2, + "metadata": {}, + "source": [ + "Inline Plotting with matplotlib" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%matplotlib inline" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 33 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "import numpy as np\n", + "from matplotlib import pyplot as plt\n", + "import math\n", + "\n", + "def pdf(x, mu=0, sigma=1):\n", + " \"\"\"Calculates the normal distribution's probability density \n", + " function (PDF). \n", + " \n", + " \"\"\"\n", + " term1 = 1.0 / ( math.sqrt(2*np.pi) * sigma )\n", + " term2 = np.exp( -0.5 * ( (x-mu)/sigma )**2 )\n", + " return term1 * term2\n", + "\n", + "\n", + "x = np.arange(0, 100, 0.05)\n", + "\n", + "pdf1 = pdf(x, mu=5, sigma=2.5**0.5)\n", + "pdf2 = pdf(x, mu=10, sigma=6**0.5)\n", + "\n", + "plt.plot(x, pdf1)\n", + "plt.plot(x, pdf2)\n", + "plt.title('Probability Density Functions')\n", + "plt.ylabel('p(x)')\n", + "plt.xlabel('random variable x')\n", + "plt.legend(['pdf1 ~ N($\\mu=5$, $\\sigma=2.5$)', 'pdf2 ~ N($\\mu=10$, $\\sigma=6$)'], loc='upper right')\n", + "plt.ylim([0,0.5])\n", + "plt.xlim([0,20])\n", + "\n", + "plt.show()" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "display_data", + "png": "iVBORw0KGgoAAAANSUhEUgAAAYQAAAEZCAYAAACXRVJOAAAABHNCSVQICAgIfAhkiAAAAAlwSFlz\nAAALEgAACxIB0t1+/AAAIABJREFUeJzs3XlcVNX/x/HXICCioLihgIigoqKCu4YLiooram64L+Q3\nt1LLtNytLOunlYb1NXM3l8wFLUVzQa3ctVJx30DcAQUXEIb7+2N0viIoizPcGfg8e8yjuTN37n3P\nMN7PnHPvPVejKIqCEEKIfM9C7QBCCCFMgxQEIYQQgBQEIYQQT0lBEEIIAUhBEEII8ZQUBCGEEIAU\nBJEDFhYWXLp0KUevdXNzY+fOnRk+t2/fPqpUqZJm3l27dgHw2WefMWTIkByt05S0a9eO5cuXqx3D\nYOzs7Lhy5YraMYSBSEHIJ9zc3LC1tcXOzo4yZcowaNAgHj58mOs5NBoNGo0mw+eaNGnCmTNn0sz7\nzIQJE1iwYAEAV65cwcLCgtTU1BxlWLJkCQUKFMDOzg47Ozvc3d0ZPHgw58+fz9HysmPLli3069dP\nn6NJkyY5XtbAgQMpWLCg/n3Y2dmxdu1aQ0VNx8/Pj4ULF6Z5LCEhATc3N6OtU+QuKQj5hEaj4ddf\nfyUhIYFjx45x5MgRPv3003TzpaSkqJAuZ17nnEpfX18SEhKIj49nx44dFCpUiDp16nDq1CkDJjQu\njUbD+PHjSUhI0N+6d+9u1PWJvE0KQj7k5OREmzZt9Bs/CwsLvvvuOypVqoSnpycACxYsoFKlSpQo\nUYJOnTpx48aNNMv47bff8PDwoFSpUowbN06/cb548SItWrSgZMmSlCpVir59+3L//v00rz106BBe\nXl4UL16cwYMHk5SUBEB4eDjlypXLMPO0adP0v6ybNm0KQLFixbC3t2fv3r2UKFGCkydP6ue/ffs2\nhQsXJiYmJsPlPcur0Whwd3dn3rx5NGvWjGnTpunnOXDgAG+88QYODg74+PiwZ88e/XN+fn5MmTKF\nxo0bY29vT0BAgH5diYmJ9O3bl5IlS+Lg4ED9+vW5c+eO/nULFy7kzJkzDB06lP3792NnZ0fx4sU5\ncuQIjo6OaQrd+vXr8fHxyfA9vMzAgQOZPHmyfvrFz9XNzY3Zs2fj7e1NsWLFCAoK0v8NAEJDQ/Hx\n8aFo0aJUrFiRbdu2MXHiRPbt28fIkSOxs7Pj3XffBdJ2H96/f5/+/ftTunRp3NzcmDFjhv69LFmy\nhMaNG/PBBx9QvHhx3N3dCQsL069zyZIleHh4YG9vj7u7OytXrszWexaGIQUhH3n2jzMqKoqtW7dS\nq1Yt/XOhoaEcPnyYiIgIdu3axYQJE1i7di03btygfPnyBAUFpVnWxo0bOXr0KMeOHSM0NJRFixbp\nn5s4cSI3btzg9OnTREVFpdnIKorCypUr2b59OxcvXuTcuXMZtlRe9Pyv03379gG6DVB8fDxNmzYl\nKCiIFStW6OdZtWoVLVu2pESJEln+fN588039sqOjo+nQoQNTpkwhLi6OWbNm0bVr1zQFZtWqVSxZ\nsoTbt2/z5MkTZs2aBcDSpUuJj4/n2rVrxMbGMn/+fGxsbPTvQ6PRUKVKFebPn0+jRo1ISEggNjaW\nunXrUrJkSbZt26Zfx/LlyxkwYMBLM2fUSnpVt9yz59euXcu2bdu4fPky//77L0uWLAF0xXrAgAHM\nnj2b+/fvs3fvXv3GvUmTJsybN4+EhATmzp2bbrnvvPMOCQkJXL58mT179rBs2TIWL16sf/7QoUNU\nqVKFmJgYxo0bR3BwMAAPHz5k1KhRhIWFER8fz/79+7NdBIVhSEHIJxRFoXPnzjg4ONCkSRP8/PyY\nMGGC/vmPPvqIYsWKUbBgQX766SeCg4Px8fHB2tqazz//nP379xMZGamff/z48RQrVoxy5coxevRo\nVq1aBYCHhwf+/v5YWVlRsmRJxowZk+aXtUajYeTIkTg7O+Pg4MDEiRP1r80sf0b3n+nfv3+a5Sxf\nvlzfosiqsmXLEhsbC8CKFSto164dbdq0AaBly5bUrVuX3377Tf8+Bg0aRMWKFbGxsaFHjx78/fff\nAFhbWxMTE8P58+fRaDTUqlULOzu7V76n59/Hs8IWGxvL9u3b6d27d4Z5FUVh1qxZODg44ODgQOnS\npfWPZ9ad9u6771KmTBkcHBzo2LGjPvvChQsJDg7G398f0LUmn7UaX5YZQKvVsmbNGj7//HMKFy5M\n+fLlef/999PsQC9fvjzBwcFoNBr69+/PjRs3uH37NqBraZw4cYLHjx/j6OhItWrVXplfGIcUhHxC\no9EQGhpKXFwcV65cISQkhIIFC+qff75L4Vmr4JnChQtTokQJoqOjM5zf1dWV69evA3Dr1i2CgoJw\ncXGhaNGi9OvXL123zcte+zoaNGhAoUKFCA8P58yZM1y8eJHAwMBsLSM6Olrforh69Spr167Vb2wd\nHBz4888/uXnzpn7+MmXK6O8XKlSIBw8eANCvXz8CAgIICgrC2dmZ8ePHZ3nfTJ8+fdi8eTOPHj3i\n559/pmnTpjg6OmY4r0aj4YMPPiAuLo64uDj9xjUrff0vZn92gMG1a9fw8PB46etetuy7d++SnJyc\n5nvj6uqa5jvz/DptbW0BePDgAYULF2bNmjX897//xcnJiQ4dOnD27NlM34MwPCkIAkj7D93JySnN\noYQPHz4kJiYGZ2dn/WPPtxYiIyP1z02YMIECBQpw8uRJ7t+/z/Lly9MdDfTia52cnHKc9XkDBgxg\nxYoVLF++nO7du2NtbZ2t5W7YsEF/1I+rqyv9+vXTb2zj4uJISEhg3LhxmS7H0tKSKVOmcOrUKf76\n6y9+/fVXli1blqX34eLiQsOGDVm/fj0rVqzItJWT0S/2woUL8+jRI/3080UsM+XKlePChQsZPveq\nQlOyZEmsrKzSfG8iIyNxcXHJ0npbt27N9u3buXnzJlWqVMkThxibIykIIp1evXqxePFi/vnnH5KS\nkpgwYQINGzbE1dVVP8+sWbO4d+8eUVFRzJ07l549ewL/+8Vnb29PdHQ0//d//5dm2YqiMG/ePKKj\no4mNjWXGjBnp9k9kplSpUlhYWHDx4sU0j/ft25f169fz008/0b9//ywtS6vVcvnyZd555x327t3L\n1KlT9cvavHkz27dvR6vVkpiYSHh4eJpfvC/rPtm9ezcnTpxAq9ViZ2eHlZUVBQoUSDefo6Mj165d\nIzk5Oc3j/fv354svvuDkyZO8+eabL83+svX7+PiwZcsW4uLiuHnzJt98802mn8OzZQUHB7N48WJ2\n7dpFamoq0dHR+l/rjo6O6T7zZwoUKECPHj2YOHEiDx484OrVq3z99df07ds303Xfvn2b0NBQHj58\niJWVFYULF87w8xLGJwVBpPvl5+/vzyeffELXrl1xcnLi8uXLrF69Os08nTp1ok6dOtSqVYsOHTow\nePBgAKZOncqxY8coWrQoHTt2pGvXrmmWr9Fo6NOnD61bt8bDw4NKlSoxadKkl2Z5/vFnz9na2jJx\n4kR8fX1xcHDg0KFDgO7Xbe3atbGwsKBx48avfL/Pju4pWrQozZs358GDBxw+fBgvLy9A90s9NDSU\nzz77jNKlS+Pq6srs2bPTbIRffF/Ppm/dukX37t0pWrQo1apVw8/PL8Nf+v7+/nh5eVGmTBl9/z/o\ndm5HRkbSpUsX/c7ozD6T5/Xr1w9vb2/c3Nxo06YNQUFBme5kfvZ8vXr1WLx4MWPGjKFYsWL4+fnp\nW3SjRo3il19+oXjx4owePTrdcr799lsKFy6Mu7s7TZo0oU+fPgwaNOilWZ9Np6am8vXXX+Ps7EyJ\nEiXYt28f33///UvzCuPRGPMCOWFhYYwePRqtVstbb73F+PHj0zwfHh5Op06dcHd3B6Br165pNg5C\nZFdwcDDOzs58/PHHakd5LZUqVWL+/Pm0aNFC7SgiH7E01oK1Wi0jR45kx44dODs7U69ePQIDA6la\ntWqa+Zo1a8amTZuMFUPkI1euXGH9+vX6I2bM1fr169FoNFIMRK4zWpfRoUOHqFixIm5ublhZWREU\nFERoaGi6+eQKnsIQJk+eTI0aNRg3blyaI13MjZ+fH8OHD2fevHlqRxH5kNFaCNHR0WkOL3RxceHg\nwYNp5tFoNPz11194e3vj7OzMrFmz5PhjkSOffPIJn3zyidoxXlt4eLjaEUQ+ZrSCkJVjoWvXrk1U\nVBS2trZs3bqVzp07c+7cOWNFEkII8QpGKwjOzs5ERUXpp6OiotIdk/z82Ztt27Zl+PDhxMbGUrx4\n8TTzVaxY8aWHuwkhhMiYh4fHS88ryZBiJMnJyYq7u7ty+fJlJSkpSfH29lYiIiLSzHPz5k0lNTVV\nURRFOXjwoFK+fPkMl2XEmPnS1KlT1Y6QZ8hnaVjyeRpWdredRmshWFpaEhISQkBAAFqtluDgYKpW\nrcr8+fMBePvtt/nll1/4/vvvsbS0xNbWNt2x7kIIIXKP0QoC6LqB2rZtm+axt99+W39/xIgRjBgx\nwpgRhBBCZJGcqZwP+fn5qR0hz5DP0rDk81SXUc9UNhSNRiPnKwghRDZld9tp1C4jIcxJ8eLFiYuL\nUzuGENnm4OCgv5bH65AWghBPyfdMmKuXfXez+52WfQhCCCEAKQhCCCGekoIghBACkIIghBDiKSkI\nQgghACkIQuRJAwcOZPLkyfrps2fP4uPjg729PSEhISomS++jjz5izpw5ascwWQ0aNCAiIiJX1iUF\nQYg86MVrGH/55Zf4+/sTHx/PyJEjCQkJoW7dutjY2Oive2xobm5uODo68ujRI/1jP/74I82bN9dP\n37lzh+XLlzN06FCjZPDw8KBgwYI4OjqybNkyo6wjMytXrmT27Nn07NnzleO1vSzr2LFjmTJlSm5E\nlRPThMirnj/+/OrVq7zxxhv6aWdnZyZPnsy2bdt4/Pix0TKkpqYyZ84cPvroowyfX7JkCe3bt6dg\nwYJGWf+HH35IQEAATk5OWFrm/ubuwoULxMTE8P7773P37l0qVapEgwYNqFChQpazduzYkaFDh3Lr\n1i0cHR2NmldaCEKYCTc3N2bOnImXlxfFixdn8ODBJCUlAXD8+HFq166Nvb09QUFBJCYm6lsILVq0\nIDw8nJEjR2Jvb8+FCxfo0qULnTp1okSJEkbLq9FoGDt2LLNmzeL+/fsZzhMWFkazZs2MlsHa2hpX\nV1dVigHAqVOn+PLLLwEoWbIkFStW5OjRoxnO+7KsNjY21KlTh23bthk9r7QQhDAjK1euZPv27dja\n2tKxY0c+/fRTpkyZQufOnXnvvfcYOXIkGzdupFevXnz44YcA7Nq1i+bNm9OvXz8GDx6cZnnGPjO7\nbt26+Pn5MWvWrAwvcXrixAk8PT2ztcxLly6xYMGClz7fsGFDOnXqBMDhw4dJSkoiPj6eypUrExgY\nmL038JoZ2rVrx9atWwHdZ33jxg0qVqyY4WtelbVq1ar8888/Bsn+Soa7FIPxmElMYeYy+56BYW45\n5ebmpsyfP18/vWXLFsXDw0PZu3ev4uTklGbeN954Q5k8ebJ+2s/PT/nxxx/TLXPSpEnKwIEDX7ne\nP//8U/Hz81NKly6ttGrVSvn555+Vu3fvKnv27FGGDx/+yrw7d+5UTp48qRQtWlS5c+eOsmDBAsXP\nz08/j5WVlXL27Nk0r0tJSVF8fX3104MHD1bOnz//yowvs379ev19b29vJS4uLkuvO3v2rNK9e3fF\nz89PKVKkiNKhQwfl+++/z1GGZzZv3qx06tQpR1knTpyoDB48+KWvfdl3N7vbTukyEiKLDFUSXke5\ncuX0911dXbl+/TrXr1/H2dk5zXzly5dP9+s/o+ucvzhPRlavXs3XX3/N9evXGTVqFIsWLaJKlSrM\nmDGD//znP5m+3svLiw4dOjBz5sx0GRwcHEhISEjz2P79+ylfvrw+3/79+1/6qzozz1oKz9YVHh6e\n6WtiY2MZOnQoy5YtY/fu3fj7+7NixYrX2vF97949lixZwooVK3KUNT4+HgcHhxyvP6uky0gIMxIZ\nGZnmvpOTE2XLliU6OjrNfFevXs3SRjSjIvGiOXPm6Odr37497du3z2ZqmD59OrVr1+b9999P83jN\nmjU5e/YsderU0T8WFhZGQEAAoNs3UqNGjTSvyWp3zYoVK9i0aRM///wzAA8fPszSvoR58+YxYsQI\nbGxsAEhKSsLW1jZHGUBX1GbOnMmPP/5IkSJFuHr1qr7gPZNZ1tOnT9O/f/9Ms78uKQhCmAlFUfju\nu+/o0KEDhQoVYsaMGQQFBdGoUSMsLS2ZO3cuw4YNY/PmzRw+fBh/f/90r39Gq9WSnJxMSkoKWq2W\npKQkLC0tKVCgQLr1ZqVoZMbDw4OePXsyZ84catasqX+8Xbt27Nmzh969e+sf27ZtG0FBQQD89ttv\n+Pv7s2nTJn2furu7O59//nmm63Rzc9P/qn/06BF37tyhRYsWgO48DY1Gw+LFi9O9LiEhgWrVqgG6\nncJeXl5YWVmlmSerGQC+/fZbunfvTmJiIocOHeLx48eUL1+eixcv4u7ujkajeWXWxMREjh07xvLl\ny7O0vtchXUZCmAmNRkPv3r1p3bo1Hh4eVKpUiUmTJmFlZcX69etZsmQJJUqU4Oeff6Zr164Zvv6Z\nTz75BFtbW7744gtWrFihLzDGNGXKFB49epQmR//+/dmyZQuJiYmA7ryEyMhINm3axJYtW7C1teXO\nnTvpfqFnRePGjblx4wbffPMNEydOZPXq1frlXLt2jcaNG2f4umHDhrF9+3bWrVvHjh07mDlzZg7e\nrc4ff/zBmDFjqFevHk5OTjRq1EjfcuvevTt///13plk3b95M8+bNKVOmTI5zZJVcD0GIp0z9e1ah\nQgUWLlyo/+WYV0ycOJHSpUszatQoVqxYwenTp41anJ48eUKtWrX4999/M2wRmZqGDRuyaNEifasl\nI4a6HoJ0GQkhVPX8xv/QoUMMGDDAqOuztrbm1KlTRl2HIR04cCDX1iUFQQhhMubOnat2hHxNuoyE\neEq+Z8JcySU0hRBCGJQUBCGEEIAUBCGEEE9JQRBCCAFIQRBCCPGUFAQhhBCAFAQhhBBPSUEQQggB\nSEEQQgjxlBQEIfKggQMHMnnyZP302bNn8fHxwd7enpCQEBWTpffRRx8xZ84ctWOYjAYNGhAREaHK\nuqUgCJEHaTSaNMNMf/nll/j7+xMfH89//vMfgoODcXNzw97enlq1ahEWFmbwDG5ubjg6OvLo0SP9\nYz/++CPNmzfXT9+5c4fly5e/1tXIMvP3338zduzYNI9t3LiRzz77jJkzZ+bKdQYy8uDBA6ZMmcKC\nBQuYPXu2foiJsWPHMmXKFFUyyeB2QuRRz49hc/XqVd544w0AkpOTcXV1Ze/evbi6uvLbb7/Ro0cP\nTpw4ke5KXq8rNTWVOXPm8NFHH2X4/JIlS2jfvj0FCxY06Hqf+eqrr/jjjz8oWrSo/rH79+/zySef\ncPToUQAaNWpE27ZtKVmypFEyvMy7777L1KlTKV++PF5eXnTr1o3y5cvTsWNHhg4dyq1bt3B0dMzV\nTNJCEMJMuLm5MXPmTLy8vChevDiDBw8mKSkJ0F1qsnbt2tjb2xMUFERiYqK+hdCiRQvCw8MZOXIk\n9vb23Lhxg6lTp+Lq6groLotZoUIFjh07ZtC8Go2GsWPHMmvWLO7fv5/hPGFhYTRr1syg633ee++9\nl+ZaxQB79+5Nc20Bb29vdu/ebbQMGbl06RLXr1/XF+Dt27fr79vY2FCnTh22bduWq5nAyC2EsLAw\nRo8ejVar5a233mL8+PEZznf48GEaNWrEzz//zJtvvmnMSEKYtZUrV7J9+3ZsbW3p2LEjn376KVOm\nTKFz58689957jBw5ko0bN9KrVy8+/PBDAHbt2kXz5s3p168fgwcPTrfMW7duce7cOby8vAyet27d\nuvj5+TFr1iw++eSTdM+fOHECT0/PbC0zO9czBtKN9nnt2jWKFSumny5WrBjnz5/PVobXzbZr1y6K\nFSvG8uXLuXfvHnZ2dgwcOFA/X9WqVfnnn38Mkik7jFYQtFotI0eOZMeOHTg7O1OvXj0CAwOpWrVq\nuvnGjx9PmzZtZOhhYdI001//2sIAytScfc81Gg0jR47E2dkZ0F1p7J133qF169akpKQwatQoALp2\n7Uq9evXSrzeDf1/Jycn06dOHgQMHUrly5QzX+9dffzFx4kQiIiLw9vZmyJAhtGjRglOnTrFmzRrm\nzZv3yswff/wxvr6++nzPe7YxfJ5Wq6VZs2b88ccfAAQHB/PRRx/pLz2ZnesZP8vw4jptbGz009bW\n1jx48CBLyzp37hyTJk3izp07HDlyBD8/P9q3b6/fB5LVbLdu3eLkyZOsXr0agCZNmuDr60ulSpUA\nsLOz48aNG1nKZEhGKwiHDh2iYsWKuLm5ARAUFERoaGi6gvDtt9/SrVs3Dh8+bKwoQhhETjfkhlSu\nXDn9fVdXV65fv87169f1ReKZ8uXLpysAL24YU1NT6devHzY2Nq888mj16tV8/fXX1KhRg7CwMEJC\nQhg+fDi1a9fmyy+/zDSzl5cXHTp0YObMmen+/Ts4OJCQkJDmsf379+u7TxRFYf/+/fpikBMvfg52\ndnbExMTopx8/fpylvvrY2FiGDh3Kli1bsLGxoXPnzixdujTN/omssre3p0aNGvppV1dXtm/fri8I\n8fHxODg4ZHu5r8toBSE6OjrNl9fFxYWDBw+mmyc0NJRdu3Zx+PDhdF9YIURakZGRae47OTlRtmxZ\noqOj08x39erVV25EFUUhODiYO3fusGXLlldeW3jOnDn6f5vt27enffv22c49ffp0ateuzfvvv5/m\n8Zo1a3L27Fnq1KmjfywsLIyAgABAt2/k+Q0nZL/L6MXtioeHB0eOHNFP3717l9q1a2f6HubNm8eI\nESP0rYukpCRsbW1zlM3Ly4t9+/bpH7ewsCA1NVU/ffr0afr3759pJkMzWkHIysZ99OjRzJw5U39V\nn1d1GU2bNk1/38/PDz8/PwOkFMJ8KIrCd999R4cOHShUqBAzZswgKCiIRo0aYWlpydy5cxk2bBib\nN2/m8OHD+Pv7p3v9M8OGDePMmTPs2LEj0yN8DPFDzcPDg549ezJnzhxq1qypf7xdu3bs2bOH3r17\n6x/btm0bQUFBAPz222/4+/uzadMmAgMDgex3Gb24XWnatCnjxo3TTx87dowvvvgC0J2/odFoWLx4\ncbrlJCQk6HdGnzp1Ci8vL6ysrNLMk9Vsvr6+TJgwQT998eJF/TYuMTGRY8eO5ehw2PDwcMLDw7P9\numeMVhCcnZ2JiorST0dFReHi4pJmnqNHj+r/8Hfv3mXr1q1YWVnp//DPe74gCJEfaTQaevfuTevW\nrbl+/TqdO3dm0qRJWFlZsX79eoYMGcKkSZNo164dXbt2zfD1oGs9/PDDD9jY2FCmTBn98z/88AO9\nevUyWv4pU6awfPnyNAWmf//++Pj4kJiYiI2NDXfu3CEyMpJNmzYRGRmJra0td+7cwd3dPUfrDAkJ\n4eeffyYqKorp06czZswY7O3tGTduHJ9++impqamMGzeO0qVLA7odzi/7DIYNG8amTZuIiIjg2rVr\nzJw5M0eZAAoWLMi0adOYMmUKqampjBgxAg8PDwA2b95M8+bN0/xtsurFH8vTp0/P3gIUI0lOTlbc\n3d2Vy5cvK0lJSYq3t7cSERHx0vkHDhyorFu3LsPnjBhTCD1T/565ubkpO3fuVDuGwU2YMEH55ptv\nFEVRlOXLlysTJkxQJUdSUpJSrVo1JSUlRZX1P9OgQQPl1KlT2XrNy7672f1OG62FYGlpSUhICAEB\nAWi1WoKDg6latSrz588H4O233zbWqoUQZmTGjBn6+4cOHWLAgAGq5LC2tubUqVOqrPt5Bw4cUG3d\nmqdVxKQ928cghDGZ+vesQoUKLFy4kBYtWqgdRZiYl313s/udloIgxFPyPRPmylAFQYauEEIIAUhB\nEEII8ZQUBCGEEIAUBCGEEE9JQRBCCAHIBXKE0HNwcJDxtIRZMtRAeHLYqRBC5FFy2KkQQogckYIg\nhBACkIIghBDiKSkIQgghACkIQgghnpKCIIQQApCCIIQQ4ikpCEIIIQApCEIIIZ6SgiCEEAKQgiCE\nEOIpKQhCCCEAKQhCCCGekoIghBACkIIghBDiKSkIQgghACkIQgghnpKCIIQQApCCIIQQ4ikpCEII\nIQApCEIIIZ6SgiCEEAKQgiCEEOIpKQhCCCEAKQhCCCGekoIghBACkIIghBDiKaMWhLCwMKpUqUKl\nSpX44osv0j0fGhqKt7c3tWrVok6dOuzatcuYcYQQQryCRlEUxRgL1mq1eHp6smPHDpydnalXrx6r\nVq2iatWq+nkePnxI4cKFAThx4gRdunThwoUL6UNqNBgpphBC5FnZ3XYarYVw6NAhKlasiJubG1ZW\nVgQFBREaGppmnmfFAODBgweULFnSWHGEEEJkwmgFITo6mnLlyumnXVxciI6OTjffxo0bqVq1Km3b\ntmXu3LnGiiOEECITRisIGo0mS/N17tyZ06dPs3nzZvr162esOEIIITJhaawFOzs7ExUVpZ+OiorC\nxcXlpfM3adKElJQUYmJiKFGiRLrnp02bpr/v5+eHn5+fIeMKIYTZCw8PJzw8PMevN9pO5ZSUFDw9\nPdm5cydOTk7Ur18/3U7lixcv4u7ujkaj4dixY3Tv3p2LFy+mDyk7lYUQItuyu+00WgvB0tKSkJAQ\nAgIC0Gq1BAcHU7VqVebPnw/A22+/zbp161i2bBlWVlYUKVKE1atXGyuOEEKITBithWBI0kIQQojs\nM5nDToUQQpgXKQhCCCEAKQhCCCGekoIghBACkIIghBDiKSkIQgghACkIQgghnpKCIIQQApCCIIQQ\n4ikpCEIIIYBsFITExESSkpKMmUUIIYSKXloQUlNTWb9+Pd27d8fZ2ZkKFSpQvnx5nJ2d6datGxs2\nbJDxhYQQIg956eB2TZs2pUmTJgQGBuLj40PBggUBSEpK4vjx42zatIk//viDvXv3Gj+kDG4nhBDZ\nlt1t50sVoc7yAAAgAElEQVQLQlJSkr4IvExW5jEEKQhCCJF9Bhvt9NmGfseOHemeW7p0aZp5hBBC\nmL9MdypPnz6dYcOG8fDhQ27evEnHjh3ZtGlTbmQTQgiRizItCHv27MHd3R1vb2+aNGlCr169WLdu\nXW5kEwaUkgKHD8P69bBrFzx8qHYiIYSpybQgxMXFcfjwYTw8PLC2tiYyMlL6883I7dswejSUKgWD\nB8OyZTBlCpQpA506wT//qJ1QCGEqMi0IjRo1IiAggG3btnH48GGio6Px9fXNjWziNa1bB9WqgVYL\nJ07obhs3wh9/wM2b4O8PAQHw4Ye6FoQQIn/L9JrKV69epXz58mke27NnD82aNTNqsOfJUUbZoygw\ndaquNbBuHdSp8/J579yBXr3AwkLXnVSkSO7lFEIYl8EOO7148SIeHh6vfHFW5jEEKQjZM3UqbNgA\nO3fquooyk5ICQ4ZAZCT8+isUKmT8jEII4zNYQejZsycPHz4kMDCQunXrUrZsWVJTU7l58yZHjhxh\n06ZN2NnZsXr1aoOFf2lIKQhZFhKiu+3dC6VLZ/11Wi306wePH+taFRYyypUQZs9gBQHgwoULrF69\nmj///JOrV68CUL58eRo3bkyvXr1wd3d//cRZCSkFIUv+/BPefBP274ec/GmePAE/P+jQASZMMHg8\nIUQuM2hBAHj8+DHfffcd+/btw8LCgsaNGzNs2DAK5WK/ghSEzN29Cz4+sGABtG2b8+VER0O9evDT\nT9C8ueHyCSFyn8ELQvfu3bG3t6dv374oisLKlSu5f/8+a9eufe2wWSUFIXNBQeDsDLNnv/6ytm6F\n4cPh33/Bzu71lyeEUIfBC0K1atWIiIjI9DFjkoLwauvW6bp4/v7bcDuEBw+GggXh++8NszwhRO4z\n2FhGz9SuXZv9+/frpw8cOECdVx3HKHJVQgK8+y4sWmTYo4O++go2bdLtjxBC5A+ZthCqVKnCuXPn\nKFeuHBqNhsjISDw9PbG0tESj0fDvv/8aP6S0EF5q/Hjd2ciLFxt+2StWwDffwKFDctSREObI4F1G\nV65ceeUC3NzcsryynJKCkLFz5+CNN+DkSd1QFIamKODrq+s+eustwy9fCGFcBi8IpkAKQsa6d9ed\nhfzhh8Zbx+HD0LkzXLggJ6wJYW4Mvg9BmKbjx3XnHbz7rnHXU68eNGgA331n3PUIIdQnLQQz1bEj\ntG4N77xj/HWdOAEtW+paCXIYqhDmQ1oI+cCBA7phq//zn9xZX40aupFR587NnfUJIdQhLQQz1LIl\n9OypG5Aut5w7p9vBfP48FCuWe+sVQuSctBDyuIMHdRvlgQNzd72VK0O7dnKimhB5mbQQzEy3btCk\nCYwalfvrPnkSWrWCy5fBxib31y+EyB6TayGEhYVRpUoVKlWqxBdffJHu+Z9++glvb29q1qyJr69v\nrpzoZq7On4c9e9Q7J6B6dd1hrsuXq7N+IYRxGbWFoNVq8fT0ZMeOHTg7O1OvXj1WrVpF1apV9fPs\n37+fatWqUbRoUcLCwpg2bRoHDhxIG1JaCAAMHaq7xsHHH6uXYe9e3b6LiAgoUEC9HEKIzJlUC+HQ\noUNUrFgRNzc3rKysCAoKIjQ0NM08jRo1omjRogA0aNCAa9euGTOS2bp1C9asgZEj1c3RpAk4OMAL\nf0YhRB5g1IIQHR1NuXLl9NMuLi5ER0e/dP6FCxfSrl07Y0YyW999pxviOjtXQTMGjQY++ABmzVI3\nhxDC8IxaEDQaTZbn3b17N4sWLcpwP0N+9+QJ/PCD8c9KzqpOnXQX0jl2TO0kQghDsjTmwp2dnYmK\nitJPR0VF4eLikm6+f//9lyFDhhAWFoaDg0OGy5o2bZr+vp+fH35+foaOa7LWrQMvL3hu14uqLC1h\n2DCYNw8WLlQ7jRDimfDwcMLDw3P8eqPuVE5JScHT05OdO3fi5ORE/fr10+1UjoyMpEWLFqxYsYKG\nDRtmHDKf71T29YWxY6FLF7WT/M+dO7pzEy5cgBIl1E4jhMiISe1UtrS0JCQkhICAAKpVq0bPnj2p\nWrUq8+fPZ/78+QB8/PHHxMXFMWzYMGrVqkX9+vWNGcnsHDsGUVG6sYtMSalSEBiouzCPECJvkBPT\nTFxwMFSsCB99pHaS9A4fhh49dK0EOQRVCNNjUi0E8XpiYmD9etO9OE29erqjnrZsUTuJEMIQpCCY\nsEWLdN0ypUqpneTlRo6EkBC1UwghDEG6jExUaqquq2j1ajDl3SpJSeDqCn/8AZUqqZ1GCPE86TLK\nI3btAnt7XbeMKStYEPr1k8NPhcgLpIVgooKCdMNEjBihdpLMnTkDfn66o6GsrNROI4R4RloIecDd\nuxAWBr17q50ka6pU0Z2TsHmz2kmEEK9DCoIJWr5cd97BS07aNklDhsCCBWqnEEK8DukyMjGKohum\n4r//haZN1U6TdY8fg4uL7kS68uXVTiOEAOkyMnv794NWq9t/YE4KFdJ1ccmZy0KYLykIJmbBAt2J\naNkYKNZkDBmiKwhardpJhBA5IQXBhNy/Dxs2QP/+aifJmZo1oWxZ2LZN7SRCiJww6vDXIntWrYKW\nLcHRUe0kOfds53Jevc7R3Ud3uRh7kVsPbxHzKAaNRoOVhRUOhRxwK+aGWzE3bK1s1Y4pRI7ITmUT\nUrcufPoptGmjdpKcS0jQnbkcEaFrLZi7Gwk3+PXcr4RdDONw9GHik+KpVKISjoUdKWGrG/c7WZtM\nzOMYrty7QtT9KCqXqIxvOV9aebSiTcU22FjaqPwuRH6V3W2nFAQTcfw4dO4Mly6Z/8ihQ4aAu7tp\njtCaFYkpiayLWMf8o/M5efskARUDaF+pPQ1dGuLu4I6F5uU9rU+0Tzh+4zh/RP7Bb+d/4/jN43So\n3IHhdYfT0KVhtq4iKMTrkoJgpkaM0I0cOnWq2kle36FD0KsXnD8PFma0lyohKYGQQyF8feBrfMr4\nMLTuUDpW7ohVgZyffn3zwU1WnVhFyOEQStqWZLzveLpU6SKFQeQKKQhm6NEj3TH8f/+t624xd4oC\nPj7w1Vfg7692mswla5P59tC3zPxjJi3dWzK56WSqljLs9Uq1qVo2n9vM9D3TsbKwYmbLmbSo0MKg\n6xDiRVIQzNCyZbpRTfPSdQVCQuDPP3U7yk3Zzks7eWfrO7gWdeWrgK+oVqqaUdeXqqTy86mfmbBz\nAnWd6vJNm29wsnMy6jpF/iUFwQw1aQJjxsCbb6qdxHDi4qBCBd3V1EqWVDtNeglJCby37T1+v/Q7\nc9rMIdAzMFe7cR4nP2bGvhnMPzqfGS1mMKT2EOlGEgYnBcHM5OWRQvv1g9q1dcXOlPwZ+Sf9N/an\nWflmfNPmG+wL2quW5dTtU/Td0JcKxSqwoOMC/ZFLQhiCDF1hZn78EQYMyHvFAP53ToKp1PJUJZWP\n93xM15+7Mrv1bBZ1WqRqMQDwKu3FgeADVChWAZ/5Puy+vFvVPCJ/kxaCipKSoFw5XV97XrzamKLo\nhsZevBjeeEPdLPcT79NvQz9iH8eytvtaytqZ3kkS2y5sY2DoQMY2Gst7jd6TLiTx2qSFYEZCQ6F6\n9bxZDEA3HtNbb6k/LHbEnQjqLahH+aLl2TVgl0kWA4CAigEcfOsgP534iQEbB5CYkqh2JJHPSEFQ\n0Q8/6LpV8rIBA2DjRt04TWrYcWkHfkv8mNhkIt+2+xbrAtbqBMki16Ku/DH4D55on9B0cVNuJNxQ\nO5LIR6QgqOTiRfjnH+jSRe0kxlW6tG58ppUrc3/dy/5ZRp/1ffilxy8M8BmQ+wFyyNbKllVdV9Gx\nckd8F/lyLuac2pFEPiEFQSULF+qOwrHJB8Pc5Ha3kaIozNg7gym7p7B7wG6aljejKw09pdFomNxs\nMpOaTqLZkmYcvHZQ7UgiH5CdyipITtadkbxrF1Q17AmxJik1VTe20fr1usNQjUlRFN7b9h67r+xm\na5+tJru/IDt+Pfcrg0IHsazzMtpWaqt2HGFGZKeyGfj1V/DwyB/FAHTjGQUHG7+VkKqkMvTXoey/\ntp/dA3bniWIA0KFyB0KDQhkYOpDQM6FqxxF5mLQQVNCuHQQFme+FcHLi2jXdBXSioqBwYcMvPyU1\nhUGhg4i8H8mvvX7FrqCd4VeisqPXj9J+ZXu+bfst3b26qx1HmAFpIZi4yEg4eBC6dVM7Se5ycdGd\ni/Dzz4Zf9hPtE4J+CeL2w9ts7bM1TxYDgDpOddjWdxvvhr3LyhMq7KUXeZ4UhFy2aJFuaGjbfHhR\nrSFDdGdmG1JKagq91/XmifYJm4I25fmrlXmX8WZHvx188PsHLPl7idpxRB4jXUa5SKsFNzfdPgRv\nb7XT5L6UFN3O9N9/By+v119eqpLK4NDBXE+4zuZemyloWfD1F2omzt49i/8yf75o+QV9avZRO44w\nUdJlZMLCwsDJKX8WAwBLSxg0yDCtBEVRGLV1FBdiL7Ch54Z8VQwAPEt6sr3fdsb+PpZ1EevUjiPy\nCGkh5KKOHXWXyQwOVjuJei5fhnr1dPtSXqfbbMLOCWy7uI1d/XdR1Kao4QKamb9v/k3AigAWBS6i\nfeX2ascRJkZaCCbq0iU4cEC3/yA/q1ABGjZ8vQvnfL7vczae2UhYn7B8XQwAfMr4sLnXZgaFDmLH\npR1qxxFmTgpCLvn+exg4MH/uTH7RyJG6K6rlpNEXciiEH4//yI7+OyhVuJThw5mh+s71WddjHb3W\n9WLf1X1qxxFmTLqMcsGjR1C+vO5wU3d3tdOoLzUVPD1h6dLsDYu99O+lTNo9ib0D91LBoYLxApqp\nHZd20Htdb37t/Sv1neurHUeYAJPqMgoLC6NKlSpUqlSJL774It3zZ86coVGjRtjY2DB79mxjRlHV\n6tXQoIEUg2csLGD4cF0rIat+ifiFD3d+yO/9fpdi8BIt3VuyqNMiOq7qyL+3/lU7jjBDRmshaLVa\nPD092bFjB87OztSrV49Vq1ZR9bnxGu7cucPVq1fZuHEjDg4OvP/++xmHNOMWgqJAnTowYwa0lWFo\n9OLidAXy9GkoU+bV8249v5WBoQPZ1ncbPmV8ciegGfv51M+M2TaG3QN2U7lEZbXjCBWZTAvh0KFD\nVKxYETc3N6ysrAgKCiI0NO04LKVKlaJu3bpY5cXrRz514ADEx0NAgNpJTIuDA/Tokfn4Rnuu7GHA\nxgFs7LlRikEW9fDqwSfNP6HV8lZE3o9UO44wI0YrCNHR0ZQrV04/7eLiQnR0tLFWZ7JCQnTdIxay\n+z6dESNg/nzd6K8ZORR9iO5ru7Oq6yoalWuUu+HM3OBagxnTcAwtl7Xk5oObascRZsLSWAs29PVg\np02bpr/v5+eHn5+fQZdvDNeuwdat2esrz09q1oSKFeGXX9Ifjnvi1gkCVwWyMHAh/u7+6gQ0c6Mb\njiYhKYHWy1sTPjCc4oWKqx1JGFl4eDjh4eE5fr3RCoKzszNRUVH66aioKFxcXHK8vOcLgrmYO1c3\noqmDg9pJTNfYsTBlim7012e/Ic7HnKfNT234ps03dPTsqG5AMzep6STik+Jp+1NbdvTbkWcH/hM6\nL/5Ynj59erZeb7SOjLp163L+/HmuXLnCkydPWLNmDYGBgRnOa647jF8lPl53VbTRo9VOYtratYPE\nRNi9WzcdeT+SVstbMd1vOkHVg9QNlwdoNBq+bPUlPo4+dFzVkcfJj9WOJEyYUc9D2Lp1K6NHj0ar\n1RIcHMxHH33E/PnzAXj77be5efMm9erVIz4+HgsLC+zs7IiIiKBIkSJpQ5rhUUazZ8ORI693Rm5+\nsXChrtto8dqbNF3clOH1hjO6oVRSQ9Kmaum/sT/3Eu+xoecGrAtYqx1J5ILsbjvlxDQjSE7WHVK5\ncaPukFPxaklJ4OoZS9FRfvSt3Y0pzaaoHSlPStYm021tNwoWKMiqrqsoYFFA7UjCyEzmsNP8bM0a\nqFRJikFWPSEB68FtsY4MYHLTyWrHybOsClixptsaYh/HMmTzEFKVVLUjCRMjBcHAFAVmzdLtLBWZ\ne/jkIe1XtqelVy2il37J9euGPTpNpGVjacPGoI2cuXuGMWFjzKrlLYxPCoKBbd+uuxCMnJWcucSU\nRDqv6YxHcQ8WvvkdA/pr+PprtVPlfUWsi7Clzxb2Ru5lavhUteMIEyL7EAxIUcDXF959V3cYpXi5\nJ9onvLnmTewK2rGiywoKWBTg2jXdxYNOn4bSpdVOmPfdfnibpoubElwrmA98P1A7jjAC2Yegot9/\n143R07272klMW7I2maBfgrAuYM2yzsv0OzddXHSFdNYslQPmE6ULl2ZH/x18d+Q7/nvkv2rHESZA\nWggGoijQuLFuOIbevdVOY7q0qVr6bujL/cT7GV76MipK10o4exZKyeUOcsXF2Is0W9KMmS1n0rdm\nX7XjCAOSFoJKdu6EmBjo2VPtJKYrVUllyOYh3H54m3U91mV4HeRy5XSfYR4eDd3keBT3YFvfbYzd\nPpYNpzeoHUeoSFoIBvBs38GIEdCnj9ppTJM2Vctbm9/iUtwltvTeQmHrwi+dNzISatWCiAhwdMzF\nkPnc0etHabeyHd+3/543q76pdhxhANJCUMHGjfDwoexIfpmU1BT6b+xP5P3ITIsBgKsr9OsHH3+c\nSwEFAHWc6rC1z1aG/zacNSfXqB1HqEBaCK8pJQWqV4dvvoE2bdROY3qStcn0Wd+H+KR4NvTcQCGr\nQll63d27UKUK7N+vO8lP5J5/b/1LwIoA/q/V/8k+BTMnLYRctnAhODvLBXAykpSSRI9fevA45TEb\ngzZmuRgAlCwJ778PEyYYMaDIUE3Hmuzsv5PxO8az6PgiteOIXCQthNfw8KHu1+vmzTJMxYseJz+m\n+9ruWBewZnW31TkaTO3RI6hcGdat012TWuSuczHnaLmsJR81/ohh9YapHUfkgLQQctGMGdCihRSD\nF8U9jqP1itbYF7RnTbc1OR5Z09YWPv1Ud6Jfqgy7k+sql6jM7gG7mbV/Fp/s+cQkf5QJw5IWQg6d\nPas7sujff8HJSe00piM6Ppo2P7XBv4I/XwV8hYXm9X5zpKbqzu8YPBjeestAIUW23HxwkzYr2uBb\nzpe5befKKKlmRIa/zgWKottn0LYtjBmjdhrTcebuGdqsaMOwusMY5zvOYJdRPX5ct8M+IgJKlDDI\nIkU23U+8T+c1nSlpW5IVXVZkeA6JMD3SZZQLfvkFbtyAkSPVTmI6/or6i+ZLmzPNbxrjG4836DW1\na9WCHj1kB7OaitoUJaxPGABtfmpD7ONYlRMJY5AWQjbFxuouDr9qFTRponYa07Dsn2WM3T6WpZ2X\n0raScYZ5vXcPvLx0n3vTpkZZhcgCbaqWD37/gF/P/crmXpvxLOmpdiTxCtJlZGR9+0Lx4jB3rtpJ\n1JeqpDJh5wTWRqxlc6/NVCtVzajr27RJ10X3zz/wwlVWRS5beGwhE3ZNYEWXFbTyaKV2HPESUhCM\naMMGGDcO/v4bCr/6ZNs878GTB/Rd35d7iff4pccvlLQtmSvrHTAA7OwgJCRXVideYe/VvfRY24PJ\nTSczvN5wg3YTCsOQgmAkd+7ouop++UV3dFF+dvL2Sbqv7U4T1yaEtAvJ1Qu237sHNWrAkiXg759r\nqxUvcSnuEp1Wd6J22dp81+67TIclEblLdiobgVar6yoaNEiKwZK/l9B8aXM+9P2QHzr+kKvFAKBY\nMVi0CPr31+3YF+pyd3DnQPABLDQW1P+xPhF3ItSOJF6DtBCyYPp02L0bduwAS0vVYqjqUfIjRmwZ\nwYFrB1jbfS3VS1dXNc+0aRAenr//JqZm8fHFjNsxjtmtZ9Pfu7/acQTSZWRw27frWgZHj0KZMqpE\nUN2R60fov6E/dZzq8H377ylirf4eXa0W2rWD2rXh88/VTiOeedadWLtsbb5t+y3FCxVXO1K+Jl1G\nBnTmjK5rYuXK/FkMkrXJTN09lfYr2zOl2RSWd1luEsUAoEABWLFCdxjqTz+pnUY8U710dY7+5ygl\nC5Wkxvc1+O3cb2pHEtkgLYSXuHkTGjWCqVNh4MBcXbVJOHr9KEM2D6FMkTL8GPgjTnamOT7HqVPQ\nvDmsWaP7vzAde67sYVDoIPzc/Pgq4CuK2RRTO1K+Iy0EA3jwADp00HUV5bdiEJ8Uz6ito2i/sj3v\nNniX33r/ZrLFAHQnq61Zo7s40cmTaqcRz2vm1ox/h/2LjaUN1eZVY8W/K1Q/WlC8mrQQXhAfr+ub\n9vKC//4X8suh1alKKqtOrGL8jvEEeATwZasvKWFrPgMHrV4N772n2+dTXd393SIDB68dZPiW4RS2\nKsy8dvOo4VhD7Uj5guxUfg337+sGUfPxgXnzwCKftJ/2XNnD+9vfx0JjwVcBX9HYtbHakXJk1Spd\nUQgLA29vtdOIF2lTtcw/Op9p4dMI9Axkmt80XOxd1I6Vp0lByKHoaAgMhDfe0A1LkR9aBkeuH2H6\nnumcvH2Sz/0/p4dXj9cerlpta9fqBh1cu1bGPDJV9xLv8cUfX/DDsR8YUnsIH7zxgVm1Rs2J7EPI\ngaNHoWFD6NYtfxSDg9cO0n5lezqv7kyARwCnR5wmqHqQ2RcDgO7ddUcfdeumO4FNmJ5iNsX4vOXn\n/DP0H+Iex1Hp20qMCRvDtfhrakfL9/J1C0FRYOlS+OADmD8f3nzT4KswGcnaZDac2cDcg3OJio/i\nQ98PGVxrcJ4d1/7MGejYUdcF+H//BzY2aicSLxMdH81X+79i8d+L6VKlC2MajVH9xMe8QrqMsigm\nBoYO1W04Vq7UjY+TF91IuMHivxfz/ZHvcXdwZ1SDUQR6BmJpkfdP742N1f2NIyJ0f+OaNdVOJF4l\n5lEM8w7PY/7R+bgVc+PtOm/TvVp3ClkVUjua2ZKCkInUVF2XwocfQs+eurNc89qvxwdPHrDxzEaW\n/7ucQ9GH6Fq1KyPrj8SnjI/a0XKdosDy5fD++zBkiO4iOzJ0tmlLSU3h13O/8t8j/+XI9SP09OpJ\nz+o9aezaOE90a+YmKQivsG8fjB8PKSnw7bfQoIEBwpmI2MexbDm/hc3nNrPtwjYauzamb82+BHoG\nYmtlq3Y81V2/rvvb794Nn36qG6xQxkAyfZfjLrPyxErWnFpDzOMYulfrTrdq3Wjo0jBftHJflxSE\nF6Smwu+/w2efwbVrMHmybjgKcz+kNCU1haPXjxJ+JZytF7Zy7MYxWlRoQcfKHeno2ZHShUurHdEk\n/fknTJoEV6/qWol9+4Kt1EuzcPrOadZGrGX96fVE3o/E392fNh5taO3RmnJFy6kdzySZVEEICwtj\n9OjRaLVa3nrrLcaPH59unnfffZetW7dia2vLkiVLqFWrVvqQOSgIkZG6fuMFC3RdBGPHQq9e5vur\nMOZRDEdvHOXI9SPsi9zHX1F/Ub5oefzc/Gjt0Rr/Cv7S15oNf/wBM2fC/v3Qu7fuwjt16uT9I8zy\nihsJN9h+cTthF8P4/eLvFLEugq+rL77lfHmj3Bt4lfLCqoCV2jFVZzIFQavV4unpyY4dO3B2dqZe\nvXqsWrWKqlWr6ufZsmULISEhbNmyhYMHDzJq1CgOHDiQPmQW3lRysu5KZr/9BqGhutZAly7w1ltQ\nr575/ENPTEnkfMx5zsac5czdMxy/eZyj148SlxhHrTK1qFO2Do1dG9OkfJMcX6UsPDwcPz8/wwY3\nU5GRsHCh7qS2x49156I8Ox/Fzi7z18tnaVg5+TwVReFszFn+jPyTv6L+4q9rf3H13lU8S3ri7eiN\nt6M3NR1r4lnSEyc7p3y1HyK7BcFov5cPHTpExYoVcXNzAyAoKIjQ0NA0BWHTpk0MGDAAgAYNGnDv\n3j1u3bqFo6PjK5edlATnzukGNjtxAv76C44cATc3aN0a5szR/YM2tdaAoigkPEkgOj6aa/HXiIqP\nIup+lO7/8VGcjznP9YTrVHCogGcJTzxLeNKtajc+9/+cisUrGuyLLBux/3F11V3vYto03RFnoaEw\nYwYcOwaVK+u+R7Vq6YYyqVYN7O3Tvl4+S8PKyeep0WioUrIKVUpWIbh2MKC7fsfJ2yf55+Y//HPr\nHzac2cCF2AvEJcbh7uBOxeIV8XDwoJx9OZzsnHCyc8LZ3pmyRcrm65a20TaZ0dHRlCv3v349FxcX\nDh48mOk8165dy7Ag9OkDUVG6282buo1/tWq6cWvGjdONTFrMwIMpalO1PNE+SXdL0ibxKPkRD548\n0N8SkhLSTN9LvEfM4xjuPrqb5mZdwBonOyfKFS1HOXvdrb5zfbpW7UrF4hWp4FBBdpapQKOBqlV1\ntw8/1P3oOHZMt89h7174/ntdwbCzg3Ll/neLiNCdAFe8+P9uxYpBoUK6m42N+e+vMke2VrbUd65P\nfef6aR5/8OQBl+IucSH2AhdjL3L1/lX+uvYX1xOu62+2VraUKFSC4oWK628ONg4UL1ScYjbFKGxd\nmMJWhSlsXRhbK1v9/Wf/ty5gjXUBa6wsrLAuYI2lhaXZXG/aaFuerH4ALzZnXva6f7xbYV0/lTIF\nFVwLpoJG4baSyk5F4ffrqSjrFFKVVBTl6f/JeDqjx7SpWpJTk9Nt+BVFoaBlQf0f+Pk/dGHrwhSx\nLoKdtR1FrIukuW9f0B4XexdKFS5FSduSlLQtSYlCJShhWwIbyzx2jGseVbCg7kdGo0b/eyw1VXfZ\nzqgoXVdTVBQcPKg7ei029n+3uDhd99Pjx/DkiW5ZzwpEwYK6azk8f7O0zPixFwvJ8/80Xud+dubL\nbWfP6kYOMJ4iQM2nt/8p/vTmhUJygVieWMaSbBlLXIFYblnGklwgjmTLWJILRKMt8BCtxUNSLHT/\nfzattXhEisVDFM0TUjXJpGqeoGiSUSxS0KRaYaFYoVGssFCs9f+3UKwADRrFArBAgwYUCzRYpH1c\neTqNxdPnNekf5+kfT9H9X0P2/5hGKwjOzs5ERUXpp6OionBxcXnlPNeuXcPZ2Tndsjw8PDg1foex\noui/7R8AAAmSSURBVL5S4tP/8prp06erHSHPOHDg1Z9lYqLuFheXS4HM3Pnzee+7qZCMluRcX6+H\nh0e25jdaQahbty7nz5/nypUrODk5sWbNGlatWpVmnsDAQEJCQggKCuLAgQMUK1Ysw+6iCxcuGCum\nEEKIp4xWECwtLQkJCSEgIACtVktwcDBVq1Zl/vz5ALz99tu0a9eOLVu2ULFiRQoXLszixYuNFUcI\nIUQmzOLENCGEEMZn0sc/hIWFUaVKFSpVqsQXX3yhdhyz5+bmRs2aNalVqxb169fP/AUijcGDB+Po\n6EiN50ZCjI2NpVWrVlSuXJnWrVtz7949FROal4w+z2nTpuHi4kKtWrWoVasWYWFhKiY0H1FRUTRv\n3hwvLy+qV6/O3Llzgex/P022IGi1WkaOHElYWBgRERGsWrWK06dPqx3LrGk0GsLDwzl+/DiHDh1S\nO47ZGTRoULoN1MyZM2nVqhXnzp3D39+fmTNnqpTO/GT0eWo0Gt577z2OHz/O8ePHadOmjUrpzIuV\nlRVff/01p06d4sCBA8ybN4/Tp09n+/tpsgXh+RPbrKys9Ce2idcjPYQ516RJExwcHNI89vzJlQMG\nDGDjxo1qRDNLGX2eIN/RnChTpgw+PrrRjIsUKULVqlWJjo7O9vfTZAtCRietRUdHq5jI/Gk0Glq2\nbEndunVZsGCB2nHyhOfPrHd0dOTWrVsqJzJ/3377Ld7e3gQHB0sXXA5cuXKF48eP06BBg2x/P022\nIJjLmX3m5M8//+T48eNs3bqVefPmsW/fPrUj5SkajUa+t69p2LBhXL58mb///puyZcvy/vvvqx3J\nrDx48ICuXbsyZ84c7F4YjCsr30+TLQhZObFNZE/ZsmUBKFWqFF26dJH9CAbg6OjIzZs3Abhx4wal\nS8uw46+jdOnS+g3XW2+9Jd/RbEhOTqZr167069ePzp07A9n/fppsQXj+xLYnT56wZs0aAgMD1Y5l\nth49ekRCQgIADx8+ZPv27WmO7hA5ExgYyNKlSwFYunSp/h+iyJkbN27o72/YsEG+o1mkKArBwcFU\nq1aN0aNH6x/P9vdTMWFbtmxRKleurHh4eCifffaZ2nHM2qVLlxRvb2/F29tb8fLyks8zB4KCgpSy\nZcsqVlZWiouLi7Jo0SIlJiZG8ff3VypVqqS0atVKiYuLUzum2Xjx81y4cKHSr18/pUaNGkrNmjWV\nTp06KTdv3lQ7plnYt2+fotFoFG9vb8XHx0fx8fFRtm7dmu3vp5yYJoQQAjDhLiMhhBC5SwqCEEII\nQAqCEEKIp6QgCCGEAKQgCCGEeEoKghBCCEAKgsjj3NzciI2NVTtGGtevX6d79+6vnCc8PJyOHTtm\n+JwpvieRN0hBECZJURSDjHppamMLpaSk4OTkxNq1a3O8DFN7TyLvkIIgTMaVK1fw9PRkwIAB1KhR\ng6ioKIYPH069evWoXr0606ZN08/r5ubGtGnTqFOnDjVr1uTs2bMAxMTE0Lp1a6pXr86QIUPSFJWv\nvvqKGjVqUKNGDebMmaNfZ5UqVRg0aBCenp706dOH7du34+vrS+XKlTl8+HC6nI0aNSIiIkI/7efn\nx7Fjxzh8+DBvvPEGtWvXxtfXl3PnzgGwZMkSAgMD8ff3p1WrVly9epXq1avr19+0aVPq1KlDnTp1\n2L9/v3658fHxdOjQgSpVqjBs2LAMC+SK/2/vbkKiWuM4jn/PGGVN9EKbkkCjRdZMTGNmBSqCNlBQ\nlIYRQWUUVEzboEUYJEELmU0EESLZGzNIY4twIRGkm8KIMgwrONaiiNBqKJWYmd9dpAdtvN26cLmB\n/8/qnPPM8zYHzv85z5k5z7VrbNy4kXA4zNGjR8lms1PSP3/+THFxsdeWvXv30tLS8lvnxcwg//l/\nqo35Ra7ryufz6cGDB96x4eFhSVI6nVZVVZX6+vokSUVFRbpw4YIk6eLFizp8+LAk6cSJEzp79qwk\n6c6dO3IcR0NDQ+rt7dXatWs1MjKiL1++KBAI6PHjx3JdV7NmzdKzZ8+UzWa1fv16HTp0SJJ0+/Zt\n7dy5M6edsVhMjY2NkqS3b99q1apVkqRUKqV0Oi1J6urqUl1dnSSptbVVy5cv914b4LqugsGgJGlk\nZERjY2OSpBcvXqi0tFSSdO/ePeXn58t1XWUyGW3ZskXt7e1e34eGhtTf36/t27d7dR47dkxtbW05\n7e3q6tLmzZt18+ZNbd269XdOiZlhZv3fAcmYyQoLC6cs7xmPx7l8+TLpdJp3797R39/vja5ra2sB\nKCkp4datWwB0d3eTTCYB2LZtG4sXL0YSPT091NbWMnfuXC9vd3c3O3bsYMWKFQQCAQACgQA1NTUA\nBINBBgcHc9pYX19PJBLhzJkzJBIJ73nAp0+f2L9/P69evcJxHNLptJcnEomwaNGinLK+fftGNBrl\nyZMn5OXl8fLlSy+trKyMoqIi4PvIvqenh7q6OuD7lNrdu3d59OgRpaWlAIyOjrJ06dKcOmpqakgk\nEkSjUZ4+ffrT79/MbBYQzB/F7/d7267r0tzcTG9vLwsXLqShoYGxsTEvfc6cOQDk5eVNufhqmqkV\nx3GmHJfkzcVPlAPg8/mYPXu2tz253AkFBQUsWbKEvr4+EokEly5dAuD06dNUV1eTTCZ5/fo1VVVV\nXp558+ZN299YLMayZcu4evUqmUyG/Pz8KW2e3F6fL3eG98CBA5w7d27asidks1meP3+O3+9neHiY\ngoKCn37ezFz2DMH8sVKpFH6/nwULFvD+/Xs6Ozv/MU9lZSU3btwAoLOzk48fP+I4DhUVFXR0dDA6\nOsrXr1/p6OigoqLiXz+43rNnD+fPnyeVSnl3LKlUyrvYtra2/nIfJ0b1bW1tZDIZL+3hw4cMDg6S\nzWaJx+OUl5d7aY7jUF1dTXt7Ox8+fAC+L6j+5s2bnDpisRiBQIDr16/T0NAwbZAzBiwgmD/M5FFx\nKBQiHA5TXFzMvn37plwQf8wzka+xsZH79+8TDAZJJpMUFhYCEA6HOXjwIGVlZWzatIkjR44QCoVy\n6vxx/+9+0bN7927i8Tj19fXesZMnT3Lq1ClKSkrIZDJe3ulWqprYP378OFeuXGHdunUMDAwwf/58\nL33Dhg1Eo1HWrFnDypUr2bVr15S8q1evpqmpiUgkQigUIhKJeIuhTBgYGKClpYXm5mbKy8uprKyk\nqalp2j4ZY6+/NsYYA9gdgjHGmHEWEIwxxgAWEIwxxoyzgGCMMQawgGCMMWacBQRjjDGABQRjjDHj\nLCAYY4wB4C+6RXPdrA827gAAAABJRU5ErkJggg==\n", + "text": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 2, + "metadata": {}, + "source": [ + "C-extensions via the Cython magic" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Cython (see [Cython's C-extensions for Python](http://cython.org)) is basically a hybrid between C and Python and can be pictured as compiled Python code with type declarations.\n", + "Since we are working in an IPython notebook here, we can make use of the very convenient IPython magic: It will take care of the conversion to C code, the compilation, and eventually the loading of the function.\n", + "Also, we are adding C type declarations; those type declarations are not necessary for using Cython, however, it will improve the performance of our code significantly." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%load_ext cythonmagic" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 29 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%%cython\n", + "import numpy as np\n", + "cimport numpy as np\n", + "cimport cython\n", + "@cython.boundscheck(False) \n", + "@cython.wraparound(False)\n", + "@cython.cdivision(True)\n", + "cpdef cython_lstsqr(x_ary, y_ary):\n", + " \"\"\" Computes the least-squares solution to a linear matrix equation. \"\"\"\n", + " cdef double x_avg, y_avg, var_x, cov_xy,\\\n", + " slope, y_interc, temp\n", + " cdef double[:] x = x_ary # memoryview\n", + " cdef double[:] y = y_ary\n", + " cdef unsigned long N, i\n", + " \n", + " N = x.shape[0]\n", + " x_avg = 0\n", + " y_avg = 0\n", + " for i in range(N):\n", + " x_avg += x[i]\n", + " y_avg += y[i]\n", + " x_avg = x_avg/N\n", + " y_avg = y_avg/N\n", + " var_x = 0\n", + " cov_xy = 0\n", + " for i in range(N):\n", + " temp = (x[i] - x_avg)\n", + " var_x += temp**2\n", + " cov_xy += temp*(y[i] - y_avg)\n", + " slope = cov_xy / var_x\n", + " y_interc = y_avg - slope*x_avg\n", + " return (slope, y_interc)" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "building '_cython_magic_cf6c91cb1e11de8d2dbe7d9178e469df' extension\n" + ] + }, + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "C compiler: /usr/bin/clang -fno-strict-aliasing -Werror=declaration-after-statement -DNDEBUG -g -fwrapv -O3 -Wall -Wstrict-prototypes -I/Users/sebastian/miniconda3/envs/py34/include -arch x86_64\n", + "\n" + ] + }, + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "compile options: '-I/Users/sebastian/miniconda3/envs/py34/lib/python3.4/site-packages/numpy/core/include -I/Users/sebastian/miniconda3/envs/py34/include/python3.4m -c'\n" + ] + }, + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "clang: /Users/sebastian/.ipython/cython/_cython_magic_cf6c91cb1e11de8d2dbe7d9178e469df.c\n" + ] + }, + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "/usr/bin/clang -bundle -undefined dynamic_lookup -L/Users/sebastian/miniconda3/envs/py34/lib -arch x86_64 /Users/sebastian/.ipython/cython/Users/sebastian/.ipython/cython/_cython_magic_cf6c91cb1e11de8d2dbe7d9178e469df.o -L/Users/sebastian/miniconda3/envs/py34/lib -o /Users/sebastian/.ipython/cython/_cython_magic_cf6c91cb1e11de8d2dbe7d9178e469df.so\n" + ] + } + ], + "prompt_number": 30 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "import numpy as np\n", + "\n", + "x_ary = np.array([x_i*np.random.randint(8,12)/10 for x_i in range(100)])\n", + "y_ary = np.array([y_i*np.random.randint(10,14)/10 for y_i in range(100)])" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 31 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "cython_lstsqr(x_ary, y_ary)" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "pyout", + "prompt_number": 32, + "text": [ + "(1.1399825800539194, 2.0824398156005444)" + ] + } + ], + "prompt_number": 32 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 2, + "metadata": {}, + "source": [ + "Running Fortran Code" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "There is also a convenient IPython magic command for compiling Fortran code. The Fortran magic uses NumPy's [F2PY](http://www.f2py.com) module for compiling and running the Fortran code. For more information, please see the ['Fortran magic's documentation'](http://nbviewer.ipython.org/github/mgaitan/fortran_magic/blob/master/documentation.ipynb)." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%install_ext https://raw.github.com/mgaitan/fortran_magic/master/fortranmagic.py" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "Installed fortranmagic.py. To use it, type:\n", + " %load_ext fortranmagic\n" + ] + } + ], + "prompt_number": 17 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%load_ext fortranmagic" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "javascript": [ + "$.getScript(\"https://raw.github.com/marijnh/CodeMirror/master/mode/fortran/fortran.js\", function () {\n", + "IPython.config.cell_magic_highlight['magic_fortran'] = {'reg':[/^%%fortran/]};});\n" + ], + "metadata": {}, + "output_type": "display_data" + } + ], + "prompt_number": 18 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%%fortran\n", + "SUBROUTINE fortran_lstsqr(ary_x, ary_y, slope, y_interc)\n", + " ! Computes the least-squares solution to a linear matrix equation. \"\"\"\n", + " IMPLICIT NONE\n", + " REAL(8), INTENT(in), DIMENSION(:) :: ary_x, ary_y\n", + " REAL(8), INTENT(out) :: slope, y_interc\n", + " REAL(8) :: x_avg, y_avg, var_x, cov_xy, temp\n", + " INTEGER(8) :: N, i\n", + " \n", + " N = SIZE(ary_x)\n", + "\n", + " x_avg = SUM(ary_x) / N\n", + " y_avg = SUM(ary_y) / N\n", + " var_x = 0\n", + " cov_xy = 0\n", + " \n", + " DO i = 1, N\n", + " temp = ary_x(i) - x_avg\n", + " var_x = var_x + temp**2\n", + " cov_xy = cov_xy + (temp*(ary_y(i) - y_avg))\n", + " END DO\n", + " \n", + " slope = cov_xy / var_x\n", + " y_interc = y_avg - slope*x_avg\n", + "\n", + "END SUBROUTINE fortran_lstsqr" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "\tBuilding module \"_fortran_magic_a044885f2b0c0feac78a230b6b714e2b\"...\n", + "\t\tConstructing wrapper function \"fortran_lstsqr\"...\n", + "\t\t slope,y_interc = fortran_lstsqr(ary_x,ary_y)\n", + "\tWrote C/API module \"_fortran_magic_a044885f2b0c0feac78a230b6b714e2b\" to file \"/var/folders/bq/_946cdn92t7bqzz5frpfpw7r0000gp/T/tmp3y_jxtl_/src.macosx-10.5-x86_64-3.4/_fortran_magic_a044885f2b0c0feac78a230b6b714e2bmodule.c\"\n", + "\tFortran 77 wrappers are saved to \"/var/folders/bq/_946cdn92t7bqzz5frpfpw7r0000gp/T/tmp3y_jxtl_/src.macosx-10.5-x86_64-3.4/_fortran_magic_a044885f2b0c0feac78a230b6b714e2b-f2pywrappers.f\"\n" + ] + } + ], + "prompt_number": 22 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "import numpy as np\n", + "\n", + "x_ary = np.array([x_i*np.random.randint(8,12)/10 for x_i in range(100)])\n", + "y_ary = np.array([y_i*np.random.randint(10,14)/10 for y_i in range(100)])" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 23 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "fortran_lstsqr(x_ary, y_ary)" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "pyout", + "prompt_number": 25, + "text": [ + "(1.1313508052697814, 3.681685640167956)" + ] + } + ], + "prompt_number": 25 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 2, + "metadata": {}, + "source": [ + "Running code from other interpreters: Ruby, Perl, and Bash" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "To use any interpreter that is installed on your system:" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%%script perl\n", + "print 'Hello, World!';" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "Hello, World!" + ] + } + ], + "prompt_number": 44 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Or use the magic command for the respective interpreter directly:" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%%perl\n", + "print 'Hello, World!';" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "Hello, World!" + ] + } + ], + "prompt_number": 45 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%%ruby\n", + "puts \"Hello, World!\"" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "Hello, World!\n" + ] + } + ], + "prompt_number": 46 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%%bash\n", + "echo \"Hello World!\"" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "Hello World!\n" + ] + } + ], + "prompt_number": 47 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%%script R --no-save\n", + "cat(\"Goodbye, World!\\n\")" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "\n", + "R version 3.0.2 (2013-09-25) -- \"Frisbee Sailing\"\n", + "Copyright (C) 2013 The R Foundation for Statistical Computing\n", + "Platform: x86_64-apple-darwin10.8.0 (64-bit)\n", + "\n", + "R is free software and comes with ABSOLUTELY NO WARRANTY.\n", + "You are welcome to redistribute it under certain conditions.\n", + "Type 'license()' or 'licence()' for distribution details.\n", + "\n", + " Natural language support but running in an English locale\n", + "\n", + "R is a collaborative project with many contributors.\n", + "Type 'contributors()' for more information and\n", + "'citation()' on how to cite R or R packages in publications.\n", + "\n", + "Type 'demo()' for some demos, 'help()' for on-line help, or\n", + "'help.start()' for an HTML browser interface to help.\n", + "Type 'q()' to quit R.\n", + "\n", + "> cat(\"Goodbye, World!\\n\")\n", + "Goodbye, World!\n", + "> \n" + ] + } + ], + "prompt_number": 55 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "def hello_world():\n", + " \"\"\"This is a hello world example function.\"\"\"\n", + " print('Hello, World!')" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 7 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%pdoc hello_world" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 9 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%pdef hello_world" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + " \u001b[0mhello_world\u001b[0m\u001b[0;34m(\u001b[0m\u001b[0;34m)\u001b[0m\u001b[0;34m\u001b[0m\u001b[0m\n", + " " + ] + } + ], + "prompt_number": 10 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%psource math.mean()" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "Object `math.mean()` not found.\n" + ] + } + ], + "prompt_number": 15 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "from math import sqrt" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "ename": "ImportError", + "evalue": "cannot import name 'mean'", + "output_type": "pyerr", + "traceback": [ + "\u001b[0;31m---------------------------------------------------------------------------\u001b[0m\n\u001b[0;31mImportError\u001b[0m Traceback (most recent call last)", + "\u001b[0;32m
\n", + "I would be happy to hear your comments and suggestions. \n", + "Please feel free to drop me a note via\n", + "[twitter](https://twitter.com/rasbt), [email](mailto:bluewoodtree@gmail.com), or [google+](https://plus.google.com/118404394130788869227).\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "# Key differences between Python 2.7.x and Python 3.x" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "Many beginning Python users are wondering with which version of Python they should start. My answer to this question is usually something along the lines \"just go with the version your favorite tutorial was written in, and check out the differences later on.\"\n", + "\n", + "But what if you are starting a new project and have the choice to pick? I would say there is currently no \"right\" or \"wrong\" as long as both Python 2.7.x and Python 3.x support the libraries that you are planning to use. However, it is worthwhile to have a look at the major differences between those two most popular versions of Python to avoid common pitfalls when writing the code for either one of them, or if you are planning to port your project." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Sections" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "- [Using the `__future__` module](#future_module)\n", + "\n", + "- [The print function](#The-print-function)\n", + "\n", + "- [Integer division](#Integer-division)\n", + "\n", + "- [Unicode](#Unicode)\n", + "\n", + "- [xrange](#xrange)\n", + "\n", + "- [Raising exceptions](#Raising-exceptions)\n", + "\n", + "- [Handling exceptions](#Handling-exceptions)\n", + "\n", + "- [The next() function and .next() method](#The-next-function-and-next-method)\n", + "\n", + "- [For-loop variables and the global namespace leak](#For-loop-variables-and-the-global-namespace-leak)\n", + "\n", + "- [Comparing unorderable types](#Comparing-unorderable-types)\n", + "\n", + "- [Parsing user inputs via input()](#Parsing-user-inputs-via-input)\n", + "\n", + "- [Returning iterable objects instead of lists](#Returning-iterable-objects-instead-of-lists)\n", + "\n", + "- [Banker's Rounding](#Banker's-Rounding)\n", + "\n", + "- [More articles about Python 2 and Python 3](#More-articles-about-Python-2-and-Python-3)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### The `__future__` module" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Python 3.x introduced some Python 2-incompatible keywords and features that can be imported via the in-built `__future__` module in Python 2. It is recommended to use `__future__` imports it if you are planning Python 3.x support for your code. For example, if we want Python 3.x's integer division behavior in Python 2, we can import it via\n", + "\n", + " from __future__ import division\n", + " \n", + "More features that can be imported from the `__future__` module are listed in the table below:" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
| feature | \n", + "optional in | \n", + "mandatory in | \n", + "effect | \n", + "
|---|---|---|---|
| nested_scopes | \n", + "2.1.0b1 | \n", + "2.2 | \n", + "PEP 227:\n", + "Statically Nested Scopes | \n", + "
| generators | \n", + "2.2.0a1 | \n", + "2.3 | \n", + "PEP 255:\n", + "Simple Generators | \n", + "
| division | \n", + "2.2.0a2 | \n", + "3.0 | \n", + "PEP 238:\n", + "Changing the Division Operator | \n", + "
| absolute_import | \n", + "2.5.0a1 | \n", + "3.0 | \n", + "PEP 328:\n", + "Imports: Multi-Line and Absolute/Relative | \n", + "
| with_statement | \n", + "2.5.0a1 | \n", + "2.6 | \n", + "PEP 343:\n", + "The “with” Statement | \n", + "
| print_function | \n", + "2.6.0a2 | \n", + "3.0 | \n", + "PEP 3105:\n", + "Make print a function | \n", + "
| unicode_literals | \n", + "2.6.0a2 | \n", + "3.0 | \n", + "PEP 3112:\n", + "Bytes literals in Python 3000 | \n", + "
\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## The print function" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to the section-overview](#Sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Very trivial, and the change in the print-syntax is probably the most widely known change, but still it is worth mentioning: Python 2's print statement has been replaced by the `print()` function, meaning that we have to wrap the object that we want to print in parantheses. \n", + "\n", + "Python 2 doesn't have a problem with additional parantheses, but in contrast, Python 3 would raise a `SyntaxError` if we called the print function the Python 2-way without the parentheses. \n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### Python 2" + ] + }, + { + "cell_type": "code", + "execution_count": 3, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Python 2.7.6\n", + "Hello, World!\n", + "Hello, World!\n", + "text print more text on the same line\n" + ] + } + ], + "source": [ + "print 'Python', python_version()\n", + "print 'Hello, World!'\n", + "print('Hello, World!')\n", + "print \"text\", ; print 'print more text on the same line'" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### Python 3" + ] + }, + { + "cell_type": "code", + "execution_count": 2, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Python 3.4.1\n", + "Hello, World!\n", + "some text, print more text on the same line\n" + ] + } + ], + "source": [ + "print('Python', python_version())\n", + "print('Hello, World!')\n", + "\n", + "print(\"some text,\", end=\"\") \n", + "print(' print more text on the same line')" + ] + }, + { + "cell_type": "code", + "execution_count": 3, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "ename": "SyntaxError", + "evalue": "invalid syntax (
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Integer division" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to the section-overview](#Sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "This change is particularly dangerous if you are porting code, or if you are executing Python 3 code in Python 2, since the change in integer-division behavior can often go unnoticed (it doesn't raise a `SyntaxError`). \n", + "So, I still tend to use a `float(3)/2` or `3/2.0` instead of a `3/2` in my Python 3 scripts to save the Python 2 guys some trouble (and vice versa, I recommend a `from __future__ import division` in your Python 2 scripts)." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### Python 2" + ] + }, + { + "cell_type": "code", + "execution_count": 4, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Python 2.7.6\n", + "3 / 2 = 1\n", + "3 // 2 = 1\n", + "3 / 2.0 = 1.5\n", + "3 // 2.0 = 1.0\n" + ] + } + ], + "source": [ + "print 'Python', python_version()\n", + "print '3 / 2 =', 3 / 2\n", + "print '3 // 2 =', 3 // 2\n", + "print '3 / 2.0 =', 3 / 2.0\n", + "print '3 // 2.0 =', 3 // 2.0" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### Python 3" + ] + }, + { + "cell_type": "code", + "execution_count": 4, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Python 3.4.1\n", + "3 / 2 = 1.5\n", + "3 // 2 = 1\n", + "3 / 2.0 = 1.5\n", + "3 // 2.0 = 1.0\n" + ] + } + ], + "source": [ + "print('Python', python_version())\n", + "print('3 / 2 =', 3 / 2)\n", + "print('3 // 2 =', 3 // 2)\n", + "print('3 / 2.0 =', 3 / 2.0)\n", + "print('3 // 2.0 =', 3 // 2.0)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Unicode" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to the section-overview](#Sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Python 2 has ASCII `str()` types, separate `unicode()`, but no `byte` type. \n", + "\n", + "Now, in Python 3, we finally have Unicode (utf-8) `str`ings, and 2 byte classes: `byte` and `bytearray`s." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### Python 2" + ] + }, + { + "cell_type": "code", + "execution_count": 2, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Python 2.7.6\n" + ] + } + ], + "source": [ + "print 'Python', python_version()" + ] + }, + { + "cell_type": "code", + "execution_count": 3, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### Python 3" + ] + }, + { + "cell_type": "code", + "execution_count": 6, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Python 3.4.1\n", + "strings are now utf-8 μnicoΔé!\n" + ] + } + ], + "source": [ + "print('Python', python_version())\n", + "print('strings are now utf-8 \\u03BCnico\\u0394é!')" + ] + }, + { + "cell_type": "code", + "execution_count": 8, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Python 3.4.1 has
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## xrange" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to the section-overview](#Sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + " \n", + "The usage of `xrange()` is very popular in Python 2.x for creating an iterable object, e.g., in a for-loop or list/set-dictionary-comprehension. \n", + "The behavior was quite similar to a generator (i.e., \"lazy evaluation\"), but here the xrange-iterable is not exhaustible - meaning, you could iterate over it infinitely. \n", + "\n", + "\n", + "Thanks to its \"lazy-evaluation\", the advantage of the regular `range()` is that `xrange()` is generally faster if you have to iterate over it only once (e.g., in a for-loop). However, in contrast to 1-time iterations, it is not recommended if you repeat the iteration multiple times, since the generation happens every time from scratch! \n", + "\n", + "In Python 3, the `range()` was implemented like the `xrange()` function so that a dedicated `xrange()` function does not exist anymore (`xrange()` raises a `NameError` in Python 3)." + ] + }, + { + "cell_type": "code", + "execution_count": 5, + "metadata": { + "collapsed": false + }, + "outputs": [], + "source": [ + "import timeit\n", + "\n", + "n = 10000\n", + "def test_range(n):\n", + " return for i in range(n):\n", + " pass\n", + " \n", + "def test_xrange(n):\n", + " for i in xrange(n):\n", + " pass " + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### Python 2" + ] + }, + { + "cell_type": "code", + "execution_count": 6, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Python 2.7.6\n", + "\n", + "timing range()\n", + "1000 loops, best of 3: 433 µs per loop\n", + "\n", + "\n", + "timing xrange()\n", + "1000 loops, best of 3: 350 µs per loop\n" + ] + } + ], + "source": [ + "print 'Python', python_version()\n", + "\n", + "print '\\ntiming range()'\n", + "%timeit test_range(n)\n", + "\n", + "print '\\n\\ntiming xrange()'\n", + "%timeit test_xrange(n)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### Python 3" + ] + }, + { + "cell_type": "code", + "execution_count": 7, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Python 3.4.1\n", + "\n", + "timing range()\n", + "1000 loops, best of 3: 520 µs per loop\n" + ] + } + ], + "source": [ + "print('Python', python_version())\n", + "\n", + "print('\\ntiming range()')\n", + "%timeit test_range(n)" + ] + }, + { + "cell_type": "code", + "execution_count": 5, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "ename": "NameError", + "evalue": "name 'xrange' is not defined", + "output_type": "error", + "traceback": [ + "\u001b[0;31m---------------------------------------------------------------------------\u001b[0m\n\u001b[0;31mNameError\u001b[0m Traceback (most recent call last)", + "\u001b[0;32m
\n", + "
\n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### The `__contains__` method for `range` objects in Python 3" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Another thing worth mentioning is that `range` got a \"new\" `__contains__` method in Python 3.x (thanks to [Yuchen Ying](https://github.com/yegle), who pointed this out). The `__contains__` method can speedup \"look-ups\" in Python 3.x `range` significantly for integer and Boolean types.\n" + ] + }, + { + "cell_type": "code", + "execution_count": 3, + "metadata": { + "collapsed": false + }, + "outputs": [], + "source": [ + "x = 10000000" + ] + }, + { + "cell_type": "code", + "execution_count": 4, + "metadata": { + "collapsed": false + }, + "outputs": [], + "source": [ + "def val_in_range(x, val):\n", + " return val in range(x)" + ] + }, + { + "cell_type": "code", + "execution_count": 5, + "metadata": { + "collapsed": false + }, + "outputs": [], + "source": [ + "def val_in_xrange(x, val):\n", + " return val in xrange(x)" + ] + }, + { + "cell_type": "code", + "execution_count": 7, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Python 3.4.1\n", + "1 loops, best of 3: 742 ms per loop\n", + "1000000 loops, best of 3: 1.19 µs per loop\n" + ] + } + ], + "source": [ + "print('Python', python_version())\n", + "assert(val_in_range(x, x/2) == True)\n", + "assert(val_in_range(x, x//2) == True)\n", + "%timeit val_in_range(x, x/2)\n", + "%timeit val_in_range(x, x//2)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Based on the `timeit` results above, you see that the execution for the \"look up\" was about 60,000 faster when it was of an integer type rather than a float. However, since Python 2.x's `range` or `xrange` doesn't have a `__contains__` method, the \"look-up speed\" wouldn't be that much different for integers or floats:" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "code", + "execution_count": 6, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Python 2.7.7\n", + "1 loops, best of 3: 285 ms per loop\n", + "1 loops, best of 3: 179 ms per loop\n", + "1 loops, best of 3: 658 ms per loop\n", + "1 loops, best of 3: 556 ms per loop\n" + ] + } + ], + "source": [ + "print 'Python', python_version()\n", + "assert(val_in_xrange(x, x/2.0) == True)\n", + "assert(val_in_xrange(x, x/2) == True)\n", + "assert(val_in_range(x, x/2) == True)\n", + "assert(val_in_range(x, x//2) == True)\n", + "%timeit val_in_xrange(x, x/2.0)\n", + "%timeit val_in_xrange(x, x/2)\n", + "%timeit val_in_range(x, x/2.0)\n", + "%timeit val_in_range(x, x/2)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Below the \"proofs\" that the `__contain__` method wasn't added to Python 2.x yet:" + ] + }, + { + "cell_type": "code", + "execution_count": 8, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Python 3.4.1\n" + ] + }, + { + "data": { + "text/plain": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### Note about the speed differences in Python 2 and 3" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Some people pointed out the speed difference between Python 3's `range()` and Python2's `xrange()`. Since they are implemented the same way one would expect the same speed. However the difference here just comes from the fact that Python 3 generally tends to run slower than Python 2. " + ] + }, + { + "cell_type": "code", + "execution_count": 3, + "metadata": { + "collapsed": false + }, + "outputs": [], + "source": [ + "def test_while():\n", + " i = 0\n", + " while i < 20000:\n", + " i += 1\n", + " return" + ] + }, + { + "cell_type": "code", + "execution_count": 4, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Python 3.4.1\n", + "100 loops, best of 3: 2.68 ms per loop\n" + ] + } + ], + "source": [ + "print('Python', python_version())\n", + "%timeit test_while()" + ] + }, + { + "cell_type": "code", + "execution_count": 6, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Python 2.7.6\n", + "1000 loops, best of 3: 1.72 ms per loop\n" + ] + } + ], + "source": [ + "print 'Python', python_version()\n", + "%timeit test_while()" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Raising exceptions" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to the section-overview](#Sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "\n", + "Where Python 2 accepts both notations, the 'old' and the 'new' syntax, Python 3 chokes (and raises a `SyntaxError` in turn) if we don't enclose the exception argument in parentheses:" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### Python 2" + ] + }, + { + "cell_type": "code", + "execution_count": 7, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Python 2.7.6\n" + ] + } + ], + "source": [ + "print 'Python', python_version()" + ] + }, + { + "cell_type": "code", + "execution_count": 8, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "ename": "IOError", + "evalue": "file error", + "output_type": "error", + "traceback": [ + "\u001b[0;31m---------------------------------------------------------------------------\u001b[0m\n\u001b[0;31mIOError\u001b[0m Traceback (most recent call last)", + "\u001b[0;32m
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### Python 3" + ] + }, + { + "cell_type": "code", + "execution_count": 9, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Python 3.4.1\n" + ] + } + ], + "source": [ + "print('Python', python_version())" + ] + }, + { + "cell_type": "code", + "execution_count": 10, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "ename": "SyntaxError", + "evalue": "invalid syntax (
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Handling exceptions" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to the section-overview](#Sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Also the handling of exceptions has slightly changed in Python 3. In Python 3 we have to use the \"`as`\" keyword now" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### Python 2" + ] + }, + { + "cell_type": "code", + "execution_count": 10, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Python 2.7.6\n", + "name 'let_us_cause_a_NameError' is not defined --> our error message\n" + ] + } + ], + "source": [ + "print 'Python', python_version()\n", + "try:\n", + " let_us_cause_a_NameError\n", + "except NameError, err:\n", + " print err, '--> our error message'" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### Python 3" + ] + }, + { + "cell_type": "code", + "execution_count": 12, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Python 3.4.1\n", + "name 'let_us_cause_a_NameError' is not defined --> our error message\n" + ] + } + ], + "source": [ + "print('Python', python_version())\n", + "try:\n", + " let_us_cause_a_NameError\n", + "except NameError as err:\n", + " print(err, '--> our error message')" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## The next() function and .next() method" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to the section-overview](#Sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Since `next()` (`.next()`) is such a commonly used function (method), this is another syntax change (or rather change in implementation) that is worth mentioning: where you can use both the function and method syntax in Python 2.7.5, the `next()` function is all that remains in Python 3 (calling the `.next()` method raises an `AttributeError`)." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### Python 2" + ] + }, + { + "cell_type": "code", + "execution_count": 11, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Python 2.7.6\n" + ] + }, + { + "data": { + "text/plain": [ + "'b'" + ] + }, + "execution_count": 11, + "metadata": {}, + "output_type": "execute_result" + } + ], + "source": [ + "print 'Python', python_version()\n", + "\n", + "my_generator = (letter for letter in 'abcdefg')\n", + "\n", + "next(my_generator)\n", + "my_generator.next()" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### Python 3" + ] + }, + { + "cell_type": "code", + "execution_count": 13, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Python 3.4.1\n" + ] + }, + { + "data": { + "text/plain": [ + "'a'" + ] + }, + "execution_count": 13, + "metadata": {}, + "output_type": "execute_result" + } + ], + "source": [ + "print('Python', python_version())\n", + "\n", + "my_generator = (letter for letter in 'abcdefg')\n", + "\n", + "next(my_generator)" + ] + }, + { + "cell_type": "code", + "execution_count": 14, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "ename": "AttributeError", + "evalue": "'generator' object has no attribute 'next'", + "output_type": "error", + "traceback": [ + "\u001b[0;31m---------------------------------------------------------------------------\u001b[0m\n\u001b[0;31mAttributeError\u001b[0m Traceback (most recent call last)", + "\u001b[0;32m
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## For-loop variables and the global namespace leak" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to the section-overview](#Sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Good news is: In Python 3.x for-loop variables don't leak into the global namespace anymore!\n", + "\n", + "This goes back to a change that was made in Python 3.x and is described in [What’s New In Python 3.0](https://docs.python.org/3/whatsnew/3.0.html) as follows:\n", + "\n", + "\"List comprehensions no longer support the syntactic form `[... for var in item1, item2, ...]`. Use `[... for var in (item1, item2, ...)]` instead. Also note that list comprehensions have different semantics: they are closer to syntactic sugar for a generator expression inside a `list()` constructor, and in particular the loop control variables are no longer leaked into the surrounding scope.\"" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### Python 2" + ] + }, + { + "cell_type": "code", + "execution_count": 12, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Python 2.7.6\n", + "before: i = 1\n", + "comprehension: [0, 1, 2, 3, 4]\n", + "after: i = 4\n" + ] + } + ], + "source": [ + "print 'Python', python_version()\n", + "\n", + "i = 1\n", + "print 'before: i =', i\n", + "\n", + "print 'comprehension: ', [i for i in range(5)]\n", + "\n", + "print 'after: i =', i" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### Python 3" + ] + }, + { + "cell_type": "code", + "execution_count": 15, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Python 3.4.1\n", + "before: i = 1\n", + "comprehension: [0, 1, 2, 3, 4]\n", + "after: i = 1\n" + ] + } + ], + "source": [ + "print('Python', python_version())\n", + "\n", + "i = 1\n", + "print('before: i =', i)\n", + "\n", + "print('comprehension:', [i for i in range(5)])\n", + "\n", + "print('after: i =', i)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Comparing unorderable types" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to the section-overview](#Sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Another nice change in Python 3 is that a `TypeError` is raised as warning if we try to compare unorderable types." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### Python 2" + ] + }, + { + "cell_type": "code", + "execution_count": 2, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Python 2.7.6\n", + "[1, 2] > 'foo' = False\n", + "(1, 2) > 'foo' = True\n", + "[1, 2] > (1, 2) = False\n" + ] + } + ], + "source": [ + "print 'Python', python_version()\n", + "print \"[1, 2] > 'foo' = \", [1, 2] > 'foo'\n", + "print \"(1, 2) > 'foo' = \", (1, 2) > 'foo'\n", + "print \"[1, 2] > (1, 2) = \", [1, 2] > (1, 2)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### Python 3" + ] + }, + { + "cell_type": "code", + "execution_count": 16, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Python 3.4.1\n" + ] + }, + { + "ename": "TypeError", + "evalue": "unorderable types: list() > str()", + "output_type": "error", + "traceback": [ + "\u001b[0;31m---------------------------------------------------------------------------\u001b[0m\n\u001b[0;31mTypeError\u001b[0m Traceback (most recent call last)", + "\u001b[0;32m
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Parsing user inputs via input()" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to the section-overview](#Sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Fortunately, the `input()` function was fixed in Python 3 so that it always stores the user inputs as `str` objects. In order to avoid the dangerous behavior in Python 2 to read in other types than `strings`, we have to use `raw_input()` instead." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### Python 2" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
Python 2.7.6 \n", + "[GCC 4.0.1 (Apple Inc. build 5493)] on darwin\n", + "Type "help", "copyright", "credits" or "license" for more information.\n", + "\n", + ">>> my_input = input('enter a number: ')\n", + "\n", + "enter a number: 123\n", + "\n", + ">>> type(my_input)\n", + "<type 'int'>\n", + "\n", + ">>> my_input = raw_input('enter a number: ')\n", + "\n", + "enter a number: 123\n", + "\n", + ">>> type(my_input)\n", + "<type 'str'>\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### Python 3" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
Python 3.4.1 \n", + "[GCC 4.2.1 (Apple Inc. build 5577)] on darwin\n", + "Type "help", "copyright", "credits" or "license" for more information.\n", + "\n", + ">>> my_input = input('enter a number: ')\n", + "\n", + "enter a number: 123\n", + "\n", + ">>> type(my_input)\n", + "<class 'str'>\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Returning iterable objects instead of lists" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to the section-overview](#Sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "As we have already seen in the [`xrange`](#xrange) section, some functions and methods return iterable objects in Python 3 now - instead of lists in Python 2. \n", + "\n", + "Since we usually iterate over those only once anyway, I think this change makes a lot of sense to save memory. However, it is also possible - in contrast to generators - to iterate over those multiple times if needed, it is aonly not so efficient.\n", + "\n", + "And for those cases where we really need the `list`-objects, we can simply convert the iterable object into a `list` via the `list()` function." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### Python 2" + ] + }, + { + "cell_type": "code", + "execution_count": 2, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Python 2.7.6\n", + "[0, 1, 2]\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "**Some more commonly used functions and methods that don't return lists anymore in Python 3:**\n", + "\n", + "- `zip()`\n", + "\n", + "- `map()`\n", + "\n", + "- `filter()`\n", + "\n", + "- dictionary's `.keys()` method\n", + "\n", + "- dictionary's `.values()` method\n", + "\n", + "- dictionary's `.items()` method\n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Banker's Rounding" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to the section-overview](#Sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Python 3 adopted the now standard way of rounding decimals when it results in a tie (.5) at the last significant digits. Now, in Python 3, decimals are rounded to the nearest even number. Although it's an inconvenience for code portability, it's supposedly a better way of rounding compared to rounding up as it avoids the bias towards large numbers. For more information, see the excellent Wikipedia articles and paragraphs:\n", + "- [https://en.wikipedia.org/wiki/Rounding#Round_half_to_even](https://en.wikipedia.org/wiki/Rounding#Round_half_to_even)\n", + "- [https://en.wikipedia.org/wiki/IEEE_floating_point#Roundings_to_nearest](https://en.wikipedia.org/wiki/IEEE_floating_point#Roundings_to_nearest)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### Python 2" + ] + }, + { + "cell_type": "code", + "execution_count": 5, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Python 2.7.12\n" + ] + } + ], + "source": [ + "print 'Python', python_version()" + ] + }, + { + "cell_type": "code", + "execution_count": 6, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "data": { + "text/plain": [ + "16.0" + ] + }, + "execution_count": 6, + "metadata": {}, + "output_type": "execute_result" + } + ], + "source": [ + "round(15.5)" + ] + }, + { + "cell_type": "code", + "execution_count": 7, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "data": { + "text/plain": [ + "17.0" + ] + }, + "execution_count": 7, + "metadata": {}, + "output_type": "execute_result" + } + ], + "source": [ + "round(16.5)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### Python 3" + ] + }, + { + "cell_type": "code", + "execution_count": 4, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Python 3.5.1\n" + ] + } + ], + "source": [ + "print('Python', python_version())" + ] + }, + { + "cell_type": "code", + "execution_count": 5, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "data": { + "text/plain": [ + "16" + ] + }, + "execution_count": 5, + "metadata": {}, + "output_type": "execute_result" + } + ], + "source": [ + "round(15.5)" + ] + }, + { + "cell_type": "code", + "execution_count": 6, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "data": { + "text/plain": [ + "16" + ] + }, + "execution_count": 6, + "metadata": {}, + "output_type": "execute_result" + } + ], + "source": [ + "round(16.5)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## More articles about Python 2 and Python 3" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to the section-overview](#Sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Here is a list of some good articles concerning Python 2 and 3 that I would recommend as a follow-up.\n", + "\n", + "\n", + "**// Porting to Python 3** \n", + "\n", + "- [Should I use Python 2 or Python 3 for my development activity?](https://wiki.python.org/moin/Python2orPython3)\n", + "\n", + "- [What’s New In Python 3.0](https://docs.python.org/3.0/whatsnew/3.0.html)\n", + "\n", + "- [Porting to Python 3](http://python3porting.com/differences.html)\n", + "\n", + "- [Porting Python 2 Code to Python 3](https://docs.python.org/3/howto/pyporting.html) \n", + "\n", + "- [How keep Python 3 moving forward](http://nothingbutsnark.svbtle.com/my-view-on-the-current-state-of-python-3)\n", + "\n", + "**// Pro and anti Python 3**\n", + "\n", + "- [10 awesome features of Python that you can't use because you refuse to upgrade to Python 3](http://asmeurer.github.io/python3-presentation/slides.html#1)\n", + "\n", + "- [Everything you did not want to know about Unicode in Python 3](http://lucumr.pocoo.org/2014/5/12/everything-about-unicode/)\n", + "\n", + "- [Python 3 is killing Python](https://medium.com/@deliciousrobots/5d2ad703365d/)\n", + "\n", + "- [Python 3 can revive Python](https://medium.com/p/2a7af4788b10)\n", + "\n", + "- [Python 3 is fine](http://sealedabstract.com/rants/python-3-is-fine/)\n" + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": { + "collapsed": false + }, + "outputs": [], + "source": [] + } + ], + "metadata": { + "anaconda-cloud": {}, + "kernelspec": { + "display_name": "Python 2", + "language": "python", + "name": "python2" + }, + "language_info": { + "codemirror_mode": { + "name": "ipython", + "version": 2 + }, + "file_extension": ".py", + "mimetype": "text/x-python", + "name": "python", + "nbconvert_exporter": "python", + "pygments_lexer": "ipython2", + "version": "2.7.12" + } + }, + "nbformat": 4, + "nbformat_minor": 0 +} diff --git a/tutorials/markdown_syntax_highlighting/README.md b/tutorials/markdown_syntax_highlighting/README.md new file mode 100644 index 0000000..0148ea1 --- /dev/null +++ b/tutorials/markdown_syntax_highlighting/README.md @@ -0,0 +1,209 @@ +[Sebastian Raschka](http://sebastianraschka.com) + +last updated: 05/28/2014 + +
+I would be happy to hear your comments and suggestions. +Please feel free to drop me a note via +[twitter](https://twitter.com/rasbt), [email](mailto:bluewoodtree@gmail.com), or [google+](https://plus.google.com/118404394130788869227). +
+ + +# 5 simple steps for converting Markdown documents into HTML and adding Python syntax highlighting + +
+
+ +In this little tutorial, I want to show you in 5 simple steps how easy it is to add code syntax highlighting to your blog articles. + +There are more sophisticated approaches using static site generators, e.g., [nikola](https://github.com/getnikola/nikola), but the focus here is to give you the brief introduction of how it generally works. + + +All the files I will be using as examples in this tutorial can be download from the GitHub repository [/rasbt/python_reference/tutorials/markdown_syntax_highlighting](https://github.com/rasbt/python_reference/tree/master/tutorials/markdown_syntax_highlighting) + +
+
+ +##1 - Installing packages + +The two packages that we will use are + +- [Python-Markdown](http://pythonhosted.org/Markdown/) + +- [Pygments](http://pygments.org) + +Just as the name suggests, Python-Markdown is the Python package that we will use for the Markdown to HTML conversion. +The second library, Pygments, will be used to add the syntax highlighting to the code blocks. +Conveniently, both libraries can be installed via `pip`: + + + pip install markdown + +and + + pip install Pygments + + +(For alternative ways to install the Python-Markdown package, please see [the +documentation](http://pythonhosted.org/Markdown/install.html)) + + +
+
+ +##2 - Writing a Markdown document + +Now, let us compose a simple Markdown document including some Python code blocks in any/our favorite Markdown editor. + + +--> [**some_markdown.md**](https://github.com/rasbt/python_reference/blob/master/tutorials/markdown_syntax_highlighting/some_markdown.md) + +
+
+##This is a test
+
+Code blocks must be indented by 4 whitespaces.
+Python-Markdown has a auto-guess function which works
+pretty well:
+
+ print("Hello, World")
+ # some comment
+ for letter in "this is a test":
+ print(letter)
+
+In cases where Python-Markdown has problems figuring out which
+programming language we use, we can also add the language-tag
+explicitly. One way to do this would be:
+
+ :::python
+ print("Hello, World")
+
+or we can highlight certain lines to
+draw the reader's attention:
+
+ :::python hl_lines="1 5"
+ print("highlight me!")
+ # but not me!
+ for letter in "this is a test":
+ print(letter)
+ # I want to be highlighted, too!
+
+
++
+ +Note that the syntax highlighting does not only work for Python, but other programming languages. + +So in the case of C++, for example: + + :::c++ + #include
+
+ +## 3 - Converting the Markdown document to HTML + + +After we created our Markdown document, we are going to use Python-Markdown directly from the command line to convert it into an HTML document. + +Note that we can also import Python-Markdown as a module in our Python scripts, and it comes with a rich repertory of different functions, which are [listed in the library reference](https://pythonhosted.org/Markdown/reference.html). + +The basic command line usage to convert a Markdown document into HTML would be: + + python -m markdown input.md > output.html + +However, since we want to have syntax highlighting for our Python code, we will use Python-Markdown's [CodeHilite extension](http://pythonhosted.org/Markdown/extensions/code_hilite.html) by providing an additional `-x codehilite` argument on the command line: + + + python -m markdown -x codehilite some_markdown.md > body.html + +This will create the HTML body with our Markdown code converted to HTML with the Python code blocks annotated for the syntax highlighting. + + +
+
+ +##4 - Generating the CSS + +If we open the [**body.html**](https://github.com/rasbt/python_reference/blob/master/tutorials/markdown_syntax_highlighting/body.html) file now, which we have created in the previous section, we will notice that it doesn't have the Python code colored yet. + + + +What is missing is the CSS code for adding the colors to our annotated Python code block. But we can simply create such a CSS file via `Pygments` from the command line. + + pygmentize -S default -f html > codehilite.css + +Note that we usually only need to create the [**codehilite.css**](https://github.com/rasbt/python_reference/blob/master/tutorials/markdown_syntax_highlighting/codehilite.css) file once and insert a link in all our HTML files that we created via Python-Markdown to get the syntax coloring + + +
+
+ + +## 5 - Insert into your HTML body + + +In order to include a link to the [codehilite.css](https://github.com/rasbt/python_reference/blob/master/tutorials/markdown_syntax_highlighting/codehilite.css) file for syntax coloring in our converted HTML file, we have to add the following line to the header section. + + + +`` + + +Now, we can insert the HTML body ([body.html](https://github.com/rasbt/python_reference/blob/master/tutorials/markdown_syntax_highlighting/body.html)), which was created from our Markdown document, directly into our final HTML file (e.g., our blog article template). + + +[**template.html**](https://github.com/rasbt/python_reference/blob/master/tutorials/markdown_syntax_highlighting/template.html): + +
+
+
+
+
+
+
+
+
+
+
+
+ <-- converted HTML contents go here
+
+
+
+
+
+
+If we open our [**final.html**](https://github.com/rasbt/python_reference/blob/master/tutorials/markdown_syntax_highlighting/template.html) file in our web browser now, we can the pretty Python syntax highlighting.
+
+
+
++
+ +## Useful links: + + +- [Python Markdown package documentation](http://pythonhosted.org//Markdown/) + +- [The CodeHilite documentation](https://pythonhosted.org/Markdown/extensions/code_hilite.html) + +- [pygments.org](http://pygments.org) + +- [languages supported](http://pygments.org/languages/) by Pygments + + diff --git a/tutorials/markdown_syntax_highlighting/body.html b/tutorials/markdown_syntax_highlighting/body.html new file mode 100644 index 0000000..e768c67 --- /dev/null +++ b/tutorials/markdown_syntax_highlighting/body.html @@ -0,0 +1,26 @@ +
This is a test
+Code blocks must be indented by 4 whitespaces. +Python-Markdown has a auto-guess function which works +pretty well:
+print("Hello, World") +# some comment +for letter in "this is a test": + print(letter) +
In cases where Python-Markdown has problems figuring out which +programming language we use, we can also add the language-tag +explicitly. One way to do this would be:
+print("Hello, World") +
or we can highlight certain lines to +draw the reader's attention:
+print("highlight me!") +# but not me! +for letter in "this is a test": + print(letter) +# I want to be highlighted, too! +
This is a test
+Code blocks must be indented by 4 whitespaces. +Python-Markdown has a auto-guess function which works +pretty well:
+print("Hello, World") +# some comment +for letter in "this is a test": + print(letter) +
In cases where Python-Markdown has problems figuring out which +programming language we use, we can also add the language-tag +explicitly. One way to do this would be:
+print("Hello, World") +
or we can highlight certain lines to +draw the reader's attention:
+print("highlight me!") +# but not me! +for letter in "this is a test": + print(letter) +# I want to be highlighted, too! +
+ + + + +## Introduction + +[[back to section overview](#sections)] +
+ +Matrices (or multidimensional arrays) are not only presenting the fundamental elements of many algebraic equations that are used in many popular fields, such as pattern classification, machine learning, data mining, and math and engineering in general. But in context of scientific computing, they also come in very handy for managing and storing data in an more organized tabular form. +Such multidimensional data structures are also very powerful performance-wise thanks to the concept of automatic vectorization: instead of the individual and sequential processing of operations on scalars in loop-structures, the whole computation can be parallelized in order to make optimal use of modern computer architectures. + + +
+ + + +
+
+ + +
+**Note:** +This article originated from an older article with containing a cheat sheet that was just about MATLAB matrices and NumPy arrays. Since then, I added a couple of more rows and doubled the width of the cheat sheet by adding those two other languages R and Julia. Instead of making further modifications, I wanted to keep this old article as is - for future reference and for people who may only be interested in this slimmer version: +[Moving from MATLAB matrices to NumPy arrays - A Matrix Cheatsheet](http://sebastianraschka.com/Articles/2014_matlab_vs_numpy.html). +
+ + +
+
+ + + +### Language overview + +[[back to section overview](#sections)] +
+ +Before we **[jump to the actual cheat sheet](#cheatsheet)**, I wanted to give you at least a brief overview of the different languages that we are dealing with. + + +All four languages, MATLAB/Octave, Python, R, and Julia are dynamically typed, have a command line interface for the interpreter, and come with great number of additional and useful libraries to support scientific and technical computing. Conveniently, these languages also offer great solutions for easy plotting and visualizations. + +Combined with interactive notebook interfaces or dynamic report generation engines ([MuPAD](http://www.mathworks.com/discovery/mupad.html) for MATLAB, [IPython Notebook](http://ipython.org/notebook.html) for Python, [knitr](http://yihui.name/knitr/) for R, and [IJulia](https://github.com/JuliaLang/IJulia.jl) for Julia based on IPython Notebook) data analysis and documentation has never been easier. + +
+
+ + + +# MATLAB/Octave + +[[back to section overview](#sections)] +
+ +
+ + + + +
+ +[MATLAB](http://www.mathworks.com/products/matlab/) (stands for MATrix LABoratory) is the name of an application and language that was developed by [MathWorks](http://www.mathworks.com/index.html?s_tid=gn_logo) back in 1984. One of its strengths is the variety of different and highly optimized "toolboxes" (including very powerful functions for image and other signal processing task), which makes suitable for tackling basically every possible science and engineering task. +Like the other languages, which will be covered in this article, it has cross-platform support and is using dynamic types, which allows for a convenient interface, but can also be quite "memory hungry" for computations on large data sets. + +Even today, MATLAB is probably (still) the most popular language for numeric computation used for engineering tasks in academia as well as in industry. + +#### GNU Octave + +
+ + + + +
+ +It is also worth mentioning that MATLAB is the only language in this cheat sheet which is not free and open-sourced. But since it is so immensely popular, I want to mention it nonetheless. And as an alternative there is also the free [GNU Octave re-implementation](http://www.gnu.org/software/octave/) that follows the same syntactic rules so that the code is compatible to MATLAB (except for very specialized libraries). + +
+ +* This [image](http://commons.wikimedia.org/wiki/File:Matlab_Logo.png) is a freely usable media under public domain and represents the first eigenfunction of the L-shaped membrane, resembling (but not identical to) MATLAB's logo trademarked by MathWorks Inc. + +
+
+ + + + + + +# Python NumPy + +[[back to section overview](#sections)] +
+ +
+ + +
+ +Initially, the [NumPy](http://www.numpy.org) project started out under the name "Numeric" in 1995 (renamed to NumPy in 2006) as a Python library for numeric computations based on multi-dimensional data structures, such as arrays and matrices. Since it makes use of pre-compiled C code for operations on its "`ndarray`" objects, it is considerably faster than using equivalent approaches in (C)Python. + + +Python NumPy is my personal favorite since I am a big fan of the Python programming language. Although similar tools exist for other languages, I found myself to be most productive doing my research and data analyses in [IPython notebooks](http://ipython.org/notebook.html). +It allows me to easily combine Python code (sometimes optimized by compiling it via the [Cython](http://cython.org) C-Extension or the just-in-time (JIT) [Numba](http://numba.pydata.org) compiler if speed is a concern) with different libraries from the [Scipy stack](http://www.scipy.org/) including [matplotlib](http://matplotlib.org) for inline data visualization (you can find some of my example benchmarks in this [GitHub repository](http://github.com/rasbt/One-Python-benchmark-per-day)). + +
+
+ + + +# R + +[[back to section overview](#sections)] +
+ +
+ + +
+ +The [R](http://www.r-project.org) programming language was developed in 1993 and is a modern GNU implementation of an older statistical programming language called [S](http://stat.bell-labs.com/S/), which was developed in the [Bell Laboratories](http://stat.bell-labs.com) in 1976. Since its release, it has a fast-growing user base and is particularly popular among statisticians. + +R was also the first language which kindled my fascination for statistics and computing. I have used it quite extensively a couple of years ago before I discovered Python as my new favorite language for data analysis. +Although R has great in-built functions for performing all sorts statistics, as well as a plethora of freely available libraries developed by the large R community, I often hear people complaining about its rather unintuitive syntax. + +
+
+ + + +# Julia + +[[back to section overview](#sections)] +
+ +
+ + + + +
+ + +With its first release in 2012, [Julia](http://julialang.org) is by far the youngest of the programming languages mentioned in this article. a +While Julia can also be used as an interpreted language with dynamic types from the command line, it aims for high-performance in scientific computing that is superior to the other dynamic programming languages for technical computing thanks to its LLVM-based just-in-time (JIT) compiler. + +Personally, I haven't used Julia that extensively, yet, but there are some exciting benchmarks that look very promising: + + + + + + +C compiled by gcc 4.8.1, taking best timing from all optimization levels (-O0 through -O3). C, Fortran and Julia use OpenBLAS v0.2.8. The Python implementations of rand_mat_stat and rand_mat_mul use NumPy (v1.6.1) functions; the rest are pure Python implementations.
+ +Bezanson, J., Karpinski, S., Shah, V.B. and Edelman, A. (2012), “Julia: A fast dynamic language for technical computing”. +(Source: [http://julialang.org/benchmarks/](http://julialang.org/benchmarks/), with permission from the copyright holder) + + +
+
+
+ +
+ + + +# Cheat sheet + +[[back to section overview](#sections)] + + +
+ + + + +### Cheat sheet overview + + +- [Creating matrices](#creating) + +- [Accessing matrix elements](#accessing) + +- [Manipulating shape](#manipulating) + +- [Basic matrix operations](#basic_ops) + +- [Advanced matrix operations](#advanced_ops) + +
+
+
+
+ + +**If you are interested in downloading this cheat sheet table for your references, you can find it [here on GitHub](https://github.com/rasbt/python_reference/blob/master/tutorials/matrix_cheatsheet_only.html)** + +
+
+ + + +
|
+ Task + |
+
+ MATLAB/Octave + |
+
+ Python + NumPy + |
+
+ R + |
+
+ Julia + |
+
+ Task + |
+
|
+ CREATING + MATRICES + |
+ |||||
|
+ Creating
+ Matrices |
+
+ M>
+ A = [1 2 3; 4 5 6; 7 8 9] |
+
+ P>
+ A = np.array([ [1,2,3], [4,5,6], [7,8,9] ]) |
+
+ R>
+ A = matrix(c(1,2,3,4,5,6,7,8,9),nrow=3,byrow=T) |
+
+ J>
+ A=[1 2 3; 4 5 6; 7 8 9] |
+
+ Creating
+ Matrices |
+
|
+ Creating + an 1D column vector + |
+
+ M>
+ a = [1; 2; 3] |
+
+ P> + a + = + np.array([1,2,3]).reshape(1,3) +
|
+
+ R>
+ a = matrix(c(1,2,3), nrow=3, byrow=T) |
+
+ J>
+ a=[1; 2; 3] |
+
+ Creating + an 1D column vector + |
+
|
+ Creating
+ an |
+
+ M>
+ b = [1 2 3] |
+
+ P>
+ b = np.array([1,2,3]) #
+ note that numpy doesn't have P> + b.shape +(3,) +
|
+
+ R>
+ b = matrix(c(1,2,3), ncol=3) |
+
+ J>
+ b=[1 2 3] |
+
+ Creating
+ an |
+
|
+ Creating
+ a |
+
+ M>
+ rand(3,2) |
+
+ P>
+ np.random.rand(3,2) |
+
+ R>
+ matrix(runif(3*2), ncol=2) |
+
+ J>
+ rand(3,2) |
+
+ Creating
+ a |
+
|
+ Creating
+ a |
+
+ M>
+ zeros(3,2) |
+
+ P>
+ np.zeros((3,2)) |
+
+ R>
+ mat.or.vec(3, 2) |
+
+ J>
+ zeros(3,2) |
+
+ Creating
+ a |
+
|
+ Creating
+ an |
+
+ M>
+ ones(3,2) |
+
+ P>
+ np.ones((3,2)) |
+
+ R>
+ mat.or.vec(3, 2) + 1 |
+
+ J>
+ ones(3,2) |
+
+ Creating
+ an |
+
|
+ Creating
+ an |
+
+ M>
+ eye(3) |
+
+ P>
+ np.eye(3) |
+
+ R>
+ diag(3) |
+
+ J>
+ eye(3) |
+
+ Creating
+ an |
+
|
+ Creating
+ a |
+
+ M>
+ a = [1 2 3] |
+
+ P>
+ a = np.array([1,2,3]) |
+
+ R>
+ diag(1:3) |
+
+ J>
+ a=[1, 2, 3] |
+
+ Creating
+ a |
+
|
+ ACCESSING + MATRIX ELEMENTS + |
+ |||||
|
+ Getting
+ the dimension |
+
+ M>
+ A = [1 2 3; 4 5 6] |
+
+ P>
+ A = np.array([ [1,2,3], [4,5,6] ]) |
+
+ R>
+ A = matrix(1:6,nrow=2,byrow=T) R>
+ dim(A) |
+
+ J>
+ A=[1 2 3; 4 5 6] |
+
+ Getting
+ the dimension |
+
|
+ Selecting + rows + |
+
+ M>
+ A = [1 2 3; 4 5 6; 7 8 9] |
+
+ P>
+ A = np.array([ [1,2,3], [4,5,6], [7,8,9] ]) |
+
+ R>
+ A = matrix(1:9,nrow=3,byrow=T)
|
+
+ J>
+ A=[1 2 3; 4 5 6; 7 8 9]; |
+
+ Selecting + rows + |
+
|
+ Selecting + columns + |
+
+ M>
+ A = [1 2 3; 4 5 6; 7 8 9] |
+
+ P>
+ A = np.array([ [1,2,3], [4,5,6], [7,8,9] ]) |
+
+ R>
+ A = matrix(1:9,nrow=3,byrow=T)
|
+
+ J>
+ A=[1 2 3; 4 5 6; 7 8 9]; |
+
+ Selecting + columns + |
+
|
+ Extracting
+ rows and columns by criteria |
+
+ M>
+ A = [1 2 3; 4 5 9; 7 8 9] |
+
+ P>
+ A = np.array([ [1,2,3], [4,5,9], [7,8,9]]) |
+
+ R>
+ A = matrix(1:9,nrow=3,byrow=T)
|
+
+ J>
+ A=[1 2 3; 4 5 9; 7 8 9] |
+
+ Extracting
+ rows and columns by criteria |
+
|
+ Accessing
+ elements |
+
+ M>
+ A = [1 2 3; 4 5 6; 7 8 9] |
+
+ P>
+ A = np.array([ [1,2,3], [4,5,6], [7,8,9] ]) |
+
+ R>
+ A = matrix(c(1,2,3,4,5,9,7,8,9),nrow=3,byrow=T)
|
+
+ J>
+ A=[1 2 3; 4 5 6; 7 8 9]; |
+
+ Accessing
+ elements |
+
|
+ MANIPULATING + SHAPE AND DIMENSIONS + |
+ |||||
|
+ Converting |
+
+ M>
+ b = [1 2 3]
|
+
+ P>
+ b = np.array([1, 2, 3]) |
+
+ R>
+ b = matrix(c(1,2,3), ncol=3) |
+
+ J>
+ b=vec([1 2 3]) |
+
+ Converting |
+
|
+ Reshaping
+ Matrices |
+
+ M>
+ A = [1 2 3; 4 5 6; 7 8 9] |
+
+ P>
+ A = np.array([[1,2,3],[4,5,6],[7,8,9]]) P>
+ B = A.reshape(1, total_elements) |
+
+ R>
+ A = matrix(1:9,nrow=3,byrow=T)
|
+
+ J>
+ A=[1 2 3; 4 5 6; 7 8 9] |
+
+ Reshaping
+ Matrices |
+
|
+ Concatenating + matrices + |
+
+ M>
+ A = [1 2 3; 4 5 6] |
+
+ P>
+ A = np.array([[1, 2, 3], [4, 5, 6]]) |
+
+ R>
+ A = matrix(1:6,nrow=2,byrow=T) |
+
+ J>
+ A=[1 2 3; 4 5 6]; |
+
+ Concatenating + matrices + |
+
|
+ Stacking |
+
+ M>
+ a = [1 2 3] |
+
+ P>
+ a = np.array([1,2,3]) |
+
+ R>
+ a = matrix(1:3, ncol=3) |
+
+ J>
+ a=[1 2 3]; |
+
+ Stacking |
+
|
+ BASIC + MATRIX OPERATIONS + |
+ |||||
|
+ Matrix-scalar |
+
+ M> A
+ = [1 2 3; 4 5 6; 7 8 9] |
+
+ P>
+ A = np.array([ [1,2,3], [4,5,6], [7,8,9] ]) #
+ Note that NumPy was optimized for |
+
+ R>
+ A = matrix(1:9, nrow=3, byrow=T) R>
+ A + 2 |
+
+ J>
+ A=[1 2 3; 4 5 6; 7 8 9]; |
+
+ Matrix-scalar |
+
|
+ Matrix-matrix |
+
+ M> A
+ = [1 2 3; 4 5 6; 7 8 9] |
+
+ P>
+ A = np.array([ [1,2,3], [4,5,6], [7,8,9] ]) |
+
+ R>
+ A = matrix(1:9, nrow=3, byrow=T) |
+
+ J>
+ A=[1 2 3; 4 5 6; 7 8 9]; |
+
+ Matrix-matrix |
+
|
+ Matrix-vector |
+
+ M>
+ A = [1 2 3; 4 5 6; 7 8 9] |
+
+ P>
+ A = np.array([ [1,2,3], [4,5,6], [7,8,9] ]) |
+
+ R>
+ A = matrix(1:9, ncol=3) |
+
+ J>
+ A=[1 2 3; 4 5 6; 7 8 9]; |
+
+ Matrix-vector |
+
|
+ Element-wise |
+
+ M> A
+ = [1 2 3; 4 5 6; 7 8 9] |
+
+ P>
+ A = np.array([ [1,2,3], [4,5,6], [7,8,9] ]) #
+ Note that NumPy was optimized for |
+
+ R>
+ A = matrix(1:9, nrow=3, byrow=T)
|
+
+ J>
+ A=[1 2 3; 4 5 6; 7 8 9]; |
+
+ Element-wise |
+
|
+ Matrix
+ elements to power n |
+
+ M> A
+ = [1 2 3; 4 5 6; 7 8 9] |
+
+ P>
+ A = np.array([ [1,2,3], [4,5,6], [7,8,9] ]) |
+
+ R>
+ A = matrix(1:9, nrow=3, byrow=T) |
+
+ J>
+ A=[1 2 3; 4 5 6; 7 8 9]; |
+
+ Matrix
+ elements to power n |
+
|
+ Matrix
+ to power n |
+
+ M> A
+ = [1 2 3; 4 5 6; 7 8 9] |
+
+ P>
+ A = np.array([ [1,2,3], [4,5,6], [7,8,9] ]) |
+
+ R>
+ A = matrix(1:9, ncol=3) |
+
+ J>
+ A=[1 2 3; 4 5 6; 7 8 9]; |
+
+ Matrix
+ to power n |
+
|
+ Matrix + transpose + |
+
+ M> A
+ = [1 2 3; 4 5 6; 7 8 9] |
+
+ P>
+ A = np.array([ [1,2,3], [4,5,6], [7,8,9] ]) |
+
+ R>
+ A = matrix(1:9, nrow=3, byrow=T)
|
+
+ J>
+ A=[1 2 3; 4 5 6; 7 8 9] |
+
+ Matrix + transpose + |
+
|
+ Determinant
+ of a matrix: |
+
+ M>
+ A = [6 1 1; 4 -2 5; 2 8 7] |
+
+ P> A
+ = np.array([[6,1,1],[4,-2,5],[2,8,7]]) |
+
+ R>
+ A = matrix(c(6,1,1,4,-2,5,2,8,7), nrow=3, byrow=T) |
+
+ J>
+ A=[6 1 1; 4 -2 5; 2 8 7] |
+
+ Determinant
+ of a matrix: |
+
|
+ Inverse + of a matrix + |
+
+ M>
+ A = [4 7; 2 6] |
+
+ P>
+ A = np.array([[4, 7], [2, 6]]) |
+
+ R>
+ A = matrix(c(4,7,2,6), nrow=2, byrow=T) |
+
+ J>
+ A=[4 7; 2 6] |
+
+ Inverse + of a matrix + |
+
|
+ ADVANCED + MATRIX OPERATIONS + |
+ |||||
|
+ Calculating
+ the covariance matrix |
+
+ M>
+ x1 = [4.0000 4.2000 3.9000 4.3000 4.1000]’ |
+
+ P>
+ x1 = np.array([ 4, 4.2, 3.9, 4.3, 4.1]) |
+
+ R>
+ x1 = matrix(c(4, 4.2, 3.9, 4.3, 4.1), ncol=5) |
+
+ J>
+ x1=[4.0 4.2 3.9 4.3 4.1]'; |
+
+ Calculating
+ the covariance matrix |
+
|
+ Calculating |
+
+ M>
+ A = [3 1; 1 3] |
+
+ P>
+ A = np.array([[3, 1], [1, 3]]) |
+
+ R>
+ A = matrix(c(3,1,1,3), ncol=2) |
+
+ J>
+ A=[3 1; 1 3] |
+
+ Calculating |
+
|
+ Generating
+ a Gaussian dataset: |
+
+ %
+ requires statistics toolbox package |
+
+ P>
+ mean = np.array([0,0]) |
+
+ #
+ requires the ‘mass’ package |
+
+ #
+ requires the Distributions package from
+ https://github.com/JuliaStats/Distributions.jl |
+
+ Generating
+ a Gaussian dataset: |
+
+ +
+(Thanks to Keith C. Campbell for providing me with the syntax for the Julia language.) + + +
+ + + + +
+
+
+
+ +### Alternative data structures: NumPy matrices vs. NumPy arrays + +[[back to section overview](#sections)] + +Python's NumPy library also has a dedicated "matrix" type with a syntax that is a little bit closer to the MATLAB matrix: For example, the "
* " operator would perform a matrix-matrix multiplication of NumPy matrices - same operator performs element-wise multiplication on NumPy arrays.
+Vice versa, the "`.dot()`" method is used for element-wise multiplication of NumPy matrices, wheras the equivalent operation would for NumPy arrays would be achieved via the " * "-operator.
+**Most people recommend the usage of the NumPy array type over NumPy matrices, since arrays are what most of the NumPy functions return.**
\ No newline at end of file
diff --git a/tutorials/matrix_cheatsheet_only.html b/tutorials/matrix_cheatsheet_only.html
new file mode 100644
index 0000000..8d9762c
--- /dev/null
+++ b/tutorials/matrix_cheatsheet_only.html
@@ -0,0 +1,1206 @@
+
+
+
+
+ |
+ Task + |
+
+ MATLAB/Octave + |
+
+ Python + NumPy + |
+
+ R + |
+
+ Julia + |
+
+ Task + |
+
|
+ CREATING + MATRICES + |
+ |||||
|
+ Creating
+ Matrices |
+
+ M>
+ A = [1 2 3; 4 5 6; 7 8 9] |
+
+ P>
+ A = np.array([ [1,2,3], [4,5,6], [7,8,9] ]) |
+
+ R>
+ A = matrix(c(1,2,3,4,5,6,7,8,9),nrow=3,byrow=T) |
+
+ J>
+ A=[1 2 3; 4 5 6; 7 8 9] |
+
+ Creating
+ Matrices |
+
|
+ Creating + an 1D column vector + |
+
+ M>
+ a = [1; 2; 3] |
+
+ P> + a + = + np.array([1,2,3]).reshape(1,3) +
|
+
+ R>
+ a = matrix(c(1,2,3), nrow=3, byrow=T) |
+
+ J>
+ a=[1; 2; 3] |
+
+ Creating + an 1D column vector + |
+
|
+ Creating
+ an |
+
+ M>
+ b = [1 2 3] |
+
+ P>
+ b = np.array([1,2,3]) #
+ note that numpy doesn't have P> + b.shape +(3,) +
|
+
+ R>
+ b = matrix(c(1,2,3), ncol=3) |
+
+ J>
+ b=[1 2 3] |
+
+ Creating
+ an |
+
|
+ Creating
+ a |
+
+ M>
+ rand(3,2) |
+
+ P>
+ np.random.rand(3,2) |
+
+ R>
+ matrix(runif(3*2), ncol=2) |
+
+ J>
+ rand(3,2) |
+
+ Creating
+ a |
+
|
+ Creating
+ a |
+
+ M>
+ zeros(3,2) |
+
+ P>
+ np.zeros((3,2)) |
+
+ R>
+ mat.or.vec(3, 2) |
+
+ J>
+ zeros(3,2) |
+
+ Creating
+ a |
+
|
+ Creating
+ an |
+
+ M>
+ ones(3,2) |
+
+ P>
+ np.ones((3,2)) |
+
+ R>
+ mat.or.vec(3, 2) + 1 |
+
+ J>
+ ones(3,2) |
+
+ Creating
+ an |
+
|
+ Creating
+ an |
+
+ M>
+ eye(3) |
+
+ P>
+ np.eye(3) |
+
+ R>
+ diag(3) |
+
+ J>
+ eye(3) |
+
+ Creating
+ an |
+
|
+ Creating
+ a |
+
+ M>
+ a = [1 2 3] |
+
+ P>
+ a = np.array([1,2,3]) |
+
+ R>
+ diag(1:3) |
+
+ J>
+ a=[1, 2, 3] |
+
+ Creating
+ a |
+
|
+ ACCESSING + MATRIX ELEMENTS + |
+ |||||
|
+ Getting
+ the dimension |
+
+ M>
+ A = [1 2 3; 4 5 6] |
+
+ P>
+ A = np.array([ [1,2,3], [4,5,6] ]) |
+
+ R>
+ A = matrix(1:6,nrow=2,byrow=T) R>
+ dim(A) |
+
+ J>
+ A=[1 2 3; 4 5 6] |
+
+ Getting
+ the dimension |
+
|
+ Selecting + rows + |
+
+ M>
+ A = [1 2 3; 4 5 6; 7 8 9] |
+
+ P>
+ A = np.array([ [1,2,3], [4,5,6], [7,8,9] ]) |
+
+ R>
+ A = matrix(1:9,nrow=3,byrow=T)
|
+
+ J>
+ A=[1 2 3; 4 5 6; 7 8 9]; |
+
+ Selecting + rows + |
+
|
+ Selecting + columns + |
+
+ M>
+ A = [1 2 3; 4 5 6; 7 8 9] |
+
+ P>
+ A = np.array([ [1,2,3], [4,5,6], [7,8,9] ]) |
+
+ R>
+ A = matrix(1:9,nrow=3,byrow=T)
|
+
+ J>
+ A=[1 2 3; 4 5 6; 7 8 9]; |
+
+ Selecting + columns + |
+
|
+ Extracting
+ rows and columns by criteria |
+
+ M>
+ A = [1 2 3; 4 5 9; 7 8 9] |
+
+ P>
+ A = np.array([ [1,2,3], [4,5,9], [7,8,9]]) |
+
+ R>
+ A = matrix(1:9,nrow=3,byrow=T)
|
+
+ J>
+ A=[1 2 3; 4 5 9; 7 8 9] |
+
+ Extracting
+ rows and columns by criteria |
+
|
+ Accessing
+ elements |
+
+ M>
+ A = [1 2 3; 4 5 6; 7 8 9] |
+
+ P>
+ A = np.array([ [1,2,3], [4,5,6], [7,8,9] ]) |
+
+ R>
+ A = matrix(c(1,2,3,4,5,9,7,8,9),nrow=3,byrow=T)
|
+
+ J>
+ A=[1 2 3; 4 5 6; 7 8 9]; |
+
+ Accessing
+ elements |
+
|
+ MANIPULATING + SHAPE AND DIMENSIONS + |
+ |||||
|
+ Converting |
+
+ M>
+ b = [1 2 3]
|
+
+ P>
+ b = np.array([1, 2, 3]) |
+
+ R>
+ b = matrix(c(1,2,3), ncol=3) |
+
+ J>
+ b=vec([1 2 3]) |
+
+ Converting |
+
|
+ Reshaping
+ Matrices |
+
+ M>
+ A = [1 2 3; 4 5 6; 7 8 9] |
+
+ P>
+ A = np.array([[1,2,3],[4,5,6],[7,8,9]]) P>
+ B = A.reshape(1, total_elements) |
+
+ R>
+ A = matrix(1:9,nrow=3,byrow=T)
|
+
+ J>
+ A=[1 2 3; 4 5 6; 7 8 9] |
+
+ Reshaping
+ Matrices |
+
|
+ Concatenating + matrices + |
+
+ M>
+ A = [1 2 3; 4 5 6] |
+
+ P>
+ A = np.array([[1, 2, 3], [4, 5, 6]]) |
+
+ R>
+ A = matrix(1:6,nrow=2,byrow=T) |
+
+ J>
+ A=[1 2 3; 4 5 6]; |
+
+ Concatenating + matrices + |
+
|
+ Stacking |
+
+ M>
+ a = [1 2 3] |
+
+ P>
+ a = np.array([1,2,3]) |
+
+ R>
+ a = matrix(1:3, ncol=3) |
+
+ J>
+ a=[1 2 3]; |
+
+ Stacking |
+
|
+ BASIC + MATRIX OPERATIONS + |
+ |||||
|
+ Matrix-scalar |
+
+ M> A
+ = [1 2 3; 4 5 6; 7 8 9] |
+
+ P>
+ A = np.array([ [1,2,3], [4,5,6], [7,8,9] ]) #
+ Note that NumPy was optimized for |
+
+ R>
+ A = matrix(1:9, nrow=3, byrow=T) R>
+ A + 2 |
+
+ J>
+ A=[1 2 3; 4 5 6; 7 8 9]; |
+
+ Matrix-scalar |
+
|
+ Matrix-matrix |
+
+ M> A
+ = [1 2 3; 4 5 6; 7 8 9] |
+
+ P>
+ A = np.array([ [1,2,3], [4,5,6], [7,8,9] ]) |
+
+ R>
+ A = matrix(1:9, nrow=3, byrow=T) |
+
+ J>
+ A=[1 2 3; 4 5 6; 7 8 9]; |
+
+ Matrix-matrix |
+
|
+ Matrix-vector |
+
+ M>
+ A = [1 2 3; 4 5 6; 7 8 9] |
+
+ P>
+ A = np.array([ [1,2,3], [4,5,6], [7,8,9] ]) |
+
+ R>
+ A = matrix(1:9, ncol=3) |
+
+ J>
+ A=[1 2 3; 4 5 6; 7 8 9]; |
+
+ Matrix-vector |
+
|
+ Element-wise |
+
+ M> A
+ = [1 2 3; 4 5 6; 7 8 9] |
+
+ P>
+ A = np.array([ [1,2,3], [4,5,6], [7,8,9] ]) #
+ Note that NumPy was optimized for |
+
+ R>
+ A = matrix(1:9, nrow=3, byrow=T)
|
+
+ J>
+ A=[1 2 3; 4 5 6; 7 8 9]; |
+
+ Element-wise |
+
|
+ Matrix
+ elements to power n |
+
+ M> A
+ = [1 2 3; 4 5 6; 7 8 9] |
+
+ P>
+ A = np.array([ [1,2,3], [4,5,6], [7,8,9] ]) |
+
+ R>
+ A = matrix(1:9, nrow=3, byrow=T) |
+
+ J>
+ A=[1 2 3; 4 5 6; 7 8 9]; |
+
+ Matrix
+ elements to power n |
+
|
+ Matrix
+ to power n |
+
+ M> A
+ = [1 2 3; 4 5 6; 7 8 9] |
+
+ P>
+ A = np.array([ [1,2,3], [4,5,6], [7,8,9] ]) |
+
+ R>
+ A = matrix(1:9, ncol=3) |
+
+ J>
+ A=[1 2 3; 4 5 6; 7 8 9]; |
+
+ Matrix
+ to power n |
+
|
+ Matrix + transpose + |
+
+ M> A
+ = [1 2 3; 4 5 6; 7 8 9] |
+
+ P>
+ A = np.array([ [1,2,3], [4,5,6], [7,8,9] ]) |
+
+ R>
+ A = matrix(1:9, nrow=3, byrow=T)
|
+
+ J>
+ A=[1 2 3; 4 5 6; 7 8 9] |
+
+ Matrix + transpose + |
+
|
+ Determinant
+ of a matrix: |
+
+ M>
+ A = [6 1 1; 4 -2 5; 2 8 7] |
+
+ P> A
+ = np.array([[6,1,1],[4,-2,5],[2,8,7]]) |
+
+ R>
+ A = matrix(c(6,1,1,4,-2,5,2,8,7), nrow=3, byrow=T) |
+
+ J>
+ A=[6 1 1; 4 -2 5; 2 8 7] |
+
+ Determinant
+ of a matrix: |
+
|
+ Inverse + of a matrix + |
+
+ M>
+ A = [4 7; 2 6] |
+
+ P>
+ A = np.array([[4, 7], [2, 6]]) |
+
+ R>
+ A = matrix(c(4,7,2,6), nrow=2, byrow=T) |
+
+ J>
+ A=[4 7; 2 6] |
+
+ Inverse + of a matrix + |
+
|
+ ADVANCED + MATRIX OPERATIONS + |
+ |||||
|
+ Calculating
+ the covariance matrix |
+
+ M>
+ x1 = [4.0000 4.2000 3.9000 4.3000 4.1000]’ |
+
+ P>
+ x1 = np.array([ 4, 4.2, 3.9, 4.3, 4.1]) |
+
+ R>
+ x1 = matrix(c(4, 4.2, 3.9, 4.3, 4.1), ncol=5) |
+
+ J>
+ x1=[4.0 4.2 3.9 4.3 4.1]'; |
+
+ Calculating
+ the covariance matrix |
+
|
+ Calculating |
+
+ M>
+ A = [3 1; 1 3] |
+
+ P>
+ A = np.array([[3, 1], [1, 3]]) |
+
+ R>
+ A = matrix(c(3,1,1,3), ncol=2) |
+
+ J>
+ A=[3 1; 1 3] |
+
+ Calculating |
+
|
+ Generating
+ a Gaussian dataset: |
+
+ %
+ requires statistics toolbox package |
+
+ P>
+ mean = np.array([0,0]) |
+
+ #
+ requires the ‘mass’ package |
+
+ #
+ requires the Distributions package from
+ https://github.com/JuliaStats/Distributions.jl |
+
+ Generating
+ a Gaussian dataset: |
+
+
\n", + "I would be happy to hear your comments and suggestions. \n", + "Please feel free to drop me a note via\n", + "[twitter](https://twitter.com/rasbt), [email](mailto:bluewoodtree@gmail.com), or [google+](https://plus.google.com/+SebastianRaschka).\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "# Parallel processing via the `multiprocessing` module" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "CPUs with multiple cores have become the standard in the recent development of modern computer architectures and we can not only find them in supercomputer facilities but also in our desktop machines at home, and our laptops; even Apple's iPhone 5S got a 1.3 Ghz Dual-core processor in 2013.\n", + "\n", + "However, the default Python interpreter was designed with simplicity in mind and has a thread-safe mechanism, the so-called \"GIL\" (Global Interpreter Lock). In order to prevent conflicts between threads, it executes only one statement at a time (so-called serial processing, or single-threading).\n", + "\n", + "In this introduction to Python's `multiprocessing` module, we will see how we can spawn multiple subprocesses to avoid some of the GIL's disadvantages." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Sections" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "- [An introduction to parallel programming using Python's `multiprocessing` module](#An-introduction-to-parallel-programming-using-Python's-`multiprocessing`-module)\n", + " - [Multi-Threading vs. Multi-Processing](#Multi-Threading-vs.-Multi-Processing)\n", + "- [Introduction to the `multiprocessing` module](#Introduction-to-the-multiprocessing-module)\n", + " - [The `Process` class](#The-Process-class)\n", + " - [How to retrieve results in a particular order](#How-to-retrieve-results-in-a-particular-order)\n", + " - [The `Pool` class](#The-Pool-class)\n", + "- [Kernel density estimation as benchmarking function](#Kernel-density-estimation-as-benchmarking-function)\n", + " - [The Parzen-window method in a nutshell](#The-Parzen-window-method-in-a-nutshell)\n", + " - [Sample data and `timeit` benchmarks](#Sample-data-and-timeit-benchmarks)\n", + " - [Benchmarking functions](#Benchmarking-functions)\n", + " - [Preparing the plotting of the results](#Preparing-the-plotting-of-the-results)\n", + "- [Results](#Results)\n", + "- [Conclusion](#Conclusion)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### Multi-Threading vs. Multi-Processing" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Depending on the application, two common approaches in parallel programming are either to run code via threads or multiple processes, respectively. If we submit \"jobs\" to different threads, those jobs can be pictured as \"sub-tasks\" of a single process and those threads will usually have access to the same memory areas (i.e., shared memory). This approach can easily lead to conflicts in case of improper synchronization, for example, if processes are writing to the same memory location at the same time. \n", + "\n", + "A safer approach (although it comes with an additional overhead due to the communication overhead between separate processes) is to submit multiple processes to completely separate memory locations (i.e., distributed memory): Every process will run completely independent from each other.\n", + "\n", + "Here, we will take a look at Python's [`multiprocessing`](https://docs.python.org/dev/library/multiprocessing.html) module and how we can use it to submit multiple processes that can run independently from each other in order to make best use of our CPU cores." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "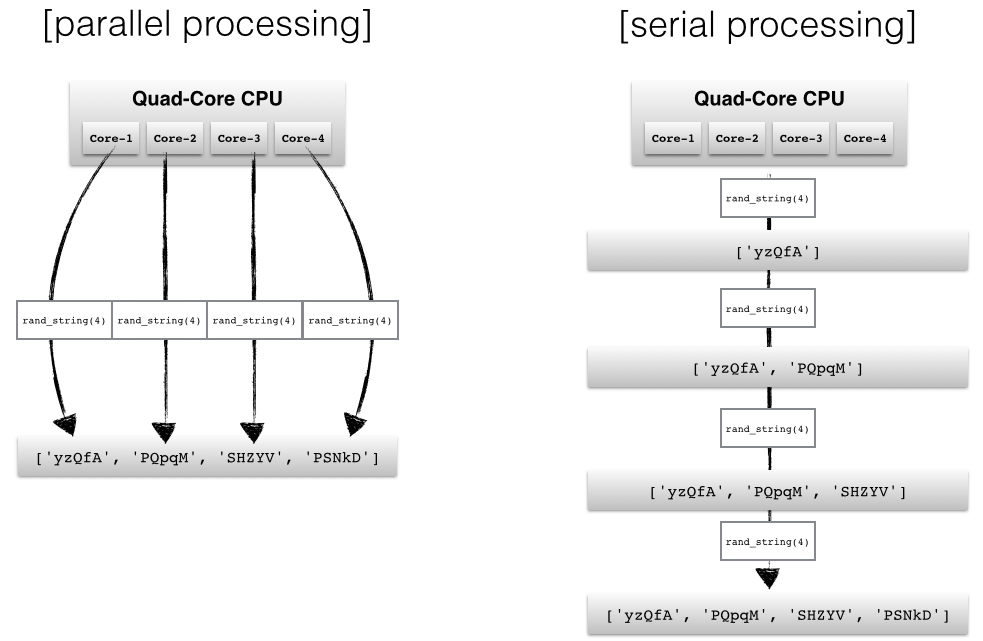" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "# Introduction to the `multiprocessing` module" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#Sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "The [multiprocessing](https://docs.python.org/dev/library/multiprocessing.html) module in Python's Standard Library has a lot of powerful features. If you want to read about all the nitty-gritty tips, tricks, and details, I would recommend to use the [official documentation](https://docs.python.org/dev/library/multiprocessing.html) as an entry point. \n", + "\n", + "In the following sections, I want to provide a brief overview of different approaches to show how the `multiprocessing` module can be used for parallel programming." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### The `Process` class" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#Sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "The most basic approach is probably to use the `Process` class from the `multiprocessing` module. \n", + "Here, we will use a simple queue function to generate four random strings in parallel." + ] + }, + { + "cell_type": "code", + "execution_count": 2, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "['BJWNs', 'GOK0H', '7CTRJ', 'THDF3']\n" + ] + } + ], + "source": [ + "import multiprocessing as mp\n", + "import random\n", + "import string\n", + "\n", + "random.seed(123)\n", + "\n", + "# Define an output queue\n", + "output = mp.Queue()\n", + "\n", + "# define a example function\n", + "def rand_string(length, output):\n", + " \"\"\" Generates a random string of numbers, lower- and uppercase chars. \"\"\"\n", + " rand_str = ''.join(random.choice(\n", + " string.ascii_lowercase \n", + " + string.ascii_uppercase \n", + " + string.digits)\n", + " for i in range(length))\n", + " output.put(rand_str)\n", + "\n", + "# Setup a list of processes that we want to run\n", + "processes = [mp.Process(target=rand_string, args=(5, output)) for x in range(4)]\n", + "\n", + "# Run processes\n", + "for p in processes:\n", + " p.start()\n", + "\n", + "# Exit the completed processes\n", + "for p in processes:\n", + " p.join()\n", + "\n", + "# Get process results from the output queue\n", + "results = [output.get() for p in processes]\n", + "\n", + "print(results)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### How to retrieve results in a particular order " + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#Sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "The order of the obtained results does not necessarily have to match the order of the processes (in the `processes` list). Since we eventually use the `.get()` method to retrieve the results from the `Queue` sequentially, the order in which the processes finished determines the order of our results. \n", + "E.g., if the second process has finished just before the first process, the order of the strings in the `results` list could have also been\n", + "`['PQpqM', 'yzQfA', 'SHZYV', 'PSNkD']` instead of `['yzQfA', 'PQpqM', 'SHZYV', 'PSNkD']`\n", + "\n", + "If our application required us to retrieve results in a particular order, one possibility would be to refer to the processes' `._identity` attribute. In this case, we could also simply use the values from our `range` object as position argument. The modified code would be:" + ] + }, + { + "cell_type": "code", + "execution_count": 3, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "[(0, 'h5hoV'), (1, 'fvdmN'), (2, 'rxGX4'), (3, '8hDJj')]\n" + ] + } + ], + "source": [ + "# Define an output queue\n", + "output = mp.Queue()\n", + "\n", + "# define a example function\n", + "def rand_string(length, pos, output):\n", + " \"\"\" Generates a random string of numbers, lower- and uppercase chars. \"\"\"\n", + " rand_str = ''.join(random.choice(\n", + " string.ascii_lowercase \n", + " + string.ascii_uppercase \n", + " + string.digits)\n", + " for i in range(length))\n", + " output.put((pos, rand_str))\n", + "\n", + "# Setup a list of processes that we want to run\n", + "processes = [mp.Process(target=rand_string, args=(5, x, output)) for x in range(4)]\n", + "\n", + "# Run processes\n", + "for p in processes:\n", + " p.start()\n", + "\n", + "# Exit the completed processes\n", + "for p in processes:\n", + " p.join()\n", + "\n", + "# Get process results from the output queue\n", + "results = [output.get() for p in processes]\n", + "\n", + "print(results)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "And the retrieved results would be tuples, for example, `[(0, 'KAQo6'), (1, '5lUya'), (2, 'nj6Q0'), (3, 'QQvLr')]` \n", + "or `[(1, '5lUya'), (3, 'QQvLr'), (0, 'KAQo6'), (2, 'nj6Q0')]`\n", + "\n", + "To make sure that we retrieved the results in order, we could simply sort the results and optionally get rid of the position argument:" + ] + }, + { + "cell_type": "code", + "execution_count": 4, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "['h5hoV', 'fvdmN', 'rxGX4', '8hDJj']\n" + ] + } + ], + "source": [ + "results.sort()\n", + "results = [r[1] for r in results]\n", + "print(results)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "**A simpler way to maintain an ordered list of results is to use the `Pool.apply` and `Pool.map` functions which we will discuss in the next section.**" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### The `Pool` class" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#Sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Another and more convenient approach for simple parallel processing tasks is provided by the `Pool` class. \n", + "\n", + "There are four methods that are particularly interesting:\n", + "\n", + " - Pool.apply\n", + " \n", + " - Pool.map\n", + " \n", + " - Pool.apply_async\n", + " \n", + " - Pool.map_async\n", + " \n", + "The `Pool.apply` and `Pool.map` methods are basically equivalents to Python's in-built [`apply`](https://docs.python.org/2/library/functions.html#apply) and [`map`](https://docs.python.org/2/library/functions.html#map) functions." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Before we come to the `async` variants of the `Pool` methods, let us take a look at a simple example using `Pool.apply` and `Pool.map`. Here, we will set the number of processes to 4, which means that the `Pool` class will only allow 4 processes running at the same time." + ] + }, + { + "cell_type": "code", + "execution_count": 5, + "metadata": {}, + "outputs": [], + "source": [ + "def cube(x):\n", + " return x**3" + ] + }, + { + "cell_type": "code", + "execution_count": 6, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "[1, 8, 27, 64, 125, 216]\n" + ] + } + ], + "source": [ + "pool = mp.Pool(processes=4)\n", + "results = [pool.apply(cube, args=(x,)) for x in range(1,7)]\n", + "print(results)" + ] + }, + { + "cell_type": "code", + "execution_count": 7, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "[1, 8, 27, 64, 125, 216]\n" + ] + } + ], + "source": [ + "pool = mp.Pool(processes=4)\n", + "results = pool.map(cube, range(1,7))\n", + "print(results)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "The `Pool.map` and `Pool.apply` will lock the main program until all processes are finished, which is quite useful if we want to obtain results in a particular order for certain applications. \n", + "In contrast, the `async` variants will submit all processes at once and retrieve the results as soon as they are finished. \n", + "One more difference is that we need to use the `get` method after the `apply_async()` call in order to obtain the `return` values of the finished processes." + ] + }, + { + "cell_type": "code", + "execution_count": 8, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "[1, 8, 27, 64, 125, 216]\n" + ] + } + ], + "source": [ + "pool = mp.Pool(processes=4)\n", + "results = [pool.apply_async(cube, args=(x,)) for x in range(1,7)]\n", + "output = [p.get() for p in results]\n", + "print(output)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "# Kernel density estimation as benchmarking function" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#Sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "In the following approach, I want to do a simple comparison of a serial vs. multiprocessing approach where I will use a slightly more complex function than the `cube` example, which he have been using above. \n", + "\n", + "Here, I define a function for performing a Kernel density estimation for probability density functions using the Parzen-window technique. \n", + "I don't want to go into much detail about the theory of this technique, since we are mostly interested to see how `multiprocessing` can be used for performance improvements, but you are welcome to read my more detailed article about the [Parzen-window method here](http://sebastianraschka.com/Articles/2014_parzen_density_est.html). " + ] + }, + { + "cell_type": "code", + "execution_count": 9, + "metadata": {}, + "outputs": [], + "source": [ + "import numpy as np\n", + "\n", + "def parzen_estimation(x_samples, point_x, h):\n", + " \"\"\"\n", + " Implementation of a hypercube kernel for Parzen-window estimation.\n", + "\n", + " Keyword arguments:\n", + " x_sample:training sample, 'd x 1'-dimensional numpy array\n", + " x: point x for density estimation, 'd x 1'-dimensional numpy array\n", + " h: window width\n", + " \n", + " Returns the predicted pdf as float.\n", + "\n", + " \"\"\"\n", + " k_n = 0\n", + " for row in x_samples:\n", + " x_i = (point_x - row[:,np.newaxis]) / (h)\n", + " for row in x_i:\n", + " if np.abs(row) > (1/2):\n", + " break\n", + " else: # \"completion-else\"*\n", + " k_n += 1\n", + " return (k_n / len(x_samples)) / (h**point_x.shape[1])" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "**A quick note about the \"completion else**\n", + "\n", + "Sometimes I receive comments about whether I used this for-else combination intentionally or if it happened by mistake. That is a legitimate question, since this \"completion-else\" is rarely used (that's what I call it, I am not aware if there is an \"official\" name for this, if so, please let me know). \n", + "I have a more detailed explanation [here](http://sebastianraschka.com/Articles/2014_deep_python.html#else_clauses) in one of my blog-posts, but in a nutshell: In contrast to a conditional else (in combination with if-statements), the \"completion else\" is only executed if the preceding code block (here the `for`-loop) has finished.\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### The Parzen-window method in a nutshell" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#Sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "So what this function does in a nutshell: It counts points in a defined region (the so-called window), and divides the number of those points inside by the number of total points to estimate the probability of a single point being in a certain region.\n", + "\n", + "Below is a simple example where our window is represented by a hypercube centered at the origin, and we want to get an estimate of the probability for a point being in the center of the plot based on the hypercube." + ] + }, + { + "cell_type": "code", + "execution_count": 10, + "metadata": {}, + "outputs": [], + "source": [ + "%matplotlib inline" + ] + }, + { + "cell_type": "code", + "execution_count": 11, + "metadata": {}, + "outputs": [ + { + "data": { + "image/png": "iVBORw0KGgoAAAANSUhEUgAAAZQAAAGUCAYAAAASxdSgAAAABHNCSVQICAgIfAhkiAAAAAlwSFlz\nAAALEgAACxIB0t1+/AAAIABJREFUeJzsnXeUVOX9/9/3Tp/ZQpG6LCxNQUXEgCJIEVgWUIkRCwqK\nDdFEjTExNiwYG/zUX+QY2/fYYn5BiFEhAZYOFiKgIUG/iqgUkaZI2d2pt/3+WJ/rnbv3ztyZuW1m\nn9c5niO7s3eeO+V5P5/OSJIkgUKhUCiUAmGdXgCFQqFQSgMqKBQKhUIxBSooFAqFQjEFKigUCoVC\nMQUqKBQKhUIxBSooFAqFQjEFKigUCoVCMQUqKBQKhUIxBSooFAqFQjEFKigUCoVCMQUqKBQKhUIx\nBSooFAqFQjEFKigUCoVCMQUqKBQKhUIxBSooFAqFQjEFKigUCoVCMQUqKBQKhUIxBSooFAqFQjEF\nKigUCoVCMQUqKBQKhUIxBSooFAqFQjEFKigUCoVCMQUqKBQKhUIxBSooFAqFQjEFKigUCoVCMQUq\nKBQKhUIxBSooFAqFQjEFKigUCoVCMQUqKBQKhUIxBSooFAqFQjEFr9MLoFD0kCQJqVQKPM/D7/fD\n4/GAYRgwDOP00igUigaMJEmS04ugUNSIogiO4+T/iIiIoohAICALDMtSI5tCcQvUQqG4CkmSIAgC\nGhoa4Pf7wbKs/J8kSYjH42AYBhzHAQBYloXX64XP56MCQ6E4DBUUimuQJAkcx0EQBNnNpYRYKcT1\nRYzrVCqFVCoFgAoMheIkVFAorkAURaRSKUiSJMdJRFFEPB6HIAjwer3weDxpf6MUGKBZkEjchQoM\nhWI/NIZCcRTi4iJxEiISx44dgyRJspVCHicIAliWhcfjkf/TC9ITgVF+xFmWhc/nkwWKCgyFYh5U\nUCiOQawJURRlMSFxkkQigVAoBL/fD47j5I2/qakJgUCACgyF4kKoy4viCFouLkEQ0NTUlCYQahiG\nSdv4lcJCxElPYNQpx0RckskkkskkACowFEohUEGh2IokSeB5HjzPg2EYecNOpVKIRqMIBoMIBoNo\namoydD2GYeD1euH1euXrmykwHo9Hjr94vV5aA0OhZIAKCsU2SG2J2sUVi8XAcRzKy8tlYSDkuoEb\nFRilBZJJYERRRGNjIwDIwqK0YKjAUCg/QQWFYjnKwDvw08bN8zyi0Sg8Hg8qKyst2ZwzCUwymYQo\nimnWi5bAkP9YloUoikgkEvL1qcBQKD9BBYViKWoXF7FKEokE4vE4wuEw/H6/bRtxPgKj/FstC4YK\nDIXSDBUUimXo1ZZEo1GIooiKigrNwDvBjgREIwJDrBOSUZbNRUYFhtJaoYJCMR11bQk55XMch2g0\nCp/Ph7Kysoyba6bfWSk0WgKTSCTkYL0RF5lynVRgKK0JKigUU9GrLUkkEkgkEohEIi1aquSC3Rsw\nEUSGYeD3+yGKIkRRBM/z4DgOkiTlLDCkHxlABYZSWlBBoZgGyYiSJAnBYFB2cZEU4MrKyqKv6yCN\nKokFI4qi7CIzKjDKGhoqMJRSggoKpWCUgXdRFAE0b57q2pJS3ByJwPh8PgAtBYY8RikQRgVGEAT4\nfD74/X4qMJSigAoKpSC0XFyiKCIWiyGVSqGsrEzebPO5drGRSWBIw0p1kaWewKRSKVlYyGNIfIcK\nDMWNUEGh5A2JIwBIE5NUKgWv14uKioq8XVylslEWIjDAT0kCQMtkBwBpnZSpwFCchgoKJWe0aksA\nyC1LvF5v1iyu1opSYEirFz2BUVtoWi4yMjuGQATG6/WmxW8oFDuggkLJCb32KdFoFDzPIxgMynUn\nZlGqm6JSILQERhRFJJNJ8DxvuNCSCIwkSXIPMyowFLuggkIxhLp9CtnYeJ5HU1MTvF4vKisrkUwm\nIQiCk0stWtQCE4vF5IaUPM8jmUzK3ZZzFRjye2UMhgoMxWyooFCyohzNq7RKksmk3D4lEAjYup7W\nABEX4sYiWWBEJPIRGHXciwoMxUyooFAykmv7FCI2haJ3nda84SnFA0BGgdHKAqMCQ7EaKigUTfTm\nlpD2KX6/nwbeLSabMGcSGI7jkEgkMk6z1BOYeDwOAHLshQoMxShUUCgtyDSaN5lMFtw+JZ/1tNaN\nLJf7NktgyPvNMAw4jmthwZA0ZSowFDVUUChpaNWWCIKAaDQKIHv7FLNcXsrr0U0rP8wQGKU7k8TS\nqMBQ9KCCQgGgX1vSGtqnGKEUEgGMCgx5rNoyNCIwyj5kVGBaH1RQKHIDR5JRRDaTTKN5rcZsS4fS\nEj2BIW36o9FozhZMKpWSkwOUAkM+V1RgShsqKK0YZW1JKpWSJycKgoCmpiZ4PJ6C2qdQigulwBAx\nUFbxi6KYt8AAkOtrqMCULlRQWimZaktisRhCoRACgUDOX/hStCxK8Z6MoEwjBtKnWRYqMEoLhsRg\nqMAUP1RQWiFatSUA5JOkEy6uTLTmLC83YURg9GbBkL9XuteA5hgd6WGmbhVDBab4cM+uQbEcvdoS\nnuflDsHl5eWu+xKrN6XWaC24kUwCY2RcMoAWAkPa+JDHq7PIKO6GCkorIdtoXq/XK8dQCsHMSnky\nrItSHJghMADSxENtwVCBcTdUUFoByvG06vYpkiShoqICyWTSlSd/0t2YbiD2QToVF0o+AqP8W6Cl\nBUMFxt1QQSlhMrVPaWpqQiAQQCgUcp2LiyAIAhoaGsCyrFzrQKBB3OIjk8AkEglZNEh35GwuMmJ1\nKwVGXQdDsRcqKCWK3twS0j6lkNG8VkM2Cp7nUV5eLt+DKIpIJBLy3HUAuidcivtRCkwgEJBHRxNX\nrCRJGV1k6j5kJEtRK02Zfj7sgQpKiaGeW6Jsn0KKF7Xap1jdJdgopM2LKIrw+/3wer1pJ1CSCeT1\netMsMJKKSjYPOg63+CCC4ff7wbJs2rhk4rKlAuNuqKCUEFq1JUDxtE9RrpNhmLTRtmrI/ZEmlbn2\nqSoGWnu6tHJcMoAWAgOgxfubi8CQFGVq4ZoHFZQSQau2JNf2KU4F5bVcceRLbxS9NiIkJZoU4dE+\nU9lxMjkjk4hmEhhixeYiMMSFSlAWWpI6GEpuUEEpcjLVlkSjUXg8HlRWVmb9cjj15SF9xIDsnYxz\nQUtgjKawUopjkJlSYIgFUojAJBIJcBwnH7yURZbFaOE6ARWUIkavtkQ5mteM2pJcyCWGko8rLt8Y\njZEMI7Lx0Crt4oO8X7kIjF6assfj0bRgqMBkhwpKkSIIAmKxmOwmUtaWaI3mzYadFehuyDbTyjAi\nm486g8yN9TmliBVzdHIRGK2/V66NCkx2qKAUGUoXF/k3wzDyaF6fz+fq0by5uLjs3MgzuU+Ur7XS\ngqFYgxWvrZbAkEMEyRIkkPdZbcFQgckOFZQiIlNtSSKRsH00rxaZLB1SUGnExeXkl1G9+ZAuAizL\nyptPsWeQtXa0YmyxWCztPVY+hgqMMaigFAF6tSXKnxca0LbS5eUGF1chkI2glFOUlbRGFx/5TpHN\nX8uCydaqX0tg4vF4Wnym1AWGCorLyTaal2EYV3YIJliVxeUkmTLIjLRxLwacWK+b6m700tCNHiKU\nVq7y70tdYKiguBi92pJ4PI5UKoVwOJz2AXUTRAiLoWdYoWTKIKMpysVDJkGzWmDIcweDwaIWGCoo\nLkTpylJ+CJXtUyoqKkzPijGr9QoAU1xcxep6UQtMtgrvUrDaWhtmC4yyjxl53Pz583H//ffbf3MF\nQD/JLoPUlhAxIR+uZDKJhoYGBAIBlJWVyadct226ZIYJz/OorKzMW0wync7cds/ZIMH9YDCISCSC\nUCgEj8cjpyhHo1G5qI7OgHGOQlxuRGD8fj9CoRAikYg8QptkYJI0f57nNT/DyhgOy7JYuXJlobdk\nO9RCcRF6Lq5oNCp33rVyNG+hGzXJ4gIgi54ZqE92xSYoatQpynrBX5K+3FpwUwylUIzE2ZQWjPq+\nBUEoSsuVCooLyNQ+hYxE1WqfYubmWsgXWTn5MRKJIBqNmrKm1kAm1wnQ7Dq0O4OslDZ2o1gt3Jni\nbERggGZPxM6dO9G+fXuEQiFL12QFxSeBJYZy9ofSKkkkEmhsbEQoFCqKQkWSuux0HUyxo3SdAJDb\n5wA/ZfaRpAxBEFqVBWMHdn3PiMAEAgGEw+G0uqyHHnoIp512GrZv346HHnoI7777bsZmqddeey06\ndeqEAQMGaP5+/fr1qKysxKBBgzBo0CA8/PDDltwTQAXFMYhVQgZGETEhG3QymURFRQUCgYChaxVK\nPtYOx3FoaGiAx+NBeXm56SY63SxbbjyRSER2lSWTSSowJQTLsggEAliwYAFWrVqFvn37oqmpCb/9\n7W/RvXt32WpVc80116C+vj7jtUeNGoWtW7di69atmD17thXLB0BdXo6gV1tCgnd+v9+QVeKU1aJ2\ncVlhlbjVInMaPdcJz/PyECr1kLFieS1bo6tND5Zl0bNnT8ybNw9As+tTrzffiBEjsHv37ozXs+ug\nQS0UmyG1CWoXVywWQ1NTE8LhMMLhsGu/WEoXV0VFhaaYmB04p6dufYjAkAyycDiclkEWi8VoBpkB\nnBYz9fPHYjGEw2H534XEUxiGwcaNGzFw4EBMmjQJn332WUFrzQS1UGyCnCRJxlZZWRmAn0beAvlV\nkiuLogrBiAgoLahSLlQsZvSaXKr7UxVz8VxrIB6PIxKJmHKtM844A3v37kU4HMby5ctx4YUXYseO\nHaZcWw21UGxAXVtCSKVSaGhogM/nsyQGYRakOr8YLCjKTygbXJLaiGAwCJZlM9ZGOH1adwKn71n9\n/NFoNM1CKYTy8nL5WhMnTgTHcThy5Igp11ZDLRSL0aotIXNLOI4ruFmi1XUZhcxYobiLTCnKytoI\noNlyttuCcXpTdxNkQJ4ZHDp0CB07dgTDMNi8eTMkSUK7du1MubYaKigWoVdbQuack/YpbrFKtISJ\n1MEYTRLIdK1816OMMykFmVI4SoHx+/2yeyyRSJRMk8tiQf19ycXldfnll2PDhg04fPgwqqurMWfO\nHLnFz6xZs/Dmm2/iueeeg9frRTgcxhtvvGH6+glUUCxAbzRvKpWSC9UikYgpX04rLBQ7srgo7oME\n+IGfgsCtZUyyG6wjtcurU6dOhv5uwYIFGX//q1/9Cr/61a8KWptRqKCYDEnfBJB2oo7FYhAEAeFw\nGIlEwvEPrx5ktDB1cVGUKcrZxiSb0eTSDZu6W4jH40VZKU8FxST0akuI28jn86GiokK3OKnQ5y4U\nst7GxsaCxwiXQr+t1kqm9y3TmGT1jHY6Jjk3yERQQiwWkzNBiwkqKCagN5pXy21k9mZrxpeWVF0D\nza4OI9X5lNLGSFGtMoss1wmHbsNt1pGZQXk7oYJSAOrRvMrAezQahSRJtriNChEoZRYXANeM5yXC\nS2JPSv9+a4C8p27a5DJR6HyQ1k62wsZiofV8Q01GkiRwHAdBEFq0T8k0pdBNForSHVdWVoZjx46Z\nti4zkCQJDQ0N8v+TwjyGYeT23nRTcidG2rer4y9OWgluc9FSC6UVkWk0b6FTCu2AbM4kNdHsLC4z\nRJNYfWRIEYF0ZlaP1i21rKNSI1P7dvJeEheaU4cFNxU20hhKK0CvtkQ5mjdb+xQrLJRcrqfMOFO7\n49wQTFcKMwAEg8E08SZuk2AwaHnWEcU61AIjiiKSyaQcewTQwj3Wmg4L1OVV4mjVlgDNA3FisRiC\nwWDaTAMj17P7C6LOOHPbF5Q0ngSa20U0Nja2eIxS9LSyjojgF1tQuLVD3kuGYeD3+9MOC+oMMisO\nC04H5dXPb2YvLzuhgmIAskmpXVyxWAwcx+U0mtfsD60Rq0Lp4gqHw5ZnceVj6Sir8kn+fS7XIO8L\ncd9ptRWhVd/ZcTqOQYSitR8WOI4ryoJiKigZIB/iY8eOpc1I53ke0WgUHo9HczSv0Wvb8SUgM+m1\nXFxqnHB56cVzzGjdotVWROmzV3fdLbVNqVTIdFgolQwyrf2g2O4BoIKii7K2hBQjqk/6fr8/rzfd\nzA9KJhFwu4srF7ErFC2fPS3KK070UpR5nk9rckkOC0asUXVhodM4HcvMFyooKjLVlsTjcVNaklht\nCRTi4rLrg0wSGTweT1axs+L1ysWlUqxf7mIjX6s9U4qyOhvQje5O9eermD9vVFAU6NWWAM0tSXLt\numsH6s22kFO/WfeVTQBSqRSi0ahcle/066nnUlGeeMlj3LghUdLRS1FWjkl245Ax5TqK1QVLBeVH\n9GpLSJfVUChkWrM2qyyUYnBxxeNxpFIpw4kMTpzW1CdecspVfh6Uc9vd5CoplGI+HethxN1JHqec\nCWMXWpZZsb4PrV5QMs0tISmsxDXiVkgOP8ldzzeLy0pXHHk9GYYxNAfGTWJIPheBQKBF111l/MVt\nJ958Kfb1Z0PL3RmPx2VXNzlQOPV+8jxftF2+W7Wg6NWWEJcMqS1pbGx0tBgxGzzP2xLYzhfSjibX\nWh23ot6QSi3jyCmcSFkm33uGYRAIBMCyrO3vp3oviMViRVmDArRiQSEfGL3aEmX7FDdUkGvB87w8\nW8VNLi7yelkxqMtt74WRjKN8AsJOF9q1VvTeT/WYZGLBmBVPU7ddKcYqeaAVCoqR9ilWj+Y1Y1Mk\nLi6l6LlhXQRJktDU1NTqBnVlyjjSailSSvGXUsRoPZOZGWTFOlwLaGWCojW3BPhpc9bLOnLTqZhk\ncfE8j/Ly8rQUZ7dAXmc3ZsXZjTIg3BriL8WKUYtQL4NMS2CMNizVagxJXV4uRl1bonRxKTdnvawj\nswUl3+spazdIhT7JPnILyWQSqVQKPp+v4C+Fm4TcLIzGX0g7d7tx8vUuRjdfpgyyfBuWUpeXi9Gr\nLSEptl6vN+/2KfmSz0apzOLKt0LfinURlPEnN9SWFAOZ3GPk8BOPx20vyKPvXf4oDwxA5o4MenNg\nqKC4FL3aklyryJ08KWdrQumGU7w6/kTakFNyQ3naJWMGfD6fXJAH0PiLVVhlHelZpMqODMokFoZh\nilpQSvITSQLvZKYGedNILUQymURFRYVjLUmMioAgCGhoaIAkSaisrLRlBG6u98lxHBoaGuR4SWvb\n5FauXIna2lrU1NTgzDPPxOuvvy6PUy4UIjDBYBCRSAShUAgejweCICAWiyEajSKZTMqdsCnuhlik\npKN2JBKR9yBJkvD9999j3LhxWLp0KQ4cOCC7zPS49tpr0alTJwwYMED3Mbfeeiv69u2LgQMHYuvW\nrabejxYl9+0XRRGJREIuUCKnDrLxkd5RTrQkyYVkMomGhgYEAgFEIhHdNZhpoeRyn6QYrKmpCWVl\nZWnjjq2IOTmN1v3Mnz8fV111FTZt2oQjR45g+/btuOOOO3DttddassGTky4RGFLTw3EcotEoYrEY\nkskkBEEoSoFxKobi1GuldHl6vV60adMG9957LwRBwLJly9ChQweMHTsWr7zyiubfX3PNNaivr9e9\n/rJly/DVV1/hyy+/xIsvvoibbrrJqluRKSmXF6kYP3bsGNq2bStvbGQCYL61EMS6MYtMG26+c1bs\nRBRFRKNR2XKyyipxgzuPrEPNDz/8gIcfflhOBSbEYjHU19fjww8/xNlnn23pmoq5IaIbcer1IULq\n9/tx7rnnYseOHZg0aRIuuOACvPvuu3K3czUjRozA7t27da+7ZMkSzJgxAwBw1lln4dixYzh06BA6\ndepkxW0AKCFBUfflIT+LRqMAUPDGZ8fGppXFZSdGNnD1ICw3rtEOli1bpvt5isfjWLBggaWCoiZb\nQ0TAWPylGDOtSg0S362oqMD555+f93X27duH6upq+d/dunXDt99+SwXFCOQEpmyfks9oXi3M/oJp\nWTzZamEyXcuODVZvEJaTa3ISMspAC5KO7iR66axunXjYmtOV1bNYzKxDUb+uVt9nyQgKkL6RxePx\ntPYpZl3XbIrBxWXnIKxiYcSIEbpfzrKyMowfP97mFWXGSP2L1+uVs42cwmlhcwNmZXlVVVVh7969\n8r+//fZbVFVVFXzdTJRUUJ5kRQEwTUysgAgUWS9pT5KPmJgdlFdfS/maOikmbrN4+vfvj5EjRyIY\nDKb93OfzoX379rjwwgsdWll2tLKNSFsR0miUjBlwW+FsKaK2kIjLq1AmT56MP//5zwCADz/8EG3a\ntLHU3QWUkIUiSRIaGhoQCoUQj8dNPelYYaGQjdotQ6a0cHoQlttdZ6+//jp+//vf44033oDX6wXH\ncRg5ciSee+65vEcIEOx0wyjdYyzLgud5eL3egqq9iwmnXV5qiFs5G5dffjk2bNiAw4cPo7q6GnPm\nzJHjZbNmzcKkSZOwbNky9OnTB5FIRDdbzExKRlBYlkWbNm3AMIzphXVmN01MpVKy+6hQF5cVm65y\nEFY+lp7bhcAsgsEg5s+fj0cffRT79u3DCSecgPbt2zu9rIJhGAY+ny9tXojV8Re3bep2ov6uRKNR\nQ4KyYMGCrI955pln8l5XPpSMoABIa2Pgxg2NZHERl4Pb4iUkWaCxsREALO+6XCqUlZXhpJNOcnoZ\npqDe2Emii1b8RdnOnTS3pOnJ+WGFy8sJ3LWjmYRbmjkqUbqPWJZtUb9QKGac8EiQNhgMOpISrIfT\ngWLKTxht567snuyWz5EebrOOjLq83EhJCorZmNU0kWRxmdkqwyx3QzKZRDKZhMfjMeV0ZNb9ET++\nJEmOduGlaJOp265eM0RKOmpB4zjOtQlF2SgpQSEbv1tcXnYN7VLed64oU4JDoZAps1XMOu0RMSYB\nbrJZAc2plWZPzaMUjta8djLQLlP8xUkrwW0WClC86dMlJSgEN7i81HPp1X5ptwheY2MjvF4vKioq\nXDOoS5KaRweLoohwOAyfzweO4+Q6iWg0Cr/fr+tmoadgd0DcXaQIViv+ohQWN3wnnMCNgpYvVFBy\nwMgbr8yQsrNQsZD5KsQCcMOXWtknTNmrSk02NwsRFzdUgVOayRR/4ThO/u60poOB+vumHLVRjJSk\noJiN0TfXqIvLyW68dlTm53t/xGLy+XwIh8Nytlk2jFaB0yyk7DhV/+LxeJBMJuX6F3X8hYzTtQI3\nWAjK53f6UFcIJSUoVrVPV15T74OXycXlFsg8GIZhXJcSTF4/o0PP9DDShVdpvbjpNWjtkENBpmFU\nbuo/RmlJSQkKwex28wQtkcqnCNCJGA/HcWhqasooeE64vLK5CAtdT6YspGQyKf+eblLuQutgIIoi\neJ5vEX8pZsuzlDK8gBIWFCssFDWkPX6+J347TG0S4E4kEq7rb6aMl2i9fla8NnrusVLapEqRTJZn\nIpGAJEl5W55ucHkRiJVerJSUoFj5oVCLVCEuLiva4WsJKNmwSfNJuxo7FsNcFSBzkJgUnio3MIp1\n5LqpKy3PQCCgW/+iLLB0I1qNIUOhkIMrKoySEhSC1e3mC+lzRSikdsQIZMP2+XwoKysz9Dx2uby0\nMszcgN4mxXEcRFGUa1/scI9RAcsNI4kZxRB/KeYqeYAKSk7XFAQBsVjMdUFt9f26dcM2I6XazkI4\nskkBzQJNal/sco85semphz0VI7nGX0iihhOoP8dmzUJxCiooBiHptmZlcVm5xnxTgs1ak9Z18s0w\nc8tJXc89pjVi18oUV0ruGIm/kM+Z05l/NIbiIqz4EpNTNWlN4kb/JrGe4vE4WJZ1ZB59JvKNl7jp\nHtQo3WN2tXgvZZyqfwkEAojFYmBZ1pHCWKuGazlFSQkKwayTNjlVA82T+Mw8uZhpoYiiKHcJdlsN\nDHG/ZZtDX8yQyma1D1/LxUKsFze9R60d9eFAL/5iR984M+fJOwEVFB3UdRuxWMw17hcCSQnmeR6B\nQKBg68lMkSM9tziOKzjDrNg2Xy0Xi3K0LlAcGUitESOFsWbGzmgMpQgoZGMkLq5kMmlp3Uahm7ey\nhsPv9zs2610Lcl8kXbnYg7yFwjAtJyCS2IvdJ+BsuO3QZAeZ3G3qwthMsTMz4i9UUFyE+kORq19W\n6eKqrKxM+3BYFUTPB3VMIhaLmb6ufDc1YtkBMJyunG0tdvDRgY8Q42IY2X2kpc+j1YFX6wTsZKNO\np7LLisFSs7rzQjweL+ox0iUlKIR8PphkIySuI/U1nGzoSCCDsEiuOtmUzMzOyhfl2sLhsNxBwKn1\n5MLOYzvx87//HDEuhtM6noaxNWMxrsc4DOkyBD6PtZ0F9DYoUvsSjUape8zFGK1/0bM+aWFjkWC0\ncNCp1iS5ioByEJadVe9GIOnKPM/LLi7ijnP7Brjhmw24btl1GNV9FKrLq3F+n/OxZvca3LX+Luw6\nvgvndDsHo6tHY1TVKPQL9rN8PWSDYhgGqVQKgUCAdk62GLM+p7nEX/RSy2lQ3kVoqX8mMrm4tK5t\nRcNJI5C2+B6PBxUVFZZbT/msjbTst2ODM6PLgCRJ+J///g/mfTgPL096Gf/57j84GD2IEdUjMKJ6\nBB4c8SC+j32PtXvWYtXOVZi3aR7KA+UY22MsxtWMw4jqESjzl5l4Vy0xskGVSufkUiioVKMVf1En\nZwCQiytZli36GEppvYMKsm02HMfh+PHj8Hq9KC8vN/RhtrtDMNDcM6yhoQGBQACRSMTyDTsXceI4\nDg0NDfD7/abES+wiJaRw2+rb8PJ/X8aqqaswsvtIMGh53x3CHXBZ/8vw3Pjn8MnVn+DP5/8Z1RXV\nePbfz+LEF07EpEWT8OTmJ/GfQ/+BKFl/2CAbVCAQQDgcRjgchsfjkTs4RKNRJJNJ8DzfKoPr+WKX\nJU2SM4LBIMLhsOxaF0URy5cvx+DBg7Fjxw58/PHHhmYB1dfXo1+/fujbty/mzp3b4vfr169HZWUl\nBg0ahEGDBuHhhx+24rbSKCkLBcg+V17p4sqlNsLuzTKXNiV2W0/KeInbOhhn43DsMKb/YzraBNtg\n1eWrUO4vB/Dj5wb6mzDLsDit42k4reNpuG3IbYhyUby39z2s2b0G1y67FseTx3Fu93MxtmYsxvQY\ng06RTpbfSzb/fT7prVSI7IEkZxCRqaurQ8eOHTF37lz89a9/xT333INBgwZhzpw5GDNmTIu/FwQB\nN998M1YioTvAAAAgAElEQVSvXo2qqioMGTIEkydPRv/+/dMeN2rUKCxZssSu2yo9QSHotf8gvv1s\nLi4j1zN7fQSlK87utNts92k0lmOGW8rs1/yT7z/BFYuvwCX9L8HsYbPBMumvay7PFfFFMKHXBEzo\nNQEAsOf4HqzdsxZLv1qKO9fdieqKaoztMRZja8ZiaNehCHit7amWrb0IYDy9tVgszVLC6/ViyJAh\niEQiePnll9G2bVu899576Natm+bjN2/ejD59+qCmpgYAMHXqVCxevLiFoNh9QChZQQHSX0yO4xCN\nRgtql27Hm5Mt20wLu2Io2WI5bmbxjsW4bc1teGLME5hy0pQWv89moWSjR2UPXHPaNbjmtGvAizw+\nOvARVu9ejQfffxA7juzA2VVny/GXPm37GEoWKVSM9dq7u7E1jFMJHE5bZFqFjZFIBOFwGHV1dbp/\nt2/fPlRXV8v/7tatGzZt2pT2GIZhsHHjRgwcOBBVVVV44okncPLJJ5t/EwpKVlDIm5Svi0vvemah\nFgG9lGC3QIQuFAohEAjY/uXP9/lEScTcD+fi9U9fx1sXvYVBnQZpX18jhpIvXtaLoVVDMbRqKGYP\nn40j8SNY/816rNm9Bk9/9DS8rFe2XkZ1H4XKQKUpz5sJo4PFlI0SWxNOCyrBaFDeyHrPOOMM7N27\nF+FwGMuXL8eFF16IHTt2mLFMXUpOUJSuFuI6kiTJlPYfVs5YKTQl2Ky1aQmdWyc+ZqMp1YSbVtyE\nA00HsO6KdRnjGgys21DahdrhopMuwkUnXQRJkrD9h+1Ys2cNXt72Mm6svxGndDhFFpgzOp0BD2tt\nSrjSPabunEze71LJHnM7agtFEARDXcKrqqqwd+9e+d979+5t4R4rLy+X/3/ixIn45S9/iSNHjqBd\nu3YmrFybkhMUAglq5+I6shMieIIgoLGxET6fL283klX3VojQOZnKDDTHNC5ffDkGdhqIpZcsNRTD\nKMTlZRSGYdD/hP7of0J/3PyzmxHn4ti4byPW7FmDm1fejIPRgzi3+7kYXT0aI6tGolewly1rIu6x\nWCwmHxpaQ+dkt9VKGW0cOnjwYHz55ZfYvXs3unbtioULF2LBggVpjzl06BA6duwIhmGwefNmSJJk\nqZgAJSgoREg4joPf7zctp9uKDVIQBDQ0NLhuEBbgjnhJvq/5xm83YsbSGbhtyG345aBfuioOpSbk\nC2FsTbN1glHA/sb9zbUvu1bhwQ8eRMdIR9l6GV41HCGftVXUkiTJVomdnZPdtrHbhfK+c/n8eb1e\nPPPMM6irq4MgCLjuuuvQv39/vPDCCwCAWbNm4c0338Rzzz0Hr9eLcDiMN954w5J7UMJIJeYwPXr0\nKDiOk1sdmNXGQBRFHD9+HG3bti34WpIkobGxUa4sL3RaHMdxiMfjqKioKHhtDQ0N8Pl8SCQSBcVL\njh07hvLy8oLcjPF4HJIkIRgMguM4eR2xWAyBQEDz2q9uexV/2PgHvDjhxeZN2iAvbH0BXxz5Ak+N\nfarF7ziOgyAICAaDed9LrnAchxSXwufHP8eaPWuwZvcafPr9pziz65kYVzMOY3qMQf/2/U3fhKPR\nKEKhkK6bS+keEwQBgDmDxTK9p1ZCsuCcqk5vamqS68skScLEiRPxwQcfOLIWMyg5C4UMwSKT2NyG\nMq6jrKJ1A+Q0aka8xKwTP2nrkkqlZL++1nU5gcPdG+7Guj3rUH9ZPfq27Zv7em1weeWCh/VgcJfB\nGNxlMO4ceieOJ4/j3W/exZo9a/DC1hfAiRzG9BiDsTVjMbr7aLQPWd9UUOkeU3ZOVrvH3NA52ShO\nrdGN+1OhuGc3Mwmv1wtBEEwv9jNjg1TOWGFZVp4O54a1kRodURQRDoddEXyXJEkWklAoJLtelIFj\nr9eLY8ljuHrp1fB7/Fh7xdq8sqYYMHCZnrSgMlCJC/pegAv6XgBJkvDVsa+wZvcavPHZG7h11a3o\n27Zvc2PLmubGll7W2q+30c7JRtxjrdXlBaQLWrG/BiUnKFaTzwdfK1OK4zjXnFCUiQHkZOk0yhNv\nJBIBx3HweDzw+XyIRqOyf/+/B/6LGctn4Pze5+OB4Q/A78s/LdxtFkomGIZB37Z90bdtX9w46EYk\n+SQ27d+ENXvW4I61d+Cbhm8wonpEc3ymx1j0qOxhy5qy9a5yW+dkNwlZKpVyxUGuEEpOUMiHww3t\n5oHCqvONUsi9plIpRKNROTHASA8hqyFjg30+n+aplqS9rty9Er9c+Us8MvIRXHLiJXJPq3yyksys\nQ3GCgDeAkd1HYmT3kZgzYg4ORQ9h7Z61WLtnLR7Z+Agq/BVy8H9EN+sbWwItB4tptXbP5MYsddRi\nRr6HxUzJCQrBiqydXNuJkEFYPp8P4XC4hWnr5JdIr1eYWevK5zrqNSkDv+rHPbXlKby07SUsunAR\nhnQZAgBZi/YyWV9uOaWaRadIJ1x+8uW4/OTLIUoiPvn+E6zZvQbPfPQMrlt6Hc7ofAZ6VPTA0Kqh\nuPLUK+W/s+rEnq1zMtB8kFC25m9tkFlCxQwVlBwxek1yyrYjJTjXe1VaTW4Z0au1Jp7nWzwuxsVw\n05qbsKdhD9ZevhZdy7um/V6raE/tdtGbqFdMLq9cYBkWAzsOxMCOA3H7mbejKdWEl7e9jAffexD/\ne/h/0wTFLtTuMZKiTg4CgP77ZCZOurzUz00FpZVhtFtrLBYDx3GGmifaTSarySmUMZxMa9rXuA9X\nLLkCvSp6YeklS1EWyO62yeZ2kVuOiJItLejdwLo96/B/t/xfnNfnPAzsONDp5cgQF2c291ixZI/l\nCnV5uRCrYihGrqksBqysrLT1Q2/kXo1YTWa+bkbnvZDah0x1Hpv3b8ZV/7wKN51xE2aePBNBb+41\nIXpuF5L2yvN8WtsRJzctq07Of/r3nzB/y3y8ddFbeOuLtyzPBMuHbJ2TScp9sbeG0WoMSQXFpdjt\n8lJujEaKAc1cn5HOtUZnq5iFkTVl6xFGXqPXP3kd96y7B8/WPYsJvSbIBY9mrFHuyOsPgPU0N1BU\nn4pLoWGiIAq4e8PdWP/Neqy6fBW6V3THos8XwcO4Z5S0HlZ2TnZTlhdpDFvMUEHJ8ZpqlJt1LsWA\nVqxP68vh5GwVPYz2CONFHrPfm41V36zCskuWod8J1s10J7NRtGoqlLUvhVaEO0GMi+G6ZdehMdWI\nlZetRJtgGwCAIAlpnwenRJM8r9HX1GjnZLe7x2iWVytHLQJu2az1vjAkXpLLDBirYzuiKKKxsTFr\nj7CjiaOY9s40CKKAd698F2Ue6+e3K4PyylMxSRDweDxFVxH+XfQ7XPrOpTip/Ul47fzX4Pf8VKcj\niAK8TMstwK33ooVe52Sjg8XcZHmWQlDe+eOqyVgZQ1HC8zwaGhpymkmvhVVrTCaTaGxslGeP271J\naL3+HMfh+PHj8Pv9cv8iLXb8sAMj/zwSJ7U7CQsvWIh2IWs7pALZ29eT4H4oFEIkEpFjUMlkEtFo\nVG5Iauco5mx88cMXGPfGOIzvOR7P1z2fJiZAs4Vidat8uyEHgUAggEgkglAoJB8EYrEYYrEYksmk\nbHWSv3ECrRgKdXm5FGUHT7M+MKSdSyKRKHgQltkfYuUGTrLM8o2XWCFyRl+zFV+vwPVLr8dDox7C\nFf2uAMdxpq9FD6P3rQ4au3Ea4vt738eMpTPw0IiHMO2UaZqPEUShKGIohaDlHlN2TmYYBizLQhAE\nxy3NeDyODh06OPb8ZlCyggKYM9dcDSnCKnRgF2D++kRRRCwWA8MwebvgzLbsjKZRS5KEpzc/jae3\nPI2FFy3EsG7DkEwmbXNJFNJ6JdumZaSw0kwWfb4Id62/Cy+f9zJGdx+t+zhBElyR5WVXYFwre4wk\neKjdY3bEybQsFLO6ozuF858mk7HqQ0By4lmWde089cbGRlcNFCNt+rMJXIJP4Ff1v8Kn332KDVdu\nQPfK7gDsd0WYlTmm3rSMFlYWiiRJeGLzE3h126v45yX/xMknZJ4fzou8nIzQGiHWCREQpzsnx+Nx\nlJVZ3xLHSkpOUJSYddomKcFmn1zMWh/JzQ+HwwXP7CBuvUIhp79sAneg6QAue+syVFdUY+30tYj4\nnfEhZ7NQ8n2fjBZWFpqRxAkcfrPmN9j23Tasvnw1upR1yfo3glT6Li+jkJ5xyiw/Ymnm2jnZKJIk\npR2yaB2KS1G6kgrZsNX1G27qEAyku5NIzYQbSKVShiZmfnTgI1z21mW4/vTrcdewuxy1qjI1hzTz\nAKFXWEliRcrTslEakg246p9Xwct6sezSZYYbP6pdXk7VZLip/QnBiKVpdudkKigupxBBISnBSneN\n2QHiQtfX2NgIlmVRWVmJhoYGU9eWD0oBJq3w9Zj+znQs3rEYdw+7G78b+rucvpCW1BjB/vb1ytRk\nUjxJxIXM9CEbm571sq9xHy5++2IM7ToU/2fM/8kpJiKIpZflZRVqS1P5XpEi2FzdY7SXVytBOQgr\nGAympSK7IS1Ub31OdQkmz00mUVZUVGStZi/3l+Ok9idh6VdL8fSWpzG6x2jU9a5DXa86VJVXpV3X\nDpzqraZ8fqXLhbhZMgWMt323DZe9cxluHHQjbh18a86nZKuzvARBwBdffAGGYXDSSSe5oqhWTT7v\nuZZ7TGuwWK6dk2kMxeXkukko24Fopbeavenks75kMqmZfuuku4j0MPN6vXLNS7b1nNzhZIR9YTxZ\n+yS+i36HVbtWof7resxeNxtdy7tiQu8JGNt9LE5re5ot95CtDsVuSMA4EAiktRshPv0N+zbglrW3\nYN7oeZjSb0pe77+VWV6LFi3CnXfeKYthJBLBU089hcmTJ1vyfIVQ6HdHaWkC6WnkmTonq7/7tFLe\npeQTQyEnbFEUTUkJNkKu6zPSrsRuiLVktIcZwct6wUvN7ek7Rjpi2qnTMO3U5sr4LQe2oP7retyz\n4R58ffRrnFtzLsb1GIfamtoW7erNxE3t69XuEGVq8ivbXsEjGx/BqxNfxeCOgxGNRvNqlmhVltey\nZctwyy23yLEGoHmznDlzJsrKyjBmzJi0x7upn5YZ6LWG0eoRp6QUXF7us0FNxOiGzfM8jh8/LqcE\nu63lvCAIcoxEb31mrc3odYg119TUhLKysjTXmxE8jAeC2HJ4lof1YGjVUDw48kG8O/1dbLxiIy7o\newHW7VmHs18/G8P+PAx/+NcfsHHfRnCCeTGtQupQ7EKURMx5fw6e/uhp1F9Wj5E1I+VOCGSWiF41\nuBbqLC+zPtv33XdfmpgQ4vE4HnjgAVOewyysFjMSAyPtj4hngcRhEokE1qxZg1deeQXBYNDQQbG+\nvh79+vVD3759MXfuXM3H3Hrrrejbty8GDhyIrVu3mn1bupSkhZILuQ7CstvlpRcvcRKSXcbzfN4C\n52E9EKSWgqKmY7jZernkxEsgSAI+Pvgxln+5HLPfm409DXswqvsojO85HuNqxhlKldWDAQM360mS\nT+KmFTfhm4ZvsHrqapwQPkH+nZHCSi1/viiJLVxehX6+4vE4du7cqfv7bdu2lZxFkgtK9xjP8wgE\nAvB4PFi/fj02btyIU045BXV1dRg/fjzGjBnTogxAEATcfPPNWL16NaqqqjBkyBBMnjwZ/fv3lx+z\nbNkyfPXVV/jyyy+xadMm3HTTTfjwww9tub+StFCM9PMiLqR4PI7y8nJDYmLnl4BkTBELIFuxopnW\nU6brkOyyQl2DHsYDXmw5kTHTWrysF2d1PQt3nXUX1k5diy1Xb8GEXhOwZvcanPXaWRj++nA8+N6D\n2PjtRkPXVuK0hcJu2QLPu+9q/u5I/Ah+/vefgxd5/OPif6SJiRpyIg4EAgiHw4hEIvB6vXK6azQa\nla0XXuRND8qTIVl6+P3+VismWrAsi9GjR+PVV1/FwIED8frrr6NTp06YN28evvvuuxaP37x5M/r0\n6YOamhr4fD5MnToVixcvTnvMkiVLMGPGDADAWWedhWPHjuHQoUO23E9JWyh6mywJIhMXl1Gfs11B\neSfjJZm+7KR7cSAQKNha8rDaLi+jawGa56ZPO2Uapp0yDbzI46MDH2Hl7pW4c/2d2HN8D0Z3H43a\nnrWGrJdMdSiWI4oIXXMNEIsh+sUXgGIEwq5ju3Dx2xdjYq+JeGjkQznHPPQKK1OpFFJ8CjzXbMnk\nWvuih9frRW1tLVasWNEiI9Lj8eDnP/95i79xYx2KE8/NMAwGDx6MwYMH4+6779b8m3379qG6ulr+\nd7du3bBp06asj/n222/RqVMnk++gJSVpoSjR6njb0NAAv9+PsrIy16UyGomXaGF1fEfZvdhoa5dM\n6/GyXkMuL0K25/OyXgytGor7h9+P96a/hy1Xb0FdrzrZejnn9XMw5/05+Ne+f2laL05aKN4lS8D8\n8AOYeBy+v/5V/vnHBz9G3cI63DjoRjw86uGCA+hKf344HAYYIOBvziCLx+Nyymu22Es25s6dizZt\n2qRlIfr9frRr1w4PPfRQQfdQKmi9vka+U0bFT319u0Sz5C0UgjIlOJdBWOrrWWmh5Dr10QrUa1J3\nC9AqVvzmm2+wcuVKSJKE8ePHo0ePHlnXrheUz2eNWqitly0HtmDVrlX4/brfY8/xPTi3x7morWm2\nXjqXdXYubVgUEZg9G0w0CgDwz5kD7oorsHTnUvx23W/x7IRnMbHXREuemhd5+L1+BINBuVCPdDko\nZI57TU0N/vWvf2H+/PlYvHgxGIbBlClTcMstt7iqm64bul7kWkNWVVWFvXv3yv/eu3cvunXrlvEx\n3377LaqqqmAHJSko6hiKKIqIRqOQJAmVlZV5WyVWWQFuFDsAaa+blmtQkiTcfvvteOWVV+Takzvu\nuANXXXUVHn/88YzXztVCKQQv68XZVWfj7Kqzcf859+Ng00Gs3r0aq3avwj0b7kGPyh7oXtEdRxJH\nwIu8rR14iXVCYOJxvPfYTNzVaSMWTl6IM7udadlzi5Iox1DI+8cwDEKhUFqxXj6deLt06YLHHnsM\njz32WNZ1OB2kd0NMJ5VKGRqFMXjwYHz55ZfYvXs3unbtioULF2LBggVpj5k8eTKeeeYZTJ06FR9+\n+CHatGlji7sLKFFBIZDKduLicksXXgJZH6l/KUTszEYQBDQ2NsLn8+kO6Hr++efx2muvyRsO4S9/\n+Qtqamowa9Ys3esbDcoDP/VRItlMhdK5rDOmnzod00+dDl7ksXn/Zvzp33/CJ999gl7P9cKYHmPk\n2EuniIVfRJV1AgBMNIqznl+MZR9tRPcTelv33NAubFQexrTmuBfbxEq3otW63shwLa/Xi2eeeQZ1\ndXUQBAHXXXcd+vfvjxdeeAEAMGvWLEyaNAnLli1Dnz59EIlE8Morr1h2Hy3WZ9sz2YwkSeA4DjzP\no6ysLO9BWErMtgJEUZRPJmVlZaZ0Ly0UpQhnS6WeN28eYrFYi5/HYjH88Y9/xMyZM3X/1mhQXim4\n5P5IFbkZJ1sv68WwbsNwOH4YkiThybFPYvXu1VixcwXuXn83aiprMLbHWIyuGo3hNcNNtV7U1gmh\nrRSAf/lGpK60VlByyfJSpyarW43kU1jpNE5bRkpymYUyceJETJyY7gZVH96eeeYZ09aWCyUpKCRL\niswBN0NM1Ncv9IOYSqWQTCbh9XpNGftphtiR1i6SJKG8vDyj643neRw8eFD394cPH5bbTmhhxOUl\nCAIkSZLfQyIwyWQSgiCkjRQodCMjzSG7lHXBladeiStPvRKcwGHzgc1Y8fUK3PXeXdhfv1+OvdTW\n1KJjpGPezwcAgQceAGIxSCwLUWrOiGIZFkwsjsgjjyA1fXpB189GviOAM7UaccvESrejZaEUe5U8\nUKKCQjaySCSieYLOFzO+GMogdzAYNLXZZKGt+kmqMkk1zYTH40FZWRkaGxs1fx8KhTIKeTaXF0lQ\nYBgG4XAYqVRKzlIim5TP55Nbihe6kWllefk8PgzvNhxndT4Ld595N47yR7F692rU76zH3evvRs82\nPTGupnlm++DOg3PenFO3347D33yOv372V5zWYSBGdx8lr5kLBgGLN2Kzug3nW1hJcJOl4BRUUFxM\nMBgEy7JpbhKzUPYJyxV1kJvjONMEpZAvJKnLISJhpBU+wzCYOXMmnn322RYxlGAwiKuvvjrjmvQq\n5ZUNMMPhsGYLD+Ua1DUWuWxkadcyUIfStbwrrhpwFa4acBU4gcOm/Zuwavcq/Gb1b7Cvad9PsZce\n4wxZL+vG9MaMpQ/hodmPYfgp06BsJJNMJi3PO7NiwJZS9IH0OSKZGiU6gZtqUIzGUNxOSQoKkHs6\nXi7XzeeapChQGeR2qjeYEnVrl1zWc99992Hjxo349NNP0dTUBAAoKytD//79cd9992VuvaJhoahb\nuuSCeiPTc8PoBZFzrUPxeXw4p/ocnFN9DuaMmIP9jfuxevdqLPtqGe5cdyd6tenV7BrrWatpvSz8\nfCHuXn+37tx3SZIsj0Wo29dbscFmKqwkok/cmq3ZUqEWiotRZqq4gVz7heVDPuKUSCQ0W+EbvU4o\nFMKaNWuwatUqvPnmmxBFERdffLGcgZJMJnX/Vm2hqAeakXiJcpPJZcPRc8PoBZELrUPRsl5W7lqJ\n21bfhgNNB2TrZWyPsXjt09fw2ievGZr7biVWtq/XQin6pEEiEReSQGP2FES3ov4sl0KnYaBEBYVg\nhQWQyzWzFQU6ZaEQS4DjuIJbu3g8HkyYMAETJkxI+7kgZA64e1mv7O4jKcqZUruVr1Wur5ue9aJM\ngSXuRzNOyUrr5aGRD2Ff4z6s3r0a//zqn/jVil8h5Avh46s/RueyzgU9T6FY0csrF0hwn+f5tIaJ\nhRZWGsVNFhF1eRURZn9wjGxm5MQNIKd+YflidJPVGm3sxHqIy4sE36203tRopcAyDANBFNJmixgp\n4DNCVXkVZgyYgRkDZuAP7/8BxxLHHBcTIP8sLysgqeBaUxDzKawsNqLRKMrLy51eRsGUtKBY8aEz\nck0SL8lWTGm3hWJ0XYD1pzcP4wEncIhGo3l3BzADckr2+XxgWRbhcLiF9UJqXsx4TYK+ICqQW3zI\nKqweAVwIpV5Yqf4sJRIJdO7s/CGjUEpSUNQdPM3cHLOJgB3xknzWZdQSsOOLKUkSuCQHTijc5WYW\npA5Fy3oh7rBYLJZ2Sg7eeSe4Sy+FOHiw4eeRJMk144bdZKFkw4rCSre5vGgMpQiwywpQxiX0mig6\nsTYjzR3tRHa5gQEYZBQTOy04recip2RCIBBoniPC8xA++gjlL7wAZtMmRNeuNXxKliDBJXpiS5aX\nEXJ93lIorNQKytMYiotRWiZWzzApJC5h5trU15IkCU1NTbrNHfUww6rTep2UwfeKsgrbmkMagVgo\nmVD6+IM/Nj30fvEFhLVrkRg2zFCGktssFDuzvKwi38JKaqGYT/F/mgxgpaDkEpfQupaZ61JCihW9\nXq9uc0c7IfUuxOXmTXhznqpoJbm8Pux//wvvpk1gJAlSLIbKRx5BdO3atAwldQBZTn2GewTF6Swv\nK8ilsNLJGjB1nREVlCLByo2UxEvUdRxGscqlQzZvJ+eqKCH1Lsrgu9GZ8nZitLAxcN99wI+ZRwwA\nz/bt8G3cCPacc5qv86OPn+d5udKfiIubTsVuiaFY+ZpkK6wkYq8WfruhLi+XY6XLi7RQMaOOw+x1\naW3e+VzLjNdM2aRT/Trl0r7eDhgwMKIn7H//C8+P1olMLIbA7NmIrV/ffC2Fj1+5iXEch2QqCQ/r\nkcfuOrWJiZIIBkzBUyCLCXVhJUlHJhMrARhyW5oBbb1SpFhhBZAuwZWVlaY1jCz0OiStNZlMukbk\nRFGEKIpy5bsSD+uRO+xmQytYbmZTTXJNIxaK0jqR/xYAu307PO+/D+FHK0V5XaULxufzgQGTtok5\n0duKF3lXWCdOQsScVO0T95hdhZVKqMurFUI+aF6v15T5JWZ9QInFBEBz886VQkWYxG8A6L5OXtZr\naB6KXRhpDsns2gXv+vWQysshqnuBJRLwP/kk4ipBUSNBgpf1yn3TtHpb2eHf16pBcZM7zm6U4gJY\nX1ipVYdCBaUIMMNCUXbAJTUcZte15Hs9ZVIAqfh2EmWzyWQyqbse17m8DFgoUnU1YsuXA7z2ukXV\nbG/Na+Cn91qrtxUJIAuCIFt4VlgvbsrwcjJdWS/z0e7CSjuagdqBOz5RFqD80po1J6SiokI2id2A\nMinA5/O1aCNvN8r4jcfjybieXIPyVtekiKKIpKDfzBIA4PVCGD7csjUoA8ixWEyus8inHX82SjHD\ny0rMLqxUi2ipWIclKyiEQjYi5ZwQ4kridU6ndq5Pq1jR6lqbbOtRN5vMFuPwMu5JGxYlEX/c8kds\n3r8ZV/7jSozvOR61NbXW9NuSYChtmPS28nq9LawXM4r3BEloVQF5LfLdxM0urHR6hIWZlLyg5It6\nTohZFk+hqId0qU9Ddp908i3qdEvacJyL48YVNyIhJLBpxiZ8fPBjrNy1EvduuBc1lTUY33M8xnYf\niwHtBpjyfEqXVy5opb/mO0wMaBZRt7i8ip18CivV31MnU5bNpOQ/UblmBEmShEQigUQioZt665Q1\nQCrNlUO6lNexm0zryYaRoDxgrYAfjh3G1MVT0b2iO5ZcvARBbxD92vfDtFOmpc00+fWaX+P72PcY\n13Mc6nrWYUyPMWgXapfXc5pRKa/OHMt1mBjgriyvUnH3AMYLK8nv5ILXErFSSlZQ8rEo1PESrdRb\nKz74RtZntLmjGV9OI6+ZsngyGAzmfA2ng/JfHvkSU96egov7XYzZw2a3cP8oZ5rcP+x+7DyyE+8e\neBeLti/Cr1f/Gqd2OBXje45HXc86nHLCKYZf83wtlEzkOkwM0B7/WyqBYaPYIWR6liXQnCr8yiuv\nQBRF2a1pdD1HjhzBZZddhj179qCmpgaLFi1CmzZtWjyupqZG3st8Ph82b95s6v2pKflPj1FBEQRB\nnjZC2BQAACAASURBVKWeqY7DikLJTJB4CWnz7nSnYKA5+N7U1ISysjJNMTECy7CQIBmuRTGT9/e+\njwmLJuCOs+7A/cPvNxRLqC6vxvUDr8eiCxfhq1lf4Xdn/g4Hmg7giiVX4OT/ORm/XvVrLP1qKZpS\nTRmvY3UvL3JCDgQCCIfDCIfD8Hg84HkesVgMsVgMyWQSHM9Rl5fNkPeGpCZHIhH069cPX3zxBT76\n6CPU1NRg1qxZePvtt7N6VR5//HHU1tZix44dGDt2LB5//HHd51y/fj22bt1quZgAJWyh5AI5/TvR\nqiSTQCktpsrKSsdPj1rB93xhGAYexgNBFMB6cruvQkT9jc/ewD0b7sFLk17CuT3OzesaIV8ItT2b\n58VLkoQvj36JlbtW4vmtz2Pm8pkY0nUI6nrWYXzP8ejTtk/a39rdy0svOymWiIGRGLnvmFNdqEvF\n1ZMPDMOgtrYWgwYNwrFjxzBv3jzU19fj7bffxoUXXpjxb5csWYINGzYAAGbMmIHRo0frioqdr3HJ\nCooRl5eReInWde14g7QyzOxam9Z18p1AmcmMJ4F5H6wfriVJEuZ+OBd/+d+/YOklS9H/hP6mXJdh\nGJzY7kSc2O5E3Pyzm9GQbMD6b9Zj5a6V+OOWPyLiizS7xnrVYXjVcEfjBcrsJH/AD6/HK1svyWRz\nyjRxe9k9tMqpOhQ3PC+pku/fvz/69zf2uTx06BA6deoEAOjUqRMOHTqk+TiGYTBu3Dh4PB7MmjUL\nM2fOLPwGMlCygkLQ22RJa3dRFHM6/Vvh8lJfTy/DzCnyCb4beYzRavlCX++UkMKtq27F5z98jjWX\nr0GnSKeCrpeJikAFJvedjMl9J0OSJGz7fhtW7FyBRzc+iu0/bEf7UHsM6DAA+xr3oaq8yrJ1ZIMU\nNir9+6QVTGsYuesm9BpD1tbW4uDBgy1+/sgjj6T9O1OG2AcffIAuXbrg+++/R21tLfr164cRI0aY\ns3ANWqWgKFu7m9FCxUxIcWA+HYytsFCyBd8LIZfAvCiKeZ0ojyaOYvqS6agIVGDZpcsQ8dnXgI9h\nGAzsOBADOw7E74f+Hj/Ef8ANy2/AruO7MOz1YehW3k0O7A/uMtjWmIa6sJFsSmQcMgkeKyvD7epr\nZRdOudv0LBQ1q1at0r1Gp06dcPDgQXTu3BkHDhxAx44dNR/XpUsXAECHDh3wi1/8Aps3b7ZUUEo+\nKK8mlUqhoaEBgUAAkUgk5y+GVRYKiZckEglUVFTk1Q7fbMwIvmfCwxirRSExLhJU5nlefs0ysevY\nLtS+UYsBHQfgLxf8xVYx0aJ9qD1ObHcipvafiq9v/BpPjnkSDBj8du1v0fv53rhm6TV447M38EP8\nB8vXote6nnwfyDCxUCgkH26IizgWiyGRSMjvQyE4nTLsBmEk2Zu5MHnyZLz22msAgNdee00z5hKL\nxdDY2Cg/x8qVKzFggDn1VHqUrIWijqEQkz6ZTBbU2p1g5hdBkiQ0NjbmXByoxiyxkyRJLsoqJPie\nrU+Zl/VmFBSSZqlsnCeKYosZ71rtxrcc2IIrllyB3535O8waNCuv9VsBSRv2sl4MrRqKoVVDcf85\n92N/436s3LUSS75cgt+t/R36tu2L8T3HY2LviRjYcaDpG58gGu/llRZ7UVTtZxsmRtFG/Z2Ix+M5\nC8pdd92FSy+9FC+99JKcNgwA+/fvx8yZM7F06VIcPHgQF110EYDmnn/Tpk3D+PHjzbsRDUpWUAhk\nUyOjcAvNljL7y0JOfYFAIOeJj1agzJMvRNyM4GH1XV7EYgN+6lgsCEJaUVgwGNSsSP7nzn/i9rW3\n49m6ZzGx10TL1p8PeoLftbwrrj7talx92tVI8kms27kOa79di2uXXYumVBNqa2pR16sOo7uPRkWg\nouB18CKfV+sVIhjqrrxaw8TcOs8dcFd2WT7Dtdq1a4fVq1e3+HnXrl2xdOlSAECvXr3wn//8x5Q1\nGqXkBUUQmk/ALMuaNgo328nbKKlUCqlUSg52m7WufCHBd1KMZXWasl5QXhRFNDY2pvnriT+ftHYn\nva78fr98auY4DvM/mo8Xt72IBZMWYFDnQRAEwXU+/2xrCXgDGNltJGp712KeZx6+Pvo1Vu5aiZe3\nvYwb62/EGZ3PkNOST2x3Yl73JkqiKc0hldaL1kwRt1svbsjyysfl5VZKWlCI7x2AqXPVC924lc0d\n3TCiF0gPvpPRqFbDMmwLC4WIGvHfNzQ0yKJAxIS0dud5Xk5xFSQBd757Jz7c/yFWX74aXSNdNavF\nrchYYvbvh//pp5F8/HEgW6GqwTHDSnq37Y2b2t6Em864CVEuig3fbMDKXStx4d8vhM/jk62XEd1G\nIOQLGbqmFe3rjVgv6mFiTsdQ3EAsFkNlZaXTyzCFkhUUsmmXl5ejsbHRNR9cpfutoqICqVRKtqLM\nun6uqMcGx+Nxy+pZlKiD8kTUSHsZ8rfxeFxOb+U4DoIgwO/3y26wo7GjmLliJiRIWH7JcrQJNbeg\nUPe6UmcsmXVq9j/8MHx/+Qv4sWMhZPFRF1opH/FFMKn3JEzqPQmSJOGzw59hxa4VeGrzU7hm6TUY\nVjUMdb2arZfuFd11r6PVvt5sN5DaetEaJsayrCPuJyf3A/VzJxIJVFU5l0JuJiUrKCzLygWBdtSO\nGEGZrmymxaRcVy6YWfmeD17WK7deUYsacZ2EQiHZnUUKK5XWxrcN3+Lity7Gzzr/DHNHzoXP40Mq\nlZItF+V/ymrxTKfmXGC+/Ra+N98EAzTPla+tzWilmFkpzzAMTulwCk7pcApuP/N2HE0cxdo9a7Fi\n5wo8svERdAh3kF1jQ7sOhc/zUyJKtiwvs1E2TSQuSkEQwHGcHC9zYhSyG4jFYgiFjFmWbqdkBQWA\nfPqxoro91+vp1XPYVXmvRm0pabXBtxoP6wEncIjFYkilUrKoEZcWiZNIkiRbFqFQSO6qu+mbTbhm\nxTW44fQbcPtZt8tzWMjfkzYjANLiMdlOzbm0gfc/+ijwY98ldu9eeFatymilWHkybhtsiyknTcGU\nk6ZAEAX8+9C/sXLXSsx+dzZ2HduF0d1Ho65XHWprajVHANsJeR8Yprn9SzAYLOh9KCa06lByDcq7\nlZIWFKvIZUMgm6Hy9G3luow2wsxkKdkVa/IyXjRGG8EHeNmaVIoJwzByejDLsnK2l9frxbp963DD\n8hvwxLlP4Pye5yMajcrJBMr0VjJGVykuRFiUg5AyDbHSuw/ZOvmxLTkTjWa1Uuzq5eVhPRjSZQiG\ndBmCe4fdi0PRQ1i1axVW7FyBu9ffjTJfGTjJ+cmjpNWLkffBTOvFLS5wIL+0YbfSKgTFKZeXsrlj\npnb4dlooVla+54IoioDU7HopLy+Xf6bMyhIEAdFoFH6/Py154cWtL+Lxfz2Ov/3ibzir6iwALQPA\nkiTJlghxdSktEqBZWImwkOfUajVO3DKJRCItHVZpnRCyWSlWtK83QqdIJ0w/dTqmnzod7+99H5ct\nvgwjq0favg4j6L0PpWK9EBElUAulSFBmkdjtViKpr8pYjtVku08ygz6bpWT168XzfLOF5GluUgg0\nb+7ki8YwDDiOQzweRzAYlLOGBFHAvRvuxYqdK7DmijXo2aZn2pqJgASDQbkAklyHzINQurvIc2az\nXjwejxyXIemwvoMHUaawTuR1ZLNSDI4AtorlO5fjlyt+iT+f/2eMrRnr2DqMooy9APkNE9PCTXUo\neq1XipGSFhQldlooPM+jsbHRUHNHO8ROmabsRPBdiXJQmN/jhyAJ8sZO3FypVEqujieFjDEuhuuW\nXoejiaNYc8WarBMTWZZFIBCQs8VIHQtxjRHLRVk/QdxtAFrUr6jTYf0vvgjwPMSyMvz4AFkmPNu3\nw7NxI4Thw1usyykLBQD+3//+Pzzw3gNYdOEiDOkypMXvnXAD5fqc+QwT08NNLi9qoRQRVmRT6YkA\nsQLyae5oxbqyBd/tXI86k4tlWCS5pGyZEOHjeR6RSEQWvkPRQ7j07UvRt21fvHr+qwh49YeM6a2D\nuFCU1ksikZCr74m4sCybFtgn/082L7nQcuZMSMOHt3ClkXsR+vWD58fHKyk0bThf5n80Hy9sfQFL\nL1mKk9qfZPvzW4Ge9aJMD8/HerEaGpQvcuy2AsrLyw0PLLJybfmmKZu9Jq30ZEFozjISJVEWk1gs\nBkmSEIlE5I3488OfY8pbUzDtlGm4Z9g9BW8KWpsQiZMQ15hSYIjvnqyZiIanVy8IvXunCYYH6TPE\nkz8mEyg3NbstFEmScP9796N+Zz1WTF2BbuXdbHtuu9EbJqZV3OqmoHwikaBpw8WA0l1hpcsr3+FT\nZkOyooD04LuT1fhqC0mZyUUGbCkzuZTCt27POlz9z6vx6KhHMe3UaZasT92+hdRGEHGTJAl+vx/B\nYFDXNZYpsE82NTJjhOf5vFvx5wov8rhl1S3YcWQH6i+rR/tQe0ufz00oY2pAS+uFuFedaM2jfu/d\nJG6FUtKCosQqQcln+JTetczCSbebmlgsJs+dAdIzuTxMcx1KU1NTi0yu1z99HfdtuA+vX/A6Rna3\nJxtJuQklk0kkEgn4/X4IgoCGhoa0rDG1a0xd8wIgrSI/EAg0CwkjQRRERKPRtD5XZh9C4lwcVy+9\nGpzIYcnFSxxv3a+HXZup2npJpVLged41w8SooBQRVrmVlAHmQCA3vz7B7LWRzCYz2s4XAgmC+/1+\n2T+szuTyMB5EY9G0TC5JkvCHD/6AhZ8tRP3UevRr36+gdeQK2WzImAPyGpJqfeUJN1PNC9kolVlj\nLMvCw3oQ8DfP4slUsV/IRns0cRRTF09FdXk1nqt7Lq1CnpI+TIwIvZ3DxJTvbSlZJ0ArEhTiojAL\njuOQTCZzipdYCSmglKTCW/QXChFaUqgGQDOTCxLg9XvlxyT5JG6svxG7ju3Cumnr0DGiPYXOKkit\nCc/zKCsrS3sNSZaX0jWWT80LaTWTrWIfaBblXK2XA00HcNFbF2Fk9Ug8NvqxnFrUl9rmZhTi9gSg\n6aa0uh1/Kb3uzu+EFmJFDIWcYHOdRa+HGWsjwXeySZkhJvmsSdkVoLy8XN5seZ5Pa6OSSCTAcRwC\nvgBIwtMP8R9w+TuXo0O4A5Zfttxw11yzUCYFZBsLXUjNi1I4yAalrhQnMSXyOKMn5q+OfoWL3roI\nMwbMwO1Dbi+ZTcoK9D7fyvdWab2o2/EXYr2UkoCoKWlBUWKGoFixcReKMvgun/wLJN8vCRnPq3S3\nkRb9yrRgURQRiUTgZb3gRR5fH/0aU/4+Bef1OQ9/GPWHvAY/FYJeUoBRcql5YVgGPq9P7jsGQJ7z\nog7uk5iSkRPzfw79B5e+cynuHXYvZgyYYe4LZCFObq5GnlfLeiECAxRuvair5oudViEoZnxgycYd\nDAbBsqwpGzfw09ry+WKpg+9mrSlXlJlc5eXl8ibo8/lk9xexxDweD8LhsByU//yHz/Gb1b/BvcPv\nxfWnX2/72kWxOUDu8/lMyYbLVvPC8zwkUcpY86K+XrYT8wcHPsANK27A/Nr5OL/P+QWtn6KP2k1J\nDg+5DBNTH2xJN4hSoaQFxSyXFynIU27cTrZu0Kt8N8u1l8t11LUuwE+ZXORkzvO8bAGQFGufz4dP\nv/8Uy3cux6vnv4oL+l5Q8LpzhayLWBdmo1XzwrBM2kRKdcW+KIry50vpKlRaL8oT89tfvI3frfsd\nXhj3AoZ1GYZkMum6Qj43UqhlQAQjl2Fi6r8HSqvtClDigkLId6PVKsgj17NifUau62Tluxp1ixmg\nZSYX2bRJJpeyXUbXSFcwEoPrl16PYVXDcF6f8zCpzyR0Le9q+dpJvCMUClnaAVqJ0pVVUVHRouaF\nrIPjOIRCobR2/EDLmpdXPnkFc/81F+9MeQcDOgzQbUNiJBXWiQPSwYMH8f777yMcDmPMmDFFfVLP\nlmShF3ehgtJKICdphmFQWVmZ9iGwo/JeCyNt5+1aF3FlEatNGXhWZnKpe3IpT+3LLl8GURTxQ/QH\nrNy5Est3LseD7z2I7hXdMbH3RJzX5zwM6jzIdAHXWpddkPb16sI7EiPh+eaRyKTDMXGfKdOReZ7H\nk1uexILPF2DpxUvRq20vQ21IjAT27bBqeJ7HbbfdhoULF8Ln88mf26effhqXXHKJ5c9vNcr3Qp0V\nSNzSyWQSX3zxBQDkLCh/+9vf8OCDD2L79u3YsmULzjjjDM3H1dfX47bbboMgCLj++utx5513FnZj\nBmgVgpLrRku64ZK55lZ/yYysz87K90zrIVlaiURCTpnWy+RKpVJpPbm0YFkWHco7YNrAabjitCuQ\n5JJ4/5v3sXznclz9j6sR5aOY0HMCzut7Hs7tcW5B2V8kC43juKzrsgotS5TUuIiiKMegiG9eXfPC\neljcteEubNy3EfWX1qNDqIOua0yrDYldqbCZeOCBB/C3v/0NyWQSyWRS/vnNN9+M6upqDB061PI1\n2JkMoDw8KOuY5syZgw8//BBdu3bF888/j0mTJqF7d/2xzYQBAwbg7bffxqxZs3QfIwgCbr75Zqxe\nvRpVVVUYMmQIJk+ejP79+5t5ay0onfQCDfKJoaRSKTQ2NiIUCulm/JhtCWS7XjKZRFNTEyKRSMbu\nxVZbKMQFSGI3Xq9XPnkRq4TEd0gtRy6bNsMwCPqDGNdnHJ6ofQJbr9uKxb9YjJryGjz14VOo+VMN\nprw5BS/95yUcaDqQ89q1Gk/ajXrAlnpdyvYt4XAY5eXlcp+n403HcdU7V2Hbd9vwjyn/QLc23eQO\nA6QYkrjRSIsX4KcNLRAIIBwOIxQKya34o9Eo4vG4nLVkNbFYDC+99JIcZ1ASj8fx+OOP27IOp1DG\nXv7+97/jpZdewoknnogPPvgAP/vZzzB9+vSs1+jXrx9OPPHEjI/ZvHkz+vTpg5qaGvh8PkydOhWL\nFy826zZ0oRbKjygD3dmKFe1yLeWyJqtRugArKioAQHe6IsMwiEQiBZ0Aidvg1C6n4tQup+K3w36L\n75u+x4qvV6B+Zz3uf/d+1FTWYFLvSTivz3kY2Gmg7vMRIQRQ8LoKRdkc0si6iBgkxASuWXUNQt4Q\n/n7h3+GFFw0NDYbmvJDsOqUVowwmk2aWQPOGb2WH3j179mSM+23bts3U53Mb6n2DZVkMHjwY999/\nPwRBwHfffWfK8+zbtw/V1dXyv7t164ZNmzaZcu1MtApBIeiZuU4HurUEKp81WZXllSmTSzldkfTt\nyjYDJh9YlkWnik64atBVuPL0KxFPxvH+3vex/OvlmL54OpJiUnaNje4xGkFvUF5nNBq1bF25IknN\nFookNU/zZFk2q1v1cOwwprw1BaeccArmj58PL+uVr5XrnBee59NcYkp3Gs/zCAQCeQf2jdC2bduM\n1lDbtm0Lfg4juKX+RRmU93g86NKlCwCgtrYWBw8ebPG3jz76KC64IHtGpFP31ioEJdOLm0+Ld6st\nFLImj8eTtWrbakgmFxkZrDwB62VyWQ3DMAgHwxjfdzxq+9RCEAR89t1nWP71cszbOA9X//NqjOg2\nAhN7T8TIziPRrU03Rzsut0CC/JnLJnJ7G/bi53/7OS7oewEeHPFgi+SQfOe8qJtZks+zGYH9THTu\n3Bmnn346tmzZ0qIdUigUwg033JDzNYsZvSyvVatWFXTdqqoq7N27V/733r170a2b9aMLSlpQtDKz\nlD8rdL66WaccpUApCyjzOVGbKXTqwslMmVx2pt8qIafy07qehtO6noY7ht+B7xq/Q/3X9aj/uh6z\nN8xG77a9ZdfYgI4DHBUWURKRTCYNFVJ+fvhzXPjmhbhl8C24efDNGa+rleVFrBetOS/qZpZ67fgz\nBfYz1Vlk4oUXXsDYsWMRi8XkWEokEsHPfvYzXHvttYavUwhOWSjq543FYmjXLvP00WzX02Lw4MH4\n8ssvsXv3bnTt2hULFy7EggUL8n4eo5S0oChRbtrKnlPZ5qvrXcsKCm07b/a6YrGYKZlcdsKyLNqF\n2uGiPhdh2oBpSAkpvP/N+1j29TJMfWcqeJGXU5JHdh8pu8bsgOd58BwvWxSZ2LRvE6a+MxWPnfsY\npp48NefnyjbnhYiLz+eT3ZUkPRnQnvNC4jSk35i6zkJpvWSiV69e+Pe//43XXnsNy5YtQ3l5Oa66\n6iqcd955rmi0aickfT0X3n77bdx66604fPgwzjvvPAwaNAjLly/H/v37MXPmTCxduhRerxfPPPMM\n6urqIAgCrrvuOsszvACAkZws+bYBUnV87NgxlJeXg2VZRKNRCIKQcxaSkqNHj5rW1bexsREA5DXl\n+6WSJAlHjx5F27Zt8xYX4ttPpVLy/ZGTrDqTSxRFuY2KGyCNO5PJZAuRI5vqp999iuVfL8eKXSuw\n/eh2jKoehUl9JmFi74mWdjcmhZQ3rb0Jv+j3C1zc72Ldx67YuQI3LL8BL058EXW96kxfi7IfFXF7\neb1eOftLPfYYQIvAvhJlYJ/EaIwG9kmnArsPJNFoVL5fOyH7EenMMHfuXAwbNgyTJk2ydR1W0WqO\nAyRwTGITZHpgIdczQ4uVJ/9CEwIKtVCUmVzKn6nFhASAnc6YUkIsJq3W88BPrrHTu56O07uejjuH\n34lDjYew/KvlWLZjGe5adxdObHciJvWehEl9JuHUDqeadm/KQkqGYTLOlH/jszdw97q7sejCRTir\n6ixTnl+NsucUqbciSSDKIL1eYF+ZjPH/2zvz6KiqdO0/VZlIJZUAgmkI+RjES6AbIQFJd2OYp5AR\nQQkgImCMtBr0gihLWxBtnJDlvYJcR4aFBklCBiEJU0tkSoKCCkhE8KabQSIIZK6qVKW+P3L38eTk\nnKpTVWeq1P6t1auXUNTZOTlnP3vv932fl69LJbtinwiRms2r+FAzKM+mubmZVsp7K42NjW7HJuSA\n7V5MHHnVHAu78+Tt27fb1TGQtGAtZUwRSPqt3e7cep6g1+vRK7wXFo1YhIWxC9FkbsJX//oKJT+X\n4IFdbdXa5GgsPioeQf7ueX2R4j2yY3LUU37D1xvw7tfvYs/sPRjSY4hb1xMLSaQwGAzMkS/Xj4oc\njZHjMfIMsM0syWKIHah3FNjXorgoCVfIqKB4GTqdjsl66dKlC1MkJsX3erJDYftgkfNqKcflyvdx\nkxPIz2UymdqllMpppOgunlrPA233LKRLCBIGJWDaf0yDzWbD6ZrTKL5YjFcOv4Lzt85j3P8bh8SB\niZh611T0NPR0+p1COyaSNsz97OrDq1H0UxEOzD2AqLAovq+UDCEfM3acRGyfF3bGGDk+c1axzzZQ\nJN+hpJmllk75Scy0s9DpBaWxsZFxYJU64Ofug8kNvpPVtRoIZXIZDAam5zbZqZAgr1aQ2noe+H1S\njYmMQUxkDFbetxK/1P2C0oulKKwqxPJ/LsfgOwa3ZY3dnYjBdwzucF12jIlUvzN/x6mUt7ZakbUv\nC2eun8H+OfvRw9DD45/BEa74mInp88Jugcx3NCYU2CdCQqr0ye5FbGBfCtTK8mL/bNQc0ssglhNS\nT9ruPIxCle9S1rWI/S6ygma3MWbHc8iLDbSJTlBQEGw2G+rq6todg6h1TKfUjkmv1yOyayQWj1iM\nRbGL0GhqRNm/ylDycwnSctLgr/dHwoAEJN6diPui7kOAPsBh9Tt792iymtr8yloaUTy7GKGBobL9\nHEDH4zdXEKp5IQsSsTUv7N0LCfKT41MS2Cc1L3JW7GsFkmnaWej0gkKaEkldjOjq96ldjc8dC8l0\nI8kJXBsVIn42m43JjiP/lm1cSIrdiLgo8eKrYT0PtP3OQ4NDkRidiOmDpsNms+G7a9+h+EIxVpWt\nwsXaixgTOQZT+09FcnQyQnQdjzLIDqXWXIvZ+bMRERKBbSnbEOgn785PyhRvd2pe2LsXIqok/Zw8\nc3yBfS2YWUoJXx0KPfLyIsgvT47qdrHfx26oxBc0JsFOKXD2c/J5cjnK5OKOl/vi8/X0IDsYOV58\nNa3n2ZBd3Ig+IzCizwistK7Ev278CwcuHcDui7ux8vBK/LHHH5mjsUHdB7XdW9hRa67F1OypGN1n\nNN6a+JasLY+dZb9JgSs1L0RciGAAbQsErh0MESxHfd3dbcOtlQwvoE3otRST9JROX4dCttHEtFCq\noDzpOujsYeA2oeJ7kEnSgBQrldra2rZ+7TyTLcnkIrb8QEcxcTeTi0wSxOmWtABmZwh5AilGbWlp\ngcFg0EwhJdB2XxsbG9vFGxpMDfiy+kuU/lyK/f/ajyD/ICQMSMCRK0dQ01CDx2Iew3N/eU7WiU0L\n9ULcmhf2YoNMpmSnQnYvABzWvJDvZNfQuFKxT2I3auwMuHU3CQkJOHz4sGYEzlM6/Q6FoMaRl9jK\nd6nHxvddfJlcXBsVT+ISQscg3Awhd+IujoLcasPnY6bT6WAMNiJlcAqSo5NhtVrx7bVvUXyxGD/e\n+BET+k7AsnuXyTou7j1Ta8Ji7zTIcanFYmFideT5c8XM0llnRGeBfS3tUDobPiUoUh0rOUMo+K4E\nfC8KETZiM8MnJlLHJbjHIEINo5ytKkmNCaC+9TwXMfeM/Kz3Rt2Le6PuxQvxL8BmbZsASTBb6gQH\nrd4zMg6SRUgWMWJqXrhmltyjMbYdjJRmllLDFjMioJ2JTi8ocsVQhL6PxB9aW1sVt53nGwsRtrCw\nMMZVlpxdk6MGT7J/xMAXd+FOJERguFX6WiykBNyP5fj7+cPfz7/dit0doRWCPH9ibPGVhggw+565\nWvPiKGtMbJdKraGl35GnaO/uegl8IsAOvpNWrmqNiytsQplccgds+cbHLp5jryjZ6ad6vR5NTU1M\nR0KtvHREgKXImHJXaIXwBgEWumdia164gX0iLI5qXriBffJvLBaL24F9d+Eet2npdyQFPiMocu0C\nCGKC70qNjQibXq+H0Whk/owrJuRYRM2eK3zZPBaLpZ1fFDkbVxs5BVioSl3s0ZgcRZ5S4UxMV40U\noQAAIABJREFUuLhS80IckrldKoGOuxcS4yI7IRKcB9y34vcE4hDQmej0giLnkReJyXhqOw9IawfR\n3NyMoKAgyTO55Ia8XCQrCQBvN0I1ahHYQW4lBJhvxS50NEZ+n1qzxQE8K6YEXKt5IUe4YswsSUEl\nO4WZBPal7lJJ4L7jzc3NbvVh0jKdXlDYyCEozc3N7arN3f0uKSArr8DAQBgMBt7gOzvFNTAwUDNi\nYrfzW89zV6nuHgd5OjYpg9y1tbX4+eefERERgd69ezv9vKOjMa3a4gBgjgal3M25UvPCDewT4WDP\nA3yCJXdgn3xHZytqBHxIUKSedNirICls5z0VO5PJhObm5nbVyXJnckmFs6Mk9kvvznGQJxDzSXJt\nT54jk8mEZ555Bjt37kRgYCDMZjNGjBiBTz75BFFR4gwh2Udj/v7+zDFXa2urZmxx2DVDcqZ5s+8F\nAN54HFtgiFg0NzdDr9ejpaUFdnv7Pi9iA/tS7JKbmpokq4vTCj4lKFLtUFpbW2EymWC32xEeHq7q\nKp+byUXGJZTJpXaFORey+rfbXbOeZx8HkWJKqa1gpI5LPPzwwzh48CBMJhMzQVVUVGDcuHH4/vvv\nXVqtksUBu4hVjqwxV2EvDpSuGeKreWHfC+KaTQL1AESbWXLT392p2OezXelMxpCADwiK1DEUq9WK\nhoYG+Pv7S2Y77+7YuP5g5HssFku7YjGlM7nEIpX1PN8RSGNjIwAwq1NXJ1Ru9bunnD9/nhET7nXq\n6+uxY8cOLF68WNR3CaUs8x2NKXlMSJ414vqg5rPGvRdsy3yy2BJT88KNvZBjRfKdxIXDHTPLztYL\nBfABQSFIISgWiwWNjY3MOTp5QNWA6w9G/owcZZGdCvB//T40VmEup/U8N1OKTHJirWBI9buUR4PH\njx8XvP+NjY3Yu3evKEERG+R25ThIiudCK5X5QpjNZiYex95pOKt54ZpZsv9fqEulUGCfu0Mhc0ln\nwmcEheCO7QJZebGD79zgnie4KnZklxQUFMRkiZCHnzzEbBNInU6H+vr6DgFLtVDCel4oO8iZFQxf\n8Z0UOPLS0ul+N+oUgh2XcGenyT0OYh8Teno0pmUxIfVY7BgYd1fLrnkB0G7hIbbmRUxgn/ue0yMv\nL4R95OUO5IEkVu/tmiVJXNciRuzILslgMDCTg5hMLim9tTxBrcQAMVYw5LgwNDRU8rqXqVOnMkct\nXIKDg/HQQw8J/lup4xJCx4TuHI1p1eYFEFfo6WnNC/mdignskyQAk8mEiooK1NfXuywoOTk5WL16\nNaqqqnDixAnExsbyfq5fv36MO0ZAQAAqKytdv4Fu0OkFhQ1ZIYh96NnHSiRGwf4uKcclBpLJ5Y4n\nl5gJVe6eJlqynueer5vNZmblSSqopYw1hIWFYf369Vi+fDkzAQNtO5dp06Zh7NixvP9O7voXMUdj\n7MZZ3LGRn8XdGJhcuFNrJUXNi9VqbScu7PtLMsssFgvWrl2LM2fOoH///ggNDUViYiL69OnjdIxD\nhw5Ffn4+MjMznf4shw4dQvfu3UXcLenwCUFhbzXF7iq4x0rcB1LqQklHkBe3paVF0JMLgOhMLqHg\nLbuniZQFhNw0Ui1UvbMhx5dGo7GdwDibUF1lwYIFGDhwIN58802cOXMGEREReOKJJzBnzhze+6zG\n6l/oaMxkMrXLoNPpdEwwWmueYVLF5zyteeGaWZLvDAsLw969e/Hee+/h3//+N44cOYIXXngBzzzz\nDF544QWHY4qOjhY9fqXmJzY+ISgEsQ8WO/iuVLGY0O6JL5NLSk8u9gqKnBWTQDYJLnqSGcQ9X9dS\nYoBQyjJfYydu8NZdURw9ejQKCwtFjY04BKi1+hc6GmtqamJsQ7SUgg7IZ0EjtJPjq4UiR2PsjDHy\nP/KO6fV6WK1WjB8/HrNmzYLNZkNDQ4MkYyXjnTRpEvz8/JCZmYmMjAzJvtsR2noaZMbZroIv+O7u\nd0kxNqFMLj5PLlfqOByNgbvl5xYQurJa1/L5OpmwHaUs8x0TKmEFw06n1srqn/y85NiGHPtYLBY0\nNzdLupNzFyX9zMTUvAQEBLQTY4vFAj8/P0ZcfvvtNyapxs/PD+Hh4QCAyZMn49q1ax2uuXbtWiQn\nJ4sa39GjR9GrVy9cv34dkydPRnR0NOLj46W7AQJQQfk/HAXfHeFO1pgYHGVyETEhL5Cfn58sq1ih\nAkIxq3Wt+oUB7p+v8wVvpa7x8Ib7xnWAdhZrUOJnYIuJ0v5YfEfIbNdoIiKBgYHMs/Prr79i9+7d\nmDBhQofv279/v8dj6tWrFwCgZ8+emDFjBiorK6mgSIWzGIqj4Luj75RrjNwjN0eZXErZuztKteSm\nnQpNPFpAivvG3skJWcG4s1rXsmOwownbWayBXeMhx8+kpphw4dZCkRR5nU6H27dv45FHHkF8fDz2\n7t2LDz74AOPGjXP7WkKL46amJthsNhiNRjQ2NmLfvn1YtWqV29dxBe0caCsA38NstVpRV1eHwMBA\nl49l5AjMm0wmNDY2wmg0CooJqQR3xypfCoiABAcHw2g0Mn5Ezc3NqK+vR0NDgyYnRSKAUt83spML\nCQlBWFgYAgICmB1mQ0MDU1jp6FkhZ+hkFaul+8YdmyPIhBocHIzQ0FAmbmY2m1FXV4fGxkbG1VcK\ntCQmXIh5bGBgIIxGI7p3746MjAwcOXIEly5dwqJFi5CVlYWTJ0+K/s78/HxERUWhvLwciYmJSEhI\nAABcvXoViYmJAIBr164hPj4ew4cPR1xcHJKSkjBlyhRZfkYuOrsaqQAKQ3ofsCc6wPPg++3bt2E0\nGiXJWqqtrYVer2fSQz3N5FIDcp7OztcXW50uN2rUv7BX6y0tLQD4rWD4+tJrBbKjk2Js7CQHkl7r\nydEYdyesJfh2m7W1tUhPT8cLL7yAyZMn48yZM9i9ezf+9Kc/iY6NaB2fEhQSbwgKCmKC76GhoW5P\nzFIJit1ux+3bt5mGWCQ+IpTJZTAYNJV6SwoCuZYg3AlELSdcLdS/sDPorFYrk0FHVu+k3cAXX3yB\nH3/8EZGRkZgxY4bTCno5kVJMuLCPTdliK/ZojCwQtdgDhmRmssWkvr4ec+bMwfLlyzF9+nS1hygb\nPiEoxMSNZPWQLAuj0ejRxFZbW9vO7dUdyHGC3W5HcHAwAgMDHWZyObLwUAOxVdzsTBir1SqpK7Aj\nPG3wJBetra1MvxAAuHDhAu6//36YTCY0NDQwHk+ffvopJk+erPj45PAzE4IrtjabzWEcSutiwrV6\naWxsRHp6OrKyspCamqr2EGXF5wTFYrEgICBAkjTWuro6j144dtvglpYWBAUFMT5c7EwuraWQEthC\n58r95B4FyZGCyy2m1JIIA+3b4ra2tmLQoEGoqanp8DmDwcAUQSqFkmLCh6OjMQCa7U7JJyZNTU2Y\nO3cuMjMzMXPmTLWHKDvaestkhBQh6fV6TdREWCwW1NfXIyQkBMHBwczOiS0mZPdCgpxqj5kNOULU\n6XRuJTOQn8loNDIpzySoT1wB3F3rkGJKNXpyiMFsNrfrsX7w4MF2dixsbDYbPv74Y4/uhyuQ7CyD\nwaBaEzaSNUaSHEjaemNjIxoaGhireC2thfnEpLm5GfPnz8fixYt9QkwAH0kbJpM3CQxLWT3r6kNN\njohMJhNTPGm326HX65mjI7YFvRYDtVKmtwql4LprfaL1Yko+x+ALFy4wR19czGYzqqqqZLGC4SKX\n07InkIxCUkRJdiXclgRqFlTyiYnZbMaCBQvw0EMPYfbs2aqMSw208dTIjE6ng9FoZI6+pMQVQSGT\nndVq7eDJRfL4ST2DzWZjDOiIVYMWkNt6nl1M6ar1idaPB4ViTX379mVaAnMJCgpCdHQ0QkNDZbGC\nIWghcUEIclxNYowER/dDqYJKtuMCEROLxYJHHnkEs2bNwty5c2Ufg5bwiRgK20GVrA6lgJuG7Ah2\nfxJHNipkjMHBwe2q09nVuEq9LFzU7EnPDepziynJi63FCnOunxl3bC0tLbjrrrvw22+/dfi3wcHB\n+P7779G7d+8O38nOkvIkDsWO52gpcQEQn2km5f0QC1tMyAKmpaUFixYtwrRp0/Doo49q6jlUAm0t\nRWRGCf8tPtixENL/wFEmF3sFy3UElttHSgi1V7DO7C1IRbYWxcTZEVxAQAAKCwuRmJjYrvhSp9Ph\n448/7iAmQEcrGO79EGsFo9UsOMC1tGUhaxy2yamU9VDk98oWE6vVioyMDEyYMMEnxQTwkR0KebjI\nCluq3H72uakQJJMrODiYaT/K58kl9qiGnWJJArVy9gpn75q0Vv8C/F79TmJRWiqm5FvBOqKxsRG5\nubk4ffo0+vbti/T0dPTs2dPl67KfD0cpuJ1FTJzBlzXmyVEhEROdTtdOTB5//HHExcUhKyvLJ8UE\noILiEeyHig8lPLnIypRMHlJOpuyjGq3VvwD8R3BaKaYkiwR2oFYN2MemLS0t7VpEu9PuQAmULKh0\ndbfP3nGS7ESbzYYnn3wSQ4cOxbJly3xWTAAfExSymiU20Z5Cjhe4bTzZNvikEp8tJuQFltpyg28y\ndTcjiO/F0RJijuDUKqbUqskjuR/s7pRSN1PzFHYAXsmCSvZuX2hBxvdOtLa2YunSpRg4cCCef/55\nTdxDNaExFInhs8EnL7JOp2MmdjliElzXVz67eTErdS1bqJMjOIvF4vSohi/uInccikyIWiy8A37v\nTkkSQ+SMM7iKkmICCKesC2WNNTc3A2gvJsuWLUPfvn2pmPwfPrFDIV5Tra2tqK2tRbdu3ST5XpIL\nT2wyXM3kUiomIZQhxbdSV9oW3xXE2ryI+R454lByHtV4irNMM/YuXo2jQrWr87lw3xlyTM1Omnn+\n+efRtWtXvPLKKx69J4sWLcKePXtw55134vTp0x3+/tChQ0hNTcWAAQMAADNnzsSLL77o9vXkxKd2\nKFLD3vHYbDameNJRJpca7XDFrtTJ+LQ+IUrZmVKKYkpAexMiG2diAnRspsbtQChnyroW7x07LZ1k\nX/r5+WHz5s147bXXMGTIEHTv3h2vvvqqx/dj4cKFeOqpp/Dwww8Lfmbs2LEoKiry6DpK4FOCIteR\nV0tLCxoaGkRncqlZwU0EhBxnkcmUvDRkhU5WZFpA7up3McWUjlbqatbnOMOdeye0AJGjYZYWxYRA\nhJjtVbdkyRL88ssv+Omnn2A2mxEVFYW4uDhs2bIFffr0ces68fHxqK6udjoWb8AnBIU89OT/pZos\n2X5bzjK5mpqaNBekJTEdMh6DwcC44JLxqp1+q3T1O18feUcrdaliYS0tLfjmm29gt9sRGxsrSfxF\nisQK9gIEaF8k7KkVjDeICXtXZ7fb8dprr8FsNqOgoAB6vR4NDQ3Yv38/7rzzTtnGotPpcOzYMQwb\nNgyRkZFYt24dhgwZItv1PMEnBIWNVKtvEpchwXelMrmkhB2TYKeP8q3U5fSQEkLtbClnK3Vi6Olp\nC4NPP/0Uzz77LLOjtdvtWLNmDR577DG3v9PVGhixkKNCd6xx2GhdTEiiAltM1q1bh+vXr2PTpk3M\nOxAaGooZM2bIOp7Y2FhcunQJBoMBJSUlSEtLw/nz52W9prv4RFAeaMuqstvtuHXrFsLDwz2aFMnL\nSgLc4eHhTIdCsish19SqPxK3Mt9ZMSW3lkFsxpi7aDlbikw4xL3ak6B+aWkpHnroISaDiBAcHIz3\n3nsPDz74oFvjk0NMnF2TW9/BFhf2GIiYaPW9YCfbEDH57//+b1y4cAEffPCBLIk01dXVSE5O5g3K\nc+nfvz+++eYbdO/eXfJxeIq2KpoUwNM4SmtrK+rr65mJGADTsIsdfCeOwp6uXuXAVet5nU6HwMBA\nGAwGxk6cHPXV19eL6pnuCqReKDg4WJNiYjabYbVaYTQaYTQamZbNFovF5b7pq1at6iAmQFuN06pV\nq1y+p+zOpEoaZBIBIS0JSLEvtyUB2x5fa++FkJi89957OHfunGxi4oyamhrmOaisrITdbtekmAA+\nfOTlDiSTKzAwEMHBwcwkSpyBuZlcWqxC9vQYSe7aDi1aqBO4Ew753XIzpLj1P+x7wv2+s2fPCl7v\nypUraGpqYhYuzlD7iJAglPhBjpH8/PyYpBWtvB/c41/yLn/00Uf49ttvsXXrVtnEZM6cOSgrK8ON\nGzcQFRWFl19+mWmLnJmZidzcXGzatInxAtyxY4cs45ACnzvyqq2tdat5EF8mFzmWIX5J/v7+zDGI\nFqvL5bSel6K2Q8veUmJSb/n+jTObj549ewo21woICMD169dFPataERMh2Jlw7GQHtuCq5aLNrg0j\nCwW73Y7Nmzfj8OHD2L59u+biPFpFW0tAGfEkGE8yWkJDQ9utzMmxFwnQm0wmAG0r1paWFlWb/nCR\nO7WVW9vBzQZylDHGfaG1KCbupC2zd3NCjsCzZs1CdnY2syIl+Pn5ISkpSbSYaLXHOsC/62Rn0ZEj\nTrncC5zBJybbt2/Hl19+iezsbComLuAzO5SWlhYm/hEUFCQq44qsSi0WC4xGI9N3QyiTi3wvWaWz\nnU2VNidko3ZygCOPMZ1OJ0n1u1zIFeAm4vLLL79gypQp+O2335gFSVBQEMLDw3H06FFe63ru92g1\neQEQf4TJ3uFarVbGCkYuF22CyWTqICY7duxAUVERdu7cqcl7qmV8TlDENsVie3IZjUbo9XqHmVx8\nK38+c0KpOuyJQQ2bFzFjYmeMAWCSA7QwPjZKeZrdunULH374IXbu3AmbzYbU1FRkZmYiIiLCocBq\n2eoF8CwepoQVDFdMACA3Nxeff/458vLyHLaloPDjM4JC2v+K6WFChIdUtZM/4/PkEmNSCAinVcp1\ndsw+89ei9bzdbmd8z/R6vWa6UhLUikmI9V3Tcn0TIG1yhStedGJhv7vk3SgoKMC2bduQn58v2JKC\n4hgqKBy4mVyAY08udyZrdnaUs5x9d5CiQlpO+Fb+fPdEjfN0QDvHSOx7QgwKySRKjjC1eL5PxESO\nXafQPXHFCoZPTHbv3o0PP/wQBQUForPqKB3xOUFx1BSLZHIZDAYmBZRPTEgAUYrJmrwgZAVGXhB3\nJ1ItW88D4tyMpcgY83R8Wlv5k3tCjjABMM+J3PfEFeQUEy7knvA1mBNKiCGZhOyU/r1792LDhg0o\nKCiA0WiUdcydHZ8TFKGmWI4yudgGj3JO1p5OpFq2ngfcP6aRsysl3/i0aAcCtD9GIpmE7BiD0tY4\nXEg8Ua14mLNWv3xicvDgQbz99tsoLCyUrPGeL+MzgsJO2SS1BIB7mVxKHYNwe4M7WpF6y5m6p5O1\nlF0p+canxYJKwPH4hKxxlEr+ANQXEy7cuAv5sy5dujD3paysDK+99hoKCws97pHkrKcJAGRlZaGk\npAQGgwFbtmxBTEyMR9fUIj4nKOxKZxIYttvtzKrF1UwupeCKC5lIAwICmCpkrU6GcqUtS+UxpnZa\ntTNcmaxd8dRSY3xqYDabYTKZEBAQgFOnTmHevHm47777cO7cOezfv99t23k2hw8fRmhoKB5++GFe\nQSkuLsaGDRtQXFyMiooKLF26FOXl5R5fV2toK/VHAUgcpLW1FXV1ddDpdDAajYzNvCNPLjWPQYi9\nR2hoKIxGIyMkdXV1aG5uRmBgoOYyuYDfX2Y5PM2k8BiTc3xS4Opk7cxTq7m5mTlOVWN8SmOxWJhj\nLoPBgNGjR+O//uu/cOPGDfTo0QNDhgxBamoqjh8/7tF14uPjHe5yioqKsGDBAgBAXFwcbt++jZqa\nGo+uqUW09wbJDImF1NXVISgoiMn2EsrkstlsmvPkIvUsRPzIRFpfX6+JQkqgfQ2MEvfPVY8xpcfn\nDp5a0XA9tVxxLxCDN4gJd3wnTpzAu+++i/z8fERERODWrVsoLi7uEFOVmitXriAqKor57z59+uDy\n5cuIiIiQ9bpK4zOCws7fJ1XZYjK5PG03Kwds63myuyJ/zm4IpXQhJXt8arQ6JgiZE7ItT4joaFVM\nSNGdlONz1MvE1cJBrYsJOQZmj+/kyZN4/vnnsWvXLmYi79atG+bNm6fImLi7Qq3NK1LgM4ICtL2k\nZrOZeanUyOTyFHYHQ27asrNVuhJFg+76XskFn8dYU1MTYy1vMplU70rJhs+oUA4cdabU6/UODRu1\nbOIJ8Kcuf/fdd1i2bBl27dqFXr16KT6myMhIXLp0ifnvy5cvIzIyUvFxyI32lmYywV5RAeiQyaXT\n6WC1WtHQ0MAUNWphgmHDFjtn4yOrdPZZOtl5NTQ0oLm5mSkMk3p8UtXoSA2Jien1eoSFhcFoNMLf\n35+JRbnSx0TO8cktJlzIYsNgMMBoNDJu2o2NjUzchTwr7NRbbxGTs2fPYunSpdi5c6dqk3hKSgq2\nbdsGACgvL0fXrl073XEX4ENZXq2trcxkUV9fz6yeyaQntxuvp0iVtsxX6+JJISVB6zs7Z+4BUmWM\neTI+bnMnteE+K0RoSWq6FsbIhi+1+ty5c1iyZAk+//xz9O/fX7Zrs3uaREREdOhpAgBPPvkkSktL\nERISgs2bNyM2Nla28aiFzwgKWV2R4DU7vkAClVrfwsshdmy7E3cr0rViVSKEq47BcnhHObueq71W\nlIZkwwUFBTFFwloopiTwicn58+eRkZGB7OxsDBw4UNXx+Qo+Iyitra1MxzigbRK0WCxM0ROxnlf7\nxeCiZI2EK4WUBK0XVHq6c5LbY4yICXFv0KqYcL2vyI5OqCpdSfh61F+8eBGLFi3C9u3bMWjQIEXH\n48toa/aUkS+++AIpKSn46KOP8Ouvv6KxsRHLli1DfX09goODGYfhhoYGmM1m1c7RCeQIhOyclKiR\n4Na68PVJZ68/SH/w4OBgTYsJaXDlzmTNjUWRSZ/bK92ddRk7W0+rYmIymTqICfB7DVBISAhTA0Tu\nN6kBkjpGxwefmFRXV2PRokXYunUrFROF8Zkdit1ux40bN5Cfn4/s7GxUVVVh1KhReO2119C3b18m\nXZjbv0SoH7jcY9WS9Tyf3YlOp2OCx1osCFTiGM4TjzEiJsSoVKti4mqCgJDZqStuwGLhE5NLly5h\n/vz5+PjjjzF06FDJrkURh88ICuGbb75BSkoKHn/8cURGRiI/Px/19fWYOnUqUlNT24kLn8W83OLi\nDdbzZKIBoJlCSjZqHMO54jEmVxdIKZEq20wuY08+I8+rV69i7ty5+OCDDzB8+HC3v5viPj4lKHa7\nHcnJyVi8eDFmzJjB/HltbS2++OIL5OXl4caNG5gyZQpSU1Nx1113ORUXKYO03pApxW7XS1Kt1d7R\nsdGCY7CjjDEAaGpqYupitPg7lqsOhk903enCyPc7vnbtGubMmYP33nsPI0aMkGzMFNfwKUEBwBQx\nClFfX489e/YgLy8PV69excSJE5GWloZBgwY5FBdPzfe0bj3vLBNJKdF1hBYdg7nHqHa7nRETpRuH\nOUOpokpyLaHFiKPnhbwnbDH59ddfkZ6ejnfeeQd//vOfZRszxTk+Jyiu0NjYiJKSEuTl5aG6uhrj\nxo3DjBkzMGTIEOj1eslqOrSeKeXqMZxQVz05uy9q3QqEJH2w409qdqXkoqSY8F1bTCYdn5jcuHED\n6enpeOuttzB69GjFxkzhhwqKSJqbm7Fv3z7k5eXhxx9/xJgxYzBjxgzcc889HomL1q3T2VYv7pz3\ny1VIyUbrViDkKJPsPgF1u1Jy4R5lqm0qyndf/Pz8OrQ9vnnzJmbPno21a9di7Nixqo2Z8jtUUNzA\nYrHg4MGDyM3NxenTpzF69GikpaVhxIgRzMvILRjkTqJ2u52x1tb6RBgQECDZMRx759La2urRJKrm\nqlosYrPNlOpKyUVLYsIHaXtssVgAABcuXMDp06cxZswYLFmyBKtXr8bEiRMluVZpaSmefvpp2Gw2\nPProo3juuefa/f2hQ4eQmpqKAQMGAABmzpyJF198UZJrdxaooHhIS0sLysrKkJOTg1OnTmHUqFFI\nS0tDXFwcIxLsyYJMolp3u1Ui7ZavkNKVtFstT4SA+/3p5epKyUWLdi9c2PfQ398fX3/9Nd566y18\n+eWXGDhwIBYuXNhukvfkOoMGDcKBAwcQGRmJe++9F9nZ2Rg8eDDzmUOHDmH9+vUoKiry9MfqtGjv\nLfQyAgICMGnSJLz//vs4duwYZs2ahYKCAkyYMAHLli3DV199BbvdzhQMkt7WJLZA0jO1pOtWq5V5\nieW0UuEWUoo1atRyrxqCu2IC/O4ETIoGAwICYLVaUV9fL1nhrbeJCXGxGDx4MJqamrB9+3asXbsW\nZ8+exV/+8hccOHDAo2tVVlZi4MCB6NevHwICApCeno7CwsIOn9PSe6pFtHdo78X4+/tj3LhxGDdu\nHGw2G44fP47c3Fy89NJLuOeeezB58mS8/fbbmDVrFp544gkmvZTd8EiNM3Q2amVKce3Uyc6lubm5\nXdqtTqfTlD0+H1KmLpOKdD6beXcz6bxRTIC2JJm5c+di6dKlSEtLAwAkJSU57copBr4GWBUVFe0+\no9PpcOzYMQwbNgyRkZFYt24dhgwZ4tF1OxtUUGTCz88P9913H+677z60traisLAQjz76KAYPHowf\nfvgB+/btw7hx45gjJXL8Y7FY0NTU1K5nvFIvvFYypYQmUZPJBAC8vWC0gpx1MK52peTDG4woSeyO\nLSZNTU2YN28elixZwogJQYpnVcx9iI2NxaVLl2AwGFBSUoK0tDScP3/e42t3JrR3VtAJOXXqFJ54\n4gmsWbMGX331FZYuXYoTJ04gISEBGRkZ2L17N8xmM4KCghASEtKhZzyfj5aUcH3DtJQgQCbRLl26\nMLUKfn5+7TyjpFihSgHxNmNnIsmFOx5j3iImDQ0NjFkr0JZhOX/+fCxevBizZs2S5brcBliXLl1C\nnz592n2G3GcASEhIQEtLC27evCnLeLwVGpRXgLNnz+LChQtITU1t9+d2ux1nzpxBTk7Dwr5tAAAY\nl0lEQVQODhw4gD59+iAtLQ1TpkxhHlyuq6vUAVpvCG7zZZtpoZCSjZaKKoUyxojAaFlMuOnVZrMZ\n8+fPR3p6Oh566CHZrm21WjFo0CAcPHgQvXv3xqhRozoE5WtqanDnnXdCp9OhsrISDz74IKqrq2Ub\nkzdCBUUj2O12VFVVITc3F3v37sWdd96J1NRUTJs2DUajkfkMV1zcsa5gX5O43Wp1khGTbSZkSKhU\nwaBWjgr5IEepZrO5Q62LlhYPfGJisVjwyCOPIDU1FY888ojsv8eSkhImbXjx4sVYuXIl3n//fQBt\nTbI2btyITZs2wd/fHwaDAevXr6eV+RyooGgQu92OCxcuIC8vD8XFxejWrRuSk5Mxffp0dO3alfkM\nmUDd6TBIDAq12q4XcC9TSqiQUg63W0DbYgJ0dK62Wq2qdaUUgk9MWlpasGjRIkydOhUZGRmafD4p\nHaGConHsdjuqq6uRl5eHPXv2wGAwIDk5GUlJSejWrZug7b6jiULrJpSAdHY0fDVAUmXSab1C31Hz\nLqW7UgrB7VkDtP3uMzIyMGbMGPztb3/T5PNJ4YcKihdht9tx+fJl7Nq1C0VFRfD390dycjKSk5PR\no0cPUT1dbDYbk6KsRRNKQL6Wx9xCSk8y6YiYaLUOxpVOkHJ3pRSCT0xsNhsef/xxjBo1CllZWZp8\nPinCUEHxUux2O2pqarBr1y4UFhbCZrMhKSkJKSkpiIiIEAxct7a2IigoiHmBtYZSwW13q9G9we7F\nk546SnmM8SVa2Gw2PPnkk/jTn/6E5cuXUzHxQjqdoOTk5GD16tWoqqrCiRMnEBsby/u5fv36ISws\njDlHrqysVHik0sHuRllQUACTyYTp06cjJSUFkZGR0Ol0OH78OO666y6EhITAZrOpnhXFh1rxCG6y\ng1BswRsy4qRu0CaHx5jdbmecl8mRa2trK55++mkMGDAAK1eu1MTzSHGdTicoVVVV0Ov1yMzMxNtv\nvy0oKP3798c333yD7t27KzxC+bl58yYKCwuxa9cu1NXVITo6Gnl5eSgoKEBsbKzmUm4B7cQjhOJR\n/v7+sFgssNlsmmjLzIfc3T6l8BgjySDsBmOtra1Yvnw5evXqhZdeeomKiRfT6QSFMH78eKeC8vXX\nX+OOO+5QeGTK8uabb+L111/HxIkTce3aNcFulGql3Gr5CImbSQcAQUFBmtrVEeQWE77rCXWlFPod\nConJypUrERYWhldffVVT95TiOj5rvaLT6TBp0iT4+fkhMzMTGRkZag9JctavX4+PP/4YJ0+eRL9+\n/ZhulK+88gquXLmCSZMmMd0o/f3921nAkICunOLC9ZTSkpgAv1ejt7S0QK/Xo0uXLoxxplSdOqWA\niIlOp1OsR72rHmNCYvLSSy8hODgYr7zyChWTToBX7lAmT56Ma9eudfjztWvXIjk5GYDzHcovv/yC\nXr164fr165g8eTLeffddxMfHyzpupblw4QLCw8PRs2fPDn/X2NiI0tJS5Obm4n//938xfvz4dt0o\nAec9XTzBG2xAhFb9SjQNc3WMSoqJs/HwZYyR40NyH+12O9asWQOz2Yz169dLsphw1s8EALKyslBS\nUgKDwYAtW7YgJibG4+tSfscrBUUMzgSFzcsvv4zQ0FAsW7ZMgZFpD75ulGlpaRg2bFgHcZGqMZaS\nxzPuIHaManZeJKt+d7tpyg0RF+LGoNPpsG7dOsTFxeHUqVOora3Fu+++K4mYiOlnUlxcjA0bNqC4\nuBgVFRVYunQpysvLPb425Xe0dcYgMUJa2dTUhPr6egBtK/V9+/Zh6NChSg5NUwQHByM1NRXbtm3D\n4cOHMWHCBHzyySeYMGECXnjhBZw4cQI6nQ5dunRBaGhou74uQkaEQnhDhb4rY9TpdMwxjtFobHdv\n6urqXLo37oxRq2JCMJlM8Pf3R1hYGLp06YLw8HCsXbsW69evx40bN5CTk8O8i54gpp9JUVERFixY\nAACIi4vD7du3UVNT4/G1Kb/T6QQlPz8fUVFRKC8vR2JiIhISEgAAV69eRWJiIgDg2rVriI+Px/Dh\nwxEXF4ekpCRMmTJFzWFrhsDAQCQkJODjjz/G0aNHkZiYiM8++wzjx4/HihUrcOzYsXYNw1yZQImT\nrJ+fn2YnQU8nam7TMPa9kco12hvEhOzw2GMku7bhw4fj4sWLmDhxIrZu3YoHHnjA4+vx9TO5cuWK\n089cvnzZ42tTfqfTBeVnzJiBGTNmdPjz3r17Y8+ePQCAAQMG4Ntvv1V6aF4H6UY5adIkWK1WHDly\nBLm5uVi5ciVGjhyJ1NRU/PWvf+3Q04U0DGNXopNJUMsV+lJb0hBxIfeGBK6bm5vdNmn0JjFhx3Xs\ndjs2bdqEH374AVu2bIGfnx8ee+wxPPbYYx53nwTE9TMhY3Pn31HE0el2KGqQk5ODP/7xj/Dz88PJ\nkycFP1daWoro6GjcfffdeOONNxQcoeeQbpQbNmxAeXk55s2bh9LSUkycOBFZWVn45z//CZvN1m51\nzu7pUl9fDz8/P82LCbEBkXqMUrT19SYxAdBOTD766COcPHkSmzdv7lBnJEUMRUw/E+5nLl++jMjI\nSI+vTfkdKigSMHToUOTn52PMmDGCnyG2EqWlpfjhhx+QnZ2Nc+fOKThK6SDdKN955x1UVFQgIyMD\nZWVlmDx5MpYsWYK9e/eipaUFgYGBqKqqwrlz55hdirN+8Wogt5hwISm3BoMBYWFhCAoKgs1mQ0ND\nAxoaGphUajbstFtvEBN2NteWLVtw9OhRbN26VTY7nZEjR+Knn35CdXU1LBYLPv/8c6SkpLT7TEpK\nCrZt2wYAKC8vR9euXRERESHLeHyVTnfkpQbR0dFOP8MOGgJggobsLBRvRK/XIy4uDnFxcWhtbcV3\n332HnJwcvPHGG7jjjjtQUVHRrm8Et1+8pz1dPEVMvxU5cdbWlxyLkXulVXdokgYOtBeTTz/9FAcP\nHsSOHTtk7WLp7++PDRs2YOrUqUw/k8GDB7frZzJ9+nQUFxdj4MCBCAkJwebNm2Ubj69CBUUh+AKC\nFRUVKo5IevR6PWJiYhATE4M9e/Zg/vz5uP/++/E///M/2L17N9LS0jB58mSEhITw9otXuj+H2mLC\nhdRsEOFgiwvBZrOpXkjJRcjZeMeOHdi9ezdycnI8akEgloSEBCYJh5CZmdnuvzds2CD7OHwZKigi\nEVNM6QgtTQByc/ToUSxevBglJSWIi4tr141y48aNiIiIaNeNkqzO2dXWznq6eIo7zbuURKfTQa/X\nw2q1IiAgAIGBgbBarYo4GLiCkJjk5uZi165dyMvL04RYU5SBCopI9u/f79G/FxM07CyMGjUKR48e\nxV133QWgbXIcPHgw/v73v+PFF19kulE+8MADHbpROhIXYnPiKaR5l9T9VqSEL+OMu3MxmUySNw1z\nBSG3g4KCAnz22WfIz8/XbJsEijx02kp5NRg/fjzWrVuHESNGdPg7q9WKQYMG4eDBg+jduzdGjRrV\noZLX1xDbjVLIGdkdcfEmMRGTYs1tGkbujRJV+nxisnv3bnz44YcoKChASEiIbNenaBMqKBKQn5+P\nrKws3LhxA+Hh4YiJiUFJSQmuXr2KjIwMpv6lpKSE8RpavHgxVq5cqfLItYPYbpSe2O4r1bzLE1wR\nE75/S+xx2PbynvQu4YNr6km+e+/evXj33XdRWFgIo9Eo2fUo3gMVFIrm4OtGmZiYiNTUVIfdKB2J\nS2cXEy7cpmFSZdMJicnBgwfx9ttvo7CwEOHh4W5/P8W7oYJC0TRiulE66+lC4g3eIiZSxx24fV3c\nzaYTEpOysjKsXbsWRUVF6Natm6Rjp3gXVFC8mJs3b2L27Nn417/+hX79+mHnzp3o2rVrh891pnbH\n3G6U06ZNQ2pqKvr27cuICzuuQIonScxEi9l2cooJF76OlGTn4igmxW5/HBoaytzHI0eOYM2aNSgs\nLOz0zeoozqGC4sWsWLECPXr0wIoVK/DGG2/g1q1beP311zt8rrO2O66trcUXX3yBvLw8XL9+nelG\nOXDgQOh0OuTn52P06NEwGo2wWq2aSrclKCkmXMQeGwp11Tx+/Dj+/ve/o7CwkLfnDsX3oILixURH\nR6OsrAwRERG4du0axo0bh6qqqg6f84V2x6QbZV5eHlNEevz4cZSWljLuBFL2dJECIiaBgYGq12qw\nxYUrvuTP2GJy4sQJPP/88ygoKJDcvsQXd96dBSooXky3bt1w69YtAG0TQvfu3Zn/ZjNgwACEh4d3\n6nbHbF566SV89NFHGD9+PC5evMjbjZJ7LKa0uGhJTLiwjw2J3X5AQADOnTuHoUOH4uzZs1i2bBny\n8/PRq1cvya/v6ztvb0abEUoKg1CF/j/+8Y92/63T6QQnwqNHj7ZrdxwdHd3p2h0T3nrrLezatQsn\nT57EH/7wB6Yb5YYNGzp0oxSy3Ze7loP0hdGK5QsX0jSMHIMZDAa0tLTg2WefxY8//gij0YjVq1fL\nFoAvKipCWVkZAGDBggUYN24cr6AAwk30KOpAdyheTHR0NA4dOoQ//OEP+OWXXzB+/HjeIy82nb3d\n8YULF9C1a1f06NGjw99ZLBYcPHgQubm5OH36NEaPHo3U1FSMHDmSd+dis9kkr+XQupgQzGYzLBZL\nu2Ous2fPYsWKFRg7diwOHTqEb7/9FvPmzcPGjRslvTbdeXsvdIfixaSkpGDr1q147rnnsHXrVqSl\npXX4TFNTE2w2G4xGI9PueNWqVSqMVhkGDhwo+HekG2VCQgJaWlpQVlaG7OxsPPvss4iLi0NaWhri\n4uKcNsVyV1y0ZkYpBJ+YnDt3Dn/729/w+eefY8CAAQCAX3/91ekCRgi68+6c0B2KF3Pz5k08+OCD\n+Pe//90ueMmu0P/5559x//33A2izHZk3bx6t0OfA7kZZUVGBESNGIC0tDX/961+ZuhW+QkFXOi5q\n3YySYDabYTabERoayvxc58+fR0ZGBj777DPcfffdso+B7ry9FyooFAoLm82G48ePIzc3F0ePHsU9\n99yDtLQ0jBkzhvH+crUK3ZvF5OLFi1i0aBG2b9+OQYMGKTKOFStW4I477sBzzz2H119/Hbdv3+4Q\nQ+HuvKdMmYJVq1ZhypQpioyRwg8VFApFgNbWVpw4cQK5ubkoKyvD4MGDkZaWhnHjxjFHVs6q0L1F\nTCwWC0wmE0JCQpgCx+rqaixYsABbt27FkCFDFBsL3Xl7L1RQKKIpLS1lzC0fffRRPPfccx0+k5WV\nhZKSEhgMBmzZsgUxMTEqjFR62N0o//nPf2LAgAFIS0vDxIkTERwcDKCjuOj1erS2tiIoKEjTNu58\nYnL58mXMmzcPn3zyCYYOHaryCCneAhUUiihsNhsGDRqEAwcOIDIyEvfee28H+/3i4mJs2LABxcXF\nqKiowNKlS1FeXq7iqOXBbrfjzJkzyMnJwYEDB9CnTx+kpqZiypQpjGX7+fPncccddzDBfal7ukgF\nn5hcvXoVc+fOxfvvv99pFgQUZVC+iTfFK6msrMTAgQPRr18/BAQEID09HYWFhe0+U1RUhAULFgAA\n4uLicPv2bdTU1KgxXFnR6XQYOnQo1qxZg6NHj+Lll1/Gzz//jLS0NDz00EPYuHEjpk6diu+//x6h\noaEwGo3o0qULU8xYX1/PmCyqCZ+YXLt2DfPmzcN7771HxYTiMlRQKKIgdiaEPn364MqVK04/c/ny\nZcXGqAbsbpRHjhzBggULsGbNGgwfPhzvv/8+PvvsM9TW1sLf3x/BwcEwGo0IDg6G3W7vIC5KHhYQ\nB2a2mPz666+YO3cu3nnnHYwcOVKxsVA6D7QOhSIKsXUX3ElRCwaMSnHq1ClkZmZiy5YtuP/++5lu\nlPPmzWO6USYmJqJ79+4d2vk2Nja63DDMXUhdDVtMbty4gblz52LdunX4y1/+Ist1KZ0fKigUUURG\nRuLSpUvMf1+6dAl9+vRx+JnLly8jMjJSsTGqTVBQED744AOkpKQAaPOaWr58OZYtW8Z0o3zkkUeY\nbpRJSUno2bNnB3FpamqSzRmZT0xu3ryJuXPnYu3atbjvvvskuQ7FN6FBeYoorFYrBg0ahIMHD6J3\n794YNWqUw6B8eXk5nn766U4ZlPcEdjfKgoIC2Gw2JCUldehGybaAkUpc+LpW3r59G7Nnz8aqVasw\nadIkKX9Uig9CBYUimpKSEiZtePHixVi5ciXef/99AEBmZiYA4Mknn0RpaSlCQkKwefNmxMbGqjlk\nTcPXjTIhIQEpKSno06cPIxzsniXuigufmNTV1SE9PR3PP/88pk2bJtvPSfEdqKBQKBrBWTdKwL2e\nLlarFU1NTe3EpKGhAenp6fjP//xPJCUlKfYzUjo3VFAoFA3irBslIK6nC5+YNDY2Ys6cOXjiiScw\nY8YM1X5GSueDCgqFonG43SgnTpyItLQ0REdH84qLzWZjMsXMZjNCQkIYMWlqasLcuXORkZGBBx54\nQPKx5uTkYPXq1aiqqsKJEycEjzzFuC5QvA9ah0LxCkpLSxEdHY27774bb7zxRoe/P3ToEMLDwxET\nE4OYmBi8+uqrKoxSHoxGI9LT05GTk4P9+/dj+PDhWLduHSZOnIhXXnkFZ86cAdCWZUYKKYE2s0eg\nrY/Jjh07cP36dcyfPx8LFy6URUwAYOjQocjPz8eYMWMEP2Oz2ZhY2w8//IDs7GycO3dOlvFQlIWm\nDVM0D5mA2LYvKSkp7TLMAGDs2LEoKipSaZTKEBISgpkzZ2LmzJmC3SjNZjNWrFiBvXv3IigoCE1N\nTdi+fTueeuopDB48GDabDbW1tQgPD5d8fNHR0U4/w3ZdAMC4LnB/nxTvg+5QKJpHjO0L4HvtYIOD\ng5Gamopt27bh8OHDmDBhAt58802mC+Xp06cBACNGjEBYWBhef/11PPXUU9i5cyeioqKQnZ2tyrjF\nuC5QvBO6Q6FoHr4JqKKiot1ndDodjh07hmHDhiEyMhLr1q1T1HJdbQIDA9GrVy9UVFRgy5YtCA8P\nR3Z2NpYvX46GhgY888wzeOyxx6DT6bBgwQLU1dW57SUm1G1x7dq1SE5Odvrvfck9wdeggkLRPGIm\noNjYWFy6dAkGgwElJSVIS0vD+fPnFRiddvjHP/6BjRs3YubMmQCASZMmwWq1orS0FImJie3uY1hY\nmNvX2b9/v0fjFOO6QPFO6JEXRfOImYCMRiMMBgMAMD3jb968qeg41Wbnzp2MmBD8/f2RlJSkyq5A\n6Ahy5MiR+Omnn1BdXQ2LxYLPP/+csauheDdUUCiaR8wEVFNTw0xglZWVsNvt6N69uxrDVQ0tHCXl\n5+cjKioK5eXlSExMREJCAoC2HiuJiYkA2kRuw4YNmDp1KoYMGYLZs2fTgHwngdahULwCZ7YvGzdu\nxKZNm+Dv7w+DwYD169fjz3/+s8qjplB8CyooFAqFQpEEeuRFoVAoFEmggkKhOGHRokWIiIjA0KFD\nBT+TlZWFu+++G8OGDcOpU6cUHB2Foh2ooFAoTli4cCFKS0sF/764uBgXLlzATz/9hA8++ABLlixR\ncHQUinaggkKhOCE+Ph7dunUT/PuioiIsWLAAABAXF4fbt2+jpqZGqeFRKJqBCgqF4iF8lfyXL19W\ncUQUijpQQaFQJICbLKmFmhAKRWmooFAoHsKt5L98+TIiIyNVHBGFog5UUCgUD0lJScG2bdsAAOXl\n5ejatSsiIiJUHhWFojzUHJJCccKcOXNQVlaGGzduICoqCi+//DJaWloAtFXpT58+HcXFxRg4cCBC\nQkKwefNmlUfsHmK7Lfbr1w9hYWHw8/NDQEAAKisrFR4pRavQSnkKhQIAqKqqgl6vR2ZmJt5++21B\nQenfvz+++eYbn/NKoziH7lAoFAoAcd0WCXQdSuGDxlAoFA3jrEr/0KFDCA8PR0xMDGJiYvDqq6/K\nPiadTodJkyZh5MiR+PDDD2W/HsV7oDsUCkXDLFy4EE899RQefvhhwc+MHTsWRUVFor7P026LAHD0\n6FH06tUL169fx+TJkxEdHY34+HhR/5bSuaGCQqFomPj4eFRXVzv8jCvHT552WwSAXr16AQB69uyJ\nGTNmoLKykgoKBQA98qJQvBqdTodjx45h2LBhmD59On744QdJvldIpJqamlBfXw8AaGxsxL59+xya\nZlJ8CyooFIoXExsbi0uXLuG7777DU089hbS0NLe/S0y3xWvXriE+Ph7Dhw9HXFwckpKSMGXKFEl+\nFor3Q9OGKRSNU11djeTkZJw+fdrpZ2lKL0VN6A6FQvFiampqmOOpyspK2O12KiYU1aBBeQpFwzir\n0s/NzcWmTZvg7+8Pg8GAHTt2qDxiii9Dj7woFAqFIgn0yItCoVAokkAFhUKhUCiSQAWFQqFQKJJA\nBYVCoVAokkAFhUKhUCiSQAWFQqFQKJLw/wG01aU/LOMn1wAAAABJRU5ErkJggg==\n", + "text/plain": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Sample data and `timeit` benchmarks" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#Sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "In the section below, we will create a random dataset from a bivariate Gaussian distribution with a mean vector centered at the origin and a identity matrix as covariance matrix. " + ] + }, + { + "cell_type": "code", + "execution_count": 13, + "metadata": {}, + "outputs": [], + "source": [ + "import numpy as np\n", + "\n", + "np.random.seed(123)\n", + "\n", + "# Generate random 2D-patterns\n", + "mu_vec = np.array([0,0])\n", + "cov_mat = np.array([[1,0],[0,1]])\n", + "x_2Dgauss = np.random.multivariate_normal(mu_vec, cov_mat, 10000)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "The expected probability of a point at the center of the distribution is ~ 0.15915 as we can see below. \n", + "And our goal is here to use the Parzen-window approach to predict this density based on the sample data set that we have created above. \n", + "\n", + "\n", + "In order to make a \"good\" prediction via the Parzen-window technique, it is - among other things - crucial to select an appropriate window with. Here, we will use multiple processes to predict the density at the center of the bivariate Gaussian distribution using different window widths." + ] + }, + { + "cell_type": "code", + "execution_count": 14, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "actual probability density: 0.159154943092\n" + ] + } + ], + "source": [ + "from scipy.stats import multivariate_normal\n", + "var = multivariate_normal(mean=[0,0], cov=[[1,0],[0,1]])\n", + "print('actual probability density:', var.pdf([0,0]))" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Benchmarking functions" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#Sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Below, we will set up benchmarking functions for our serial and multiprocessing approach that we can pass to our `timeit` benchmark function. \n", + "We will be using the `Pool.apply_async` function to take advantage of firing up processes simultaneously: Here, we don't care about the order in which the results for the different window widths are computed, we just need to associate each result with the input window width. \n", + "Thus we add a little tweak to our Parzen-density-estimation function by returning a tuple of 2 values: window width and the estimated density, which will allow us to sort our list of results later." + ] + }, + { + "cell_type": "code", + "execution_count": 15, + "metadata": {}, + "outputs": [], + "source": [ + "def parzen_estimation(x_samples, point_x, h):\n", + " k_n = 0\n", + " for row in x_samples:\n", + " x_i = (point_x - row[:,np.newaxis]) / (h)\n", + " for row in x_i:\n", + " if np.abs(row) > (1/2):\n", + " break\n", + " else: # \"completion-else\"*\n", + " k_n += 1\n", + " return (h, (k_n / len(x_samples)) / (h**point_x.shape[1]))" + ] + }, + { + "cell_type": "code", + "execution_count": 16, + "metadata": {}, + "outputs": [], + "source": [ + "def serial(samples, x, widths):\n", + " return [parzen_estimation(samples, x, w) for w in widths]\n", + "\n", + "def multiprocess(processes, samples, x, widths):\n", + " pool = mp.Pool(processes=processes)\n", + " results = [pool.apply_async(parzen_estimation, args=(samples, x, w)) for w in widths]\n", + " results = [p.get() for p in results]\n", + " results.sort() # to sort the results by input window width\n", + " return results" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Just to get an idea what the results would look like (i.e., the predicted densities for different window widths):" + ] + }, + { + "cell_type": "code", + "execution_count": 17, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "h = 0.1, p(x) = 0.016\n", + "h = 0.2, p(x) = 0.0305\n", + "h = 0.3, p(x) = 0.045\n", + "h = 0.4, p(x) = 0.06175\n", + "h = 0.5, p(x) = 0.078\n", + "h = 0.6, p(x) = 0.0911666666667\n", + "h = 0.7, p(x) = 0.106\n", + "h = 0.8, p(x) = 0.117375\n", + "h = 0.9, p(x) = 0.132666666667\n", + "h = 1.0, p(x) = 0.1445\n", + "h = 1.1, p(x) = 0.157090909091\n", + "h = 1.2, p(x) = 0.1685\n" + ] + } + ], + "source": [ + "widths = np.arange(0.1, 1.3, 0.1)\n", + "point_x = np.array([[0],[0]])\n", + "results = []\n", + "\n", + "results = multiprocess(4, x_2Dgauss, point_x, widths)\n", + "\n", + "for r in results:\n", + " print('h = %s, p(x) = %s' %(r[0], r[1]))" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Based on the results, we can say that the best window-width would be h=1.1, since the estimated result is close to the actual result ~0.15915. \n", + "Thus, for the benchmark, let us create 100 evenly spaced window width in the range of 1.0 to 1.2." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "code", + "execution_count": 18, + "metadata": {}, + "outputs": [], + "source": [ + "widths = np.linspace(1.0, 1.2, 100)" + ] + }, + { + "cell_type": "code", + "execution_count": 19, + "metadata": {}, + "outputs": [], + "source": [ + "import timeit\n", + "\n", + "mu_vec = np.array([0,0])\n", + "cov_mat = np.array([[1,0],[0,1]])\n", + "n = 10000\n", + "\n", + "x_2Dgauss = np.random.multivariate_normal(mu_vec, cov_mat, n)\n", + "\n", + "benchmarks = []\n", + "\n", + "benchmarks.append(timeit.Timer('serial(x_2Dgauss, point_x, widths)', \n", + " 'from __main__ import serial, x_2Dgauss, point_x, widths').timeit(number=1))\n", + "\n", + "benchmarks.append(timeit.Timer('multiprocess(2, x_2Dgauss, point_x, widths)', \n", + " 'from __main__ import multiprocess, x_2Dgauss, point_x, widths').timeit(number=1))\n", + "\n", + "benchmarks.append(timeit.Timer('multiprocess(3, x_2Dgauss, point_x, widths)', \n", + " 'from __main__ import multiprocess, x_2Dgauss, point_x, widths').timeit(number=1))\n", + "\n", + "benchmarks.append(timeit.Timer('multiprocess(4, x_2Dgauss, point_x, widths)', \n", + " 'from __main__ import multiprocess, x_2Dgauss, point_x, widths').timeit(number=1))\n", + "\n", + "benchmarks.append(timeit.Timer('multiprocess(6, x_2Dgauss, point_x, widths)', \n", + " 'from __main__ import multiprocess, x_2Dgauss, point_x, widths').timeit(number=1))" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Preparing the plotting of the results" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#Sections)]" + ] + }, + { + "cell_type": "code", + "execution_count": 20, + "metadata": {}, + "outputs": [], + "source": [ + "import platform\n", + "\n", + "def print_sysinfo():\n", + " \n", + " print('\\nPython version :', platform.python_version())\n", + " print('compiler :', platform.python_compiler())\n", + " \n", + " print('\\nsystem :', platform.system())\n", + " print('release :', platform.release())\n", + " print('machine :', platform.machine())\n", + " print('processor :', platform.processor())\n", + " print('CPU count :', mp.cpu_count())\n", + " print('interpreter:', platform.architecture()[0])\n", + " print('\\n\\n')" + ] + }, + { + "cell_type": "code", + "execution_count": 25, + "metadata": {}, + "outputs": [], + "source": [ + "from matplotlib import pyplot as plt\n", + "import numpy as np\n", + "\n", + "def plot_results():\n", + " bar_labels = ['serial', '2', '3', '4', '6']\n", + "\n", + " fig = plt.figure(figsize=(10,8))\n", + "\n", + " # plot bars\n", + " y_pos = np.arange(len(benchmarks))\n", + " plt.yticks(y_pos, bar_labels, fontsize=16)\n", + " bars = plt.barh(y_pos, benchmarks,\n", + " align='center', alpha=0.4, color='g')\n", + "\n", + " # annotation and labels\n", + " \n", + " for ba,be in zip(bars, benchmarks):\n", + " plt.text(ba.get_width() + 2, ba.get_y() + ba.get_height()/2,\n", + " '{0:.2%}'.format(benchmarks[0]/be), \n", + " ha='center', va='bottom', fontsize=12)\n", + " \n", + " plt.xlabel('time in seconds for n=%s' %n, fontsize=14)\n", + " plt.ylabel('number of processes', fontsize=14)\n", + " t = plt.title('Serial vs. Multiprocessing via Parzen-window estimation', fontsize=18)\n", + " plt.ylim([-1,len(benchmarks)+0.5])\n", + " plt.xlim([0,max(benchmarks)*1.1])\n", + " plt.vlines(benchmarks[0], -1, len(benchmarks)+0.5, linestyles='dashed')\n", + " plt.grid()\n", + "\n", + " plt.show()" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "# Results" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#Sections)]" + ] + }, + { + "cell_type": "code", + "execution_count": 26, + "metadata": {}, + "outputs": [ + { + "data": { + "image/png": "iVBORw0KGgoAAAANSUhEUgAAAowAAAIACAYAAAAIQT11AAAABHNCSVQICAgIfAhkiAAAAAlwSFlz\nAAALEgAACxIB0t1+/AAAIABJREFUeJzs3Xl8TPf6B/DPmZBVElmERGWzFQlFaFGyqL3V2kKqIfYt\nqm7Rlt4mVAVxKbXUls3FrSD2NRJL0LpibWuXxBZLbElIJJHz+8NvzjVmJjPImJP4vF8vr3bOnDnn\nmfPM8uScZ75fQRRFEUREREREWiiMHQARERERyRsLRiIiIiIqEQtGIiIiIioRC0YiIiIiKhELRiIi\nIiIqEQtGIiIiIioRC0YCAOzduxcKhQKxsbGvvA2FQoEBAwaUYlRln5+fHzw8PPRePyQkBAoF35a6\npKenQ6FQYPLkycYORW/u7u7w9/c3dhikhaE+v8ria7U0lYXnHxMTA4VCgX379hk7FFnjN5MMXb58\nGUOHDsW7774LKysr2Nvbo379+ggJCcHevXsNtl9BECAIwmtvQ+6URZlCoUBqaqrGdebMmSOt8zpF\nNKB+TGJiYjB37lyt65aFYygXZelYGSq37u7u0mtVoVDAzMwMHh4eGDJkCK5du1bq+yvPDPl6Kkuv\n1ZeVnp6O8PBwnDx5Uus6xn7+e/fuxeTJk/Hw4UO1+5TvTWPHKHcVjB0AqTp69Ch8fX1hZmaGfv36\noUGDBsjLy8P58+exa9cu2NjYwM/Pr9T36+vri7y8PFSo8Pa8JMzNzREdHY2mTZuq3RcdHQ1zc3Pk\n5+eX+odITEwMMjIyMGbMGLX7li5disWLF5fq/sojd3d35Ofnw8TExNih6O38+fMG+0KqUaMGIiIi\nAAA5OTlITk5GVFQUtm3bhlOnTsHBwcEg+y1PytrrSU7S09MxZcoUeHp6olGjRir3yeW9unfvXkyZ\nMgUDBgyAra2tyn3BwcEICgpCxYoVjRRd2fD2VAdlxOTJk5Gfn4/ff/8d3t7eavffunWrVPeXk5MD\na2trCIIAU1PTUt223HXr1g2rV6/G7NmzVZ77f//7X/z555/4/PPPsWrVKoPsW1vhYKiCPTc3F5Uq\nVTLIto2lrL1eDfllZGtri88//1y6PWzYMDg5OWH+/PmIjo7GuHHjXnsfys+K8qqsvZ7kSNvEcXI6\ntppiVCgUsopRrnhJWmYuXLgABwcHjcUiAFStWlVtWWJiItq3bw87OztYWFigUaNGGs9SKXuojh8/\njg4dOqBy5crSX4OaehhFUcRPP/2ENm3awNnZGWZmZnBzc8PIkSNx7969V3p+Z86cgUKhwNdff63x\n/qCgIJiZmeHu3bsAgKtXr2LgwIFwc3ODubk5qlatilatWiEuLu6V9v+8AQMG4P79+9iwYYPK8ujo\naDg5OeHjjz9We4yy12X//v1q9+nTr+ju7o79+/dLfT3Kf8rtaephVC7LyspCv3794OjoiEqVKuGj\njz7C8ePHVdZ9vl/ot99+Q9OmTWFpaYnRo0dL6yxbtgxNmjSBpaUlKleujA4dOuDgwYMa401OTkaX\nLl3g4OAACwsL1KxZE4MHD5byo/Tbb7/hww8/hI2NDaysrPDBBx9g3bp1atvbunUrfH19UaVKFVha\nWsLNzQ09evTAhQsXpHX0ybmmvqjnl23ZsgXNmjWDhYUFXFxcMGHCBDx9+lQtnnXr1qFRo0awsLCA\nm5sbpkyZgsTERL1aEb755hsoFAqcPn1a7b6HDx/CwsIC3bp1k5Zp6mHctWsXevfuDU9PT1haWsLO\nzg4dOnTQ+Pp6We3btwcAXLp06aX3pXwtp6WloWfPnrC3t5fOyrx4Cfz5fy8+v6NHj6Jbt26oUqUK\nzM3N8e6772LatGlquVDuLzMzE0FBQbC3t4eVlRU6duyo8tooyYABA2BhYYEnT55Iyw4fPgyFQgEH\nBweVQmH79u1QKBSIj4+XlmnqYVQuO3z4MHx9fVGpUiU4OjpiyJAhePTokVoMKSkpaNWqFSwtLVGt\nWjWMHj0aubm5GuN99OgRvvvuO9SsWRPm5uZwdnZG//79ceXKFWmdJ0+ewMLCAiEhISqPHTZsGBQK\nBb766iuV5b1794atrS2Ki4t1Hq8LFy4gODhY+mz38PDAhAkT8PjxY5X1dL0fY2JiEBAQAOBZDl58\nLeh6r65duxbvvfceLC0tUatWLSxbtgwAkJGRgZ49e8LBwQE2NjYIDg5WO5Znz57FyJEj0aBBA+mz\nx8fHB8uXL1dZLyQkBFOmTAEAeHh4SDEql2n7XM/KysKoUaNQo0YNmJmZwdXVFaGhoWrff8rHJycn\nY9asWVJO69atWyrfVXLBM4wyU6tWLWzbtg0JCQkqXzbaLFmyBMOHD0fLli3x/fffw8rKCrt27cKI\nESNw6dIlzJw5U1pXEARcuXIFbdu2RWBgIHr16qX2Bnz+zNeTJ08wa9Ys9OzZE926dYOVlRWOHDmC\n5cuXIyUlBampqS991qRevXpo1qwZVq1ahcjISJXiKDs7Gxs3bkTnzp3h4OCAoqIitGvXDjdu3MCo\nUaNQp04dPHz4ECdPnkRKSgr69ev3Uvt+8Xk2btwY7733HqKiohAYGAjg2WWp1atXY9CgQa90RkjX\nJce5c+fiu+++Q1ZWFn7++Wdpeb169XRuo2PHjnBwcMDkyZORmZmJ+fPnw9fXF4cPH0aDBg1U1t2w\nYQOuXLmCkSNHYuTIkbCxsQHwrMiJjIzE+++/j4iICGRnZ2PJkiXw9/fHxo0b0alTJ2kbixcvxogR\nI1CjRg2MGjUKbm5uyMjIwJYtW3D9+nXpMuf333+PadOmoVOnTpg6dSoUCgXWr1+PXr16Yf78+Rg5\nciQAYN++fejatSsaNmyIiRMnonLlyrh+/Tr27NmDS5cuoXbt2i+dc03Hatu2bVi4cCFGjBiBwYMH\nY8OGDZg1axbs7Ozw3XffSev99ttvCAoKQu3atREeHg4TExPExsZi8+bNeuUyJCQEkZGRiIuLQ2Rk\npMp9a9aswZMnT1S+6DX1SMXGxuLBgwcICQnBO++8g2vXrmHZsmVo27YtkpOT8eGHH5YYQ0mUhZaj\no+NL70sQBOTm5sLX1xcffvghIiIicPv2bQDPXsMvFku///475s+fj2rVqknLtm7diu7du6NOnToY\nN24c7O3tcejQIfzwww84ceIE1qxZo7K/R48eoU2bNmjRogUiIiJw+fJlzJ07F59++in+/PNPnT8G\na9u2LWJjY3Hw4EGpgNmzZw8UCgUePHiA48ePo0mTJgCApKQkjQWuppyfOHECn3zyCQYOHIgvvvgC\nycnJWL58ORQKhcof5n/88Qc++ugj2Nra4ttvv4WtrS3+85//aPxjrLCwEB06dMChQ4fQq1cvjB8/\nHufPn8eiRYuwa9cuHD16FNWrV4eZmRlatWqF5ORklccrn1dSUpK0TBRF7N27F23atNF5rFJTUxEQ\nEAB7e3uMGDEC1atXx4kTJzBv3jwcPHgQ+/btQ4UKFfR6P/r6+mLixImYNm0ahg0bhtatWwNQP7mh\n6dhu2bIFv/76K0aNGgV7e3ssW7YMQ4cOhYmJCcLCwtCuXTtERETgyJEjiIqKgrm5OZYuXSo9ft++\nfThw4AC6du0KDw8PPHr0CGvWrMGQIUNw584dfPvttwCA4cOHIycnBwkJCfj555+l90TDhg21HqOH\nDx+iZcuWuHTpEgYNGoQmTZrg2LFjWLRoEZKSknDkyBG1qzYTJ05Efn4+RowYAVNTUyxatAghISGo\nVasWWrZsWWJOygSRZOXw4cOiqampKAiCWLt2bXHAgAHiokWLxDNnzqite+PGDdHMzEzs27ev2n1j\nxowRTUxMxMuXL0vL3NzcREEQxOXLl6utn5ycLAqCIMbGxqosz8/PV1t3+fLloiAI4po1a1SWC4Ig\nDhgwQOdzXLBggSgIgrht2zaV5cuWLRMFQRATEhJEURTFkydPioIgiJGRkTq3+TL69+8vCoIgZmVl\nib/88otoYmIiXrt2TRRFUVy5cqUoCIL4119/ifHx8WrHJDo6WhQEQdy3b5/adn19fUUPD49XWvZi\nbJqW9ejRQ2V5amqqqFAoxI4dO0rL0tLSREEQRFNTU/Hs2bMq6589e1YUBEFs3bq1WFhYKC2/ceOG\nWLlyZdHd3V18+vSpKIqiePXqVdHU1FRs0KCB+PDhQ7U4i4uLpRgEQRAnTZqkts5nn30m2tjYiLm5\nuaIoiuLYsWNFQRDEO3fuaHzuoqh/zpXPc/LkyWrLKlWqJGZkZKis7+XlJTo7O0u3CwsLRRcXF7Fa\ntWrigwcPpOW5ubmip6enxveCJs2aNRNdXFyk46b04YcfilWqVFE5zm5ubqK/v7/Keo8ePVLb5q1b\nt0RHR0exc+fOOvev3G69evXErKws8c6dO+Lly5fFqKgo0dbWVjQ1NRX/+uuvl96Xr6+vKAiC+M9/\n/lPn/tPS0kQnJyexdu3a4r1790RRFMW8vDyxatWqoq+vr9qxmTNnjigIgrh37161/b2Y98jISFEQ\nBHHnzp0647h27Zraa9Hf31/89NNPRRsbG3HmzJnS8iZNmogNGzZUebymzy9BEEQTExPxyJEjKsu7\ndOkiVqxYUeWYtmjRQjQzMxMvXLggLSsoKBCbN2+u9lpdsmSJKAiC+M0336hsd+vWraIgCGJwcLC0\nbOrUqaIgCNJ2MzIypHUEQRBv3boliqIonjp1ShQEQZw9e7bOY9WwYUOxXr160ntTKSEhQRQEQYyJ\niRFFUf/3o7bvD1HU/V69cuWKtPzOnTuiubm5KAiCOGfOHJXtdO/eXTQ1NVU55ppe08XFxaKfn59o\na2ur8v4LCwsTBUFQ+2wQRc2f6xMnThQFQRAXLVqksq7y++v594by8U2aNFHZ5/Xr10UzMzMxKChI\nbZ9lES9Jy8wHH3yA1NRU9O/fH9nZ2YiJicHIkSNRv359+Pr6Ii0tTVp37dq1KCgowMCBA5GVlaXy\n7+OPP0ZxcTESExNVtu/g4PBSQ0eYmZkBAJ4+fYoHDx4gKytL+qv8yJEjr/Qcg4KCYGpqqnaqPi4u\nDg4ODtKlYOUlsKSkJNy5c+eV9lUSQRDw+eefo2LFitLlx+joaDRv3hz169cv9f29rgkTJqjcbtKk\nCdq1a4fExES1y0hdunRB3bp1VZZt3LhR2s7zvZLOzs4YMGAAMjIycOLECQBAfHw8CgsLERYWJp2d\nfJ7ybMHKlSshCAL69eun9hr85JNPkJOTg8OHDwMAKleuDODZ67aoqEjjcyyNnH/22WdwdXVVWebn\n54ebN29Kxyk1NRWZmZkICQlRaYC3srLC8OHD9d5X//79kZmZid27d0vL0tLScOjQIQQFBensSbW0\ntJT+Pzc3F3fv3oVCoUDz5s3xxx9/6B3H2bNnUaVKFTg5OaFmzZoYNGgQnJycsHHjRum1/LL7EgRB\nZ+/jw4cP8fHHH6OoqAhbt26FnZ0dAGD37t24ffs2QkJCcO/ePZXXhfIs9q5du1S2ZWJigi+//FJl\nmfKz5uLFizqPQfXq1VGnTh3prJuyF7xjx47w9fXFnj17AAAPHjzAyZMnpbOQurRo0QLNmjVTi6uo\nqAjp6ekAgNu3b+P333/Hp59+ilq1aknrVaxYEWPHjlXbZkJCAkxMTFTOeANA586d0ahRI+m9Cjw7\ncwpAel5JSUmoUKECwsPDIQiCdPZR+V9dz+v06dM4ffo0goKCkJeXp5Ib5eV0ZW4M/Rn82WefoUaN\nGtJtR0dH1KlTBxUqVMCoUaNU1v3www9RWFgoHXNA9TWdn5+Pu3fv4u7du2jXrh2ys7Nx7ty5V44t\nISEBTk5OGDp0qMryYcOGoUqVKkhISFB7zMiRI1Xe8y4uLqhTp45er9+ygAWjDHl5eSE6Oho3b95E\neno6YmNj0bp1axw4cACffvopCgsLATzrBwSAjz76CE5OTir/2rdvD0EQpMtISjVr1nypX2quWbMG\n77//PiwtLWFvby99IQHA/fv3X+n52dnZ4eOPP8bGjRuRk5MD4FlPS0pKCvr06SO94dzc3DBp0iTs\n2rULzs7O8PHxwTfffIOjR4++0n41sbe3R9euXaVfLicnJ8t2LMnnL1s/v+zp06fIyMhQWV6nTh21\ndZV/bLx4+RqAVFRcvnwZwP8uZzZu3LjEmM6cOQNRFPHuu++qvQYHDx4MQRCkH2qFhoaicePGGDly\nJBwcHNClSxf88ssvyMrKkrZXGjn39PRUW6a8fK7svVQeixeLakDzsdNG0x8/cXFxEEVRr5aJS5cu\noU+fPrCzs4ONjY1U9G3fvh0PHjzQOw4PDw8kJiYiMTER+/fvx8WLF3H+/Hl07NjxlfdVpUoVjX8s\nKBUVFaFXr164ePEi1q1bh9q1a0v3KT+bBg4cqPa6qFevnsbPJhcXF7UfHryYt+LiYty8eVPlX3Z2\ntrS+v78/jh49itzcXBw6dAj5+fkICAiAv78/UlJSUFhYiL1796K4uFjvglGf15PyffPuu++qravp\nfZuWlgYXFxe1X+sCz96fOTk50vvCx8cH1tbWUkGYlJQEHx8feHp6wtvbWyqEk5KS4ODgoPYr5Rcp\ncxMWFqaWm6pVq+Lx48dSbgz9Gazp2NrZ2cHZ2VmtJUj5x8jz/dO5ubkYN24cXF1dYWlpKb2mv//+\newCv/h0FPMtR3bp11S7vm5iYoHbt2ionb0p6Pvb29mo932UVexhlztXVFcHBwQgODkbr1q1x8OBB\n/Pe//0XLli2lJu4VK1bA2dlZ4+Nf/BHG83+R6bJ+/Xr06dMH77//PubNm4caNWrA3NwcRUVF6Nix\no16N1dr069cP69evx5o1azBo0CCsWLECoiiif//+Kuv9+OOPGDhwILZu3YoDBw5g2bJliIyMxIQJ\nEzB9+vRX3v/zBg4ciE6dOmHIkCEwMzNDUFCQ1nVLKra1nTUzhpfJ8+sQRRGCIGDHjh1ah81QFqP2\n9vb473//iwMHDmD37t3Yv38/xo4di7CwMGzbtg0ffPABgNfPeUnDd4hafsX5quzt7dG5c2ds2LAB\njx49gpWVFVasWIH69etrHK7pebm5uWjTpg3y8vIwduxYeHt7w9raGgqFAtOmTVPrWyuJlZVViQXQ\nq+xL12to5MiRSExMxPLly9WG+lIe51mzZuG9997T+HgXFxeV2/rk7cqVK2pfyiEhIYiKigLw7Gzc\n4sWLsX//fhw6dEg665iXl4evv/4av//+O5KSkmBiYgJfX98Sn9/LxGVIFSpUQOvWrVUKRmVvrL+/\nPzZt2gRRFLFv3z589NFHOrenjHncuHEqf1A8T1mcAYb9DNZ2bPU95p9//jm2bt2KYcOGoU2bNnBw\ncICJiQm2bt2KOXPmvNZ31KvQFvebeJ28CSwYy5DmzZvj4MGDuH79OoD/nQlxcHDQ+6/ll7FixQpY\nWFggOTkZ5ubm0vKzZ8++9rY7d+4MR0dHrFixQioY69WrBx8fH7V1PTw8EBoaitDQUDx58gQdOnTA\nzJkzMW7cOKl5+XW0b98e77zzDhITE9G3b98Sz6rY29sDgMZfiaelpUmX8EvyqmPx/f3333j//ffV\nllWoUAFubm46H688M/znn3+q/SHx999/A/jfX8jKM2/Hjx9XucT2ojp16mDnzp2oUaOGxrMrL1Io\nFPD19ZW+rE+fPo2mTZti6tSp2LJli7SeoXPu7u4OQPNr+WUvY/Xv3x8bNmzAmjVrUKdOHVy+fBkz\nZszQ+bg9e/YgMzMT0dHRan8oTZw48aVieNP7ioyMxLJly/DNN99oPCOv/GyytLQs1c8mZ2dntTab\n5wtPPz8/CIKAPXv24PDhw9K+GzZsCEdHR+zZswfJyclo0qRJie/zl6V8PynP3j1P+d56nqenJ3bu\n3ImHDx+qnWX8+++/YWtrq/I6DwgIwLZt2xAfH48bN25Il6nbtm2LuXPnYt26dXj48KFex1qZG4VC\noXdudL0fjTHg9YMHD7Blyxb0798fCxcuVLnvxXYH4OU/dz09PXH27Fk8ffpUpRAsKirC+fPnNZ5N\nLO94SVpmdu/erXH4j7y8POzatQuCIEhnbAIDA2FmZoawsDDk5+erPebhw4coKCh45ViUb5Ln4xFF\nEVOnTn3lbSpVqFABn3/+OQ4cOIBVq1bh4sWLal9k2dnZ0uV3JTMzM6kwef5yw9mzZ6XLQi9LEAQs\nWLAA4eHh+Oabb0pcV/lh+3zPGgCsXr0amZmZeu2vUqVKJQ5LpO2D7flfvAPAsWPHkJiYiLZt2+p1\nRrFr164QBAGRkZEqZ0OVhYS7u7t0Cbpnz54wNTXF5MmTpbYBTYKDgwE8Kzo0/TX//Lihmi7L1K1b\nF+bm5lIuXybnr6NZs2ZwdnZGTEyMyuXY3Nxc/Prrry+1rS5dusDR0RFxcXGIi4uDQqHAF198ofNx\nyvfXi8dt165dr9wf/Cb2lZCQgG+//Rbdu3eXBgt/UYcOHeDk5ITp06drzFleXp7W4WZKYmZmhoCA\nAJV/z/+h4ujoCG9vb2zZsgVHjx6VCiJBEODv74/4+Hj8/fffpf4HdtWqVfHBBx9g48aNKsMAFRQU\nYM6cOWrrd+vWDcXFxWpn6LZv344TJ06ga9euKsuV8YaHh8Pc3BytWrUCALRp0wYmJiYIDw9XWa8k\njRs3hpeXF3799VeNl1WLiope+v2o/LXwm7z0amJiAkEQ1F7TmZmZWLZsmdrn6MvG2K1bN9y5c0ca\n5kdp6dKlyMrK0msUk/KGZxhlZuzYsbh37x66du0KLy8vWFpa4urVq1i1ahUuXLiA/v37Sz1o1atX\nx6JFizB48GDUq1cPwcHBcHV1xZ07d3D69Gls3LgRZ86cUfsBgL569eqF9evXIyAgAMHBwSgsLMSG\nDRuQl5dXKs+1f//+mDdvHkaMGAETExO1L9mkpCQMHToUPXv2RJ06dVCpUiWkpqZi+fLl+OCDD1R6\npurXrw83NzeNH4D6+OSTT/DJJ5/oXK9u3br46KOPsHjxYoiiiEaNGuHEiRPYsGEDatWqpfbhCqhf\njmjRogW2bt2K0NBQtGjRAiYmJmjbti2qVKmicX2lK1euoEOHDvjkk0+kYXWsrKzUhnTRpk6dOhg/\nfjxmzpyJNm3aIDAwEDk5OViyZAkeP36M1atXSx+y1atXx88//4xRo0bB29sb/fr1g6urK65fv45N\nmzYhKioKjRo1go+PD8LDwxEeHo733nsPvXr1grOzMzIzM5Gamort27dL4+INHjwY169fR/v27eHq\n6oq8vDz89ttvePTokdTv9zI5fx0mJiaYNWsW+vbti+bNm2PQoEEwMTFBTEwMHBwckJ6ervcZiQoV\nKiAoKAjz589Hamoq2rVrp7VF5HmtW7dGtWrV8PXXXyM9PV0a2uTf//43vL29NY7v+KpeZV+aXod3\n797FF198AUtLS3To0AH//ve/Ve6vVq0aPvroI1haWiIuLg6fffYZ6tati4EDB6JmzZp48OABzp49\ni4SEBGzYsAFt2rQpcX+vIiAgAD///DMEQVApoAICAqRxFw1xRWb27Nnw8/NDq1atMGrUKGlYHU0n\nAEJCQhAbG4sZM2YgPT0drVu3xsWLF7Fw4UJUq1YN06ZNU1n/vffeg52dHc6cOQN/f3+p19PGxgY+\nPj74448/4OLiorEnV5MVK1YgICAADRs2xMCBA1G/fn08fvwYFy9eREJCAqZPn45+/frp/X5s0KAB\nrK2tsXDhQlhaWsLW1hZVq1Z95bnT9XktWFtbo3379vj3v/8NCwsL+Pj4ICMjA0uWLIGnp6dan2WL\nFi0APBta7PPPP4e5uTm8vb019nQDz34cGB8fj1GjRuHYsWN47733cPz4cURFReHdd99V+xHi6z6f\nMuGN/iabdNq1a5c4atQosVGjRqKjo6NYoUIF0dHRUQwICBCjo6M1PubgwYNit27dRCcnJ9HU1FR0\ncXERAwICxNmzZ6sMi+Pu7q42rIdScnKyqFAo1IZFWLp0qVi/fn3R3NxcdHZ2FocNGybeu3dP6xAU\n+gyr8zxvb29RoVCI7du3V7svLS1NHD58uFivXj3RxsZGtLKyEuvXry+GhYWJ2dnZavvWNlTNi0JC\nQkSFQiHevXu3xPXi4+M1HpObN2+KvXr1Em1sbMRKlSqJnTt3Fs+ePSv6+fmpxaBp2ePHj8VBgwaJ\nVatWFU1MTESFQiEN56CM7Xn9+/cXFQqFmJWVJQYHB4sODg6ipaWl2LZtW/HYsWMq62oawuJFS5cu\nFRs3biyam5uLNjY2Yvv27cWUlBSN6+7atUts166daGtrK5qbm4s1a9YUhw4dqnbstm7dKnbo0EG0\nt7cXzczMRFdXV7Fz587i4sWLpXXWr18vdu3aVXznnXdEMzMzsUqVKqKfn5+4fv16lfj1yXlJQ3Vo\neu7h4eGiQqFQG1IjPj5ebNiwoRRzWFiYNLRIfHy81mP4IuXwQgqFQly1apXGdTS9/06dOiV27NhR\ntLOzE62trUV/f38xJSVF4+tAG3d3d9Hb21vnei+zL02vW1H83zFWKBSiIAhq/158fn/++af4xRdf\niNWrVxdNTU3FqlWriq1atRKnTp0qDcGjz/5Kej2/aPPmzaIgCGKtWrVUll+4cEEUBEE0MzMT8/Ly\n1B73Mp9p0dHRKu9bpf3794stW7YUzc3NxWrVqomhoaHin3/+qfE5PHr0SPzuu+9ET09P6dj069dP\nZZiZ5/Xo0UNUKBTi1KlTVZZPmjRJVCgU4hdffKH9oGiQkZEhDh8+XHR3dxdNTU1FBwcH0cfHR5w4\ncaI0zNjLfAZv27ZNbNKkiTQsjvK18LLvVW2vBU3HPCsrSxw8eLDo4uIimpubiw0bNhSXLVsmxsTE\naMzPzJkzRU9PT7FixYqiQqGQ9q8tn3fu3BFHjhwpvvPOO2LFihXFGjVqiKGhoWqff9oeX9LzKYsE\nUSwvpS9R+RQSEoK4uLg33sD9tvrXv/6F8ePH4/fff0fz5s2NHQ4RkSywh5GoDDBGU3l5V1hYqHa5\nMDc3FwsWLICjo6M0KwgREbGHkahM4IWA0nfp0iV06tQJQUFBcHd3R2ZmJmJjY5GRkYFFixbpHHSb\niOhtwk9xJtkgAAAgAElEQVREIpnTNAcxvT4nJye0aNECK1euxO3bt1GhQgU0bNgQM2fORM+ePY0d\nHhGRrLCHUYf33nsPJ0+eNHYYRERERDr5+vpi7969pb5dFow6CILAy4EyoxzGheSFeZEn5kWemBd5\nUX7Xl4e8GKpu4Y9eqMx5fvJ5kg/mRZ6YF3liXuSJedGOBSMRERG91cLCwowdguyxYKQyJyQkxNgh\nkAbMizwxL/LEvMiL8jI086Idexh1YA8jERERlRXsYST6f4b49Re9PuZFnpgXeWJe5Il50Y4FIxER\nERGViJekdeAlaSIiIioreEmaiIiIyADK+tiLbwILRipz2GMiT8yLPDEv8sS8yMvkyZMBMC8lYcFI\nRERERCViD6MO7GEkIiIq38rTdz17GImIiIjIKFgwUpnDHhN5Yl7kiXmRJ+ZFnpgX7VgwEhER0VuN\nc0nrxh5GHcpTXwMRERGVb+xhJCIiIiKjYMFIZQ57TOSJeZEn5kWemBd5Yl60Y8FIRERERCViD6MO\n7GEkIiKisoI9jEREREQGwLmkdWPBSGUOe0zkiXmRJ+ZFnpgXeeFc0rqxYCQiIiKiErGHUQf2MBIR\nEZVv5em7nj2MRERERGQULBipzGGPiTwxL/LEvMgT8yJPzIt2LBiJiIjorca5pHVjD6MO5amvgYiI\niMo39jASERERkVGwYKQyhz0m8sS8yBPzIk/MizwxL9qxYCQiIiKiErGHUQf2MBIREVFZwR5GIiIi\nIgPgXNK6sWCkMoc9JvLEvMgT8yJPzIu8cC5p3VgwEhEREVGJ2MOoA3sYiYiIyrfy9F3PHkYiIiIi\nMgoWjFTmsMdEnpgXeWJe5Il5kSfmRTsWjERERPRW41zSurGHUYfy1NdARERE5Rt7GImIiIjIKFgw\nUpnDHhN5Yl7kiXmRJ+ZFnpgX7VgwEhEREVGJ2MOoA3sYiYiIqKxgDyMRERGRAXAuad1YMFKZwx4T\neWJe5Il5kSfmRV44l7RuLBiJiIiIqETsYdSBPYxERETlW3n6rmcPIxEREREZBQtGKnPYYyJPzIs8\nMS/yxLzIE/OiHQtGIiIieqtxLmnd2MOoQ3nqayAiIqLyjT2MRERERGQULBipzGGPiTwxL/LEvMgT\n8yJPzIt2LBiJiIiIqETsYdSBPYxERERUVrCHkYiIiMgAOJe0biwYqcxhj4k8MS/yxLzIE/MiL5xL\nWjcWjERERERUIvYw6sAeRiIiovKtPH3Xs4eRiIiIiIyCBSOVOewxkSfmRZ6YF3liXuSJedGOBSMR\nERG91TiXtG7sYdShPPU1EBERUfnGHkYiIiIiMgoWjFTmsMdEnpgXeWJe5Il5kSfmRTsWjERERERU\nIvYw6sAeRiIiIior2MNIREREZACcS1o3FoxU5rDHRJ6YF3liXuSJeZEXziWtGwtGIiIiIioRexh1\nYA8jERFR+VaevuvZw0hERERERsGCkcoc9pjIE/MiT8yLPDEv8sS8aMeCkYiIiN5qnEtaN/Yw6iAI\nAibOmGjsMIjKBCcbJ4wZPsbYYRARvbUM1cNYodS3WA65tXUzdghEZULGngxjh0BERAbAS9JU5pw7\nes7YIZAGzIs8sSdLnpgXeWJetGPBSEREREQlYg+jDoIgYPHRxcYOg6hMyNiTgZ8m/GTsMIiI3loc\nh5GIiIjIADiXtG4sGKnMYa+cPDEv8sSeLHliXuSFc0nrxoKRiIiIiErEHkYd2MNIpD/2MBJRWcS5\npHXjGUYiIiIiKhELRipz2CsnT8yLPLEnS56YF3liXrRjwUhERERvNc4lrRt7GHVgDyOR/tjDSERk\nXOxhJCIiIiKjYMFIZQ575eSJeZEn9mTJE/MiT8yLdiwYiYiIiKhELBipzKnrU9fYIZAGr5uXgoIC\nDBo0CO7u7rCxsUHjxo2xY8cOtfWmTJkChUKBpKQkaVlRURFGjx4NZ2dnODg4oGvXrrhx44bG/fz+\n++9o164dHBwc4OTkhMDAQNy8eVOvbRUVFaFPnz6ws7NDp06dkJOTIz1u2rRpmDNnzmsdA0Pw8/Mz\ndgikAfMiT8yLdiwYiUgWioqK4Orqiv379yM7OxtTp05FYGAgMjIypHUuXbqEtWvXwsXFReWxCxcu\nxIEDB3Dq1CncuHEDdnZ2GD16tMb9PHjwAMOHD0dGRgYyMjJgbW2NAQMG6LWt9evXw8TEBHfv3oWt\nrS2WLFkCAEhLS8PmzZsxZsyY0j4sRPQGcC5p3VgwUpnDXjl5et28WFpaIiwsDK6urgCALl26wMPD\nA8eOHZPWCQ0NxYwZM1CxYkWVx/7111/o0KEDqlSpAjMzMwQGBuKvv/7SuJ+OHTuiR48eqFSpEiws\nLDBq1CgcPHhQr22lp6fD19cXCoUCfn5+uHz5MgDgyy+/xOzZs6FQyO8jlT1Z8sS8yAvnktZNfp9u\nREQAbt26hfPnz6NBgwYAgPj4eJibm6NTp05q67Zv3x7bt29HZmYmHj9+jJUrV6Jz58567Wf//v3w\n8vLSa1teXl5ISkrCkydPkJycDC8vLyQkJMDJyQktWrQohWdNRCRPFYwdANHLYg+jPNX1qYuMPRm6\nV9RDYWEh+vbti5CQENSpUwc5OTmYNGkSEhMTNa7fo0cPbNq0CdWrV4eJiQkaNmyIBQsW6NzPqVOn\n8OOPP2LTpk16batz5844cOAAmjdvjhYtWqB3795o27YtEhMTMWnSJKSkpMDLyws///yz2llQY2FP\nljwxL/LEvGj31p5h3LZtG9q0aQNra2vY2tqiWbNmSE5ONnZYRG+94uJiBAcHw9zcHPPnzwfwrL8o\nODhYulwNQGVg2nHjxiEnJwf37t3Do0eP0K1bN41nIp938eJFdO7cGfPmzUOrVq303lZERAROnjyJ\nX3/9FRERERgxYgT++OMPpKamYt++fSgoKEBUVFRpHQ4iIll4KwvGxYsX47PPPkOzZs2wYcMGxMfH\nIzAwEHl5ecYOjfTAHkZ5Ko28iKKIQYMG4c6dO1i3bh1MTEwAAElJSZg3bx6cnZ3h7OyMq1evIjAw\nEJGRkQCAHTt2YMCAAahcuTJMTU0RGhqKI0eO4N69exr3k5GRgXbt2uGHH35A3759Ve7Td1unT5/G\n4cOHMWTIEJw+fRpNmzYFAPj4+ODUqVOvfSxKC3uy5Il5kSfmRbu37pJ0eno6vvrqK8yaNQtffvml\ntLx9+/ZGjIqIAGDEiBE4e/YsEhMTYWZmJi3fs2cPioqKADwrKps1a4Y5c+ZIZ/4aNmyI2NhY+Pr6\nwsLCAgsXLkT16tVhb2+vto/r168jICAAoaGhGDp0qNr9+mxLFEWMHj0av/zyCwRBgKenJ+bPn4+C\nggLs27cPPj4+pX1oiMiAOJe0bm/dGcaoqChUqFABw4cPN3Yo9IrYwyhPr5uXjIwMLFmyBCdPnkS1\natVgbW0Na2trrF69Gvb29nBycoKTkxOqVq0KExMT2NnZwdLSEgAwZ84cKBQK1KxZE05OTtixYwcS\nEhKkbXt5eWH16tUAgGXLliEtLQ3h4eHSPmxsbKR1dW0LAGJiYuDt7Y3GjRsDALp37w4XFxc4OTnh\n/v37GgtRY2FPljwxL/KiHFaHedFOEA0xQ7WMBQQEIDs7G6Ghofjxxx9x5coVuLu7Y+zYsRg5cqTa\n+oIgYPHRxUaIlKjsydiTgZ8m/GTsMIiI3lqCIMAQpd1bd4bxxo0buHDhAiZMmICJEydi9+7daNeu\nHUJDQzFv3jxjh0d6YA+jPDEv8sSeLHliXuSJedHurethLC4uRk5ODmJjY/HZZ58BeHYKOj09HRER\nESp9jUrRYdFwdHEEAFhUskCNujWky2/KL0nefnO3r567Kqt4ePt/tzMuZWDv3r3SZR3lhy9vG+/2\niRMnZBUPb/O2nG+XxfeL8v/T09NhSG/dJekWLVrgyJEjyM7OhpWVlbR8zpw5+Prrr5GZmYmqVatK\ny3lJmkh/vCRNRGRcvCRdSho0aGCQA0lERERlE+eS1u2tKxi7d+8O4NlYa8/bsWMHatSooXJ2keSJ\nvXLyxLzI0/OXrUg+mBd54VzSur11PYydO3eGv78/hg0bhqysLHh4eCA+Ph67d+9GTEyMscMjIiIi\nkp23rocRAHJycvDdd99h7dq1uH//PurVq4dvv/0Wffr0UVuXPYxE+mMPIxGVRYbq+zMGQz2Xt+4M\nIwBYW1tj/vz50jy1RERERKTdW9fDSGUfe+XkiXmRJ/ZkyRPzIk/Mi3YsGImIiOitxrmkdXsrexhf\nBnsYifTHHkYiIuPiOIxEREREZBQsGKnMYa+cPDEv8sSeLHliXuSJedGOBSMRERERlYg9jDqwh5FI\nf+xhJCIyLvYwEhERERkA55LWjQUjlTnslZMn5kWe2JMlT8yLvHAuad1YMBIRERFRidjDqAN7GIn0\nxx5GIiqLOJe0bjzDSEREREQlYsFIZQ575eSJeZEn9mTJE/MiT8yLdiwYiYiI6K3GuaR1Yw+jDuxh\nJNIfexiJiIyLPYxEREREZBQsGKnMYa+cPDEv8sSeLHliXuSJedGOBSMRERERlYg9jDqwh5FIf+xh\nJCIyLvYwEhERERkA55LWjQUjlTnslZMn5kWe2JMlT8yLvHAuad0qGDuAsiBjT4axQ6Dn3Lp0C+YP\nzY0dBr3g1qVbaNa4mbHDICIiA2APow7laX5JIiIiUleevuvZw0hERERERsGCkcoc9pjIE/MiT8yL\nPDEv8sS8aMeCkYiIiN5qnEtaN/Yw6lCe+hqIiIiofGMPIxEREREZBQtGKnPYYyJPzIs8MS/yxLzI\nE/OiHQtGIiIiIioRexh1YA8jERERlRXsYSQiIiIyAM4lrRsLRipz2GMiT8yLPDEv8sS8yAvnktaN\nBSMRERERlYg9jDqwh5GIiKh8K0/f9YZ6LhVKfYvl0KSZk4wdAlGZ4GTjhDHDxxg7DCIiKmUsGPXg\n1tbN2CHQc84dPYe6PnWNHQa94NzRc7j98Laxw6AX7N27F35+fsYOg17AvMgT86IdexiJiIjorca5\npHVjD6MOgiBg8dHFxg6DqEzI2JOBnyb8ZOwwiIjeWhyHkYiIiIiMggUjlTnnjp4zdgikAfMiTxxX\nTp6YF3liXrRjwUhEREREJdKrh/Hp06cAABMTEwBAZmYmtm7dinr16qFVq1aGjdDI2MNIpD/2MBIR\nGZdRexi7dOmC+fPnAwByc3PRrFkzjB8/Hr6+voiNjS31oIiIiIjeFM4lrZteBWNqair8/f0BAOvX\nr4e1tTVu376NZcuW4V//+pdBAyR6EXvl5Il5kSf2ZMkT8yIvnEtaN70KxtzcXNjZ2QEAdu3ahW7d\nuqFixYrw9/fHxYsXDRogERERERmXXgVjjRo1kJKSgtzcXOzcuRPt2rUDANy7dw+WlpYGDZDoRZzl\nRZ6YF3nirBXyxLzIE/OinV5TA3799dfo168frKys4ObmhjZt2gAA9u/fj4YNGxo0QCIiIiIyLr3O\nMA4bNgyHDx9GVFQUDh48KP1aumbNmvjxxx8NGiDRi9grJ0/MizyxJ0uemBd5Yl600+sMIwD4+PjA\nx8dHZdnHH39c6gERERERvUmcS1o3vc4wiqKIBQsWoEGDBrCwsMDly5cBANOnT8eaNWsMGiDRi9gr\nJ0/MizyxJ0uemBd5UQ6rw7xop1fBOHfuXEydOhVDhgxRWe7i4iKNz0hE9DoKCgowaNAguLu7w8bG\nBo0bN8aOHTvU1psyZQoUCgX27NkjLevUqROsra2lf2ZmZlr7q1euXKmyrpWVFRQKBY4fPw7g2RdH\nxYoVpfttbGyQnp4OACgqKkKfPn1gZ2eHTp06IScnR9rutGnTMGfOnFI8IkRE8qFXwbho0SIsXboU\nX331FSpU+N9V7CZNmuDPP/80WHBEmrBXTp5eNy9FRUVwdXXF/v37kZ2djalTpyIwMBAZGRnSOpcu\nXcLatWvh4uICQRCk5du3b0dOTo70r2XLlggMDNS4n759+6qsu3DhQtSsWRONGzcG8GyWhKCgIOn+\n7OxsuLu7A3g2Dq2JiQnu3r0LW1tbLFmyBACQlpaGzZs3Y8yYMa91DAyBPVnyxLzIE/OinV4F45Ur\nV+Dt7a22vGLFisjLyyv1oIjo7WNpaYmwsDC4uroCeDbDlIeHB44dOyatExoaihkzZqBixYpat5Oe\nno4DBw6gX79+eu03JiZGZV1RFLVOq5Weng5fX18oFAr4+flJ7TlffvklZs+eDYVCr49UIqIyR69P\nNw8PD6Smpqot3759O+rXr1/qQRGVhL1y8lTaebl16xbOnz+PBg0aAADi4+Nhbm6OTp06lfi4uLg4\ntGnTRio8S5KRkaFWXAqCgM2bN8PBwQFeXl749ddfpfu8vLyQlJSEJ0+eIDk5GV5eXkhISICTkxNa\ntGjxis/UsNiTJU/MizwxL9rpVTCOHz8eoaGhWLlyJYqLi3Ho0CGEh4dj4sSJGD9+vKFjNKiOHTtC\noVDgn//8p7FDIaL/V1hYiL59+yIkJAR16tRBTk4OJk2ahLlz5+p8bFxcHEJCQvTaj7K4dHNzk5YF\nBgbi7NmzyMrKwtKlSzFlyhT85z//AQB07twZHh4eaN68Oezs7NC7d29MmTIFM2fOxKRJk+Dr64tR\no0ahsLDwlZ43ERkH55LWTa+CccCAAZg8eTK+++475OXloV+/fli2bBl++eUX9OnTx9AxGszq1atx\n6tQpAFDphyJ5Yw+jPJVWXoqLixEcHAxzc3PpR3Xh4eEIDg5WOWuo6bJxSkoKbt26hZ49e+q1r7i4\nOPTv319lWb169VCtWjUIgoAWLVpgzJgxWLt2rXR/REQETp48iV9//RUREREYMWIE/vjjD6SmpmLf\nvn0oKChAVFTUqzx1g2BPljwxL/LCuaR107vhZsiQIbhy5Qpu3bqFzMxMXLt2DYMGDTJkbAZ1//59\n/OMf/+CvGolkRBRFDBo0CHfu3MG6deukSQKSkpIwb948ODs7w9nZGVevXkVgYCAiIyNVHh8bG4se\nPXroNWXpwYMHkZmZqXdx+aLTp0/j8OHDGDJkCE6fPo2mTZsCeDZmrfIPUSKi8kKvgvHp06d4+vQp\nAKBKlSooLi7GsmXLcPDgQYMGZ0jffPMNvL290bt3b2OHQi+JPYzyVBp5GTFiBM6ePYtNmzbBzMxM\nWr5nzx789ddfOHnyJE6cOAEXFxcsWbIEI0eOlNbJy8tDfHy83pejY2Nj0bNnT1hZWaks37hxI+7f\nvw9RFHHkyBHMmzcPn376qco6oihi9OjR+OWXXyAIAjw9PZGSkoKCggLs27cPNWvWfPWDUMrYkyVP\nzIs8MS/a6VUwdunSRbo0lJubi2bNmmH8+PHw9fVFbGysQQM0hJSUFKxYsQILFiwwdihE9P8yMjKw\nZMkSnDx5EtWqVZPGQVy9ejXs7e3h5OQEJycnVK1aFSYmJrCzs1Mp9jZs2AA7OzuNH/heXl5YvXq1\ndDs/Px/x8fFql6MB4LfffkPt2rVhY2OD/v3747vvvkNwcLDKOjExMfD29paG4unevTtcXFzg5OSE\n+/fvY+jQoaV0VIiI5EEQtY0f8ZwqVapgz549aNiwIeLi4hAREYFTp05h5cqVmD17dpm6/FJQUIDG\njRujR48emDJlCgBAoVDg+++/l24/TxAELD66+E2HSSU4d/QczzLK0Lmj52D+0Bw/TfjJ2KHQc/bu\n3cuzJjLEvMiLIAgQRbFc5EX5XEqbXmcYc3NzYWdnBwDYtWsXunXrhooVK8Lf3x8XL14s9aAMaebM\nmXjy5AkmTZpk7FCIiIhIBjiXtG4VdK8C1KhRAykpKfjkk0+wc+dOaf7oe/fu6dVcLhdXrlzBTz/9\nhOXLlyMvL09l0PH8/Hw8fPgQ1tbWaoPvRodFw9HFEQBgUckCNerWkM5wKX8Zyttv9raSXOLh7bqo\n61MX+5fsV/kLXfmLQ9427m0lucTD237w8/OTVTxv++3w8PAy+35R/r9yClND0euS9OLFixEaGgor\nKyu4ubnh2LFjMDExwdy5c7Fx40YkJSUZNMjSsnfvXgQEBJS4zokTJ1TmoOUlaSL9ZezJ4CVpIiIj\nMuol6WHDhuHw4cOIiorCwYMHpaEuatasiR9//LHUgzKUxo0bY+/evSr/kpOTAQDBwcHYu3evrH7d\nSJpxHEZ5Yl7k6cWzJiQPzIs8MS/a6XVJGng2tpiPj490u7CwEB9//LFBgjIUW1tbtGnTRuN9bm5u\nWu8jIiIiepvpdYZx7ty5WLdunXR74MCBMDc3R506dXDuHM8q0JvFX0jLE/MiT8p+J5IX5kWemBft\n9CoY582bB0fHZz/62L9/P+Lj47Fq1So0btwYX3/9tUEDfBOKi4s1DqlDRERE5R/nktZNr4Lxxo0b\n8PT0BABs3rwZPXv2RO/evREeHo7Dhw8bNECiF7FXTp6YF3liT5Y8MS/ywrmkddOrYLSxscGtW7cA\nALt370bbtm0BABUqVEB+fr7hoiMiIiIio9PrRy/t27fHkCFD0KRJE1y8eBGdOnUCAPz999/w8PAw\naIBEL2KvnDzV9amLjD0Zxg6DXsCeLHliXuSJedFOrzOM8+fPx4cffoisrCysXbsWDg4OAIDU1FR8\n/vnnBg2QiIiIiIxLr4G732YcuFt+OJe0PHEuaXnaWw7mxi2PmBd54VzSuul1hhEAbt68icjISIwY\nMQJZWVkAgJSUFKSlpZV6UERERERvCueS1k2vM4ypqakICAiAp6cn/vzzT5w7dw6enp4ICwvDhQsX\nsGrVqjcRq1HwDCOR/jg1IBGRcRn1DOPXX3+NMWPG4Pjx4zA3N5eWd+zYESkpKaUeFBERERHJh14F\n47FjxxASEqK2vFq1atJwO0RvCsf7kyfmRZ44rpw8MS/yxLxop1fBaGFhgXv37qktP3fuHJycnEo9\nKCIiIiKSD70Kxk8//RSTJ09WGaQ7LS0NEyZMQI8ePQwWHJEm/IW0PDEv8lTWf/FZXjEv8sS8aKdX\nwRgZGYn79++jSpUqePz4MT788EPUqlULlStXxtSpUw0dIxEREZHBcC5p3fQqGG1tbXHgwAFs3LgR\n06dPx5gxY7Bz507s378flSpVMnSMRCrYKydPzIs8sSdLnpgXeeFc0rrpNTUg8Oxn2gEBAQgICDBk\nPEREREQkM3qdYQwJCcGcOXPUls+ePRuDBw8u9aCISsJeOXliXuSJPVnyxLzIE/OinV4F444dO+Dv\n76+2PCAgAFu3bi31oIiIiIhIPvQqGB88eKCxV9HS0lLjcDtEhsReOXliXuSJPVnyxLzIE/OinV4F\nY+3atbFlyxa15du2bUOtWrVKPSgiIiKiN4VzSeum11zSsbGxGD58OMaOHYu2bdsCABITE/Hzzz9j\nwYIFGDhwoMEDNRbOJU2kP84lTURkXIaaS1qvX0n3798f+fn5+PHHHzF9+nQAQPXq1TFnzpxyXSwS\nERERkZ6XpAFg2LBhuHbtGm7evImbN2/i6tWrGD58uCFjI9KIvXLyxLzIE3uy5Il5kSfmRTu9x2EE\ngMuXL+Pvv/+GIAioV68ePD09DRUXEREREcmEXj2M2dnZGDhwINavXw+F4tlJyeLiYvTo0QNRUVGw\ntrY2eKDGIggCJs6YaOwwiMoEJxsnjBk+xthhEBG9tQzVw6hXwThgwAAcOnQIS5YsQYsWLQAAhw4d\nwrBhw9CqVStERUWVemByYagDT0RERPIQHh5ebuaTNlTdolcP46ZNm7B06VL4+vrC1NQUpqam8PPz\nw9KlS7Fhw4ZSD4qoJOwxkSfmRZ6YF3liXuSFc0nrplfBmJeXBwcHB7Xl9vb2yM/PL/WgiIiIiEg+\n9Lok/dFHH8HGxgYrVqyAlZUVACA3Nxf9+vVDdnY2EhMTDR6osfCSNBERUflWnr7rjdrDePr0aXTo\n0AGPHz9Go0aNIIoiTp8+DUtLS+zcuRNeXl6lHphclKcXEREREakrT9/1Ru1h9Pb2xoULFxAZGYmm\nTZvCx8cHkZGRuHjxYrkuFkme2GMiT8yLPDEv8sS8yBPzop3OcRgLCgrg6uqKPXv2YMiQIW8iJiIi\nIqI3hnNJ66bXJel33nkHu3btQv369d9ETLJSnk5TExERUflm1EvSo0ePRkREBAoLC0s9ACIiIiKS\nN70KxpSUFGzcuBHvvPMO2rZti08++UT617VrV0PHSKSCPSbyxLzIE/MiT8yLPDEv2uk1l7SDgwO6\nd++u8T5BEEo1ICIiIiKSF716GN9m7GEkIiKissJQdYteZxiVLl26hDNnzgAA6tWrh5o1a5Z6QHI0\naeYkY4dAVG442ThhzPAxxg6DiEhSnuaSNhS9zjDevXsXAwcOxObNm6FQPGt7LC4uxscff4zo6GiN\n0waWF4IgYPHRxcYOg55z7ug51PWpa+ww6AX65iVjTwZ+mvDTG4iIgGc9WX5+fsYOg17AvMiL8qxc\neciLUX8lPXjwYFy6dAkHDhxAXl4e8vLycODAAaSlpWHw4MGlHhQRERERyYdeZxgtLS2RmJiIli1b\nqiw/fPgw2rZti8ePHxssQGPjGUai0sUzjEQkN+Xp9wpGPcPo6OgIKysrteWWlpZwdHQs9aCIiIiI\nSD70Khh/+OEHjB07FteuXZOWXbt2Df/4xz/www8/GCw4Ik3OHT1n7BBIA+ZFnjiunDwxL/LEvGin\n16+k586di/T0dLi7u6N69eoAgOvXr8PCwgK3b9/G3LlzATw7DXrq1CnDRUtERERUyjiXtG56FYw9\nevTQa2McxJveBP5CWp6YF3kq67/4LK+YF3lRDqnDvGinV8HIsYmIiIiI3l569TASyQl75eSJeZEn\n9mTJE/MiT8yLdiwYiYiIiKhEnEtaB47DSFS6OA4jEZHhGHUcRiIiIqLyir/V0E1rwWhiYoLbt28D\nAAYOHIjs7Ow3FhRRSdgrJ0/MizyxJ0uemBd5mTx5MgDmpSRaC0YLCwvk5OQAAGJiYpCfn//GgiIi\nIp2VIUAAACAASURBVCIi+dA6rE7Lli3RrVs3NGnSBAAwZswYWFhYqKwjiiIEQUBUVJRhoyR6Dsf7\nkyfmRZ44rpw8MS/yxLxop7VgjIuLw6xZs3Dx4kUAwN27d2FqaqoyOLeyYCQiIiKi8kvrJelq1aph\n1qxZ2LBhA1xdXbFq1Sps2bIFmzdvlv4pbxO9SeyVk6c3kZeCggIMGjQI7u7usLGxQePGjbFjxw4A\nwN9//w0fHx/Y29ujcuXKaNWqFVJSUqTHdurUCdbW1tI/MzMzNGzYUON+Vq5cqbKulZUVFAoFjh8/\nLq1z7NgxtGnTBtbW1qhWrRrmzZsHACgqKkKfPn1gZ2eHTp06Sa09ADBt2jTMmTPHEIdGK/ZkyRPz\nIk/Mi3Z6/Uo6PT0djo6Oho6FiKhERUVFcHV1xf79+5GdnY2pU6ciMDAQGRkZqF69OuLj43H37l3c\nv38fffr0Qc+ePaXHbt++HTk5OdK/li1bIjAwUON++vbtq7LuwoULUbNmTTRu3BgAkJWVhU6dOmHE\niBG4d+8eLl26hPbt2wMA1q9fDxMTE9y9exe2trZYsmQJACAtLQ2bN2/GmDFjDHyUiOhlcS5p3fQe\nVmfLli1o3bo1HBwc4OjoCF9fX2zdutWQsRFpxF45eXoTebG0tERYWBhcXV0BAF26dIGHhweOHTsG\nW1tbeHh4QBAEPH36FAqFAs7Ozhq3k56ejgMHDqBfv3567TcmJkZl3dmzZ6Njx44ICgpCxYoVYWVl\nhXfffVfatq+vLxQKBfz8/HD58mUAwJdffonZs2dDoXizo5mxJ0uemBd54VzSuun1ybVs2TJ0794d\ntWrVwowZMzB9+nR4eHigW7duWL58uaFjJCLS6NatWzh//jwaNGggLatcuTIsLCwwc+ZMrF27VuPj\n4uLi0KZNG6nwLElGRoZacfnHH3/Azs4OrVq1QtWqVdG1a1dcvXoVAODl5YWkpCQ8efIEycnJ8PLy\nQkJCApycnNCiRYvXfMZERMahV8E4Y8YMzJ49G9HR0Rg8eDAGDx6MmJgY/Otf/8KMGTMMHWOp2rlz\nJwICAuDs7Axzc3PUqFEDvXv3xpkzZ4wdGumJPYzy9KbzUlhYiL59+yIkJAR16tSRlj948AAPHz5E\nnz590KtXL40zHsTFxSEkJESv/SiLSzc3N2nZ1atXERsbi3nz5uHKlSvw8PBAUFAQAKBz587w8PBA\n8+bNYWdnh969e2PKlCmYOXMmJk2aBF9fX4waNQqFhYWvdwD0xJ4seWJe5Il50U6vgvHKlSvo2LGj\n2vKOHTsiPT29tGMyqPv376NZs2ZYsGABdu/ejYiICPz111/44IMPpDMERCRvxcXFCA4Ohrm5OebP\nn692v6WlJaZPn47z58/j9OnTKvelpKTg1q1bKv2NJYmLi0P//v3Vtt+9e3c0bdoUZmZmCAsLw6FD\nh6QfuERERODkyZP49ddfERERgREjRuCPP/5Aamoq9u3bh4KCAg5HRkRlil4FY40aNbBr1y615bt3\n71b5q7ss6NOnD2bMmIHu3bujdevW+OKLL7B+/Xrk5ORovXxF8sIeRnl6U3kRRRGDBg3CnTt3sG7d\nOpiYmGhc7+nTpyguLoalpaXK8tjYWPTo0UNtuSYHDx5EZmamWnGp7dfVLzp9+jQOHz6MIUOG4PTp\n02jatCkAwMfHB6dOndJrG6+LPVnyxLzIE/OinV4F4/jx4/HVV19h8ODBiI6ORnR0NAYNGoSvvvoK\n48aNM3SMBmdvbw8AWr94iEg+RowYgbNnz2LTpk0wMzOTlicmJuLEiRN4+vQpsrOz8Y9//AN169ZF\nrVq1pHXy8vIQHx+v9+Xo2NhY9OzZE1ZWVirLBwwYgISEBJw8eRKFhYX48ccf0bp1a1hbW0vriKKI\n0aNH45dffoEgCPD09ERKSgoKCgqwb98+1KxZ8/UOBBGVGs4lrZteBeOwYcPw22+/4cyZMxg3bhzG\njRuHc+fOIT4+HsOGDTN0jAbx9OlTFBQU4MKFCxg2bBiqVq2KPn36GDss0gN7GOXpTeQlIyMDS5Ys\nwcmTJ1GtWjVpnMRVq1bhwYMHCAoKQuXKlVG3bl3cuXMHmzZtUnn8hg0bYGdnp/EsgpeXF1avXi3d\nzs/PR3x8vNrlaADw9/fHtGnT0KVLF1StWhWXL1/GqlWrVNaJiYmBt7e3NBRP9+7d4eLiAicnJ9y/\nfx9Dhw4thSOiG3uy5Il5kRfOJa2bIGrqCH8L+Pj44NixYwAANzc3bN26FfXr11dbTxAELD66+E2H\nRyU4d/QcL0vL0P+1d+dxVdX5/8Bf5yoICAgpICAi4C6oCDlp7pOm5JKaS5qKmlt9zZrMMiaFHFya\nEXNpcsnUxiWdxFzGVFJBEZVwRQwtA1FUFGUUcGH7/P7oxx2vcBeNy/3ce1/Px4NHnOWe8768vfHm\nc97nfAzNy+X9lxE9I7oaIiLg91+AvMwmH+ZFLoqiQAhhEXkpfy9VflxrLRjT09ORn5+PS5cu4R//\n+AdycnKQmJhYoSeTBSNR1WLBSESyMVaRZQosGI3o7t27aNSoEYYPH44vv/xSY5uiKHjhlRdQz+v3\nmW7sHe3h08xHPZJSfhmOy1zmsmHLOSdzsH7FegD/u/xT/hc9l7nMZS6bYllRFBw8eFCaeJ5mufz7\n8qfWrFu3jgWjMZXPQfvk3eAcYZQPL0nLiZek5RRvAZfYLBHzIhdektaveueoklROTg7S09N51yIR\nEZEV4lzS+ukdYSwqKkLnzp3xzTffoFkz8x/VGThwIEJCQhAUFARnZ2dcvHgRixYtws2bN5GcnKzx\nCA6AI4xEVY0jjERExmOsEcaa+nawtbVFRkYGFEWp8pObQocOHbBlyxYsXLgQRUVF8PHxQffu3TFz\n5kyD5pUlIiIisjYGXZIePXo0Vq1aZexYqsWMGTOQkpKCvLw8FBYWIj09HV9++SWLRTPC5zDKiXmR\n0+ON8SQP5kVOzIt2ekcYAeD+/ftYv3494uLiEBISop71QAgBRVGwZMkSowZJRERERKZj0F3ST94x\nVH55urxgLL8V3RKxh5GoarGHkYjIeEzWwwhwiJaIiIgsV2RkJOeT1uOpHquTm5uL48eP4+HDh8aK\nh0gv9srJiXmRE//glxPzIhfOJa2fQQVjfn4+hgwZAnd3d3Ts2BHXrl0DAEyePJkVOREREZGFM6hg\n/PDDD5GdnY2TJ0/C3t5evb5v376IjY01WnBEleEsL3JiXuRk7rNWWCrmRU7Mi3YG9TDu2LEDsbGx\naNu2rcbzGJs3b47ffvvNaMERERERkekZNMKYl5eHunXrVlifn5+PGjVqVHlQRLqwV05OzIuc2JMl\nJ+ZFTsyLdgYVjKGhodixY0eF9StXrkTHjh2rPCgiIiKi6sK5pPUz6DmMSUlJePnllzFs2DCsX78e\nEyZMwLlz55CcnIxDhw4hJCSkOmI1CT6Hkahq8TmMRETGY6znMBo0wtixY0ckJSWhqKgIAQEB2L9/\nP7y9vXHs2DGLLhaJiIiI6CmewxgUFIRvvvkGaWlpOH/+PNavX4+goCBjxkZUKfbKyYl5kRN7suTE\nvMiJedHOoLukAeDBgwfYuHEjfv75ZwBAixYtMGLECI3H7BARERGR5TGoh/HkyZPo27cvHjx4gKCg\nIAghkJaWhlq1amHXrl0WfVmaPYxEVYs9jERExmPSHsaJEyeiU6dOuHr1Kg4dOoTDhw/jypUr6NKl\nCyZNmlTlQRERERFVF85ap59BBWNaWhpmz56N2rVrq9fVrl0bs2bNwrlz54wWHFFl2CsnJ+ZFTuzJ\nkhPzIhfOJa2fQQVjs2bN1PNHP+769eto1ozTgRERERFZMq09jHfu3FF/f/ToUUyfPh2zZs1Chw4d\n1Ouio6Mxf/589O3bt3qiNQH2MBJVLfYwEpFsjNX3ZwrGei9a75KuV69ehXUjR46ssG7AgAEoLS2t\n2qiIiIiISBpaC8YDBw5UZxxEBruQcgHNQtkKIRvmRU7x8fHo1q2bqcOgJzAvcmJetNNaMPIHRkRE\nRNaAc0nrZ9BzGAHg0aNHSEtLw82bN1FWVqaxLSwszCjByUBRFHy84GNTh0FkMdyd3TFt8jRTh0FE\nZJGM1cNoUMF44MABjBw5Ejk5OZVuf7KAtCSW1AhLREREls2kD+6eMmUKXnnlFWRkZKCwsBD379/X\n+CKqTnxOlpyYFzkxL3JiXuTEvGhn0FzS165dw8cffwxfX19jx0NEREREkjHokvTQoUPRv39/vPHG\nG9URk1R4SZqIiIjMhUl7GPPy8vD666+jefPmCAoKgo2Njcb20aNHV3lgsmDBSEREZNkiIyMtZj5p\nkxaMW7ZsQXh4OB4+fAgHBwcoiqKxPT8/v8oDkwULRvnwOVlyYl7kxLzIiXmRS/nvekvIi0lvepk+\nfTreeust5Ofno6CgAPn5+RpfRERERGS5DBphdHZ2xqlTpxAQEFAdMUmFI4xERESWzZJ+15t0hHHQ\noEGIi4ur8pMTERERkfwMeqxOQEAAIiIicPjwYbRu3brCTS9/+ctfjBIcUWUsocfEEjEvcmJe5MS8\nyIl50c6ggnH16tVwcnLCkSNHkJSUVGE7C0YiIiIyV5xLWj+D55K2VpbU10BERESWzaQ9jERERERk\nvQy6JD116tQKz1583JIlS6osIBlFfBZh6hDoMZcvXYZvAKeplA3zIqfqyIu7szumTZ5m1HNYGvbK\nyYl50c6ggjE1NVWjYCwqKkJ6ejpKS0sRHBxstOBk4ftn/hKUycM6D+EbypzIhnmRU3Xk5fL+y0Y9\nPhGZnkEFY3x8fIV1Dx8+xLhx49ClS5eqjolIp2ahzUwdAlWCeZET8yInjmLJiXnR7pl7GO3s7BAR\nEYHo6OiqjIeIiIioWlnKPNLG9IduesnNzeXUgFTtLqRcMHUIVAnmRU7Mi5wqu3JHphMVFQWAedHF\noEvSCxcu1OhhFELg2rVr2LBhA8LCwowWHBERERGZnkHPYWzUqJFGwahSqeDm5oYePXpg5syZcHJy\nMmqQpqQoClakrDB1GERE0rq8/zKiZ7A9icyXJT1z2VjvxaARxszMzCo/MRERERGZBz64m8wOe7Lk\nxLzIiXmRE3vl5MS8aGfQCKMQAps3b8b+/ftx8+ZNlJWVqbcpioIdO3YYLUAiIiIiY+Jc0voZ1MP4\nwQcf4PPPP0f37t3h6emp0c+oKArWrFlj1CBNiT2MRES6sYeRSB4m7WH85ptvsHHjRgwZMqTKAyAi\nIiIiuRnUw1hWVmYVUwCSeWBPlpyYFzkxL3Jir5ycmBftDCoYJ0yYgPXr1xs7FiIiIiKSkEGXpO/e\nvYsNGzYgLi4OrVu3ho2NDYDfb4ZRFAVLliwxapBEj+PcuHJiXuTEvMiJcxbLiXnRzqCCMS0tDW3b\ntgUApKenq9eXF4xERERE5ioyMpLzSeth0CXp+Ph49dfBgwfVX+XLRNWJPVlyYl7kJHteli1bhtDQ\nUNjZ2WHs2LHq9ZmZmVCpVHByclJ/RUf/707sRYsWISAgAM7OzvDw8MDYsWORn5+v9Tz379/HW2+9\nBTc3N7i4uKBr164a20+ePIkuXbrAyckJ9evXV185KykpwfDhw+Hq6oo+ffponGPu3LlYtGjRM71v\n9srJhXNJ68cHdxMRkcl4e3vjk08+wbhx4yrdfu/ePeTn5yM/Px8RERHq9QMGDEBKSgru3buH9PR0\nZGVlaRSUT5o4cSL++9//Ij09HXl5efj888/V23Jzc9GnTx9MmTIFd+7cwaVLl9CrVy8AQGxsLGrU\nqIHbt2+jTp06WLlyJQAgIyMDO3fuxLRp06rix0AkPYMuSRPJhD1ZcmJe5CR7XgYOHAgASElJwdWr\nVytsLysrQ40aNSqs9/f319hHpVLB09Oz0nOkp6dj586dyM7OhqOjIwBoPPkjJiYGvXv3xuuvvw4A\nsLGxQfPmzQH8PtLZtWtXqFQqdOvWDampqQCAd955BzExMVCpnm3chb1ycmJetOMIIxERmZy2Bw37\n+vrCx8cH48aNw+3btzW2bdy4EXXq1IGbmxvc3Ny0jvYlJyfD19cXs2bNgpubG1q3bo3Y2Fj19uPH\nj8PV1RUvvvgiPDw80L9/f1y5cgUAEBgYiAMHDuDRo0c4ePAgAgMDsW3bNri7u6NDhw5V9O6J5MeC\nkcyO7D1Z1op5kZO55OXJGyjd3NyQkpKCrKwsnDhxAvn5+Rg5cqTGPiNGjMDdu3dx8eJF/Pzzz1r7\nCa9evYpz587BxcUF169fx7JlyzBmzBhcuPD7z+bKlStYt24dlixZgqysLPj5+alHG8PCwuDn54f2\n7dvD1dUVw4YNw6efforPPvsMERER6Nq1K95++20UFxc/1ftlr5ycmBftrK5g/O677/Dqq6+iYcOG\ncHBwQPPmzfHxxx+joKDA1KEREVmtJ0cYa9eujXbt2kGlUsHd3R3Lli3Dvn37UFhYWOG1jRs3xkcf\nfYRvvvmm0mPb29vDxsYGf/3rX1GzZk106dIF3bt3x969ewEADg4OGDRoEEJCQlCrVi3Mnj0bSUlJ\n6htc5s2bhzNnzmD58uWYN28epkyZguPHj+PEiRNISEhAUVERvv766yr+iVB14lzS+lldwbhw4ULY\n2Nhg/vz52LNnD6ZMmYIvv/wSPXv2NMrci1T1ZO/JslbMi5zMJS+GPqKtrKys0vXFxcVwcHCodFvr\n1q0BVCxKy89Zvl2f1NRUHD16FBMmTEBqaipCQkIAAKGhoTh79qxBxyjHXjm5lD9Sh3nRzuoKxl27\nduHf//43RowYgS5dumDatGlYsmQJjh8/zqFoIqJqVlpaiocPH6KkpASlpaV49OgRSkpKkJycjAsX\nLqCsrAy3b9/GO++8g+7du8PJyQkA8NVXX+HWrVsAgPPnz2P+/PkYPHhwpefo2rUrGjZsiHnz5qGk\npARHjhxBfHw8Xn75ZQDA2LFjsW3bNpw5cwbFxcWYM2cOOnfurD4X8HuxOXXqVCxduhSKosDf3x+J\niYkoKipCQkICAgICjPyTIjItqysY69atW2FdaGgoAODatWvVHQ49A3PpybI2zIucZM/LnDlz4ODg\ngAULFmD9+vWwt7fH3Llz8dtvv6FPnz5wdnZGUFAQ7O3tsWnTJvXrkpKSEBQUBCcnJwwcOBCjR4/G\ne++9p94eGBio3r9mzZrYvn07du/eDRcXF0yaNAn/+te/0LRpUwBA9+7dMXfuXLzyyivw8PDAb7/9\nho0bN2rEuXbtWgQFBanvrh40aBC8vLzg7u6OvLw8TJw48aneNwco5MS8aKcIXofF8uXL8dZbbyEl\nJQXt2rXT2KYoClakrDBRZFSZCykXzOYymzVhXuRUHXm5vP8yomdofwYiVRQfH8/LnxKyhLwoimKU\nFjurLxizs7MRHByM4OBgdQP041gwEhHpxoKRSB7GKhit+sHdBQUFGDBgAGxtbbFmzRqt+62ZvQb1\nvOoBAOwd7eHTzEf9F3v55R4uc5nLXLbWZTvYAfjf5bzyERouc9lcliMjI9XrZYjnaZbLv8/MzIQx\nWe0I44MHDxAWFobU1FQkJCSgVatWle7HEUb58NKnnJgXOfGStJws4dKnJSkflbOEvHCEsQoVFxfj\ntddew8mTJxEXF6e1WCQiIiIiKywYy8rKMHLkSMTHx2PXrl1o3769qUOip8RRLDkxL3JiXuRk7qNY\nlop50c7qCsa3334b3333HSIiImBvb49jx46pt/n4+MDb29uE0RERERHJx+qew7hnzx4oioLo6Gh0\n7NhR42v16tWmDo8MIPtz5awV8yIn5kVOfN6fnJgX7axuhDEjI8PUIRAREZFEOJe0flZ7l7SheJc0\nEZFuvEuaSB7Gukva6i5JExEREdHTYcFIZoc9WXJiXuTEvMiJvXJyYl60Y8FIRERERDqxh1EP9jAS\nEenGHkYiebCHkYiIiMgIIiMjTR2C9FgwktlhT5acmBc5MS9yYq+cXKKiogAwL7qwYCQiIiIindjD\nqAd7GImIdGMPI5k7Y/X9mQJ7GImIiIjIJFgwktlhT5acmBc5MS9yYq+cnJgX7VgwEhERkVXjXNL6\nsYdRD/YwEhHpxh5GInmwh5GIiIiITIIFI5kd9mTJiXmRE/MiJ/bKyYl50Y4FIxERERHpxB5GPdjD\nSESkG3sYieTBHkYiIiIiI+Bc0vqxYCSzw54sOTEvcmJe5MReOblwLmn9WDASERERkU7sYdSDPYxE\nRLqxh5HMHeeS1q9mlR/RAl3ef9nUIRARScvd2d3UIRCRkXGEUQ9L+qvDUsTHx6Nbt26mDoOewLzI\niXmRE/Mil/Lf9ZaQF94lTURERGQEnEtaP44w6sERRiIiIjIXHGEkIiIiIpNgwUhmh8/JkhPzIifm\nRU7Mi5yYF+1YMBIRERGRTuxh1IM9jERERGQu2MNIREREZAScS1o/FoxkdthjIifmRU7Mi5yYF7lw\nLmn9WDASERERkU7sYdSDPYxERESWzZJ+17OHkYiIiIhMggUjmR32mMiJeZET8yIn5kVOzIt2NU0d\ngDmI+CzC1CHQYy5fuoy45DhTh0FPYF7kxLzIiXmRy8uvvGzqEKTHHkY9FEXBipQVpg6DiIiIjOTy\n/suInhFt6jCqBHsYiYiIiMgkWDCS2bmQcsHUIVAlmBc5MS9yYl7kxB5G7VgwEhEREZFOLBjJ7DQL\nbWbqEKgSzIucmBc5MS9y6tatm6lDkBYLRiIiIrJqh+IOmToE6bFgJLPD3h85MS9yYl7kxLzIJfHH\nRADsYdSFBSMRERER6cTnMOrB5zASERFZtkmhkziXtB4cYSQiIiIinVgwktlh74+cmBc5MS9yYl7k\nxB5G7VgwEhERkVXr9FInU4cgPfYw6sEeRiIiIsvGuaT14wgjEREREenEgpHMDnt/5MS8yIl5kRPz\nIif2MGrHgpGIiIjoGS1btgyhoaGws7PD2LFjNbbt378fzZs3R+3atdGjRw9kZWVpbP/www9Rr149\n1KtXDx999JHO8xh6LAAaxyopKcHw4cPh6uqKPn36ID8/X71t7ty5WLRokUHvkwUjmR3OwSon5kVO\nzIucmBc5Pctc0t7e3vjkk08wbtw4jfW5ubkYPHgwoqOjkZeXh9DQUAwbNky9fcWKFdi+fTvOnj2L\ns2fPYufOnVixovJ7Jp7mWAA0jhUbG4saNWrg9u3bqFOnDlauXAkAyMjIwM6dOzFt2jSD3icLRiIi\nIrJqf2Qu6YEDB2LAgAGoW7euxvrY2FgEBgZi8ODBsLW1RWRkJM6cOYOLFy8CANatW4fp06fDy8sL\nXl5emD59OtauXVvpOZ7mWAA0jpWZmYmuXbtCpVKhW7du+O233wAA77zzDmJiYqBSGVYKsmAks8Pe\nHzkxL3JiXuTEvMilKuaSfvLO5LS0NLRp00a97ODggMaNGyMtLQ0AcP78eY3trVu3Vm970h85VmBg\nIA4cOIBHjx7h4MGDCAwMxLZt2+Du7o4OHToY/P4sqmAMDw+Hn5/fU78uPj4eKpUKhw49+18YRERE\nZL0URdFYLiwshLOzs8Y6Z2dndQ9hQUEB6tSpo7GtoKCg0mP/kWOFhYXBz88P7du3h6urK4YNG4ZP\nP/0Un332GSIiItC1a1e8/fbbKC4u1vn+LKpgnDVrFr7//ntTh0FGxt4fOTEvcmJe5MS8yOlZehjL\nPTnC6OjoiHv37mmsu3v3LpycnCrdfvfuXTg6OlZ67D96rHnz5uHMmTNYvnw55s2bhylTpuD48eM4\nceIEEhISUFRUhK+//lrn+7OIgvHRo0cAAH9/f40hWSIiIqLq8OQIY6tWrXDmzBn1cmFhIS5duoRW\nrVqpt58+fVq9/cyZMwgMDKz02FV1rNTUVBw9ehQTJkxAamoqQkJCAAChoaHqG2a0qdaC8eLFixg4\ncCA8PDxgb28PX19fDB06FKWlpQCAW7duYfLkyWjQoAHs7OzQokULrFq1SuMYa9euhUqlwuHDhzFk\nyBC4urqqr8FXdkl69uzZaNeuHerUqQM3Nzf8+c9/xvHjx6vnDZNRsPdHTsyLnJgXOTEvcnqWHsbS\n0lI8fPgQJSUlKC0txaNHj1BaWoqBAwfi3LlziI2NxcOHDxEVFYW2bduiadOmAIDRo0cjJiYG165d\nQ3Z2NmJiYhAeHl7pOZ7mWAAqPZYQAlOnTsXSpUuhKAr8/f2RmJiIoqIiJCQkICAgQOf7rNaC8ZVX\nXsH169exfPly7Nu3D/Pnz4ednR2EELh37x46deqEPXv2ICoqCrt370a/fv0wZcoULFu2rMKxRo4c\niYCAAGzduhXz589Xr3+yws/Ozsa7776LHTt2YN26dXB3d0eXLl1w7tw5o79fIiIikt8fmUt6zpw5\ncHBwwIIFC7B+/XrY29sjOjoa9erVw9atWxEREYHnnnsOKSkp+Pbbb9WvmzRpEvr164egoCC0bt0a\n/fr1w8SJE9XbAwMDsWnTJgB4qmMBqHAs4PcBt6CgIAQHBwMABg0aBC8vL7i7uyMvL6/C/k+qtrmk\nc3Nz4e7ujh07dqBv374Vts+ZMwdz587FuXPnNKrciRMnYtu2bcjJyYFKpcLatWsxbtw4vPfee1i4\ncKHGMcLDw5GQkICMjIxKYygtLYUQAoGBgejduzc+//xzAL//RdGjRw/Ex8ejS5cuGq/hXNJERESW\njXNJ61dtI4z16tWDv78/PvzwQ3z11Vf45ZdfNLbv2bMHL7zwAho1aoSSkhL1V69evXD79m2cP39e\nY/+BAwcadN4ff/wR3bt3R7169WBjYwNbW1tcvHhR/ewiIiIiItKtZnWeLC4uDpGRkZg5cyZu374N\nPz8/fPDBB5g8eTJu3ryJS5cuwcbGpsLrFEXB7du3NdZ5enrqPd/JkycRFhaGPn364Ouvv4anpydU\nKhXefPNNPHz40OC418xeg3pev0+3Y+9oD59mPuo73Mr7ULhcfctXLlzBSyNfkiYeLv++/HhPgnjy\nNAAAFuBJREFUlgzxcJmfF5mX+XmRbzk+Ph6nT5/Gu+++q14G/nfntKzL5d9nZmbCmKrtkvSTzpw5\ng2XLlmH16tXYvXs3oqKiULNmTSxevLjS/Zs2bQpHR0f1Jelff/0V/v7+Gvs8eUk6IiICixcvxt27\nd1GjRg31fr6+vggICMCBAwcA8JK0ubmQcoGPpJAQ8yIn5kVOzItcyi9Jx8fH/6FH68jAWJekq3WE\n8XFt2rTBwoULsXr1aqSlpaF3795YunQpfHx84ObmViXnuH//foUpbw4cOIArV67ovRuI5MX/ycqJ\neZET8yIn5kVO5l4sGlO1FYxnz57FtGnTMHz4cAQEBKC0tBRr166FjY0NevTogYCAAGzevBmdO3fG\ne++9h6ZNm6KwsBDp6elITEx8pgdy9+nTB4sXL0Z4eDjCw8Nx8eJF/O1vf4O3t7dRqm8iIiIyP4fi\nDgEzTB2F3KrtphdPT0/4+voiJiYGAwYMwIgRI3Djxg3s2rULwcHBcHZ2RlJSEsLCwrBgwQL07t0b\n48ePx86dO9GjRw+NYz356JzH1z++rVevXliyZAmOHDmCfv36Ye3atfjXv/6Fxo0bVziGtmOSfB7v\n/SF5MC9yYl7kxLzIpSrmkrZ0JuthNBfsYZQPe3/kxLzIiXmRE/Mil0mhkyCEYA+jruOyYNSNBSMR\nEZFlKy8YLYHZP4eRiIiIiMwTC0YyO+z9kRPzIifmRU7Mi5zYw6gdC0YiIiKyan9kLmlrwR5GPdjD\nSEREZNk4l7R+HGEkIiIiIp1YMJLZYe+PnJgXOTEvcmJe5MQeRu1YMBIRERGRTiwYyezwYbdyYl7k\nxLzIiXmRk7k/tNuYWDASERGRVTsUd8jUIUiPBSOZHfb+yIl5kRPzIifmRS6cS1o/FoxEREREpBOf\nw6gHn8NIRERk2TiXtH4cYSQiIiIinVgwktlh74+cmBc5MS9yYl7kxB5G7VgwEhERkVXjXNL6sYdR\nD/YwEhERWTbOJa0fRxiJiIiISCcWjGR22PsjJ+ZFTsyLnJgXObGHUTsWjERERESkEwtGMjucg1VO\nzIucmBc5MS9y4lzS2rFgJCIiIqvGuaT1q2nqAMzB5f2XTR0CPebypcvwDfA1dRj0BOZFTsyLnJgX\nuTw+lzRHGSvHx+roYazb0+nZ8QMtJ+ZFTsyLnJgXuZT/rreEvBirbmHBqAcLRiIiIstmSb/r+RxG\nIiIiIjIJFoxkdvicLDkxL3JiXuTEvMiJedGOBSMRERFZtdmzZ5s6BOmxh1EPS+prICIiIsvGHkYi\nIiIiMgkWjGR22GMiJ+ZFTsyLnJgXOTEv2rFgJCIiIiKd2MOoB3sYiYiIyFywh5GIiIjICCIjI00d\ngvRYMJLZYY+JnJgXOTEvcmJe5BIVFQWAedGFBSMRERER6cQeRj3Yw0hERGTZLOl3PXsYiYiIiMgk\nWDCS2WGPiZyYFzkxL3JiXuTEvGjHgpGIiIisGueS1o89jHpYUl8DERERWTb2MBIRERGRSbBgJLPD\nHhM5MS9yYl7kxLzIiXnRjgUjEREREenEHkY92MNIRERE5oI9jERERERGwLmk9WPBSGaHPSZyYl7k\nxLzIiXmRC+eS1o8FIxERERHpxB5GPdjDSEREZNks6Xc9exiJiIiIyCRYMJLZYY+JnJgXOTEvcmJe\n5MS8aMeCkYiIiKwa55LWjz2MelhSXwMRERFZNvYwEhEREZFJsGAks8MeEzkxL3JiXuTEvMiJedGO\nBSMRERER6cQeRj3Yw0hERETmgj2MREREREbAuaT1Y8FIZoc9JnJiXuTEvMiJeZEL55LWjwUjmZ3T\np0+bOgSqBPMiJ+ZFTsyLnJgX7Vgwktn573//a+oQqBLMi5yYFzkxL3JiXrRjwUhEREREOrFgJLOT\nmZlp6hCoEsyLnJgXOTEvcmJetONjdfRo27Ytzpw5Y+owiIiIiPTq2rWrUW7eYcFIRERERDrxkjQR\nERER6cSCkYiIiIh0YsFIRERERDqxYNRiz549aN68OZo0aYIFCxaYOhz6/xo1aoTWrVsjODgY7du3\nN3U4VmvcuHHw8PBAUFCQet2dO3fQs2dPNG3aFL169eLzzEygsrxERkaiQYMGCA4ORnBwMPbs2WPC\nCK3PlStX0L17d7Rq1QqBgYFYsmQJAH5eTE1bXvh50Y43vVSitLQUzZo1w48//ghvb288//zz2LRp\nE1q0aGHq0Kyen58fTpw4geeee87UoVi1w4cPw9HREaNHj0ZqaioAYMaMGahXrx5mzJiBBQsWIC8v\nD/PnzzdxpNalsrxERUXByckJf/nLX0wcnXW6ceMGbty4gbZt26KgoAAhISH4/vvvsWbNGn5eTEhb\nXrZs2cLPixYcYaxEcnIyGjdujEaNGsHGxgbDhw/H9u3bTR0W/X/8G8f0OnfuDFdXV411O3bswJgx\nYwAAY8aMwffff2+K0KxaZXkB+Jkxpfr166Nt27YAAEdHR7Ro0QLZ2dn8vJiYtrwA/Lxow4KxEtnZ\n2fDx8VEvN2jQQP0PiUxLURS89NJLCA0NxapVq0wdDj0mJycHHh4eAAAPDw/k5OSYOCIqt3TpUrRp\n0wbjx4/npU8TyszMxKlTp/CnP/2JnxeJlOflhRdeAMDPizYsGCuhKIqpQyAtjhw5glOnTuGHH37A\nF198gcOHD5s6JKqEoij8HEliypQpyMjIwOnTp+Hp6Yn333/f1CFZpYKCAgwePBiLFy+Gk5OTxjZ+\nXkynoKAAr732GhYvXgxHR0d+XnRgwVgJb29vXLlyRb185coVNGjQwIQRUTlPT08AgJubGwYOHIjk\n5GQTR0TlPDw8cOPGDQDA9evX4e7ubuKICADc3d3VBcmbb77Jz4wJFBcXY/DgwRg1ahReffVVAPy8\nyKA8L2+88YY6L/y8aMeCsRKhoaH45ZdfkJmZiaKiImzevBn9+/c3dVhW7/79+8jPzwcAFBYWYt++\nfRp3g5Jp9e/fH+vWrQMArFu3Tv0/YDKt69evq7/ftm0bPzPVTAiB8ePHo2XLlnj33XfV6/l5MS1t\neeHnRTveJa3FDz/8gHfffRelpaUYP348Zs6caeqQrF5GRgYGDhwIACgpKcHIkSOZFxN5/fXXkZCQ\ngNzcXHh4eODTTz/FgAEDMHToUGRlZaFRo0bYsmULXFxcTB2qVXkyL1FRUYiPj8fp06ehKAr8/Pyw\nYsUKde8cGV9iYiK6dOmC1q1bqy87z5s3D+3bt+fnxYQqy8vcuXOxadMmfl60YMFIRERERDrxkjQR\nERER6cSCkYiIiIh0YsFIRERERDqxYCQiIiIinVgwEhEREZFOLBiJiIiISCcWjERWKjMzEyqVCidP\nnqz2c69du7bC9GjWIjc3FyqVCocOHXrmY2zfvh1NmjSBjY0Nxo0bV4XRERFVjgUjkRXo1q0bpk6d\nqrGuYcOGuHHjBtq0aVPt8QwfPhwZGRnVfl5LMX78eAwZMgRZWVlYvHixqcPRa+XKlejevTtcXFyg\nUqmQlZVVYZ+8vDyMGjUKLi4ucHFxwejRo3H37l2NfbKystCvXz84OjrCzc0N06ZNQ3FxscY+qamp\n6Nq1KxwcHNCgQQPMmTOnwrkSEhIQEhICe3t7BAQEYMWKFVX7hoksEAtGIiulUqng7u6OGjVqVPu5\n7ezsUK9evWo/ryXIy8vDnTt30KtXL3h6ej7zSG1RUVEVR6bdgwcP0Lt3b0RFRWndZ8SIETh9+jT2\n7t2LPXv24OTJkxg1apR6e2lpKV555RUUFhYiMTERmzZtwnfffYf3339fvc+9e/fQs2dPeHp6IiUl\nBYsXL8bf//53xMTEqPfJyMhAWFgYOnXqhNOnT2PmzJmYOnUqYmNjjfPmiSyFICKLNmbMGKEoisbX\n5cuXRUZGhlAURZw4cUIIIcTBgweFoijihx9+EMHBwcLe3l507txZXL16Vezfv18EBQUJR0dH0a9f\nP3Hnzh2Nc3z99deiRYsWws7OTjRt2lQsWrRIlJWVaY1pzZo1wtHRUb08e/ZsERgYKDZt2iT8/f2F\nk5OTePXVV0Vubq7O9xYVFSV8fX1FrVq1RP369cXo0aM1ti9YsEAEBAQIe3t7ERQUJNavX6+xPTs7\nW4wYMULUrVtXODg4iLZt24qDBw+qty9fvlwEBAQIW1tb0bhxY7Fq1SqN1yuKIlauXClee+01Ubt2\nbeHv71/hHMnJyaJdu3bCzs5OBAcHi127dglFUURCQoIQQoiioiIxdepU4eXlJWrVqiV8fHzERx99\nVOn7Lc/R41/lx9m6dasIDAxUHyM6Olrjtb6+viIyMlKMHTtWuLi4iKFDh1Z6jjFjxoi+ffuKzz//\nXHh7ewtXV1cxduxYcf/+fS1ZMNxPP/2k/vf3uPPnzwtFUURSUpJ6XWJiolAURVy8eFEIIcTu3buF\nSqUSV69eVe+zfv16YWdnJ/Lz84UQQvzzn/8UderUEQ8fPlTv87e//U14e3url2fMmCGaNm2qcf43\n33xTdOjQ4Q+/PyJLxoKRyMLdvXtXdOzYUYwfP17k5OSInJwcUVpaqrVg/NOf/iQSExPF2bNnRWBg\noOjYsaPo3r27SE5OFikpKcLPz09MmzZNffyVK1cKT09PsXXrVpGZmSl27twp6tevL5YtW6Y1psoK\nRkdHRzFo0CCRmpoqjh49Knx9fcWkSZO0HuO7774Tzs7OYvfu3eLKlSsiJSVFfPHFF+rtH3/8sWje\nvLnYu3evyMzMFBs3bhS1a9cW//nPf4QQQhQUFIjGjRuLTp06icTERJGRkSG2b9+uLhhjY2OFjY2N\n+OKLL8Qvv/wili5dKmxsbMTOnTvV51AURTRo0EBs2LBBXLp0ScycOVPY2tqKrKwsIYQQ+fn5ws3N\nTQwdOlSkpaWJvXv3iubNm2sUev/4xz+Ej4+POHz4sLhy5YpISkoSa9eurfQ9FxUVqYurbdu2iZyc\nHFFUVCRSUlJEjRo1RGRkpPjll1/Ehg0bhKOjo1i6dKn6tb6+vsLZ2Vn8/e9/F5cuXRK//vprpecY\nM2aMqFOnjpg4caJIT08X+/btEy4uLmLevHnqfaKjo4Wjo6POr8TExArH1lYwrl69Wjg5OWmsKysr\nE46OjuqfxSeffCICAwM19rl586ZQFEXEx8cLIYQYNWqU6Nu3r8Y+ycnJQlEUkZmZKYQQonPnzuL/\n/u//NPbZsmWLsLGxESUlJZX+TIiIBSORVejWrZuYOnWqxjptBeO+ffvU+yxbtkwoiiJOnTqlXhcZ\nGanxi9vHx6fCqNqiRYtEy5YttcZTWcFoZ2cn7t27p14XHR0tGjdurPUYCxcuFM2aNRPFxcUVthUU\nFAh7e/sKRcu0adNEWFiYEOL3QtfJyUncvn270uOXF9mPCw8PF506dVIvK4oiPv74Y/VySUmJcHBw\nEBs2bBBCCLFixQrh4uIiCgsL1fusX79eo2B85513xJ///Get7/NJt27d0ni9EEKMGDGiwjEiIyNF\ngwYN1Mu+vr6if//+eo8/ZswY0bBhQ40R4gkTJoiXXnpJvXznzh1x6dIlnV8PHjyocGxtBWN0dLTw\n9/evsL+/v7+YP3++OoYn32NZWZmoWbOm+Pbbb4UQQvTs2bNCzi5fviwURRHHjh0TQgjRtGlTMWfO\nHI19EhIShKIo4saNG3p/PkTWqqapL4kTkVxat26t/t7d3R0AEBQUpLHu5s2bAIBbt27h6tWrmDhx\nIiZPnqzep6Sk5KnP6+vrq9GP5+npqT5PZYYOHYolS5bAz88PL7/8Mnr37o3+/fvD1tYW58+fx8OH\nD/Hyyy9DURT1a4qLi+Hn5wcAOHXqFNq0aYPnnnuu0uOnp6fjzTff1Fj34osvYseOHRrrHv951ahR\nA25ubuq4f/75Z7Rp0wYODg7qfV544QWN14eHh6Nnz55o2rQpevXqhbCwMPTp00cjbn3S09PRt2/f\nCrFGRUWhoKAAjo6OUBQFoaGhBh2vZcuWGuf39PTE8ePH1cuurq5wdXU1OL6qIoTQuf1pfmZE9HRY\nMBKRBhsbG/X35b+AH78xRlEUlJWVAYD6vytWrEDHjh2r7LxPnqcyDRo0wIULF7B//378+OOPeP/9\n9xEVFYXjx4+rX7dr1y40bNhQ63n0FSCVebIo0Re3vnMEBwcjMzMTe/fuxf79+zFmzBi0adMGcXFx\nT1UAaTvP48eoXbu2QceqWVPzV8OT72nu3LmYN2+ezmPs2bMHL774okHnq1+/Pm7duqWxTgiBmzdv\non79+up9kpKSNPbJzc1FaWmpxj43btzQ2CcnJ0e9Tdc+NWvW5I1YRDrwLmkiK2Bra/tMo376eHh4\nwMvLC7/++iv8/f0rfBlbrVq1EBYWhpiYGPz0009IS0tDUlISWrVqhVq1aiEzM7NCTD4+PgCAdu3a\n4ezZs7h9+3alx27RogUSExM11iUmJqJVq1YGx9eyZUukpqbi/v376nXHjh2rsJ+joyMGDx6Mf/7z\nn/jPf/6DAwcO4NKlSwafp0WLFjhy5EiFWH18fAwuEh+nr1CdMmUKzpw5o/MrJCTE4PN16NABBQUF\nOHr0qHrd0aNHUVhYqP5DpGPHjvj555+RnZ2t3icuLg61atVSn6tDhw44fPgwHj16pLGPt7c3fH19\n1fvExcVpnD8uLg7PP/+8SZ4YQGQuOMJIZAUaNWqE5ORkXL58GbVr10bdunWr7NhRUVGYOnUqXFxc\n0KdPHxQXF+PkyZO4du0aPvrooyo7z5PWrl2L0tJStG/fHo6Ojti8eTNsbW3RpEkTODo6Yvr06Zg+\nfTqEEOjcuTMKCgpw7Ngx1KhRAxMmTMCIESMwf/58DBgwAPPnz4eXlxfOnTsHZ2dndOvWDR988AGG\nDBmCkJAQ9OzZE3v27MHGjRuxbds2g2McMWIEIiIiMG7cOMyaNQvZ2dmIjo7W2CcmJgZeXl5o06YN\nbGxssGHDBtSpUwcNGjQw+Dzvv/8+nn/+eURFReH111/HTz/9hJiYGL2jgNroGxV92kvSN27cwI0b\nN3Dx4kUAQFpaGu7cuQNfX1+4urqiRYsW6N27NyZNmoSVK1dCCIFJkyahX79+aNKkCQCgV69eaNWq\nFUaPHo2FCxciNzcXM2bMwMSJE+Ho6Ajg9593VFQUwsPD8de//hUXLlzAggULEBkZqY5l8uTJWLZs\nGd577z1MnDgRR44cwbp16/Dtt98+5U+JyMqYrn2SiKrLxYsXRYcOHYSDg4NQqVTqx+qoVCqNm15U\nKpXGTSD//ve/hUql0jjW8uXLhZubm8a6TZs2qR8d4+rqKjp37iw2b96sNZ41a9Zo3BUbGRkpgoKC\ndO7zpO+//1506NBBuLi4iNq1a4v27dur74Aut3TpUtGyZUtRq1Yt4ebmJnr16iV+/PFH9farV6+K\nYcOGCRcXF+Hg4CDatWuncTPJ8uXLRePGjYWNjY1o0qSJ+OqrrzSOryiK2Lp1q8a6Ro0aiYULF6qX\njx8/Ltq1aydq1aol2rZtK3bu3ClUKpX6PKtWrRLt2rUTTk5OwtnZWXTr1k0cPXpU6/u+deuWxuvL\nxcbGiqCgIGFraysaNmwo5s6dqzMubcLDw0W/fv001lWWn6cxe/Zs9WOAVCqV+r/r1q1T75OXlyfe\neOMN4ezsLJydncWoUaPE3bt3NY6TlZUl+vbtKxwcHETdunXFtGnTRFFRkcY+qampokuXLsLOzk54\neXmJTz/9tEI8CQkJ6pz4+/uLFStWPPN7I7IWihDP0MRDRERERFaDPYxEREREpBMLRiIiIiLSiQUj\nEREREenEgpGIiIiIdGLBSEREREQ6sWAkIiIiIp1YMBIRERGRTiwYiYiIiEin/wdr/eoLv1plMAAA\nAABJRU5ErkJggg==\n", + "text/plain": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "# Conclusion" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#Sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "We can see that we could speed up the density estimations for our Parzen-window function if we submitted them in parallel. However, on my particular machine, the submission of 6 parallel 6 processes doesn't lead to a further performance improvement, which makes sense for a 4-core CPU. \n", + "We also notice that there was a significant performance increase when we were using 3 instead of only 2 processes in parallel. However, the performance increase was less significant when we moved up to 4 parallel processes, respectively. \n", + "This can be attributed to the fact that in this case, the CPU consists of only 4 cores, and system processes, such as the operating system, are also running in the background. Thus, the fourth core simply does not have enough capacity left to further increase the performance of the fourth process to a large extend. And we also have to keep in mind that every additional process comes with an additional overhead for inter-process communication. \n", + "\n", + "Also, an improvement due to parallel processing only makes sense if our tasks are \"CPU-bound\" where the majority of the task is spent in the CPU in contrast to I/O bound tasks, i.e., tasks that are processing data from a disk. " + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [] + } + ], + "metadata": { + "kernelspec": { + "display_name": "Python 3", + "language": "python", + "name": "python3" + }, + "language_info": { + "codemirror_mode": { + "name": "ipython", + "version": 3 + }, + "file_extension": ".py", + "mimetype": "text/x-python", + "name": "python", + "nbconvert_exporter": "python", + "pygments_lexer": "ipython3", + "version": "3.6.3" + } + }, + "nbformat": 4, + "nbformat_minor": 1 +} diff --git a/tutorials/not_so_obvious_python_stuff.ipynb b/tutorials/not_so_obvious_python_stuff.ipynb new file mode 100644 index 0000000..cf683b0 --- /dev/null +++ b/tutorials/not_so_obvious_python_stuff.ipynb @@ -0,0 +1,4151 @@ +{ + "cells": [ + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[Sebastian Raschka](http://sebastianraschka.com) \n", + "\n", + "- [Link to this IPython Notebook on GitHub](https://github.com/rasbt/python_reference/blob/master/tutorials/not_so_obvious_python_stuff.ipynb) \n", + "- [Link to the GitHub repository](https://github.com/rasbt/python_reference) \n", + "\n" + ] + }, + { + "cell_type": "code", + "execution_count": 1, + "metadata": {}, + "outputs": [], + "source": [ + "%load_ext watermark" + ] + }, + { + "cell_type": "code", + "execution_count": 2, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "last updated: 2018-06-09 \n", + "\n", + "CPython 3.6.4\n", + "IPython 6.2.1\n" + ] + } + ], + "source": [ + "%watermark -d -u -v" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[More information](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/ipython_magic/watermark.ipynb) about the `watermark` magic command extension.\n", + "\n", + "([Changelog](#changelog))" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "# A collection of not-so-obvious Python stuff you should know!" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "I am really looking forward to your comments and suggestions to improve and \n", + "extend this little collection! Just send me a quick note \n", + "via Twitter: [@rasbt](https://twitter.com/rasbt) \n", + "or Email: [bluewoodtree@gmail.com](mailto:bluewoodtree@gmail.com)\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "# Sections\n", + "- [The C3 class resolution algorithm for multiple class inheritance](#c3_class_res)\n", + "\n", + "- [Assignment operators and lists - simple-add vs. add-AND operators](#pm_in_lists)\n", + "\n", + "- [`True` and `False` in the datetime module](#datetime_module)\n", + "\n", + "- [Python reuses objects for small integers - always use \"==\" for equality, \"is\" for identity](#python_small_int)\n", + "\n", + "- [Shallow vs. deep copies if list contains other structures and objects](#shallow_vs_deep)\n", + "\n", + "- [Picking `True` values from logical `and`s and `or`s](#false_true_expressions)\n", + "\n", + "- [Don't use mutable objects as default arguments for functions!](#def_mutable_func)\n", + "\n", + "- [Be aware of the consuming generator](#consuming_generator)\n", + "\n", + "- [`bool` is a subclass of `int`](#bool_int)\n", + "\n", + "- [About lambda-in-closures and-a-loop pitfall](#lambda_closure)\n", + "\n", + "- [Python's LEGB scope resolution and the keywords `global` and `nonlocal`](#python_legb)\n", + "\n", + "- [When mutable contents of immutable tuples aren't so mutable](#immutable_tuple)\n", + "\n", + "- [List comprehensions are fast, but generators are faster!?](#list_generator)\n", + "\n", + "- [Public vs. private class methods and name mangling](#private_class)\n", + "\n", + "- [The consequences of modifying a list when looping through it](#looping_pitfall)\n", + "\n", + "- [Dynamic binding and typos in variable names](#dynamic_binding)\n", + "\n", + "- [List slicing using indexes that are \"out of range](#out_of_range_slicing)\n", + "\n", + "- [Reusing global variable names and UnboundLocalErrors](#unboundlocalerror)\n", + "\n", + "- [Creating copies of mutable objects](#copy_mutable)\n", + "\n", + "- [Key differences between Python 2 and 3](#python_differences)\n", + "\n", + "- [Function annotations - What are those `->`'s in my Python code?](#function_annotation)\n", + "\n", + "- [Abortive statements in `finally` blocks](#finally_blocks)\n", + "\n", + "- [Assigning types to variables as values](#variable_types)\n", + "\n", + "- [Only the first clause of generators is evaluated immediately](#generator_rhs)\n", + "\n", + "- [Keyword argument unpacking syntax - `*args` and `**kwargs`](#splat_op)\n", + "\n", + "- [Metaclasses - What creates a new instance of a class?](#new_instance)\n", + "\n", + "- [Else-clauses: \"conditional else\" and \"completion else\"](#else_clauses)\n", + "\n", + "- [Interning of compile-time constants vs. run-time expressions](#string_interning)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## The C3 class resolution algorithm for multiple class inheritance" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "If we are dealing with multiple inheritance, according to the newer C3 class resolution algorithm, the following applies: \n", + "Assuming that child class C inherits from two parent classes A and B, \"class A should be checked before class B\".\n", + "\n", + "If you want to learn more, please read the [original blog](http://python-history.blogspot.ru/2010/06/method-resolution-order.html) post by Guido van Rossum.\n", + "\n", + "(Original source: [http://gistroll.com/rolls/21/horizontal_assessments/new](http://gistroll.com/rolls/21/horizontal_assessments/new))" + ] + }, + { + "cell_type": "code", + "execution_count": 3, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "class A\n" + ] + } + ], + "source": [ + "class A(object):\n", + " def foo(self):\n", + " print(\"class A\")\n", + "\n", + "class B(object):\n", + " def foo(self):\n", + " print(\"class B\")\n", + "\n", + "class C(A, B):\n", + " pass\n", + "\n", + "C().foo()" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "So what actually happened above was that class `C` looked in the scope of the parent class `A` for the method `.foo()` first (and found it)!" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "I received an email containing a suggestion which uses a more nested example to illustrate Guido van Rossum's point a little bit better:" + ] + }, + { + "cell_type": "code", + "execution_count": 4, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "class C\n" + ] + } + ], + "source": [ + "class A(object):\n", + " def foo(self):\n", + " print(\"class A\")\n", + "\n", + "class B(A):\n", + " pass\n", + "\n", + "class C(A):\n", + " def foo(self):\n", + " print(\"class C\")\n", + "\n", + "class D(B,C):\n", + " pass\n", + "\n", + "D().foo()" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Here, class `D` searches in `B` first, which in turn inherits from `A` (note that class `C` also inherits from `A`, but has its own `.foo()` method) so that we come up with the search order: `D, B, C, A`. " + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Assignment operators and lists - simple-add vs. add-AND operators" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Python `list`s are mutable objects as we all know. So, if we are using the `+=` operator on `list`s, we extend the `list` by directly modifying the object. \n", + "\n", + "However, if we use the assignment via `my_list = my_list + ...`, we create a new list object, which can be demonstrated by the following code:" + ] + }, + { + "cell_type": "code", + "execution_count": 5, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "ID: 4486856904\n", + "ID (+=): 4486856904\n", + "ID (list = list + ...): 4486959368\n" + ] + } + ], + "source": [ + "a_list = []\n", + "print('ID:', id(a_list))\n", + "\n", + "a_list += [1]\n", + "print('ID (+=):', id(a_list))\n", + "\n", + "a_list = a_list + [2]\n", + "print('ID (list = list + ...):', id(a_list))" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Just for reference, the `.append()` and `.extends()` methods are modifying the `list` object in place, just as expected." + ] + }, + { + "cell_type": "code", + "execution_count": 6, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "[] \n", + "ID (initial): 4486857224 \n", + "\n", + "[1] \n", + "ID (append): 4486857224 \n", + "\n", + "[1, 2] \n", + "ID (extend): 4486857224\n" + ] + } + ], + "source": [ + "a_list = []\n", + "print(a_list, '\\nID (initial):',id(a_list), '\\n')\n", + "\n", + "a_list.append(1)\n", + "print(a_list, '\\nID (append):',id(a_list), '\\n')\n", + "\n", + "a_list.extend([2])\n", + "print(a_list, '\\nID (extend):',id(a_list))" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
\n", + "" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "## `True` and `False` in the datetime module\n", + "\n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\"It often comes as a big surprise for programmers to find (sometimes by way of a hard-to-reproduce bug) that, unlike any other time value, midnight (i.e. `datetime.time(0,0,0)`) is False. A long discussion on the python-ideas mailing list shows that, while surprising, that behavior is desirable — at least in some quarters.\" \n", + "\n", + "Please note that Python version <= 3.4.5 evaluated the first statement `bool(datetime.time(0,0,0))` as `False`, which was regarded counter-intuitive, since \"12am\" refers to \"midnight.\"\n", + "\n", + "(Original source: [http://lwn.net/SubscriberLink/590299/bf73fe823974acea/](http://lwn.net/SubscriberLink/590299/bf73fe823974acea/))" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "code", + "execution_count": 3, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Current python version: 3.6.4\n", + "\"datetime.time(0,0,0)\" (Midnight) -> True\n", + "\"datetime.time(1,0,0)\" (1 am) -> True\n" + ] + } + ], + "source": [ + "from platform import python_version\n", + "import datetime\n", + "\n", + "print(\"Current python version: \", python_version())\n", + "print('\"datetime.time(0,0,0)\" (Midnight) ->', bool(datetime.time(0,0,0))) # Python version <= 3.4.5 evaluates this statement to False\n", + "\n", + "print('\"datetime.time(1,0,0)\" (1 am) ->', bool(datetime.time(1,0,0)))" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
\n", + "" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "## Python reuses objects for small integers - use \"==\" for equality, \"is\" for identity\n", + "\n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "This oddity occurs, because Python keeps an array of small integer objects (i.e., integers between -5 and 256, [see the doc](https://docs.python.org/2/c-api/int.html#PyInt_FromLong))." + ] + }, + { + "cell_type": "code", + "execution_count": 8, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "a is b True\n", + "c is d False\n" + ] + } + ], + "source": [ + "a = 1\n", + "b = 1\n", + "print('a is b', bool(a is b))\n", + "True\n", + "\n", + "c = 999\n", + "d = 999\n", + "print('c is d', bool(c is d))" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "(*I received a comment that this is in fact a CPython artifact and **must not necessarily be true** in all implementations of Python!*)\n", + "\n", + "So the take home message is: always use \"==\" for equality, \"is\" for identity!\n", + "\n", + "Here is a [nice article](http://python.net/%7Egoodger/projects/pycon/2007/idiomatic/handout.html#other-languages-have-variables) explaining it by comparing \"boxes\" (C language) with \"name tags\" (Python)." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "This example demonstrates that this applies indeed for integers in the range in -5 to 256:" + ] + }, + { + "cell_type": "code", + "execution_count": 9, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "256 is 257-1 True\n", + "257 is 258-1 False\n", + "-5 is -6+1 True\n", + "-7 is -6-1 False\n" + ] + } + ], + "source": [ + "print('256 is 257-1', 256 is 257-1)\n", + "print('257 is 258-1', 257 is 258 - 1)\n", + "print('-5 is -6+1', -5 is -6+1)\n", + "print('-7 is -6-1', -7 is -6-1)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### And to illustrate the test for equality (`==`) vs. identity (`is`):" + ] + }, + { + "cell_type": "code", + "execution_count": 10, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "a is b, False\n", + "a == b, True\n" + ] + } + ], + "source": [ + "a = 'hello world!'\n", + "b = 'hello world!'\n", + "print('a is b,', a is b)\n", + "print('a == b,', a == b)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "We would think that identity would always imply equality, but this is not always true, as we can see in the next example:" + ] + }, + { + "cell_type": "code", + "execution_count": 11, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "a is a, True\n", + "a == a, False\n" + ] + } + ], + "source": [ + "a = float('nan')\n", + "print('a is a,', a is a)\n", + "print('a == a,', a == a)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
\n", + "" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Shallow vs. deep copies if list contains other structures and objects\n", + "\n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "**Shallow copy**: \n", + "If we use the assignment operator to assign one list to another list, we just create a new name reference to the original list. If we want to create a new list object, we have to make a copy of the original list. This can be done via `a_list[:]` or `a_list.copy()`." + ] + }, + { + "cell_type": "code", + "execution_count": 12, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "IDs:\n", + "list1: 4486860424\n", + "list2: 4486860424\n", + "list3: 4486818632\n", + "list4: 4486818568\n", + "\n", + "list1: [3, 2]\n", + "\n", + "list1: [3, 2]\n", + "list2: [3, 2]\n", + "list3: [4, 2]\n", + "list4: [1, 4]\n" + ] + } + ], + "source": [ + "list1 = [1,2]\n", + "list2 = list1 # reference\n", + "list3 = list1[:] # shallow copy\n", + "list4 = list1.copy() # shallow copy\n", + "\n", + "print('IDs:\\nlist1: {}\\nlist2: {}\\nlist3: {}\\nlist4: {}\\n'\n", + " .format(id(list1), id(list2), id(list3), id(list4)))\n", + "\n", + "list2[0] = 3\n", + "print('list1:', list1)\n", + "\n", + "list3[0] = 4\n", + "list4[1] = 4\n", + "print('\\nlist1:', list1)\n", + "print('list2:', list2)\n", + "print('list3:', list3)\n", + "print('list4:', list4)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "**Deep copy** \n", + "As we have seen above, a shallow copy works fine if we want to create a new list with contents of the original list which we want to modify independently. \n", + "\n", + "However, if we are dealing with compound objects (e.g., lists that contain other lists, [read here](https://docs.python.org/2/library/copy.html) for more information) it becomes a little trickier.\n", + "\n", + "In the case of compound objects, a shallow copy would create a new compound object, but it would just insert the references to the contained objects into the new compound object. In contrast, a deep copy would go \"deeper\" and create also new objects \n", + "for the objects found in the original compound object. \n", + "If you follow the code, the concept should become more clear:" + ] + }, + { + "cell_type": "code", + "execution_count": 13, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "IDs:\n", + "list1: 4486818824\n", + "list2: 4486886024\n", + "list3: 4486888200\n", + "\n", + "list1: [[3], [2]]\n", + "\n", + "list1: [[3], [2]]\n", + "list2: [[3], [2]]\n", + "list3: [[5], [2]]\n" + ] + } + ], + "source": [ + "from copy import deepcopy\n", + "\n", + "list1 = [[1],[2]]\n", + "list2 = list1.copy() # shallow copy\n", + "list3 = deepcopy(list1) # deep copy\n", + "\n", + "print('IDs:\\nlist1: {}\\nlist2: {}\\nlist3: {}\\n'\n", + " .format(id(list1), id(list2), id(list3)))\n", + "\n", + "list2[0][0] = 3\n", + "print('list1:', list1)\n", + "\n", + "list3[0][0] = 5\n", + "print('\\nlist1:', list1)\n", + "print('list2:', list2)\n", + "print('list3:', list3)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
\n", + "" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Picking `True` values from logical `and`s and `or`s" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "**Logical `or`:** \n", + "\n", + "`a or b == a if a else b` \n", + "- If both values in `or` expressions are `True`, Python will select the first value (e.g., select `\"a\"` in `\"a\" or \"b\"`), and the second one in `and` expressions. \n", + "This is also called **short-circuiting** - we already know that the logical `or` must be `True` if the first value is `True` and therefore can omit the evaluation of the second value.\n", + "\n", + "**Logical `and`:** \n", + "\n", + "`a and b == b if a else a` \n", + "- If both values in `and` expressions are `True`, Python will select the second value, since for a logical `and`, both values must be true.\n" + ] + }, + { + "cell_type": "code", + "execution_count": 14, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "2 * 7 = 14\n" + ] + } + ], + "source": [ + "result = (2 or 3) * (5 and 7)\n", + "print('2 * 7 =', result)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
\n", + "" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Don't use mutable objects as default arguments for functions!" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Don't use mutable objects (e.g., dictionaries, lists, sets, etc.) as default arguments for functions! You might expect that a new list is created every time when we call the function without providing an argument for the default parameter, but this is not the case: **Python will create the mutable object (default parameter) the first time the function is defined - not when it is called**, see the following code:\n", + "\n", + "(Original source: [http://docs.python-guide.org/en/latest/writing/gotchas/](http://docs.python-guide.org/en/latest/writing/gotchas/)" + ] + }, + { + "cell_type": "code", + "execution_count": 15, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "[1]\n", + "[1, 2]\n" + ] + } + ], + "source": [ + "def append_to_list(value, def_list=[]):\n", + " def_list.append(value)\n", + " return def_list\n", + "\n", + "my_list = append_to_list(1)\n", + "print(my_list)\n", + "\n", + "my_other_list = append_to_list(2)\n", + "print(my_other_list)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Another good example showing that demonstrates that default arguments are created when the function is created (**and not when it is called!**):" + ] + }, + { + "cell_type": "code", + "execution_count": 16, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "1528560045.3962939\n", + "1528560045.3962939\n" + ] + } + ], + "source": [ + "import time\n", + "def report_arg(my_default=time.time()):\n", + " print(my_default)\n", + "\n", + "report_arg()\n", + "\n", + "time.sleep(5)\n", + "\n", + "report_arg()" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
\n", + "\n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Be aware of the consuming generator" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Be aware of what is happening when combining `in` checks with generators, since they won't evaluate from the beginning once a position is \"consumed\"." + ] + }, + { + "cell_type": "code", + "execution_count": 17, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "2 in gen, True\n", + "3 in gen, True\n", + "1 in gen, False\n" + ] + } + ], + "source": [ + "gen = (i for i in range(5))\n", + "print('2 in gen,', 2 in gen)\n", + "print('3 in gen,', 3 in gen)\n", + "print('1 in gen,', 1 in gen) " + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Although this defeats the purpose of a generator (in most cases), we can convert a generator into a list to circumvent the problem. " + ] + }, + { + "cell_type": "code", + "execution_count": 18, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "2 in l, True\n", + "3 in l, True\n", + "1 in l, True\n" + ] + } + ], + "source": [ + "gen = (i for i in range(5))\n", + "a_list = list(gen)\n", + "print('2 in l,', 2 in a_list)\n", + "print('3 in l,', 3 in a_list)\n", + "print('1 in l,', 1 in a_list) " + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
\n", + "\n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "## `bool` is a subclass of `int`\n", + "\n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Chicken or egg? In the history of Python (Python 2.2 to be specific) truth values were implemented via 1 and 0 (similar to the old C). In order to avoid syntax errors in old (but perfectly working) Python code, `bool` was added as a subclass of `int` in Python 2.3.\n", + "\n", + "Original source: [http://www.peterbe.com/plog/bool-is-int](http://www.peterbe.com/plog/bool-is-int)" + ] + }, + { + "cell_type": "code", + "execution_count": 19, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "isinstance(True, int): True\n", + "True + True: 2\n", + "3*True + True: 4\n", + "3*True - False: 3\n" + ] + } + ], + "source": [ + "print('isinstance(True, int):', isinstance(True, int))\n", + "print('True + True:', True + True)\n", + "print('3*True + True:', 3*True + True)\n", + "print('3*True - False:', 3*True - False)\n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
\n", + "" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## About lambda-in-closures-and-a-loop pitfall" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Remember the section about the [consuming generators](#consuming_generator)? This example is somewhat related, but the result might still come as unexpected. \n", + "\n", + "(Original source: [http://openhome.cc/eGossip/Blog/UnderstandingLambdaClosure3.html](http://openhome.cc/eGossip/Blog/UnderstandingLambdaClosure3.html))\n", + "\n", + "In the first example below, we call a `lambda` function in a list comprehension, and the value `i` will be dereferenced every time we call `lambda` within the scope. Since the list comprehension has already been constructed and evaluated when we `for-loop` through the list, the closure-variable will be set to the last value 4." + ] + }, + { + "cell_type": "code", + "execution_count": 20, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "4\n", + "4\n", + "4\n", + "4\n", + "4\n" + ] + } + ], + "source": [ + "my_list = [lambda: i for i in range(5)]\n", + "for l in my_list:\n", + " print(l())" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "However, by using a generator expression, we can make use of its stepwise evaluation (note that the returned variable still stems from the same closure, but the value changes as we iterate over the generator)." + ] + }, + { + "cell_type": "code", + "execution_count": 21, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "0\n", + "1\n", + "2\n", + "3\n", + "4\n" + ] + } + ], + "source": [ + "my_gen = (lambda: n for n in range(5))\n", + "for l in my_gen:\n", + " print(l())" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "And if you are really keen on using lists, there is a nifty trick that circumvents this problem as a reader nicely pointed out in the comments: We can simply pass the loop variable `i` as a default argument to the lambdas." + ] + }, + { + "cell_type": "code", + "execution_count": 22, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "0\n", + "1\n", + "2\n", + "3\n", + "4\n" + ] + } + ], + "source": [ + "my_list = [lambda x=i: x for i in range(5)]\n", + "for l in my_list:\n", + " print(l())" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
\n", + "" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Python's LEGB scope resolution and the keywords `global` and `nonlocal`" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "There is nothing particularly surprising about Python's LEGB scope resolution (Local -> Enclosed -> Global -> Built-in), but it is still useful to take a look at some examples!" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### `global` vs. `local`\n", + "\n", + "According to the LEGB rule, Python will first look for a variable in the local scope. So if we set the variable `x = 1` `local`ly in the function's scope, it won't have an effect on the `global` `x`." + ] + }, + { + "cell_type": "code", + "execution_count": 23, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "in_func: 1\n", + "global: 0\n" + ] + } + ], + "source": [ + "x = 0\n", + "\n", + "\n", + "def in_func():\n", + " x = 1\n", + " print('in_func:', x)\n", + "\n", + "\n", + "in_func()\n", + "print('global:', x)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "If we want to modify the `global` x via a function, we can simply use the `global` keyword to import the variable into the function's scope:" + ] + }, + { + "cell_type": "code", + "execution_count": 24, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "in_func: 1\n", + "global: 1\n" + ] + } + ], + "source": [ + "x = 0\n", + "\n", + "\n", + "def in_func():\n", + " global x\n", + " x = 1\n", + " print('in_func:', x)\n", + "\n", + "\n", + "in_func()\n", + "print('global:', x)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### `local` vs. `enclosed`\n", + "\n", + "Now, let us take a look at `local` vs. `enclosed`. Here, we set the variable `x = 1` in the `outer` function and set `x = 1` in the enclosed function `inner`. Since `inner` looks in the local scope first, it won't modify `outer`'s `x`." + ] + }, + { + "cell_type": "code", + "execution_count": 25, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "outer before: 1\n", + "inner: 2\n", + "outer after: 1\n" + ] + } + ], + "source": [ + "def outer():\n", + " x = 1\n", + " print('outer before:', x)\n", + "\n", + " def inner():\n", + " x = 2\n", + " print(\"inner:\", x)\n", + " inner()\n", + " print(\"outer after:\", x)\n", + "\n", + "\n", + "outer()" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Here is where the `nonlocal` keyword comes in handy - it allows us to modify the `x` variable in the `enclosed` scope:" + ] + }, + { + "cell_type": "code", + "execution_count": 26, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "outer before: 1\n", + "inner: 2\n", + "outer after: 2\n" + ] + } + ], + "source": [ + "def outer():\n", + " x = 1\n", + " print('outer before:', x)\n", + "\n", + " def inner():\n", + " nonlocal x\n", + " x = 2\n", + " print(\"inner:\", x)\n", + " inner()\n", + " print(\"outer after:\", x)\n", + "\n", + "\n", + "outer()" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
\n", + "" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## When mutable contents of immutable tuples aren't so mutable" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "As we all know, tuples are immutable objects in Python, right!? But what happens if they contain mutable objects? \n", + "\n", + "First, let us have a look at the expected behavior: a `TypeError` is raised if we try to modify immutable types in a tuple: " + ] + }, + { + "cell_type": "code", + "execution_count": 27, + "metadata": {}, + "outputs": [ + { + "ename": "TypeError", + "evalue": "'tuple' object does not support item assignment", + "output_type": "error", + "traceback": [ + "\u001b[0;31m---------------------------------------------------------------------------\u001b[0m", + "\u001b[0;31mTypeError\u001b[0m Traceback (most recent call last)", + "\u001b[0;32m
\n", + "
\n", + "However, **there are ways** to modify the mutable contents of the tuple without raising the `TypeError`, the solution is the `.extend()` method, or alternatively `.append()` (for lists):" + ] + }, + { + "cell_type": "code", + "execution_count": 29, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "tup before: ([],)\n", + "tup after: ([1],)\n" + ] + } + ], + "source": [ + "tup = ([],)\n", + "print('tup before: ', tup)\n", + "tup[0].extend([1])\n", + "print('tup after: ', tup)" + ] + }, + { + "cell_type": "code", + "execution_count": 30, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "tup before: ([],)\n", + "tup after: ([1],)\n" + ] + } + ], + "source": [ + "tup = ([],)\n", + "print('tup before: ', tup)\n", + "tup[0].append(1)\n", + "print('tup after: ', tup)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### Explanation\n", + "\n", + "**A. Jesse Jiryu Davis** has a nice explanation for this phenomenon (Original source: [http://emptysqua.re/blog/python-increment-is-weird-part-ii/](http://emptysqua.re/blog/python-increment-is-weird-part-ii/))\n", + "\n", + "If we try to extend the list via `+=` *\"then the statement executes `STORE_SUBSCR`, which calls the C function `PyObject_SetItem`, which checks if the object supports item assignment. In our case the object is a tuple, so `PyObject_SetItem` throws the `TypeError`. Mystery solved.\"*" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### One more note about the `immutable` status of tuples. Tuples are famous for being immutable. However, how comes that this code works?" + ] + }, + { + "cell_type": "code", + "execution_count": 31, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "(1, 4, 5)\n" + ] + } + ], + "source": [ + "my_tup = (1,)\n", + "my_tup += (4,)\n", + "my_tup = my_tup + (5,)\n", + "print(my_tup)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "What happens \"behind\" the curtains is that the tuple is not modified, but a new object is generated every time, which will inherit the old \"name tag\":" + ] + }, + { + "cell_type": "code", + "execution_count": 32, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "4486707912\n", + "4485211784\n", + "4486955152\n" + ] + } + ], + "source": [ + "my_tup = (1,)\n", + "print(id(my_tup))\n", + "my_tup += (4,)\n", + "print(id(my_tup))\n", + "my_tup = my_tup + (5,)\n", + "print(id(my_tup))" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
\n", + "\n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## List comprehensions are fast, but generators are faster!?" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\"List comprehensions are fast, but generators are faster!?\" - No, not really (or significantly, see the benchmarks below). So what's the reason to prefer one over the other?\n", + "- use lists if you want to use the plethora of list methods \n", + "- use generators when you are dealing with huge collections to avoid memory issues" + ] + }, + { + "cell_type": "code", + "execution_count": 33, + "metadata": {}, + "outputs": [], + "source": [ + "import timeit\n", + "\n", + "\n", + "def plainlist(n=100000):\n", + " my_list = []\n", + " for i in range(n):\n", + " if i % 5 == 0:\n", + " my_list.append(i)\n", + " return my_list\n", + "\n", + "\n", + "def listcompr(n=100000):\n", + " my_list = [i for i in range(n) if i % 5 == 0]\n", + " return my_list\n", + "\n", + "\n", + "def generator(n=100000):\n", + " my_gen = (i for i in range(n) if i % 5 == 0)\n", + " return my_gen\n", + "\n", + "\n", + "def generator_yield(n=100000):\n", + " for i in range(n):\n", + " if i % 5 == 0:\n", + " yield i" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### To be fair to the list, let us exhaust the generators:" + ] + }, + { + "cell_type": "code", + "execution_count": 34, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "plain_list: 10.8 ms ± 793 µs per loop (mean ± std. dev. of 7 runs, 100 loops each)\n", + "\n", + "listcompr: 10 ms ± 830 µs per loop (mean ± std. dev. of 7 runs, 100 loops each)\n", + "\n", + "generator: 11.4 ms ± 1 ms per loop (mean ± std. dev. of 7 runs, 100 loops each)\n", + "\n", + "generator_yield: 12.3 ms ± 1.82 ms per loop (mean ± std. dev. of 7 runs, 100 loops each)\n" + ] + } + ], + "source": [ + "def test_plainlist(plain_list):\n", + " for i in plain_list():\n", + " pass\n", + "\n", + "\n", + "def test_listcompr(listcompr):\n", + " for i in listcompr():\n", + " pass\n", + "\n", + "\n", + "def test_generator(generator):\n", + " for i in generator():\n", + " pass\n", + "\n", + "\n", + "def test_generator_yield(generator_yield):\n", + " for i in generator_yield():\n", + " pass\n", + "\n", + "\n", + "print('plain_list: ', end='')\n", + "%timeit test_plainlist(plainlist)\n", + "print('\\nlistcompr: ', end='')\n", + "%timeit test_listcompr(listcompr)\n", + "print('\\ngenerator: ', end='')\n", + "%timeit test_generator(generator)\n", + "print('\\ngenerator_yield: ', end='')\n", + "%timeit test_generator_yield(generator_yield)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
\n", + "" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Public vs. private class methods and name mangling\n", + "\n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Who has not stumbled across this quote \"we are all consenting adults here\" in the Python community, yet? Unlike in other languages like C++ (sorry, there are many more, but that's one I am most familiar with), we can't really protect class methods from being used outside the class (i.e., by the API user). \n", + "All we can do is indicate methods as private to make clear that they are not to be used outside the class, but it really is up to the class user, since \"we are all consenting adults here\"! \n", + "So, when we want to mark a class method as private, we can put a single underscore in front of it. \n", + "If we additionally want to avoid name clashes with other classes that might use the same method names, we can prefix the name with a double-underscore to invoke the name mangling.\n", + "\n", + "This doesn't prevent the class users to access this class member though, but they have to know the trick and also know that it is at their own risk...\n", + "\n", + "Let the following example illustrate what I mean:" + ] + }, + { + "cell_type": "code", + "execution_count": 35, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Hello public world!\n", + "Hello private world!\n", + "Hello private world!\n" + ] + } + ], + "source": [ + "class my_class():\n", + " def public_method(self):\n", + " print('Hello public world!')\n", + "\n", + " def __private_method(self):\n", + " print('Hello private world!')\n", + "\n", + " def call_private_method_in_class(self):\n", + " self.__private_method()\n", + "\n", + "\n", + "my_instance = my_class()\n", + "\n", + "my_instance.public_method()\n", + "my_instance._my_class__private_method()\n", + "my_instance.call_private_method_in_class()" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
\n", + "" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## The consequences of modifying a list when looping through it" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "It can be really dangerous to modify a list when iterating through it - this is a very common pitfall that can cause unintended behavior! \n", + "Look at the following examples, and for a fun exercise: try to figure out what is going on before you skip to the solution!" + ] + }, + { + "cell_type": "code", + "execution_count": 36, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "[1, 3, 5]\n" + ] + } + ], + "source": [ + "a = [1, 2, 3, 4, 5]\n", + "for i in a:\n", + " if not i % 2:\n", + " a.remove(i)\n", + "print(a)" + ] + }, + { + "cell_type": "code", + "execution_count": 37, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "[4, 5]\n" + ] + } + ], + "source": [ + "b = [2, 4, 5, 6]\n", + "for i in b:\n", + " if not i % 2:\n", + " b.remove(i)\n", + "print(b)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
\n", + "**The solution** is that we are iterating through the list index by index, and if we remove one of the items in-between, we inevitably mess around with the indexing. Look at the following example and it will become clear:" + ] + }, + { + "cell_type": "code", + "execution_count": 38, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "0 2\n", + "1 5\n", + "2 6\n", + "[4, 5]\n" + ] + } + ], + "source": [ + "b = [2, 4, 5, 6]\n", + "for index, item in enumerate(b):\n", + " print(index, item)\n", + " if not item % 2:\n", + " b.remove(item)\n", + "print(b)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
\n", + "" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Dynamic binding and typos in variable names\n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Be careful, dynamic binding is convenient, but can also quickly become dangerous!" + ] + }, + { + "cell_type": "code", + "execution_count": 39, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "first list:\n", + "0\n", + "1\n", + "2\n", + "\n", + "second list:\n", + "2\n", + "2\n", + "2\n" + ] + } + ], + "source": [ + "print('first list:')\n", + "for i in range(3):\n", + " print(i)\n", + " \n", + "print('\\nsecond list:')\n", + "for j in range(3):\n", + " print(i) # I (intentionally) made typo here!" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
\n", + "" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "## List slicing using indexes that are \"out of range\"" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "As we have all encountered it 1 (x10000) time(s) in our lives, the infamous `IndexError`:" + ] + }, + { + "cell_type": "code", + "execution_count": 40, + "metadata": {}, + "outputs": [ + { + "ename": "IndexError", + "evalue": "list index out of range", + "output_type": "error", + "traceback": [ + "\u001b[0;31m---------------------------------------------------------------------------\u001b[0m", + "\u001b[0;31mIndexError\u001b[0m Traceback (most recent call last)", + "\u001b[0;32m
\n", + "
\n", + "" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "## Reusing global variable names and `UnboundLocalErrors`" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Usually, it is no problem to access global variables in the local scope of a function:" + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "def my_func():\n", + " print(var)\n", + "\n", + "\n", + "var = 'global'\n", + "my_func()" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "And is also no problem to use the same variable name in the local scope without affecting the local counterpart: " + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "def my_func():\n", + " var = 'locally changed'\n", + "\n", + "\n", + "var = 'global'\n", + "my_func()\n", + "print(var)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "But we have to be careful if we use a variable name that occurs in the global scope, and we want to access it in the local function scope if we want to reuse this name:" + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "def my_func():\n", + " print(var) # want to access global variable\n", + " var = 'locally changed' # but Python thinks we forgot to define the local variable!\n", + "\n", + "\n", + "var = 'global'\n", + "my_func()" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "In this case, we have to use the `global` keyword!" + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "def my_func():\n", + " global var\n", + " print(var) # want to access global variable\n", + " var = 'locally changed' # changes the gobal variable\n", + "\n", + "\n", + "var = 'global'\n", + "\n", + "my_func()\n", + "print(var)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
\n", + "" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Creating copies of mutable objects\n", + "\n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Let's assume a scenario where we want to duplicate sub`list`s of values stored in another list. If we want to create an independent sub`list` object, using the arithmetic multiplication operator could lead to rather unexpected (or undesired) results:" + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "my_list1 = [[1, 2, 3]] * 2\n", + "\n", + "print('initially ---> ', my_list1)\n", + "\n", + "# modify the 1st element of the 2nd sublist\n", + "my_list1[1][0] = 'a'\n", + "print(\"after my_list1[1][0] = 'a' ---> \", my_list1)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
\n", + "In this case, we should better create \"new\" objects:" + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "my_list2 = [[1, 2, 3] for i in range(2)]\n", + "\n", + "print('initially: ---> ', my_list2)\n", + "\n", + "# modify the 1st element of the 2nd sublist\n", + "my_list2[1][0] = 'a'\n", + "print(\"after my_list2[1][0] = 'a': ---> \", my_list2)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "And here is the proof:" + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "for a, b in zip(my_list1, my_list2):\n", + " print('id my_list1: {}, id my_list2: {}'.format(id(a), id(b)))" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
\n", + "" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "## Key differences between Python 2 and 3\n", + "\n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "There are some good articles already that are summarizing the differences between Python 2 and 3, e.g., \n", + "- [https://wiki.python.org/moin/Python2orPython3](https://wiki.python.org/moin/Python2orPython3)\n", + "- [https://docs.python.org/3.0/whatsnew/3.0.html](https://docs.python.org/3.0/whatsnew/3.0.html)\n", + "- [http://python3porting.com/differences.html](http://python3porting.com/differences.html)\n", + "- [https://docs.python.org/3/howto/pyporting.html](https://docs.python.org/3/howto/pyporting.html) \n", + "etc.\n", + "\n", + "But it might be still worthwhile, especially for Python newcomers, to take a look at some of those!\n", + "(Note: the the code was executed in Python 3.4.0 and Python 2.7.5 and copied from interactive shell sessions.)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### Overview - Key differences between Python 2 and 3" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "\n", + "- [Unicode](#unicode)\n", + "- [The print statement](#print)\n", + "- [Integer division](#integer_div)\n", + "- [xrange()](#xrange)\n", + "- [Raising exceptions](#raising_exceptions)\n", + "- [Handling exceptions](#handling_exceptions)\n", + "- [next() function and .next() method](#next_next)\n", + "- [Loop variables and leaking into the global scope](#loop_leak)\n", + "- [Comparing unorderable types](#compare_unorder)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### Unicode..." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to Python 2.x vs 3.x overview](#py23_overview)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "#### Python 2: \n", + "We have ASCII `str()` types, separate `unicode()`, but no `byte` type\n", + "#### Python 3: \n", + "Now, we finally have Unicode (utf-8) `str`ings, and 2 byte classes: `byte` and `bytearray`s" + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": { + "code_folding": [] + }, + "outputs": [], + "source": [ + "#############\n", + "# Python 2 #\n", + "#############\n", + "\n", + ">>> type(unicode('is like a python3 str()'))\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### The print statement" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to Python 2.x vs 3.x overview](#py23_overview)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Very trivial, but this change makes sense, Python 3 now only accepts `print`s with proper parentheses - just like the other function calls ..." + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "# Python 2\n", + ">>> print 'Hello, World!'\n", + "Hello, World!\n", + ">>> print('Hello, World!')\n", + "Hello, World!\n", + "\n", + "# Python 3\n", + ">>> print('Hello, World!')\n", + "Hello, World!\n", + ">>> print 'Hello, World!'\n", + " File \"
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### Integer division" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to Python 2.x vs 3.x overview](#py23_overview)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "This is a pretty dangerous thing if you are porting code, or executing Python 3 code in Python 2 since the change in integer-division behavior can often go unnoticed. \n", + "So, I still tend to use a `float(3)/2` or `3/2.0` instead of a `3/2` in my Python 3 scripts to save the Python 2 guys some trouble ... (PS: and vice versa, you can `from __future__ import division` in your Python 2 scripts)." + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "# Python 2\n", + ">>> 3 / 2\n", + "1\n", + ">>> 3 // 2\n", + "1\n", + ">>> 3 / 2.0\n", + "1.5\n", + ">>> 3 // 2.0\n", + "1.0\n", + "\n", + "# Python 3\n", + ">>> 3 / 2\n", + "1.5\n", + ">>> 3 // 2\n", + "1\n", + ">>> 3 / 2.0\n", + "1.5\n", + ">>> 3 // 2.0\n", + "1.0" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### `xrange()`" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to Python 2.x vs 3.x overview](#py23_overview)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + " \n", + "`xrange()` was pretty popular in Python 2.x if you wanted to create an iterable object. The behavior was quite similar to a generator ('lazy evaluation'), but you could iterate over it infinitely. The advantage was that it was generally faster than `range()` (e.g., in a for-loop) - not if you had to iterate over the list multiple times, since the generation happens every time from scratch! \n", + "In Python 3, the `range()` was implemented like the `xrange()` function so that a dedicated `xrange()` function does not exist anymore." + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "# Python 2\n", + ">>> python -m timeit 'for i in range(1000000):' ' pass'\n", + "10 loops, best of 3: 66 msec per loop\n", + "\n", + " > python -m timeit 'for i in xrange(1000000):' ' pass'\n", + "10 loops, best of 3: 27.8 msec per loop\n", + "\n", + "# Python 3\n", + ">>> python3 -m timeit 'for i in range(1000000):' ' pass'\n", + "10 loops, best of 3: 51.1 msec per loop\n", + "\n", + ">>> python3 -m timeit 'for i in xrange(1000000):' ' pass'\n", + "Traceback (most recent call last):\n", + " File \"/Library/Frameworks/Python.framework/Versions/3.4/lib/python3.4/timeit.py\", line 292, in main\n", + " x = t.timeit(number)\n", + " File \"/Library/Frameworks/Python.framework/Versions/3.4/lib/python3.4/timeit.py\", line 178, in timeit\n", + " timing = self.inner(it, self.timer)\n", + " File \"
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### Raising exceptions" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to Python 2.x vs 3.x overview](#py23_overview)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "\n", + "Where Python 2 accepts both notations, the 'old' and the 'new' way, Python 3 chokes (and raises a `SyntaxError` in turn) if we don't enclose the exception argument in parentheses:" + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "# Python 2\n", + ">>> raise IOError, \"file error\"\n", + "Traceback (most recent call last):\n", + " File \"
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### Handling exceptions" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to Python 2.x vs 3.x overview](#py23_overview)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "\n", + "Also the handling of exceptions has slightly changed in Python 3. Now, we have to use the `as` keyword!" + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "# Python 2\n", + ">>> try:\n", + "... blabla\n", + "... except NameError, err:\n", + "... print err, '--> our error msg'\n", + "... \n", + "name 'blabla' is not defined --> our error msg\n", + "\n", + "# Python 3\n", + ">>> try:\n", + "... blabla\n", + "... except NameError as err:\n", + "... print(err, '--> our error msg')\n", + "... \n", + "name 'blabla' is not defined --> our error msg" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### The `next()` function and `.next()` method" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to Python 2.x vs 3.x overview](#py23_overview)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "\n", + "Where you can use both function and method in Python 2.7.5, the `next()` function is all that remains in Python 3!" + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "# Python 2\n", + ">>> my_generator = (letter for letter in 'abcdefg')\n", + ">>> my_generator.next()\n", + "'a'\n", + ">>> next(my_generator)\n", + "'b'\n", + "\n", + "# Python 3\n", + ">>> my_generator = (letter for letter in 'abcdefg')\n", + ">>> next(my_generator)\n", + "'a'\n", + ">>> my_generator.next()\n", + "Traceback (most recent call last):\n", + " File \"
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### In Python 3.x for-loop variables don't leak into the global namespace anymore" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to Python 2.x vs 3.x overview](#py23_overview)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "This goes back to a change that was made in Python 3.x and is described in [What’s New In Python 3.0](https://docs.python.org/3/whatsnew/3.0.html) as follows:\n", + "\n", + "*\"List comprehensions no longer support the syntactic form `[... for var in item1, item2, ...]`. Use `[... for var in (item1, item2, ...)]` instead. Also note that list comprehensions have different semantics: they are closer to syntactic sugar for a generator expression inside a `list()` constructor, and in particular the loop control variables are no longer leaked into the surrounding scope.\"*" + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + ">>> from platform import python_version\n", + ">>> print 'This code cell was executed in Python', python_version()\n", + "'This code cell was executed in Python 2.7.6'\n", + ">>> i = 1\n", + ">>> print [i for i in range(5)]\n", + "'[0, 1, 2, 3, 4]'\n", + ">>> print i, '-> i in global'\n", + "'4 -> i in global'" + ] + }, + { + "cell_type": "code", + "execution_count": 61, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "This code cell was executed in Python 3.6.4\n", + "[0, 1, 2, 3, 4]\n", + "1 -> i in global\n" + ] + } + ], + "source": [ + "%%python3\n", + "from platform import python_version\n", + "print('This code cell was executed in Python', python_version())\n", + "\n", + "i = 1\n", + "print([i for i in range(5)])\n", + "print(i, '-> i in global')" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### Python 3.x prevents us from comparing unorderable types" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to Python 2.x vs 3.x overview](#py23_overview)]" + ] + }, + { + "cell_type": "code", + "execution_count": 101, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Couldn't find program: 'python2'\n" + ] + } + ], + "source": [ + ">>> from platform import python_version\n", + ">>> print 'This code cell was executed in Python', python_version()\n", + "'This code cell was executed in Python 2.7.6'\n", + ">>> print [1, 2] > 'foo'\n", + "'False'\n", + ">>> print (1, 2) > 'foo'\n", + "'True'\n", + ">>> print [1, 2] > (1, 2)\n", + "'False'" + ] + }, + { + "cell_type": "code", + "execution_count": 67, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "This code cell was executed in Python 3.6.4\n" + ] + }, + { + "ename": "TypeError", + "evalue": "'>' not supported between instances of 'list' and 'str'", + "output_type": "error", + "traceback": [ + "\u001b[0;31m---------------------------------------------------------------------------\u001b[0m", + "\u001b[0;31mTypeError\u001b[0m Traceback (most recent call last)", + "\u001b[0;32m
\n", + "
\n", + "" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "## Function annotations - What are those `->`'s in my Python code?\n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Have you ever seen any Python code that used colons inside the parantheses of a function definition?" + ] + }, + { + "cell_type": "code", + "execution_count": 14, + "metadata": {}, + "outputs": [], + "source": [ + "def foo1(x: 'insert x here', y: 'insert x^2 here'):\n", + " print('Hello, World')\n", + " return" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "And what about the fancy arrow here?" + ] + }, + { + "cell_type": "code", + "execution_count": 15, + "metadata": {}, + "outputs": [], + "source": [ + "def foo2(x, y) -> 'Hi!':\n", + " print('Hello, World')\n", + " return" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Q: Is this valid Python syntax? \n", + "A: Yes!\n", + " \n", + " \n", + "Q: So, what happens if I *just call* the function? \n", + "A: Nothing!\n", + " \n", + "Here is the proof!" + ] + }, + { + "cell_type": "code", + "execution_count": 17, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Hello, World\n" + ] + } + ], + "source": [ + "foo1(1,2)" + ] + }, + { + "cell_type": "code", + "execution_count": 16, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Hello, World\n" + ] + } + ], + "source": [ + "foo2(1,2) " + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "**So, those are function annotations ... ** \n", + "- the colon for the function parameters \n", + "- the arrow for the return value \n", + "\n", + "You probably will never make use of them (or at least very rarely). Usually, we write good function documentations below the function as a docstring - or at least this is how I would do it (okay this case is a little bit extreme, I have to admit):" + ] + }, + { + "cell_type": "code", + "execution_count": 18, + "metadata": {}, + "outputs": [], + "source": [ + "def is_palindrome(a):\n", + " \"\"\"\n", + " Case-and punctuation insensitive check if a string is a palindrom.\n", + "\n", + " Keyword arguments:\n", + " a (str): The string to be checked if it is a palindrome.\n", + "\n", + " Returns `True` if input string is a palindrome, else False.\n", + "\n", + " \"\"\"\n", + " stripped_str = [l for l in my_str.lower() if l.isalpha()]\n", + " return stripped_str == stripped_str[::-1]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "However, function annotations can be useful to indicate that work is still in progress in some cases. But they are optional and I see them very, very rarely.\n", + "\n", + "As it is stated in [PEP3107](http://legacy.python.org/dev/peps/pep-3107/#fundamentals-of-function-annotations):\n", + "\n", + "1. *Function annotations, both for parameters and return values, are completely optional.*\n", + "\n", + "2. *Function annotations are nothing more than a way of associating arbitrary Python expressions with various parts of a function at compile-time.*\n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "The nice thing about function annotations is their `__annotations__` attribute, which is a dictionary of all the parameters and/or the `return` value you annotated." + ] + }, + { + "cell_type": "code", + "execution_count": 19, + "metadata": {}, + "outputs": [ + { + "data": { + "text/plain": [ + "{'x': 'insert x here', 'y': 'insert x^2 here'}" + ] + }, + "execution_count": 19, + "metadata": {}, + "output_type": "execute_result" + } + ], + "source": [ + "foo1.__annotations__" + ] + }, + { + "cell_type": "code", + "execution_count": 20, + "metadata": {}, + "outputs": [ + { + "data": { + "text/plain": [ + "{'return': 'Hi!'}" + ] + }, + "execution_count": 20, + "metadata": {}, + "output_type": "execute_result" + } + ], + "source": [ + "foo2.__annotations__" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "**When are they useful?**" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Function annotations can be useful for a couple of things \n", + "- Documentation in general\n", + "- pre-condition testing\n", + "- [type checking](http://legacy.python.org/dev/peps/pep-0362/#annotation-checker)\n", + " \n", + "..." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Abortive statements in `finally` blocks" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Python's `try-except-finally` blocks are very handy for catching and handling errors. The `finally` block is always executed whether an `exception` has been raised or not as illustrated in the following example." + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [ + "def try_finally1():\n", + " try:\n", + " print('in try:')\n", + " print('do some stuff')\n", + " float('abc')\n", + " except ValueError:\n", + " print('an error occurred')\n", + " else:\n", + " print('no error occurred')\n", + " finally:\n", + " print('always execute finally')\n", + "\n", + "\n", + "try_finally1()" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
\n", + "But can you also guess what will be printed in the next code cell?" + ] + }, + { + "cell_type": "code", + "execution_count": 21, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "do some stuff in try block\n", + "do some stuff in finally block\n", + "always execute finally\n" + ] + } + ], + "source": [ + "def try_finally2():\n", + " try:\n", + " print(\"do some stuff in try block\")\n", + " return \"return from try block\"\n", + " finally:\n", + " print(\"do some stuff in finally block\")\n", + " return \"always execute finally\"\n", + "\n", + "\n", + "print(try_finally2())" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "Here, the abortive `return` statement in the `finally` block simply overrules the `return` in the `try` block, since **`finally` is guaranteed to always be executed.** So, be careful using abortive statements in `finally` blocks!" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
\n", + "" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Assigning types to variables as values" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "I am not yet sure in which context this can be useful, but it is a nice fun fact to know that we can assign types as values to variables." + ] + }, + { + "cell_type": "code", + "execution_count": 22, + "metadata": {}, + "outputs": [ + { + "data": { + "text/plain": [ + "'123'" + ] + }, + "execution_count": 22, + "metadata": {}, + "output_type": "execute_result" + } + ], + "source": [ + "a_var = str\n", + "a_var(123)" + ] + }, + { + "cell_type": "code", + "execution_count": 23, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "0.0
\n", + "
\n", + "" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Only the first clause of generators is evaluated immediately" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "The main reason why we love to use generators in certain cases (i.e., when we are dealing with large numbers of computations) is that it only computes the next value when it is needed, which is also known as \"lazy\" evaluation.\n", + "However, the first clause of an generator is already checked upon it's creation, as the following example demonstrates:" + ] + }, + { + "cell_type": "code", + "execution_count": 24, + "metadata": {}, + "outputs": [ + { + "ename": "ZeroDivisionError", + "evalue": "division by zero", + "output_type": "error", + "traceback": [ + "\u001b[0;31m---------------------------------------------------------------------------\u001b[0m", + "\u001b[0;31mZeroDivisionError\u001b[0m Traceback (most recent call last)", + "\u001b[0;32m
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Keyword argument unpacking syntax - `*args` and `**kwargs`" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Python has a very convenient \"keyword argument unpacking syntax\" (often referred to as \"splat\"-operators). This is particularly useful, if we want to define a function that can take a arbitrary number of input arguments." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### Single-asterisk (*args)" + ] + }, + { + "cell_type": "code", + "execution_count": 27, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "type of args:
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Metaclasses - What creates a new instance of a class?" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Usually, it is the `__init__` method when we think of instanciating a new object from a class. However, it is the static method `__new__` (it is not a class method!) that creates and returns a new instance before `__init__()` is called. \n", + "More specifically, this is what is returned: \n", + "`return super(
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Else-clauses: \"conditional else\" and \"completion else\"" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "I would claim that the conditional `else` is every programmer's daily bread and butter. However, there is a second flavor of `else`-clauses in Python, which I will call \"completion else\" (for reason that will become clear later). \n", + "But first, let us take a look at our \"traditional\" conditional else that we all are familiar with. \n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### Conditional else:" + ] + }, + { + "cell_type": "code", + "execution_count": 34, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Hello, World!\n" + ] + } + ], + "source": [ + "# conditional else\n", + "\n", + "a_list = [1,2]\n", + "if a_list[0] == 1:\n", + " print('Hello, World!')\n", + "else:\n", + " print('Bye, World!')" + ] + }, + { + "cell_type": "code", + "execution_count": 35, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Bye, World!\n" + ] + } + ], + "source": [ + "# conditional else\n", + "\n", + "a_list = [1,2]\n", + "if a_list[0] == 2:\n", + " print('Hello, World!')\n", + "else:\n", + " print('Bye, World!')" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Why am I showing those simple examples? I think they are good to highlight some of the key points: It is **either** the code under the `if` clause that is executed, **or** the code under the `else` block, but not both. \n", + "If the condition of the `if` clause evaluates to `True`, the `if`-block is exectured, and if it evaluated to `False`, it is the `else` block. \n", + "
\n", + "### Completion else\n", + "**In contrast** to the **either...or*** situation that we know from the conditional `else`, the completion `else` is executed if a code block finished. \n", + "To show you an example, let us use `else` for error-handling:" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### Completion else (try-except)" + ] + }, + { + "cell_type": "code", + "execution_count": 36, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "first element: 1\n", + "no error in try-block\n" + ] + } + ], + "source": [ + "try:\n", + " print('first element:', a_list[0])\n", + "except IndexError:\n", + " print('raised IndexError')\n", + "else:\n", + " print('no error in try-block')" + ] + }, + { + "cell_type": "code", + "execution_count": 37, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "raised IndexError\n" + ] + } + ], + "source": [ + "try:\n", + " print('third element:', a_list[2])\n", + "except IndexError:\n", + " print('raised IndexError')\n", + "else:\n", + " print('no error in try-block')" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "In the code above, we can see that the code under the **`else`-clause is only executed if the `try-block` was executed without encountering an error, i.e., if the `try`-block is \"complete\".** \n", + "The same rule applies to the \"completion\" `else` in while- and for-loops, which you can confirm in the following samples below." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### Completion else (while-loop)" + ] + }, + { + "cell_type": "code", + "execution_count": 38, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "0\n", + "1\n", + "in else\n" + ] + } + ], + "source": [ + "i = 0\n", + "while i < 2:\n", + " print(i)\n", + " i += 1\n", + "else:\n", + " print('in else')" + ] + }, + { + "cell_type": "code", + "execution_count": 39, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "0\n" + ] + } + ], + "source": [ + "i = 0\n", + "while i < 2:\n", + " print(i)\n", + " i += 1\n", + " break\n", + "else:\n", + " print('completed while-loop')" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### Completion else (for-loop)" + ] + }, + { + "cell_type": "code", + "execution_count": 40, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "0\n", + "1\n", + "completed for-loop\n" + ] + } + ], + "source": [ + "for i in range(2):\n", + " print(i)\n", + "else:\n", + " print('completed for-loop')" + ] + }, + { + "cell_type": "code", + "execution_count": 41, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "0\n" + ] + } + ], + "source": [ + "for i in range(2):\n", + " print(i)\n", + " break\n", + "else:\n", + " print('completed for-loop')" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Interning of compile-time constants vs. run-time expressions" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "This might not be particularly useful, but it is nonetheless interesting: Python's interpreter is interning compile-time constants but not run-time expressions (note that this is implementation-specific).\n", + "\n", + "(Original source: [Stackoverflow](http://stackoverflow.com/questions/15541404/python-string-interning))" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Let us have a look at the simple example below. Here we are creating 3 variables and assign the value \"Hello\" to them in different ways before we test them for identity." + ] + }, + { + "cell_type": "code", + "execution_count": 42, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "hello1 is hello2: True\n", + "hello1 is hello3: False\n" + ] + } + ], + "source": [ + "hello1 = 'Hello'\n", + "\n", + "hello2 = 'Hell' + 'o'\n", + "\n", + "hello3 = 'Hell'\n", + "hello3 = hello3 + 'o'\n", + "\n", + "print('hello1 is hello2:', hello1 is hello2)\n", + "print('hello1 is hello3:', hello1 is hello3)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Now, how does it come that the first expression evaluates to true, but the second does not? To answer this question, we need to take a closer look at the underlying byte codes:" + ] + }, + { + "cell_type": "code", + "execution_count": 43, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + " 3 0 LOAD_CONST 1 ('Hello')\n", + " 2 STORE_FAST 0 (s)\n", + "\n", + " 4 4 LOAD_FAST 0 (s)\n", + " 6 RETURN_VALUE\n" + ] + } + ], + "source": [ + "import dis\n", + "def hello1_func():\n", + " s = 'Hello'\n", + " return s\n", + "dis.dis(hello1_func)" + ] + }, + { + "cell_type": "code", + "execution_count": 44, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + " 2 0 LOAD_CONST 3 ('Hello')\n", + " 2 STORE_FAST 0 (s)\n", + "\n", + " 3 4 LOAD_FAST 0 (s)\n", + " 6 RETURN_VALUE\n" + ] + } + ], + "source": [ + "def hello2_func():\n", + " s = 'Hell' + 'o'\n", + " return s\n", + "dis.dis(hello2_func)" + ] + }, + { + "cell_type": "code", + "execution_count": 45, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + " 2 0 LOAD_CONST 1 ('Hell')\n", + " 2 STORE_FAST 0 (s)\n", + "\n", + " 3 4 LOAD_FAST 0 (s)\n", + " 6 LOAD_CONST 2 ('o')\n", + " 8 BINARY_ADD\n", + " 10 STORE_FAST 0 (s)\n", + "\n", + " 4 12 LOAD_FAST 0 (s)\n", + " 14 RETURN_VALUE\n" + ] + } + ], + "source": [ + "def hello3_func():\n", + " s = 'Hell'\n", + " s = s + 'o'\n", + " return s\n", + "dis.dis(hello3_func)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "It looks like that `'Hello'` and `'Hell'` + `'o'` are both evaluated and stored as `'Hello'` at compile-time, whereas the third version \n", + "`s = 'Hell'` \n", + "`s = s + 'o'` seems to not be interned. Let us quickly confirm the behavior with the following code:" + ] + }, + { + "cell_type": "code", + "execution_count": 46, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "True\n", + "False\n" + ] + } + ], + "source": [ + "print(hello1_func() is hello2_func())\n", + "print(hello1_func() is hello3_func())" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Finally, to show that this hypothesis is the answer to this rather unexpected observation, let us `intern` the value manually:" + ] + }, + { + "cell_type": "code", + "execution_count": 47, + "metadata": {}, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "True\n" + ] + } + ], + "source": [ + "import sys\n", + "\n", + "print(hello1_func() is sys.intern(hello3_func()))" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "
\n", + "
\n", + "
\n", + "
\n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "# Changelog" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### 06/09/2018\n", + "- pep8 spacing\n", + "- fixed minor typos\n", + "- fixed minor markdown formatting\n", + "- fixed broken page jumps\n", + "\n", + "#### 07/16/2014\n", + "- slight change of wording in the [lambda-closure section](#lambda_closure)\n", + "\n", + "#### 05/24/2014\n", + "- new section: unorderable types in Python 2\n", + "- table of contents for the Python 2 vs. Python 3 topic\n", + " \n", + "#### 05/03/2014\n", + "- new section: else clauses: conditional vs. completion\n", + "- new section: Interning of compile-time constants vs. run-time expressions\n", + "\n", + "#### 05/02/2014\n", + "- new section in Python 3.x and Python 2.x key differences: for-loop leak\n", + "- new section: Metaclasses - What creates a new instance of a class? \n", + "\n", + "#### 05/01/2014\n", + "- new section: keyword argument unpacking syntax\n", + "\n", + "#### 04/27/2014\n", + "- minor fixes of typos \n", + "- new section: \"Only the first clause of generators is evaluated immediately\"" + ] + }, + { + "cell_type": "code", + "execution_count": null, + "metadata": {}, + "outputs": [], + "source": [] + } + ], + "metadata": { + "kernelspec": { + "display_name": "Python 3", + "language": "python", + "name": "python3" + }, + "language_info": { + "codemirror_mode": { + "name": "ipython", + "version": 3 + }, + "file_extension": ".py", + "mimetype": "text/x-python", + "name": "python", + "nbconvert_exporter": "python", + "pygments_lexer": "ipython3", + "version": "3.6.4" + }, + "latex_envs": { + "LaTeX_envs_menu_present": true, + "autoclose": false, + "autocomplete": true, + "bibliofile": "biblio.bib", + "cite_by": "apalike", + "current_citInitial": 1, + "eqLabelWithNumbers": true, + "eqNumInitial": 1, + "hotkeys": { + "equation": "Ctrl-E", + "itemize": "Ctrl-I" + }, + "labels_anchors": false, + "latex_user_defs": false, + "report_style_numbering": false, + "user_envs_cfg": false + }, + "toc": { + "base_numbering": 1, + "nav_menu": {}, + "number_sections": true, + "sideBar": true, + "skip_h1_title": false, + "title_cell": "Table of Contents", + "title_sidebar": "Contents", + "toc_cell": false, + "toc_position": {}, + "toc_section_display": true, + "toc_window_display": false + } + }, + "nbformat": 4, + "nbformat_minor": 1 +} diff --git a/tutorials/numpy_nan_quickguide.ipynb b/tutorials/numpy_nan_quickguide.ipynb new file mode 100644 index 0000000..acbbeed --- /dev/null +++ b/tutorials/numpy_nan_quickguide.ipynb @@ -0,0 +1,770 @@ +{ + "metadata": { + "name": "", + "signature": "sha256:b2597ea4263c11dd6774b227e7a3a5626197c4863e6895002657fd55d02b55d9" + }, + "nbformat": 3, + "nbformat_minor": 0, + "worksheets": [ + { + "cells": [ + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to python_reference](https://github.com/rasbt/python_reference)]" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%load_ext watermark" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 1 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%watermark -v -p numpy -d -u" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "Last updated: 31/07/2014 \n", + "\n", + "CPython 3.4.1\n", + "IPython 2.1.0\n", + "\n", + "numpy 1.8.1\n" + ] + } + ], + "prompt_number": 2 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[More information](https://github.com/rasbt/watermark) about the `watermark` magic command extension." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 1, + "metadata": {}, + "source": [ + "Quick guide for dealing with missing numbers in NumPy" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "This is just a quick overview of how to deal with missing values (i.e., \"NaN\"s for \"Not-a-Number\") in NumPy and I am happy to expand it over time. Yes, and there will also be a separate one for pandas some time!\n", + "\n", + "I would be happy to hear your comments and suggestions. \n", + "Please feel free to drop me a note via\n", + "[twitter](https://twitter.com/rasbt), [email](mailto:bluewoodtree@gmail.com), or [google+](https://plus.google.com/+SebastianRaschka).\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 2, + "metadata": {}, + "source": [ + "Sections" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "- [Sample data from a CSV file](#Sample-data-from-a-CSV-file)\n", + "- [Determining if a value is missing](#Determining-if-a-value-is-missing)\n", + "- [Counting the number of missing values](#Counting-the-number-of-missing-values)\n", + "- [Calculating the sum of an array that contains NaNs](#Calculating the sum of an array that contains NaNs)\n", + "- [Removing all rows that contain missing values](#Removing-all-rows-that-contain-missing-values)\n", + "- [Convert missing values to 0](#Convert-missing-values-to-0)\n", + "- [Converting certain numbers to NaN](#Converting-certain-numbers-to-NaN)\n", + "- [Remove all missing elements from an array](#Remove-all-missing-elements-from-an-array)\n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 2, + "metadata": {}, + "source": [ + "Sample data from a CSV file" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#Sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Let's assume that we have a CSV file with missing elements like the one shown below." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "
" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%%file example.csv\n", + "1,2,3,4\n", + "5,6,,8\n", + "10,11,12," + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "Writing example.csv\n" + ] + } + ], + "prompt_number": 3 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "The `np.genfromtxt` function has a `missing_values` parameters which translates missing values into `np.nan` objects by default. This allows us to construct a new NumPy `ndarray` object, even if elements are missing." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "
" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "import numpy as np\n", + "ary = np.genfromtxt('./example.csv', delimiter=',')\n", + "\n", + "print('%s x %s array:\\n' %(ary.shape[0], ary.shape[1]))\n", + "print(ary)" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "3 x 4 array:\n", + "\n", + "[[ 1. 2. 3. 4.]\n", + " [ 5. 6. nan 8.]\n", + " [ 10. 11. 12. nan]]\n" + ] + } + ], + "prompt_number": 4 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 2, + "metadata": {}, + "source": [ + "Determining if a value is missing" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#Sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "A handy function to test whether a value is a `NaN` or not is to use the `np.isnan` function." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "np.isnan(np.nan)" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "pyout", + "prompt_number": 5, + "text": [ + "True" + ] + } + ], + "prompt_number": 5 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "It is especially useful to create boolean masks for the so-called \"fancy indexing\" of NumPy arrays, which we will come back to later." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "np.isnan(ary)" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "pyout", + "prompt_number": 6, + "text": [ + "array([[False, False, False, False],\n", + " [False, False, True, False],\n", + " [False, False, False, True]], dtype=bool)" + ] + } + ], + "prompt_number": 6 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 2, + "metadata": {}, + "source": [ + "Counting the number of missing values" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#Sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "In order to find out how many elements are missing in our array, we can use the `np.isnan` function that we have seen in the previous section. " + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "np.count_nonzero(np.isnan(ary))" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "pyout", + "prompt_number": 7, + "text": [ + "2" + ] + } + ], + "prompt_number": 7 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "If we want to determine the number of non-missing elements, we can simply revert the returned `Boolean` mask via the handy \"tilde\" sign." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "np.count_nonzero(~np.isnan(ary))" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "pyout", + "prompt_number": 8, + "text": [ + "10" + ] + } + ], + "prompt_number": 8 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 2, + "metadata": {}, + "source": [ + "Calculating the sum of an array that contains `NaN`s" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#Sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "As we will find out via the following code snippet, we can't use NumPy's regular `sum` function to calculate the sum of an array." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "np.sum(ary)" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "pyout", + "prompt_number": 9, + "text": [ + "nan" + ] + } + ], + "prompt_number": 9 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Since the `np.sum` function does not work, use `np.nansum` instead:" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "print('total sum:', np.nansum(ary))" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "total sum: 62.0\n" + ] + } + ], + "prompt_number": 10 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "print('column sums:', np.nansum(ary, axis=0))" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "column sums: [ 16. 19. 15. 12.]\n" + ] + } + ], + "prompt_number": 11 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "print('row sums:', np.nansum(ary, axis=1))" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "row sums: [ 10. 19. 33.]\n" + ] + } + ], + "prompt_number": 12 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 2, + "metadata": {}, + "source": [ + "Removing all rows that contain missing values" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#Sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Here, we will use the `Boolean mask` again to return only those rows that DON'T contain missing values. And if we want to get only the rows that contain `NaN`s, we could simply drop the `~`." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "ary[~np.isnan(ary).any(1)]" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "pyout", + "prompt_number": 14, + "text": [ + "array([[ 1., 2., 3., 4.]])" + ] + } + ], + "prompt_number": 14 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 2, + "metadata": {}, + "source": [ + "Convert missing values to 0" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#Sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Certain operations, algorithms, and other analyses might not work with `NaN` objects in our data array. But that's not a problem: We can use the convenient `np.nan_to_num` function will convert it to the value 0." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "ary0 = np.nan_to_num(ary)\n", + "ary0" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "pyout", + "prompt_number": 15, + "text": [ + "array([[ 1., 2., 3., 4.],\n", + " [ 5., 6., 0., 8.],\n", + " [ 10., 11., 12., 0.]])" + ] + } + ], + "prompt_number": 15 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 2, + "metadata": {}, + "source": [ + "Converting certain numbers to NaN" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#Sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Vice versa, we can also convert any number to a `np.NaN` object. Here, we use the array that we created in the previous section and convert the `0`s back to `np.nan` objects." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "ary0[ary0==0] = np.nan\n", + "ary0" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "pyout", + "prompt_number": 16, + "text": [ + "array([[ 1., 2., 3., 4.],\n", + " [ 5., 6., nan, 8.],\n", + " [ 10., 11., 12., nan]])" + ] + } + ], + "prompt_number": 16 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 2, + "metadata": {}, + "source": [ + "Remove all missing elements from an array" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#Sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "This is one is a little bit more tricky. We can remove missing values via a combination of the `Boolean` mask and fancy indexing, however, this will have the disadvantage that it will flatten our array (we can't just punch holes into a NumPy array)." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "ary[~np.isnan(ary)]" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "pyout", + "prompt_number": 17, + "text": [ + "array([ 1., 2., 3., 4., 5., 6., 8., 10., 11., 12.])" + ] + } + ], + "prompt_number": 17 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Thus, this is a method that would better work on individual rows:" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "x = np.array([1,2,np.nan])\n", + "\n", + "x[~np.isnan(np.array(x))]" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "pyout", + "prompt_number": 21, + "text": [ + "array([ 1., 2.])" + ] + } + ], + "prompt_number": 21 + } + ], + "metadata": {} + } + ] +} \ No newline at end of file diff --git a/tutorials/python_data_entry_point.ipynb b/tutorials/python_data_entry_point.ipynb new file mode 100644 index 0000000..ffbd6ad --- /dev/null +++ b/tutorials/python_data_entry_point.ipynb @@ -0,0 +1,1801 @@ +{ + "metadata": { + "name": "", + "signature": "sha256:cf3234a0ec4044266603b3f223b30b7b1595acce96b566935292bc57addfbdc5" + }, + "nbformat": 3, + "nbformat_minor": 0, + "worksheets": [ + { + "cells": [ + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[Sebastian Raschka](http://sebastianraschka.com) \n", + "\n", + "- [Open in IPython nbviewer](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/tutorials/python_data_entry_point.ipynb?create=1) \n", + "\n", + "- [Link to this IPython notebook on Github](https://github.com/rasbt/python_reference/blob/master/tutorials/python_data_entry_point.ipynb) \n", + "\n", + "- [Link to the GitHub Repository pattern_classification](http://nbviewer.ipython.org/github/rasbt/pattern_classification/blob/master/python_howtos/)" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%load_ext watermark" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 1 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%watermark -a 'Sebastian Raschka' -v -d -p numpy,scipy,matplotlib,scikit-learn" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "Sebastian Raschka 10/07/2014 \n", + "\n", + "CPython 3.4.1\n", + "IPython 2.1.0\n", + "\n", + "numpy 1.8.1\n", + "scipy 0.14.0\n", + "matplotlib 1.3.1\n", + "scikit-learn 0.15.0b1\n" + ] + } + ], + "prompt_number": 2 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[More information](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/ipython_magic/watermark.ipynb) about the `watermark` magic command extension." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "I would be happy to hear your comments and suggestions. \n", + "Please feel free to drop me a note via\n", + "[twitter](https://twitter.com/rasbt), [email](mailto:bluewoodtree@gmail.com), or [google+](https://plus.google.com/+SebastianRaschka).\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 1, + "metadata": {}, + "source": [ + "Entry point: Data " + ] + }, + { + "cell_type": "heading", + "level": 3, + "metadata": {}, + "source": [ + "- Using Python's sci-packages to prepare data for Machine Learning tasks and other data analyses" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "In this short tutorial I want to provide a short overview of some of my favorite Python tools for common procedures as entry points for general pattern classification and machine learning tasks, and various other data analyses. " + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 1, + "metadata": {}, + "source": [ + "Sections" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "- [Installing Python packages](#Installing-Python-packages)\n", + "\n", + "- [About the dataset](#About-the-dataset)\n", + "\n", + "- [Downloading and saving CSV data files from the web](#Downloading-and-savin-CSV-data-files-from-the-web)\n", + "\n", + "- [Reading in a dataset from a CSV file](#Reading-in-a-dataset-from-a-CSV-file)\n", + "\n", + "- [Visualizating of a dataset](#Visualizating-of-a-data)\n", + "\n", + " - [Histograms](#Histograms)\n", + "\n", + " - [Scatterplots](#Scatterplots)\n", + "\n", + "- [Splitting into training and test dataset](#Splitting-into-training-and-test-dataset)\n", + "\n", + "- [Feature Scaling](#Feature-Scaling)\n", + "\n", + " - [Standardization](#Standardization)\n", + " \n", + " - [Min-Max scaling (Normalization)](#Min-Max-scaling-Normalization)\n", + "\n", + "- [Linear Transformation: Principal Component Analysis (PCA)](#PCA)\n", + "\n", + "- [Linear Transformation: Linear Discrciminant Analysis (LDA)](#MDA)\n", + "\n", + "- [Simple Supervised Classification](#Simple-Supervised-Classification)\n", + "\n", + " - [Linear Discriminant Analysis as simple linear classifier](#Linear-Discriminant-Analysis-as-simple-linear-classifier)\n", + " \n", + " - [Classification Stochastic Gradient Descent (SGD)](#SGD)\n", + "\n", + "- [Saving the processed datasets](#Saving-the-processed-datasets)\n", + "\n", + " - [Pickle](#Pickle)\n", + "\n", + " - [Comma Separated Values (CSV)](#Comma-Separated-Values)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 2, + "metadata": {}, + "source": [ + "Installing Python packages" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top]](#Sections)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "**In this section want to recommend a way for installing the required Python-packages packages if you have not done so, yet. Otherwise you can skip this part.**\n", + "\n", + "The packages we will be using in this tutorial are:\n", + "\n", + "- [NumPy](http://www.numpy.org)\n", + "- [SciPy](http://www.scipy.org)\n", + "- [matplotlib](http://matplotlib.org)\n", + "- [scikit-learn](http://scikit-learn.org/stable/)\n", + "\n", + "Although they can be installed step-by-step \"manually\", but I highly recommend you to take a look at the [Anaconda](https://store.continuum.io/cshop/anaconda/) Python distribution for scientific computing.\n", + "\n", + "Anaconda is distributed by Continuum Analytics, but it is completely free and includes more than 195+ packages for science and data analysis as of today.\n", + "The installation procedure is nicely summarized here: http://docs.continuum.io/anaconda/install.html\n", + "\n", + "If this is too much, the [Miniconda](http://conda.pydata.org/miniconda.html) might be right for you. Miniconda is basically just a Python distribution with the Conda package manager, which let's us install a list of Python packages into a specified `conda` environment from the Shell terminal, e.g.,\n", + "\n", + "
$[bash]> conda create -n myenv python=3\n", + "$[bash]> source activate myenv\n", + "$[bash]> conda install -n myenv numpy scipy matplotlib scikit-learn\n", + "\n", + "When we start \"python\" in your current shell session now, it will use the Python distribution in the virtual environment \"myenv\" that we have just created. To un-attach the virtual environment, you can just use\n", + "
$[bash]> source deactivate myenv\n", + "\n", + "**Note:** environments will be created in ROOT_DIR/envs by default, you can use the `-p` instead of the `-n` flag in the conda commands above in order to specify a custom path.\n", + "\n", + "**I find this procedure very convenient, especially if you are working with different distributions and versions of Python with different modules and packages installed and it is extremely useful for testing your own modules.**" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 2, + "metadata": {}, + "source": [ + "About the dataset" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top]](#Sections)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "For the following tutorial, we will be working with the free \"Wine\" Dataset that is deposited on the UCI machine learning repository \n", + "(http://archive.ics.uci.edu/ml/datasets/Wine).\n", + "\n", + "
\n", + "\n", + "\n", + "**Reference:** \n", + "Forina, M. et al, PARVUS - An Extendible Package for Data\n", + "Exploration, Classification and Correlation. Institute of Pharmaceutical\n", + "and Food Analysis and Technologies, Via Brigata Salerno, \n", + "16147 Genoa, Italy.\n", + "\n", + "Bache, K. & Lichman, M. (2013). UCI Machine Learning Repository [http://archive.ics.uci.edu/ml]. Irvine, CA: University of California, School of Information and Computer Science.\n", + "\n", + "" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "The Wine dataset consists of 3 different classes where each row correspond to a particular wine sample.\n", + "\n", + "The class labels (1, 2, 3) are listed in the first column, and the columns 2-14 correspond to the following 13 attributes (features):\n", + "\n", + "1) Alcohol \n", + "2) Malic acid \n", + "3) Ash \n", + "4) Alcalinity of ash \n", + "5) Magnesium \n", + "6) Total phenols \n", + "7) Flavanoids \n", + "8) Nonflavanoid phenols \n", + "9) Proanthocyanins \n", + "10) Color intensity \n", + "11) Hue \n", + "12) OD280/OD315 of diluted wines \n", + "13) Proline \n", + "\n", + "An excerpt from the wine_data.csv dataset:\n", + " \n", + "
1,14.23,1.71,2.43,15.6,127,2.8,3.06,.28,2.29,5.64,1.04,3.92,1065\n", + "1,13.2,1.78,2.14,11.2,100,2.65,2.76,.26,1.28,4.38,1.05,3.4,1050\n", + "[...]\n", + "2,12.37,.94,1.36,10.6,88,1.98,.57,.28,.42,1.95,1.05,1.82,520\n", + "2,12.33,1.1,2.28,16,101,2.05,1.09,.63,.41,3.27,1.25,1.67,680\n", + "[...]\n", + "3,12.86,1.35,2.32,18,122,1.51,1.25,.21,.94,4.1,.76,1.29,630\n", + "3,12.88,2.99,2.4,20,104,1.3,1.22,.24,.83,5.4,.74,1.42,530" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 2, + "metadata": {}, + "source": [ + "Downloading and saving CSV data files from the web" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top]](#Sections)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Usually, we have our data stored locally on our disk in as a common text (or CSV) file with comma-, tab-, or whitespace-separated rows. Below is just an example for how you can CSV datafile from a HTML website directly into Python and optionally save it locally." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "import csv\n", + "import urllib\n", + "\n", + "url = 'https://raw.githubusercontent.com/rasbt/pattern_classification/master/data/wine_data.csv'\n", + "csv_cont = urllib.request.urlopen(url)\n", + "csv_cont = csv_cont.read() #.decode('utf-8')\n", + "\n", + "# Optional: saving the data to your local drive\n", + "with open('./wine_data.csv', 'wb') as out:\n", + " out.write(csv_cont)" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 3 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "**Note:** If you'd rather like to work with the data directly in `str`ing format, you could just apply the `.decode('utf-8')` method to the data that was read in byte-format by default.\n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 2, + "metadata": {}, + "source": [ + "Reading in a dataset from a CSV file" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top]](#Sections)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Since it is quite typical to have the input data stored locally, as mentioned above, we will use the [`numpy.loadtxt`](http://docs.scipy.org/doc/numpy/reference/generated/numpy.loadtxt.html) function now to read in the data from the CSV file. \n", + "(alternatively [`np.genfromtxt()`](http://docs.scipy.org/doc/numpy/reference/generated/numpy.genfromtxt.html) could be used in similar way, it provides some additional options)" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "import numpy as np\n", + "\n", + "# reading in all data into a NumPy array\n", + "all_data = np.loadtxt(open(\"./wine_data.csv\",\"r\"),\n", + " delimiter=\",\", \n", + " skiprows=0, \n", + " dtype=np.float64\n", + " )\n", + "\n", + "# load class labels from column 1\n", + "y_wine = all_data[:,0]\n", + "\n", + "# conversion of the class labels to integer-type array\n", + "y_wine = y_wine.astype(np.int64, copy=False)\n", + "\n", + "# load the 14 features\n", + "X_wine = all_data[:,1:]\n", + "\n", + "# printing some general information about the data\n", + "print('\\ntotal number of samples (rows):', X_wine.shape[0])\n", + "print('total number of features (columns):', X_wine.shape[1])\n", + "\n", + "# printing the 1st wine sample\n", + "float_formatter = lambda x: '{:.2f}'.format(x)\n", + "np.set_printoptions(formatter={'float_kind':float_formatter})\n", + "print('\\n1st sample (i.e., 1st row):\\nClass label: {:d}\\n{:}\\n'\n", + " .format(int(y_wine[0]), X_wine[0]))\n", + "\n", + "# printing the rel.frequency of the class labels\n", + "print('Class label frequencies')\n", + "print('Class 1 samples: {:.2%}'.format(list(y_wine).count(1)/y_wine.shape[0]))\n", + "print('Class 2 samples: {:.2%}'.format(list(y_wine).count(2)/y_wine.shape[0]))\n", + "print('Class 3 samples: {:.2%}'.format(list(y_wine).count(3)/y_wine.shape[0]))" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "\n", + "total number of samples (rows): 178\n", + "total number of features (columns): 13\n", + "\n", + "1st sample (i.e., 1st row):\n", + "Class label: 1\n", + "[14.23 1.71 2.43 15.60 127.00 2.80 3.06 0.28 2.29 5.64 1.04 3.92 1065.00]\n", + "\n", + "Class label frequencies\n", + "Class 1 samples: 33.15%\n", + "Class 2 samples: 39.89%\n", + "Class 3 samples: 26.97%\n" + ] + } + ], + "prompt_number": 4 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 2, + "metadata": {}, + "source": [ + "Visualizating of a dataset" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top]](#Sections)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "There are endless way to visualize datasets for get an initial idea of how the data looks like. The most common ones are probably histograms and scatter plots." + ] + }, + { + "cell_type": "heading", + "level": 3, + "metadata": {}, + "source": [ + "Histograms" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top]](#Sections)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Histograms are a useful data to explore the distribution of each feature across the different classes. This could provide us with intuitive insights which features have a good and not-so-good inter-class separation. Below, we will plot a sample histogram for the \"Alcohol content\" feature for the three wine classes." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%matplotlib inline" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 5 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "from matplotlib import pyplot as plt\n", + "from math import floor, ceil # for rounding up and down\n", + "\n", + "plt.figure(figsize=(10,8))\n", + "\n", + "# bin width of the histogram in steps of 0.15\n", + "bins = np.arange(floor(min(X_wine[:,0])), ceil(max(X_wine[:,0])), 0.15)\n", + "\n", + "# get the max count for a particular bin for all classes combined\n", + "max_bin = max(np.histogram(X_wine[:,0], bins=bins)[0])\n", + "\n", + "# the order of the colors for each histogram\n", + "colors = ('blue', 'red', 'green')\n", + "\n", + "for label,color in zip(\n", + " range(1,4), colors):\n", + "\n", + " mean = np.mean(X_wine[:,0][y_wine == label]) # class sample mean\n", + " stdev = np.std(X_wine[:,0][y_wine == label]) # class standard deviation\n", + " plt.hist(X_wine[:,0][y_wine == label], \n", + " bins=bins, \n", + " alpha=0.3, # opacity level\n", + " label='class {} ($\\mu={:.2f}$, $\\sigma={:.2f}$)'.format(label, mean, stdev), \n", + " color=color)\n", + "\n", + "plt.ylim([0, max_bin*1.3])\n", + "plt.title('Wine data set - Distribution of alocohol contents')\n", + "plt.xlabel('alcohol by volume', fontsize=14)\n", + "plt.ylabel('count', fontsize=14)\n", + "plt.legend(loc='upper right')\n", + "\n", + "plt.show()" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "display_data", + "png": "iVBORw0KGgoAAAANSUhEUgAAAmgAAAH8CAYAAABl8FOBAAAABHNCSVQICAgIfAhkiAAAAAlwSFlz\nAAALEgAACxIB0t1+/AAAIABJREFUeJzs3Xt4lPWd///nhFPKSWJSQKOAgCfwANIKiJVYrKgUraxo\n8YBs3Srqr3zXw5dK6SrUA1irq19166pVCtha4roIS5VWDtJYuxRQlIMgUAggtSEYOQiJQH5/3JNh\nEmbCBCbkJnk+risXc58+877vucO88vnc9wxIkiRJkiRJkiRJkiRJkiRJkiRJkiRJkiRJkuqRbwEf\nH6Xnmg/ccpSeK+yWARelqa0bgNlx0/uBzmlqG2AH0CmN7aXia8BMoAT43WFsn+5jUNUk4MHD3HY+\n/h5IhyWjrguQjsAY4PdV5n2SZN61wJ+AM45CXQDl0Z9UrAe+XXulJHWkb+ydom3siP78nSBoXFJl\nvbOABSm2daj/k14BBtawzmTmc3B4aEXwehxN1wBtgeOB647yc6eiJudyOrc9HJ1I7TxK1QiC/zek\no86ApmPZO8AFQCQ6fQLQGOjBgXP7BKALhw4IdamcA/twtKXjeY8jCDbnAH8E/hu4uRbqaXSYbSZz\nNINDdToCqwmChdKjrn6fJElAU2AX0DM6fS3wEkHPyHlx81ZHH+cBG+O2Xw/cAywlGF56FWgWt/y7\nwAfA58C7wNnV1PIdguHTEuBpKvfOdAHmAluBImAqQagBmALsA74k6IW6Nzo/H9gSbe8doFs1zz0C\nWAtsB9YB18ct+wGwAtgGvAV0iM5fQBAIdkafd2g17SfTicS9FfcQ9KZVWM+BHsLzgUXAF9F1fhGd\nX8iB3rjtQJ/ofr0LPEFw7B7k4B6N/cCPCPa/CPg5B96cxxEc36r1NgIeBvYCu6PP+f/i2qvoVTwO\nmAz8I7oPY+PaHgEUAI8RHNt1wGUkdybBOfE5wZDv4Oj88UApUBat458TbHs+8F50208Jzq8mVY5B\nKjUD/JDgfNgOLOfA706y+gBeBp4B/ie63V+o3PN6AfBXgnN1IdA3btk8gnMwkQzgJ8CaaLuLgJNS\naHM+8DOC47+dYMg7O7os/jzaAfSOzk/2e0B0/dsI/p/4PLqvFcdkN8F5siO6LcAVBMduO7CJ4HyX\nJFUxF/jX6ONnCN7gHqoy78Xo4zwqB7S/EbzZtAeyCP4Dvy26rCfwGfBNgje44dH1myaoIYfgP+sh\nBG/+/wp8xYE3pi7AAII31RyCwPXvVeqoOsQ5AmgR3ebfgfcT7z4tCMLOqdHpdhwIc1cRDO+eTvBm\nOJYg8FRI1xBn1YDWOTr/9Oh0/P69R3AdGUBzDryBdkzQ1giC43hndH4miQPaHKANcDKwigPB+AES\nB7SK50gUHuKPyWSC3sAW0fpWxa0/giBU3UJwfowENpNYE4IQch9BD+/FBOfLaXF1Tk6yLQR/bJwf\nrbsjwXn6fw6j5qEEgaJXdLoLQVA5VH2TCALyNwjO76nAb6PLjicINTdE6/s+QZDJii6vLqD9X+BD\nDpy7Z0fbO1Sb8wnO664E58Q8YEJ0WaLzKJXfgxlAa4Jz6B8cGEa/mYOHOLcA/aKPj+NAyJUkxXkA\neD36+AOCN52BcfOWAjdFH+dxcECL7216FPhl9PEvCf5Kj/cxiS92Hw78ucq8jSR/Y/oesKRKHdVd\ng9aG4E2kVYJlLQjezIYQXGwe780qNWQQ9DieHJ2urYCWGZ1f0esRv3/vEPRs5aTQ1ghgQ5X1RnBw\nQLs0bvp24O3o43EcOqBVvQat4pg0IujZir9m8dboNhV1fBK3rHl027Yc7FsEb+rxfkNw7iaq81D+\nlQPnd01qnk3Q21jT+iYBz8ctuxxYGX18E8EfOfH+zIEh7uoC2sdU7qmrkEqbP4lbdjvBuQ6Jz6NU\nfg8uiFv+O+DH0ccjODigbSA4rq0T1C6ljdeg6Vi3ALiQ4K/rrxMMdb1H8B9uFtCd6q8/ix+K2w20\njD7uSDB08Xncz0kE17RVdSJBz0S8+CDYjmD4dBNBb9cUDgzJJJIBTCTo1fiCIOCUc3CogeCN5jqC\nHpxPCYahKnquOgJPxdVfHJ2fW81zx1vOgaGifodYN15F+9sSLLuFoGdmJcHQ1aBDtLXxEMurrlNI\n8HqkKtl1aDkEPUvxAbGQyscu/tz5MvpvSw52IgfvxwZSfx1OI3hdtxCcDw+T+Pw5VM0nEfx+pFpf\nxXEsJ+hNrhD/e3Ji9DmSbVudk6up51BtJvu9TSSV34Oqr2WLatr7J4JhzvUEvXl9qllXOmwGNB3r\n/kIwzPBDDgxbbCcIK7dG/63aC1OdijfsQoI3wqy4n5Yk/hiETznw1zgEQ17x048QXGd2VrTWm6j8\nu1c1JNwAXEkwLHoccEq0zWQXPv+BoBepPUGvxAtx+3BrlX1owcG9E8l0J+i1a0XlIaFDuZrgDX1V\ngmVrCHotv07QY/kaQc9fsqCUyoX8Hao8rhhq3EXQs1WhfQ3a3kowvNqpSttVg3gqKs6P+NevYw3a\n+iXBsGZXgvNhLIn/7z5UzRujbaRaX7Ih23ibo+vGS3XbZPUcSZuJXtMj+T1I1N4igl7wrwPTgWkp\ntCPVmAFNx7rdBP9h3k3lnrKC6Lx3athexZvUCwS9UudH57Ug6O1J9Jf6LIIwczXBNTyjqBwGWhKE\nhe0Ef7X/3yrbf0YwNBu/filBD1QLgoCXTFuCa2xaELw57yIIgwDPEQwFVVyTdhyVbwao+ryHq+KY\ntQP+P+B+go9ASeRGgjc2CHqDygmGmIqi/x5OPfdy4Bq0URwI0e8TDEmfTLDvVWuqbv/3EbzxPkzw\nenQE7iK4/qqm/kLQKzOaoIcrj+AGlFdT3L4lQS/mlwTDl7cfZs0vEhyr8whes64EAe5Q9VV3R+Sb\nBD18wwjO/euiNf5P3DrJtn+R4MaPrtF1ziG4/uz3R9BmovPoUL8HVcX/MfQZQc9jxU0ZTQj+gDqO\n4Hjv4MDvmySpiooeqh5x84ZG5/0wbl4elYdOql77VfVi7YEEw3AVd8/9juRDKQMJeowq7uKMv/am\nG0GI3EFw7dndVeq4kqCX7/PoshYEf5lvj9Z4U3RfEl0v1p5gmKUkuv1cKl+DdCPBhdhfRJ/zxbhl\nt0X363OCz+KqqU4cuGNuJ8Gb2f9Q+ZowqHycp0TX2wF8RLDvFcYTXKC9jeDmgZs5eHi66rz9BKFw\nLUEP0mNU/sPzGYL9Ww38C8FxrFjeh+A12wY8GddexXFuE633HwTH7qcceONOVFuy1wiCc2A+weu0\njCBUVzjUTQLfIhgS3hF9zvFVnjv+eaurGYLX/ONoWx8C56ZQ38tUvh4zj8rnbz+C87uE4M7L+Ou5\nDnUX51iCO2C3A//LgWHMmrRZ9bWoOI8+J/gDC6r/Paj6usXvbxOCc7o42mYTglC6LdrW/1apTZIk\nSZIkSZIkSZIkSZIkqc4dE99Xdu6555YvXbq0rsuQJElKxVIq37hWY8fEx2wsXbqU8vJyf6r8PPDA\nA3VeQxh/PC4eE4+Lx8Xj4jGpyx8O3CF92I6JgCZJktSQGNAkSZJCxoB2DMvLy6vrEkLJ43Iwj0li\nHpfEPC6JeVwO5jGpPcfETQJAeXRMV5IkKdQikQgcYcZqnJ5SJEkNyfHHH8/nn39e12VIdSorK4tt\n27bVStv2oEmSaiwSieD/y2rokv0epKMHzWvQJEmSQsaAJkmSFDIGNEmSpJAxoEmSJIWMAU2SJClk\nDGiSJEkhY0CTJDUonTp1Ys6cOXVdRlJjxozhqaeequsyBPTu3ZsVK1bUyXP7QbWSpLTIz59NcXFZ\nrbWfnd2UoUMHHnE7kUik4nOqat0zzzzDpEmTWLZsGcOGDePll1+udv2ioiKmTJnC2rVra62mDz74\ngKlTp/KLX/wiNu+NN95g586drF27lpycHO64445K2+zfv5+srCwyMg7063znO99h2rRpsemFCxcy\nZ84cxowZU2u1JzN9+nRWrFhBRkYGubm53HTTTdWuX7XWnTt38vOf/5yTTz6Z7du3c/fddxOJRLj3\n3nu5//77ee21147GblRiQJMkpUVxcRm5uYNrrf3Nm2fWWtu1JTc3l3/7t39j9uzZ7N69+5DrT5o0\niUGDBtGsWbNaqeeJJ56goKCA4447LjavpKSE6667jpKSEpo1a0ZOTg6DBg2iY8eOsXU2bNjAL3/5\nSy644AIikQjTp0/n0ksvjS3fv38/999/PxdccEGt1F2dL774ggcffJDFixcD0LdvXy6//HJycnIS\nrp+o1lGjRvHAAw/QsWNHunfvzjXXXEPHjh0ZPHgwI0eO5LPPPqNdu3ZHZX8qOMQpSaqXNm7cyJAh\nQ2jbti05OTmMGjXqoHUmTpxI165dad26Nd27d2f69OmVlj/66KOcdNJJtG7dmjPOOIO5c+dWO7+q\nq6++mquuuors7OyUan7rrbfo379/Dfc0dXfffTdXXXVVpXlt2rRh8eLFZGZmEolE2Lt370Gfjt+s\nWTO+973v0alTJ1q3bk2TJk0488wzY8vz8/O55JJL6uTbJRYsWEC3bt1i0+eeey7z5s1Lun7VWtet\nW8enn34aC6R/+MMfYo8zMzPp1asXs2fPrsU9SMweNElSvbNv3z6++93vcskll/DKK6+QkZER62GJ\n17VrVwoKCmjfvj3Tpk3jxhtvZM2aNbRv355Vq1bx7LPPsmjRItq3b09hYSF79+5NOr86qQaXjz76\niNNPP71G+7pu3TpeeOGFpMv79OlTKZQlqqV79+4AFBQUkJeXR6dOnSotP/HEE2OP//M//5O77ror\nNl1UVESjRo34+te/zq5du2pUe3VS3a9NmzbRpk2b2Pw2bdrwySefJNwmUa1z586lTZs2TJkyhZKS\nElq1asWIESNi25x55pksXbo0PTtVAwY0SVK9s3DhQrZs2cJjjz0Wu24q0fDbNddcE3t87bXXMmHC\nBBYuXMiVV15Jo0aNKC0tZfny5WRnZ9OhQwcA1qxZk3B+dVK95q0iIFTYt28f/fv3p6CgAIBbbrmF\nMWPG0LVr19g6nTt3ZsKECSm1X10tr7/+Ovn5+Tz++ONJt922bRtbt26tNAT7+uuvc+uttzJ58uSU\nawBYvXo1P/3pTykqKmLRokXk5eUxaNAgRo4cCaS+XyUlJWRmZsammzZtys6dOxOum6jWzz77jGXL\nlvHqq68C8K1vfYt+/fpx6qmnAtCqVSu2bNlSo31LB4c4JUn1zsaNG+nYsWOli9oTmTx5Mj179iQr\nK4usrCyWLVtGcXExEPSuPfnkk4wbN4527doxbNgwtmzZknR+dVLtQcvKymLHjh2x6ffeey823FZe\nXs57771XKZwdjmS1DBkyhBdeeIHLL7+c9evXJ1znd7/7XaWhzb/85S/07t076ZeGJ7Nt2zZGjhzJ\n5MmTmTdvHgMGDGDq1KmxcFYTrVq1qvTcu3fv5vjjjz9ovWS1tm7dmrPPPjs23aFDB/7whz/Eprdv\n305WVlaN6zpS9qBJkuqdk08+mcLCQvbt20ejRo0SrrNhwwZuvfVW5s6dS9++fYlEIvTs2bPSm/ew\nYcMYNmwYO3bs4LbbbuPHP/4xkydPTjo/mVR70M455xxWrVpFr169gOCatIEDgztX33///UpBokJN\nhzir1jJr1iweeeQR3n33XVq2bEnbtm157bXXuPfeew9qa968eQwfPjw2/de//pUvv/yS2bNn8+67\n77J7925mzJjBlVdeWe1+Pvvss9x5552xnq/S0lKaN29+WPvVpUsXFi1aFJu/detWzjvvvIPWT1Tr\nG2+8Qffu3fnTn/4UWy8jI4P9+/fHpleuXFlpn48WA5okqd7p3bs3J5xwAvfddx/jx48nIyODJUuW\nVBrm3LVrF5FIhJycHPbv38/kyZNZtmxZbPnq1avZtGkT/fr1o1mzZmRmZlJeXp50fiL79u3jq6++\nYu/evezbt4/S0lIaN26cNDReccUVvPPOO1x//fUAzJ49m+9///tAEKQGDBhwUACq6RBn1VobNWpE\nXl5ebNnGjRs555xzAFi7di2dO3eOhbpPPvmEr33ta7Ftf/SjH8Uejxs3jkgkEqttxIgRRCKRhB8t\nsmPHjtiF/cuXL6d79+40adKk0jqp7tdFF13E6NGjY9NLlizh0UcfPaj+RLVeddVV7Nmzh5/85Cex\nZWvXrmXcuHEA7NmzhyVLljBlypRD1pFuBjRJUlpkZzet1Y/CyM5umvK6GRkZzJw5k1GjRtGhQwci\nkQg33HBDpYDWrVs37rnnHvr27UtGRgbDhw/nwgsvjC0vLS1lzJgxrFy5kiZNmtCvXz+ef/55ioqK\nEs5P5MEHH+RnP/tZbHrq1KmMGzeO+++/P+H6w4cPp0ePHuzZs4cdO3ZQWFjIjBkzKCwspHnz5hQV\nFdG5c+eUj0NVzzzzDNOmTWPjxo2MHz+eu+66i8suu4x169bx9NNPs2HDBsaOHRv7CI2hQ4fyq1/9\nip49ewJw/PHHk5ube1C706ZNY8aMGUQikdjHVGzcuDEWNKu6/fbbmTFjBitWrGDTpk1MnDjxsPep\nRYsWjB49moceeoj9+/czevRo2rZtm7D+ZLVWvCb79+/nzjvvpEuXLgDMnDmTiy++mPbt2x92fYfr\n6HxS35Err4tbdyVJidX0miOlbuzYsbRt25bs7GxWrlzJww8/XNcl1VhZWRk9e/bkww8/TNpbeCzo\n06cPL730UqWP8YiX7Pcg2uN4RBnLgCZJqjEDWu0bNWoUN998c+x6NIWPAc2AJkmhYkCTajeg+TEb\nkiRJIWNAkyRJChkDmiRJUsgY0CRJkkLGgCZJkhQyBjRJkqSQMaBJkiSFjAFNkiQpZAxokiRJIWNA\nkyQ1KJ06dWLOnDl1XUZSY8aM4amnnqrrMgT07t2bFStW1MlzN66TZ5Uk1Tuz8/MpKy6utfabZmcz\ncOjQI24nEolUfBVPrSorK+P2229nzpw5bNu2jS5dujBhwgQuu+yypNsUFRUxZcoU1q5dW2t1ffDB\nB0ydOpVf/OIXsXm/+c1v2LJlCwsXLuTqq6/m+9///kHbJVtn5syZbNq0iT179tCxY0eGDBlSa7Un\nM336dFasWEFGRga5ubncdNNNCdfr0qULmzZtok2bNjz22GMMHz4cgDfeeIOdO3eydu1acnJyuOOO\nOwC49957uf/++3nttdeO2r5UMKBJktKirLiYwbm5tdb+zM2ba63t2rB37146dOjAggUL6NChA7Nm\nzeLaa6/lo48+omPHjgm3mTRpEoMGDaJZs2a1UtMTTzxBQUEBxx13XGzemjVrKC4u5p577mHr1q2c\neuqp9O7dm1NOOeWQ6zRu3JhVq1Zx7733AvAv//IvXHrppbRs2bJW6k/kiy++4MEHH2Tx4sUA9O3b\nl8svv5ycnJyD1r3vvvsYOHAgJ554Io0bBxGopKSE6667jpKSEpo1a0ZOTg6DBg2iY8eODB48mJEj\nR/LZZ5/Rrl27o7ZP4BCnJKme2rhxI0OGDKFt27bk5OQwatSog9aZOHEiXbt2pXXr1nTv3p3p06dX\nWv7oo49y0kkn0bp1a8444wzmzp1b7fx4zZs354EHHqBDhw4ADBo0iFNOOYUlS5Ykrfmtt96if//+\nR7Lb1br77ru56qqrKs1bvnw5P//5zwHIycmha9eusbBzqHW2bt3K22+/TVlZGQAtWrSgadOmtVZ/\nIgsWLKBbt26x6XPPPZd58+YlXLdp06Z06NAhFs4A2rRpw+LFi8nMzCQSibB3797YF6BnZmbSq1cv\nZs+eXbs7kYA9aJKkemffvn1897vf5ZJLLuGVV14hIyPjoNAB0LVrVwoKCmjfvj3Tpk3jxhtvZM2a\nNbRv355Vq1bx7LPPsmjRItq3b09hYSF79+5NOv9QPvvsM1avXk337t2TrvPRRx9x+umn12hf161b\nxwsvvJB0eZ8+fSqFsorwUeGKK67gzTffjC3bsmULXbt2TWmdHj16sH//fr75zW9y6623cumll6Yt\noKW6XxVDlhXatGnDJ598knCbv/71r5SWlrJ9+3ZOO+00rrzySoDYa1JQUEBeXh6dOnWKbXPmmWey\ndOnSNOxRzRjQJEn1zsKFC9myZQuPPfYYGRnBYNEFF1xw0HrXXHNN7PG1117LhAkTWLhwIVdeeSWN\nGjWitLSU5cuXk52dHesJW7NmTcL51fnqq6+44YYbGDFiBKeddlrS9UpKSmjVqlVset++ffTv35+C\nggIAbrnlFsaMGVMpQHXu3JkJEyYcsoYKVa+/a9KkCWeddRYAs2bN4hvf+AY9evRIeZ377ruPCRMm\ncO+99/Lkk0+mXMfq1av56U9/SlFREYsWLSIvL49BgwYxcuTIGu1XSUkJmZmZsemmTZuyc+fOhOsO\nGDCAq6++GoAePXpw0UUXxcLd66+/Tn5+Po8//nilbVq1asWWLVtS3q90cYhTklTvbNy4kY4dO8bC\nWTKTJ0+mZ8+eZGVlkZWVxbJlyyiO3ujQtWtXnnzyScaNG0e7du0YNmxYrOco0fxk9u/fz0033URm\nZibPPPNMtfVkZWWxY8eO2PR7770Xu16tvLyc995776DerZqq2oNWoaSkhEmTJjF16tSk21ZdZ/Xq\n1cyfP58//vGPzJw5k4ceeog///nPh6xh27ZtjBw5ksmTJzNv3jwGDBjA1KlTY+GsJlq1alVpn3bv\n3s3xxx+fcN34nsSsrCzmz58fmx4yZAgvvPACl19+OevXr4/N3759O1lZWTWu60jZgyZJqndOPvlk\nCgsL2bdvH40aNUq4zoYNG7j11luZO3cuffv2JRKJ0LNnz0pv9sOGDWPYsGHs2LGD2267jR//+MdM\nnjw56fyqysvLueWWWygqKuL3v/990loqnHPOOaxatYpevXoBwTVpAwcOBOD999/n7LPPPmibmg5x\nJrqDtby8nIkTJ/Liiy/SsmVLNmzYcNCNDFXXWb9+PTNnzmRo9M7aSy65hF//+tcUFBQk7K2M9+yz\nz3LnnXfGer5KS0tp3rz5Ye1Xly5dWLRoUWz+1q1bOe+88w5af+rUqcyYMYNp06YBsGvXLho3bsys\nWbN45JFHePfdd2nZsiVt27bltddei934sHLlytjdnkeTAU2SVO/07t2bE044gfvuu4/x48eTkZHB\nkiVLKgWHXbt2EYlEyMnJYf/+/UyePJlly5bFlq9evZpNmzbRr18/mjVrRmZmJuXl5UnnJ3L77bfz\n8ccf8/bbb6d0Z+YVV1zBO++8w/XXXw/A7NmzYx9nMWvWLAYMGMCMGTNi105BzYc4E9X69NNPM3To\nUPbs2cPChQvZvXs3HTt2ZO3atXTu3JlIJJJwnVNOOYVly5bFgmNpaSl9+vQBYMSIEUQiEV5++eWD\nnm/Hjh2xC/uXL19O9+7dadKkSaV1Ut2viy66iNGjR8emlyxZwqOPPgpQqf5OnTrFeui+/PJLioqK\n+Pa3v82CBQvIy8uLHZuNGzdyzjnnALBnzx6WLFnClClTDllHuhnQJElp0TQ7u1Y/CqNpdnbK62Zk\nZDBz5kxGjRpFhw4diEQi3HDDDZUCWrdu3bjnnnvo27cvGRkZDB8+nAsvvDC2vLS0lDFjxrBy5Uqa\nNGlCv379eP755ykqKko4v6oNGzbw/PPPk5mZSfv27WPzn3/+eYYNG5aw7uHDh9OjRw/27NnDjh07\nKCwsZMaMGRQWFtK8eXOKioro3LlzysehqmeeeYZp06axceNGxo8fz1133cWHH37IXXfdFQtukUiE\nwsJCAIYOHcqvfvUrdu3alXCd/v3789RTT/HII4/QokUL2rRpw8033wzApk2bku7n7bffzowZM1ix\nYgWbNm1i4sSJh71PLVq0YPTo0Tz00EPs37+f0aNH07Zt20r19+zZkwsvvJBXXnmFJ598kg0bNvDq\nq6/SvHlzLrvsMtatW8fTTz/Nhg0bGDt2LJdeeikQfMbbxRdfXOn1O1pq/5P60qM82V8nkqSjLxKJ\nJO010pEZO3Ysbdu2JTs7m5UrV/Lwww/XdUk1VlZWRs+ePfnwww8POawbZn369OGll16q9DEe8ZL9\nHkSHkY8oYxnQJEk1ZkCrfaNGjeLmm2+OXY+m8DGgGdAkKVQMaFLtBjQ/ZkOSJClkDGiSJEkhY0CT\nJEkKGQOaJElSyBjQJEmSQsaAJkmSFDIGNEmSpJAxoEmSJIWMAU2SJClkDGiSpAalU6dOzJkzp67L\nSGrMmDE89dRTdV2GgN69e7NixYo6ee7GdfKskqR6J/+NfIp3Ftda+9ktsxl61dAjbicSiVR8FU+t\nu/HGG5kzZw67du0iJyeHW265hbFjxyZdv6ioiClTprB27dpaq+mDDz5g6tSp/OIXv4jNe+ONN9i5\ncydr164lJyeHO+64I+Vtu3TpwqZNm2jTpg2PPfYYw4cPr7Xak5k+fTorVqwgIyOD3NxcbrrppoTr\nJav1N7/5DVu2bGHhwoVcffXVfP/73wfg3nvv5f777+e11147avtSwYAmSUqL4p3F5H4jt9ba37xo\nc621XVvGjBnDiy++SGZmJqtWraJ///706tWLyy67LOH6kyZNYtCgQTRr1qxW6nniiScoKCjguOOO\ni80rKSnhuuuuo6SkhGbNmpGTk8OgQYPo2LHjIbcFuO+++xg4cCAnnngijRsf/VjxxRdf8OCDD7J4\n8WIA+vbty+WXX05OTs5B6yaqdc2aNRQXF3PPPfewdetWTj31VHr37s0pp5zC4MGDGTlyJJ999hnt\n2rU7qvvlEKckqV7auHEjQ4YMoW3btuTk5DBq1KiD1pk4cSJdu3aldevWdO/enenTp1da/uijj3LS\nSSfRunVrzjjjDObOnVvt/Kq6d+9OZmZmbLpx48a0bds2ac1vvfUW/fv3P5zdTcndd9/NVVddVWle\nmzZtWLx4MZmZmUQiEfbu3ZvwC8ATbQvQtGlTOnToUCfhDGDBggV069YtNn3uuecyb968hOsmqnX5\n8uX8/Oc/ByAnJ4euXbvGwl5mZia9evVi9uzZtbgHidmDJkmqd/bt28d3v/tdLrnkEl555RUyMjJi\nb7rxunarePzEAAAgAElEQVTtSkFBAe3bt2fatGnceOONrFmzhvbt27Nq1SqeffZZFi1aRPv27Sks\nLGTv3r1J5ydzxx138Otf/5rS0lKeeeYZzjvvvKTrfvTRR5x++uk12td169bxwgsvJF3ep0+fSsEq\nUfjq3r07AAUFBeTl5dGpU6eEbSXa9q9//SulpaVs376d0047jSuvvLJG9SeT6n5VDFlWaNOmDZ98\n8knCbRLVesUVV/Dmm28Cwf5t2bKFrl27xrY588wzWbp0aVr2qSYMaJKkemfhwoVs2bKFxx57jIyM\nYLDoggsuOGi9a665Jvb42muvZcKECSxcuJArr7ySRo0aUVpayvLly8nOzqZDhw5AMCSWaH4y//Ef\n/8Gzzz7LO++8wzXXXMN5553H+eefn3DdkpISWrVqFZvet28f/fv3p6CgAIBbbrmFMWPGVAoQnTt3\nZsKECSkeGZJef/f666+Tn5/P448/XqNtBwwYwNVXXw1Ajx49uOiiiyoFpmRWr17NT3/6U4qKili0\naBF5eXkMGjSIkSNHAqnvV0lJSaVeyqZNm7Jz586E6yar9ayzzgJg1qxZfOMb36BHjx6xbVq1asWW\nLVsOWUe6OcQpSap3Nm7cSMeOHWPhLJnJkyfTs2dPsrKyyMrKYtmyZRQXBzc6dO3alSeffJJx48bR\nrl07hg0bFutdSTS/OpFIhLy8PIYOHcpvf/vbpOtlZWWxY8eO2PR7770XuxasvLyc9957r1I4OxyJ\nesEAhgwZwgsvvMDll1/O+vXrU942vncuKyuL+fPnH7KGbdu2MXLkSCZPnsy8efMYMGAAU6dOjYWz\nmmjVqlWlunbv3s3xxx+fcN3qai0pKWHSpElMnTq10jbbt28nKyurxnUdKXvQJEn1zsknn0xhYSH7\n9u2jUaNGCdfZsGEDt956K3PnzqVv375EIhF69uxZ6c1+2LBhDBs2jB07dnDbbbfx4x//mMmTJyed\nfyhfffUV2dnZSZefc845rFq1il69egHBNWkDBw4E4P333+fss88+aJuaDnFW7QWbNWsWjzzyCO++\n+y4tW7akbdu2vPbaa9x7770HtVV126lTpzJjxgymTZsGwK5du1K6Fu3ZZ5/lzjvvjPV8lZaW0rx5\n88Pary5durBo0aLY/K1btyYcRq6u1vLyciZOnMiLL75Iy5Yt2bBhQywYr1y5sk7uTDWgSZLqnd69\ne3PCCSdw3333MX78eDIyMliyZEmlYc5du3YRiUTIyclh//79TJ48mWXLlsWWr169mk2bNtGvXz+a\nNWtGZmYm5eXlSedXVVRUxJw5cxg8eDCZmZm8/fbb5Ofn8/bbbyet+4orruCdd97h+uuvB2D27Nmx\nj3yYNWsWAwYMYMaMGZWu86rpEGfVWhs1akReXl5s2caNGznnnHMAWLt2LZ07d44Fs6rbdurUKdbr\n9eWXX1JUVMS3v/1tAEaMGEEkEuHll18+qIYdO3bELuxfvnw53bt3p0mTJpXWSXW/LrroIkaPHh2b\nXrJkCY8++uhB9VdX69NPP83QoUPZs2cPCxcuZPfu3XTs2JE9e/awZMkSpkyZcsg60s2AJklKi+yW\n2bX6URjZLZP3PFWVkZHBzJkzGTVqFB06dCASiXDDDTdUCmjdunXjnnvuoW/fvmRkZDB8+HAuvPDC\n2PLS0lLGjBnDypUradKkCf369eP555+nqKgo4fyqIpEIzz33HLfffjvl5eWcdtppTJkyhW9+85tJ\n6x4+fDg9evRgz5497Nixg8LCQmbMmEFhYSHNmzenqKiIzp07p3wcqnrmmWeYNm0aGzduZPz48dx1\n111cdtllrFu3jqeffpoNGzYwduxYLr30UgCGDh3Kr371K3r27Jlw2wsvvJBXXnmFJ598kg0bNvDq\nq6/GesI2bdrEsGHDEtZx++23M2PGDFasWMGmTZuYOHHiYe9TixYtGD16NA899BD79+9n9OjRsTtl\n4+tPVmtBQQF33XVXLHxGIhEKCwsBmDlzJhdffDHt27c/7PoO19H5pL4jV55szFySdPRFIpGk1zLp\nyIwdO5a2bduSnZ3NypUrefjhh+u6pBorKyujZ8+efPjhh0mHmI8Fffr04aWXXqr0MR7xkv0eRHsc\njyhjGdAkSTVmQKt9o0aN4uabb45dj6bwMaAZ0CQpVAxoUu0GND9mQ5IkKWQMaJIkSSFjQJMkSQoZ\nA5okSVLIHM2A9hLwGfBR3LxxwCbg/ejPZUexHkmSpFA6mh9U+zLwNBD/XRjlwBPRH0nSMSIrKyvp\nl25LDUVtfkfn0QxofwI6JZjvb7gkHWO2bdtW1yVI9VoYrkH7EbAU+BXQpo5rkSRJqnN1/V2cvwR+\nFn38IPA4cEuiFceNGxd7nJeXF/tiV0mSpLo0f/585s+fn9Y2j/bwYidgJnB2DZf5TQKSJOmYUB++\nSeCEuMdXU/kOT0mSpAbpaA5x/hboD+QAG4EHgDygB8HdnH8DbjuK9UiSJIXSsXIHpUOckiTpmFAf\nhjglSZJUhQFNkiQpZAxokiRJIWNAkyRJChkDmiRJUsgY0CRJkkLGgCZJkhQyBjRJkqSQMaBJkiSF\njAFNkiQpZAxokiRJIWNAkyRJChkDmiRJUsgY0CRJkkLGgCZJkhQyBjRJkqSQMaBJkiSFjAFNkiQp\nZAxokiRJIWNAkyRJChkDmiRJUsgY0CRJkkLGgCZJkhQyBjRJkqSQMaBJkiSFjAFNkiQpZAxokiRJ\nIWNAkyRJChkDmiRJUsgY0CRJkkLGgCZJkhQyBjRJkqSQMaBJkiSFjAFNkiQpZAxokiRJIWNAkyRJ\nChkDmiRJUsgY0CRJkkLGgCZJkhQyBjRJkqSQMaBJkiSFjAFNkiQpZAxokiRJIWNAkyRJChkDmiRJ\nUsgY0CRJkkLGgCZJkhQyBjRJkqSQMaBJkiSFjAFNkiQpZAxokiRJIWNAkyRJChkDmiRJUsgY0CRJ\nkkLGgCZJkhQyBjRJkqSQMaBJkiSFjAFNkiQpZAxokiRJIWNAkyRJChkDmiRJUsgY0CRJkkLGgCZJ\nkhQyBjRJkqSQMaBJkiSFjAFNkiQpZAxokiRJIWNAkyRJChkDmiRJUsgY0CRJkkLGgCZJkhQyBjRJ\nkqSQMaBJkiSFjAFNkiQpZAxokiRJIWNAkyRJChkDmiRJUsgY0CRJkkLGgCZJkhQyBjRJkqSQMaBJ\nkiSFjAFNkiQpZAxokiRJIWNAkyRJChkDmiRJUsgY0CRJkkLGgCZJkhQyBjRJkqSQMaBJkiSFjAFN\nkiQpZAxokiRJIWNAkyRJChkDmiRJUsg0rusCJFU2Oz+fsuLitLbZNDubgUOHprVNSVLtMaBJIVNW\nXMzg3Ny0tjlz8+a0tidJql0OcUqSJIWMAU2SJClkDGiSJEkhY0CTJEkKGQOaJElSyBjQJEmSQsaA\nJkmSFDIGNEmSpJAxoEmSJIWMAU2SJClkDGiSJEkhY0CTJEkKGQOaJElSyBjQJEmSQsaAJkmSFDIG\nNEmSpJAxoEmSJIWMAU2SJClkjmZAewn4DPgobt7xwB+B1cAfgDZHsR5JkqRQOpoB7WXgsirz7iMI\naKcBc6LTkiRJDdrRDGh/Aj6vMu9K4NfRx78GvncU65EkSQqlur4GrR3BsCfRf9vVYS2SJEmhUNcB\nLV559EeSJKlBa1zHz/8Z0B74O3AC8I9kK44bNy72OC8vj7y8vFouTTq02fn5lBUXp7XN5YsXMzg3\nN61tSmGWnz+b4uKytLaZnd2UoUMHprVNKZn58+czf/78tLZZ1wFtBnAz8Gj03+nJVowPaFJYlBUX\npz1MLV2wIK3tSWFXXFxGbu7gtLa5efPMtLYnVadqx9H48eOPuM2jOcT5W+DPwOnARuCfgYnAdwg+\nZuPb0WlJkqQG7Wj2oA1LMv+So1iDJElS6KXag9YhybqR6DJJkiSlSaoBbT2Qk2B+NvC3tFUjSZKk\nI74GrQWwJx2FSJIkKXCoa9Cejnv8CPBllW3PB5amuyhJkqSG7FAB7ey4x2cC8R9UUwYsBn6R7qIk\nSZIaskMFtLzov5OAUcD22ixGkiRJqX/MxojaLEKSJEkHpBrQvgb8H2AA0JbKNxeUA+ekuS5JkqQG\nK9WA9ixwNZBP8G0A8V9q7hecS5IkpVGqAe17wLXAH2uxFkmSJJH656B9CRTWZiGSJEkKpBrQHgPu\nJvhqJ0mSJNWiVIc4LwG+BVwGrAD2Elx7Fon+e2WtVCdJktQApRrQioHpSZZ5k4AkSVIa+TlokiRJ\nIXOkX5YuSZKkNEu1B+2jBPPir0Hzg2olSZLSJNWA9l9VppsAPYALgP9Ia0WSJEkNXKoBbVyS+aOB\nDukpRZIkSXDk16C9DtyYjkIkSZIUONKA9i2CbxmQJElSmqQ6xDmTAzcFEP33BKAnML4W6pIkSWqw\navJBtfEBbT+wDBgD/KEW6pIkSWqw/KBaSZKkkEk1oFXoDHQj6E1bCaxLe0WSJEkNXKoBrTXwEjCE\nYHgTghsM/gv4AbAj/aVJkiQ1TKnexfkUcDZwMdA8+vNtgm8QeKp2SpMkSWqYUg1oVwI/BN4ByqI/\n86PzvlcrlUmSJDVQqQa0rxHcyVnVNiAzfeVIkiQp1YD2Z+BBoEXcvJbAz6LLJEmSlCap3iRwFzAb\n2AwsJfg8tLMJvkVgYO2UJkmS1DClGtA+Ak4FrgfOjM6bDLwC7K6FuiRJkhqsVAPaI8AG4D+rzB8J\n5AL/ls6iJEmSGrJUr0G7CViSYP4S4Ob0lSNJkqRUA9rXga0J5hcD7dJXjiRJklINaBuB/gnmfwvY\nlL5yJEmSlOo1aM8B/w40BeZE510CTAAerYW6JEmSGqxUA9rjQA7B1zo1i84rjU7/vBbqkiRJarBS\nDWgAY4CHgW7R6ZX4JemSJElpV5OABrATWFgbhUiSJCmQ6k0CkiRJOkoMaJIkSSFjQJMkSQoZA5ok\nSVLIGNAkSZJCpqZ3cUoJzc7Pp6y4OK1tNs3OZuDQoWltM911Ll+8mMG5uWlrr6HLfyOf4p3pPY+y\nW2Yz9Kr0nkeSVNsMaEqLsuLitAeVmZs3p7U9SH+dSxcsSFtbguKdxeR+I73n0eZF6T+PJKm2OcQp\nSZIUMgY0SZKkkDGgSZIkhYwBTZIkKWQMaJIkSSFjQJMkSQoZA5okSVLIGNAkSZJCxoAmSZIUMgY0\nSZKkkDGgSZIkhYwBTZIkKWQMaJIkSSFjQJMkSQoZA5okSVLIGNAkSZJCxoAmSZIUMgY0SZKkkDGg\nSZIkhYwBTZIkKWQMaJIkSSFjQJMkSQqZxnVdgCRJDVV+/myKi8vS1l52dlOGDh2YtvZUdwxokiTV\nkeLiMnJzB6etvc2bZ6atLdUthzglSZJCxoAmSZIUMgY0SZKkkDGgSZIkhYwBTZIkKWQMaJIkSSFj\nQJMkSQoZA5okSVLIGNAkSZJCxoAmSZIUMgY0SZKkkDGgSZIkhYwBTZIkKWQMaJIkSSFjQJMkSQoZ\nA5okSVLIGNAkSZJCxoAmSZIUMgY0SZKkkDGgSZIkhYwBTZIkKWQMaJIkSSFjQJMkSQqZxnVdgKRj\n0+z8fMqKi9Pa5vLVi8n9Rm5a21R65efPpri4LK1tLl68nNzcwWltszY05H3X0WdAk3RYyoqLGZyb\n3jD15vsL0tqe0q+4uCztgWLBgqVpba+2NOR919HnEKckSVLIGNAkSZJCxoAmSZIUMgY0SZKkkDGg\nSZIkhYwBTZIkKWQMaJIkSSFjQJMkSQoZA5okSVLIGNAkSZJCxoAmSZIUMgY0SZKkkDGgSZIkhYwB\nTZIkKWQMaJIkSSFjQJMkSQoZA5okSVLIGNAkSZJCpnFdFxC1HtgO7AO+As6v02okSZLqUFgCWjmQ\nB2yr4zokSZLqXJiGOCN1XYAkSVIYhCWglQNvA4uAH9ZxLZIkSXUqLEOc/YAtwNeBPwIfA3+KX2Hc\nuHGxx3l5eeTl5R296uqZ2fn5lBUXp7XN5YsXMzg3N61tfrh4MTz3XFrbrI06FW6LFy/nuR0z09rm\n6vV/4bSzT05vmx+v5rQzTgt9m4uX/Y3c3MFpbbM2LF78Ybr/+2Dx4uXHxL7r6Js/fz7z589Pa5th\nCWhbov8WAf9NcJNA0oCmI1NWXJz2kLJ0wYK0tgdQvmvXMVGnwm3Xrn1pf1Nd8P6bXPyNPultc+EC\nLv7GxeFvc8GHaW2vtuzaVZ7+133B0rS2p/qjasfR+PHjj7jNMAxxNgdaRR+3AC4FPqq7ciRJkupW\nGHrQ2hH0mkFQzyvAH+quHEmSpLoVhoD2N6BHXRchSZIUFmEY4pQkSVIcA5okSVLIGNAkSZJCxoAm\nSZIUMgY0SZKkkDGgSZIkhYwBTZIkKWQMaJIkSSFjQJMkSQoZA5okSVLIGNAkSZJCxoAmSZIUMgY0\nSZKkkDGgSZIkhYwBTZIkKWQMaJIkSSFjQJMkSQoZA5okSVLIGNAkSZJCxoAmSZIUMo3rugBJte83\nM/J58/0FaW2zZONnDL79X9PaZm34R+Ealr75XFrbXLFkIW++lZ3WNpcsWUGbtxaGvs3Cwk/T2h5A\n4d+X8WZBel+jwr8vS2t7x4rFiz/kufQeSrKzmzJ06MD0NqpDMqBJDcCuPdu5/sIL0trmCy9/ktb2\nak1pKZdm56a1yfzdZWQff35a29y9+41jos3S0qVpbQ+glC/JPiu9r1HpX79Ma3vHil27ysnNHZzW\nNjdvnpnW9pQahzglSZJCxoAmSZIUMgY0SZKkkDGgSZIkhYwBTZIkKWQMaJIkSSFjQJMkSQoZA5ok\nSVLIGNAkSZJCxoAmSZIUMgY0SZKkkDGgSZIkhYwBTZIkKWQMaJIkSSFjQJMkSQoZA5okSVLIGNAk\nSZJCxoAmSZIUMgY0SZKkkDGgSZIkhYwBTZIkKWQMaJIkSSHTuK4LkHRs+rRkG88VvJnWNtd8WpjW\n9pR+20o+5c2C59Lb5vZP09qeVB8Y0CQdlrKMveSelZ3WNks/Kk1re0q/vRllZJ+Vm94255SltT2p\nPnCIU5IkKWQMaJIkSSFjQJMkSQoZA5okSVLIGNAkSZJCxoAmSZIUMgY0SZKkkDGgSZIkhYwBTZIk\nKWQMaJIkSSFjQJMkSQoZA5okSVLIGNAkSZJCxoAmSZIUMgY0SZKkkDGgSZIkhYwBTZIkKWQMaJIk\nSSFjQJMkSQoZA5okSVLIGNAkSZJCxoAmSZIUMgY0SZKkkGlc1wXUJ7Pz8ykrLk5rmytWr6bbaael\ntc3lixczODc3rW02VMv+XshzBW+mtc3ff7yYnII2aW3z0+3b0tpebSne9gVvvrUwrW2u27ye11en\n9zUqKfsire1JUlUGtDQqKy5Oe/BZumABgy++OO1tKj2+pJTcs7LT2ubOOXvS3mbZnL1pba+27N1b\nTvbx56e1zTJe4uunp/d47vvg2Dieko5dDnFKkiSFjAFNkiQpZAxokiRJIWNAkyRJChkDmiRJUsgY\n0CRJkkLGgCZJkhQyBjRJkqSQMaBJkiSFjAFNkiQpZAxokiRJIWNAkyRJChkDmiRJUsgY0CRJkkLG\ngCZJkhQyBjRJkqSQMaBJkiSFjAFNkiQpZAxokiRJIWNAkyRJChkDmiRJUsg0rusCUvX555+ntb1W\nrVrRuPExs/uSJKkBOWYSyopp09LW1u7SUjpefjmnnnpq2trUsWHZ3wt5ruDNtLX36fZtaWtLx46v\nynbzt8XpO48Avtz2aYNts+zLL9LaHsCWzct44T+vT2ubH65eRJuTc9La5uKPf0+bgvS1me72AL7a\n9TdgcFrbzM+fTXFxWVrbXL16Baed1i2tbWZnN2Xo0IFpbTNVx0xA63fiiWlra/XmzZSXl6etPR07\nvqSU3LOy09Ze2Zy9aWtLx46M/fvp2Tp95xHAnH17G2yb8/fVxu9RKf3P65LWFpd88g7ZZ+Wmtc09\nc3amtc10twewtuDDtLYHUFxcRm5uekPfggVLufji9La5efPMtLZXE16DJkmSFDIGNEmSpJAxoEmS\nJIWMAU2SJClkDGiSJEkhY0CTJEkKGQOaJElSyBjQJEmSQsaAJkmSFDIGNEmSpJAxoEmSJIWMAU2S\nJClkDGiSJEkhY0CTJEkKGQOaJElSyBjQJEmSQsaAJkmSFDIGNEmSpJAxoEmSJIVMWALaZcDHwCfA\nj+u4lmPG/I8+qusSQsnjcrCdO/bUdQmhtGb9+rouIZQ8LomVflla1yWEzvz58+u6hHorDAGtEfAM\nQUjrBgwDzqzTio4RBpHEPC4H22VAS2jthg11XUIoeVwSM6AdzIBWe8IQ0M4H1gDrga+AV4Gr6rIg\nSZKkutS4rgsAcoGNcdObgN5VV3ooPz9tT1i6dy83ffvbaWtPkiQpnSJ1XQDwTwTDmz+MTt9IENB+\nFLfOGqDLUa5LkiTpcKwFuh5JA2HoQdsMnBw3fTJBL1q8I9pJSZIk1UxjgqTZCWgKfIA3CUiSJNW5\ny4FVBEOZY+q4FkmSJEmSJCk8XgI+A+I/uGoosBzYB5xXzbb1+cNtj+S4rAc+BN4HFtZSfXUh0TF5\nDFgJLAVeB45Lsm1DO1dSPS7rqZ/nCiQ+Lg8SHJMPgDlUvvY1XkM7X1I9Luupn+dLomNS4R5gP3B8\nkm0b2rlS4VDHZT3181yBxMdlHMG18+9Hfy5Lsu0xdb58C+hJ5R09AzgNmEfyINKIYDi0E9CE+nfd\n2uEeF4C/kfyX5liW6Jh8hwOf5Tcx+lNVQzxXUjkuUH/PFUh8XFrFPf4R8GKC7Rri+ZLKcYH6e74k\nOiYQBNW3SL7fDfFcgUMfFw6x7FiX6Lg8ANx9iO1qfL7U9QfV/gn4vMq8j4HVh9iuvn+47eEelwph\n+PiUdEt0TP5I8FccwP8CJyXYriGeK6kclwr18VyBxMdlR9zjlsDWBNs1xPMlleNSoT6eL4mOCcAT\nwOhqtmuI5woc+rhUqI/nCiQ/Lofa3xqfL3Ud0A5Xog+3za2jWsKmHHgbWMSBz5ZrCH4A/D7B/IZ+\nriQ7LtAwz5WHgULgZhL3LDbU8+VQxwUa1vlyFcFr/2E16zTEcyWV4wIN61yp8COCSwV+BbRJsLzG\n58uxGtDK67qAEOtH0P16OXAnQXdsfTcWKAN+k2BZQz5Xqjsu0HDPlQ7AJODfEyxvqOfLoY4LNJzz\npTnwE4JhqwqJekca2rmS6nGBhnOuVPglcArQA9gCPJ5gnRqfL8dqQEvlw20bqi3Rf4uA/yboVq3P\nRgBXADckWd5Qz5URVH9coOGdK/F+A3wzwfyGer5USHZcoOGcL10IrhNaSnAt1UnAYqBtlfUa2rmS\n6nGBhnOuVPgHQQArJ7iGM9H+HpPnSycS3yUyD+iVZJuG8OG2naj5cWnOgQt+WwDvApemvbK604nK\nx+Qygjtbc6rZpiGeK6kcl/p+rsDBx+XUuMc/AqYk2KYhni+pHJf6fr50IvH/t5D8gveGeK7ES3Zc\n6vu5AgcflxPiHt9F4lGLY+58+S3wKcEwzEaC62W+F328G/g78GZ03ROBWXHb1ucPtz3c49KZ4EX/\nAFhG/TouiY7JJ8AGDtza/B/RdRv6uZLKcanP5wokPi6vEfyn+gHwXxz4y7+hny+pHJf6fL5UHJNS\ngmPyz1WWr+NAEGmI50pNj0t9Plcg8e/QZILr8pYC04F20XUb0vkiSZIkSZIkSZIkSZIkSZIkSZIk\nSZIkSZIkSVJt2Q8MCVl740j+IZoQfCDkfuC8I3ye2jAJmFnXRUgKr2P1q54k6VhW8bUwkpSQAU2S\njr4Iyb9oWpIMaJK4DPgTsA0oBt4CzjjENicCrwBbgV0EXymVF7f8NoKvMykl+Oqpf0nQRjaQD+wk\n+I66ql/sfjbwNvBltK6Xgdap7VIlpwMFBF+TthL4TnR+JFrjPVXWP5VgaLRHgrZOiy47q8r8Wwm+\nGLpRdPoi4H858NVsTwBNqqlxPvB0lXmTqDwMOp/ga7seJzge/wBGAZnAc0AJwVd8DavSTi7wKsHr\nuw34H6BrNbVICgEDmqTmBAHim0B/4AuCYJAsULQA3gE6AFcB3YEH4pZfTRA2nogue4ogWHy3Sjv3\nA/8NnAP8DngJODnuOWYD26N1XQ1cEF2npn4OPAmcC/wReIMgYJYDL3Lwdwz+gCBwfpCgrdXAXzk4\nTN4Q3Yd9BIHoTWAxQci7hSA0TaimxkRDnonm3UDw+pwPTIzu1xvAcoJr7X5NcIwqvguwOTCPIORe\nBPQBthAE369VU48kSQqZFsBeoF/cvPiL+n9IEJyOJ7F3CYJPvJcJeuni23s4broRQU/c9XHPURKt\npUL/6Hado9PjSO0mgfgvJY4QfFnxg9Hp9gRfetw7ro7NwB3VtPsjYH3cdAeCYNYnOv1w9Dni3Qzs\nIejtgoN7x+YB/6/KNlXXmU9wbOP9g+DLmSs0Jui1rHitfkAQKuM1Iuj5HIqk0LIHTVIX4DcEw31f\nEAzJZXCgN6uqnsBSguGyRM7g4CDxLtCtyrwP4x7vIxgibBudPjP6HLvi1nmPIHBVbedQ3ot7XE4w\n9FjRxt8Jhvx+EJ2+DMgiGL5N5ncEPXDfik4PA9YBf4mr/S9VtnkXaMqRDS2WU/mYQRDQ4kPqXuBz\nDhzHXsApwI64nxKgDQeCrqQQalzXBUiqc/8DFBJcR7WZICytIAgUyRzOBe5Vh+u+SrA8/o/GZM9x\npHc/Rqq08SJBQP1XgqD2OkFQTeYfBEOlNxD0Ct5A5UBXTs1r359gm0RDzImOWXXHMYNgqPa6BG19\nnqQWSSFgD5rUsGUTXET/CDCXYGiuNdX/8baE4Lqx7CTLVwIXVpl3IcF1UqlaQXCTQMu4eRcQ/J+1\nsgbtAPSNexwhuH4rvo2Ka91uJ7hOLpXr3KYSDBH2IrhhYGrcspUEw53xgetCgqHUtUnaKyLolYt3\nLkceRhcT9NoVE/Tyxf8Y0CRJCqkMgh6hVwjeyPsDCwnCxPC49eKvQWtOMBxaQBA8OgNXcuAuzqui\n27T4JV4AAAGISURBVN9BcEfkj6LTg5K0V+FvwN3Rx18j6M17nSAAXUQQHvPj1h9HategbQD+iSCI\nPkVwwXzVMDSO4NqtZAGqqq8RhLoPOHg480SCO1N/STDcOYjgwvzH4taZROXry26N1jU4WucTBEOR\nVa9Bq3qn5zKCmy3ibeHANXRfAz6ObnsRwXDnRcAv8E5OSZJC7WKCoLOb4Bqn/7+dO8alIIrCAPwX\norMDEbtQaVmDTqLQWILECnRqYgtKa6DS60VCKyrFPxPjee8pn/B93eTe3LlTzck95569tFZpUYCW\nfLZueE3rxO7TH//oOG2v8Z4WqR/NvPOnAC1pYDa22XhJT7Y2JuNn+V6TNbWdpmsP0hqwsc3G/py5\nW8OeTpesN+t6WP9kzthuGri9pXVu5/masrxKcjN5XktykZ6kPaffNjtn3kWChywP0JLWo10meRr2\n85imdRedgAIA/Ao7aS3X5qo3AgDw362nQdltejsTAIAVO0xbU9ylaVsAAAAAAAAAAAAAAAAAAAD+\nrA8xusMnb/cPHAAAAABJRU5ErkJggg==\n", + "text": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 3, + "metadata": {}, + "source": [ + "Scatterplots" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top]](#Sections)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Scatter plots are useful for visualizing features in more than just one dimension, for example to get a feeling for the correlation between particular features. \n", + "Unfortunately, we can't plot all 13 features here at once, since the visual cortex of us humans is limited to a maximum of three dimensions." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Below, we will create an example 2D-Scatter plot from the features \"Alcohol content\" and \"Malic acid content\". \n", + "Additionally, we will use the [`scipy.stats.pearsonr`](http://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.pearsonr.html) function to calculate a Pearson correlation coefficient between these two features.\n" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "from scipy.stats import pearsonr\n", + "\n", + "plt.figure(figsize=(10,8))\n", + "\n", + "for label,marker,color in zip(\n", + " range(1,4),('x', 'o', '^'),('blue', 'red', 'green')):\n", + "\n", + " # Calculate Pearson correlation coefficient\n", + " R = pearsonr(X_wine[:,0][y_wine == label], X_wine[:,1][y_wine == label])\n", + " plt.scatter(x=X_wine[:,0][y_wine == label], # x-axis: feat. from col. 1\n", + " y=X_wine[:,1][y_wine == label], # y-axis: feat. from col. 2\n", + " marker=marker, # data point symbol for the scatter plot\n", + " color=color,\n", + " alpha=0.7, \n", + " label='class {:}, R={:.2f}'.format(label, R[0]) # label for the legend\n", + " )\n", + " \n", + "plt.title('Wine Dataset')\n", + "plt.xlabel('alcohol by volume in percent')\n", + "plt.ylabel('malic acid in g/l')\n", + "plt.legend(loc='upper right')\n", + "\n", + "plt.show()" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "display_data", + "png": "iVBORw0KGgoAAAANSUhEUgAAAloAAAH4CAYAAACSZ0OSAAAABHNCSVQICAgIfAhkiAAAAAlwSFlz\nAAALEgAACxIB0t1+/AAAIABJREFUeJzs3Xd8VfX9x/HXvdnjZhHDkgAWUZZKRSgKNVZ/FWdxoCJD\n1F9ba7WoiFAriqIdgtYfoJQKMmTJUlRcdaCMypA9NMgKJJA9bnZy7/n9cZKQSBJuQm5Oxvv5eNxH\n7tmfGyN55/v9nu8BEREREREREREREREREREREREREREREREREREREREREREREWnmBgPfW12EiIiI\nSHPwZ+Cjn6w7WMO6u7xcixvIBZxAGvB5Ha8ZBxxv+LIsu46IiIg0c1cCWYCtbLk9cARIAuyV1rmB\ndl6uxQ1cUPY+ChgJpADPenh8HApaIiIi0oT4A3lA37Llu4C3gHXAzyutiy97H0fVkHEUGAfswgxs\ny4CASttvBnYCmcBGoE8ttVQOWuXuAAqAyLLl+4H9QA5wCPhd2fqQsv1cmC1iOZjBsD/w37LrJwEz\nAL9K5/8nkAxkA7uBXmXrA4BpwDHgFDALCKzlOiIiIiLV+hJ4rOz9TMww8+JP1s0pex9H1aB1BPgW\nM2xEYoag35dt64sZYq7AbDEbXba/fw11VBe0/IAS4Pqy5RuBrmXvf0nVkHg1Z7Y0/RwzbNmBzmX1\njS3bdj2wDQgrW76I06Hpn8B7QAQQCrwP/LWW64iIiIhU6zlgddn7ncDPMENI+bpdwKiy93GcGbTu\nrbT8D8zWH8q+vvCTa32PGZCqU13QAjgJDK/hmHeBP9VQW3Ue4/Tn+hXwAzCA092kYIbC3J/UMhA4\nXIfriEgLYz/7LiIi1foGGITZInUeZpfcfzHHb0Vidqd9U8vxpyq9L8BsAQKzBWkcZrdd+et8zDFf\nnvIrqymjbPkGzBa09LLz3Qi0qeX47sCHmGEtG3ip0v5fYrbWvY7Z8jYbcJRdLxj4rlLdHwPRdahb\nRFoYBS0Rqa9vgXDgt5jjqMAce5SEOQYqCXOskqeMsq8JmMEmstIrFHinDuf6DVAKbMEcN7UKeBmI\nKTvfR5weyG9Uc/wszO7Cbpif8S9U/fdyBtAP6IkZysYDqZiBsWeluiM43cVY3XVEpIVT0BKR+irA\nHKv0BFVbrjaUrfu6jucrDz5vAg9hjpGyYQ4kv4nTLV61HRsFjMBscfo7ZquSf9krDbOb8Qbg15WO\nTcZsrQqrtC4Uc9B6PnAx8AdOB6V+mN2GfmXbCzEHuRtltb+G2boF0LHStaq7joiIiEiN/ooZMi6r\ntG5Y2brfVloXh9lSVe4I5lincs8BCystX4/ZGlV+19871By0Ks+jlQ58Adzzk30exuyqzCy7zhKq\njgObixnEMjAHtg8GDpSd8xvgeU6HyV9hjj9zYrZivY3ZZQhm69lLmN2o2ZitYo/Uch0RkXNyEbCj\n0iub0wNQRURERKSB2DEHlnayuhARERGRlubXmGM3RERERFqFxhwMfw/muAgRERGRVsF29l0ahD+Q\niHnbc2r5yp/97GfGoUOHGqkEERERkXNyCHPaF481VovWDZiT+KVWXnno0CEMw9CrEV/PPfec5TW0\ntpe+5/qet4aXvuf6nreGF+YTMOqksYLWcGBpI11LREREpElojKAVAlzH6eeEiYiIiLQKvo1wjTz0\nrK8mIy4uzuoSWh19zxufvueNT9/zxqfvefPQWIPha2KU9XmKiIiINGk2mw3qmJ0ao0VLRETEUlFR\nUWRmZlpdhjQTkZGRZGRkNMi51KIlIiItns1mQ79vxFM1/bzUp0WrMScsFREREWlVFLREREREvERB\nS0RERMRLFLREREREvERBS0REpAmaP38+gwcPtroMOUcKWiIiIsLMmTPp168fgYGB3H///XU6dsyY\nMQQEBOBwOIiKiuLaa69l3759DVLX0aNHueaaawgJCaFHjx588cUXte4/YcIEoqOjiY6OZuLEidXu\n8/XXX2O325k0aVKD1FgbBS0REZFquN2wfj2U3+XvdMJ331lbkzd17NiRSZMm8cADD9T5WJvNxoQJ\nE3A6nSQlJREbG1vnsFaT4cOHc/nll5ORkcFLL73EnXfeSVpaWrX7zp49mzVr1rB79252797NBx98\nwOzZs6vsU1JSwtixY/nFL35RPl2DVyloiYhIq5ScDMeOnV4+cADy8k4vFxTAqlUwf74ZsiZNgr17\nq57jp1Mt1WeqruPHj3P77bcTExNDdHQ0jz76aLX7jR07ltjYWMLDw+nXrx8bNmyo2LZlyxb69etH\neHg47dq1Y9y4cQAUFhYycuRIoqOjiYyMpH///qSkpFR7/ttuu43f/OY3tGnTpu4fopLAwECGDRvW\nIC1a8fHx7Nixg+eff56AgABuv/12LrnkElatWlXt/gsWLODJJ5+kQ4cOdOjQgSeffJL58+dX2eeV\nV15hyJAhXHTRRY0yt5qCloiItEqHD5vh6dgx2LkTXnoJEhNPbw8JgSlTYMMGuPdeuPRSGD266jne\nfBPKe7Kys+HppyE93fMaXC4XN998M127duXYsWMkJiYyfPjwavft378/u3btIjMzk3vvvZdhw4ZR\nXFwMmCHs8ccfJzs7m8OHD3P33XcDZvDIycnhxIkTZGRkMHv2bIKCgmqtqb7ho/y4vLw8li5dyoAB\nAyq2bdiwgcjIyBpfmzZtqvac+/bt44ILLiAkJKRi3aWXXlpjiNu/fz+XXnppxfIll1xSZd9jx44x\nb948Jk2a1GgT2OoRPCIi0ioNHAglJfDII+by3/8O3bvX7Rw33QR/+YsZsr78En7xC4iK8vz4LVu2\ncPLkSaZOnYrdbrZ9XHnlldXuO2LEiIr3TzzxBC+++CI//PADffr0wd/fn4MHD5KWlkZ0dDT9+/cH\nwN/fn/T0dA4ePEifPn3o27fvWWuqT3eaYRhMmzaNmTNnkpOTQ5cuXdi8eXPF9kGDBtXrEUi5ubmE\nh4dXWRcWFkZi5URcy/5hYWHk5uZWLP/pT3/ixRdfJCQkBJvNpq5DERERbwoLO/0+NLTqtrw8s8Vr\n0CBYsgR27YKFC6vu07EjPPUUzJtn7j9iBNTld/fx48fp3LlzRciqzbRp0+jZsycRERFERkaSnZ1d\nMVZp7ty5xMfH06NHD/r378/atWsBGDVqFNdffz333HMPHTt2ZMKECZSWltZ6nfq09NhsNsaPH09m\nZiZHjx4lICCAhT/9ZnmgV69eOBwOwsLC2LhxIw6Hg5ycnCr7ZGVlEVb5P1wloaGhVfbPzs4mtOw/\n7AcffEBubi7Dhg0DzM+prkMREREv2bULpk0zW7LGjzdDVULC6e1BQXDHHTBmDDgcZjdi795Vz5Gd\nDW+8YYYxwzBbteqiU6dOJCQk4HK5at1v/fr1TJ06lRUrVpCVlUVmZibh4eEVQaFbt24sWbKE1NRU\nJkyYwJ133klBQQG+vr48++yz7Nu3j02bNvHhhx+eNQDVt5WnvJZOnToxffp0pkyZUhF61q9fj8Ph\nqPG1ceNGwOwqdDqd5OTkcNVVV9GzZ08OHz5cpVVq165d9OrVq9oaevXqxc6dO6vs27vsP9qXX37J\ntm3baN++Pe3bt2f58uW89tpr3HbbbfX6vJ5S0BIRkVYpPNwcU9WrF/zyl/DHP5rjssrZ7TB48OkW\nKocDLr+86jkWLjS7C596yhzjtWxZ3cZoDRgwgPbt2zNx4kTy8/MpLCysdryS0+nE19eX6OhoiouL\neeGFF6q03CxatIjU1NSyzxWOzWbDbrfz1VdfsWfPHlwuFw6HAz8/P3x8fKqtxeVyUVhYSGlpKS6X\ni6KioioB0G63880331R77E9bhq677jq6devGrFmzABg8eDBOp7PG11VXXVXtebt3785ll13G888/\nT2FhIatXr2bv3r3ccccd1e4/evRoXn31VZKSkkhMTOTVV19lzJgxAEyZMoWDBw+ya9cudu7cya23\n3srvfvc75s2bV+25GoqCloiItEpdukDPnqeXBwyAut5w99BDp7sLO3Y0W7fqcg673c4HH3zAjz/+\nSGxsLJ06dWL58uUAVcYQDRkyhCFDhtC9e3e6dOlCUFAQsbGxFef59NNP6d27Nw6Hg8cff5xly5YR\nEBBAcnIyw4YNIzw8nJ49exIXF8eoUaOqrWXKlCkEBwfzj3/8g0WLFhEUFMRLL70EmF2cDoeDPn36\nVHtsdeOdxo8fz/Tp0ykpKfH8G1KNZcuWsW3bNqKiovjLX/7CqlWrKu6MLG8pK/f73/+eW265hT59\n+nDJJZdwyy238Lvf/Q4wuxVjYmKIiYmhbdu2BAUFERISQkRExDnVdzbeHwVWO6OxRv2LiEjrZbPZ\nGu0us5Zo8eLF7N+/vyJ4tXQ1/byUhck6ZScFLRERafEUtKQuGjJoqetQRERExEsUtERERES8REFL\nRERExEsUtERERES8REFLRERExEsUtERERES8REFLRERExEsUtERERJqg+fPnM3jwYKvLkHOkoCUi\nItLKFRcX8+CDD9KlSxfCwsLo27cvn3zyicfHjxkzhoCAABwOB1FRUVx77bXs27evQWo7evQo11xz\nDSEhIfTo0YMvvvii1v0nTJhAdHQ00dHRTJw4sWJ9amoqw4cPp2PHjkRERDBo0CC2bNnSIDXWRkFL\nRESkJmlp8PLL8Nhj5hOjKz1kuSUpLS0lNjaWb775hpycHF588UXuuusujh075tHxNpuNCRMm4HQ6\nSUpKIjY2lvvvv79Bahs+fDiXX345GRkZvPTSS9x5552kpaVVu+/s2bNZs2YNu3fvZvfu3XzwwQfM\nnj0bgNzcXAYMGMD27dvJzMzkvvvu46abbiIvL69B6qyJgpaIiLReO3bACy/A3/4GP/5YdVteHjz4\nIKxYAdu2mYHrn/888xwuFyQnm/vXw/Hjx7n99tuJiYkhOjqaRx99tNr9xo4dS2xsLOHh4fTr148N\nGzZUbNuyZQv9+vUjPDycdu3aMW7cOAAKCwsZOXIk0dHRREZG0r9/f1JSUs44d3BwMM8991zFg6pv\nuukmunbtyvbt2+v8eQIDAxk2bFiDtGjFx8ezY8cOnn/+eQICArj99tu55JJLWLVqVbX7L1iwgCef\nfJIOHTrQoUMHnnzySebPnw9A165deeyxx2jbti02m43f/va3FBcXEx8ff8511kZBS0REWqfNm+Gh\nh2DtWli9GsaMgUOHTm//7js4eRLatYPISPPr8uVQWnp6n5Mn4e674dZb4Ve/Mlu96sDlcnHzzTfT\ntWtXjh07RmJiIsOHD6923/79+7Nr1y4yMzO59957GTZsGMXFxYAZwh5//HGys7M5fPgwd999N2AG\nj5ycHE6cOEFGRgazZ88mKCjorHUlJycTHx9Pr169PP4s5c8GzMvLY+nSpQwYMKBi24YNG4iMjKzx\ntWnTpmrPuW/fPi644AJCQkIq1l166aU1hrj9+/dz6aWXVixfcsklNe67c+dOiouL6datm8efsT4U\ntEREpHWaNw/8/CAmxgxRRUVQQ0tJjZ5+Gk6cMM8REQGvvAJ793p8+JYtWzh58iRTp04lKCiIgIAA\nrrzyymr3HTFiBJGRkdjtdp544gmKior44YcfAPD39+fgwYOkpaURHBxM//79K9anp6dz8OBBbDYb\nffv2xeFw1FpTSUkJI0aMYMyYMXTv3t2jz2EYBtOmTSMyMpKwsDA2bdrE8uXLK7YPGjSIzMzMGl81\nfebc3FzCw8OrrAsLC8PpdHq0f1hYGLm5uWfsl5OTw6hRo5g8efJZvx/nSkFLRERaJ5cLbLbTyzZb\n1TFYl18O7dvDqVOQmWl+vesu8PU9vc++fdCmjfne39/8WrlV7CyOHz9O586dsdvP/ut42rRp9OzZ\nk4iICCIjI8nOzq4YqzR37lzi4+Pp0aMH/fv3Z+3atQCMGjWK66+/nnvuuYeOHTsyYcIESiu3yP2E\n2+1m1KhRBAYGMnPmTI8/h81mY/z48WRmZnL06FECAgJYuHChx8eX69WrFw6Hg7CwMDZu3IjD4SAn\nJ6fKPllZWYSFhVV7fGhoaJX9s7OzCQ0NrbJPQUEBt9xyC1deeSUTJkyoc411paAlIiKt0113QUEB\nZGVBejrY7XDLLae3h4TA3LkwbBj06wdPPQWPP171HLGxkJ1tvne5wDDM1jEPderUiYSEBFxnGWS/\nfv16pk6dyooVK8jKyiIzM5Pw8PCK7rpu3bqxZMkSUlNTmTBhAnfeeScFBQX4+vry7LPPsm/fPjZt\n2sSHH35YYwAyDIMHH3yQ1NRUVq1ahY+Pj8efo/z48s80ffp0pkyZUhF61q9fj8PhqPG1ceNGwOwq\ndDqd5OTkcNVVV9GzZ08OHz5cpVVq165dNXZp9urVi507d1bZt3fv3hXLRUVFDB06lNjY2IpB8t6m\noCUiIq3T//wP/P3v0KsXXHEFvPEGVPqlDEB0tBmwXnsN7rkHfho+XnzRbMlKS4PUVLjzTijrtvPE\ngAEDaN++PRMnTiQ/P5/CwsJqxys5nU58fX2Jjo6muLiYF154oUrLzaJFi0hNTQUgPDwcm82G3W7n\nq6++Ys+ePbhcLhwOB35+fjUGqD/84Q98//33vP/++wQEBJyx3W63880331R7bHnIKnfdddfRrVs3\nZs2aBcDgwYNxOp01vq666qpqz9u9e3cuu+wynn/+eQoLC1m9ejV79+7ljjvuqHb/0aNH8+qrr5KU\nlERiYiKvvvoqY8aMAcwu0TvvvJPg4OCKAfKtgSEiIuJtXv19k5VlGNu3G8aPPxqG213nwxMSEoyh\nQ4cabdq0MaKjo42xY8cahmEY8+fPNwYPHmwYhmG4XC7jgQceMMLCwoz27dsbL7/8stG1a1fjiy++\nMAzDMEaOHGnExMQYoaGhRu/evY01a9YYhmEYS5cuNS666CIjJCTEaNu2rTF27FjD5XKdUcPRo0cN\nm81mBAUFGaGhoRWvJUuWVNQYFhZmZGRkVPsZxowZY0yaNKnKunfeecfo0KGDUVxcXOfvyU9ri4uL\nM4KCgoyLL7644jMbhmF88803RmhoaJX9n3rqKSMqKsqIiooyJkyYULF+3bp1hs1mM0JCQqp8xg0b\nNpxxzZp+XgDjLLnmDLaz7+JVZXWLiIh4j81mO6PVRTy3ePFi9u/fz0svvWR1KY2ipp8Xmzmmr07Z\nSUFLRERaPAUtqYuGDFoaoyUiIiLiJQpaIiIiIl6ioCUiIiLiJQpaIiIiIl6ioCUiIiLiJQpaIiIi\nIl6ioCUiIiLiJQpaIiIiTdD8+fMZPHiw1WXIOVLQEhEREUaOHEn79u0JCwvjggsuqNMs8GPGjCEg\nIACHw0FUVBTXXnst+/btq1cdS5YsoXPnzoSGhnLbbbeRmZlZ476TJk2iT58++Pn58fzzz1fZdurU\nKW699VY6duyI3W4nISGhXvWcKwUtERGRWqTlpzF3+9wWP7P8n//8Z44cOUJOTg4ff/wxM2bM4JNP\nPvHoWJvNxoQJE3A6nSQlJREbG8v9999f5xr27dvHQw89xOLFi0lOTiY4OJiHH364xv0vvPBCpk6d\nyk033VQ+a3sFu93OjTfeyKpVq+pcR0NS0BIRkVbt66Nfszdlb43b5++czyv/fYU9KXtq3KewtLDe\n1z9+/Di33347MTExREdH8+ijj1a739ixY4mNjSU8PJx+/fqxYcOGim1btmyhX79+hIeH065dO8aN\nG2fWVVjIyJEjiY6OJjIykv79+5OSklLt+Xv16kVgYGDFsq+vLzExMXX+PIGBgQwbNqxeLVqLFy/m\n1ltvZdCgQYSEhDBlyhRWr15NXl5etfuPHj2aIUOG4HA4zgjCMTExPPTQQ/Tr16/OdTQkBS0REWm1\nCkoKmLxuMlO+noLbcJ+xPSUvhZX7VxLiH8LrW16vtlWrxFXCiFUj+PLIl3W+vsvl4uabb6Zr164c\nO3aMxMREhg8fXu2+/fv3Z9euXWRmZnLvvfcybNgwiouLATOEPf7442RnZ3P48GHuvvtuABYsWEBO\nTg4nTpwgIyOD2bNnExQUVGM9Dz/8MCEhIfTq1YtnnnmGn//85x5/lvLvTV5eHkuXLmXAgAEV2zZs\n2EBkZGSNr02bNgGwf/9+Lr300orjLrjgAgICAoiPj/e4jqZGQUtERFqtNT+sIbckl8OZh9l0fNMZ\n2xfuWojbcNM2pC3bT26vtlXr00OfciDtANM3T8fldtXp+lu2bOHkyZNMnTqVoKAgAgICuPLKK6vd\nd8SIEURGRmK323niiScoKirihx9+AMDf35+DBw+SlpZGcHAw/fv3r1ifnp7OwYMHsdls9O3bF4fD\nUWM9b7zxBrm5uXz++ec888wzbNmyxaPPYRgG06ZNIzIykrCwMDZt2sTy5csrtg8aNIjMzMwaX+Wf\nOTc3l/Dw8CrnDgsLw+l0elRHU6SgJSIirVJBSQGzv5tNeEA4gX6BzNg8o0qrVkpeCkv3LMXX7ktO\nUQ5FrqIzWrVKXCW8sfUN2oa2JdGZyLqj6+pUw/Hjx+ncuTN2+9l/HU+bNo2ePXsSERFBZGQk2dnZ\npKWlATB37lzi4+Pp0aMH/fv3Z+3atQCMGjWK66+/nnvuuYeOHTsyYcIESktLa72OzWYjLi6OYcOG\nsXTpUo8+h81mY/z48WRmZnL06FECAgJYuHChR8dWFhoaSnZ2dpV12dnZtYbDpk5BS0REWqU1P6wh\nLT8Nl+HC1+7LgbQDVVq1sguz+cX5v6BP2z70OK8HAzsNpE1wmyrn+PTQp6TmpRLqH0qwXzAztsyo\nU6tWp06dSEhIwOWq/Zj169czdepUVqxYQVZWFpmZmYSHh1eEvm7durFkyRJSU1OZMGECd955JwUF\nBfj6+vLss8+yb98+Nm3axIcffuhxACopKSEkJMTjz1JeS6dOnZg+fTpTpkwhJyenon6Hw1Hja+PG\njYA5TmzXrl0V5zx06BDFxcV07979rNf/6WD4psLX6gJERESskF2YzaVtT48H6ujoSHp+esXyhW0u\n5PWbXq/1HPN3zqfEXUJavtmydDjzMJsTN3Nlp+q7/35qwIABtG/fnokTJ/L8889jt9vZvn37Gd2H\nTqcTX19foqOjKS4u5u9//3tFiAFYtGgR119/Peeddx7h4eHYbDbsdjtfffUV0dHR9OzZE4fDgZ+f\nHz4+PmfUkZqayhdffMEtt9xCYGAgn3/+OStWrODzzz+v2Mdut7Nu3Tp++ctfnnH8T8euXXfddXTr\n1o1Zs2YxYcIEBg8e7FH334gRIxg4cCAbNmygb9++TJo0iTvuuKPGwFdaWkppaSkul4uSkhIKCwvx\n9/evaCEsLCysaMErLCyksLCwyoD/1sAQERHxNm/9vtmTvMf47/H/VnnlFObU6RwJCQnG0KFDjTZt\n2hjR0dHG2LFjDcMwjPnz5xuDBw82DMMwXC6X8cADDxhhYWFG+/btjZdfftno2rWr8cUXXxiGYRgj\nR440YmJijNDQUKN3797GmjVrDMMwjKVLlxoXXXSRERISYrRt29YYO3as4XK5zqghNTXVuPrqq42I\niAgjPDzcuOKKKyrOUV5jWFiYkZGRUe1nGDNmjDFp0qQq69555x2jQ4cORnFxcZ2+H0uWLDFiY2ON\nkJAQY+jQoUZmZmbFtoceesh46KGHKpbvu+8+w2azVXktWLCgYnv5OrvdXvHVEzX9vAB1nuPD2+1s\nEcAcoBdmcQ8A31baXla3iIiI99hsthY/D5Y3LV68mP3799dpEtPmrKafl7LuyTplJ28HrQXA18Bb\nmN2UIUDlUW4KWiIi4nUKWlIXzSVohQM7gAtq2UdBS0REvE5BS+qiIYOWN+867AqkAvOA7cCbQLAX\nryciIiLSpHjzrkNf4OfAI8BW4DVgIvBs5Z0mT55c8T4uLo64uDgvliQiIiLimXXr1rFu3bpzOoc3\nuw7bAf/FbNkCGIQZtG6utI+6DkVExOvUdSh10Vy6Dk8Bx4HyWcauA+r+hEkRERGRZsrbE5Y+CiwG\n/IFDwP1evp6IiMgZIiMjm+zM4dL0REZGNti5rP6pU9ehiIiINAtNretQREREpFVT0BIRERHxEgUt\nERERES9R0BIRERHxEgUtERERES/x9vQOIiLW2rMHZs+GvDy4+Wa4/XbQbf4i0kis/tdG0zuIiPf8\n+COMHm2+9/OD3Fx46im4+25r6xKRZknTO4iIVPbVV1BUBG3aQFgYhIfD8uVWVyUirYiCloi0XH5+\nULnV3OUy14mINBIFLRFpuYYMgagoOHkSUlIgPx9++1urqxKRVkRjtESkZUtKgpUrzfFZ//M/cMUV\nVlckIs1UfcZoKWiJiIiIeECD4UVERESaEAUtERERES9R0BIRaaEyCjKsLkGk1VPQEhFpgRKyE7ht\n2W3Ep8dbXYpIq6agJSLSAs3ZPodEZyKzt822uhSRVk1BS0SkhUnITuCTHz/hwjYXsj5hvVq1RCyk\noCUi0sLM2T4HAF+7L3abXa1aIhZS0BIRaUEyCjL4z6H/AJCen46BwfqE9SQ5kyyuTKR10oSlIiIt\niNtwcyjjEKXu0op1PnYfukV1w27T39Yi50Izw4uIiIh4iWaGFxEREWlCFLRE5KxOOk9S4iqxugwR\nkWZHQUtEalXsKuZ/P/hflu5danUpIiLNjoKWiNTqo4MfcSL7BHO3zyWvOM/qckREmhUFLRGpUbGr\nmFlbZ9EmuA35pfmsOrDK6pJERJoVBS0RqdFHBz8ioyCDEP8QIgIj1KolIlJHCloiUqOV+1fiNtyk\n5aWRW5RLTnEOXx/72uqyRESaDc2jJSI1yizIJK+kagtW25C2+Pn4WVSRiIh1NGGpiIiIiJdowlIR\nERGRJkRBS0RERMRLFLREREREvERBS0RERMRLFLREREREvERBS0RERMRLFLREREREvERBS0RERMRL\nFLREREREvERBS0RERMRLFLREREREvERBS0RERMRLFLREREREvERBS0Skkh8zfuSZL5/BbbitLkVE\nWgAFLRGRSmZtncXK/SvZfGKz1aWISAugoCUiUiY+PZ71CetpE9yGGVtmqFVLRM6ZgpaISJnZ22Zj\nt9mJDIzKmcCEAAAgAElEQVTkYPpBtWqJyDlT0BIRAQ6mH+TTQ59S6i4lOS+Z3JJcZmyZYXVZItLM\n+VpdgIhIUxDkF8Qf+v0BA6NiXXhAuIUViUhLYLP4+oZhGGffS0RERMRiNpsN6pid1HUoIiIi4iUK\nWiIiIiJeoqAlIiJncLldHM06anUZIs2egpaIiJzhP4f+w+h3R5NRkGF1KSLNWmMEraPAbmAHsKUR\nriciIueg1F3KzK0zSc9PZ9HuRVaXI9KsNUbQMoA4oC/QvxGuJyIi5+DzQ5+TnJfM+eHns3TvUrVq\niZyDxuo6tHoaCRER8UB5a1aIXwj+Pv643C61aomcg8Zq0foc2Ab8thGuJyIi9bTz1E5O5Z6isLSQ\n1LxUDMPgg/gP9NxHkXpqjJnhrwJOAucB/wG+B9aXb5w8eXLFjnFxccTFxTVCSSIizVtiTiJHs45y\nVexVDXren7f/OWvvXVtlXYBvAHab7p2S1mfdunWsW7funM7R2F16zwG5wCtly5oZXkSkHsZ9Oo7N\niZv5eMTHOAIcVpcj0io0xZnhg4HyfwFCgF8De7x8TRGRFi0+PZ71CespKC1g5f6VVpcjIrXwdtBq\ni9lNuBPYDHwIfObla4qItGizt83GbrPTJqgNb+18C2eR0+qSRKQG3g5aR4DLyl69gb95+XoiIi1a\nfHo8Xx79klD/UAwMnEVOtWqJNGGNMRheREQayJHMI3R0dKR8fGtHR0eOZB2xuCoRqYnV81tpMLyI\niIg0C01xMLyIiIhIq6WgJSIiIuIlCloiIiIiXqKgJSIiIuIlCloiIiIiXqKgJSIiIuIlCloiIiIi\nXqKgJSIiIuIlCloizZRhGDzz5TPsSdZz2kVEmioFLZFmavvJ7bx74F1e+/Y19IQFEZGmSUFLpBky\nDIPXt75OeGA4u1N2s/PUTqtLEhGRaihoiTRD209uZ0/yHqKCovCz+zFzy0y1aomINEEKWiLN0Btb\n3yCnOIdTuacochWx6cQmdiXvsrosERH5CV+rCxCRuru5+80M7DSwyrrIwEiLqhERkZrYLL6+oe4O\nERERaQ5sNhvUMTup61BEmrRTuad44tMnKHYVW13KOUnOTcZtuK0uQ0QamYKWiDRpC3Yu4P0f3ueT\nHz+xupR6cxY5Gbl6JJ/9+JnVpYhII1PQEpEm61TuKd79/l06ODrw+tbXm22r1qoDq0hyJjFz60xK\n3aVWlyMijUhBS0SarAU7F+A23IQHhpORn9EsW7WcRU7e2vEWHcI6kJyXzOeHPre6JBFpRApaItIk\nJecms2zvMkrcJZx0niS/NJ+ZW5pfi9CqA6soLC0k0DeQEL8QtWqJtDKa3kFEmiQ/Hz/+cMUfcLld\nFeuC/IKa3cSsy/ctx2W4SMtLAyCnKIetiVvPmJ5DRFomTe8gIuJFp3JPkV+SX2VdbHgsvnb9nSvS\n3NRnegcFLREREREPaB4tERERkSZEQUtERETESxS0RERERLxEozFFWqv9++HoUWjXDvr2BZvVQzZF\nRFoeBS2R1mjZMnjlFTNcGQaMHAljx1pdlYhIi2P1n7C661CksTmdcN11EBEB/v7gckFaGixfDl26\nWF2diEiTpbsOReTscnLMlix/f3PZxwd8fSE729q6RERaIAUtkdambVuIiYHUVLPbMDMTAgOha1er\nKxMRaXEUtERaG19fmDkTfvYzOHUKzjsPXn8dwsKsrky8ICE7gd9/+HuKXcVWlyLSKmmMlkhr5naD\nXX9vtWTPfPkM7+x9h6n/M5WhPYZaXY5Is6YxWiJSNwpZLdqxrGN8dugzOoZ15I1tb6hVS8QC+ldW\nxFPp6TBlCtx/P8yYAQUFdT9HXp45Nsrtbvj6RH7ize1vYrPZcAQ4yCzI5KP4j6wuSaTVUdehiCcK\nCsy5phISIDgYcnPhmmtg6lTPJ/qcPx9mzTLfd+sG//d/EB3ttZKldTuRc4IbF9+I3WbHbrNTWFrI\n+WHn89GIj/C1awpFkfqoT9eh/m8T8cT330NiojmLOkBoKHz9tTklQkTE2Y/futUcgB4dbQ5GP3gQ\nnn/ebBmTZqfUXcpzXz3Hnwb8ibahba0up1ohfiE8PfjpKusCfAKwWf73tUjroqAl4gkfH3MqBMM4\nPZt6+XpPHDpkHuPnZy5HRcGePd6pVbzu80Ofs3zfciICIxh/1Xiry6lWZFAk9/S+x+oyRFo9jdES\n8USPHtCzJ5w8ac6inpwMQ4eCw+HZ8W3bmgGtfGxWTg7ExnqvXvGaUncpM7fOpG1oW1YdWEVybrLV\nJYlIE6agJeIJPz9zrqk//MF8fM2f/wwTJ3p+/NVXww03QEqKGdRCQuC557xXr3jN54c+Jzkvmcig\nSNyGm4W7Flpdkog0YVZ31mswvLQehgHx8eZA+u7dPW8Nkyaj1F3K0GVDOZp1lBD/EErdpRiGwScj\nPyEmJMbq8kTEyzQYXqQps9ngoousrkLOgWEY3HThTeSV5FWss9vUMSAiNVOLloiIiIgHNDO8iIiI\nSBOioCUiIiLiJQpaIiIiIl6ioCUiIhiGweI9i8ktzrW6FJEWRUFLRETYfnI7f/3mr6zYt6JRrrc/\ndT9uQw9Xl5ZPQUtEpJUzDIPXt75OoF8g83bOw1nk9Or1jmYd5f4197MhYYNXryPSFChoiYi0cttP\nbmdP8h7ah7anoLSAlftXevV6c76bQ05hDjM2z1CrlrR4CloiIq1YeWtWqVFKXkke/j7+zNkxx2ut\nWkcyj/DZ4c/oGtmVo1lH1aolLV5tM8NfDtQ2m+j2Bq5FREQaWYm7hEDfQC6OvrhiXYBPAGn5aTgC\nGv4xUXO2z8GGDR+7D0F+QczYPINBsYOw2+wUFkJGBnToYO6bnAyhoeajQUWaq9pmN11H7UHrmga4\nvmaGFxFpJfJL8rlx8Y3kFudit9kxMPCx+bDwtoV0b9Od7dvh//4Ppkwxn+P+l7/Agw/CVVdZXbmI\nqT4zw9e2c0cg8VwKKuMDbANOALf8ZJuClohIK5Jfko/L7apYttlshPqHVix/9RW8+qr5/uGH4YYb\nGrtCkZo19EOl3wTaAF8BnwAbgNJ61DUW2A80fBu0iIg0K8F+wbVuv/BCcLnAxwd69YL8fAgKMp/J\nLtIc1TYY/kYgDvgauB34FngX+B0Q6+H5zy87zxysf4C1iEi9/PADJCWdXt60CQoKrKunpTp5Ev73\nf80xWX/8I0yYYLZqbdtmdWUi9Xe2uw4LgI+BPwH9gHGAH/A6sNWD8/8TGA/o/l0RabaOHzfHCyUl\nwUcfwZw54PTuVFOtUmEhPP44xMXBmjWQk2O2avXrZ3VlIvVXW9dhdQ5jhqzXAf+z7HszkALswGwZ\nq9bkyZMr3sfFxREXV+OuIiKWuO46MAz4/e/N5TffhJgYa2tqibp2NV8DBsCoUWC3wyOPqNtQrLNu\n3TrWrVt3Tufw5Me3ur/bsjAHuI/DDF/V+SswCnNcVyAQBqwCRlfaR4PhRaRZ+OgjmDXLfD979ukp\nCKRhZWfDM8/AFVeYrYYJCTB5sjlOS8RqDX3XYbkXgePA0rLle4CfYbZUPUQtrVWVXA08ie46FJFm\n6LPPYNky+OtfYc8eWLIE/v53aNvW83Mk5yZzXsh52G2aJ7o2mzbB4cMwYoTZivjvf8OVV8Ill1hd\nmYj3gtZu4Kc/4juBy4BdwKUenONqzNavW3+yXkFLRBrVtye+JSIwosoEnWdz8CA4HNCunbm8dav5\niz8gwLPjnUVObn/ndsYNHMeQC4fUo2oRaQrqE7Q8+dMqH7i7bF87cBdQWLbN05T0NWeGLBE5R0cy\nj1DsKra6jGajqLSIZ758hhe+fqFOz9i78MLTIQvMbi1PQxbA6gOrSXImMXPrTErd9ZklR0SaK0+C\n1gjMsVYpZa/RwEggCHjEe6WJSG3yivP43/f/l+X7lltdSs1KSmD+fHjsMXjtNfM2MqvqKChgbfxa\nsguzOZh+kM0nNjfKpZ1FTubumEuHsA4k5yXz+aHPG+W6ItI0WH0vh7oORepp4a6FTN00lTZBbVh7\n71pC/JvgA+EmTTJHkQcFmffuX3ghLFgA/me7abmBGIYZ9P71L4qMEm7+dRruCy6giFJiw2NZdPsi\nr4+ZWrBzATO3zqRdaDucRU5C/UN575738LXX9aZvEbGat7oORaSJySvOY+72ubQLbUdeSR7vfv9u\nrfsXu4obv8vK6YRPPjH73CIjza+HD8P+/Y1Xw4YNMHMmREaytruNlPxUOH6cAN8A9qXsa5RWrWX7\nlpGdY5Cck0aRq4hEZxL/XL6FUvUgirQKCloizdCqA6vIL80n0DeQiMAI5myfQ15xXo37v7DuBaZu\nnNqIFWK2JlXXYt2Yrdi7d5uTMPn5cdK/mG4lDiIy84kIiKBbVDdO5Jzweglv3foWD0cvp/vexbz+\nq8Vcn7WSnAP9cWsaZ5FWQW3XIs3Q+z+8j9twk5KXAkCpu5RvT3zLtRdce8a+R7OO8tnhz7BhY8xl\nY2jvaN84RYaFwZAh8PHHZtdhURH87GfQs+c5ndYwwO02n4UHp5+LV6327c2dDYM/pnbhjwdC4bLL\n4O5Z51RDXbR3tOex+2CRDzz3J/Nb8MLLjdd7KiLW8iRoBQJ3AF0q7W8AL3ipJhE5iwVDF1DkKqqy\nLiIwotp93/zuTWzYMDCYt3MeTw9+ujFKND37rJksduyALl3gwQfrdrteNdavh3Xr4M9/NkPXSy/B\n9debcy2d4aabzO7LnTvNacYdDnjqqXO6fn24XOZz/ADy8swHJStoibQOngzo+hRzJvjvAFel9a80\nwPU1GF7Ei45kHuHulXcTHRyNgUFGfgbv3fNe47VqeUFpKUybBrm5ZmNVZCQ88UQtrVqlpWbQKyqC\n3r0hovpA6k2vvQaZmebzEleuhI0bzc+g2c5FmhdvTVi6F+hdn4I8oKAl4kX/2PAP3t79No4AB2BO\nNfD7y3/PowMetbiyc1NQAHfdZb5fsQICA62t52wOHDAb9spbsXbvhj599Aw/kebGW0Hr38BMzBni\nG5qClpzJ5YJvvzUfetarF3TubHVFzVZiTiJJzqQq684PO79Zt2gVF5vdhUFB5o+Ky2V2I/r5WV2Z\niLR03gpaB4BuwBGgfFCIwZmP5akPBS2pyuWCcePM2/LtdrM/6J//hF/8wurKpInYsAH++1+zu9Aw\nzC64a66BAQOsrkxEWjpvBa0uNaw/WpcL1UBBS6pavx4ef9ycc8lmMwfiBAaad66JlDGM091uld+L\niHhTQ09YGlb2NaeGl0jDy842W7LKf3MGB0N6euPOvSRNXuVg1VxDVkJ2As98+UydnrkoIs1PbUFr\nadnX7Zh3HFZ+bfNyXdJa9expBq28PPOWsuRkGDiw+f42FanBnO1zWLFvBRsSNlhdioh4kdW/vdR1\nKGdatw5efNFs3RowwHxvwS35It5yLOsYw1YMw9/Hn3ah7Vg+bLnXn7koIueuPl2Hmhlemp64OPPl\ndputWyItzNwdc7HZbEQERnAs6xgbEjbwy86/tLosEfEC/RaTpkshS1qghOwE3vv+PVxuFyl5KeSX\n5jN983SN1RJpodSiJSLSiPzsfjzQ9wEqD5sI9Q+1sCIR8aba+hmjznJsRgNcX2O0REREpFlo6Okd\nyu823A6kAQfLXmll60VEpIF8cfgL3t71ttVlNGtbtsD+/eZ7w4D334eMhmgSEDkHtQWtLkBX4D/A\nzUCbstdNZetERFqU/JJ8Pvzhw0a/brGrmJc3vswb294go0DJoL78/eGvfzXD1uLF8NlntTxsXKSR\neDLaeCDwUaXlj4ErvVOOiLRECdkJNIdhAu8eeJenv3yafSn7GvW6Hx38iIyCDErdpSzZs6RRr92S\nXHYZPPkkTJgA77xjPhMzPNzqqqS18yRoJQHPcLqF6y9AohdrEpEW5KTzJCNWjeDbE99aXUqt8orz\neHP7m/jYfJi1bVajXbfYVcwbW9/AEeAgKiiKJXuWqFWrngwD9u49vZyo31TSBHgStIYDMcC7wOqy\n98O9WZSItBzzds4jJT+FGVtmNOkpDN77/j3ySvLoENaBzSc2N1qr1scHPyYhO4G8kjyyCrNIz09X\nq1Y9rVkD334LixbBlClmN2JCgtVVSWunmeFFxGtOOk8ydNlQooKjSMtLY/oN0xnYaaDVZZ0hrziP\nGxffiIFBoG8g6fnpDOw0kJk3zvT6tf97/L+sT1hfZV2fmD7ccOENXr92S5OSAgEBp7sLDx+Gzp01\nTksaTkPPDP9/wFjgg2q2GcCtdbmQiLQ+83bOw8DA1+5LgG8AM7bMYMD5A5rc42aSnElEBUVR5CoC\noG1oW7ILsyl2FePv4+/Vaw/sNLBJhs/mKCam6vIFF1hTh0hltaWyyzGncYirZpsBfN0A11eLlkgL\nVVBSwJBFQygoLagIVm7DzcLbFnJx9MUWVyciUnf1adHyZOdQoABwlS37AIFAXl0uVAMFLZEWLDUv\nlWJXccWy3WanXWi78n+sRESalYaesLTcF0BQpeVgNI+WiHjgvJDz6BjWseLV3tG+wUOWYRhM+moS\n3yVpHmURaXo8edZhAJBbadmJGbZEpDbFxbB8OcTHw0UXwbBh5oyKYjp2DFauhIICuOEGuPzyep1m\nT8oe3j3wLoczDrPo9kVqLRORJsWToJXH6fFaAP0wuxJFpCZuN0ycCF9/bYartWthxw54+WWwN62B\n4JZISID77oO8PPP78f778OqrMGhQnU5jGAavb3kdR4CD+PR4tiRuYcD5A7xUtIhI3XnyL/5jwHJg\nQ9nrHeBRbxYl0uydOAEbNkD79hAdDe3awTffaAbFch9+CE6n+X2JiTHvyZ87t86n2ZOyh+0nt9Mm\nqA0BvgFM3zy9WcxALyKthyctWluBHsBFmHcb/gCUeLMokWavtBQqd2HZbObL5ar5mNakpKTq98du\nN9fV0ayts8guysaNORHqdye/Y2vSVvp37N9QlYqInBNPghaYIasn5t2GPy9bt9ArFYm0BLGx0KOH\n+TyQ0FDIzYU+faBTJ6sraxquvx6WLYP0dPD1NbsQ77oLgH/+95/c1P0murfpftbT3HjhjWeEqvOC\nz/NKySIi9eHJqNHJwNVAL2AtcANmF+KdDXB9Te8gLZfTCTNnwvffm6Hrj38Eh8PqqpqOnTthzhwo\nKoLf/AZuuokDad8zbMUw4rrENcqs7CIideGtebT2ApcC28u+tgUWA9fVsb7qKGiJSIWxH4/l2xPf\n4jbczBs6j94xva0uSUSkgrfm0SqfrLQUCAdSAPV/iEiDOpB6gE0nNhEdEo2v3Zd/bfuX1SWJiJwz\nT4LWViASeBPYBuwANnmzKBFpff617V/kFuWSUZCBGzdfHf2KfSn7rC6rQmJOIs4ip9VliEgz48lg\n+IfLvv4L+BQIA3Z5rSIRaZV6t+1NdHB0xbLNZsPX7un9Ot7lcrt49ONH6duuL5OunmR1OSLSjNT1\nX7EjXqlCRFq93/78t2es23xiM27DXfFQaqt8c+wbjmUd40TOCR7o+wAdwzpaWo+INB+aolpEmqQ9\nyXv440d/ZNNxa0cquNwupm+eTmhAKADzds6ztJ6zMQyDf2z8B0nOJKtLEREUtESkiZq1bRb5JflM\n3zwdt+G2rI5vjn3DiZwTOPwdtAluw/s/vE9izpkz/GcVZrExYaMFFVa1JXEL83bMY872OVaXIiJ4\nFrQGYo7LKhcG6GFiIuI1e5L3sDVxK10ju3Ik84ilrVofxH9AqVFKSl4K6fnplLhL+PLol2fsN3/n\nfB7/9HFS81ItqNJkGAbTN08nMiiStfFrOZFzwrJaRMTkyVwQOzFngy//k9IH8+7Dvg1wfc2jJSJn\neHjtw+w4tYPo4GiyCrNoH9qeZXcus2SsVlFpEQWlBVXWhfqHVhmon5afxi1Lb6GwtJCRl4xk3MBx\njV0mYI5pe+SjR2gb2paUvBRu7n4zz179rCW1iLRE3ppHC06HLDDn1PKpy0VERDyVVZjFgdQDFJUW\nkZiTSF5xHiedJy1rnQnwDSAiMKLK66d3Qy7avQiX20XbkLas2LeClLyURq+zvDUrvzSf9IJ0AFYf\nWF1tN6eINB5P7jo8AvwJmIWZ4v4AHPZmUSJSD243FBRAcHDVBzY3MxGBEXx535ldc7Ym+pnS8tNY\nuncpYQHmCIsSVwlv7377nFq1XG4XPva6/z17eYfL6RbVrWLZbrPjMvQgcxEreRK0HgKmA8+ULX8B\n/M5rFYlI3e3eDU8+CZmZ0LYtvPIKXHSR1VXVW1MNVdWJT4/H4e+g1F1KqbsUR4CD71O/P6dzjvts\nHINjB3NHzzs8PsZms/HEwCfO6boi0vCs/tdMY7REzlVODtx6KxgGhIWZYSskBN5/HwICrK5O6mhP\n8h5GvzeaqMAoPrz3Q4L8gqwuSUTK1GeMVm0tWhOAfwAzqtlmYHYniojVEhKgqAiiy2ZVj4yE1FQ4\ndQo6d7a2NqmzWdtmEegbSE5RDh/Ef8Bdve6yuiQROQe1Ba39ZV+/q2abmqFEmoo2bczxWSUl4Odn\nhi6AiAhr65I6e2XTK3x66FN6RPegsLSQ2dtmc0v3W9SqJdKMqetQpCVYuBBmzgS73exCnDgRbrvN\n6qqkDpxFTi5+/WIyCzLpeV5PfOw+OIuc/O3av/Gbi39zxv55eZCWdrrR8vhxM1s7HI1cuEgr0tBd\nhx/Uss0Abq3LhUTEi0aPhoEDISkJYmOha1erK5I6Wrl/JRGBEYT6h/KrLr/i191+DUCP6B7V7n/w\nILz6KkyeDD4+MGkSPPooXHFFIxYtImdVWyqLO8ux6xrg+mrREpFWz1nk5MYlNxLkG4SBQVFpEWvv\nXYsjoPbmqU2b4G9/M9+PHw+//GUjFCvSijV0i9a6cylGREQ8s3L/SpJzk4kINMfVZRdm8+737zL6\n0tG1Htex4+n355/vzQpFpL48mUerO/BXoBcQWLbOAC7wVlEicia34abUXYq/j79Xzl9QUqBB1xa5\nIPICHun/SJV1XSNq7/49dszsLhw/Hnx9zS7EyZPhAv3LLNKkeNL8tRF4DngVuAW4H/MRPJM8ODYQ\n+BoIAPyBNcCfK21X16GIh/617V/sT93P9BumN/i5tyZu5YWvX2DZncsI8Q9p8PNLw0tIMF+DBpnL\n334L7dpBly6WliXSonnrWYdBwOdlJz4GTAZu8vD8hcA1wGXAJWXvB9WlQBExn/+3cNdCNh3fxL6U\nfQ16bsMwmLFlBvHp8bz3/XsNem7xntjY0yEL4Be/UMgSaYo8CVqFmC1YPwKPALcDdfmTN7/sq3/Z\neTLqUqCIwNK9Sylxl+Bn92PWtlkNeu5tSds4kHaATuGdeHP7m+QV5zXo+RuCs8jJ2vi1VpchIlJn\nngStx4BgzJng+wEjgfvqeI2dQDLwFacnQhURD2QVZvH2rreJCooiKjiKzSc2N1irVnlrlr+PP0F+\nQeSV5DXJVq1le5fx9BdPczD9oNWliIjUiSeD4beUfXUCY+pxDTdm12E48CnmtBHryjdOnjy5Yse4\nuDji4uLqcQmRluuDHz4gpyiH8vGMBaUFLNqziL9d+7dzPvfelL3sSt5FsG8waXlpuNwuFuxawPA+\nw7HbPPk7zDsMw2DBrgXc0eMO3IabBbsW4GP34d/f/Zupv55qWV0i0rqsW7eOdevWndM5PBnQdQXw\nNNCF08HMwBxzVVeTgAJgWvl5NBhepHbJuckczTpaZV1MSAxdI899UtKCkgJ2J++usi7QN5BL2l5S\nPujTElsTt3Lfe/cxbuA43IabN7e/yXkh55GSm8KSO5ZwYZsLLatNRFqv+gyG92TneOBJYC9m61S5\nox4cGw2UAlmYg+o/BZ4HvijbrqAlIlUYhsGYNWPYn7ofX7svLreLIL8g/H38Sc9P59qu16pVS0Qs\n0dATlpZLBd6vT0FAe2AB5jgtO/A2p0OWiMgZtiVtY3/KftqGtuVo1lH8fPyI8Y8BoIOjA2kFabgN\nt6VdmyIinvIklf0auBtziofisnUGsLoBrq8WLWn68vLMZwi2aQNRUVZX06KVt2btTt5NeEA4xS7z\nn5xPRn5CWECYxdWJSGvnrRat+4CLyvat3HXYEEFLpGnbvRvGjoXCQjAMcxruO+6wuqoWq8RdQkRA\nBH1i+lSs8/fxJ6MgQ0FLRJolT1LZD8DFmK1YDU0tWtJ0uVwwZAgUF0NYmPk1KwuWL4fOna2uTkRE\nGpm3ZobfBPSsT0EizZrTCdnZZsgC8PcHux0SE62tS0REmg1Pug4HYk44egQoKltX3+kdRJoPhwPC\nwyEn53SLltsNHTtaXVn19u83Q2DnztC9u9XViIgInjV/dalh/dEGuL66DqVpay5jtObMgX//G2y2\n03UOG2Z1VV5V6i7Fhg0fu4/VpYhIK+GtebS8SUFLmr6mftdhUhIMHWrW5+trtrxlZ8Mnn0BEhNXV\nec2L37yIv48/T131lNWliEgr4a0xWiKtW0gIXHhh0wxZAJmZ4ONjhiwwx5LZbGbYaqFO5Jzg/R/e\nZ/WB1ZzKPWV1OSIiNVLQEmnuYmMhKMi8I9IwID3dbMlq397qyrzmrR1vAZjPQdy5wOJqRERqpqAl\n0tw5HDBjhjlg/+RJaNvWXPb3t7oyrziRc4K18WuJDo6mTXAbVn+vVq26Wrl/JW9tf8vqMkRaBU/u\nOhSRpq5XL/jwQygpabEBq9w7e98htzi34qHXziInK/at4NEBj1pcWfOQV5zHjM0zKHYXM7THUKKC\nmmiXuEgLoaAl0lLYbC0+ZAHc0fMO+nfsX2VdbHisRdU0P6sPrCa/NB8bNhbtXsSfBvzJ6pJEWjTd\ndSgi0krkFedx4+Ib8ff1x26zk1OUw9p716pVS8RDuutQRERq9N7375GUm0RGQQZp+Wmk5aexZM8S\nq8sSadHUdSiyezfMnGk+cmfIEBg1ynzUjjSawtJC7DY7/j4tv+vTShe1uYiJV02ssu7i6Istqkak\ndVBNjXIAACAASURBVFDXobRuhw/DyJGnxzfl5MDDD8ODD1pdWasy/j/jiQyM5OnBT1tdiohIjdR1\nKFJXGzZAUZE5GWloKERGwurVVlfVqvyY8SPrjq5jzfdrOOk8aXU5IiINSkFLWjd/f3OSz3KlpRAY\naF09rdDsbbPxsflgYDBv5zyryxERaVAKWtK6/frX5gSfiYlw6pT5XMOHH7a6qlbjx4wfWXdsHW2C\n29AmuI1atUSkxdFgeGndoqLg7bfN7sLcXLj6arj8cqurajWW/397dx7eVJm2AfxON0oX9pYdyiK4\n4KAzKrgO4uCKAi4o7jKDCOqgoqOAo4xjcUcQUEQ20UEQFAVFEYGyyCebskkFpOy0hQLdaGmb5nx/\n3DmmxXTPyUnb+3ddvUjSJOfNaWmePO/zPu/2T3HaeRonc08CAHIKcvB54ud49JJHbR2X0+VESJD+\nPIpI1akYXkRsk5yVjNRTqcVuaxndEjGRMTaNCNhweAPe/L838XG/jxEaHAqAcXhEBBel5ucDY8dy\nDUWrVv4bl8twYeORjX9o1ioi/lOZYnh9ZBMR2zSPbo7m0YGz+bVhGHhn3TvYkrIFS/YsQe9OvQEA\nl14KjBrFIGvjRm4rWdae3fmF+XDA8XuwVlUr963EM0ufwUf9PsI5Mef45DlFxHrKaInURobBFZe/\n/ca0zDXXqHcYgHWH1uGxxY8hqk4U6gTXwaIBi34PlPbvBx57jPf74gsgOLj05xq5bCQiQyMx6qpR\nVR5XoasQd8y7AzvTduJv7f+G8TeMr/JzikjFqb2DiJTPxInAk08CEyYAzz0HvPBC8dWXtZBhGJi4\nfiLqhNRBVFgUTuSewJI9SwAwkzVtGtC5M9C4MbB0aenPlXQyCUuTlmLhzoU4knWkymNbtX8VDmYc\nRFzDOKw9tBaJxxKr/Jwi4h8KtERqm/R0LgCIiQFatgSaNQO++w7Ys8fukdlqc8pmbE7ZDKfLiWOn\njiHPmYcpm6YAAObM4XTha68Br7wCfPopcOBAyc/1waYPEIQgtqz4uWotKwpdhZiwfgIiwiIQ5AhC\nsCMYkzdOrtJzBhrNbEhNphotkdomN5ed8M25r6AgXs7JsXdcNmvXsN0fpuQiQyMBAHfeCYSE8DQ1\nbw5MmgTUrev9eZJOJmHZ3mWIiYyBy3Bh4c6FeOjCh9AiukWlxnUw8yBST6UityAXWXlZAIBtR7ch\nKy8L0XWiK/WcgSTpZBLiV8Xjvd7vaQsmqZEUaInUNrGxQMeOwK5d7ISfmcn5sA4d7B6ZrRqEN0DP\ndj29fq9OneLXSwqyAODTXz5FbkEuTuSeAACcKjiFz3Z8hse7PV6pccU1iMOah9b84XZ3rUi1N2XT\nFKzavwqLdy1G33P62j0cEZ+z+3+qiuFF7JCWBowZA2zfzgBr5EigdWu7R1UjHMk68oemq63qtULT\nqKY2jShwJZ1Mwp3z7kRUnSiEBoXiq7u/UlZLAlpliuEVaInUZCdOAAUFrMeyYFXhgYwD+Dzxcwzr\nNqzGZFjEf577/jms2LcCsZGxSMlKwagrRymrJQFNqw5FhFwu4NVXgeuuA26+GXj4YU4R+tiUjVMw\nZdMUbE3d6vPnlpptf/p+fL3ra+QX5uNI1hHkFOTg3Y3vwuly2j00EZ+y+yOoMloiVli8GPj3v7mP\nY1AQ93Hs04e3lVdhIXsafPUV26I/9hhwxRW/f3tf+j70n9cfDocDXZt2xfu931dWS8rtZO5JLN+7\nHAY87wF1guvgpk43IcihHIAEJnWGFxHasYNL5MyVhfXqAVu2VOw5ZswAJk/mfpAZGcDw4Qy8unQB\nAEzdNBUOhwMxETH4OflnbE3diq7Nuvr4hUhN1bBuQ9x27m12D0PEcvrYIIFlyxagXz/uefL446wx\nkopr1w5wOj1NSLOzK76q8OuvuSqxbl0Gak4nu8mD2ayFuxbCAQdO5J5AjjMHkzZMqvAws/OzkZKd\nUuHHiYhUF8poSeBITeX0lMPBN/gffwT+9S9g6lS7R1b93HILsHo1sHYtpw5btgSeeqpizxEVxdWJ\nERG8bhi8DWyieUvnW+AyXL/fvUlEkwoPc/yP47Ht6DbMvm22potEpEayu6BCNVrisXIl8OyzXCEH\n8I09JYUBQ2mNi8Q7lwvYvZv7x3TsWPFzuGEDs4oFBfxZtGgBzJrFqUQfOJJ1BP3m9IPT5cS468fh\nyrZX+uR5Kys9HWjQgJcNg7Ol5nUREUA1WlLdRUWxANswmNXKywPCwv7YLVLKJyiIm/NV1sUXAx9+\nyKxYeDhXMPooyAKAGT/PgAEDEWERmLB+Ai5vc7ltWa3cXG79+MgjwCWXMJ7ctw948UVbhiMiNYgC\nLQkcF14IXH01sHw5Ay2Hg5sdW9D/Scqpc+eqBWslOJJ1BAt3LkTjiMYIdgRj78m9+OHAD7ZlterW\nZc/Wl14CoqO53U58vC1DEZEaRoGWBI6gIO7Yu2YNi+A7dwbOPdfuUYkFlu5ZivzC/N+3qXEaTiz4\ndQHOb3o+osKiEBLk/z9NHTsCbdtyPcawYQy4RESqSjVaIuJ3+YX5SD+dXuy28OBw/GPRP3BDxxvw\n0IUP+XU8hsHpwk2bgAcfBN5+m+syunXz6zBEJMCpRktEqoWw4DDERsYWu23F3hXYdXwXUrJTcPu5\ntyO6jv9SSqdPc4FlfDwzWS+8ACQkKNASkapTRktEbFfoKsQd8+7A8dzjyC3IxdCLh+LBCx60e1gi\nIsVor0MRqZZW7V+FgxkHER0Wjfrh9TH95+nIysuye1giIlWmQEtEbDd7+2wUuApwIvcETuWfQsbp\nDHyf9L3dwxIRqTJNHYqI7ZKzkpGRl1Hsttb1WiMyLNKmEYmI/FFlpg4VaImIiIiUg2q0RERERAKI\nAi0RERERi6iPlogdEhOBzz/n5X791AFfarSkJG7QbW6VuXkz0KULtzoSqemU0RLxt+3bgYEDgS+/\n5Nff/w5s22b3qER8ZtcuYOpUwOXi9alTgYce4s5ay5cD48YBx4/bO0YRf9HnCZGi1qwB5s4FgoOB\nvn3ZGrxuXd8e45NP+A7UtCmvHzsGzJ7NfR5FaoBWrRhsTZ4MtGjBX/E+fYAHHuD333vP8+svUtMp\n0BIxrVkDPPkkN7feuxf44AOgXTvgiSeYdXL4aJGu08ljmIKCgIIC3zy3SACIiABGjwbuvJPXp01j\nIrfo90VqC00dipjmzgXCwji/kZ8PhIYCubn8+L12re+Oc9ttDLZOnABOnmSQdfvtvnt+kQCwdClQ\nvz4QEwOMGQN8+CH/Kz3wADBiBH/9RWoDBVoippAQwDCAjAwGWeZtAPDrr747ziWXAOPHA+3b8zjX\nXQfExfnu+UVstmUL8NVXwNtvAxMnAunpwDXXcErx9tuBu+4CwsPtHqWIf6hhqdRMBQXAzp283KkT\nM1Vl2bQJGDqU04Y5OXzM+ecD2dnAf/8L3Hij78a3YwcwaBBw+jSDu0aN+JG/ZUvfHUPEJoYBZGUB\n9erxem4uyx7L899QJJCpM7zUHoWFwIIFLPxo1w7o399TtH7qFAMmMwvVsSPnLMy/+qXZvBmYPh34\n+msgOprvDJdfDrzxhm/Xoj/xBLBuHdCkCa8nJwP33svbxWfy84Hdu4HzzuP11FTO2iqeFZHKqEyg\npWJ4qZ5eeYWBVlgY303XrGEwFRICzJwJ/PIL0KwZ77tzJzBlCvD002U/7wUXAO+8A7z8Mh8XHs53\n6SAfz7JnZ3umJwGOOzPTt8cQpKQAr74KPPYYZ2dHjQLuuEOBloj4j9WBVmsAswDEAjAATAHwjsXH\nlJouPR1YuJCBVFAQ5ym2bGFgdN557I5Yp45nlWDdupwOrIh69YCLL/b92E033QT8/DMDLJeLaZbr\nrrPueLVUmzbAiy9yMSkADB6s02yHHTuAhg2B5s15fc0a4KKLVKcltYPVxfAFAJ4EcB6A7gAeBXCO\nxceUms7p5L9mIOVwMOAybz//fNY+uVz8ys0F/vQne8Zakr59gWee4bKs2Fhm6Lp1s3tUNVJ0tOdy\nTIx946jNDh1iNjE5mZ+RPvyQSV2R2sDqjFaK+wsAsgEkAmjh/lekcho3ZlCydi3fRXNymLro3Jk1\nVqtXM7hKTGSReY8ewIMP2j3q4hwOLr266y67R1KjHT3KN/jBg4Gzzwb+8x8mQLt3t3tktcu11/Iz\nz8MP8/q0aZ7yRJGazp81WnEALgSwzo/HlJrI4QBeew14/30GVu3aAY8/Dhw8CAwZwuxW06Zs1PPg\ng8Cjj/qu2ahUK+HhXGPQowevv/gikJfnm+fet8/TlcMwgP371aWjNPn5nsuFhfaNQ8Tf/BVoRQGY\nD2AYmNn63ejRo3+/3KNHD/Qw/yKKlCYiwlN4Y/riC/41b9GC14OCgO+/ZyW01Er16nmCLIALUH2h\nsBAYO5aZsQEDuIlAUhJngBXT/9HixcCiRcxk/fQTs4yvvspZc19xuRjwBgfzutOpTaul6hISEpCQ\nkFCl5/DHn4RQAF8B+AbAuDO+p/YO4jsff8x3P3NJWWYmC+bnzrV3XFIjpaczYDhwgK3aXnoJiIy0\ne1SBadcuoEEDT2C1YQPQtatv+2otWsRqgeHDWTnwwgvcu71LF98dQ6Qy7R2sLoZ3AJgGYAf+GGSJ\n+Nb117Pa+cgRruvPzWU/LREL1K8PtG3Ly2efrSCrNJ06Fc9eXXxx5YOsnBzv16+7ji30XnqJAfC5\n53r6p4nYyeqM1hUAVgHYCrZ3AIARAL51X1ZGS3zr6FFOIWZnA1dfDVx4od0jkgDmdDkRElTx+SXD\n4HThzp3AU09xL7/LLwfuvtuCQcrv8vL42WnoUOAvf2ErvU2b2PYO4Nah99/PywsWaOpQfE+d4UVE\nysnpcuLBLx7E0IuH4rLWl1XosYWFnJHu04eZrPR04NtvgTvvDNwarV27uOHB448zAJk3j9N5vXp5\n7pOfzzqzAQOYhUpMZH3VQw9xAW8gSEwE4uO5BiY1lUFukyb8bPXCC6zDS03lz2X4cE/NlogvBOLU\noYh9DAP47DN+xB08mFW4Im7Lk5bjp+SfMO7HcXAZrgo9NjiY2StzurBBA3bqCNQgC+CKyMxM4K23\ngDlzgGXLmBUqKiyMXVJuuombKbz8MnDsGLN3geKcczgluHkzu/ybbSJWr+b3HnmEU4enT3MXLn2W\nF7sp0JKaa/58ftzdv597Ij76KD8OS63ndDkxccNExEbGIulkEtYeXGv3kHzqwIHiAcaBAwyiRoxg\nV/b//Y/7pHvLUt11F/vp/uc/nIl3uYB//tN/Yy/LggXc6OHZZ9n4dNMm3n799Vxl+sgjwJIlzGZN\nn87rqan2jllqNwVaUnPNn8+/vPXq8R0lPx9YvtzuUUkAWJ60HMnZyYiuE426oXXxzrp3KpzVClSG\nAUycyCDDMIDZs4E33+R055dfsii9fXt+39xMoajERK4nad2aNWhXXOHZr91ueXn8zDRmDMc1ahQD\nR4DZxOuu42t64w3uM79pE3DZZb5tIyFSUXYnulWjJda57z52lWzQgNcPH2ZD00GDbB3W7xITWdgT\nEsJinzZt7B5RrXHHp3dg5/GdiAiNAABk52dj2i3T0K1V9d0Gae9eBhSRkaxXGjKExeFt27Km6fBh\nYMIEBilRUazF6tqV2StTfj4fFxPDTFbLlpw2nDHD2q0/fSk5GbjnHu4rf9VVwKefBvaUrlQvKoYX\nKWrNGi4JA/hxvkED9toyd7a10+bNfEcrKOD1yEjOg9Tk1uJJSUyVtGlje1C54fAGZOZlFrvtz83/\njIZ1G/p9LLm53AM9P58xt2Hw1zUsjD13vTEM1iCZmab8fE4Hbt/O9gZr17ILfnQ0t7956ikGG7m5\nnscUFPC2M1fmpaUxCzZoEO+7dSu/7r3XunPgKzk5wHPPsU+Xw8Hz+tRTwM032z0yqSkUaImcafNm\nYOlSdpLv18/TNd5ujz3G4vzGjXk9JYXFMcOH2zsuq3z0EdMpwcGMEp5/Hujd2+5R2W73bk5z9e7N\nFYHh4SxQ37qVjTZvvZUz32faupU7UMXH81c7Ph646CJmcxYtYgF75868/c03uc/6wIE1P7MzezbP\n47XXArfcAjz9NAPSsWO5K5dIVSnQEqkuBg1iAUz9+rx+9CjfGUaNsndcVkhO5vxUw4ZAaCjf+bKz\nge++Y8olALlcXKFnzjrn5DC7FB7u+2MtWAAsXMhs1N69bLfQti1w6BCzU+3be3/c7NnAihVMhrZo\nwRh9+XLgnXeYlZo/n1N/2dlsLTdgQM1vdVBYyK/QUAaVZg2a+mmJr6i9g0h10bcv21hnZrIJk2EA\nN9xg96iskZbGKCU0lNfDw/l609MtP3RGBoMR0+7dDGjKsmMHp5ySkxlkvfgiE6MVtWEDa6MAvuQl\nSzh9V9TNN/MUxcQwqFq+nBmrF14oOcgCgNtvZyJ0zx7gH//g4/73P2DyZPa9GjuWv2JRUZz2qw5B\nVtHP3YZR8dYMwcGccjUzdyEhCrLEfgq0ROxw443A6NFMXZx1FjBuHPDnP9s9Kmu0bs13uwMHmKbZ\nv5+ZLD/M5eTnM/OzcCGDrJdeYvBRli5d2KPpqafYgq19e/aWqqjMTCYpDx9mEPTVV56yPIAZl9df\nBy64gEFCfj4XyO7cyculva74eODKK9kk9d//ZiYnPp5ZrEGDuDFCSTVepqNHuZLPdPiwfX2nEhPZ\ntys/n2OYMYNrRUSqO8X6InZwOJjKqEqVbk4Og5Z69TwbaftSQQHnskJCWKRf1rt2SerX58Zzs2d7\n5nX69PHtjsIliIlhcPXww7w+alT5V8/99a/Au+/yct++lXv511zDoOGRR/iyZ8woXnN14ABP7403\neqYr776b6wb+9S9OA3bs6Ll/YSEDsj17OK35z3/yenAwn9/8NXA4ipfAmY878/qiRVyY+/zzzPS9\n/TYzYd7aIZgZJvM8nPmcVdWpE4vXx4zhXvC//spgt7oq6ZxL7aMaLZGS/Pgjl281asR3WrNgJxAk\nJXHDt4wMpkXuvht44gnfVTunp7PB62+/8d31ssuYeqlMcLR3L9MuTZowmgCAEyeYrrB4X5c9e5gl\nSUnhm1zr1sz09O9f+uPM6cL27Zl0nD+f2aIzF6wmJrIbOcAfQ1ISAwaTYTCTNXcuA6EJE0qOiQ8d\n4rTiWWfx+saN7ODerx/3Udy5k48fO7ZiP4ajR9l8dPRoBp5z5vDH+8gj/HG8/TaQkMAVhi+9xA2y\nvVm8mK/3ySeZBRs9mm0U/vSn8o+lLE4nXy/ARbiBsu1PRZV2zqV6U42WiK98+SVXBs6Zw3e3hx4C\nsrLsHpXHv//N9EeTJvxLPns2C4J8ZeJEbo4XE8P0xurVjDYqIyeneNrFrFQ+s1jJAiEhDFCio7ma\n74cfyhekpKQwgBo8mNmm/v0ZcxaVl8dfjU8+8UwBfv558fvMmwesW8euIkOHMqOWlub9mK1aeYIs\ngOONiWEgNGMGa7bMLX8qIjaWq/BGjuSPdeVKxr0As1NXXcXLhsH9A0tyzTUMFl55hWNp04ZTrL5i\nGMCsWUyeNmjAgC4/n78ms2YxeVtdlHbOpfZRoCXizcSJ/GvftCmXdB08yGAjUCQleTJs5nyEWXXt\nC7t28V3d4eBXaCiLnCqjXTumJtLS+M559CjfTf1Qo9W8Odsg5Oaym8bs2eWbOmzfnu0QzGmy669n\nPVRR5jRXQgKzMIWFf+zOcdFFzKjVrw/87W+s+WpYzlZdDgfL9jIyGGiYwZ+5pqAi+vRhbdqSJcXH\nsGkTMH488NprQPfuHGvRmq0zX+/w4Uz0/vorg9DKziZ788svXATwt7/xfG3dym2CevfmuM09DUsS\naJ39SzrnUvso0JKa6+RJ4PvvueysPBXQReXn/7GgorTqZH/r3JmvD/CsYfdlE9BzzmFfAMPg/FJB\ngWeOrKIiIoD33mP6o7AQ6NaNqSArloP9+ivwzTfAzz8DhoHQUGaxOnfmos4PPvDel6qyoqI88W5c\n3B+DoPbtPR08AE6zeavTST+djj0n9vzh9jZtGPhkZvKrY8eKzw6npzMxW78+A0Jzo2iA/z7/PEvo\nnnyS055mojE5mRtQm7/2337LxG7PnuwoP368Zyb41ClmujIyeP2XXxjgVkSXLuz5dfPNQIcOXCAw\naxbj8/ffLz2bN3f7XIxYNqJiB7SYec5vu43ZTvOcS+2jYnipmQ4fZkrCDEZatuT8S3nrrPr04XxP\n/fqeFtyBtAfJf//LGipz199BgzjX5CuPPcYCp23bGGxdey27Z1ZWmzaMcqw0bx67fwIc8z33IOnm\nJ7B3L+tlIiJY9/Ptt74psjanC6OiPFN7ISHsV1VR73z+LH7c/i2+TOuF0DvuBHr1gmEAM2cyK3f0\nKDMis2YB553H7Nw115T9vJmZXAgQFsbAyNyOZsEC3n799Z5VhkFB3LXKvB4by9f4yivApZcy6Lrq\nKmDYMMbd8fEMqM4/n+e2dWsGbXffDUyaxGahFWXG3gMHMoNYUMBgdcMG1tZ5k1OQg/c2voesvCzs\nvGAnOjfpXPED+9jRo5wyHjOGP7eGDT3nXGofFcNLzTRyJLBsmWf5VHIy8OCDDCDKw+kEpk3jczRu\nzELzzvb/AS8mP59b2kRHezrM+5LLxYA1JITLwAK5rfipU4w86tdnVFFYCKSl4cCbn2LasjiMHMmp\nrzlz+O2qxIwAT3tMDNs19O7NrEvdusD//V/F26EdWrcUt83ph4Ig4KU9bdD7UAQwZgyMXtdi5Ehm\nsY4c4WxxYSGn0Dp14tRdeX4k5ibM9erxV+U//2ErM3Pab9IkZqiuuIJrFF5/nf996tXj8cy9EMeN\n4yyw+TiXq/jUoWHwcdu3A88846n9qqjsbAarp09zGvHQIX5eev5574X6s7fNxtv/9zaCg4JxWevL\nMPa6sZU7sI+deX6KXk9N5X9ZM7BMTg6MncGkbCqGFzGlphZv4x0ayo+Z5RUSwneyTz/ltFegBVkA\nI4a4uLKDLJeLxUm33sptflauLN/zBwUxTdG8eWAHWQBTN4bhqXR3F9+3ikpHgwZcTffRR3zpPXpU\n7VCGwQL12bMZhCxcyOm46OjK9Zyd/t1rMGCgPupgUrs0FETUAebNg8PB2p6BA4ERI/hjSElhhqe8\nQRbAacFGjRgz33ors09FA4CbbgKmTOHrGDWKdWHm9OqqVUwCt23LBK85Sw38sT5rxw6WMnbuzAzf\nZ595vrd+ffkaxQLcNevGG5kBeuYZLhD4xz+Kr+Y05RTkYMqmKagfXh+NIxpj9f7V2Jm2s3wHstiZ\n56fo9U8+YYbQ6eTC5mef5a+w1EwKtKRmuvxyfjR2Ojn/kJ/P2qDqIi+PU2HjxrHGrKzMr9PJfVbG\njeM6fFeRwuB58/hXPT2dH52ffpq9A779lvdfsKD4O2h1FBPDSOTYMU/X+fBwBLWPw7BhLKz+9FO2\nbKhqywCHg4s+N21ioPXddwzkKtMj6VDmIXxt7EKTvBBEuUKQFpKPJQ1P/F7sFRPD4+XlcfFmnTqs\ngyosLN/zu1ysb6pbl1ObkyYBW7YUv09cHBO2H3zAONVsfXHoEKda4+M57RgezuDSm1OnOGv79NP8\n9+KLOeX45ZecQpswgWMvjyuu4H7rUVEMTv75TwbH3grvv/ntG6RmpyIrPwtpOWnILsjGzC0zy3cg\nGw0dyozdbbfxZzJ6tG9rByWw2P0xVVOHYg2nkw2H5s/nO+DAgfxYHOiZGYBjf/RRFqYEB3tqsEpq\nwuNyAc89x2nO4GC+C996K+dxHA7uv3LokGdfwdRUvoObzaWcThbAvPGGb5eRVZLZFbx3b878ZmZy\nym/gwDLq5w8c4HnYuZM1eWPGAF26YO5cbk/TtClPzQsvlP9NvzQzZzJr07Nn5VuYzdk+B68vfxlB\n+/YDhgsuh4ErT9TD+EcX/V4T6HQyo9WhA/D3v3OFYHAwX2pZx8zKYgD1yCPMZG3fzvqu++/33OfE\nCWay2rdnSd7DDzPYAfhZJSqKlwsL+Xmlbl3vxyp6X8NgI9R//pPXx44t3rrCm/x8T0LS5fL0ti3N\ngYwD2HFsR7Hbmkc1R9dmXUt/YABYuZLF/y1aMNjSVkHVgzaVFjmTmdkJgACi3H76ie+MsbGenXGP\nH2d7CW8RQlISpwRjYvg63fVJ+OorPsfgwXyHNVM5Bw+y6OW88/iObRicVv34Y+/zMzZYtIhTWc8+\ny2zKJZcwXixXMFOkBfeePUzmvfwyp8DGj2fi6667qja+zz5jJmvECAYRf/kLg5eKBlsuw4WCwgJg\nbxLw+QKgoADBN/VGyIXFFzb89BNw4YWeX4cdO3zXKPSNNzg12L8/g6OXX+ZrqmqGZd06PhfAzzh9\n+pR83337mAGLj+ev6fjx/NW9556qjSFQrV3LPSlHjeKHiPBwts5QsBX4KhNo6ccqNVt1CrBMeXkc\nt/mubQZDBQXeAy3z/uZrNR9rrssfMoRfR47weerV4/fN+zscno32AsTNNzOT9eSTnDYqd5AFFJvD\n69CBs6NmpmTYsOKzqpVhGMzejBnD8riXX+bsrMtV8enDIEcQ6oTUAc46B3i25PYZRbfBDAnxbTf2\nhg09RebmxtZVTfzu2MFWdGPHMsAdOZIJ1Z49vd8/Lo71bc8952mvNnRo1cYQyFJTOV3Yvj2D9Y8+\n4n9jBVo1kzJaIoEmM5PphfR0Ng9KT+dczvjx3u+fl8c19QcP8t0sI4M9r2bO9Lzz797NzpphYWzV\nMGoU54nq1eP8UosWrNAtaV7Iz8zNmFNTuZAwPt77/ntSddu3M5v02GOcae/UidOHVQm28vM5M222\ndjt2jL96RXuKncnl8mS9Jk+2ZvtOkarS1KFITXHwIAs4Dh3ivNSwYaV3bDx2jPfftYudH8tqRZ2R\nwTm1lSsZbA0ZwpSCr2vYtm/nBnmNGnGX5nJ8ZDcMThmefz4zWV99xfp+q3qc1gbm7LCZLcrNivrO\nPwAAG6tJREFUZXxutpUzp/natuV59ncpo8vFzxFpaczWLVvGjGFZ3eBF/E2BlpTfsWN8s23ZMmCy\nGDVSZibTMk2bBt6yorfeYhbLMDiNeM01fDfOyGDW6+9/r1pks2gRG6sWFvKd+4oreMxyzK+lpnpK\n1Mzrftixp8bav58Zwn//m1mm//yHvbMGDOCKwRde4Mz08ePAiKEZ6BJ7lL3TzAUUfhjfrFnAv/7F\n2fEvvuCvzW23+eXwIuWmQEvKZ+pULkUKCuJH2kmTWCwgvrVqFYtTzOLsMWMq38XR144cYW+CJk04\ntqwstnw45xy+uaanc7+Vxx+v3PO7XGztHRHBSl/DYLQ0aRIr2/0sJ4f1/+Z01JEjnMaqzCbN1clv\nv7HhKcBmqi+9xM9VV1/Nha1BQUyERkdzunDvjBVwPvc82se5EFInhEscL7vM3hchEkDUsFTKtmUL\nG+s0asQ32fR0VmOKb2Vk4Pd25E2a8N+RIwOnK2FWFt9lzexSVpanUD4ykmNetKjyz+90cm7KLN43\nC+4ruuekj+zYwdN/4ABnY0eO5PYxNdnp01xRaLZJW7aMq/sKCoBevTxrIQYPdtdknTyB9lNHof25\ndRHStAl7Kzz3nE9/Zmeutwig9RcillGgVdscPMg3PXNKqFEjroGv6lKsmsYwgKVLOafyzjssHqmI\n1FRmsiIieD0igtdTU30/1spo25bBVFoa34Wzs/nGanbTLygo3lm/osLC2CA2JYXPlZ7O5z/3XN+M\nv4Iuuog7MD36KMvR7rvPlsSaX4WHcxHBV18Bt9zC/rQPP8xf6ZdfZrsxgNkshwP83TQMhES7Swki\nIxkJVWRHhTKYgR/A9RmPPmpb7C3iNwq0aptWrRhEmJ3AT57ktGF1bINgpblz+Wl+6VIWjzzwAIOF\n8oqN5TnNyeH1nBxer0ihkcvF9uPLlnH/FF8KDwfefZdThadOAd27M/JISWH3+Ozsyk8bmuLjOUeV\nm8t6n0mTbC20Ktows6zmmTVFgwb8LOV0siZr2DD2QR02jJm9Ypo2ZcSVm8vrp07xA1lMjM/GM3gw\n8M03nK586SX216rp07ciqtGqjaZMYZ1WcDALVSZNYsMh8bjmGgZGZlYnOZmNb3r3Lv9zJCSwAtls\nsBQfz5V35eFyMfWwZIlniu+tt4BLL63oKym/9HSmPzIyWJdz4YXWHcvPDh3ipsT33cfTOXMm6/TN\n9gM1kdPJX5nTp5nJevFFLizt16+UB33/PX/vXC4GWa+8wlo7H/rxR/5X6NKFTy9SnagYXsovNZX1\nQq1aadWhNz16sL7I7HSZnMwlW6W1t/YmI4NZombNSm8idKb16zmvYmbGsrP5xrd0acWOLwCAvXu5\nss3cUHrVKhbG19TPF04nPxe0aMGs0b59DC67d+c0YqnS0/n3oXlzn6+U3b2bmay77uI+iGUGfiIB\nRp3hpfyaNtV6+dL07w9Mm8bN206fZiFL9+4Vf5769SsWYJlOnCje7T0ykgGb0+m95UJaGtMXu3Zx\na52nnvI0SRK0a8cvU6As/rRKSAi7sM+axa7v06d7pg3L1KCBZb87ixezMWq3bvx68012EtH0odRk\nymiJeONyscfUihUschk6lPuE+FJSEp8/LIzLwJo183xv3z42OYqIYMbx6FF2cpw69Y/Pk5/PzvAH\nDjAwzMxksDV9esX3hJEa5f33ORs8cGBgZI4Mo3gz1DOviwQ6TR2KVBc7dgCDBjFbZhgM5j78sPi+\nIytWcP7n1Cm2SX/9de+Fyb/+yiV15vcMgw1pP/uMU8NSK+3axWm6889nc/6XX9avg0hVaepQxDA8\nnc4D2ZQpzJq1aMHrycncmfiJJzz3ufpqFhUVFHhqxbwJC+NzuVx83eY5KO0xVti1iy29DYOFQOeU\nvElyRRiGYf5xk3JyOoG33/ZMFy5fzutvvqkMkoi/Bfi7kUgFfPcdC1O6dQOGD2cTzkBl9q0yhYR4\nb2bqcJQdMMXFMSBLSWERc3IycPPNPl2W75XLBWzezMzbqlXsJD9vHncmHjiQm1ZX0dFTR3Hv5/ci\n/XQFWmsIQkKAceM8NVk9e3JjgtKCrMJCPubYMV5PTua+h2qxJ1I1CrSkZkhM5Pr94GCu1EtICOy1\n4zfcwKBo82Z268/KYp1WZQQF8V30+eeBO+9k34JRo6xNXbhc7BcwaBD7jd1zD3uyNWvGL8MAPvqo\nyoeZtWUWfjz8Iz7Z/okPBl27mE35S7p+puBgLhgYORLYupW/QmedFfjJYZFAp6lDqRm2beObv9mq\nIiYG+OEHe8dUmsaNOSVoNjSNjKzc6kRTSIh/q503bGCrcbP9RHIyNxA0+yUEB/P1VcHRU0cxf8d8\nxDWIw0dbPsKALgPQIFwrKa3Upw9/lKNGMWa//nq7RyRS/emzitQMDRp4apMABjBNmtg7ptLMmcP6\nrMsu41d0NBsLlcXlYkCTkuJ5rXY4cQJZIS5PuqN1awZWJ06wD1N+PnDrrVU6xKwts+AyXIgIjUCB\nq0BZLT9ITmYLt1atgJUrPdOIIlJ5CrSkZujRgwUpqan8cjr5sby6KM8691On2ISob18Wmz/7bJWz\nRpWV2DQI/S76DUedGRx7QQGL97t0YRH8G29UqaP4idwTmLt9LvIL85GSnYL8wnzM2jILWXkBXHdX\nzRUWcpVi//7Ae+9xE4T4eNVoiVSV3etP1N5BfMfpBNau9bRDCOS17D/8wBWGISF8hwsJAWbMADp3\nLvkx48ax7ql5cwY3yclsTHrvvf4bt9uwb4bh6y3zMWRrGIbvasy+XW+8walEH8gtyMX3Sd+j0Cj8\n/baQoBBc2+FahAX7eTVlLZKeXrxX6ZnXRWo79dESqU7Wr2c7hNBQ7klSVjuEQYPYQsHcFuX4cWby\nxoyxbIjeWiskHkvE/QvuR6OIRsg4nYGFfecitklby8Yg5fTjj1wEUr8+cPvt1q86FamF1EdLpDq5\n5BJ+lVfHjsDPP7OeCwDy8rgszEKvr30d7Rq0Q//z+v9+2+SNkxEcFIyw4DC4DBc+2j0fw5sMt3Qc\nUobFi7kZdHAwM7sLFwL/+x8b4YqIrVSjJVJdDBkCnHsuK5SPHWO/sAEDyv/43FwWqZfTocxDmL9j\nPt7d8C5O5Z8CAOxP348fDnI15/Gc4zAMAwsSFyA7P7tCL0V87P33GYDHxnKRRWoqu5SKiO2U0RKp\nLurV4/6Fe/ZwtV/79uXbyzA3lz2vli9nwf2DD3LvxjKK76f/PB0OOHCq4BQW/LoA9/7pXrSs1xLT\nbpkGA54p/5CgEESERlTxxUmV5OUV/11wOCoUVIuIdRRoiVQXq1YBM2eyEP7uu8s/bTh5MrBsGRuJ\nulwM1jp0KLVJ0qHMQ/h619doEtEEBa4CTP1pKvqd3Q+RYZHo2qyrb16P+M4ddwDvvstpw7w89pO7\n/HK7RyUiUKAlUj2sWwc8/TTbezscwIgRzGD07Fn2Y9evZ8H9kSOevlebN5caaM3aMgsnT5/8fdVf\nZl4mFu5ciAHnV2CqUvznoYeA8HBgyRIWww8dCrTVAgWRQKBAS6Q6WLiQgZW51t7l4orF8gRaERHA\nL78U30vFLKgvQa/2vdC5cfFWE+fFnlfRUYu/BAVxG6R77rF7JCJyBgVaItVBeDj7bZmczrI3ryt6\n3+BgZsIMg5eLPpcXF7e8GBe3vLgKAxYREUCrDkXsUVgIjB8PXHEFe2HNnl36ljp33QWEhbFJaXIy\nMxgPPFC+YxUUAA0b8piGwaxYXp5PXoaIiJROGS0RO3z8MTBrFptKulzAW2/xcq9e3u9/1lm8/8KF\nDJh69y69izzATFZBAbNhqakskHa5uE+i2fRUREQspUBLxJcOHQK2bgWiooBLL2URujcJCbyP+f2w\nMG4fVFKgBbCdwxNPlG8cc+dyy56CAiAjA2jZkv+GhrLX0smTFXpZIiJSOQq0RHzlp5+46bPTySm6\nP/8ZmDCBQdSZYmKAxETP9YICoHFj34xjwwbuO9i4MQOrAwe4l+IFF7BOKy0NaNLEN8cSEZFSqUZL\nxFfi4xnQNG3Kr40bgRUrvN936FCuBkxJ4VeLFuyN5QuJiQz0wsIYWHXowH0Rf/gBWLOGDUxvvdU3\nxxIRkVIpoyXiK8ePM3gCPF3XMzK83zcujtN769dzFeBll/mubsrMVhkGx3H4MMfVsSNvLyhgwHXL\nLb45noiIlEgZLRFfufxy7kHocgE5OQygunQp+f4xMcBNN7FxqC+L03v1Arp3ZwH8sWPA6dMsnG/W\njF916rCvloiIWM7qjNZ0ADcBOArgfIuPJWKvESO4v9zKlSx0j4/nJtD+FhrK1hE//8xpwgULmMEC\nmOVyOplRq+2OHwd272Yn9bPPLnPvRxGRyrD6L8uVALIBzIL3QMswSusdJFIduVzFu7DbOQ6nE8jM\nBAYP5hSiywVccgkwdqz3Iv3aYvt24NFHGRib7TKefz4wfm4iErAc/EBWodjJ6ozWagBxFh9DJLAE\nwpv1vHkMpgoK2BR18mQW3YeEAJ06cVrTSsnJwLZtrA3r1q3kNhd2ef55Bp1NmvDfhQuBa6/llKuI\niA+pGF6kptm0CXj9dXaDDwsDVq8G3nkH+O9//XP87duBIUPYfd4w2FZi0qTAyaAZBrN7sbG8HhTE\nacNjx+wdl4jUSLYHWqNHj/79co8ePdCjRw/bxiJSI/zyC7M05l6IjRsD69b57/hjxjCYadqU//70\nE/Ddd5yeCwQOB9C1KxvLxsZy+tDh8KzKFBFxS0hIQEJCQpWewx/Vn3EAFkE1WiL+8fXXwAsvAM2b\nM4A4fpxThbGxDL4GDwb++lfrjt+rF49rBnpHjgDDhgH332/dMSsqNZVd9vfsYUbr2WeBfv2q/rxr\n1gDvvsuVnn37AvfeGxhTySLiE4FYoyVSexUWAosX8828Y0fghhusr40CGOgsWsQpxKAgICuLgU9o\nKMf09NPA+++zc70VLr+cx2/WjNmioCDgT3+y5liV1bQpN/LOyGAdmS+mNbduBZ56intLhoRw5Wdw\nMHDPPVV/bhGptqwOtD4B8FcAjQEcBPACgBkWH1PEfoYBjB7NQCsoiAHOhg28zeo2AmFhwMSJDLRy\nclgflZYGREby+7m5wLJl1gVazzwDnDrF/Rzr1mV27YILrDlWaQyD7RsyMtgdv1Gj4t93OIAGDXx3\nvIQETtnWr+85/qJFCrREajmrA60BFj+/iLUKC4HPP2dPqtatgfvuY4+sshw+DCxZwqxOUBDfgL/5\nBnj4YW7wbLWQEK72A4A5c7gK0FRYWL7XUFmRkdxrsbDQU2jub4YBvPoqe4gFB3Mac8IE4HwL2/lF\nRPC4poICT3ArIrWWigdESvP663zDTkgApk0DHnmE02FlyctjkGHW5zgc/CrPY31tyBAGekeOMABs\n1Mg39UhlCQ62rwnoxo0MkGNi2MKhsJAtHax0yy081uHDPNdOJ/e0FJFaTTVaIiXJyeGbddOmDBoM\nA9i1i/2h/vKX0h/bpg3Qti2wdy8QHc2moR07MivmbxdcAMyaxWAxLIxb/jRt6v9x+FNqKoM8M9Ct\nV8/TsNWq4vTYWODjj5m5PH2aCw46d7bmWCJSbSjQEimJOQ1kZmXMN26Xq+zHhoZy9dmbbwKJiZzG\nGz6cU3p2OOssftUWHTrw37w8ThumpXHfSatXAMbEBNbqShGxnd2be6m9gwS2kSNZaxUVxQxX69bM\nWkRE2D0yKcuCBcBrrzFgbtOGTVubN7d7VCJSjVWmvYMCLZHS5OcD06dzBV/btqzRatLE7lFJeZ0+\nzRWQDRuqn5WIVJkCLRERERGLVCbQ0kc8EREREYso0BIRERGxiAItsZ/LBZw4wQaPIiIiNYgCLbHX\n3r1snnn99UDPnsCKFXaPSERExGdUDC/2cbmAW29lc8kmTdg+IScHmD/fP9vUiIiIVICK4aV6ycpi\nt26zXYK5V9zevfaOS0RExEcUaIl9oqIYXJ06xetOJ/eki4mxd1wiIiI+okBL7BMcDIwZw6aSaWnA\n8ePAwIHaH05ERGoM1WiJ/VJTOV3YuHHt2o9PRESqFXWGFxEREbGIiuFFREREAogCLRERERGLKNAS\nERERsYgCLRERERGLKNASERERsYgCLRERERGLKNASERERsYgCLRERERGLKNASERERsYgCLRERERGL\nKNASERERsYgCLRERERGLKNASERERsYgCLRERERGLKNASERERsYgCLRERERGLKNASERERsYgCLRER\nERGLKNASERERsYgCLRERERGLKNASERERsYgCLRERERGLKNASERERsYgCLRERERGLKNASERERsYgC\nLRERERGLKNASERERsYgCLRERERGLKNASERERsYgCLRERERGLKNASERERsYgCLRERERGLKNASERER\nsYgCLRERERGLKNASERERsYgCLRERERGLKNASERERsYgCLRERERGLKNASERERsYjVgdb1AH4FsBvA\nsxYfS8ohISHB7iHUOjrn/qdz7n865/6nc149WBloBQOYCAZb5wIYAOAcC48n5aD/mP6nc+5/Ouf+\np3Pufzrn1YOVgdYlAH4DsA9AAYA5APpYeDwRERGRgGJloNUSwMEi1w+5bxMRERGpFRwWPvdt4LTh\nIPf1ewF0A/B4kfv8BqCDhWMQERER8ZU9ADpW5AEhFg0EAA4DaF3kemswq1VUhQYrIiIiIhQCRn5x\nAMIAbIaK4UVERER85gYAO8EpwhE2j0VERERERERERKRipgNIBbCtyG2NACwFsAvAdwAa2DCumszb\nOb8DwC8ACgH82Y5B1XDezvkbABIBbAHwOYD6NoyrJvN2zv8Lnu/NAJaheL2oVJ23c24aDsAF/n0X\n3/F2zkeDtc8/u7+u9/+warSSfs8fB/+mbwfwmr8HVZorAVyI4gN+HcC/3JefBfCqvwdVw3k752cD\n6ARgBRRoWcHbOe8FTyuVV6Hfc1/zds6ji1x+HMBUv46o5vN2zgEGtN8C2AsFWr7m7Zy/COApe4ZT\nK3g751eDCaJQ9/WYsp7En3sdrgZw8ozbbgHwofvyhwD6+nE8tYG3c/4rmEEUa3g750vBT/gAsA5A\nK7+OqObzds6zilyOApDmv+HUCt7OOQCMhefDs/hWSefcyjZNtZ23cz4EwCtgI3YAOFbWk9i9qXRT\nMC0H979NbRyLiD8MBLDY7kHUEvEADgB4AMoi+kMfcBprq90DqWUeB6fJp0HlN/5wFoCrAPwIIAHA\nRWU9wO5AqyjD/SVSU40CkA9gtt0DqSVGAWgDYCaAt+0dSo0XAWAkOJVlUqbFeu8BaAfgAgDJAN6y\ndzi1QgiAhgC6A3gGwKdlPcDuQCsVQDP35eYAjto4FhErPQjgRgD32DyO2mg2gIvtHkQN1wHsmbgF\nrM9qBWATgFgbx1QbHIUnSTEV3GNYrHUIXNQEABvAspDGpT3A7kBrIZjWh/vfL2wcS22kT5z+cT34\nyacPgNM2j6W2OKvI5T7giiyxzjaw9KOd++sQuNhGH56t1bzI5X7wvgpUfOsLAD3dlzuBDdmP2zec\n4j4BcAScOjkI4CFwVcr3UHsHq5x5zgeCCw4OAsgFkALgG9tGVzN5O+e7AeyHZwn2u7aNrmbyds7n\ng286mwF8BmVWfM0853nw/D0vKgladehr3n7PZ4E1cVvAAEB1zr7l7fc8FMBH4N+XTQB62DU4ERER\nERERERERERERERERERERERERERERERERERERERERkRpqHyrXw2gmgNsqcP84eG+Q2APAokocv7Ky\n/XgsAPgaQD0/H9MXugK4we5BiNQ2dneGFxHfq+yeodV1r1F/j/smAJl+OlaID5/rQnAbKBHxIwVa\nItXXAgAbAWwHMKiE+9wPdo3eDHaRBpiJWu6+/XsArYvc/yoAPwDYA092ywHgDTB7tRVA/zLGZYAZ\nn68A/ApufOsAO1kX3dx5EICxZzx2MIDXi1x/EMAE9+Wn3GPYBmCYl+P2QPFM2kR4tvjaB2AM2Jl/\nI7g9zHcAfnMf0/QMgPXguRldwuvbB2YM4wAkApgC/gyWAAj3cv+ZACaD+6LtBAM1AAgGz6t5vIeL\nvI7VAL50P28QgDfB170FwGPu+/0FQIL79XwLz76xCQBeBbDOfbwrwG7WLwG4030O7ijhtYmIiIhb\nQ/e/dcE3YfP6XjAQOA98ozWnEc0trhYBuM99+SEwYAMYEMx1Xz4H3DoIYMD1HRgsxYLbCTVF6VOH\nue7vB7kfexuASDCwCXbf7wf3GItqUuS4ALAYwGVgULHV/VojwQCkq/s+WUWOWzTQmgAGmgDPiRlQ\njXU/V6T7eCnu268F8L77cpD7ua708vrM8xsHoADAn9y3z4X3TcNnuF8HAHQEt/KoAwZWo9y31wED\nsTj368gG0Nb9vSEAPoXng3FDMHBaC89mtncCmOa+vAIM4ABOFS51X34AwDtexiciFvJlWlpE/GsY\nuHclwKzUWWB2BGBQ1BN8gz7hvi3d/W/3Io/7GJ4MkgHPxu6J8OybdgWA2e7vHwWwEsAlKH0D2/Vg\n5gfgfmFXgHsOLgdwM5jpCgXwyxmPSwP3yesGBmVngwHFMACfgwEc3JevAjM85bXQ/e82MMg65f7K\nA1AfDLSuhWcD6kgwMFpdynPuBYM2gPuexZVwv0/d//4Gvr6z3cc6H8Dt7u/Vcx/PCZ6//e7brwGz\ngi739ZMAuoBB6vfu24LBPdlMn7v//anImBzQRvIifqdAS6R66gG+AXcHcBrMYpw5bWWg5DfWkm7P\n93Ifb89TVl1U0e87ilyfCmZxEgFML+Gxc8DpyV/hCRjOHEPR5zQ5Ubwcou4Z389z/+tC8dfpgudv\n4SvgVGB55RW5XOjlmCUxx/4YPBknUw8wACzqzPPvAIPUy8oYVyH0d17EVqrREqme6oGZjdNgdqT7\nGd83wOzRHfBMHZpTi2sB3OW+fA+AVWUcazU4NRUEIAbMJK0v9RHMeMW5H9MfnqzQegCtANwNZrq8\nWQBm3AaAQZc5hr7wTB32xR8zTfsBnAsgDJwm7VnC83sLMg2wxmqg+/kBoCX4eqvKAf4cHAA6AGgP\nBpFLAAyFJxDqBCDCy+OXgtOe5pRrQ/fjY+D5uYeCr700mQCiK/UKRKTS9ElHpHr6FsAjAHaAdVj/\n5+U+OwDEg1N9heA00kAAj4N1Q8+AU4EPFXmM4eXyAgCXgtN0RpHHxcF7ZssA640mglNhy+GZkgQ4\njdYVQEYJry3dPfZzwEJvgNN5M+EJ8D6AZ9rQHMNB93NvB6f0firh+Q14f51L3cc0z2UWgHsBHPPy\neG+XvV03bzvgHns9MGjKB7N7ce5xOsBz2s/L+KaCQdhWsCZsCoB3wSnHd8BpzxBwocGOEo4PMOv5\nHHguxwCY5+W+IiIiUs0tAnC13YPwoxkAbrV7ECJiD00dioi/NACzbzlgdkVERERERERERERERERE\nRERERERERERERERERERERKSW+H/CqQ+gYvnblgAAAABJRU5ErkJggg==\n", + "text": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "If we want to pack 3 different features into one scatter plot at once, we can also do the same thing in 3D:" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "from mpl_toolkits.mplot3d import Axes3D\n", + "\n", + "fig = plt.figure(figsize=(8,8))\n", + "ax = fig.add_subplot(111, projection='3d')\n", + " \n", + "for label,marker,color in zip(\n", + " range(1,4),('x', 'o', '^'),('blue','red','green')):\n", + " \n", + " ax.scatter(X_wine[:,0][y_wine == label], \n", + " X_wine[:,1][y_wine == label], \n", + " X_wine[:,2][y_wine == label], \n", + " marker=marker, \n", + " color=color, \n", + " s=40, \n", + " alpha=0.7,\n", + " label='class {}'.format(label))\n", + "\n", + "ax.set_xlabel('alcohol by volume in percent')\n", + "ax.set_ylabel('malic acid in g/l')\n", + "ax.set_zlabel('ash content in g/l')\n", + "\n", + "plt.title('Wine dataset')\n", + " \n", + "plt.show()" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "display_data", + "png": "iVBORw0KGgoAAAANSUhEUgAAAcwAAAHMCAYAAABY25iGAAAABHNCSVQICAgIfAhkiAAAAAlwSFlz\nAAALEgAACxIB0t1+/AAAIABJREFUeJzsvXmUVOWd//++t/a9u4GmbZoGBAkIyiIKImvUqLjGNWZQ\nJzHKccwYRyff4zITz/klmWRmgsZ8Z8YlUb/GmBhHR8ERTVBBFkUkCCJEoJutm26g1+rabtVdnt8f\n5XO5VV1VXcutqltVz+ucPgdqufXcqnuf9/P5PJ8FYDAYDAaDwWAwGAwGg8FgMBgMBoPBYDAYDAaD\nwWAwGAwGg8FgMKoNLtOThBBSqoEwGAwGg2EEOI5LqY18qQfCYDAYDEYlwgSTwWAwGIwsYILJYDAY\nDEYWMMFkMBgMBiMLmGAyGAwGg5EFTDAZDAaDwcgCJpgMBoPBYGQBE0wGg8FgMLKACSaDwWAwGFnA\nBJPBYDAYjCxggslgMBgMRhYwwWQwGAwGIwuYYDIYDAaDkQVMMBkMBoPByAImmAwGg8FgZAETTAaD\nwWAwsoAJJoPBYDAYWcAEk8FgMBiMLGCCyWAwGAxGFjDBZDAYDAYjC5hgMhgMBoORBUwwGYws2bx5\nM6ZNm1aSz1q2bBmee+65knwWg8HIDiaYjJrlZz/7GVasWJHw2FlnnZXysVdffRWLFy/Gl19+WZKx\ncRwHjuOyeu3EiRPxwQcfFHlEpfscBsOoMMFk1CxLly7FRx99BEIIAKC7uxuSJGHXrl1QFEV9rL29\nHUuWLCnnUDPCcZx6DtXwOQyGUWGCyahZ5s2bB1EUsWvXLgBxl+vy5csxderUhMemTJmCpqYmbNy4\nEePHj1ffP3HiRKxevRqzZs1CXV0dvvWtbyEajarP/+///i9mz56N+vp6XHTRRdizZ0/asaxfvx7T\npk1DXV0d/v7v/x6EEFWc2tvb8fWvfx2jR4/GmDFjsHLlSvj9fgDAbbfdhmPHjuHqq6+Gx+PBL37x\nCwDATTfdhDPOOAN1dXVYunQp9u3bp37WunXrMGPGDHi9XrS0tGD16tUjjjnd5zAYjK8gDEaVs3z5\ncvLEE08QQgi59957yfPPP08effTRhMfuvPNOQgghGzZsIC0tLep7J06cSObPn0+6u7tJf38/mT59\nOnn66acJIYTs3LmTNDY2ku3btxNFUciLL75IJk6cSKLR6LAx9PT0EI/HQ15//XUiSRJ54okniNls\nJs899xwhhJC2tjby3nvvkVgsRnp6esiSJUvI/fffnzCO999/P+GYL7zwAgkGgyQWi5H777+fzJ49\nW32uqamJbNmyhRBCyODgINm5c2fGMcdisbSfw2BUI+k0kVmYjJpm6dKl2LRpEwBgy5YtWLJkCRYv\nXqw+tnnzZixdujTt+++77z40NTWhvr4eV199tWqZPvvss1i1ahXOP/98cByH22+/HTabDdu2bRt2\njHXr1mHmzJm4/vrrYTKZcP/996OpqUl9fvLkybj44othsVgwevRo/MM//AM+/PDDjOf1t3/7t3C5\nXLBYLHjsscewe/duBAIBAIDVasXevXsxNDQEn8+HOXPm5DxmBqMWYYLJqGmWLFmCLVu2YGBgAD09\nPZg8eTIuvPBCfPTRRxgYGMDevXsz7l9qhc3hcCAYDAIAjh49itWrV6O+vl796+zsRHd397BjdHV1\noaWlJeExrev35MmT+Na3voWWlhb4fD7cdttt6OvrSzsmRVHw0EMPYcqUKfD5fJg0aRI4jkNvby8A\n4PXXX8e6deswceJELFu2TBXEdGPu6urK4ptkMKofJpiMmoQQglgshunTp8Pv9+OZZ57BRRddBADw\ner1obm7Gs88+i+bmZkyYMCHr49LI1tbWVjz66KMYGBhQ/4LBIG655ZZh72lubkZHR0fC2LT/f+SR\nR2AymfDFF1/A7/fjpZdeUoOStJ9Jefnll7F27Vq8//778Pv9OHz4cMKe6Lx58/Dmm2+ip6cH1113\nHW6++easxpxt1C6DUa0wwWTUHIqiIBqNQhRF2Gw2zJ49G0888QTOO+88DAwMQBAELFy4EI8//nhG\nd2wqqCjdddddePrpp7F9+3YQQhAKhfD222+rFqiWK6+8Env37sUbb7wBSZLwq1/9CidOnFCfDwaD\ncLlc8Hq9OH78OP793/894f1jx45Fe3t7wuttNhsaGhoQCoXwyCOPqM+JooiXX34Zfr8fJpMJHo8H\nJpMpqzEnfw6DUWswwWTUDIQQSJKEgYEBEELA8zx4nsfixYvR29uLBQsWQBRFCIKAefPmobe3Fxdc\ncAEEQYAkSSCEZLSytLmT5513Hn7961/j+9//PhoaGnDWWWfht7/9bcr3jRo1Cv/93/+Nhx56CKNH\nj0ZbWxsWLVqkPv/YY49h586d8Pl8uPrqq3HDDTckjOPhhx/GT37yE9TX1+Pxxx/H7bffjgkTJmDc\nuHGYOXMmLrzwwoTX/+53v8OkSZPg8/nw7LPP4uWXX85qzMmfw2DUGhl9LJmihRiMSoK6YBVFweDg\nIOrr69XHeD6+bpQkCaIowuFwqO/RujIBwGQywWKxwGw2w2QyMTclg1GFcGlubHOpB8JglBpFURCL\nxYZZiFQgzWazKppakqvtEEKgKAoEQVAfYwLKYNQOTDAZVQt1wUqSBI7jEkQxEokgGo2C53kIgqC6\naAkhkGUZPM8PE790AhqJRNTHmYAyGNULc8kyqhJFUSCKIhRFSRA6WZbh9/thNpvhdDohSRJ4nldf\nL0mSegye5xOEbyTxo+5b+pkAE1AGoxJhLllGTUAtRFEUASRahbFYDKFQCADgcrkSxIvneZhMJiiK\nAofDAUVRIMsyZFlGLBYDEBc/+pdKQOlj1JJNZYGazWb1jwkog1FZMMFkVA3JLlgqRoQQhMNhiKII\nj8eDQCAwolDRCFqLxQIAugkoFXMmoAxG5cEEk1EVpHPBSpKEUCgEk8kEr9ebMrhHS7pdCK2AUtdr\nJgHNFEQ0koBaLJYEIWYwGMaACSajokkWHa0YRaNRRCIROJ1OWK3WlOKjfSxbcdIKXzEENBwOq++j\nAkojeZmAMhjlgwkmo2LR5lZqrUpFURAOhyHLMrxer1rJJvm9eomP3gIKnLZoZVlWiybQfVYmoAxG\neWCCyahIknMrtS7YYDAIi8UCr9c7olVZDFIJKN0DlSQJ0WgUHMflJKDAaQuURvJyHJewB8oElMEo\nLkwwGRVFutxKQggEQYAgCHC5XLBarWUe6Wm04gggo4Cazea0+6ipBJQWX6DPMwFlMIoHE0xGxZDJ\nBUsLhKdzwRqJTAJKA5ei0ShkWc4Y/MMElMEoLUwwGRWBJEkIh8OIxWJwu93DcittNhscDkdFCkKy\ngIbDYZjN8VuTFoOn+5e5CqgoikxAGQydYILJMDRaFyyF4zgQQhCJRFQBpfmSuRx3pBSTckFdzVQ0\nky3QXARUa22nE1CaxsIElMHIDBNMhmHJVN4uGAyC5/msciuTqTRRGMmFywSUwSgNTDAZhiO5vB0V\nRI7jIMsyhoaG4HA4YLPZdJ/QK6F8cqkElBACi8UCq9XKBJTBABNMhsGgk7Ysy8PK20UiESiKAq/X\nq7or9aRSxSCVgGpzQBVFyUtABUFIKArBcZyaA2o2m7MqSM9gVBNMMBmGYaTcSjrpF0MsqwltcA+Q\nv4ACGLaXGovFEI1G1eeYgDJqCTbzMMpOptxKbXk7nucRiUR0/+xqn+SzEdDkIgrZunCpgGotULoH\nygSUUW0wwWSUlUy5laFQSHXBmkwmtURcodAoW/rvWiOTgEajUVVA6WIi3aJCK6D0+4zFYmopQGqd\nMgFlVAtMMBllg1qVyS5YURQRCoVgsVgSci4ZxSGdgEajUbUQwkgWKP1/LgJq1LQeBiMdTDAZJacS\ny9vVElRAJUlS94xTWaBMQBm1BhNMRklJl1upLW/n8/nY5GkgsnXhMgFlVDtMMBklIV1uJXC6vJ3d\nbofdbk/rgtXuPTLKR7KA0hxQ+vsSQvISULqfrRVQGoXLBJRhBJhgMopOunQRQgjC4TBEUcyrvF2+\nZBJeJsinyTaCmPbupL9fJgFNl36Sqg4ujZIWBAGiKMJutzMBZZQVJpiMokI7bwwODqK+vl638nbF\ngAUX6UMmAaVpQck5oJkElO5tUwFNlQfKBJRRCphgMopCcmCP9vFYLIZwOJxzeTvmkq1MMgkodb+O\nJKD0OBStBUoF1GQyqfufTEAZxYAJJkN30uVWUhesJEnweDysYk+NohVQKnyZBBQYbv2ncuEqigJB\nENT/awOIqCuYwSgENmMxdEW7b5U8qQ0NDcFiscDn87HJiwHgtPCNJKC0xnA6yzGTgNJrkVqg1IXL\nrkFGrjDBZOjCSLmVANQo2HLDXLvGJZWAyrKsFoJPZYHmKqAUJqCMXGGCySiYTLmVtLwdgIILETCh\nqz2oeHIcB7vdntDKTJIktY4tE1BGKWCCycib5NzK5PJ2wWAQNpsNbrcbg4OD5Rwqo0rI1AuUCSij\n2DDBZOTFSH0ro9FoQm4lsw5zg31X2ZGNgGbbCzSVgEYikYQiC0xAaxsmmIycSVeIQJZlhEIhcBxX\n0eXtyt3yi03EieTye2QSUFEUIQhCTgKq3YunAhqLxWC1WtUoXCagtQMTTEbWaF2w2skEyL68XSFo\nrdhCjp/O2mUTXvWht4ByHKc24GYWaO3BBJORFSPlVoqimDG3krlkGUZALwGli8VUFih9Pa23ywS0\nemCCyRiRdC5YSZIQCoVgMpkMU96OwciFXAU03TWeyoWr9cYATECrASaYjLRkyq2MRqOIRCJwOp2w\nWq0Ve/NX6rhLQbn3cstBKgHVFlGgKVLRaDTnPdBUAqot5Vdr33UlwgSTkRJaND0QCMDr9SbkVobD\nYciyDK/Xq04sI6GXS5Yeh00ujFKQ3MpMW0CeCmi+QUQ0kpeiDSJK1RKNUX6YYDIS0K6EFUVR9yyB\nuAs2GAzCYrEkiGglwvZTKwcjLZCo8NlsNgCpLdCReoFqj0NJFtDkUn5MQI0BE0yGSrIL1mQyqbU9\nBUGAIAhwuVwFV+wpN2ziYehFsgWqFdBoNKqrgGr3QJmAlgcmmAwAqQN7qBUWCAQAICcXbDJ6u2T1\nwkjWC6PyKaaASpKUUFWLCWjpYYJZ42TKraQ3p9lshsPhYDcko+bJdYFVbAEVBAGKoqiFFJiAFhcm\nmDVMptxKWtEEQNWKpSiKiEQiatAG7ZnIrE5GsSiGgNJ/i6LILNAiwwSzRslU3i4YDILneXi9Xt2K\nphupcIF2dW61WtX/R6NRAPHoR5YrxygFyQJKA+2o+5UQkrWAardLaK3nZAGlaSxMQPODCWaNkS63\nEojnloXDYTgcDthstoS9TKPcXIUKL21FRgiB1+tVgyqocNJauMlJ62yVXj6MdP0VG57nwfN8goBS\nCzSVgKa7F5iAFgcmmDVEJhdsKBSCJEkZy9tVOqIoIhQKJeTNybKsTjr0O7FYLOB5Pm93GaN6KbV4\nUwGlXX9SCSjP8xBFUfWI5GuBauvgsms7NdU5MzKGkRxhl5xbaTab4fP5ht0kRo1uzYXktBitSGYi\n036TIAjqal/rvmWTDKOYJAsovQ5pHiiAYUUUshXQWCymbkvQz6DXP7u24zDBrHKSXbBaq1Jb3o4m\nYlcbiqIgFApBURQ1LYZWaskVrYDabLaE1X6qyYrV1mUUG7qtQnOjR7omCxVQ6sKtVQFlglnF0P26\nZBdsKhGpFHKxVLWVidxu97Cw/ELRrvZpgQdt42IqsKxWKKNYUJcsJd01meuiTiug9F6hAqrduqg1\nAWWCWYVocysBJNwUdB8vlYikohJdslrrOVVlomLc2NpaoXSyyqdtlJEwSlSzkaikAKRU12Q+AkrP\nN5WA0mPQQKVqF1AmmFUG7U9JCFH3Hujj1VTeLh00gCnX4vDJxygUukJPbhslSVJC0W6jB1kYYUyV\nJFKlJpfvJp2A0i0baj0WIqCCIECWZdhstmFRuNVAdZwFA8DpDiPRaFTds6SPBwIBiKIIn8+Xk1ga\nKX9yJCRJgt/vB8dxeYtlsSZmOhHZbDY4nU510UKt4VAopBaLoBMZg1FMtPufDocDLpcLdrsdPM9D\nkiSEw2GEQiEIgqBu7aQ7TjoXbigUwj333INDhw6V7LyKCbMwq4B0gT3A6YvWbrfDbreXbaVeTNdu\nJQYwpYvAlSRJdZlp01iqZYVe6VSztZvOK5LrvnyygPb09FTEPZkNTDArnFS5lRzHqYE9oijC7Xar\nYej5foZR0cMFawS0Akr3OzmOYxG4jIwUU8AzCWi6fflUc0U4HIbT6SzKGEsNu+sqGG3x5eQoWCqi\nXq+3ILHU82bUW3hlWYbf7weQWyeVSnAz0yhEu90Op9MJh8OR4CoLh8Oq693o58KoDqiAal24tCIY\nDSakrltt3jcNvssWQRAwf/58zJ49G2effTYefvjhYa/ZuHEjfD4f5syZgzlz5uAnP/mJbueZCWZh\nViCZcitp6LfZbM4qCrZU6C28tIxfpbhgC4H+xnTvuRoicHOBLQiMSSoLlFbEEkURt9xyC/r7++Fw\nOPDuu+9i6dKl8Pl8Ix7Xbrdjw4YNcDqdkCQJixYtwpYtW7Bo0aKE1y1duhRr164tyrmlg1mYFQa1\nHlOJJQ0csdvtuk2aRrTG6H6lx+OperFMRaqVPhVTumdNA4iyrWpkdIyyADDaHqaRxkPnI9oO8NVX\nX8XPf/5zxGIx/PKXv8S4ceNwwQUX4MknnxzxWNSFS6/hhoaGYa8px3XNLMwKIVNupTZB3+fzIRaL\nqa+rJmjwAc/zKcv45YMRFwS5kqmEX3INXFbmjFEq7HY7LrroIjgcDrz//vuIRqP45JNPEAwGR3yv\noiiYO3cu2tvbcc899+Dss89OeJ7jOHz00UeYNWsWxo0bh1/84hfDXlMMmGBWALRQsizLw6xKmltZ\nLNekUQoXUBcsz/PqvgkjNckCqi2XRssCsgCi6sBIFiYwvPIQheM42O12LF26NKvj8DyPXbt2we/3\n47LLLsPGjRuxbNky9fm5c+eio6MDTqcT77zzDq677jocOHBAr9NIP66ifwKjIGhuZbJYKoqCYDCI\nWCwGr9ebIJbVYDVRtK5mvTqpjPT9VMt3R6GJ6toAItqpRZtrxwKIGMWgEEH3+Xy48sorsWPHjoTH\nPR6P6ra94oorIIoi+vv7CxpnNjDBNCjUqtQWP6YXniiK8Pv9MJlMFZNKkY+Iy7KMoaEhNdq3FG3H\njLRaLwbaSi92uz0hWZ1GOtIIXFZAYThGs+iMRqrvJ9drqLe3V21cH4lEsH79esyZMyfhNSdPnlSP\nu337dhBCUu5z6g1zyRqQTH0rI5EIotFoxtxKPS3MclmrNHhF28yaoT+Zcu2A+IRV7hJ+6dx8tU4l\nLGZEUcw5ra27uxt33HEHFEWBoii47bbbcPHFF+OZZ54BAKxatQqvvfYannrqKZjNZjidTrzyyivF\nGP4wMl75pBJ+kSpD2xhWK5ayLCMUCoHjOLhcrowTiCiKiEQi8Hq9BY8nHA6D4zg4HI6CjkPTXTwe\nT8bX0UVBLBaD2+0eZlWGQiGYTCbY7XZdxqLt5gLEBYL2ASwXgiDAZDIVlD+rB8FgEA6HIyGIqBxN\ntI3yfRhtLHS7wu12l3soKuFwGDabTV2ADQwM4J577sHbb79d5pHlBpfmomYWpkFIzq3UCmKu5e2M\namGOdBztosDr9aYNHmDruNJBBTFVCT+6sEsulVbN3gDmks2NcDhc8GLbSDDBNAC0jJ0sywmCSDuP\niKKoW8BLuRhpkil1zVsmuvmRKQKXlfArLUYU7+Qx0frO1ULlzsBVgDa3kk469GKTJEl1P6azttKh\ntxVWTHHJdl9WT4w2yVQyqRoWJ7eLYk20a5dwOJxTWTyjwwSzTCTnVvI8r1Zl0XbesFqtZZ1k9Prs\nVCJOU2OAePg4s0Yqm1ov4ccYbmFWU+F1gAlmWaDl7ZIDe/TqvFEJ+3yiKCIYDObsgq2Ec2PESReB\nq22inU0AkZFcj2wsucEEk5E3mQJ7aMFim80Gr9drmBuhGO5dWp2oVC5YhjFIJaDpSviVK4WFkT/V\n3toLYIJZMjLlVlIBMZlMuvj7jWiF0R6dgUAAQPldsOm+HyN+d9VKpgAiWgvZZDKp+XiMRIxqYSYH\n/bA9TEZOpMut1O7hOZ1OtaqPkdBLQGjZNdrJoJAbvdDxaD/biBNOraINIAKGR+CKojisiDzD2IRC\nIWZhMrIjm9xKm80Gh8NRlDZMRliBai1ojuMKvnnKfT6M0kEFVJIkmM1mNTCORuCWI4DICPeUUUn1\n3UQiEYwePbpMI9IfJphFgu5JpitvRyvZFGMPzyg3NM0vJYTA4/Go7lgGI1cylfBLjsCl4mqU+6BY\nVIJ4szxMRkaS+1Yml7cLBoPgeX5YbmUx9s70uKHyHRft0UmbHLN9QUa+pLqOWQCR8UhnYbI9TEZK\nkl2w2ouH9nMsVTHxck0O2jxSl8uVkJOnBywoh5GKTE20BUEAIUS1PmuhhJ9RYHuYjJSMlFspSVLG\n8nbFsjALJZdxURcsbcelzSM1ktAlj6USXFulxii/Vb5oBdRms+lWws9I14qRxgKktzCZYDJUtC7Y\n5MAe6pY0m83w+XwlvbhLfSPRc7VYLHC73Ya6kRn5Uy2/Y6oSftoAIlbCrzgwlyxDJVNupba8nc1m\nG/FYRrUws/mMXM+VwSgn9F7VCmgllvAzmhcglYXJaskyAMTdj0NDQ6rbR5tbmc4tmQ16uVmKWQOW\nkkspP+1iopCxFdu1ayTXMaM0ZFPCjwoofd4oAmqUcaSDVfqpcbSBPZIkJaw+RVFEKBTKyy1ZjAu/\nmBO/1gVrpFJ+jOqj1AKVLgJXkiQAcRFgEbjDYVGyjARSuWDpfghNztdGhuYKPZ7RLUwa8VupLlhm\nRTJyQbu/KUkSXC5XQhNtoDw9QAkhhu/wI8ty3k0kjAgTzCzIlFtppPqoyRRjTzSbiN906LkgYDDK\nRaYauLXcRDvdvV1N9zsTzBHIlFtJU0kcDkdOLarSoaflo7eFSYsumEymkkf8phqPnsdjMLIhnSCM\n1ES7EgKIikE1Lo6ZYGYgU3m7cDgMSZLUSjZGRE9hGRoaKlnRhVJBA7RkWVZdbtV4kzNKB50nStFE\n22jXajoXsZHGWChMMFOQKbdSW97OZrPp6m4xmoVJXbAAdKl7a6S9Q0II/H4/rFar2kJKuz8NgAV0\nMAomUwm/XJpoVypGud/1gglmEulyK4Hh5e1oyS290FtQCjmW1gULIOf9ymJSyHnRvFEAcLlcMJlM\nEEVRXQwIgqC+jv67XAEdjDhGs6QKIVMJv+QauDRdLd25G+0aSf6djDY+PTDOLGgA8ilvZ9SLopAJ\nhrYeowsDGshgBAo5L23eKABYrVb139rjU5dackm1Uu1HVYs4VBvFEO5MAUSRSARA5gWbka8VURQr\nMoo+E0wwkblvpTbfMDnYpRg3TzktTLo3K4piwsJAr+jWcrpktRazx+PB4OBgytcljzE5oCPVfhTd\n/6w2dxqj9IzURBuAan0adbFOqbbC6wATzIzl7WhuZbp8QyPtySWT68SdqfVYpZNsMedLpv0orTtN\nK6AMRiFkWrDROcooNXCTF9V0+6qaqGnB1CYeJ+dWZlverhr2MKmg2O12XdJjikku35G2WbfWYtYz\nsCqbfDxtS6lKoZr2DfWi3Ivj5AUb7TerXfQbqYl2tVX5AWpUMDPlVoqiiGAwCJvNNmJ5OyOXs8tG\nWLSCkikKVi8hL6VFrigKgsEgAKRs1g3oPwFm4741ymRWKZRbpJIx0m9GBZTneVU4y9lEO5WFyVyy\nFU6m3MpIJIJoNJp1CoXRk+gzjY0KCsdxVeeCpfvONEe2HJNctu7bbKIhGcYSKqOQLFCZInDL0USb\n7WFWMIQQiKIIURRVS4AiyzJCoRA4jsu5vJ1RXbKZbgRqRWfrgjXyXm0ygiCorqB8a/oWg1STmSRJ\nKaMhmTuUoQfaa07PJtrpSL5uq615NFAjgknFMhQKgRCS4FcvZP+u0ia1fKxoPdFDeNMdQ1t9KZ+2\naqWG4zhYLJaU5dRkWVbP0wjBHIw4RlrI5HMflbqJdrX1wgRqQDBpojot20Tz7tKlUORCMVyyxdrD\n1O7p5VMk3sgWpjZlJNdWY0aYAJPLqdGCCRzH1UQ1GEb+5Hsd0GtOrybadH5ge5gVDr0w6B9dyYdC\nIXWCrab9u1RoA5ny2dMz8uScnDKSy1iNugjQCmhyMEc5qw8xqpdUe+7JTbRzXbQxC7NC0Vpbsiwj\nEAjA6XTCarUWJAbFsDAVRdH1WJFIBIIglMUFm2pMen1f2jzZfM4t3e+u52+gF5n2oorhSjMSRnKD\nGolify+5BK3R1ySPRxAEjB07tmhjLAc1sTSlk2A0GoUsy/B6vbp03ShGMIyegkLdKz6fryCxNFLQ\nDx1LMBjU5dwqEepGs9vtcLlc6t473acPh8OIxWKQZdkwv1s1UMviTRdlNpsNTqcTLpcLZrMZiqKo\ngXbawEog98IFgiBg/vz5mD17Ns4++2w8/PDDKV9333334ayzzsKsWbPw2Wef6XJ+2VITFqYoiggE\nAsNWRHqh142k180oiiLC4TAAwOPxVNVNLkkSgLhoOJ3Oqjq3fGDVhxjlQBu0BsTnHLpIe/rpp/Gb\n3/wG06ZNgyRJWLBgARobG0c8pt1ux4YNG+B0OiFJEhYtWoQtW7Zg0aJF6mvWrVuHtrY2HDx4EJ98\n8gnuuecebNu2rWjnmUxN3D20vJ3eZZqMVkuWRsHSlBG93HNGKVwQjUbVwCWXy1XzYpmKZEvA6XTC\nbDar6SuhUAiCIECSJGZ9VjBGs3bpnqbdbsd9992HV155BR6PBx9++CGmTp2Kc889Fw888MCwZgfJ\n0CAhKr4NDQ0Jz69duxZ33HEHAGD+/PkYHBzEyZMni3NSKagJwfR4POp+pd6ThFHclTQKNhaLwev1\nqpFv1QDtMhKJROB2u9XH9DhutaN13zqdTtjtdvA8n+C+pVsVtfB9MIqDVsB5nsfMmTPh8/nwn//5\nn+jt7cU9iSOEAAAgAElEQVSvf/1rTJ06dUTvnqIomD17NsaOHYvly5fj7LPPTnj++PHjGD9+vPr/\nlpYWdHZ26n9CaagJlyz9IY0ibunId3zajiq0nB91XVY6xapIZKTVeanIJZCD53lD3yvlwGhWndGh\naSVmsxnz58/H/PnzR3wPz/PYtWsX/H4/LrvsMmzcuBHLli1LeE3ydVnK36QmLMxiUk4RppGiNOpX\n66YsZk5nqY4jiiL8fr+6EGD7b/qSKpDDYrGoAXJAfDuDlpIsB0ykUlMJ30shxdd9Ph+uvPJK7Nix\nI+HxcePGoaOjQ/1/Z2cnxo0bV9A4c6GmZiCju2RzORZ1U0ajUXi9XkOVgSsUuhAIBoNwu91lqwdb\na1ABtdvtahS5yWSCJEkIh8Oq+5btfzKSSSXgueZh9vb2qn1qI5EI1q9fjzlz5iS85pprrsFvf/tb\nAMC2bdtQV1dX0tSVmnDJUrSF1o04AWcrmFoXbLrKNka0MIGv3ClHjoD/85/BdXaCzJgB5dJLga82\n9+lCgKb/pNrzoOPR6zc04rVQbmjxBG35Ppr/WavVh2i1MEZ25FpLtru7G3fccQcURYGiKLjttttw\n8cUX45lnngEArFq1CitWrMC6deswZcoUuFwuvPDCC8UafkpqQjCTK/rrPdmWarVNy/zRC7GQZsjl\ngOM48Nu3w/z44/EHHA5wu3aBX7sW0r/9G+SmJgQCgYwLAT1JdR0YfZ+7XGj3P1n1ofJjtEV/qvHQ\n/pzZcs4552Dnzp3DHl+1alXC///jP/4jv0HqALuqC6RUllyyC3YksdR74tflWKII+3/9F+DzAc3N\nQH09MG4cEA4Dzz2HoaEhNRm/2JOBkSabSkS7/+lyueBwOGAymSDLMsLhsHqtMvdt7UK9FNVEzQlm\nJVoQkiTB7/cDQM6dOPQK1tED/sgRIBQCNPsaBIBUXw98/DE8X6U9MCoPVn2otBjdwqzW37hmXbJ6\nHruYFmY0GlXDs3NxwRrpZqJwFgugOT9CCERJAhQFVpsNyLLEXSUuemoJPasPGUkYjDSWSqAav6+a\nEEwtRhZMCj1eoe3H9EKvguTKhAlQGhthGhiAUlcHURRh4nmY+/qgfOMbQJn2varxxjYS2uLxANI2\nMqYCyn6LyqNWAqKq/wwrCDpRyLKMoaEhEELg8/nyFkvDWWI8j/APfgBZUaAcOQLLqVMwnzgBTJgA\n5bbbyjYsNkGXFlZ9qHCMvsirVgFlFqbBjgcAgUAgr/6OxULPcxQnToT/8cfh+eILoKcH8uTJIPPm\nASXOI9U7WpqRH9m4b4F4bdFaSV+pRJLvpWg0WlW54ZSaEMxK2MMkhKgdRlwuly4Xm5EsTG36gWfc\nOHAtLcjXyWuk82LoS7L7lraLUhRFbRtVrvQVtsDKHhp3UW3UhGBqMeJkK8sygsGgunrWu/1YuYnF\nYgiFQmrZNTbpMLKFpibY7XYQQlQLVJIkRKNR8DyfIKC1cm0ZzeWZvJgwomDS4hv0e8vnWqk5wdSb\nQgNiqJhQFyxNH9FrbOVsy0VL3AmCALfbrRZeMDJGXFAVg6gUhc1s/MIX2omYiifdA2XVh4xLrlV+\nSsGHH36ILVu2YPTo0bDZbHA4HLDb7XA4HCCE4JxzzhmxLm3NCaZRXLK0d2UsFkuIgq2WCVtRFIRC\nITVwied5NSLSaNSaq21QGMTuk7txYcuFsJpSu/4r4RocqfoQIUSNvNXDfWuk78Ro12zyeEKhkOEE\nk/bO9Pv96OvrU3vDAvEi7o899hjGjRuX0QCqCcEs5h5mPmhdsHq2rEqmXBamttat0+nUPaFZj/PS\nHsNIE08paBtoQ3+0H51DnTiz/sy0r6u070W7/2mz2RLSV6LRqPp8Ie7bSvtOyoURXbITJ07EDTfc\ngGnTpqGpqSnt6zLNxzUhmMmU08KkLli73a5WQynkeEYjU6EFNtmUn0FhED3hHjS7mtE22IYWb0ta\nKzMVoizi81OfY27TXMP/njzPp3TfiqIIQRDA83xC8QSjn4+RSbYwC2ntVSy++OILvPbaaxg9ejTO\nPvtszJs3D62traivr4fH48nKcKkZwSxmGkE2Aqd1wbrdbljSVLUxdJeREZ43SqEFRnraBtpgN9th\n4k0gICNamcns69uHNw68gVHOUZjom1i8gepMrtWHjF4H1Wgu2WSMaGGuXLkSK1euxIkTJ/DWW29h\n9erViEQimDdvHi6//HLMmTMHHo8n4zGME2ZVIoqxhzkSiqIgEAhAkiR4vd60YmlURjpHRVEwNDQE\nRVHg9XrTimUpBbyUpBuKgYYI4LR16bHGJ4U6ax3aBtsQk7PbWxZlER8e+xAemwcfHv3QUL9BrmiL\nxzudTjidTpjNZsiyjEgkgnA4DEEQEorHG12kykWq68CIgkkIgSiKaGpqwl133YU1a9bgz3/+My65\n5BL88Y9/xHnnnYddu3ZlPEbNmQGlDvoRRRHBYDCtC7aY4yuFezfX89MDI01asRjw4x/bsGpVDC0t\np7/r9etN6OricccdYhlHlwi1Lim5Wpn7+vYhEAug1duKo0NHcXToaNGtzFKJVDbuWyAef1BL6Su5\nkJxW4vV6yzia1FgsFnz88ceQZRl2ux1utxvnnHMOFi1apC6iMlEzFmYxK+mnOh51wQaDQbjdbjgc\njoq9yVIJb6Wfn16LCasV+MY3JPz0pzZ0dsbPf/16E9asMePyy6X8xzc4CH7bNpjeew/8gQOAlP+x\nAECQBARiAUTlKHojveofIQQnQidGfD+1Lsc4xgAAfFZfxVuZ6aDuW6vVCofDAZfLpe7H0xgEur1S\nrvJ9Rrd2BUEw3B4mjX594IEHsHTpUnzzm9/E8uXLMX78eDQ1NWHZsmX4+OOPMx6jJi3MYh9PURQE\ng0EAUFMqcjme0S1MQgiCwaDqgs220EKlBzSlY/FiGQDw05/asGCBjE8/5fHP/xzD2LH5nSvf3g77\nmjXgTSbAYgG3fTuUlhZIN98M5Nn+zG62Y1nrMhAMHxOHke8Jal3We+sBAHX2upJZmeVGW0zE4XAA\ngFo8odzVh4xAKvGmueVGgv4uV111FR555BFcffXVAIANGzbgnXfewaJFi/Dwww/jlVdeSX+MkozU\nQBTbJSuKIvx+P8xmc9aRV8kYTVS05yjLMvx+v5oSU46qRMN+Q1kGIpG8Ng1jsRgikYha7BvI7/tf\nvFhGY6OCd94x4bvfFfMWS8RisL79NpT6epBx40AaG6FMmADu+HHwKbrR5wLHceA5ftjfSItIURax\n8ehGAMCp8Cn1T5KlqrUyk9GmIFHXHe39SZtnS5KEcDisFo+v5ebZRoySpTz//PM477zz1P8vX74c\n69atwxVXXIFYLJbRLVuTFmYxXLLaqjaF1ILV0wLW+1ypOyrX3pxFQ5LAr1sH7s9/BhcOgzQ3Q7nx\nRpDZs0d8KyFEdanRkn3aYt+iKOZkLaxfb0JfH4cbb5Tw619b8eij0YQ9zWzhurrAxWIgSatz0tgI\n065dUBYuzPmYhaIQBbMaZ0FUhu/HavdEa5V0+5/FrD5kJJdsqrEYUTDpGL/xjW/gv/7rv3DNNdeg\nrq4OW7ZsQX19vbroydTEvmYEs1h7mPS4wWAQhJCCrS4jui3pgiAcDheUMqL3ufGvvAJ+/XqQ5maQ\nUaOAoSGYfvlLyP/4jyAzZ6Z9HyEEkiSB4zh4PB713wDUKMlckt3pniV1w44dS/DTn9ryFk2jYTPb\nsGzCsrJ8ttHuhZEodfUho2LESj+UX/7yl/jhD3+I22+/HYIg4Pzzz8fLL7+MWCyGJ554Am63O+17\na0YwKXqvyqgbj+f5YVVtyo0eAkVL3AEoalWinBkYAL9hA0hrK0AXKD4fiKKAe/PNtIIpyzICgQA4\njoPNZht2PsnFvtNZC2azGRzHQRQ5bNtmStizpHuaH35owt/8TW7BOqS5GbBY4i5mTU4Y19MDefHi\nnI5VLRjpnsqVTNWHaKnIXKsPGW0Rkc7CNKpg2mw2/OpXv0r53KJFizK+tyYFU692XNFoFJFIBAB0\nE8tCi7nrCW3oa7VaIUmSYcSS4zhwp06B8PxpsaTU1YE/cgQKIUDS76EtdC/LclYpPqmsBUmS1N/d\nZDLh4Yelr7wKp49HRTNnrFbErroK9jVrwA0NAVYruEgEyrhxUDT7LozSopcLVM/qQ0ZeSEQikYyW\nWrmh0c3agv7ZUHOCSSnkBqBWF40SHRoaMtSeAiVf8dUuBuh+LHUnFXKOuhYu8PnA0ZB+7ZiCQZAx\nYxIeS+6aYrFYVKs5F7TWAnVT00hJPUutKZMnQ/jud2E7fBhcIADS2grlzDPjlmeJMJoVU41US/Wh\ndFGyRrUwAeS9bVYzgqltEVQI2sLibrdbvYiNngqSLYQQhEIhyLJctijYbCBjx0KZPRv8rl0gLS0A\nz8erCPT2Qlm16vTrNCkwuab4ZIL+7jS4K9VkV8heFamvhzJ2rC5jzRcjTs7VjHZBBiCl+7ZS9j0F\nQTBcWomWoaEhiKIIi8UCs9kMi8WSVQW2mhFMLfnUldVaXcWOEi2X+NIuKiaTCV6v1/ATpnLnncCL\nL4L/y1/i7lmzGcqtt4JceCGA0p5PpsmulhsdVwvl8CAlu2+pR0P6qohFOBw2xDWV7rsxorCLooit\nW7fivffeQywWA8dxkCQJ48aNwwMPPDDi71zTgpktI1ldeluY5SC5kXW6LiqFumSBwicf9ft2uaD8\n3d9BGRgAAgGgsVFN7Kcl+9KdDx1HsUi1VyVJUtrgoeTxEULQH+lHg6OhaGOsBIy41VEOtB4Ns9mM\ncDgMm82W8poySveVcn++FnodHThwAHfddRe+853vYOrUqRBFEeFwGGPGjMnqODUjmNofLxeB07pg\n01kpRnXJZnOsbLuoGJr6+vgfEj0BI3WFSfe43kKaaq9KkiQ1hQVAwl4VEC8QsKNnBy6deCncVuMG\nT5QCI028RoAQkuCxoI+Vq/pQ8qKGWsNGJBQKYcGCBXjkkUdSPj/StVYzgpkPmXo7JmPUCyQTNHiJ\n5o9muqnKvbeaDZWy/8pxnLpnkipSEgA+P/U5wmIYB/oPYG7T3DKPmGF00m0JSJJUti0BIy50LBYL\n/H4/3njjDcyYMUOtF+z1erPac61JwRxp8qcTL23HNdLEa9TqPJmORS1nesGU6uLWw7ULDF+gFLJf\nWc4bO5X1eaz/GAaEAYxxjMFfT/4Vra5W+Oy+ign4qEaM5BrOZn4oZfUhavFqMcp3RaG/HyEEbW1t\nePDBBzFq1CgAQE9PD6699lo8+eSTajeadDDBTEI78fp8vqx++EqwvrRQyzmXEn5GOsfksZSjxVgx\n2du3F3WOOrgcLoRJGIeHDmOmeSYLHmKo5LogLGX1ISMtLij0nObOnYt9+/YBON0fk1b0AkZON6kZ\nwcxmD5MKSaZAkWJTTAuTlrcTRdHQLstsSZUvmuv7jcap8Cn4o340uZsAAKMco3AscAzTx0yHy+5K\nGTykzf1kMEaiGNWHtAiCYIxa0xp27doFQgjGjBmDDRs2oLGxUW3dZrVa0dzcnFWhhZoRzExohSSf\nWqlGDfrRQi1nnueztpyLgZ6VlsLhcNZu82KNQ08IIdhzag9sJhticgyEi4+PgOBg/0HMaZqTNnhI\nj4nOiBjRWik3en8n2VQfoguzVO7b5PFQ75WR6OrqAsdx8Pv9+PnPf45Ro0ap+dmdnZ148MEH8fDD\nD0OSJNatJBntZJmPCzbT8YwEHZceLkujnSNtw1MJ+aLZEpWjsJgssJltkIkMnsQtRp/Nh4gcGfb6\nkYKHjJZmUMnUinBnW31ImxKVDPXSGQVCCFasWKH+f+/evWlfO5KxVNOCOVLuYTnHpgc0vDsYDFZu\nykgSkiSpq15aaalQjDIZ2s12LG1dimg0CgA5ubUyTXQ08nakic6oyIoMc21OVWUnU0EOmhKlTXPh\nOE7NLDAKHBcvEcrzPGRZVv9NnwOyL7JQM5seyRNELBZT21UVGihSDOtLjy4jkUgEhBD4fL6CxVKv\ncyzkONFoFIFAQC1nVUmTfqmhE53NZhvW5DgUCqlNjmkRaqMyGB3E5s7NkJTcur5UM+Vc4FHXrd1u\nh9PpVCPsZVlGe3s7Fi1ahGeeeUa9vrKlo6MDy5cvx4wZMzBz5syU3UQ2btwIn8+HOXPmYM6cOfjJ\nT36S07iB+KLRYrEkeF5y2fuvuWWbts+hXu2q6ApGD/S4EWjKCF0VVnowSHKwkiiKalu1Qo+bjF7V\niIxGuspDqYp88zxfdBFViAKeG/m6PNB/AN2hbnQGOjHRN7GoY8qEkRcV5UJbocput6O1tRU/+9nP\n8Oabb2LLli0YM2YMFi5ciEsvvRS33XYbGhsb0x7LYrHgiSeewOzZsxEMBnHeeefh0ksvxfTp0xNe\nt3TpUqxduzbnsdL7+fDhw6ivr0ddXZ1qaQqCAKvVmtU8WdkzaY6IooihoSHVxVDpQpIKaoXRCLBK\nR1EUBAIBtTMMdfsUOoEZWQzTjU0Ugfb24c/19nLo68s9zcBms8HpdMLpdMJsNqtuNpqDXKyKLbIi\n46mdT6F9oD3j6waEAZwInUCLuwX7eveV3co08jVjBKxWKxYvXozLLrsMd911F44ePYq7774bbW1t\nI3YHampqwuzZswEAbrcb06dPR1dX17DX5Xs90t/uySefxJ49ewCcNiR+9KMfYcOGDVkdv/oUIw20\nxZPb7c45/WAk9HbJ5nM8WmwhEonA4/HoHtZdDpesJEnqAsftdhdlgUNXyZVgQXAc0NPD44svTn8P\nvb0cdu/mwfP5jz/ZzUa3KGglKFo6US/37d7evdh9ajfWta/LeLz9ffthN9thMVkQk2PoDHQW/NnV\ngNG8H8njoQ0q6uvrcf311+Opp57CpEmTsj7ekSNH8Nlnn2H+/PkJj3Mch48++gizZs3CihUr1HzK\nbGhra8N7772HzZs3Y+vWrfjiiy+wc+dO+P1+7NmzR/VYjXR914xLlrpgabBPuQVOz+MpioJgMJjW\nzWy0GywbotEoIl1d8OzaBcuRI0BLC5QlS4Ayt7wqBn19cQtx6tREt/7u3TymTFFAHQVmMzBvnowd\nO0z44gseY8cSfP45j7lzZVpKNy37+/bDY/Wg2dOc8XXa4CGe52G1WtPWKM1nH1lWZLzd/jbGe8bj\n0OAhtA+2Y0r9lGGvGxAG0B3sRp21DgBQb6/Hvt59aPG0wMzXzLRVkRSSVhIMBnHjjTfiySefHJYX\nOXfuXHR0dMDpdOKdd97BddddhwMHDmR13K6uLjz77LPo6OjA888/jxdeeAGxWAyBQADXXnutat2O\ndD3XjIUJVKdLRRRF+P1+tT+nViyNWrIvE3S/MnroEBp+/nPYXn8d3Jdfgl+7FuaHHgL35ZdFH0Op\niUSA114z48svT/92n3xixvr1ZiTHTVDR7OjgsGMHjzlzRhZLQRKwvXs7Pun+BArJba+dBg9pgzx4\nns87eGhv7170hHrgtXnhtXrxTvs7Kd+3v28/nJbTkZZWkxWiLJbNyjTSotNoY0km3yhZURRxww03\nYOXKlbjuuuuGPe/xeNTjXnHFFRBFEf39/Vkde8mSJXj11Vfxhz/8AQcOHMD+/ftx+PBh9Pb24rnn\nnlP3V1nx9RSU2yLU43jZVrnRq3arXmQ6N2opA0Dd2rXgRBGktRUAQEaNAvx+8M88A+5nPzOWC3Vg\nAHxbGzAwAJxxBpTJk4EcJoyWFoLbbhPx0ksWXH+9hJ4eMz76yIy7746hIUV3r8FBDjwPKArQ1cWj\noSGzCB7sPwgAGBKG0DHUgQm+CTmdHoW6r/Ntmk2ty3p7XOEbHA0prcyQGEJfpA8ykTEUHYJFtIA3\n8VCg4MjgkbIG/zBSk+ySHTduXE7vJ4TgzjvvxNlnn437778/5WtOnjyJxsZGcByH7du3gxCChlQ3\nSJrjy7KMBQsW4M0330Rvb6+6EOQ4DldffXVWIs8E04DHG4lydOUo9jlq26g5eR78nj0gLS2JL/L5\nwHV0gOvuBkaPLujzdNuTPX4c5nfeATGbQex2cB0d4D//HNJVVwE+X9bHGT8+LppPP22FLPO4//4w\nGhqGL4LonuUFF8jweKC6Z2fOTC2agiTg81OfY5RjFKJyFDtP7sR47/isIlRHIlOOXqq6t9S61Ao2\ntTK/f9731UnXZXHhkomXAADCkTBsVpt6jesxboZ+pFqM51O4YOvWrfjd736Hc889F3PmzAEA/Mu/\n/AuOHTsGAFi1ahVee+01PPXUUzCbzXA6nXjllVdy+gyz2Yy7774be/fuxZQpU2A2myFJEvr6+nDJ\nJZcwwUyH0YM8si0On02VG6OfK3C6ebXaRi0aBXgekCRwnZ3gjhwBJAmkqQlwOACj1MCVZZg2bYJS\nV6dalMTrBdfbC37nTijLl8fPhRC1sXUmjh8/LQanTvFoakp8PhKJ72tq9yzpnuaRIxwmThz+Ox/s\nPwgFCsy8GWbejO5Ad0FWZiZG6pDxwaEP4sE7Q50Ad/raPOI/gs5AJ8Z7x6vHspnjQWuySYbNbCvJ\norBSMJLHKBW0F20uLFq0aMTUvHvvvRf33ntvIUPDp59+qkbJ5kNNCWaxLrJSiZIRKxPliva7IuR0\n8+qEGr42G5R588C//DK4cDguRhYLuPZ2EI8HxGYzxiJgcBCnAidgPWM8tLYkaWgAv28fuGgU/LFj\nACFQWlogL1yIdBuO27aZsGWLCQ8+GMXgoISXX7bBaiWYNu30JOJwAIsWydAGQNM9zVSXgta6pHjt\nXl2tzHSk6pDxNzP/BqFoSI1I1FqfY12pg7mMIg6pWlgx0luYRqr0A5ye+8866yysWbMG5557Lux2\nO2w2GywWCzweT1bHqSnBpBjdJZuqy0hKYSnh2PQ+R1qyj5DUzavJhReCe/bZuKUZjYIoCuByAS0t\nMG/ZAlx6qW5jSSbbfV+FA14Wd2B04ARW1i09/UQsBtPevVCcTpAzzogfs68P5rfegnTjjcP2N/ft\n47Fliwnf/W58z9LtVvDtbwv44x9d+Nu/FdHcfPp7T5UtlO5yaB9shz/mh0wSizyExBC6gl1o8bSk\nfmMR4DgOY9xjMMY9Rs3vpIXjZVlGVIgmFE4wgkgy8sOIgklpaGjAgw8+iIULF6pGByEEzz77bFbX\nHBNMA6IdnzYQRq/KROVGURQ1stfpdKa+UP1+kHPOAbHZwA0NAS5X3CUrCOB37y6qYGbLQdKL4zYB\nXf796HadgzMs8QAEfv9+EJ8PZMyY0y9uaAC6usAfPgxlxoyE45x1loK77oolbHm2tChYtUpEQ0P+\n12mzuxnnNp6LUDSE6WMSK6bU2eryPm6h5BI8ZOT7tFwYxeoGUkfJ0kBEI/Kd73wH9913H06dOoVY\nLAZJkqAoStbfZ00KJkWvC69YAkwDYaxWq1qzMR/0sjD1KP+nKApisZiaIJ8Wlyu+VzluHIg24m5g\nAJgyPG8vVwr9zRSi4J1D78B35kyIB/+KD45vwkr7AnCKAtjtIKmCkhwOoKcHhwYPoc5WhwZHXGAt\nltTxQaNGFfa7+Ww+bDi6AQPCAJZOWAq7eeR91Gxp62+D3WLXxUpNFzwkSfHKPpFIpOrallUTqVyy\nRhXM888/H1u3bgUArFixQs3Lz5bKN1dygP6wxbrh9OyJKYoiAoGAWros3zEbZXKhbmVJkmCz2TKL\nJQBy7rkgLhfg959+MBYDF4lA/vrXy255HOw/iO5gN3x1Y9Ew5yLsGmfC8VlnQrriCkhXXhl3IScj\nCIjVe/H0Z0/jzQNvFn2M+3r3oXOoE2ExjO1d2xOeU4iC3+75LTqGOnI+riAJONB/oGjl6mjgEI20\npJYo3cOn2xOKopTsOjCaVWeUsaSCVvoxGoIg4Mc//jEeeOAB3H333QCADz74ANdeey2A7ObvmhJM\nLXpahXpevLR/JS00rkcZv3KLC02DicVisFqt2UU8ulxQfvjD+L87OuLpJD09kG+/HSSpIHOpodYl\ndWvyZgtsDY14394FMm4cyIQJcQu5r+/0mwYHAasV21wDGIoOYceJHegKDK+VqecY3zjwBnw2H8Y4\nxmDNwTUQJEF9vm2gDTtO7MD6w+tTvj/TNXPMfwzggJgcQ1eweOcADK9763K5YDab1W484XAYgiCo\ntW8ZpSWVeAuCYKh+mJQTJ07g/fffx2effYaxX1UMmzFjBnp7ewEwwcyIEQN/ZFnG0NAQgHgvRD1C\n6fUS83zPT3tOdA822+OQKVMgP/EElIcegvzAA5D+7/8FueyynMegNwf7D+KY/xgsJgvCYhhhMQy3\nxY2dJ3eiO9gN2GzxPMxRo8B1dYHr6gLcbkSu+Abe6vgzGp2NsJqsePfQu+ox2wfa8Xb727qNkVqX\nPpsPDosDITGkWpkKUfBu+7todjfj4MBBHBs6lvIYqa4dQRLQNtCGOlsdfDYfvuz7sqhF0ZMnZI7j\nhtW95XkeoiiqlYf0rHvLyA8jxlpIkoTRo0fj4MGDarvDI0eO5GQN19QeZvKNZyTBFEURwWAQDocD\nsiyXXej0QHtOeafBWCwgZ5+t/+A05Pod9UX60OprHfZ4i6cF/ZF+nOE+A6iri4tmOBzPw3S58EnH\nFgRiAdR762E327HjxA5cfublaHI34e22t9ER6MD5TeejzlJYQI7WuqTfObUyL2i+AMeGjqE71I0J\nvgkQFRHvHX4P35313ayOTa1LE2+CCSb4o350BbvQ6h3+fRQbbeoKkLlpdqrKQ7lgJPE1kks21ViM\n9F1pqa+vx+LFi/H0009DFEWsW7cOL7zwAm666aasj1FTgmlEaBcV2knFYrGojZ+NRC6ioi3bR88p\n+flSjSUT+R5jYctCLGxZmN2Lv1q9xuQY3mp7C06zEzu6d2Bu01zVylw0fhG6Q91wWVz4sONDXHvm\ntXmNi/LX3r9if99+1NvrIQRPu2F7I73YdnwbPj/1OerscVEe7RiNtoE2HBs6NqLoqdal/bSgUyuz\n2dAFM4IAACAASURBVN0MM+GAUCgexZTkkjs8eBjjveOLWjhdGzxks9kSgoe0lYdoa79cRccoImVk\njDZvUQghGDVqFG699Va1duwzzzyDlStX4qabbso6z7ZmBdMIFiZtn0QIgc/nU38wvSJS8x1XIYxU\nts8oKT2lnvy2d23HidAJ+AU/jg0dg9fmxRjnGGzt3IquQBfq7HXwWD347ORnuLDpQoxxjBn5oGmw\nm+345te+mfK5QCygWpdA/HtwWpxZWZndwW4IsoABYSDh8agcRe+hPRh3tB+QJEBRQBobocycCdhs\n6Iv0YcPRDVjWugxn1p+Z93nlSq5NsysBI9w7WlJZmDRtyEhwHIeDBw9i06ZNeOyxx9THu7u78frr\nr+OGG27IynJngqkjuRwvoXZqAVGwpSKb7yvXsn1GpRjXRkyOwcpbcTJ0Ei3eFvSEe3DOmHNgM9lw\nPHQcsxpnAQCsvBWbOzbj+qnXA7IMxGLxsno5fJeT6ydjcv3klM89t/s5iIqI44Hj6mN+wY+QGMKJ\n4Ak0uZtSvg8AznCfkWBdUrj+AXh27gEZPfZ0FYX+fvC7dkGZPx+7T+6G1WzFzpM70eprLUt7rlTu\nW1o4gaYVVFLqipHHZzRRj0QiGBoawhtvvIEPPvgAt9xyC7q7u9Ha2op3330Xf/rTn3DDDTeoi6hM\n1JRgFtPXnssFHI1G1WoYqRo9F6OqTrGh+5V2mw32EfYrjXZDlYJZY2fhD/v+gNHO0Vgyfgm6Al1Y\n1LIIHx//GC7r6Zy1Rlcjdp/8DMu67Wjdcyhei7auDvLXvw5l2rSsP09WZGzv2o4F4xYk/BbXnHUN\nLp10uuhDd7Ab/7P/f3Dd1Osw2pm5oL3dbE+Zy8kfbwM8dYklh+rrwfX0oO9UvE5ss6cZ3cFuHPMf\ny9rKLOZ1QoOHkuveiqIIQRCG7X0aad/QSCS7MhVFMZy1HggEsGbNGrz22mvo7e3FI488gmAwCJ7n\nceTIESxbtizrY9WUYGrR++LPtiVXOByGKIo5l7grZFzFRhAECN3d8L31FqxbtgCKAmXhQigrVwJf\n9ZnTczxGcetq6evjhhUakCRgaChe5Gfj0Y3oCnbBa/OiJ9SDUY5R+P2+34OAwG11IxALqO+LHGvH\np8c70HrGxYDVCgSDML/6KsRvfxsky6INu07uwnOfP4d6Rz2mjTottGOcia7ezR2bYTfb0T7QjvnN\n85MPkxVcOAySqmYfz+PzE7vgtMb3cevt9XlZmcW+hrMJHqIWqcViYcKZASOmlHg8Hlx++eUQBAH9\n/f249dZbcfLkSUiShEmTJmHSpEkAkFVWQk0LZildsrTEHcdxI5a4M6KFmWpMdAEgBYNoWL0a/IkT\nwNixAMeB//hj8H/9K6THHweyLGxcauj50By+fF1xsRjw059acc01EpYtk786JvDUUxa4XMC1t57A\nnw7/CS6LC26LG53BTsxxzYGkSPj6hK9jxhhNqbxwGPzmF+EeMy4ulgDgdoMoCkwbN0LKQjBlRcba\ntrWwmW1Yc3ANvtbwtZTn1THUgcODhzG5bjIODR5CZ6Azr8o9SkMDuFOnEssVEYLe2CA6xACavRMB\nxC3UAWEgJyuzHKQKHgqHw6r7ttDgoUIwmqWbPB4j1pF1OBxobW3Ffffdh2AwiPb2dowdOxZOpxM8\nz+c0ZmPZzkWm2Gkl6RBFUa2d6na7s3JZGLGoghZFURAIBKAoCnwHDoA/fhwYNy7uljOZgOZmoK8P\n/KZN8TdEIuC2boX5f/4Hpo8/BgQh8wcUGfr7h0IhCIKgVpERBEEtDp4tVivwf/5PDK+sGcJb7/lV\nsYxGOdx2m4gtHVtwPHgcMpExGB3EgDCAL3q+AAAcHDiIqQ1T1b+v8Y2YSkahyZLU1cTrBX/yZLxr\ntIZtx7fhqP9owmO7Tu5Cb7gXrd5WHPUfxf7+/SnHvaljEzxWDziOg9vqxqZjm7I+Zy1k4kRwkgR8\nVfMYogiupwe7vGFIFh6DwqD6x3M8/nLyL0XN3dQbKoo2my2hWXs0GlUrD4miqFugXiVD2/QZCfq7\nbN++Hffddx/uvvtu3HzzzVixYgVmzJiB3/zmNwCgdtHJRM1amID+e5ipLLBM6RWZjqUneluYNGCJ\nlrjjDxyIpxIk43AA+/YB550H009/Cq63F+A4mCUJprVrIf/TPw1z2WY7lkKh+1aKosDlcqkiSdMQ\nBEHIyZJoaiLwXf44fv4hjz+++P9h9iyCH/wghiGpD13BLnzjjOtx4iSPyWfKkBQJ/qgfV0+5Gu1/\nrUNXF6d2JCFuN3hChgkjQiGQ+vp495avCMaCeGbXMziz7kz808J/AsdxqnVZ74gLrtfmTWllUuuS\nRss22BvytzI9HsgLFoA/eBBcXx9gsUCZPh0NjjPgkKMJLz15kgNRTJAVWXXLyjLwl7/wOOccJTkb\nxTBQSypd3VtaOJ4+X6zgIaNbmJFIxHAuWbqv+uKLL8LlcuGTTz5Rn9MujplLNgXa1k3FdMmOlF6R\nzRj1QO+biwYsaVfaaGqKz3rDXww0N4N/7jkgEACZMAHkq+4Apv5+8M8/D+Whh/IaRyHfjyzLCIVC\nccvK7VZdsjQNQRRF9dxSddAwm83DvteD/QdxIPAZ/C4eQdsBzJ59JiwW4Ghf3Pqz2Ai6ugBR5PC1\naWb4bD7s2+PAiX0TsGSWePpAXi+kmTNh+vxzoLU1bq3HYuBOnYJ0/fUJn/nBkQ8gKiK+7PsS+/v3\nY9qoaaety68KK9Tb61UrU7uXualjE8y8GWExrD5m5s3YdGwTvj3j27l/qT4flHnz4kL/lajPSvGy\ngBfYuNGMznoZkycTyDKwZYsJNltWPbYNx0hNs5ODh4wkdsXAiJ1KqEdv0qRJ8H21bSBJkjon5xKk\nVHOCSdEz15Eej0LTK8xmsyHSK/RK8qf1O5MXAMqFF4L//e/jhdLpPlYwCPA8lLlzYX77bZDx4wF8\ntRgAgMZG8Hv2QBkaArzegseXLdT1arPZIIpi2t+GBoKk6qCRKgn+93tfwfFjdjgswJTrX8Tat34C\nsxlYtORcVahungr84Q8WNJ9S4HQCe/Y6ccft4rBOJeKll4KYTDB98QU4AMRqhbRiRTyv8SuCsSDW\ntK1Bo6MRgVgA//3lf+PhBQ9jbdtaSIqErkAXbOZ4II5M5AQrMybHwHP8sKjY0c7R6vN5M8Lk4/EA\ny5ZJ2LjRDEWR0dXFw2YD5s8f3gTbaIFdI6ENHqJNs+k1I4rxRZE297Pc84IeJFuYwWDQcC5Zeh3V\n19fjpZdeQkdHB2bOnKl6/BYuXKjWlh2JmhbMYrhk6YTscDhG7MhRirHpcVPSoAcgTU/OhgbI//zP\nMD3+OHD8q/w+txvyQw+ld7nScUkSEAqBO3Ag3ii6pQVo0b+xcbJ7nHaEyZZkS0LrhvtrzwG8+eln\ncCvjMHOGgkORXfjHe/fij/8ZF7hly+K3mdMH3Hk78ItfxK3XH/wglrKtF6xWiJddBv6SS8BFIiBe\nL2CxIBIB1q4144YbJHxw5ANE5ShsZhsCgzZ8dGo/9k7Zi2mjpkGQBAzFhjCrYRZEicNAP4cJXjtk\nIsPMmWE1WXGh89tweqL4n2PP46ZpN2GUY1TCEKJSNMXA9MHjARYvlvCnP8W/l5tvltKmmRpBVPK9\nF7XuW+r6o+JJXf65Ns02mks2GSN2KqHfFyEE06dPh9/vx7p16xCLxXD48GGsXr0aY8eOzSolpmYF\nU29ol5FYLFaylJFsKUR8tQUWJElKe0GRadMgPfUUcOQIOEUBmTgxvq9JSNy6HBiI51dQ+vtBJk0C\n19MD/v/9v3ioKcfFX79gAZQbbhjRWsl28lCjeSVJtY61xblTVSrJ9J0l72P977G18LmsmNYiA4TA\nBBP+1P07/PCHP0JnZ+Le7u7dJrUOwaefmnDxxXHLav3h9ZhUNwlT6jVRsE4niGbysViA7m4e//5k\nDAe/Frcu+/o47NptwsSznVjbthbfn/t9nAqdQhNpwsUTL0Y9NwFr15oxo0mBmY97VA4d4rBpkxlN\n52/DR8c/wij7KNw0PbGeZralwvJBluPfQ0MDQSTC4dAhDpMnx7/viBgBAYHTYqxJFyhMvOn+ZzZN\nsyup8lAyRnXJyrKM733ve/je976Hrq4u8DyP+vr6hDz4bL7zyvxVCoBe9HpacbQpsqIo8Hq9BYul\nUSzMaDSKQCAAh8OR3arRZAImTwY566zTQUAcB+Wuu8CJIrjOTmBgAPzx4+AkCfKtt4L/7W/jLtkJ\nE+J7duPHg9u6Fdzu3bqckzaaN9e95Gw42H8Qn53aiekT6sGbAfDxAJrPTn2GPu4LnHNOAIIgQBRF\nfPQRj7/8hceqVTHce28M7e083n/fhJ5QLz47+Rk2HdsEWUkfqWc2A9//fgyHuffxl/29OHJyEB/t\n6UHr9ONwe0XsPLETa9vWwmqyos5eh62dW+HxEFxzjYTdu3ns2cOrYnnp5WFs7ftfnFl3JrYe34q+\nSF/az82WmBzDlo4tUEj6rQ5BAF56yQyLBbjkEhnLl0vYt8+Ebdt4dHZy2Hly57DendUIXXTRtmVO\np1NdyIXDYYTDYUSjUcO3LUtetBoxSpamjO3btw/3338/br31Vlx77bW46aabsHnz5pyOVXOCSdFL\nlCRJwtDQUMJ+llHGRsn1WNQii0Qi8Hg8KasR5XS8KVMg/eu/Qrn6apDJkxFdsQLSv/4rOELiM6h2\nRcrzQEMDuI8/LugzgdO/jdlsVt2werO/fz9cFhf8UT8GhUH4o34ExAA8Ng+OhY8hEnFCFE04ckTB\nxx/LuPHGITgcMfT1KVi5Mi6ab+z4FC6LCwPCANoH2jN+ntkM3H5jHcZ034qeT76Bb89fimvPWYav\n++ZiRXg8jm5ei8ajPfDKFpwKn4rXrfUC11wjYfNmE95914wrr5TQIf8FQ9EheKwemDgTPjjyQcHf\nxb6efXj30Lto629L+5ojR3goCnWRxd2z06fL+POfzRgS+3HUfxRdwS5dBLySoO5+u90Ol8uldvdJ\nbpqdTepDqUg1rxjRwqSxKj/60Y/Q0NCA119/HR9//DHuvPNOPProo2hvj99z2cyTxvEblphCRUm7\nJ0bTEnLZEysVuYqEtiB8qv3KvPdQGhuh3HJL3G0dicDu9cb3OzkO6O0F9+WXcbetzxd34brduX+G\nBjrRpCs/qBdXTbkKV025atjjoizCYrLgnXdMOHjQirvvFvF3f0dgsViweTOHDRtM+MEPgrj02uN4\n7dAXaHWMR0SKYEvnFoz/2njwXPqFV0PwIkzyLwfHAbZ9Cla27IJ9w+t43xaFYrHAtP8gcPgIfAvm\nYmvnVrR6W9Hbe/p4HV0i3va/rQb9jHWNxftH38cE3wRc0HxBXt9DTI5hc+dmjHGOwcaOjZjSMCXl\nOXztawqmTlXw6ac8PvnEhAkTFBw4wOPee2P4MrIPNpMNPMdjz6k9WDJ+SV5j0ZtS7xtmCh6SpHj+\nqiAIhgkeMnpaCR3f0aNHsXr1aoweHb/ur732Wvzbv/1bTouQmrUwC4GmjESjUXi9Xlit1rIXc9fj\nONQi43keHo8nQSz1uim13xNpbQV6e8F/8AHQ1xevADA4CH7DBrUtVjbH0UIIQSQSQSgUgtvtHlks\nZRn855/D8tJLML/yCriDB/M+N0ogFsDq7av/f/bOOzyqMu3/n+ec6ZNeCCSELk2kKUVpiiBYcRXr\nWtay6trWrrvr+1vd9XV91XVVXNeyVlzLqmtnFekdQQQB6SUkkIT0STKZmVOe3x8nZ1JISAJBRsP3\nunJBJmee85x6P9+7fG/2V++n3+htGGnf89JLlqTaypVuFi/28tvfQnKyl/Xl3+JSnEQiEZzSSXF1\nMdtLtx9wbLar9ocfFJ591sWdd0Z4/vkQTjRWP/glxcl+vouvJtGbTCQpHk0P49u+i/yqfJZsyGXR\nIgcXXaRzxRUan337HbvzK6NxQlVRKQ4W88b6N6jRag7pmH8o+oGgFiTNl0ZZTVmzLFMIy4kwcqRJ\nUZFg5UqVUaNMFH8ZORU5JHuSSfIksa9qH8XB4qNuDGIBtvvW4/HgdDqjnqxYbJodDAaJO8zFbnvD\nvofGjh3Lyy+/zOLFi/nhhx/49NNP8Xg8JCUlNdjuYOhwDPNwY5jNdeSIlbjjoY71YzGyBkhJQRgG\naBq43da/kQiyUycra9YwrLhoK9GW2lchBFLTUF54AWXNGkyvF2GaqAsXop9+OvL88w/5sJbmLmVz\nyWZm75qN1+mlyxAdY2Nvfve7eHw+uOuuCKmpkuJgKdsrtpOVkIUiFEzTJNmTzJK8JXSLs+ooHQ4H\nW8q2sKlkE1O7Tufvf3dy++0RBg603Ey3TM/j+wVhlmxzkzLAh0atgk68F/btwdV7EF8uKmPaCIO0\nNIkQoPX4ksJNGitDe0lPl+giSCAcoFqrZnXBasZlj2vT8drs0masSZ6kg7JMsAQMVBVcLsnWrQp6\nlsUu7fvV6/CyoWgDozNGH8ol+NnCPj9NJQ/ZKlU/VvJQU8w7FqXx7Dk+9dRTXHfdddxxxx1omobX\n6432xmwtOpzBtHEoBq5+yYi7hY4ch4v6AguHi4Mdp83IWpPd255zAqCyEuLjMceOtRKCIhFkr16Q\nmYkoKLBYZytv5sZava1aLW7YgFi7FrN7d+scCQGGgfvrrwmPGEF1Zhpzds3hzN5n4lJdrTukSCXz\ncubRP7U/83PmMzJzJMmeZMJJW4CTcDgkfr91PVbmryRiRiioLkDX65p9BCNB8mvyyRK9iOg1fLXt\nK8oj5ZzU6SQefTSbhAQFsI7P4XEw4iQdrUsXXK56Ta0jEYRSTviUa6g4HoqLJdu3C/r0kVw3/EpG\neTRcLujZExbmLmBA6gC8Ti/bSrdxUueTELT+GtvsMsWbQlWkijhXHHmBPLaXbqdvat8Dti8oEKxZ\no3DyyTrJyTB3WQVLvtvN6EFpaIYV1vA7/eRV5lGSUBJzL+BYQlO6t03VC/9Ybcti0SVrIycnh1df\nfbXBZ+Fw28qnOqxLti0Gs7Gbz+PxtLkU4WjhYA+IlJKqqqpoucWP1T0lep7cbqSiQFIScuhQ5MiR\nVrasqlriBq188A41uce5Zo0VK62/fe2+1S1b+K7gO1blr2JzyeZWH9/S3KUYGDgVJ2Vhy81YsDON\n/67awl33l3PCCSYvveTkh8IdfJf/HZcOuJSTnJeT/9WVnJ19KZcOuJQrjr+CBJHJU0/FMW99HiFC\npHhTWJK3BIcjRDAYJBQKoes6Zno6IiMdV1Vpg3mIggKMESMQApKSoHdvi11u26agF/Wkf3pfzjjx\nOBI9CQgE/VL70S2hG0KINh2vZmgszF1IxIywrXQbc3fPZUvJFmqMGhbkLjggYzYchu++Uxg92iAl\nxTr1fY4vJ96RyM7cCCE9FP1JcCcQiARaPZcjhViqfWxpLnbykNfrjSYPAQckD7WHaEtTc7HrnGMR\n9913H/v3749KYpaWlnL33Xe36b3d4Rhm4wvc0g1YPwkmMTGxWRfHkRJCaA80NY5hGFRWVrapgXW7\nLwrcbuSYMSiLFlmJPrZ7e+9ezBNPPGiXk8ZCEYfkSlZVK1Wz8dhSUkWEVfkb6Z7YnSV5S+if2r9F\nlmmzywxfBuXhcuKccazauZNO+VWccYakVGzlwguH8MEHKn9+ez6pA4rYE9jD+CETKNrl4M1/SO65\nS8clNf42w8ugwRE2h+eRHJ9GvNvHnsAeyowyMuMy2bPHJCNDI2QYhM48E/8HH6Dk5Fj1foDZqxfG\nKXWMUwjo1UuyerUABCeeaCIErC5Yjd9Zl9WY5k1j3f519IjrQZza8otPCMGEbhMwpcnKfSsJRAKk\nelMZlz0Oh+I4gKm63TB5stGgdWaPpO7cdVZ3TLOhB96uUTyGQ0P95CFo2DS7psaKVbe37m11dXXM\nZcmCtahet25dA/drSkoKCxYsaNNxd2iG2RIOlgRzpNFexqmp44xEIgQCgWgK+9FcPZvnn495wgmI\n3FzIy4M9ezCPOw7zkksO+j07S7m6uvqQS1+0E0+0JPzqn2dNAyH4Nk1DIol3xVMdqW4V67LZpSpU\n9lXtI84Vh8+jknnSt3RLT2ZzyWZq9CDDJm3Fl7mLvql9WLBnATV6kPPP1xHH/Zdb/7KeRx7xcfzx\nOsefupF1W6pYsSgBw7Q6iizdu5Svv3byz3/6cTgsFuHMymLzpVOZNTaLqtNPp+qyywhdcgnSVWfg\npYQdOwRJSZCcLNm5U1BQVcjOsp2E9BAFVQUUVBVQHCymPFzO1tKtrTqHDsXB0IyhdE/sjiENxnW1\nDGVWfBaDOw1u8t5qypEhRJvC1cdwCBBCREtXfD5f1FNmJw/VL11pzbunOYYZiy70SCRCly5dWLly\nJeXl5VRVVbFu3bo2q7F1OIZZHweLydki461lLrHqkoU6himltJo9h0Jt6p7SnjjgPHm9mDffjJmX\nhygpQSYlWW7Zg8CWGYtEIgdl/S3NQxswADl2LGLJEoSiWFzINCmZdiarItvJiLP0JdN96SzZvYAB\noXicSalNat8apsGKfSswTZOtZVsprCpEVVSkS7Kjej17K3shkewu382THy4jJSset8OFXqOzKn8V\nieHj+b74e4qMbYjyQZw6McSHexZw6ohkVi2DeXNVTpuYwuzVu/BvKeChu9JrtSEEiqrwdcEC8tQ8\nThz2C+Id8Wi6Tigcrk38UNm920lNjaCwUDB5ssnOnYJ9e7xM6jGZrVsVvB5Jt+5118VD214kawrW\n4HV4UYSCz+FjVf6qJstt2oJYcoXGCtpLgakp9tm4abb996aaDTSHWHXJejwebrzxRn73u98xdepU\nqqurmTt3LnfffXebxjlmMJtoyRUMBtE0rU0Sd7Hqkq2vo2hnkB6OkTlii4KuXS0d2aYQCiHy8pBe\nL2bnzlTW9l1sbW/RZqEomFdfjTF6NHL9eoTLhTl4MN+GtsB+cCrWgsL3w1bKvpnNrsJPGVIdhzF+\nPJEbb2wguKAqKneMuANDGoT1MNVaNWAl+jpVhSSvlbpeUFWAu/MuNqzqRaLLoFNqJz7dsJDtSwKU\nSy/H9Y8wwL2GP89II21iBX6vgz7Dy/n2W5UX3xG4vZKbL1pNSsqZ0X3vKNvB3sq9uFQXy/ctjxoq\n+yVYXm65Njt1MnnjjQRKShQuucRg9+5E9m9L4rNXXdx9d4T+qXVxrVAb+pWW1JSwpXQLXeOs65fi\nTSGnIofC6kIy/K0TtY5ldATD3drkofqt7po6L+FwuK6LUQxBURQuu+wy+vXrx+zZs0lJSeGVV16h\nX79+bRqnwxnMluKVlZWVKIrStMh4KxCrD5etRnREu6fs34/Yvh38fuTxxzfte2sjxNdfo77+Omga\nUtfRe/TAfccdhFrT4SQvD2XRIsTu3ciUFOSYMchBgxom+QgBxx2H3qMHiqIQ0kOsXbUWzdTYV7UP\nZfdu1EULMOK8LM02GVSajjp/Pq7qaiL/8z8NdpfkSTpgCh995EDT4KKLdEDy2vevocmaaFurnj09\nrNhehanM5vQ+Yxg6PEzO/sWcKH/Non/fzkMPhYlPgF4lKp9tcJLpNZlyXL1OMdJk9q7ZJLoTiXfF\ns2LfCk7JOoUUb0r0JZiW5iA11WLlf/hDDY884kHTTAYO1Hn5ZT933x2if38JbciMrY81BWtwKS4M\naUDtesqlutqFZR7D0UFTbcts42m3LWuOcMSqDq6UkuHDhzN8+PBDHqPDGcz6qH/BNU2jqqoKj8fT\nZBZsa8Y6UnM7HNjKIC6X65COq1VzMk2Uf/4T5ZNPrKp0KSE1Ff2hh6y6hdaM0dT+1q5Fff55yMjA\ncDqtPpV5eXiefJLwQw8d/Mt79qC+9hrS50Omp0MwiPL++5iBAHLMmGa/5lJdnNfnPIRiGRvX239E\n1PQGNQ4HCorqQGZmoq5ebbHeFjqrTJ2q8/zzLt5/38HEcwr5fPMcNucVcuYJHspLnXyxWMXTdR8R\nzcTlEtRUu5k9FwZ6thIqOwUtEGb1WoW1yx387X8jfPihg9deEtxySwSns45d2o2gVaGybO8yzulz\nDuGwlWRjn3chBGlpCn/4g8Ett/j5/HPJH/5QRY8eEYLBhq2nWrvwM0yDQDiAS3VRGamMfu5UnIT0\nEGE9HG0zdgyHj6OxIG/svrXZp6Zp0aTIDz/8kPT09DZn2ufm5nLVVVexf/9+hBDccMMN3H777Qds\nd/vtt/Pf//4Xn8/H66+/zrBhww7pOOz4rBDikPqTdniDafd4bI+4XrvXKR4G6kv3AYdtLA8GMX8+\nyn/+A1lZdZkbJSU4HnoI/ZVX2sY0S0tRFi6EbdtQVq1CqiqGw4Gh67hcLpSMDNi9G3XTJhg1qtlh\nlHnzkH5/XYeU+Hikx4Myfz7G8OHNditWhELX+K6oqmql5+8uR2ZkgK6AYSCKC6360IoK1M8+Q7/i\nioNm83q9cPPNEZ5/3sXT/6ihUHOS7s2kavMpdA+cx8B++5m54Q3iZVeWVqqs+RaS/WnksJR/vTCQ\nNd/4WbjQwT33hElJgeuv1/jnP538/e8ufnNzKMoubWT4MlixbwUnZ53CG//ozIgRBuPG1Ul/LVqk\n8vnnDsAyoJs3exk82IGUdS9B2x1b/6XS3L2jKioX9r+w2eP/OSBWnulYgc0+bebpdDoJBoM8/fTT\nrFmzhsmTJzN16lSmTJnCoEGDDnrunE4nf/vb3xg6dChVVVWceOKJTJ48mQEDBkS3mTVrFtu3b2fb\ntm2sXLmS3/zmN6xYseKQ5n64zRdikzv/iLAzwxISEo5KEkxzOByG2Vi6rz3joU2No370kVXsV/9m\nTE2FkhLEhg2t38GePTjuugvl7bdRakUFxLp1mCUluFQVdetWxOzZKGvWEPfQQ5aEXlMwTURODiQn\nN/y8tt0YJXXC3i2dF7NHD0tgQUrE7t2Qnw+KglBVlD17cL75JtT2Cm0OXi+cc47OwvK3cQgnYxy3\nwQAAIABJREFUfbNS+TbyLiY6gyZsJiXNoEwrZG/lXvIq91KuFzNqbAVF+i66dJFRYwnWKb7+eo0R\nIwxyq3expWQLZaEy8gJ55AXyyK/KJxAOsGrfN1x5pcbs2Q4WL7auy6JFKm+/7SA/X/Dww2FefrmG\nVatU/v1vJ4pi6Zba9Xv2C7GmpuYn0zmjIyCWjLftflVVlZtuuokvv/ySIUOGcNttt7Fz506mTZvG\nww8/fNAxOnfuzNChQwErJ2HAgAHs27evwTaffvopV199NQCjRo2ivLycwsLCNs+3uLiYZcuWsXr1\najZt2sTOnTupqKho0xgdjmHaN5sd0HY4HMTHx7fLTdje8niHMpYdhz3i8cr6KC+3dGCbQmVlg1/r\nJyE1npv66qtWX8yuXa2ElS5dUPfuxb15s2WQ9+wBrxfp8SBdLpxPPonp8x3oYlUUi/WFQg3FD6S0\nsnBaWXcKoF1+Oe4//YnC3Ahp4UrUOC8iEMDs25dAVn/mf7+aE9bMo+vY5mN1OTmCZ2fmInouJrE8\nk20/qHTrk0u44isW/etMHpg0hFdfdbFjo0K3PibdRJAbi+eQ/dfXcbg8GFOmYIweHe0PqqowZoxB\neSiVvql9MUyDiT0mNthnsieZ9HjJ7bdHePZZF4sXq+wq38mqpH8w87Y/M2CANdb/+39h/vQnN+np\nktNPN6LXyC5BUFU16oKzC94bZ08eqXsslozDMbQODoeDadOmMW3atDY3pNi9ezffffcdoxp5jvbu\n3Ut2dnb0965du5KXl0dGRssJZfY9tHPnTp599lnWrFlDTU0NmqZRUFDAhRdeyN///veoLWjx+Fp9\nND8j2MXuqqq2q8Td0S4taS4Oe9B5SYnYuBGxZAkYBvLkk5FDh7bYvLk+zOHDURYsgM6d6z40DDBN\nZO/ezX/RMBDr1iE2bgSPB/Htt9C7N6Zpomkajl69EEVFUFaGKCtDxsVZRjA1FZmejgwGUd98E72J\nmKQ5ZgzKF19YAu/2seTnW7066zeyrsXWrQp9+jT0Hus67EwazYD77sP1hyepzK8mPsNEDB5MxeBT\nmL3Y4MuBm/lhS4C7xpzd5H2UkyN48UUXzpFvkmo4SM2AzVskeiCVFaG3GOecwt//msGoUTpZqQoO\nM8I92x7AM3sNjp4uHMLEMX8+2jnnoN17LyWlCqmp1rU0pEFZqAxTmrjCnenXNf2A/aenS4YNM5g7\nVyUw7EVM+V+K4yYBpwGQmGgZzebWO/XjV407Zxyp4vdj+GnCXkzZEEK0OmO2qqqK6dOn88wzzzRZ\nltL4/dXa+8w2mHPnzmXTpk0sWrTogDkDrY69djiDaTdojY+PJxwOx6yLqa3GNxQKRVuNtTqtW0qU\nV15B+fRTy10pBMyejTluHOZddx1gNJubk3nRRSjLlkFBgWWMwmEoLcU87zzIzGxuwqj/+7+I77+3\n9qPriPXrMRQFwzRxKgpKWhpy5EjEsmXI6mpLVKB7d2SPHmCaEB+P2LOnSZF2OWIEZnk5ysqVIATS\nMCxjPG1aU6eB775TWb5c4corNSu2vX0XK17chleJYF7fH98zD1H47H9YUtWHof0VVi9VCA3+AZcn\nzC5RwdbSrfRLPTBFPSdHYfy0rTy1az7JnmSqjQApnWHDRoXkroXsLplN5y7nsjD/Kz7+61hCHywk\nfv4aIumd2LJfpX9/EwUT5xdfoJ15Nn//aBgjRxpMnWqwLG8ZTsXFunUqz6xbwfO3nHvA/hctUlm7\nVuH8azfzm7lLSE/qxIvfvcj47PGoinXOEhMP+FqzqF9+YMewmmOfsZot2VbEEtONtbnUv8bBYPCQ\ndGQ1TePCCy/kiiuu4Pwmmh5kZWWRm5sb/T0vL4+srKxWjW2aJoqikJKSwsiRI6P7sxd3bb1HO5zB\ndDgcJCYmIoRod4N5NBimXTdq68E2FdRubl5i2zaUzz5rmKwjpSVVN368pe3aGmRnoz/1FMp776F8\n+y0kJmJcdRXyjDOa3FwIgfjiC8Tq1aCqiP37kQ4HUgjU+fNRMjMRigLbtiFTUjAnT0bs2mUZ39p5\nCtOEYBDZqVPTEjGKgpwyBePkk62Ypd/frJC7EHDRRRrvv+9i5kwnl6X8l73Pz6aXatLreDfizZWY\nffuSOTCBkh1BFi1KpNfAEPMT19Ip6KEqozufbvuUe1LuOeBlNn68wff7KxheM7zWwMCqTSoThpgk\nJnalU9cgQ7NX88TSf/HS5/E8vGMONekuPJkSw5C1axaFUAjEsuXceuvxPP20i4BezOakteRu7AZV\nkHbSGoqCo0n31bHMRYtU5sxRuf12jac3vkKXDEEgP4ltZj6LchdxWvfTWnd9m8Ex9nkM9XEonUqk\nlFx33XUMHDiQO+64o8ltzjvvPJ577jkuvfRSVqxYQVJSUqvcsQDLly/nyy+/RAjBmjVruO222zjl\nlFOi9+PQoUM57rjjWj3fDmcwoWGLr/Ye98eMYR5Kh44G+/jmmwM1yYSw3KNLlhxgMA86p27dMO+9\nl9ZKOquzZiH27wcpkR4PZlUVSmWldQyRiBV7NE1EYSHGtGkoq1ejLF2K7NLFmm8kgigpwWhJqSMh\noUllHhv28SgKXHKJzkczCih48lXSHRV06qIg1oOsZc2lY89h/+JN9BC5LMvbRVVKFWkDR5Gc1ond\nFbubZZmDOw1mcKfB0d+DJ9e1+zRMg0eXP8pJA1MJmR9TtEpFL4WasErvXtbcKioEFYWCgk0e1s1y\ncOedEW5+diU5YR89UhQmTjQojXhYkreEX/T9Re1xQWmp4PbbNSrU7SzJW0JGQhqpHpO9Jf4DWCZA\nRbiC5XnLmdp7aquuYWN0RPbZkdGY7R4Kw1y6dClvvfUWgwcPjpaKPProo+zZsweAG2+8kbPOOotZ\ns2bRp08f/H4/r732WqvHN02TiooKEhMTGTx4MKWlpXzxxRc4HA5ycnL49a9/zXHHHYdhGK3KoO2w\nBtO+2LHqkm0Juq5TVVWF2+1usWSk2eNUlIZF/A2/1E4zbWb4/HxLjCAxEdM0EZEIwulEUttYOj4e\n4uKQqorYsQPjjjuQbrdVciIsSW/tmmtQmmGxrZpDo2M0Tei++mOSqvcS7NwNmQgIEOXl6JVhFu4r\nJv33tzCgy3aemf0UhSWT6BTvxQskuBKaZZmNUX8Rvr5oPUXBIrondmdPxR52nj+QEauXsKlUsmWL\nQpdMSdE+ncwEeLnyVH410cBwl1Cd8C2e/G5kZEicDkhX01lbsJaxXceS7ktHCDj/fKsG9+mFr6EI\nBUUouNzQMzOOvZV7+Xz750zrW+ei/nLnl7z7w7scl3IcmZ5mXOltOLeN2eeRFv7uCIgll2xj2CGh\ntmDs2LGt6pzy3HPPHdKcJkyYwIQJEw74vLS0lJR6uQytLTfp0Mu8WJWza2mscDhMZWUlPp8Pr9d7\nyA+QHDnSshK14gaA9XsohBw//pDGbA2EEJjJycjaF6gQAsU+BkVBdumCPP54ZPfuVr1kOAx+P+Y9\n96DPnIn+zDNU/fWv6Gee2W6GXddh5kwnWflr6NTLi8MBubkKUoKMj6d8WwlD+gQYebJghdiLzAyS\nlB1g6fpC8qvyCRthfij5odWi5WCxy8+2f0aKx3pw03xpfOrNQV4wif6phfgCBcwr3onHXcRn2b/h\nV3/OpksXyd/fyUU3YMTpu1m/J48Fa/P4fv/3bCvbxvbS7Q32kRvIZf6e+ZjSpDhYHP0pCZXw1DdP\nRe+x8lA5X+z4Ar/Lz3+2/Kddzml9NCf8Xb/tlF0IbyNWjEOszCPW0BTDjDXhdTtL9+mnn+bdd98F\nrEbSJ598MjfffDMFBQVtGq9DMkwbPzWDWV/ntrl4ZVvmJXv3xrzwQpQPP6xzyxoG5qRJyCaUNNrz\n+LQTT8SxaRNqZSXC7bYMtaZBWlpdKYiUiKoqzHpp5mL/fpS33sKfk4OiqohRozAvu+ygbteWICW8\n/bYTr0un+3EOxA6VzGSd/P1O8vIUsrsaZCRrJE/qg4Fl2C7sZxXraxrUL9/1OlvvkqrPLgF8Th/F\nwWKWXj6V1J7nsejlhbx13Oekhi/jj7dfRpcuJjNnOvGVjeD9m0/A44Gyk+CpZxRq1NcY0quIrPiG\nyRCp3lQenfAokrrrtmlXBasq/ovf42RXxS56JfXio41fUVFp0i8jk+8Kv2NXxS76ph3Y/PlwsG6d\nQs+eJgkJdewzEnGxZYtg8OBI1H0LRN26x1CHWD8fsWgwbYO+dOlSrr32WoqKivj222957733mDFj\nBnPmzOGKK66IJge1hA5tMH9KaByvbJcYkBCYV16JHDUKsXw56Dpy1KgD9VbbEXZsq3rMGBK3bLGy\nV/PzLSOZmYkoKLCyXktKrHrHIUOQJ51kfTkvD/XJJ5F+P2ZWluX2W70aCgsx//CHQ+4PpSiCCRMi\ndO+uIMsGIkM1qPv2keWDSFBHFBrIzM4YtTHdQemDGJQ+CLAaKAshcChte5RMafLZ9s8IG2H2Vu6N\nfq6ZGq+v/C/KwkfxXfkD5dsGUhTeyDMv1PCXhzxkZ5tcfLGBx2Mda1oqnH/NFt79top0fzrL9y4n\nOyE7+qLwOX1M6NbQJVUc+Iz08oF09rtYuGchajiZjzfNoltqJ4QQuB1uPt7+Mfel3XdI57M5xMdL\n5s1zMHGiTkKC1Vlt3jwH/ftbajH1dUvt5CFbPMF23x6LfbZ/7sWhoimGGWu9MO35xcXFkZeXx9y5\ncxkyZAhDhw4lHA4TfxCVrqbQIQ3mkYphHimGaccrbSWWQ9G5bXZeQiD79UO2QrX/cI9PSklVbacR\nz/DhUFEBs2ZZtZEAfftinHyyJeCuaZinnIIcNSoqiqDMnYtUFEvEIBIBVUV27YrYvRuxbRuyf/9D\nmpMQgu7da6tbzj4bdccODF1Hzc/HLTQwJMYJJ1j7bPRC+Hjbx6hC5YJ+F7R5v+OzxxPSG3YF2b1b\nYdZKN5NOyeONnJVMO60LO4vy2blwLg89dAEPPxxuoOqnmzobAksZ1j8Zn9NHbiCXvMo8shOyaQpF\nwSJ2VP5A3+wsivc72FqZw9bd/8LrM/B5LKqc7k1n3f517Czf2a4ss1cvCRjMm+dg1CiDVausspm+\nfevcsE3plqqqGm0mbSvLHIt9xh5ikWHaC6xLLrmEd955h61btzJjxgzAegZTmqjJPhg6pMG0EcsG\n00Zb+3LGKgzDoKqqKsoSFLvsY+RIq3m0y4Xs1cv6t5kxxK5dTeq2CiGgqAjaaDCbetnKnj0JTZ+O\n9/nnISkJMzkZc/BgZEICjlmz0C+9NMpki4PFfLPvG4QQTOg2gVRvaqv3rSrqAcwPoCIVxnXWefjj\nfzNsiCApQeEEfydcrtmcnzKeefOSuOKKupjzjrIdBCKBqIFMcCewLG8ZFw+4uMnjW7F3BW6HG69H\nISFBUlTgYXH5u/RJ7U5hdZ3cWFAL8tHWj7g/7f5WH1Nr0KuXpLraZP58lb59GxrLpmDHPht3zaif\neXuk2WesdOCItVhq4/nEYvNo+508depUJk2aFBUoqKys5M9//jOZtXXirb2+HdpgQmzHBeyswrb0\n5WwKrTLkFRWIhQtRVq8Glwtz7FhLcq5egM4Wq28rGisQBQKBuj8mJyMba742A9mtm1UK43QipLTY\nJrUdpVJbb6xaglpejnbaaVaNZz2IvDxEfn60Q8n8PfNxKA4kkoV7FraJZUp5oNdbSktEoFIWkDx0\nMd2TugHgVJ0kJZro2XP55cS6rFbd1Fmat7SBoU50J1q6so1YpilNCqsL+b7oezr5O1EaqKGkTJCS\nopBUmUX/xGFkp6ZFt49EItEG2u2JqirYtUshOVmSl6fQt6/ZbPi5KXWXxk2P7WfkGPs8+ggGg6Sn\nH6g2dbQhhKC8vJxvvvmGXbt2RRsK+Hw+zjvvvDaN1SEN5pGswzwUg9IYdgcVKeUhN3tuE6qrUWfM\nsAr809PBMFA+/BC5bRvm9dcfVjzTZsj1FYgOlYnLlBTU5cvBNHG6XJg9e1oC6127tsql3FqIQADZ\nRDcTUwgqKotIoGuUXWbFZ4GEFftWtJplFhYK3n/fwbXXatESEynhvfcc9O5tssk1j6BeTWmoNPod\nVVGZvWs2p3Y/NdqdJL8q32pUrVlZrtF5SpNNJZsaGMz/7vgvm0s2kxWfRTAoKStVyOhk4nbHcUHc\nVJKCw5iSPZSk2paewWCw3T0a9WOWffua7NwpGsQ0m0JL5VJHm312VDT1/MYiw7RZ8DPPPMPGjRv5\n5JNPOPvss1m3bh1CCCZOnNikFF9z6JAG00YsumTteKWdJdgeD3pL8xKrViGKiqwyDhs9eiDWr4ed\nO+FgerDNwE7WiEQih82Q7Tmq77+POXiw5ZoNBFDXrUOOHo1xxx0tJ/wEAlBcbP0/Le2gWbVGdjaO\n9esbbiMl24z9fJD/Kbf26Rdll6pQQVhtwVrLMjt1kmRnS154wcVNN0Xwei1jWVws+MUvTIr3pHNa\n9mkHSByqQkU369yxXeO7cs3ga5rch0ut+24gHGDO7jnops4vj/8lwf1dSB8oCQQEWVnWfVFdbRly\nw5AcgrpZq/DNNw1jlnZMc/lylSlTjIN/uQU0Ffs0DOMA9mmLJrRlsRwrrtBYmUd9xHrSj33O3n33\nXTZt2sTEiRN5//33UVWVCy+8sM3tvo4ZzBhyydo1aT6fD4fDQWWjTh9HCmLTJmRjQVEhrDZWeXlR\nAfXWni87uUdK2W4ZvcpHH1mKO4mJyG7d0CoqEIqC027PEwpZ7cBWrbJcyqeeCjbr3LYNZe3aqAtX\nSIk5aFBDsfh6MAYMQN20yZLs8/ksgYVABV+ll1FgKszLmVfHLmvR2de5VSxz+XKV1FTJ2WfrfPGF\ng3/8w0VCgmT1aoUnngjjcMCkHpMwDANPMz07bQghiHO1vDpetGeRJdDucPHVrq+4+oSryc8XPPmk\nixtu0BgwwMTvt9YHjz/u5sILNQYNanHYNmP8eOOA1qi9ekm6dTs8Y9kU7Dh5Y/YZDoePsc92QFPG\nOxaTfuw5xsfHEwgEUFWVuXPnMnr0aDZu3HjMYLYVscAwm2JjhtF+L5EW55WUBLt2NTWxA7JCW4Kd\n3ONwOPA100arzefJNBG5uZYCkDUAonZesrISsXMnyocfIubORQSDYJqozz2HccklmDfdZBnL9PQo\nC5WGgbJ+PYrXi4yLO3CO8fFoY8fievVV1A0bQFXZPCybvT2d9EzuzUdbP8Lv9LOvsmHfPl3qbCza\nyPhuzYs+ZGaavPmmk/JywaWXasydq7J5s8qvfqWxYYPK8uWW9mt7IRAOMDdnLhn+DFRFZXX+aqb0\nnEKXLp35zW80/vEPJzfcoJGeLnniCRdnnKEzcqTZUovPQ0JzTobDdD60iCPJPo+hDoei9HOkYV/L\niy++GF3XufHGG3n88cdxuVwMHz78WFlJa9A4htmero62Gszm2NiPyX7N0aOt2GAkUtfXsrISPB5k\nvc7nLc3JTu7xer2H1jatogJlwQLEd98hk5ORkyYhBw601H86dbKCYPVvcNNEGIblUl6wwNqfzRpD\nIdQPPoCUFGQ90XbAKkdxOlEKCizh+caoqcHz3nvgcGBOnoyJ5KvqeaSs1XCn9qaLvwvju43nlKxT\nDviq13Fwf2b37pKrrtKYMcPJffe58Xhg2DCTlStVli+HBx6ItO2ctYDFuYsxpYlTtZK36rPMvn1N\nfvMbjf/7P+uaX365Fu2J+XNGS+yzvmRfLLHPWHLJ/lQYpo177rkHgOnTpzN69GjKyso44YQT2jxO\n7NwNRwFHIumnLTAMg4qKChRFIT4+/og9nC0a3549MS+5xCrN2LPH+tE0jBtuaDXDDIfDVFVV4ff7\nW9S2bRKlpaj/8z8o770HhYWIdetQ//xnxKxZAJjnn48oLrZk8sDqpbl3L+aIEYhvv0VEIhAXZxn9\nykpLOUhVUWbPbnp/QlgCCbVzD4fDUWk2ddMmRFkZZGSAorBdlLHXb5IUUVByc8mIy2DlvpUoQiHO\nFdfgp76YeXPo1k3Stat1PYJBQVaWyZ49CikpkuTkuuukadEpNoB9ClqCHbvM8Ndlu3bydWJ1/moK\nqixJsJSUuv1lZsZOeAJ+nAx2m3263W58Ph8+nw9VVdF1nWAwSDAYxDAMTNOMqfBNLCIWY5hNoWvX\nrodkLKGDMsz6qC9i0F5jtQZ2vNLr9TYZq/qx46tyzBiMoUOt/pIOh9Vzsr7mW3Pfq+dOblGuzzAQ\nO3bgKC+3xArqJbUoX3yBKCmpc7sCUtNQ33kHfcwY5NixGMGg5XqNRBCGgT52LI6rrkK99VbrC/n5\niPLyaFavBEhLQ0Qi1rm0r7GUiEgEo1MnQqEQRn4+vjlzUFevxvB6kQkJmIpiNcBG8iVbScIDLgPK\nK3CrbjRD49uCbxmXPa6NZxpycwVr1qhMm6azcoXCrlnb+NPxy3EUV7Lpb73Rhg9j6Gk+5s1T2bfP\n6tE5b57KiScaBIOCf/7Tyd13Rw6qBrhwoUplygbCRjhaX6nrkJ8v6JSps7pgNaOTzuWJJ1xcfrlG\ndraMumcHDDj8TO/2wo/NqJpin7bObSQSOarsM5YYZlOIRZdse6PDG8z2RktGTkpJKBQiFAoRFxeH\nswWj1B4PSauNr9/fwAXb0jhtSu7ZsQPHU09BSQlew0A4nXDddcjTT7fGXrkSmZbW8Du150Zs3448\n6SRL6ODUU6G4mBqnExkXh8Pns4TiFy1ChEJ1OrSGgaipQWoask8fa4zav4maGsxevTCSkxH795P8\n5JPIQADS01EiEcTy5UjDINK5M0UiSIlSTUQxCBhVSG8yZiAXKSWbije12WCGQvDFFw4uvFAnPV1i\nzF/K8ZWL2LAqlXGTvPQWG1n83Ba+1S/m9DM9vPKKwt13u8nKknTvbjJzpouLLtIaGMuqKnjhBRe/\n/nWExET4978dGAZ8+fo4Tpl4PNOn6+TuEfzxITcXX6wzZZzOwjmJPLbYxZln6lE37BVXaDzyiIv/\n+Z8I9dYtHRY2+1QUJRpisCX7Onrss6n30jGD+TNF/Qvd3nJ2B4OUkurqagzDaLG+MtYfvtYk90QR\nDOJ49FGL8XXtiqlpKJqG44UXMDIzLSPt8VjWpDGkbMh03W4r7lhTY/0NMH/xC9THH7cSgMBikqZp\nZdXGxSFTU5FZWYi8PAD0zEwqvV6ElMQtWWIJvNdq0wq3G+P443EsWIA7N5cuvXrxB/NUZFExUtep\nmXgnSmpqlGm0FXPnOsjMlKSnSz79Vw1/HLKYcGom8xa5WLBMkJnppVv2Xr547ntE6mTS0yVbtyoE\ngyb33efhvvvCDBnSkAH6/dCjh8ljj7l54IEww4cb/O53bmpqBDs3dGL3QI0HHnBz+ukGl58fQUoo\n3OtE12HcOMtYlpfDRx85Oe00naSkY67HptCYfdZPHKof+3Q4HDH//B4JhMPhFgnATx0d0mDWx4/V\nkqu+NFxrmz23l7u4vY7RHqd+ck9LpQ8AYu1aK65YS1sEWBJ4Hg/iq6+QAwZgTp6M+uqrSL+/znUa\nCIDPd3CNWCmhshLZs6c1fm6u1ZQ6MxPZrx+istJimt27IzMy0HWdyspKPG43kUgE9fvv63pyCmGd\na7cbo1cvFLcbNS8PFTCzs9GmT8ed0dkqjg/XddVoi7LMlCk6ug7vvOPkriv3kbgUzAyFCy/Q2b9f\n8NVXDpxdUunv2Mozz5yF2w29ext8/71KaiqMGnWgu1QIuPBCqz7zscfcjBxpEBcHKSkmnTub3Hqr\nhwkTDO65J4IQ1vY33aTxyitOZsxwcdVVGs884+Lkkw3OPtsa50hkyf5U0dQzKERds2yggWD8kWKf\nseSSbW4usZQkdSRwzGC2c6ywqbEOO3s0hmB3TWmNO9mGCASibLABvF5EUREAcuJEzI0bUVatst7o\nUoLXi3HvvRarbDymEMjiYtTXXkPs3AnFxYjKSuTAgcg+fawxNA2qqqJSdnbc2O/z4aquRguH0ZIS\nmBP6nsnKIAQCU0qkaYLbTfjSSzH79LEy42rl+xzUtZ6yX5K2skxrGIbDAe++6yAz0ySjmwu51Pq8\noEBh7VqFK67Q+GZeiLxIPHv2CKqqBD6fwqWXWqpAM2c6ueoq7QCdBttozp+v8vzzTk4/3WDUKJ0H\nH/SQmmqd+/vuc/Pgg2FSUiyh+euu03jqKRcPPODm3HP1qLE8hrbjGPuMfa9Ye6BDGszGF/ZIuWSl\nlITDYWpqatpkYOqP157M8HBgx15tub62uCNl9+51RrC+OzwQwJw40frF6cS8807ktm2WulBcHHLI\nkCbF1msnhOcf/4D9+y2DGBeHWLTIUidyuZCJiYiyMsxf/hLp9xMOhaipqSEhJwfXG28g8vOJNwyW\ndhM8599IikxlNNno4TCUlSESEtD69wePB11KFF2PvhShYW2fy+VqkmHUL4yvf29Mm6bz4osuMLM4\nOzWVwh/K+GZLOhMmGDgUk0SthLlVF9G3r8nGjSpSSkaONBk1yuCVV5y8+WbTRnPePBWXC849V+f9\n9x289ZaDSZMMQiHB5s1WFq6uC2rToQgEoLTUmteOHUqDqqJYQCwxqragNeyzuXvjp4LG10ZK2SGy\niH/e/LkVaM+btX5dpx2vDIfDJCQk/KR9+zartN2WbY3dyX79kIMHI3JyrNijla4JcXGYkyfXbSgE\nsm9f5NSpyLFjLSO4cCHq3Xfj+NWvUJ58EnbvtjbNyUHZs8equxQCkpMxJ0ywYpVbt4LDgfHLX2Kc\ncw7BYJBwOExiSQnu//s/qKrCzMzEzOzMu3IdflNlZvgbjDmzcX72Ge5VqxBZWXh0HZfLFX3xGYaB\npmnR8pP6usE2u/B6vVHdXHuREQwGCYVC0f6O8fFw440R1m90MmPvdL7ZEMfpfXNIC+0lf1U+C/Rx\nJJ3Sh7w8heOOMznzTINXX3VQUCC47jqNxER5QLh37lyVzz938MADES65RCcUEhgGJCYOCHCYAAAg\nAElEQVRKgkGJyyWZPFnn9dedmCbs2CF4/HE3Y8YYvPxyiIQEyYwZLmr7N/9kjdWRwOGei9bcG5qm\ntSphMNavSazP73BxzGAegfIN0zSjHTlaLLU4CGKBYRqGQSAQQFGUlpN7moOiYNxzD+YllyDCYZTS\nUoxRo9AfeeSgXUaUd99F/dvfEKWl4POhrF6N4/e/h507EZWVRANyNpKSkN26ITUNUVKC8q9/Yd57\nL+TlkZCQgOPLL5EOBzIhASkEqz0l7E9Q6BbxUuDSWNkvDm3CBGrOOANz1y6URx9F1jYvdrvdUeOp\nKErU5aZpWrROz4bNMNxuN36/H6/Xi6IoaJpGdXU1S5ZofPedyZgxGjvKUsk/61c4rr2M/6jT2X3O\njeT0HE9CItx6q8bUqQZZWSY9e0p27lRwOOCCC/QG5bFVVTB/vmUsMzIkn3/u4E9/CnPHHRrz56ts\n26Zy++0Rtm9XuOmmCN9/r/DLX3rp08fk7LP1qHu2sFDwt7/9dBd2PwU0dW/YdZ/V1dUEg0EikQiG\nYcQ0Y2survtzR4d0ydbHkTCYgUAg2srqcG+io/nQNE7uOazibY8H86KLMC+6iGBtRslBVUHKylD+\n8x/o2rVOO61zZygsRPn3v+GqqywXr2GAqlJMkC3VOYxb+p312aZNSMPAuX07rrIyjBdeQOzciVlr\naUxM3vJsIUm6kaEykkx4e4DGqHBnXChInw+Zk4O2fDmRkSNxOp1RN5vD4YieC9vdBtbiQgjRwHUL\nFsOwhdSllPTrZ/Loo07Ky+G22wLMmuXlnr/14OSTDdatUBk2LIIQJuefr2AY8PbbToYONVi7VmHI\nEIPGsr9xcfCnP4Wxd3nNNRqlpVYnkPvvj7BggYNnn3XxxBNhgkHBJ584OP54ky5d6oy87c694IJj\nccwfE83FPkO1LgTb7d+ezRiOBAzDiNm5tSc6pME8Uish+yZvbfZoS2iveR7KoiBUG/M7lNhre8xH\n5ORY/2ksNJqairJuHTI1lcipp+KbOxdqaviPZwPz/PsZWCVJSc3G8PmsTMVQCFasQF+2DCM7G3X1\naqTfzzeOQgpFkK66H6FpJPmTyBFVrFLyGW1mIRQF4fHgKyrC6fdH40/BYDBqNO3WUk6nM2o467/0\npJTR7Nn6sc/9+5243SqZmVBe7sHrhYoKE683xC23GLjd9grei8NhSdZJCQ6HQXO3RP13VWmp4PHH\nrRrLiRMNJk0yePBBN7/9rYdBg0wuu0wjK8vk6actI+50wsKFDu68M9yebUWPoY2oH/t0u91RyT77\n3oO6ZtZHO/bZ2HjbTSN+7vj5LwlaQHslxFRXVxMKhbB79MXK3NqK+sfSOPb6o87H7286szYUwqZY\nkenTkampFORvY2FyBQ4Jn/cTVusv07QeaJ/PctGuXIlx1lmg6xiBcma6N6OZOiWRMopS3BS5NAwk\nbzk3YWAxL2kY0KVLVDotLi4uek7sMqHKykpCoVA0E9LlcuFyuXA6naiqGjWk9V23BQWCW27RuO02\njTlzXJSUOJkxQ8c03Xi9KlKamKYRdc8JYeBwyGaNZWNs3aowdaplLMFKMr711gimacnqJSVJkpLg\njjsifPKJkw8+cHLHHceMZVM4mh4e2zNhxz5tA9VUXPxoIxZ7YR4JdEiGCQ1rHA/nhrMTYoQQJCYm\nEggEYuIGro/WHmP9Y2mvtlzNocUEh969kV27Ivbvh06drA8NA4qKMG680WJteXmIcJhPzj0OVfHQ\naXsBs3tWcPY+k7TKSkhNRZomolbmjj590O67D+2fLzKsUGWgIwGZ3Q3zhMGo8+ZClQOvJx5Dmqil\nZeD3Y554YoN5CSGiRtFmk7quRxt+12efjbtj2IlC48bphMOCt97y0q+fyf79go0bVS64wACcaJrV\nF9U2zPXdc3Z25cHYxSmnNBSgLSgQvPuuk7/8xXLbzpzp5MorNTZtUnC5rOvwzTcqZ54ZW8LrsZTk\ncrTnYdcI26EBm31qmkYoFPrRM28bX5tYFl5vT3RYg2lDCNEgYaMtsJs9u93uaLzyxxJCaG8YhkFl\nZWV0RXuwB+5wX2St+q6iYNx3H47HHoO8vKh6j3nWWcgpUyzpu/x8CtQaFih76GL4UeMTEeUBvuge\n5uodQavHZ3U10u9HjhplZSb274/2v49xVU0NitcbZaui16U4XnkFsWs/kI/s2hX917+G+HgrI3fJ\nEtA0zBEjrA4qtdfaNpB2jNfOoq2pqUFV1egLzmbqpq4TLKlm5rvxZHTROfdcjfJywWuveTBNwZgx\nMnqO7LFt46zrerTms7X9HAsLBW+84eScc3SOP96+z3UeftiF3w9//GMYVSXqnm1sNINBKC4WdOvW\n8D7csUOQnS1jqgylI6FxXLyp2GdrFlfthWMG8xgOCjueZaeJxzJa25bL5/PhbkIkoP44PyoyM9Gf\nfhqxZQsEAlY9p92+yzCQ8fF84stFMSQqApGaSkagnNndqjg7Vyc1FEKmpCAHDsQYMYKamhpM08Qf\nH4/SKHNG9u2L9pe/IAoLkUJYnUqEQP33v3G89FKde3jmTIwzz0S/666GgUOIao663e6ogbOzH4WU\neGfPxvPZZwT3VHORJ43O916C4RxPaqrJNdeEeOstN/37h/D7G2ZI2gux+sxW1/VW9XOMi5P84hc6\nffvWLQpV1aq3rO+GveOOCE8/7SI7WzJoUN22u3crzJjh4p57wvTubc1p40brs/vvD9OzZ2x5Uzoi\nmot9Hkn2eYxhdjAcqktWSkkwGETTtCZLRn5KDPNwhRUOFW06LkVpVhB+b1YiXybsJy6iUOQNIUyJ\n7JpKRUDjs7EZXB3ujzFsGPrll1MtJYoQ+P3+5l8YioLs0qVunjk5OF56yRKFt8+NaaLOmoU5Zgzm\nyScf9BjthCCPx4PyzjtW55X0dPz9OxFfE8TxzNMIaaJOmkSXLnD99SY+n0I4HImWoQBUVyskJAi+\n+cZJcrKVZWuPHQpJPv9cZfLkGpzOA1Vl/H7RwFgCHHecyWOPhYmLq/ssKQnuvTdC43fewIEm118f\n4ckn3dxzT5hQSDBjhovf/jbSIYxlLLmFWzuXo8E+O4LwOnRgg2mjLS/v1sT4fgypvcMZq764QjAY\nRNf1NtWKtmc7tMOBaZpoSCaPuxbHooUo67dCRYX1t159SL/saiInX4HhchEMBnHWrr7bMm9l2TIr\n9ll/IaEoSLcb9auvDmow60PU1OD85BNkdra1KJES0xGPDsg336Rq5EgcLhfvvuulc+cIU6Y4cbut\nF96cOYLNmwW//nWItDTJzJkeLrjAZNAgS//h1VddpKVJEhLc1NRY5+WVVxxcckkQj6dOVWbFChea\npnD66SZC0MBY2mjufXfiiSYQ4f/9P8v78OCDkZhqAXYMzaM5177NPu2ylcNlnx0lS/aYwWylgbPj\nla2J8bXn3I7EOI0N/9E2fq3Cjh0os2Yh9u4l0rMn4fHjyeySyQ0T7kT9JBdKIsjkZEsdqKACXppL\nTd8zqU5IwOPxHJrbXNMOcLsC1me2JE4tiouhcXcyXbdEBZIrihp2XRECRVVRkpIQeXl4DIOIaTJt\nWgWvv+7D4YApU0wWL3axfr3K9ddHcLtd9OplcuWVIX7/ey+GAWPGWKIGF11ksH274NFH3fzylxpZ\nWYKXX07g1lvDuFwGixYJvvpK4eabqwmHlTaJxduoXyVlJwodw08L9V37UMc+bQMKrWefjRfNx7Jk\njyEKO17ZUowPYtcla49lmmark3uO1JyiY9TUINavR2zahExMRA4bBt27H7j9ypWoTz2FdDrRvV7E\njh0kLVpE5b33WlZp925kr17RsUlPR+blYX76Kb4bbohK27UV5kknwRtvWCzTNpxSImpq0G0NXKwp\nfPaZyuDBspaNWZ998olKcrJk4olJDUQWogiFrLIXjwdd00hP93LLLYKXXlKZNw9SUzWuv74ar1dF\nCMsF268f/PGPOldc4WHXLgfz5lWwaZPgscd8uN0mvXrp9OwpAAfPPedm+HCDxYsd3H13mNRUVwOx\n+NYmDtkxywcfjBAMEnXP2jHNI4VY8GTEUsZ7e5+P+uzTlvNsin3agvEH2/exGObPHPbFP5gBkFJS\nU1NDJBIhPj6+VS/eo1E72VpomtZqw3/EUV2N8s47iP37kQkJiF27UFaswDz/fOTo0XXbaRrqP/+J\nTElBq3UpuRITkUVFeN59FzlypGWIqDv3hmkivV68u3ejH6KxBJADBmCcfTbq558j3W7LHVtdQ2jQ\nMJTx46PbORwwYYLBvHkqNTUKJ5xgMmeOitcLp55qgpJIYMRpJCz/GpmVZRlNXUcpKCB0+eXUaBo+\nnw+Hw0FiIvTvr1JUpNCvn0JysolhWIXrVka3k4ULfVx6qcEHHzg499wEuneXeL0m999fQ3a2iaZJ\nzjnHYPFiL59+6uDeeyO1lTl1YvEtJQ7Z2LpVicYs69ywVkzzd78LH5A9+3PE0TbaPwYOxj5ramqA\nhuyzKYbZ2U7I+xnjmHBBMwbOdlvaMb5DZSlHYm5thT1GMBgkLi7u6BtLw8D10UcoK1ZYbk+3Gzp3\nRmZmonz+ueXHtLFvH7Kykkjtg+xyuSyR9tRU1K1b0bxeqI292A+5NE2cmobMzj68eQqBfuedaI8+\nijz5ZOTQoeRd+3tmDnmcsuo6F29OjiVDN3mywZdfqtx4owshYOpUA0WBLVsErzlvIDhuMqKwEJGf\njygqoub886k84wyk9LN8uTXe3LkKW7YI7rxTY9UqlQ8+8OL1+oiPj0dRvLz+upuEhDC33FLKww9X\nkpursG6dyn33mfTr54rq3S5dquJ2m5xwgsa//qUQCJiNDk1EE5L8fn/0nrC9Kfa5zMgwuPvucIOY\n5Yknmtx2W4T09J+/sYwl/JiM22afHo8Hn89nJa7V00MGGjQiCAaDeL3eVo9/7bXXkpGRwQknnNDk\n3xcsWEBiYiLDhg1j2LBhPPLII+1yXIeLDsswbTRllOyaRKfT2WbB8VhjmLZyj9UlI/6wM2EP+/hC\nIVwvvoh47z1wuRBFRYi1azHHjrVKRkwTkZsbzYw1HA6kpqEADqezrk2YriNcLmpOOAE1IQElPx8z\nJQWhKDjCYaShY5511mEda+0BY44ejVnLejOAET8IPv5Y4fzzDQIBwddfq5x9tkF6upWBWlFh9bac\nMsUgL8/6+/TLJI7ONxMpvxxKS6mJj0f3eIjz+6msVJg3T2XJEgWnE264QWf3bkE4DJs3K8yeLTnj\nDJN589x07gzTpwu2b3fw3nsORo7U2LRJ5YEHBK+8EiIpycHy5S7mzHFw110RUlIMPvpIYcYMJzff\nHCIujmhyR1OtyoDoC1DXdVQ1TNeuKpFIQ9fcwIHHkn46Cpq7P0zT5J133uHJJ59k8ODBSCmZOnVq\nq1yz11xzDbfddhtXXXVVs9tMmDCBTz/9tN2Ooz3Q4RlmY0QiEQKBQFSO6mjE+NprLDteaes+xoJr\nSSxejLJ5MzI1FRISrMbMHo/FNvVa4e/aB1PTNAJeL/Tti6O83GKWtWo5orAQTj2VuC5dMP78Z4zM\nTNSCAkR+PpoW4YnLevK9P3hEFi8DB0pGjTKZOdPBJ5/YxlLyyScq6enwyCMaui753e+cfPKJyvTp\nerR8VCYmUp2RgenzERcXh6IoJCbCHXdo7Nih0L27ZPduwZtvOrjzTp3779ewF+5TpxpcfLHByy+r\n3HabG69X4a9/NVmxIozTqTBpUjxbtgT5+muDm26qJDXVwOFQuegihUGDBMuXew6Q69N1/QDhDtuQ\n2uzT1sqtqamJtkqLFUm2I41YiKPGGuzz4fF4uPrqq3nnnXdwuVx8/PHHZGRkMHXqVJ5//vmDjjFu\n3DiSa5uyN4dYvL86LMNsHMO0+9OFw+FWxyuPNA7HYDbO6rXbjR3NOQEoS5dipKdbaq1bt1oV9G63\nJSmTmwt+P7J796hrMC4+HuW88xC//z2sXg1+P6JzZ+SgQRiXXoqu6wSTk3E/8QRKaSkyHGaDu4IV\nKx6jaOO73Ou9F5fTFVXcaa+XX/0SDKfTMpZeb50bdvp0g8cecyJEXQ9se2WuKMoBCVeJifDXv0a4\n/34Xc+eqPPCARo8e1nkeP96s3Y+1bffukupqwfjxBj16SPLzBf37C1wuyYYNifzxjxq6bmmOGoaB\nw+Hg7LOdtfGn5sXigQMWVo0TQ+zvtTVx6BgOD/YzFwvGu/4iQlEUBg8eTFpaGg8++CC9e/dm7ty5\nbN++/bD2IYRg2bJlDBkyhKysLJ588kkGDhzYHtM/LBx9qxAjqKqqQkp52BqqseCSjUQi0booOzYV\nC/MCLFeqqmJ07gyBABQUWBmolZWIykqMG24gWPtCjo+Px7l69f9n78vDoyizr0/1lnSnO4EgBAIo\niCjIIIuiiMMuOySBEQkBGUGcICoKIqiMOrgN+KmMooPLoPwYBYEkQNgyLmyiAoKyKKAICBjZIUl3\n0mtVfX80b1Fdqeq1qqua9Hken0cjVL1dqX7Pe+8991zo3nwTbHY22MxM6CoqwFIUfOPHw5OSAtfl\n2onRaPT/GZbFmm/noLGtMc64zuCE8wRuSr0JnosX4d23D4aqKuhatIDuT3+CLoxa7s6dOng8wJ//\nfCUKO3CAwrx5Bjz1lA9uN1BaqkebNgy6dmW5muU33+jxz3968MMPenz6qQEjR7pBUTUwGo2S/aDH\nj1/52U8/6dCihbi366BBDLp0cWPhQgMWL9bj6FEdRozwoUMHFjQNGAx6GAxX0mdXlI9OjuD4Bwgy\nqoz43ZL3hIxsEkvdRuo4FA2S0V3igLSVZGRkYMSIETFfr3Pnzjh58iQsFgs2bNiAvLw8/PLLLzKs\nNDbUecIk6SiKomC1WmP+gqqZkiVRssvl0kyULARz++2gysr8Ip+OHYFLl4Bz5/ytGlOmwOl0wnfh\nAtKbNoWOpqG7rJBlL0dkDACcOwf2v/+Fa+pUpKWlBZguHLpwCL9e+hXXpV8HHaXDqsOr8Oy142Cd\n/zZQXQ2aogCvF97sbDgmTYKhfn0YjUbJDb5dOwZLl/qfY5cuDFau1GPDBj3+8hcf11bxxx8svvpK\nj5tv9uHs2cs1y8tp2FataGzZwmLxYgZjx6bAZhPvB923z5+GfeopL+rXZ/Gvf/nDySFDxEnzmmuA\n4cNpvPeeAY0bs+jUyb8W4VmPuL7wzeKJWppl2YA5n3q9HjRNcz64/OiT1Dz5h0m+mxGJPn0+n2p+\npknEB2IHGSIolAs2kpYBMGjQIEyePBkXL15EZmambPeIBtrbUeMEiqK4SAxAxOIerYGIe2iaVtSy\nL9brsL17A3v3Qnf8ONCgAeB2gzIYQLduDWbBAqTo9bAZjWDuvBPsn/7kN0/PyAj43fjq1YPuwAFY\njUboeJ+TZVms+mUVbCYbKIpCvZR6OFF5HD8vehV/0lmB667jivapJ05Av3EjnCNGcOQhjLwAf+p1\n9Ggfli41oKoK+OorHa6/nsGQIf6D1vHjFE6epNCnD420NCA9nQ2oWXq9XnTq5ES9emmwWsXdlP74\nw0+Wjzzi49Kwjz/uxb/+ZUSDBiy6dq0tsDl1CliyxIDcXBrffqvDZ5/p0L9/cCEOP70KgGsbIOlv\nUt80GAxcypg/aSXUnE8SfYr5mfJ7+hIpdauVKFcr65CC3MYFZ86cQaNGjUBRFHbu3AmWZVUnS6AO\nE6bb7UZ1dTWsViunIpUDakSYCeXcY7PBN3UqXN98A+Px40BmJuhTp+A7fBjUtdfCaDIBLAvd1q3w\nOp3QMQz4n8ZbUwP9iRMwnDgB3UsvgenZE0yPHkBqakB0CfifX7pPjyL397g5MzdA4cZmZ8O4YwdQ\nUABQVC3y4JNnWpoOw4b5sGiRAddey+KWWxh89pkObdqw2LxZh0GDaI4g+a1obrcbbrcbaWlp6NBB\n2nqwSRMWTz/tDZhHmZEBTJ3qhdgc8lOngPfeMyI314dOnVh07MhgwQL/VzkUafLBJzhiEq/T6bj6\nNz/6FEvdAv5aOYk8+UQo5mfK7+kjkWcy+kw8SEWYkXjJjh49Glu2bMH58+fRvHlzzJ49m3MbKiws\nRFFRERYsWACDwQCLxYJPP/1U1s8QLYK+qawmil7KgAgXdDodKisrkZaWJksKk2w8GYJpGEpdK1zL\nPrIBxtqDKcd1SNtOvXr14L14EdTs2dBfdx0MlzdYhmHAut2gKitBeb2gTp0C07AhvC4XTD/8AN25\nc2BvvBFsu3bA2bNg27WDb8oUzN+zAN/98R0sRnLSZYE//oB3/x78/WQr3FivlX/iicnkb18pL4d3\n/vxaeUz+pHv/Jq9HSYkF119P4cQJPW68kcXBgzrQtD8tmp0d+DUhqXGfzxcw+DcaMAywdasOXbsy\nHHl+/rkORiMLs5nCHXf4iauqCigu1qOggEakvxqfz4eamhrOQlBomSac80neMX70yd8qSHZD7HPz\nhUNEoSslHHI4HFEp1eUEGfydKnZyiSNInVgLbjrkveD3XQ4cOBBfffVVQmUPgoGSeOnqbITJTycB\n2pQwh4owxcQ9iQS32w3X2bOoZzJBd3mjJv/AZALldsM3aRJ0r78O9rffYLp0Cfpz58A2a+avf5pM\nQIsW0B04AOrgQdzb9l4MbnWl91L35ZfQf30G1JF6aO6tge7EPuDYMdA9eoC6dAlM586iXrH8up/d\nzmLVKh3atPHgtttc6NAB+M9/rKiqAtq0obB3L4WsLJZzvCPuUAzDxEyWgL/t1OGgsGiRAfff70Nq\nKtC5M4MPPzSgZ88r0WR6OjB+fOQDoMnsTiKc8t8z/Dmf/OiT/EMUt0Q0JJW6NZlMAeRJhEOJlrat\naxCLMENZ510tqNOEKfbvclxX6ZRsNC0wqtYwz5+Hbs0a6LZvB2s0Ar16gbnzTjgB2K67DrqUFLBu\nN9jL7Q4URYFyOMA2agR3kyZwPf88rEePQrdsGegmTYDWrf1M4l8QYDJB98svyGrfHllpWf6fX7gA\n09pvwDZuA5gaQ7dnD6D3/1y/cyfoW24BnZMTcukHD+rRvj3QrZsRgBHnzjGoVw+w2bzo378a331n\nxrp1egwaBBgMlH/2JRVijFgEoCi/8GfdOj0WLTIgJ4fGJ5/o0bMng9tvj808wOPxwOVycbZ8Ugg6\n5/MyuRqNRi5SJBGnkDzFhEPkv4lwiNRJ3W43AP+hSk3hkNZrh1qBFgMOJZA8xkFbZgOhQMQ9Ho9H\nNcu+iFBVBf1LL0G3ZQvYjAzAZAJbVIS0995DutUKvdkMesAAsCdPgrXbQQH+VOyFC6jp2xcupxPW\n8+dh0OvB3nCDfy6VYANjadp/bR50R46ABfzNiw0b+kVE114LNisLbHo6fM88AzRsGHL5t9/OoFs3\nPzE5ncDWrUbccw+Fxx7TIzvbioEDWTAMjW3bXFyvq9zRPiFNs5nFO+8YcMcdsZOl2+2Gy+WKuBRB\nlLFmsxk2m40Ty7lc/s9fXV0Nn88XYL1nMpk4wiMqXY/HE1ALJdcmQ5BJ6pFvx0Z8nYVGC3UBWiJu\n4VrqClkCdTjC5EMzPYoCCNdFnHv0en3E4h61PiP11VegLl70kxXLwsMwoJo1g/HQIeh//hnMzTeD\nvvNOUKmp0G/aBF15OZhmzeDIyYEvPR3133wTuvJygKJA2e2gysvBZGRccQ6oqQF0OjAdOwbe2GAA\nBYD7xFYr2JtuAqqqwNar589hRgizGRg9+kqN0G9AYMLgwTTsdi9X33M6nbVaNmLd7C5eBM6epWAw\nAAcO6HD77YyoICgUyNBwr9fLOQ1FCzHLtCt13/B6PgFctuALVN0ShBIOydXzmURsSKZk6xi0HGGS\nzcLhcCAlJQWpqamqvZyRfj5q/36w6en+OpjH499gDQb4dDrg2DH4brzR/+c6dwbduTN8rH+wNVgW\n9RYsAHX2LHDttQAu/46qq0Ht3Qu2WTP/Wkwm+AoLaw2jZNq0AZua6jdyJ/1hLAvqwgXQw4dH/fmF\nwSMxBEhPtwT49AZT3UZKUhcugKtZdunCcOlZUtMMF3LXV4UI1fPJfwZ8kiXkyU/hCtcm5jhEni9p\nheGPopILWjxIJ6Ee6ixhKlXDJJAjhUL+vsfj4WTbUQ1ChopR9DXXgP35Z3hSU2E0GKA3GPzm6SwL\n3+U6H191WV1d7RebVFZC9+uvYJs1438IsLfcApw/D9+DDwIGA9jWrQGxKQkWC3wPPwzD/PnAxYuc\nHJzu1s1v9C4DgtUAxXoSSVM/EbYEM0wgYBjg448NATVLUtMsKfGrYsMBSw4iQMT1VZYFzpwJbJkB\ngMpKv+5K7PGH6vnkq27JQYNPmr7LvsJerzcss3gx4RBR3cr1PVQTWk7JkgxBXUCdJUw+5K5hygX+\nWC6tOPdE+qxc3brB8PnnMNWrB11qKsCyYCoqAIsFVa1aweB0clGB0+nkxCXU2bNgKapWvRImE+Dz\ngW3f/oq5qgTY9u3hfe016PbtA5xOsNdf7x80LYNlm9vthsfjqeU0JIZI3HaE749OBzzwgA98ExVS\n07zsuRESwTxsw8HZs8Arrxjx0EM+tG3r/91XVACvvmrEoEE0uncPXVPkHyD4g4rJnE9h5Ol2u7lM\nCkndRiocSjoOxQdEqV8XoP4OrCLI5i939MW/brQg4h4AspAlf+OJB0j6z5OdjYyHHoJ+8WKwFy/6\nwxWrFeyMGbA2aQKfz8cJQMgmSNM09JdHfeHECcBiATIz/ezxxx9+h6DqaqBevdALSU+XLaIkn4v0\nWEZTAwyWWpRK3Yo5jlGU+M+FCIjao0zlZ2UBkyf78O9/G/DQQz40acLi1VeN6NqVCYssa6890FKP\nX5sk72hKSgpMJlOtnk8ps3h+9EmeL9/vlu84lDSLjw1k+hGB3C4/WkadJkw+lCDMaMEX9yRiMZ1l\n2QAze6pXL3i7dAFz+DBYgwG44QZQRiOoyz2XpKbGsqw/6nC5kLp6NQwnTkB//Cn8LeMAACAASURB\nVDhgMPhJ1mCA7tIl0O3bwzh9OphevUDn5/v/Pxn5xTBgs7L8P1Pgc5G0ply+w1KpW6FoJprUIk3T\nqK6u5qL2WNCmDYvJk32YO9cf1Q8fTiMnJ/K+TyH4BKfT6bgsA03TqKqqkuz5FJrFE9IUkidFUQHC\noUjN4oXkoBa0lJIVoqamJkmYdQlaehF9Ph/sdjtSU1ORmpqKyspKWa4rZx9msEiVT/aEVFiWBZ2S\nArZdO24Tk4rUjEYjqG+/haGsDL727eFr2hT648dhOHYMOpqGb9gwUA0aADQN/RdfgLVawXbpAn1R\nkd/uhqIAsxl0167QHTkC6tdfwWZng7n7bn9bSpSINa0ZDmJJ3QohdO+RA40bX3l/WreWN1tBbASt\nViuXlpXq+eSbxfN7Pvk1ULHUrTCyjZdw6GqDkLyThFlHoHRKNlKQtBxf3KPVlhcxEJs+vpKXbHrA\nFeVjqEhN/7//AZmZMKSkAE2agG3QAOylS2B8PnjdbsDrhV6ng65JE+jXrQN78CDYtDSgeXP/BU6e\nhOnJJ8HedBPYRo1A/fgj9N99B19hIZjbb4/4c9E0jZqa4KO55EY0qVsCMfeeWEFqlsOH07jxRoZL\nz5KaZrQI1ubCJzi+45DYM+ALh4Sp21Bm8fy/JxQOcc5TKkPrEWYkPrKJjDpNmARKtYJE8mdJU7ZS\n4h6lnX7EbPpIykxSCStRU6MuXQK/X4LyeqGjKLAGA0wAaJ3OTyAADGfOgG3QAFRWFiiWBSgKup9+\n8tc7dTp/v2ZaGtiaGuiXLAHTqVNIsRAfSkRqkSKS1C3Z8EO590SCykpwNUuShiU1zcmTfWjTJrr3\nKhLPXTl7PqXM4sWEQ0LSTQqHkhFmEpC/hhnJfYMNr06ECNPlcsHpdMJqtQaku4Rkya+p8QUdQjDt\n20P/7bdgmzTx/yA1FaxOB3g8QEYGt3GxDgeQkgKvxQLfZSs1Pcsi9fRpsDab35qHwGLxt5ecORPY\nqhIESkRqckAqdetwOAAgIDKSY3M3mYDBg+mAIdqkphltYMHvCY2mHhxrzydfOCQ0i+dH9zRNcwcP\n4jKUFA4FIkmYdQxynxjDJTliRsCv94lBLsKUO8IkqVWv18vN4OR7iPKFFJGQDzNkCHTffQfq1Cmw\nl2dmsunpoDwerocTdjt0Fy6AvuceGH/+GYbUVP/G6fGAoSiwNTVgmjUDRdYB+EVGYUaJ/NFcWu4x\nI5GX1+sFRVEwm81gGEYWwwQCsxkBZEkQS2QZbU+oGGLp+QzHLJ6kZ/mq20iEQ3JAK+IjoHaEmVTJ\n1hGQX7oaUZxQ3CP1RZPrCyj3F1ksMiYnffLlJs/V4/FElCZkmzSB7/nn/Ybte/f6Z2jOmgVQFPTr\n14M6ccL/Zx55BEzHjtAvXgzq5EmgSRN/Oi4rC7qTJ8E2bAiaYeD1+WA4dw5o2RJ0ZiaC0V+sbSPx\nBj9Ss9ls3O9ZCdWtXOslczeVEk9F0vMZyixeuC8khUO1kaxh1jHEW/QjJu6Jx9rkvE5VVRU33JVT\nwl4my1BK2LDukZ0NurAQwsYFpmdPgKYBvZ4zIKBHj4bum2+g270bYBjQ+fnQ/fgj9MeOQa/TASwL\nulEjVN93H7w1NbU2TbKxKW0dJzdCRWpyqm7lAFEa6/X6uNk7Buv5JKlb/rQV4+WJOUK7Pv4hMFLh\nEHm+VwuBitUw69evr+KK4ockYSoAKZKLh7gn2JrkADlRm83mmJSwUYOiavdYWixg7r4bzN13X1ln\nbi6oI0f8jkHp6WDbtIHZYEBqkE2TbHJqDy0OB5G2ucSiupVrvdXV1XFVGgvBfwah5nzye0KNRmOA\ncEgsdQuIC4eEZvHRCoe0rJIlpZa6gCRhIj4p2VDinnisLdbreDweOJ1ObpMGgith9Xq9Ymm3kKAo\nsDfcUKv3UmzTJJ8LuDJOSstDjGMlH6UNE4SQ00BBTgSb88lXxZI6JVDbLJ6MMhMKgMQOKDRNJ7xw\nSGwPqampgTUc26mrAHWaMJWqYQqvR8Q9/BRmvBHLPUmvHCnuu1yumJWwWgGpsZK2Ea/XG5VJeryg\nBPkombol61WzLScckNStTqeDx+Phnm2onk++4jaYXR8h32iFQ1pTyifbSpKQLe3BJ0wi9zebzVFH\nA2pGmCS16vP5kJ6ezl0jViVsWPD5/J6xaWmKWN2JrVeMOIinL6mFqdWLF4+eUDlTt1roYY0EZL1m\nszlgvcEicKFwSJi6jdRxiES2YmbxWjiwAbXXkVTJ1jEo9SJGIu4JBrnaQaK5L+nts9ls0Ol08Pl8\nXO2H/4UmbRiyNMwzDHQbN/odf9xuIDUV9KBBYHr1innSCEGw0VyAdL3L5XKBpumAqCuslJrTCd3O\nnaB+/hnIygJ91121ZngGAyF3OQ0JQiGW1K1We1ilwCdL4Xrl7PkMRzgkNIsn7VpagNg6nE5nUiVb\nFyA8vclZWPd6vfB4PFx/YrRQK8IUSyOTZnij0cilZUkkQtO0bG0Yui++gKGkBEzTpv6JzW439MuX\n+0eD9ekT07VJejnc0VxA8E1NKBYRvd7FizA+9xyoU6f8LgAeD/TLlsE7axbYdu1C3l8rPaFSxMGP\nwMn7EOwwojUEI0shou355EefZJ+REg7xzeLJ9ckzjUU4pBSSbSV1EHIRE6mJMQyDjIyMmAmEWHzF\niki+XKRHlJ9GJnUX0hwPXPFYJetzOp2xKy09Huj/9z8wzZr5yQUAUlLAZmdDv2EDmO7dI7K244Oo\nlGMldyFxBOvzoygK+qVL/Q5DxOsWAKqqYPzXv+BZsEAy3UzI3ev1qk6WQohF4KT2S0QtpManZVFL\nrJF7JD2f0ZjFk2dMPJqJSE0t4ZBYUJGsYdZByEGYJCojL7oWN4pQUTTxhOWnkaWUsKSnzmq1BmwW\nYUVdUqiq8lvgCVPYKSmAywU4HEAUPV9KtbmE7PPT65GxZQvYRo0C/2J6OlBeDurYMbCtW4uuN1yf\nVbVBInASCaWlpXEEqhXDBDHIneaOtOcTqG0WT9qzxMzi+Y5DUj2fhDzj+YyTKdkkIgZf3ENRFDwe\njyzXlSvyDfUFIhu0y+XiekRDKWFNJhMXgVIUFVHUJQmbzR9BCknT4/GTZhTy9XiM5gIk6p4eD5jL\ndT8drvTqcbVYiX5dUh+TtYdVIfAjYX7kHip1q6YbDqlhKxW5R9LzGcosntQw+c9Kyize5XIBgKRw\nKFaIHbj5frtXO+rGp5SAWA0zGgjNx71er2aK9EKIvfB8JSxJI4fyhA2mfAx10g66YaakgO7bF4a1\na/1p2cvkqSsvhy8vL+J0rBqjuQh0Oh1SUlOBnj2Rsm0b6MaNwbAsaK8XVHU1dBYLfNdeCx3vd8K3\njlOrBSkSkIMWTdOikbAUccTLMEEMSpOlGIL1fAKBhwgx5yAAnF9wsJ5PvjhLKBxS6hlr2VRBbtRp\nwuQjGsIkRMM3H9fCuoJdSwiGYbg0cnp6Onc/oR0YEJ0SVrhhiokkhBsmM3AgfAD0X3zht8EzGODL\nyQHTr19En1crbQ10QQF0Bw9C/8cf0KWmgvJ4wFIUqqdNg8vjATweLk3ncrm4+Y9a34SEVoLhuA2F\nEk8pnbrVgoCKf6AMdYigKAperxdmsxm6y2Ptgqluyc/EhENXu+NQPJAkzMuIlJiI24qYc4/cRghK\nXYumadjtdhiNxrA9YWPdaMTaFIRpKqPRCAwZ4re6q6ry1/sibNLXVFtDw4bwvvYadN98A+rAAaBJ\nE9A9esDUpAmMlyN5vtsQ2UC1WgcH5Jk4Iiaekoq65NikXS6X6KBqNSF1iOATHDlMkTplMLP4YMIh\nfs8nEQ7FahbP3yfqApKEGQXEiIYPpaNCOa7Fr7mmXh7WzFfCCj1hiZhDzo0mLLVp/fr+k3AE19VC\nFFELViuY/v2B/v0DfkzeFRJFGAwGecRTCoKfNparJhxJ1BXpOyhUG2uFLMVAvhPEopE8C6meT75Z\nPL+WSa4VquczUuFQMsKsw4imhilGNEpCiWhVaKggJe7hi2WUrqeJ1T0JaYRrz8aPhLUURQRDKLeh\nqMVTCoFkVkiKXYk1yJm6TSS1MQH5nGlpaQGlj2h6PsnfC9csPhrhkFb1GkqgThMmH+EQk1DcE8u1\n1AI5aUejhI0XQjWH8z09+WKZRBrNBYTnNhSqTYFf61Iaak0ciTZ1yxckJYLaGAje6hKq55PfsiJl\nFh9O6lZ4WOMLh4R9tV6vV/2SRxxR5wmTkFswkotU3CN3SlYuQwWS+uMPfBZTwmpFLEMQqu5pMBjg\n9XoTZjQXEHnaWG21qVYmjoSbutXr9dxQ50R5JyLpCw2351NoFi+06yPetcLULWkTAwKFQ6Rkw7Is\nDh48iKZNm0bUgzlhwgSsW7cOjRo1wv79+0X/zJQpU7BhwwZYLBYsWrQInTp1Cvv6SkP7x3CVwTAM\n7HY7GIaJWAmrlSiTfAYAsFgsHFmSLw+fLD0eD+fcoQWyFIJEG2lpaUhPT4fRaOQ2RoZh4Ha7uY1A\niyBRj8fjgdVqjbouSdoUrFYr9xx8Ph8cDgccDgcXWcnxHPgTR7Q0noukblNTU2G1WmGz2bi2LofD\nwQmnSJpSy4jFRIEcplJTU2Gz2WC1WrlDpN1uh8Ph4IzdSeuJyWTiDhb80WPEK1rq+uSA6vP58Nhj\nj+HWW2/FsWPHsGTJEly8eDHkWsePH4+ysjLJ/79+/Xr8+uuvOHz4MN5//3089NBDET0LpZEkzMsQ\ni+RomkZVVRXnZhPJDEsl1xUJyGfgRx7BlLAk6kmERmRCkKmpqUhPT4fZbOayAXa7HU6nU1M9sSRt\nLHc9jUQDFosFNpsNqampnDAn1udAUp/CCR5aBKnHAf6shMVike05KAm5HYeEh0qSmeE/B5JVMplM\nXCaJXwIhJgvkIEpAItG0tDRs2bIFy5YtQ2ZmJpYvX46WLVviz3/+Mw4ePCi5tu7du6N+EKeu0tJS\n/PWvfwUA3HHHHaioqMCZM2difiZyQfu7osKQSsnGKu7hX1eO9UUD4Wew2+2SSthEq/+JiWWE8nl+\nqk6s7hlPyNGGEQ5CpW4jmbKixoSUWECeMUVRnEhNCdWtnFD6GUeiPhbO+RS2rZDMFP/dTUtLQ/v2\n7bFgwQK4XC5s2bIF2dnZUa+3vLwczXmey82aNcPvv/+OrKysqK8pJ7T/LYgTCDERFWk44h4tg3wh\nyGcgn40Mx+VMAnhK2ESp9YQjlgmmsuTXduKxWcbLmk+IcNSmUi0roZ6x1hCs1UULhgli4Kth49Ey\nFKrnkzwHfm8mv+eTpGv5bSvV1dWc8XpqaioGDBgQ8zqFAYKW9iTtfxPiCL5FnJbGcpG1hfPikGjR\n4/HUUsKaTCZ4PB44HA7ui0MINN5K2GhADjORjOYCJPo9KypA//orYDBA16YNjBaL34mnvBxsWhrY\ntm1lGVodjzaMcBHulBXS2K6pPtYg4A8CCOcZx9swQQxq2PMJITWuTUyFrdPp4Ha7OSIF/FHn4cOH\nZU2ZNm3aFCdPnuT++/fff0fTpk1lu36sSBImD0RyTSziYoFchBnJOsgpm6bpWkpY4upBok2i0gT8\nJ10AmpsmwQc5CMQ6mouiKKR8/TXS/u//wNI0wDCgzWZ4GzUCdfQoKL0eeooC27Ah6CeeABvDl1Ur\nylIxSPW98hvkhYIwLSLWA4mShglS0AJZCiHVykVEgETrkJqayv2ZI0eO4IMPPsDcuXNlW0dOTg7e\nfvtt5OfnY/v27ahXr55m0rFAkjBBURS3sQHyjn6SC+HUQ4knrE6nE/WE5fdYkuHW5Asr3CzJBqKV\nIbX8+l+svx/q8GEY/vMf/7ityyRmOHAApq1b4RsyBLTJBB/DgDp/HtRrr8E7Zw4MJlPE99Raa04w\nkFQdMfe2WCycalLYIK+l+rbcfaHxSN1qkSzFQJ6DyWTi9kadToe1a9di9uzZ6NWrF7766issXboU\nHTp0CPu6o0ePxpYtW3D+/Hk0b94cs2fP5g7shYWFGDx4MNavX48bbrgBaWlp+OijjxT5fNEi6G+c\n1ZqcTAHU1NSgqqoKKSkp8Hg8qFevnizXraqqks3H9NKlS0GHUROrPpPJxNVvpJSwxLjAYrHU+sLy\nxTKkXhHv5ngh5K7/6d9/H/pvvwXbuDH3M93mzYDdDrZzZ27IM8uyYE+cQPXMmXBfe21EpKEpH9sw\nwBd9kbYj/v8j74PP5+Ps09TORhCyJMYaSoOfuuVnZCJJ3SYKWRLwD6pEREXTND777DPMnz8fZ86c\nwdmzZzFw4EAMHToU+fn5mjhgywFK4oPU+QiTYRguxSfXDEsgfuYFRAlrsVi4jYN8uQFEpIQVO2GT\naFQNZaESo7moCxfAClTPlMcDVqcDLqeogcvyeYMBZgAp6ekcafA9N8VIIxHFMsEmjpCWFWGdi5+N\nULreJ4Qaqe5YU7dXA1kCwNmzZ/Hqq6/i3XffRadOnVBeXo7169dj+/btGD16tJpLjgu0/41WGGTk\nlJabm6UIU0wJG8oTNhIlrHCGHyEN4WQRJchTqZQm0749DD/9BJaXSWAaNoTu2DEw/OyC1wtQFNjr\nrpMkjerq6oDaD0ljJuKmGO54rnBad5Q8UPFNFNRKdUeauiX2con4XvDJ8vTp0xgzZgzmz5/Pue80\nbdoUDz74oGprjTfqPGGSl0FuVavc1+ODr4Qlat5gnrByRGlC0hBTWMqVplMypcn06AH2889BlZeD\nveYa/7xNsxlsZiYotxus0wk4naCqquDLz/ePFuNBqs+RryxMBLFMsDaMcBBLy0q00AJZiiGU6pZl\nWaSmpmqq/isFsrcAtSPLMWPGYN68ebj99tvVXKKqqPM1TKIGY1kWly5dQv369WXZ6KqrqzmZe6zg\n10PJRsdPJUuRZTyEJ/yIizzHWERDcRnNdeEC9GvW+GuZKSlg+vQB3bEj9Js2Qbd/P9hrrgEzYACY\nzp2BEOvnpzTNZjO3Wfp8Ps00xwuhdKsL/0BFTDJinbJC3uVEqQsD4JyzTCYT592q1XcCuPIusywb\nQJYXLlxAfn4+5syZg+7du6u8yvhAqoaZJMzLhAkAFy9e1DRh6vV62O126PV6LoUmpYRVo5YWi2iI\nWPMl0hgmqdQV+X/kOXi9XtWa44WI98QRvnE3saWLVEiWiGRJDn78Fih+FO7z+TTzTgDStexLly5h\n1KhReOmll9CrVy/V1hdvJAlTAmSDB0KrUSMB6V0ym80xX8tut8NgMHCn1XCUsFqopZFn6/V6g56u\ng6k0tYpI1LtChSVJ68a7dUcLfaH8AxWJwoOpjxPNng8QJ0shpFS3arRzSZFlRUUF8vPz8dxzz+Hu\nu++O23q0gCRhSoBPmBUVFbDZbLKQDD/yiBWVlZUcmYSrhNUa8ZCIi3+6JuTpdDoD/D+1jliiNH4U\nHm3EFQ202BcaqmWFCMwSjSzJYTXc758wM0PTdFwEVOTeZLINnyyrqqqQn5+Pp556CgMHDlTs/lpF\nsq0kzpBL9ENeZjJaKZQSlqIoTXrCSomGXC4XV+NKBLFMrFEaXywTL2cZraY0Q7WsELFMIihLgejI\nEohMQCXn90OKLB0OBwoKCjB9+vQ6SZbBkCRMHuTunRTOlYsEfCUsOXGHUsJqwa80HFAUxUUQKSkp\n3Ow+rToNESgRpYXTuhOL0jRRUpp89bHb7YbL5YLJZOIOVfGKuKKFy+WC1+uVpf4eTHUrVzpfiiyr\nq6tRUFCARx99FEOHDo3pc1yN0O43KE7gv3DxMhsIBZZl4XA4wLIs0tPTOeVaMCWsFv1KpSA1mouf\nmhIzgFaTPONBPGJReCwbZaKZKACB9b94tqzEAjnJUggpwwRCdtEcJKTI0ul0YuzYsSgsLMTw4cNl\n/RxXC+p8DZO9PPIK8ItryDTyWEEs6KxWa0R/j2GYWkpY0s9luuxpKlTCai3VFgyRbOLhioaUhtou\nLdEoTePSniMzwklpKtGyEguUJMtQEFPdhkrd8u0x+Wt2uVwYO3Ys7rvvvjrh2BMKSdGPBJQiTI/H\nA7fbDZvNFvbf8fl8cDgcSElJ4VKr5DTIr/WRFCZ56RNhQ4xVvSu1OShpCB7Ke1ctBDtIUBQluiFq\nGVKbeDh/L9aWlXivWcn1hKPEFiN4t9uNcePGYeTIkbjvvvs0VQZRC0nCDAIy5srhcHDKx1hBopJ0\ngVNMsD9PhrGKKWGpy8bHxNsV8MvQTSaT5mp9QpB6LEkBxbq5SEUZcvazJUpfqPAgQT57oqRh5XzO\nkbasxLJmLZGlEFKqW3LA4Le7eDwejB8/HkOHDsWECRM0vY/EE0nCDAJCmHKaDZB6SziE6XK54HQ6\nQ3rCEpcfwO+Byz9Zq2GCHQ6CNffLdX1ykJCrn42smWVZTSqOxUDWzDAM9Hq9JtKVocCvpcndBhWq\nZSUWsYyWyVIMDMPA6XRyh+9Dhw5h69atGDBgAP7f//t/6NevHwoLCzX3fqiJZFtJEJDUZ7xFP2ST\n83q9IT1hxezMyL+TtJTb7YbT6dSMUEbu0VxiCMfbNZJnQQ4lWm3PEQP/UELmhfLTlVoTUJE1B5uS\nEitCtaxEc8BMlKyDEF6vFwzDwGazce/1b7/9htzcXDAMg+bNm2PTpk3o3r17wmgh1EIywoQ/LUE2\nHbnceYi6MSMjQ/T/85Ww4XrChqOEFUtLqSGUIf2KZF6hGht0pKIhpT1WlQAh+FCHEn6tT21PU6XJ\nMtS9ozEJSFSyFBNS0TSNyZMno1OnTujduzfWrl2LNWvWwOFw4KeffpL19/Hzzz8jPz+f+++jR4/i\nxRdfxJQpU2S7hxJIpmSDgBAm33g4VpChzmIDqYkS1mAwcGlKKU/YWCZ3SLnrKCmUAbTpKhNKNKTE\n7E2lEW0Er4aAikDpFH2kCKY0JSIvqTYMrUPMoo+maUyZMgVt27bFk08+GfBZqqurkZaWpth6GIZB\n06ZNsXPnTjS/PKhdq0imZMNArGYDwmuJnTeklLBinrBEaRutElbKXUc4BFlO9afso7kYBtTp0wDL\ngm3cGIhyrWLN4PzxZAzDcBF8ImyIsUTDoZ6FUnVPrZElEJ5JAMn6kHR3IkDK/H3atGm44YYbapEl\nAEXJEgC++OILtGrVSvNkGQxJwoQyNUwxECVsWloaF3kF84QVKtpiAV9JKrYxyKEylbv3jzpxAvpP\nPgF18aKfMNPTQY8ZA7ZVq9iuy/u8pK5lMBjg8Xjg9XpVMUaPBHKaqAvfC1LrI9kWucRk4aaO1YTQ\nJICmaS51TFEUnE6nZmrAwUAO2kKynDFjBrKzs/HMM8+osv5PP/0UBQUFcb+vnEimZHGlKB6t2YAY\nWNY/XzMzM5NT1jmdTthstgBXGzElbDxP4fxNMtp5lorUd6qqYJw7F6zJBJC0tt0Oym6Hd8YMoEGD\nmG8hNFFQyxg9EsRz4oiw7kmeg9FojOhZJAJZCiGsswpVt0q1rMQKMZMNhmEwa9YspKWl4eWXX1bl\n+Xs8HjRt2hQHDhxAw4YN437/SJFMyYYBJSJMvqQ7GiWs0uCrTKOxphNuLHJtHrr9+wG3G2jU6MoP\nbTagqgq6H34AE+O4IbFomKLCM0aPlDDkQrxrw+RZpKSkSNrThSKMRBVSCUVJFEWJev4KyxtqDg+Q\nIst//OMfMBqNeOmll1Rb24YNG3DrrbcmBFkGQ5IweZC7rQTwmyFQFAWbzSabElYphCIMYYShaAvG\nhQuASA2UTUkBde5c1Jfl99GFSnfzjdGjJQy5IHttOEKEU/cUEka8h1XLgXAUvEq0rMQKMbJkWRYv\nv/wyvF4v5s2bp2okvHTp0qvCci9JmLhCbnISJk3TAPwbDfniKaGEVRKhCINhGBgMBkXSbOx11wFb\nttT6OeV0gmnRIrprxhANByMMpSMMrU0ckap78glDr9dzE0fkMAKJB6Jpd5HK0AgPmUq270iR5dy5\nc1FZWYm3335bVbKsrq7GF198gQ8++EC1NcgF9b99VyF8Ph/sdjsoilJcCRsv8AmDiIV0Oh1X/5U7\n2mLbtAGblQXq99/96liKAnXmDNj69cF06BD59Xi14VijYSnCkHP8EoHaxu+hIEYYHo8HTqcTALj/\nViuNHS74bWXRvh/8DA0QnykrZHKJkCznzZuHU6dO4b333lO9xpqWlobz58+ruga5kBT9AFwDc7De\nyXDBV8LW1NRwqVgxJWwiNkILo2FFfV2rqqD/3/+g27ULYFkwnTqBHjgQqF8/osvES3Qit2goESeO\n8EVJRqMx4N3QqlBG2IOtxPvBV6YHM0ePBOS7KCTL+fPn49ChQ1i4cGHCvDdaQ9K4IAiIGpBhGFRW\nVqJ+hBsycIUAXS4Xp4StrKzkNmg1lbByIdRoLqGvq2zRFumNjWKTVVN0IuU0FCraSkS/UuAKWYqJ\nkhQ9WMUANb6LfNvCaA9WYml6lmXx3nvv4YcffsCiRYuSZBkDkoQZBOTl5beCRALypfP5fAHinqqq\nKuj1em6iCJC4qsFIR3OJRVuxmqJHCn60Q2aJqgWho4xUtJWomQciWgunDi88WAGxG+ZHA60cXCOd\nsiJFlgsXLsQ333yDjz/+WBO17kRGkjCDQEiY9evXj8hmjChh+cbXfLIg0xL0ej08Hg93Ak8UspRj\nNBe/15OIhZTsb9SiPR+BMNoioiGDwQC32x1THU0NREKWQgi9XePxbpD7aoEshRD2ewoFZcTCUUiW\nixcvxsaNG7FkyRJNCQcTFUnCDAJCmABw8eLFsAmT1DyNRmNQT1gSoZExYjqdThPpqFBQalOJNlUZ\nLrSqOhYDP9oic05NJpOmnYb4kFvBG4+ZllolSyHETEVYlg2wcGRZFkuWLMH69euxbNkyzR0OExVJ\nwgwC8iUFgEuXLiEjIyPkl9Pr9cLhcMBsNnOyeam2EUKWFosFer1emTqfzIjHaC5yHzmFIaHqrFoE\nv581JSUlptpWPKF0u0uoaCua55EoZCkEadsxGo1gGAYPPfQQvF4vWrVqhOtllgAAIABJREFUhQMH\nDqC0tFS1/u2rEUnCDAI+YVZUVMBmswWt05EeKzFPWKG4J1g9Sgt1PjGoNZpLLFUZiQw/EVWlwWra\nclnTKYF4H0zkqHsSsqQoKmEs+oArKW/+sz5z5gzeeecdlJaW4vTp0+jUqRNycnIwfPhwXH/99bKv\noaKiAhMnTuTGf3344Yfo2rWr7PfRCpLWeDKAEKDb7Y7IE1ZqyoHQWUdofK1GdKFm7U/Y3xiuQTz/\nYCKXWX08EMoJJ5g1nZotGmr0hvL7PaMZFJ7oZGk2mwMOJjt27MD+/fuxZ88eUBSFjRs3orS0FOvW\nrcOjjz4q+zoee+wxDB48GEVFRdx3si4iGWEiMMKsrKxEWlparVMzSZvRNB3S5o5shHq9PuovJz/y\njJcQQqu1P7FaDp88XS4XGIaBxWJJGLKMxURdSjQUj5q4FqP4UEPTE9H8HZAWU61btw7vvfceVq1a\nJcugiFCorKxEp06dcPToUcXvpRUkI8wg4D8bMXs8vhI2PT09QAkrJEu5WhmEtnRKm4BrcSMkkLIf\n449eSpQWHSB4v2I4iKfTEB9afUf43xV+3ZNv4UiyOIn0joiR5WeffYYFCxbEjSwB4NixY2jYsCHG\njx+PvXv34tZbb8Wbb74Ji8USl/trCYlxHI8z+IRJ0zSqqqpgMBgC2kZomq5FlmTTMpvNstb+yIZg\ntVphs9m4OY5VVVWorq6Gx+OJevA1aRvxeDywWq2a2gjFQNLY5DBiMBhgMpng8XhQVVWFmpoaeDwe\n2afOyAWSzjKbzbKkvMkzMJvNsNlsXATldDpht9vhdDq5+ni0ICnvRHhHKMpvjG6xWGCz2TgBHjmk\nkMlBWn0/gCsHKiFZbty4EfPmzcPKlSuRnp4et/X4fD58//33mDx5Mr7//nukpaVhzpw5cbu/lpCM\nMAXgk1y0SlglRRBCE3CSto1mgkYsZuRqQqr2x09jkzofPzWnNpROeQvrfEQ0JDVtJhwkqusQqVmS\nsgiAiOueakCKLLdu3Yq5c+eitLQUGRkZcV1Ts2bN0KxZM3Tp0gUAcM899yQJsy5DLCVLNhmr1cq9\nuAzDgKZpUBQl6gkbb8EJOU1LTdAIRp78VoZEapIPVvsTS82pNY5LCDXaXWIVDSWq6xB5t4Vp2HBm\nnap5uOKn6vlk+fXXX+PFF19EaWlpVLadsaJx48Zo3rw5fvnlF9x444344osv0K5du7ivQwtIin4u\ng5gKOBwOrjZptVrDUsJqzZkllG8n2VASyZ4PiF7BG0wkE4/0otZqf+E8D2H2IVHeESmyDPV3yLPw\ner2KTBUJBam69o4dOzBr1iysXr1a1eHLe/fuxcSJE+HxeNCqVSt89NFHcY9044lkH2YIkDpgZWUl\nWJblzAuCKWHj0dgfK8T611iW5eYUanXdQsiVzhQziOdvjnI/D1L70wpZCiE1RYOUHaRaorSIaMhS\n7BpyTxUJBSmy3LVrF2bOnImVK1eicePGst83CWkkCTMEXC4X7HY7GIbhRAOhlLDxbuyPFWROocFg\nAMMwmjFKCAWl0pliUyPkeh6JmM4kz4NkTfiHCS3V+cQgB1mKXTPWqSKhQBT4QrLcs2cPpk2bhpKS\nEmRnZ8d8nyQiQ5Iwg4BlWZw7dy4gcjGbzaLiHhLpaNHUOxiEpCPlMqSlzZHUkuMVocllEM9PZyZS\nb6jQNo5f99Sa0xAf8ZoAJOWBHG3dk5ClsB6/f/9+TJkyBcXFxWjWrJmcHyGJMJEkzBBwOp1cEzxN\n05xwhP9FiJcSVk6ESzr8k3S8jBKCQe0ILZZZlolc+5MqMcjt+SsX1BqXJ3wekdY9pcjywIEDmDx5\nMpYvX44WLVoo+AmSCIYkYYYAIQqn0wm3243U1FSOLPibNzFQTwSQzTvS0VxCsoh3ZKE10olklmUi\nGntH6oSjBZEMENpaMF6QEtlJ1cXJuklJh+DQoUMoLCzEp59+ilatWsX7YyTBQ5IwQ8Dj8cDn84mm\n5WiaBoCEqUUB8m3eYmShpPye3+6iRdLhb45CsiBZCi2LwIQg4rVoa39iIhklRVT8dWuBLIUIVfck\n77eQLA8fPoyJEydiyZIlaN26tYqfIAkgSZghsXv3brRu3TrgS04UjgACBCFaqfFJQSkFL7+3UYm0\nnFrptWghJE9CFiaTSdNzTgnkft5SIiq56+JaJUsxCCfOAOCcmch35ujRoxg/fjz++9//ok2bNmou\nN4nLSBJmEHi9XkyaNAn79u3DXXfdhdzcXFgsFowZMwbFxcW44YYbAshCywKIeCl4pSKtaMmTqDMT\nYRPkg086BoMhru0IsSAepKPEeDL+uon7ViKA1CxJynrJkiVYvnw5evfujdLSUixdurTOmgFoEUnC\nDANerxdbt27FvHnzsHnzZowcORIFBQXo2rVrQG1GKk2pNnmqNZorVmOAWCZ3qAmpdWtdgSwlOFH6\nnsEmioR7jUQlS+G6a2pqsHLlSrz//vs4evQosrOzkZubi9zcXNx2222KvCMtWrRAeno6d7jduXOn\n7Pe4WpAkzDCxcOFCzJo1C0uXLgVFUSgqKsKOHTvQuXNn5OXl4a677gpQyIqlKdWw2NLKaC4pYwCp\n0VNSI4y0jkgOJ1pSIGvhcBKNaEhKKKN1SEXyp06dQkFBARYsWIAOHTpgx44dWL16Nfbv349169Yp\n8l60bNkSu3fvRmZmpuzXvtqQJMwwcOTIEQwdOhSrV6/GjTfeyP2cpmls374dRUVF2LZtG2655Rbk\n5uaiR48eAZulWuSpNes1AjGXIb4xgNfrjbu/qhyIheTliLSiRaxjxZRAOKKhRCVLlmXhcDhq1YhP\nnz6NgoICvPXWW7j99tvjtp6WLVti165daNCgQdzumahIEmaYIJuYFBiGwa5du1BUVITNmzejTZs2\nyMvLQ+/evWul5eSs8YlB7V7FSCBMU5JxZGTzVjtNGS5IJC8HyQsPWEoaxKuVro8EUgpTn8/HWTkm\nCqSch86dO4f8/Hy88cYbuPPOO+O6puuvvx4ZGRnQ6/UoLCzEgw8+GNf7JxKShKkAGIbB3r17sWLF\nCmzcuBEtW7ZEXl4e7r77bm6kEKAMeSaymwwxUjAajdxcUbWNEsKBkhNHlDSIT9S0t9frRU1NDfc+\nJMI7AkiT5YULFzBq1CjMnTsX3bt3j/u6Tp06hSZNmuDcuXPo168f5s+fr8o6EgFJwlQYLMvixx9/\nRFFRET7//HM0bdoUubm5GDBgANLS0gL+nHBjjJQ8td6rKAUpkhczStDaxhjPtLecBvGJSpZCYZKa\nqexIIEWWly5dwqhRo/Diiy+id+/eKq8SmD17NqxWK5544gm1l6JJJAkzjmBZFj///DOKiopQVlaG\nhg0bIjc3FwMHDgyYlC5VvwkWVSRaryJBuEYKUhujWgpkEhGrNUA5FoN4OdPH8UQoYZJWnIbE1iXm\nmFRRUYH8/Hw8++yz6Nevnyprq6mpAU3TsNlsqK6uRv/+/fH888+jf//+qqxH60gSpkpgWRZHjhxB\ncXEx1q1bh4yMDOTk5GDIkCGoV69ewJ8LpS7VgsIxGkRqvUYQb5chIbRWIxZrV5FKUyY6WYZba1XL\naUhsHWLveFVVFfLz8zFz5kwMGjQoLmsRw7FjxzB8+HAA/qzDmDFj8PTTT6u2Hq0jSZgaAMuyOH78\nOIqLi7FmzRpYLBYMGzYMQ4cORWZmZsBgaiF56nQ6+Hw+mM1mzYo2xCBXRCwVVShl/q01P1sxSHn+\nkqhYa6rpUIhVxcv/3vh8vriNryPZE4qiAsjS4XAgPz8fU6dOxbBhwxS5dxLKIEmYGgPLsigvL0dJ\nSQlKS0uh0+kwbNgwDBs2DA0bNgwgT4fDETCTU8sOMnzwI2I5lbBSdWC5UnKJaKJOonG32w2GYaDX\n62EymTRX45OC3C0v/Gg81nFtoe4jRpbV1dUYPXo0Jk+ejBEjRshyryTihyRhahgsy+LMmTNYuXIl\nVq1aBa/Xy5Hnhx9+iB9//BGffPIJdDqdqIOMFskzXm0MkRolhHO9aNLHWgB//Bw/na10NB4r4tEf\nqoRoSOpg5XQ6UVBQgIkTJ2LkyJGyfYYk4ockYSYIWJbF+fPnUVxcjDlz5kCn02HChAm455570Lx5\n84DIU6vkqZbrUKzqUqVM6+MBl8slKkySGj0V7YFCbqhhpiCHaEiKLF0uF8aOHYuxY8eioKBAsc+Q\nhLJIEmYCweFwYNSoUWAYBu+++y42bdqEkpISVFRUYODAgcjNzUWLFi0CNjs+UajpXapkr2IkiPRA\nkcjq43BVvGIHCjXT+1pwHormQCFFlm63G3/9619xzz334L777kuYdyiJ2kgSZgLhueeeQ3l5Od59\n992ACK2yshJr1qxBSUkJzp49i379+iE3NxetW7cO+HIKHXXi1fCtVYs+oPaBQmxGYaKZesei4lXb\nIJ6QpZb6Q8MRDRExGMuyAWTp8Xgwfvx4DB06FBMmTLgqyXLv3r0wm80BtqFXK5KEmUDwer0hNy27\n3Y7169ejqKgI5eXl6NOnD/Ly8tC2bVtJ8lTKFEBr7RehIHwmgN/jNpHSsOSZ0zQti4o3ngbxWiRL\nIaREQ8TSkf/MvV4vJk6ciD59+mDSpEkJ8w5FAqfTienTp6O8vByvvvrqVU+aScK8ilFTU4OysjIU\nFRXh6NGj6NWrF4YPH4527doFkJcS5JmoFn2AX5hEIkuGYUDTtOpGCeFA6ZYX4XsiZ/9rojoPkXmt\nhDD37NmDo0ePYuDAgfj73/+Obt264ZFHHtHsOxMLjh8/jjlz5mDevHl45ZVX8Msvv+D5559H27Zt\n1V6aYkgSZh2By+XCZ599huLiYhw8eBA9evRAXl4eOnbsWIs8Y53pmYjtFwRiG7faRgnhQColqBTE\nngl5LpE+k0QlS+EBBQC2bduGd955Bxs3bkRWVhYefvhhDB8+HC1btlR5tcrg4YcfhtVqxdy5c/HE\nE0+gvLwczz77LG6++eaE+t6HiyRh1kF4PB5s3LgRK1aswP79+9GtWzfk5uaiS5cuMZNnIitKw3HB\nieckkXCh9gEllv7XRCZLsdQ3TdOYMmUKWrVqhY4dO2L16tUoLS3FDTfcgG3btin2u6FpGrfddhua\nNWuGNWvWKHIPPhiGgU6ng91ux9y5czF27Fi0adMGjz32GE6dOoXnnnsO7dq1S6jvfzhIEmYdh8/n\nw5YtW7BixQrs3r0bd9xxB3JycnDnnXcGbHbhzPSUGoqbCCAq3kiESXIY5scKqQZ5tRDMB1moLr3a\nyJJhGEydOhXXXXcdZs2aFUCihw8fRps2bRRb0xtvvIHdu3fDbrejtLRUsfuwLBvwO3Q6nXjrrbfg\n9Xrx97//HQAwY8YM7NmzB/PmzUO7du0UW4saSBJmEhxomsZXX32F4uJibN++HZ07d0ZeXh7uuuuu\ngIhLbCyZwWCA2+1GampqQvnZAvKoeIO1ISilDNa6mUIwdSlJZ15NZDljxgxcc801mD17dlx/F7//\n/jvuv/9+zJo1C2+88YZiESafLFesWIH09HQMGDAA58+fx4gRIzB27Fj87W9/AwC88MILmDhxIrKz\nsxVZi1qo84Q5YcIErFu3Do0aNcL+/fsBABcvXsSoUaNw/PhxtGjRAsuXLw8wRK8LoGka27dvR1FR\nEbZt24ZbbrkFubm56NGjR0BvHNlAPB4PAGgiRRkulJo4IrfLkBhI6ls4LkqrELarEHUpsenT+vqB\n4GQ5a9YsWCwWvPLKK3H/LCNHjsQzzzyDqqoqvPbaa4oT5s6dO/HSSy9h3bp1WL16NYYOHYoffvgB\nS5cuxeTJk9GiRQvu75DU7dUCKcK8ej5hCIwfPx5lZWUBP5szZw769euHX375BX379sWcOXNUWp16\n0Ov1uOuuuzBv3jzs2LEDhYWF2LZtG/r3749JkyahrKwMbrcbpaWlmDBhAiwWC9LT05GSkgKapuFw\nOOBwODgPU62BbH5KjOcijf9msxk2m40bGl5TUwO73Q6n08lFXNGApL4ThSwB/zMhmQiWZWE2m7ms\nRFVVFWpqauDxeKJ+JkqDHK58Pl9AnZhhGMyePRsGgwEvv/xy3H8Xa9euRaNGjdCpUyfFnx1FUVi9\nejXGjRuHRx99FM8++yxGjx6NkpISdOrUCQDw66+/AgD3nb+ayDIY6kyECQC//fYbhg0bxkWYbdq0\nwZYtW5CVlYXTp0+jV69eOHTokMqr1AYYhsG+ffuwYsUKLF++HJcuXcKsWbMwbtw4jhgAdVKU4UKt\niSNy2BYmcp2Y1CyFoqpEUCGLHa5YlsXLL78Mh8OBf/3rX6qs9ZlnnsF///tfGAwGuFwuVFVV4S9/\n+QsWL16syP1effVVGI1GTJ06FQCwatUqjBkzBhs2bIDVasWoUaPw1VdfISsrK6HezXBR5yNMMZw5\ncwZZWVkAgKysLJw5c0blFWkHOp0OHTp0QFpaGnw+Hz766CNUVFRg2LBhGDduHEpKSlBdXc0RpMVi\n4aIsUnOz2+1caiveZy8ikmFZNu7juUiUlZqaCpvNxm2+LpcLdrsdNTU1HJGKgU+WiRJZEni9XlGy\nBPzvlMlkQlpaGtLT02E0GuHz+WC32zWRpZAiy1dffRWXLl1SjSwB4JVXXsHJkydx7NgxfPrpp+jT\np49iZAkABoMBu3fv5v47Ly8PgwYNQn5+PgwGA9atW4fGjRsn1LspB+o0YfJBRmclcQUffPABli1b\nhq+//hrDhg3D7Nmz8fXXX+Oll17Cb7/9huHDh2PMmDFYvnw5qqqqRFOUhDwdDkfcyJPck6IoTfSH\n6vV6pKSkwGq1wmq1Qq/XS6YoSZrbZDIllE0fENnQaoqiYDKZRFP8ahy0pMhy3rx5KC8vxzvvvKOZ\nKBiA4u/05MmTsW/fPjzwwAOoqqpCaWkprrnmGhQWFqKsrOyqd/qRQp1PyW7evBmNGzfGqVOn0Lt3\n72RKlofq6mp4vV5JIRTLsjhy5AiKi4uxbt06ZGRkICcnB0OGDAn4O0IVJQDFTL8TSSQjTFHq9XrQ\nNJ2QCuRIyDIYYp04Ew3cbjc8Hk8tspw/fz4OHTqEhQsXql5eUAp8RSz5d6/XC6PRCI/Hg7y8PDRs\n2BC7du3CqlWrsHbtWhw9ehTz589XeeXKos6rZIHahDljxgw0aNAAM2fOxJw5c1BRUVEnhT9ygGVZ\nHD9+HMXFxVi7di3MZjOGDRuGoUOHIjMzMy5jyRK97kdaR4iyNFpHnXhDLrIUgpAnOVQoMcJOiizf\ne+89fP/991i0aJGqU3eUBFG2Hj58GCaTCRkZGdxB1+PxwGQycdkamqaxefNmPP/881i1alWAQvZq\nRJ0nzNGjR2PLli04f/48srKy8MILLyA3Nxf33nsvTpw4UWfbSpQAy7IoLy9HSUkJSktLodPpuIHY\nDRs2VIQ8iaF3SkpKwkVnwsZ+sf5XrbbwKEWWYuBnKeQwiCd9uVarNYAsFy5ciG+++QYff/zxVUuW\nNE1Dr9dj165duPfee9GyZUt07doVgwcPxl133QUAnDALAM6dO4eFCxdixIgRdSIdW+cJMwl1wLIs\nzpw5g5UrV2LVqlXwer0YOnQocnNza4kGop3pqYW5itEilAuOmMsQP52tJghZqjHOTcogPlwvZCmy\nXLx4Mb788kssXbo0oYwWosHBgwfx5ptv4v7770dmZianhid92ELwCfRqR5Iwk1AdLMvi/PnzWLVq\nFVatWoWamhoMGjQIOTk5aN68eVQzPRPVdg2IPDqLh1FCuFCTLIWI1CCe2CMKyXLJkiVYt24dli9f\nnnAHr3BQU1OD9957D1OnTgXDMJg2bRoWLlyIY8eO4ZprrsFPP/2E0tJSlJeXIy8vD3fffbfaS1YN\nScJMQnO4ePEiSktLUVJSgoqKCgwcOBC5ublo0aJFWORJTAnqAlkKIUaeSgmphIjGjzdekEpnE9GQ\n1NqXLVuGkpISrFixIuHUyeHC6XTi888/R+/evWGz2eByuTBy5Eh4PB6sW7cOBoMBBw4cQFFREXJz\nc9GhQwe1l6wakoSpEYhZ9K1YsQL/+Mc/cOjQIXz33Xfo3LmzyquMPyorK7F27VoUFxfjzJkz6N+/\nP3Jzc9G6dWtR8vR4PGAYhmvZSBTbNUB+wlFSSCWElslSCKFBPPkZOWCR51JSUoJPPvkEJSUlAaYc\nVxNIzRIAcnJyYDQaUVxcDI/Hg0mTJuHs2bNYsWIFzGYz7HY7bDabyitWF0nC1Ai++uorWK1WjBs3\njiPMQ4cOQafTobCwEK+//nqdJEw+7HY71q9fj6KiIpSXl6NPnz7Iy8tD27ZtQVEUFi1ahI4dO6Jd\nu3bcdJVoZ3rGG/EgHLnFMQSJRJZCuN1uuFwuzixh0qRJyM7OxnXXXYcvvvgCq1evhsViUXuZikA4\necTtdmPIkCHIzs7G4sWL4fP58MADD+DIkSPYvHkzdDqd5sRl8UaSMDUEYXsLQe/evZOEKUBNTQ3K\nyspQVFSEo0ePIjs7G7t27cLq1atx0003cX9OjoHYSkOOaSmRQiiOIc8kUvJMZLIUrp1lWezduxcf\nfvghSktLQVEU8vLyMHz4cPTp00ex+qXL5ULPnj25Vpbc3Fz885//VOReBD/99BN3sOT3WNI0jf79\n+6Np06ZYvHgxaJrGjh070K1bN0XXkyhIWuMlkZCwWCwYMWIEPvnkE3Tv3h27du1C7969UVhYiOee\new7ff/89108mtF3zer2oqqpCdXU1l8JVC3xVZjwJR6fTcS5DNpuNq+Pxn0uoc3Eik6XX6621doqi\ncO7cORw5cgS//vorvv76a7Ru3RovvPACZs2apdhaUlNTsWnTJuzZswf79u3Dpk2bsG3bNsXut3Ll\nSnTt2hVff/01KIriUvXEJOPLL7/EyZMnMXToUOj1eo4sk3GSNOqGRjiJhAbDMJg8eTK+//57/PDD\nD2jQoAE8Hg82btyIRYsWYe/evejWrRvy8vLQpUsXjjxJ4zWJOp1OZ9x7GvmjxfiqTDVAyDMlJSUg\nInc6nZJG6IlOlmJK3k2bNuGNN97A6tWrkZ6ejvT0dEyfPh3Tp09X/FBF0r4ejwc0TSMzM1OR+9A0\njeHDh+OVV17BpEmT8Pbbb6Nnz55gWRYGg4HLwmzatAlr164N+LtaychoEckIMwnNQ6fT4bbbbsOX\nX36JBg0aAABMJhMGDhyI//znP/j222+Rk5ODZcuWoXfv3pgxYwa2bdsGmqYlPUvjYfit5GixWBGO\nEbrL5Up4srRYLAFr37p1K+bMmYOVK1eKmpQo/TtiGAYdO3ZEVlYWevfujZtvvln2e5AI8syZMzhy\n5AgyMzMxYMAAlJWVcZEmIU0AGDp0KIBkZBkOtPMNTgJA8qWVwsSJEyWVewaDAX379sW7776L7du3\nY+TIkVizZg369OmDqVOnYsuWLQGjx+Ix01M4hFhLZCmE2KHC4/HA7XaDoij4fD7QNK32MsOGVMvO\n119/jRdffBErV65ULLILBZ1Ohz179uD333/H1q1bsXnzZtnvQeaPFhQUoHHjxtiyZQvefvtt3H//\n/Vi7dm0AafKRjCxDIyn6iTOEFn2zZ89GZmYmHn30UZw/fx4ZGRno1KkTNmzYoPZSEx40TWP79u0o\nLi7Gtm3b0L59e87FhC/skNtNR605nHKBL04iKW21jRLChdQszh07dmDWrFlYtWoVGjVqpOIKr+DF\nF1+E2WzG9OnTZbne66+/jm7duuHOO+8EAPztb3/DuHHj8Oc//xkAMG/ePEyfPh2rVq3CsGHDZLnn\n1YqkSjaJOg2GYbBr1y4UFRVh8+bNaNOmDXJzc9GnT58A79lgbjrhkOfVQpbCeqsaU0QihRRZ7tq1\nCzNnzsTKlSvRuHFj1dZ3/vx5GAwG1KtXD06nEwMGDMDzzz+Pvn37xnxtr9eLY8eOoXXr1vj0008x\nevRoTJs2DTqdDq+99hoA4I8//sDIkSMxZswYTJ48OeZ7Xs1IEmYSSVwGwzDYt28fVqxYgS+//BIt\nW7bkrMD4jeuRWtGRodUANDGHM1KITe4QQzyNEsKFFFnu2bMH06ZNQ0lJCbKzs+O+Lj7279+Pv/71\nr2AYBgzD4L777sOTTz4Z83X5Hq+7d+/G448/jvHjxyM/Px9DhgzBTTfdhE6dOuHjjz/GhAkTMH78\n+JjvebUjSZhJJCEClmXx448/oqioCJ9//jmys7ORl5eHAQMGIC0tLeDPBYuwACQ0WUYrTpIiz3BM\n8+WCFFnu378fjz76KIqLi9G8eXPF16EGCFl6PB5s3boVffr0wdatW/Hmm28iJycHBQUFeP/993Hh\nwgWYzWbMnDkTwJXRXkmII0mYSQRAzKLvySefxNq1a2EymdCqVSt89NFHyMjIUHml8QPLsvj5559R\nVFSEsrIyNGzYELm5uRg4cCDS09MD/pyQJAC/oMNisSTcRiSnkjdc03y5IGW+f+DAATz00ENYsWLF\nVT+7kaZpDB48GDfddBPeeustuN1u7NixA/Pnz8ef//xnPPbYY7X+fKKpnuONJGEmEQAxi77PP/8c\nffv2hU6nw1NPPQUAdXagNsuyOHLkCIqLi7Fu3TpkZGQgJycHQ4YMCWhHqKysBICAqRdqpycjgZJt\nL7GO4AoFKbI8dOgQCgsL8emnn6JVq1Yx30freOqpp+BwOPD2229zP3O73di5cydeeOEFTJs2DYMG\nDQJQ2yYvCXEkCTOJWpCy6AP8LiHFxcX4+OOPVViZtsCyLI4fP47i4mKsXbsWqampyMnJQbdu3XDf\nfffh0UcfRUFBASiKEp3pqUXy5BsqxKPtRcq6UGiUEC7IDFQhWR4+fBgTJ07EJ598ctUOOhaS3syZ\nM9GpUyfk5+ejuroaaWlpnAXesWPH0LJlSxVXm5hIWuMlERE+/PBDDB48WO1laAIURaFFixZ44okn\nsHHjRnzwwQc4f/48evbsiSZNmsDpdOLcuXNgWRZ6vR6pqamw2WxdQm3+AAAgAElEQVQcEblcLtjt\ndtTU1ASkcNVCvMkSEDdK8Hq9URlISJHl0aNHMXHiRCxevPiqJUtixsFHu3bt8MYbb+DgwYNc3T0v\nLw9bt27lyFJNW8irCUlrvCRq4eWXX4bJZEJBQYHaS9EcKIoCwzBYvHgxpk+fjgcffBAlJSUoLCyE\n1+vF0KFDkZubi8aNG0Ov13Pjx0h60u1217Kii2fkqQZZCkGMEoh1IYk83W43dDpdUOtCKbI8ceIE\nxo8fj48++ght27aN58eJG/i1x6effhoURWHgwIEYOnQoampqkJ+fj8LCQmzYsAHNmjVDjx49uL+b\naHV1rSKZkq3DEEvJLlq0CB988AG+/PLLq3aQbqwYOnQo7r77bjz++OPcz1iWxYULF7Bq1SqsXLkS\nNTU1GDRoEHJyctC8eXPJgdixTBCJFMR9yOfzadJ9SGggIeyBJWSZmpoaYDxRXl6OMWPG4IMPPqgT\nQ4/vvfde3HDDDUhLS8PHH3+Mxx9/HDk5Odi9ezd++eUXGAwGTJkyBUBS4BMtkjXMJGpBSJhlZWV4\n4oknsGXLFlxzzTUqr067cLlcIQ8TFy9eRGlpKUpKSlBRUYEBAwYgNzcXLVu2rEWe8RhLpnWyFELY\nxkN+RszjybM5deoUCgoKsGDBgjoxFq+kpAQ7d+7EnDlzkJubC7PZDLfbjR49euD+++9H/fr1uT+b\nJMvokSTMJAIgZtH3z3/+Ex6Ph/PZvPPOO/Hvf/9b5ZUmPiorK7F27VoUFxfj7Nmz6NevH3Jzc9G6\ndeu4kKfQ11ZL4qNw4PP5UF1dDYPBAIZhsGjRIvz222/c/Ni33noLd9xxh9rLjBuqq6uxYMECnDt3\nDnPnzsXTTz+NtWvX4vXXX0f//v3VXt5VgSRhJpGEBuBwOLB+/XoUFRXh999/R58+fZCXl4e2bdsG\nEBnfwzUWVWmikyXDMHA4HFwalmVZHDp0CB9//DFWrFgBl8uFe++9FyNGjEDPnj0D6ppy4uTJkxg3\nbhzOnj0LiqLwt7/9jUt7KgWhGpZvNjB79mwcPXoU//d//4fHHnsM9erVw+zZsxVdT11CkjA1ADLx\ngaRJkj1RdRs1NTUoKytDUVERjh49il69eiEvLw9/+tOfavm4EuL0er1hz/S8WsiSpGEJLly4gFGj\nRmHOnDlo3Lgx1wLlcrmwb98+RdZy+vRpnD59Gh07doTD4cCtt96KVatWKSYw4qdTnU4nZ9lI9oxj\nx47hgQceQHV1NRo3bozVq1cDSDr4yIUkYaoIp9MJk8kkWk84deoUmjRposKqktAS3G43PvvsMxQV\nFeHgwYPo0aMH8vLy0LFjx1rkSYgzGHlerWR56dIljBo1Ci+++CJ69+4d8HfsdrvkCDi5kZeXh0cf\nfVQW43Qh+KQ3bdo03HXXXRgyZEiturndbseuXbu455CsWcqHJGGqiLKyMjz77LNo0qQJHnnkEfTt\n2xd6vR4//PADhg0bht9++63WbLok6i48Hg82btyIoqIi7N27F926dUNeXh66dOkiSZ5kLBlJ25K+\nxkQly+rqaphMpgCyrKysxKhRo/Dss8+iX79+qq3vt99+Q8+ePfHTTz/BarUqdp+nnnoK+/fvR2lp\naS0iFGankmQpL5LGBSqiW7du2Lx5M2bNmoVPPvkEJ0+eBOBXvA0ePBgGg4FrZqdpOqGG9caKCRMm\nICsrC+3bt+d+9uyzz6JDhw7o2LEj+vbtyz2vugKTyYSBAwfiP//5D7799lvk5ORg2bJl6N27N2bM\nmIFt27ZxDexkILbNZgsYiO31ejmRTCJBiiyrqqowevRoPP3006qSpcPhwD333IM333xTdrLcuXMn\nZ7V48eJFHDx4EPPnz4der4fH4wHgz0QAtYc9J8kyPkgSpsLYvXs3Hn74YQwYMAD/+9//8MUXX3Dk\nuHbtWgwfPhyAv1WhpqaGa3YnYFlWdWcYJTF+/HiUlZUF/GzGjBnYu3cv9uzZg7y8vDotZjAYDOjb\nty/effddbN++HSNHjsSaNWvQp08fTJ06FVu2bOF6FimKwrJly0BRFCwWC1iWRXV1Nex2O5ee1TII\nWRqNxgCydDgcKCgowBNPPMF5oqoBr9eLv/zlLxg7dizy8vJkvfaFCxewfft2pKSkwOFwIDMzEyzL\n4tKlSwDA9Z1++eWXqKqqkvXeSYSPJGEqiIqKCsybNw9t2rRBWVkZjh8/jhYtWqBRo0b4+eefUVNT\ng759++KPP/7AzJkz0b17dwwbNgzFxcVgGIZLuxB3Ga1veNGge/fuAb1jAALqUA6HI9kTehl6vR49\ne/bE/PnzsX37dowbN44zzJ88eTLy8/OxbNkyrqZpNpths9lgNptFyVNLBzE+WfJrddXV1RgzZgwe\neeQRDBs2TLX1sSyL/9/enYdFWe6PH3/PMAgI6FEEkkVFTMUQJLM0wJVSE0WNREFRj0tRudExNNPQ\nFC0vK06mFi5paulRuMRCMpKKzJXjLqKyBGpFAsoAsg337w+aJ1D0fPslDMv9ui6vZobHee4Jn/k8\n9/b5TJs2jR49etRIWPEwnD59GisrK2bPns3Ro0d57bXXKCoqYvDgwfj7+3P16lWKi4tZtGgRH374\nYZ0OA0sPJifO6pCZmRkFBQUMHDgQCwsLjIyM6NOnD+bm5uzcuZMhQ4ag1Wr54IMPyMvLIzk5mejo\naH7++WfUajW3b98mISGB/v37Y21trbzv6tWr6dWrFwMHDqyzZfSGtmjRIj777DNatmzJ0aNHDd2c\nBsfIyAhPT088PT0pKyvD19eXzMxM2rVrx7x58/Dz82Pw4MGYmJgotSlNTU2VZABFRUX31PQ01Fzn\n/YLlnTt3mDhxIjNmzGDs2LEGaZve4cOH2b59O25ubnh4eACwcuVKhg0b9rfe99atW7z77ru0adOG\ntWvXYmFhgZmZGStWrGD58uXodDqmTJmCnZ0dxcXF7N27F7VaLVfYG4hc9FOHKisrefPNN4mNjcXd\n3Z09e/awbds2AgIC6Nu3LytWrKCyspJDhw4xceJEHnvsMeXvxsfHs3v3bn777Teys7MZO3Ys4eHh\naLVaZsyYgb+/P/7+/kovoTFfPA+qmrJq1SpSU1PZsmWLAVrW8JWXlxMUFIRWqyUmJoYWLVpw9uxZ\n/vOf//Dtt9/i5OTE6NGj8fHxUbYmwP0LP9d3ZZW7h2H15y0pKWHSpEkEBgYSFBRUL20xBJ1Ox9mz\nZ9mwYQMWFhasXr2aS5cusXHjRjQaDeHh4coNjpWVFSYmJsq+XKnuyEU/BqBWq4mIiOD8+fPMmjWL\nyMhIvLy8uHLlCqmpqXh6emJubs6JEyfo3r078Oek/rJly/Dw8OCrr74iJiaG9PR08vPzOXnyJCYm\nJhQVFfHrr78qQ7Z6+qHcpiIwMJATJ04YuhkNVl5eHlZWVsTExGBqaoparaZXr16sWLGCI0eOsGjR\nIlJSUhg5ciTBwcFER0crvUt9ZRULCwtlNe2dO3fQarXcuXOHioqKOv23pB8m1mg0NYJlaWkpU6ZM\nYdy4cU22AIB+MZaRkRFubm7MmjWL/Px85s2bR7du3Zg5cyYqlYrZs2dTUFCAnZ2dksRfBkvDkQGz\nDlWfd+zbty8vvfQS9vb2GBsbExERgampKW3atEGj0XDz5k0ATExMuH37NqmpqcyaNYuKigqcnZ05\ncuQIhYWFpKSkkJaWxqlTp+jbt69yB6qnVqtRqVSNOmheuXJFebxv3z5lCEy6l62tLevXr681t61K\npcLV1ZXw8HAOHz7M8uXLyczMZMyYMQQFBbFr1y4KCgpqBE99WbK6Dp7Vg6WpqakSLMvKypg2bRqj\nRo0iODi4UY+c3E/1fZbXrl1Dq9Xi6urKwoULKSwsZO7cuTg7OzN58mRcXFywsrJS/q5MSmBYcki2\nnlRWVt7TG4SqPJkfffQR69atw9nZmVWrVlFSUsKSJUuU1aNnzpzB19eX7OxsJk2ahJubG/Pnz6eg\noIDnnnuO6OhobGxsOHjwIFevXmXQoEE1MpCsWbMGc3NzXnrppXr9zP8XteW0jYuLIzU1FSMjI5yd\nnVm/fj02NjaGbmqTIYQgLS2NvXv3EhcXR6tWrRg1ahQjRozgH//4R41jdTqdstdT37v5u5VV9MFS\nH6T171NeXs706dMZNGgQISEhTTJYVrdmzRri4+NxdHTE2dmZRYsWkZGRwTvvvINWq2XLli1KKkSZ\nwad+ycQFDcj9NhknJyfj4OCAra0tM2fOpG3btgwaNIitW7fi7u7O888/z7Jly5g7dy6PP/44J06c\nYMyYMVy7do0tW7awbds23N3dSUxMZOnSpYwePRqdToePjw8hISGMGzdOmf+QG50lqApeWVlZ7N27\nl/3792NqasqoUaPw9fWlbdu29y1L9v8bPO8XLCsqKnjxxRfp168fs2bNavLBctOmTWzdupXo6Gjm\nz5/P119/jZ+fH+vXryc9PZ2NGzcSFhZG69atDd3UZkkGzAZKv/ji7uB1/vx5Nm/ezLlz53jllVcY\nPXq00vucOXMmdnZ2zJw5k1atWvHyyy+zatUqhg0bxtixY0lMTCQ8PJzvv/+ec+fOMXHiRJKTk5UE\nCSqVikuXLvHWW2+xePFiXF1dDfTppYZECMH169eJjo5m//79qFQqRo4cyciRI7G2tv7bNT31wVKt\nVmNmZqYcp9PpePnll+nVqxehoaFNMlhWX9VaXl7OV199haenJzt27CAxMZF///vfDBs2DE9PTzZu\n3Kj0KGXP0jDuFzDl7LGB6eeP7ubq6sp7772nPC8vLycnJwcPDw9leDIuLo4DBw5w9uxZbGxslBJH\nWVlZdOvWDYADBw7g6uqKRqNRepeVlZWkp6dTXFwsg6WkUKlUODg4MHv2bGbNmkVOTg7R0dG8+OKL\nlJeX4+vri5+fH4888ghqtVrJ86oPnmVlZRQXF9daluxBwXLOnDk89thjzSZYGhkZMXr0aPLy8khK\nSmLlypV07NiRgQMHcv78eXJzc5V5SxksGxb522igqi8Y0i/5f++99/D390ej0ZCSksJvv/1Gz549\nsbS05NSpU9jb2wNVQ7uurq7k5+fz/fffK9mE9EpKSjh58iRFRUW88847HDlypN4/X0NUW5o+vTVr\n1qBWq8nLyzNAy+qfSqXC1taWkJAQZYtTq1atmD17NiNGjGDt2rVkZWUhhFCCp7m5OZaWlhgbG1Ne\nXk5BQQFFRUWUlpbWGiwrKysJDQ3FycmJsLCwJhks4c8tXxs2bGDChAm8+uqrnDlzhrZt22JsbMx/\n//tf1qxZw82bN4mNjcXKyqrRpTRsLuSQbCNRfWhGP/94/fp17O3tycrKIiQkBDc3NxwdHfnggw9I\nTExU8l7qt6Lo73Szs7Px9vbmhRdeoHPnzkRFRbF582acnJzIy8vDycmpxl1xcxkWSkpKwsLCguDg\n4Bp7QrOzs5kxYwapqakkJycrBbabq/z8fPbt20dMTAz5+fkMHToUPz8/nJyc7qnpWVZWRmlpKUII\nNBoNV65cwc7Ojnbt2vH666/Trl07li5d2iSDZfVr6Pr160yePJl58+Zx8eJFPv/8c7744gtl/lif\nN7Znz54yKUEDcL8h2QcSUoNVWVkphBCioqJCCCFERkaGWLBggViyZIk4ffq0EEKIiIgIERAQUOO4\nyspKERcXJ4YMGSLKy8uFEEIEBweLjRs3ioyMDOHn5ycuXLgghBDizp07Dzx3U5SRkSFcXV1rvObv\n7y/OnDkjOnXqJHJzcw3Usobp1q1bYvv27WLMmDHC09NThIeHi1OnTonCwkKRl5cnXnrpJZGamiq0\nWq3Iz88XoaGhwtLSUri5uYlnn31W3Lhxo17aOXXqVGFjY3PP77auVL9GkpKSxLp168S7776rvBYZ\nGSmeeOIJcerUKSHEn9ea/jqVDOt+MVHOYTZS+hsg/fxnp06dWLlyZY1jsrOz8fX1BVAWFpWXl5OY\nmEi3bt2U/Z8eHh7k5+fj4ODAxYsXlfnPjz76iKysLJYsWVJjL1hzuvvdt28fDg4OuLm5GbopDVLr\n1q0JCgoiKCiIwsJC4uLiiIiIICsrC5VKhaWlJdbW1qjValq0aKH0Jq9du4ZaraZHjx64ubkpKfDq\nytSpU5k1axbBwcF1do7afPPNN7zyyis4ODhQXl6Ol5cXTz75JLNnz1ZWBh86dEjJwiRXrjdsMmA2\nEfoMP9UvuHXr1imP9dlBsrKyiImJYfLkyQCkpKSQkZGBl5cXt27d4plnnuHmzZukpKQQHR1NVFRU\njWAZHx9Ply5d6NKlSz19MsMpLi4mIiKCb775RnntATefzZ6FhQXjxo1j7NixjBs3jmvXrmFtbc3w\n4cMZMGAAY8aMITY2ltLSUnbu3IlaraakpISEhAQyMzPrtG3e3t51fo7qVCoVSUlJREZGEh8fT+fO\nnfnXv/7F7t27EULw1FNPERoaSnBwMObm5vXWLunvkQGzibh7jlH/xa7vDer/a2JiwvDhwzly5AjT\npk0jIyMDX19fBg0aRLt27VCr1Xz88cdotVoCAgLo0aOHsgBBrVZz6dIlvvzyS9auXYtOpyMnJwdb\nW9sa59fXamzs855paWlkZmbi7u4OVGVl6d27N8ePH5eJFO6joqKCSZMmUVJSQlJSEiYmJpSWlnLw\n4EFWrlyJVqslISFB+bdhamqqjII0NdnZ2cTFxTFhwgQ6d+7M4sWLWblyJZs3b0YIgaenJ+3atZNz\nlk2FIceQpbp18+ZNsWHDBrFjx44ar/fr10906tRJxMbGCq1WK4Somo/Rz63MmzdPLFiwQAghREJC\ngpg2bZqIiooSQghx4cIFUVJScs+5Vq9e3WjmZmqbw9STc5j/W3l5uXj33XfvO/9tSA/63daVqKgo\n4eLiIuLi4oQQQhQUFIiwsDCRlpZWr+2Q/hoZMCUhhBA6nU7odLoar+kXKOTk5IgZM2aIoKCgGq9X\nt2DBArF161YhhBBDhgwRmzdvFr///rsIDw8XQ4cOFY6OjuLNN98Ut2/fFkIIsW/fPtGqVSshRNWC\nhrvP3ZCMHz9etG/fXrRo0UI4ODiIzZs31/i5k5OTDJiNmCECphBC7NixQ/Tq1Uvs2bNHCCGUa6Ap\nL55r7O4XE+WQbDOjHwoTf2QY0idr/+yzzzh48CB2dnaEhoYC1MhAJP4YNnryySc5ePAgLVq0QAjB\n1KlT+fTTT0lOTmbHjh1YWVkxePBgfH19eeqpp/j000+V97t7QUND267y+eefP/Dn6enp9dQSqSkJ\nDAxECMGSJUvw9vZW1gTIYdjGp+F8W0n1Sp9hSKVSUVFRweXLl7GwsGDhwoVKqbHqAU4/j5mVlcXW\nrVu5cOECy5cvp7i4mJSUFMaPH4+VlRUlJSW0bt2aoqIiABITE5k6dSrZ2dksWrSI33//XXnPu+c9\nJamuTJgwgaeffprLly/j6OhY7/VVg4KCOHToEDY2NnIlbCMme5gSGo2Gt99+W3kualmEoL/I27Rp\ng4mJCQMGDKBfv35UVFRw9epVevfuDcDFixfp2rUrxsbG/Pjjj1hZWdGhQwf27NnD8ePHsba2Bqq2\nrHh7eyvbNfTvf7+qLpL0d/yv0YP6YGtrC9R+fUmNg+xhSvd40MUcHBzMzZs38fHxAaqC7aOPPsrF\nixcRQrB69WoqKyvx9vZm06ZNjBs3DoDjx4/j5eUFVG1lSUpK4vr16+Tm5jJy5Eh++uknfvnlF2WI\nGKqCp0wRJjU1Mlg2XjJgSn+JPi1f9UCmTxv35JNPYmtry2uvvQZUJX6fMmUK+fn5XL16lcGDBwNV\nwdPW1pann36aL7/8kpMnT/Ltt9/St29f3nrrLQoLC5WN7dWHbYUQzWofZG25bcPDw3FwcMDDwwMP\nDw+lZqokSXVPDslKf4l+6LR6IHN2dlaGvIqLi2nZsiU//PADBQUFdO3alfPnz3Pu3Dm8vb2BqoLY\nHTp0oHXr1nz22WfMnTuXsLAwvLy8WLhwIebm5nzyyScMGDCANWvWKEWNq1e+EH8k/W7KastOo1Kp\nCA0NVRZSSZJUf5r2N45UL6pXVmnZsiU6nY7+/ftz9uxZ5Rhra2s++eQTtm7dyv79++nTpw95eXmk\npaUREhICQE5ODmZmZkyaNImrV69y/vx5MjIygKr6oNu2bePixYtNIinC/4W3tzdt2rS55/Xm1MuW\npIak6X/rSHVOrVbXWPmnf9ylSxcqKytxdXXl/fffJyUlha+//ponnngCDw8Pdu/eTc+ePWnVqhW5\nublkZGQwcOBA2rdvz+3bt8nIyMDFxYUffviBTZs2cePGDaZPn87rr79OSUnJPe3YuXMn77//fr19\nbkP58MMPcXd3Z9q0ady6dcvQzZGkZkMGTKlO6XuCTz31FO+//74S1CwtLfnuu+8YNmwYABkZGdy4\ncUNZNfvFF1/Qv39/bt26RWRkJHFxcQwaNIjY2FiysrK4ffv2PecKCAhg/PjxQFUvTKfTNblFQyEh\nIWRkZHD69Gnat2+vzBdLklT35BymVC+qJ4d/5JFHgKqgqB/KzcnJUSqnAMTExDBx4kSuXLmCg4MD\nQ4YM4YMPPuDkyZNYWlpy/fp1bG1tleQHOTk5BAQEcODAASoqKtBoNE1yv1v1HLbTp09n5MiRBmyN\nJDUvMmBK9aK25PD65AkAzz33HL1798bW1pbi4mKOHTtGVFQU5eXlnDhxgsjISF5++WV0Oh2XLl3C\nzs4OqFq1q1ar2bt3L0ZGRpiamrJ9+3aioqKYOHEirq6u9OvXr94/b1355ZdfaN++PVB1U1F9Ba0k\nSXVLDslKBlHbXjRbW1t0Oh0tW7bkzJkzODo6Ym9vj4eHB7NnzyYpKYnCwkJcXFyUxTD6QLxnzx4C\nAwMB2L59OyYmJpiZmREcHFyjzNndGvJeT312mtTUVBwdHdm8eTNhYWG4ubnh7u7O999/3yzmbCWp\noXjgDtoHJaGVpLqmH269ceMGH3/8Md988w0eHh5ERETQunVrpZd669YtPDw8OHbsGGq1mv79+xMd\nHU337t2JiIigoqKCN954A41Go7xnfn5+rStQpYYpPj6euXPnotPpmD59OmFhYYZuktSEqe6TXUIO\nyUoNlr73aGdnx9KlS1m6dClarRZLS0ugajhWo9Gwc+dOXFxcsLGxYc+ePdjY2NC9e3e0Wi1qtRpj\nY+N7hoRXrVpFcXExNjY29OrVS5kL1Afh2gpyS4ah0+l49dVXSUhIwN7enj59+jBq1ChcXFwM3TSp\nmZFDslKDp9/nKYRQgiVUpeUDuHDhAiNGjABg165dPPvss0DVytvff/+djh07olarlfnOoqIirl27\nRkpKCm3btmX58uXs2rWrRo7Pu7fKSIZz/PhxunTpQqdOnTA2Nmb8+PHs27fP0M2SmiHZw5QavP+V\npOCjjz6isrKS0tJSTExMGDt2LADnzp2joqICd3d34M9yZYcPH8bExISwsDCeeeYZHn30UZYuXUpA\nQABZWVmsXbuWnJwcfH19ef7552vMt8qeZ/27fv06jo6OynMHBweOHTtmwBZJzZXsYUqNWvU0eSYm\nJmzfvp3u3bsrVU8sLS3p2rUr8GdChZMnT9KmTRtlSO/06dMMGDCA06dPs2zZMh5//HFCQkKIiYlR\ncrX++uuvVFRU1Oh56vd6Npep/tpy20JVIgUXFxdcXV3rZG5RJiuXGgrZw5Qatbt7f/rSYGq1msDA\nQMrKyjA2NlYW+xQXF3PhwgXy8vJwcHAA4ODBg4SFhbFx40ZOnz7NmTNnmDt3LkZGRvz0008MGzaM\nFStWcObMGTp27EhgYCDDhw+vsS2mOZRsqi23bWJiIrGxsZw9exZjY+Ma9U4fFnt7e7Kzs5Xn2dnZ\nyu9OkuqT7GFKTUb10mD6nmeLFi1qHHP8+HE0Gg2PPPIIL7zwAqGhoeTl5eHl5UVZWRkrV65ky5Yt\nXLp0ifT0dDp06EBmZiYJCQkMGzYMPz8/ysrKOHfuHHPmzOGTTz6hsLAQlUql9DSrz7k2JbXltl2/\nfj0LFy7E2NgYQKl3+jA98cQTXLlyhczMTMrKyti1axejRo166OeRpP9FBkypSbq7CLX+cVxcHB06\ndGDZsmWMGDGCLl26sH//fszMzOjSpQvx8fG4urry9ttv8+OPPzJx4kQuXbqEk5MTYWFh+Pv7k5mZ\nyRtvvEHnzp359ttvCQsL45dffkGlUnHnzh1l2Lb6+Zta8NS7cuUKP/zwA3379mXgwIGcPHnyoZ9D\no9Gwdu1ahg4dSo8ePQgICJArZCWDkEOyUrOgD15eXl4YGxvj6OjIlClTahzj4+NDaGgofn5+9OnT\nhwkTJmBtbU1ycjJubm4YGRlx6tQpEhMTWbBgAZ6ensyZM4fc3FysrKz44osv2L17N8XFxbzwwgtM\nnjxZqemp74Hq64k2leHbiooK8vPzOXr0KCdOnGDcuHGkp6c/9PMMHz6c4cOHP/T3laS/QvYwpWZl\n1KhRyhevPo+t3uOPP05CQgJTpkzh559/pqysjLy8PJKSkpTi12lpadjZ2eHi4kJpaSk6nQ4rKyti\nYmJYvHgx0dHRREREcOzYMW7cuIFarSY2NlYZttVoNEqw1Gq1aLXa+v0f8JA5ODgoq5L79OmDWq0m\nNzfXwK2SpLohA6bUbN29NaSyshKNRsOYMWOIiorCxcUFc3NzfHx8GDRoEAA9e/YkKysLIQQmJiYY\nGRlx584dYmNjEULg7e3Npk2byMvL48svv+Tq1avMmTOHt956Cx8fH9atW6ek4ktJSWHfvn0UFRXV\n+2d/WEaPHs2hQ4cAuHz5MmVlZVhZWRm4VZJUN2TAlKQ/6Pd7Vl+wY21tzfz585VFLY8++iidO3fG\n39+fyMhIMjMzMTMz48iRIxw9elSZA9VoNAwcOJDvvvuOrl274u/vT2hoKPHx8eTm5nLixAnmzZvH\nkSNHMDc3N9hn/iv0uW0vX76Mo6MjW7Zs4Z///Cfp6en07G0pmCkAAAFASURBVNmTCRMmsG3bNkM3\nU5LqjMwlK0kPoN+Ocrevv/6aH3/8kQEDBjB48GCmTp1KYGAgQ4cOrXHciBEjePHFFxkxYgRGRkYM\nGTKExYsX06JFC4KDg9HpdIwcOZIFCxYoFVgkSTKs++WSlSTpr7nfheQDJAM/AeFAR8AFOAB0/uMY\neyANMPnj+EigDdAeaBzdTElqxuQqWUn6a/SjLuo/HuufJwC9AU+gK3ADmA/k/PEHwA84A5QCfYFC\nIL9eWi1JkiRJDURtyWWdATf+XCvwX2Ay0BrYAOj3ScgbV0lqBOSFKkkPh36PihrQV6ROq/ZzDbAL\n2EtVD7MboE9DVFEfDZQk6e+RJRck6eGqvlCu+nxnJXAYKKcqqLYDJgC3gdR6a50kSZIkNXAqal8w\nZFLfDZEkSZKkxkKFnA6RpEbn/wFnNYMMVynu2gAAAABJRU5ErkJggg==\n", + "text": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 2, + "metadata": {}, + "source": [ + "Splitting into training and test dataset " + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top]](#Sections)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "It is a typical procedure for machine learning and pattern classification tasks to split one dataset into two: a training dataset and a test dataset. \n", + "The training dataset is henceforth used to train our algorithms or classifier, and the test dataset is a way to validate the outcome quite objectively before we apply it to \"new, real world data\".\n", + "\n", + "Here, we will split the dataset randomly so that 70% of the total dataset will become our training dataset, and 30% will become our test dataset, respectively." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "from sklearn.cross_validation import train_test_split\n", + "from sklearn import preprocessing\n", + "\n", + "X_train, X_test, y_train, y_test = train_test_split(X_wine, y_wine,\n", + " test_size=0.30, random_state=123)" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 9 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Note that since this a random assignment, the original relative frequencies for each class label are not maintained." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "print('Class label frequencies')\n", + " \n", + "print('\\nTraining Dataset:') \n", + "for l in range(1,4):\n", + " print('Class {:} samples: {:.2%}'.format(l, list(y_train).count(l)/y_train.shape[0]))\n", + " \n", + "print('\\nTest Dataset:') \n", + "for l in range(1,4):\n", + " print('Class {:} samples: {:.2%}'.format(l, list(y_test).count(l)/y_test.shape[0]))" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "Class label frequencies\n", + "\n", + "Training Dataset:\n", + "Class 1 samples: 36.29%\n", + "Class 2 samples: 42.74%\n", + "Class 3 samples: 20.97%\n", + "\n", + "Test Dataset:\n", + "Class 1 samples: 25.93%\n", + "Class 2 samples: 33.33%\n", + "Class 3 samples: 40.74%\n" + ] + } + ], + "prompt_number": 10 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 2, + "metadata": {}, + "source": [ + "Feature Scaling" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top]](#Sections)" + ] + }, + { + "cell_type": "heading", + "level": 3, + "metadata": {}, + "source": [ + "Standardization" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Another important procedure is to standardize the data prior to fitting the model and other analyses so that the features will have the properties of a standard normal distribution with \n", + "\n", + "$\\mu = 0$ and $\\sigma = 1$\n", + "\n", + "where $\\mu$ is the mean (average) and $\\sigma$ is the standard deviation from the mean; standard scores (also called ***z*** scores) of the samples are calculated as follows:\n", + "\n", + "\\begin{equation} z = \\frac{x - \\mu}{\\sigma}\\end{equation} \n", + "\n", + "Standardizing the features so that they are centered around 0 with a standard deviation of 1 is especially important if we are comparing measurements that have different units, e.g., in our \"wine data\" example, where the alcohol content is measured in volume percent, and the malic acid content in g/l. " + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "std_scale = preprocessing.StandardScaler().fit(X_train)\n", + "X_train = std_scale.transform(X_train)\n", + "X_test = std_scale.transform(X_test)" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 11 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "f, ax = plt.subplots(1, 2, sharex=True, sharey=True, figsize=(10,5))\n", + "\n", + "for a,x_dat, y_lab in zip(ax, (X_train, X_test), (y_train, y_test)):\n", + "\n", + " for label,marker,color in zip(\n", + " range(1,4),('x', 'o', '^'),('blue','red','green')):\n", + "\n", + " a.scatter(x=x_dat[:,0][y_lab == label], \n", + " y=x_dat[:,1][y_lab == label], \n", + " marker=marker, \n", + " color=color, \n", + " alpha=0.7, \n", + " label='class {}'.format(label)\n", + " )\n", + "\n", + " a.legend(loc='upper left')\n", + "\n", + "ax[0].set_title('Training Dataset')\n", + "ax[1].set_title('Test Dataset')\n", + "f.text(0.5, 0.04, 'malic acid (standardized)', ha='center', va='center')\n", + "f.text(0.08, 0.5, 'alcohol (standardized)', ha='center', va='center', rotation='vertical')\n", + "\n", + "plt.show()" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "display_data", + "png": "iVBORw0KGgoAAAANSUhEUgAAAmQAAAFXCAYAAAAMF1IiAAAABHNCSVQICAgIfAhkiAAAAAlwSFlz\nAAALEgAACxIB0t1+/AAAIABJREFUeJzs3Xl4VNX5wPHvTPY9gRBAdkWtgFaUhpZKResO7guKWBHr\nXpqfWhWtggu0KloVEHFhE2QTEaQoIAgiRQmyb5LIGgIh+77Mdn5/nAlJyMIkmcmdybyf57lPZu7c\nufedgbx57znnngtCCCGEEEIIIYQQQgghhBBCCCGEEEIIIYQQQgghhBBCCCGEEEIIIYQQQrS4AKMD\nED7pa8AM7HTztkIIIYQQrVoxUORcHEBptef3GBhXUw1Cf47Kz5AGLAD6NWIfLwOz3R2YgccRQmju\nznfrgAcbeL07NfNRBrAMuKoRxxgB/NCE2BqrpY4jGslsdACixUQCUc7lCDCk2vN51bYLbPnQmiyd\nqs/we+AXdKK50sighBCGczXfuUq5uF2M8xgXAd8CXwL3N+F4Qgg/cYiqomUQcAx4FjgBzAJigf8C\nmUAu+kyvU7X3r6PqbHEEsAGY4Nz2IHBdE7ftAawHCtHJ7H3qb1kahG4VO90kYHO15+8BR4EC4Gfg\nMuf664AKwII+o93mXP8AsNcZwwHg4Wr7ikd/L3lAjjNWk/O1s4Av0N/ZQWDUGY4jhGgZ1fOdGRgN\n/Apko1vV45yvhQJznOvzgGQgARgP2IAy9O/wxDqO0R3dQnZ6I8fT6NaySpXHLgT2ALc411/g3L/N\neYxc5/rB6JxRgM5jY6vtq754QReG04Dj6Pz+mjO2+o4jhDDI6QWZFfg3EIT+JW8D3Op8HAksRJ/p\nVVoLjHQ+HoEuNh5EFyePoluumrLtj8Cb6Fa6P6KT0Kf1fIZB1F2QXQnYgTDn83vRCdcMPIUuOoOd\nr42tY/83oAtDgD8BJcDFzuf/Bj5Aj70McMaIc99bgBedsfdAF3PXNHAcIUTLqJ7vkoCN6BOoIGAq\nMNf52iPAV+i8ZwL6olu7oGYeq0t36i7IznauP9/5/A6gg/PxXeiu1fbO5/dTuyvxcqC38/GF6OLu\nZhfi/RKdq8KAdsAmqk4u6zqO8ALSZSlAJ4yx6MKsHH3W9KXzcTHwL3RiqM8R9NmYQhceHak6U3N1\n267o8V9j0Gdv/0MnG1Pdu6nXced7Yp3PP0OfPTqA/wAhVCVHUx37/xqdwEG3gK1CF2agi8mO6ORr\nd8YI8Dt069k4Z+yHgE+Auxs4jhCi5T2CPnE6js53r6CLpAD073db4Fx0ftqGbkWq1JTf4ePOn22c\nPxdR1WK2EEgF+jew/+/RLWkAu4D5VOXi+uJtD1wPPIluDcsC3qVmPhJeSAoyAfoX1lLteTjwIXAY\n3Ur1PboJvL5f5OpN8qXOn5GN3PYsdCFYXu31ulrAzqQTOjnlO5//A90FmY8uzGLQxVN9rgd+QndJ\n5qFbzNo6X5uA7m5YhW4Be865vpsz/rxqy/PUX5QKIYzRHX2yWfl7uhd9EpWAHh6xEl30pANvUHNM\nravjyKqrHOpR2TX4F3ThVHn8PlTll7r0R7fOZaJz2CPVtq8v3m7o1r8T1Y4zFd1SJryYFGQCaiea\np4HzgER0AXM5nm/lOYE+iwyrtq5rE/ZzK7r7sAwYCDwD3IluMYtDF5iVn+P0zx2CHgf2JjpBx6Fb\nzCq3L0YXeOcAN6G7QK9Ej+045Ny+colGDyQG3TonhDDeUfS4zuq/q+Ho/GMDXkV3EQ5A//7+xfm+\nphRjoPPRSWA/ulD6CHgCnevigN3Un49Ad6cuATqjc9hUqv5u1xfvUfS41bbVPmMMusuzOZ9FeJgU\nZKIukeiCpgCdOMY2vLlbHEEPun8ZfXb3B3SCcSV5mNBnomPR49NecK6PQietbPS4sTHoQqlSBvqM\nuTIhBjuXbHQRdT1V48BwxtPTuX0hutvSjh5MW4S+MCIM3f3Rh6opOE6edhwhhDGmoodgVJ7stUOf\nXIEel3oh+ve3CN2laXe+dhJ9InYmlb/j7YG/oXPO8851Eeh8lo3+2/sAOk9UOokuvIKqrYtEt3BZ\n0CfIw6jKifXFm4Fuxf8POgeanbFXDr2o6zjCC0hBJqB20fMuurDIRg+A/aaObaq/9/TXmrrtvehC\nLAd9VdACanalnv6+s6ia9ycZfaZ4ObDauc0K55KC7n4tQ589Vvrc+TMHXQwWAX9Hj+3IRc9XtLTa\n9j3RV38Wob+X99HduQ50sXYx+grLLPSZcGXxd/pxhBDGeA89NnUV+qTqR3ShA3qw/efoE9G96CvE\nZ1d73x3ovPBuA/vPR7ek70S3xN0BzHS+thd423nMDHQxtqHae9egx4tloLsoAR5Ht4IVAi+hc2Kl\nhuL9C/rkcq8z5s+pupigruMIUa8AdD/7MqMDEYZaQMu0zgkhhBCG8tYWsiR0ZS993f6lH7pp3Yzu\nLrwJPX5CCCGEEC2sM7rL6QqkhczfDEF3KZagZ92XGa6FEEIIg3yOnuDucqQgE0IIIYQf8Lb7Fg5B\nDzLchr6CpE7nnHOOOnDgQEvFJIQw3gH0RRU+T/KXEH7pjDnM28aQDUCPGzqEvgHsldRxy5kDBw6g\nlPL6ZezYsYbHIHFKrK0hTlybcsAnSP7y31glTv+MUynXcpi3FWQvAF3Q9wK8G/iOqon5hBBCCCFa\nJW8ryE4nV1kKIYQQotXztjFk1X3vXHzWoEGDjA7BJRKn+/lKrL4Sp2h5vvR/w1dilTjdy1fidJWv\n3spFOftkhRB+wGQyge/mq9NJ/hLCz7iSw7y5hazR2rRpQ15entFhtApxcXHk5uYaHYYQfkPyl3tJ\nDhO+xlfPOOs8wzSZTMiZp3vIdym8iT+0kMnvnHvJ9ym8iSs5zNsH9QshhBBCtHpSkAkhhBBCGEwK\nMiGEEEIIg0lBJoQQQghhMCnIDDZz5kwGDhxodBhCCNFokr+EcB8pyPzM5MmT6devH6GhoTzwwANG\nhyOEEC6T/CVaM78vyCwWmDEDysr084wMmD/f2Jg8qVOnTrz00kuMHDnS6FCEEG6wahXs26cfOxww\nezbk5xsbk6dI/hKtWasvyA4cgHXrqp6vW6fXVQoMhOJiePllOHwY/vlPiI6uuY+KCp3oKlUWb42R\nlpbGbbfdRkJCAvHx8YwaNarO7ZKSkujatSsxMTH069ePDRs2nHotOTmZfv36ERMTQ4cOHXj66acB\nKC8vZ/jw4cTHxxMXF0diYiKZmZl17v/WW2/l5ptvpm3bto3/EEKIFmWx6BNEq1U/P3kS/vvfmtu0\nbQvjx8PevfDuu7B/P4SF1dymes6y2/V+G0PylxCe1+oLspAQmDkTvvtOLzNn6nWVzGZ44gkICoJR\no+CGG/RS3dy5MHGiLsqOH9fbHzvmegx2u50hQ4bQo0cPjhw5Qnp6Ovfcc0+d2yYmJrJjxw7y8vIY\nNmwYd955JxZn9kxKSuLJJ5+koKCAgwcPMnToUABmzZpFYWEhx44dIzc3lw8//JCw0zPyaWTCRCG8\nn9kMBw/C669Dero+YTzdpZfC//0fPPccbNwIL71UM8cdOqRz28mTuhh7+21YtMj1GCR/CdEyWn1B\n1rkzjBsH77yjl3Hj9LrqMjN1oQWQnFy7BeyeeyArC55/Hl54Ae6+u/Y+GpKcnMyJEyeYMGECYWFh\nhISEMGDAgDq3vffee4mLi8NsNvPUU09RUVHB/v37AQgODiY1NZXs7GzCw8NJTEw8tT4nJ4fU1FRM\nJhN9+/YlKiqqwZicswYLIbxYYCA8+yzk5MCjj8LgwTBkSM1tHA5Yv14/rizgquvRA265BUaP1ktJ\nCdxxh+sxSP4SomW0+oIMICWl7seguyNfekknqKVLdaH15ps1twkNhb/+VXcJ5OTAVVc17vhpaWl0\n69YNs/nMX/dbb71Fr169iI2NJS4ujoKCArKzswGYNm0aKSkpXHDBBSQmJrJ8+XIA7rvvPq699lru\nvvtuOnXqxHPPPYfNZmvwOHKGKYRvyMmpGhO2e3dV92WlWbMgN1e3ej33nO6+PHmy5jbXXw/Z2fDL\nL/DQQxAc7PrxJX8JIRqi6lLX+uRkpe6/X6m0NL3cf79eV11aWtVju12pY8dqvp6ertSIEUp99ZVS\nL7yg1Dvv6O1ctXHjRpWQkKBsNlut12bMmKEuu+wypZRS69evVwkJCWr37t2nXo+Li1Nr1qyp9b5F\nixap0NBQVVpaWmP94cOHVa9evdS0adMajOnFF19UI0aMqPf1+r5jIYwAtKa/wPV+xtOVlCj14INK\nLVumlNWq1PjxSk2YUHObzEylysurnh87ppTDUfXcZlPqjTeUGjNGqcWL9f4yMlz/7n0xfyklOUx4\nF1zIYa2+hax3b/jXv3TLV+fO+nHv3jW3qd79aDZDp041X9+wQXdb3nijbk0rKdFXY7qqf//+dOzY\nkdGjR1NaWkp5eTkbN26stV1RURGBgYHEx8djsVh49dVXKSwsPPX6nDlzyMrKAiAmJgaTyYTZbGbt\n2rXs2rULu91OVFQUQUFBBAQE1BmL3W6nvLwcm82G3W6noqICu93u+ocRQrSY8HA9VGLIkKruy9O7\nG9u1qzlmrFMnqN6jl56uuzX/+U+49Vbdffnjj67HIPlLCNGQeitQb3X06FF1yy23qLZt26r4+HiV\nlJSklFJq5syZauDAgUoppex2uxo5cqSKjo5WHTt2VG+++abq0aPHqTPM4cOHq4SEBBUZGan69Omj\nli5dqpRSat68eer8889XERERqn379iopKUnZ62nCGzt2rDKZTDWWV155pdZ23vxdCv+Dn7aQeQtf\ny19Keff3KfwPLuQwXx0Z6fx8NZlMJhlb4CbyXQpv4hzE7av56nSSv1qAfJ/Cm7iSw1p9l6UQQggh\nhLeTgkwIIYQQwmBSkAkhhBBCGEwKMiGEEEIIg0lBJoQQQghhMCnIhBBCCCEMJgWZEEIIIYTBpCAT\nQgghhDCYFGQGmzlzJgMHDjQ6DO9nt8OxY5CZCTLZoxBeQfKXEO4jBZkfsVgsPPjgg3Tv3p3o6Gj6\n9u3LihUrjA7rzPLz4YEH4Pbb9U39xo/XN+cTQvgNn81fQrhICjKAAwdgxAi46ir4xz8gN9foiDzC\nZrPRtWtX1q9fT2FhIePGjeOuu+7iyJEjRofWsLffhn379F2U27WDL7+Eb74xOiohvEN5Ofz733D1\n1XDnnbB5s9EReYTP5i8hXNT6CzKlYNkyuOMOvSxbVrPLq6AAHn0UUlIgIAC+/x6eeqp2t1heHmzc\nCNu3N6l1Ji0tjdtuu42EhATi4+MZNWpUndslJSXRtWtXYmJi6NevHxs2bDj1WnJyMv369SMmJoYO\nHTrw9NNPA1BeXs7w4cOJj48nLi6OxMREMjMza+07PDycsWPH0rVrVwAGDx5Mjx492Lp1a6M/T4va\nuxeio8FkArNZ/zvt3290VEK0jMOHdY668UZ47TUoLq75+oQJsGiR/v04eRL+/nc4eLDmNnY7bNsG\nP/6oW5wbSfKXEJ7X+guy1avhlVcgJ0cvL78Ma9ZUvZ6SohNc27YQHAzt28OePboAq5SaqrvLnnoK\nHnpIt6LZbC6HYLfbGTJkCD169ODIkSOkp6dzzz331LltYmIiO3bsIC8vj2HDhnHnnXdisVgAneye\nfPJJCgoKOHjwIEOHDgVg1qxZFBYWcuzYMXJzc/nwww8JCws7Y1wnT54kJSWF3r17u/xZDNGzJxQV\n6ccOh/7ue/QwNiYhWkJ+vs4527dDWRksXQovvFBzm9WrdctxSAjExIDVqouvShYLJCXBww/Dk0/q\nVrTDh10OQfKXEC2j9RdkK1boRBUZqZfQ0JrdXWFh+uyxstXLZtNnmtUTwmuv6W6Bdu10wfb99/Dd\ndy6HkJyczIkTJ5gwYQJhYWGEhIQwYMCAOre99957iYuLw2w289RTT1FRUcF+Z2tQcHAwqampZGdn\nEx4eTmJi4qn1OTk5pKamYjKZ6Nu3L1FRUQ3GZLVauffeexkxYgTnnXeey5/FEP/4B3TtCllZernq\nKt1aIERrt2ePPhlp107npA4ddEt9aWnVNhERUFFR830REVWPV67U72nfXu+nuFh3cbpI8pcQLaP1\nF2SRkTVbs6xWva5Sr14waBBkZMDx45CdDY8/XrMgO34cKhOEyaR/1tGkXp+0tDS6deuG2Xzmr/ut\nt96iV69exMbGEhcXR0FBAdnZ2QBMmzaNlJQULrjgAhITE1m+fDkA9913H9deey133303nTp14rnn\nnsPWQAuew+HgvvvuIzQ0lMmTJ7v8OQzTrh189hnMnAnz58Prr0NgoNFRCeF5oaH6ZLFyCIXNprvs\ng4Kqtnn2WV2gHT+ul/PPh8svr3o9I0PnrcrcFRmpr1h2keQvIURDVF3qXH/ggFIDByp1ySV6GThQ\nr6vOZlNq1SqlZs9W6qefau/jmWeUuvRSpQYPVuqaa/TjzZvrjKEuGzduVAkJCcpms9V6bcaMGeqy\nyy5TSim1fv16lZCQoHbv3n3q9bi4OLVmzZpa71u0aJEKDQ1VpaWlNdYfPnxY9erVS02bNq3OWBwO\nhxoxYoS68sorVXl5eb0x1/cdC6XyyvLUv9b/S1ntVqND8RtAa5rrpN7PWIvVqtQjjyjVt6/OO5dc\nolRdv9t79+r89dVXSp2WE9SGDfq9116rc9gllyj10ksuf/e+mL+UkhwmvAsu5LDW30J29tkwZ45u\n9Xr8cd3ScvbZNbcJCNBXKA0fDv37197HCy9A3756wGxBgR6H0a+fyyH079+fjh07Mnr0aEpLSykv\nL2fjxo21tisqKiIwMJD4+HgsFguvvvoqhYWFp16fM2cOWVlZAMTExGAymTCbzaxdu5Zdu3Zht9uJ\niooiKCiIgICAOmN57LHH+OWXX/jqq68ICQlx+TOIKvN3z2fG9hmsObjmzBsL0RyBgTBxIrz0Ejz4\nILz3np4C5nQXXKDz14031mzdBxgwQOe+vDydw37/e3jmGZdDkPwlhGhIvRWoxzgcShUWKlVR0aS3\nHz16VN1yyy2qbdu2Kj4+XiUlJSmllJo5c6YaOHCgUkopu92uRo4cqaKjo1XHjh3Vm2++qXr06HHq\nDHP48OEqISFBRUZGqj59+qilS5cqpZSaN2+eOv/881VERIRq3769SkpKUna7vVYMhw8fViaTSYWF\nhanIyMhTy9y5c2tt69Hv0oflleWpAZ8MUAOmDVCDPxssrWQtBH9tIXOnigqlioqa9FZfy19KSQ4T\n3gUXcpjJ46nHM5yfryaTyURd60XjyXdZt6k/T2X6tum0j2xPRnEG464Yx7U9rzU6rFbPpMc/+Wq+\nOp3krxYg36fwJq7ksNbfZSmEm+SX5zNj+wwcykFOaQ4Wu4VJyZOwOVyfAkUIIYSoizdeqhYKfA+E\nAMHAUuB5QyMSArDYLVzf83qsduupdRHBEdgcNgLN3virJIQQwld4axdAOFCKLhg3AP9w/qwkTf4e\nJt+l8CbSZSkay5Xv85vUbyi3lXPrBbe2UFTCX7mSw7z1tL5y1sNgIABonTeXFEIIYYgyaxlvbnwT\nu8POVWdfRVRIw5PRCuFp3jqGzAxsB04Ca4G9xoYjhBCiNflq/1cUW4opt5Xzxb4vjA5HCK9tIXMA\nFwMxwEpgELCu+gYvv/zyqceDBg1i0KBBLRWbEMLD1q1bx7p164wOw2MkfxmrzFrG1C1TiQnR86FN\n3zad2y+4XVrJhNs0JYf5wpiMl4Ay4K1q62QMhofJdym8iYwhE43V0Pe5cM9CxqwdQ2xoLKCvoB59\n2WhGXDyiBSMU/sRXx5DFAzYgHwgDrgZeMTQiIYQQrUa78Hb85bd/qbHurKizDIpGCM0bzzgvBGah\nx5GZgdnAhNO2aTVnmDNnzmTatGn88MMPRodSgy9+l6L1khYy7+St+Qt88/sUrZevTgy7C7gEPYbs\nImoXY6IZhg8fTseOHYmOjubss89m/PjxRockhBAukfwlWjNvLMgMcbzoOEnfJGGxW4wOxaOef/55\nDh06RGFhId988w2TJk1ixYoVRoclhGgGpRRj145lR8YOo0PxKMlfojXzm4Js8b7FLN63uN7XZ2yb\nwfLU5az4tf5f7qMFRym3lTfp+Glpadx2220kJCQQHx/PqFGj6twuKSmJrl27EhMTQ79+/diwoWo+\n3OTkZPr160dMTAwdOnTg6aefBqC8vJzhw4cTHx9PXFwciYmJZGZm1rn/3r17Exoaeup5YGAgCQkJ\nTfpMQoiWkV6YzovfvYjdYa/z9R0nd7Bo3yLe+emdervpSq2lpBemN+n4kr+E8Dy/KMiKLcW899N7\nvPfTe5RYSmq9frzoOF/t/4pO0Z14f/P7dbaSVdgqeGTZI8zaPqvRx7fb7QwZMoQePXpw5MgR0tPT\nueeee+rcNjExkR07dpCXl8ewYcO48847sVh0PElJSTz55JMUFBRw8OBBhg4dCsCsWbMoLCzk2LFj\n5Obm8uGHHxIWFlZvPI8//jgRERH07t2bF198kUsuuaTRn0kI0XKmbZvGwj0LWX9kfa3XlFJM2TyF\n6JBo9mTuYcuJLXXu46MtH/HY8scafe9VyV9CtAy/KMgW71tMqa2UUltpnRMAztg2A4UiOiSa3NLc\nOlvJvk79mhPFJ/h056fkl+c36vjJycmcOHGCCRMmEBYWRkhICAMGDKhz23vvvZe4uDjMZjNPPfUU\nFRUV7N+/H4Dg4GBSU1PJzs4mPDycxMTEU+tzcnJITU3FZDLRt29foqLqn09nypQpFBcXs3r1al58\n8UWSk5Mb9XmEEC3nWOExlqcsJyEigYmbJtZqJdtxcgfbTmyjbVhbggODmZw8uVYrWXZpNgt2LyCt\nII1VB1Y16viSv4RoGa2+ICu2FDNt6zRiQ2OJDY1l2tZpNVrJThSd4PO9n2N1WDlRdIJSWymTkyfX\nOIussFUwZfMU2oa3xWK3sGD3gkbFkJaWRrdu3TCbz/x1v/XWW/Tq1YvY2Fji4uIoKCggOzsbgGnT\nppGSksIFF1xAYmIiy5cvB+C+++7j2muv5e6776ZTp04899xz2GwNnwWbTCYGDRrEnXfeybx58xr1\neYQQLWf6tukoFLGhsRwrPFarlWzK5ikUVBSQUZxBua2cTcc2sfXE1hrbzNk5B7uyExsWWyu/nYnk\nLyFahjfOQ+ZWi/ctJqMkg5iQGAAKygtYvG8x9/32PgBCAkMYlTgKRdUZZUhASI19fJ36Nfnl+XSI\n6kCgOZBPd37K0D5DT00qeCZdunTh6NGj2O12AgIC6t3uhx9+YMKECXz33Xf07t0bgDZt2pw62+3Z\nsydz584F4IsvvuCOO+4gNzeXsLAwxowZw5gxYzhy5Ag33HAD559/PiNHjjxjbFarlbZt27r0ObyV\nzWGj1FpKdEi00aEI4Vbpheks3reYAHMAmSWZlNpKmbhpIpd3vxyzSRdIN/S8gcROiTXe1yaszanH\nla1jbcLaEBQQREZRBqsOrOKGc29wKQbJX0K0jFZfkHWN6crDlz5cY12XmC6nHrcJa8ODlzzY4D6W\n/LIEh3KQWawHmlodVjYc3cCQ84a4FEP//v3p2LEjo0eP5pVXXsFsNrN169Zazf5FRUUEBgYSHx+P\nxWLh9ddfp7Cw8NTrc+bM4dprr6Vdu3bExOhbfpjNZtauXUt8fDy9evUiKiqKoKCgOhNnVlYWa9as\n4cYbbyQ0NJTVq1fz+eefs3r1apc+h7f6dMenrDm4htm3zT71R0qI1iDAHMCIi0dgV1XdlJFBkTW2\nueWCWxrcx7rD66iwV5BXlgeAXdlZ+stSlwsyyV9CiIaoutS3vrmKK4pVVklWjcVmtzVqH0ePHlW3\n3HKLatu2rYqPj1dJSUlKKaVmzpypBg4cqJRSym63q5EjR6ro6GjVsWNH9eabb6oePXqoNWvWKKWU\nGj58uEpISFCRkZGqT58+aunSpUoppebNm6fOP/98FRERodq3b6+SkpKU3W6vFUNWVpa6/PLLVWxs\nrIqJiVG/+93vTu3jdJ76Lt2tsLxQXTb9MnXxBxerH478YHQ4wkOA1jTDZ72f0ROsdmut/FViKWnU\nPnwtfynlOzlM+AdcyGG+OvO18/PVJDMzu4+vfJfTt03ng80fEBoUSsfIjsy/Y760krVCMlO/aCz5\nPoU38dWZ+oVwSVFFETO2zyAuLI6YkBgO5R1iY9pGo8MSQgghGk0KMuGzvj34LYXlhRRbiskty8Xi\nsDBn5xyjwxJCCCEazVe7AKTJ38N84bssqigirTCtxrrY0FjOijrLoIiEp0iXpWgs+T6FN3Elh/lq\ngpOE5mHyXTZPWkEas3fO5vnLnq/8RRTNIAWZaCz5PoU3kTFkQhjk460f8+mOT0lOl1nEhRBCnJkU\nZEK42ZH8I6z4dQVtwtowcdNEOUsXQghxRq1qYti4uDjpHnKTuLg4o0PwWdO2TcNkMtEmrA0pOSkk\npyfTv3N/o8MSXk7yl3tJDquilJL/Wz7AV/+F6hyDIYTRjhYcZcjcIYQGhmI2mSmyFHFx+4uZc9sc\nSYjN4A9jyITwhIV7FnIw7yCjLxttdCh+zZUc1qpayITwBnf3uRuHcpx6HhMSI8WYEKLFlVhKeH/z\n+5RaShl+0XA6R3c2OiTRAF/9KyFnmEL4EWkhE6Lx5uycw3ub3sOMmRvOvYGxg8YaHZLfkqsshRBC\nCD9UYinh460fExsaS9vwtixPXc6xwmNGhyUaIAWZEEII0cosS1lGRlEGeWV5ZJZkkl+ez6zts4wO\nSzTAV7sApMlfCD8iXZZCNM7+7P3szdpbY12XmC70O6ufQRH5N7+bqV8I0TpJQSaE8GUyhkwIIYQQ\nwgdIQSaEEEIIYTApyIQQQvgspRT55flGhyFEs0lBJoQQwmdtSt/EHQvvkKJM+DwpyIQQQvgkpRST\nNk0irTCNhXsWGh2OEM0iBZkQQgiftCl9Eym5KXSP7c6nOz6loLzA6JCEaDIpyIQQQvicytaxkIAQ\nggOCsdgtLNizwOiwhGgyKciEEEL4nN2Zu9mXvQ+7w05OaQ4O5WD+7vnYHLYWj2Vz+mayS7Nb7HhW\nu7XFjiWT7gmwAAAgAElEQVRajq9OtCgTKwrhR2RiWHE6q91KSk4KiqrvMjQwlHPizqn8/9IiiiqK\nuOGzG7jq7Kta5ObduWW53L/kft677j3Ojjvb48cT7iETwwohhGiVggKC6J3Qmz4JfU4tPdv0bNFi\nDGDR3kWUWEtYnrqctII0jx9v3q557M3ay8dbPvb4sUTLkoJMCCGEaIKiiiKmb59OfHg8JkxM3zbd\no8fLLcvls12fcXbc2aw5tIaDeQc9ejzRsqQgE0IIIZpg0d5FlFhKCDAHEBMaw7KUZR5tJZu3ax5W\nh5XQwFBMmKSVrJWRgkwIIYRogp2ZO4kKiaLUWkqFvYKo4Cj2Zu31yLFKLCXM2z0Pu8NOVkkWDuVg\n5YGVpBeme+R4ouX56iBZGRQrhB+RQf3C39kddjalb6pxhaXJZCKxUyKhgaEGRiZc4UoO88YE1wX4\nFEgAFPARMPG0bSShCeFHpCATQvgyXy3IOjiX7UAksAW4BdhXbRtJaEL4ESnIRJPt2QNvvw3Z2TBo\nEDzxBISEGB2V8DOu5LDAM+wjAbgT+BPQHd1idQRYD3wOZDY3yDpkOBeAYnQhdhY1CzIhhBCiYenp\n8Oij4HBAWBh89hmUlsKLLxodmRC1NDSofxqwEN1KNRW4H3gA+BCIcr72iYfj6w70BTZ5+DhCCCFa\nm61bobwc2rTRBVlCAnz9tdFRCVGnhlrIJgI76li/D/gOeB24yBNBOUUCi4AkdEtZDS+//PKpx4MG\nDWLQoEEeDEWImvLL83nt+9d49YpXiQiOMDqcVmfdunWsW7fO6DA8RvJXCwkJgerdw1arLsyE8LCm\n5DBvHZMRBPwX+AZ4t47XZQyGMNSHP3/IhI0TeGXQK9x70b1Gh9PqyRgy0SRlZfDAA5CSAmZnh9DY\nsXDjjcbGJfxOcwf172rgNYXnWsdMwCwgB3iyvuP7bULbvRumT9eJ5pZb4Jpr4PRbhZSX6+2Ugj59\n5IzQzfLL8xk8dzCB5kBMmFg+bLm0knmYFGSiyUpKYNkyyM+Hfv30IkQLa+6g/spTiMedP2c7d+bp\n5oA/AsOBncA257rngRUePq73S0mBhx7SjwMDYfNm3QQ/ZEjVNgUF8PDDcPiwft6tG3z0EcTGtni4\nRrLarWw+vpkBXQa4fd8Ldi/AYrfQJqwNGcUZLPllibSSCeGtIiLg7ruNjkKIM2poUP9h53IN8Cy6\nxWwn8JxznadscMZ1MXpAf1+kGNNWrgSLBeLjdYEVEQHz59fcZvp0OHBAD15NSICDB+Fj/7u9xsoD\nKxn19ShSc1Ldut/CikJmbJ+B1W4loziDClsFH235iDJrmVuPI4QQwr+cadoL0K1il6ELJdAtWK2l\n68C3BATU7J5USq+r7uhRCK02a3NoqF7nR6x2K+8nv4/FYeGjLR8x4ZoJbt3//b+9H6ujarbs0MBQ\nFNIFJYQQoulcKchGAjOAGOfzfPT0F6KlDR6sW8QyMnQhZrfDyJE1t7nkEvj+e4hx/nOVlcGll7Z8\nrAZaeWAl2aXZdI3pyrrD60jNSeXctue6Zd/RIdE89rvH3LIvIYQQolJjWrpinNvneyiWxvDfQbGH\nDumirKxMF2j9+9d83WaD8ePhv//VLWiDB+tJEIOCjIm3hVntVm6adxNltjIigiPILsnmT93+5PZW\nMtGyZFC/EMKXuevWSR2A8UAn4DqgF/AH9MSxRpGEdiYlJfpnhH9d/ZeSk8Jfv/orFbaKU+vahrfl\ny6FfEhIot0vxVVKQCSF8mbsKshXoLst/oqe6CEJf/dinmfE1hyQ0IfyIFGRCCF/mSg5r6CrLSvHA\nAsDufG4FbM2KTIhWKr0wnSX7lhgdhhBCCB/jSkFWDLSt9vz3QIFnwhHCt03ZPIVXvn+F9ML0Rr83\ntyyXV79/FZtDzneEEMLfuFKQPQ0sA84GNqIniP27J4MSwhcdyjvEtwe/JcAcwMztMxv9/jk75zBn\n5xzWHFzj/uCEEEJ4NVcKsi3An9Dzjz0C9Kbum44L4dc+2foJJky0i2jH0v1LG9VKlluWy7zd82gX\n0Y5JyZMot9RsJbNJo5kQQrRqrhRkB4GHgN3o2fot6Bt/CyGcDuUdYnnqckKDQim3lVNmLWtUK9mc\nnXOwO+y0CWvDyZKTjBy3hu++06/t3w+jRkFFRcP7EEII4btcmRjWCgwCEoFHgQr0FBhCCKfs0mx+\n2/63OJQDgM5RnbHYLS69N7csl9k7ZqNQZJdmU2Gr4GS3ScyY9WdSUgLZsAGSkiA42JOfQAghhJFc\nuYx8G/p+ks8CtwN3AUuc64wil42LWlJTITISOnbUz5OT4aKLat5JyhtllmTyydZPatyOqbQgghNf\n/p2czGAuvxw6d4bo6Jr3kfcnMu2FEMKXuWsessqCDOAq4H2gDdCuOcE1kyQ0Ucu338K8efpGBbt3\nw9y58O9/Q4cORkfWeHv3wvDhcO65kJ4ObdvCrFn6nvL+SAoyIYQvcyWHudJlOaba49XANcD9TQ9L\nCM+4+mp9e8+HH9bPp071zWIM4MsvYcoUXVyGhUGnTvqnEEKI1qmhQf0XOH8eBy6ptrQFlns4LiHq\npJTiaMHRel8PCKh6bK7nf/eB3APM3z3fzZG51wsvwMGDuquyf3+IjwdTa2kfEkIIUUtDBdlTzp9v\n17G85eG4hKjT9ozt3PPFPRzOP1zrtTVrdDfl1KnwxBPwz3/CyZO19/HOT+/w+obXmzR5a0tJSYGN\nG3WX6yuvQFQULFtmdFRCeB+L3cLm9M2eO4DdDg6H5/YvhJOvnnPLGAx/s2sXas0a/mr5nB+CT3DX\nhcMY9+dxNTbZv18P6u/kvAb4p5/gt7+t2dW3J3MPI5aMAOCW39zCP//0zxb6AI1nsVRdWWm3g1IQ\n6Mogg1ZIxpCJ+izZt4TxG8bzxV1f0DWmq/t2bLHAm2/CV1/p5vaHH4YHHpCmatEkzb2X5e3AbQ0s\nopVLK0hzeeoGj9q0Cf76V7Z/9RE7j2+j56FCVu1fXquV7Pzzq4oxgN//vva4qw9+/oBAcyDxEfGN\nnry1pVWf5iIgwH+LMSHqY7FbmPLzFMqsZXyy9RP37nz6dFi8WI8XiI2F99/XVw4ZKD9fp8NK+/fD\n4cOGhSPcrKGC7Ebn8iAwDbjXuXwCjPR8aMJIpdZSHvzqQRbuWWh0KPDBB6jgICb3KiYoMIQAixVT\nfgGfbGlcAt6TuYfvj3xPYEAgJZYSSiwlTbrFkRDCO3yd8jV5ZXl0jenKil9XNDi+tNE2bNBzzQQE\nQFCQXpKT3bf/Jigs1HXh+vW6GHvtNcjJMTQk4UYNFWQjgAeAYKAXusXsdvStk2SKylZu6S9LySjO\n4JOtn1BiKWmRY06erOcSA8jNhddf170GlJVxJNLG7rAi7CbICrahHHbWHllLQbnr97kvs5Xxp65/\n4sKEC+mT0IfLu19OeFC4Zz6MEMKjKlvHokKiCDDrq3nc2krWsSOUlVU9t1oNv2y7a1ddhE2YAP/4\nB/zf/8GllxoaknAjVzpBugAZ1Z6fBNzYUS+8Tam1lI+2fERCRAIFFQV8+cuXDL9ouMeP+7vfwauv\n6tsEzZgBV1zh7La79Va6TXiTJTsvwGGzgs0Ob79F0AV9iAmNcXn//c7qR7+z+nnuAwghWkxKTgpF\nFUVU2CvIL88HIDk9GZvDRqDZDf37o0bB9u1VVwadcw4MHdr8/TZTeXnV49JS4+IQ7ufK6MTJwHnA\nXOf2Q4FUYJQH4zoTGRTrQfN2zeM/P/2HDpEdKLeVY7VbWT5sORHBER4/9ooVukl+wAB4/nnnSocD\nPv8clizRg8Iee0xXb8JvyKB+A2Rnww8/6Md//CMkJBgbjxFyc2HbNt1d+bvfGT4Z4IEDMHasbhlr\n1w5eegkefVTnS+Hd3DVTP+hB/AOdj9cDXzY9LLfwjYTmo4Z+PpRfc38lKCAIAKvdyutXvc7V51zt\n0ePm5uqpKiIjISMDxozRM9ULIQVZC0tPhxEj9C8l6EHtM2boPjNhmJISOHQI+vTRz48c0b0IlbeL\nE97LHQVZILAb+I2bYnIX709oPqywopAya1mNdfHh8afGaXjKP/+pp6m46y59JdFHH8EHH8hNtYUU\nZC3uX//SLdLt2+vnJ0/CDTfAyy8bGpaoaU/mHjalb2JkX7nOztu549ZJNmA/0A044p6whLeLDokm\nOiS6xY87erSeABX07PS9ekkxJoQhcnN1N12l4GDIyzMuHlGLUooJGyewI2MH15xzDZ2jOxsdkmim\nhq6yrNQG2AN8ByxzLl95MijhnyqLsfqeCyFayBVX6NHjZWX6Z2kpXHml0VGJarae2MqerD0EBQQx\nbes0o8MRbuBKF8Cgetavc18Yjeb9Tf5CCLeRLssWphTMmwczZ+rH992nF5ml3isopXhg6QOk5qYS\nExJDdmk2i4cullYyL+bOQf3exvsTmhDCbaQgE6LKluNb+MuSvxAToqf9ySvP485ed/LqFa8aHJmo\nT3NvnVTpD8BmoBiwAg6gsLnBCeEuJ4tP4m9/4BwOPWdb5W1TDh3Sz+UeyEK0fmaTmVvOv4VB3Qcx\nqPsg+nXsxzep32Bz2IwOTTSDKwXZZGAYeu6xUPStlKZ4MijRumzP2I5DeaZSyC3L5e4v7uZ/af/z\nyP69ldmsh/SMGQNr1+q5ia68Uq8XQrRufTv2Zfyfx/OvP/+LcVeOo9RaSom1hNUHVhsdmmgGV9N3\nKhAA2IEZwHUei0i0Kvuz9/Pwsof5Me1Hj+x/7q65HC86zqRNkzxW9Hmryy6Da66B//xH/7zsMqMj\nEkK0tB+O/MDRgqMkRCQwefNkaSXzYa4UZCVACLADeBN4itYzlkN42NSfp1JsKWZSsvsLptyyXObu\nmkv32O4czDvIxrSNbt2/tzt0CFat0oXYqlVV3ZdCCP/gUA4mbppIWFAYUSFRnCw5Ka1kPsyVguwv\nzu3+BpQCndE3GReiQfuz97Ph6AZ6xPXgQO4Bt7eSzd01F5vDRnBAMKFBoX7VSuZwwDvvwMMPw3PP\n6Z//+Y+MIRPCnySnJ5OSm4LFbiGrJAuLzcIn29x4g3XRony1pUuuUvIBT654kp/SfyI+PJ788nw6\nRXVi7u1zMZuaP9DJ5rBxzexrKKooOnUjYYvdwoxbZnBR+4uavX9fUFEBISH1P29N5CpL4WvKrGWE\nBXn23pf55fnsPLmzxrqo4Cj6duzr0eOKxmvutBe7GnhNAUb+1ZOEVpeyMti/HwID4Te/0T8NUlhR\nyI3zbqTEUnJqXXBAEHMvHkd3YvVNKqObdzeA9MJ0ym3lp56bTCa6xXTz+C2eRMuTgkz4khNFJ7h/\nyf18dONHdI/tbnQ4wgs0tyDr7vz5uPPnbOf29zqfP9eM2JpLEtrpsrJ0v9WJE7rfqm9feO89CA01\nLCSL3YLdYddPHA5M48cT+s23ulCMjISpU+GccwyLT/gOKciEL3l9w+t8vPVj7ul9D+P+PM7ocIQX\naO48ZIedyzXAs+gWs53oQuwadwQo3Oi99yA9Hdq1g4QE+PlnWLTI0JCCA4IJCwrTy4+bCV2+UscW\nHw/FxXKjYiFEq3Oi6ARLfllCzzY9WXVwFYfzDxsdkvARrgzmMQHVL6j/I63nTLX1OHQIIiL0Y5NJ\n3xj4iBfdDz4jQ9+CpXKirJgYOHrU2JiEEKKJfs39lc92flZr/awds3AoB8EBwZgw8ckWGWQvXONK\nQTYSPRHsEecyxbnOU6YDJ2l4DJs43cUXQ2GhLnrsdrBaoU8fo6Oq0rOnLsasVh1jbi5ceKHRUQkh\nRJP858f/MGHjBI4VHju1Lr88n6X7l6KUIrs0G4Vi5YGVZBRnGBip8BWNaemKcf4s8EQg1QxE36bp\nU6C+v9gyBuN0JSV6/oNNm/Tz22+HZ5/1rqnbZ86EDz7Qj3v2hHff1V2sQpyBjCET3mR35m5GLBmB\nyWTixvNuZMzlYwCwO+xsz9heY3JWs8lM3459T10NLvyTu24uHoqed6w7UPk/SgGevItpd2AZUpA1\njlKQl6cHzTfzCkaPKSnRV4O2aeORYjG9MJ2l+5fy+O8eP/PGwmdIQSa8yePLH2fbiW3EhcWRU5rD\nF0O/oHN0Z6PDEl7MXTcXXwrchL6xeLFzKWnwHcIYJpMudLy1GAM9zi0+3mMtdx9t+YjJyZPZddJz\nPd7+MvmsEKK23Zm7+eHIDwSYAyiyFJFVmsX7ye8bHZZoBVxpQ+0EXOvpQBrr5WpX6A0aNIhBgwYZ\nFovwDkcLjvLNr98QGRzJBz9/wJTBU9x+jP3Z+3nxuxeZdesswoPC3b5/oa1bt45169YZHYbHSP7y\nXRW2Cq7scSUKRYWtgl9zfmV/zn6PHjO/PJ/Y0FiPHkO4V1NymCtdAB8Bk9FTXrSU7kiXpWiksWvH\nsuLXFbSLaMfJkpNMv2k6F7Z374UDSSuS+Drla1654hWGXTjMrfsW9ZMuS+GNpv48lQ9//pDQoFCW\n3bOM+PB4tx8jOT2ZF9a8wKK7FklR5sPc1WU5ENgCpKCvfKycj0wIr3G04ChL9y/FbDaTX55PqbWU\nKZvd20K2L2sfG49upGtsVz7e8jGl1lK37l8I4Tvyy/OZvWM2CZEJ2B125uyc4/ZjKKWYtGkSRwuO\nMm/3PLfvX3gXVwqy64Fz0ZPB3uhcbvJgTPOAjcB5QBrwgAePJSo5HHD4MKSm6qkpfEyFrYKrz7ma\nAV0G0L9zf67reR3dYru59RhTt0wlwBxAeFA4xdZilvyyxK37F0L4jvm751NhryDQHEhsaCzzd88n\nuzTbrcfYfHwzv2T/Qo+4HszeMZv88vwGty8oL6hxhafwLY3pAkhAX3FZychZPaXJ352sVnjhBfj+\nez3YvkcPmDIF4uKMjsxrHMg9wG0LbiM4MBgzZsrt5bQLb8eK4SvkcvYWIF2Wwts8uPRB9mTtOfU8\nwBzAa1e8xpU9rnTL/pVS/OXLv3Ao/xBxYXFkFGcwsu9IHuv3WJ3bO5SD4YuHc0X3K3jo0ofcEoNw\nH3dNe3ET8DZwFpAJdAP2Ab2bGV9zSEJzp4UL4Y03oEMHfaXmyZMweLDf3drIarcSFBBU52vFlmJ+\nTPsRRdX/u5CAEAZ2G4jZ5EVzvbVSUpAJf7Mncw/3fXkfIQEhmEwmLHYL0SHRrBy+ss489cORH/j7\nN38nIjiCr+/9mugQL77a3g+5ksNcObUfB/wB+BboC1wB3Nfc4IQXOXBAz11WORVFZCTs9+xVQ97m\nYN5BklYk8ektnxIXVtUyaLfrryUyOJKrz7kaux0CAgwMVAjhF86PP5/Zt86udRJYV4u8QzmYuGki\nUSFRlNnKWLB7gbSS+SBXTu2tQLZz2wBgLdDPk0GJFnbeeWCz6XFkSukbf/fqZXRULeqjLR+xN2tv\nrYGzn3wC8+bpryUrC/72N92AKIQQnhRoDqR3Qm/6JPQ5tZzb9tzKlpYaNhzdwOH8w0SHRBMXGses\nHbMorCg0IGrRHK4UZHlAFPAD8BkwET05rGgtbr4ZrrtOVxxZWboYGzXK6KhazMG8g3x36DvOjjub\nOTvnkFeWd+q1oUNh40aYOFEPs7vuOmjf3sBghRDiNHN3zcXqsJJblkuxpZiiiiJWHVhldFiikVwZ\nkxEBlKOLt3uBaHRhluPBuM5ExmA0k1KKjOIMOkZ1rFwBJ07oAf6dO/tVv9zo1aNZd3gd7SLakVGc\nwQMXP1Dj1kupqfDUUxAeDvPn62F2omXJGDLhDtml2USHRBMcEGx0KG6VUZxR6wrMLtFdiAiOMCgi\ncTp3zUM2BrCjuy5nolvInm1mbMJgPx77kbu/uLvqMm2TCc46C7p186ti7FjhMVYdWIXdYSerJAu7\nw868XfNOzTGWlQVvvgl33qnvg17ZfSmE8C12h51H//soM7fPNDoUt+sQ2YHfxP+mxiLFmO9x5Yxz\nG3owf3W7qH8W/ZYgZ5jN4FAOhn0xjG0Z23j00kd58g9PGh0SuzN3E2QO4vz481v0uOW2cpLTk6n+\n/ykoIIj+nfoTYA5g2jR9682bb4b8fBg3Dp55RrotW5q0kInmWn1gNc98+wwRwREsH7acmNAYo0MS\nfqS50148BjwOnAMcqLY+CvgfuvvSKJLQmmFj2kb+/s3fiQ+Pp6C8gGXDPHPLD1fZHXZuW3AboYGh\nzLtjnldNI6FUzS7K05+LliEFmWgOu8PO7QtvJ688j1JrKQ9d8hAPX/pws/drc9jIKsmqGvohRD2a\n22U5Fz0r/1fAEKpm6b8EY4sx0QyVl0eHBYURFBCEXdmZvWO2oTGtPbSW40XHOZh3kI1pGw2N5XSn\nF19SjAnhe9YeWkt6YTrRIdG0CWvDpzs+paC8oNn7/WLvF4xYOoIya5kbohT+rqGCrAA4DLwInHQ+\n7gEMB+QOpz5qb9Zefs39FavdSnZpNkoplu5fisVuMSQeu8POpORJhAeHExoUyqRNk3AohyGxCCFa\np4V7F2JTNrJLsymsKKTYUszqg6ubtc8yaxlTt0zleOFxlqUsc1Okwp+5cr6/HT3vWHfga2Apepb+\nGzwX1hlJk38TOZSDowU173oVHBDMWVFnGRJP5biOdhHtAMgqyWLSDZO4rOtlhsQjvJN0WbYCJSXw\nyy8QFKSn1glsuVuO5ZTmUGQpqrGuQ2QHQgND63nHmS3YvYC3fnyLmJAYlFL8d9h/CQsKa26oopVy\n10z9CrABtwGTnMu25gYnjGE2meke293oME7Zn7Of9pFVI+TbR7YnJSdFCjIhWpOTJ+GhhyAzU09A\nffHFenK/0KYXRI3RNrwtbcPbum1/la1jMSExhAWFkVGUwbKUZdzV+y63HUP4H1fOODcB7wEvoMeQ\nHQJ2A308GNeZ+OcZphB+SlrIfNzo0fDdd/ry5Mo5D59+GoYNMzqyJllzcA3PfPsMIQEhAFgcFs6O\nPZvP7/rc4MiEt3JXC9lI4BFgPLoY6wEYOwpcCCGE7zhyBCKc82KZTLq78ujRht/TGAUFMHUq/Por\n9O4NDz+sZ3L2kMu7X87Su5fWWCfzfonmcqUg2wP8vdrzQ8AbnglH+DuL3cLo1aN5ZsAzcim5EK1F\n376wYIEuyhwOfe/cC900laXVCk88ocenRUTA1q26MJs4EcyemUIn0BxIl5guHtm38F8N/W9dDtwJ\n1HWaEQ4MRQ/yF8JtVvy6gmX7l7XK2bSF8FtPPAEDBuixZFlZuqvy+uvds+8DB/T9zTp0gOho6NgR\nkpP1sYTwIQ0VZA+gZ+P/GT0z/yrgW+fjLcAFwP2eDlD4D4vdwvub3+es6LNYsn8JJ4pOGB2SEMId\nIiJ0i9W338LatXr8mLtar8zmmvczq3zsodax1qbYUswb/3sDq91qdCh+r6H/sZno+1j2Aq4GXkLP\nSXY1uhh7GcjycHzCj6z4dQW5pblEh0SjlJJWMiFaE5MJYmMhMtK9+z3nHLjkEn2hQE4OZGTAlVdC\nQoJ7j9NKLd63mGlbp7HywEqjQ/F7vnrVkv9dpdTK2Rw2bvjsBtKL0gkLDMOu7KBgxfAVNabFEP5J\nrrIUDSorg7lzdddlnz4wdKie70w0qNhSzODPBmN1WIkNjWXp3UsJCpDvzRPcdZWlMIJSsGuXPuPr\n2RO6tO4BpEophl04jHJb+al1AaYASQ5CiDMLC4MHHzQ6Cp+zeN9iSm2ldIjsQEZRBisPrGTIeUOM\nDstv+eoZZ+s+w1QKXn8dFi+GgAC97o034PLLjY1LCINIC5nwFla7lf9b8X8888dnvGqS7cYqthRz\n3ZzrKLeVExIYQqm1lA4RHfjqnq/kRNgDpIXMV+3erYuxhAQ9MLW0FMaM0YNhZaCqEEIYZuWBlaw6\nsIq40DjG/Xmc0eE0WbmtnEHdB9W4j3F4UDgWu0UKMoM0VK3tauA1BVzk5lgao3WfYX7/PTz3HLTT\n93dEKT1Qdf16j052KIS3khYy4Q2sdis3zbuJUlspZdYyFt650KdbyUTLaW4L2Y1ujUa4rmdPfUVS\nSYkuwLKy4NxzpRirR0ZxBmkFafQI+R2hoVUXcR09Cp07N71R0WK3EBwQ7L5AhRA+beWBlWSXZtMh\nqgPl1nI+2fKJT7eSCe/S0J+qw9WWMvScZH2AUuc64SmdOsGECXpG64wMOPtsePtto6PyWu/+9C5P\nr3qar1eXMGYMFBfr6yGefx6OHWvaPostxdyx8A52Z+52b7BCCJ9kd9h5P/l9iq3FpBemU2GvYOn+\npRwtcOMtoM4gozijxY4lWp4rY8juAiYA3zufTwaeAeQuqp502WX6ZrwVFfoKIqMoBZ9/Dl98AaGh\n8MgjesZtL3Ew7yDfHfoOu8MOlyymV/F93HOPvhbitdega9em7XfR3kXsy97HlM1TmDJ4inuDFkL4\nHJPJxF8v+WuNK8FNJhMRQS1zD8s9mXt45L+PMOe2OdJN2kq5MiZjJ3AVeqJYgHbAGmQMmX/4/HN9\nxWd0tL7/nMUCH38MFxn5z19l9OrRrD20lujQaCw2C2/89mvGj9UJct68ps1BWVRRxOC5gwkJDCG/\nLJ/pN0/nwvZuuu+eC2wOG6Dvlyc0GUMm/N3fvv4bKw+sZGivodJN6oNcyWGujK4xUXNG/pwz7VS0\nIkuWQFSUrmxiY3U36rffGh0VoFvHvj3wLTGhMQSYAsgsKOTJjxYzfjzcfDOnui8b64t9X1BuKyc0\nMJTAgEA++PkD9wffgHHrx/HWxrda9JhCCO+1J3MPm45tomebnqw6uIrD+YeNDkl4gCsF2QpgJTAC\nfX/Lr4FvPBiT8Cbh4bplrJLD0eDFBS155v9L1i/EhcVhc9iw2C2Em+PodfkeLrpIzxH5+99DefmZ\n91Od1W5l9o7Z2Bw2skuycSgHG9M2kpKT4pkPcZqjBUf5OvVrvvzlS7mXpxACgA9+/oBAcyCB5kBM\nJhOfbPnE6JCEB7jS0mUCbgMuQ0938QPwpSeDcoE0+XvSjz/Czz9DfLy+Q8A//qGLMqUgLg5mz4aO\nHSjvd3QAACAASURBVGu9bXfmbt758R0+vPFDn+1ucygHW45vocJeUWP9pR0vJSzI82P5xqwdw4pf\nVwBw629u5fmBz3v8mL5AuiyFvzpWeIzbF9yOyWTChAmHcmA2m1k+bDltwtoYHZ5wkSs5zFcTnCQ0\nT/n8c31XAJMJ7Hb4zW/gmWfgf//Tg/oHD9bFWEoKzJypp+YYMgSuuorHv36C1QdXM/G6iVx37nVG\nfxKfc7TgKHcsvIP48HgUityyXJYMXULHqNrFr7+Rgkz4K6UURwqO4FCOU+sCzYF0ie5S+XshfIC7\nZuq/HXgdaF9tZwqIbk5wwgspBZMmQZs2EBKi16WmQn4+/O1vVdsdPgwjR4LVqm/g+7//sSvvFzYX\nbOasqLOYvHkyV51zVb2tZEopSSR1WLhnISXWEgLK9e2yii3FLNq7iFH9RxkcmfA7SsGWLXremC5d\n4NJLjY7Ib5lMJrmq0k+4UpC9CQwB9nk4FmE0pfQ0G9UvTTSZ9LrqVq3St3Pq1Ek/Lyrig/+9R+BF\n7YgKiSKjOIPVB1bX2Uq2/sh65u+ez/s3vC9F2Wnu6n0Xf+zyxxrrusS07pvKCy/1/vswa1bV84ce\ngocfNi4eIfyAK4P6M5BizD+YzXDDDXDypC64srP1AP6+fWtuZzLpxWlfWAnfR2RhtVvJKM6g1FrK\nlJ+n1Brgb3fYeefHd/jhyA8kpyc3O9yNG6GoSD92OGDNGv3TV3WN6cofuvyhxtI5urPRYQl/c+IE\nfPqpHkPaoYP+OW2azgdCCI9pqIXsdufPn4EFwBKg8i6kCljswbiEUUaP1nOOrV+vb9f01FPQvn3N\nba69Vg/sP3kSAgNJUFbG90mCSy45tUl4UHitFrD1R9ZzrPAY0aHRTE6eTGKnxGa1kqWmwoIFegLY\nTz+FtDT44x/1UDchRBMVFuqTs0Dnn4fAQH0CVlioizMhhEc09NdwJrrwqtzu9FGoD3giIBfJoFij\nHTwIc+boQf3XXw+DBjW4ud1h546Fd5BbnktUcBTpRenc0+cenv3js00OQSl9XcHixXqYy9tvG3tT\nA+E5Mqi/BZWVwW23QUGBnnswLw/attV366gcWyqEaJTmDuof4c5gRCtz9tl65lUXbUzbyC85vxAZ\nHEmZtYwTRSd4fcPr3HbBbfRs07NJISil60HQF4RWny5NCNFEYWEwZQq8+CL8+iucdx6MGyfFmBAe\n5soZZxdgInoeMoD1QBLQxNs2u+Q64F0gAPgEeOO01737DFPUcqLoBFtObAH0rYle+f4VlFLcdP5N\nTLhmQpP2+cEH+oLPl1/WXZfbtum7PEkrWesjLWRCCF/mrnnIVgOfAXOcz+91Llc3J7gGBAD70ffP\nTAc2A/dQ88ICSWg+bOKmiczZOYd2Ee3ILMlk3u3zmtRKtmOHPnkPC9OtZcnJkJhY43oD0UpIQSaE\n8GXuupdlO2AGYHUuM4GEZsbWkETgV+Cw83jzgZs9eDzRgnLLcpmzcw6hgaGU28qx2W18+POHTdrX\nb39b1RpmMkH//lKMCSGE8E2uzEOWw/+3d97hUdTbG3/TQxJCJwgooIhIFSxw8QooFgQbdhQVRRQF\nRRFFBTEXBPwpAopYL8qjVDsgFpQqFpqUG7ooCqQQEtLLJtn9/fHuOJuYskl2M7vJ+3mefTKzZebs\nZPfs+Z4K3AVgMWjd3Q7Am/XPrQAcddk/BqCXF88napD4zHi0a9gOhXYmfDUMa4jsgmyLpRJCCCGs\nxR2D7D4AcwHMcu7/BO9WWLrly4+Njf17u3///uhfQZVfnSQ9HYiPB5o185ly9S7Nu2DZLcusFkP4\nOOvXr8f69eutFsNrSH+J2kJRERDE4SJwONgL0tivy1RFh/ligKc3gFgwsR8AngFgR/HEfuVgVMSW\nLRwKXlDAb8nTTwM33GC1VMLJTz8BZ5wBtG7Nf8+qVcCAASpIKAvlkAnhexw/ztHHsbFAo0ZsT1lU\nBNxrZVMsH8VTOWQfAGjost8IwHtVF6tCtgE4G0BbAKEAbgOwwovn832OHWPPr8WL2Yy1Imw24Kmn\nuExp2pSNXmfMoLdM+AS5uewqcOwYm6CvWUNFJkStYutW4Lrr2LH5qafM0RqiVtCqFf+1EycCr70G\nbNsG3HRTxa8TpeNOyLIbgDSX/VMAepbxXE9QCGAMgG/Bisv5qMujm377jYO8s7K4P38+Z8y1Lmek\nzqlT/MVv7qy9CAtj5+2EBKBlS+/LLCpkwAD+feghesXee6/4CFEh/J6//gLGjgVCQrgoNFYdr7xi\ntWTCg9x6K/0Fx44Br7/Of7WoGu54yAIANHbZbwwaSt7kawDnAGgPYIaXz+XbzJ/P4d6tWvGWkQEs\nWlT+axo1AiIj+VwAyMvjX2MYuLAchwP44w9u2+1AWlr5zxfC74iLY7fm6GiOX2rRAvjhB3743eHk\nSaZa3HwzY2L6kvgcDgfDlO3aAddfzz6QqalWS+W/uGOQvQLgZwBTAbzg3K5aJ09ReTIygNBQcz84\n2DS0yiI0FJg1i16x5GS2s3/+eSpEPyQ1lU4/g7/+8v+u/IsWAXv2AEuW0Es2aZJmNws/IDsb+Phj\n4N13gZ07gQMHaDD16sXEIde0iKgo/mIbBlhuLo0zd3rT2GzAqFHA2rVUAF9+SW+b4vo+RXw8sHs3\nBzncfz8n6K1cabVU/ou7SbKdAVwGVkCuBbDXaxK5R91Jil2xAvjPfzhTzuGgMfbSS8Bll1X82pwc\nIDHRzCPzU776ivp42jS+nWnTaF+efbbVklWdI0f4bwkPp928bx9w7rn0lgW7k0hQx1BSvw+QkwMM\nH840ioAA09iKiAAaNKDh1KoV8NFHzF8tLAQefZQFRgEBvM2YYcbry2P/fp6rWTPuOxxcXH76afnp\nGqLGsdupw8raF6S6syxdw5QJYB8ygEZZYwByTHoCQzGXtWq89loqwkWL+CkfO9Y9YwygojzzTM/I\naSGDBtEOvftu7sfG+rcxBgBt2/LvkiX0jI0ezS4lzz0HjBtXK/5toraxcSPw++9m6kNiInDoEHDJ\nJdxv1oyJRMnJ9MYHBzPTe8MGfrg7dwbOOce9c4WG8pfd+HU3+im4RguET1DS+JIxVnXKM8h+Rfk9\nwdp5WJa6hd1Ot/8HH3B76FBgzJh/fpoDAoDbb+etDtO9u5k6546x4nAUt3FL7vsKN9xAB+i0aXT/\n9+snY0z4KEYuqkG9egwhGo2obDZ+yVyrU4KD3fOIlaRtW34Z1q41vW1DhpgeMyFqIT74E+UW/uny\nd2X5cv4SN2tGJXbiBMvC67jhVRr79tFgefxxLsg3buR+o0alP7+wkDlZDz0EtGnD3OLPPqP3yReN\nsoQE4IEHuL18uXsrTIeD7zMkhPs2G7d98f15AoUsfYC//uLCMSCAsfaUFCAmhi5e44P32GPAnXd6\n5nyFhUxIOnwY6NiRrnK5X4Sf4qnh4gB7j50NINzlvo1VE8sj+KdCc+Wpp4AffwQaOyPDaWlAt27A\nvHnWyuWDbN/Ov+efz78rVnCIeHk1Chs3Av/9L+3bRYuACRN4eX2NU6fYw+eii5g206oVw5cV/e6s\nW0fnwaRJdLDGxtLb9q9/1YjYNY4MMh9h9262rUhNZQb3mDFM7k9MZKmdL37JhPABqptDZjASwKMA\nTgewA+yk/zOY5C+qSkwM21kY5OebfcNEMQxDzOC66yp+Td++9Iy9+Sbw4IO++zuxdi1lvf12FqHN\nmAH8+Sd/21zJzATeeYdev4gIoEkT5lZPnEhHwtlns9BNCK/SrRv7ILqiD16VyLZlIzI00lIZXFM5\njG1/Sfeojbjj/x0L4CIARwBcCqAHgHQvylQ3uOceungSExmzCg2lmyRbg7Y9QVwcHZBXXsmirz//\ntFqi0rnxRjNKXa8eo9gljTGAbeUiIlhdumULx5WMHcuuA4cPAyNGKJojhL+wI2EHbvroJqTnWfdT\n+t13wFtv0cNeUMDWFV9/zT7kRveSRYvofff3NkP+gjsqPA9ArnM7HMB+sGmrqA5Nm3IUUmwsvWU5\nOfw1vu02jTiqJoWFwBtvMEz5yCPsj/PGG+73oyyP5GRz26jErw4lV55lrUQDA+npy8kBpk7le3r2\nWX50+vUDJk9mD829VjekEUKUi8PhwOtbXsfhU4exbM8yy+S4+GIWzb7+OjB9OnNQBwxgDcWgQSyQ\nnT+f6c2uwRzhPdxxRH4O4D7QUzYAHJ0UDGCQF+WqCP/NwSjJRx/R3XHaaWZy/yWXADNnWi2ZX1NQ\nYCa8l7ZfWRwOHmP0aOCWW6i4Fi6kh2ratJpx6cfFMaQZHc2Uw7w8nn/CBH5cgoKYA3366d6XpaZR\nDlkdJTOTRQPNm9NNXAvYkbADI1eORON6jZFTkINVd6xCg/AGlsiSng4MG8btzz9nUazdDlxwAQeH\n9+tHo6x+fUvEq1V4arj4ENAIiwXwHID/ArihmrIJg6NH+Utq/KJHRvpufK2qHD8ObNoEHDxYY6cs\naXyVZYzl5rq3/9lnNMCefpqjQrp1Y/7XM8/UjDFm9AOeMIF1H337MoH/zjtZH+Jw1F5jTNRR1q8H\nBg4E7rgDuPpqxur9HMM7FhoUirDgMNiKbNX2kuXn04gyKKnDyqKgAJgzB+jRA+jQgV2Y7Hb2RmzQ\ngPVme/ZoHnxNUtmsk/UAVgCweV6UOkrXruzjU1jIX9X0dKCnN2e31zDr1nG0yvjxXIq99ZbVEv1N\nXh6LxD79lMbV55/TA/Xll3TlJyUBDz/Mav/LL6chNnEi7eXAQPbsramVY3Q0MHcuDUEjfHnnncC2\nbXw8OxvYvLlmZBHC66SmMiYfHs70jqAg4MknGbP3Yw6kHMCvib/C4XDgZM5J2B12LPnfEtiKqv6T\nungxdYPdzmyX0aO5Bq6INWuAsDCmO0ydSp33zTfA+++zy8gvv7Be4+67ZZTVFP4aAqg9Ln+7nd+m\nhQu536sXQ5i1wT1vs3GqQHg4M9YLC9m7aMkS4Kyz3D5MQgK9UEabi3372Dw1LIz7H37IFV6vXtTX\nr75Kg6Vx4+LHOXAAOOMMigIAu3bRuJk8maOMWrYEbr2VxQDTp/N3YN06KqjWrRkm3LSJyumrr+ix\nGjqUhQOexnX8iN1uTp5xffzuuyn3kiX04L37LqtKjcbptQmFLOsYcXHAyJH8EhokJwNLl5pjLvwQ\nW5ENcSfi4Pr/DwsOQ+dmnY3PeKXJywOmTOG6PimJC7Urrqj4dcbwg6Ag7hcWMmS5YwfQvj0Xm3Y7\nDbPevVU0VF082YfM16h9Ci07mz7kBg1qT41xSgpw1VX0oWdk0BKKjmbMrRKl8mvWcBU4bRpXfrNn\nc0VnVCP+9hvrIe67j4bSmWfSICupQN5+m6vA2FhWE337LXOvXn2V0ZHsbOr/9983fwfsdlZBZmUx\ngT4tjX9796bS+/Zb5pRt2WI2JD9yhI7O7t2rdtl272Zq4aRJLL6dO5cG59VXF3/ewYNsit6yJZXr\nr79yHmZERNXO68vIIKtjnDxJF3RUFBd0OTmMzX37be1YrHqY339n1TXgfnNpUbN4KodM1ASRkRwg\nXpox5nCw0+miRXTReEqZOxw85pVX8rZwoXvHTk1lR+7+/bkcKys3rGFDdj49coQKNSGBmqOS/dbq\n1aOhMXIk2z706FF8pF379oxmzJrFRfSDD1J3p7pMWz1xgrOKW7emF+yLL2jgff89Q5L9+jEUeegQ\njTOAnrC77mJeVvfu9MzNmUMH5uLFfDv33cffjilTaEQdOUKPW2YmV6zz59MWBfi8BQsqvsRdutAg\nnDKFBmNSEnDppf98XocONMYAfmzOP792GmOiDtK0Kb/sWVn84uTl0W0tY+wfxMdzgfrAA8yAMcKX\ndYnUVC6ki4q4HxfH8Ku/IYPM13E4gJdf5sTpV16hITR7tmeOvWoVrRhjiO/s2byvInnGjWNcLzyc\nFsioUTS8SpKVRQUaFcVvSmgo444pKZUSs0ULhg5TU1kDsX8/HW0GOTlm2LKwENi6lbenn6Yuj49n\nMnxcnGnAhIXRVb9rFzB4MI+7dCmNv/HjmSA/ejSnBDz3HI236dOBH36gE3PECE4C+OMP/luGDuVY\n0kceoeH4739zlRoSQk/XH38w/yw6umIHaGAg21p8/DH/HRMm8PVPPKE2daIOMXAgEzrnz+ffvn2t\nlsgn2bSJdQ/XXsvFYFYW21vWJaKiqCNnz2aE4cUX2bjA35BB5uskJACffEKvUqtW/Lt0Kd0m1WXN\nGlogrrc1a8p/TXo6S29iYmhtNG5Mi6g0L1lwMC2fnj0Z4+vdm4kJwe4MiADf+9q1yP9xG0KC7AgN\npQ2Yn188t3f+fIYpX36Z4cg33gA6d2aI79576TG74w56wlav5kqqWzd+aSdMoHNwxgzq+3fe4Vub\nMYNvc+ZMOvoCA5kON2wYbddrrwUuvBB49FGmtFxxhWloJSTQoLPb6WGLieH5+/Th/E3DHrXb+byS\nTRftduanORymF+6GG/jvMRwEqbmpmP7DdBTa1bFRWExmJj34mza5X+LnLk2aAJ06lT24VuDWW4G2\nPQ8hrzAP4eFc+BkLz7pCaCgXvj/9xPf/xBNVTxmxEhlkvk52Nq0BI/MyKIj7nnCVNG7MxHsDm61i\nxVevHs9fUMB9w7tWWighMpIJVidO0JBLTKRy7dy5Ytl+/ZWvffZZtH3pIUw49TQa1LejVSsONHAN\nWY4YYeaMtW9Pg6xJk+Jpaj168L5p0xgNGT2aBlhwMN9SkyZ8XqdOXFn9/jsfu/DCf4q2aRO/8Js3\n863Pn88E+3HjGJJcvZres1deYQh08WJeru3bgWPHqDBOnmT4c/Nm081uEBfH+774grKuXk2j7oUX\nzOcs3L0QC3YuwPo/1ld8LYXwFklJdA8/+SS/AHfdxe+6+BuHg4tBg/x8z6ynDTJ/Xo8HZvwLS4d2\nBZ55Bns2Z+HLL83Hv/iC6Ra1nYMHqbODguhXKKlX/QEZZL7OGWfQK5acTIPpxAlaDK1bV//Yw4cz\nhhYfz1t0NF1KgGkxPPsswwVG4lNYGN1CqanMsE9MZIJTp06ln2P8eMb8Bg5kj4l589zr0Boby29W\ns2ZID4tBq4NrsXj0j5g5kwn0rh3yIyKKJ7FGRvLtTJxIUe+7j2+jUyczWT8wkKFKV1EcDtNYGjWK\nx330UdP2NDA6XB88SEOpb1+muGzYwEs4ZAiPf/iwmZ63dSubLe7ezct1770sRnjuObNa1KBbN+aE\nBAXxfbRoQaNx1y4+npKTgiVxS9AsshnmbpmLIrsfah5RO3jzTeqk5s35ZThyxKwYFwCoq559ljog\nP5+5oeVlhqSmMn3OaDWxaRMXfaVy5Ag+mTkCqcjFe2ekIHPN12j74RQsXw6sWMGWPl9/7fkxyTk5\n1GUGx46512rDWyQlMeIxcSJzeU+doh42Qrf79plRB1/GX6uW6laV0vHjLCM8dIgNYiZPrlqA/PBh\nGllJSbQiHnjA7OVw8CCwcyctiauvBpYt46c5JITPb9aMnq0HH6Traft2ytO8OTPiDQ+ep+jdmx68\noCCG9JISEfz8JOD66/8Wp7xKov/9j+Ib5d8rV/LSnX122a9xOGgzXn01cM01VKBz5tDrVdKGPHSI\nDgGAHrC0NNZH/Pgj8xmMBq5HjlCGJ56gvPHxvLTr1tE4fOWVf7bnABiqvOYavnbECPNcS5cC8+Ne\nw8LdCxETFYPEzETMGDADl591uZsX1j9RlaWPMnIk+8k0cHaaT0nhimPaNGvlqgGys5lCYai+jIzi\nua2uHDjAkFpeHlMfxo4tW385HNQ5u3ZxHbt4cdkzbjOXf4TBa0YgLDgM6UGFeDjpDAw/UA8nVvyC\nEffz67JggRkB8BRHjnAxOWYMM2kmTeLC16o0P4eD+t74WbTZ2Apo9Wo6befPpw7u0cMa+QC1vRCu\nJCcz2SAvj+6W9HROtp44kUudkSMZBwwKogFot9OlFB9P6yMggK4bm40Nr7p18668Y8YwnhcTQ5kz\nMpi5f453x6jm5pp9ykrbB7hyfe455ig4HLx8U6eyf8/LLzPqGxHBItQbb+RvU8OGwOOPc25cYiJf\nv3IlB/zOmmVGfB0OXt6+fbnqvOACKuRLL+WlP71DKq768CogAAgJDEGWLQttG7bF57d9jqBADxvF\nPoQMMh/lvfe48oiJ4Yf3xAl+uG+o/cNc3n2XC7Fx41ihHRtLL4zRL9GV/Hzmn+bl8fJcdFH5x3Y4\nqJKTkqg/ylK37y95Cm9unoeYgCjkBRQh31GIVft7YvX9a7FoEZ8zfDhw3XXVeael47oofewxs+2P\nLzFrFhe/Dz/8z7ZBNY3aXgiT7dtZftO0KX/9Y2Lo07bbuYyw22k11K/PZV9aGl+XmMjAfGAgH7fb\naUV4mylTgPPO4/nz87lE9LIxBvzT+Cq5D9CZ2KMHFd2991LM0aOpOGfPZpL/tm1UUKGhtHkNZXDF\nFWaY8uabaXe6tqoICGCU+qWXGJWeNImXvWNHGoAFRQW4psM1GHjWQAxoNwDXn3M9Lj79YhQ5FLYU\nFnDXXVx1JCfTO3bPPd759fdB7rmHKnXMGH6nR40q2xibMoVFPS+/zKHdW7eWf+wff+QCrEcP2rxl\ndcr/OvB3FEXWQ5IjC+n2XOSgAEsvH4zVq2kwvvUWF37emOLhqhujojx//Oqybx9Tkf/1L1as+0Pl\nqb+uOGvPCrOmWLuWcTRDY+Tn0/2zcSNXuAsWmP7e5GS6Z1q2ZDZqRgYfa9KEfvrRo7nkqIjMTCZ8\n2e3Mjm/YsPJy5+czXujjnQ7nz2deiNGc1uhWUlVFtWwZU3H69+cqtLb0Cq4q8pD5OAUF/JC6W0Ft\nJca198CXav9+1jOEhPBHv7TMjbQ06oahQ6nGDhygWi2rm35qKsNrkydzcbdgAVWw0fjVlbzCPBTk\nZgM/bgIyMoFO5yL07J7IyQn4uz4rNZXr7JJpF2lppko2pva5q6KPH+dC8667uICcMoUtfyry/NUU\nNhsN5YceolH71VdsWTR9unW6VCFLd0hJ4bclN5dxonPP9cxxrcDoahoayuSDmBjzsdxcLul++41K\ns6iImuT22/ntuvtufkMNpTp5MvDzzyz527iRvnaAGmfmTOaflUdKCt1HxrKkSRMu9arRHMbhML9M\nrts1TclzG/u7dgFTn87GVQUrcf+NqQi48MLSyzQrIC2NnrGMDP4rpk3zz546nkQGmag2NhtXSStX\n0kU9ZgwruauIMfVjxAiud6OiuHjyRDptXh4DFQD1S36+ue8JbDYaK/feyyKlJUvY5sfd1L/0dGDv\nXnqfAP70BAZWaiKe13G9hqXt1zQyyCri5Ema+ElJ/EUNCaE/2VfM/Mqwezd95vn5/AY3acJOpa6/\n5JmZzHQ8eZLv0XXoYXw8Q5g2G2NtRmuKw4eZD1JUROugUSOeY82a8rtmz57N5CfDI5eURCNxypQq\nvb3vv2ci6YgRFGXmTCa9d+lSpcNVmaIi2qr3388k2wMHqMzGjgWmPJOLET+NQOSx/WjaPBD1o0DL\nqoJ8GqM8OyiI/7rx49l1f+hQTor5/HM6Mf3B+eAtZJCJajN3Lt1NzZuz+d+pU9T3ffpU6XCffkr1\n2qcP1ea8eRxc4umKRm9hTBWJjKTueeGFqgUxhHu4o8PqsIoHPWNJSSwTAeiaeP11GjK+xIkTnNdz\n8CDzqJ5+uvjQXYDl54DZETAhgRpjzBjzOfXr00tmTKt2pWVLGnQlSUvj0s/1fCdOmF34yyIpqbiP\nPDycr6sivXvT7fz222Zj1Y4dq3y4KhMUBAwaxLL0O+5gWPHxx6nnBzf6CZ1DDyG3RyscPAR0bpGL\nkDlzKjTIVqzgCtPoxJ+Rwby0gADasBdcULeNMSE8woYNrAYNDjbzYrdsqbJBdtNN5nZoKPWAP9Gm\nDX9OfvmFql/GmPX4dmKOt8nJKZ6bFBLie7NpbDbma/3wA8OOGzYwh6tkc6ycnOIGUGAgjSZXsrOZ\nR9arF1tVrFhR8fnPOovu/bQ0um+Sk2nAljQIS9KnDz1pBQVcjWZnc55QFYmK4mpu1SoqkAkTrDNS\n+vRhbte8ecD119ObNXo0MODf+QgIDEBEBNCtKxASEcr/WQXekMGD+e97/nngmWf+GTmv6FILIdyg\nWbPikwQKCz3fD8JPcDjo2U9IYIX4smXsfiSspW4bZJdcQpdHejp/EdPTORPHl/jrLybYx8SwrKVF\nC9ZYHztW/HnXXEMDzHCxOBz/zBqdOZOxv+bNaWRNncreYwCV0+7drMZ0NUobNqTXsFEj5oO1b083\nf0WJEtdcw7rt9HSGBoYNYwyuihQW0gDq2pWhQneGdHuDdetoGK5Zw35C06fTSAwNBQJ69qAnMCUF\ngfm5vF6DBlWY7BYaSkfmrl38d99xh5L4hfA448ZR7yUmmrqsDrTnKI2CAl6CF16gN37KFKYLC2vx\nV7XvuRyMn3+mwZGTw3Lte+7xrYq+Y8dYVm50QrXb6aVavrx4fpjdzmXOJ5/QUzZqFN04rlx5Jf8a\nreHj45n8dOutbElvNIZt3pw1065FAcY5KnttHA7eqnlNv/qKfb4mTGBy5uTJrEM477xqHbbSZGXR\ng3XppbRR//qLkdtZs5xG1L597PaanEyD/5FH/tmKvwQZGaxYOu88DjmPiGD40tO9dv0Z5ZDVQRwO\nhhTj4xlf69mz+sdMSuKiMyyMGemuPWeE8CJK6q8NOBzswbVihWmQGQ1dK+tGGTaMmZyNGpmTq//z\nH3qxZs2igRcQwFyvyy7jhO2qkJnJRC/Dq+cBjJGZRpiyoMC9CUzeIDWVdjvAGgmgerJ89RUv17Bh\nfF8vvcTirxpou+Y3yCCrg8yaxbgaQH310EOs6hHCD5FB5stkZbFr34ED/OUdNarsplVG89bff2dO\n1xVXVM3jFBdHpWaz8ZjdunES96xZnEBrlAdlZTE0aihDd3A42JTnm284Qyg4mKvPOXMq7Oq/7LuQ\ntwAAFKBJREFU8uBKNI9ojl6te5X7PF+gsJD1Fbm5vLVrxxS/wEBe1h9+oC0bEMBi1iNHmJRfHiXb\nIlnZ0sNXkUFWxzh6lFnzTZvSVVxYyJXQN9/g7wZbQvgRqrL0VYqKGCrctYvxrh07aMy8/XbpcarA\nQJbbVZbVq2lw5eXRq3b//Zy8uns3PVe9ezOBqWtXhjqLiniuzEzmPrmL0QV18WKzz1nXrjzeuHGc\nbluGCykzPxP/t+n/0LheY3x222cIDvTtj+S6dfwbG0tvVmws/33nn88ahuXLGWUePJgDhQcPrtgg\nK2l8yRgTdZ7MTOoRQx8GB/OLkZkpg0zUWnwoWaoOcfQovVUtWnAabYsWnIZ99KjnzrF1Ky2C9HSu\nLt9+mz0aTjsNuOoqJkLl5LDPWL9+TLhPTmaORZ8+7nXiN9i1i22qIyNpeAUF0cCsX58JUqmpZb70\n470fI78wHwlZCVj3xzoPvHHvcvnlZoVnvXpspHj++Xysfn0myX77LRsuDhrEKkwhRCVp04ZfqJMn\nubBLTqae9GKH5PS8dK8dWwh3kEFmBYGB/ywR9EDiezE2bODxoqJoOURH091v8M03tBiGDWNl6aWX\n0v3z3XfAq6+W32OsJMnJPJeRIBsQwPhdZibPXcaKNjM/E+/vfB+N6jVCREgE5m6Zi0J7YTXetPcp\nOR2mZOsNIxoM0BZWZEqIKhAZyd6K7dpxQdexI4uvvJQ4mp6Xjls+vgWbj3lh6KMQbiKDzApat2aF\nT0ICs7kTEuiVat3ac+eIjjZbwAOMpzVowO2kJCbzR0aaTa7Gj2f4smHDysfMjHkZDge38/KoOO12\nZqiHhpb6so/3foyUnBTkFOSgyF6Ew6cOY+3vayv5RmuIwkKGgJcsoUewFNLS6JS8/Xam0W3fXrk0\nPCGEC2eeyTSIX35hnxtP6kcAdocddgdXT8v2LMOf6X/itc2vQfl9wip8O2GnthIYyJyrJUsY2uvY\n0Zw86yluvJEzd44fp4EVHs7upQDvA8zBXvXrs7IyJaVqIYEzz6SBN3Uqz3XZZcBjj7FMvZz2z/WC\n6+Gqs64qdl+ALyZQFRby/fzyC43MoCCORCoRj4yO5ohPI2fshRfMSy2E8C3+b9P/ITI0Evd0vwcf\n7PoAbRq0wcGUg9hyfItfFBiJ2ocP/vq5haqU3CE1lWHI/Hx64Nq25f0JCcCQIbQgwsJYVelwMPnJ\n6Jl15AhnvyUlMd/svvsqbo1fWMhjNWhQuzLTN29m59aYGL6v/Hy+z02bfKtnXS1GVZbCkxzPOI4h\ny4YgKDAIt3S6BUvjliImKganck+hTYM2WHjjQt9cHAq/RVWWdZ3GjYsPXDNo1gzo3p0eusBAjkJ6\n/33TGDt5kv1+MjJ43969jMc99VT55wsOrp0D0bKzeZ0MBR0aymSxwsIyw7FCCN9lwc4FAIAiexHe\n3PomosOjkZKTAgcc2HtyL+JOxKFrTFdrhRR1Dl8zyG4BEAugI4ALAfxqqTS1lQ8+ALZtY2wtN5c5\nX66zMbdtYwv6kyfpOatXj1MAnnyydnm+3KVzZxqmp04x7y4lxWwZIoTwK45nHMfyA8vRJKIJHHYH\nCooKMOeqOWhUzyw+6tCkg4USirqKr8Vb/gdgCICNVgtSq1mzhuFKI6k/JIQdTQ3i43kzwipZWXU7\nGSomhv3cWremZ2zAAA6xFEJ4lz17WPl94IDHDrk0bikybZk4lXMKaXlpyCvKw97kvejSvMvft9Ag\nLbZEzeNrHrL9VgtQJ2jcmF3/jckARUW8zyAqivfl5dEjZrfTgKvLdOkCLF1qtRRC1B3efRd45x1z\nZNyTT3LubkUUFjLFokGDUttkXHvOtTivRfEhuO0atfOU1EJUGV8zyERlcTiAtWu5imzQgH3FTj+9\n/NeMGQOMHMnkfgBo2RK4+Wbz8bQ0s6+YMRzcZqub4UohRM0TH0+DrGlT5qbabMArr7CptdG+pzT2\n7gUef5zpBRERwIsvMr3AhQ5NOigkKXwSKwyy7wC0KOX+ZwGsdPcgsbGxf2/3798f/fv3r65c/smK\nFWw3ERrKleF337F3T4vSLrGTc85hQv+WLVR2ffsWV3ItWrAVxqlT3A8OZiGAhiyKGmL9+vVYv369\n1WJ4DemvCjh1iu1ljMru0FAuEg3PV2nYbBxJl5fHNIOsLHrVli8vHgEQogaoig7z1V/XdQCeQNlJ\n/SobN7j+enbENzrrHz/OJq933FH1Y/7yC3DllWbPLZuNA8I3q4u1sAa1vahjZGZStxUU0ABLTeXf\n5cvLLqY5doye/mbNzPtSUoC33qL+EsJC3NFhvpbU70ptUb7epaTXKiCg+vN6UlMZxgwL4/GaNqX3\nzVd+RAoKOAs0Lo7GohCidlG/PkclNWjA1IoWLbhfXmVzo0b0ouXmct9mYy6sMY1ECB/H13LIhgB4\nDUBTAKsA7ABwtaUS+Tp33snxRPn5NFSiooDqhD9sNmDfPnrHevY0e275iuGTnc2JA/v2cb99e868\nq+tFB0LUNjp1Ar78krrHnRYzkZFAbCzw/PMMV9rtwLhxXFxWFbtdzZ9FjeGvXii5/A0cDiqtb7+l\nUTJihDlbsrLk5QEPPQTs3g388QcVYdu2HLE0YQJwyy0eFb1KvPEGMH++OeIpMZHDI8ePt1Yu4VUU\nshRuEx8PHD1Kr1qbNlU7xrZtwHPPsRdj9+7AjBnFQ6FCVBJ3dJi/KjgpNG+wYgVnUp52Gl39+/cz\nbDlnDjBwoNXSkfHjgZ9/NpN009KYHzJvXtmvKSxk9VVREeeG1qtXM7IKjyGDTNQYCQnMRQsJYcTh\nxAnqjQ8+sFoy4cdodFJdIykJ2LqVlUkXX8w8jMqQlsa/AQGcZZmSwu0XX2RT1C5dPC5ypenShW0+\n7Hbu5+SUn7Cbk8M2H3v2cL91a/Y2atLE+7IKIfyPAweK92Zs3pwLupwcttIQwkvIIPMncnOB5GQa\nE0ZVpcHvvzNcaQwKb90aWLCgcrMlu3dnvkRiIt3+gOktmziRFU5WsGMHsGEDDcxBg6gwv/+ej116\nKTB8eNmvXbIE2LWL7yMggCOh5s0DJk+uEdGFEH5Gw4bUeUb+WF4e0zbCw62WTNRyZJD5C9u3A088\nQeUQHAxMmwb062c+PncuDTaj/9jRo8BHHwEPPOD+Obp3Z1LsM89QIZ12GnDmmTRkjFFKNd2HbN06\nc6h5URHwySfAhx+yv5DdTuO0PJn+/JMJwcZzIiKYHyeEEKXRrRtTNL75hnojIIC9HpXcL7yMDDJ/\nIDeXxhjAxNKcHODZZ4GVK023enJy8RVccDBDjpVl8GAaYXffzWMHBTGHonNna5rCvvEGjSijivL4\ncWD1avf7rHXrBqxaRWMuMJAexB49vCevEMK/CQxkLu0111CHdujAam4hvIxMfn8gOZltLYycMGOk\nkRFWBNjqIiODrS/y8pjI3qdP1c537rmsqkxPZ17aGWdYN0w7P7/4PLqAgMq14BgyBLjxRl7DpCRO\nJRg50vNyCiFqD4GBQK9eTJGQMSZqCH+tWqpbVUrZ2ZzhFhZGYywvj56eFSvMUuzCQuC114DPPqMB\nM2oUB/FWx6uVm8tzN25snbv+vfeY8xUdTUPMbmfIsrJKMjOTXrIGDTT+yQ9RlaUQwp9R24vaxIYN\nDFMaw74nTWJ4sbZjtwMLFwJff80S9NGjgfPOs1oqUcPIIBNC+DMyyGobqakMU8bEqEmhqFPIIBNC\n+DMyyKxg0yY2ECwqAoYOBS6/3GqJhPB7ZJAJIfwZGWQ1zdatwMMPm0O5c3OBl19mrywhRJWRQSaE\n8Gfc0WGqsvQkX3zBdhONGrG5YHg4k+yFEEIIIcpBBpknCQ9nqNKgqIjeMiGEEEKIcpBB5kluv51d\n4RMSOH4IYINVIYQQQohy8NecDN/NwfjtN858LCwErruOTVaFENVCOWRCCH9GSf1CiFqBDDIhhD+j\npH4hhBBCCD9ABpkQQgghhMXIIBNCCCGEsBgZZEIIIYQQFiODTAghhBDCYmSQCSGEEEJYjAwyIYQQ\nQgiLkUEmhBBCCGExMsiEEEIIISxGBpkQQgghhMXIIBNCCCGEsBgZZEIIIYQQFiODTAghhBDCYmSQ\nCSGEEEJYjAwyIYQQQgiLkUEmhBBCCGExMsiEEEIIISxGBpkQQgghhMXIIBNCCCGEsBgZZEIIIYQQ\nFiODTAghhBDCYnzNIHsZwD4AuwB8BqCBteJUj/Xr11stgltITs/jL7L6i5yi5vGnz4a/yCo5PYu/\nyOkuvmaQrQbQGUB3AAcBPGOtONXDXz4sktPz+Ius/iKnqHn86bPhL7JKTs/iL3K6i68ZZN8BsDu3\nNwNobaEsQgghhBA1gq8ZZK7cB+Arq4UQQgghhPA2ARac8zsALUq5/1kAK53bEwH0BHBTGcf4DcBZ\nnhdNCOGjHAbQ3mohPIT0lxB1D7/UYcMB/Agg3GI5hBBCCCHqJAMB7AHQ1GpBhBBCCCFqCitCluVx\nCEAogFTn/s8AHrZOHCGEEEIIIYQQQgghfJSpYPPYnQDWADjdWnHKxV+a3d4ChouLwIIKX2MggP2g\nF3WCxbKUx3sAkgD8z2pBKuB0AOvA/3kcgEetFadMwsEWODsB7AUww1pxPIa/6DDpL88g/eVZpL98\niPou248A+K9VgrjBFTDbi7zovPkiHQF0AD/kvqbQgsDKtLYAQsAP97lWClQOlwDoAd9XaC0AnOfc\njgJwAL57TSOcf4MB/ALg3xbK4in8RYdJf1Uf6S/PUyv1ly/3ISuPTJftKAAnrRLEDfyl2e1+cDqC\nL3IRqNCOACgAsBTA9VYKVA4/ADhltRBukAj+MABAFugFaWmdOOWS4/wbCv64pZbzXH/BX3SY9Ff1\nkf7yPLVSf/mrQQYA0wD8BeAe+O6qrSRqdls1WgE46rJ/zHmf8AxtwVXxZovlKItAUPkmgR6QvdaK\n4zH8TYdJf1UN6S/v0ha1RH/5skH2Heg2LXm71vn4RABnAFgAYLYF8rlSkawA5bUBWFzj0pm4I6cv\n4rBagFpMFIBPAIwFV5q+iB0MT7QG0BdAf0ulcR9/0WHSX95F+st71Cr9FVxDAlWFK9x83mJYv2qr\nSNbhAAYBGOB9UcrF3WvqaxxH8aTn08FVpqgeIQA+BbAQwBcWy+IO6QBWAbgAwHprRXELf9Fh0l/e\nRfrLO0h/+Qhnu2w/AuBDqwRxA39rdrsOwPlWC1GCYHDsRFswDu/LSbEA5fT1pNgAAB/Aeu9yRTQF\n0NC5XQ/ARlhvGHgCf9Fh0l/VR/rL80h/+RCfgB+YnaCF3NxaccrlEIA/Aexw3t6wVpwyGQLmOeSC\nCZNfWyvOP7garKT5DcAzFstSHksAxAPIB6/nvdaKUyb/Bl3pO2F+NgdaKlHpdAXwKyjnbgBPWiuO\nx/AXHSb95RmkvzyL9JcQQgghhBBCCCGEEEIIIYQQQgghhBBCCCGEEEIIIYQQQgghhBBCCCGEEEII\nIYQQQgghhBBCCCGEEEIIIYQQQgghhBBCCCGEEEIIIYQQQgghhBBCCCFEnaA/gJXO7WsBTPDSec4H\n8GoZjx0B0LiMx74HUL+c4z4GoF7VxaqULO4wHMBc5/aDAO6qpjyAKVMYgI0AAj1wTCGElwm2WgAh\nhN+yEqZx5mm2O2+l4Sjj/ssAHACQWc5xxwL4EEBu1UVzS5ayCARgL+Oxt6spi4EhUz6AHwDcAOAz\nDx1bCOEltHISom7RFsB+AO+DxssiAFcC+BHAQQAXOp93EYCfAPzqfKxDKccaDtO7EwPgcwA7nbd/\nlfL8NwBsBRAHINbl/gud59gJYDOAKBT3xDUBsNr5uncBBJTx3u4AsNy5HQlglfOY/wNwK4BHALQE\nsA7AGufz3ixDpiPO/e0AdgM4xw1ZPgewzfnYSJf7swDMhHld7gWv/WYAfVyeFwvgCQCnAdjhcisE\ncDqAZgA+AbDFeTNeW55MKwAMhRBCCCF8irYACgB0Bn+4twGY73zsOtCoABj2C3JuXw4aAkBxQ2k4\nTINsGYBHnduBAKJLOXcj598g0CjqCiAUwGEwRAnQGAsqcZ7XAExybg8CPUylhQn3udx/E4B3XB4z\nwph/lHhtSZm6uDxvtHP7IdDQqUgW41j1QCPQ2LcDuNm5fRqAP0EjKgTAJucxAeB50CBzZTSApc7t\nxQAudm6fAWCvGzKFATgOIYTPo5ClEHWPPwDscW7vAfOuAHpY2jq3GwL4AEB7MAQWUsExLwUwzLlt\nB5BRynNuAz1HwaBh0sl5fwLM8GRWKa+7BMAQ5/ZXAE6VIUNLAKnO7d2gV+pFAF+Chk9plCZTnPMx\nI8z3K4Ab3ZBlLBgeBOjROhv0ZBUB+NR5fy/Q8Etx7i9D6d5HgMbX/TCNsMsBnOvyeH3QE1ieTPmg\ngRwOIK+M8wghfAAZZELUPfJdtu0AbC7bhk6YCob1hgBoA2C9G8ctK5QIAO1A788FANLBkGk43M/B\nKu/YpXEIQA8AgwG8AL6XqW7KZGBcpyIU15WlydIfwAAAvUHDZ53LsfJgvk9HideX9b5OA/BfsHAi\nx+W5vWD+v1wp7/oEoPK5bkKIGkY5ZEKI0ogGEO/cvteN568BQ3sAw38lQ5bRALJBz1kMgKtBI+EA\naHxc4Hyea6jUYCOYHwbn6xqhdOJhhupOAw2hRaCnrIfz/kwX2UqTqSLKkiUa9EzlAegIGmalsQVA\nP6ecIQBugWksGUZVMICPATwF4DeX166GGRYGgO4VyAQwZFmE4ka4EMIHkUEmRN2jpLfEUcr2SwBm\ngOG6oDKe43DZHguGLXeDeWmuoTUA2AUmqO8HjSQjhFgAhg3ngknv38L0nBnH/g+AvmAocQiYg1Ua\nm2AWJXQFk+Z3AJgMeskA5pV9AxqQZclUEndk+QY0pPaC1+3nEq83SACT9392nm9Piec5wGT98wFM\ngZnY3wI0xi5wyr0HbJNRnkwADVFXWYQQQgghvEp/sGpSmEyHmV8mhBBCCFEjVNQYti5hNIatbP6d\nEEIIIYQQQgghhBBCCCGEEEIIIYQQQgghhBBCCCGEEEIIIYQQQtQQ/w9tumyoeHsVsgAAAABJRU5E\nrkJggg==\n", + "text": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "" + ] + }, + { + "cell_type": "heading", + "level": 3, + "metadata": {}, + "source": [ + "Min-Max scaling (Normalization)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top]](#Sections)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "An alternative approach to standardization is the so-called Min-Max scaling (sometimes also referred to as \"normalization\"). \n", + "In this approach, the data is scaled to a fixed range - usually 0 to 1. \n", + "The cost of having this bounded range - in contrast to standardization - is that we will end up with small standard deviations, for example in the case where outliers are present.\n", + "\n", + "The equation to calculate the \"normalized\" score is:\n", + "\n", + "\\begin{equation} X' = \\frac{X - X_{min}}{X_{max}-X_{min}} \\end{equation}" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "minmax_scale = preprocessing.MinMaxScaler(feature_range=(0, 1)).fit(X_train)\n", + "X_train_minmax = minmax_scale.transform(X_train)\n", + "X_test_minmax = minmax_scale.transform(X_test)" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 19 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "f, ax = plt.subplots(1, 2, sharex=True, sharey=True, figsize=(10,5))\n", + "\n", + "for a,x_dat, y_lab in zip(ax, (X_train_minmax, X_test_minmax), (y_train, y_test)):\n", + "\n", + " for label,marker,color in zip(\n", + " range(1,4),('x', 'o', '^'),('blue','red','green')):\n", + "\n", + " a.scatter(x=x_dat[:,0][y_lab == label], \n", + " y=x_dat[:,1][y_lab == label], \n", + " marker=marker, \n", + " color=color, \n", + " alpha=0.7, \n", + " label='class {}'.format(label)\n", + " )\n", + "\n", + " a.legend(loc='upper left')\n", + "\n", + "ax[0].set_title('Training Dataset')\n", + "ax[1].set_title('Test Dataset')\n", + "f.text(0.5, 0.04, 'malic acid (normalized)', ha='center', va='center')\n", + "f.text(0.08, 0.5, 'alcohol (normalized)', ha='center', va='center', rotation='vertical')\n", + "\n", + "plt.show()" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "display_data", + "png": "iVBORw0KGgoAAAANSUhEUgAAAmgAAAFXCAYAAAAWK7KsAAAABHNCSVQICAgIfAhkiAAAAAlwSFlz\nAAALEgAACxIB0t1+/AAAIABJREFUeJzs3Xd4VGXawOHfTHpITwhEFMHGAjYUg8vKGv10YQE7KAqs\niK6KK7LoutgQLOyquGuhCCgdARERVBARBJFFCb1Ii5SQhPTeJzNzvj/eSS9MwkzOTOa5r2uuzJxz\n5pxnBvLmOW8FIYQQQgghhBBCCCGEEEIIIYQQQgghhBBCCCGEEEIIIYQQQgghhBBCCCHaPC+9AxBu\naR1gBA44+FghhBBCCI9SBBTaHlagpMbrB3WMq6XiUJ+j8jMkAZ8BvZtxjsnAYkcHpuN1hBCKo8u7\nLcCjTezvQu3yKA34GritGdcYBfzUgtiaq7WuI86TUe8ARKsJAoJtj0RgcI3Xy2oc5936obVYCtWf\n4UbgKKrguVXPoIQQurO3vLOXZudxobZrXA18D3wJPNyC6wkhPNQpqpOYOCAZ+CeQCiwEwoBvgAwg\nB3Un2KnG+7dQfTc5CtgGTLUdexIY0MJjuwJbgQJU4TaDxmue4lC1ZnVNA3bWeP0BcAbIB3YBN9m2\nDwDKARPqjnevbfsjwGFbDCeAx2ucKwr1veQC2bZYDbZ9FwBfoL6zk8DYc1xHCNE6apZ3RuAF4Dcg\nC1XrHm7b5w8ssW3PBeKBaGAKYAZKUb/DHzZwjS6oGrS6lR7PoWrTKlVeuwD4Fbjbtr277fxm2zVy\nbNsHocqMfFQ5NqnGuRqLF1SiOBc4iyrf37DF1th1hBAuom6CVgH8G/BB/dJHAPfYngcBK1B3gpU2\nA6Ntz0ehko9HUcnKk6iarZYc+zPwDqoW7w+oQmlRI58hjoYTtFsBCxBgez0cVQAbgWdRSaivbd+k\nBs4/EJUoAvwRKAautb3+N/ARqu+mly1GbOfeDbxii70rKrn7UxPXEUK0jprl3ThgO+qGygeYBSy1\n7XsC+ApV7hmAXqjaMKhdjjWkCw0naJfYtnezvR4CdLQ9vx/VFNvB9vph6jc93gz0tD2/CpXs3WVH\nvF+iyqoAoD2wg+qbzYauI1yQNHEKUAXIJFSiVoa6q/rS9rwI+BeqoGhMIupuTUMlIjFU38nZe2xn\nVP+xV1F3d/9DFT6Ghk/TqLO294TZXn+Kuru0Av8F/KguLA0NnH8dqkAHVUO2AZWogUouY1CFscUW\nI8ANqNq1N22xnwI+AYY1cR0hROt7AnUjdRZV3r2GSpq8UL/fkcDlqPJpL6qWqVJLfofP2n5G2H6u\npLpGbQWQAPRp4vw/omraAA4Cy6kuixuLtwPwZ2A8qrYsE3if2uWRcAOSoAlQv8CmGq8DgdnAaVQt\n1o+oKvPGfrFrVuGX2H4GNfPYC1CJYVmN/Q3VkJ1LJ1RhlWd7/Q9Uk2UeKlELRSVTjfkz8AuqCTMX\nVaMWads3FdU8sQFVQzbBtv1iW/y5NR4v0niSKoTQRxfUzWfl7+lh1E1VNKo7xXeoJCgFeJvafXLt\n7YdWU2XXkMqmxL+gEqnK619JdfnSkD6o2rsMVBn2RI3jG4v3YlTtYGqN68xC1aQJNyIJmoD6Bc9z\nwBVALCqhuRnn1wKlou4yA2ps69yC89yDam4sBfoBzwNDUTVq4aiEs/Jz1P3cfqh+ZO+gCuxwVI1a\n5fFFqITvUuBOVJPprai+Iadsx1c+QlAdk0HV3gkh9HcG1S+05u9qIKr8MQOvo5oU+6J+f/9ie19L\nkjNQ5VE6cAyVOM0B/oYq68KBQzReHoFqfl0NXIgqw2ZR/Xe7sXjPoPq9Rtb4jKGoJtLz+SyilUmC\nJhoShEpw8lEFyaSmD3eIRFQn/smou7/fowocewoTA+pOdRKqf9tLtu3BqEIsC9Xv7FVU4lQpDXVH\nXVlA+toeWaik6s9U9yPDFs9ltuMLUM2cFlTn3ELUQIsAVHPJlVRP+ZFe5zpCCH3MQnXZqLz5a4+6\n2QLVr/Uq1O9vIaoJ1GLbl466MTuXyt/xDsDTqDLnRdu2dqjyLAv1t/cRVDlRKR2ViPnU2BaEqgEz\noW6YH6K6TGws3jRULf9/UWWg0RZ7ZVeNhq4jXJAkaALqJ0HvoxKNLFSH2m8bOKbme+vua+mxw1GJ\nWTZq1NFn1G56rfu+C6iedygedSd5M7DRdsx62+M4qrm2FHV3Welz289sVHJYCDyD6huSg5ovaU2N\n4y9DjS4tRH0vM1DNv1ZU8nYtagRnJupOuTIZrHsdIYQ+PkD1bd2Ausn6GZX4gOq8/znqxvQwagT6\n4hrvG4IqF95v4vx5qJr2A6iauiHAAtu+w8B/bNdMQyVn22q8dxOqv1kaqkkT4ClULVkBMBFVJlZq\nKt6/oG42D9ti/pzqwQkNXUcI5qGy94ON7B8O7Ef95/4fai4Z4bk+o3Vq74QQQgiP1g81FLixBO33\nqLZyUHcfv7RGUMJl9EZVxRtRzYulwDW6RiSEEEJ4iC40nqDVFI6aYE94jsGoJshi1KoAMgO3EEII\n0Uq6YF+C9g9UPx4hhBBCCI/iqusu3oKatfkPDe289NJLtRMnTrRuREIIPZ1ADdJwe1J+CeGRml2G\nueIozquBj1FDn3MbOuDEiRNomuYyj0mTJukeg8Tj3jFJPE0/sG+KA7cg5Zd7xeOKMUk87hWPprWs\nDHO1BK0zsAoYgZqxXQghhBDC47R2E+cy1DxVUahlfCZRPVnebNSkfuGoRV5BTbwXixBCCCGEB2nt\nBO3Bc+x/zPZwK3FxcXqHUIvEc26uFpPEI/Tiav/WrhYPuF5MEk/TXC2elnLXpWc0W5uuEMIDGAwG\ncN/yqi4pv4TwMC0pw1x1FGeLREREkJvb4LgC0Uzh4eHk5OToHYYQHkPKL8eSMky4O3e9I23wDtRg\nMCB3po4h36VwJZ5Qgya/c44l36dwJS0pw1xtFKcQQgghhMeTBE0IIYQQwsVIgiaEEEII4WIkQRNC\nCCGEcDGSoOlswYIF9OvXT+8whBCi2aT8EsJ5JEHzMNOnT6d37974+/vzyCOP6B2OEELYTcov4Uk8\nPkEzmWD+fCgtVa/T0mD5cn1jcqZOnToxceJERo8erXcoQggH2LABjhxRz61WWLwY8vL0jclZpPwS\nnqTNJ2gnTsCWLdWvt2xR2yp5e0NREUyeDKdPw8svQ0hI7XOUl6uCr1JlMtccSUlJ3HvvvURHRxMV\nFcXYsWMbPG7cuHF07tyZ0NBQevfuzbZt26r2xcfH07t3b0JDQ+nYsSPPPfccAGVlZYwYMYKoqCjC\nw8OJjY0lIyOjwfPfc8893HXXXURGRjb/QwghWpXJpG4YKyrU6/R0+Oab2sdERsKUKXD4MLz/Phw7\nBgEBtY+pWWZZLOq8zSHllxCtr80naH5+sGAB/PCDeixYoLZVMhrhb38DHx8YOxYGDlSPmpYuhQ8/\nVEna2bPq+ORk+2OwWCwMHjyYrl27kpiYSEpKCg8+2PCypLGxsezfv5/c3Fweeughhg4dislWmo4b\nN47x48eTn5/PyZMneeCBBwBYuHAhBQUFJCcnk5OTw+zZswmoW0LXIRM4CuH6jEY4eRLeegtSUtQN\nZF3XXw9//ztMmADbt8PEibXLuFOnVNmWnq6Ss//8B1autD8GKb+E0EebT9AuvBDefBPee0893nxT\nbaspI0MlXgDx8fVryB58EDIz4cUX4aWXYNiw+udoSnx8PKmpqUydOpWAgAD8/Pzo27dvg8cOHz6c\n8PBwjEYjzz77LOXl5Rw7dgwAX19fEhISyMrKIjAwkNjY2Krt2dnZJCQkYDAY6NWrF8HBwU3GZJvV\nWAjhwry94Z//hOxsePJJGDQIBg+ufYzVClu3queVCV1NXbvC3XfDCy+oR3ExDBlifwxSfgmhjzaf\noAEcP97wc1DNlxMnqgJrzRqVeL3zTu1j/P3hscdUE0J2Ntx2W/Oun5SUxMUXX4zReO6v+91336VH\njx6EhYURHh5Ofn4+WVlZAMydO5fjx4/TvXt3YmNjWbt2LQAjR46kf//+DBs2jE6dOjFhwgTMZnOT\n15E7UCHcQ3Z2dZ+yQ4eqmzsrLVwIOTmqVmzCBNXcmZ5e+5g//xmysuDoUfjrX8HX1/7rS/klhGgO\nrSENbY+P17SHH9a0pCT1ePhhta2mpKTq5xaLpiUn196fkqJpo0Zp2ldfadpLL2nae++p4+y1fft2\nLTo6WjObzfX2zZ8/X7vppps0TdO0rVu3atHR0dqhQ4eq9oeHh2ubNm2q976VK1dq/v7+WklJSa3t\np0+f1nr06KHNnTu3yZheeeUVbdSoUY3ub+w7FkIPQFv6i9zoZ6yruFjTHn1U077+WtMqKjRtyhRN\nmzq19jEZGZpWVlb9OjlZ06zW6tdms6a9/bamvfqqpq1apc6Xlmb/d++O5ZemSRkmXAstKMPafA1a\nz57wr3+pmrELL1TPe/asfUzN5kqjETp1qr1/2zbVzHnHHaq2rbhYjfa0V58+fYiJieGFF16gpKSE\nsrIytm/fXu+4wsJCvL29iYqKwmQy8frrr1NQUFC1f8mSJWRmZgIQGhqKwWDAaDSyefNmDh48iMVi\nITg4GB8fH7y8vBqMxWKxUFZWhtlsxmKxUF5ejsVisf/DCCFaTWCg6loxeHB1c2fd5sn27Wv3OevU\nCWq2AKakqGbQl1+Ge+5RzZ0//2x/DFJ+CSGao9EM1VWdOXNGu/vuu7XIyEgtKipKGzdunKZpmrZg\nwQKtX79+mqZpmsVi0UaPHq2FhIRoMTEx2jvvvKN17dq16g50xIgRWnR0tBYUFKRdeeWV2po1azRN\n07Rly5Zp3bp109q1a6d16NBBGzdunGZppIpv0qRJmsFgqPV47bXX6h3nyt+l8Dx4aA2aq3C38kvT\nXPv7FJ6HFpRh7trT0vZ5azMYDNI3wUHkuxSuxNYp3F3Lq7qk/GoF8n0KV9KSMqzNN3EKIYQQQrgb\nSdCEEEIIIVyMJGhCCCGEEC5GEjQhhBBCCBcjCZoQQgghhIuRBE0IIYQQwsVIgiaEEEII4WIkQRNC\nCCGEcDGSoOlswYIF9OvXT+8whBCi2aT8EsJ5JEHzICaTiUcffZQuXboQEhJCr169WL9+vd5hCSHE\nOUn5JTyNJGgAJ07AqFFw223wj39ATo7eETmF2Wymc+fObN26lYKCAt58803uv/9+EhMT9Q5NCNFS\nZWXw73/D7bfD0KGwc6feETmFlF/C07T9BE3T4OuvYcgQ9fj6a7WtUn4+PPkkHD8OXl7w44/w7LO1\njwHIzYXt22HfPrBamx1GUlIS9957L9HR0URFRTF27NgGjxs3bhydO3cmNDSU3r17s23btqp98fHx\n9O7dm9DQUDp27Mhzzz0HQFlZGSNGjCAqKorw8HBiY2PJyMiod+7AwEAmTZpE586dARg0aBBdu3Zl\nz549zf48QohWcvq0KqPuuAPeeAOKimrvnzoVVq4EgwHS0+GZZ+DkydrHWCywdy/8/DPk5TU7BCm/\nhGh9bT9B27gRXnsNsrPVY/Jk2LSpev/x46rAi4wEX1/o0AF+/VUlZJUSEuC++1Ti9te/qlo2s9nu\nECwWC4MHD6Zr164kJiaSkpLCgw8+2OCxsbGx7N+/n9zcXB566CGGDh2KyWQCVOE3fvx48vPzOXny\nJA888AAACxcupKCggOTkZHJycpg9ezYBAQHnjCs9PZ3jx4/Ts2dPuz+LEKIV5eWpMmffPigthTVr\n4KWXah+zcSO0bw9+fhAaChUVKhmrZDLBuHHw+OMwfryqZTt92u4QpPwSQh+tmaDNA9KBg00c8yGQ\nAOwHejnkquvXq4IrKEg9/P3h22+r9wcEqLvLyloxs1ndidYsIN54QzUjtG+vErgff4QffrA7hPj4\neFJTU5k6dSoBAQH4+fnRt2/fBo8dPnw44eHhGI1Gnn32WcrLyzl27BgAvr6+JCQkkJWVRWBgILGx\nsVXbs7OzSUhIwGAw0KtXL4KDg5uMqaKiguHDhzNq1CiuuOIKuz+LEKIV/forFBaqsicgADp2VDX5\nJSXVx7RrB+Xltd/Xrl318+++U+/p0EGdp6hINYnaScovIfTRmgnafGBAE/sHApcBlwOPAx855KpB\nQbVruyoq1LZKPXpAXBykpcHZs5CVBU89VTtBO3sWKgsMg0H9bKAKvjFJSUlcfPHFGI3n/rrfffdd\nevToQVhYGOHh4eTn55OVlQXA3LlzOX78ON27dyc2Npa1a9cCMHLkSPr378+wYcPo1KkTEyZMwNxE\nDZ/VamXkyJH4+/szffp0uz+H29M0+P57ePVV+PBD9W8thCvz91c3j5VdLsxm1RXDx6f6mH/+UyVs\nZ8+qR7ducPPN1fvT0lS5VVl2BQVBcrLdIUj5JYRn6ELjNWizgAdqvD4KdGjkWK0hDW4/cULT+vXT\ntOuuU49+/dS2msxmTduwQdMWL9a0X36pf47nn9e066/XtEGDNO1Pf1LPd+5sMIaGbN++XYuOjtbM\nZnO9ffPnz9duuukmTdM0bevWrVp0dLR26NChqv3h4eHapk2b6r1v5cqVmr+/v1ZSUlJr++nTp7Ue\nPXpoc+fObTAWq9WqjRo1Srv11lu1srKyRmNu7Dt2a0uXqv8DN96o/g0HDtS03Fy9oxJ2ALRGygJ3\n1OhnrKeiQtOeeELTevVS/2evu07TGvrdPnxYlV9ffaVpdcoEbds29d7+/VUZdt11mjZxot3fvTuW\nX5rWRssw4bZoQRnmSn3QOgFJNV4nAxee91kvuQSWLFG1Yk89BZ9+qrbV5OWlRkCNGAF9+tQ/x0sv\nQa9eqgNufr7qx9G7t90h9OnTh5iYGF544QVKSkooKytj+/bt9Y4rLCzE29ubqKgoTCYTr7/+OgUF\nBVX7lyxZQmZmJgChoaEYDAaMRiObN2/m4MGDWCwWgoOD8fHxwcvLq8FYxowZw9GjR/nqq6/w8/Oz\n+zO0CR9/DBEREBUFMTHq37OBfwchXIa3t6rtnTgRHn0UPvgAHnmk/nHdu6vy6447atf+A/Ttq8q+\n3Fz1f/7GG+H55+0OQcovIfThrXcAdRjqvG4045w8eXLV87i4OOLi4ho/a+fOqnBrqbAwmDNH9d3w\n81ODCZrBaDTy9ddf88wzz9C5c2cMBgPDhw+nb9++GAwGDLamhwEDBjBgwACuuOIK2rVrx/jx46tG\nLAF89913PPfcc5SUlNClSxeWL1+On58f6enpjBkzhuTkZIKCghg2bBgjR46sF0diYiJz5szB39+f\njh07Vm2fM2dOo51+2xSLpXbTUOU24XK2bNnCli1b9A7DaZpVfvn6wj33tPxiBgOMHq0SOJOpdhcP\nO0j5JUTzOaIMq5sQOVsX4Gvgqgb2zQK2AMttr48CN6MGFtRlqzGszWAw0NB20Xxt8rucORM++UT1\nJywvh8BAWLZMdZ4WLs2WBLR2eeUsUn61Avk+hStpSRnmSjVoXwFPoxK0G4E8Gk7OhGiZJ55QydmW\nLWpalaeekuRMCCGES2rNO9JlqBqxKFTiNQmobG+abfs5HTXSsxh4BGhsBkK5A3Uy+S5bR0ZxBisP\nr2RM7zFVTUWiPqlBE80l36dwJa5eg2ZPJ4GnnR6FEC5kwb4FzN83nz6d+nD9BdfrHY4QQggX4Uqj\nOIXwKGlFaaw6sopg32Bm7Jwhd/tCCCGqSIImhE4W7V+EVbMS3S6ag+kH2ZMqawoKIYRQJEETQgdp\nRWksP7QcL6MXeWV5lFnKpBZNCCFEFVcaxSmExyipKOGWrrdgsVbPwxbuH65jREKIlig3l2OymAj2\na3r9UCGay11HRckoKCeT71K4EhnFKZqrtb7Pt7e9zcnck8waPEtGYotGtaQMkyZOnS1YsIB+/frp\nHYYQQjSbp5dfaUVpfHn0S3an7uZA+gG9wxFtjCRoHmbEiBHExMQQEhLCJZdcwpQpU/QOSQgh7OJq\n5dfCfQuxalZ8vHykD6lwOEnQbM4WnmXct+MwWUx6h+JUL774IqdOnaKgoIBvv/2WadOmsX79er3D\nEkKcB03TmLR5EvvT9usdilO5UvlVWXsWGRhJZEAke1P3Si2acCiPSdBWHVnFqiOrGt0/f+981ias\nZf1vjf+yn8k/Q5m5rEXXT0pK4t577yU6OpqoqCjGjh3b4HHjxo2jc+fOhIaG0rt3b7Zt21a1Lz4+\nnt69exMaGkrHjh157rnnACgrK2PEiBFERUURHh5ObGwsGRkZDZ6/Z8+e+Pv7V7329vYmOjq6RZ9J\nCNE6UgpSeOWHV2oNKqlpf/p+Vh5ZyXu/vNdoLU5JRQkpBSktur6UX/V9c/wbSipKyCvLI7s0G5PV\nxGe/fqZLLKJt8ogErchUxAe/fMAHv3xAsam43v6zhWf56thXdArpxIydMxqsRSs3l/PE10+wcN/C\nZl/fYrEwePBgunbtSmJiIikpKTz4YMMLK8TGxrJ//35yc3N56KGHGDp0KCaTimfcuHGMHz+e/Px8\nTp48yQMPPADAwoULKSgoIDk5mZycHGbPnk1AQECj8Tz11FO0a9eOnj178sorr3Ddddc1+zMJIVrP\n3L1zWfHrCrYmbq23T9M0Zu6cSYhfCL9m/Mru1N0NnmPO7jmMWTsGs9XcrGtL+dWw+3vez+dDP2fB\nXQtYcNcCVgxZwbO/f1aXWETb5BEJ2qojqygxl1BiLuGLI1/U2z9/73w0NEL8QsgpyWmwFm1dwjpS\ni1JZdGAReWV5zbp+fHw8qampTJ06lYCAAPz8/Ojbt2+Dxw4fPpzw8HCMRiPPPvss5eXlHDt2DABf\nX18SEhLIysoiMDCQ2NjYqu3Z2dkkJCRgMBjo1asXwcGND/meOXMmRUVFbNy4kVdeeYX4+PhmfR4h\nROtJLkhm7fG1RLeL5sMdH9arRdufvp+9qXuJDIjE19uX6fHT69WiZZVk8dmhz0jKT2LDiQ3Nur6U\nXw0L8QuhW1S3Wo+owChdYhFtU5tP0IpMRczdM5cw/zDC/MOYu2durVq01MJUPj/8ORXWClILUykx\nlzA9fnqtu8xyczkzd84kMjASk8XEZ4eaV42dlJTExRdfjNF47q/73XffpUePHoSFhREeHk5+fj5Z\nWVkAzJ07l+PHj9O9e3diY2NZu3YtACNHjqR///4MGzaMTp06MWHCBMzmpu+SDQYDcXFxDB06lGXL\nljXr8wghWs+8vfPQ0AjzDyO5ILleLdrMnTPJL88nrSiNMnMZO5J31FuVYsmBJVg0C2EBYfXKt3OR\n8ksIfbT5iWpXHVlFWnEaoX6hAOSX5bPqyCpGXjMSAD9vP8bGjkWj+o7Tz8uv1jnWJawjryyPjsEd\n8TZ6s+jAIh648gHC/MPsiuGiiy7izJkzWCwWvLy8Gj3up59+YurUqfzwww/07NkTgIiIiKq74csu\nu4ylS5cC8MUXXzBkyBBycnIICAjg1Vdf5dVXXyUxMZGBAwfSrVs3Ro8efc7YKioqiIyMtOtzCCFa\nV0pBCquOrMLL6EVGcQYl5hI+3PEhN3e5GaNBJUwDLxtIbKfYWu+LCIioel5ZexYREIGPlw9phWls\nOLGBgZcPtCsGKb+E0EebT9A6h3bm8esfr7XtotCLqp5HBETw6HWPNnmO1UdXY9WsZBSpjqsV1gq2\nndnG4CsG2xVDnz59iImJ4YUXXuC1117DaDSyZ8+ees0EhYWFeHt7ExUVhclk4q233qKgoKBq/5Il\nS+jfvz/t27cnNDQUg8GA0Whk8+bNREVF0aNHD4KDg/Hx8WmwIM3MzGTTpk3ccccd+Pv7s3HjRj7/\n/HM2btxo1+cQQrQuL6MXo64dhUWrbtYM8gmqdczd3e9u8hxbTm+h3FJObmkuABbNwpqja+xO0KT8\nEkI0h9aQxrafr6LyIi2zOLPWw2wxN+scZ86c0e6++24tMjJSi4qK0saNG6dpmqYtWLBA69evn6Zp\nmmaxWLTRo0drISEhWkxMjPbOO+9oXbt21TZt2qRpmqaNGDFCi46O1oKCgrQrr7xSW7NmjaZpmrZs\n2TKtW7duWrt27bQOHTpo48aN0ywWS70YMjMztZtvvlkLCwvTQkNDtRtuuKHqHHU567sUoiWAtjTB\nVKOf0RkqLBX1yq9iU3GzzuFu5ZemSRkmXAstKMPcdV0K2+etTZZKcRz5LoUrkaWeRHPJ9ylciSz1\nJMR5SC1MZV3COr3DEEIIIdp+HzQh7DUjfgbf/vYt18dcT4egDnqHI4QQwoNJDZoQwOm802w4uQEM\nsHB/8ycjFkIIIRxJEjQhgE92f4LBYCC6XTSrjqwivShd75CEEEJ4MEnQhMc7nXeatQlrCfAOwGQx\nUVJRIrVoQgghdCV90ITHSy1M5YrIK7BiBdTcePll+TpHJYQQwpO567D1BoepR0REkJubq0M4bU94\neDg5OTl6hyEAk8XE7F2zeaL3E/h6+eodji48YZoNKb8cS8ow4UpaUoa1qRo0+WUUbdG3Cd8yfed0\nLgq56Jyzxgv3JeWXEKIm6YMmhAszWUzM3DmTcP9wZu6aicli0jskIYQQrUASNCFc2LcJ35JTmkP7\ndu3JLc1l3XGZSFcIoY9iUzFWzap3GB5DEjQhXJTJYmLGzhloaOSX5aOhMWPXDKlFE0K0Ok3TeGrt\nUyzav0jvUDxGm+qDJkRbUlJRwlXRV1FqLq3aFuAdQLGpGN8AzxwsIITQx46UHRzIOMCpvFPc1/0+\ngv2C9Q6pzXPXUVENjoISQrRNnjCKUwhXpWkaI1aNIDE/EZPFxFM3PMWoa0fpHZZbkcXShRBCCOFQ\nO1J2cDznOGH+YYT6hzJv7zwKywv1DqvNkwRNCCGEEI1avH8xJrOJnNIcik3F5JXl8f3J7/UOq81r\n7SaDAcD7gBfwCfB2nf1RwBKgI6p/3LvAggbOI00EQngQaeIUQj+JeYnkltWeRLlLWBfC/MN0isj9\ntKQMa80Czws4BtwGpAA7gQeBIzWOmQz4AS+ikrVjQAfAXOdcUsAJ4UEkQRNCuDNX74MWC/wGnAYq\ngOXAXXWOSQVCbM9DgGzqJ2dCCCGEEG1aa06z0QlIqvE6GehT55iPgR+As0AwcH/rhCaEEMJTma1m\nDBjwMnop1YUsAAAgAElEQVTpHYoQVVozQbOnTv8lYB8QB1wKfA9cA9QbLjJ58uSq53FxccTFxTkg\nRCGEK9iyZQtbtmzROwynkfLLtUzZOoUAnwD++Yd/6h2KaCMcUYa1Zp+OG1F9zAbYXr8IWKk9UGAd\nMAX4n+31JmACsKvOuaQPhxAeRPqgCWc5k3+GISuGYDQYWT1sNR2DOuodkmiDXL0P2i7gcqAL4As8\nAHxV55ijqEEEoAYHdANOtlJ8QgghPMzcPXMxYMCqWWUZI+FSWjNBMwNPA98Bh4HPUCM4n7A9AP4F\n9Ab2AxuBfwI5rRijEEIID3Em/wzf/vYtkYGRRAZGsurIKtKK0vQOSwig9Seq/RZVK3YZ8G/bttm2\nB0AWcAeq39lVwNJWjk8IIYSHWPHrCoorisktyyWvLI/C8kK+OPyFLrHkluby1ra3sFgtrXbN1MJU\n0ovSW+16onlksXQhhBAeaUiPIfz+wt/X2tY5tLMusSw9uJR5e+fRO6Y3t11627nfcJ40TePlH17G\nz8uPjwZ/5PTrieaTBE0IIYRH6hLWhS5hXfQOg5zSHD49+ClRgVFM3zmdW7re4vQpP/ak7uFgxkEA\nDqYf5KoOVzn1eqL5ZC1OIYQQQkfLDi6jwlJBZGAkKQUpbD612anX0zSN6fHT8fXyxdvozUe7pAbN\nFUmCJoQQQugktzSXhfsXggGyS7IxWU1M3zndqX3R9qTu4VDmIcL9w4kMiGTn2Z0cTD/otOuJlpEm\nTiGEEEIn5ZZyBlw2ALO1elXDIN8gLJoFL5zTzLnx5EYsVgvpxWqAgNli5vsT30szp4tx14kfZaJH\nITyITFQrhOOYrWZMFlOtbX5efrLUlRO1pAxz1wJPCjghPIgkaEJXp07B4sVQVAR//jPccoveEQk3\nIwmaEKJNkgRN6CYpCUaMgNJS8PYGkwleew0GDdI7MuFGXH2pJyGEEMK9bNigas46dIDISAgKgoUL\n9Y5KeABJ0ITQWWphKmO+GUOZuUzvUIQQ52IwgNSAilZgb4LWHfgz0B/4nfPCEcLzzN83n+9Pfs83\nx7/ROxQhRF233QaBgZCeDrm5UFCgmjyFcLKm2kO7AuOBgUAKcNZ2fAxwIfAN8B5w2rkhNkj6cLTU\n6tUwbZrqT3HHHfDcc+DrW/uY4mL46ivIyoLeveH3v2/4XOK8pRamcvfyuwnwCcDb6M03D32Dv7e/\n3mG5HOmDJnSVkAALFqimzkGD4PbbVU2aEHZy9CCBFcDHwBagos4+H+AW4DHg/uZc0EGkgGuJHTvg\nb3+DsDDw8VF3hKNHw9NPVx9TWgqPPgpHj4KXl6rKnzABhg7VL24dWTUrU/83lceue4zIwEiHn/9f\nP/2L1UdX0yGoA2lFaUz4wwSG9Bji8Ou4O0nQhBDuzNGDBO4Hvqd+coZt2wb0Sc5ES+3Yoe76AgLU\naKSwMNhcZ0mRX35Rd4udOkHHjhARoWrcPPQPytbErczbN4/FBxY7/NzpRems+HUF5ZZyUgpSKDYV\nM3PnzHrzEwkhhPA8Ta0kcB+goTK+yp/YngOscmJcwhnCw8FSY/mQsjKIiqp9THl57ap7b2+1TdM8\nrkrfqln5cMeHRAZE8tmhzxhx9QiiAqPO/UY7+Xn78Xzf57Firdrma/TF0GYqioQQQrRUUwnaHahk\nLBroC/xg234LsB1J0NzP3XervmWnTlXXpI0fX/uYXr1Uh9isLPUzNxfuuguMnjfgd2viVpLyk+gQ\n1IGM4gyWHFjC32/8u8POH+YfxvCrhzvsfEIIIdoOe27Vvwf+AqTaXscAC4E/OSsoO0gfjpYqLoZt\n29Rki717Q0xM/WMSEuA//1F91Pr1U/3W/PxaP1YdWTUrQ1YM4bec3wjyDcJsNWPVrKwfsd6htWjC\nPtIHTQjhzlpShtmzWPpFQFqN1+lA5+ZcRLiQdu2gf/+mj7n8cpg1q3XicVEWq4XYTrF0j+petc3L\n6EWFpaEumUIIIYRj2ZPNTQeuAJbajn8ASADGOjGuc5E7UNHmlZnL2JO6h74X9dU7FN1JDZoQwp05\na6mnscAs4BrgamA2+iZnQniE1UdX88y3z5CYl9ii929N3MrPST87OCohhBCtwZ4ETQP2AOtQE9d+\nBwQ7MyghPF1pRSmzd8/GZDExd+/cZr+/3FzOGz++wRtb38BsNTshQiGEEM5kT4L2OPA5qhYN1CoC\nq50WkRCCNcfWUGQq4sKQC1n/2/pm16KtS1hHXlkemcWZbDixgdLS6n2aRq3XQgghXI89CdrfgJuA\nAtvr46ipN4QQTlBaUcrsXbMJ9lUV1Rpas2rRys3lzNw5kxD/EIL8gnhv+3SeGGPmt99UcrZwIXz0\nkbOiF0II4Qj2jOIstz1qvkd6uArhJAk5CVixYqowUVpRirfBmwPpB7BqVoyGc99TrUtYR2pRatXS\nVKklZ7j9zg1MmjSQrl2hsBBefNHZn0IIIcT5sCdB+xF4GQgEbgeeAr52ZlBC6OnAAThxAu65R73+\n4gvo1g2uvLJ1rn91h6v5cdSPLX5/mbmM319YY4H7CEhJLeG33yAvD8aNg5degokToWtXBwQshBDC\n4ewZ8mlELYpeOTHtd8An6FuLJsPUhdPk5KgapttvB6tVLVc6ZYpaltQdaRrMmweffAIdOqgEdOJE\neOQRvSOzn0yzIYRwZ86aqHY4sAyYU2PbYOCb5lxICHcREQH//jc8/LB6vXBh48lZTmkO4f7hlb98\nLik/H5KTYe5cmDABLrhALSQhhBDCddkzSGAa8BPQo8a2N5wTjhCtI60ojTe3volVsza4f+NGtRRp\nYCD88EODh5BTmsPQFUOJT4l3YqTnLywMHn8c3n0XnnxSLSSRnQ1mmX1DCEAt7Sa1msLV2JOgnQIe\nRU21cb9zwxHCyYqL4ddfmb/5PRbvX8xPiT/VO2TbNtWs+dFH6vH997B9e/1TLTu4jOTCZKbHT3f5\nwn3XLnjgARg0SDXf+vvD2bN6RyWEa/jvz/9l1m4nLW+XnQ0zZsDrr6uCxcXLCuE67GmX2Qv0AqJQ\nTZ0HUIMFrnZiXOcifTjchMVqwcvopXcYysmTMGYMqeVZ3H3tMbwjo4i54npW3P95rdGRZjMUFama\nJ1Ad64OCwLtGh4Cc0hwGLx1MsF8wOSU5zBg0g9hOsa38gTyH9EETzpJWlMZdy+/CaDCy9qG1RAQ4\nsLNpXh6MGAGpqeDjAxUVqp/B/frXdWRmQvv2jb8WjuWspZ5SbT+zgAGAFWil8WzCneWU5nDfivtI\nyk/SOxRl4kQoLGTB5cVoPl6En80l8ezherVo3t7VyRmo5951emsuO7iMCmsFvl6++Hn7MW3HNJev\nRRNC1Ldw30I0TcNitbDkwBLHnnzbNkhLUx0/27eH8HD4+GPHXqMFNE31s/3sM/V6zRqYPBksFl3D\nEnXYk6ANrPHcAjxv5/saMgA4ilpsfUIjx8Shau0OAVtaeB3hApYeXMrBjIPM2zuv1a5ZVARLl1YX\nNKdOwfr1tp0nT5LWPoAVEaloGMj2s1BaXsy0+GmN9kVriMliYuWRlaBBdkk2FquFw5mHOZx52PEf\nSAjhNGlFaXx59EsiAyMJDwhn2aFl5JTmOO4CdTMeLy9Vi6YzgwFeeQW2bIHhw+Gbb1SC5uUijR1C\naWoU5wfAOBqe80wD7mzmtbyA6cBtQAqwE/gKOFLjmDBgBtAfSEY1qwo3lFOaw9KDS7kk/BLWJqxl\ndK/RXBR6kdOv6+sLR4/Ce++pecxee011kAege3e0E/u4PycGs2aB4hLoM5Dgzr9TtV92Vj77GH2Y\ne+dcys3ltbZfHnm5Yz+MEMKplh9aTl5ZXtUNWn55PisPr+Tx6x8/xzvt1KcPBAer9kM/PzVL9KOP\nOubc5ykiAm68EVauhL59pXnTFTX1J6k3sAtVo9WQLc281u+BSahaNIAXbD/fqnHMU0BH4NVznEv6\ncLi46fHTWbh/IR2DOpJRlMHAywcyKW5Sq1zbZILHHoPcXHj+efjjH207UlLgb39T/UEA/vpXdaBw\nedIHzc2YzWoodHo6/O53KlFxQYczD3M673StbZeGX0q3qG6Ou8jJk2qQQHY23HqrqrJygaqqNWtU\nzdnzz6sb2rg4NZBIOEdLyrDWLPCGoGrG/mp7PQLoA4ytccx7gA/QEwhG1eItbuBcbb+Ac2NFpiL6\nL+lPaUUpPl4+WKwWNDTWPbSODkEdnH79U6fUSMXiYrj5Zhg/vkZ5WFGhErTgYNUfRLgFSdDciNUK\nzz0HW7eq1wYDPPMM/OUv+sYlqlitauLqe+5RNWc5ObBihbpndYHcsU1y9ES1B5vYp9H8UZz2lEg+\nwHXA/6GWlvoZ+AXVZ62WyZMnVz2Pi4sjLi6umeEIZ/H39mfKrVMwW6sn2jIajIT6hzr92jk5MGkS\nPP00xMbCm2+qPrlPPmk7wMcHOnd2ehzi/GzZsoUtW7boHYbTtOny69Ah+N//ICZGJWcVFaoGadgw\n1QdB6M5orNH1AwgNs3D44vH8lvs3x9YeejBHlGFNZXNdzvHe08281o3AZKqbOF9EjQh9u8YxE4AA\n23GglpRaD6ysc662fQcqWkzT4MwZuPhi9dpkUi0LMTH6xiXOj9SguZFffoG//726U5OmqabOH35Q\nNdfC5Ww+tZknv3mSAZcN4IM/f6B3OG2So6fZOH2OR3PtAi5HJX6+wAOoQQI1rQFuQg0oCEQ1gcrQ\nOGE3g6E6OQN1wy7JmRCt6He/UxMHZmVBebmaZuLaa9U24XIsVgsf7viQDkEd2J68nSOZR879JtEq\n7Jku4/eoEZfFQAWq1qugBdcyA0+jFls/DHyGGsH5hO0BagqO9ajJcHcAHyMJmhBCuI+wMJg9G7p3\nV7Vn//d/MHWqunsSLmdr4laSC5IJ8QvBy+DFrF1OWlFBNJs9vzG7gWHACtTIzr8A3agehamHtt1E\nIIQODh2Cnj3V31GrVU1X0qPHud/XGqSJUwjnuGf5PRzJOkKgTyAaGjmlOSy4awG3dL1F79DaFGet\nJACqk74XaqLa+VT3IxOiVTRnItmW0DSNf//0b87kn3HqdVyVxQILFsCcOer5++/D8uWybKAQbd2o\na0fx6s2v8o++/+CpG57Cx+jDskPL9A5LYF+CVgz4AfuBd4BnaTt3ssJNvLTpJVYerjtWxHHiU+JZ\nuH8hc3bPcdo1XJmXl5rU99gxuPtuNRr25ZelVUqItu6u393FQ1c9xENXPYTFaiHMP4y9qXs5lnVM\n79A8nj0J2l9sxz0NlAAXAvc5MyghajqWdYwNJzYwc+dMSitKHX5+TdOYHj+d8IBwNpzYQGJeosOv\n4Q4CAqqnhuvQQWZEEMKT5JXlsXj/YiIDI/H28pa+aC7AngTtNFAK5KOmv3gW+M15IQlR26xds/D1\n8qXQVMhXx+oO/D1/8SnxHM06SmRAJAaDgY/36L+YcWuzWlWzZnk5LFwIp0+r5k5p4hTCMyw/tJwi\nUxEWq4UA7wA2n94stWg6sydBuwO1eHkuUGh7tGQUpxDNdizrGNvObCMyMJJQv1Bm757t8Fq06fHT\nKTGXkFWShaZpfHXsK4+sRbvsMpg4Ua3R9/rrcOGFekckhKiwtM7i6pnFmVwacSkhfiGE+Ydxafil\nJOUntcq1RcOaWkmg0vvAPcAh1BQboq2yWODXX1U1SvfuLjFv0YJ9Cyg0FWIoVp2hCssL+XbPZ9xL\nd5VJXH7+C5T3ubBPrdmzjQZj5Ygbj2E0wp13Vr9u1w4GDdIvHiEE/Hj6R2btnsXiexbjbbTnz3XL\nTbx5olPPL5rPnr9CPwK3okZwugoZpu5oJhM8+yzEx6u/1hERao2kTp10DWtv6l5Si1KrNxw/Ts/3\nlnBxsY9KKEeMUOv8iTZNptkQnsZitTDk8yEczzrO+wPe5/ZLb9c7JHEenLVY+o3A68BmwGTbpgH/\nbc6FHEwKOEf74guYMgUuuEAN3cvMhD594AMXWvbDaoXbblMdo4KCVIKWmanmh+jZU+/ohBNJgiY8\nzeZTm/nn9/8k0DeQUL9QVj2wyum1aMJ5nDUP2htAEeAPBNkesqBaW5OSouZaqGzaa9cOEl2sH1ZZ\nGRQUVDe9enmpR0aGvnEJIUQLWDUrf1//dw5lHKq13WK18GH8hwT6BhLiF0JqUSqbT23WKUqhF3vS\n8RhA6lbbuiuvVDVUZrNKevLzIS5O76hqCwhQC22ePQtRUVBqGyxwySX6xiWEEC2wPWk73/32HaUV\npcwaPKuq7+uOlB0kZCcQ5BtEpjmTcnM5n+z5RJo5PYw91W3vAJtQa2i6CmkicDRNgxkzYNEi9fyG\nG+DttyHYxSpLExNh3DiVpPn4wKuvQv/+TrlUQXkBIX4hTjm3aB5p4hRtjVWzMmzlMM4WnqXcXM7c\nu+ZydYerATUn2YH0A7WOD/YNpldMLz1CFQ7grD5oRUAgqv9Z5XhfDdDzL5cUcM5SUqIGDISGuu40\n8lYr5OWppk4nzaZ6LOsYY9aO4dN7PyUmOMYp1xD2kwRNtDXbzmxj/PrxdAjqQHZpNldFX8XswbM9\nbgS5p3BGHzQj0N/20x/V9ywYfZMz4UyBgRAW5rrJGVSPMnXiVPezds3iTP4ZFuxb4LRrZBZnsvb4\nWqedXwjhuqbHT6fAVMDZwrPkleax4tcV7Evbp3dYwoWcK0GzAjNaIxAhXEXl5LiXRVzG6qOrSS1M\nPfebWuDjPR/zyuZXSClIccr5hRCua8TVI5h882Se7/s8F4ZcSIB3QL3BAo5k1az866d/yeSzbsSe\nUZwbgSG0neYFIZo0a9csvL288fHyQUNzSi1aamEqa46uwYiReXvnOfz8QgjXNviKwTx41YNcGX0l\neWV5dIvqxrJDy5y2csCO5B0s2r/II5eyc1f2JGhPAitQfdBkqSfRtOJiNRLUTSUXJLMtaRuappFd\nkg3AN8e/Ib8s36HXmb9vPhoaHYI68PXxr6UWTQgP9dGuj/D28ibYL5jMkkw2nNjg8GtYNSvT4qcR\nERDB+t/Wcyb/zDnfY7G60tz0nsmeBC3IdpwP0gdNNCYnBx57DG6+Gfr1UxPfuqGYoBjm3zWfOXfM\nYdbgWXxy5yd8cucnBPs5bjRramEqXxz+gkCfQMot5ZSby6UWTQgPdCTzCFsTt1JhqSCtMI3SilI+\n2vURjh5EsiNZTdsRERABwCd7Pmny+DP5Z7hvxX3klOY4NA7RPPZOS3wX8EfU6M0fga+dFpFwT2++\nCfv3Q8eOahToW2+p1bevuUbvyJrFy+jFldFXntc5Ku88vYxeDe5PKUzh0ohLq44L8wsjryzvvK4p\nhHA/HYI6MOXWKbW2BfoEOnQkp6ZpTIufRom5hKySLADWHF3DY9c9RufQzg2+Z+6euRzMOMjSg0t5\nOvZph8Uimsee/wVvATcAn9qOHwbsAl50YlznIsPUXc0f/6hGgPr4qNdnz8KECXD//frGpYOp26di\nMpt4+Y8v19peXg5+fo2/Fo2TaTaEaBlN05ixc0atbhpGg5ERV4/gotCL6h1/Jv8MQ1YMIdQ/lGJT\nMd889E1VzZtouZaUYfbUoA0CrqV6sfQFwD70TdCEq4mJUctFhYeriW4NBjXbv4dJK0rji8NfYNWs\njLp2FJ1C1GLzmgYTJ8Ltt6vH7t0wZ46aG9hbltcTQjiJwWBoVi3Y3D1zMWDA39uf/LJ8qUXTkT19\n0DQgrMbrMNs2IapNmqSWiMrMhPR0uOUW1R/NwyzavwirZgWo1a/MYFALICxdqhZoeO89GD9ekjMh\nhOtIL0rnm4RvsGgWMosyMVvNLD24lCJTkd6heSR7qtseRDVzbrG9vhl4AVjupJjsIU0ELmBf2j7C\n/MPoEtZFbcjKgqNH1ULr11yjJpT1IGlFady9/G7CA8IxYCCrJIsvH/iyqhYN4OuvVc3Zn/4EY8fq\nGKybkSZO4Sq2Jm6lZ/ueRAZG6h2Kw5WZy/g56eeqm0wAb6M3f+j8B7yNcjd5Ppy11BPABah+aBoQ\nD6Q1KzLHkwJOZyaLiUFLB9EltAtz7pjjMsuTaJqmWyzT46czc+fMqvU7C8oLeLTXozzX9zlANWu+\n9x6MHg2LF8NDD6nmTnFukqAJV5BZnMnApQMZ0n0IE26aoHc4wo04qw9a5UmzbMdfYXtsbc6FRNuy\n7vg6cktzySvNY3/6fq7teK3eIVFSUcKYtWOYcusULgy5sNWvP/iKwVWLHVe6KER1wtU0+P57eOUV\n+N3voFs3mD1btQRLM6cQ7mHRgUVYrBZWHV3Fw9c+TMegjnqHJNowe7K5t4EHgMNUDxQAuMMpEdlH\n7kB1ZLKYGLx0MGarmTJzGT3a9+DjOz7WvRZt6cGlvLr5VUZePZJJcZN0jUU4ltSgCb1lFmdy57I7\nCfUPJac0h/u63+ewWrRZu2Zx88U30719d4ecT7geZyyWDnAP0A0YiErKKh/CQ607vo7M4kx8vHwI\n8g1id+pu9qfv1zWmkooS5uyeQ+fQzqxNWCvrzQkhHGrRgUWUWcrQ0Aj2C2blkZWkFZ1/b5+TuSeZ\nuXMm7/3ynsMnqBXuzZ4E7QTg6+xAhPs4ln2M6HbRoIEBA9GB0RzLOqZrTKuPrqa4oph2vu0wYJCZ\n+YVoi0wmsOizBNHpvNNEB6pyz9vgTVRgFL/l/Hbe552zew7+3v7sTd3LgfQDDohUtBX2VLetAq4B\nNgHltm0a8IyzgrKDNBGIKiaLiT8t/hNZJVn4e/tj0Sxomsa64eu4IPgCvcMTDiBNnB6utBReew02\nblTT+YwZAw8/rOavcWMnc0/ywMoHiG4XTU5pDldFX8XswbN17y4iHM9ZgwS+sj0qSxQDMg+acCFG\ng5Fnf/8sFZaKqm0Gg4FgX8etnymE0NG0aWqUTceOYDar15dcolYwcWNzds+hqLyoagqL/yX9jwPp\nB7imo3stkSecw54EbYGzgxDifHgbvbmz2516hyGEcJYdOyAsTM2t6Ourfu7Z49gE7dQp+PRTKCmB\nP/8Z+vVz3LkbcXnE5QztObTWtsbW8BWep6kEbS0qOVsLlNTZF4gaKPAwavCAEC7laNZRlhxYwhu3\nvCHNBUK4u44d1VJygYFqzhqLRW1zlKQkGDVKJWfe3rBhA0yZAv37O+4aDXj0ukeden7h3poaJPAI\ncBVqYfSDwAbge9vz3UB3VILWHAOAo0AC0NT45BsAM3BvM88vBADTdkzji8NfsOvsLr1DEUKcr3/8\nA4KCICNDLSV31VVw112OO/9330FRkUr6oqLUaigLFjju/EK0QFM1aBnAq7ZHR+Bi2/ZEWraSgBcw\nHbgNSAF2ovq2HWnguLeB9bSdTsGiFR3KOER8SjzhAeFMi5/GwrsXSi2aEO6sa1dYsQIOHFBNnL17\nq5+OYrXWfm0w6DZa1J3llObwyg+v8PZtbxPsJ32Az5e9iyWmATtsj5ZO/BIL/AacBipQa3k2dAs0\nFlgJZLbwOsLDfbTzI7yN3kQERHAk64jUognRFkREQFwc9O3r2OQM1JprAQGqhi43FwoLYfhwx17D\nAyw7uIwNJzbwxZEv9A6lTWjN1aw7ATVnD022bat7zF3AR7bXMlpUNMvhzMNsPr2ZCmsF6cXpFJYX\nMj1+ut5hCSFcWdeu8MknKgG89lrV/+xOGXjUHDmlOXx68FMuDLmQeXvnUVheqHdIbq81VwG0J9l6\nH3jBdqyBJpo4J0+eXPU8Li6OuLi484uurTl2TC32mJcHAwbAkCFq5FMbF+QbxNjYsbW2hfqH6hSN\naKktW7awZcsWvcNwGim/XNDvfgdvv91ql8sry6OwvJCLQi9qtWs607KDy6iwVhAZGEl6UTpfHPmC\nUdeO0jss3TiiDGvNjjk3ApNRAwUAXgSsqP5mlU7WiCkKNXr0r6i+ajXJRI9NSUqChx6Cigrw81Od\nX59+Gh55RO/IhGgRmahWtDWvbHqFI1lH+Pz+zzEa3PvmOa8sjz8t/hNl5jJ8vHwwWUyE+IWwYcQG\n2vm20zs8l+DoiWoPNrFPA65uzoVQo0EvB7oAZ1ELsD9Y55hLajyfD3xN/eRMnMtPP0FxMXSytSD7\n+KgOtpKgCSGE7k7nnWbDyQ1YNStbE7cS1yVO75DOi7fRm7GxYzFbzVXbfIw+bp946q2pBM3RC6Kb\ngaeB71AjNeeiRnA+Yds/28HX81w+PrWXQLFa1TbRJKtmJb8sn/CAcCwWtaIMUOu5EEKcr092f4LB\nYKCdTzs+3PEhf7z4j26dzAT5BjHympF6h9HmNPU/4nSNRylqTrQrUc2Op1t4vW+BbsBlwL9t22bT\ncHL2CGodUNFct9yi5vJJTYXMTCgogL/+Ve+oXN7Kwyt5ZM0jbNlawb/+pVqIKyrgzTfhf/9r+XnP\n5J/hza1vYtWs5z5YCNGmVdaeRQZEEuwbTFJ+ElsTt7ZqDFIWuQd7Uvb7gXhgaJ3nwlVFRcGiRTBy\nJAwcCO+9B3c4ukK0mTZvhldfhfffV0PZXUxpRSmzds3iZO5JCqI34O0Nb7wBr7+uuvH16dPyc3+8\n+2OWHFjCjuQdjgtYCOGWfjj1A2armaySLDKKMzBrZtYlrGu162uaxrhvx7XqNUXL2NNh7QBqctnK\nv6rtgU00vw+aI0knW3eyapUatu7rq6qk2rdXa95FROgdWZUVv65g6vapBPkGEeAdwPK71jD8QdUs\n/Pnn4O/fsvMm5iUy9POheBu96RLWhSX3Lmn1pgyz1Vy1GLO7kkECoq2wWC2UmktrbfPz8sPHq3W6\noexL28fDqx8mJiiGrx78Cl8vB88pJxrUkjLMnr8UBmpPGpvd3IsID/fxx2qh46goiIlRNWhbW7dK\nvymVtWehfqEE+QaRUZzJk//ZwI03qpqzqVNVXtkSn+xRfU0iAiJIyElo9Vq0wvJChqwYwrGsY616\nXSFEw7yMXgT5BtV6tFZypmka0+On086nHdml2az/bX2rXFe0jD0J2npUx/5RqH5h61B9yYSwj9lc\nf7c1kf4AACAASURBVA62JpZRMVlMTg6oth8TfySnNIfiimKySrIoKLRw2PgpEybACy+o8RU7dzb/\nvGfyz7Dm2Bo0TSOrJIuSihKmxU+jNWtPVh5eyaGMQ8zeLWNwhPB0+9P3sz9tPxEBEQT7BjNz58xW\nL2+F/eypCTOgFi2/CTW9xk/Al84Myg7SRODKioth3jxISIAePUDTVC1acDCUl6tOXcuXq9q0OpIL\nknn868eZd9c8OgZ1bJVwSytKOZN/pta2dj7BXBh6AaAGwbZkjt+k/CSWHFxSKyEL9QvlqRueapW1\nQQvLCxm0dBC+Xr7kl+Wz5N4ldIvq5vTrOoM0cQpx/p5e9zQ/nPqhap3MwvJC/vun/zLg8gHneKc4\nXy0pw9y1wJMCzlWZzfDEE7Bvn1rbrrRUrZ3Xty9s2qSaOseMgUsvVQMH/vtfKCmB/v3h73/n9Z/f\nYuH+hTx+3eNMuGmC3p/Grc3fO5+Zu2bSMagjmcWZ9L2oL//t/1+9w2oRSdCEOH+7z+4ms6T2MtdX\nRV9Fp5C6qy4KR3NWgnYf8BbQocbxGhDSnAs5mBRwriohAUaMUAMBDAZVe5aRAatXwwUXVB936BCM\nHg3t2qnBA5mZJA+/g/tC1hPqH0pBeQGrh61utBYtITuBruFd3b7zu7OYLCb6L+5PXlkeft5+WDUr\nJouJ1cNW0yWsi97hNZskaG5E02DtWli3DoKC4NFHoZt71twK4SiOXkmg0jvAYNSkskK0TN0mvfh4\n1Q8tKEi9joxk3tFlaLFhKqEos7Jw38IGa9EyijMYtXoUL//xZQZePvC8Q9O02uHVfe2OvI3evHHr\nG7X6lxgwEN0uWseohEdYuRLeegsCA8Fkgp9/hiVL4OKL9Y5MCLdiT8+aNCQ5E/bq2hWuvlpNkpub\nq3727Qsd69SEhYSoTMgmy1LIN5HZqkN9cRZWzcqXR78ktzS33iUW7V9EblkuM+Jn1FpapKU++EC1\ntoKq7HvuOTW3rzszGozc1Pkmbu16a9Xjlq63EOgTqHdooq1btkz9foeFQXS06sLwww96RyWE22mq\nBu0+289dwGfAaqDydlxDZvkXDfH2VpPRLlwIx4+rQQIPP1y/SmrAAPjsMzh5EoBQP2/e/7+pWLtd\nUXWIl8GrqjNrpYziDFYeXknn0M5klGSw5uga7ul+z3nNLXbvvTBxokrONm5Uc/qG6NmAL4Q78/Kq\ndfOFpslaaUK0QFMNOQtQiVjlcXU7Tei58nbb7sPhKYqL1Z11aSn07g2XXHLOt7y7/V1W/LqCDkEd\nKCwv5FjWMd65/Z3zXgdu926YPBkuu0wtvCBci/RBcyNr18KkSWp+GrNZdWNYurTBUdtCeAoZxSna\nNKtmpf+S/uSW5uJl9KKgrIDE/ES6/3975x3fVL2/8SdtKVRKKRQoRUAU8IpeQVQU1xUcCG4cFxVx\n4BYU11UQL0uQC7/rBBVQFBQXoiguBJEKOBAVyhCqjMrsLt07+f3x5NykJWmTNOOkfd6vV17NSU7O\n+SQ9+eb5fta3XS+sHbnW5/BdZiYwfjzQty+wfj1w221czlSYBwm0MGPtWmD5crqib7wR6No11BYJ\nEVICJdC6AHgJ7IMGAGsAjAGw35sT+ZnGP8AJlxRXFKOiugLV1mrc+smtyCvLQ0V1BR456xHcdPJN\nPh3z//6PRWZXXgns3Qs88wwwc6bCnGZCAk0IEc4ESqB9A+AdAIvs28Ptt4u9OZGf0QDXxFm9ZzUe\nX/k4EmMTUVZVhiprFb4c/qVPXrSqKqbOudsWoUcCTQgRzgRqLc72AN4EUGm/LQCgWn0RUt7e/DYq\nrZXILc1FSWUJ8krz8O0e3yrFaosxiTMhhBChxhM19y0o0N61738DWCBwYQDtqg/NQINBaSmQmkrF\ncsIJplIu+/L3Ib88v8Zj3eK7ITY6NkQWiUAiD5oQjZ+KCtaWGEX/FRXsY94YCFSIsxuAWQD627d/\nAPAAgL3uXhAENMAFmsxM4O67gfR0LkbZty8bhrVoEWrLRC1yc5mPfcMNXDN02zYgIwO44IJQW+Y/\nJNCaMCUlQFkZ0KZN+HeQFnUyaxbb5918M3DgACvrZ8wAEhJCbVnDCdRKAmkArvDBHmEWVq8Gli1j\nZ+9bbvFs2ZUXXgAOHgQSE9nH6JdfgI8+AoYPD7y9witiYoCUFAq1AQPYxP2xx0JtlRANxGYDXn8d\neO01bv/971y7Nz4+tHaJgHHrrayo37+fwZtbbmkc4sxXPMlBewuA8zeiDYA3AmOO8DsrVgD/+heX\nVvrmG66Lt2tX/a/bs4frZAKctUZFAWlpATVV+EZMDGeaX38NjBtHcXbKKaG2SogG8sMPwJw5/IXu\n0AHYvJnuFG+wWulODvelQZoIcXHAfffxX2+xqN2RJwKtN4DDTtt5AE4NjDnC77z9NoVW27Yc5MrK\nuIhxffTtCxQWchZbXc3SxpNPDry9wid273bkaqxbx98lIUxPRQWbD65bd6SISk3l36go/lq3aQNs\n3Oj5sXNy6IK54grgoouAV1+tucKBMB3797PF0R13sL/xokVN+1/miUCzAGjrtN0WgNbtCBdq52zY\nbExUqo9Ro4D+/Tn7zMoC/vlP4PLLA2NjkPj0U2DLFt6vqgLmzmVYMNw5eBCYPp3LVS1eDOzbx8bt\nQpiKkhLgqaeAc84Bhgxh4uSddwIPPEC377BhvJgNjJUHjF/ooiLvGt5Om0aR16EDvXDz59M1I0zL\nV19RU199Nf9927c3jjHaVzxJWLsFwHgAi+37Xw9gGhj6DBVKsvWUlSsZ94qOpiqJjgbeesujZZVg\ns3HB86ioRtG1dcsWR36W4UR84glTFaf6hM3GmWenTgxznn8+C3CjooBNm5iXFu6oSKARMGECl4Ey\nPPl//cX4/LHHciKZmcmL1QhjVlXxC7p2LSeVrVtzVtWtm2fnGzSIf5s359+DB4EHH2SikzAlNltN\nn0Lt7XAmUEUCbwH4FcAF4HqcQwH87q1xws/YbMCqVfwFTkriit8xMUfud/HFrLz87DM+f/PNnokz\ngN+Mtm3r3y9MOPlkirMJExj1XbQo/MUZwH9Tly6MFv3wA7BjB5ereuop4OyzQ22dEHbWrAHat+eX\nLjYWKC/nIurGL/BRR3GmYRAVxWU+fv+dgu5vfwNatfL8fN26cVbWvLkj5t+pk9/ejvA/tcVYYxFn\nvlLX26/9y2zsa0z9Qul4bJozUGfmzgXmzeMAV1UF9OnDxxpL05gAUFXFybkR5hw/vv60uuRkoFcv\nFrNardS5gwc7JuVmo7ycIjQtjRGj4cMbxyAnD1oj4Oqr6ZGPi+MEMzWVf3v2pIcsPZ3xrTFj/HO+\nvXuBe+4BDh9mHu3FFwNTpnDMFCLI+NuD9hscYswVx3pzIuFHKiuZT9GhA2eZNhubX23cCJx5Zqit\nMy3z5/PvokXMbZgxA3j+eU7q3VFcDDz5JPMhFi8GDh1yRE7MSGkpnQ0AI0aehgjWrQOys/kbarUC\nb7wBnHeeZx1ZhPCIceOAhx6iELPZmIt26qn8Qlqt/GLdc4//zte1K7BkCbBzJ71zPXo0jtmKaDKE\n69XaNGegBqWl/PVMTHQk/GdlAf/9L3DuuXW/tgmTm8vJuxHWzMqqW5wZfP45nZPt2wMvv+w6kmwG\nSkvZUeXss4FrrwWefpqXyAMPHLlvTg6L5y69lNtr11KsjhhBEZqWxtYdR3m/tGlAkAetkbB7N9tl\ntGzJZMnoaE44q6vVBNtHrDYrIiye1PuJUBKoHDSAvc96AnD+Bq3x5kTCj8TEsE38qlVMnC0qcuR1\nNKasSj9TO53OE3FmtfI3BWD4sKDAvAKtRQsWxfXpw0vg3/923/IuIoLh2oIC9v+cO5dC7rnn+PwH\nH5hHnIlGxHHHHZkD26wZb8JrtmZuxdNrnsZbV7+F5lGhy7tYt47pgzfcwDFz3jzgtNOADz8E7rqL\nUeyUFODjj+lIlRb3DE9k912gGFsBYDKArwFMCqBNwhMmTwZuvJG/opmZjGvdfTfdHmqC5Tdef50e\npcWLgZtuYrjTXz0v09Md961W/hsbgsXCBrWGPm/eHDjxRNf7tmkDPPMM8M47jsjTyy/zMoqP50A6\nYQLwxx8Ns0kIEThe2fAKNh7aiC/++CKkdpx0EmtA3nuP40haGh8DgKuuAj75hAW5v/3W8HGuKeGJ\nq2UrgH4AfgRwCoATAEwHqzlDRdMNEdTmrruArVuBdu0cXbNnzAAuDOVa9o2HXbtY+GV4zbZto+hp\niJOyooLeuFGjqKlPP509NCsrgccf94/dnrB1K8UZwEKIn35iHtpDDzHnLiGBK4R17hw8m9yhEKfw\nicJCzjKOOooJlZ70gAwTtmRswchlI9EquhUiLZH4/KbPQ+pFy811dDBZvJhjps3GBgM//EAv2sKF\nQPfuITMxpPgyhnlytZYBKLXfbwFgBwClDpuFPXscPcoiIijSDhwIrU2B4MABxt2WLKGKCBLdu9cM\naZ50kmtxVlxcs6FiRgarRg0qKhwzx1dfZV70uHFMG+zVi+EBfxWveUJaGnvCTZ3KtniHDzPEOXQo\nQ6OtWrGxrxnEmRA+kZYGXHcdMHo0lcPYscx1ayS8+suriIqIQsvolsgvy/eLF835p8Nqrdk3uC6s\nVnrjjz6aqSOffsrHN29mf+LYWB5bwR3v8ESg7QNz0D4BsBLAMnABdV8YDAq8PwE84eL54QBSAGwG\n8D24zJSoi5NOYum6zUZFEBHR+KYou3YxvjhjBlvm33QT445+pqKi5vhtVEN6wk8/Mfw5fjxTAx9/\nnGvUP/UU/y0pKfx9OHQIuOwyYPZs5nulpgIdOzJZP5itOzp3ZseBPn0Y7pwxg6t7bd3K50tL2WJP\niLBl2jTmI7RrR9WwahVvjYAd2Tuwdu9aVFurkVmUibLqMsz7bR6sNt8VUGkpJ2crVlBIzZoFvPmm\nZ69dsYIC7PnngWefZbhz7VqOidHRbKB9zTXMynFO7RB1423IYACAOADLAVR4+dpIAKkALgJwAMAG\nADcC2O60z1lgE9x8UMxNAtDfxbEUIjDIymJ37N27KdJuvx24997GVSjw+OPAd98xOQqgq+qGG4BH\nH/XqMJ9/zvzkE0+kaHrrLQ4axmHffZeDzCOPsMpx/Hh6uWrnNKelARs2cHJusVCIxcbSmTlrFv+O\nGcNw6C23sJ4D4CA1ezbvX3gh8NprHLzefZe68+67WYTr727a6el8j0ZibloacMwxNY9ZXU3de/Ag\n80gWLeKM+LXXuOJXqFGIU3hNI15JoKC8AD/u+7HGYzHNYnBe1/OM74pPHDxIUZWTwx6REyZ4ltBf\nVcWbsW9xMaPKmzYxwNO9O8exb75hJ6hGsDCN1wSyitMg2cv9nTkDwE44vG/vA7gKNQWa8xW3HoAC\nLPXRvj1/TTMzGYsz1EYw2buXArF9e+8StPbvZ4+idu3cxw4BCrJ9+xyLJ8fGcgTxkrZtOakeO5ZL\nPRUU1KxUvO46Pj9unKMnmKtFF9q0oV4sL6dLf+FCeqMSE7k2fVISJ+qXXOIQZ1aro6dYaSlnmOef\nz0Hrhx9Y2/HLL3xbU6cCEyfyX7lyJQXf3XczVNCihSONpriY3Qrq46uvmIYzcSI9ebNnc5bboYNj\nn8hIxzk7dWJC78CBQG/5sEW40qcPO00nJlI9WCxMhGoExDWPwyU9LvH7cTt25MeVk8OJmafVllFR\nNVdlMcalvn0dj1ks7BUcLpSWUtt7O976k2BmTB4NhksN9tsfc8cdAL4MqEWNhchIqgJ34uyPP+hx\nOuss4I47/BseXLWKC6mPHcv1hV56ybPXJScD11/P191+OxWDO6+CzcYkKWMUyM31yaW0aRMF1bhx\nFEh79wL5+Y7no6MphLZvp2Pysss4UM2d68gn27SJr502jSlxzz1HcQYwPDBiBL/E2dnML1u5kuJs\n6lS+3TvvpBY1wgnPPAO88gpXN/jnP9lr+Pff6b1bsoTetcsu4/HfeAOYM8fR+uP++z1Lx7v1Vg66\nQ4bQpgkTKDL37Km534knOlbCsVg40zVrSxEh6mXcOH7ZsrI4ZtxzD8dA4RIjrBkZyWF86VKGLpsq\nixbxc7Ba+VsxalRAMmvqJJgrEXrj0x8IYCSAc9ztMGnSpP/dHzBgAAY0hhWhA0FBAX/JS0v5q7x1\nK93877/f8CVPKivpdomN5S95dTXjYoMH192CvrqaKuGoo3irrqbaufRS130hjMUm8/Ic970NEdls\nGDHCgiuuoOOuqoqiybkXWmYmPVm33cbk1ueeY+J8ZiaXBBw0iDkWTz7JcnFjEYc1a/g7cOut9FZN\nmODIxxg5kp64NWuAf/yDJedXXklXf24uX/fxx/wYN2ygbfPnU7wtXAjMnOkIB4wcSfseeoivvffe\nmnlrJSUUmbXXF42I4Ee3ZQsdCHFxnMn2789CAQCoqK7Artxd6NW+l3efa4BITk5GcnJyqM0IGBq/\nahGI/o1t23JWk5dHV1Cw3R9hRmUlx4a777YhJsaCadPY36ypMmIEm30/9hg1/p130g/iKf4Yw4KZ\n09EfzCkbbN8eB8AKYEat/XoD+Ni+3043x1IOh6ds2gTcdx/DiAaZmeyfkJjYsGPn5TGO53yc7Gx6\nw+papbuwkLG/jh1rvm7mTK6QUJvp04GPPnLsf+gQ1cldd9Vv4549nEnv3IndtmPxVq/p2JDXA9XV\nbCPxwgsOkfbJJxQzV17JgoFnn2VeVqdOdEBWVNBzFhFBT9SUKewTPH48cPnl1KUVFY7lUHfvZhiz\nooKeucmTj6zy37CBYu7wYXrMunTh6jfff0/xlpbGHp7nnEMP3KJFFIjnnkuh9uabtKl5c4q8Sy45\nMoywfj3FptXKMGpeHo+3dKnD1sXbFuPF9S9i2Q3LkHBUQv2fa5BRDlojJSODF/TmzRxHpk5lMz/h\nEqvVMYZYrdS0/tS1m5M/wKwfnsechNsQec212F3YHp06OUKd27cDJ5zQuFKc6+LPP5mTHBXFn6CG\ndGkJVJsNf/ELuBpBNwDRAIaBFaHOdAXF2c1wL86EN7RqRQ+VUd9cWcm//phNtm7NckAjzlZUVLOK\ntLiYIdDly2vG4mJjmaWenc2Zc3Exv/Huqk/vu4+un6ws3vr2BW6+uX77jGZje/cCiYloX3UA16wa\nhcfuL0GfPnTWJThpkauvpjgDKFzGjaOZ27Y5Fmr44guaMnMml/pr3ZoCydCVhuCxWin4WrZkHtuG\nDcBffx1p4rHHMhF/4UL+Ln37Ld3qp57KGoi+fSmqPv2UHrSJE3m+zp35+PnnAw8/TE9fjx7ARRcd\neY6iIv7uPfccP2abjeFUw9bSylLM/WUuCsoL8M6Wd+r/XIXwBzYbf/22bGFCZGGhw2UtjmDHDmrZ\n0lKOL3Pncp7tDpuNhVElJdwuLeX+7lpd2JKT8eLbo7E2dxO++/g5YMQIrF2ajSlTWNH+5ZecmBYW\n+v+95eXVvR0K9u7luPnAA/ytmDatZhFwSgr/J4EkmAKtCsBocCWC3wF8ABYI3GO/AcAEsKXHqwA2\nAvg5iPY1To47js2tMjJYzpeTwysuNta34333Ha/UV1+l2+eFF5jYdegQt//+d8bs9uxx9B6aMIHx\nwcceY8JVTg5f16ULbbJaGUM0EqBqEx9PBTN/Pl1Gc+Z4lhx14ABjgQkJQEQE8iMT0C2hAAN77MOk\nSdSSzr3LXJGezkFp4kRWfVZV0YvlnGDfuvWRevfAAQq0e+9lHhngumS9bVuKsagofuGXLuW/KzWV\nbzMlha71YcOYnzZ4MF3vkybx4z77bL6HrCyKLlcz2wsvpL033UQRd801/DiXLuXzn/3xGQrKC3B0\nq6Px3tb3kFPifQGGEF5jNJHt0IFfxrg4fsH+/DPUlgWV2q3Z3LVqO/54TgonTODwuXt3/Un3+/Zx\n7MrNpce/dt6pM5vemIbN8WVItB2FWb0KUJ2ZgRGJXyMxkd77d97h0O/vCkybjW1+3nuP26tXs3Df\n8CWEih9/5KR40CCmnJSVcWWZr77iuDxzZs1el4EgXB2VChF4g83Gqy0jg4KtTx/fjvPRRww3RkZy\nFOnYkd/auDjmtP33v1QaVVX8dlksHFEOHaJvPC6OYi4xkSonPp7TOufSRH+Snc0M+zZtGCesqqI4\n/PRTr5IJsrMdEeLKSjr86iuWLSpiW42hQ/nW0tJ4q51qZLMBCxYwJ81qpcgaOpQD6/33c5+xYzlb\nHjOG3rYLL+SAWVzMwbdHDwq91as5gDqvOVpQwFlfQQHzSR55hHZ8/DGLAIbfWorL370cFosFMc1i\nkF6UjhG9R+DBMx/0+PMJBgpxNkKqquh6jo3lGGCsd/bmm5zoNQEKCylGxo3jUPnll0wTdreiiNXK\nPFaA40ZCPdkIVivw4ov0zJ9zDo/raqi12Wy484FjsD06Hwm2FkhvVo4Zm9rjgmFj8XniHZg7l6kg\nr7wSmHU0Dx+md7C6mkLo6af5eZiNQ4eYrgLwp9Cby9TsIU4RKiwWulqGDvVdnAF06cTHc8ablET3\n0tq1PP6CBRRCHTrQE5aTQ/EF0FfcrJmj2jQ9nVnzFguLBAK1/Eq7dqzcysmhOM3O5pTIm0xP1Ezf\na9bMs04msbHAtdc63lq3bkeKM4ADZ0oKP9pZszg7e/ppOjkvuYR69sEH+TZOO41Vn3v2MK/tr7+4\nCsE991DYXXCBoxOJQUQE/0Xl5Wyd8cwztH/OHHrmUjJSUFBRgPzyfKQXpcNqs2LFriZcuiWCR1QU\nEzjz8/n9zMxkMqexiGMToFUrfncnTOAC40uWsHeiK4yw5gknMLVhxgzHEOuO8nJ+tACHP3fNt7dk\nbsFPSdWosFbiUEQJim0VmN09B8nWf2DpUtrWpw/+F+70N/Hx9FQdPMh/vxnFGVAz+r5vn/v9/EW4\nzkg1Aw0FAwcycclIXjp0iK3yr76a6iAqylFWuGMHp0M9ezJZqryc4q1NGyZFTJ1KBVMfVVUMq+bk\nUI2cfLL3dm/eTJHYubPpEpArK/nRGBHnw4epdT/6iLPq775jwcCiRRR5vlBUBAwfzgH+ySdrdhqw\n2WwoqSypsX+zyGaIjoz27WQBQh60RkxqqqMfYr9+5l8v02bjlyo62m/Lfzz+OIMMEybwI3DF9u0c\nGyZOpBdrzhxO4AyPWm2sVobmOnTghG/uXE7u/vOfIz/irOIsrPtrDSfOv/4GtGiB2CuuRVSze9Gr\nF4MlVitz2IYMOfJtp6c7arhsNgoZb2rQVq/me3v0Ub6v887jqgNmIjWVAnXcOHoux49nusnAgZ69\n3pcxLFwHPHMMcKtWsVFMq1b8Tx1zTKgtahi7d1MJFBez5cX559d8fvZslq3HxXEa1bw5Q5tJScxJ\ne+21ms9dey1Xzd23j9M3Y63QiAjg5ZfdTxUNqqv5jV23jt/6iAiOOEYmvw9UVtKRZwxQzlWXoaD2\n+SsqqHPT0oAnRhXhgn0LcPdFexB5yslUWc2aeX2OXbuAf/2L7/2qq9gKL9yqsCTQhCkoLKQresMG\nfonuvZd9HBvwhfryS3rOBg1iPdWUKe49SLWrOIG69ezvv9PjZgy9O3a47mTUEMrLmY5xww0sUpo7\nl3P3yZM9e73Nxtdceinf9+HDjHKPHu3TcBcwysrYBqlHD26np9O++sLMBhJoweTTTxmLio6ml6dl\nS+ZjuUt0DzUVFQwltGlzZKMsgB6mESPoM4+K4v5TpzIr3aC6mgLu22+Z6DRqlONqNXqgrVrF5+6/\nn94zm40uoFtucaiRpCSOGEaY0x0bNvA4iYncr7ycM9d163yeZS9a5KiTyM11LOfkq3eqIVRU0I5R\no9ixf8UK9u+dOBH4z9OVGPLRnWiXuQ1tkpqjTfMyxjynTav3x2D/fsci54cOcaB77DHH0i39+plv\ndlofEmjCFEyYwFLupCSO+1lZTPI691yfDldYyDnn2LH0QK1ezR6LXq5iF3IOHOBYWlTEsXTyZLWd\nq00wlnoSBgsW0HNmxKYOHKA4GTEipGa55PvvGdsqK6NAe/55hgudWbGC3y5DYBYW8j06C7TISAqt\nfv04zXEu5zGeq+0Vs1h43vbtHaWPNhunH9XVrsWigdG2wxAk0dFUNZWVPocWrruOM9SJE2nCpZeG\nRpwBfDujRzPk0Lcv23lMm0anZJeSVPRrtQNlR3dEaqoFLXpaEbNyJZWWcxVALaqqeIyLL2bK4bJl\njBz178+PccqUmqsnCCG84Ndf+f2zWOg+sVj4xfVRoLVqxeHYGOIGDnSdq2p2OnWi92vjRoYnJc78\ng8mD/SbGVedrdw1mQkl2NhMcIiMpkIqLWQ5Yu4bZaj2yO7+r7enT2Wr/0UfZr+G33+q34aSTOJgd\nPszzpqczEaoucQbQFx8dXfN1/fo1KO+jRQt6rDZt4uHc5W8Eg7w8R2Pa775jGl+HDkwavvUWGywW\nIKYF8PeTPV9yKSqKjt3lyxkJTk119D8DOJ84uq4F1oQQ7uncmRNHgOOhzdbgqEntn5FwSz8wQpQl\nJRxrPvmES9yJhiOB5is330xXRF4eMyJbtWL/A7Oxdy/FlzGliY+n3bUXcRw0iBWVmZl8T0VFRzaD\n/fVXNs9q355umchI+rUBnuODD9jdf+zYmg13kpJYn52YyBDqwIEMn9ZHYiJz25KS+LoBAygQG0B2\nNt3vw4cz5DdrVuh0tRH5/fVXCsUJE1jFGRMDRJ74NzbuTU9HVOFhqskBA+gBrYeEBMci7337ajYr\nhN8YN46znKwsjpVnnVUzytAEqaxkoGPyZGa1TJ3qqBwVDSPMtPr/CH0Oh83GX9Plyxnqu/12953w\nQ8n+/UzWN3qBlZYy5Lhy5ZFumdRUhjVLSljuftFFNadzX33F2KBRnmOEKtevZxPZOXM4eJWXU+y9\n917N5ZxMwJIl1JVDh/JjmDGD/7pQlHVXVNBbVlxMz97NN1PXPvGE/WPPz2fhxZ49rHG/7bZ6xFB9\n4QAAF2dJREFUKxpsNjZT3L6djtJnnmHq2jXXBOUtBQzloAmvKS3lF3zFCo5L//pX/Z1dPSE/n9n3\nMTGc5TV0TWPRJFCRQFOirIxJ9BUVdJPUkZeERYvoKjIS66dNYzzNW/78kyoiLo6KIjOTJUILF9J7\nGBnpCD8ePMgWHEOHen+e0lIKvp9/ZkjhgQfq7V1ms9mwOWMzeif2Nr4IYcGOHfzdAIC3367ZY62w\nkI5Zd9uuMFY6uP56es6yszmHGD48/EInzkigCa+ZPh348ENOKMvL+QV64w3fWvUI0UAk0JoKxcUM\nJe60L1faujU9LXVlu//1F/3OXbt67tWqqGC16p49FGKXX85f+6lT6dfu3p0ZrklJDJFarQ6v3KFD\nLE/yJsnrzz9ZzLBuHQXo8cfTJdS+PTPn61AnKekpuOuzuzDvink4paO5ep25Y8cOfpQPP0zn5fff\nUzvHx/Pjvf9+rpZ17rn0/G3YwIKCcBZaviKBJrzmkks4fjhPGseMqb+9jxABQFWcTYWPPuKvu5Ht\nnZVFofTii+5fc8wx3vVps1pZXLBmjWP5pi1bmHM2aBBFYlwc96uuZnOtmTP5eFUVRZWxgrgnlJSw\npLGggPcjIigMTz+dbqDNm7lWiQtsNhte3vAyCsoLMPvn2XjtitfCwotWWEhxdtppvLVs6ejS3awZ\nterEifx3l5TQIRAGb0sIc9CmDavrmzd3FDy1bh3QU1ptVkRYlNot/IOupHAkI6NmBWRMDD1W/mTX\nLq7f2akTQwQdO9KblpPDc8fGsmTnrLN4O3CACuKii9ixcMGCusOutdm3j+LMuYS9spKKxWars+Jz\nc8ZmbDy0Ed3bdkdKRgpSMlIa/v6DQL9+FGYGV11V07l57LHsj7ZzJ5cU9ebjFKLJM3YsJ4/p6byd\neCInlwHCarPi/i/ux7q96wJ2DtG0kActHDntNIb8jLbzBQVcbsmfVFRQKBkuGyN/zWjP8eGHbExr\nFAy88w49bk8/7dv5WrfmYGq1Mu/MqD7NyWHOiJslmgzvWVRkFCIsEWgW0cz8XrTSUoZxS0qYP+im\nQmHJEkZ9J02ic7RtW5/bLQnR9DjlFI6TmzZxEnveeX5f6dtqYwl4hCUC6/evx7q965BZnImzu5wt\nT5poMLqCwpGBA5k4X1DgWGD43nv9e44ePRhCzchg2DI9ne4co9nsjz+yUjMqireYGD7mKx07Mkya\nnc1jJSbSG/fgg2yy46b3WVZJFn7P+h2V1ZVIL0pHRXUFtmdvR2Zxpsv9Q05JCVcpHzeOCWg33sgf\nkFpUVnLlrenTqccnT2YTSKUuCeEFXbuyIeDFF/tdnAHA8z89jxd/ehFWmxWzfp6F+Bbx2Je/D2v+\nWuP3c4mmh0ldDPWiJFuAHiartf6Gr76Snc0w5s6dFGdjxjgS9adPZ3JUu3Y8f0YG2/Q/8YTj9StX\nAvPmsYLquutYAVrfEk2bNrE1SNeuPKcHlFWV/W8mCwAWWBDTzMPOrsHmk0/oZTSaWx4+zPf6zjuh\ntcvkqEhAmI2Mogxc9T6LoP79j39jUvIkJMYmorCiEAkxCVjyzyXyoon/oSpOETxWrQKGDXOs3Xnq\nqQx7tmvH5zdsoFfPCJNWVtJrNGxYaO0ONQsWsOWJIdDKytieZPnykJpldiTQhNmY+f1MLPl9CQCm\nWhRXFiM2mkv/FVUUYd4V83BuV+UkCKIqThEcDh9mzK1XL1ZsFhdThDm3wVi+nHlklZV8LjKSHrem\nLtBOPZVexJISNp3NzWXTMiFE2JBRlIGPt3+MhKMSAAB78/di/HnjEd/C0ciwR9seoTJPNBIk0IT3\n7NtH4WV4y9q1Y6sPo88awLBoUZGjyMBmY2VoU6d3b+aePfsscwivvBJ46KFQWyVE48NmA774Avj6\naxYh3XEHS6P9wOJti5Fflg8b6Am1Wq3IK8vD8N7D/XJ8IQAJNAFwIMvOpphKSKi/2Va7dsx9q6xk\nO4yyMr7GeZ3I5s1ZCQrwOZuN+wo20LzkklBbIUTjZvFi9maMieFYtW4d8O679S9uXlrKCEB2Nqus\nTz/9iF0G9xiME9qdUOOxbvHd/Gi8EBJooqyM/YJ++IEi6sILgSlT6l7zMSmJ1ZUvvsjQpc3GVQOc\nQ5yGeDOeB9hGQwghgoGxdtpRR3H7wAEgORm46Sb3rykvZ+7s1q3ctlhcrojSM6Eneib0DIzdQtiR\nQGvqLFgArF1L0WWzsfLyhBO4MHddDB/OBrWHDgFduhzZy6tnTw5uhnctMrLupaiEEMKfGJ575+36\nqsh//JELoSclOcav555jKoJZ+yqKRotqgJs6KSmcYRqDV4sWjtljfRx3HJdfctVoNTKSt5YtuepA\nVJQ5Q5xWq5qLCdEYuf12ID+fza4zMpiHNmBA3a8xJpSGGGvWjCFPjREiBMiD1tTp3p0tMYw16srK\nKLwaQlUVG+h26eJYHSAurv7ZazCxWoHZs4H33uP2iBEMbZjJRiGE7wwdyrSLb77h+HPLLTXXUnPF\nKadwkpqby4lrbi4wZIjGBRESwtVnqz5C/qKgALj/fq4pBAAnncQ+XS1b+na8/Hzmp/30ExvOdujA\nkGlmJsOmDzzgN9MbxAcfMIG4fXtuZ2UBTz3FQV2YDvVBE0Fj+3ZgxgyOCeeeyyrrGB8aX5eUcBL4\nyy+crD7yCFdnEU0SNaoVvlFZSYFmsTB3rCErExgrDCQmAmlpXK8oIYEeqkmT3C7ZFHRGj2Z4N97e\ntyg3F+jfn+0v6iI9ndVdnTs7XisCjgSaCDsefhhYs4bRieJiLqa7eHHNYirRZPBlDJPfVjDP4sQT\nmUv2zTfAp5+y4skXtm9nzhnA3I2ICOZvpKVxkDILHTuyYsugvNyx8Ls73n+f1Vx33cWk4V9/DayN\nQojwpLiYbT2SkhiN6NAByMsDtm0LtWUijFAOWlMiIwM4eJCDRu1cjMJCYORIYM8ebsfEcB3NXr28\nO0evXqyCys+npykqioPTzp3ASy/RixYKrFZg2TLg55/53q+/Hvj+e3rELBY+NnKk+9enpQHPP0+v\nWXQ0P6/HHwdWrGAxhBBCGERFcVypruZ9m41jkBkLpYRpkUBrKixfzuWZLBYOFP/+N3DZZY7nP/uM\nnf6NHImcHIb7Xn/du/OMGgWkptITZ7XSK9WpE/MxjDy3UPDyy8CbbzIBuKKCoYf58x0z2jPPZCKx\nOw4epDfQ6A/XqhUFb0FBzQa9QgjRvDlzbufP5wSuuppNb3v3DrVlIoyQQGsKHD7M5rOxsRQoZWVc\nbuiss5gXAVCQOVcqxcTwMW+Ji6OomzcPePVVJsdaLBQygwb55/14S3U1m1YmJjry6/bupSC9+GLP\njtGlC2fBZWX8DA8fpjAzql+FEMKZ++4DevQANm/mxHfoUHnQhFcoB60pkJ1NcdGiBbeNv1lZjn36\n9+ff0lIWDeTlARdc4Nv5oqKAu+8Grr6a58jM5Oxx9Gjf30NDMBKyazeatFo9P0aXLvQ6FhXxPbVo\nwZCnyu+FEK6wWDgpfewx4MYbHeOuEB4SrlVRqoLyhqIihjMtFobmioroVfrii5oVRZ99xjyx0lLg\niitYhVTXkk/1YbNRnFVVMe8tlGLmmWdYXdqyJb1g7duzB1pdYU1XFBay4jMxUQNuEFEVpxAinAmH\nNhuDAbwAIBLA6wBmuNjnJQBDAJQAuA3ARhf7aIDzll9/5UyutJTCYuZM4IwzQm1V8KiqAt56i/3Z\nOnViU9r6mlYK0yCBJoQIZ8wu0CIBpAK4CMABABsA3Ahgu9M+lwIYbf97JoAXAfR3cazGOcDZbOzN\nlZcHHH+8/5salpczr6xtW3l/RFghgSaECGd8GcOCWSRwBoCdANLs2+8DuAo1BdqVABba768HEA8g\nEUBGcEwMITYbE/m/+IKhwIgI4L//Bc4+23/naN6c3iMhhBBCmJpgJgUdDWCf0/Z++2P17dM5wHaZ\ng99+Az7/nLlR7dtTTD31lBbpFUIIIZogwfSgeao0arsAXb5uklPD0wEDBmDAgAE+GWUacnMdnjOA\nyezp6ayobEiivhBhSHJyMpKTk0NtRsBodOOXEKIG/hjDgpnT0R/AJLBQAADGAbCiZqHAHADJYPgT\nAHYAOB9HhjgbXw7HX38Bw4ZRmMXEsPqxVy9g4cL6XytEI0c5aEKIcMbsa3H+AqAngG4AogEMA7Cs\n1j7LANxiv98fwGE0hfwzADjmGOA//2G1YXo68Le/sdJSCCGEEE2OYM9Ih8DRZmM+gOkA7rE/N9f+\ndzboZSsGcDuA31wcp/HOQK1WVlvGxITaEiFMgzxoQohwxuxtNvyJBjghmhASaEKIcMbsIU4hhBBC\nCOEBEmhCCCGEECZDAk0IIYQQwmRIoAkhhBBCmAwJNCGEEEIIkyGBJoQQQghhMiTQhBBCCCFMhgSa\nEEIIIYTJkEATQgghhDAZEmhCCCGEECZDAk0IIYQQwmRIoAkhhBBCmAwJNCGEEEIIkyGBJoQQQghh\nMiTQhBBCCCFMhgSaEEIIIYTJkEATQgghhDAZEmhCCCGEECZDAk0IIYQQwmRIoAkhhBBCmAwJNCGE\nEEIIkyGBJoQQQghhMiTQhBBCCCFMhgSaEEIIIYTJkEATQgghhDAZEmhCCCGEECZDAk0IIYQQwmRI\noAkhhBBCmAwJNCGEEEIIkyGBJoQQQghhMiTQhBBCCCFMRrAEWlsAKwH8AWAFgHgX+3QBsBrANgBb\nATwYJNsaTHJycqhNqIHsqR+z2SR7RKgw2//abPYA5rNJ9tSN2ezxlWAJtLGgQDsewCr7dm0qATwM\n4CQA/QGMAtArSPY1CLNdDLKnfsxmk+wRocJs/2uz2QOYzybZUzdms8dXgiXQrgSw0H5/IYCrXeyT\nDmCT/X4RgO0AOgXeNCGEEEIIcxEsgZYIIMN+P8O+XRfdAPQFsD6ANgkhhBBCmBKLH4+1EkBHF4+P\nB71mbZweywXz0lwRCyAZwFQAn7jZZyeA7j5ZKYQIR3YB6BFqI/yExi8hmh6mHcN2wCHekuzbrmgG\n4GsADwXDKCGEEEIIMxIZpPN0BQsEvgcwGkAagG9q7WMB8CaAvQAmB8kuIYQQQogmS1tQkNVus9EJ\nwBf2++cCsIKFAhvtt8HBNVMIIYQQQgghhBBCiDDETE1uB4P5c38CeMLNPi/Zn08BK1EDSX32DLfb\nsRkML/cOsT0G/QBUAbjGBPYMAL21W8HilFDa0w7ActCLvBXAbQG25w2wqnpLHfsE83quz55gX8/+\nwixjmMavhtljEKzxy1ObBqBpjmFmG788sSlcxzC3zATwuP3+EwD+42KfjgBOsd+PBZAK/ze5jQSr\nr7qBxQybXJzjUgBf2u+fCeAnP9vgrT1nAWhtvz/YBPYY+30L4HMA14bYnnjwB7GzfbtdiO2ZBGC6\nky05AKICaNN54KDlbjAJ5vXsiT3BvJ79iRnGMI1fDbfH2C8Y45enNjXlMcxs45cnNnl1TYfDWpxm\naXJ7BnhxpoGrHrwP4Ko6bF0Pfnnq6/kWSHt+BJDvZE9nBA5P7AGABwAsAZAVQFs8tecmAB8B2G/f\nzg6xPYcAxNnvx4GDW1UAbVoLIK+O54N5PXtiTzCvZ39ihjFM41fD7QGCN355alNTHsPMNn55YpNX\n13Q4CDSzNLk9GsA+p+399sfq2ydQg4on9jhzBxyziVDZczT4hX7Vvm0LsT09wfDTagC/ABgRYnte\nA5c6Owi6wccE0B5PCOb17C2Bvp79iRnGMI1fDbcnmOOXpzZpDHOPmccvwINrOpDhE2+oq8mtMzbU\n/aWIBWc3Y8BZqD/x9MtYu/lvoL7E3hx3IICRAM4JkC2AZ/a8AK7DagM/J382SvbFnmYATgVwIYCj\nwNnNT2DOQijseRL0ogwAG5muBNAHQGEA7PGUYF3P3hCM69lbzD6GafyqG7ONX4DGMH9gxvEL8PCa\nNotAu7iO5zLAgS8dbHKb6Wa/ZqCrdxHcr0DQEA6AibwGXeBwK7vbp7P9sUDgiT0AkxBfA+Pddble\ng2HPaaBbHGB+whDQVb4sRPbsA0MCpfbbGnAwCcTg5ok9ZwOYZr+/C8AeAH8DZ8ahIJjXs6cE63r2\nFrOPYRq/Gm5PMMcvT23SGOYeM45fgHnHMJ+YCUe1yFi4TrC1AHgLwPMBtCMKvOC6AYhG/Um2/RHY\npERP7OkK5gz0D6Ad3tjjzJsIbBWUJ/acAPbniwRnn1sAnBhCe54DMNF+PxEc/NwtieYvusGzJNtA\nX8+e2BPM69mfmGEM0/jVcHucCfT45alNTX0M6wZzjV9A4xzD3GKmJrdDwOqqnQDG2R+7x34zmG1/\nPgV0PQeS+ux5HUzSND6Tn0NsjzPBGOA8secxsApqCwLXnsVTe9oB+Ay8draACcCB5D0wV6QCnImP\nRGiv5/rsCfb17C/MMoZp/GqYPc4EY/zy1KamOoaZbfzyxKZwHcOEEEIIIYQQQgghhBBCCCGEEEII\nIYQQQgghhBBCCCGEEEIIIYQQQgghhBBCCCGEEEIIIYQQQgghhBBCCCGEEEIIIYQQQgghhBBCCCGE\nEEIIIYQQQgghREgYAOAz+/0rADwRoPOcBuBFN8+lAWjr5rlvALQKhEE+kAaHnd/74Xi3AZhlv/8g\ngBF+OKYQQgghGgED4BBooWIPXAu0CwC87KdzRPrhGO7s9JXb4BBorQD87MdjCyFCTESoDRBChJRu\nAHYAeBNAKoB3AAwCPTx/AOhn3+8MAD8A+M3+3PEujnUbHIIhEcBSAJvst7Nc7P8KgA0AtgKY5PR4\nP/s5NgFYDyAWNYVgAoAV9te9BsDi5r3dBOBTp/e5HcA8++u+BtDC/twpAH4CkALgYwDx9seTATxv\nt3GMffs5+/Z2u51Lwc/paafzLgXwi/08d7mxrcj+dwqAjfbbAQBv2B+/2f7eNwKYA8dYfTv4f1oP\n4Gyn4xUCyAFwkpvzCSGEECKM6AagEvxht4DCYr79uStBsQHQQ2N4kS4CsMR+fwAcwuk2OATaB2DY\nDaC4iHNx7jb2v5EAVgM4GUA0gF1gSBOgOIusdZ6XADxlv38pACtce6a2Oz1uvM/eTvYNt9/fDOA8\n+/3JoCiD3abZTsdbDWC6/f6DAA6CQjQawD6n92P8jQGwxWnb2YNWWMvW1nY7+gLoBWAZHJ/3K2D4\nMgnAX6BAbQZgHfhZGEwGcB+EEI2CqFAbIIQIOXsAbLPf3wbmbQH0AHWz348H8BaAHgBsoECoi4Gg\nFwiggCpwsc8w0MMUBYqPE+2PHwLwq/1+kYvXnQdgqP3+lwDy3NjQCUCu0/YeUATBfvxuoHBsDWCt\n/fGFAD50es0HtY65zP53q/2WYd/eDaCL3ZYxAK62P94FQE/UHX60gJ7LZ0GP2WhQoP5if74FgHTQ\ni5kMesoM25w9mQcBHFfHeYQQYYQEmhCi3Om+FUCF031jjHgawCpQGB0DCoX6cBd6BIBjATwK4HQA\n+WCItQUo/jyhrmO7w/l9VsMR4qzruMVujmHFkZ9bFOjpuxBAfwBloNfN1XmcmQRgLygODRYCeLLW\nflfVY6sFnn9+QgiToxw0IYQnxIEeGoB5UPWxCo5wWySODHHGgeKnAAwTDgHFRSroTTvdvp9zaNVg\nDZhfBvvr2sA1B8FwoDss9vPnATjX/tgI1BSf3ghBC/i+8kBxdgIo1OriClDQjXF6bBWA6wC0t2+3\nBdAVzDs7377dDMD1tY6VBFaKCiEaARJoQojaXhebi/szwfyr30DB5Gofm9P9MWCYczMYqutV6xwp\nYDhvBxjeW2d/vBIMfc4CiwSMZH7nY08G8A8wxDgUzMtyxTo4hF5d7/NWAP9nt6k3mLjv7jXOj7s6\n3nLQk/Y7+Hn9WMfrAeBhMBT7M/h5TAJz554CCyFS7H87gmHOSfZjrgPD0c42nAFHqFYIIYQQwpQM\nAPBqqI0IEnFgdakQQgghhOkxU6PaQPIgHEUZQgghhBBCCCGEEEIIIYQQQgghhBBCCCGEEEIIIYQQ\nQgghhBBCCCFc8v8g+URU3Pf4WQAAAABJRU5ErkJggg==\n", + "text": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "" + ] + }, + { + "cell_type": "heading", + "level": 2, + "metadata": {}, + "source": [ + "Linear Transformation: Principal Component Analysis (PCA)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top]](#Sections)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "The main purposes of a principal component analysis are the analysis of data to identify patterns and finding patterns to reduce the dimensions of the dataset with minimal loss of information.\n", + "\n", + "Here, our desired outcome of the principal component analysis is to project a feature space (our dataset consisting of n x d-dimensional samples) onto a smaller subspace that represents our data \"well\". A possible application would be a pattern classification task, where we want to reduce the computational costs and the error of parameter estimation by reducing the number of dimensions of our feature space by extracting a subspace that describes our data \"best\".\n", + "\n", + "If you are interested in the Principal Component Analysis in more detail, I have outlined the procedure in a separate article \n", + "[\"Implementing a Principal Component Analysis (PCA) in Python step by step](http://sebastianraschka.com/Articles/2014_pca_step_by_step.html)." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Here, we will use the [`sklearn.decomposition.PCA`](http://scikit-learn.org/stable/modules/generated/sklearn.decomposition.PCA.html) to transform our training data onto 2 dimensional subspace:" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "from sklearn.decomposition import PCA\n", + "sklearn_pca = PCA(n_components=2) # number of components to keep\n", + "sklearn_transf = sklearn_pca.fit_transform(X_train)\n", + "\n", + "plt.figure(figsize=(10,8))\n", + "\n", + "for label,marker,color in zip(\n", + " range(1,4),('x', 'o', '^'),('blue', 'red', 'green')):\n", + "\n", + " plt.scatter(x=sklearn_transf[:,0][y_train == label],\n", + " y=sklearn_transf[:,1][y_train == label], \n", + " marker=marker, \n", + " color=color,\n", + " alpha=0.7, \n", + " label='class {}'.format(label)\n", + " )\n", + "\n", + "plt.xlabel('vector 1')\n", + "plt.ylabel('vector 2')\n", + "\n", + "plt.legend()\n", + "plt.title('Most significant singular vectors after linear transformation via PCA')\n", + "\n", + "plt.show()" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "display_data", + "png": "iVBORw0KGgoAAAANSUhEUgAAAl4AAAH4CAYAAACbjOPoAAAABHNCSVQICAgIfAhkiAAAAAlwSFlz\nAAALEgAACxIB0t1+/AAAIABJREFUeJzs3Xl4VOX5//H3TPaEEAIRQSSACxZ3LWKlWqm2ihWXWqks\noli/rUur/pC2iFVErUuLWhRQUFFUBNz3fQERFwIICCgQZE1YkpCVLCSZmd8f94SZhCyTZDKT5fO6\nrlzMds6558zhzD3385znAREREREREREREREREREREREREREREREREREREREREREREZFGKwL6BmE9\no4GP/O7/EkgHCoFLgPeBq4KwnZbyPjAmBNuZDLwQgu20F1XHURFwcQttww0c4b39BHBHC22nLYkD\n3gHygZfCHEttap5vQiEVOw4dId6uiDTBVmA/0K3G4yuxk35qM9fv/8XRWnwG3BTibS4Crg3xNhvr\nLlp/4rWI1rMfax5HLXGst8b/P2DnjXPCtO0xwFLAGabt++uLfUatIZamGgu4sMStADv3X+j3fGdg\nKrDN+5pNwP84+DtjEZALRLdotB1UWz7A5GAeYDMw0u+xE7BflZ4gbaO1/QpLBX4I8TaDtS9bUnM+\np1CdF5qzHx0E91is7Thq6vojmxlLS6lrn3nqeLxKS76fPsBGLOFprJaKq7Wd4xrrKyAR6ALMBl4G\nkrAk6jNgAHC+9zVnADnAIL/l+3rvZ9Fy1V+RdmML8C8gze+xh4DbqV7xSgKex/5jbfUuU3WyOQr4\nAiv9ZwPzvY8v9q5jH/ZLaXgt26+57AK/5/x/7XfDmhcKvLH+G/iyxmuvw07IecB0v+fG+r32J+zX\nXQnW1BjNwVWUP2NfqIXAOuAU7+O3Yb/2qh6/tMY2lgBTsF99m4Gh3ufuAyqBUu9+eKyW/RALzMVO\naHne93iI9zn/+OrbDkA/bL8XAp8AM/BVsYYAO2psdyu+ysVkqle8XgF2YZ/NF8Cxfs/NwZq+3sc+\n35rVjyuAZTUeGwe85b0dgx1n24Dd3nXF+r32EmAV9nlvwk76de3Hwd5t5WP77Qy/9SzCjpWvsM/8\nSGwf/oTto83AKGo3CPgG+zx2AtOAKO9z/sdREfA1tR/rw7zvI88bwwl+698K/BP43vueakte/f8P\nzAHu9d4eAmQAtwJ7vPGN9Vuuvv3bBXgX+7+ci/2/6uW37CKq77OaFbcXarz3v+Or/PzJu81F3tc2\ndAzN8MZSCHxbY1v/8763AmwfHQfcjVXoy73bvgY7D92B7c89wHNYlYZa4voCuNr73h7BPpdN2DF0\nDbDduw7/rgcXYlWgAu/zd/k9t927/iLve/gF1c830PDxeQ/2f7oQa6KsWUmq8iPVK1GR2DnzZA6u\nvF2D7xz2E/CXOtZJLfEmeNd1KvB/2PETX8/yAJOAt7HvhXcaeK1Ih7cFOBdYD/wMiMC+nFOpnng9\nD7yB/afsA2zATmZgidZE7+1o7ERTpaGmkkCXXQDMw748BmAnvMU1Xvs2dsLtjX2pnO99bizVTyxb\nqJ4oLPR7L8OxL7Sfe+8fiW8fXA708N7+I/Yle6jfNsqxBMkBXA9k1rGN2lznjT/Wu/wp2K/Lmss2\ntJ1vgP9iJ+VfYl8Wz3ufG8LBiZf/vphM9cRrLPZ5R2Ffgiv9npuDfZFUfYnE1FhvHHbSP8rvsWXY\nfsO7vjexJKCT973f731ukHfd53rvHwYc471dcz92xb48R2NfOiOwZCLZ+/wi7At5gPf5JGyfHO19\n/lCqJwP+TvXG4sSO+R+AW/yer3kc1TzWT8G+xE/DPqurvMtUJW9bge+wpKfm/qttnc9iX9Jgn2UF\n9plFABcAxd73B/Xv367A77FjrRNW3XjDb5uLqL7PaqsS1Xzvfb2xzsE++6r3M5b6j6EcYKD3PczF\n96PtfGA5vgTqGHz/9+7Cd0yDHQ/p3hgSgNf8nq8ZV6w3pgosAXNgyWwGvsT6t9ixW5VsnI0lfWCJ\n827shwHYcVGzqXEsvvNNIMdnOvb/JBY7vh+gdndi+6jKhdgPQP/3WRXH77AfYQC/wo6NU6idf7yR\n2DFegO37Bdhx15BN2Hs8Gjs/dQ9gGZEOqyrx+hd2Yh6K/eqKwJd4RWC/Mn/mt9xfsJME2C/MWVT/\n1VylocQrkGUjsP/MR/s9dy8HV7z8k7aXgAne22MJPPH6iMD7f63EV1Yfi51Aq8R7Y6o6AS2k/r5J\n13BwRaS2+OrbTir2heJfOXqBpide/rp4t1OVDM7x/tXnBezLAuyzK8SXWO6j+nFxBlZ9AjseHq5j\nnTX34xisUuLva+xLter1k/2eS8C+CC/Dvogb4/8Br/vdbyjxegJfolRlPXCW3/JjG9hmzcTLv+JV\nQvUv/D1YotjQ/q3pZCwZqFJzn9WmrsSrbz3L1DyGngWe9Hv+Aqyqg3fdG4DTObgSOJnqx+ln2A+Q\nKv2x84WzjrjGYpXxKid4X3OI32M5wIl1vI+pWLUMau/jNRbf+SaQ4/N2v+duAD6oY7tH4vs/BPAi\nvostaovD3xvAzXU8NxY7b+RhFbSv8X22H+NL2OtyJlaxrfpcV2H/VySI1Mer/fFgJ7LR2Anhear3\nWUjBfglu83tsO75k6Z/e16cBa7EkIlCBLHsI9kvMP2nIqOV1u/1ul2Bfso11OFaar81VWLKV5/07\nnurNAjW3D1ZRqFJf/6QXsKRvAVbB+g9190epazuHYV+gZX7P76Bp/U8igAexX7IF2Bct2LEA9l5q\nJnE1zcPXd3AUdvIvwz7PeGAFvn35gd+66/sMqrZd5TDsWPS3zft4Ff84i7Fm0Oux5rl38VXTaurv\nfX4Xtg/uo+5moNr0Acbje4952HurK7bG2kv1fk4l2HHQ0P6Nx5Lbrdj7+gKrlPkfJ02Ny385J/Uf\nQ2DJYpVSfP9fPse6C8zwvmYWvi/2mnpy8LkpEl81umZctW0XLOmoLZbTsQQpC6vEXkfgx0Egx6f/\n/2f/7db0E5aYXox9hhdh/8dqcwGW8O3FPv/fNRDzt1gV7hDsB+zn3sf31oi1NldjCVqR9/4r+BJL\nCRIlXu3TduwX8QVU/1UP9uuvguq/GlPxJT97sApYL+yk9DiBX4kVyLLZWN+e3n6P9aZl7KB681iV\nPtiv879izQfJWKIYaFLTUKfwSqw6chx24htG44e42OWNzb+Sk+q37WKq99WIoPqvfH+jsBP8udiX\nclWzRWOSuE+96z8Ja2Kp+pLIwb5gjsX2YzJWDalqVqrrM4CD92Mm9tn460P15teay3wMnIc1Xa0H\nnqpjW09gzYtHYfvgXzTu/LcdS9aS/f46UX0IhMZeLBDI6xvav+OxpHIQ9r7O5uBO9A1tp67n/R8f\nTfOOoWlYM+Sx3nj/UcfrdnLwuamS6slVcy7KmIc12x6O7ceZ+I6DhtYbyPHZGPOxHzOXYMdmbVXM\nGKy59b9YJTwZ64vZlB9gn2LNvnX18YrDug+cg51/dmHH10nUXTGUJlDi1X5di/0HKq3xuAvrB3If\n9sXRB+soXdXfYDh2UgL7RejB90t8D1Yir0t9y/pv/3WsiSEOa/IcQ/0nvaZewfY01ln4VO/yR2En\n8gTv9nKw/wPXYBWvQDW0H4ZgTR4R2C/HCux9N8Y2rF/MZKxCeQaWwFXZiDVT/M77/B3U3beoE9a8\nnIu995rNDYHs2wrs1+9D2Mn/E+/jbizZmYov8euFJUNgV1Vdgx2LTu9zVVWpmvvxfexLeSRW5bgC\nOz7erSPW7tiXVoI3vmLq3s+dsM+ixLvOGxp4vzVjewqrrFU1/yVg/XLqqmg0JNBjuqH92wn7P16A\nJep31VxBANtp6Hiu2k5Tj6GBWKUpCtv/ZdT9Oc3Hzkd9vdu8H6scN+Wqx9p0wqpG5dhnOQrfuSfb\nu5269sUHNO74bMgCLBG6HmtqrE209y/HG9sF+D77xnoB+yH0GvZ/0IlVzm73rvdSLMkdgCVbJ3lv\nf0nrHhuxzVHi1X5txjr7VvFPbG7CvqQ2Y/+pXgSe8T43ECtVF2FXrd2MNWOAJQHPYSeuy2vZZn3L\n+m//b9iv5t3e9c3HToS1xVp131PL7Ya8iiWY87D+FK9jScMPWL+jb7wxHI9diVTb9mqL6VHs/edi\nX4g19cCSlALvthZRe3+rhrYzGku49mL9gV7Ct58KgBux5DID6wfk3wTjv+7nsUQuE6vsfVNjO4Hu\n03lYxeMVqn8RTsCaoL71xvUJ9gUF1gn/Gqwzdj62L6oucKi5H3Ox5HI89kXzd+99/z5L/nE6sS/p\nTGwfnUXdCdXfsS/ZQqzauaCB9zyZ6sf6CuwK2eneeNKxL6PGVF/q2+f1rae+/TsV+wGTg/Xn+aCW\ndTUU4wNY4p6HXVlZ2zJNOYaq7nfG9nkudj7Iwa7krW25Z7D/K4ux81MJ1ftp1nduqOs1/m7EqtGF\nWJ9F/4plCXa++Mob6+k11r+Xxh2fDf2/2o19Zmdw8OCxVcsVYefRl73bGYnvauLa1LfNcuA3WGX4\nE+xYWool7Eux4/kZ7HyS5f3bgx3zo1C+INKu/IfArrbp6F6i9oqGiIiISJ2OwfoMOLByfzYaqK82\nA7FmDyfWFFCKlf9FREREAjYQa6qpau6cUP/LO6xhWKfuYqx5QFcXiYiIiIiIiASiVc9JddJJJ3lW\nr14d7jBEREREArEaG8i4Tq36KoXVq1fj8Xj0F8K/u+66K+wxdLQ/7XPt847wp32ufd4R/gigH26r\nTrxERERE2hMlXiIiIiIhosRLqhkyZEi4Q+hwtM9DT/s89LTPQ0/7vHVq1Z3rAY+3zVRERESkVXM4\nHNBAbhUZmlBEREQkXLp27UpeXl64w2g3kpOTyc3NbfiFtVDFS0REpJ1zOBzo+zR46tqfgVS81MdL\nREREJESUeImIiIiEiBIvERERkRBR4iUiIiISIkq8REREpFWaM2cOZ511VrjDCColXiIiIiLA9OnT\nGThwILGxsVxzzTUtsg0lXiIiIlKr0lJYutR3f+dOSE8PXzwtrVevXtx555386U9/arFtKPESERHp\noDZtAv9xVVeuhMpK3/3CQpg5Ez7+2JKuf/0Ltm2rvo6aw1k1ZbiwHTt2cNlll9G9e3dSUlK46aab\nan3dLbfcQmpqKklJSQwcOJAlS5YceC4tLY2BAweSlJREjx49GD9+PABlZWVceeWVpKSkkJyczKBB\ng8jKyqp1/b///e+55JJL6NatW+PfRICUeImIiHRQ338Pt99uydfHH8Njj0F+vu/5Qw+F++6DadPg\nuutg5Ej4zW98z3s8cP/9sGaN3c/IsPVVVAQeg8vlYtiwYfTr149t27aRmZnJyJEja33toEGDWL16\nNXl5eYwaNYrhw4dTXl4OWFI2btw4CgoK2Lx5M1dccQUAzz33HIWFhWRkZJCbm8usWbOIi4urN6aW\nHGxWiZeIiEgHddllcPbZcNVV8MQTlmSlpAS+vMMBF18MDz4IH3wAd9wBv/0tREUFvo60tDR27drF\nlClTiIuLIyYmhsGDB9f62tGjR5OcnIzT6eTWW29l//79bNiwAYDo6GjS09PJyckhPj6eQYMGHXh8\n7969pKen43A4OOWUU0hMTGzgfbXcxD5KvERERDqwrl3tX48HahaC9uyx5sWbboJZs2D+fPj00+qv\nOeEEuPJKePxxOOUUOOecxm1/x44d9OnTB6ez4ZTkoYce4thjj6VLly4kJydTUFBATk4OALNnz2bj\nxo0MGDCAQYMG8d577wEwZswYzj//fEaMGEGvXr2YMGEClf7tqbVQxUtERESC7pNPLJmaNQtGjLBm\nQv+mxs6d4frr4bzz4LDDrCLWp0/1dWRkwEsvWRNkWpqv2TFQvXv3Zvv27bhcrnpf9+WXXzJlyhRe\neeUV8vPzycvLIykp6UCSdNRRRzFv3jyys7OZMGECl19+OaWlpURGRjJp0iTWrVvH119/zbvvvsvz\nzz9f77ZU8RIREZGgO+QQS6YOO8wSr+HDISbG93xcHJx+uu/+YYfB0Uf77ns8MGOGNVXecgvcdptV\nvhrTx+v000+nZ8+e3HbbbZSUlFBWVsbXX3990OuKioqIjIwkJSWF8vJy7rnnHgoLCw88P3fuXLKz\nswFISkrC4XDgdDpZuHAha9asweVykZiYSFRUFBEREbXG4nK5KCsro7KyEpfLxf79+xtMCBtLiZeI\niEgHdfLJlkxVOeecg5sb6+NwwD33+JoXTzjBOug3po+X0+nknXfeYdOmTaSmptK7d29efvll7/od\nB6pPQ4cOZejQofTv35++ffsSFxdHamrqgfV89NFHHH/88SQmJjJu3DgWLFhATEwMe/bsYfjw4SQl\nJXHssccyZMgQxowZU2ss9957L/Hx8fznP/9h7ty5xMXFcd999wX+ZgLQcrW04PC0ZDuriIhIR+Bw\nOFq031JHU9f+9CaJ9eZWqniJiIiIhIgSLxEREZEQUeIlIiIiEiJKvERERERCRImXiIiISIgo8RIR\nEREJESVeIiIiIiGixEtEREQkRJR4iYiISKs0Z84czjrrrHCHEVRKvERERKTDKy8v59prr6Vv3750\n7tyZU045hQ8//DDo24kM+hpFRESk/di6FebMgcJCOP98OO88m6SxnamsrCQ1NZXFixeTmprKe++9\nxx//+EfWrFlDnz59grYdVbxEREQ6Ko8HPvsMJk2CqVNhz57qz+/cCWPHwvvvw9KlcPvt8MYbB6+n\nogJ274aysiaFsWPHDi677DK6d+9OSkoKN910U62vu+WWW0hNTSUpKYmBAweyZMmSA8+lpaUxcOBA\nkpKS6NGjB+PHjwegrKyMK6+8kpSUFJKTkxk0aBBZWVkHrTs+Pp677rrrwMTbF154If369eO7775r\n0nuqixIvERGRjurVV2HCBPjkE5g715Ks3Fzf859/bpWuQw+Frl0hKQmee676On74AS68EC65BH77\nW/jii0aF4HK5GDZsGP369WPbtm1kZmYycuTIWl87aNAgVq9eTV5eHqNGjWL48OGUl5cDlpSNGzeO\ngoICNm/ezBVXXAHAc889R2FhIRkZGeTm5jJr1izi4uIajGvPnj1s3LiR4447rlHvpyFKvERERDqq\np56CLl0gJQV69oSsLFi8OPDlKyrg//0/KC2F7t0hOtqqYtnZAa8iLS2NXbt2MWXKFOLi4oiJiWHw\n4MG1vnb06NEkJyfjdDq59dZb2b9/Pxs2bAAgOjqa9PR0cnJyiI+PZ9CgQQce37t3L+np6TgcDk45\n5RQSExMbeFsVjB49mrFjx9K/f/+A30sglHiJiIh0VG43OJ0HP1blnHOsyrVnj1XCCgrg6qt9z+/d\na4916WL34+Nt+e3bAw5hx44d9OnTB2fNOGrx0EMPceyxx9KlSxeSk5MpKCggJycHgNmzZ7Nx40YG\nDBjAoEGDeO+99wAYM2YM559/PiNGjKBXr15MmDCBysrKOrfhdrsZM2YMsbGxTJ8+PeD3ESglXiIi\nIh3VFVdATo4lT1lZ0Lkz+FebDjsMnn0Wfvc7+MUv4P774fe/9z3fpYtVuUpK7H55uSVehx4acAi9\ne/dm+/btuFyuel/35ZdfMmXKFF555RXy8/PJy8sjKSkJj8cDwFFHHcW8efPIzs5mwoQJXH755ZSW\nlhIZGcmkSZNYt24dX3/9Ne+++y7PP/98rdvweDxce+21ZGdn89prrxERERHw+wiUEi8REZGO6tpr\nrY/XMcfAkCEwezb06FH9NX37wuTJ8PDDdlWj/xWNsbHw739bp/qcHMjPh3Hj4PDDAw7h9NNPp2fP\nntx2222UlJRQVlbG119/fdDrioqKiIyMJCUlhfLycu655x4KCwsPPD937lyyvU2cSUlJOBwOnE4n\nCxcuZM2aNbhcLhITE4mKiqozobrhhhtYv349b7/9NjExMQG/h8bQcBIiIiIdldNpVS9vR/QmOfts\nePNNa1489NBGJV0WgpN33nmHm2++mdTUVBwOB6NHj2bw4ME4HA4c3kRv6NChDB06lP79+5OQkMC4\nceMOXIEI8NFHHzF+/HhKSkro27cvCxYsICYmhj179nDDDTeQkZFBp06dGDFiBGPGjDkojm3btvHk\nk08SGxtLD7/k88knn6yzs39TtIaBOCKA5UAGcFGN5zxVJUQRERFpGofDgb5Pg6eu/elNEuvNrVpD\nU+MtwA+AjggRERFp18KdeB0O/A54mtZRfRMRERFpMeFOvP4H/ANwN/RCERERkbYunJ3rhwFZwEpg\nSF0vmjx58oHbQ4YMYciQOl8qIiIiEjKLFi1i0aJFjVomnM179wNjgEogFugMvAZc5fcada4XERFp\nJnWuD67mdK5vLf2qzgb+jq5qFBERCTolXsHV1q9qrKIjQkRERNq11lLxqosqXiIiIs2kildwtZeK\nl4iIiMgBc+bM4ayzzgp3GEGlxEtEREQEuPLKK+nZsyedO3fmiCOO4L777gv6NpR4iYiISL3S96bz\n2g+vhTuMFjdx4kS2bNlCYWEhH3zwAdOmTePDDz8M6jaUeImItBHrstYxZ9WccIch7dCb698kszCz\n1uc8Hg8Pf/Mw9395P3v27alzHWWVZU3e/o4dO7jsssvo3r07KSkp3HTTTbW+7pZbbiE1NZWkpCQG\nDhzIkiVLDjyXlpbGwIEDSUpKokePHowfP97iKivjyiuvJCUlheTkZAYNGkRWVlat6z/uuOOIjY09\ncD8yMpLu3bs3+X3VRomXiEgb4PF4mPL1FB799tE6vyBFmmJn0U7uXnQ3M5bNqPX5NVlrWLFzBU6H\nk+dXP1/ra3JLc/n9gt+zIWdDo7fvcrkYNmwY/fr1Y9u2bWRmZjJy5MhaXzto0CBWr15NXl4eo0aN\nYvjw4ZSXlwOWlI0bN46CggI2b97MFVdcAcBzzz1HYWEhGRkZ5ObmMmvWLOLi4uqM58YbbyQhIYHj\njjuOO+64g1NPPbXR76k+SrxERNqAFbtWsC5rHVERUTyz8plwhyPtyLMrnyXCGcEnmz9hS96Was95\nPB4eX/Y4kRGRpCSk8NqPr9Va9Zr7/VzSc9OZuXxmo7eflpbGrl27mDJlCnFxccTExDB48OBaXzt6\n9GiSk5NxOp3ceuut7N+/nw0bLNmLjo4mPT2dnJwc4uPjGTRo0IHH9+7dS3p6Og6Hg1NOOYXExMQ6\n43n88cfZt28fn376KXfccQdpaWmNfk/1UeIlItLKeTwepqdNJzoympT4FN7Z+I6qXhIUO4t28vaG\ntzkk4RCcOHnqu6eqPb8maw1fbf+KSGckxeXFFO0vOqjqlVuay/y18zki+QiWbF/S6KrXjh076NOn\nD05nwynJQw89xLHHHkuXLl1ITk6moKCAnJwcAGbPns3GjRsZMGAAgwYN4r333gNgzJgxnH/++YwY\nMYJevXoxYcIEKisr692Ow+FgyJAhDB8+nPnz5zfq/TREiZeISCu3YtcKlmUuw+PxULC/gKLyImav\nnB3usKQdeHbls5RUllBWWUZsVCzvp79freq1b/8+fpn6S4495FgGHDKAM/ucSXREdLV1zP1+Li63\ni5jIGCKcEY2uevXu3Zvt27fjcrnqfd2XX37JlClTeOWVV8jPzycvL4+kpKQD42kdddRRzJs3j+zs\nbCZMmMDll19OaWkpkZGRTJo0iXXr1vH111/z7rvv8vzztTeZ1lRRUUFCQkKj3k9DwjlJtoiIBMDp\ncHLxMRdXe6xXYq8wRSPtSbmrnJMOPenA/dSkVLJLsumX3A+AwamDGZxae7MfQElFCa+sewWXx0VO\nSQ4ej4fF2xazLX8bfbr0CSiG008/nZ49e3Lbbbdx991343Q6+e677w5qbiwqKiIyMpKUlBTKy8t5\n8MEHKSwsPPD83LlzOf/88znkkENISkrC4XDgdDpZuHAhKSkpHHvssSQmJhIVFUVERMRBcWRnZ/PZ\nZ59x0UUXERsby6effsorr7zCp59+GtD7CJQSLxGRVu7Unqdyas/gdvAVAbj713c3a/nYyFimDp1K\nhbviwGMOHPRM7BnwOpxOJ++88w4333wzqampOBwORo8ezeDBg3E4HFWjwTN06FCGDh1K//79SUhI\nYNy4caSmph5Yz0cffcT48eMpKSmhb9++LFiwgJiYGPbs2cMNN9xARkYGnTp1YsSIEYwZM+agOBwO\nBzNnzuSGG27A4/HQv39/XnjhBU477bRm7KGDacogERGRdk5TBgWXpgwSERERaQOUeImIiIiEiBIv\nERERkRBR4iUiIiISIkq8REREREJEiZeIiIhIiGgcLxERkXYuOTn5wHhY0nzJyclNXra1fwoax0tE\nRETaBI3jJSIiItKKKPESERERCRElXiIiIiIhosRLREREJESUeImIiIiEiBIvERERkRBR4iUiIiIS\nIkq8REREgujZlc+SVZwV7jCklVLiJSIiEiRrs9Yy5espPLvy2XCHIq2UEi8REZEgmbl8JglRCby+\n/nV279sd7nCkFVLiJSIiEgRrs9ayNGMp3Tt1x+Px8Nyq58IdkrRCSrxERESCYObymex37Wdf+T6i\nIqJ4ad1LqnrJQSLDHYCIiEh70CW2C2ccfsaB+xHOCArKCujRqUcYo5LWpt4ZtFsBj8fjCXcMIiIi\nIg1yOBzQQG6lpkYRERGREFHiJSIiIhIiSrxERKTD+W7Xd7y67tVwhyEdkDrXi4hIh+L2uLn/y/vZ\nUbiDc444h65xXcMdknQgqniJiEiHsmT7Erblb8PtcTN/zfxwhyMdjBIvERHpMNweN48tfYy4qDi6\nxnXlxTUvkluaG+6wpANR4iUiIh3Gku1L+DH7R9weN8XlxeSX5avqJSGlPl4iItJheDwehh49tNpj\niTGJYYpGOiINoCoiIiISBBpAVURERKQVUeIlIiIiEiJKvERERERCRImXiIiISIgo8RIREREJESVe\nIiIiIiGixEtEREQkRJR4iYiIiISIEi8RERGREFHiJSIiEibF5cVohpaORYmXiIhIGFS6Kxn75lje\n3vB2uEOREFLiJSIiEgafb/6c9TnreXzZ45S7ysMdjoSIEi8REWkT3B43C7csxO1xhzuUZqt0VzIt\nbRrd4rvo5n7IAAAgAElEQVSRW5rLB+kfhDskCRElXiIi0iYszVjKrR/fyrcZ34Y7lGb7fPPn7N63\nm8SYRBJjElX16kCUeImISKvn9riZljaNSncl09Omt/mq13PfP0eFu4Kc4hxKK0rZtW8Xi7ctDndY\nEgKR4Q5ARESkIUszlpK+N50+SX1I35vOtxnfMrj34APPz18zn4ToBC4+5uIwRhm4e4bcQ+H+wmqP\nHdX1qDBFI6GkxEtERFq1qmpXVEQUbo+bqIgopqdN5xeH/wKnw0lBWQHTl00nyhnFb474DfFR8eEO\nuUFHdj0y3CFImKipUZokIwPWrvXdX74ccnLCF4+ItF97S/aSW5oLQH5ZPgA5JTlkF2cD8PK6l6lw\nVVBcUcyb698MW5wigXCEO4AGeDSwXOu0Zg08+CBMnAhFRfD443D33XDEEeGOTEQ6koKyAi6cdyEJ\n0Qm43C4q3ZW8P/r9JlW9Kt2VRDrVECRN53A4oIHcShUvaZITToAJEyzxuv9+mDxZSZeIhN7L614m\nqziLvNI8CvcXsqtoF2/+2Piq14/ZPzL85eEUlxe3QJQiPkrtpcmK/c5PpaXhi0NEOq4jux7JDafd\ncOD+5tzNvLn+TUaeMLKq+hCQGctmsDZrLa//+DpjThrTEqGKAGpqlCZatgwee8wqXcXF8J//wJ13\nws9+Fu7IRKSjcrldXP7y5WzK3cRzv3+OU3ueGtBy67LWMfatsSTFJFHhruD9Ue+TEJ3QwtFKe6Sm\nRmkxfftan64jj4QTT4Q77oDDDgt3VCLSkX2x7QsyCjNIjElkRtqMgCeffmL5E0Q6IomLiqOkooTX\nf3y9hSOVjkyJlzTJIYdU79M1YAB07hy+eESkfSmpKGnUCPUut4tpS6cRHx1P17iufL/ne1buXtng\ncptyN7Fk+xJcHhfZxdlUuCqYs2oOFa6K5oQvUif18RIRkVbn5XUv8/iyx3lrxFv0TOzZ4Ou/2vEV\n63PW0ymmEyUVJRSVFzFr+SxmXTSr3uV6durJ/87/X7XHYiJjiHBGAJbQfbn9S87uc3aj+oyJ1KW1\nH0Xq4yUi0sEU7S/iwnkXUrC/gBHHjWDiWRMbXCajMIPlO5dXe6xbXDfO6nNWs2L5bPNn/OOTf/DM\nJc9wco+Tm7Uuaf8C6eOlxEtERFqVOavm8Piyx+kW343cklzeHPFmQFWvYHO5Xfzh5T+wce9Gzkw9\nk1nDZrWKqpfL7TpQkZPWRZ3rRUSkTSnaX8Ts72aTGJOIAwcuj4s5q+aEJZZFWxexo3AHeaV5LN62\nmNV7VoclDn+fbf6Mv7zzlzY/SXhHpsRLWpzL5bvtdttfR7JrF7z/vu/+8uWwalX44hFpzTbs3YDD\n4aCkvIS80jyinFF8t+u7kMfhcruYljaN0opSisqLyCrOatSVki2h0l3Jo0sfJS0zjW92fBO2OKR5\n1LleWtSPP8Izz9h4X3FxMGMG9O4Nl14a7shCJzoa3nrLBpnt0wemTrUxz0TkYAMPG8jiaxaHOwwy\nizIp3F/IzqKdOB1Oyl3lrMlaQ+H+QpJik8IS0+ebP2fXvl10ievCtLRpnNH7DJwO1U/amvA3VtdP\nfbzaOI8HnnwSNm60ISjy8+GuuywJ60j27oWxY+32Qw/BMceENRwRCcDbG97m3sX30qNTD/LL8unZ\nqScLLl8QlmSn0l3JZS9dRuH+QjpFd2LPvj1MHTqVX6b+MuSxSN3Ux0vCzuGAP//ZEq+vvoJ//KPj\nJV0AW7b4bq9dG744RCRwz6x8hgpXBTklOVS6Klmfs54VO1eEJZavtn/FT7k/UVpZSnZJNqWVpTz9\n3dNhiUWaRxUvaVFutzUvZmTYyPbbt8M990BCB5qNY8MGuPdea15MSYHbb4c//AHOOy/ckYlIfVbt\nXnXQpNknHnoiiTGJIY8luzj7oM79ybHJ/Pywn4c8FqmbhpOQsFu/Hp5/3pKO2FhrduzRAy65JNyR\nhU5FhXWwT021+3v3QkQEdOkS3rhERCS4lHhJq+B2g9PbqF31cbaCoXBERESCSn28pFVw+h1lDoeS\nLhEJr11Fu9hesD3cYUgH1dq/AlXxEhGRoPF4PFz/7vXsK9/H3MvmtoqR6KX9UMVLRETEz5qsNXy3\n6zs25m5kaebScIcjHVC4E6/ewEJgHbAWuDm84YiISHvl8XiYkTaDyIhIYiJimLZ0WlhHopeOKdyJ\nVwUwDjgO+AXwV2BAWCMSEZF2qara1S2uG11iu6jqJWER7sRrN1A1a90+4EfgsPCFIyIi7dUXW7+g\n0lPJrqJd7Crahcvt4vMtn4c7LOlgWlOvwr7AF1j1a5/3MXWuFxGRoHB73Lg97mqPOR1OzXcoQRNI\n5/rWMkl2J+BV4BZ8SRcAkydPPnB7yJAhDBkyJJRxiYhIO6EkS4Jt0aJFLFq0qFHLtIaKVxTwLvAB\nMLXGc6p4SUCKiqBTJ98YYUVFkBj6WT1ERKQDawvDSTiA2cAPHJx0iQRsxgx49lkbGX/1avjb32Df\nvoaXExERCaVwV7zOBBYD3wNVpa2JwIfe26p4SUCKimw+SIDsbJg4EY4/PrwxiYhIx9IW+ngtIfxV\nN2kHEhNh+HB48EHo3x+OOy7cEYmIiBxMSY+0C6tXwxNPwL/+BS6Xr9lRgs/lgsJC3/2iIqisDF88\nIiJtiRIvaReWL4fbboNf/ALuvdeaG4uLwx1V+7R0qe3rvDzIz7dm3W++CXdUIiJtQ7j7eDVEfbya\nwOOBkhJISLD7ZWUQGWl/IsGwYAG8847dvvBCGDnSd0WpiEhH1RauapQW8P338Pe/Q26uJV333AMf\nfRTuqKQ9GTrUmhsLC+GCC5R0iYgEqrWfLlXxaqKXX4YPPoDYWPjZz+Cmm8CpNLvJZs+GAQNg8GDr\n0/TII3DzzZCcHO7IQi8/H+64w/aF0wmLF8N993XMfSEi4q8tXNUoLeTii+GFF+z2vfcq6WquX/8a\nJk+2fmPvvQcnnghduoQ7qvDYvh3OPBOuuMIqXU4nbNumxEtEJBCqeLVDVc2Lhx4KPXrAokVWkeja\nNdyRtW2rV1ulp0cPePJJNa+JiEh1qnh1UFu3wuGHw/XXWzXC6YQ1a+Dss8MdWdtVVGRDVJx0klV3\nvvnGmtpEREQao7X/ZlfFS1qFqVOhc2e45hrYssUqiv/7n5rXRETEJ5CKlxIvkQCUltqFClXNi6Wl\nEBcX3phERKR1UeIlIiIiEiIax0ukjVuxwsZjAxsYd9EiTc8jItKWKfGSDi8/Hx54wDfF0LJl8Nxz\n4Y2pypYtNv9kbq4ND/L669bMKSIibZOuapQOLynJOsnfdRcMGwZPPw2TJjVtXZWVUFAA3brZ/bw8\niI+HmJiGl/V4fH3IqlrYL78c3G64+mobN2zGDEhMbFpsIiISfqp4SYfncMB111mS9PDDNt1S//5N\nW9fKlfDPf8KePValuu02m1S6IatW2ZWS5eWWaE2bBp9+aglYWZm9xu1WM6OISFunxEvanKws+6uS\nnm4JS3MsXw7791vCNXeur9mxsU47DS67DG64Aa69Fn77W/jVrxpe7oQT7CrJf//bhqnYtctGh58/\n32J78UW45BJrdiwsbFpsIiISfkq8pM1ZvdoSkKwsu3333ZCR0fT15eXB9OnWvPjQQ3DUUfDUU01f\n3xlnQEWFVafOOiuwZSIi4NZbrWK2aBHcfrslYieeaMlY587wxz9ak2NCQtNjE2lPVu9ezas/vBru\nMEQaRcNJSJv0zjs2bQ9Yx/jjj2/e+oqLfQmNxwMlJU1LcHJzYeJEq3TFxVln+Pvvt+mb6uN2W/KX\nkQGdOtn922+H6OjGxyDSEbg9bka9NorNeZv58MoP6RqnOdEk/DSchLRbqam+2927N399/kmWw9H0\nqlJ2NlxwgXWKv/BC+MMfrL9XQ1avtubFu++2al58PCxc2LQYRDqCb3Z8w0+5PwEw9/u5YY5GJHCq\neEmbs3o1TJliHde3bIG337ZJwIORgIWTy2VNjlW3nU5NxC1Sm6pqV2ZRJvFR8RTuL+S9Ue+p6iVh\np4qXtEo//GBjZVX55BPYuTPw5SMiLOk6/ni46CIYMcKXsLRl/u8hIkJJl0hdvtnxDWuy1uDxeCgu\nL6agrEBVL2kzNI6XhFxkJDz6KNxyi3WQr+oHFaia/bnOPTe48YlI6+byuDjviPOqPZYYrQHupG1o\n7b+p1dTYTm3cCOPH2+2nn26487mIiEhrp6ZGabXS0323t28PXxwiIm3JGz++wTc7vgl3GNIMSrwk\n5BYtsubFp5+2keIffdT6fYmISN0Kygp46OuHuP/L+6l0axqLtkqJl4TcySf7xrbq398GCD3yyINf\nt2SJDW4KNq7Vhx+27JQ5BQX13xcRCaeX171MhbuC3ft28/nmz8MdjjSREi8JuS5dqvfp6tu39kmk\nMzNtENHcXJsceuFCGxG+JVRW2hyLn3vPZW+8YXMnqouhiLQGBWUFPLf6OZLjkkmITmBa2jRVvdoo\nXdUordYVV1jic/XVcNhhMHWqjQbfEiIj4c474Y47LOnav9+qchrSQURag5fXvUxuaS4e7NfgT3k/\n8dnmzzj/qPPDHJk0lhIvabXcbhsJHmwS7LKylku8AA4/HH7+c/j4YxgzBlJS6n6txwM7dvhG0C8v\nt2ZRXZ0pIi2hc0xnLup/UbXHoiKiwhSNNEdr/z2v4SQ6sJkzbWT6yZPhrbfgiy9sEuuWmiT6jTfg\ngw/gxhutunbVVXDOObW/NicHxo2Dv/0NTjkF7r3XkrA//7l5MXg8lsB19Q7AXVpqCWhrnxjb5bK/\nqrklS0shNlYVQxHpWDSchLRpv/qVJV1xcTY6/Z//bHMYtoTKSti82ZoXTz7ZOvyvW3dwH6+MDNi3\nz6phkybZ3IqXXQZJSXDttc2PIz0dbr3VhtgoLbX3/9FHzV9vS/voI0s+y8shPx/+8Q+b2klEWt4b\n699g1e5V4Q5DAtTaf4+q4iWtyvz5sHSpJWbLl8M118DRR1sCdsYZwdnGwoVW7YuMtHXeeKPN29ia\nud3wyCO+hPHXv4aRI1XxEmlpeaV5DH1xKH2S+rDg8gU4Ha38ZNHOqeIlEmQjRsAJJ1jH/+uvt+Ri\n2jR4/HFLyOqTkwNffum7v2ZN9YFkq/ziF1BSAoWFcPHFrT/pAovxT3+ypuHdu+Hyy5V0iQBUuiuZ\ntHASWcVZLbL++Wvn4/a42ZK3ha93fN0i25DgagOndJHQefVV3wTe5eU2uGtOju95hwOOOcb6MyUk\n2GTdxxxjzY5VY47VZf9+mD3bJgVfswYefNAuGPBX1bx4/vnWh2zSpLYxsn9+vsV6xRVw9tm+ZkeR\nju7zzZ+zYO0Cnl/9fNDXnVeax9zv59I1riuxUbE8tvQx3B530LdTnx0FO3hs6WOodSpwSrxE/Jx4\noiVbX30F991nyUNysu/5r76CJ5+0CteNN1qysW+fNTcOHVr/unv1snU+9piNT3bbbVY985edbYnc\njTdax/6rr4affgr++wy2lSth8GAYPdr6qKWkwKZN4Y5KJLwq3ZVMS5tG94TuvPrDq0Gves1fO5/8\nsnyKy4vxeDysy17HV9u/Cuo2GjJr+SxmLp/Jmqw1Id1uW6bhJET89O9vCdHEidaRf948iIjwPe9w\nWH+ufv3s7+23GzfIam6u7/bu3QcnXqmp1mRX5de/btr7CDX/OB0OuOWW8MUi0lp8vvlzdu/bTY/E\nHuzZt4fnVz/P3wf/PWjrj4+K57wjz6v2mMvjCtr6G7I1fysfb/6YxJhEHl/2OE9c+ERVHyephxIv\nET/l5fDKK9CjBxQXw3ffwWmn+Z4fPNh32+GAX/4SnngCbr7Zhk/49lvr5zRy5MHr3rjRmhfvv9+G\ni/jXv2yZs85q+fcVChUVEBVV932RjsTldjEtbRplrjKyirNweVzMWzOPq066iu4J3YOyjbEnjw3K\neprq6RVP48DBIfGHsGLnCtZkreHEQ08Ma0xtQWtPTXVVo4TUzJlQVGTNZT/9ZH2VHn4YutdxnnS7\nrXP97t3WL2v2bOujVdvck6WlNuhq//52PzPTkrfkZN/AsIWFkJjY9jqmezw25dIll8CZZ8KqVTYJ\n+tSpdnWmSEdT4apg5vKZFFcUH3jM6XBy9UlXc2intj/S8tb8rVw8/2ISohNwOpzkl+VzZu8zmXXR\nrHCHFlaBXNXY2k/vSrwkpIqKrImxqnkxP9/mlty2zSbp/vOf7Qq+Tz6xf88915KvK6+0ZR98EI47\nLvDtvfeeDQx7993WV+z22+Gmm6yvWVuzdav1eRs0yCp/t98Oxx4b7qhEpCVsyt3EsyufrdaZPyUh\nhfFnjA9jVOGnxEskSPbvt0mzU1IssZo3zzrK9+plSca0adCtm13peNdd1oQYCLfbqmzLl1sz5/Dh\nVjVqq954A555Bi66CP7yl3BHIyISWkq8RIJo/34bnwrsqsbevW2oiVtvtWSrXz9LwKKi7KrEQGVl\n+Ua9f/nllp2PsiWtWmVTOo0aBQsWWOJ15pnhjkpEJHSUeIkE0SefwLPPQkyMNQXecos1N+7bB506\n2WvcbuvLFejcitnZ1iQ3bJj1+dq61Zod21ry5fFYRXD4cGte3LoVZs2yPnLq4yUiHYUSL5Eg2bjR\nrka87z5rbrznHpscu6oC1lRLlsDevda86HZbM92vfuXrgN+WeDzVLwqoeV9EpL1T4iUSJB4PFBRY\nR3uwZke3u+1VpkREpOVorkaRIHE4fEkXWHNjMJKuVausqRIsufvqK0voRCR8Vu1axYqdK8IdhrRT\nSrxEwmjlSrjzTku+nn8eXnrJ+oiJSHi43C7uWnQXkxZOotJdGe5wpB1S4iUSRmPHwvHH20j3ixdb\nH7JAO+aLSPAt3LKQnUU7ySrJ4uOfPg53ONIOKfESCbOqwVodDnVGFwmnqml+4qPjSYhKYEbaDFW9\nJOiUeImE0dy5Nh/kiy/CGWdYs2NJSbijEumYFm5ZyNb8rUQ6I4lwRrCjcIeqXhJ0GmFHJIwGDIBL\nL7X5Gf/0J2tuDHTUexEJrl3Fu+if4hvLpWtcV/bs2xPGiKQ9au0NGxpOQkRERNoEDSchEkT+wzx4\nPBr2QaQjURFAgkWJl0gAMjJsTsb8fEu6XnwRXngh3FEFz4oV8M03vvtvvmlTGIkILNqyiH9+8s8m\nJV97S/by3a7vWiAqaauUeIkEoFcvOO00uOMOmyD722+tb1Z70aWLva9vvrEJrj/+GOLjwx2VSPhV\nuit55NtH+Pinj1mTtabRy89YNoP/9+H/Y1/5vhaITtoiJV4iAXA4YNQo2LYNPvwQ/vlPSEoKd1Sw\naxcsXeq7v3IlbN/e+PUceSRMnmzzUb74oo0nlpwctDBF2qyFWxayq2gX8dHxPL7s8UZVvTIKM3h3\n47vsK9/H6z++3oJRSluixEs6lJrnzED7aVU1L/bpAxdcAP/9rzU7hltZGcyYAV9/bc2FDz/c9JHv\nly3z3V6/PjjxiYRThauiWctXuisPjOvVLa4bK3auaFTV65mVzwDQLb4bs7+braqXAEq8pIOZPx/m\nzbPb+/bBhAlWIcrKgs8+871u9Wr44Qff/cxMS2zuuw9uuMHG3HrrrdDGXpt+/axS9cAD9u8dd8Ax\nxzR+PR9+aENZPP88TJ1qzY4//hjsaEVCZ9XuVfzxlT9SWtH0Obi+3PYlG/dupLi8mN37dlNYXsiT\nK54MaNmMwgze2fgOyXHJRDmj2FehqpcYjeMlHcrvfgf/+pclXT/8ACecAL17W+I1b55VkA4/HKZM\ngdtu8y13+OFWTXJ6f6qMGnVw9Sxc8vJ8t3Nzm7aOwYPh9NOteTE5GR58EA49NDjxiQTTB+kfsHvf\nbq455Zo6X+PxeJi2dBrrstfx9oa3ueL4K5q0rSO7Hsn9595f7bFD4g8JaNm1WWuJjYhl336rcsVG\nxLIscxlXnXRVk2KR9kPjeEmHk5kJ119vt99+2zdNz5498H//Z7cfeMDmUAyFykrYvduSO7BEyukM\nrA/Z2rWWJN1xB0RHW9Xr5pth4MAWDVkkLMoqyxg2bxj7yvfx7qh3SYlPqfV13+36juveuY6k2CRc\nHhfvj3qfuKi4Fotrf+V+YiJjWmz90nZoHC+RGvbts2rWkCGQmmpNj1V27/bd3rYtdDFt2gQTJ0J6\nuiVdt99efWiH+vTpA3fdBT/7GRxxBNxzj3WUF2mP3t34LoVlhbg9buZ+P7fW13g8HmakzSAqIoq4\nqDj2le/j7Q1vt1hMe0v2culLl5K+N73FtiHtixIv6VDefNOaF2+91fprffut9fFKT7eE7IEH4Omn\n4fXX4fPPQxPTz34Gf/ubxXTVVXD22TB0aGDLJibC0Uf77vftq6sRpX0qqyxj5vKZJMYm0jWuKy+t\nfYmckpyDXvdD9g8s27kMt8dNdnE2la5Knln1DG5Py4x4/OKaF9m4d2PAfb9E1NQoHYrLZc14Vc2L\nLhdERMD+/bBjBxx1lD2+Z489nlJ7S0bQ5eVZ0gXwyCPVk6nGrqdLF9/7y8tTIibtw+s/vs7ETyfS\nObYzAAVlBfz1tL9yyy9uqfa64vJiVuxaUe2xuMg4Bh42sKoZKGj2luxl2PxhJEYnkleax7w/zOPo\nbk38zyvtQiBNjUq8pMPzT048HhsmIpTJSlXz4tln21WK06fDpEmNT748Hrtw4PjjYeRI+OILmDvX\nrlCMjm6Z2MPtq6+gc2erYno88Npr1owcqoRZQictM420zLRqj/0s5Wf85ojfhCkieGzpY7zw/Qv0\n6NSD7OJszko9iynnTQlbPBJ+SrxEGlBebsNDXH01nHWW9flau9aaIYP847hOmZmwfDlccondX7oU\nIiPh5z9v/Lry862jfXm5VfHuvdf6srVXa9bYxQUTJtg4ZGvWwL//DZ06hTsyae+Ky4s5f+75lFWW\nEeWMwu1xU+mu5M0Rb9I7qXe4w5MwUeIlEoBt2+DOOyEuDqKi7Iu7S5dwR9V0b71l/dR++UtLSEKV\nQIbL999bpQ8scVbSJaHg9rhZsXMFFW7fIK1Oh5NTe55KdEQ7LTFLgwJJvDSOl3R4qakwYICN/v7n\nP7ftpGvRIrsw4P77YdYsS0RGjmy/yZfH4xtxPyICtmyxZkeRluZ0ODmt12nhDkPaoPquajwR+BbI\nAJ4E/Hu9pNW6hEgb4/FYcpKZac2Lr75qI7i3RR4PbNhgzYsnnGCVu6wsqGjerCn1qhrRv8qyZbBz\nZ8ttr6bXXrPmxfnz7X0/+GDT5qqUDsjttslOcw6+MlKkJdX3O/gr4F5gKXAt8CfgYmATsBI4pcWj\nU1OjtLDycpvr8JprrNJVNQn2X/7SfqtEwZSebmOH3XyzXSE6Y4YN4hqqscRyciA21te8uH27DUTr\n1EA5Up/CQhu/5fvv7RfLJZfYFS46cKSZmtvH63us6lXl18BTwJXAEyjxEhFg40YYP95uT52qAVyl\nDfj3v21Qvx49LPHatct+MVx8cbgjkzauuSPXewD/SUsWApcBc4F2fJ2UiDSG//yQTZ0rUiSkvv/e\nxiFxOKzKFRVlk7eKhEB9idd/gWNrPPY9cA6gKdZFhO++s+bFqVNtEvFHH4XVq8MdlUgDjjoKiors\ntsdjHSFVqpUQae29WNTUKK2XxwMffWS9yw87DIYP73BjGeTnW5XriCPs/k8/wSGHWDFBpNXKyYHr\nrrMrQVwuGDwY/vvf9jvSsISMxvESaUkzZ8JTT9lop5WVcMwx8Mwz1ttbpJ3YkLOBo7sdjdPRzjqe\n799vvxSioqzapY71EgTN7eMlInWprLQkq3t3++vZEzZtsrY3kXZiR8EOxr45lsXb2ugYK/WJiYFj\nj7W5uZR0SQg1dLRFAONCEYhIm+J2W1Nj1Qnb4bC/ysrwxiUSRM+sfIb8/fk8tvQx3B53uMMRaRca\nSrxcwKhQBCLSpkRHw9ChsHs37NtnI5V26QInntjwsiJtwI6CHbyX/h79uvRjR8GO9ln1EgmDQOqr\nS4DpwFnAqX5/Ih3bnXfayKu9esGQIdb02MLzDW3daoO+VklPb9HNSQf2zMpncOAgwhlBfHS8ql4i\nQRJI5/pF2JheNf06uKHUSp3rRfw8+qhdSThxInz6qU2Z8+ijHe5iSmlhZZVlXDD3AvaV76vqLIwD\nB3MuncOAQwaEOTqR1ktXNYq0M5WVNl7WkiWQnGxXwPfoEe6opD0qLi+mwu2b6NOBg6TYpHqWEJFg\nXdXYBfgfsML79zDVR7QXkRCJjLQLscCqXF27hjceab8SohPoEtvlwJ+SruDzeDwsWLuA0orScIci\nIRRI4vUMUAgMB/4IFAHPBmn7Q4H1QDowIUjrFGnVSkvhoYdsnl6wflpPPhnYsu+/b1PMzZwJffrA\nAw9U7/NVn6oLMf3vi0j4pGWmce/ie3ln4zvhDkVCKJDE60jgLmAz8BMw2ftYc0VgnfaHYlMTjQTU\neUDavdhYG/rrjjts2K977oGTTgpsWacT7rvP+vOPHw8DBlRPpuozbx7MmWOvLyiAW2+Fbdua/DZE\npBk8Hg/T06YTGxnLrOWzVPXqQAJJvEqxKxqrnAmUBGHbg4BNwFagAlgAXBKE9Yq0ag4HjBljF0De\ndRdcdRWcfnpgyw4d6uvTFRkJf/yjjQMZiEsusXkUH3vMkr6BAyFV092LhEVaZhrrc9bTs1NPCvcX\nqurVgQSSeF0PzAC2ef+mex9rrl7ADr/7Gd7HRNq9TZtgyxZLfN55x9fs2JISE+Ef/7CrIbduhdGj\nLQkUkdCqqna5cVNaWUpURBRPLH9CVa8OIpDEqxA40e/vZKyfV3PpckXpkEpL4f774W9/g+nTrfL0\nyCMtv92CAnjwQfjd72xS66pmRxEJrdLKUqIjounbpS9dYrvQM7EnvRJ7kV2SHe7QJAQiA3jNa8Ap\nQIHfY68AP2/mtjOB3n73e2NVr2omT5584PaQIUMYMmRIMzcrEl5xcTB1KiR5LxIbMyY0Fa/33rMm\nzb8X/IsAACAASURBVNGjbbD9u++GHTvU3CgSavFR8cy+ZHa4w5AgWLRoEYsWLWrUMvU1NAzAOr1P\nAf7ufa0H6Az8AziuSVH6RAIbgHOBnUAa1sH+R7/XaBwvkSBxu31TSlbd19zAIiLBE8g4XvVVvPoD\nF2Fjdl3k93gR8OfmBgdUAn8DPsKucJxN9aRLpE65udY5vSpx2LsXunULb0ytXc0kS0lXO1NebmOT\nREbCUUdBRES4IxKRWgTStfYM4JuWDqQOqnhJraZMsWEZ/vpXWLECpk2DGTOsA7lIh5ObC9dfD9u3\nWylz0CCb4iDQS15FJCiCNXL9Ddjo9VWSsUFVRcLmb3+DzEybo3rqVPjXv5R0SQf22GN2mewhh9gg\ncd98A6+8Eu6oRKQWgSReJwL5fvfzgFNbJhyRwMTFwbBh9kO/a1c4+uhwRxQ8paX2vqrs2QMVFXW/\nXoRNmyAhwW47HBAVBZs3hzcmEalVIImXA/CfEa4r1idLJGyWLbNpc/79b/u+mTEjPFPg5ObamFhV\nNm6EomYOtpKWBhMnWr+1zEyYMAHWrGneOltKzZ4A6hkQJieeaJeqejz2H6Giwjepp4i0KoEkXg9j\nfbzuBf7tvT2lJYMSaUh6Otx5p021c9dd9lhJMOZTaKSffoJJk6yVZ906m/5nx46Gl6vP2WfDeefB\n2LFwww02/MOprbDG7PHAvffC8uV2f9s2+PvfA587Uvy43ZCd3fSs/a9/tQHhsrLsb9gwuPTS4MYo\nIkER6LjVxwG/9t7+HPihZcI5iDrXS6u3ZAn85z92+9574eSTm7/OzEzrKw020GlrvWJzwwZ7z7//\nPbz1Fvzf/8GvfhXuqNqY3FwYNw7Wr7ds9uqr4cYbGz+tgNttSVdkpB0wmpZAJOSC1bkerHmxGJsu\nKBvo16zIRNqR5GTf7apBUZsjM9MuFrj5Zqt63X67NTu2RsccY3NNzpkDp5yipKtJ/vMf+OEH6xjf\nrRs8+yx8+WXj1+N02kSeKSlKukRasUASr8nAP4GJ3vvRwNyWCkikLfnhB3jgAav6TJhgzZ7+fb6a\nYu9euPJK+O1v4Q9/gAsvhPz8hpcLh23bYO5cm4B7xQpfs6M0wurVNiidw2HVKo/H2tJFpF0KZMqg\n32NTBq3w3s8EdOG+CNCpk008fdJJdj8qyq64bI4TT6x+/+KLm7e+luLxwFNP+ZoXzzrLRjU48USI\njg53dK2cx2OzlX/5pfXrKiuDww7zXSHSq1d44xORFhNIPToNGASsxBKwBKyD/Yn1LRQk6uMl0oq5\nXNUHSK95X+owb54NcBodbVcj7tkDffpY1WvIEJtFPTKQ38Ui0po0d8qgKq8As7BBVP8C/Al4urnB\niUjbVzPJUtIVoKeftgHoYmOtT1ZUlHWWO+88G5RO8zmJtFuBJF5TgPOwORr7A3cCn7RkUCIi7ZrL\nZclWFacT+va1qxVEpF0L5GfVeGAd8Hfvn5IuEZHmuOIKG7ersNCGgOjcGU4/Pbwx5efbFSLnngtj\nxthYISISdIH08ZoMDMemClqANT3uacGY/KmPl4i0P243zJ8PCxdaU+P111vFK5yuu84uTe3Wzfqd\nRUfDq6+23kHkRFqhQPp4NWawl5OAPwKXAxnAuU2OLHBKvEREWlpxsU2Z0KOHbwyw7Gx46CE488zw\nxibShgRzAFWALGA3sBc4pOlhiYhIqxITY1dRVs3G7vFYP7T4+PDGJdIOBZJ43QgsAj4DUoD/IzRD\nSYiISChERtq0RXv3ws6dsGsX/PKXvgHqRCRoAmlqfAB4CVjVwrHURk2N0iG43b4RBKoOec36IiG3\nYgX8+KNNX3TuuRpLTKSRgt3HKxyUeEm7t3YtvPyyzc8YFQUzZ0K/fnDBBeGOTEREGkOJl0gb4HbD\n1KmQm2sXkO3aBXff3fyph0REJLSUeIm0ES4XXHqp3X7hBZszWURE2hYlXiJtgNttzYtbtljCVVoK\nd95pF5qJiEjbEezhJESkBaxbB9u2wT33wMSJNoXfp5+GOyoREWkJqniJtAIul2+CabfbrmjUVY0i\nIm2LmhpFRKRtcLvtyhKXCw4/3De+ikgbEkjipUFaREQkvMrLbYLur76yUu/JJ8Mjj0BCQrgjEwk6\n/aQQkdByueD772HZMigqCnc00hrMmweLF0P37jZ464oV8NRT4Y5KpEWo4iUioVNRAePHw7ffWlNS\nUpJ9waamhjsyCaacHFiwwAanO/ts+6vP+vV2GW9Vx8b4eBtBX6QdUsVLRELnww9hyRJfZaOwEB58\nMNxRSTDl58PVV8Ozz8L771ui/frr9S9z9NGwf7/Nl+XxQEkJHHNMaOIVCTFVvEQkdDIzrdJVVdlI\nSIDt28Mb0/9v787D4yzLxY9/k7SBpFu6ktKCFBDqwtofHjZttYissvxYBPQcKLIcWSxiRUSh4FFc\nWBXwICJiQUEKqAgiBQmLgGWvIK0sXWhp2ibd27RpMnP+uDNOW0rX5H1nMt/Pdc01eSdvZu5Mlrnn\nee7nftS+nnwS5syBbbeN46amaFR37LFrnjdpUvzsBw2Ck0+Gl1+GiRPjd2P33eGMM5KPXUqAiZek\n5Hz0ozGi0dIS/TMWLnRTys6mtTW/0ztEot3SsuY5t90GN90USVY2C6eeCj/5CUybFscf+pAbdKvT\ncqpRUnI++Uk46yxobIxRkWHDYMyYtKNSe9p3X+jeHebNi6nkhgY47rj85+fPjxGwvn1hm22gXz+4\n/Xaor4cdd4SddjLpUqdmHy9JW27hQnjzzXjBHTp0w91fm5qipqdXLzvFdkZvvRUjWo2NMHIknHJK\nvkPwtGnwhS9EjV9OQ0OMgg0dmkq4UnuxgaqkjjdlCvz3f0cy1doKhxwCY8faAFPr1twc9V6NjbE/\n1oIFkbDff799u1T03KtRUmhpgX/8A154AZYtW++pTU35j7NZWLFiA/d92WUxetWvX4xiPPRQrFyU\n1qWyEm64AYYMgblzo0v9TTeZdKlkOJEudXbNzTB6dCRd5eVRW3PLLflVZ6tpbIzV/2PHRn3zrbdG\nInbeeeu5/5kzoWfP+Dg3yjVvXrt/G+pEdtghmqZms041q+Q44iV1dn/4QzQszfXOamiAq65a56l9\n+8KXvwyXXgrf+Q78859w2mkbuP/dd4+MLZuNJK+sLAqkpQ0x6VIJMvGSOruZM2OVWO5Frnt3mD79\nA08/4IAYuHr1VRg1Kk5fr8svjwaYc+dGkf0FF8Ree6WkpSWKxufMWbOVgiStxalGqbPbbTe44441\ne2d9+tPrPDWbjenFPn1i4dmPfgRXXBEzQx+of38YNy6KpKuroaqqQ76NgjVvHpx7biSzra1ROH7R\nRS4uKAZz5sQbk4ED1zn1LnUE/zNInd3IkbGFS0NDvNDsu2/UfK3DggXRXP6KK2Jx4plnwuOPb8Rj\n5GrHSi3pgtjyaOrUSED794fx4zfySVOqHnsMjj46kuZjjoF77007IpWIQp9gt52E1F6WLYtNqu2d\n1b4OPzxq27beOo7fey8y1rPOSjcufbBly+Dgg+ONQlVV/PwWLYp6yG22STs6FTHbSUjK69YNampM\nutrbzjvHizZAJhPXQ4akF482rLExfla5EdrKyvi7mDs33bhUEky8JGlLfOtb0Ytq3ry4HHkkHHRQ\n2lEVhmnT4KWXYpugQjJgQCRdixfH8bJlUf84aFC6cakkFPpbX6caJRW+lSujuH7rrWG77RxVBLj5\n5lipUVERq2qvvx723jvtqPImTYoVuEuXxs/tyith//3TjkpFzi2DJGlzrVoFd94JL74I228fDc56\n9047quLwxhuxoKNv30i6liyJ6wkTCispbW7Ob1201VZpR6NOYGMSL9tJSNK6fO978MAD0SLjuedg\n4kT49a9Lc+Xmpqqvj5WuXdpeYnr0gNmzYfnywtoaqLIyWklICbLGS5LWtmwZ/OlPUFsbCxJqa2HG\nDHjttbQjKw5Dhqy50WdjY9TBVVenG5dUAEy8JGlthTQdVox22CH2nVq2LFYK9ukD11zj8yphjZck\nrdt3vwv33x+jNCtWxP6Tt9+e79elDWtqipWDuVovqZOzuF6SNldLC/z2t9EOYfvtY+PKXr3SjkpS\nATPxkpS++fNjI+2JE6NW6rLLSm8TbW2+TMZ9L1U07FwvKX3f/CY880y0YmhsjL3x6uvTjkqF7pVX\nYjumT3wiWlPMnp12RFK7MPGS1HFWrIipum22iUaavXrFFN4bb6Qd2ebLZuH3v4dzzomu9e+8k3ZE\nxW3lSmhtXfO2hgY477xoblpbC5Mnx8buuS2ZpCJm4iWp41RWRmPKlSvjOJuNF9mePdONa0vccUcU\n3k+aBI89FrVfjsZsuiVLIrk64AA48EAYPz7/ubfeigS9Z89YCTlgAEydmt8TUypiJl6SOk55eYwK\nLVoU04v19TBiBOy1V9qRbb4774xp05qaSAiWLIEnn0w7quLzgx/As8/GiFbPnnH84ovxuV69IkHP\njXCtXBkjpoXUfFXaTK7vldSxDj8cdtwxpov69o0RjmIulq6oiO2EcrLZuE2bZuLE+H0oK4uR0WwW\n/vlPGDYMhg6FY46B++7L9/76znfiPKnImXhJ6ngf+UhcOoNTT43RmaamSMD69oXhw9OOqvjU1kZ9\nXJ8+kXQB9OsX12VlsSjjoINg3jzYeWfYZZf0YpXake0kpM4ok4G//AWmTInRpsMOyzewbG6G666D\nhx6KZqCjR8Mhh6QbbzHJZuHRR6O+q3dv+OIXYdCgtKMqPpMnw9lnxwKM1tZYvXjdddC1a9qRSZvN\nPl5Sqfr+96NYuaIiXtQOOihGacrL4Sc/gV/9Cvr3jxGbJUvg5z9Pr+4qk4GnnoI5c+DDHy7u+i9t\nmoaG2P+yuhr23tvu9ip6G5N4+VsudTYNDbHVTW1tJF6ZDNTVwbRpMfr12GMxvVNZGZfFi+Hvf+/Y\nhCebjcLp6dNh4EDYb7+YTspkYk+/hx+Oc8rL4YIL4OSTOy4WFY5+/WKxhVRCTLykzmblykhgcgXs\nuY9zLR1yjUyrquI4m+34rXBuuSVG1XJOPBHGjInppkceiT5f5eUxDXr99VFYnYtPkjqRIl5aJGmd\namth112jdcPy5TGFt+22MdoF8LWvRbI1e3ZchgyBI47ouHjmz4dbb42pzYEDowXDPffAjBnRILOi\nIp8kdu0asTU1dVw8kpQiR7ykzqaiIkaNrr466meGDYOvfz0amQLsvnts/vzCC1FcP3x4x/ZHWro0\nrnP1OxUVcVmyJFaqVVVFctazZ4zE7bJL9MiSpE7I4npJHWvVKjj++BiB6907mqn27An33hsJ35Qp\nsYn2zJmRFI4dm28rIElFxFWNUjFobo72BI2NsNtusOeeaUfU/mbNiiL6N96AnXaKRCs39anOZfZs\n+OlP43r//aPvmS0iVCJMvKRCt2oVnHtuTPtB1Dp9+9tw1FGbd3/NzfDqq3G9227FvSeiis+iRfCF\nL+QXbyxdCsceC5dcknZkUiJMvKRC97e/wVe/GgXxZWXRTLKpKfpalW3in2dTE3zlK/D665HA1dTA\nL34Bgwd3TOzS2urq4KKLYgEFRA+5hgZ45hl7dKkkbEzi5apGKU3Ll0eheS7JqqyMtg+trZt+X/fd\nF6NdAwbECsKFC+Haa9s3Xml9Kiry2/9AfFxWtulvIqROzLcgUpp22y2SrQULotC8sTFWGW7O6MB7\n78XX5V7kqqvh3XfbN15pfYYNgw99CKZOjbqu5mY4/XQ3EZdW44iXlKbaWvjZz2D77WOU69BDo/B8\nc+y1V9xHS0t0hF+0KPa/k5JSXR3T26efDp/9bKxQPfvstKOSCkqhj/9a4yVtrGwWbr4ZfvnLSLw+\n/Wm44go7wBeT3P87p+akomRxvVSKmptj5MuEq7jcfTfccEP8/I46Kpre5mr+Hn44itR33x322Sft\nSCV9ADfJlkpRZWXaEWhTPf00/OhH0Lcv9OgB48fH/plnngnnnAMvvZQvUr/oomhIK6koWeMlSWl7\n7rkoQN9qq7ju1QuefBKefz5Wqm67bVz69ImVqplM2hFL2kwmXpLUHiZPjqLyz38efvjD92/0vWpV\nJFEvvvj+z/Xvv2YLkRUroi1IU1P0ZMvVfOVWCpp4SUXLqUZJ2lJz5sBZZ0VyVV0Nv/tdbAL+P/8T\nn1++PHYoeO21SKS23RZ+/vP8npTHHgsPPhhtGMrKorXI6NGx80BVVbQZ6dYtNhP/zGdsRioVMYvr\nJWlLPfxw7EW5zTZxnOvY/txzkWj98pdw440wcGAkVnPmwGGHRbuFnOXL4dlnI3nbe+989/cpU+AH\nP4iv2W8/+NrXIgmTVHAsrpekJGy9dbSCyHVqX7UqbstNEU6bFosecsdVVTB9+pr3UV0NI0e+/753\n3RVuu61Dw5eUHGu8JGlL7bcffOQjsXvA7NmxE8Ho0flEa4898ltBZTKxefRee6Ubs6RUONUoSe2h\nqSnqtBoaYqpw9V0DWlujXcR998Xx8OHw3e/aa03qZGygKkmFZMmSSMJ69bI7vdQJmXhJUhLeeisK\n46uq4KCDoKYm7YgkpaDQE68fA0cAzcDbwGnAorXOMfGSVNhefDFaRTQ3R3H94MFw++3Qu3fakUlK\n2MYkXmkW1z8CfAzYA/gXcHGKsUjS5rn++ug2v+22MGgQzJwZtV6StA5pJl4TgFz75b8Dg1OMRZI2\nz+LFsdVPTnl51HK1p9ZWmDs3en1JKmqF0k5iFPBQ2kFIBe/552Na6ytfgSeeSO5x586FCy6AQw6B\n886LlgkKhx0W7SNWrIiEq7wcDjhg/V+TyUBdHdx1V/xM12fWLDjhBDjyyOhaf++96z93zBg46SS4\n7rqISVJB6egarwlA7Tpu/xbwQNvHlwB7A/9/HedlL7vssn8fjBgxghEjRrRziFIByWTgX/+KeqEP\nf3jNdgMvvxzb0nTtGiviVq6Eq66K1gQdqaUFTj45moDW1MCiRdGB/a67okloMclm4f77oyFpNgtf\n+lIkNVuywrClBW6+Gf70p2iCev756/+ZZLPRsf7BB+PnXV4eifSoUes+/0tfgjffjP0cm5sjybv9\n9ugbtrrFi+HEE2NboerqOD74YLjyys3/3iStV11dHXV1df8+vvzyy6GAi+sBTgXOAEYC63prZnG9\nSkdzc4xWPPtsvBjX1sYLem4bmrFj4c9/zm8ls2BBNOa88caOjWv69HhB798/f9u8efHiv+uuHfvY\n7W3CBLj44tgDsawMFi6EK66Aww9PLoY334RTTonns7w8utzPnw9//St0777muZlM9AOrrc0nh3Pm\nwGWXvT/mp5+GCy/M/35kMnHu008XX4IsFalCL64/BBgDHMW6ky6ptPzxj/DUU/HC2b9/TOddfXX+\n8126xGhJTiYTo18draoqaoxaW/OPm8kUZ/PPRx6JrXu6dYtRoaqquC1JS5dGMX5527/f3IbX06bF\naGdTU/7c3Ibai9oWfLe2xu9ALhlfXdeu+W2LID+aVlHRYd+KpE2XZuL1U6A7MR35MnBTirFI6Zs+\nPV6EcyMb3brB22/nP3/88fHiWl8fIxmZTExDdbQBA2I6bu7c2BKnvj5GW7bbruMfu7316hUjTDnN\nzTH6laSdd46RrcbGiGXu3Lj9tNPgv/4Ljjkm+oLlfP/7kTw1NMRI4wknwLBh77/fvfaCoUPjZzR3\nblxOOy2Z5FzSRkt7qnFDnGpU6fjzn+Hb347RjPLySHCOPDKmlXKmTIni6pYWOPpo2H33ZGLLZqMY\nfOpU2H77KPIuL5S1OZtgxgw49dSYYoRIum67DYYMSTaOd96JqePp06NebvLkuO7SJRKyQYPgnnvy\n5y9YEF/TqxfstNMH16QtWwbjx0fytdde8LnP2SFfSlChN1DdGCZeKh2ZTBTLjx8fx3vsEVONSY/I\ndHb19fD445FMjhgRU3lpevBBuPzy/PRhrjZr4sTiTG6lEmbiJRWjhQtjxWKu+Fqd26RJcPrp0K9f\njHg1NMSo4t13px2ZpE1k4iWp8C1dGrVL/fu/f1VfIclmYwHEI4/ElN+Xvww77tg+933rrbGCtaIi\nWnbceGP73bekxJh4SSpsTz0V7R1aWqII/Mor4cADk3nsRYuixqqmJkaYNmTcuGhKWl0dRflVVfDb\n30ZtVntoaIiYBg1ad/uHXCF+jx5OP0sFysRLUuFavDi6vufaOyxbFsnFQw9FctGRXnstOvCvXBlJ\n36mnRhPT9Tn00Igv10bjvffgG9+IHmcdbcaM2LFg7twYeRs9OrrTSyoohd7HS1Ipq6+PvlTdusVx\nt26RBNXXd+zjZrORMLW2Qt++cbnttkjG1qe8fM0+atlscj2yvvnN/HRsTQ1ccw28/noyjy2pXZl4\nSUrHgAHR6iDXMLSpKZKbXOf1jtLSEqsGc9N1XbrE425o/8lRo2LhQ2NjJId9+8KnPtWxsUJ+G6m+\nfeO4sjKet2nTOv6xJbW7LmkHIKlE1dTEdj2XXhrTjGVl8L3vReF6R+raNQrXZ86MlYTNzTF6tcMO\n6/+6Y4+N2B59FHr3hi9+seOTRIikcLvtYsSrd+8Yqctk0m+DIWmzWOMlKV3z58cI1DbbQJ8+yTzm\ntGlRM9XQEMdf/zocd1wyj705Jk+Gc86JUcGWlkj6zjvP5qhSgbG4XpI+SG6VYK9ehd3GImfJktg5\nYGNXYea88kp0wc9mY7uhPffsuBilEmfiJUml7JVX4Oyz17ztf//X5EvqIK5qlKTNtWwZvPlmFNMX\nq7vuiunI/v3jUlYWt0lKjcX1krS2116D88+PmqpMJmrAjj8+7ag23bpmDJxFkFLliJckrS6TgQsv\njCL2fv2ipuqqq6K+Ki3LlsFLL0VC2Nq68V93wglxfkNDXDKZuE1SahzxkqTVLV0KCxbEKkuIvlnl\n5TBrFgwZknw8s2fDmWdGO4lMBvbZB669NuLakGHD4Gc/i62NILrdDxvWsfFKWi8TL0laXffu0S9r\n0aJY8djcHAnPoEHpxHP11fl2G9ksPPss/OEPGz/1OWyYyZZUQJxqlKTVlZdHstOlS0zPLVwIY8ak\nM9oF0XMs1+6irCzimjEjnVgkbTFHvCRpbR//ODzwQGyE3adPfrueNOy5J9x/P1RXx8hba2vEJ6ko\n2cdLkgrZkiWxqfcLL8TxSSfB6NExMgdReP+b38C770aSdvTR+c9JSpQNVCUVtnnzYipv0KAY0dG6\nZbNR8N+1K/Tokb+9uRnOOCNWO1ZWxvGJJ0aiJilxJl6SCte4cXDDDTE60707/PSnMHRo2lEVl5de\nis70AwZE/VdraySzTzxhIiulwM71kgrTlCmRaPXuHb2ympqigN03WpumtTUS19xm2bnrTCa9mCSt\nl8X1kpI3c2YkDF27xnFNTfSram6GrbZKN7Zi8rGPQW1t9Birro4eZCNHFsem31KJMvGSlLzBg2NU\nZtWqSL4WLoSBAzeuKajyqqvhllvgxhth+vTo13XGGWlHJWk9rPGSlI477ojpxrKyKBi/4QbYdde0\no5KkzWZxvaTC1tAQq/UGD4aqqrSjScbbb8MvfgGLF8PBB8PnP5+vzZJU1DYm8XKqUVJ6+vWLS6mY\nNQtGjYIVK2Ja9bnnog/XySenHZmkhLiqUZKS8sQT0RB1wIBYUFBTA3femXZUkhJk4iVJSamoWPM4\nm7XLvFRi/IuXpKSMGBF7P9bXQ2Nj1HmNGpV2VJISVOgVnRbXS+pcZs2Krv2LF0fPrZEj045IUjtx\nVaMkSVJC3DJIkiSpgJh4SZIkJcTES5IkKSEmXpIkSQkx8ZIkSUqIiZckSVJCTLwkSZISYuIlSZKU\nEBMvSZKkhJh4SZIkJcTES5IkKSEmXpIkSQkx8ZIkSUqIiZckSVJCuqQdgCSpQDQ2wrhxUF8P++8P\nRxwB5b4/l9qTiZckCZYsgVGjYNYsqKyECRNg9mw466y0I5M6Fd/KSJJg4kR47z0YOBD69oX+/eFX\nv4JsNu3IpE7FxEuSBJnMmsdlZZF0mXhJ7cqpRkkS7LNPjHTNmQNbbw1Ll8Ipp1jjJbWzsrQD2IBs\n1ndbkpSM996Dm2+O5Gv//SPxqqhIOyqpaJSVlcEGcisTL0mSpHawMYmXY8iSitfadUmSVOBMvCQV\nn+efh0MPhf/4DzjjDJg3L+2IJGmjONUoqbjMng3HHQddu0K3bpF0ffSj0fpAklLkVKOkzmfyZGht\nhR49YsXdgAHw2mvQ1JR2ZJK0QSZekopLTU0kXrn6rhUroKoKttoq3bgkaSOYeEkqLnvsAZ/7XLQ8\nmDs3trq59FL7TUkqCtZ4SSo+mUxscTN/PuyyC+y8c9oRSZJ9vCRJkpJicb0kSVIBMfGSJElKiImX\nJElSQky8JEmSEmLiJUmSlBATL0mSpISYeEmSJCXExEtSaclm4yJJKTDxklQaWlrgmmvggANg+HAY\nN84ETFLiTLwklYY77ohLz56w9dZw3XXw6KNpRyWpxJh4SSoNTzwBPXpA166w1VZQWQnPPJN2VJJK\njImXpNLQvz+sWJE/bm6Gfv3Si0dSSTLxklQazjknRrzq6+MyeDCcdFLaUUkqMevdQbsAZLMWv0pq\nLw0N8PzzUFEB++0XiZgktZOysjLYQG5l4iVJktQONibxcqpRkiQpISZekiRJCTHxkiRJSoiJlyRJ\nUkLSTrwuBDJAn5TjkCRJ6nBpJl7bAZ8FpqcYgyRJUmLSTLyuAb6R4uNLkiQlKq3E6yhgJjAppceX\nJElKXJcOvO8JQO06br8EuBg4eLXbPrDZ2NixY//98YgRIxgxYkT7RCdJkrQF6urqqKur26SvSaNz\n/ceBx4DlbceDgVnAJ4C5a51r53pJklQUimXLoKnAMGD+Oj5n4iVJkopCsWwZZGYlSZJKQiGMeK2P\nI16SJKkoFMuIlyRJUkkw8ZIkSUqIiZckSVJCTLwkSZISYuIlSZKUEBMvSZKkhJh4SZIkJcTE0598\n7gAABUlJREFUS5IkKSEmXpIkSQkx8ZIkSUqIiZckSVJCTLwkSZISYuIlSZKUEBMvSZKkhJh4SZIk\nJcTES5IkKSEmXpIkSQkx8ZIkSUqIiZckSVJCTLwkSZISYuIlSZKUEBMvSZKkhJh4SZIkJcTES5Ik\nKSEmXpIkSQkx8ZIkSUqIiZckSVJCTLwkSZISYuIlSZKUEBMvSZKkhJh4SZIkJcTES5IkKSEmXpIk\nSQkx8ZIkSUqIiZckSVJCTLwkSZISYuKlNdTV1aUdQsnxOU+ez3nyfM6T53NemEy8tAb/UJPnc548\nn/Pk+Zwnz+e8MJl4SZIkJcTES5IkKSFlaQewAa8Ae6QdhCRJ0kZ4Fdgz7SAkSZIkSZIkSZIkSZIk\nSZ3becAbwGvAD1OOpZRcCGSAPmkHUgJ+TPyOvwrcB/RKN5xO7RBgMvAmcFHKsZSC7YDHgdeJ/+Hn\npxtOSakAXgYeSDuQElEDjCf+l/8T2DfdcDbfp4EJQNe24/4pxlJKtgMeBqZi4pWEz5Jv7/KDtova\nXwXwFrAD8T/lFeAjaQZUAmrJr/LqDkzB5zwpXwPuBP6YdiAl4nZgVNvHXSjiN9C/Az6TdhAl6B5g\nd0y80nAMcEfaQXRS+xFvKHK+2XZRcn4PjEw7iBIwGHiUGLxwxKvj9QLe2ZgTi6GB6oeBTwHPAXXA\n/0s1mtJwFDATmJR2ICVqFPBQ2kF0UoOAd1c7ntl2m5KxA7AX8PeU4ygF1wJjiHIRdbwhwDzgNuAl\n4Bagel0ndkkwqPWZQAxHr+0SIsbexFzpPsQI2I7JhdZpre85vxg4eLXbCr3RbrH4oOf8W+TfkV4C\nNAO/SSqoEpNNO4AS1p2of/kqsDTlWDq7I4C5RH3XiHRDKRldgL2Bc4HngeuI0fRL0wxqc/0ZGL7a\n8VtA35RiKQUfB+YQU4xTgVXANGBAijGVilOBvwFbpxxHZ7Yva041XowF9knoCvwFGJ12ICXi+8TI\n7lRgNrAM+HWqEXV+tcTznXMg8KeUYtliZwGXt328CzAjxVhKkTVeyTiEWPXVL+1AOrkuwNvElFcl\nFtcnoYx40b827UBK1HCs8UrKk0SeAjCWIu7C0BUYB/wDeBGHTZP2DiZeSXgTmE5MDbwM3JRuOJ3a\nocTKureIES91rAOJOqNXyP9+H5JqRKVlOK5qTMoexDSjbYEkSZIkSZIkSZIkSZIkSZIkSZIkSZIk\nSZIkSZ3RUWx5w9ShwLPACuDCLY5IUkmpSDsASUrQxcAC4I1N+JoK1tzjsQx4BphPJF/Ptlt0kjq9\n8rQDkKSNcCXwldWOx5IfbRoDTCS6RY9d7Zz/bLvtFWLLmv2AI4EfE93TdwT2BJ4j32m6pu1r64gt\nbp4Hzl8rlnnAC8Q+ppIkSZ3OnkQylPM6MAg4GLi57bZyYk+6TwIfI7YFym13lUuobgOOXe1+JrWd\nD7EnbG4/wceBGzYQ02U41ShpE3VJOwBJ2givAAOAgW3XC4BZwAVE8vVy23ndgJ3brn9HTAcCLFzt\nvsrarnu1XZ5qO74duGe18+5u1+9AkjDxklQ87gGOA2qBu1a7/Urg52udey75BGtt2Q+4fe3zl21q\ngJK0IdZ4SSoWdwMnEclXbmTqL8AoYoQLYvqxP/BX4HjyU429266XAD3bPl5EjJwd2Hb8JdacztyQ\nD0rsJEmSOoVJwGNr3XZ+2+2TgL8BQ9pu/0/gH8Q05S/bbtufqA97kSiu34NYlZgrru/Vdt7jwN4f\nEEMt8C75xG0G0H0LvidJkiRJkiRJkiRJkiRJkiRJkiRJkiRJkiRJkiRJkrQe/we9FR4c7KQSlAAA\nAABJRU5ErkJggg==\n", + "text": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 4, + "metadata": {}, + "source": [ + "PCA for feature extraction" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "As mentioned in the short introduction above (and in more detail in my separate [PCA article](http://sebastianraschka.com/Articles/2014_pca_step_by_step.html)), PCA is commonly used in the field of pattern classification for feature selection (or dimensionality reduction). \n", + "By default, the transformed data will be ordered by the components with the maximum variance (in descending order). \n", + "\n", + "In the example above, I only kept the top 2 components (the 2 components with the maximum variance along the axes): The sample space of projected onto a 2-dimensional subspace, which was basically sufficient for plotting the data onto a 2D scatter plot.\n", + "\n", + "However, if we want to use PCA for feature selection, we probably don't want to reduce the dimensionality that drastically. By default, the `PCA` function (`PCA(n_components=None)`) keeps all the components in ranked order. So we could basically either set the number `n_components` to a smaller size then the input dataset, or we could extract the top **n** components later from the returned NumPy array.\n", + "\n", + "To get an idea about how well each components (relatively) \"explains\" the variance, we can use `explained_variance_ratio_` instant method, which also confirms that the components are ordered from most explanatory to least explanatory (the ratios sum up to 1.0)." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "sklearn_pca = PCA(n_components=None)\n", + "sklearn_transf = sklearn_pca.fit_transform(X_train)\n", + "sklearn_pca.explained_variance_ratio_" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "pyout", + "prompt_number": 13, + "text": [ + "array([0.36, 0.21, 0.10, 0.08, 0.06, 0.05, 0.04, 0.03, 0.02, 0.02, 0.01,\n", + " 0.01, 0.01])" + ] + } + ], + "prompt_number": 13 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "" + ] + }, + { + "cell_type": "heading", + "level": 2, + "metadata": {}, + "source": [ + "Linear Transformation: Linear Discriminant Analysis (MDA)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top]](#Sections)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "The main purposes of a Linear Discriminant Analysis (LDA) is to analyze the data to identify patterns to project it onto a subspace that yields a better separation of the classes. Also, the dimensionality of the dataset shall be reduced with minimal loss of information.\n", + "\n", + "**The approach is very similar to a Principal Component Analysis (PCA), but in addition to finding the component axes that maximize the variance of our data, we are additionally interested in the axes that maximize the separation of our classes (e.g., in a supervised pattern classification problem)**\n", + "\n", + "Here, our desired outcome of the Linear discriminant analysis is to project a feature space (our dataset consisting of n d-dimensional samples) onto a smaller subspace that represents our data \"well\" and has a good class separation. A possible application would be a pattern classification task, where we want to reduce the computational costs and the error of parameter estimation by reducing the number of dimensions of our feature space by extracting a subspace that describes our data \"best\"." + ] + }, + { + "cell_type": "heading", + "level": 4, + "metadata": {}, + "source": [ + "Principal Component Analysis (PCA) Vs. Linear Discriminant Analysis (LDA)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Both Linear Discriminant Analysis and Principal Component Analysis are linear transformation methods and closely related to each other. In PCA, we are interested to find the directions (components) that maximize the variance in our dataset, where in LDA, we are additionally interested to find the directions that maximize the separation (or discrimination) between different classes (for example, in pattern classification problems where our dataset consists of multiple classes. In contrast two PCA, which ignores the class labels).\n", + "\n", + "**In other words, via PCA, we are projecting the entire set of data (without class labels) onto a different subspace, and in LDA, we are trying to determine a suitable subspace to distinguish between patterns that belong to different classes. Or, roughly speaking in PCA we are trying to find the axes with maximum variances where the data is most spread (within a class, since PCA treats the whole data set as one class), and in LDA we are additionally maximizing the spread between classes.**\n", + "\n", + "In typical pattern recognition problems, a PCA is often followed by an LDA." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "If you are interested, you can find more information about the LDA in my IPython notebook \n", + "[Stepping through a Linear Discriminant Analysis - using Python's NumPy and matplotlib](http://nbviewer.ipython.org/github/rasbt/pattern_classification/blob/master/dimensionality_reduction/projection/minear_discriminant_analysis.ipynb?create=1)." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Like we did in the PCA section above, we will use a `scikit-learn` funcion, [`sklearn.lda.LDA`](http://scikit-learn.org/stable/modules/generated/sklearn.lda.LDA.html) in order to transform our training data onto 2 dimensional subspace:" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "from sklearn.lda import LDA\n", + "sklearn_lda = LDA(n_components=2)\n", + "transf_lda = sklearn_lda.fit_transform(X_train, y_train)\n", + "\n", + "plt.figure(figsize=(10,8))\n", + "\n", + "for label,marker,color in zip(\n", + " range(1,4),('x', 'o', '^'),('blue', 'red', 'green')):\n", + "\n", + "\n", + " plt.scatter(x=transf_lda[:,0][y_train == label],\n", + " y=transf_lda[:,1][y_train == label], \n", + " marker=marker, \n", + " color=color,\n", + " alpha=0.7, \n", + " label='class {}'.format(label)\n", + " )\n", + "\n", + "plt.xlabel('vector 1')\n", + "plt.ylabel('vector 2')\n", + "\n", + "plt.legend()\n", + "plt.title('Most significant singular vectors after linear transformation via LDA')\n", + "\n", + "plt.show()" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "display_data", + "png": "iVBORw0KGgoAAAANSUhEUgAAAmgAAAH4CAYAAAD+YRGXAAAABHNCSVQICAgIfAhkiAAAAAlwSFlz\nAAALEgAACxIB0t1+/AAAIABJREFUeJzs3Xl4lPXV//H3JCErIQQCBJGwqCi4ohie0qrRPlW0Wlof\nEBVxrxUtUtSfuAIutCq4ggsuLSBFEEXrjktBQBQoCioUREB2QkJCErJnZn5/nBlmErJMQiYzST6v\n65ors973mXsmM2fOdwMRERERERERERERERERERERERERERERERERERERERERERERCbICoGcjbGcE\nsNDv8i+BTUA+MAT4ELi6EfYTLB8CI5tgPxOB15pgPy2F931UAPwuSPtwAb09518A7g/SfpqTOOA9\n4AAwL8SxVKfq501TSMPeh44m3q+IBNHPQCnQscr132JfDmlHuH3/L5hw8Tkwuon3uRi4oYn3WV8T\nCP8EbTHhcxyrvo+C8V4Px/8fsM+N80K075HACiAiRPv31xN7jcIhloa6Flhaw22LgWLsx2we8B9g\nHBBdzX1nAOVAamMHKIdrzm84CZwb2AJc4XfdydivVHcj7SPcftWlAeubeJ+NdSyD6Uhep6b6vDiS\n4+igcd+L1b2PGrr9qCOMJVhqOmbuGq73Cubz6QH8iCVG9RWsuMLtM66xuIFbgXZY4nUHcDlW1feX\nAPwf9v9wVVMGKNKSbQXuA1b6XTcFuJfKFbQkYBawD/v1fB++D6VjgS+wJocs4HXP9Us82ziIld+H\nVbP/qo+d63ebf/WgI9askeeJ9REq/+pzAX/CPrhzgWl+t13rd9/NgBMown4VRnN4VeaP2AdNPrAO\n6O+5/m7gJ7/rf19lH8uAyUAOlvQO9tw2CajAfokWAM9WcxxigdlAtif+lUAnz23+8dW2H4Be2HHP\nBz4FnsNXFcsAdlTZ78/4KiETqVxBmw/swV6bL4B+frfNwJrcPsRe36rVlOHAqirXjQX+5Tkfg73P\ntgF7PduK9bvvEGAN9nr/BFxAzcdxkGdfB7Dj9gu/7SzG3itfYq/5Mdgx3Iwdoy3AlVQvHfgKez12\nA1OBNp7b/N9HBcByqn+vX+x5HrmeGE722/7PwF3Ad57nVF2S6/8/MAN42HM+A9gJ3A5keuK71u9x\ntR3f9sD72P9yDvZ/1c3vsYupfMyqVvBeq/Lc78RXSbres8/FnvvW9R56zhNLPvB1lX095Xluedgx\nOhF4EKv4l3n2fR32OXQ/djwzgZlYQkE1cX0BXON5bk9ir8tP2HvoOmC7Zxv+XR5+i7Uo5Hlun+B3\n23bP9gs8z+F/OLwiVdf78yHsfzofaxqt2prh9V9PLF5R2GfmaRxeybsO32fYZuCmGrZJNfH6W8Th\nFevuQGGVWK7GXqMRwPe17EtE6mEr8GtgA3ACEIl9iadROUGbBbyN/VLqAWzEPvTAErJ7POejsQ8k\nr7qaaAJ97FxgDvYl0xf7YFxS5b7vYh/M3bEvnws8t11L5Q+grVROKBb5PZdh2BffGZ7Lx+A7BkPx\nle8vw76Mu/jtowz7MHMANwO7athHdf7kiT/W8/j+QGI1j61rP18Bj2Mf3r/EvlRmeW7L4PAEzf9Y\nTKRygnYt9nq3wb4sv/W7bQb2heP9sompst047MvhWL/rVmHHDc/23sGShbae5/5Xz23pnm3/2nP5\nKOB4z/mqx7ED9iU7AvtyuhxLOpI9ty/Gvrj7em5Pwo7JcZ7bu1A5afB3uieWCOw9vx4Y43d71fdR\n1fd6f+zL/kzstbra8xhvkvcz8A2WHFU9ftVt8x/YlznYa1mOvWaRwIXYl2aS5/bajm8H4A/Ye60t\n8Ab2v+21mMrHrLqqU9Xn3tMT6wzstfc+n2up/T2UDQzwPIfZ+H7cXYA1p3kTrePx/e9NwPeeBns/\nbPLEkAC85Xd71bhiPTGVY4maA0t6d+JLwH+DvXfjPds4B0sOwRLsvdgPCLD3RdUmzmvxfd4E8v7c\nhP2fxGLv779RvQewY+T1W+yHov/z9MZxEfZjDeBs7L3Rn+r5x1tVTZ9bXwCP+l3+HPvRnoj92Di9\nhu2JSD14E7T7sA/wwdivuEh8CVok9qv1BL/H3YT984L9Yp1O5V/hXnUlaIE8NhJLSo7zu+1hDq+g\n+Sd387C+ElC/BG0hgfdP+xZf5/BrsQ9ar3hPTJ399lFb36nrOLzCUl18te0nDfvi8a9EvUbDEzR/\n7T378SaNMzyn2ryGfamAvXb5+BLQg1R+X/wCq2aBvR+eqGGbVY/jSKzy4m859uXrvf9Ev9sSsC/M\nS7Ev7Pr4C7DA73JdCdoL+BIqrw3AWX6Pv7aOfVZN0PwraEVUTgwysYSyruNb1WlY0uBV9ZhVp6YE\nrWctj6n6HvoH8JLf7RdiVSI8294IDOTwyuJEKr9PP8d+qHj1wT4vImqI61qs0u51suc+nfyuywZO\nqeF5PI1V36D6PmjX4vu8CeT9ea/fbaOAj2rY7zH4/ocA/olv0Eh1cfh7G7ithtv8462qpgTtdXyv\nXRpWUe3jufwOdowkiNQHrfVwYx94I7APjllU7lORgv2y3OZ33XZ8SdVdnvuvBH7Ako1ABfLYTtiv\neP/kYmc199vrd74I+zKur6OxJoHqXI0lZbme00lUbo6oun+wCoVXbf2nXsOSw7lYRewxau4vU9N+\njsK+aEv8bt9Bw/rHRGK/kH/CKk5bPdeneP66OTzZq2oOvr6NV2JfEiXY6xkPrMZ3LD/y23Ztr4F3\n315HYe9Ff9s813v5x1mINb/ejDULvo+vOldVH8/te7BjMImam5+q0wPrs5Prdzq6ltjqaz+V+2EV\nYe+Duo5vPJYE/4w9ry+wypv/+6Shcfk/LoLa30NgSaVXMb7/l39j3RSe89xnOr7ErqquHP7ZFIWv\nul01rur2C9ZcWF0sA7FEZR9W2f0Tgb8PAnl/+v8/+++3qs1YAvs77DW8BPsfq86FWGK4H3v9L6pH\nzIE42rNtsCT0B3xJ73zs/z1c+1W2CErQWpft2C/sC6lcJQD7NVlO5V+hafiSpEysotYN+/B6nsBH\nngXy2Cys71F3v+u6Exw7qNws59UD+8V4K9ZskYx9KAWa/NTVub0Cq7aciFUCL6b+U3/s8cTmXxlK\n89t3Ib5mG7AkzL9q4O9K7Ivg19iXt7e5pD7J3mee7Z+KNe14v0yysS+ifthxTMaqK97mrJpeAzj8\nOO7CXht/Pajc7Fv1MZ8A52NNZhuAl2vY1wtYs+ax2DG4j/p9Lm7Hkrpkv1NbKk8NUd9BD4Hcv67j\neweWfKZjz+scDh8MUNd+arrd//oRHNl7aCrW/NnPE+//q+F+uzn8s6mCyknYkQwumYNVhY7GjuOL\n+N4HdW03kPdnfbyO/egZgr03q6uKxmDNvI9jlfVkrK9oYw1k6I41YXqrbldjFfI9ntPTWBJ+USPt\nT6qhBK31uQFrWiiucr0T66cyCfuC6YF1+Pb2hxiGfXiB/cJ04/tln4mV5mtS22P9978Aa9qIw5pa\nR1L7h2NDR+y9gnV6Pt3z+GOxD/wEz/6ysf+N67AKWqDqOg4ZWFNLJNbhuBx73vWxDeu3MxGreP4C\nS/S8fsSaRy7y3H4/Nfd9aos1a+dgz/2vVW4P5NiWY7+mp2BfEp96rndhSdHT+BLEbljSBPAqdnzP\nw451N3xVrqrH8UPsy/sK7Bf7cOz98X4NsXbGvtwSPPEVUvNxbou9FkWebY6q4/lWje1lrFLnbXZM\nwPoN1VQhqUug7+m6jm9b7H88D0voJ1TdQAD7qev97N1PQ99DA7DKVRvs+JdQ8+v0OvZ51NOzz79i\nleiGjPKsTlusClWGvZZX4vvsyfLsp6Zj8RH1e3/WZS7WP+9mrImzOtGeU7YntgvxvfY1cWCfBbF+\np6rxxWPJ/L+waU4+xD5jemP9LE/1nE7Cktpwnluy2VOC1vpswTote/knQKOxL7Mt2C+nfwJ/99w2\nACunF2D/vLdhzSdgycJM7ANuaDX7rO2x/vv/M/YrfK9ne69jH5jVxeq97K7mfF3exBLROVh/jwVY\ncrEe6xf1lSeGk7CRV9Xtr7qYnsGefw7V989IxZKZPM++FlN9f7C69jMC+9Dcj/VXmofvOOUBt2BJ\n6E6sn5J/04//tmdhCd8urFL4VZX9BHpM52AVlPlU/sIchzV9fe2J61N8fVhWYQnaU1jSvhjfQI2q\nxzEHS0LvwL6Q7vRc9u9T5R9nBPZlvgs7RmdRc+J1J/ZlnI9VT+fW8ZwnUvm9vhobETzNE88m7Eur\nPtWc2o55bdup7fg+jf3Qycb6Q31UzbbqivFvWIKfi40kre4xDXkPeS+3w455DvZ5kI2NXK7ucX/H\n/leWYJ9PRVTuR1rbZ0NN9/F3C1bdzsf6VPpXQIuwz4svPbEOrLL9/dTv/VnX/9Ve7DX7BYdP0ut9\nXAH2OfqGZz9X4Bs9XR03VrUv9jyfIuyzPtJz+zTsue/F/ifn4xs5fjVWXVyHNQHvw5L3Z7AfI+1r\n2a80Y3/HXmj/IbsTsS+Wbz2nwYc/TFqJx7BOxlK7eVRfIREREWmQs7Bhwf4J2gR8v9akdTkeG1Xl\nwJoZsgje8jrN2QCsuSUCa9ooxpodRESkhQj1CIylVD9ku6XO2Cy1S8SaNY/CKqtTsLmdpLJUrFm2\nI9Z8eTOwNqQRiYhIi9OTwytoP2NfOK+i9m0RERFpZcKhUtUTW4bEO3lnZ3xz1TyMzX9TafLPY445\nxr15c21TKImIiIiEjc3UPLVQtcJxFOc+fKNcXsH6IlWyefNm3G53qztNmDAh5DHoeet563nreet5\n63nredfvRN1T1hwmHBO0rn7n/4AWZRUREZFWJtSDBF7HJsVLwTo7T8Am8zwNq6BtxWaeFxEREWk1\nQp2gXVHNdX+v5joBMjIyQh1CSOh5ty563q2Lnnfr0lqfd0OEwyCBhnB72nRFREREwprD4YB65lyh\nrqCJiIhICHXo0IHc3NxQh9EiJCcnk5OTU/cdA6AKmoiISCvmcDjQd2rjqOlYNqSCFo6jOEVERERa\nNSVoIiIiImFGCZqIiIhImFGCJiIiIhJmlKCJiIhIszJjxgzOOuusUIcRVErQRERERGoxbdo0BgwY\nQGxsLNddd12T7FMJmoiIiNSL2w1Ll4LLZZdLS+Grr0IbUzB169aNBx54gOuvv77J9qkETURERCop\nKICNG32Xt2yB7GzfZacTPvkEnnoKiovhkUcsQfOfAqzqdGANmWptx44dXHrppXTu3JmUlBRGjx5d\n7f3GjBlDWloaSUlJDBgwgGXLlh26beXKlQwYMICkpCRSU1O54447ACgpKeGqq64iJSWF5ORk0tPT\n2bdvX7Xb/8Mf/sCQIUPo2LFj/Z9EAylBExERkUp27oSHH4YffrDkbOJE2LrVd3tUFNx/P+zbB5dd\nBklJ8Je/gMNvKtYFC+CNN+x8SYlt4+efA4/B6XRy8cUX06tXL7Zt28auXbu44orqlvCG9PR01q5d\nS25uLldeeSXDhg2jrKwMsORt7Nix5OXlsWXLFoYPHw7AzJkzyc/PZ+fOneTk5DB9+nTi4uJqjakp\nJ/RVgiYiIiKV9O0L/+//wT33wJgxcPPNcOaZ9dvGuefCokXw2mvw4IPQoQOkpQX++JUrV7Jnzx4m\nT55MXFwcMTExDBo0qNr7jhgxguTkZCIiIrj99tspLS1lo6cEGB0dzaZNm8jOziY+Pp709PRD1+/f\nv59NmzbhcDjo378/iYmJtcbkcDTdAkxK0EREROQw/rlKu3aVb6uosGbNzp2tSpaXB08/XbkZs0MH\nGD/ebv/hBxg9GiLqkXXs2LGDHj16EBHAg6ZMmUK/fv1o3749ycnJ5OXlke1pk3311Vf58ccf6du3\nL+np6XzwwQcAjBw5kgsuuIDLL7+cbt26MW7cOCoqKmrdjypoIiIiEjLeZs177rFE7NFHYd063+2R\nkXD++TB2LMTFWXPnL35RuYmzpASefRbOOAOOOgrefLN+MXTv3p3t27fjdDprvd/SpUuZPHky8+fP\n58CBA+Tm5pKUlHQomTr22GOZM2cOWVlZjBs3jqFDh1JcXExUVBTjx49n3bp1LF++nPfff59Zs2bV\nui9V0ERERCRk2ra1itegQXDqqXDXXZWraA4HnHWWryIWE2MJmr8FCyA11apof/ubNXdu2RJ4DAMH\nDqRr167cfffdFBUVUVJSwvLlyw+7X0FBAVFRUaSkpFBWVsZDDz1Efn7+odtnz55NVlYWAElJSTgc\nDiIiIli0aBHff/89TqeTxMRE2rRpQ2RkZLWxOJ1OSkpKqKiowOl0UlpaWmfieKSUoImIiEglnTtX\n7nN2yinQvXv9tnHZZb5mzQ4drJrWu3fgj4+IiOC9997jp59+Ii0tje7du/OGZ9SBw+E4VM0aPHgw\ngwcPpk+fPvTs2ZO4uDjS/Dq7LVy4kJNOOonExETGjh3L3LlziYmJITMzk2HDhpGUlES/fv3IyMhg\n5MiR1cby8MMPEx8fz2OPPcbs2bOJi4tj0qRJ9Tsg9dR0tbrG5W7KdmAREZGWyuFwNGnfqpaspmPp\nSSbrlXOpgiYiIiISZpSgiYiIiIQZJWgiIiIiYUYJmoiIiEiYUYImIiIiEmaUoImIiIiEGSVoIiIi\nImFGCZq0Di5X5UXiREREwpgSNGnZ8vLgtttg4ED49a/h3/8OdUQiInKEZsyYwVlnnRXqMIJKCZq0\nbBMnwvLltm6Jw2Er/27aFOqoRESkmSgrK+OGG26gZ8+etGvXjv79+/Pxxx8Hfb9K0KRl++orS84i\nIiA+3po6160LdVQiIs1fQYEtsDlmDLzyCpSVhTqioKioqCAtLY0lS5aQn5/PI488wmWXXca2bduC\nul8laNKydewIRUV23u22U/v2oY1JRKQ52LgR/vpXeOgh+O67yreVl8Of/wyzZsHq1fDCC3DvvYf3\n9XW5ICvLkrkG2LFjB5deeimdO3cmJSWF0aNHV3u/MWPGkJaWRlJSEgMGDGDZsmWHblu5ciUDBgwg\nKSmJ1NRU7rjjDgBKSkq46qqrSElJITk5mfT0dPbt23fYtuPj45kwYcKhBdh/+9vf0qtXL7755psG\nPadAKUGTlm3CBPtVl5lpp0GD4Fe/CnVUIiLhbeNGuP56ePtt+OADuOkmS8S8Nm2CDRsgNRWSk+3v\nkiWQne27z4EDcMMNcPHF1gd46tR6DdZyOp1cfPHF9OrVi23btrFr1y6uuOKKau+bnp7O2rVryc3N\n5corr2TYsGGUeSp6Y8aMYezYseTl5bFlyxaGDx8OwMyZM8nPz2fnzp3k5OQwffp04uLi6owrMzOT\nH3/8kRNPPDHg59IQStCkZUtPh7lz4cEH4amn4MknISoq1FGJiIS3+fOtSpaa6usm8tpr9dvG44/D\n999Dp07WmjFjBnzxRcAPX7lyJXv27GHy5MnExcURExPDoEGDqr3viBEjSE5OJiIigttvv53S0lI2\nbtwIQHR0NJs2bSI7O5v4+HjS09MPXb9//342bdqEw+Ggf//+JCYm1hpTeXk5I0aM4Nprr6VPnz4B\nP5eGUIImLV9aGlx4oVXOlJyJiNStosIGVnlFRFjC5nXccXDCCbB3L+Tm2t9zzoGUFN991q616prD\nYZ+9DodV5gK0Y8cOevToQURE3anKlClT6NevH+3btyc5OZm8vDyyPdW8V199lR9//JG+ffuSnp7O\nBx98AMDIkSO54IILuPzyy+nWrRvjxo2joqKixn24XC5GjhxJbGws06ZNC/h5NJQSNBEREans97+3\n5sicHEvASkrgsst8t7dpA9OmwdVXwxlnwC23wKRJlZO6nj19fc+8fYC7dw84hO7du7N9+3acTmet\n91u6dCmTJ09m/vz5HDhwgNzcXJKSknB7mlOPPfZY5syZQ1ZWFuPGjWPo0KEUFxcTFRXF+PHjWbdu\nHcuXL+f9999n1qxZ1e7D7XZzww03kJWVxVtvvUVkZGTAz6OhlKCJiIhIZaedBs89B/37wymnwBNP\nWIXMX2KizTP5zDPW1yw6uvLt99xjTZvZ2bBvH5x7Lpx/fsAhDBw4kK5du3L33XdTVFRESUkJy5cv\nP+x+BQUFREVFkZKSQllZGQ899BD5+fmHbp89ezZZWVkAJCUl4XA4iIiIYNGiRXz//fc4nU4SExNp\n06ZNjYnXqFGj2LBhA++++y4xMTEBP4cjofYeEREROdyZZ9qpoY4+Gt54wwYUxMVZs2gAzZVeERER\nvPfee9x2222kpaXhcDgYMWIEgwYNwuFw4PBU6wYPHszgwYPp06cPCQkJjB079tCIS4CFCxdyxx13\nUFRURM+ePZk7dy4xMTFkZmYyatQodu7cSdu2bbn88ssZOXLkYXFs27aNl156idjYWFJTUw9d/9JL\nL9U4aKExOOq+S1hyu7Vsj4iIyBFzOBzoO7Vx1HQsPclkvXIuNXGKiIiIhBklaCIiIiJhRgmaiIiI\nSJhRgiYiIiISZpSgiYiIiIQZJWgiIiIiYUYJmoiIiEiYUYImIiIiEmaUoImIiEizMmPGDM4666xQ\nhxFUStBEREREanHVVVfRtWtX2rVrR+/evZk0aVLQ96kETURERBqksKyQ51c+j9PlDHUoQXXPPfew\ndetW8vPz+eijj5g6dSoff/xxUPepBE1ERESqtXbvWr7c/mWNt7+94W2eWfkMX2z7osb7lFaUNnit\nzx07dnDppZfSuXNnUlJSGD16dLX3GzNmDGlpaSQlJTFgwACWLVt26LaVK1cyYMAAkpKSSE1N5Y47\n7gCgpKSEq666ipSUFJKTk0lPT2ffvn3Vbv/EE08kNjb20OWoqCg6d+7coOcUKCVoIiIichiny8nE\nxROZuHgipRWlh91eWFbIK9+8QvvY9kxdMbXaKprb7ebWD2/l9R9er//+nU4uvvhievXqxbZt29i1\naxdXXHFFtfdNT09n7dq15ObmcuWVVzJs2DDKysoAS97Gjh1LXl4eW7ZsYfjw4QDMnDmT/Px8du7c\nSU5ODtOnTycuLq7GeG655RYSEhI48cQTuf/++zn99NPr/ZzqQwmaiIiIHGbJtiXszN9JbkkuH276\n8LDb397wNoXlhXRO6MzOgp3VVtFW7V7F6t2reWn1SxSVF9Vr/ytXrmTPnj1MnjyZuLg4YmJiGDRo\nULX3HTFiBMnJyURERHD77bdTWlrKxo0bAYiOjmbTpk1kZ2cTHx9Penr6oev379/Ppk2bcDgc9O/f\nn8TExBrjef755zl48CCfffYZ999/PytXrqzX86kvJWgiIiJSidPl5NkVzxIfHU9SbBLPr3q+UhWt\nsKyQl1a/hAMHeSV5OF3Ow6pobrebqSumkhCdQGF5Ie9seKdeMezYsYMePXoQEVF3qjJlyhT69etH\n+/btSU5OJi8vj+zsbABeffVVfvzxR/r27Ut6ejoffPABACNHjuSCCy7g8ssvp1u3bowbN46Kiopa\n9+NwOMjIyGDYsGG8/nr9q4L1oQRNREREKlmybQk/5f5EhCMCl9vF3oN7K1XRCsoK6J/an/5d+9O3\nU1/O7HYmx3Q4hpKKkkP3WbV7FRuyN9A+tj1JMUn1rqJ1796d7du343TWPgBh6dKlTJ48mfnz53Pg\nwAFyc3NJSko61O/t2GOPZc6cOWRlZTFu3DiGDh1KcXExUVFRjB8/nnXr1rF8+XLef/99Zs2aFVBs\n5eXlJCQkBPxcGiIqqFsXERGRZie7OJtTu5x66HK3xG7kluQeupzaNpVnLnym1m3MWjuLUmcp+4v3\nA1BQWsCnmz9lyAlDAoph4MCBdO3albvvvpsHH3yQiIgIvvnmm8OaOQsKCoiKiiIlJYWysjIeffRR\n8vPzD90+e/ZsLrjgAjp16kRSUhIOh4OIiAgWLVpESkoK/fr1IzExkTZt2hAZGXlYHFlZWXz++edc\ncsklxMbG8tlnnzF//nw+++yzgJ5HQylBExERkUqG9RvGsH7DjmgbYwaO4apTrqp03bEdjg348RER\nEbz33nvcdtttpKWl4XA4GDFiBIMGDcLhcOBwOAAYPHgwgwcPpk+fPiQkJDB27FjS0tIObWfhwoXc\ncccdFBUV0bNnT+bOnUtMTAyZmZmMGjWKnTt30rZtWy6//HJGjhx5WBwOh4MXX3yRUaNG4Xa76dOn\nD6+99hpnnnlmA49MYBxB3XrwuBs6ZFdERER8HA5Hg6fBkMpqOpaeZLJeOZf6oImIiIiEGSVoIiIi\nImFGCZqIiIhImFGCJiIiIhJmlKCJiIiIhBklaCIiIiJhRvOgiYiItGLJycmH5hSTI5OcnNxo22qu\nr4jmQRMREZFmQfOgiYiIiLQAStBEREREwkyoE7S/A5nA937XdQA+BX4EPgHahyAuERERkZAJdYL2\nD2BwlevuxhK0PsDnnssiIiIirUY4DBLoCbwHnOy5vAE4B6uspQKLgROqPEaDBERERKRZaCmDBLpg\nyRmev11CGIuIiIhIkwv3edDcntNhJk6ceOh8RkYGGRkZTRORiIiISC0WL17M4sWLj2gb4drEmQHs\nBboCi1ATp4iIiDRTLaWJ813gGs/5a4B3QhiLiIiISJMLdQXtdWxAQArW32w88C/gDSAN+Bm4DDhQ\n5XGqoEn4crng3Xdh7VpIS4PhwyE+PtRRiYhIiDSkghbqBK2hlKBJ+Hr8cZg3D6KjoawMTjkFXnoJ\n2rQJdWQiIhICLaWJU6T5KiyE+fOhSxdISYGuXWH9eli3LtSRiYhIM6IETaQxOZ32N8Lzr+Vw2Kmi\nInQxiYhIs6METaQxJSbCOefAnj1QUACZmZCaCv36hToyERFpRtQHTaSxFRfD9Onw7bc2SOC226BT\np1BHJSIiIaJBAiIiIiJhRoMERERERFoAJWgiIiIiYSbc1+KU5m7PHnjvPSgthf/9X+jbN9QRiYiI\nhD31QZPg2b0bRo6E3FybdiIqCp57Ds44I9SRiYiINBn1QZPwsmABHDgA3brZhK2RkTajvoiIiNRK\nCZoET3GxJWVeUVF2nYiIiNRKCZoEz29+Y38PHICDB+00ZEhoYxIREWkG1AdNgmv5cpu0tawMLr0U\nhg61pY9b7h6dAAAgAElEQVRERERaCU1UKyIiIhJmNEhAREREpAVQgiYiIiISZpSgidRl1y646SY4\n5xy48UbYuTPUEYmISAunPmgitSkrg2HDIDMT2reHvDxISYE334SYmFBHJyIizYD6oIk0th07YN8+\n6NQJ2rSx5Cwry64XEREJEiVoIrVJSACn005gf10um4B31y4oKQltfCIi0iKpiVOkLk89Bf/8J7jd\nNofbL38Ja9dachYbC48/DunpoY5SRETClOZBEwkGtxu++gq2b4eOHeGRR2zx97ZtbXUElwvefx8S\nE0MdqYiIhKGGJGhRwQlFJEiys+G116wf2C9/CRddFPyVCRwOGDTITps22cCBlBS7rW1bi2nPHiVo\nIiLSaJSgSfORlwfXXgt790J0NCxcaB34r7uu6WLo1Mn+eps3vX3QvNe3ZgUF8OST8O230KMH3HUX\ndOsW6qhERJolDRKQ5uOrr2y6i65dramxY0f4xz+aNob27WHCBGvazM62v+PHQ3Jy08YRbtxuuPNO\nePddS9RWrIA//tGOj4iI1JsqaNJ8uFyVL0dE+EZXNqXBg6F/f2vW7NoVunRp+hjCzYED8M03djwc\nDoiPtwR240Y444xQRyci0uwoQZPmY+BAq2Dt22eTxBYWwjXXhCaWLl2UmPmLjra/TidERVlFzenU\nZL4iIg2kUZzSvOzcCdOnW5J29tlw+eU2J5mE3tSpMGOGr7I5aBA884xeHxFp9TTNhkgorFkDc+ZY\n1eiyy+DMM0MdUWi43fDvf8P69XDUUXDJJb7KmohIK6YETaSqhQttZGFhIVx4oXVkb8xmtzVr4Oab\nfVN9OJ3w3HOtN0kTEZHDaC1OEX9r1sADD0B5uc1XtmCBNcM1pjfftL8pKXaKioLXX2/cfYiISKuj\nBE1artWrraKVkGALnXfoAIsWNe4+qk6S610OKlDLlsGDD8ITT8Du3Y0bm4iINFsaxSktV1KSJUxe\nxcWNP/Jy2DD49FNb2cDhsITwiisCe+yHH9ocalFRUFEBH38Ms2drdKiIiKgPmrRghYVwww3w0092\nOSYGnn8eTj21cffz3Xcwd67N0zZs2OHzfhUWWoVsxQpLvu65B447Di69FHJzrfkVYNcu6yN35ZWN\nG5+IiISU1uIU8ZeQYCsNfPGFLcl0+umQltb4+znlFDvVZPx4WLzYmlg3bICbboL5861qFlGll0Eo\nJt4VEZGwowqaSDCVldl8YKmpvr5p+/bBo4/aSgRPPGGJZFmZNXX+85+2jqWIiLQYqqCJhJuoKBug\nUF5uc4K53dYUGhdnfdViYqzvWWKiVdaUnImICKqgiQTfvHkwebIlZ2639VF7/nlL3EREpMXTRLUi\n4WrFChtM0LmzTZirGfZFRFoNJWgiUje3G9auhcxM6N3bRpSKiEjQqA+aiNTO7YYpU+CNN3wjSO+/\n39bNFBGRsKEKmkhr8uOPMGIEdOoEkZFQWgoFBTYNSGOuUSoiIodoLU4Rqd2BAzayNDLSLsfE2Nxr\nBQWhjUtERCpRE6dIOCoshMcfh+XLbWDBPffASScd+XaPOcbmY1uzBoqK7Hz//jaJroiIhA1V0ETC\n0YQJ8P77lkBt2wajRtnEtkeqY0dbTSE/31YycDisepadfeTbFhGRRqMKmki4cTptearUVOvIHxNj\nqw98/z107Xpk2y4rs7VJzzrLBgxERNi2162zSp2IiIQFVdAk/BQWwldfwddfQ3FxqKNpehERttJA\nWZld9q4+EB9/5NsuLLRVDfLybD/ebSckHPm2RUSk0aiCJuElOxtuvNHXnNejB7z8MiQlhTYusERm\n2TKLrXdvOPPM4OzH4YA774SHH7Z9gi30PnDgkW33v/+FW2+1heN/+gmSk61qNmiQbV9ERMKGptmQ\n8PLXv8Lbb1vzHlgydPXVcNttoY3L7baE6d137bzDYf3CbrghePtcs8ZWH+jQAc4//8hXH/jDHyAr\nyxKz3FzYvRvGjrXnEaXfaiIiwaKJaqX5274dYmN9l2NiYOfO0MXjtWWLddrv0sWaBisqYPp0uOwy\nW+g8GE47zU6NweWCHTssfrAkrawMevVSciYiEobUB03CS3q6Tf/gclln+ZISW1w81A4etLnDvLPv\nR0VZFa2wsObHbNpkFaprroFZs+z5hEpEBPTtC/v32+WyMqsE9u4duphERKRGStAkvIwcCb/7na0T\nuW8fDB8OQ4ce2Tb37oU//tH6cA0dan2x6uuYY6xStn+/dbLPzISePW1G/urs2mXNn199ZdNkPPOM\n9aULpb/9zZqO9+2zQQJ33gn9+oU2JhERqZb6oEl4KimxCtWRLj/kcsEVV8DWrZCSYvN/tWlj/dza\nt6/ftrZsgQcfhJ9/hhNPtLnKvE2GVb31liVE3mkxSkstls8+O6Knc8QqKqwfWmIitG0b2lhERFoJ\n9UGTlsO/H1qgXC5rckxI8DVF5uRYcubf9yo720YxDhhQv+337g0zZwZ236goa0L0cjotMQRbbmnd\nOnuOp57atH3AoqKOfC41EREJOiVo0jJ895012eXmWrPjk0/CCSfY3GEOh/W5io729W0LVsd+r3PO\nsebE3bstKaqogPvvt2Txppts9n6Xy/rXPfNMw0ZoFhXBtGmwerU1t/7lL0q+RERaCDVxSvNXUGD9\n1lwuaNfOkrS4OHjvPatSzZsHTzxhFS2326abuPdeS9yCKTsb5s+3Kt7ZZ9vs/TffDGvXWnOr220J\n3IQJMGRI/bbtdtsAhKVL7TkXFtqcZvPm1Tzp7Pr1NhI1Kgp+/3sNEBARaSJq4pTWaccO6+PVsaNd\nTk62flZ791plafhwG8G4ebM1df7P/wQ/OQNLwkaNqnzdzp2+BMrhsKbYhqyxWVAAX35pFTOHw/qT\nZWVZElbdBLpr1lgsFRWW3L39NvzjH3DssfXft4iIBJ1GcUrz17GjNVuWl9vl0lL7m5zsu88pp1jl\nbNAgX/+0UBgwwCp8brfF63bDSSfVfzvefmveqTu8SzZ5+7lVNWOGJXKpqZbUlZZadU9ERMKSEjRp\n/rp0gdGjrSkxK8umkLj77vBYHqqqO++0JHHvXkvU/vxn+OUv67+d+HgYMcKmzNi3z6pwp55ac7JX\nWlo5MY2I8CWyIiISdtQHTVqOn36yPl1pada0Gc4KC21gQE0Vr0C4XPDxxzZAIi0NLr205tGvn31m\nSWt8vD2urAxeeCE8JgEWEWnhGtIHTQmaSKC8zZJHuiZmqHzyCcyda82j115rlTwREQk6JWgiwbJm\nDYwbZyMze/eGKVOgR49QRyUiIs2AEjSRYMjJsWkwIiJ8yz116WKrBURGhjq66nmnFAnlgAgREQEa\nlqDp01ukLlu32vQU7drZSMiUFOuU7114PJy43fDaazbw4H/+Bx55xPqbiYhIs6J50ETqkpxs01k4\nnVYxKynxVdMCVVEBc+bYxLKpqTZhbbdujR/r4sXw9NOWREZGwoIFFv+ttzb+vkREJGjCuYL2M/Ad\n8C2wMrShSKvWuzeMHGlTeGRl2YLr991nqxUE6plnLHHauBEWLoTrrrNpNhrbihU2CCA62hK0pCRY\ntqzx9yMiIkEVzhU0N5AB5IQ4DhGbrywjAzIzLWGrzzJJLpdNCtuliyVPSUm2nf/8B37zm5of53bb\nXGUxMYGvfJCaatU6r5ISu66l2rcPHnoIfvjBplYZP15LWIlIixDOFTRovoMYpKVxOODkk+F//7f+\nCYB3SSeXy3ddXR34N2609UV/+Uu4+GJbwqku+fmQng69etlEuJmZNu/ZmDH1izdcZWdbn7o//Qle\necWSzzFjYOVKm/9t40ZrOj54MNSRiogcsXBOgLYAeYATmA687HebRnFK8/Lyy/Dii1YNKyuz/mez\nZ1ffj6242JKz4mLrP3bggDVZvvtuzQuhz59vC8KDVeiuuQY6dYL+/a0/WnNXWGgrJ+zaZU3LhYVW\n0Vy2zJ6nt8KYnW0T8J52WkjDFRHx19IWS/8lsAfoBHwKbACWem+cOHHioTtmZGSQkZHRtNGJ1MeN\nN1pT41dfWVPnyJE1DzLYs8eqQN7Eqn17Szx27oTjjz/8/hs3wuTJlsxFR9t933674Wttetf1bOgU\nIoWFsGSJNc8OGABHH92w7fj77js7Lt7m2sREWLTIqpBOpzUdu1x2vqYkVkSkiSxevJjFixcf0TbC\nuYLmbwJwEPCUCFRBkxYsJwcuvNASs+hoW70gNxfeew86dz78/p98Ag884LvN7bZkxjtgoD4WL7Y+\nXd7m0kmTLPErLbU1Tjt2rD1xKyiwARA//2yX4+Jg+nTo169+cVS1YgXcdpvvOTqdNmDj+uvh1Vft\nOTsc1hw8YULgffZERJpAS6qgxQORQAGQAJwPPBjSiEQaqqwMvvzSKkunngrdu9d+/w4dYOxYeOop\nu+x222Lw1SVnAEcdZdWjigpLyPLyoGvX+idnW7b41utMTYVVqyzx+93v4MEHbfudOsGzz9bcD+/d\nd2073ilE9u+HJ5+0PmNH4tRTbRDA5s2WtJaUwOWXW5+zM86wfaamwtlnKzkTkRYhXD/JegFve85H\nAf8E/uZ3uypo0jyUlsKoUfD995Y4REXBc89Z37C6bNoE27dbE2F1TZv+XnrJkqCoKOswP21a/atW\n779v1bMuXeyyy2XNqnFx0Lat/d2/35pe//Wv6hOhqVNh5kxLEAGKimyC37ffPvy+9ZWfD7NmWUxn\nnAH/939aKUFEmgUt9SQSbj780KpQXbtaQpOXZ5WeN95o/H3t2mVNoT161G8SXa9ly+Avf7H4HA7r\nB1dUZOc7dfLdLzMTPvvMEq+qVq2yhDQpCdq0sWbI666zaUpERFopLfUkEm4OHLC/3mpTXJz1MQuG\nbt3gpJMalpwB/OIXNjLSO0VHSQnccYev+RQsaUtIsIpadc480/qARUZacjd8ONx0U8PiaQybN8MH\nH8Dy5ZWnORERCXPh2gdNpGU49VRfshITYyMsL7kk1FFVLzISHn/c5hXLy4O+fSEtzZLMV16x2yMj\nYcqU2psWL7kkPJ7j55/Dvff6Fo4/91x49FE1i4pIs6AmTpFg+/RTS3zy8+HXv7ZloprbVBBbt1py\n2bNn5ebOcOV2WzUwKsoGPbjdVhV8/nmr8omINKGWNIpTpOX4zW/s5J0Kojnq1ctOzUV5uTXHegcr\nOBxW/cvLC21cIiIBUq1fpKk01+SsOYqOtkly9+61vmcFBda02bdvqCMTEQmIEjQRaRzl5bbc1Dnn\nwPnn28S6ofS3v9lku/v22eCMp5/2zc8mIhLmmutPevVBEwk3L7xga4526mSjPvPyrM9Xenpo42rO\nTcsi0iJomg0RCZ1//9u3PFV8vCVFy5eHOiolZyLSLGmQgIgcmS+/hI8/hm3bKi9W7nLZslUiIlJv\nStBEWiqXC+bMsSWcEhLg1lvh9NMbdx+ffmpzjUVF2VxvO3ZYX7ToaFvRYMiQxt2fiEgr0Vxr/+qD\nJlKXGTNsYfOkJEuanE5bJ7NPn8bbxxVXwO7dvmWffv4ZzjoLLr7YFi6vacUBEZFWRPOgiYjPO+9A\ncrL1BwNLpJYsCSxB27/fkrndu+FXv4Lf/a76GfidzsrXR0fb6gkXXdQ4z0FEpJXSIAGRlio21reG\nJthoxtjYuh9XUADDhsFDD8GLL8I119j56lx5pa2QcOCALYweG2uT8oqIyBFRE6dIS7V0qS127nZb\nf7ROnWD2bEhJqf1xH35oTZeRkdCmDZSW2jZ27Tq8ydLttvt7+7ndcIMmgxURqaIhTZxK0KT52LfP\n1oTs1Al69w51NM3DmjWweLElVkOGBLaO5qxZcMstlUdjFhfDqlVKvkREGkAJmrRcX34Jd91lyYLT\nCTfeCDfdFOqoWqatW+G00+x8ZKQ1k7Zvbwlaly6hjU1EpBnSRLXSMlVUwH33QUyMNc917AivvAKb\nNoU6spapVy949FGrurVpY8f8wQeVnImINCGN4pTwd/AgFBZCaqpdjoqyyk5WFhx3XGhja6lGjYLz\nzrN5zbp1U9OmiEgTU4Im4a9dO0vO9u+36llxsV3fo0do42rpjj/eTi1JcTHs2WPTjyQnhzoaEZEa\nqYlTwl9EBDz9tC0blJkJJSUwaZJVdkQC9d//2nxuI0bAhRfC/PmhjkhEpEYaJCDNh9MJublWUYuO\nDnU0dSsogGnTYMMGOOEEW2rJO+O+NC23G377W3tN2reHsjJ7L73+OhxzTKijE5EWTisJSMsWGVn3\nHF7hoqICRo+GH36wzvbr1sHGjTa4IUr/dk2uqMj6LHr7MUZHW2V2xw4laCISltTEKRIMO3ZYk1pq\nKiQm2t///hd27gx1ZK1TfLw1kefl2eXycpuyRc3kIhKmlKCJBENkpDWr+XO7q1/PUoLP4YAnnrDq\nZXa2NW/edptGAYtI2FJbi0gwHH00nH02LFpkzWllZZCRYddLaJx0Erz7ri1ZlZwMnTuHOiIRkRpp\nkIBIsJSVwZtv+gYJDB3aPAY3iIhIo9JSTyIiIiJhRqM4RaT5cLlg/Xqb+qJPH5uEWEREACVoIlKT\nPXts9Ya0tMafv83lgvHjYeFCG1ARGwvPPQcnnti4+xERaabUxCkih3vtNZtkNyLC+s09+yycemrj\nbX/pUhg71hZgj4iwUZWpqdZnT0SkhWlIE6fG/ItIZT/9BFOn2khH78TAd95pVa/qlJXZjPyTJsE7\n79iKD3XJyrKpL7zTjrRrZ6MrRUQEUBOniFS1e7clTm3a2OV27WDvXigstEl3/blccPvt8NVXNsfY\nggXw/ffwwAO178M7/1hZme0nOxsGDGj851KT5cvh669t8tpLL9USXCISdlRBE6mL223JSU0VpJYm\nLc2ec2mpXc7NhU6dbMmqqjZtgpUroWtXm1csNdXmGtu/v/Z9nHwyjBtnM/tnZsLxx8NDDzX+c6nO\nggU2Se3cuVYpvP56e31FRMKIEjSR2uzYAZddBuecA7/+tVVeWrqePeGee2x0ZVaWLZP05JPWJFlV\neblV27y3ORx2qqioez9Dh8KSJfDZZzBrVtOtszptmi2Y3qULHHUU/PwzLFvWNPsWEQmQmjhFauJy\nwV/+Yn2juna1Ksudd1oFxrvodk3KyuDTT62SdPLJ0L9/08TcWH7/ezjvPF/n/ZiY6u933HG2OsL2\n7ZCQYEndGWdYxS0Q0dFNP3lvaWnlplqHw14vEZEwogRNpCb5+ZZ4eJOxhAQoLobNm2tP0MrL4dZb\n4Ztv7HJEBNx/PwwZEvyYG1O7dnX3zYqJgRdfhGeeseNy/vkwenR4rzk6ZIg1b7Zvb69nfHzT9n8T\nEQmAEjQRf9nZ8Pnn1kQ3aJDNz1VcDHFxNjrR6ax7QtVVq2DNGqu6ORxQUgKTJ8Pvfld9M2Fzl5IC\nDz8c6igCN3as9adbvNhey7Fj7bUSEQkjStBEvPbuhauvtiTN4bDK0A03wPTpcPCgJWdXXmkd2mtT\nWFi5X1Z0NBw4YI+P0r9cyLVpA7fcYicRkTClbwsRr7lzIScHunWzy1lZ8N138MYb1nzXqRP061d3\nFeyUU6zpLyfHmkX374eMDCVnRyIz05qcjz7aqpkiIi2cvjFEvPLyKidRMTF2XffudgpUly5Wffvr\nX2HfPrj4YrjrrsaPt7WYPh1efdWWhGrf3paE6t071FGJiARVc+0Qo6WepPF5lx9KTLRkIDcX7r4b\nhg0LdWSt15o18Mc/Wj+3qChrfu7Rw6qdIiLNhJZ6EjkSZ50FEybYyMWYGBuN+H//F+qoWrcdO6xJ\n2VvZ7NDBmpv1A01EWjg1cUrrU1hoE69+/bVNl3H33b6lhy65xE4SHrp3t2SsosKStNxcOOaYljka\nVkTET3P9lFMTpzTcHXfYFAsdOtjozNhYmD+/6Wayl/p5+WU7efugPf889OoV6qhERALWkCZOJWjS\nupSV2fxmqam+Ksy+ffDoo3DuuaGNTWqWlWWjOLt1s4RaRKQZaUiCpiZOaV2iomwerPJym5/M7bYl\nnTR1Q3jr1Cnw5aNERFoADRKQ1iUiAsaMsbnJdu+202mn2fqRcmQKCmD1ali3zpJeERFpMDVxSuu0\nYoVNQtu5MwweXPNi4BKY7dvhppts3jinE371K3j8cU3OKyKC+qCJSKjcfDOsXWsDLdxu2LPH1ue8\n6KJQRyYiEnKaB01EQmPbNluAHHyDL3btCl08IiLNnBI0ETly/fvbHGXeOcscDjjhhFBHJSLSbNWW\noJ0CfA3sBF4Ckv1uWxnMoESkmbnrLjj1VJuyJDsbbrzR+qGJiEiD1NYe+iXwMLACuAG4Hvgd8BPw\nLdA/6NHVTH3QRMKN221VtJgYSEgIdTQiImGjsedBSwQ+9pyfAqz2XL6qIcGJSAvncNjqDCIicsRq\nS9DcQBKQ57m8CLgUWEDl5k4RERERaUS19UF7HOhX5brvgPOwJE1EREREgkDzoImIiIgEkeZBExER\nEWkBlKCJiIiIhJm6ErRIYGxTBCIiIiIipq4EzQlc2RSBiIiIiIgJpMPaU0AbYB5Q6Hf9N0GJKDAa\nJCAiIiLNQkMGCQRy58XYnGhVnVufHTUyJWgiIiLSLAQrQQtHStBERESkWQjWNBvtsWbO1Z7TE9gK\nAyIiIiISBIEkaH8H8oFhwGVAAfCPYAYFDAY2AJuAcUHel4iIiEhYCaTcthY4NYDrGksksBH4X2AX\nsAq4Aviv333UxBlk32V+R5+OfYiNig11KCIiIs1asJo4i4Gz/C7/Ciiqz07qKR34CfgZKAfmAkOC\nuD+pIrsom5vfv5k3178Z6lBEpIGWLIE3Pf/CLhe8+CJs2BDamEQkcFEB3OdmYBa+fme5wDVBiwi6\nATv8Lu8EBgZxf62Gy+0irySP5LjkWu83+7vZHCw7yMvfvMwfTvgDCdEJTRShiDSWk06Ce++15Gzv\nXtizB3r0CHVUIhKoQCpo+cApfqfTsH5owaK2yyB5c/2bXPev6yh3ltd4n+yibOb9MI+jEo+iqLyI\ntze83Sj7rnBVMG/dPJwuZ6NsL5jWZ61n7d61oQ5D5Ih06ACPPAKvvQaffgrjx0NcXKijEpFABVJB\newvoD+T5XTcfOCMoEVm/s+5+l7tjVbRKJk6ceOh8RkYGGRkZQQrnyFW4KigqL6JdTLuQxVBcXsyL\n/3mR7KJsPtn8Cb/t89tq7zf7u9k43U7aRLahfWz7Rquifb7lcx7+4mE6x3fm3F6hnEKvdi63i/GL\nxlPmLGPB8AVERQTyLyISflwumDMHjjoKSkvh/fdh2LBQRyXSOixevJjFixcf0TZq+/bpC/TDmjYv\nxTq3uYF2QDB7jv8HOA7oCewGhmODBCrxT9DC3UurX2LFzhXM+P0Mb0fBJvfej+9RUFpAx/iOPLfq\nOc4/5nzaRLY57H6Lf15MhauC3fm7AYiMiGRt5loGdR/U4H1XuCqYunIq0ZHRTF05lbN7nE1kRGSD\ntxdMS7ctZduBbeCAf2/5N+cfe36oQxJpkI8/tmbNp5+G4mJr7jzuODjttLofu2IFnHACJCWB2w3/\n/jdkZEBkeP7bioSdqoWjBx98sN7bqC1B6wNcgiVol/hdXwD8sd57ClwF8GdgITai81Uqj+BsVnKL\nc5n93WxKKkpYuWslA49u+u503upZUmwScW3i2Htwb41VtAXDF+Byuypdd6RVpM+3fE5mYSZHJR7F\njrwdLNm2JCyraC63i6krpxLXJo4IRwTTVk3jvN7nqYomzdL558N550FsrDVtTpkCCQEWwjdtgtmz\nrYn09ddh82YYNEhNpCJNqbZvnn95Tr8AvmqacA75yHNq9l7/4XXKXeXEt4nn2RXPMrvb7Cavoi3Z\ntoSc4hzi2sRRWF6I0+Xkn9//s9oELcIRQYQjkK6JgfFWzyIdkZRUlOBwOMK2irZ021J+yvmJlPgU\nHA4H2/O2q4omzVZUlJ282rYN/LEjRlgT6VVXWRPpk08qORNpaoGUBkZhFawDnsvJ2GoC1wcrqJbC\nWz3rENeBNhFt+HH/jyGpop3d42zmDZ1X6brEmMQm2XdOcQ5JMUmHkrGE6ATi2sSRV5pHh7gOTRJD\noLYc2EL3JF/3x+5J3dlyYEsIIxKpv61brWlzkKdXwrJlkJZmp/oo8kymVFEBzvAf2yPS4gRSylmD\njdys67qm1Cwmqp3z/RweXfYobaPtp2theSEZPTJ45sJnQhyZiLRUW7fChAlw002WWP397/Dww/VL\n0F55BTZuhIkTYcECWLkSHn008CZSEaksWIulrwXOBXI8lzsAXwAn12dHjaxZJGgFpQXszK88ALVD\nXAe6tO0SoohEpDXYuhVuu83OP/dc/atn330HxxxjCZnbbQnamWdCROP1fhBpVRqSoAXSxPkE1gft\nDc/GhwGT6htca5QYk0jfTn1DHYaItDLbt1c+X98E7ZRTfOcdDhioqcJFmlwgCdosYDVWRQP4A7A+\naBGJiEiDrVplzZrPPWdNnBMmQHw8nH56qCMTkfoItNx2FnAs8A+gE9AW2BqsoALQLJo4JTScLmfY\njRAVaSoFBZCXB0cfbZe3b4eOHdV/TCSUgrVY+kTgLuAez+VoYHZ9diItw782/otVu1aFOoxaLd+x\nnGveuYYKV0WoQxEJicREX3IG1ryp5Eyk+QkkQfsDMAQo9FzeBTTNHA0SNvJK8nhs2WNMWjopbNfT\ndLldPP3103yz5xsWb10c6nBEpBaFhfDhhzYIAazSt2JFaGMSCSeBJGilgP/U8vot1gq9se4NKlwV\n7CrYxeKfF4c6nGp9uf1LtuZupVNCJ6aunBq2iaSI2ES4CxfCzJmWnD3wgC1JJSImkARtPjAdaA/c\nBHwOvBLMoCS85JXkMXPtTJLjkolvEx+WyY//Mk3tYtqxu2A3i7YuapRtz1gzg9lr1aov0pgSE20p\nqXffhVtvhWuvtfU+RcQEkqBNBt7ynPoADwDPBjMoCS9vrHuDnOIcCstsmahNOZvCroq2atcq1met\npy97kyAAACAASURBVLSilKyDWZRUlPDSNy8d8XZzi3N5afVLvLj6RfJL8xshUpHKVq60pj6w5r45\nc2wtzNYgN9d3fts2X3OniAQ2zcYdwFzgkyDHImEqoU0Cg48dXOk6N+H1SXpcx+N4ZnDlFRoaYzkr\n71qqAPN+mMcfz/jjEW9TxF+PHjB9uiUnWVnw7bdwySWhjir4srKsWXP0aBgwAO6/3xZ2v/zyUEcm\nEh4CGfI5EZucNhdL1OYDmUGMKRCaZkOCLrc4l9/O+S2JMYm43W6Kyov4cMSHtItpF+rQpIXJzIQb\nb7Tzc+ZY819L53LBhg3Qr59dLiiAnBxLWEVammBOs3EicCvQFViC9UMTadHmrZtHVlEWWYVZZBdl\nk1mYyZvr3wx1WNLCuN3w0Uc2FUZ8PCxZEuqIoLzcd97trny5sURE+JIzsKRUyZmITyBNnF77gL3A\nfmyyWpEW7YyuZ/DA2Q9Uuu7ETieGKBpprtxuWy6ppsuffmrNmi+/DEVFcO+90K0bnHZa08cK1vR4\n773w0EOQmgqzZtmUGLfcEtjjMzMt2YqPt8vbttlcbI561Q5EJJB/mVuAy4DOWPPmPEK/1JOaOFuR\n1btXc2LnE4mNig11KCL1UlEB48fDzTdbkrJ+PSxYAPfd50tYSkuhrMzXrJmTA+3bh3Zh8oULYd48\n6NMHdu+20ZbtAmzZnz3bFlufOBH++1946imYPBm6dg1qyCJhrSFNnIHc+W9YUramATEFixK0VmLv\nwb0MmTuEMQPHcOXJV4Y6HJF6W7TI5vq64gp47TW4887QVccC5XbDiBHWL2zKFDj++MAf63LBiy9a\ns210NEyaBCecELxYRZqDYPVBu4fwSs4kSNxuN4u2LsLldtV95yYyc81MisuLeXn1yxSVF4U6HJF6\nO/dcSE+HadNg+PDmkZzNmgUpKXDNNfDYY7BnT+CPj4iw5wvWdy0tLThxirR0ISyiS7hZuWslt39y\nO8t3LA91KIBVz97e8Dbd2nXjYPlB3tnwTqhDEqm39eth+XI47zx46y2bNT+cZWfb6MpHHoGhQy2p\n/OijwB+/ejU8/TQ8/jgMHmxNnUX6bSVSb82126aaOBuZ2+3mqgVX8d2+7zix04nM+b85RDhCm78/\ntuwx3vrvW3Rp24Xi8mKcLicfjPiA+DbxIY1LJFAVFTbP15/+ZJWzRYss2XnssfDuNF/XwIbaLFpk\n/c1OOMGaO+fPhwsusH51Iq1VsPqghSMlaI1sxc4V/PnDP9OlbRcyD2by1OCn+FXar0Ia0yVzLmFX\nwa5Dl6Mjo5l20TQGHDUghFGJ1E9ZmfXFqumyiLR8StCkQbzVsy0HtpAcm0xeaR5HJx7N60NfD3kV\nTUREpLkL1iABaeH2F+9nf/F+IoggryQP3HCg9AD7CveFOjQREZFWSRU0ERERkSBSBU1ERESkBVCC\nJiIiIhJmlKCJiIiIhBklaCIiIiJhRglaE9i0fxOlFaWhDkNERESaCSVoQZZfms+N793I/PXzQx2K\niIiINBNK0ILsjXVvkFOcwyvfvEJhWWGowxEREZFmQAlaEOWX5jNzzUy6tu1KYXkhb294O9QhiYiI\nSDOgBC2I5v0wjxJnCTFRMbSPba8qmogckbw8ePBByM+3yytWwMsvhzYmEQkOJWhB9NFPH+Fyucg8\nmEleSR4Hyw6yaveqUIclIs1Uu3bQsyfcfz989hlMmwYZGaGOSkSCQUs9BVFJRQnlzvJK17WNbutd\n8iFslVSUUFBaQKeETqEORUSqcLvhz3+G7dth4kQ444ym2W9pKWzeDP362eW9ey2Wrl2bZv8izZmW\negozsVGxJMYkVjqFe3IG8OJ/XuSWD2/B5XaFOpSQO1BywBaQFwkTK1daE+dpp8HMmb7mzmDbswf+\n+ldYtcqSs3vvhfXrm2bfIq1R+GcL1WsWFbTmKLsom0vmXEKps5QnL3iSjJ4ZoQ4ppEZ/NJroiGie\nuOCJUIciQl4ejBkD990Hxx4Ls2ZBVhbceWfT7H/jRt++Ro2Ciy5qmv22BhUVUFICbdva5fx8Ox+h\nMkqLoAqaHLHZ383G6XaSGJPIsyuebdVVtPVZ6/l659cs3b6UH/f/GOpwREhKghdegOOOA4cDrr4a\nbr21affv1Uk9IBrV0qVWlSwogJwcuOsuq1ZK66UErZVzupzsKdgDWPVs3g/z6BDXgcToRHbk7WDJ\ntiUhjjB0XvjPC0Q5oohwRDD9P9NDHY4IAHFxvvMOR+XLwZSZaQnEqFEwZQo88wz85z9Ns2+wCpPL\n7/diWVnT7bspZGRYf8JbboHRo+HXv4aBA0MdlYSSErRW7oNNH3DNO9dQWFbIJ5s/oaSihLySPPYX\n7afCXcH8da1zBQRv9axDfAc6xndUFU1avZgYuOYaa9Y8/nh44AFISDjy7VZUwKRJ1scNYMsWePxx\nG4Dg78034dln/397dx4fVXk2fPwXwr4IAoJbFXADt7rXWvsU61a1iisuuFXt8uhbbRVr0aoUVLTq\nS8Wi4oLIi9rivgEqrRFbEAUEREEQBAKigpGwk23eP+4Jk4RQSJjknMz8vn7ymTNnzpy5DofBK/d1\nLyFJKygIpd4FC7b/8+MiJwdOPx1WrgzlzZ/9LOqIFDX7oGWxotIiTn/2dJasWkK/Y/tx3v7nsXzd\n8krHtGnahrbN227hDJnr7n/fzd9n/Z3mjZsDYWTrJQdfwg3H3BBxZFLmGTcORo+GK6+EYcPg17+G\nH/2o8jEbNsDAgZCbG1rzjj8eeveOJt66UFAQWiiPPx7WrYOpU0Pi2qZN1JEpHWrTB80ELYu9+tmr\nDHx3IO1atKOotIgxF42hVdM0/EqcAdYWrWXlhpWV9rVr3s4/H6kW1q6t3NpW9TmEvnVjxoRWunPP\nrf48y5bBr34Vtl95JbM60E+ZAosWwTnnhNbDp58OI3UPPDDqyJQODhLQNisqLWLoh0Np06wNzRs3\nZ13xulotRVWWKCMTk+VWTVux2w67VfoxOVMc5OfDoEGpPlhjxsCLL0YbU1XLlqX6pxUVQZ8+8MYb\n4fnLL4eWoYoWLIBJk+DQQ8P1lJc7KyoogAEDQqvZwQenyp2Z4ogjQnIGodx58cUmZ9nOBC1LTV4y\nmeVrl7O+ZD0r1q0gkUjUqr/ZXe/dxSNTHqmDCCVVZ9ddoUkTuOOOkOy88AIcc0zUUVW2Zk0YRDB5\nMsyYERKpYcPCCghjxsD116eOLSkJfc5+/etUAnbvvZv3QcvLg+OOg0suCf3fCgtDi5OUqSxxZqmS\nshKWrlpaaV/LJi1rtHpAfmE+Z48+myaNmvD6Ra/TvkX7dIcpqRqlpXDmmWF76FDYY49o46nOvHmp\nROy++0JJ8r33wrQgVTvAr1sHLVtu+TmEhK3iPN9Vn8dJYWFqSpLS0nA99iXLbpY4tc0aN2rMnu32\nrPRT06Wdhn80nEY0orismGc+fqaOIpVU1ZtvQseO0L07PP54PKecWFmhC+crr8Dnn8Mf/wijRm0+\nv1fVZKzqc9g8GYtrcvbNN2EprnnzQnI2eHDoTybVlAmaaiW/MJ835r1Bh5YdaN+iPc98/AwF6wui\nDkvKeIsWhbLmoEFw991hAfXRo9P7GWVl8NxzoTM/hJLlCy9se5+vGTNCifO++0KMTz8NF1wQRmbe\neiv85z/pjTdOOnUK85jdfnuY02zVKrjiiqijUkMU099BtsoSZ8Tu+fc9jJg+gh2a7wDAqg2r6HtM\nX6487MqII5My3/r1qQlqS0vDT9Om6Tt/IhH6jM2fH5Z2GjQIDjooJBrb0nJVWBiWoNp77/B8wQJo\n3x7atUudv7Aw9byoCIqL0zOvWhyUlsJFF4XS5qBBdvaX02yoHs1ZMYf8wvxK+/ZqvxfdduwWUUSS\n0imRgPvvh3ffhZNOCmW7dJUVFy6E224L85rtsksY1bnffiGpaejKy5qrVoU/t2HDwrXus0/UkSlK\ntUnQGtdNKMp03Tt2p3vH7lGHIamOrF0LS5aE7UWLQmtQulq4unSBq64KfdKaNYMDDoDzz0/PuaP2\n3XdhMt0//Sm0ajZtCtOnm6Cp5mxBkyRVUlYGN94I++8Pv/gFPPoofPFFKNela3LYoqLUvF9DhkDX\nrjU/R0lJKI2Wl3vXrg0DDOI6gEDZyxKn0mr4R8M5Ze9T2KXNLlGHIqmezZsX+pDl5IRy5/z5qT5l\n26uoKJQ1W7cOE7Q++WQod+65Z83O8+ab8K9/Qf/+sHEj3HJL6Cd3+OHpiVNKF0ucSpvZy2dz/6T7\nyS/M5/aet0cdjqR6VrEkl5OTvuQMwgCCnXcOyzbl5oaf2bNrnqCdeGJIJH/3u9CSdsIJJmfKHLag\nqVrXjbuOifkTySGHF89/kd132D3qkCRlqeJiGDEiDCJo1Qq++greeiusKrByJVx6aThu9OhUuVOK\nEyeqVVrMXj6biYsn0qlVJ3LI4YlpT0QdkqQs1jhZ67n11jBlxy23hIl6CwtDZ/wLL4STTw6lzvXr\nIw1VShsTNG3mkamPsL5kPas2rqJxbmNemvPSZlNqSFJ9yckJoz7bt4frrgtLRZ16KnzyCRx7bGhZ\nu/rqMNBg/vyoo5XSwz5o2kzbZm05oNMBdGzREYDcRrmsLV4bcVSSskkiEVrDypd9ys8Py0U1aQKT\nJoUE7Uc/Cj8QRpf+5jfRxSulmy1o2sw+7fchvzCfPx/3Z4aeNpQhpwzZNOdZbfv+rd64mrLENq4T\nIynrffop/P73UFAQlpo680z43vfCklM9eoQpP+yKrEzmIAFVsrZoLac+fSrfbfiOKw69gt8d/btN\nr5UlyrjmjWu47JDLOHr3o7f5nKVlpVz0wkWcsd8Z9Dm4T12ELSkDPfccjB0bpuNo0wYGDAgjPhOJ\nMFBgF2cAUgPhIAFttxdnv8i6knXs2mZX/vHJP1ixbsWm195f8j4TFk1g8KTBm7WG/beEecKiCcxZ\nMYfHpj3GuuJ1NY5p4cqFjJwxssbvk9Sw9eoVpuT44oswlUZubtifk2NypsxngqZN1hat5fFpj9Ou\neTua5DahtKyUUTNHAaH1bMjkIbRv2Z4F3y1gYv7ETe9bvXE1fV7sw9JVSzc7Z2lZKQ9+8CDtWrRj\nbfFaXp7zco3jGjJ5CPdNvI+FKxfW+tokNSxFRXDXXWEQQJ8+YbRmQUHUUUn1xwRNm8z8eiYbSjZQ\nsK6AL1d9SUlZCXkL84DQeja/YD5tm7WleZPmPDj5wU2taM9/+jyTl05mxPQRm51zwqIJLC5cTJum\nbWjbrC2PTn20Rq1on634jPcWvUezxs14fOrj6bhMSQ3AokXQti307QsXXBAmoZ06tfpjCwtT28XF\nYcknqaGzD1oD8X7++3yx8gsuPOjCOv2c6jryN8ppxKUvXcqUL6fQqmlYLXlN0Rqe7PUkB+x0AKc+\nfSpNGzdl9cbVvHT+S+y2w26b3nvFK1cwbdk0WjZpuel9dx1/Fz/f9+fbFM/1b17PxPyJtG/RnhXr\nVjD6vNF0addl+y9UUkZYtAhuuy30T5sxAz78EPbbD3r3Dv3XTj89feuHSrXlWpwZqrSslHNGn8Oy\nNcsYc9EYOrTsUO8xTFg0gZUbVlbad9RuRzF23lge+vAhdm6zM1+v+Zpe+/Xilv+5ZdMxiwsXU7ih\nsNL7urTrQptmbbb6mXO/ncu5o8+lTbM2NMppRMG6As7sfiZ3Hn9nei5KUkZ4992woHt+PpSWwksv\nwd/+BjvsEEaClvddk6Jigpahxs8fT79/9oMcuOTgS7j2B9dGHRIA64vXc/KokyncUEjT3KaUJkoB\nGNtnLJ1bd97u88/8eiZPTHuCBKl7vUfbPeh7TN/tPrekzFFcDGefHbaPPDK0oh1xROi3ZnKmODBB\ny0DlrWffbfiOFo1bULixkNcvfD2SVrSqikuLeXvB2xSXFm/a1yinET/t+tNNpVBJqkvFxXD33SER\nO+wwuOkm2GknOOAAuPPOMEWHFLVMSdD6A1cBy5PP+wHjqhyTNQna+PnjuXbctbRr3g6AgvUFXH3k\n1bVuRftz3p/5+b4/5/BdD09nmJIUiS+/DJPX/uY3YdTnV1+FpaCWLw+T3f7lL6m1PKWo1CZBi+Nf\n2wTwf5M/Wa95k+ac0+OcSvs6t6pd+XDWN7N4/tPnmfvtXEadPar8L4wkNVi77gq//W3Y7t0b9t03\nNZntp5/WX3JWWAhNm0Lz5mGetq+/hk6dwmv+U6vaiGOCBvFs2YvEsXscy7F7HJuWcz304UO0btqa\nuQVzmbx0co1WA5CkuOvRI7WdkxPKnPVl7FiYPh1atIBDDoHnn4drrgkDFgYODMmbVBNxHXz8W2AG\n8ATQLuJYMsKsb2bx4dIP6dCyA81ym/Hg5Adrva6mJKmy3r1ht93gP/8JLXqHHgqPPAInnWRyptqJ\nqgXtbWDnavbfAjwMDEg+HwjcD1xZ9cD+/ftv2u7Zsyc9e/ZMd4wZ5eEpD1O4sZBE8r+py6bywdIP\n+MHuP4g6NElq8Bo1gpNPhrfeCmXX8ePD5LrHHx91ZIpCXl4eeXl523WOuJcSuwCvAQdV2Z81gwTS\n5dXPXuWbtd9U2ndcl+PYq/1eEUUkSXVvxAjo2hV+8pOwwsB994XSY8eO6f2cefPCZLm9e8Mdd4Q5\n2Dp2DKNKDzkkvZ+lhidTRnHuAixLbv8eOBK4qMoxJmiSpK1avBhuvRXOPz+0avXoAVddlf6O+//8\nZ5jS47XXQjK4dGlIDJ99FoYMscyZ7TIlQRsJHEIYzfkF8Gvg6yrHmKBJkrbJ7Nnwhz9Au3YwcmT1\nydmLL4YRoAceGOZWe/xxuPDC8J6aKC6GJk22/Ly+LVwIe+4ZrjmRCEtjdekSXTzZqjYJWhwHCVwK\nHAx8HziTzZMzSZK2ydq18NhjocyYSMATT6Re+/ZbWLAgbO+9NwwaBB99FOZTKyys3SS3VZOxKJOz\nsjJ48MFQ5k0kwuODD4b9ir84tqBtC1vQJElbNXRoKC9edRVMmwYXXBAmr/2f/4Gbb4bTToMzzgjH\nTp0K/ftD27YhmcmECW5Xrw4l3vnzoVu30D+uzdaXQlaaZUoLmiRJaXHllak+Z4cfHuYrGz4cLr88\nTIFRnpwVF8Prr4fkpbQU5syJNOy0ad0a9kqOBdtrL5e+akhM0CRJGat8Zv9yO+5Y/fZjj0GzZqGP\nWr9+YX3Pb79NvV5xu6QklEBrauVKeOqpkABCSALHVV3I8L94/vnQnw6gqAgefhhWrdry8eVlzfnz\n4dFHQzm3vNyp+DNBkyRlhYKCUNa8/PKQ3IwcCe+8E1678ELo2zeUNQ8+GP76V+jQIbz21Vdw3XUh\nOSopgXvugeeeq/nnt2wZkqXBg8MyVHfcERZ231bduoUF4GfODI9r1kCrVls+PpEIKxsMHAi77BIe\nW7QwQWso7IMmScoK69bB5Mlw3HHh+ZIlIWk7+OCtv3fqVLj33tDpv3v3ML9ZbfqoFRXBJZeEWG67\nDY48smbvnzw5ldg99lhYd1TxZx80SZK2oGXLVHIGsPvu25acAXz/+6Gf2sqV0KtX7QcQLFiQasF6\n991UuXNbFBXBmDGhH9n69TB3bu1iUMNggqZY+HL1l9yedztlCcd/S4qX8rLmYYeFEZF3353qC1YT\n33wTWr9uugleeCH0HxsxYtvf/8gjITkbNQpuvDGUOb/7ruZxqGGwxKlYGPjuQEbNHMWw04fRs0vP\nqMORpE1WrIDRo+FXvwotZ1OnhvJor141O08iAV9+GRZVh9AiVli47f3QCgrCFCDlZc0VK9K/ZJXq\nRqasJLAtTNAyyNJVSznrH2fRJLcJnVt15vnez9Mox8ZdSVJmsA+aGqThHw0HYMfmO5JfmM+ERRMi\njkiSpGiZoClSS1ct5YXZL1CWKGP5uuWsK1nHkMlD7IsmKTa+/jqUI8uNHAnz5oXt0tIwt9nq1dHE\npsyVAQtZqCFrlNOIPgf3qZSQtWma/nVI3pr/Fvt22Jcu7bqk/dySMttLL4UkrV+/sFbns8+GVQcG\nDoSXX4YNG8Ikt1I62QdNGa9gfQGnjDqFo3c/mgdOeSDqcCQ1MCUlcP/98O9/h4lhBw6E5cvD4upt\n24alo5o2jTpKxZl90KRqPPvxs5QmSpm4ZCKzl9dibLykrNa4Mfz4x2G7SZMwf9p774XnxcWwaFF0\nsSlzmaApoxWsL+Dpj5+mQ8sO5Obk8siUR6IOSVIDM3lyWBrqvvvgwAPhvPPCMksvvADXXw8DBlRe\nq1NKB/ugKaM9+/GzrClaQ/PGzWnRpAV5i/KYvXw2PXbqEXVokhqIFSvCskz77AM33BDWs+zTJ5Q1\nf/CDMK9Z+/ZRR6lMYwua6t2C7xbw1/f/Wi+ftWrjKvbfaX86tepE51ad2b/j/ny15qt6+WxJmeG0\n00JyBqHcee21qYXUIZQ8v/gizPBf3j16/HiYNKn+Y1XmsAVN9W7oB0MZ9/k4Tuh2Agd2OrBOP6vf\nj/vV6fklZbbSUvjkk9SanYWFoZzZrVvl4zp1gilTYONG2GMPePrpsBSTVFu2oKlezf12LhMWT6BN\nszY89OFDUYcjKYMlEmFR8XJFRTVbnBzCWpeDB8Pbb4fk7JZbQiJWVevWYZ3Nl1+GIUNCcla+pJNU\nGyZoqlfDpgwjNyeXji078uHSD5n1zayoQ5KUoaZPD4uKr1wZWrYGDIC33qrZOTp2DInXww/DxRfD\n0UeHQQLVmTQpzIfWsiWMG5cqd0q1YYKmejPv23m8Of9NAL5d/y1ri9faiiapzhxyCPzwhyFJ69s3\n9Bs7+eSan6d169R2586QU81sVrNmhbLmAw/AE0/Axx/D2LG1j11yolrVm3nfzuPpj5+m4r3r3Loz\nVx95dYRRScpkGzfCueeG7ZEjYccda/b+Vavg5ptDy9lxx4US58UXwwknVD6urCyUQ8sHD6xZA7m5\nYcSnVJuJak3QJEkZaePGMOt/hw6hVDl5cihXtmu37edYvz5MSnviiaHlbOlSWLYMjjii7uJW5jFB\nkyQp6ZNP4F//gmuuCcnVM8+EKTF+8pOoI1O2MUGTJCkNJk0KU2u0ahXKl+++GxK7RvbcVi24Fqck\nSWnw8cdh9YA1a+BvfwujP0tKoo5K2cQWNEmSqkgkYNgweOMN6NoV7rnHDv+qPVvQJElKg0QiTGwL\nocRZVhZtPDXx5Zep7bIymDu38uurVtVvPKodEzRJkqp4+OEwWnP0aDjooFDu3Lgx6qi2bsMGuPVW\nePPNkJwNHQqXXQavvx5eHz8ebrqp5isqqP5Z4pQkqYpp06BHj1DWTCTg/ffDXGjVTVIbN8uWQb9+\nYc3Q/feH//3fML1I69ZhVQWXoap/ljglSUqDww5L9TnLyQkrEjSE5AzCagflCdgxx0CXLvDTn8L8\n+SHJNDlrGEzQJEnKEGVl8NBDYcTpAw/AK6/AoEFhFOqAAWGh9/JyZ3kfO6jc507xYIImSVKGKCkJ\nLX/9+0O3bqG0OXt2KGseemh4nDMnrIhwzTWhHJpIhPVDR4yIOnpV1EAabDdjHzRJkrbDuHFhEETX\nrmEd0QEDKi8Mr/RxJQFJkrRNEgk477wwOnXwYNh776gjylwOEpAkSVtVXtbcYw+4/HK4665Q7lR8\nNI46AEmSVL+++QYWLkyVNVu1CgMJLrss6shUzhKnJElZKJGoPHVI1edKH0uckiRpm1RNxkzO4sUE\nTZIkKWZM0CRJkmLGBE2SJClmTNAkSZJixgRNkiQpZkzQJEmSYsYETZIkKWZM0CRJkmLGBE2SJMVC\nUVFqu6QEysqiiyVqJmiSJClys2ZB376wenVIzv7yF3j99aijik5DXdjBtTglScogiQSMHAkffABt\n20KLFvDHP0KTJlFHtv1qsxanCZokSYqFkhI466yw/dRT0L59tPGkiwmaJElqkMrLmqWlsOuuMGMG\n3HkntGkTdWTbrzYJmn3QJElS5ObNg5ycUNa84go4/HCYODHqqKJjC5okSVmgoAAGD4Y//CG0Sr37\nLnz+OVx5ZdSRpSQSIUmrut3Q2YImSZKqteOOsPfecMstYXTk8OFw4olRR1VZxYQsU5Kz2mqol28L\nmiRJNZRIhBaz5cth0CA48MCoI8oOtqBJkqQtmjAhdMI/6ih49NEw55jiyQRNkqQsUFAQpq4YOBD+\n9KfQCf/JJ6OOSltiiVOSpCxRVARNm4btRAKKi1PPVXecB02SJClm7IMmSZIarKptL9ncFmOCJkmS\nYuGuu2Dy5LC9eDHccANs3BhtTFFpHHUAkiRJAL17w4ABsGQJvPYaXHYZNGsWdVTRsAVNkiTFwj77\nhGWeRoyAHj3guOOijig6JmiSJCkWFi8OU4GccQbMmpUqd2ajqBK084BPgFLgsCqv9QPmAXOAk+o5\nLkmSFJHhw0NZ85e/hNtuC8latvZBi2qaje5AGTAMuAGYlty/P/AMcCSwGzAe2Dd5bEVOsyFJUoYp\nLYXc3C0/b6ga0jQbc4C51ezvBTwLFAMLgc+Bo+ovLEmSFJWqyVgmJGe1Fbc+aLsCSyo8X0JoSZMk\nScoadTnNxtvAztXsvxl4rQbnqbaW2b9//03bPXv2pGfPnjU4pSRJUt3Iy8sjLy9vu84R9VJP71C5\nD9ofk493Jx/HAbcDVcdx2AdNkiQ1CA2pD1pFFQN+FbgAaAp0BfYBPogiKEmSpKhElaCdBeQDRwNv\nAGOT+z8FRicfxwJXs4USpyRJUqaKusRZW5Y4JUlSg9BQS5ySJEmqwARNkiQpZkzQJEmSYsYETZIk\nKWZM0CRJkmLGBE2SJClmTNAkSZJixgRNkiQpZkzQJEmSYsYETZIkKWZM0CRJkmLGBE2SJClmTNAk\nSZJixgRNkiQpZkzQJEmSYsYETZIkKWZM0CRJkmLGBE2SJClmTNAkSZJixgRNkiQpZkzQJEmSYsYE\nTZIkKWZM0CRJkmLGBE2SJClmTNAkSZJixgRNkiQpZkzQJEmSYsYETZIkKWZM0CRJkmLGBE2Sr5zp\nqgAABzhJREFUJClmTNAkSZJixgRNkiQpZkzQJEmSYsYETZIkKWZM0CRJkmLGBE2SJClmTNAkSZJi\nxgRNkiQpZkzQJEmSYsYETZIkKWZM0CRJkmLGBE2SJClmTNAkSZJixgRNkiQpZkzQJEmSYsYETZIk\nKWZM0CRJkmLGBE2SJClmTNAkSZJixgRNkiQpZkzQJEmSYsYETZIkKWZM0CRJkmLGBE2SJClmTNAk\nSZJixgRNkiQpZkzQJEmSYsYETZIkKWZM0CRJkmLGBE2SJClmTNAkSZJixgRNkiQpZkzQJEmSYsYE\nTZIkKWZM0CRJkmLGBE2SJClmokrQzgM+AUqBwyrs7wKsBz5K/jxU75HFWF5eXtQhRMLrzi5ed3bx\nurNLtl53bUSVoH0MnAVMqOa1z4FDkz9X12dQcZetf7G97uzidWcXrzu7ZOt110bjiD53TkSfK0mS\nFHtx7IPWlVDezAOOjTYUSZKk+pdTh+d+G9i5mv03A68lt98BbgCmJZ83BVoB3xH6pr0MHACsrnKO\nz4G90hyvJElSXZgP7B11EDXxDpUHCdT0dUmSpIwThxJnxVa8jkBucrsbsA+woN4jkiRJykJnAfmE\nKTW+AsYm958DzCL0QZsKnBZJdJIkSZIkSVJDdS8wG5gBvAi0rfBaP2AeYQqPk+o/tDqVrRP7bum6\nIbPvd0X9gSWk7vHPIo2m7v2McE/nATdFHEt9WgjMJNzjD6INpU4NB74mzIVZrj1hUNlc4C2gXQRx\n1bXqrrs/mf/d/h6hL/knhOrYtcn9mX7Pt3Td/cnge34iqX5zdyd/APYHpgNNCEnL58Sjf126dAf2\nZfNBE12o/IXPNFu67ky/3xXdDlwfdRD1JJdwL7sQ7u10oEeUAdWjLwj/08p0PyZMQl7x362/AH9I\nbt9E6t/1TFLddWfDd3tn4JDkdmvgM8J3OtPv+Zauu0b3vKH9T+1toCy5PRnYPbndC3gWKCb8Jvo5\ncFR9B1eH5hB+08g2W7ruTL/fVdXldDhxchThXi4k3Nu/E+51tsiG+/weYRqlis4AnkpuPwWcWa8R\n1Y/qrhsy/55/RfhFC2ANoQK2G5l/z7d03VCDe97QErSKrgDGJLd3JTQblltC6g8j02XjxL7Zdr9/\nSyjrP0HmlQIq2o0weKhcpt/XihLAeGAK8MuIY6lvnQnlP5KPnSOMpb5ly3cbQsv4oYTGlWy6510I\n1/1+8vk23/M4JmhvE5qBq/6cXuGYW4Ai4Jn/cp5EXQVYR7bluqv6klDrPpTQbPoM0KZuw0y72lx3\ndRra/a5oS38GZwAPE5LwQ4BlwP0RxVgfGvI93F4/InyPTwGuIZTEslGC7Pl7kE3f7dbAC8B1bD7x\nfCbf89bA84TrXkMN73lUa3H+Nydu5fXLgVOB4yvsW0pIVMrtntzXkGztuqtTlPyBsBrDfMLccdO2\n+I74qc11Z8L9rmhb/wweJ7UKRyaqel+/R+WW0ky2LPm4HHiJUO59L7pw6tXXhD47XwG7AN9EG069\nqXidmfzdbkJIzv4fYXUgyI57Xn7do0hdd43ueRxb0P6bnwE3EvqlbKiw/1XgAsJSUV0JSUqmjoTK\n1ol9K153Nt3vXSpsn0VmDwqZQriXXQj39nzCvc50LUm1fLcijErO5Ptc1avAZcnty0j9zyzTZcN3\nO4dQyvsU+GuF/Zl+z7d03Rl9z+cBi6h+WombCR2M5wAn139odSpbJ/bd0nVDZt/vikYSpl+YQfhH\nLJP7akAo8X1GuLf9Io6lvnQldCieTvg+Z/J1P0vomlFE+G7/gjB6dTyZO+UCbH7dV5Ad3+1jCQP7\nplN5aolMv+fVXfcpZMc9lyRJkiRJkiRJkiRJkiRJkiRJkiRJkiRJkiRJNdML6LGd5+gOTCJMon3D\ndkckKavlbv0QScp4/YDvgNk1eE8uldcQzAEmAgWEJG1S2qKTlHUa2lJPklTRIODqCs/7k2q9upGw\nBNiM5P5ylyb3TSfM7P1D4HTgXsKM390Iixm/nzzuRVIznecBg4EPgWurxLKcsFxV8XZekyRJUoN2\nCCFpKvcJsBthPcthyX2NCIsS/xg4gLCUVPvka+WJ15PA2RXOMzN5PMCfCUkZwDvA37YS0+1Y4pS0\nnRpHHYAkbYfpQCfCIsSdCGXKpcDvCUnaR8njWgF7Jx9HE8qQACsrnCsn+dg2+fNe8vlTwHMVjvtH\nWq9AkqphgiapoXsOOBfYGfh7hf2DgEerHPt/SCViVSW2sL/q8WtrGqAk1ZR90CQ1dP8ALiQkaeUt\nXW8CVxBazCCUPXcC/gWcR6rEuWPycTWwQ3K7kNASd2zy+SVULqNuzZYSQEmSpKwyE/hnlX3XJvfP\nBP4DdE3uvxT4mFAeHZ7cdwyh/9pUwiCB7xNGYZYPEmibPO4d4LAtxLAzkE8qwVsMtN6Oa5IkSZIk\nSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSao3/x9Cx3v+AzwHUgAAAABJRU5ErkJggg==\n", + "text": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 2, + "metadata": {}, + "source": [ + "Simple Supervised Classification" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top]](#Sections)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 3, + "metadata": {}, + "source": [ + "Linear Discriminant Analysis as simple linear classifier" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top]](#Sections)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "The LDA that we've just used in the section above can also be used as a simple linear classifier." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "# fit model\n", + "lda_clf = LDA()\n", + "lda_clf.fit(X_train, y_train)\n", + "LDA(n_components=None, priors=None)\n", + "\n", + "# prediction\n", + "print('1st sample from test dataset classified as:', lda_clf.predict(X_test[0,:]))\n", + "print('actual class label:', y_test[0])" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "1st sample from test dataset classified as: [3]\n", + "actual class label: 3\n" + ] + } + ], + "prompt_number": 15 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Another handy subpackage of sklearn is `metrics`. The [`metrics.accuracy_score`](http://scikit-learn.org/stable/modules/generated/sklearn.metrics.accuracy_score.html), for example, is quite useful to evaluate how many samples can be classified correctly:" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "from sklearn import metrics\n", + "pred_train_lda = lda_clf.predict(X_train)\n", + "\n", + "print('Prediction accuracy for the training dataset')\n", + "print('{:.2%}'.format(metrics.accuracy_score(y_train, pred_train_lda)))" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "Prediction accuracy for the training dataset\n", + "100.00%\n" + ] + } + ], + "prompt_number": 17 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "To verify that over model was not overfitted to the training dataset, let us evaluate the classifier's accuracy on the test dataset:" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "pred_test_lda = lda_clf.predict(X_test)\n", + "\n", + "print('Prediction accuracy for the test dataset')\n", + "print('{:.2%}'.format(metrics.accuracy_score(y_test, pred_test_lda)))" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "Prediction accuracy for the test dataset\n", + "98.15%\n" + ] + } + ], + "prompt_number": 18 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "**Confusion Matrix** \n", + "As we can see above, there was a very low misclassification rate when we'd apply the classifier on the test data set. A confusion matrix can tell us in more detail which particular classes could not classified correctly.\n", + "\n", + "
| predicted class | \n", + "\t\t||||
| class 1 | \n", + "\t\tclass 2 | \n", + "\t\tclass 3 | \n", + "\t||
| actual class | \n", + "\t\tclass 1 | \n", + "\t\tTrue positives | \n", + "\t\t||
| class 2 | \n", + "\t\tTrue positives | \n", + "\t\t|||
| class 3 | \n", + "\t\tTrue positives | \n", + "\t|||
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "" + ] + }, + { + "cell_type": "heading", + "level": 3, + "metadata": {}, + "source": [ + "Classification Stochastic Gradient Descent (SGD)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top]](#Sections)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Let us now compare the classification accuracy of the LDA classifier with a simple classification (we also use the probably not ideal default settings here) via stochastic gradient descent, an algorithm that minimizes a linear objective function. \n", + "More information about the `sklearn.linear_model.SGDClassifier` can be found [here](http://scikit-learn.org/stable/modules/generated/sklearn.linear_model.SGDClassifier.html)." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "from sklearn.linear_model import SGDClassifier\n", + "\n", + "sgd_clf = SGDClassifier()\n", + "sgd_clf.fit(X_train, y_train)\n", + "\n", + "pred_train_sgd = sgd_clf.predict(X_train)\n", + "pred_test_sgd = sgd_clf.predict(X_test)\n", + "\n", + "print('\\nPrediction accuracy for the training dataset')\n", + "print('{:.2%}\\n'.format(metrics.accuracy_score(y_train, pred_train_sgd)))\n", + "\n", + "print('Prediction accuracy for the test dataset')\n", + "print('{:.2%}\\n'.format(metrics.accuracy_score(y_test, pred_test_sgd)))\n", + "\n", + "print('Confusion Matrix of the SGD-classifier')\n", + "print(metrics.confusion_matrix(y_test, sgd_clf.predict(X_test)))" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "\n", + "Prediction accuracy for the training dataset\n", + "99.19%\n", + "\n", + "Prediction accuracy for the test dataset\n", + "100.00%\n", + "\n", + "Confusion Matrix of the SGD-classifier\n", + "[[14 0 0]\n", + " [ 0 18 0]\n", + " [ 0 0 22]]\n" + ] + } + ], + "prompt_number": 22 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Quite impressively, we achieved a 100% prediction accuracy on the test dataset without any additional efforts of tweaking any parameters and settings." + ] + }, + { + "cell_type": "heading", + "level": 3, + "metadata": {}, + "source": [ + "Decision Regions" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "sgd_clf2 = SGDClassifier()\n", + "sgd_clf2.fit(X_train[:, :2], y_train)\n", + "\n", + "x_min = X_test[:, 0].min() \n", + "x_max = X_test[:, 0].max() \n", + "y_min = X_test[:, 1].min() \n", + "y_max = X_test[:, 1].max() \n", + "\n", + "step = 0.01\n", + "X, Y = np.meshgrid(np.arange(x_min, x_max, step), np.arange(y_min, y_max, step))\n", + "\n", + "Z = sgd_clf2.predict(np.c_[X.ravel(), Y.ravel()])\n", + "Z = Z.reshape(X.shape)\n", + "\n", + "# Plots decision regions\n", + "plt.contourf(X, Y, Z)\n", + "\n", + "\n", + "# Plots samples from training data set\n", + "plt.scatter(X_train[:, 0], X_train[:, 1], c=y_train)\n", + "plt.show()" + ], + "language": "python", + "metadata": {}, + "outputs": [] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 2, + "metadata": {}, + "source": [ + "Saving the processed datasets" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top]](#Sections)" + ] + }, + { + "cell_type": "heading", + "level": 3, + "metadata": {}, + "source": [ + "Pickle" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top]](#Sections)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "The in-built [`pickle`](https://docs.python.org/3.4/library/pickle.html) module is a convenient tool in Python's standard library to save Python objects in byte format. This allows us, for example, to save our NumPy arrays and classifiers so that we can load them in a later or different Python session to continue working with our data, e.g., to train a classifier." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "# export objects via pickle\n", + "\n", + "import pickle\n", + "\n", + "pickle_out = open('standardized_data.pkl', 'wb')\n", + "pickle.dump([X_train, X_test, y_train, y_test], pickle_out)\n", + "pickle_out.close()\n", + "\n", + "pickle_out = open('classifiers.pkl', 'wb')\n", + "pickle.dump([lda_clf, sgd_clf], pickle_out)\n", + "pickle_out.close()" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 24 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "# import objects via pickle\n", + "\n", + "my_object_file = open('standardized_data.pkl', 'rb')\n", + "X_train, X_test, y_train, y_test = pickle.load(my_object_file)\n", + "my_object_file.close()\n", + "\n", + "my_object_file = open('classifiers.pkl', 'rb')\n", + "lda_clf, sgd_clf = pickle.load(my_object_file)\n", + "my_object_file.close()\n", + "\n", + "print('Confusion Matrix of the SGD-classifier')\n", + "print(metrics.confusion_matrix(y_test, sgd_clf.predict(X_test)))" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "Confusion Matrix of the SGD-classifier\n", + "[[14 0 0]\n", + " [ 0 18 0]\n", + " [ 0 0 22]]\n" + ] + } + ], + "prompt_number": 26 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
" + ] + }, + { + "cell_type": "heading", + "level": 3, + "metadata": {}, + "source": [ + "Comma-Separated-Values" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top]](#Sections)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "And it is also always a good idea to save your data in common text formats, such as the CSV format that we started with. But first, let us add back the class labels to the front column of the test and training data sets." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "training_data = np.hstack((y_train.reshape(y_train.shape[0], 1), X_train))\n", + "test_data = np.hstack((y_test.reshape(y_test.shape[0], 1), X_test))" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 21 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Now, we can save our test and training datasets as 2 separate CSV files using the [`numpy.savetxt`](http://docs.scipy.org/doc/numpy/reference/generated/numpy.savetxt.html) function." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "np.savetxt('./training_set.csv', training_data, delimiter=',')\n", + "np.savetxt('./test_set.csv', test_data, delimiter=',')" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 22 + } + ], + "metadata": {} + } + ] +} \ No newline at end of file diff --git a/python_true_false.ipynb b/tutorials/python_true_false.ipynb similarity index 96% rename from python_true_false.ipynb rename to tutorials/python_true_false.ipynb index b2786a2..fa8c446 100644 --- a/python_true_false.ipynb +++ b/tutorials/python_true_false.ipynb @@ -1,6 +1,7 @@ { "metadata": { - "name": "" + "name": "", + "signature": "sha256:f6d59feb844af096c0581919fa052e8f29f4fa0aaa81752f5e50241ab084a0e9" }, "nbformat": 3, "nbformat_minor": 0, @@ -11,7 +12,12 @@ "cell_type": "markdown", "metadata": {}, "source": [ - "Sebastian Raschka, 03/2014 \n", + "[Sebastian Raschka](http://sebastianraschka.com) \n", + "last updated: 05/03/2014\n", + "\n", + "- [Link to this IPython Notebook on GitHub](https://github.com/rasbt/python_reference/blob/master/benchmarks/python_true_false.ipynb) \n", + "- [Link to the GitHub repository](https://github.com/rasbt/python_reference) \n", + "\n", "Code was executed in Python 3.4.0" ] }, @@ -392,8 +398,8 @@ "\n", "Possibly the best explanation of shallow vs. deep copies I've read so far:\n", "\n", - "*** \"Shallow copies duplicate as little as possible. A shallow copy of a collection is a copy of the collection structure, not the elements. With a shallow copy, two collections now share the individual elements.\n", - "Deep copies duplicate everything. A deep copy of a collection is two collections with all of the elements in the original collection duplicated.\"***\n", + "***\"Shallow copies duplicate as little as possible. A shallow copy of a collection is a copy of the collection structure, not the elements. With a shallow copy, two collections now share the individual elements.\n", + "Deep copies duplicate everything. A deep copy of a collection is two collections with all of the elements in the original collection duplicated.\"*** \n", "\n", "(via [S.Lott](http://stackoverflow.com/users/10661/s-lott) on [StackOverflow](http://stackoverflow.com/questions/184710/what-is-the-difference-between-a-deep-copy-and-a-shallow-copy))" ] diff --git a/tutorials/running_cython.ipynb b/tutorials/running_cython.ipynb new file mode 100644 index 0000000..2c21a11 --- /dev/null +++ b/tutorials/running_cython.ipynb @@ -0,0 +1,1002 @@ +{ + "metadata": { + "name": "", + "signature": "sha256:c53d1aaf41825ecf8aae8e9e7691d07f984de379cd765b3cabd973cfb29cc420" + }, + "nbformat": 3, + "nbformat_minor": 0, + "worksheets": [ + { + "cells": [ + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[Sebastian Raschka](http://sebastianraschka.com) \n", + "\n", + "[Link to this IPython notebook on GitHub](https://github.com/rasbt/python_reference/blob/master/tutorials/running_cython.ipynb)" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%load_ext watermark" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 1 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%watermark -d -m -v -p cython" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "06/07/2014 \n", + "\n", + "CPython 3.4.1\n", + "IPython 2.1.0\n", + "\n", + "cython 0.20.2\n", + "\n", + "compiler : GCC 4.2.1 (Apple Inc. build 5577)\n", + "system : Darwin\n", + "release : 13.2.0\n", + "machine : x86_64\n", + "processor : i386\n", + "CPU cores : 2\n", + "interpreter: 64bit\n" + ] + } + ], + "prompt_number": 2 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[More information](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/ipython_magic/watermark.ipynb) about the `watermark` magic command extension." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "I would be happy to hear your comments and suggestions. \n", + "Please feel free to drop me a note via\n", + "[twitter](https://twitter.com/rasbt), [email](mailto:bluewoodtree@gmail.com), or [google+](https://plus.google.com/118404394130788869227).\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "
\n", + "
\n" + ] + }, + { + "cell_type": "heading", + "level": 1, + "metadata": {}, + "source": [ + "Using Cython with and without IPython magic" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 1, + "metadata": {}, + "source": [ + "Sections" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "- [Bubblesort in regular (C)Python](#bubblesort-cpython)\n", + "- [Bubblesort implemented in Cython](#Bubblesort-implemented-in-Cython)\n", + " - [Naive Cython implementation - auto-guessing types](#Naive-Cython-implementation---auto-guessing-types)\n", + " - [Cython with explicit type-declarations](#Cython-with-explicit-type-declarations)\n", + "- [Speed comparison](#Speed-comparison)\n", + "- [How to use Cython without the IPython magic](#How-to-use-Cython-without-the-IPython-magic)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "" + ] + }, + { + "cell_type": "heading", + "level": 2, + "metadata": {}, + "source": [ + "Bubblesort in regular (C)Python" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#Sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "First, we will write a simple implementation of the bubble sort algorithm in regular (C)Python" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "def python_bubblesort(a_list):\n", + " \"\"\" Bubblesort in Python for list objects. \"\"\"\n", + " length = len(a_list)\n", + " swapped = 1\n", + " for i in range(0, length):\n", + " if swapped: \n", + " swapped = 0\n", + " for ele in range(0, length-i-1):\n", + " if a_list[ele] > a_list[ele + 1]:\n", + " temp = a_list[ele + 1]\n", + " a_list[ele + 1] = a_list[ele]\n", + " a_list[ele] = temp\n", + " swapped = 1\n", + " return a_list" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 3 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "python_bubblesort([6,3,1,5,6])" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "pyout", + "prompt_number": 4, + "text": [ + "[1, 3, 5, 6, 6]" + ] + } + ], + "prompt_number": 4 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 2, + "metadata": {}, + "source": [ + "Bubblesort implemented in Cython" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#Sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Next, we will speed things up a little bit via [Cython's C-extensions for Python](http://cython.org). Cython is basically a hybrid between C and Python and can be pictured as compiled Python code with type declarations. \n", + "Since we are working in an IPython notebook here, we can make use of the very convenient ***IPython magic***: It will take care of the conversion to C code, the compilation, and eventually the loading of the function. " + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%load_ext cythonmagic" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 5 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "First, we will take the initial Python code as is and use Cython for the compilation. Cython is capable of auto-guessing types, however, we can make our code way more efficient by adding static types." + ] + }, + { + "cell_type": "heading", + "level": 4, + "metadata": {}, + "source": [ + "Naive Cython implementation - auto-guessing types" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#Sections)]" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%%cython\n", + "def cython_bubblesort_untyped(a_list):\n", + " \"\"\" Bubblesort in Python for list objects. \"\"\"\n", + " length = len(a_list)\n", + " swapped = 1\n", + " for i in range(0, length):\n", + " if swapped: \n", + " swapped = 0\n", + " for ele in range(0, length-i-1):\n", + " if a_list[ele] > a_list[ele + 1]:\n", + " temp = a_list[ele + 1]\n", + " a_list[ele + 1] = a_list[ele]\n", + " a_list[ele] = temp\n", + " swapped = 1\n", + " return a_list" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 6 + }, + { + "cell_type": "heading", + "level": 4, + "metadata": {}, + "source": [ + "Cython with explicit type-declarations" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#Sections)]" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%%cython\n", + "import numpy as np\n", + "cimport numpy as np\n", + "cimport cython\n", + "@cython.boundscheck(False) \n", + "@cython.wraparound(False)\n", + "cpdef cython_bubblesort_typed(np.ndarray[long, ndim=1] inp_ary):\n", + " \"\"\" The Cython implementation of Bubblesort with NumPy memoryview.\"\"\"\n", + " cdef unsigned long length, i, swapped, ele, temp\n", + " cdef long[:] np_ary = inp_ary\n", + " length = np_ary.shape[0]\n", + " swapped = 1\n", + " for i in xrange(0, length):\n", + " if swapped: \n", + " swapped = 0\n", + " for ele in xrange(0, length-i-1):\n", + " if np_ary[ele] > np_ary[ele + 1]:\n", + " temp = np_ary[ele + 1]\n", + " np_ary[ele + 1] = np_ary[ele]\n", + " np_ary[ele] = temp\n", + " swapped = 1\n", + " return inp_ary" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 7 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 2, + "metadata": {}, + "source": [ + "Speed comparison" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#Sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Below, we will do a quick speed comparison of our 3 implementations of the bubble sort algorithm by sorting a list (or numpy array) of 1000 random digits. Here, we have to make copies of the lists/numpy arrays, since our bubble sort implementation is sorting in place." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "import random\n", + "import numpy as np\n", + "import copy\n", + "\n", + "list_a = [random.randint(0,1000) for num in range(1000)]\n", + "list_b = copy.deepcopy(list_a)\n", + "\n", + "ary_a = np.asarray(list_a)\n", + "ary_b = copy.deepcopy(ary_a)\n", + "ary_c = copy.deepcopy(ary_a)" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 8 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "import timeit\n", + "\n", + "times = []\n", + "\n", + "times.append(min(timeit.Timer('python_bubblesort(list_a)', \n", + " 'from __main__ import python_bubblesort, list_a').repeat(repeat=3, number=1000)))\n", + "\n", + "times.append(min(timeit.Timer('python_bubblesort(ary_a)', \n", + " 'from __main__ import python_bubblesort, ary_a').repeat(repeat=3, number=1000)))\n", + "\n", + "times.append(min(timeit.Timer('cython_bubblesort_untyped(list_b)', \n", + " 'from __main__ import cython_bubblesort_untyped, list_b').repeat(repeat=3, number=1000)))\n", + "\n", + "times.append(min(timeit.Timer('cython_bubblesort_untyped(ary_b)', \n", + " 'from __main__ import cython_bubblesort_untyped, ary_b').repeat(repeat=3, number=1000)))\n", + "\n", + "times.append(min(timeit.Timer('cython_bubblesort_typed(ary_c)', \n", + " 'from __main__ import cython_bubblesort_typed, ary_c').repeat(repeat=3, number=1000)))\n", + "\n" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 9 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%matplotlib inline" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 10 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "from matplotlib import pyplot as plt\n", + "import numpy as np\n", + "\n", + "bar_labels = ('(C)Python on list', \n", + " '(C)Python on numpy array', \n", + " 'untyped Cython on list', \n", + " 'untyped Cython on numpy array', \n", + " 'typed Cython with memoryview on numpy array')\n", + "\n", + "fig = plt.figure(figsize=(10,8))\n", + "\n", + "# plot bars\n", + "y_pos = np.arange(len(times))\n", + "plt.yticks(y_pos, bar_labels, fontsize=14)\n", + "bars = plt.barh(y_pos, times,\n", + " align='center', alpha=0.4, color='g')\n", + "\n", + "# annotation and labels\n", + "\n", + "for b,d in zip(bars, times):\n", + " plt.text(max(times)+0.1, b.get_y() + b.get_height()/2.5, \n", + " '{:.2} ms'.format(d),\n", + " ha='center', va='bottom', fontsize=12)\n", + "\n", + "t = plt.title('Bubblesort on 1000 random integers', fontsize=18)\n", + "plt.ylim([-1,len(times)+0.5])\n", + "plt.vlines(min(times), -1, len(times)+0.5, linestyles='dashed')\n", + "plt.grid()\n", + "\n", + "plt.show()" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "display_data", + "png": "iVBORw0KGgoAAAANSUhEUgAAA9sAAAHtCAYAAAAJNW1FAAAABHNCSVQICAgIfAhkiAAAAAlwSFlz\nAAALEgAACxIB0t1+/AAAIABJREFUeJzs3XmcHGWd+PHPAHIIgkREiAIRZSN4gBoRECGC+HM1KIgu\nCgIRZAOIgCeKRyZqVJBlvd0YhHgArgcIiYu7HBkximJQAQGDIAkg900MASHz++NbRdfUVHfP9HSm\nZvJ83q9XvzJdXf30862q7tT3OapAkiRJkiRJkiRJkiRJkiRJkiRJkiRJkiRJkiRJkiRJkiRJkiRJ\nkiRJkiRJGtf6gJuHuO5UYBVw2BDXn5etL40XvcQxu3XN9agyleF9/yQlYK26KyBJ0jg2lTjBLj4e\nBW4CzgBe1IXP6F+N6w+37DrtB8ysuxJtTAZOBS4FHiSOh3Z1PhT4I7ACuBOYC2zWZN2JwPeAe7L1\nfw+8vUtla+T6Gdl36gRM1iVJkiSgkWz/ADgoexwBfJVIcB5iZL1wfcDfhlmXQ4e4/jzGV8/2PMZ+\nfacDTwJLgIuJ+n66xfofyNa5FHgvMAt4BPgz8PTSuhOIY+Fhoof3vcDC7P3TR1j2eNHL2O3Z7gHW\nZWQdWUuJ/SVJkiQlbypx8v/Bitfen712wgjK78Nke6Ps33lEIjuWbQpsnP39Slon25sB/wB+SyRq\nuWnZ+z5eWv+UbPmbC8vWAn4H3AtsOIKyh+MZI3jvSPUydpPtbljK+Eu2exh47EkqcBi5JEmrxx3Z\nv48Vlk2l+bzOeTRPfp8PnE8MTX4IODdbVqWHSPRvIIa0LwGOHXq12RL4FnBLVve/A3OAZ5fWmwD8\nJzFk/lEi4VsMfLi03jrAicB1hfXOBV5SWm8SjWHXBwJXEqMDvkb04B6axVYcsj+UhoX9gF8Dy4me\n3UXAWyrWW5p9zouAnxM9yA8CPwaeM4TPAXggex8MTHKb1WsDIr7i0OMFRAPLu0vrHwTcmNUttyp7\n/wTgTSMou5lVwJnA3sR2ewS4IHttIvAfwJ+A+4l9ey3wUQafX07PynodcXzcBKwkjs2qfbgW0SBw\nc1buNUT8zbwMOA+4r1CPj1TUY15WjwnENI97iP11PnHcA8wArs/KuZ7qY6XKVAZ/t4vL3pPVayVx\nrH2k9P68ESF/zyoGNyxMyeK8JyvnL8BJwNoV9TkAuCqLYxnR6PP6ijoCrJeVc222/gPEft6pRYzv\no/Gdzr/zLya+L3/P6ncH0XjwJqRErVN3BSRJWgNsSGMu7AZEIjmbOCn+acX6zeZ1Vi3fiOjh/i3w\nMeBfgGOAXYCXA3eV1n8/sAXwX0RydBAxrH0C8Jk2cWwNXE6cH3yHSIq2A44mEqUpNJLJHwOvJRLz\nq4m4dwD2JOYt584C3gH8H/ANIql5X/Y5ryWStaL9snp8M3s8TJz8r5WtX0wUf9MmnmOArxNJ0ywi\nAZ4O/IxIquYW1u0Hnksk3OcSCdhO2XobA/+vzWcN16uyfy+veO13wDuJ4d4riG02kZiuULUuxL75\ncQdltzOFSNy+TSTeuZcB+xPb6ibgacC/Al8EtgWOqijr88D6xDHzOHFczSMaEYr78jTgOOCXREL/\nHOLYqRrlMSVb77FsnTuJBPlkYEeqGxZ+AdwKfIo4vo8jksvziOPj9Ky844CfEN+5pRXlVKn6Dh+V\nxXA60YBzSFa/24BzsnUOIRqv7iF+O3L3Zv++mdjWNxDfr/uB3Yjv9E7AvxXec2BW7l+J0QBPEgny\nvhV1fBqxPXYlrgfwVeCZwJFEI9UeRMNX0QnAs4hj4k5iWz6LSKxXEb89y4gGuinAzsD/VGwXSZIk\nqampDL5AWv74M3GSXrV+VW/ePAb3bPdly04rLd8vW/6tirIfIhKz3NOIBOtxIpls9XnnEyfPE0vL\nXwn8k8bFvjbJ3vv1ijiK9snWO6e0/GVZeZcVlk3K1n2MuNBYWVV9W9mU6M2+gcZQdIhh0DcSSfwm\nheVLs/LLFxz7era8vC/bmULrYeTziSRovYrX8iHjL8ye50PSv1Cx7tNpXDegk7JbWZWVs1fFa+s3\nec/3gCeIBp/c9KysKxnY0TOR6AE9u7BscrbuRQwcHfDyQn2Kvb2/Jo7t8kiJ/87WL9Z9Ho3RAEX/\nkS1fxsBj5aXZ8s/T3lQGf7fzZbcxcPj9BsDdDG4sWkr1MPL1ie9lH4N760/IPmPP7Pk6RM/yHQw8\nvjckGkXKdczn9u9TKvcZxPZYWBHPvQy+0N5bqP7+SElzGLkkSSM3hxii+XpiXuyJxMnohYx8fmk/\n0VtY9DMiidyvYv2zgNsLz/9J9JitQ6Nnq8omRN0vIJKXzQqPZcSJ+huydR8lkuJdgG1alLl/9u/s\n0vKriYRwd6JHrOjnxPDikdqHSES/SiTduUeyZRsR+6vo70RPZlGebAwlOR2O/CJlj1W8trK0znDW\n7WT9Vq6iOgFcWfh7XWLkxGbECIa1iAaCsm8SiXjuduI4Lm7bt2b/nsbAHtg/ZmUXE/DNiR7ZC4jG\nraL8mNufwb5cer4o+/e7DDxWriEaZUa6788kjrvco0QD2HZDfP8+RKzzaGzn/HFhtk7+3XwlMRJi\nHtHwlvsH0eNc9m5i5McfSuWuR1zkb3cGN9p8j0aPe+7B7N83Ue+8fmlMcRi5JEkj91cGJiT/Qwxt\n/S0xXPRdIyj7QaIXrOx6IjHZgDh5Ly6vWheaz/OG6FHsIa5c/d4m69yU/fs40aP2FWJe7XVE/D9j\n4HZ4PtETWVWn64jGgucTc21zN7So43DksV7b5LOL6+SqhinndSs3CoxUPoR7PQYnxeuX1imuW1Ze\nd7hlt9Nsf6xDTGs4FHgBg+eob1rxnqrtez+wVeH5ttm/f6lY93oaSSW03sd/IZL1qmO+XI8Hsn+r\n7mn/ICPf982Oq6GWu3327xlNXu8nknFoxFvVYFW1L7cnjol7WpS9GdEQ1aqcy4gkfDpwMHFbuouJ\nEQZV338pCSbbkiStHlcQvWKvKyxrdQ/eddq8XqWb98nOk6XvEz18VYpJ/Rxi2PmbiSGsbycuxPbf\njKxxYahJ4OrQ6mrn7S54Nly3Z2U+l8HJ2HOJIbm3F9bNl5fly4rJ0HDKbqfZ/jiN2N8/BD5LNAj9\nk+hZPZnq0ZPNtm+3t207zb43q6t+I72Kfv75H2bwNQ5yQ92fVWVfTfUdFXLlXuxmx8R04EvE3P3X\nAh8CPkE0zH2jw/pJ45rJtiRJq886DBxSeX/274SKdbetWAbRQ/gcBl8IbXsiwXm0tHyHijLyZa1u\nI3YjkYSsx9BvP3QncSG17xDJ1feJRPtUYn7u34grJe9ADMkt16mf6t7EKv0Mr3Eh74V/CQPnneaf\nDUO/rdrqcAVxEardKuqxC9EzmSc1dxDJ9K4V5eyS/bu4w7I7dQgxeqN8lfDhzm0vy/fb9gw+NsrH\ndv56eb42xFXle6h3Hw9Xs+M770leQfvvZr5NXlTxWtW1EG4gesUXtvj84bg2e5xKTE35HTENxmRb\nSXLOtiRJq0c+Z7h4Jd+biTmr5YsR7UYjaarysdLz/Ymk5mcV6x7MwB7QdYmLID1B3PqpqHhyfR8x\n/P1twKsryu1h4BXXy3N+V9FIqPPGhPOyf8v3dX4JcUGlRQwcQt7K8qwOVcOTq1xEzFN9P4MvkPZ+\nYg7tRUMsa3U4n2goOZaB52P7EkOBzyqtfw4xXHtaYdnaRCwPMPBqz8MtuxNPMPg8ckPiWBuJC4jj\n8oOl8l9BzLEvHrP5Rcb2JW47leuhccydx0DdHA0yUuW6LKd6aPn/ErF+jOrjfwMax/hionFmOnFV\n8dxGVF8h/nvExeya9WwP9bZ3mzL4eHiIuOjbBlRPgZDWePZsS5I0cq+kcYuh9YgT/38n5jZ/srDe\ncuLCRe8lrsD8S+IiSdOJC1HtWFH2vUQCPLGw/jFEr3Jvxfo3EL1J/5V93kHElbE/w8ChxjB4eOzR\nRAKcz7/8E3ECvS2RHH83K2dyVpdziV6sB4ieyKOInsRfZeVdDPyIuNXUpsTFz7Ygbv21gri10lBd\nnr3vm0Ri+U9iTvzSJus/RNzz+RvE9phH49Zf2xK39HqkyXs7tTGNmPIruu9J4xg4n0aDxL3EradO\nJbbTD4lGkg8Rc1zLF/H6InELtbOJIdy3E6MIXkkcT/8orDvcsjvxE2Ib/hC4hEjK3sPQG0+Kisfh\nEmKfHUv04p5L9Ly+jzgeX1567/HEsfir7H13EQ0SbyAaFcqjGkZ7yHor5bpcDhxBfMf+QjRgXUB8\nVw4lGteWEHO3byKS6RcRjW/7Ed/bJ4nh5mcRIxy+ky2bTuybSQxM8r9CNP59ibhy+0Ji+svWxP3V\nH6X6avRlhxENLfmt4P5JHPtvIKaWVF2sT5IkSWpqTxq3I8pv+fUEkQj/hOorMm9I3N/5XiJB+iXR\nq30mg+d2LiSS10nEifZD2eM8Bg87n5q9/1Cit/MG4orRS7LnZVWfB9Gzdkr2vkeJRPoq4orm+dDU\nCUTC98fs9RXZ553G4J6wtYmk97qsPvcSJ+QvLq03ida3yuohEoJbiW2cx9rOfsTtoZZnj0VEw0HZ\nzVQP0Z06jM+axMDbvz1J49hoVsZhRBL5KHHcnM7g2yrlJhKNIPdk6y8mEvBmhlN2lVU0vyjXBsRx\nsjQrfwmxn/di8O2lphPx71FRTn6MF/UAJ2VlryTmFL+LuPVc+dZfELeSO49IJlcSDUAfZnAy2+yY\nn0rz/dPsuBhKGa3KrarLs4nfjftoHDvFWF9MTNW4jUhe7ySO508wuMf77cT3diVxN4FeGrcMLN+e\nKx8hcQWN78mS7LOKV+xvFc+ORIPWX7P3P0T8PnyAuP2gJEmSJElrpA8RyfbOdVdEkiRJkqTx5mlE\nb3XRRsQIgrtxGqk0avyySZIkSWuOFwAXEhfVWwpsSUwp2Ia4LsMTtdVMkiRJkqRxagJxIb1lxHz6\nR4jrFpTnaktazcbSFRklKRk77rhj/1VXXVV3NSRJkobiKmCnuisx3nifbUmqwVVXXUV/fz/9/XEH\nlvzvNf0xc+bM2utg3MZt3MZt3MZt3MN7UH1rSrVhsi1JGjVLly6tuwq1MO60GHdajDstqcatzphs\nS1LNZs6cWXcVJEmS1GXl2wJIkkZHb29vLwBTp06ttSKj6ZnPfCaTJk2quxqjzrjTYtxpMe60pBr3\nrFmzAGbVXY/xxgukSVI9+rM5UJIkSWNaT08PmDsOm8PIJUmjpq+vr+4q1MK402LcaTHutKQatzpj\nsi1JkiRJUpc5FECS6uEwckmSNC44jLwz9mxLUs3yC6VJkiRpzWGyLUk1y67wmYRU57oZd1qMOy3G\nnZZU41ZnTLYlSZIkSeoyx91LUj2emrPd09OD87clSdJY5ZztztizLUmSJElSl5lsS5JGTapz3Yw7\nLcadFuNOS6pxqzMm25JUs5kzZ9ZdBUmSJHWZ4+4lqR7eZ1uSJI0LztnujD3bkiRJkiR1mcm2JGnU\npDrXzbjTYtxpMe60pBq3OmOyLUmSJElSlznuXpLq4ZxtSZI0LjhnuzP2bEtSzXp7e+uugiRJkrrM\nZFuSajZr1qy6qzBqUp3rZtxpMe60GHdaUo1bnTHZliRJkiSpyxx3L0n1eGrOdk9PD87fliRJY5Vz\ntjtjz7YkSZIkSV1msi1JGjWpznUz7rQYd1qMOy2pxq3OmGxLUs1mzpxZdxUkSZLUZY67l6R6eJ9t\nSZI0LjhnuzP2bEuSJEmS1GUm25KkUZPqXDfjTotxp8W405Jq3OqMybYkSZIkSV3muHtJqodztiVJ\n0rjgnO3O2LMtSTXr7e2tuwqSJEnqMpNtSarZrFmz6q7CqEl1rptxp8W402LcaUk1bnXGZFuSJEmS\npC5z3L0k1eOpOds9PT04f1uSJI1VztnujD3bkiRJkiR1mcm2JGnUpDrXzbjTYtxpMe60pBq3OmOy\nLUk1mzlzZt1VkCRJUpc57l6S6uF9tiVJ0rjgnO3O2LMtSZIkSVKXmWxLkkZNqnPdjDstxp0W405L\nqnGrMybbkiRJkiR1mePuJakeztmWJEnjgnO2O2PPtiTVrLe3t+4qSJIkqctMtiWpZrNmzaq7CqMm\n1bluxp0W406Lcacl1bjVGZNtSZIkSZK6zHH3klSPp+Zs9/T04PxtSZI0VjlnuzP2bEuSJEmS1GUm\n25KkUZPqXDfjTotxp8W405Jq3OqMybYk1WzmzJl1V0GSJEld5rh7SaqH99mWJEnjgnO2O2PPtiRJ\nkiRJXWayLUkaNanOdTPutBh3Wow7LanGrc6YbEuSJEmS1GWOu5ekejhnW5IkjQvO2e6MPduSVLPe\n3t66qyBJkqQuM9mWpJrNmjWr7iqMmlTnuhl3Wow7LcadllTjVmdMtiVJkiRJ6jLH3UtSPZ6as93T\n04PztyVJ0ljlnO3O2LMtSZIkSVKXmWxLkkZNqnPdjDstxp0W405LqnGrMybbklSzmTNn1l0FSZIk\ndZnj7iWpHt5nW5IkjQvO2e6MPduSJEmSJHWZybYkadSkOtfNuNNi3Gkx7rSkGrc6Y7ItSZIkSVKX\nOe5ekurhnG1JkjQuOGe7M/ZsS1LNent7666CJEmSusxkW5JqNmvWrLqrMGpSnetm3Gkx7rQYd1pS\njVudMdmWJEmSJKnLHHcvSfV4as52T08Pzt+WJEljlXO2O2PPtiRJkiRJXWayLUkaNanOdTPutBh3\nWow7LanGrc6YbEtSzWbOnFl3FSRJktRljruXpHr0n3TySQBsvvHmHH/U8TVXR5IkqZpztjuzTt0V\nkKRUbbP3NgAsu2RZzTWRJElStzmMXJI0alKd62bcaTHutBh3WlKNW50x2ZYkSZIkqcscdy9J9eif\ns3gOEMPIZ390ds3VkSRJquac7c7Ysy1JNbvsosvqroIkSZK6zGRbkmq26OJFdVdh1KQ6182402Lc\naTHutKQatzpjsi1JkiRJUpc57l6S6vHUnO0ZU2bQ399fc3UkSZKqOWe7M/ZsS5IkSZLUZSbbkqRR\nk+pcN+NOi3GnxbjTkmrc6ozJtiTVbPfX7153FSRJktRljruXpHp4n21JkjQuOGe7M/ZsS5IkSZLU\nZSbbkqRRk+pcN+NOi3GnxbjTkmrc6ozJtiRJkiRJXea4e0mqh3O2JUnSuOCc7c7Ysy1JNbvsosvq\nroIkSZK6zGRbkmq26OJFdVdh1KQ6182402LcaTHutKQatzpjsi1JkiRJUpc57l6S6vHUnO0ZU2bQ\n399fc3UkSZKqOWe7M+OxZ/vDwM01fv48YH6Nnz9UU4FVwIQ26y0FPrS6K7MG6wO+WnclJEmSJI0t\nQ0m2+4CvreZ6rA5TgQXAPcAK4HoiKdpmiO+fRCSrrygt788eY92vgS2A+7Pn04FHKtYbL/GMVfsB\nH6+7EtJ4kepcN+NOi3GnxbjTkmrcQzQBOA9YTnTovavN+h8A7gAeAr4DrDvEsiYRedojhccnCq/3\nAv8svPZw9p5RNx57todiBnAxkWi/HXgRcAQR7yeHWVZ5uMR4GT7xT+Duuisxhq3TpXIeBP7RpbJU\nv3WbLO/W8VJp99fvvjqLlyRJGg3fAFYCmwMHA98Cdmiy7v8DTgT2IjpDtwVmDbOsjYFnZI/iPVT7\ngXMKr21MJOyjrl2yPQ/YA3gf0XrwJNEqcCODhx5vl62zU/Z8Vfa+nxPJyFJiQxU9F/gh0ft6P9ET\n/cLSOh8F7iRaJb4LbNSmzs8jerC/DrwH+CVwC/Ab4Nis3hsSLRwHlN67D/A4sVP/li37fRbLpYX1\neoDjgduyep8BbFB4fT3gy1m9HwUuB15TeH1qVuZewO+I7fN74OUt4jqK6J3PvT4r48TCsh8Ac0uf\nMSH7+4ws7lXZ49OF920AzCFalW4lhuq30gtcAxxG7NflWflPA96flXEv8KXS+9YFTs5e/wdwBfCG\nwut5nd8I/IEYkXAZcZzsBVxNHAcXAJsW3tcDfCord2W23lsKr0/Kyn0nsR9XAMdk8bY6Bn4DnFp6\nfWNin+6XPe9j4MiPdjH+lsH7bBXwnOz504HHgN1obg/iuHmUOMZOI7Z9ro/4gfo80eB0F7EvWjUU\nTSe27V7An4l9eikDWwF7if1e9b7yOsM9NpYCM4nt8QjRyln8jTmDwdM31iK+2yc0iWktopX0b8Q+\nvwH4CAO3w7ys3BOJ7/MtxA9++Xj5d+K7dE4WwwpiO00vlHVoFls5YT8LOL9JHQHYY589Wr28Rpk6\ndWrdVaiFcafFuNNi3GlJNe4h2BB4G3FOvoIYZXs+cEiT9Q8DTifymweBz9A4rxpqWc1y2R6G3kE6\niTjvm06cB95H5F2vInKKBxh4rv9CIr98kDjP/mGrwtsl28cRieIZxJDkLbNKnE4kskWHA38E/lRY\nNgv4GbAj8G3ge8Ars9eeDiwkNuAewC7ECfbFNBLXfwM+S2zolwNLgA/SetjzO4gT+y82ef1hIgk6\nO6tzOYb5RI/wztmy/0fE/rbCeq8lWlb2Bg4E9ieS79wpWd3fQzQ+XAP8Iiun6PNEY8IriB17Vou4\nFgKTiSQQIjG9N/s3t0e2XtmviYRkRVaHLWgkkT3EEI6riG18clb/XVrUBeLA3Bd4E7Ft3kE0rOxE\nNAS8l9gm+xXecyax7d4FvJhoPJkPvKxUdi+RmL2aSKp/RIxIOCKL9yVEYpY7gWgg+Ej22nnAucRx\nV/QFohFme+CnROLU6hj4PpFwFb+sBxDb8efZ8/Iw/HYxLmTgPtuT+KLmy3YjRiVcQbXnAhcCVxLb\n+ojss75QWu9gotFgV6KR6QTiWG1lPeBjxI/NrsAzgf9q854qkxj+sQHx3b6WOA5nEt+P/bPXvk00\nwhS/Q/sQjRTfb1KPtYgE+h3E6JZPACcx+LdrT+K4eQPxnc73d/F4OR9YH1gMvJn4/n+FaKTaK1v/\nR9lnvrVQ9iZZnKc3qaMkSdKa4F+AJ4hO2dxVxPlwlR2y13NXE+d1mw6jrGVEJ8gZwLMKy/uJc9H7\niM6Ro4ZQ/52JRPqdxDneScQ53ouJvO612XqfJfK6ZxLn5SO+dtPCikK2IE7kX509Xxv4O9FbmFtF\nnIgWXUTjxPhwoqepaG0igXxH9vw3Tcr4G819k2iBaOeVRFIzMXu+KZFEvSl7PonqOdvziB1bTMC+\nndULoiXmMeDdhdfXIg6Wz2bPp2Zl71NYZ7ds2USau51GwvQrIlF/JCv/haX355+RXyBtOtVztpcy\nOMm/gYHzHsp6iW31jMKyHxM9qMXhtgtptAS9gBgZsVWprJ8RvbDFOhe3Sz6qYqfCspkM7GH9O4On\nByykcaxNysr4QGmddsfAs4h9uVfhPRczMAEtfj+GEuMbGbjPHiJa8vIyPwf8H83NJhqdig4jevTX\nz573EQ0sRf9HY9RDlenENtqusOygrNxcL0Pr2R7usQFxHP5vqey5xHGeu4aBowL+m0hwh+OLNL6r\nEN/nuxg4MmAS1cdLlXMYuF2/RjSG5I4mvrdVDZv9cxbP6Z+zeE7/SSef1J+KhQsX1l2FWhh3Wow7\nLcadllTjpv01nl5LdJwWHUl1RyBEblQc/fk04vxr6yGUtSGRo61FdET+mEiAc9sT+WoP0YF0O5FE\nV5mUfe6WhWXFfBTgJ0QnNERH2hwi0W6r03mIdxJDvg8nhrO+kUhUyknb5aXnv6WRyLwSeD6DE8AN\niDH7EL1R364oozzUvGiowwaupDHc9QtEYnEfA0+Um7mOgQfcHTQaHl5AHCzFZGcVsS3K8wyuLpUB\nccDc3uRzfwm8jugpfRXRy3o00RLzEuKgbfbeZvpL9SAr49lt3ncLA/fd3USS/kRpWV7OK4j9cl2p\nnPWAS0rLivXJ551fU1qW9/BvTHw5ysnlIhrHWm5x6Xm7Y+A+4ot7MDGceCLRINBLtaHEuCh7nu+z\nX2Wv5Y1KU4H/aVI+xI/Hb0vLfk0MXX4h0XpXtU/voLHNmnkM+GvpPesSLXcPtnlv0VCOjbtK9emn\n+veiOKJkLtGgdzLRiPQWBveOlx1F9KRvTfy2PI3Bc3b+TDS6lJWPl7WJnv8DiWNhPWL7FP8TmUtM\ngZhIfI8OJ36UV1VV7syZZ7LZxM148OYH+fK6X2annXZ6anhafgGWNe15bqzUZ7Se/+lPfxpT9XF/\nr97n7u+xUR/39+p9nhsr9XF/d/d5/vfSpUsZouXEeXnRJlR39lWtv0n27yNDKOsfxPkWxHnmscR5\n64bZa8Wpt5cTPdVvp/WQ77sKfz9a8TzvSPoo0YF6BdHB+x/EyNaOVfVsQyQyDxInsD8l5loW5WPf\niz5L4wT2W8Q85W0rHs/M1rm/SRmtbv11AoNbJ5o5hkYv4ZVEr2JuEs17tstzR3tpJIMvy973/NI6\nPyBaXQCmMvi2XM0+r2hGVt+9iQQBYud+nOjFLTZMlD9jOtUH+83E8N2iZvs818vgHs6vM7jl6oc0\neh4PJHp9JzN4f+f7qlxniC9GOVE5ihh6DfFFXEU0QhR9jsaxNonm27bVMQDRqvUgkVh9kMGjKorb\naigxQnzp8332oazsFURDzUpaz9f+KTEdoygf1ZAPranaf/Nofcu66Qw+PqYycH98msZxlzuS6jnb\nRc2OjR8Xnt/M4EaMI4gGj9wE4sfuNcQ0g2W0dmC2/jHEyIhtifntxd+PeQzeLpOoPl5OzOpzMPE9\n35boXS/HdgUxMuQlDB4tUJRkz7YkSRp/aN+znY/uLXaKfp+YFljlLAaed+9No/NxuGU9hzjnekaT\n108keqerTMreu1Zh2a3E9NziZ59U8d7XEOea2zYpe0ChzTxOdQ/4/xLzn48GphFj5ct2LT3fhUZL\nw5XEBryPSGCKj7wX7fomZbTa2T/J6vyxJq9vUvj7bOKCascS80SLrRKPZ/+uXVFGq8+/KXtv8fLC\naxNxlHs8h6uPOHE/mMYJfh8xzHnP7O9mHqc6ltHyR6LXd0sG7+/yMJHheJjoQSxfznl3Yv5vO62O\nAWgkYtOI7X52i7KGGmMfA/fZY8QIkU/Ser42xHdiFwaO3tid2L83tXgfjPwWb/fQuJBbbqeqFTuQ\nD/Mp2oWYPHKhAAAgAElEQVSB35n7ibn4RxDzrr/bpszdie36TeJaEn8jfnM63Q67ExfnO4sYOXAz\n0bBSLm8u0XhxBDGS4a+0cdlFl3VYJUmSpDHhH8R52meIa3PtTsybbnZtne8R50rbEyOkP0XjPLxd\nWTsT52BrEdM+v0rkRnkH0FuzMnuydY+jzcVqhyA/934HkTtA5Kz9NBnBCENLtpcSldwG2KzwQU8S\nCfYXiIsQXVrx3v2JIZzbET15exFX6YY4Yb2LCHwPoid4D+LCXXkrxleIIb7FMvILlzVzGzHX8lii\n12rPrO67EvMpi1dBfpDoXTuVGKJdTFbuJloq3kgkGMWhDK2Gqf+D6LU/GfhX4gD6FjGc+ptt6t7O\nEmKbvZuByfZUYt5AX4v3LiXm9L6e2I8btFh3OFfwaycv5wZin88jhr9vC0whLmy2f+U7h+5LWTnv\nJC6o8BniS1m+kniVVscARE/zT2lcpK88gqO4rYYaYx+xz55BYwhMH7FfL2fgcOuybxJDlL9JHFtv\nJr6DX6Mxv7rZ/hvpPl1I9C6fRPTCH8Hgq7mPxC5EI9l2RI/5IcB/ltaZS6NnuaqBr2gJ0Tv9xqzM\nTxG/MZ1uhyXE9+c1xBSXr1N9z8ZziHlCRxNXQ29r0cWLOqzS+FMefpgK406LcafFuNOSatxDdAyR\nY9xNnDMX76a0NZEM54nq/xIXZV5I5Ck3MfACyK3K2paY9vkwMaLyUQbeh/tAorPjYaJz5gs0T/ph\naB0x+TpTiKmOjxB57HG0uK3YUJLtU4les+uIRK948af8lj7Nxqn3EifjVxFDoKcTPdoQG2UPorfp\nx8TGm0cMIc8vcPajrIzZRFLyYmIYaDvfIi6y9WwiUfpLVsf1GHzV5jOIeZflk+IniI33XuICXD/L\nllcNoygvO5EYXnom0dv5EuKE/67Se8qGsqP7iP32y+z5MqKB4SYGz9culvcb4iJc5xAH7UdafEa7\noSJD2QZVy95DbJNTiP09n0iKlzapc7Nl5XK/SiTcpxBfuLcSc32vKb2nmWbHQO4HRHL3B+JYalWX\nocT46+w9vyq8t48YedDXop4Q+/hficT/j1mdz2bg0Jah7p+ydtv+L0QC+e/Ed3pvYjhPf2n9To6N\nfmLOS76dP0Mkx+eW3tdHDO3po/39EucQvyFnE6MFts4+o119abLsc1k5FxLfv0eovoPAcuI3bSXD\nv4CbJEnSePUA0cG0EdEhUZwjfQvR0XRbYdl/Eh0UmxCdOMVr6LQq64dEwr0R0Qk1ncZ1niCuw7RZ\n9nnbEx0kzSwlzsGLvdNbEbcfzh1CYwj7iUSDwTOIDuKWd5wZaU/Xq4lhks9n4IaDqPDbGXyyPNYc\nSCShWzLwystKh8dA/W4meufbNaZtQPzWHEs0HI1VFxL/qcxosU7/nMVxXbwZU2YQ06EkSZLGnp6e\nHujeyNdkdHo18nWJKwl/lkimy4n2eLABkVydRFxYzCQrPR4D40cPMVLleOJicmO1x3hT4nYV+zD4\n/vGSJElKyFCGkVc5iOhyn8DgK1mPFycSw2LvpXH/a6XFY2D82Ia45eC7iaH6T9Zbnab+SFzw4+OM\n/IKIa6RU57oZd1qMOy3GnZZU41ZnOu3Znpc9Wuk0kR8tvTS/X7LS0IvHwFhRvlVe2VLG/m8KVF8w\nra3dX1++mL4kSZLGO8fdS1I9npqzveySZcz+6OyaqyNJklTNOdudGQ89RZIkSZIkjSsm25KkUZPq\nXDfjTotxp8W405Jq3OqMybYkSZIkSV3muHtJqodztiVJ0rjgnO3O2LMtSTW77KLL6q6CJEmSusxk\nW5JqtujiRXVXYdSkOtfNuNNi3Gkx7rSkGrc6Y7ItSZIkSVKXOe5ekurx1JztGVNm0N/fX3N1JEmS\nqjlnuzP2bEuSJEmS1GUm25KkUZPqXDfjTotxp8W405Jq3OqMybYk1Wz31+9edxUkSZLUZY67l6R6\neJ9tSZI0LjhnuzP2bEuSJEmS1GUm25KkUZPqXDfjTotxp8W405Jq3OqMybYkSZIkSV3muHtJqodz\ntiVJ0rjgnO3O2LMtSTW77KLL6q6CJEmSusxkW5JqtujiRXVXYdSkOtfNuNNi3Gkx7rSkGrc6s07d\nFZCkVC27ZFndVZAkSdJq4rh7SapHf39/PxDzoPK/JUmSxhrnbHfGYeSSJEmSJHWZybYkadSkOtfN\nuNNi3Gkx7rSkGrc6Y7ItSTWbOXNm3VWQJElSlznuXpLq0e88bUmSNB44Z7sz9mxLkiRJktRlJtuS\npFGT6lw3406LcafFuNOSatzqjMm2JEmSJEld5rh7SaqHc7YlSdK44JztztizLUk16+3trbsKkiRJ\n6jKTbUmq2axZs+quwqhJda6bcafFuNNi3GlJNW51xmRbkiRJkqQuc9y9JNXjqTnbPT09OH9bkiSN\nVc7Z7sw6dVdAklL1iVM+Ufm3JEnScG2+8eYcf9TxdVdDBSbbklSTbfbepvLvNdmSxUuYPGVy3dUY\ndcadFuNOi3GnZSzHveySZXVXQSXO2Zakmk07clrdVZAkSVKXOe5ekurRP2fxnLrrIEmS1hDLLlnG\n7I/OXi1lO2e7M/ZsS5IkSZLUZSbbkqRRs2TxkrqrUAvjTotxp8W405Jq3OqMybYkSZIkSV3muHtJ\nqodztiVJUtc4Z3vssWdbkmo2f878uqsgSZKkLjPZlqSaLZi7oO4qjJpU57oZd1qMOy3GnZZU41Zn\nTLYlSZIkSeoyk21J0qiZPGVy3VWohXGnxbjTYtxpSTVudcZkW5IkSZKkLjPZliSNmlTnuhl3Wow7\nLcadllTjVmdMtiWpZtOOnFZ3FSRJktRl3itNkurhfbYlSVLXeJ/tsceebUmSJEmSusxkW5I0alKd\n62bcaTHutBh3WlKNW50x2ZYkSZIkqcscdy9J9XDOtiRJ6hrnbI899mxLUs3mz5lfdxUkSZLUZSbb\nGq++Diys8fP7gK/V+PlagyyYu6DuKoyaVOe6GXdajDstxp2WVOPuggnAecByYCnwrjbrfwq4FXiQ\nOOffofT6O4Hrs/JuBHbvYl27xmRbQ7UU+FDdlSjpH8I6bwMuBR4gvoxXA58Dnj3Ez5gKrCJ+IMqf\nPZTPlyRJklL3DWAlsDlwMPAtBifQubcARwGvJc7BLwe+X3h9H+CLwGHARtl6f1sttR4hk20N1VhM\nLNvNG5kN/Aj4A/BmYHvgeOD5wNFd/iytGdai+ndx3dGuyJpq8pTJdVehFsadFuNOi3GnJdW4R2hD\nogPsU8AK4NfA+cAhTdZ/MbCI6OxbBZzFwMR8Vva4Int+B3B7k7KmZ593GtH5diOwG/Ae4BbgLuDQ\nwvpvAq4FHgZuY4SdjSbba74+Bg93ngfML63zDeDzwD3EQfclGglmH7BNtmwV8CTwdOIgPKBU9j7A\n40TP8aRs/XcRX5hHieEe+5TeswPw86y8u4CzgecUXl8bOBW4P3v8Z7aslZ2BjwMfzh6/IYaiLCRa\n075SqN8rS+89MtsO2xG94mTPVwFnlOrVbJsBbAp8N6vzCuAiBv5QTAceAfYC/kz0vF+a1auVrYlh\nOA9nj58Czy283gtcQwyvuSlb5zzgWS3KnJTF97asnv8gfmheX1hnKoN7+fP3vaK0zhuJRo4VwGVZ\n/fYiRhY8AlxAbJ/cPOKY/CRwZ7bOGcD62euHAvcyOOk9i/ixbuaDwFXEtr0NmAtsUnh9evZZ/0rs\ng5VEo8xSYGZWhwdotKZ+EfhLFtfNwMnAeqVt0ex4WqdFPSVJktZU/wI8QSS6uauIpLrKJcCuxLn4\n04ge7Auz19YmzrU2B/5KnN9/jcY5Y5Wds8+bAJxDdMa9AngB8G5ieurTs3W/A/w7sHFWv0vLhQ2H\nyfaar2q4c9Wyg4kkeVfgWOAE4MDstf2JRGUWsAWwJZFsnA0cXirncCJpuqew7BTgy8CORCJ3PjAx\ne21LIhm7GngVsDcxHOR8Gonrh4D3Egf+LsSX7KCKGMrxLKf5vOqHiITq/5rE8D0iUc0bE3YgYj8+\ne95D620GkUC+ihgKszOxzX7BwB+D9YCPEUnfrsAzgf9qEddaxLZ5NpHYvo7Ylj8rrTcJeAfwVuAN\nwMuJnv52ZhP76mXA74EfEq2Rw9ULvB94NZFU/4hIpI/I6v0SIpkt2hN4KZGUH5DV++TstR8Rsb+1\nsP4mwH7A6S3q8SSxz3YgjpmdGXxMrJ/V7chsvWXZ8g8C1xE/6Cdly5YTLaEvAo4hGjQ+kb22lNbH\n0xMt6pmMVOe6GXdajDstxp2WVOMeoY2Izp+iR4BnNFn/CqLDaglx/nwAcV4G0SH3tGzZ7sBOxHnu\nJ1t8/s1Zef3EOeVE4DPAP4nc5HHghdm6jxNJ9sZEvvDHIcTXlMl2mnoYPCz6WiJBuhH4MdEDvHf2\n2gNE0vIIcHf2gOglfAONxHlTIhn6TqnsbwI/AW4gEp9baQzjPhr4E9ELvYToXTyMSIryHsITiKSr\nWMadbWLcjkiWn2yz3lyi5z3vndyeSBC/Q/RSPpAtz+N+pPDeVttsO2BfooFgURbXIcQX9+BCGesA\n7wMWE73RpxLJaDN7EwnpQUTP8ZXZ368gktRiudOzz/0t8O1C3Vo5jRhlcBORYE4gGkmG61PEkJ1r\niMaDXYkfyd9ndf5uRX2eIBLZ64ik9URgBrAB0eN8FgMT2YOIH8Gft6jHV4iRGbcQjTonAv9WWmdt\norHkcmJfLs+W9xH742/E9oCY7395Vt6FwBcYeIGPVsdTU9OOnNbqZUmSpPFsOXEOXLQJA8+ri44l\nzhOfR5xTfYboYV6fGCkL0XlyF3Afcf76phaff1fh7/z995SWbZT9fUBW1lLiXHCXFuW25bBGQbTy\nXF1adgcxPKOVK4lk6jAi6TiIOOAvLK13eemzfkckIRAJ9R4M/rL1E0M7/kr0KFeVsVWLulU1KFS5\ngBhC/zZiWMnhWdnXtXlfu222PZGsF+v9MLG9ti8se4yIsVjGukQP94MVn7s9MSfllsKym7NlO9AY\n6rKMgdt0KPsTBsZ0R/bvUN7Xqpy8ceaa0rJyuVcTrZe53xLb4gVEo8FcooFhIhHv4UTSvqpFPfYi\nGnJeRPyor020hm5Bo8HmCaLBp6ifaAApezvR+PMC4kd5bQY2Wg7reDpz5plsNnEzAC4+62K2mrzV\nU3PB8pZzn68Zz/NlY6U+Pnd/+9z97fM1c3/39fUBMHXq1BE9z/9eunQpXXADkXe+kMZQ8h2J87sq\nbyTOo/J52N8lRl7uQJwL3taNSjWxmBg5uTYxSvNHxBTOjnjRpzXfJcSJ/vsLy84iWpf2zZ4vJBKh\n4wrrzCPm+Obr3Ey0IJ1WKv8Yoqd5MpF8X0hjGMckoldwL6JlKPd9orfy7dn6jxLzqsvuJg70B5qU\n8TxiGHWVLxNDlicQQ0RaOYUYfvJG4O9Z/fOhyVOJBHYzYu51rtk2m0AMG38LcC7RGlfsXV9EJOAf\nIXqev8bAITTNPi93HLGtyl/6W4n5xN8getsPIHrAc1WfVTSJ2FdTiB+x3CpiP51LNIr0EUPY78te\n3w5YUnhfVf3fTmMYeO4o4LM0rgo/j7hw3Z6FdfLGlpfR+DG+ghhGfz6RnE9mYGNF0TbE/Oo5xA/2\nfUTjzjlZvLfQfLtUHe+7AL8itu8viMaQtxK938XYWh1PRf1zFs9pUnVJkqThWXbJMmZ/dCizBoev\np6cHRpY7nkN0ZryXGJG5gBj5eH3Fup8nrjB+AHHNnoOJkbLPJTqvZhHX23kz0WlyAXH+WZ6iCHGu\nd0RWHkTCfwMDz91uJaaC/p4YAbmAGD15BHEe9/xhR5txGPma7x4aw7xzOzL8q4s/TvVFyc4mkt5j\niQTjzIp1di383UMMEc+/WFcS83dvIZK94mM5caDf0aSMVjGcTcw1PrbJ68WLZJ1OJO3vI3orf1h4\n7fHs33YXZCu7nvh+7VZYtjERa7te83blTiQSydy22bKRlDsU+XCb4vG0UxfLfymNi1NAJLeP0xjC\nDdG7PZ348VtE80QbogHgacAHiN7lGxl4Ibnheg2RPM8mjtubqL6YXavjKXl5C3xqjDstxp0W405L\nqnF3wTFEZ9vdwA+Ijpc8H9iaGJH5vOz554jOnKuJTrfjicQ7n/f9WSIxvoE4/72S5tcmanb9qmbe\nTXS6PERMBz24xbptmWyv+S4lWn72JXoBTyMO5GLL1FCGXC8lejYnEr2WuQeJ+cqnAr9kYGKUO4r4\ngkwmepy3Iu6tB9ETuwnw30QCvS1xBew5NOZOfAX4aKmMLdrU9wqih/FLwH8QidI2RM/r92lc6Azi\ni7ooW//HNObsQgzH7gemEb2w+cXCmm2zfNlfid7XOcTFG15K/LA8RDQEdOoi4ofnLKKXdkr295VE\nb/vqdCPR8tdL9Gi/gdYXoxiudYirf+9A4/6J36YxtwaiVXQLYq5/y3nQNFotP0C0SL6Lgft9uJYQ\nyfpBxHF6NHGBtKrPbXY8SZIkpegB4qLLGxGdFcXOiFuIUYb58PAVRA/4FkSeMIW4nk/uCaJTY1Pi\nYssn0OggK/sukcPkbmRwJ9pWxJ2L/knkTROyz311trxjJttrvjMKj0VEsnceA1t0hnLF8k8TB+JN\nDLzIQP4Z69I8+fkYcXGsPxEJ2v405mDcQSTCq4ihuX8mLr+/kpjPDJEsn0n0GP42W3ZWk88qf+47\niaEqPycuaPY1okWtfMXvZjH8nRiSMpuY45tfyXoo2+w9RNJ/AdGzuj4xtPix0nvK2o06eCvRy7yQ\naEy5nZhb0qweQy233ev/JLbntsTtE2YS86GH0lrYblv1E4011xJxnQtcTDSyFC0nEtiVxND0Vq4h\nkusPZuUeTgzBH07rZtECovHmy0T8exPfi6r3t/tOJKs45y0lxp0W406Lcacl1bjVGedsqxsOJJLX\nLYkkKDeJ6nnAY9GJNG7ppNE3j4HXCGjlQqIFdMbqrNAIDeV4emrO9vw589l3xlBClyRJqjbG52wn\nyZ5tjcQGRC/nScRw35WtVx+TNiTupXccMVxdY9emxIXn9mHs7quOjqcFcxestgqNNanOdTPutBh3\nWow7LanGrc6YbGskTiSu9nwvcaGCKsO9ENto+wYx33kRMb9a9Wg29L3oj8D3iKHrq/ticJ3yeJIk\nSRLgUABJqstTw8hnTJmBtwGTJEkj4TDysceebUmSJEmSusxkW5I0alKd62bcaTHutBh3WlKNW50x\n2Zakmk07clrdVZAkSVKXOe5ekurR7zxtSZLULc7ZHnvs2ZYkSZIkqctMtiVJoybVuW7GnRbjTotx\npyXVuNUZk21JkiRJkrrMcfeSVA/nbEuSpK5xzvbYY8+2JNVs/pz5dVdBkiRJXWayLUk1WzB3Qd1V\nGDWpznUz7rQYd1qMOy2pxq3OmGxLkiRJktRlJtuSpFEzecrkuqtQC+NOi3GnxbjTkmrc6ozJtiRJ\nkiRJXWayLUkaNanOdTPutBh3Wow7LanGrc6YbEtSzaYdOa3uKkiSJKnLvFeaJNXD+2xLkqSu8T7b\nY48925IkSZIkdZnJtiRp1KQ6182402LcaTHutKQatzpjsi1JkiRJUpc57l6S6tF/0skn1V0HSZK0\nhth84805/qjjV0vZztnujBtMkurR39/fD0Bvby+9vb311kaSJKkJk+3OOIxckmo2a9asuqswavr6\n+uquQi2MOy3GnRbjTkuqcaszJtuSJEmSJHWZQwEkqR5PDSPv6ekh/1uSJGmscRh5Z+zZliRJkiSp\ny0y2JUmjJtW5bsadFuNOi3GnJdW41RmTbUmq2cyZM+uugiRJkrrMcfeSVI9+52lLkqTxwDnbnbFn\nW5IkSZKkLjPZliSNmlTnuhl3Wow7LcadllTjVmdMtiVJkiRJ6jLH3UtSPZyzLUmSxgXnbHdmnbor\nIEmp+sQpnwDgsosuY4999qi5Nqqy+cabc/xRx9ddDUmSNA6ZbEtSTbbZexsAFp24iEO+eEjNtRkd\nSxYvYfKUyXVXY8iWXbKsK+X09fUxderUrpQ1nhh3Wow7LcYtteecbUmSJEmSusxx95JUj/45i+cA\nMGPKDPK/NbYsu2QZsz86u+5qSJJUK+dsd8aebUmSJEmSusxkW5I0apYsXlJ3FWqR6n1ZjTstxp0W\n45baM9mWpJpNO3Ja3VWQJElSlznuXpLq0e887bHPOduSJDlnu1P2bEuSJEmS1GUm25KkUeOc7bQY\nd1qMOy3GLbVnsi1JkiRJUpc57l6S6uGc7XHAOduSJDlnu1P2bEtSzebPmV93FSRJktRlJtuSVLMF\ncxfUXYVR45zttBh3Wow7LcYttWeyLUmSJElSlznuXpLq8dSc7RlTZuD87bHJOduSJDlnu1P2bEuS\nJEmS1GUm25KkUeOc7bQYd1qMOy3GLbVnsi1JNZt25LS6qyBJkqQuc9y9JNXD+2yPA87ZliTJOdud\nsmdbkiRJkqQuM9mWJI0a52ynxbjTYtxpMW6pPZNtac3xdWBhjZ/fB3ytxs9fHVYBb2vxXJIkSe1N\nAM4DlgNLgXe1WPclwP8C9xDnXmXLgUcKjyeAr3axrl1jsi11binwoborUdI/hHXeBlwKPED8WF0N\nfA549hA/Yyrxwzeh4rOH8vnj2RbAgiGu28ea1/gwYpOnTK67CrWYOnVq3VWohXGnxbjTYtwapm8A\nK4HNgYOBbwE7NFn3ceCHwBFNXt8IeEb22AJ4FPhRNyvbLSbbUufGYmLZ7sIVs4kfoz8Abwa2B44H\nng8c3eXPWhPdTfwH0FXz58zvdpGSJEljxYZEZ8+ngBXAr4HzgUOarH8DcCZw3RDKfjtwF7CoyevT\ns887jehouhHYDXgPcEv23kML678JuBZ4GLiNEXasmWwrRX0M7nGcB8wvrfMN4PPEEJa7gC/RSDD7\ngG2yZauAJ4GnE1/MA0pl70MkaM8GJmXrv4v4UXgUuD5bp2gH4OdZeXcBZwPPKby+NnAqcH/2+M9s\nWSs7Ax8HPpw9fgPcSgw9Pxj4SqF+ryy998hsO2xH9IpDY2jPGaV6NdtmAJsC383qvAK4iIGtmtOJ\n4UB7AX8met4vzerVytbE0KSHs8dPgecWXu8FrgHeCdyUrXMe8Kw25ZaVh5F/mhjhsBK4g4gN4nja\nA3hf9p5VWR0rLZg71M7y8c8522kx7rQYd1qMW8PwL8RQ7xsLy64CXtyFsg8DvtdmnZ2zz5sAnEN0\nPL0CeAHwbmIq5tOzdb8D/DuwcVa/S8uFDYfJtlJUNdy5atnBRJK8K3AscAJwYPba/kRr1yxi+MqW\nRPJ4NnB4qZzDiUT+nsKyU4AvAzsSCef5wMTstS2By4jh3a8C9iaGy5xPI3H9EPBe4sdgFyLJPagi\nhnI8y2k+tPkhInH8vyYxfI9IVPPGhB2I2I/PnvfQeptBJKGvAt5C/PCtAH4BrF9YZz3gY0TivSvw\nTOC/WsS1FrFtnk0McX8dsS1/VlpvEvAO4K3AG4CXEz39nTqA2A9HAy8EpgG/y147DricaIjYInvc\nNoLPkiRJGq82Ijo6ih4hhoGPxDZE58Z326x3c7ZOP5FoTwQ+A/yTOA9/nDiXI/v7xUSy/RDwx5FU\n0GRbCj0MHhZ9LdEjeiPwY6IHeO/stQeI3uxHiKHFd2fL5xKJXJ44b0okd98plf1N4CfEMJnjiR7m\nfBj30cCfiF7oJUQP72FEcpr3OJ8AnFwq4842MW5HJMtPtllvLtHzvl72fHvg1VkMq4jYoRH3I4X3\nttpm2wH7Eg0Ei7K4DiF+zA4ulLEO0SO8mOiNPpVIopvZG3gp0djwB+DK7O9XED3kxXKnZ5/7W+Db\nhbp1YhuiN/siIpG+ktivEP+hPE40JuTbqeoCH8lxznZajDstxp0W49YwLCfO94o2YeA5ZCcOAX4F\nLGuz3l2Fvx/N/r2ntGyj7O8DiKHkS4mRrLuMpILrjOTN0hqsn+hZLrqDuKhDK1cSCeJhwBeIpO8+\n4MLSepeXPut3RFILkVDvweAfoH5iuMtfiZ7SqjK2alG3qgaFKhcQQ+jfRgy1OTwru928mXbbbHsi\n4SzW+2Fie21fWPYYEWOxjHWJHu4HKz53e+B2Yt5N7uZs2Q40hv8sY+A2Hcr+bOVHRA/2zcQVM39B\nbLshz+k+c+aZbDZxMwAuPutitpq81VPJaD7c2uf1Pl8/G3SRDxvMT7J87nOf+9znPl+Tn+d/L126\nlC64gcg7X0hjKPmORAfISBxKTF/spsXAfsSo0fcT53tNpwK2k+IFjqRLiMTx/YVlZxEtbvtmzxcS\nSeBxhXXmEXN883VuJoZkn1Yq/xiip3kykXxfCHwye20S8Deix7Wv8J7vAxsQF3m4kGhh+3BF3e8m\nvvwPNCnjecQw6ipfJq7qOIEYNtPKKcQw6zcCf8/qf3r22lQigd2MmHuda7bNJhDDxt8CnEv0mBd7\n1xcRCfhHiJ7nrzFwWFGzz8sdR2yr8g/hrcAXiYaDXqKl8qWF16s+q2wVsU/ObfJ8PaJ3/PVZ+Q8T\nowBWUL09ivrnLJ4DwIwpM8j/XtMtWbxkXPVuL7tkGbM/OpLZBqGvr++pE5mUGHdajDstxp2Wnp4e\nGFnueA7RMfNeYvThAmK64PVN1l8f2JZIyDfIlj1WeH03Yurjc4B/tPjc6cT572uz5y8kkv+1Cuvc\nSkx7/D3wb1ndHsre90niQsIdWav9KtIa5x4aw7xzOzL8q4s/TvVFyc4mkt5jiYT1zIp1di383UMM\nEc9/bK4k7i94C5GYFx/LiS//HU3KaBXD2cTVII9t8vomhb9PJ5L29xHDan5YeC3vuW13Qbay64nf\nnN0KyzYmYh3K1SZblTuRGNad2zZbNpJyh+Ix4H+ADxJz0V9MI77HGeLooWlHTlstlZMkSRojjiGS\n5ruBHwBH0Tj33ZoYffi87PkkouPiz8S57aMMTsoPJS6I2yrRhubXamrm3USH2kPE1MeDW6zblsPI\nlYBtXCgAACAASURBVKJLiV7efYmWrRnEl/vmwjpDGXK9lBjufRaRWN2bLX+QmK98KvBLYp502VHZ\nZ/+Z+PHZirjfIERP7JHAfxPzsu8lksd3EBfkWk5cOfzjpTK2IHqhm7mC6LH+UhbvucRc4+cTLXd/\nJS4WQVbuomz9c7LPzC0jfqSmES1/K4gfumbbLF/2V+JCZnOIH6+HiAuUPUQ0BHTqImL4+lnEiIIe\nosf6SqJ3eXWZTjQ4XEFsnwOJ4yAfAr+UaADZhtg+99Hkx33fGftWLV4jjade7W5KsRcEjDs1xp0W\n49YwPUBcYLjKLQwcabiU9p3CRw3xc7/LwAuo3cjgDqPiNMx/HWK5Q2LPtlJ0RuGxiEj2zmNgIjSU\nK5Z/mvhy3sTACy/kn7Eugy+MlvsY0Rv6J+KCavsTc4wheq1fQwxZ/gWRTH+duL1UPnzmP4ge89OJ\ni31BJJvtfIy4/dUriFuLXUskpncz+IrfzWL4OzCTSJTvpHF186Fss/cQyekFxDzw9Ymh6o+V3lPW\nbtTBW4kRCwuJxpTbifk2zeox1HJbeYBopLiMGC6+PzHPPb9Ix6lE8n0dcXy0mk8vSZKkNYxztqXV\n40Aied2SSJJzk4jh4FOIK2ePZScSyfGL6q7IGqo/lXnaRc7ZTotxp8W402LcaenCnO0kOYxc6q4N\niAT7JOLWUitbrz4mbUg0ChwHfK7eqkiSJEnjk60TUnf1Eon2r4ihzctLr08ihp2/irHbsz2PGGp+\nPnG/be8PvXok2bM93nSrZ1uSpPHMnu3OOGdb6q5eYp7z3gxOtCEu+LA2YzfRhrjw1/rEUHgT7VEw\nf878uqsgSZKkLjPZlqSaLZi7oO4qjJoli5fUXYVa9PX11V2FWhh3Wow7LcYttWeyLUmSJElSlznu\nXpLq8dSc7RlTZuD87bHJOduSJDlnu1P2bEuSJEmS1GUm25KkUeOc7bQYd1qMOy3GLbVnsi1JNZt2\n5LS6qyBJkqQuc9y9JNXD+2yPA87ZliTJOdudsmdbkiRJkqQuM9mWJI0a52ynxbjTYtxpMW6pPZNt\nSZIkSZK6zHH3klQP52yPA87ZliTJOdudsmdbkmo2f878uqsgSZKkLjPZlqSaLZi7oO4qjBrnbKfF\nuNNi3Gkxbqk9k21JkiRJkrrMcfeSVI+n5mzPmDID52+PTc7ZliTJOdudsmdbkiRJkqQuM9mWJI0a\n52ynxbjTYtxpMW6pPZNtSarZtCOn1V0FSZIkdZnj7iWpHv0nnXxS3XVQG5tvvDnHH3V83dWQJKlW\nztnujBtMkurR39/fX3cdJEmS2jLZ7ozDyCVJoybVuW7GnRbjTotxpyXVuNUZk21JkiRJkrrMoQCS\nVA+HkUuSpHHBYeSdsWdbkmrW29tbdxUkSZLUZSbbklSzWbNm1V2FUZPqXDfjTotxp8W405Jq3OqM\nybYkSZIkSV3muHtJqsdTc7Z7enpw/rYkSRqrnLPdGXu2JUmSJEnqMpNtSdKoSXWum3GnxbjTYtxp\nSTVudcZkW5JqNnPmzLqrIEmSpC5z3L0k1cP7bEuSpHHBOdudsWdbkiRJkqQuW6fuCkhSqj5xyifq\nrsKoW3bTMrZ5wTZ1V2PUGXdajDstxp2WZnFvvvHmHH/U8TXUSGOZybYk1WSbvdM7SVm5yUq2mWLc\nqTDutBh3Wox7oGWXLKuhNhrrHHcvSfXon7N4Tt11kCRJXbDskmXM/ujsuqux2jhnuzPO2Zakms2f\nM7/uKkiSJKnLTLYlqWYL5i6ouwqjZsniJXVXoRbGnRbjTotxpyXVuNUZk21JkiRJkrrMZFuSNGom\nT5lcdxVqYdxpMe60GHdaUo1bnTHZliRJkiSpy0y2JUmjJtW5bsadFuNOi3GnJdW41RmTbUmq2bQj\np9VdBUmSJHWZ90qTpHp4n21JktYQ3mdbVezZliRJkiSpy0y2JUmjJtW5bsadFuNOi3GnJdW41RmT\nbUmSJEmSusxx95JUD+dsS5K0hnDOtqrYsy1JNZs/Z37dVZAkSVKXmWxLUs0WzF1QdxVGTapz3Yw7\nLcadFuNOS6pxqzMm25IkSZIkdZnJtiRp1EyeMrnuKtTCuNNi3Gkx7rSkGrc6Y7ItSZIkSVKXmWxL\nkkZNqnPdjDstxp0W405LqnGrMybbGms2Ae4Eth3m+44Dzut+dZpaCnxoFD9Pa7BpR06ruwqSJEnq\nMpNtjTUfAS4G/lZa/jbgUuABYDlwNfA54NnZ698GdgNeVXrfqsLjYeD3wP7DqM904JGK5f3ZQxqx\nfWfsW3cVRk2qc92MOy3GnRbjTkuqcXfBBKJjbDnRafWuNut/CrgVeBBYCOxQeO1YYDGwEjiz2xXt\nJpNtjSXrAkcy+EszG/gR8AfgzcD2wPHA84Gjs3VWAj8G3ldR7nuBLYhE/KpsvVd3ue5aM6zTZPnT\nRrUWkiRJa5ZvEOfrmwMHA99iYAJd9BbgKOC1RJJ+OfD9wut/Bz4LnLG6KtstJtsaS14PbED0YOd2\nBj4OfDh7/IZo5VpIfFG/Ulj3fOAAYO1SuQ8CdwNLgBnEF31f4gv8OPCc0vqziaR8T+JLvCGN3vFP\nF9bbAJgDPJTV6cOlcrYmWvAezh4/BZ5beL0XuAZ4J3BTts55wLNo7aVE7/8K4D6icWLjwuvzgPlE\ng8RtwP1ZHBu0KHNqFt9ewO+AfxCjAF5eWGc6g3v58/dNKK3zRuAvWTnnZ/U7ELiB2B/zgPUK5fQR\nP7pfyep7P3AK0JO9/mliW5X9moHHQNkXs3qsAG4GTi59bm9W7nRiHzxKY38fA5xLtMDOJn4vv0OM\nuliRxfKRQh33oPXxJNKd62bcaTHutBh3WlKNe4Q2JEapfoo4h/o1cX54SJP1XwwsInrAVwFnMTAx\nPy97/31D+Ozp2eedRoyQvZEYDfse4BbgLuDQwvpvAq4lzstvY4TTRk22NZbsQfReF4dnH0wkO19r\n8p6HCn8vJr7M5aHkRU9mj/WAXxEJVvELtlb2/HQisT+B+FHYInucmq3XA3yASKJeTiRxpwC7FMo5\nnxjmPhV4HTAR+FmpPpOAdwBvBd6QlTW7Rf03hP/f3v3HyVXWhx7/bBOQSEgUY6SUXwoYgYoUA0UE\nuxq9SBtEpbZeEEU0DVqFXkGgomaDIN7Scqml2hh6+SU/XqJVb7b+aED2agWlQX4pGIEmIRQsWCGA\nwOXX3j++Z7JnZ2d2zk5mztmZ5/N+vfaVOWfOzDzfndmT832e5/sM3yVOAAcQU+IPZmLP3qHESWkR\nkeS+g0i+W/kscCqwP3ECu7zAY+q9APgYMT1oEbCQSFqPIU60byd6LD9U97hjsn8PIjpF/oz4/UMk\nua9i/Hu7AHgd8V418zhxMn0VkTy/Gzij7piXZ/uPAl5DdMYALAOGgd8lemN/izjpvit7vjOAT2TP\nD/B9Jv88SZIkpeiVwLNEoltzK5FUN3ItcY23JzG78H3AtxscN9BgXyMHZq+3PXAlMWN2f2B34D3A\nBcALs2P/kbgGnZO173v1TzYVzaZMSlXYk+hhqt93D5Egt/IwMaq6J/Cj3P7aH+ILiERyO2JkGCIJ\n+gBwbrZ9GJEgfxl4hkhqR4mR8XrfBb6Q3b6AWKRtUfbai4gR6FfkYjqaOMm8ibE/3JmMHzH+EmPJ\nWyNHEyeDY4lRY4gTwnXZa9Vq3TcR029GiRH9q7M2fW6S54bocfy/2e0ziV7FHYH7WzwubyYxnf+u\nbPsKomNiPjFiDdERsQg4P/e4+xnrEPgFcWL+GPC/iOlC3wGOJ0bcyW6vofGId81Zudv3AucQPZT5\nGQpbE7/Ph+oeexUTOzGW1T3fa4lOhdpxk32eRLq1bsadFuNOi3GnJdW4t9Bs4po67zHimryRG4FL\niGvY54hrrkUNjiu6ftK67PkgEu0ziOvcZ4DVxMzEPYg1oZ4mkuzbievpmwu+RkMm25pOtiOmcuQN\nULzXCuIPeW7dvsuIacuziCnMJxOJMsClxEjyQUSSfDwxNeXhFq8zSvxB5t3P2IJte2Xb+c6Dddm+\nvRlLtjcwfmr2A0RS2sxeRM/cb3L7biCm2OzNWLJ9B+NPQA9QrE49H9MD2b/zmVqy/f8YS7QhOip+\nyViiXduXnw40yvgOErLtzxAn6MeBlcSJ8i+I3tFjgeUt2vLH2fG7Z88zg4kzeu5jYqINkcjXO4FY\nA2AX4vO0FTHFqWZKn6eLll3EvB3nsfamtez3B/ux84KdN/8nXpum5rbbbrvttttuT//tbdgGgJGR\nEQAGBwd7ert2e/369XTA44wveYS4Xm+0CDHEAmiLgJ2Ia8hjiWvnfYiSv5qiOUI+v6g9/qG6fbOz\n20cBnyQGqG4DTmfiNWphU0lipG67mpjCm6/fOJ8YKdye6H1qZRMxqlobSXye+IP9DpGI/6rBY76S\nPe50YgR1MWMj38cRU9jre97WZfvPy+27jugFOzH7OYVIyvI2En+8f0/UCx9FjIDXNHu9mvOIadlv\nyO3bmpjqfiTwz0THwkuIuvSaRq+VN0icxOYxlhTvRiTvC4np/e8lRvJn5x73FqLjova4Ru0/hXhP\nXp7b9zniJFqbFn4d0THxvtwxi4jexjnESXom0TlxKvFeXkVM7W92oj6IKBUYIt7/R4jf0V8zlnAP\n0fj38jyRqP9Tbt+fEr/bk4kSg0eJz9Y76mKb7POUN7pizQoAli5cSu12v1u7Zm2SowLGnRbjTotx\np6VZ3Buu3cDZp05WCdjbBgYGoP3ccVviOnEfxqaSX0ZcF3+iwfHDxPVlvoz0YeLa8Ce5fZ8hEvLJ\nZoUeR+QSh2bbexAzKPODLxuJ67zrc/tmAB8lZlnWX88XZs22ppO7mfhhvoL4A/1Ik8fkR7FfTCR5\nd9Ud80siaWyUaEOMmP4JUSf8AOMTo6eZuOBaEXcS0693ze17Rbbvjjaer+YOIjHMJ7wHE3/Ld+b2\ndeNryR4iprDnE+n9OvTcA0wceT+ISFYfz7afJZLd44mT6tdonmgDvD57/NnATUQ5wm5b0MZDiMXj\nvgDcQnym9mDi73qyz5MkSVJqfkMMYJxJXEseQgwKXdbk+NuIa6n5xDXuscSgSy1RnwFsk+2bQZSK\ntnO9Xm8rYg2hucT09ccoVsralMm2ppMfEAuE5XvNbiQWHjsX+BsigdqVGIm9jPGLfh1IjPDeNMXX\nXU0sBvZpIpnLW0/8Mb+ZGL2dbEXv/JT31cSJ4nKirndhdvsmYhS3XZcTMV5KLNz1BmJF9K8x/rvJ\nuzFrpbZK+TlEknkUsehYp+xIzGRYQIwqn0LUa+ddSLz3i4kFLCazllj9/Wiio+NDxEJo7VpLLKbx\nVmJdgE8xfoZBzWSfp+SlOAoCxp0a406Lcacl1bg74MPEdfSDxAzUExgbKNqFSGx3yrbPIq67biNG\ntE8irjtrdd+1Vc1PIxY4e5KJC+DWjDJxYGSyQan3EDNYNxHrIh0zybEtmWxrOrmGmEZevwDC6USS\ntD8xTfpnxLSSB4F/yB33NuCrxAjoVF1M9GbVf8f39dlrXJm93scneY76P+YjidHg64gp2vcTK3E3\nOz6/v5kniUW35hAdEd8gvs7g+BbP2+y1Wr1uft+viRPOW4iT3weJmpZWJ7Ai7RklTrwziLqYLxGJ\ndX2yvY5YwG0DYwu5NTNMdNKcT9S5LyIS4PrXLToLYAUxRfwK4ne/C9EB1MjFNP48SZIkpehhovRu\nNjHT8KrcffcSMyfvy7afIK4zdyBGmRcC/5I7fojIY/M/ZzZ53UsYPzhyNxNHwXcmrvmfAQ4nylfn\nErMur2cLWLOt6eYsov51qr1Is4hR6COIRGiqvkiMfh7WxmO15fL17q3cQcxqOKerLdoyRT5P1mwn\nxLjTYtxpMe60WLOtqXA1ck035xLTRvJfY1XEEmKEd6qJ9lxiVexjie9PVjWKrDr/UmJ6+S7EKPN0\n1NbnafGSxV1rkCRJkqph74RSN0KsiH0h4+u/Va4iI9vPE9PyP0bUrk9HIxT/PI2mMpotSVK/c2Rb\njTiyrdQNVt0AAfDGAsf0whoTg1U3QJIkSdNDL1y8SpL6xNo1a6tuQiWMOy3GnRbjTkuqcas9JtuS\nJEmSJHWY8+4lqRrWbEuS1Ces2VYjjmxLUsVWrVhVdRMkSZLUYSbbklSx4ZXDVTehNKnWuhl3Wow7\nLcadllTjVntMtiVJkiRJ6jCTbUlSaRYsXFB1Eyph3Gkx7rQYd1pSjVvtMdmWJEmSJKnDTLYlSaVJ\ntdbNuNNi3Gkx7rSkGrfaY7ItSRVbvGRx1U2QJElSh/ldaZJUDb9nW5KkPuH3bKsRR7YlSZIkSeow\nk21JUmlSrXUz7rQYd1qMOy2pxq32mGxLkiRJktRhzruXpGpYsy1JUp+wZluNOLItSRVbtWJV1U2Q\nJElSh5lsS1LFhlcOV92E0qRa62bcaTHutBh3WlKNW+0x2ZYkSZIkqcNMtiVJpVmwcEHVTaiEcafF\nuNNi3GlJNW61Z2bVDZCkVG24dkPD25IkqbfMnzO/6iZoGnJFOUmqxujo6CgQK3zWbve7kZERBgcH\nq25G6Yw7LcadFuNOS6pxuxp5e5xGLkkVW7ZsWdVNkCRJUofZOyFJ1RhNZTRbkiT1Nke22+PItiRJ\nkiRJHWayLUkqzcjISNVNqIRxp8W402LcaUk1brXHZFuSJEmSpA5z3r0kVcOabUmS1BOs2W6PI9uS\nVLGhoaGqmyBJkqQOM9mWpIotX7686iaUJtVaN+NOi3GnxbjTkmrcao/JtiRJkiRJHea8e0mqxuaa\n7YGBAazfliRJ05U12+1xZFuSJEmSpA4z2ZYklSbVWjfjTotxp8W405Jq3GrPzKobIEmpOuOvzgDg\nkDcfsvl2v9twzwZW37i66maUzrjD/DnzOemEkypskSRJ5XHevSRVY3TFmhVVt0Eq1YZrN3D2qWdX\n3QxJ0hRZs90ep5FLkiRJktRhJtuSpNKsXbO26iZUwrjTkmpNp3Gnxbil1ky2JUmSJEnqMOfdS1I1\nrNlWcqzZlqTeZM12exzZlqSKrVqxquomSJIkqcNMtiWpYsMrh6tuQmlSreE17rSkWtNp3Gkxbqk1\nk21JkiRJkjrMefeSVI3NNdtLFy7F+m2lwJptSepN1my3x5FtSZIkSZI6zGRbklSaVGt4jTstqdZ0\nGndajFtqzWRbkiq2eMniqpsgSZKkDnPevSRVw+/ZVnKs2Zak3mTNdnsc2ZYkSZIkqcNMtiVJpUm1\nhte405JqTadxp8W4pdZMtiVJkiRJ6jDn3UtSNazZVnKs2Zak3mTNdnsc2Zakiq1asarqJkiSJKnD\nTLYlqWLDK4erbkJpUq3hNe60pFrTadxpMW6pNZNtSZIkSZI6zGRbmv7mAr8EXjHFx50IfL3zzWlq\nPXByia/XbbsBzwP7N9lWGxYsXFB1Eyph3GkZHBysugmVMO60GLemaHviuvRx4prxv09y7LuBnwOb\ngF8B/wTsmLv/I8Aa4Cngoi60tWNMtqXp7+PANcC/1+1/J/A94GHixHUbcBbw0uz+LwEHAwfUPe75\n3M+jwL8B75hCe44DHmuwfzT76Vf3AjsAtxY8fj391fkgSZLUrr8nkuP5wDHAF4G9mxz7Q+ANxIDT\nrsATwHm5+/8D+Azwv7vV2E4x2Zamt62BJUzstTsb+ArwE+CPgL2Ak4CXAx/KjnkKuBr48wbP+0Ei\ncTyASB6vBn6/w23vN88DDwLPFTy+nzse2pZqDa9xpyXVmk7jTotxawq2JQaJPkUkzj8Evgkc2+T4\njcQ1F8QK6M8BD+Tu/3r2+P8q8NrHZa93HjFAdTcxGPV+YiDlP4H35o7/Q+BnxIDUfWzhwInJtjS9\nvRmYRYxg1xwI/CVwSvZzPXFSuo7oKfzb3LHfBI4CZtQ97yPESWwtsJRIzI8ADgWeBl5Wd/zZRFL+\nB0Qv4raMjY5/OnfcLGAFMe1nY9a+vF2IE+Sj2c/XgN/J3T8E3E5MH7onO+brwEuY3KuJ0f8niBPv\nRcCc3P0XA6uIDon7gF9nccxq8bx5uzF+GvlWwOeJ3tWniBP2Odl9I0RP7LnZYyZN0BcvWTyFZkiS\nJPWUVwLPEoluza3APpM85hDievVR4vrxtAbHFP0qsgOz19seuJIYsNof2B14D3AB8MLs2H8E/oy4\njtyH8dfgU2ayLU1vbyBGr/OjpMcQ08b/rsljNuVuryES4/qp5HnPZT8vAH5AJLn5Hr7fyrYvJBL7\nvyCS2h2yn7/OjhsA/gdxMvs94H8CfwUclHuebxLT3AeBNxL1N9+oa89uwLuAI4H/lj3XZF/Muy3w\nXeJkfAAxJf5gJk4tOpSYrrQI+NPsuJMmed5WTgTenj3XHtm/P8/ueweR1C8nfke/PdkTHbH0iC1o\nRm9JtYbXuNOSak2ncafFuDUFs4nrtLzHgO0mecy/Ai8CdgKeIQYw6hWdRbgOuCQ7/ivE9eeZ2fOu\nJgaa9siOfZpIsucQ19Q3F3yNhky2peltT2LEtH7fPRSbzvwwcTLbs25/rSfwBcSUnu2IkWGIpPr9\nuWMPIxLkLxMnpUeJk9WD2c8TuWO/C3yBqC+/gOjBXJTdt4gYgT6a6EC4Kbu9P/Cm3HPMJKb8/BT4\nEVF7vojmjiZ6I48lpv18n+iRfCfjF5XbBJxAjOavJqbOT/a8rewC/IL4z+A+4AbiRA7xe3+O+N3X\nfk+SJEkpepzxMw4h6rEbrQFU737iWvW9De4rOrL9n7nbT2b/PlS3b3Z2+yhiKvl6YqbiQWyBmVvy\nYEldtx3jTxAQJ5aiJxeI5Hhu3b7LiKnVs4gpOicTiTLApcRI8kFEsns8MZX74RavM0os0pZ3P2ML\ntu2Vbec7D9Zl+/ZmbJrOBsaffB8gFtNoZi9iNP03uX03ENO392ZsYbk7GN8D+gBbVqd+MZG0/wL4\nF+BbwLeZQq32RcsuYt6O8wCYNXsWOy/YefNIYK3Wtd+2a/umS3vK2r7m8muSeH9bvd/bsA0wVvNY\nGyHqt+3zzz+f/fbbb9q0p6zt2r7p0h7f7+5u1/ZNl/b4fnd2u3Z7/fr1dMAviLxzD8amkr+GGFgp\nYivGD+7UdGN9nDXEzMUZwEeJkfBd2n2yqVywSyrf1UQ9cH4BifOBDxB1J88UeI5NxCJpX862nye+\nMuE7RCL+qwaP+Ur2uNOJmuTFjI18H0dMYa+f+rMu259fLfI6ogb7xOznFCaesDYCnyNWqRwiehRf\nnbu/2evVnAcsJKbc12xNnJSPBP6ZSIxfQtSl1zR6rbzdiER9ITESX78N0Qt6GDFC/i4i6X8LcfJv\n9PvIG12xZkWTu/rX2jVrk5xabNxhw7UbOPvUyapC+sPIyMjmC9eUGHdajDstAwMDsGW545XE9dEH\niVmNw8DrgDsbHHs0Udq4kVgD51LiGuvE7P4ZRAK+jFj7ZwlRE95o1udxxHXzodn2HkTyn5/hvZEo\nB/w34E+ytm3KHvdJYgHitjiNXJre7mZicnoFUaf8kSaPyY9iv5hIUu+qO+aXROLYKNEGWEmcbJYS\nI8DX5O57mokLrhVxJ1Ejs2tu3yuyfXe08Xw1dxAJ8+zcvoOJ81v+BN6N3s/HiUXePkysCv8mYrEN\naP/31NdSTDjBuFOT4oU4GHdqjFtT9GFiRuWDxADQCYxdp+1CzGrcKdvem1gn6HFiKvcNwKm556qt\nan4ascDZk8AZTV630VfTTnZN+B5iwGQTUZZ4zKRRtWCyLU1vPyAWCMv3JN5ILDx2LvA3wOuJBHaQ\nmB6eX/TrQOJkdNMUX3c1sar3p4lR4bz1wDbESunzmHxF7/yU99XENPPLgdcSI8SXZ227borty7uc\niPFS4HeJEe4VRBKc/27yTs/k+RixavpeRC/pMcSJ+b7s/vVZW3Ykfk9NrVqxqsNNkyRJmlYeJhaQ\nnU3MFrwqd9+9xOBQ7Rrqk8DO2bEvJ2ZaPpU7fojIY/M/ZzZ53UsYP/vxbiYOhuxMJPfPAIcTs0fn\nEuWG1xeKrgmTbWl6u4Y4udQv5HU6kejtT0yT/hkxZflB4B9yx70N+CoxtWaqLiam6NR/x/f12Wtc\nmb3exyd5jvrexCOJBSmuI2q07yfqYpodn9/fzJPEVO45REfEN4jvUzy+xfM2e63JXje//SgR+4+J\nDoN9iRN07T+DTxMn73uYWHc/zvDK4RbN6B+pfu+ycaclX/OYEuNOi3FLrblAmjS9PU2sxv1+xk/l\nhkiivzrJY2cBf8z4OmUo3sn229lr1q+GDjEV6MN1+xrVs7yxbnsj0avZzPLsJ+9iJo6u1/spMdLe\nzPsb7Gv0WnnrGd/zWb99YfbTzI+B/Sa5X5IkSX3MBdKk6W8u8XVVBzN+WnQrJxJTy9/ZxuvtTUz7\nfhexwrY6b/MCaUsXLiXFxdKUnlQWSJOkftOBBdKS5Mi2NP1tAnZo43Gfz36m6pvAAcSorYm2JEmS\n1AZrtiXVGyRWOz+pxXHSlKVaw2vcaUm1ptO402LcUmsm25JUscVLFlfdBEmSJHWY8+4lqRqj1mkr\nNdZsS1Jvsma7PY5sS5IkSZLUYSbbkqTSpFrDa9xpSbWm07jTYtxSaybbkiRJkiR1mPPuJaka1mwr\nOdZsS1Jvsma7PY5sS1LFVq1YVXUTJEmS1GEm25JUseGVw1U3oTSp1vAad1pSrek07rQYt9SaybYk\nSZIkSR3mvHtJqsbmmu2lC5di/bZSYM22JPUma7bb48i2JEmSJEkdZrItSSpNqjW8xp2WVGs6jTst\nxi21ZrItSRVbvGRx1U2QJElShznvXpKq4fdsKznWbEtSb7Jmuz2ObEuSJEmS1GEm25Kk0qRaw2vc\naUm1ptO402LcUmsm25IkSZIkdZjz7iWpGtZsKznWbEtSb7Jmuz2ObEtSxVatWFV1EyRJktRhM6tu\ngCSlasO1GwAYXjnMvnvsW3FryrHhng3suvuuVTejdMYd5s+ZX2FryjMyMsLg4GDVzSidcafFj+0c\ndgAAB3pJREFUuKXWTLYlqSK16bSfPe2zyUytTfUixbglSUqP8+4lqRqjo6OjQNRB1W5LkiRNN9Zs\nt8eabUmSJEmSOsxkW5JUmlS/n9S402LcaTHutKQat9pjsi1JFVu2bFnVTZAkSVKHOe9ekqoxap22\nJEnqBdZst8eRbUmSJEmSOsxkW5JUmlRr3Yw7LcadFuNOS6pxqz0m25IkSZIkdZjz7iWpGtZsS5Kk\nnmDNdnsc2Zakig0NDVXdBEmSJHWYybYkVWz58uVVN6E0qda6GXdajDstxp2WVONWe0y2JUmSJEnq\nMOfdS1I1NtdsDwwMYP22JEmarqzZbo8j25IkSZIkdZjJtiSpNKnWuhl3Wow7LcadllTjVntMtiWp\nYsuWLau6CZIkSeow591LUjX8nm1JktQTrNlujyPbkiRJkiR1mMm2JKk0qda6GXdajDstxp2WVONW\ne0y2JUmSJEnqMOfdS1I1rNmWJEk9wZrt9jiyLUkVGxoaqroJkiRJ6jCTbUmq2PLly6tuQmlSrXUz\n7rQYd1qMOy2pxq32mGxLkiRJktRhzruXpGpsrtkeGBjA+m1JkjRdWbPdHke2JUmSJEnqMJNtSVJp\nUq11M+60GHdajDstqcat9phsS1LFli1bVnUTJEmS1GHOu5ekavg925IkqSdYs90eR7YlSZIkSeow\nk21JUmlSrXUz7rQYd1qMOy2pxq32mGxLkiRJktRhzruXpGpYsy1JknqCNdvtcWRbkio2NDRUdRMk\nSZLUYSbbklSx5cuXV92E0qRa62bcaTHutBh3WlKNW+0x2ZYkleaWW26pugmVMO60GHdajDstqcat\n9phsS5JK88gjj1TdhEoYd1qMOy3GnZZU41Z7TLYlSZIkSeowk21JUmnWr19fdRMqYdxpMe60GHda\nUo1b7XH5dkmqxi3Aa6puhCRJUgG3AvtV3QhJkiRJkiRJkiRJkiRJkiRJkiRJkiRJkiRJnfFW4OfA\nXcBpTY75fHb/rcDvldSubmsV96uAG4CngJNLbFe3tYr7GOJ9vg34IbBveU3rqlZxH0nEfTNwE/Cm\n8prWVUX+vgEOAJ4F3llGo0rQKu5BYBPxft8MfLK0lnVXkfd7kIj5p8BIKa3qvlZxn8LYe3078Vl/\nUWmt655Wcc8DvkMsePpT4LjSWtZdreJ+MfB14pz+Y2Cf8pomSdKYGcDdwG7AVsR/yHvVHfOHwLey\n278P/KisxnVRkbhfCiwEzqJ/ku0icb8OmJvdfivpvN/b5m6/Oju+1xWJu3bc94Bh4KiyGtdFReIe\nBP5Pqa3qviJxvwj4GbBTtj2vrMZ1UdHPec1i4JruN6vrisQ9BJyT3Z4H/Bcws5zmdU2RuM8FPpXd\nXkB/vN9d4/dsS1L3HEj8p7UeeAa4ihjhy3sbcEl2+8fExdrLSmpftxSJ+yFgTXZ/vygS9w3EiB/E\n+70Tva9I3L/J3Z4N/KqUlnVXkbgBPgp8lfjM94Oicffb18sWifto4GvAfdl2Sp/zmqOBK7vfrK4r\nEvcDwJzs9hwi2X62pPZ1S5G49wKuy26vJRLzl5bTvN5jsi1J3fM7wMbc9n3ZvlbH9HoCViTufjTV\nuD/A2KyGXlY07rcDdwLfBk4soV3dVvTv+0jgi9n2aAnt6rYicY8CBxPTTL8F7F1O07qqSNx7AtsT\nicga4NhymtZVUzmvvRA4jOhw6HVF4l5JTKG+n/isn1RO07qqSNy3MlYScyCwK71/3dI1vT7VQZKm\ns6IX1vUjQL1+Qd7r7W/XVOJ+I3A88PoutaVMReP+RvZzKHAZMf2wlxWJ+3zg9OzYAfpjtLdI3D8B\ndgaeAA4n3vdXdrNRJSgS91bA/sAiIvG8gSgVuauL7eq2qZzXjgD+FXikS20pU5G4P0FMsx4EdgdW\nA68BHutes7quSNyfA/6WsRr9m4HnutmoXmayLUnd8x/EBWfNzoxNL2x2zE7Zvl5WJO5+VDTufYkR\nkbcCD5fQrm6b6vv9A+L64yXEtMteVSTu1xLTMCFqOg8npmb2cj1zkbjzyca3gS8QI76/7m7TuqpI\n3BuJqeNPZj/fJ5KvXk62p/L3/W76Ywo5FIv7YODs7PY9wDqiE3FN11vXPUX/vo/Pba8D/r3L7ZIk\naYKZxH/AuwFb03qBtIPojwWzisRdM0T/LJBWJO5diHq4g0ptWXcViXt3xkZ198+O73VT+ZwDXER/\nrEZeJO6XMfZ+H0jUf/a6InG/ilgsagYxsn07vT+FvujnfC7ReTartJZ1V5G4zwOWZbdfRiSl25fU\nvm4pEvfc7D6AJcDFJbVNkqQJDicWELkb+Mts39Lsp+aC7P5biUSkH7SKewdiFGgTMbp7L7FwVq9r\nFfeFxAVp7Wtybiy7gV3SKu5Tia/GuZkY2T6g7AZ2SZG/75p+Sbahddx/TrzftwDX0z+dS0Xe71OI\nFclvpz/WJoBicb8PuKLkdnVbq7jnAauI/7tvJxaH6wet4n5ddv/PicUf59Y/gSRJkiRJkiRJkiRJ\nkiRJkiRJkiRJkiRJkiRJkiRJkiRJkiRJkiRJkiRJkiRJkqRq/X+gw5RTr4p5twAAAABJRU5ErkJg\ngg==\n", + "text": [ + "
\n", + "
\n", + "As we can see from the results above, we are already able to make our Python code run almost as twice as fast if we compile it via Cython (Python on list: 332 \u00b5s, untyped Cython on list: 183 \u00b5s). \n", + "However, although it is more \"work\" to adjust the Python code, the \"typed Cython with memoryview on numpy array\" is significantly as expected." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 1, + "metadata": {}, + "source": [ + "How to use Cython without the IPython magic" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#sections)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "IPython's notebook is really great for explanatory analysis and documentation, but what if we want to compile our Python code via Cython without letting IPython's magic doing all the work? \n", + "These are the steps you would need." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### 1. Creating a .pyx file containing the the desired code or function." + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%%file cython_bubblesort_nomagic.pyx\n", + "\n", + "cimport numpy as np\n", + "cimport cython\n", + "@cython.boundscheck(False) \n", + "@cython.wraparound(False)\n", + "cpdef cython_bubblesort_nomagic(np.ndarray[long, ndim=1] inp_ary):\n", + " \"\"\" The Cython implementation of Bubblesort with NumPy memoryview.\"\"\"\n", + " cdef unsigned long length, i, swapped, ele, temp\n", + " cdef long[:] np_ary = inp_ary\n", + " length = np_ary.shape[0]\n", + " swapped = 1\n", + " for i in xrange(0, length):\n", + " if swapped: \n", + " swapped = 0\n", + " for ele in xrange(0, length-i-1):\n", + " if np_ary[ele] > np_ary[ele + 1]:\n", + " temp = np_ary[ele + 1]\n", + " np_ary[ele + 1] = np_ary[ele]\n", + " np_ary[ele] = temp\n", + " swapped = 1\n", + " return inp_ary" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "Writing cython_bubblesort_nomagic.pyx\n" + ] + } + ], + "prompt_number": 16 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### 2. Creating a simple setup file" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "%%file setup.py\n", + "\n", + "import numpy as np\n", + "from distutils.core import setup\n", + "from distutils.extension import Extension\n", + "from Cython.Distutils import build_ext\n", + "\n", + "setup(\n", + " cmdclass = {'build_ext': build_ext},\n", + " ext_modules = [\n", + " Extension(\"cython_bubblesort_nomagic\",\n", + " [\"cython_bubblesort_nomagic.pyx\"],\n", + " include_dirs=[np.get_include()])\n", + " ]\n", + ")" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "Writing setup.py\n" + ] + } + ], + "prompt_number": 17 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
\n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "####3. Building and Compiling" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "!python3 setup.py build_ext --inplace" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "running build_ext\r\n" + ] + }, + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "cythoning cython_bubblesort_nomagic.pyx to cython_bubblesort_nomagic.c\r\n" + ] + }, + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "building 'cython_bubblesort_nomagic' extension\r\n", + "/usr/bin/clang -fno-strict-aliasing -Werror=declaration-after-statement -DNDEBUG -g -fwrapv -O3 -Wall -Wstrict-prototypes -I/Users/sebastian/miniconda3/envs/py34/include -arch x86_64 -I/Users/sebastian/miniconda3/envs/py34/lib/python3.4/site-packages/numpy/core/include -I/Users/sebastian/miniconda3/envs/py34/include/python3.4m -c cython_bubblesort_nomagic.c -o build/temp.macosx-10.5-x86_64-3.4/cython_bubblesort_nomagic.o\r\n" + ] + }, + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "In file included from cython_bubblesort_nomagic.c:352:\r\n", + "In file included from /Users/sebastian/miniconda3/envs/py34/lib/python3.4/site-packages/numpy/core/include/numpy/arrayobject.h:4:\r\n", + "In file included from /Users/sebastian/miniconda3/envs/py34/lib/python3.4/site-packages/numpy/core/include/numpy/ndarrayobject.h:17:\r\n", + "In file included from /Users/sebastian/miniconda3/envs/py34/lib/python3.4/site-packages/numpy/core/include/numpy/ndarraytypes.h:1761:\r\n", + "\u001b[1m/Users/sebastian/miniconda3/envs/py34/lib/python3.4/site-packages/numpy/core/include/numpy/npy_1_7_deprecated_api.h:15:2: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1m\r\n", + " \"Using deprecated NumPy API, disable it by \" \"#defining\r\n", + " NPY_NO_DEPRECATED_API NPY_1_7_API_VERSION\" [-W#warnings]\u001b[0m\r\n", + "#warning \"Using deprecated NumPy API, disable it by \" \\\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m" + ] + }, + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "In file included from cython_bubblesort_nomagic.c:352:\r\n", + "In file included from /Users/sebastian/miniconda3/envs/py34/lib/python3.4/site-packages/numpy/core/include/numpy/arrayobject.h:4:\r\n", + "In file included from /Users/sebastian/miniconda3/envs/py34/lib/python3.4/site-packages/numpy/core/include/numpy/ndarrayobject.h:26:\r\n", + "\u001b[1m/Users/sebastian/miniconda3/envs/py34/lib/python3.4/site-packages/numpy/core/include/numpy/__multiarray_api.h:1629:1: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1m\r\n", + " unused function '_import_array' [-Wunused-function]\u001b[0m\r\n", + "_import_array(void)\r\n", + "\u001b[0;1;32m^\r\n", + "\u001b[0mIn file included from cython_bubblesort_nomagic.c:353:\r\n", + "In file included from /Users/sebastian/miniconda3/envs/py34/lib/python3.4/site-packages/numpy/core/include/numpy/ufuncobject.h:327:\r\n", + "\u001b[1m/Users/sebastian/miniconda3/envs/py34/lib/python3.4/site-packages/numpy/core/include/numpy/__ufunc_api.h:241:1: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1m\r\n", + " unused function '_import_umath' [-Wunused-function]\u001b[0m\r\n", + "_import_umath(void)\r\n", + "\u001b[0;1;32m^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:18981:32: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function\r\n", + " '__Pyx_PyUnicode_FromString' [-Wunused-function]\u001b[0m\r\n", + "static CYTHON_INLINE PyObject* __Pyx_PyUnicode_FromString(const char* c_str) {\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:19132:33: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function\r\n", + " '__Pyx_PyInt_FromSize_t' [-Wunused-function]\u001b[0m\r\n", + "static CYTHON_INLINE PyObject * __Pyx_PyInt_FromSize_t(size_t ival) {\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:16538:1: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function\r\n", + " '__pyx_add_acquisition_count_locked' [-Wunused-function]\u001b[0m\r\n", + "__pyx_add_acquisition_count_locked(__pyx_atomic_int *acquisition_count,\r\n", + "\u001b[0;1;32m^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:16548:1: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function\r\n", + " '__pyx_sub_acquisition_count_locked' [-Wunused-function]\u001b[0m\r\n", + "__pyx_sub_acquisition_count_locked(__pyx_atomic_int *acquisition_count,\r\n", + "\u001b[0;1;32m^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:17017:26: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function\r\n", + " '__Pyx_PyBytes_Equals' [-Wunused-function]\u001b[0m\r\n", + "static CYTHON_INLINE int __Pyx_PyBytes_Equals(PyObject* s1, PyObject* s2...\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:17298:32: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function\r\n", + " '__Pyx_GetItemInt_List_Fast' [-Wunused-function]\u001b[0m\r\n", + "static CYTHON_INLINE PyObject *__Pyx_GetItemInt_List_Fast(PyObject *o, P...\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:17312:32: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function\r\n", + " '__Pyx_GetItemInt_Tuple_Fast' [-Wunused-function]\u001b[0m\r\n", + "static CYTHON_INLINE PyObject *__Pyx_GetItemInt_Tuple_Fast(PyObject *o, ...\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:17491:38: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function\r\n", + " '__Pyx_PyInt_From_unsigned_long' [-Wunused-function]\u001b[0m\r\n", + " static CYTHON_INLINE PyObject* __Pyx_PyInt_From_unsigned_long(unsi...\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:17538:36: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1mfunction\r\n", + " '__Pyx_PyInt_As_unsigned_long' is not needed and will not be emitted\r\n", + " [-Wunneeded-internal-declaration]\u001b[0m\r\n", + "static CYTHON_INLINE unsigned long __Pyx_PyInt_As_unsigned_long(PyObject *x) {\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:17644:48: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function\r\n", + " '__pyx_t_float_complex_from_parts' [-Wunused-function]\u001b[0m\r\n", + " static CYTHON_INLINE __pyx_t_float_complex __pyx_t_float_complex_fro...\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:17654:30: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function '__Pyx_c_eqf'\r\n", + " [-Wunused-function]\u001b[0m\r\n", + " static CYTHON_INLINE int __Pyx_c_eqf(__pyx_t_float_complex a, __pyx_...\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:17657:48: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function '__Pyx_c_sumf'\r\n", + " [-Wunused-function]\u001b[0m\r\n", + " static CYTHON_INLINE __pyx_t_float_complex __Pyx_c_sumf(__pyx_t_floa...\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:17663:48: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function '__Pyx_c_difff'\r\n", + " [-Wunused-function]\u001b[0m\r\n", + " static CYTHON_INLINE __pyx_t_float_complex __Pyx_c_difff(__pyx_t_flo...\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:17675:48: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function '__Pyx_c_quotf'\r\n", + " [-Wunused-function]\u001b[0m\r\n", + " static CYTHON_INLINE __pyx_t_float_complex __Pyx_c_quotf(__pyx_t_flo...\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:17682:48: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function '__Pyx_c_negf'\r\n", + " [-Wunused-function]\u001b[0m\r\n", + " static CYTHON_INLINE __pyx_t_float_complex __Pyx_c_negf(__pyx_t_floa...\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:17688:30: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function\r\n", + " '__Pyx_c_is_zerof' [-Wunused-function]\u001b[0m\r\n", + " static CYTHON_INLINE int __Pyx_c_is_zerof(__pyx_t_float_complex a) {\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:17691:48: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function '__Pyx_c_conjf'\r\n", + " [-Wunused-function]\u001b[0m\r\n", + " static CYTHON_INLINE __pyx_t_float_complex __Pyx_c_conjf(__pyx_t_flo...\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:17705:52: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function '__Pyx_c_powf'\r\n", + " [-Wunused-function]\u001b[0m\r\n", + " static CYTHON_INLINE __pyx_t_float_complex __Pyx_c_powf(__pyx_t_...\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:17764:49: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function\r\n", + " '__pyx_t_double_complex_from_parts' [-Wunused-function]\u001b[0m\r\n", + " static CYTHON_INLINE __pyx_t_double_complex __pyx_t_double_complex_f...\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:17774:30: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function '__Pyx_c_eq'\r\n", + " [-Wunused-function]\u001b[0m\r\n", + " static CYTHON_INLINE int __Pyx_c_eq(__pyx_t_double_complex a, __pyx_...\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:17777:49: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function '__Pyx_c_sum'\r\n", + " [-Wunused-function]\u001b[0m\r\n", + " static CYTHON_INLINE __pyx_t_double_complex __Pyx_c_sum(__pyx_t_doub...\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:17783:49: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function '__Pyx_c_diff'\r\n", + " [-Wunused-function]\u001b[0m\r\n", + " static CYTHON_INLINE __pyx_t_double_complex __Pyx_c_diff(__pyx_t_dou...\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:17795:49: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function '__Pyx_c_quot'\r\n", + " [-Wunused-function]\u001b[0m\r\n", + " static CYTHON_INLINE __pyx_t_double_complex __Pyx_c_quot(__pyx_t_dou...\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:17802:49: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function '__Pyx_c_neg'\r\n", + " [-Wunused-function]\u001b[0m\r\n", + " static CYTHON_INLINE __pyx_t_double_complex __Pyx_c_neg(__pyx_t_doub...\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:17808:30: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function '__Pyx_c_is_zero'\r\n", + " [-Wunused-function]\u001b[0m\r\n", + " static CYTHON_INLINE int __Pyx_c_is_zero(__pyx_t_double_complex a) {\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:17811:49: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function '__Pyx_c_conj'\r\n", + " [-Wunused-function]\u001b[0m\r\n", + " static CYTHON_INLINE __pyx_t_double_complex __Pyx_c_conj(__pyx_t_dou...\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:17825:53: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function '__Pyx_c_pow'\r\n", + " [-Wunused-function]\u001b[0m\r\n", + " static CYTHON_INLINE __pyx_t_double_complex __Pyx_c_pow(__pyx_t_...\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:18247:27: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1mfunction '__Pyx_PyInt_As_char' is\r\n", + " not needed and will not be emitted [-Wunneeded-internal-declaration]\u001b[0m\r\n", + "static CYTHON_INLINE char __Pyx_PyInt_As_char(PyObject *x) {\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:18347:27: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1mfunction '__Pyx_PyInt_As_long' is\r\n", + " not needed and will not be emitted [-Wunneeded-internal-declaration]\u001b[0m\r\n", + "static CYTHON_INLINE long __Pyx_PyInt_As_long(PyObject *x) {\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:2955:32: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function\r\n", + " '__pyx_f_5numpy_PyArray_MultiIterNew1' [-Wunused-function]\u001b[0m\r\n", + "static CYTHON_INLINE PyObject *__pyx_f_5numpy_PyArray_MultiIterNew1(PyOb...\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:3005:32: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function\r\n", + " '__pyx_f_5numpy_PyArray_MultiIterNew2' [-Wunused-function]\u001b[0m\r\n", + "static CYTHON_INLINE PyObject *__pyx_f_5numpy_PyArray_MultiIterNew2(PyOb...\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:3055:32: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function\r\n", + " '__pyx_f_5numpy_PyArray_MultiIterNew3' [-Wunused-function]\u001b[0m\r\n", + "static CYTHON_INLINE PyObject *__pyx_f_5numpy_PyArray_MultiIterNew3(PyOb...\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:3105:32: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function\r\n", + " '__pyx_f_5numpy_PyArray_MultiIterNew4' [-Wunused-function]\u001b[0m\r\n", + "static CYTHON_INLINE PyObject *__pyx_f_5numpy_PyArray_MultiIterNew4(PyOb...\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:3155:32: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function\r\n", + " '__pyx_f_5numpy_PyArray_MultiIterNew5' [-Wunused-function]\u001b[0m\r\n", + "static CYTHON_INLINE PyObject *__pyx_f_5numpy_PyArray_MultiIterNew5(PyOb...\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:3909:27: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function\r\n", + " '__pyx_f_5numpy_set_array_base' [-Wunused-function]\u001b[0m\r\n", + "static CYTHON_INLINE void __pyx_f_5numpy_set_array_base(PyArrayObject *_...\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m\u001b[1mcython_bubblesort_nomagic.c:3997:32: \u001b[0m\u001b[0;1;35mwarning: \u001b[0m\u001b[1munused function\r\n", + " '__pyx_f_5numpy_get_array_base' [-Wunused-function]\u001b[0m\r\n", + "static CYTHON_INLINE PyObject *__pyx_f_5numpy_get_array_base(PyArrayObje...\r\n", + "\u001b[0;1;32m ^\r\n", + "\u001b[0m" + ] + }, + { + "output_type": "stream", + "stream": "stdout", + "text": [ + "39 warnings generated.\r\n", + "/usr/bin/clang -bundle -undefined dynamic_lookup -L/Users/sebastian/miniconda3/envs/py34/lib -arch x86_64 build/temp.macosx-10.5-x86_64-3.4/cython_bubblesort_nomagic.o -L/Users/sebastian/miniconda3/envs/py34/lib -o /Users/sebastian/github/python_reference/tutorials/cython_bubblesort_nomagic.so\r\n" + ] + } + ], + "prompt_number": 18 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### 4. Importing and running the code" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "import cython_bubblesort_nomagic\n", + "\n", + "cython_bubblesort_nomagic.cython_bubblesort_nomagic(np.array([4,6,2,1,6]))" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "pyout", + "prompt_number": 19, + "text": [ + "array([1, 2, 4, 6, 6])" + ] + } + ], + "prompt_number": 19 + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "heading", + "level": 2, + "metadata": {}, + "source": [ + "Speed comparison" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[back to top](#Sections)]" + ] + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "from cython_bubblesort_nomagic import cython_bubblesort_nomagic\n", + "\n", + "list_a = [random.randint(0,100000) for num in range(100000)]\n", + "\n", + "ary_a = np.asarray(list_a)\n", + "ary_b = np.asarray(list_a)\n", + "\n", + "times = []\n", + "\n", + "times.append(min(timeit.Timer('cython_bubblesort_typed(ary_a)', \n", + " 'from __main__ import cython_bubblesort_typed, ary_a').repeat(repeat=3, number=1000)))\n", + "\n", + "times.append(min(timeit.Timer('cython_bubblesort_nomagic(ary_b)', \n", + " 'from __main__ import cython_bubblesort_nomagic, ary_b').repeat(repeat=3, number=1000)))" + ], + "language": "python", + "metadata": {}, + "outputs": [], + "prompt_number": 20 + }, + { + "cell_type": "code", + "collapsed": false, + "input": [ + "from matplotlib import pyplot as plt\n", + "import numpy as np\n", + "\n", + "bar_labels = ('Cython with IPython magic', \n", + " 'Cython without IPython magic')\n", + "\n", + "fig = plt.figure(figsize=(10,8))\n", + "\n", + "# plot bars\n", + "y_pos = np.arange(len(times))\n", + "plt.yticks(y_pos, bar_labels, fontsize=14)\n", + "bars = plt.barh(y_pos, times,\n", + " align='center', alpha=0.4, color='g')\n", + "\n", + "# annotation and labels\n", + "\n", + "for b,d in zip(bars, times):\n", + " plt.text(max(times)+0.02, b.get_y() + b.get_height()/2.5, \n", + " '{:.2} ms'.format(d),\n", + " ha='center', va='bottom', fontsize=12)\n", + "\n", + "t = plt.title('Bubblesort on 100,000 random integers', fontsize=18)\n", + "plt.ylim([-1,len(times)+0.5])\n", + "plt.vlines(min(times), -1, len(times)+0.5, linestyles='dashed')\n", + "plt.grid()\n", + "\n", + "plt.show()" + ], + "language": "python", + "metadata": {}, + "outputs": [ + { + "metadata": {}, + "output_type": "display_data", + "png": "iVBORw0KGgoAAAANSUhEUgAAAx0AAAHtCAYAAAB8lBOjAAAABHNCSVQICAgIfAhkiAAAAAlwSFlz\nAAALEgAACxIB0t1+/AAAIABJREFUeJzt3XucXHV5+PHPlHCTbKKRW7GRYJGbaERStZqmK4WqFRGo\nYBEhEcQLYhVRqDd2twUs4uWnojWJXIVSa0UgaaTltoRYkVIQNWoQJAFERUUSQiBgkt8fzxl2djK7\nO2cns9/97nzer1demzlz5pzvzDOz+33OeZ4zIEmSJEmSJEmSJEmSJEmSJEmSJEmSJEmSJEmSJEmS\nJEmSJKkD9AP3NbluN7ARmNvk+hcX60saWj/NfwbHWi/xGX5+4nFIbfdHqQcgSYl1E3/0a/89AdwL\nXAjsswX2samN65fddkqHAz2pBzGCvYFPAzcCjxLvh5HGfDxwJ7AO+BWwENhxiHV3Ay4FflOs/7/A\nm0uOcVvgH4mJ9JPAPcDHgElDrP8K4HpgDbAa+DYwcwttOxfj9XOyidbGNpVIXP5yi4xGkiS1TTcx\nsbwMeGvx70TgC8SkcDWtHYXsB35ecizHN7n+xeR1puNixv945wEbgBXERH0jcOYw659arHMj8A6g\nD3gM+BHwrLp1pxHvhTXERPEdwE3F4+eVGONVxWMWAicAXy1uX9Rg3VcSycPPgPcDHyASiTXA/i1u\nOxf9NP8ZHGtbAdu08PgZjPwelSRJ40A38Uf7gw3ue19x3wda2H4/Jh2Ti58XExP68ew5wJTi/wcy\n/IRuR+Bx4FagUrP80OJxH6lb/1PF8jfULPsj4HvAb4Edmhjf3xTbOK9u+aeL5X9et/w24ozNH9cs\n241Ipv+rxW03q0Jzz61d+hm/SUerZtDc2bjxZmvirJokSR2jm6GTjjcX972nwfqN+i4uZvMkoJ+Y\n8OwBXE1MAFcDVxbLGo1lLpHw3E2Ueq0ATmlyfxATzH8B7gfWA78A5gM71a03DfgcUUr2BDHxvR34\nUN16k4AzgB/XrHclmx8pn8HABOgtwP8RZ4suYuCIfv2/ZhKsw4HvAGuJswjLgMMarLey2M8+wH8S\nR/MfBb4B7NLEfurNYvik4x3F/cc2uO8eYHndsgeJmNZ7W7Gdo5oY02XFus+rW/4nxfIv1Szbk4Gz\nFvW+SiSAta9LmW0PpZuB9/B7iffMkwxMil9OvG/vJhK2NUQ8D2+wrYuLbU0h3s+/Jt5/y4rt1HsO\n8Vx/S7xXbiISx34aJx1l31cvAa4rxvww8dmZBGwPfIb4nD0B3EzzZZm9bN7TUV22F3AO8b55Evg+\n8Pqa9bpp/Jmq7195S/Hc1jCQJP9tg7FsBXwCWFU8j7uI92SjMULzv2eqj98P+GzxfP4AzCnufwPx\nmlVLDlcB3wRe2GCMyljuNZqStKXswEAd/vbEhPps4g/hNxusP1QddqPlk4mJz63APxCTiZOJ0pcD\niMlUrfcBuwJfISZDbyXKvaYR9fbDeT7wXeL3+wVEQvFCInF6DTGRXlOs+w3gL4iJww+I570fUR/+\n6ZptXk5MPv6bmHj+MTGh/G7x+O/XjeHwYhxfLv6tAX5PHNX/C2KSXfU/Izyfk4HzgZ8QpUsVohTp\nKuBdDJ5QbyImzDcRSdHVwEuL9aYArx1hX2X9WfHzuw3u+x7wd0SJ1TriNduNmNg3WhciNt9oYp8P\nEhO8Wg8CDxXbaHZ8JxCT8iWj2PZIPgA8F1hA9Lk8UCw/nHj//xsxudyRSFCuJJK3Kxps67+ISX5f\nsf4HiaRyDyJhgDhy/l/FGC8lPmsHEEnC7xpss+z76k+I9//XiRi9lihXq06mJxEJwk5E0n4VsC+t\n9WtcAjxFnCHblnhNryJev1VEQncqkfxcWfyDgdcE4Czgo0Qfz8eL8R5ZPIdTiM9n1fnFc7+x2OfO\nxO+G+xo8jzK/Z6ouJz4L5xXb+xXxu+Ya4vfPOcRBgucBfwX8KVEWKEnShNBN46OFG4m6/L2GWL/R\nEfqLaXymYyNxhK/W4cXyf2mw7dXEBLVqa2KS+BSDj0I32t/VxB/z3eqWHwg8zcAR56nFY89v8Dxq\nHVKsVz8ZfEmxvaU1y2YU664nGrLrNRrvcJ5DTKDuZqBEC6CLgb6EqTXLVxbbr2/MPp+BI8dljHSm\nYxFxtqBRmUi1lGrP4na1VOuTDdZ9FgN9RSN5jMZJBEQp1YM1t08rttso2aqWUr1jlNseSnex3d/S\nuJm+vs8FItn9KZufGbqYxu/R6hnId9YseyeNy4yqiUHtmY7Rvq/qzw7cXiz/Vt3yalnmXzOyXoY+\n03FN3brV9+M5NctmMPR79GXFfWc1uO9bxO+Z6vN/UbHukrr19ife4xvqxtjs75na53Mjm1/A6LPF\nfUNdeEETiFevkqQwHzi4+HcoUU60I3GEsJVGcoijev9ct+wqYtLTqKzkcuLIctXTDJRyvHGY/Uwl\nxn4NkaDsWPNvFXE0sjoReoJIDl4J7D7MNo8ofp5dt/wHxKR7NnFEu9Z/EiVhrTqEmKR+gcFHbx8r\nlk0m4lXrF8B/1C27qfi5J1tWdQK9vsF9T9atU2bdkfbZaBvV7Tyrbt0y+yyz7ZFcSiQe9dbV7e+5\nxFnGm4gzA5MbPOZzdbcbxfNwomTnM3Xr/gvxfqk1mvfVg2x+xvM7xc8v1i1f1mB8o/H5utu3E+Nt\ndrvHEr97LmXw74Idic9uF/H5h/i90WifPwKuZXDPUpnfM7X+H5sfdHi0+PlmrL6Z8Ew6JCn8jDgS\ndyNxtO88or57D+DcFrf9KFEeUu8nRE399g2WN1oXNu8DqbU3MTl4R7G/+n97ESUTEJOFDxBHMu8j\nJhdfAA6q2+YexFHORmP68RBjatS3MBrV7dYfAR9u341q96vlNfXJUauqE+hGZzq2q1unzLoj7XOo\nBtzt6rZRdp9ltj2Sod4DOxMlV78mJtC/Id6b7yImyM9u8Jj6mDaK5wuAXzI4iYB4n9c/fjTvq0bf\n8/H7Ie6rLm/1/dbovfxIie3uS/w++Cmb/y74KvF6V3t6qs+30cGC+liW+T0z3HYgzmLdSZR5/Y44\nYPE+PPMxIZlVStLQbiNKLV5Ts2y4Gu1JI9zfyJb8/oDq0civEfXgjTxR8//5RJnEG4ja6jcTdd5f\nB45pYRxlJqdb2nBXx6oMc99oPFRs83lsPkF8HnFU96GadavL61WX1fdSDLXPRtuobufBunXL7LPM\ntkfS6D1QIfoi9iGOet9OlPhsIPpL3krjg6FDfUa2dDyHM9z7aqj7Wh1fq9utEK/d64bZ1o+HWD7S\ndqH53zNVjd4TjxC9RH9BnIGaQ5zZ6iNKAG8dxfg0Tpl0SNLwJhFlCFWPFD+nNVj3BUNs4znEEcX6\nhvF9iSOD9X+g92uwjeqy4S79eQ8xydiWOGPTjF8RjaAXEBO+rxEJx6eJq0/9nLiqzX7ADxuMaRPN\nf9tz2S9Cu7f4uT8DJTW1+4a0l0K9DTgJeFWDcbySOGpcnWj9kpjgN7rsbLXE5fYm93ks0dhcmwRM\nJ5rVr6pbl2J8FzbY5yYixqPZ9mi8pPjXV/yr9c7NVy/l58SktYvB5VTbEp/L2mby8f6+KmO4z9Pd\nRD/PA8TZjuFUP8P7ED0ster7s0bze2Y4G4mrV91c3H4x8b78OANlX5oALK+SpKFVa79rJ2b3EbXj\nh9St+yoGJo+N/EPd7SOIMoRGE7ljGXzEeRviKjV/ABbXrVs76fgdURp2JPEt1PUqDL5CV32N/kYG\nEotqUlVtkq3/zon9ifKzZTS+OlAja4sxPKfJ9a8jLvH5PjZv+H0fMbm8rslttcPVRMJ4CoP/nr6R\nKFe5vG79K4gr8tROpLYinsvv2byJt5FqQ3/9d8dUb9fu814ikTmKzb+n4yjgBgaX/ZXZ9mhUj7bX\nzz32Jz4PjSbQzSapVxGv5Wl1y9/D4IMGMP7fV2VUy8kalVx9rfh5Do3ne7WXS15U/Hw/g8+kvJhI\nXEb7e2Ykjca9gughavb3hDLhmQ5JCgcycCnXbYmrubyTqAn/eM16a4mr6rwD+Ffi6NwLictt3gXM\nbLDt3xJ/oHerWf9k4ixDb4P17yauVvWVYn9vJa5c849sXoJTX2rxHiIRWEo0kH6fmHC8gEgSLim2\ns3cxliuJ2vbfE2de3k0c5b2l2N71wL8Tl399DlFzvStxydx1wN83GP9Qvls87svEpOVponxi5RDr\nrwZOJy7T+z3ida9e2vQFRB9AfZNwq6Yw8JyqV+b5SwbeA1czkJj9lvheg08Tr9O/EcniaUQPzP+r\n2/Y/E5P9fyWu2vMQcVbpQOL99Hjd+v1EuckM4rsQIF63xcRlY6cSr9+fE+VJX2PzSxC/nziafwvR\n8FwhJtaw+QS97LbL+jHxXjudSHjvJhLvdxIXJjiwwWOaLSW6qNjOmUTCV71k7puJ5Kt2vpPifdUu\nvyPOPPwd8TwfJt5Hi4iEs7f4933iMrm/JBLQA4nv/Kj28PyY6LV5J/Fevoq4/O/JwB3F+rWJR7O/\nZ0byVeIz89/Ee3x74ntFdii2K0nShPGXxBH+DQxcKvcPRELwHzSeCO3AwJeQPU5M3l9JTHzqa6dv\nIibxM4g/5KuLf99i83Ks7uLxxzPw5YBPEkf+3sfmGu0P4ujhp4rHPUEkFHcRtdLVLy2bRkx87yzu\nX1fs77Ns/kV6WxGTtOoXvVW/HPBFdevNYPhLzFaIBv0HiNe4+lxHUvslbmsZ+kvc7qNxuUd3iX3N\nYPBlk6uXC63+v9E25hKTrieI981XGfpI727EZOo3xfrVMxGN/B8x+Z1St3xb4J+I5/skMen8GBGn\nRl5JTCQfI3qUvk18f0kjZbddr5vhX+vnE0lsdXJ8K/Am4hKr9ZdlHer9DRGP+pKx5xCvffXLAW8k\nLhtb/QzWa/V91WjMMPLnYKRtDLXdocbyZ8TY17L55YEheiOuJRKUJ4krTP0nm5e0/VEx5lXFej8g\nkpnqN9LXv6eb+T0z0vM5gkjkHyj2+TARryMarCtJkqQt7DlEYlb/vRPSWFtEJKxj2bgvSZKkMXAU\nUUpXfzllqV22a7DsJUTye/UYj0UTkFmrJEmS3k2UxS0mStT2YaAE69VE6ZQkSZIkjdqfEb0fvyIu\noPFbog/tgJSD0sThmQ4pgZkzZ2666y4PGkmSpCzcxdAXoGiKSYeUxqZNm7bkF1FrrPT29tLb25t6\nGBol45c345ev1LGrVCr4d3f0KpUKtJg3+OWAklTCypUrUw9BLTB+eTN++TJ2MumQJEnShNbT49Wn\nUzPpkKQS5s2bl3oIaoHxy5vxy1fq2FmWl549HVIa9nRIkqQs2NMhSWOsv78/9RDUAuOXN+OXL2Mn\nkw5JkiRJbWV5lZSG5VWSJCkLlldJkiRJI7CRPD2TDkkqwbrkvBm/vBm/fKWOXV9fX9L9y6RDkiRJ\nUpvZ0yGlYU+HJEljpFKp4N/d0bOnQ5IkSdK4Z9IhSSWkrktWa4xf3oxfvoydTDokSZI0ofX09KQe\nQsezp0NKw54OSZKUBXs6JEmSJI17Jh2SVIJ1yXkzfnkzfvkydjLpkCRJktRW9nRIadjTIUmSsmBP\nhyRJkjSC3t7e1EPoeCYdklSCdcl5M355M375Sh27vr6+pPuXSYckSZKkNrOnQ0rDng5JksZIpVLB\nv7ujZ0+HJEmSpHHPpEOSSkhdl6zWGL+8Gb98GTuZdEiSJGlC6+npST2EjmdPh5SGPR2SJCkL9nRI\nkiRJGvdMOiSpBOuS82b88mb88mXsZNIhSZIkqa3s6ZDSsKdDkiRlwZ4OSZIkaQS9vb2ph9DxTDok\nqQTrkvNm/PJm/PKVOnZ9fX1J9y+TDkmSJEltZk+HlIY9HZIkjZFKpYJ/d0fPng5JkiRJ455JhySV\nkLouWa0xfnkzfvkydjLpkCRJ0oTW09OTeggdz54OKQ17OiRJUhbs6ZAkSZI07pl0SFIJ1iXnzfjl\nzfjly9jJpEOSJElSW9nTIaVhT4ckScqCPR2SJEnSCHp7e1MPoeOZdEhSCdYl58345c345St17Pr6\n+pLuXyYdkiRJktrMng4pDXs6JEkaI5VKBf/ujp49HZIkSZLGPZMOSSohdV2yWmP88mb88mXsZNIh\nSZKkCa2npyf1EDqePR1SGvZ0SJKkLNjTIUmSJGncM+mQpBKsS86b8cub8cuXsZNJhyRJkqS2sqdD\nSsOeDkmSlAV7OiRJkqQR9Pb2ph5CxzPpkKQSrEvOm/HLm/HLV+rY9fX1Jd2/TDokSZIktZk9HVIa\n9nRIkjRGKpUK/t0dPXs6JEmSJI17Jh2SVELqumS1xvjlzfjly9jJpEOSJEkTWk9PT+ohdDx7OqQ0\n7OmQJElZsKdDkiRJ0rhn0iFJJViXnDfjlzfjly9jJ5MOSZIkSW1lT4eUhj0dkiQpC/Z0SJIkSSPo\n7e1NPYSOZ9IhSSVYl5w345c345ev1LHr6+tLun+ZdEiSJElqM3s6pDTs6ZAkaYxUKhX8uzt69nRI\nkiRJGvdMOiSphNR1yWqN8cub8cuXsZNJhyRJkia0np6e1EPoePZ0SGnY0yFJkrJgT4ckSZKkcc+k\nQ5JKsC45b8Yvb8YvX8ZOJh2SJEmS2sqeDikNezokSVIW7OmQJEmSRtDb25t6CB3PpEOSSrAuOW/G\nL2/GL1+pY9fX15d0/zLpkCRJktRm9nRIadjTIUnSGKlUKvh3d/Ts6ZAkSZI07pl0SFIJqeuS1Rrj\nlzfjly9jJ5MOSZIkTWg9PT2ph9Dx7OmQ0rCnQ5IkZcGeDkmSJEnjnkmHJJVgXXLejF/ejF++jJ1M\nOiRJkiS1lT0dUhr2dEiSpCzY0yFJkiSNoLe3N/UQOp5JhySVYF1y3oxf3oxfvlLHrq+vL+n+ZdIh\nSZIkqc3s6ZDSsKdDkqQxUqlU8O/u6NnTIUmSJGncM+mQpBJS1yWrNcYvb8YvX8ZOJh2SJEma0Hp6\nelIPoePZ0yGlYU+HJEnKgj0dkiRJksY9kw5JKsG65LwZv7wZv3wZO5l0SJIkSWorezqkNOzpkCRJ\nWbCnQ5IkSRpBb29v6iF0PJMOSSrBuuS8Gb+8Gb98pY5dX19f0v3LpEOSJElSm9nTIaVhT4ckSWOk\nUqng393Rs6dDkiRJ0rhn0iFJJaSuS1ZrjF/ejF++jJ1MOiRJkjSh9fT0pB5Cx7OnQ0rDng5JkpQF\nezokSZIkjXsmHZJUgnXJeTN+eTN++TJ2MumQJEmS1Fb2dEhp2NMhSZKyYE+HJEmSNILe3t7UQ+h4\nJh2SVIJ1yXkzfnkzfvlKHbu+vr6k+5dJhyRJkqQ2s6dDSsOeDkmSxkilUsG/u6NnT4ckSZKkcc+k\nQ5JKSF2XrNYYv7wZv3wZO5l0SJIkaULr6elJPYSOZ0+HlIY9HZIkKQv2dEiSJEka90w6JKkE65Lz\nZvzyZvzyZexk0iFJkiSprezpkNKwp0OSJGXBng5JkiRpBL29vamH0PFMOiSpBOuS82b88mb88pU6\ndn19fUn3L5MOSZIkSW1mT4eUhj0dkiSNkUqlgn93R8+eDkmSJEnjnkmHJJWQui5ZrTF+eTN++TJ2\nMumQJEnShNbT05N6CB3Png4pDXs6JElSFuzpkCRJkjTumXRIUgnWJefN+OXN+OXL2MmkQ5IkSVJb\n2dMhpWFPhyRJysKW6OmYtGWGIqmsj33qY6mHIElSR1h63VLmHDJn0LKdp+zM+9/9/kQj6jwmHVIi\nu//V7qmHoFFYcfsK9p61d+phaJSMX96MX75Sx27ZGcs47p+PG7Rs1Q2rEo2mM9nTIUmSJKmtTDok\nqQSPsubN+OXN+OXL2MmkQ5IkSVJbmXRIUgkrbl+ReghqgfHLm/HLl7GTSYckSZImtENPOjT1EDqe\n39MhpbFp/u3zU49BkqSOteqGVZx9+tmph5GFLfE9HZ7pkCRJktRWJh2SVIJ1yXkzfnkzfvkydjLp\nkCRJktRWJh2SVILXms+b8cub8cuXsZNJhyRJkia0RfMXpR5CxzPpkKQSrEvOm/HLm/HLV+rYLV64\nOOn+ZdIhSZIkqc1MOiSpBOuS82b88mb88mXsZNIhSZIkqa1MOiSphNR1yWqN8cub8cuXsZNJhyRJ\nkia0Q086NPUQOl4l9QCkDrVp/u3zU49BkqSOteqGVZx9+tmph5GFSqUCLeYNnumQJEmS1FYmHZJU\ngnXJeTN+eTN++TJ2MumQJEmS1FYmHZJUgteaz5vxy5vxy5exk0mHJEmSJrRF8xelHkLHM+mQpBKs\nS86b8cub8ctX6tgtXrg46f5l0iFJkiSpzUw6JKkE65LzZvzyZvzyZexk0iFJkiSprUw6JKmE1HXJ\nao3xy5vxy5exk0mHJEmSJrRDTzo09RA6XiX1AKQOtWn+7fNTj0GSpI616oZVnH362amHkYVKpQIt\n5g2e6ZAkSZLUViYdklSCdcl5M355M375MnYy6ZAkSZLUViYdklSC15rPm/HLm/HLl7GTSYckSZIm\ntEXzF6UeQscz6ZCkEqxLzpvxy5vxy1fq2C1euDjp/mXSIUmSJKnNTDokqQTrkvNm/PJm/PJl7GTS\nIUmSJKmtTDokqYTUdclqjfHLm/HLl7GTSYckSZImtENPOjT1EDpeJfUApA61af7t81OPQZKkjrXq\nhlWcffrZqYeRhUqlAi3mDZ7pkCRJktRWqZOObmAjMC3xOJpxH/DBEdbpBX7Y/qGMS/3AF1MPIoF+\n4AupB6GxY11y3oxf3oxfvoydmk06dgE+D9wDPAk8CCwBXl9iX/3kPSmdBfxLze2NwJGJxjKj2P/L\nmli3fpwri2UbgceJJOmkEvvupnGiuKn412kOBz6SehCSJEnj2aQm1pkBfAdYDfwDcBeRrBxMTMJn\ntGls483vGixL3RMzmv1vAvqI2HUB84D5wKPAN9q874no0dQD0NjyWvN5M355M375MnZq5kzHl4kj\n27OA/wB+BqwAvgS8pFjnQmBRg23fD5wKXATMAd5bbGsD8PyadV8KfI848v6/wAF12zqSOCL/ZLHN\nj9bdvxL4GDF5Xg08AHxomOc0GXgaeEXNsgeAn9TcPhhYy0BitpKB8qqVxc9vFM/n53Xb/zvgXmAN\n8C3guTX3VYBPFPt7EvgBcFjN/TNofBaj9oxFdX//Wyy/cfOnOKzHgIeLMX6CiOmbgN2L7R1Yt/5J\nwG+AF9bs6zfFuhfWrLcVcE5x36+B8xicnDwHuAR4BFgHXAfsV3P/vGJsBwE/Il7/Gxk5sd0IvBu4\nhngPrSDOyDwf+O9iO3cw8H6FOFNzBRGHdcX+5tVtdwfg0mJMDxHvqcXE+7mqn8Fn8LYhXoOVRHzv\nBd43wvglSVIbLZpfP03VWBsp6ZgGvJZIMNY1uH9N8XMB8Dpg15r7DiHKsi4F3g98l5ig7gr8MVGi\nVXUOcDox0f4dcHnNfQcC/04kPPsTZ1s+ApxSN5ZTibMwBwDnAp8CXjnE81oL3E5MTAH2BKYSk9Rd\nimXdwP8Afyhu15YOzSp+vqN4Pn9Wc98M4ChiEv/XxXhqL43wAWLy+uHi+XwLuBKYOcRYG3l58fO1\nxf5bLfNaD2wLrCIm6SfU3X8CEcd7gb8tlu1X7Pv9xe0KcCzwFPDnRHw+ALylZjsXE6/VYcVzWAdc\nC2xXs862RIznFdt5NvCVJp7Dx4n3zUwitlcQ77cvEjH4JZHwVG1XrPeG4rl8nkhaD6pZ5zNEsnw4\nkYQeCMxm8HuhvqzsEuA44v24DzAX+H0T41cmrEvOm/HLm/HLV+rYLV64OOn+NXLSsScxmfzJCOvd\nCvyUmGBVnQBcTSQRa4jJ6DriCPvDxNHpqk8ANxNHqP+RmKztVtz3QeJoch/RU/KvwKeBM+rG8F/E\nWZmfA+cX6/7VMGPuB15T/L8bWAbcVresf4jH/rb4+WjxXGpLryYRE+YfEa/LgrpxfIg4A/BvxRh7\ngFsY/szMUPv/XbH/siU+1bMP1bHuD1xfLFsIHENM/gH2Jc4IXUDErDqBrsbxsZrtLiea6e8hzgLd\nxMBzfyHwRuCdxGv9I2JyPoVIVqomEWfEbifObn2ageRwOJcAXy/2fQ6RPC4mzsD9jEhCZzLQi/IQ\nkVT8gDgrsZBI/o4p7p8MvJ1Ihm8AfgycyOD3bb0XEknWiUQyubJ4rpc1MX5JkpS5Rx55hCOOOILJ\nkyczY8YMrrjiiiHXveSSS5g1axZTp05l+vTpnHHGGWzYsAGAp556ihNPPJEZM2YwZcoUDjjgAK69\n9tqxehptMVJPR5m6/YXAycRZhmnE0ezDm3zsD2r+/8vi587ExHAfYvJY6zvEZH0ycdZiU902KB67\n0zD7vJk4Gj+JmNTeBDyr+P/VxNmM05scf61VDJ6I/5J4LhAT7D8uxl9rGfA3o9jXaFSIMy+9RGLx\nFDEhX1Dcfw1xZutI4mzBCUTp249H2G6jGNQ+932JCft3a+5fQyQW+9YsW08kCbXb2IY44zFcclW7\n74eLnz9ssGxnorxrK+KMyluIBHfbYj83Fev9KbA1kYhWVcuwhnIA8RxvGmadZ1zUcxE77rYjANtP\n3p7pe09/pua1ekTI2+Pv9t6z9h5X4/G28euk28bP26O9XVV/f39/PwDd3d1b5PZRRx0FwMMPP8yd\nd97Ja1/7WtavX8+8efM2W/+JJ55g7ty57Lfffuy7774cdthhnHzyyRxzzDG8/OUv5/nPfz7nnnsu\nu+yyC48//jhHH300CxYsYNddd91i4x3qdvX/K1euZEsZKamYRtTnfxz4ZBPr/oIoQ3kZceR+95r7\nbyImgX9fs6ybqNnfkZgIQpQn/ZyY9N8B/B/wn8CZNY87mCgD6iJq+O8jymg+O8L+ak0u9vmXxBHy\nI4plC4jk6Wpiolstr6rfx0bgzcTR8apeovzoxTXL5hWP6yKSjkeJo/+1E9OziPK0WUSJ10qiDOn/\nivu3Jibj1f3NYPBrNJz6cd5HnC26gJhE/6rBYz5FTKBfR8T048BXi/u62Txm0Pj1vpiBBPSwYgzb\nEj09VcuIROTDDH6tqoba33DPcUciyegGlhbL9iESp/2Ln2cQSeXfF+NeS7zHdyJKrGYCdwIvYKCH\nB+Ks1M/l9wJUAAAgAElEQVQYKEGrfd5HE6/t9kTP0HD8ckBJksbIu2a9i/q/u1v6ywEff/xxpk2b\nxvLly9lzzz0BmDt3Lrvtthuf/ORI02j43Oc+x0033cQ111zT8P6ZM2fS29vLEUccsdl9F198MQsX\nLuQVr3gFF110Ec997nO59NJLWbFiBT09Paxfv57zzjuP448/HoAlS5bw4Q9/mAceeIApU6Zw6qmn\nctpppw05trH4csBHiLKlU4im2nrPrlv3SqK05O0Mrp+HOKLezNWy6v0EeHXdstlEA/Djo9he1Vpi\nUv9OIhm4gziiP50o96nt52jkaeJoeRlriDMws+uWzyZKkyCSPBgoL4NotK/1VPGz7P6rfkckLY0S\nDogE4zVEmdNkohSs1X3/hHi/vapm2RQGkoCxNps4q3M5cZbkPmDvmvvvJWL88pplzyLGO5TvE8/x\noGHWUeZS1yWrNcYvb8YvX50Qu7vvvptJkyY9k3BAJArLly8f5lEDbr75Zvbfv/E049e//jV33303\nL3rRi4Z8/G233cbMmTN55JFHOOaYYzj66KO54447uPfee7nssss45ZRTWLcuWrRPPPFEFixYwJo1\na1i+fDkHHdT+qUszV696L5HZ3E4cTd6bOGr8HqJxu9ZCYsL+EgZf1QjiaPHLibMfO9J8tvQZ4mxE\nD7BXsf0PEkfjh1NpYh/9wNuIo+GbiKsNfa9Y1j/CY1cSZ1x2Ja7K1KzziLNAf0c8n38kJsCfLu5/\ngugFOYNocH5VzX1VDxfrvY7oXZhaYv/NuJs4A/Epojdjbc19q4jX6lDirEA1GR3q9a4u+xlx9mg+\n8XxfTPQ6rCbODoy1FUT8Xk28n88nziBVx7uWeA+fSyQR+xHJWIXBjeO1z/tu4qIHXyXK0/YA/oJ4\nP0mSpEQOPenQtu9j7dq1TJkyZdCyrq4uHnvssSEeMeDCCy/kjjvu4EMf2rzF9+mnn+bYY49l3rx5\n7LXXXkNuY4899mDu3LlUKhWOPvpoHnroIc4880y23nprDjnkELbZZhvuueceALbZZhuWL1/OmjVr\nmDp1KgccUH/h2C2vmaTjPqJc6jpiAnYX0Vj7JuLqRLX6iTMQ/QwuSYGYOD9FHNX+NXFGARp/oVzt\nsjuJq0H9LVHGcg5RBvOlEcbdzJfV9ROvQX/dsq0YOek4jTgbcD8DZVBD7bN22ReIxONTxPN5EwOX\nBK6qlu78L/F9Gh+r294fiHKedxDlT98aYayjcSHR43BB3fJfEAng2cSZkurlYhs99/plbyd6JK4h\nkrvtiMRpfd1j6o3mSwdH2s5ZxVi+TfT3PEac9ahd50NEOdU1xHv+LiL5frJum7WPOZ5Ior5AnN25\niDijownCa83nzfjlzfjlK3Xs3viuN7Z9H5MnT2bNmjWDlq1evZqurq4hHhGuuuoqPvrRj/Ltb3+b\nadMGf/fyxo0bOe6449huu+04//zzh93OLrvs8sz/t99+ewB22mmnQcvWro3jyN/85jdZsmQJM2bM\noLu7m1tvvXXkJ9iiLf0Fb9sTl8I9hWhCVr7OIJKEfVIPZBzZljjTcy7wuRa3ZU+HJEkJjUVPx3HH\nHcf06dM555xzGj7m2muv5fjjj2fJkiXMmjVr0H2bNm3ihBNO4P7772fJkiVsu+22Q+774osv5oIL\nLuCWW24B4J577mGvvfZi48aBi25Onz6dr3/967zqVQOV7hs2bOCLX/win/3sZ7n//vuH3P5Y9HQ0\nq0JcFejjRHPyv2+h7Wrs7QC8iDiT8vnEY0ntpcBbiUtHH0D0Ke1AXHhAHaoT6pInMuOXN+OXr06I\n3Q477MCRRx7JmWeeybp161i2bBmLFi3iuOOOa7j+jTfeyLHHHsuVV165WcIB8J73vIef/vSnXHPN\nNcMmHGU9/fTTXH755axevZqtttqKrq4uttpqtG3CzdtSScfuRKnN24ij4xuGX13j2JeIcrFlRP9F\npzuVuMjADUQPyxziYgCSJEmDfPnLX+aJJ55g55135m1vextf+cpX2Hff+GaA+++/n66uLh58ML4f\n+6yzzuKxxx7j9a9/PV1dXXR1dfGGN7wBgFWrVrFgwQLuuusudt1112fuH+p7PyqVSvVsxKBlQ7ns\nssvYY489mDp1KgsWLODyyy8fct0tZUuXV0lqjuVVkiQltKXLqyay8VReJUmSJI1Li+YvSj2EjmfS\nIUkldEJd8kRm/PJm/PKVOnaLFy5Oun+ZdEiSJElqM5MOSSoh9bXm1Rrjlzfjly9jJ5MOSZIkSW1l\n0iFJJaSuS1ZrjF/ejF++jJ1MOiRJkjShHXrSoamH0PH8ng4pDb+nQ5KkhPyejub5PR2SJEmSxj2T\nDkkqwbrkvBm/vBm/fBk7mXRIkiRJaiuTDkkqwWvN58345c345cvYyaRDkiRJE9qi+YtSD6HjmXRI\nUgnWJefN+OXN+OUrdewWL1ycdP8y6ZAkSZLUZiYdklSCdcl5M355M375MnYy6ZAkSZLUViYdklRC\n6rpktcb45c345cvYyaRDkiRJE9qhJx2aeggdr5J6AFKH2jT/9vmpxyBJUsdadcMqzj797NTDyEKl\nUoEW8wbPdEiSJElqK5MOSSrBuuS8Gb+8Gb98GTuZdEiSJElqK5MOSSrBa83nzfjlzfjly9jJpEOS\nJEkT2qL5i1IPoeOZdEhSCdYl58345c345St17BYvXJx0/zLpkCRJktRmJh2SVIJ1yXkzfnkzfvky\ndjLpkCRJktRWJh2SVELqumS1xvjlzfjly9jJpEOSJEkT2qEnHZp6CB2vknoAUofaNP/2+anHIElS\nx1p1wyrOPv3s1MPIQqVSgRbzBs90SJIkSWorkw5JKsG65LwZv7wZv3wZO5l0SJIkSWorkw5JKsFr\nzefN+OXN+OXL2MmkQ5IkSRPaovmLUg+h45l0SFIJ1iXnzfjlzfjlK3XsFi9cnHT/MumQJEmS1GYm\nHZJUgnXJeTN+eTN++TJ2MumQJEmS1FYmHZJUQuq6ZLXG+OXN+OXL2GlS6gFInWrVDatSD0Gj8Ot7\nf812q7dLPQyNkvHLm/HLV+rYzT549mZ/d3eesnOi0XSmSuoBSB1q06ZNm1KPQZIkaUSVSgVazBss\nr5IkSZLUViYdklRCf39/6iGoBcYvb8YvX8ZOJh2SJEmS2sqeDikNezokSVIW7OmQJEmSRtDb25t6\nCB3PpEOSSrAuOW/GL2/GL1+pY9fX15d0/zLpkCRJktRm9nRIadjTIUnSGKlUKvh3d/Ts6ZAkSZI0\n7pl0SFIJqeuS1Rrjlzfjly9jJ5MOSZIkTWg9PT2ph9Dx7OmQ0rCnQ5IkZcGeDkmSJEnjnkmHJJVg\nXXLejF/ejF++jJ1MOiRJkiS1lT0dUhr2dEiSpCzY0yFJkiSNoLe3N/UQOp5JhySVYF1y3oxf3oxf\nvlLHrq+vL+n+ZdIhSZIkqc3s6ZDSsKdDkqQxUqlU8O/u6NnTIUmSJGncM+mQpBJS1yWrNcYvb8Yv\nX8ZOJh2SJEma0Hp6elIPoePZ0yGlsemj53409RgkSeoIO0/Zmfe/+/2ph5GtLdHTMWnLDEVSWbv/\n1e6phyBJUkdYdcOq1EPoeJZXSVIJK25fkXoIaoHxy5vxy5exk0mHJEmSpLYy6ZCkEvaetXfqIagF\nxi9vxi9fxk4mHZIkSZrQll63NPUQOp5JhySVYF1y3oxf3oxfvlLHbtn1y5LuXyYdkiRJktrMpEOS\nSrAuOW/GL2/GL1/GTiYdkiRJktrKpEOSSkhdl6zWGL+8Gb98GTuZdEiSJGlCm33w7NRD6HgmHZJU\ngnXJeTN+eTN++UoduzmHzEm6f5l0SJIkSWozkw5JKsG65LwZv7wZv3wZO5l0SJIkSWorkw5JKiF1\nXbJaY/zyZvzyZexk0iFJkqQJbel1S1MPoeOZdEhSCdYl58345c345St17JZdvyzp/mXSIUmSJKnN\nTDokqQTrkvNm/PJm/PJl7GTSIUmSJKmtTDokqYTUdclqjfHLm/HLl7GTSYckSZImtNkHz049hI5n\n0iFJJViXnDfjlzfjl6/UsZtzyJyk+5dJhyRJkqQ2M+mQpBKsS86b8cub8cuXsZNJhyRJkqS2MumQ\npBJS1yWrNcYvb8YvX8ZOJh2SJEma0JZetzT1EDqeSYcklWBdct6MX96MX75Sx27Z9cuS7l8mHZIk\nSZLazKRDkkqwLjlvxi9vxi9fxk4mHZIkSZLayqRDkkpIXZes1hi/vBm/fBk7mXRIkiRpQpt98OzU\nQ+h4Jh2SVIJ1yXkzfnkzfvlKHbs5h8xJun+ZdEiSJElqM5MOSSrBuuS8Gb+8Gb98GTuZdEiSJElq\nK5MOSSohdV2yWmP88mb88mXsZNIhSZKkCW3pdUtTD6HjmXRIUgnWJefN+OXN+OUrdeyWXb8s6f5l\n0iFJkiSpzUw6JKkE65LzZvzyZvzyZew0kZOObmAjMC3xOJpxH/DBEdbpBX7Y/qGMWj/wxdSDSKAf\n+ELqQUiSJI1n4yHp2AX4PHAP8CTwILAEeH2JbfST94R3FvAvNbc3AkduoW3Xb2tlsWwj8DiRyJxU\nYnvdNE7mNhX/Os3hwEdSD0JjJ3Vdslpj/PJm/PJl7DQp8f5nAN8BVgP/ANxFJEIHE5PwGakGNsZ+\n12BZpU372gT0Ea9vFzAPmA88CnyjxHbaNb7cPJp6AJIkaXizD56deggdL/WZji8TR81nAf8B/AxY\nAXwJeEmxzoXAorrH/RFwP3AqcBEwB3hvsa0NwPNr1n0p8D3iqP7/AgfUbetI4mj/k8U2P1p3/0rg\nY8TEfDXwAPChYZ7TZOBp4BU1yx4AflJz+2BgLQNJ30oGyqtWFj+/UTyfn9dt/++Ae4E1wLeA5w4z\nlqE8BjxcbOcTxOv+JmD3Yp8H1q1/EvAb4IXAjcWy3xTrXliz3lbAOcV9vwbOY3By8hzgEuARYB1w\nHbBfzf3zirEdBPyIeI1uZOTkcyPwbuAaIs4riDMyzwf+u9jOHQy8pyDO1FxBxGZdsb95ddvdAbi0\nGNNDRNwXE++5qn4Gn2XbhngNVhLvqXuB940wfmXEuuS8Gb+8Gb98pY7dnEPmJN2/0iYd04DXEgnG\nugb3ryl+LgBeB+xac98hRFnWpcD7ge8Sk99dgT8mSrSqzgFOB15GnFG4vOa+A4F/JxKe/YmzLR8B\nTqkby6nEWZgDgHOBTwGvHOJ5rQVuJya9AHsCU4kJ8C7Fsm7gf4A/FLdry5JmFT/fUTyfP6u5bwZw\nFJEg/HUxnrOHGEcZ64FtgVXEJP2EuvtPIF7re4G/LZbtV4zv/cXtCnAs8BTw58Rr+AHgLTXbuZh4\nPocBLyfifi2wXc062xJxmFds59nAV5p4Dh8nYjuTeP2vIN4TXyRep18SCU/VdsV6byiey+eJxPKg\nmnU+QyS0hxOJ4oHAbAbHq76s7BLgOOI9sw8wF/h9E+OXJEmasFImHXsSE9WfjLDercBPiclb1QnA\n1UQSsYaY6K4jjt4/TBz5rvoEcDNx9PsfiYngbsV9HySOVPcRPSX/CnwaOKNuDP9FnJX5OXB+se5f\nDTPmfuA1xf+7gWXAbXXL+od47G+Ln48Wz6W29GoSMRn/EfG6LBhhHEOpnn2obm9/4Ppi2ULgGGLy\nD7AvcdbmAuJ1rU6gq6/1YzXbXU40vN9DnKm5qWZ8LwTeCLyTeD1+REzOpxDJSu1zfC+REPyQiEd3\nE8/pEuDrxb7PIRK8xcRZsp8RieJMBnpRHiKSih8QZyUWAlcWzx3ijNXbiYT1BuDHwIkMfm/VeyGR\nZJ1InIVaWTzXy5oYvzJhXXLejF/ejF++OiV2jzzyCEcccQSTJ09mxowZXHHFFUOue8kllzBr1iym\nTp3K9OnTOeOMM9iwYcMz959//vnMmjWL7bbbjre//e1jMfy2StnTUaYnYCFwMnGWYRpxpPzwJh/7\ng5r//7L4uTMx6dyHmJjW+g7QQ0w61xJHsX9Qt85DwE7D7PNm4kj/JGLCfBPwrOL/VxNnM05vcvy1\nVjF4kv9L4rmUUSHOjvQSicVTxIR8QXH/NcTZpyOJswUnEOVpPx5hu41ep9rx7UtM2L9bc/8aIrHY\nt2bZeiJJqN3GNsQZj+H6J2r3/XDx84cNlu1MlHdtRZxReQuRhG5b7OemYr0/BbYmksWqahnWUA4g\nnuNNw6zzjIt6LmLH3XYEYPvJ2zN97+nPnH6u/nL2tre97W1ve3si3K5Ktf/tiqKK/v5+ALq7u9ty\n+6ijjgLg4Ycf5s477+S1r30t69evZ968eZut/8QTTzB37lz2228/9t13Xw477DBOPvlkjjnmGLq7\nu3ne857H4YcfzvOe97xnXr92j796u/r/lStXsqWkbAaeRtT+fxz4ZBPr/oIocXkZUVu/e839NxET\nzL+vWdZN9APsSEwyIcqTfk5M+u8A/g/4T+DMmscdTJQYdRH9AfcRJTqfHWF/tSYX+/xL4uj7EcWy\nBUTydDUxia6WV9XvYyPwZuLIe1UvUdr04ppl84rHdQ0xjkbbuo84o3MBMYn+VYPHfIqYQL+OeN0/\nDny1uK+bzV9XaPyaXMxAknhYMYZtib6bqmVEIvLhIZ7PUPsb7jnuSCQZ3cDSYtk+ROK0f/HzDCLx\n+/ti3GuJ9+FORInVTOBO4AUM9NkA3EIkRdUStNrnfTTx2m5P9PUMZ9P82+ePsIokSdoSVt2wirNP\n3xIV6UN7/PHHmTZtGsuXL2fPPfcEYO7cuey222588pMjTXXhc5/7HDfddBPXXHPNoOWf+MQnePDB\nB7nooouGeCRcfPHFLFy4kFe84hVcdNFFPPe5z+XSSy9lxYoV9PT0sH79es477zyOP/54AJYsWcKH\nP/xhHnjgAaZMmcKpp57KaaedNuT2K5UKtJg3pCyveoQoWzqFaNit9+y6da8kylbezuDafIij9aM5\na/MT4NV1y2YTzcWPj2J7VWuJhOadRPnQHcTZgulEKVFtP0cjTxNH4tvld0Ty1SjhgEgwXkOUOU0G\n/q3mvqeKn2XH9xPi/faqmmVTGEgCxtps4qzO5cRZkvuAvWvuv5eIw8trlj2LGO9Qvk88x4OGWUeS\nJI2xpdctHXmlFt19991MmjTpmYQDYObMmSxfvrypx998883sv//m04xNm5r7RoLbbruNmTNn8sgj\nj3DMMcdw9NFHc8cdd3Dvvfdy2WWXccopp7BuXbRRn3jiiSxYsIA1a9awfPlyDjqo/VOX1Fevei+R\nNd1OHKnemzgi/R6icbvWQmLC/hIGXzEJ4kj0y4mzHzvSfCb2GeJsRA+wV7H9DxJH+odTaWIf/cDb\niCPtm4grGX2vWNY/wmNXEmdcdiWu+DTW7ibOQHyK6M1YW3PfKuL5HEqcFagmjEO9JtVlPyPO8Mwn\nJvwvJnodVhNnB8baCuI1fjXxnjufOBNWHe9a4n12LpFE7EckYxUGN47XPu+7iQsTfJUoT9sD+Asi\n5pogOqUueaIyfnkzfvlKHbtl1y9r+z7Wrl3LlClTBi3r6uriscceG+IRAy688ELuuOMOPvShzS+Q\nWpxlGNEee+zB3LlzqVQqHH300Tz00EOceeaZbL311hxyyCFss8023HPPPQBss802LF++nDVr1jB1\n6lQOOKD+4q5bXuqk4z6iXOo6YnJ3F9G0+ybiyke1+okzEP0MLneBaDZ+ijhi/mvijAI0/rK62mV3\nEleD+luiROYcosTmSyOMu5kvwusnXt/+umVbMXLScRpxpuF+4ozJcPts1xfyXUj0OFxQt/wXRJJ2\nNnGmpHq52Ebjq1/2dqJH4hoiAduOKOFaX/eYeqN5jiNt56xiLN8menAeI8561K7zIaKc6hrifXkX\nkSA/WbfN2sccTyRRXyDO7lxEnNGRJEkT2OTJk1mzZs2gZatXr6ara7gqeLjqqqv46Ec/yre//W2m\nTav/7uXmz3Tssssuz/x/++23B2CnnXYatGzt2jiO/M1vfpMlS5YwY8YMuru7ufXWW5vaRyty+oK3\n7YlL4Z5CNDirvc4gkoR9Ug9kHNmWONNzLvC5FrdlT4ckSWPkXbPe1fTkfbQa9XQcd9xxTJ8+nXPO\nOafhY6699lqOP/54lixZwqxZsxqu02xPxwUXXMAtt9wCwD333MNee+3Fxo0DF92cPn06X//613nV\nqwYq3Tds2MAXv/hFPvvZz3L//fcPuf3cezqaVSGuOPRxovH539MOZ8LbAXgR0Rj9+cRjSe2lwFuJ\nyzsfQPQS7UBcHECSJOkZO+ywA0ceeSRnnnkm69atY9myZSxatIjjjjuu4fo33ngjxx57LFdeeWXD\nhGPDhg08+eST/OEPf2DDhg2sX79+0CV1R+vpp5/m8ssvZ/Xq1Wy11VZ0dXWx1VbtbCUOOSQduxNl\nPG8jjry3/mprOF8iSrqWEf0Xne5U4kIANxA9LHOISyarQ6WuS1ZrjF/ejF++OiV2X/7yl3niiSfY\neeededvb3sZXvvIV9t03vhng/vvvp6uriwcfjO+wPuuss3jsscd4/etfT1dXF11dXbzhDW94Zlv/\n9E//xLOe9SzOPfdcLrvsMrbffnvOPrvxFbgqlcpmvR/D9YJcdtll7LHHHkydOpUFCxZw+eWXD7nu\nlpJTeZU0kVhelakVt6945vrvyo/xy5vxy1fq2H3tH77GLdfdkmz/ueuU8ipJGjec8OTN+OXN+OUr\ndezmHDIn6f5l0iFJkiSpzUw6JKmETqlLnqiMX96MX76MnUw6JEmSJLWVSYcklZC6LlmtMX55M375\nMnYy6ZAkSdKEtvS6pamH0PFMOiSpBOuS82b88mb88pU6dsuuX5Z0/zLpkCRJktRmJh2SVIJ1yXkz\nfnkzfvkydjLpkCRJktRWJh2SVELqumS1xvjlzfjly9jJpEOSJEkT2uyDZ6ceQscz6ZCkEqxLzpvx\ny5vxy1fq2M05ZE7S/cukQ5IkSVKbmXRIUgnWJefN+OXN+OXL2MmkQ5IkSVJbmXRIUgmp65LVGuOX\nN+OXL2Mnkw5JkiRNaEuvW5p6CB3PpEOSSrAuOW/GL2/GL1+pY7fs+mVJ9y+TDkmSJEltZtIhSSVY\nl5w345c345cvYyeTDkmSJEltZdIhSSWkrktWa4xf3oxfvoydTDokSZI0oc0+eHbqIXQ8kw5JKsG6\n5LwZv7wZv3yljt2cQ+Yk3b9MOiRJkiS1mUmHJJVgXXLejF/ejF++jJ1MOiRJkiS1lUmHJJWQui5Z\nrTF+eTN++TJ2MumQJEnShLb0uqWph9DxTDokqQTrkvNm/PJm/PKVOnbLrl+WdP8y6ZAkSZLUZiYd\nklSCdcl5M355M375MnYy6ZAkSZLUViYdklRC6rpktcb45c345cvYyaRDkiRJE9rsg2enHkLHM+mQ\npBKsS86b8cub8ctX6tjNOWRO0v3LpEOSJElSm5l0SFIJ1iXnzfjlzfjly9jJpEOSJElSW5l0SFIJ\nqeuS1Rrjlzfjly9jJ5MOSZIkTWhLr1uaeggdb1LqAUidatUNq1IPQaOw6t5V7P6nu6cehkbJ+OXN\n+OUrdeyWXb8s2b4VKqkHIHWoTZs2bUo9Bo1Cf38/3d3dqYehUTJ+eTN++Uodu0qlgn93R69SqUCL\neYNJh5SGSYckSWPEpKM1WyLpsKdDkiRJUluZdEhSCf39/amHoBYYv7wZv3wZO5l0SJIkaULr6elJ\nPYSOZ0+HlIY9HZIkKQv2dEiSJEka90w6JKkE65LzZvzyZvzyZexk0iFJkiSprezpkNKwp0OSJGXB\nng5JkiRpBL29vamH0PFMOiSpBOuS82b88mb88pU6dn19fUn3L5MOSZIkSW1mT4eUhj0dkiSNkUql\ngn93R8+eDkmSJEnjnkmHJJWQui5ZrTF+eTN++TJ2MumQJEnShNbT05N6CB3Png4pDXs6JElSFuzp\nkCRJkjTumXRIUgnWJefN+OXN+OXL2MmkQ5IkSVJb2dMhpWFPhyRJyoI9HZIkSdIIent7Uw+h45l0\nSFIJ1iXnzfjlzfjlK3Xs+vr6ku5fJh2SJEmS2syeDikNezokSRojlUoF/+6Onj0dkiRJksY9kw5J\nKiF1XbJaY/zyZvzyZexk0iFJkqQJraenJ/UQOp49HVIa9nRIkqQs2NMhSZIkadwz6ZCkEqxLzpvx\ny5vxy5exk0mHJEmSpLayp0NKw54OSZKUBXs6JEmSpBH09vamHkLHM+mQpBKsS86b8cub8ctX6tj1\n9fUl3b9MOiRJkiS1mT0dUhr2dEiSNEYqlQr+3R09ezokSZIkjXsmHZJUQuq6ZLXG+OXN+OXL2Mmk\nQ5IkSRNaT09P6iF0PHs6pDTs6ZAkSVmwp0OSJEnSuGfSIUklWJecN+OXN+OXL2Mnkw5JkiRJbWVP\nh5SGPR2SJCkL9nRIkiRJI+jt7U09hI5n0iFJJViXnDfjlzfjl6/Usevr60u6f5l0SJIkSWozezqk\nNOzpkCRpjFQqFfy7O3r2dEiSJEka90w6JKmE1HXJao3xy5vxy5exk0mHJEmSJrSenp7UQ+h49nRI\nadjTIUmSsmBPhyRJkqRxz6RDkkqwLjlvxi9vxi9fxk4mHZIkSZLayp4OKQ17OiRJUhbs6ZAkSZJG\n0Nvbm3oIHc+kQ5JKsC45b8Yvb8YvX6lj19fXl3T/MumQJEmS1Gb2dEhp2NMhSdIYqVQq+Hd39Ozp\nkCRJkjTumXRIUgmp65LVGuOXN+OXL2Mnkw5JkiRNaD09PamH0PHs6ZDSsKdDkiRlwZ4OSZIkSeOe\nSYcklWBdct6MX96MX76MnUw6JEmSJLWVPR1SGvZ0SJKkLNjTIUmSJI2gt7c39RA6nkmHJJVgXXLe\njF/ejF++Useur68v6f5l0iFJpXz/+99PPQS1wPjlzfjly9jJpEOSSnj00UdTD0EtMH55M375MnYy\n6ZAkSZLUViYdklTCypUrUw9BLTB+eTN++TJ28pK5UhrfB2amHoQkSVIT7gJemnoQkiRJkiRJkiRJ\nkgteCbgAAAKlSURBVCRJkiRJkiRJkpTS64CfAj8DzhhinS8U998FHFDysWqvVuK3EvgBcCdwW/uG\nqGGMFL99gO8CTwKnlXys2quV2K3Ez15qI8XvWOJ35g+A7wAvKfFYtVcrsVuJnz0pia2Ae4AZwNbE\nFar2rVvnb4Alxf9fAdxa4rFqr1biB3AfMK29Q9QwmonfTsAs4CwGT1z9/KXVSuzAz15qzcTvz4Gp\nxf9fh3/7xotWYgclP3t+T4e05byc+PCuBJ4G/g14U906hwGXFP//HvBsYNcmH6v2Gm38dqm538uQ\np9NM/H4D3F7cX/axap9WYlflZy+dZuL3XWB18f/vAX9S4rFqn1ZiV9X0Z8+kQ9pyngc8UHP7wWJZ\nM+vs1sRj1V6txA9gE3A9MTE6qU1j1NCaiV87HqvWtfr6+9lLq2z8TmTgjLGfvbRaiR2U/OxNGsUA\nJTW2qcn1PCI3PrUav9nAQ0QZyHVEjewtW2Bcak6z8dvSj1XrWn39Xw38Ej97qZSJ32uAE4iYlX2s\ntrxWYgclP3ue6ZC2nF8A02tuTyeOGgy3zp8U6zTzWLXXaOP3i+L/DxU/fwN8izhtrbHTymfIz19a\nrb7+vyx++tlLo9n4vQRYSJSp/r7kY9UercQO/OxJyUwC7iUasrZh5EbkVzLQkNXMY9VercTvWUBX\n8f8diCt8/HUbx6rNlfkM9TK4GdnPX1qtxM7PXnrNxO/5RO/AK0fxWLVPK7Hzsycl9npgBfEB/Uix\n7F3Fv6rzi/vvAl42wmM1tkYbvxcQv6y/D/wI45fKSPHblahfXk0crbsfmDzMYzV2Rhs7P3vjw0jx\n+yrwO+LSqvWXV/Wzl9ZoY+dnT5IkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSWrg\n/wPnZgT7CL7wcwAAAABJRU5ErkJggg==\n", + "text": [ + "
\n", + "
" + ] + } + ], + "metadata": {} + } + ] +} \ No newline at end of file diff --git a/tutorials/scope_resolution_legb_rule.ipynb b/tutorials/scope_resolution_legb_rule.ipynb new file mode 100644 index 0000000..58adb8a --- /dev/null +++ b/tutorials/scope_resolution_legb_rule.ipynb @@ -0,0 +1,1159 @@ +{ + "cells": [ + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[Sebastian Raschka](http://www.sebastianraschka.com) \n", + "\n", + "- [Link to the containing GitHub Repository](https://github.com/rasbt/python_reference)\n", + "- [Link to this IPython Notebook on GitHub](https://github.com/rasbt/python_reference/blob/master/tutorials/scope_resolution_legb_rule.ipynb)\n" + ] + }, + { + "cell_type": "code", + "execution_count": 1, + "metadata": { + "collapsed": false + }, + "outputs": [], + "source": [ + "%load_ext watermark" + ] + }, + { + "cell_type": "code", + "execution_count": 2, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "Sebastian Raschka 01/27/2016 \n", + "\n", + "CPython 3.5.1\n", + "IPython 4.0.3\n" + ] + } + ], + "source": [ + "%watermark -a 'Sebastian Raschka' -v -d" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[More information](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/ipython_magic/watermark.ipynb) about the `watermark` magic command extension." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "I would be happy to hear your comments and suggestions. \n", + "Please feel free to drop me a note via\n", + "[twitter](https://twitter.com/rasbt), [email](mailto:bluewoodtree@gmail.com), or [google+](https://plus.google.com/+SebastianRaschka).\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#A beginner's guide to Python's namespaces, scope resolution, and the LEGB rule" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "This is a short tutorial about Python's namespaces and the scope resolution for variable names using the LEGB-rule. The following sections will provide short example code blocks that should illustrate the problem followed by short explanations. You can simply read this tutorial from start to end, but I'd like to encourage you to execute the code snippets - you can either copy & paste them, or for your convenience, simply [download this IPython notebook](https://raw.githubusercontent.com/rasbt/python_reference/master/tutorials/scope_resolution_legb_rule.ipynb)." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Sections " + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "\n", + "- [Introduction to namespaces and scopes](#introduction) \n", + "- [1. LG - Local and Global scopes](#section_1) \n", + "- [2. LEG - Local, Enclosed, and Global scope](#section_2) \n", + "- [3. LEGB - Local, Enclosed, Global, Built-in](#section_3) \n", + "- [Self-assessment exercise](#assessment)\n", + "- [Conclusion](#conclusion) \n", + "- [Solutions](#solutions)\n", + "- [Warning: For-loop variables \"leaking\" into the global namespace](#for_loop)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Objectives\n", + "- Namespaces and scopes - where does Python look for variable names?\n", + "- Can we define/reuse variable names for multiple objects at the same time?\n", + "- In which order does Python search different namespaces for variable names?" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Introduction to namespaces and scopes" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### Namespaces" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Roughly speaking, namespaces are just containers for mapping names to objects. As you might have already heard, everything in Python - literals, lists, dictionaries, functions, classes, etc. - is an object. \n", + "Such a \"name-to-object\" mapping allows us to access an object by a name that we've assigned to it. E.g., if we make a simple string assignment via `a_string = \"Hello string\"`, we created a reference to the `\"Hello string\"` object, and henceforth we can access via its variable name `a_string`.\n", + "\n", + "We can picture a namespace as a Python dictionary structure, where the dictionary keys represent the names and the dictionary values the object itself (and this is also how namespaces are currently implemented in Python), e.g., \n", + "\n", + "
a_namespace = {'name_a':object_1, 'name_b':object_2, ...} \n",
+ "\n",
+ "\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Now, the tricky part is that we have multiple independent namespaces in Python, and names can be reused for different namespaces (only the objects are unique, for example:\n",
+ "\n",
+ "a_namespace = {'name_a':object_1, 'name_b':object_2, ...}\n",
+ "b_namespace = {'name_a':object_3, 'name_b':object_4, ...}\n",
+ "\n",
+ "For example, everytime we call a `for-loop` or define a function, it will create its own namespace. Namespaces also have different levels of hierarchy (the so-called \"scope\"), which we will discuss in more detail in the next section."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Scope"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "In the section above, we have learned that namespaces can exist independently from each other and that they are structured in a certain hierarchy, which brings us to the concept of \"scope\". The \"scope\" in Python defines the \"hierarchy level\" in which we search namespaces for certain \"name-to-object\" mappings. \n",
+ "For example, let us consider the following code:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 1,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [
+ {
+ "name": "stdout",
+ "output_type": "stream",
+ "text": [
+ "1 global\n",
+ "5 in foo()\n"
+ ]
+ }
+ ],
+ "source": [
+ "i = 1\n",
+ "\n",
+ "def foo():\n",
+ " i = 5\n",
+ " print(i, 'in foo()')\n",
+ "\n",
+ "print(i, 'global')\n",
+ "\n",
+ "foo()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Here, we just defined the variable name `i` twice, once on the `foo` function."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "- `foo_namespace = {'i':object_3, ...}` \n",
+ "- `global_namespace = {'i':object_1, 'name_b':object_2, ...}`"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "So, how does Python know which namespace it has to search if we want to print the value of the variable `i`? This is where Python's LEGB-rule comes into play, which we will discuss in the next section."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Tip:\n",
+ "If we want to print out the dictionary mapping of the global and local variables, we can use the\n",
+ "the functions `global()` and `local()`"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 11,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [
+ {
+ "name": "stdout",
+ "output_type": "stream",
+ "text": [
+ "loc in foo(): True\n",
+ "loc in global: False\n",
+ "glob in global: True\n"
+ ]
+ }
+ ],
+ "source": [
+ "#print(globals()) # prints global namespace\n",
+ "#print(locals()) # prints local namespace\n",
+ "\n",
+ "glob = 1\n",
+ "\n",
+ "def foo():\n",
+ " loc = 5\n",
+ " print('loc in foo():', 'loc' in locals())\n",
+ "\n",
+ "foo()\n",
+ "print('loc in global:', 'loc' in globals()) \n",
+ "print('glob in global:', 'foo' in globals())"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Scope resolution for variable names via the LEGB rule."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "We have seen that multiple namespaces can exist independently from each other and that they can contain the same variable names on different hierachy levels. The \"scope\" defines on which hierarchy level Python searches for a particular \"variable name\" for its associated object. Now, the next question is: \"In which order does Python search the different levels of namespaces before it finds the name-to-object' mapping?\" \n",
+ "To answer is: It uses the LEGB-rule, which stands for\n",
+ "\n",
+ "**Local -> Enclosed -> Global -> Built-in**, \n",
+ "\n",
+ "where the arrows should denote the direction of the namespace-hierarchy search order. \n",
+ "\n",
+ "- *Local* can be inside a function or class method, for example. \n",
+ "- *Enclosed* can be its `enclosing` function, e.g., if a function is wrapped inside another function. \n",
+ "- *Global* refers to the uppermost level of the executing script itself, and \n",
+ "- *Built-in* are special names that Python reserves for itself. \n",
+ "\n",
+ "So, if a particular name:object mapping cannot be found in the local namespaces, the namespaces of the enclosed scope are being searched next. If the search in the enclosed scope is unsuccessful, too, Python moves on to the global namespace, and eventually, it will search the built-in namespace (side note: if a name cannot found in any of the namespaces, a *NameError* will is raised).\n",
+ "\n",
+ "**Note**: \n",
+ "Namespaces can also be further nested, for example if we import modules, or if we are defining new classes. In those cases we have to use prefixes to access those nested namespaces. Let me illustrate this concept in the following code block:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 8,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [
+ {
+ "name": "stdout",
+ "output_type": "stream",
+ "text": [
+ "3.141592653589793 from the math module\n",
+ "3.141592653589793 from the numpy package\n",
+ "3.141592653589793 from the scipy package\n"
+ ]
+ }
+ ],
+ "source": [
+ "import numpy\n",
+ "import math\n",
+ "import scipy\n",
+ "\n",
+ "print(math.pi, 'from the math module')\n",
+ "print(numpy.pi, 'from the numpy package')\n",
+ "print(scipy.pi, 'from the scipy package')"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "(This is also why we have to be careful if we import modules via \"`from a_module import *`\", since it loads the variable names into the global namespace and could potentially overwrite already existing variable names)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "\n", + "
\n", + "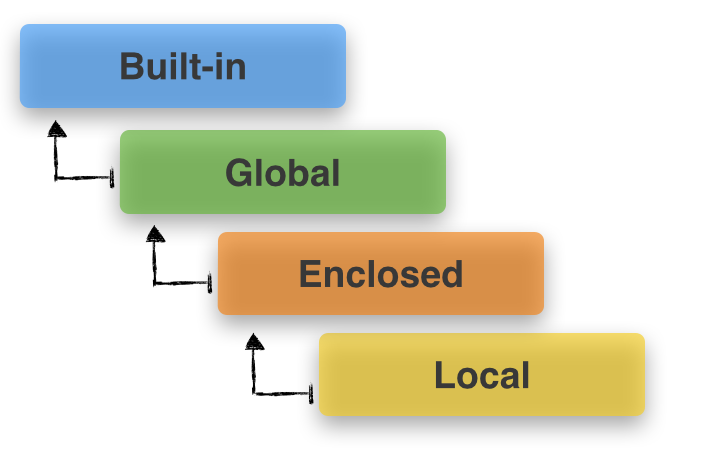\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## 1. LG - Local and Global scopes" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "**Example 1.1** \n", + "As a warm-up exercise, let us first forget about the enclosed (E) and built-in (B) scopes in the LEGB rule and only take a look at LG - the local and global scopes. \n", + "What does the following code print?" + ] + }, + { + "cell_type": "code", + "execution_count": 1, + "metadata": { + "collapsed": false + }, + "outputs": [], + "source": [ + "a_var = 'global variable'\n", + "\n", + "def a_func():\n", + " print(a_var, '[ a_var inside a_func() ]')\n", + "\n", + "a_func()\n", + "print(a_var, '[ a_var outside a_func() ]')" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "**a)**\n", + "
raises an error\n", + "\n", + "**b)** \n", + "
\n", + "global value [ a_var outside a_func() ]\n", + "\n", + "**c)** \n", + "
global value [ a_var inside a_func() ] \n", + "global value [ a_var outside a_func() ]\n", + "\n", + "\n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[go to solution](#solutions)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### Here is why:\n", + "\n", + "We call `a_func()` first, which is supposed to print the value of `a_var`. According to the LEGB rule, the function will first look in its own local scope (L) if `a_var` is defined there. Since `a_func()` does not define its own `a_var`, it will look one-level above in the global scope (G) in which `a_var` has been defined previously.\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "**Example 1.2** \n", + "Now, let us define the variable `a_var` in the global and the local scope. \n", + "Can you guess what the following code will produce?" + ] + }, + { + "cell_type": "code", + "execution_count": 2, + "metadata": { + "collapsed": false + }, + "outputs": [], + "source": [ + "a_var = 'global value'\n", + "\n", + "def a_func():\n", + " a_var = 'local value'\n", + " print(a_var, '[ a_var inside a_func() ]')\n", + "\n", + "a_func()\n", + "print(a_var, '[ a_var outside a_func() ]')" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "**a)**\n", + "
raises an error\n", + "\n", + "**b)** \n", + "
local value [ a_var inside a_func() ]\n", + "global value [ a_var outside a_func() ]\n", + "\n", + "**c)** \n", + "
global value [ a_var inside a_func() ] \n", + "global value [ a_var outside a_func() ]\n" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[go to solution](#solutions)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### Here is why:\n", + "\n", + "When we call `a_func()`, it will first look in its local scope (L) for `a_var`, since `a_var` is defined in the local scope of `a_func`, its assigned value `local variable` is printed. Note that this doesn't affect the global variable, which is in a different scope." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "However, it is also possible to modify the global by, e.g., re-assigning a new value to it if we use the global keyword as the following example will illustrate:" + ] + }, + { + "cell_type": "code", + "execution_count": 3, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "global value [ a_var outside a_func() ]\n", + "local value [ a_var inside a_func() ]\n", + "local value [ a_var outside a_func() ]\n" + ] + } + ], + "source": [ + "a_var = 'global value'\n", + "\n", + "def a_func():\n", + " global a_var\n", + " a_var = 'local value'\n", + " print(a_var, '[ a_var inside a_func() ]')\n", + "\n", + "print(a_var, '[ a_var outside a_func() ]')\n", + "a_func()\n", + "print(a_var, '[ a_var outside a_func() ]')" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "But we have to be careful about the order: it is easy to raise an `UnboundLocalError` if we don't explicitly tell Python that we want to use the global scope and try to modify a variable's value (remember, the right side of an assignment operation is executed first):" + ] + }, + { + "cell_type": "code", + "execution_count": 4, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "ename": "UnboundLocalError", + "evalue": "local variable 'a_var' referenced before assignment", + "output_type": "error", + "traceback": [ + "\u001b[0;31m---------------------------------------------------------------------------\u001b[0m\n\u001b[0;31mUnboundLocalError\u001b[0m Traceback (most recent call last)", + "\u001b[0;32m
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## 2. LEG - Local, Enclosed, and Global scope\n", + "\n", + "\n", + "\n", + "Now, let us introduce the concept of the enclosed (E) scope. Following the order \"Local -> Enclosed -> Global\", can you guess what the following code will print?" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "**Example 2.1**" + ] + }, + { + "cell_type": "code", + "execution_count": 4, + "metadata": { + "collapsed": false + }, + "outputs": [], + "source": [ + "a_var = 'global value'\n", + "\n", + "def outer():\n", + " a_var = 'enclosed value'\n", + " \n", + " def inner():\n", + " a_var = 'local value'\n", + " print(a_var)\n", + " \n", + " inner()\n", + "\n", + "outer()" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "**a)**\n", + "
global value\n", + "\n", + "**b)** \n", + "
enclosed value\n", + "\n", + "**c)** \n", + "
local value" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[go to solution](#solutions)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### Here is why:\n", + "\n", + "Let us quickly recapitulate what we just did: We called `outer()`, which defined the variable `a_var` locally (next to an existing `a_var` in the global scope). Next, the `outer()` function called `inner()`, which in turn defined a variable with of name `a_var` as well. The `print()` function inside `inner()` searched in the local scope first (L->E) before it went up in the scope hierarchy, and therefore it printed the value that was assigned in the local scope." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Similar to the concept of the `global` keyword, which we have seen in the section above, we can use the keyword `nonlocal` inside the inner function to explicitly access a variable from the outer (enclosed) scope in order to modify its value. \n", + "Note that the `nonlocal` keyword was added in Python 3.x and is not implemented in Python 2.x (yet)." + ] + }, + { + "cell_type": "code", + "execution_count": 5, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "outer before: local value\n", + "in inner(): inner value\n", + "outer after: inner value\n" + ] + } + ], + "source": [ + "a_var = 'global value'\n", + "\n", + "def outer():\n", + " a_var = 'local value'\n", + " print('outer before:', a_var)\n", + " def inner():\n", + " nonlocal a_var\n", + " a_var = 'inner value'\n", + " print('in inner():', a_var)\n", + " inner()\n", + " print(\"outer after:\", a_var)\n", + "outer()" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## 3. LEGB - Local, Enclosed, Global, Built-in\n", + "\n", + "To wrap up the LEGB rule, let us come to the built-in scope. Here, we will define our \"own\" length-funcion, which happens to bear the same name as the in-built `len()` function. What outcome do you excpect if we'd execute the following code?" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "**Example 3**" + ] + }, + { + "cell_type": "code", + "execution_count": 6, + "metadata": { + "collapsed": false + }, + "outputs": [], + "source": [ + "a_var = 'global variable'\n", + "\n", + "def len(in_var):\n", + " print('called my len() function')\n", + " l = 0\n", + " for i in in_var:\n", + " l += 1\n", + " return l\n", + "\n", + "def a_func(in_var):\n", + " len_in_var = len(in_var)\n", + " print('Input variable is of length', len_in_var)\n", + "\n", + "a_func('Hello, World!')" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "**a)**\n", + "
raises an error (conflict with in-built `len()` function)\n", + "\n", + "**b)** \n", + "
called my len() function\n", + "Input variable is of length 13\n", + "\n", + "**c)** \n", + "
Input variable is of length 13" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[go to solution](#solutions)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "### Here is why:\n", + "\n", + "Since the exact same names can be used to map names to different objects - as long as the names are in different name spaces - there is no problem of reusing the name `len` to define our own length function (this is just for demonstration pruposes, it is NOT recommended). As we go up in Python's L -> E -> G -> B hierarchy, the function `a_func()` finds `len()` already in the global scope (G) first before it attempts to search the built-in (B) namespace." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "# Self-assessment exercise" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Now, after we went through a couple of exercises, let us quickly check where we are. So, one more time: What would the following code print out?" + ] + }, + { + "cell_type": "code", + "execution_count": 59, + "metadata": { + "collapsed": false + }, + "outputs": [], + "source": [ + "a = 'global'\n", + "\n", + "def outer():\n", + " \n", + " def len(in_var):\n", + " print('called my len() function: ', end=\"\")\n", + " l = 0\n", + " for i in in_var:\n", + " l += 1\n", + " return l\n", + " \n", + " a = 'local'\n", + " \n", + " def inner():\n", + " global len\n", + " nonlocal a\n", + " a += ' variable'\n", + " inner()\n", + " print('a is', a)\n", + " print(len(a))\n", + "\n", + "\n", + "outer()\n", + "\n", + "print(len(a))\n", + "print('a is', a)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "[[go to solution](#solutions)]" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "# Conclusion" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "I hope this short tutorial was helpful to understand the basic concept of Python's scope resolution order using the LEGB rule. I want to encourage you (as a little self-assessment exercise) to look at the code snippets again tomorrow and check if you can correctly predict all their outcomes." + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "#### A rule of thumb" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "In practice, **it is usually a bad idea to modify global variables inside the function scope**, since it often be the cause of confusion and weird errors that are hard to debug. \n", + "If you want to modify a global variable via a function, it is recommended to pass it as an argument and reassign the return-value. \n", + "For example:" + ] + }, + { + "cell_type": "code", + "execution_count": 42, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "8\n" + ] + } + ], + "source": [ + "a_var = 2\n", + "\n", + "def a_func(some_var):\n", + " return 2**3\n", + "\n", + "a_var = a_func(a_var)\n", + "print(a_var)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "\n", + "
\n", + "
" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Solutions\n", + "\n", + "In order to prevent you from unintentional spoilers, I have written the solutions in binary format. In order to display the character representation, you just need to execute the following lines of code:" + ] + }, + { + "cell_type": "code", + "execution_count": 6, + "metadata": { + "collapsed": false + }, + "outputs": [], + "source": [ + "print('Example 1.1:', chr(int('01100011',2)))" + ] + }, + { + "cell_type": "code", + "execution_count": 7, + "metadata": { + "collapsed": false + }, + "outputs": [], + "source": [ + "print('Example 1.2:', chr(int('01100010',2)))" + ] + }, + { + "cell_type": "code", + "execution_count": 8, + "metadata": { + "collapsed": false + }, + "outputs": [], + "source": [ + "print('Example 2.1:', chr(int('01100011',2)))" + ] + }, + { + "cell_type": "code", + "execution_count": 9, + "metadata": { + "collapsed": false + }, + "outputs": [], + "source": [ + "print('Example 3.1:', chr(int('01100010',2)))" + ] + }, + { + "cell_type": "code", + "execution_count": 58, + "metadata": { + "collapsed": false + }, + "outputs": [], + "source": [ + "# Execute to run the self-assessment solution\n", + "\n", + "sol = \"000010100110111101110101011101000110010101110010001010\"\\\n", + "\"0000101001001110100000101000001010011000010010000001101001011100110\"\\\n", + "\"0100000011011000110111101100011011000010110110000100000011101100110\"\\\n", + "\"0001011100100110100101100001011000100110110001100101000010100110001\"\\\n", + "\"1011000010110110001101100011001010110010000100000011011010111100100\"\\\n", + "\"1000000110110001100101011011100010100000101001001000000110011001110\"\\\n", + "\"1010110111001100011011101000110100101101111011011100011101000100000\"\\\n", + "\"0011000100110100000010100000101001100111011011000110111101100010011\"\\\n", + "\"0000101101100001110100000101000001010001101100000101001100001001000\"\\\n", + "\"0001101001011100110010000001100111011011000110111101100010011000010\"\\\n", + "\"1101100\"\n", + "\n", + "sol_str =''.join(chr(int(sol[i:i+8], 2)) for i in range(0, len(sol), 8))\n", + "for line in sol_str.split('\\n'):\n", + " print(line)" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "
\n", + "
\n", + "" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "## Warning: For-loop variables \"leaking\" into the global namespace" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "In contrast to some other programming languages, `for-loops` will use the scope they exist in and leave their defined loop-variable behind.\n" + ] + }, + { + "cell_type": "code", + "execution_count": 5, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "4 -> a in for-loop\n", + "4 -> a in global\n" + ] + } + ], + "source": [ + "for a in range(5):\n", + " if a == 4:\n", + " print(a, '-> a in for-loop')\n", + "print(a, '-> a in global')" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "**This also applies if we explicitly defined the `for-loop` variable in the global namespace before!** In this case it will rebind the existing variable:" + ] + }, + { + "cell_type": "code", + "execution_count": 9, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "4 -> b in for-loop\n", + "4 -> b in global\n" + ] + } + ], + "source": [ + "b = 1\n", + "for b in range(5):\n", + " if b == 4:\n", + " print(b, '-> b in for-loop')\n", + "print(b, '-> b in global')" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "However, in **Python 3.x**, we can use closures to prevent the for-loop variable to cut into the global namespace. Here is an example (exectuted in Python 3.4):" + ] + }, + { + "cell_type": "code", + "execution_count": 1, + "metadata": { + "collapsed": false + }, + "outputs": [ + { + "name": "stdout", + "output_type": "stream", + "text": [ + "[0, 1, 2, 3, 4]\n", + "1 -> i in global\n" + ] + } + ], + "source": [ + "i = 1\n", + "print([i for i in range(5)])\n", + "print(i, '-> i in global')" + ] + }, + { + "cell_type": "markdown", + "metadata": {}, + "source": [ + "Why did I mention \"Python 3.x\"? Well, as it happens, the same code executed in Python 2.x would print:\n", + "\n", + "
\n",
+ "4 -> i in global\n",
+ ""
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "This goes back to a change that was made in Python 3.x and is described in [What’s New In Python 3.0](https://docs.python.org/3/whatsnew/3.0.html) as follows:\n",
+ "\n",
+ "\"List comprehensions no longer support the syntactic form `[... for var in item1, item2, ...]`. Use `[... for var in (item1, item2, ...)]` instead. Also note that list comprehensions have different semantics: they are closer to syntactic sugar for a generator expression inside a `list()` constructor, and in particular the loop control variables are no longer leaked into the surrounding scope.\""
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [],
+ "source": []
+ }
+ ],
+ "metadata": {
+ "kernelspec": {
+ "display_name": "Python 3",
+ "language": "python",
+ "name": "python3"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.5.0"
+ }
+ },
+ "nbformat": 4,
+ "nbformat_minor": 0
+}
diff --git a/tutorials/sorting_csvs.ipynb b/tutorials/sorting_csvs.ipynb
new file mode 100644
index 0000000..df1b182
--- /dev/null
+++ b/tutorials/sorting_csvs.ipynb
@@ -0,0 +1,757 @@
+{
+ "metadata": {
+ "name": "",
+ "signature": "sha256:f56b7081a6e5b63610100fcfa0a226c7a0184dfe0d63128614a7a68555653428"
+ },
+ "nbformat": 3,
+ "nbformat_minor": 0,
+ "worksheets": [
+ {
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[Sebastian Raschka](http://sebastianraschka.com) \n",
+ "last updated: 05/13/2014\n",
+ "\n",
+ "- Open in [IPython nbviewer](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/tutorials/sorting_csvs.ipynb?create=1) \n",
+ "- Link to this [IPython notebook on Github](https://github.com/rasbt/python_reference/blob/master/tutorials/sorting_csvs.ipynb) \n",
+ "- Link to the GitHub Repository [`python_reference`](https://github.com/rasbt/python_reference)\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "I am looking forward to comments or suggestions, please don't hesitate to contact me via\n",
+ "[twitter](https://twitter.com/rasbt), [email](mailto:bluewoodtree@gmail.com), or [google+](https://plus.google.com/118404394130788869227).\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Sorting CSV files using the Python `csv` module"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "I wanted to summarize a way to sort CSV files by just using the [`csv` module](https://docs.python.org/3.4/library/csv.html) and other standard library Python modules \n",
+ "(you probably also want to consider using the [pandas](http://pandas.pydata.org) library if you are working with very large CSV files - I am planning to make this a separate topic)."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
\n",
+ "
\n",
+ "## Sections\n",
+ "- [Reading in a CSV file](#reading)\n",
+ "- [Printing the CSV file contents](#printing)\n",
+ "- [Converting numeric cells to floats](#floats)\n",
+ "- [Sorting the CSV file](#sorting)\n",
+ "- [Marking min/max values in particular columns](#marking)\n",
+ "- [Writing out the modified table to as a new CSV file](#writing)\n",
+ "- [Batch processing CSV files](#batch)\n",
+ "
\n",
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Objective:\n",
+ "\n",
+ "Let us assume that we have an [example CSV](../Data/test.csv) file formatted like this:\n",
+ " \n",
+ "name,column1,column2,column3\n",
+ "abc,1.1,4.2,1.2\n",
+ "def,2.1,1.4,5.2\n",
+ "ghi,1.5,1.2,2.1\n",
+ "jkl,1.8,1.1,4.2\n",
+ "mno,9.4,6.6,6.2\n",
+ "pqr,1.4,8.3,8.4
\n",
+ "\n",
+ "And we want to sort particular columns and eventually mark min- of max-values in the table.\n",
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ ""
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ ""
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "##Reading in a CSV file"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[[back to top](#sections)]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Because we will be iterating over our CSV file a couple of times, let us read in the CSV file using the `csv` module and hold the contents in memory using a Python list object (note: be careful with very large CSV files and possible memory issues associated with this approach).\n"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "import csv\n",
+ "\n",
+ "def csv_to_list(csv_file, delimiter=','):\n",
+ " \"\"\" \n",
+ " Reads in a CSV file and returns the contents as list,\n",
+ " where every row is stored as a sublist, and each element\n",
+ " in the sublist represents 1 cell in the table.\n",
+ " \n",
+ " \"\"\"\n",
+ " with open(csv_file, 'r') as csv_con:\n",
+ " reader = csv.reader(csv_con, delimiter=delimiter)\n",
+ " return list(reader)"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [],
+ "prompt_number": 1
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "csv_cont = csv_to_list('../Data/test.csv')\n",
+ "\n",
+ "print('first 3 rows:')\n",
+ "for row in range(3):\n",
+ " print(csv_cont[row])"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [
+ {
+ "output_type": "stream",
+ "stream": "stdout",
+ "text": [
+ "first 3 rows:\n",
+ "['name', 'column1', 'column2', 'column3']\n",
+ "['abc', '1.1', '4.2', '1.2']\n",
+ "['def', '2.1', '1.4', '5.2']\n"
+ ]
+ }
+ ],
+ "prompt_number": 2
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "\n",
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "##Printing the CSV file contents"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[[back to top](#sections)]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Also, let us define a short function that prints out the CSV file to the standard output screen in a slightly prettier format:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "def print_csv(csv_content):\n",
+ " \"\"\" Prints CSV file to standard output.\"\"\"\n",
+ " print(50*'-')\n",
+ " for row in csv_content:\n",
+ " row = [str(e) for e in row]\n",
+ " print('\\t'.join(row))\n",
+ " print(50*'-')"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [],
+ "prompt_number": 3
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "csv_cont = csv_to_list('../Data/test.csv')\n",
+ "\n",
+ "print('\\n\\nOriginal CSV file:')\n",
+ "print_csv(csv_cont)"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [
+ {
+ "output_type": "stream",
+ "stream": "stdout",
+ "text": [
+ "\n",
+ "\n",
+ "Original CSV file:\n",
+ "--------------------------------------------------\n",
+ "name\tcolumn1\tcolumn2\tcolumn3\n",
+ "abc\t1.1\t4.2\t1.2\n",
+ "def\t2.1\t1.4\t5.2\n",
+ "ghi\t1.5\t1.2\t-2.1\n",
+ "jkl\t1.8\t-1.1\t4.2\n",
+ "mno\t9.4\t6.6\t6.2\n",
+ "pqr\t1.4\t8.3\t8.4\n",
+ "--------------------------------------------------\n"
+ ]
+ }
+ ],
+ "prompt_number": 4
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "\n",
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Converting numeric cells to floats"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "To avoid problems with the sorting approach that can occur when we have negative values in some cells, let us define a function that converts all numeric cells into float values."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "def convert_cells_to_floats(csv_cont):\n",
+ " \"\"\" \n",
+ " Converts cells to floats if possible\n",
+ " (modifies input CSV content list).\n",
+ " \n",
+ " \"\"\"\n",
+ " for row in range(len(csv_cont)):\n",
+ " for cell in range(len(csv_cont[row])):\n",
+ " try:\n",
+ " csv_cont[row][cell] = float(csv_cont[row][cell])\n",
+ " except ValueError:\n",
+ " pass "
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [],
+ "prompt_number": 5
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "print('first 3 rows:')\n",
+ "for row in range(3):\n",
+ " print(csv_cont[row])"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [
+ {
+ "output_type": "stream",
+ "stream": "stdout",
+ "text": [
+ "first 3 rows:\n",
+ "['name', 'column1', 'column2', 'column3']\n",
+ "['abc', '1.1', '4.2', '1.2']\n",
+ "['def', '2.1', '1.4', '5.2']\n"
+ ]
+ }
+ ],
+ "prompt_number": 6
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "\n",
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "##Sorting the CSV file"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[[back to top](#sections)]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Using the very handy [`operator.itemgetter`](https://docs.python.org/3.4/library/operator.html#operator.itemgetter) function, we define a function that returns a CSV file contents sorted by a particular column (column index or column name)."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "import operator\n",
+ "\n",
+ "def sort_by_column(csv_cont, col, reverse=False):\n",
+ " \"\"\" \n",
+ " Sorts CSV contents by column name (if col argument is type ) \n",
+ " or column index (if col argument is type ). \n",
+ " \n",
+ " \"\"\"\n",
+ " header = csv_cont[0]\n",
+ " body = csv_cont[1:]\n",
+ " if isinstance(col, str): \n",
+ " col_index = header.index(col)\n",
+ " else:\n",
+ " col_index = col\n",
+ " body = sorted(body, \n",
+ " key=operator.itemgetter(col_index), \n",
+ " reverse=reverse)\n",
+ " body.insert(0, header)\n",
+ " return body"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [],
+ "prompt_number": 7
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "To see how (and if) it works, let us sort the CSV file in [../Data/test.csv](../Data/test.csv) by the column name \"column3\"."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "csv_cont = csv_to_list('../Data/test.csv')\n",
+ "\n",
+ "print('\\n\\nOriginal CSV file:')\n",
+ "print_csv(csv_cont)\n",
+ "\n",
+ "print('\\n\\nCSV sorted by column \"column3\":')\n",
+ "convert_cells_to_floats(csv_cont)\n",
+ "csv_sorted = sort_by_column(csv_cont, 'column3')\n",
+ "print_csv(csv_sorted)"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [
+ {
+ "output_type": "stream",
+ "stream": "stdout",
+ "text": [
+ "\n",
+ "\n",
+ "Original CSV file:\n",
+ "--------------------------------------------------\n",
+ "name\tcolumn1\tcolumn2\tcolumn3\n",
+ "abc\t1.1\t4.2\t1.2\n",
+ "def\t2.1\t1.4\t5.2\n",
+ "ghi\t1.5\t1.2\t-2.1\n",
+ "jkl\t1.8\t-1.1\t4.2\n",
+ "mno\t9.4\t6.6\t6.2\n",
+ "pqr\t1.4\t8.3\t8.4\n",
+ "--------------------------------------------------\n",
+ "\n",
+ "\n",
+ "CSV sorted by column \"column3\":\n",
+ "--------------------------------------------------\n",
+ "name\tcolumn1\tcolumn2\tcolumn3\n",
+ "ghi\t1.5\t1.2\t-2.1\n",
+ "abc\t1.1\t4.2\t1.2\n",
+ "jkl\t1.8\t-1.1\t4.2\n",
+ "def\t2.1\t1.4\t5.2\n",
+ "mno\t9.4\t6.6\t6.2\n",
+ "pqr\t1.4\t8.3\t8.4\n",
+ "--------------------------------------------------\n"
+ ]
+ }
+ ],
+ "prompt_number": 8
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "\n",
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Marking min/max values in particular columns"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[[back to top](#sections)]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "To visualize minimum and maximum values in certain columns if find it quite useful to add little symbols to the cells (most people like to highlight cells with colors in e.g., Excel spreadsheets, but CSV doesn't support colors, so this is my workaround - please let me know if you figured out a better approach, I would be looking forward to your suggestion)."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "def mark_minmax(csv_cont, col, mark_max=True, marker='*'):\n",
+ " \"\"\"\n",
+ " Sorts a list of CSV contents by a particular column \n",
+ " (see sort_by_column function).\n",
+ " Puts a marker on the maximum value if mark_max=True,\n",
+ " or puts a marker on the minimum value mark_max=False\n",
+ " (modifies input CSV content list).\n",
+ " \n",
+ " \"\"\"\n",
+ " \n",
+ " sorted_csv = sort_by_column(csv_cont, col, reverse=mark_max)\n",
+ " if isinstance(col, str): \n",
+ " col_index = sorted_csv[0].index(col)\n",
+ " else:\n",
+ " col_index = col\n",
+ " sorted_csv[1][col_index] = str(sorted_csv[1][col_index]) + marker\n",
+ " return None"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [],
+ "prompt_number": 9
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "def mark_all_col(csv_cont, mark_max=True, marker='*'):\n",
+ " \"\"\"\n",
+ " Marks all maximum (if mark_max=True) or minimum (if mark_max=False)\n",
+ " values in all columns of a CSV contents list - except the first column.\n",
+ " Returns a new list that is sorted by the names in the first column\n",
+ " (modifies input CSV content list).\n",
+ " \n",
+ " \"\"\"\n",
+ " for c in range(1, len(csv_cont[0])):\n",
+ " mark_minmax(csv_cont, c, mark_max, marker)\n",
+ " marked_csv = sort_by_column(csv_cont, 0, False)\n",
+ " return marked_csv"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [],
+ "prompt_number": 10
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "import copy\n",
+ "\n",
+ "csv_cont = csv_to_list('../Data/test.csv')\n",
+ "\n",
+ "csv_marked = copy.deepcopy(csv_cont)\n",
+ "convert_cells_to_floats(csv_marked)\n",
+ "mark_all_col(csv_marked, mark_max=False, marker='*')\n",
+ "print_csv(csv_marked)\n",
+ "print('*: min-value')"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [
+ {
+ "output_type": "stream",
+ "stream": "stdout",
+ "text": [
+ "--------------------------------------------------\n",
+ "name\tcolumn1\tcolumn2\tcolumn3\n",
+ "abc\t1.1*\t4.2\t1.2\n",
+ "def\t2.1\t1.4\t5.2\n",
+ "ghi\t1.5\t1.2\t-2.1*\n",
+ "jkl\t1.8\t-1.1*\t4.2\n",
+ "mno\t9.4\t6.6\t6.2\n",
+ "pqr\t1.4\t8.3\t8.4\n",
+ "--------------------------------------------------\n",
+ "*: min-value\n"
+ ]
+ }
+ ],
+ "prompt_number": 12
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "\n",
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Writing out the modified table to as a new CSV file"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[[back to top](#sections)]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "After the sorting and maybe marking of minimum and maximum values, we likely want to write out the modified data table as CSV file again."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "def write_csv(dest, csv_cont):\n",
+ " \"\"\" Writes a comma-delimited CSV file. \"\"\"\n",
+ "\n",
+ " with open(dest, 'w') as out_file:\n",
+ " writer = csv.writer(out_file, delimiter=',')\n",
+ " for row in csv_cont:\n",
+ " writer.writerow(row)\n",
+ "\n",
+ "write_csv('../Data/test_marked.csv', csv_marked)"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [],
+ "prompt_number": 13
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Let us read in the written CSV file to confirm that the formatting is correct:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "csv_cont = csv_to_list('../Data/test_marked.csv')\n",
+ "\n",
+ "print('\\n\\nWritten CSV file:')\n",
+ "print_csv(csv_cont)"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [
+ {
+ "output_type": "stream",
+ "stream": "stdout",
+ "text": [
+ "\n",
+ "\n",
+ "Written CSV file:\n",
+ "--------------------------------------------------\n",
+ "name\tcolumn1\tcolumn2\tcolumn3\n",
+ "abc\t1.1*\t4.2\t1.2\n",
+ "def\t2.1\t1.4\t5.2\n",
+ "ghi\t1.5\t1.2\t-2.1*\n",
+ "jkl\t1.8\t-1.1*\t4.2\n",
+ "mno\t9.4\t6.6\t6.2\n",
+ "pqr\t1.4\t8.3\t8.4\n",
+ "--------------------------------------------------\n"
+ ]
+ }
+ ],
+ "prompt_number": 14
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "\n",
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## Batch processing CSV files"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[[back to top](#sections)]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Usually, CSV files never come alone, but we have to process a whole bunch of similar formatted CSV files from some output device. \n",
+ "For example, if we want to process all CSV files in a particular input directory and want to save the processed files in a separate output directory, we can use a simple list comprehension to collect tuples of input-output file names."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "import os\n",
+ "\n",
+ "in_dir = '../Data'\n",
+ "out_dir = '../Data/processed'\n",
+ "csvs = [\n",
+ " (os.path.join(in_dir, csv), \n",
+ " os.path.join(out_dir, csv))\n",
+ " for csv in os.listdir(in_dir) \n",
+ " if csv.endswith('.csv')\n",
+ " ]\n",
+ "\n",
+ "for i in csvs:\n",
+ " print(i)"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [
+ {
+ "output_type": "stream",
+ "stream": "stdout",
+ "text": [
+ "('../Data/test.csv', '../Data/processed/test.csv')\n",
+ "('../Data/test_marked.csv', '../Data/processed/test_marked.csv')\n"
+ ]
+ }
+ ],
+ "prompt_number": 12
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "Next, we can summarize the processes we want to apply to the CSV files in a simple function and loop over our file names:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "def process_csv(csv_in, csv_out):\n",
+ " \"\"\" \n",
+ " Takes an input- and output-filename of an CSV file\n",
+ " and marks minimum values for every column.\n",
+ " \n",
+ " \"\"\"\n",
+ " csv_cont = csv_to_list(csv_in)\n",
+ " csv_marked = copy.deepcopy(csv_cont)\n",
+ " convert_cells_to_floats(csv_marked)\n",
+ " mark_all_col(csv_marked, mark_max=False, marker='*')\n",
+ " write_csv(csv_out, csv_marked)"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [],
+ "prompt_number": 18
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "for inout in csvs:\n",
+ " process_csv(inout[0], inout[1])"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": []
+ }
+ ],
+ "metadata": {}
+ }
+ ]
+}
\ No newline at end of file
diff --git a/tutorials/sqlite3_howto/LICENSE b/tutorials/sqlite3_howto/LICENSE
new file mode 100644
index 0000000..ef7e7ef
--- /dev/null
+++ b/tutorials/sqlite3_howto/LICENSE
@@ -0,0 +1,674 @@
+GNU GENERAL PUBLIC LICENSE
+ Version 3, 29 June 2007
+
+ Copyright (C) 2007 Free Software Foundation, Inc. .
diff --git a/tutorials/sqlite3_howto/README.md b/tutorials/sqlite3_howto/README.md
new file mode 100644
index 0000000..c596dfc
--- /dev/null
+++ b/tutorials/sqlite3_howto/README.md
@@ -0,0 +1,780 @@
+## A thorough guide to SQLite database operations in Python
+
+_\-- written by Sebastian Raschka_ on March 7, 2014
+
+
+
+
+
+ +
+
+
+
+* * *
+
+#### Sections
+
+• Connecting to an SQLite database
+• Creating a new SQLite database
+ - Overview of SQLite data types
+ - A quick word on PRIMARY KEYS:
+• Adding new columns
+• Inserting and updating rows
+• Creating unique indexes
+• Querying the database - Selecting rows
+• Security and injection attacks
+• Date and time operations
+• Printing a database summary
+• Conclusion
+
+The complete Python code that I am using in this tutorial can be downloaded
+from my GitHub repository: [https://github.com/rasbt/python_reference/tree/master/tutorials/sqlite3_howto](https://github.com/rasbt/python_reference/tree/master/tutorials/sqlite3_howto)
+
+
+* * *
+
+
+
+## Connecting to an SQLite database
+
+The sqlite3 that we will be using throughout this tutorial is part of the
+Python Standard Library and is a nice and easy interface to SQLite databases:
+There are no server processes involved, no configurations required, and no
+other obstacles we have to worry about.
+
+In general, the only thing that needs to be done before we can perform any
+operation on a SQLite database via Python's `sqlite3` module, is to open a
+connection to an SQLite database file:
+
+
+
+ import sqlite3
+ conn = sqlite3.connect(sqlite_file)
+ c = conn.cursor()
+
+
+where the database file (`sqlite_file`) can reside anywhere on our disk, e.g.,
+
+
+
+ sqlite_file = '/Users/Sebastian/Desktop/my_db.sqlite'
+
+
+Conveniently, a new database file (`.sqlite` file) will be created
+automatically the first time we try to connect to a database. However, we have
+to be aware that it won't have a table, yet. In the following section, we will
+take a look at some example code of how to create a new SQLite database files
+with tables for storing some data.
+
+To round up this section about connecting to a SQLite database file, there are
+two more operations that are worth mentioning. If we are finished with our
+operations on the database file, we have to close the connection via the
+`.close()` method:
+
+
+
+ conn.close()
+
+
+And if we performed any operation on the database other than sending queries,
+we need to commit those changes via the `.commit()` method before we close the
+connection:
+
+
+
+ conn.commit()
+ conn.close()
+
+
+
+
+## Creating a new SQLite database
+
+Let us have a look at some example code to create a new SQLite database file
+with two tables: One with and one without a PRIMARY KEY column (don't worry,
+there is more information about PRIMARY KEYs further down in this section).
+
+
+
+ import sqlite3
+
+ sqlite_file = 'my_first_db.sqlite' # name of the sqlite database file
+ table_name1 = 'my_table_1' # name of the table to be created
+ table_name2 = 'my_table_2' # name of the table to be created
+ new_field = 'my_1st_column' # name of the column
+ field_type = 'INTEGER' # column data type
+
+ # Connecting to the database file
+ conn = sqlite3.connect(sqlite_file)
+ c = conn.cursor()
+
+ # Creating a new SQLite table with 1 column
+ c.execute('CREATE TABLE {tn} ({nf} {ft})'\
+ .format(tn=table_name1, nf=new_field, ft=field_type))
+
+ # Creating a second table with 1 column and set it as PRIMARY KEY
+ # note that PRIMARY KEY column must consist of unique values!
+ c.execute('CREATE TABLE {tn} ({nf} {ft} PRIMARY KEY)'\
+ .format(tn=table_name2, nf=new_field, ft=field_type))
+
+ # Committing changes and closing the connection to the database file
+ conn.commit()
+ conn.close()
+
+
+Download the script: [create_new_db.py](https://github.com/rasbt/python_reference/blob/master/tutorials/sqlite3_howto/code/create_new_db.py)
+
+* * *
+
+**Tip:** A handy tool to visualize and access SQLite databases is the free FireFox [SQLite Manager](https://addons.mozilla.org/en-US/firefox/addon/sqlite-manager/?src) add-on. Throughout this article, I will use this tool to provide screenshots of the database structures that we created below the corresponding code sections.
+
+* * *
+
+
+
+
+
+
+Using the code above, we created a new `.sqlite` database file with 2 tables.
+Each table consists of currently one column only, which is of type INTEGER.
+
+
+
+* * *
+
+**Here is a quick overview of all data types that are supported by SQLite 3:**
+
+ * INTEGER: A signed integer up to 8 bytes depending on the magnitude of the value.
+ * REAL: An 8-byte floating point value.
+ * TEXT: A text string, typically UTF-8 encoded (depending on the database encoding).
+ * BLOB: A blob of data (binary large object) for storing binary data.
+ * NULL: A NULL value, represents missing data or an empty cell.
+
+* * *
+
+Looking at the table above, You might have noticed that SQLite 3 has no
+designated Boolean data type. However, this should not be an issue, since we
+could simply re-purpose the INTEGER type to represent Boolean values (0 =
+false, 1 = true).
+
+
+
+**A quick word on PRIMARY KEYS:**
+In our example code above, we set our 1 column in the second table to PRIMARY
+KEY. The advantage of a PRIMARY KEY index is a significant performance gain if
+we use the PRIMARY KEY column as query for accessing rows in the table. Every
+table can only have max. 1 PRIMARY KEY (single or multiple column(s)), and the
+values in this column MUST be unique! But more on column indexing in the a
+later section.
+
+
+
+## Adding new columns
+
+If we want to add a new column to an existing SQLite database table, we can
+either leave the cells for each row empty (NULL value), or we can set a
+default value for each cell, which is pretty convenient for certain
+applications.
+Let's have a look at some code:
+
+
+
+ import sqlite3
+
+ sqlite_file = 'my_first_db.sqlite' # name of the sqlite database file
+ table_name = 'my_table_2' # name of the table to be created
+ id_column = 'my_1st_column' # name of the PRIMARY KEY column
+ new_column1 = 'my_2nd_column' # name of the new column
+ new_column2 = 'my_3nd_column' # name of the new column
+ column_type = 'TEXT' # E.g., INTEGER, TEXT, NULL, REAL, BLOB
+ default_val = 'Hello World' # a default value for the new column rows
+
+ # Connecting to the database file
+ conn = sqlite3.connect(sqlite_file)
+ c = conn.cursor()
+
+ # A) Adding a new column without a row value
+ c.execute("ALTER TABLE {tn} ADD COLUMN '{cn}' {ct}"\
+ .format(tn=table_name, cn=new_column1, ct=column_type))
+
+ # B) Adding a new column with a default row value
+ c.execute("ALTER TABLE {tn} ADD COLUMN '{cn}' {ct} DEFAULT '{df}'"\
+ .format(tn=table_name, cn=new_column2, ct=column_type, df=default_val))
+
+ # Committing changes and closing the connection to the database file
+ conn.commit()
+ conn.close()
+
+
+Download the script: [add_new_column.py](https://github.com/rasbt/python_reference/blob/master/tutorials/sqlite3_howto/code/add_new_column.py)
+
+
+
+
+
+
+We just added 2 more columns (`my_2nd_column` and `my_3rd_column`) to
+`my_table_2` of our SQLite database next to the PRIMARY KEY column
+`my_1st_column`.
+The difference between the two new columns is that we initialized
+`my_3rd_column` with a default value (here:'Hello World'), which will be
+inserted for every existing cell under this column and for every new row that
+we are going to add to the table if we don't insert or update it with a
+different value.
+
+
+
+## Inserting and updating rows
+
+Inserting and updating rows into an existing SQLite database table - next to
+sending queries - is probably the most common database operation. The
+Structured Query Language has a convenient `UPSERT` function, which is
+basically just a merge between UPDATE and INSERT: It inserts new rows into a
+database table with a value for the PRIMARY KEY column if it does not exist
+yet, or updates a row for an existing PRIMARY KEY value. Unfortunately, this
+convenient syntax is not supported by the more compact SQLite database
+implementation that we are using here. However, there are some workarounds.
+But let us first have a look at the example code:
+
+
+
+ import sqlite3
+
+ sqlite_file = 'my_first_db.sqlite'
+ table_name = 'my_table_2'
+ id_column = 'my_1st_column'
+ column_name = 'my_2nd_column'
+
+ # Connecting to the database file
+ conn = sqlite3.connect(sqlite_file)
+ c = conn.cursor()
+
+ # A) Inserts an ID with a specific value in a second column
+ try:
+ c.execute("INSERT INTO {tn} ({idf}, {cn}) VALUES (123456, 'test')".\
+ format(tn=table_name, idf=id_column, cn=column_name))
+ except sqlite3.IntegrityError:
+ print('ERROR: ID already exists in PRIMARY KEY column {}'.format(id_column))
+
+ # B) Tries to insert an ID (if it does not exist yet)
+ # with a specific value in a second column
+ c.execute("INSERT OR IGNORE INTO {tn} ({idf}, {cn}) VALUES (123456, 'test')".\
+ format(tn=table_name, idf=id_column, cn=column_name))
+
+ # C) Updates the newly inserted or pre-existing entry
+ c.execute("UPDATE {tn} SET {cn}=('Hi World') WHERE {idf}=(123456)".\
+ format(tn=table_name, cn=column_name, idf=id_column))
+
+ conn.commit()
+ conn.close()
+
+
+Download the script: [update_or_insert_records.py](code/update_or_insert_records.py)
+
+
+
+Both A) `INSERT` and B) `INSERT OR IGNORE` have in common that they append new
+rows to the database if a given PRIMARY KEY does not exist in the database
+table, yet. However, if we'd try to append a PRIMARY KEY value that is not
+unique, a simple `INSERT` would raise an `sqlite3.IntegrityError` exception,
+which can be either captured via a try-except statement (case A) or
+circumvented by the SQLite call `INSERT OR IGNORE` (case B). This can be
+pretty useful if we want to construct an `UPSERT` equivalent in SQLite. E.g.,
+if we want to add a dataset to an existing database table that contains a mix
+between existing and new IDs for our PRIMARY KEY column.
+
+
+
+## Creating unique indexes
+
+Just like hashtable-datastructures, indexes function as direct pointers to our
+data in a table for a particular column (i.e., the indexed column). For
+example, the PRIMARY KEY column would have such an index by default. The
+downside of indexes is that every row value in the column must be unique.
+However, it is recommended and pretty useful to index certain columns if
+possible, since it rewards us with a significant performance gain for the data
+retrieval.
+The example code below shows how to add such an unique index to an existing
+column in an SQLite database table. And if we should decide to insert non-
+unique values into a indexed column later, there is also a convenient way to
+drop the index, which is also shown in the code below.
+
+
+
+ import sqlite3
+
+ sqlite_file = 'my_first_db.sqlite' # name of the sqlite database file
+ table_name = 'my_table_2' # name of the table to be created
+ id_column = 'my_1st_column' # name of the PRIMARY KEY column
+ new_column = 'unique_names' # name of the new column
+ column_type = 'TEXT' # E.g., INTEGER, TEXT, NULL, REAL, BLOB
+ index_name = 'my_unique_index' # name for the new unique index
+
+ # Connecting to the database file
+ conn = sqlite3.connect(sqlite_file)
+ c = conn.cursor()
+
+ # Adding a new column and update some record
+ c.execute("ALTER TABLE {tn} ADD COLUMN '{cn}' {ct}"\
+ .format(tn=table_name, cn=new_column, ct=column_type))
+ c.execute("UPDATE {tn} SET {cn}='sebastian_r' WHERE {idf}=123456".\
+ format(tn=table_name, idf=id_column, cn=new_column))
+
+ # Creating an unique index
+ c.execute('CREATE INDEX {ix} on {tn}({cn})'\
+ .format(ix=index_name, tn=table_name, cn=new_column))
+
+ # Dropping the unique index
+ # E.g., to avoid future conflicts with update/insert functions
+ c.execute('DROP INDEX {ix}'.format(ix=index_name))
+
+ # Committing changes and closing the connection to the database file
+ conn.commit()
+ conn.close()
+
+
+Download the script: [create_unique_index.py](code/create_unique_index.py)
+
+
+
+
+
+
+## Querying the database - Selecting rows
+
+After we learned about how to create and modify SQLite databases, it's about
+time for some data retrieval. The code below illustrates how we can retrieve
+row entries for all or some columns if they match certain criteria.
+
+
+
+ import sqlite3
+
+ sqlite_file = 'my_first_db.sqlite' # name of the sqlite database file
+ table_name = 'my_table_2' # name of the table to be queried
+ id_column = 'my_1st_column'
+ some_id = 123456
+ column_2 = 'my_2nd_column'
+ column_3 = 'my_3rd_column'
+
+ # Connecting to the database file
+ conn = sqlite3.connect(sqlite_file)
+ c = conn.cursor()
+
+ # 1) Contents of all columns for row that match a certain value in 1 column
+ c.execute('SELECT * FROM {tn} WHERE {cn}="Hi World"'.\
+ format(tn=table_name, cn=column_2))
+ all_rows = c.fetchall()
+ print('1):', all_rows)
+
+ # 2) Value of a particular column for rows that match a certain value in column_1
+ c.execute('SELECT ({coi}) FROM {tn} WHERE {cn}="Hi World"'.\
+ format(coi=column_2, tn=table_name, cn=column_2))
+ all_rows = c.fetchall()
+ print('2):', all_rows)
+
+ # 3) Value of 2 particular columns for rows that match a certain value in 1 column
+ c.execute('SELECT {coi1},{coi2} FROM {tn} WHERE {coi1}="Hi World"'.\
+ format(coi1=column_2, coi2=column_3, tn=table_name, cn=column_2))
+ all_rows = c.fetchall()
+ print('3):', all_rows)
+
+ # 4) Selecting only up to 10 rows that match a certain value in 1 column
+ c.execute('SELECT * FROM {tn} WHERE {cn}="Hi World" LIMIT 10'.\
+ format(tn=table_name, cn=column_2))
+ ten_rows = c.fetchall()
+ print('4):', ten_rows)
+
+ # 5) Check if a certain ID exists and print its column contents
+ c.execute("SELECT * FROM {tn} WHERE {idf}={my_id}".\
+ format(tn=table_name, cn=column_2, idf=id_column, my_id=some_id))
+ id_exists = c.fetchone()
+ if id_exists:
+ print('5): {}'.format(id_exists))
+ else:
+ print('5): {} does not exist'.format(some_id))
+
+ # Closing the connection to the database file
+ conn.close()
+
+
+Download the script: [selecting_entries.py](code/selecting_entries.py)
+
+
+
+
+if we use the `.fetchall()` method, we return a list of tuples from the
+database query, where each tuple represents one row entry. The print output
+for the 5 different cases shown in the code above would look like this (note
+that we only have a table with 1 row here):
+
+
+
+
+
+
+## Security and injection attacks
+
+So far, we have been using Python's string formatting method to insert
+parameters like table and column names into the `c.execute()` functions. This
+is fine if we just want to use the database for ourselves. However, this
+leaves our database vulnerable to injection attacks. For example, if our
+database would be part of a web application, it would allow hackers to
+directly communicate with the database in order to bypass login and password
+verification and steal data.
+In order to prevent this, it is recommended to use `?` place holders in the
+SQLite commands instead of the `%` formatting expression or the `.format()`
+method, which we have been using in this tutorial.
+For example, instead of using
+
+
+
+ # 5) Check if a certain ID exists and print its column contents
+ c.execute("SELECT * FROM {tn} WHERE {idf}={my_id}".\
+ format(tn=table_name, cn=column_2, idf=id_column, my_id=some_id))
+
+
+in the Querying the database - Selecting rows section above, we would want to
+use the `?` placeholder for the queried column value and include the
+variable(s) (here: `123456`), which we want to insert, as tuple at the end of
+the `c.execute()` string.
+
+
+
+ # 5) Check if a certain ID exists and print its column contents
+ c.execute("SELECT * FROM {tn} WHERE {idf}=?".\
+ format(tn=table_name, cn=column_2, idf=id_column), (123456,))
+
+
+However, the problem with this approach is that it would only work for values,
+not for column or table names. So what are we supposed to do with the rest of
+the string if we want to protect ourselves from injection attacks? The easy
+solution would be to refrain from using variables in SQLite queries whenever
+possible, and if it cannot be avoided, we would want to use a function that
+strips all non-alphanumerical characters from the stored content of the
+variable, e.g.,
+
+
+
+ def clean_name(some_var):
+ return ''.join(char for char in some_var if char.isalnum())
+
+
+
+
+## Date and time operations
+
+SQLite inherited the convenient date and time operations from SQL, which are
+one of my favorite features of the Structured Query Language: It does not only
+allow us to insert dates and times in various different formats, but we can
+also perform simple `+` and `-` arithmetic, for example to look up entries
+that have been added xxx days ago.
+
+
+
+ import sqlite3
+
+ sqlite_file = 'my_first_db.sqlite' # name of the sqlite database file
+ table_name = 'my_table_3' # name of the table to be created
+ id_field = 'id' # name of the ID column
+ date_col = 'date' # name of the date column
+ time_col = 'time'# name of the time column
+ date_time_col = 'date_time' # name of the date & time column
+ field_type = 'TEXT' # column data type
+
+ # Connecting to the database file
+ conn = sqlite3.connect(sqlite_file)
+ c = conn.cursor()
+
+ # Creating a new SQLite table with 1 column
+ c.execute('CREATE TABLE {tn} ({fn} {ft} PRIMARY KEY)'\
+ .format(tn=table_name, fn=id_field, ft=field_type))
+
+ # A) Adding a new column to save date insert a row with the current date
+ # in the following format: YYYY-MM-DD
+ # e.g., 2014-03-06
+ c.execute("ALTER TABLE {tn} ADD COLUMN '{cn}'"\
+ .format(tn=table_name, cn=date_col))
+ # insert a new row with the current date and time, e.g., 2014-03-06
+ c.execute("INSERT INTO {tn} ({idf}, {cn}) VALUES('some_id1', DATE('now'))"\
+ .format(tn=table_name, idf=id_field, cn=date_col))
+
+ # B) Adding a new column to save date and time and update with the current time
+ # in the following format: HH:MM:SS
+ # e.g., 16:26:37
+ c.execute("ALTER TABLE {tn} ADD COLUMN '{cn}'"\
+ .format(tn=table_name, cn=time_col))
+ # update row for the new current date and time column, e.g., 2014-03-06 16:26:37
+ c.execute("UPDATE {tn} SET {cn}=TIME('now') WHERE {idf}='some_id1'"\
+ .format(tn=table_name, idf=id_field, cn=time_col))
+
+ # C) Adding a new column to save date and time and update with current date-time
+ # in the following format: YYYY-MM-DD HH:MM:SS
+ # e.g., 2014-03-06 16:26:37
+ c.execute("ALTER TABLE {tn} ADD COLUMN '{cn}'"\
+ .format(tn=table_name, cn=date_time_col))
+ # update row for the new current date and time column, e.g., 2014-03-06 16:26:37
+ c.execute("UPDATE {tn} SET {cn}=(CURRENT_TIMESTAMP) WHERE {idf}='some_id1'"\
+ .format(tn=table_name, idf=id_field, cn=date_time_col))
+
+ # The database should now look like this:
+ # id date time date_time
+ # "some_id1" "2014-03-06" "16:42:30" "2014-03-06 16:42:30"
+
+ # 4) Retrieve all IDs of entries between 2 date_times
+ c.execute("SELECT {idf} FROM {tn} WHERE {cn} BETWEEN '2013-03-06 10:10:10' AND '2015-03-06 10:10:10'".\
+ format(idf=id_field, tn=table_name, cn=date_time_col))
+ all_date_times = c.fetchall()
+ print('4) all entries between ~2013 - 2015:', all_date_times)
+
+ # 5) Retrieve all IDs of entries between that are older than 1 day and 12 hrs
+ c.execute("SELECT {idf} FROM {tn} WHERE DATE('now') - {dc} >= 1 AND DATE('now') - {tc} >= 12".\
+ format(idf=id_field, tn=table_name, dc=date_col, tc=time_col))
+ all_1day12hrs_entries = c.fetchall()
+ print('5) entries older than 1 day:', all_1day12hrs_entries)
+
+ # Committing changes and closing the connection to the database file
+ conn.commit()
+ conn.close()
+
+
+Download the script: [date_time_ops.py](code/date_time_ops.py)
+
+
+
+
+
+
+Some of the really convenient functions that return the current time and date
+are:
+
+* * *
+
+
+ DATE('now') # returns current date, e.g., 2014-03-06
+ TIME('now') # returns current time, e.g., 10:10:10
+ CURRENT_TIMESTAMP # returns current date and time, e.g., 2014-03-06 16:42:30
+ # (or alternatively: DATETIME('now'))
+
+
+* * *
+
+The screenshot below shows the print outputs of the code that we used to query
+for entries that lie between a specified date interval using
+
+
+
+ BETWEEN '2013-03-06 10:10:10' AND '2015-03-06 10:10:10'
+
+
+and entries that are older than 1 day via
+
+
+
+ WHERE DATE('now') - some_date
+
+
+Note that we don't have to provide the complete time stamps here, the same
+syntax applies to simple dates or simple times only, too.
+
+
+
+
+
+
+#### Update Mar 16, 2014:
+
+
+If'd we are interested to calculate the hours between two `DATETIME()`
+timestamps, we can could use the handy `STRFTIME()` function like this
+
+
+
+
+ SELECT (STRFTIME('%s','2014-03-14 14:51:00') - STRFTIME('%s','2014-03-16 14:51:00'))
+ / -3600
+
+
+
+which would calculate the difference in hours between two dates in this
+particular example above (here: `48`) in this case.
+And to calculate the difference in hours between the current `DATETIME` and a
+given `DATETIME` string, we could use the following SQLite syntax:
+
+
+
+
+ SELECT (STRFTIME('%s',DATETIME('now')) - STRFTIME('%s','2014-03-15 14:51:00')) / 3600
+
+
+
+
+## Retrieving column names
+
+In the previous two sections we have seen how we query SQLite databases for
+data contents. Now let us have a look at how we retrieve its metadata (here:
+column names):
+
+
+
+ import sqlite3
+
+ sqlite_file = 'my_first_db.sqlite'
+ table_name = 'my_table_3'
+
+ # Connecting to the database file
+ conn = sqlite3.connect(sqlite_file)
+ c = conn.cursor()
+
+ # Retrieve column information
+ # Every column will be represented by a tuple with the following attributes:
+ # (id, name, type, notnull, default_value, primary_key)
+ c.execute('PRAGMA TABLE_INFO({})'.format(table_name))
+
+ # collect names in a list
+ names = [tup[1] for tup in c.fetchall()]
+ print(names)
+ # e.g., ['id', 'date', 'time', 'date_time']
+
+ # Closing the connection to the database file
+ conn.close()
+
+
+Download the script: [get_columnnames.py](code/get_columnnames.py)
+
+
+
+Since we haven't created a PRIMARY KEY column for `my_table_3`, SQLite
+automatically provides an indexed `rowid` column with unique ascending integer
+values, which will be ignored in our case. Using the `PRAGMA TABLE_INFO()`
+function on our table, we return a list of tuples, where each tuple contains
+the following information about every column in the table: `(id, name, type,
+notnull, default_value, primary_key)`.
+So, in order to get the names of every column in our table, we only have to
+grab the 2nd value in each tuple of the returned list, which can be done by
+
+
+
+ names = [tup[1] for tup in c.fetchall()]
+
+after the `PRAGMA TABLE_INFO()` call. If we would print the contents of the
+variable `names` now, the output would look like this:
+
+
+
+
+
+
+## Printing a database summary
+
+I hope we covered most of the basics about SQLite database operations in the
+previous sections, and by now we should be well equipped to get some serious
+work done using SQLite in Python.
+Let me conclude this tutorial with an obligatory "last but not least" and a
+convenient script to print a nice overview of SQLite database tables:
+
+
+
+ import sqlite3
+
+
+ def connect(sqlite_file):
+ """ Make connection to an SQLite database file """
+ conn = sqlite3.connect(sqlite_file)
+ c = conn.cursor()
+ return conn, c
+
+
+ def close(conn):
+ """ Commit changes and close connection to the database """
+ # conn.commit()
+ conn.close()
+
+
+ def total_rows(cursor, table_name, print_out=False):
+ """ Returns the total number of rows in the database """
+ cursor.execute('SELECT COUNT(*) FROM {}'.format(table_name))
+ count = cursor.fetchall()
+ if print_out:
+ print('\nTotal rows: {}'.format(count[0][0]))
+ return count[0][0]
+
+
+ def table_col_info(cursor, table_name, print_out=False):
+ """ Returns a list of tuples with column informations:
+ (id, name, type, notnull, default_value, primary_key)
+ """
+ cursor.execute('PRAGMA TABLE_INFO({})'.format(table_name))
+ info = cursor.fetchall()
+
+ if print_out:
+ print("\nColumn Info:\nID, Name, Type, NotNull, DefaultVal, PrimaryKey")
+ for col in info:
+ print(col)
+ return info
+
+
+ def values_in_col(cursor, table_name, print_out=True):
+ """ Returns a dictionary with columns as keys
+ and the number of not-null entries as associated values.
+ """
+ cursor.execute('PRAGMA TABLE_INFO({})'.format(table_name))
+ info = cursor.fetchall()
+ col_dict = dict()
+ for col in info:
+ col_dict[col[1]] = 0
+ for col in col_dict:
+ c.execute('SELECT ({0}) FROM {1} '
+ 'WHERE {0} IS NOT NULL'.format(col, table_name))
+ # In my case this approach resulted in a
+ # better performance than using COUNT
+ number_rows = len(c.fetchall())
+ col_dict[col] = number_rows
+ if print_out:
+ print("\nNumber of entries per column:")
+ for i in col_dict.items():
+ print('{}: {}'.format(i[0], i[1]))
+ return col_dict
+
+
+ if __name__ == '__main__':
+
+ sqlite_file = 'my_first_db.sqlite'
+ table_name = 'my_table_3'
+
+ conn, c = connect(sqlite_file)
+ total_rows(c, table_name, print_out=True)
+ table_col_info(c, table_name, print_out=True)
+ # next line might be slow on large databases
+ values_in_col(c, table_name, print_out=True)
+
+ close(conn)
+
+Download the script: [print_db_info.py](code/print_db_info.py)
+
+
+
+
+
+
+
+## Conclusion
+
+I really hope this tutorial was helpful to you to get started with SQLite
+database operations via Python. I have been using the `sqlite3` module a lot
+recently, and it has found its way into most of my programs for larger data
+analyses.
+Currently, I am working on a novel drug screening software that requires me to
+store 3D structures and other functional data for ~13 million chemical
+compounds, and SQLite has been an invaluable part of my program to quickly
+store, query, analyze, and share my data.
+Another smaller project that uses `sqlite3` in Python would be smilite, a
+module to retrieve and compare SMILE strings of chemical compounds from the
+free ZINC online database. If you are interested, you can check it out at:
+
+
+
+
+
+* * *
+
+#### Sections
+
+• Connecting to an SQLite database
+• Creating a new SQLite database
+ - Overview of SQLite data types
+ - A quick word on PRIMARY KEYS:
+• Adding new columns
+• Inserting and updating rows
+• Creating unique indexes
+• Querying the database - Selecting rows
+• Security and injection attacks
+• Date and time operations
+• Printing a database summary
+• Conclusion
+
+The complete Python code that I am using in this tutorial can be downloaded
+from my GitHub repository: [https://github.com/rasbt/python_reference/tree/master/tutorials/sqlite3_howto](https://github.com/rasbt/python_reference/tree/master/tutorials/sqlite3_howto)
+
+
+* * *
+
+
+
+## Connecting to an SQLite database
+
+The sqlite3 that we will be using throughout this tutorial is part of the
+Python Standard Library and is a nice and easy interface to SQLite databases:
+There are no server processes involved, no configurations required, and no
+other obstacles we have to worry about.
+
+In general, the only thing that needs to be done before we can perform any
+operation on a SQLite database via Python's `sqlite3` module, is to open a
+connection to an SQLite database file:
+
+
+
+ import sqlite3
+ conn = sqlite3.connect(sqlite_file)
+ c = conn.cursor()
+
+
+where the database file (`sqlite_file`) can reside anywhere on our disk, e.g.,
+
+
+
+ sqlite_file = '/Users/Sebastian/Desktop/my_db.sqlite'
+
+
+Conveniently, a new database file (`.sqlite` file) will be created
+automatically the first time we try to connect to a database. However, we have
+to be aware that it won't have a table, yet. In the following section, we will
+take a look at some example code of how to create a new SQLite database files
+with tables for storing some data.
+
+To round up this section about connecting to a SQLite database file, there are
+two more operations that are worth mentioning. If we are finished with our
+operations on the database file, we have to close the connection via the
+`.close()` method:
+
+
+
+ conn.close()
+
+
+And if we performed any operation on the database other than sending queries,
+we need to commit those changes via the `.commit()` method before we close the
+connection:
+
+
+
+ conn.commit()
+ conn.close()
+
+
+
+
+## Creating a new SQLite database
+
+Let us have a look at some example code to create a new SQLite database file
+with two tables: One with and one without a PRIMARY KEY column (don't worry,
+there is more information about PRIMARY KEYs further down in this section).
+
+
+
+ import sqlite3
+
+ sqlite_file = 'my_first_db.sqlite' # name of the sqlite database file
+ table_name1 = 'my_table_1' # name of the table to be created
+ table_name2 = 'my_table_2' # name of the table to be created
+ new_field = 'my_1st_column' # name of the column
+ field_type = 'INTEGER' # column data type
+
+ # Connecting to the database file
+ conn = sqlite3.connect(sqlite_file)
+ c = conn.cursor()
+
+ # Creating a new SQLite table with 1 column
+ c.execute('CREATE TABLE {tn} ({nf} {ft})'\
+ .format(tn=table_name1, nf=new_field, ft=field_type))
+
+ # Creating a second table with 1 column and set it as PRIMARY KEY
+ # note that PRIMARY KEY column must consist of unique values!
+ c.execute('CREATE TABLE {tn} ({nf} {ft} PRIMARY KEY)'\
+ .format(tn=table_name2, nf=new_field, ft=field_type))
+
+ # Committing changes and closing the connection to the database file
+ conn.commit()
+ conn.close()
+
+
+Download the script: [create_new_db.py](https://github.com/rasbt/python_reference/blob/master/tutorials/sqlite3_howto/code/create_new_db.py)
+
+* * *
+
+**Tip:** A handy tool to visualize and access SQLite databases is the free FireFox [SQLite Manager](https://addons.mozilla.org/en-US/firefox/addon/sqlite-manager/?src) add-on. Throughout this article, I will use this tool to provide screenshots of the database structures that we created below the corresponding code sections.
+
+* * *
+
+
+
+
+
+
+Using the code above, we created a new `.sqlite` database file with 2 tables.
+Each table consists of currently one column only, which is of type INTEGER.
+
+
+
+* * *
+
+**Here is a quick overview of all data types that are supported by SQLite 3:**
+
+ * INTEGER: A signed integer up to 8 bytes depending on the magnitude of the value.
+ * REAL: An 8-byte floating point value.
+ * TEXT: A text string, typically UTF-8 encoded (depending on the database encoding).
+ * BLOB: A blob of data (binary large object) for storing binary data.
+ * NULL: A NULL value, represents missing data or an empty cell.
+
+* * *
+
+Looking at the table above, You might have noticed that SQLite 3 has no
+designated Boolean data type. However, this should not be an issue, since we
+could simply re-purpose the INTEGER type to represent Boolean values (0 =
+false, 1 = true).
+
+
+
+**A quick word on PRIMARY KEYS:**
+In our example code above, we set our 1 column in the second table to PRIMARY
+KEY. The advantage of a PRIMARY KEY index is a significant performance gain if
+we use the PRIMARY KEY column as query for accessing rows in the table. Every
+table can only have max. 1 PRIMARY KEY (single or multiple column(s)), and the
+values in this column MUST be unique! But more on column indexing in the a
+later section.
+
+
+
+## Adding new columns
+
+If we want to add a new column to an existing SQLite database table, we can
+either leave the cells for each row empty (NULL value), or we can set a
+default value for each cell, which is pretty convenient for certain
+applications.
+Let's have a look at some code:
+
+
+
+ import sqlite3
+
+ sqlite_file = 'my_first_db.sqlite' # name of the sqlite database file
+ table_name = 'my_table_2' # name of the table to be created
+ id_column = 'my_1st_column' # name of the PRIMARY KEY column
+ new_column1 = 'my_2nd_column' # name of the new column
+ new_column2 = 'my_3nd_column' # name of the new column
+ column_type = 'TEXT' # E.g., INTEGER, TEXT, NULL, REAL, BLOB
+ default_val = 'Hello World' # a default value for the new column rows
+
+ # Connecting to the database file
+ conn = sqlite3.connect(sqlite_file)
+ c = conn.cursor()
+
+ # A) Adding a new column without a row value
+ c.execute("ALTER TABLE {tn} ADD COLUMN '{cn}' {ct}"\
+ .format(tn=table_name, cn=new_column1, ct=column_type))
+
+ # B) Adding a new column with a default row value
+ c.execute("ALTER TABLE {tn} ADD COLUMN '{cn}' {ct} DEFAULT '{df}'"\
+ .format(tn=table_name, cn=new_column2, ct=column_type, df=default_val))
+
+ # Committing changes and closing the connection to the database file
+ conn.commit()
+ conn.close()
+
+
+Download the script: [add_new_column.py](https://github.com/rasbt/python_reference/blob/master/tutorials/sqlite3_howto/code/add_new_column.py)
+
+
+
+
+
+
+We just added 2 more columns (`my_2nd_column` and `my_3rd_column`) to
+`my_table_2` of our SQLite database next to the PRIMARY KEY column
+`my_1st_column`.
+The difference between the two new columns is that we initialized
+`my_3rd_column` with a default value (here:'Hello World'), which will be
+inserted for every existing cell under this column and for every new row that
+we are going to add to the table if we don't insert or update it with a
+different value.
+
+
+
+## Inserting and updating rows
+
+Inserting and updating rows into an existing SQLite database table - next to
+sending queries - is probably the most common database operation. The
+Structured Query Language has a convenient `UPSERT` function, which is
+basically just a merge between UPDATE and INSERT: It inserts new rows into a
+database table with a value for the PRIMARY KEY column if it does not exist
+yet, or updates a row for an existing PRIMARY KEY value. Unfortunately, this
+convenient syntax is not supported by the more compact SQLite database
+implementation that we are using here. However, there are some workarounds.
+But let us first have a look at the example code:
+
+
+
+ import sqlite3
+
+ sqlite_file = 'my_first_db.sqlite'
+ table_name = 'my_table_2'
+ id_column = 'my_1st_column'
+ column_name = 'my_2nd_column'
+
+ # Connecting to the database file
+ conn = sqlite3.connect(sqlite_file)
+ c = conn.cursor()
+
+ # A) Inserts an ID with a specific value in a second column
+ try:
+ c.execute("INSERT INTO {tn} ({idf}, {cn}) VALUES (123456, 'test')".\
+ format(tn=table_name, idf=id_column, cn=column_name))
+ except sqlite3.IntegrityError:
+ print('ERROR: ID already exists in PRIMARY KEY column {}'.format(id_column))
+
+ # B) Tries to insert an ID (if it does not exist yet)
+ # with a specific value in a second column
+ c.execute("INSERT OR IGNORE INTO {tn} ({idf}, {cn}) VALUES (123456, 'test')".\
+ format(tn=table_name, idf=id_column, cn=column_name))
+
+ # C) Updates the newly inserted or pre-existing entry
+ c.execute("UPDATE {tn} SET {cn}=('Hi World') WHERE {idf}=(123456)".\
+ format(tn=table_name, cn=column_name, idf=id_column))
+
+ conn.commit()
+ conn.close()
+
+
+Download the script: [update_or_insert_records.py](code/update_or_insert_records.py)
+
+
+
+Both A) `INSERT` and B) `INSERT OR IGNORE` have in common that they append new
+rows to the database if a given PRIMARY KEY does not exist in the database
+table, yet. However, if we'd try to append a PRIMARY KEY value that is not
+unique, a simple `INSERT` would raise an `sqlite3.IntegrityError` exception,
+which can be either captured via a try-except statement (case A) or
+circumvented by the SQLite call `INSERT OR IGNORE` (case B). This can be
+pretty useful if we want to construct an `UPSERT` equivalent in SQLite. E.g.,
+if we want to add a dataset to an existing database table that contains a mix
+between existing and new IDs for our PRIMARY KEY column.
+
+
+
+## Creating unique indexes
+
+Just like hashtable-datastructures, indexes function as direct pointers to our
+data in a table for a particular column (i.e., the indexed column). For
+example, the PRIMARY KEY column would have such an index by default. The
+downside of indexes is that every row value in the column must be unique.
+However, it is recommended and pretty useful to index certain columns if
+possible, since it rewards us with a significant performance gain for the data
+retrieval.
+The example code below shows how to add such an unique index to an existing
+column in an SQLite database table. And if we should decide to insert non-
+unique values into a indexed column later, there is also a convenient way to
+drop the index, which is also shown in the code below.
+
+
+
+ import sqlite3
+
+ sqlite_file = 'my_first_db.sqlite' # name of the sqlite database file
+ table_name = 'my_table_2' # name of the table to be created
+ id_column = 'my_1st_column' # name of the PRIMARY KEY column
+ new_column = 'unique_names' # name of the new column
+ column_type = 'TEXT' # E.g., INTEGER, TEXT, NULL, REAL, BLOB
+ index_name = 'my_unique_index' # name for the new unique index
+
+ # Connecting to the database file
+ conn = sqlite3.connect(sqlite_file)
+ c = conn.cursor()
+
+ # Adding a new column and update some record
+ c.execute("ALTER TABLE {tn} ADD COLUMN '{cn}' {ct}"\
+ .format(tn=table_name, cn=new_column, ct=column_type))
+ c.execute("UPDATE {tn} SET {cn}='sebastian_r' WHERE {idf}=123456".\
+ format(tn=table_name, idf=id_column, cn=new_column))
+
+ # Creating an unique index
+ c.execute('CREATE INDEX {ix} on {tn}({cn})'\
+ .format(ix=index_name, tn=table_name, cn=new_column))
+
+ # Dropping the unique index
+ # E.g., to avoid future conflicts with update/insert functions
+ c.execute('DROP INDEX {ix}'.format(ix=index_name))
+
+ # Committing changes and closing the connection to the database file
+ conn.commit()
+ conn.close()
+
+
+Download the script: [create_unique_index.py](code/create_unique_index.py)
+
+
+
+
+
+
+## Querying the database - Selecting rows
+
+After we learned about how to create and modify SQLite databases, it's about
+time for some data retrieval. The code below illustrates how we can retrieve
+row entries for all or some columns if they match certain criteria.
+
+
+
+ import sqlite3
+
+ sqlite_file = 'my_first_db.sqlite' # name of the sqlite database file
+ table_name = 'my_table_2' # name of the table to be queried
+ id_column = 'my_1st_column'
+ some_id = 123456
+ column_2 = 'my_2nd_column'
+ column_3 = 'my_3rd_column'
+
+ # Connecting to the database file
+ conn = sqlite3.connect(sqlite_file)
+ c = conn.cursor()
+
+ # 1) Contents of all columns for row that match a certain value in 1 column
+ c.execute('SELECT * FROM {tn} WHERE {cn}="Hi World"'.\
+ format(tn=table_name, cn=column_2))
+ all_rows = c.fetchall()
+ print('1):', all_rows)
+
+ # 2) Value of a particular column for rows that match a certain value in column_1
+ c.execute('SELECT ({coi}) FROM {tn} WHERE {cn}="Hi World"'.\
+ format(coi=column_2, tn=table_name, cn=column_2))
+ all_rows = c.fetchall()
+ print('2):', all_rows)
+
+ # 3) Value of 2 particular columns for rows that match a certain value in 1 column
+ c.execute('SELECT {coi1},{coi2} FROM {tn} WHERE {coi1}="Hi World"'.\
+ format(coi1=column_2, coi2=column_3, tn=table_name, cn=column_2))
+ all_rows = c.fetchall()
+ print('3):', all_rows)
+
+ # 4) Selecting only up to 10 rows that match a certain value in 1 column
+ c.execute('SELECT * FROM {tn} WHERE {cn}="Hi World" LIMIT 10'.\
+ format(tn=table_name, cn=column_2))
+ ten_rows = c.fetchall()
+ print('4):', ten_rows)
+
+ # 5) Check if a certain ID exists and print its column contents
+ c.execute("SELECT * FROM {tn} WHERE {idf}={my_id}".\
+ format(tn=table_name, cn=column_2, idf=id_column, my_id=some_id))
+ id_exists = c.fetchone()
+ if id_exists:
+ print('5): {}'.format(id_exists))
+ else:
+ print('5): {} does not exist'.format(some_id))
+
+ # Closing the connection to the database file
+ conn.close()
+
+
+Download the script: [selecting_entries.py](code/selecting_entries.py)
+
+
+
+
+if we use the `.fetchall()` method, we return a list of tuples from the
+database query, where each tuple represents one row entry. The print output
+for the 5 different cases shown in the code above would look like this (note
+that we only have a table with 1 row here):
+
+
+
+
+
+
+## Security and injection attacks
+
+So far, we have been using Python's string formatting method to insert
+parameters like table and column names into the `c.execute()` functions. This
+is fine if we just want to use the database for ourselves. However, this
+leaves our database vulnerable to injection attacks. For example, if our
+database would be part of a web application, it would allow hackers to
+directly communicate with the database in order to bypass login and password
+verification and steal data.
+In order to prevent this, it is recommended to use `?` place holders in the
+SQLite commands instead of the `%` formatting expression or the `.format()`
+method, which we have been using in this tutorial.
+For example, instead of using
+
+
+
+ # 5) Check if a certain ID exists and print its column contents
+ c.execute("SELECT * FROM {tn} WHERE {idf}={my_id}".\
+ format(tn=table_name, cn=column_2, idf=id_column, my_id=some_id))
+
+
+in the Querying the database - Selecting rows section above, we would want to
+use the `?` placeholder for the queried column value and include the
+variable(s) (here: `123456`), which we want to insert, as tuple at the end of
+the `c.execute()` string.
+
+
+
+ # 5) Check if a certain ID exists and print its column contents
+ c.execute("SELECT * FROM {tn} WHERE {idf}=?".\
+ format(tn=table_name, cn=column_2, idf=id_column), (123456,))
+
+
+However, the problem with this approach is that it would only work for values,
+not for column or table names. So what are we supposed to do with the rest of
+the string if we want to protect ourselves from injection attacks? The easy
+solution would be to refrain from using variables in SQLite queries whenever
+possible, and if it cannot be avoided, we would want to use a function that
+strips all non-alphanumerical characters from the stored content of the
+variable, e.g.,
+
+
+
+ def clean_name(some_var):
+ return ''.join(char for char in some_var if char.isalnum())
+
+
+
+
+## Date and time operations
+
+SQLite inherited the convenient date and time operations from SQL, which are
+one of my favorite features of the Structured Query Language: It does not only
+allow us to insert dates and times in various different formats, but we can
+also perform simple `+` and `-` arithmetic, for example to look up entries
+that have been added xxx days ago.
+
+
+
+ import sqlite3
+
+ sqlite_file = 'my_first_db.sqlite' # name of the sqlite database file
+ table_name = 'my_table_3' # name of the table to be created
+ id_field = 'id' # name of the ID column
+ date_col = 'date' # name of the date column
+ time_col = 'time'# name of the time column
+ date_time_col = 'date_time' # name of the date & time column
+ field_type = 'TEXT' # column data type
+
+ # Connecting to the database file
+ conn = sqlite3.connect(sqlite_file)
+ c = conn.cursor()
+
+ # Creating a new SQLite table with 1 column
+ c.execute('CREATE TABLE {tn} ({fn} {ft} PRIMARY KEY)'\
+ .format(tn=table_name, fn=id_field, ft=field_type))
+
+ # A) Adding a new column to save date insert a row with the current date
+ # in the following format: YYYY-MM-DD
+ # e.g., 2014-03-06
+ c.execute("ALTER TABLE {tn} ADD COLUMN '{cn}'"\
+ .format(tn=table_name, cn=date_col))
+ # insert a new row with the current date and time, e.g., 2014-03-06
+ c.execute("INSERT INTO {tn} ({idf}, {cn}) VALUES('some_id1', DATE('now'))"\
+ .format(tn=table_name, idf=id_field, cn=date_col))
+
+ # B) Adding a new column to save date and time and update with the current time
+ # in the following format: HH:MM:SS
+ # e.g., 16:26:37
+ c.execute("ALTER TABLE {tn} ADD COLUMN '{cn}'"\
+ .format(tn=table_name, cn=time_col))
+ # update row for the new current date and time column, e.g., 2014-03-06 16:26:37
+ c.execute("UPDATE {tn} SET {cn}=TIME('now') WHERE {idf}='some_id1'"\
+ .format(tn=table_name, idf=id_field, cn=time_col))
+
+ # C) Adding a new column to save date and time and update with current date-time
+ # in the following format: YYYY-MM-DD HH:MM:SS
+ # e.g., 2014-03-06 16:26:37
+ c.execute("ALTER TABLE {tn} ADD COLUMN '{cn}'"\
+ .format(tn=table_name, cn=date_time_col))
+ # update row for the new current date and time column, e.g., 2014-03-06 16:26:37
+ c.execute("UPDATE {tn} SET {cn}=(CURRENT_TIMESTAMP) WHERE {idf}='some_id1'"\
+ .format(tn=table_name, idf=id_field, cn=date_time_col))
+
+ # The database should now look like this:
+ # id date time date_time
+ # "some_id1" "2014-03-06" "16:42:30" "2014-03-06 16:42:30"
+
+ # 4) Retrieve all IDs of entries between 2 date_times
+ c.execute("SELECT {idf} FROM {tn} WHERE {cn} BETWEEN '2013-03-06 10:10:10' AND '2015-03-06 10:10:10'".\
+ format(idf=id_field, tn=table_name, cn=date_time_col))
+ all_date_times = c.fetchall()
+ print('4) all entries between ~2013 - 2015:', all_date_times)
+
+ # 5) Retrieve all IDs of entries between that are older than 1 day and 12 hrs
+ c.execute("SELECT {idf} FROM {tn} WHERE DATE('now') - {dc} >= 1 AND DATE('now') - {tc} >= 12".\
+ format(idf=id_field, tn=table_name, dc=date_col, tc=time_col))
+ all_1day12hrs_entries = c.fetchall()
+ print('5) entries older than 1 day:', all_1day12hrs_entries)
+
+ # Committing changes and closing the connection to the database file
+ conn.commit()
+ conn.close()
+
+
+Download the script: [date_time_ops.py](code/date_time_ops.py)
+
+
+
+
+
+
+Some of the really convenient functions that return the current time and date
+are:
+
+* * *
+
+
+ DATE('now') # returns current date, e.g., 2014-03-06
+ TIME('now') # returns current time, e.g., 10:10:10
+ CURRENT_TIMESTAMP # returns current date and time, e.g., 2014-03-06 16:42:30
+ # (or alternatively: DATETIME('now'))
+
+
+* * *
+
+The screenshot below shows the print outputs of the code that we used to query
+for entries that lie between a specified date interval using
+
+
+
+ BETWEEN '2013-03-06 10:10:10' AND '2015-03-06 10:10:10'
+
+
+and entries that are older than 1 day via
+
+
+
+ WHERE DATE('now') - some_date
+
+
+Note that we don't have to provide the complete time stamps here, the same
+syntax applies to simple dates or simple times only, too.
+
+
+
+
+
+
+#### Update Mar 16, 2014:
+
+
+If'd we are interested to calculate the hours between two `DATETIME()`
+timestamps, we can could use the handy `STRFTIME()` function like this
+
+
+
+
+ SELECT (STRFTIME('%s','2014-03-14 14:51:00') - STRFTIME('%s','2014-03-16 14:51:00'))
+ / -3600
+
+
+
+which would calculate the difference in hours between two dates in this
+particular example above (here: `48`) in this case.
+And to calculate the difference in hours between the current `DATETIME` and a
+given `DATETIME` string, we could use the following SQLite syntax:
+
+
+
+
+ SELECT (STRFTIME('%s',DATETIME('now')) - STRFTIME('%s','2014-03-15 14:51:00')) / 3600
+
+
+
+
+## Retrieving column names
+
+In the previous two sections we have seen how we query SQLite databases for
+data contents. Now let us have a look at how we retrieve its metadata (here:
+column names):
+
+
+
+ import sqlite3
+
+ sqlite_file = 'my_first_db.sqlite'
+ table_name = 'my_table_3'
+
+ # Connecting to the database file
+ conn = sqlite3.connect(sqlite_file)
+ c = conn.cursor()
+
+ # Retrieve column information
+ # Every column will be represented by a tuple with the following attributes:
+ # (id, name, type, notnull, default_value, primary_key)
+ c.execute('PRAGMA TABLE_INFO({})'.format(table_name))
+
+ # collect names in a list
+ names = [tup[1] for tup in c.fetchall()]
+ print(names)
+ # e.g., ['id', 'date', 'time', 'date_time']
+
+ # Closing the connection to the database file
+ conn.close()
+
+
+Download the script: [get_columnnames.py](code/get_columnnames.py)
+
+
+
+Since we haven't created a PRIMARY KEY column for `my_table_3`, SQLite
+automatically provides an indexed `rowid` column with unique ascending integer
+values, which will be ignored in our case. Using the `PRAGMA TABLE_INFO()`
+function on our table, we return a list of tuples, where each tuple contains
+the following information about every column in the table: `(id, name, type,
+notnull, default_value, primary_key)`.
+So, in order to get the names of every column in our table, we only have to
+grab the 2nd value in each tuple of the returned list, which can be done by
+
+
+
+ names = [tup[1] for tup in c.fetchall()]
+
+after the `PRAGMA TABLE_INFO()` call. If we would print the contents of the
+variable `names` now, the output would look like this:
+
+
+
+
+
+
+## Printing a database summary
+
+I hope we covered most of the basics about SQLite database operations in the
+previous sections, and by now we should be well equipped to get some serious
+work done using SQLite in Python.
+Let me conclude this tutorial with an obligatory "last but not least" and a
+convenient script to print a nice overview of SQLite database tables:
+
+
+
+ import sqlite3
+
+
+ def connect(sqlite_file):
+ """ Make connection to an SQLite database file """
+ conn = sqlite3.connect(sqlite_file)
+ c = conn.cursor()
+ return conn, c
+
+
+ def close(conn):
+ """ Commit changes and close connection to the database """
+ # conn.commit()
+ conn.close()
+
+
+ def total_rows(cursor, table_name, print_out=False):
+ """ Returns the total number of rows in the database """
+ cursor.execute('SELECT COUNT(*) FROM {}'.format(table_name))
+ count = cursor.fetchall()
+ if print_out:
+ print('\nTotal rows: {}'.format(count[0][0]))
+ return count[0][0]
+
+
+ def table_col_info(cursor, table_name, print_out=False):
+ """ Returns a list of tuples with column informations:
+ (id, name, type, notnull, default_value, primary_key)
+ """
+ cursor.execute('PRAGMA TABLE_INFO({})'.format(table_name))
+ info = cursor.fetchall()
+
+ if print_out:
+ print("\nColumn Info:\nID, Name, Type, NotNull, DefaultVal, PrimaryKey")
+ for col in info:
+ print(col)
+ return info
+
+
+ def values_in_col(cursor, table_name, print_out=True):
+ """ Returns a dictionary with columns as keys
+ and the number of not-null entries as associated values.
+ """
+ cursor.execute('PRAGMA TABLE_INFO({})'.format(table_name))
+ info = cursor.fetchall()
+ col_dict = dict()
+ for col in info:
+ col_dict[col[1]] = 0
+ for col in col_dict:
+ c.execute('SELECT ({0}) FROM {1} '
+ 'WHERE {0} IS NOT NULL'.format(col, table_name))
+ # In my case this approach resulted in a
+ # better performance than using COUNT
+ number_rows = len(c.fetchall())
+ col_dict[col] = number_rows
+ if print_out:
+ print("\nNumber of entries per column:")
+ for i in col_dict.items():
+ print('{}: {}'.format(i[0], i[1]))
+ return col_dict
+
+
+ if __name__ == '__main__':
+
+ sqlite_file = 'my_first_db.sqlite'
+ table_name = 'my_table_3'
+
+ conn, c = connect(sqlite_file)
+ total_rows(c, table_name, print_out=True)
+ table_col_info(c, table_name, print_out=True)
+ # next line might be slow on large databases
+ values_in_col(c, table_name, print_out=True)
+
+ close(conn)
+
+Download the script: [print_db_info.py](code/print_db_info.py)
+
+
+
+
+
+
+
+## Conclusion
+
+I really hope this tutorial was helpful to you to get started with SQLite
+database operations via Python. I have been using the `sqlite3` module a lot
+recently, and it has found its way into most of my programs for larger data
+analyses.
+Currently, I am working on a novel drug screening software that requires me to
+store 3D structures and other functional data for ~13 million chemical
+compounds, and SQLite has been an invaluable part of my program to quickly
+store, query, analyze, and share my data.
+Another smaller project that uses `sqlite3` in Python would be smilite, a
+module to retrieve and compare SMILE strings of chemical compounds from the
+free ZINC online database. If you are interested, you can check it out at:
+.
+
+If you have any suggestions or questions, please don't hesitate to write me an
+[ email](mailto:se.raschka@gmail.com) or leave a comment in the comment
+section below! I am looking forward to your opinions and ideas, and I hope I
+can improve and extend this tutorial in future.
diff --git a/tutorials/sqlite3_howto/code/add_new_column.py b/tutorials/sqlite3_howto/code/add_new_column.py
new file mode 100644
index 0000000..7e9fcd5
--- /dev/null
+++ b/tutorials/sqlite3_howto/code/add_new_column.py
@@ -0,0 +1,28 @@
+# Sebastian Raschka, 2014
+# Adding a new column to an existing SQLite database
+
+import sqlite3
+
+sqlite_file = 'my_first_db.sqlite' # name of the sqlite database file
+table_name = 'my_table_2' # name of the table to be created
+id_column = 'my_1st_column' # name of the PRIMARY KEY column
+new_column1 = 'my_2nd_column' # name of the new column
+new_column2 = 'my_3rd_column' # name of the new column
+column_type = 'TEXT' # E.g., INTEGER, TEXT, NULL, REAL, BLOB
+default_val = 'Hello World' # a default value for the new column rows
+
+# Connecting to the database file
+conn = sqlite3.connect(sqlite_file)
+c = conn.cursor()
+
+# A) Adding a new column without a row value
+c.execute("ALTER TABLE {tn} ADD COLUMN '{cn}' {ct}"\
+ .format(tn=table_name, cn=new_column1, ct=column_type))
+
+# B) Adding a new column with a default row value
+c.execute("ALTER TABLE {tn} ADD COLUMN '{cn}' {ct} DEFAULT '{df}'"\
+ .format(tn=table_name, cn=new_column2, ct=column_type, df=default_val))
+
+# Committing changes and closing the connection to the database file
+conn.commit()
+conn.close()
diff --git a/tutorials/sqlite3_howto/code/create_new_db.py b/tutorials/sqlite3_howto/code/create_new_db.py
new file mode 100644
index 0000000..df220da
--- /dev/null
+++ b/tutorials/sqlite3_howto/code/create_new_db.py
@@ -0,0 +1,27 @@
+# Sebastian Raschka, 2014
+# Creating a new SQLite database
+
+import sqlite3
+
+sqlite_file = 'my_first_db.sqlite' # name of the sqlite database file
+table_name1 = 'my_table_1' # name of the table to be created
+table_name2 = 'my_table_2' # name of the table to be created
+new_field = 'my_1st_column' # name of the column
+field_type = 'INTEGER' # column data type
+
+# Connecting to the database file
+conn = sqlite3.connect(sqlite_file)
+c = conn.cursor()
+
+# Creating a new SQLite table with 1 column
+c.execute('CREATE TABLE {tn} ({nf} {ft})'\
+ .format(tn=table_name1, nf=new_field, ft=field_type))
+
+# Creating a second table with 1 column and set it as PRIMARY KEY
+# note that PRIMARY KEY column must consist of unique values!
+c.execute('CREATE TABLE {tn} ({nf} {ft} PRIMARY KEY)'\
+ .format(tn=table_name2, nf=new_field, ft=field_type))
+
+# Committing changes and closing the connection to the database file
+conn.commit()
+conn.close()
diff --git a/tutorials/sqlite3_howto/code/create_unique_index.py b/tutorials/sqlite3_howto/code/create_unique_index.py
new file mode 100644
index 0000000..28f56a8
--- /dev/null
+++ b/tutorials/sqlite3_howto/code/create_unique_index.py
@@ -0,0 +1,34 @@
+# Sebastian Raschka, 2014
+# Creating an index on a column with unique! values
+# Boosts performance for data base operations.
+
+import sqlite3
+
+sqlite_file = 'my_first_db.sqlite' # name of the sqlite database file
+table_name = 'my_table_2' # name of the table to be created
+id_column = 'my_1st_column' # name of the PRIMARY KEY column
+new_column = 'unique_names' # name of the new column
+column_type = 'TEXT' # E.g., INTEGER, TEXT, NULL, REAL, BLOB
+index_name = 'my_unique_index' # name for the new unique index
+
+# Connecting to the database file
+conn = sqlite3.connect(sqlite_file)
+c = conn.cursor()
+
+# Adding a new column and update some record
+c.execute("ALTER TABLE {tn} ADD COLUMN '{cn}' {ct}"\
+ .format(tn=table_name, cn=new_column, ct=column_type))
+c.execute("UPDATE {tn} SET {cn}='sebastian_r' WHERE {idf}=123456".\
+ format(tn=table_name, idf=id_column, cn=new_column))
+
+# Creating an unique index
+c.execute('CREATE INDEX {ix} on {tn}({cn})'\
+ .format(ix=index_name, tn=table_name, cn=new_column))
+
+# Dropping the unique index
+# E.g., to avoid future conflicts with update/insert functions
+c.execute('DROP INDEX {ix}'.format(ix=index_name))
+
+# Committing changes and closing the connection to the database file
+conn.commit()
+conn.close()
diff --git a/tutorials/sqlite3_howto/code/date_time_ops.py b/tutorials/sqlite3_howto/code/date_time_ops.py
new file mode 100644
index 0000000..ddb8547
--- /dev/null
+++ b/tutorials/sqlite3_howto/code/date_time_ops.py
@@ -0,0 +1,69 @@
+# Sebastian Raschka, 03/2014
+# Date and Time operations in sqlite3
+
+import sqlite3
+
+sqlite_file = 'my_first_db.sqlite' # name of the sqlite database file
+table_name = 'my_table_3' # name of the table to be created
+id_field = 'id' # name of the ID column
+date_col = 'date' # name of the date column
+time_col = 'time'# name of the time column
+date_time_col = 'date_time' # name of the date & time column
+field_type = 'TEXT' # column data type
+
+# Connecting to the database file
+conn = sqlite3.connect(sqlite_file)
+c = conn.cursor()
+
+# Creating a new SQLite table with 1 column
+c.execute('CREATE TABLE {tn} ({fn} {ft} PRIMARY KEY)'\
+ .format(tn=table_name, fn=id_field, ft=field_type))
+
+
+# 1) Adding a new column to save date insert a row with the current date
+# in the following format: YYYY-MM-DD
+# e.g., 2014-03-06
+c.execute("ALTER TABLE {tn} ADD COLUMN '{cn}'"\
+ .format(tn=table_name, cn=date_col))
+# insert a new row with the current date and time, e.g., 2014-03-06
+c.execute("INSERT INTO {tn} ({idf}, {cn}) VALUES('some_id1', DATE('now'))"\
+ .format(tn=table_name, idf=id_field, cn=date_col))
+
+
+# 2) Adding a new column to save date and time and update with the current time
+# in the following format: HH:MM:SS
+# e.g., 16:26:37
+c.execute("ALTER TABLE {tn} ADD COLUMN '{cn}'"\
+ .format(tn=table_name, cn=time_col))
+# update row for the new current date and time column, e.g., 2014-03-06 16:26:37
+c.execute("UPDATE {tn} SET {cn}=TIME('now') WHERE {idf}='some_id1'"\
+ .format(tn=table_name, idf=id_field, cn=time_col))
+
+# 3) Adding a new column to save date and time and update with current date-time
+# in the following format: YYYY-MM-DD HH:MM:SS
+# e.g., 2014-03-06 16:26:37
+c.execute("ALTER TABLE {tn} ADD COLUMN '{cn}'"\
+ .format(tn=table_name, cn=date_time_col))
+# update row for the new current date and time column, e.g., 2014-03-06 16:26:37
+c.execute("UPDATE {tn} SET {cn}=(CURRENT_TIMESTAMP) WHERE {idf}='some_id1'"\
+ .format(tn=table_name, idf=id_field, cn=date_time_col))
+
+# Database should now look like this:
+# id date time date_time
+# "some_id1" "2014-03-06" "16:42:30" "2014-03-06 16:42:30"
+
+# 4) Retrieve all IDs of entries between 2 date_times
+c.execute("SELECT {idf} FROM {tn} WHERE {cn} BETWEEN '2013-03-06 10:10:10' AND '2015-03-06 10:10:10'".\
+ format(idf=id_field, tn=table_name, cn=date_time_col))
+all_date_times = c.fetchall()
+print('4) all entries between ~2013 - 2015:', all_date_times)
+
+# 5) Retrieve all IDs of entries between that are older than 1 day and 12 hrs
+c.execute("SELECT {idf} FROM {tn} WHERE DATE('now') - {dc} >= 1 AND DATE('now') - {tc} >= 12".\
+ format(idf=id_field, tn=table_name, dc=date_col, tc=time_col))
+all_1day12hrs_entries = c.fetchall()
+print('5) entries older than 1 day:', all_1day12hrs_entries)
+
+# Committing changes and closing the connection to the database file
+conn.commit()
+conn.close()
diff --git a/tutorials/sqlite3_howto/code/get_columnnames.py b/tutorials/sqlite3_howto/code/get_columnnames.py
new file mode 100644
index 0000000..f02142e
--- /dev/null
+++ b/tutorials/sqlite3_howto/code/get_columnnames.py
@@ -0,0 +1,24 @@
+# Sebastian Raschka, 2014
+# Getting column names of an SQLite database table
+
+import sqlite3
+
+sqlite_file = 'my_first_db.sqlite'
+table_name = 'my_table_3'
+
+# Connecting to the database file
+conn = sqlite3.connect(sqlite_file)
+c = conn.cursor()
+
+# Retrieve column information
+# Every column will be represented by a tuple with the following attributes:
+# (id, name, type, notnull, default_value, primary_key)
+c.execute('PRAGMA TABLE_INFO({})'.format(table_name))
+
+# collect names in a list
+names = [tup[1] for tup in c.fetchall()]
+print(names)
+# e.g., ['id', 'date', 'time', 'date_time']
+
+# Closing the connection to the database file
+conn.close()
diff --git a/tutorials/sqlite3_howto/code/print_db_info.py b/tutorials/sqlite3_howto/code/print_db_info.py
new file mode 100644
index 0000000..285a635
--- /dev/null
+++ b/tutorials/sqlite3_howto/code/print_db_info.py
@@ -0,0 +1,96 @@
+# Sebastian Raschka 2014
+# Prints Information of a SQLite database.
+
+# E.g.,
+#
+"""
+Total rows: 1
+
+Column Info:
+ID, Name, Type, NotNull, DefaultVal, PrimaryKey
+(0, 'id', 'TEXT', 0, None, 1)
+(1, 'date', '', 0, None, 0)
+(2, 'time', '', 0, None, 0)
+(3, 'date_time', '', 0, None, 0)
+
+Number of entries per column:
+date: 1
+date_time: 1
+id: 1
+time: 1
+"""
+
+import sqlite3
+
+
+def connect(sqlite_file):
+ """ Make connection to an SQLite database file """
+ conn = sqlite3.connect(sqlite_file)
+ c = conn.cursor()
+ return conn, c
+
+
+def close(conn):
+ """ Commit changes and close connection to the database """
+ # conn.commit()
+ conn.close()
+
+
+def total_rows(cursor, table_name, print_out=False):
+ """ Returns the total number of rows in the database """
+ cursor.execute('SELECT COUNT(*) FROM {}'.format(table_name))
+ count = cursor.fetchall()
+ if print_out:
+ print('\nTotal rows: {}'.format(count[0][0]))
+ return count[0][0]
+
+
+def table_col_info(cursor, table_name, print_out=False):
+ """ Returns a list of tuples with column informations:
+ (id, name, type, notnull, default_value, primary_key)
+ """
+ cursor.execute('PRAGMA TABLE_INFO({})'.format(table_name))
+ info = cursor.fetchall()
+
+ if print_out:
+ print("\nColumn Info:\nID, Name, Type, NotNull, DefaultVal, PrimaryKey")
+ for col in info:
+ print(col)
+ return info
+
+
+def values_in_col(cursor, table_name, print_out=True):
+ """ Returns a dictionary with columns as keys
+ and the number of not-null entries as associated values.
+ """
+ cursor.execute('PRAGMA TABLE_INFO({})'.format(table_name))
+ info = cursor.fetchall()
+ col_dict = dict()
+ for col in info:
+ col_dict[col[1]] = 0
+ for col in col_dict:
+ c.execute('SELECT ({0}) FROM {1} '
+ 'WHERE {0} IS NOT NULL'.format(col, table_name))
+ # In my case this approach resulted in a
+ # better performance than using COUNT
+ number_rows = len(c.fetchall())
+ col_dict[col] = number_rows
+ if print_out:
+ print("\nNumber of entries per column:")
+ for i in col_dict.items():
+ print('{}: {}'.format(i[0], i[1]))
+ return col_dict
+
+
+if __name__ == '__main__':
+
+ sqlite_file = 'my_first_db.sqlite'
+ table_name = 'my_table_3'
+
+ conn, c = connect(sqlite_file)
+ total_rows(c, table_name, print_out=True)
+ table_col_info(c, table_name, print_out=True)
+ # next line might be slow on large databases
+ values_in_col(c, table_name, print_out=True)
+
+ close(conn)
diff --git a/tutorials/sqlite3_howto/code/selecting_entries.py b/tutorials/sqlite3_howto/code/selecting_entries.py
new file mode 100644
index 0000000..0ba8e19
--- /dev/null
+++ b/tutorials/sqlite3_howto/code/selecting_entries.py
@@ -0,0 +1,51 @@
+# Sebastian Raschka, 2014
+# Selecting rows from an existing SQLite database
+
+import sqlite3
+
+sqlite_file = 'my_first_db.sqlite' # name of the sqlite database file
+table_name = 'my_table_2' # name of the table to be queried
+id_column = 'my_1st_column'
+some_id = 123456
+column_2 = 'my_2nd_column'
+column_3 = 'my_3rd_column'
+
+# Connecting to the database file
+conn = sqlite3.connect(sqlite_file)
+c = conn.cursor()
+
+# 1) Contents of all columns for row that match a certain value in 1 column
+c.execute('SELECT * FROM {tn} WHERE {cn}="Hi World"'.\
+ format(tn=table_name, cn=column_2))
+all_rows = c.fetchall()
+print('1):', all_rows)
+
+# 2) Value of a particular column for rows that match a certain value in column_1
+c.execute('SELECT ({coi}) FROM {tn} WHERE {cn}="Hi World"'.\
+ format(coi=column_2, tn=table_name, cn=column_2))
+all_rows = c.fetchall()
+print('2):', all_rows)
+
+# 3) Value of 2 particular columns for rows that match a certain value in 1 column
+c.execute('SELECT {coi1},{coi2} FROM {tn} WHERE {coi1}="Hi World"'.\
+ format(coi1=column_2, coi2=column_3, tn=table_name, cn=column_2))
+all_rows = c.fetchall()
+print('3):', all_rows)
+
+# 4) Selecting only up to 10 rows that match a certain value in 1 column
+c.execute('SELECT * FROM {tn} WHERE {cn}="Hi World" LIMIT 10'.\
+ format(tn=table_name, cn=column_2))
+ten_rows = c.fetchall()
+print('4):', ten_rows)
+
+# 5) Check if a certain ID exists and print its column contents
+c.execute("SELECT * FROM {tn} WHERE {idf}=?".\
+ format(tn=table_name, cn=column_2, idf=id_column), (123456,))
+id_exists = c.fetchone()
+if id_exists:
+ print('5): {}'.format(id_exists))
+else:
+ print('5): {} does not exist'.format(some_id))
+
+# Closing the connection to the database file
+conn.close()
diff --git a/tutorials/sqlite3_howto/code/update_or_insert_records.py b/tutorials/sqlite3_howto/code/update_or_insert_records.py
new file mode 100644
index 0000000..ee461ec
--- /dev/null
+++ b/tutorials/sqlite3_howto/code/update_or_insert_records.py
@@ -0,0 +1,35 @@
+# Sebastian Raschka, 2014
+# Update records or insert them if they don't exist.
+# Note that this is a workaround to accommodate for missing
+# SQL features in SQLite.
+
+import sqlite3
+
+sqlite_file = 'my_first_db.sqlite'
+table_name = 'my_table_2'
+id_column = 'my_1st_column'
+column_name = 'my_2nd_column'
+
+# Connecting to the database file
+conn = sqlite3.connect(sqlite_file)
+c = conn.cursor()
+
+
+# A) Inserts an ID with a specific value in a second column
+try:
+ c.execute("INSERT INTO {tn} ({idf}, {cn}) VALUES (123456, 'test')".\
+ format(tn=table_name, idf=id_column, cn=column_name))
+except sqlite3.IntegrityError:
+ print('ERROR: ID already exists in PRIMARY KEY column {}'.format(id_column))
+
+# B) Tries to insert an ID (if it does not exist yet)
+# with a specific value in a second column
+c.execute("INSERT OR IGNORE INTO {tn} ({idf}, {cn}) VALUES (123456, 'test')".\
+ format(tn=table_name, idf=id_column, cn=column_name))
+
+# C) Updates the newly inserted or pre-existing entry
+c.execute("UPDATE {tn} SET {cn}=('Hi World') WHERE {idf}=(123456)".\
+ format(tn=table_name, cn=column_name, idf=id_column))
+
+conn.commit()
+conn.close()
diff --git a/tutorials/sqlite3_howto/code/updating_rows.py b/tutorials/sqlite3_howto/code/updating_rows.py
new file mode 100644
index 0000000..5c4f762
--- /dev/null
+++ b/tutorials/sqlite3_howto/code/updating_rows.py
@@ -0,0 +1,50 @@
+# Sebastian Raschka, 2014
+# Updating rows in an existing SQLite database
+
+import sqlite3
+
+sqlite_file = ''
+table_name = ''
+column_name_1 = ''
+column_name_2 = ''
+column_name_3 = ''
+value_1 = 'hello world'
+value_2 = 12345
+
+conn = sqlite3.connect(sqlite_file)
+c = conn.cursor()
+
+
+# A.1) Updating all rows for a single column
+
+c.execute('UPDATE {dn} SET {cn1}={v1}'.\
+ format(dn=table_name, cn1=column_name_1, v1=value1)
+
+
+# A.2) Updating all rows for 2 columns (same for multiple columns)
+
+c.execute('UPDATE {dn} SET {cn1}={v1}, {cn2}={v2}'.\
+ format(dn=table_name, cn1=column_name_1, cn2=column_name_2,
+ v1=value1, v2=value2)
+
+
+
+
+# B.1) Updating specific rows that meet a certain criterion
+# here: update column_1 with value_1 if row has value_2 in column_2
+
+c.execute('UPDATE {dn} SET {cn1}={v1} WHERE {cn2}={v2}'.\
+ format(dn=table_name, cn1=column_name_1, v1=value1)
+
+
+# B.2) Updating specific rows that meet multiple criteria
+# here: update column_1 with value_1
+# if row has value_2 in column_2
+# and if row has value = 1 in column_3
+
+c.execute('UPDATE {dn} SET {cn1}={v1} WHERE {cn2}={v2} AND {cn3}=1'.\
+ format(dn=table_name, cn1=column_name_1, v1=value1, cn3=column_name_3)
+
+
+conn.commit()
+conn.close()
diff --git a/tutorials/sqlite3_howto/code/write_from_sqlite.py b/tutorials/sqlite3_howto/code/write_from_sqlite.py
new file mode 100644
index 0000000..f3f41a4
--- /dev/null
+++ b/tutorials/sqlite3_howto/code/write_from_sqlite.py
@@ -0,0 +1,102 @@
+import sqlite3
+
+def create_col_index(db_name, table_name, column_name, index_name):
+ '''
+ Creates a column index on a SQLite table.
+
+ Keyword arguments:
+ db_name (str): Path of the .sqlite database file.
+ table_name (str): Name of the target table in the SQLite file.
+ condition (str): Condition for querying the SQLite database table.
+ column_name (str): Name of the column for which the index is created.
+
+ '''
+
+ # Connecting to the database file
+ conn = sqlite3.connect(db_name)
+ c = conn.cursor()
+
+ # Creating the index
+ c.execute('CREATE INDEX {} ON {} ({})'.format(index_name, table_name, column_name))
+
+ # Save index and close the connection to the database
+ conn.commit()
+ conn.close()
+
+
+
+def drop_col_index(db_name, index_name):
+ '''
+ Drops a column index from a SQLite table.
+
+ Keyword arguments:
+ db_name (str): Path of the .sqlite database file.
+ table_name (str): Name of the target table in the SQLite file.
+ condition (str): Condition for querying the SQLite database table.
+ column_name (str): Name of the column for which the index is dropped.
+
+ '''
+
+ # Connecting to the database file
+ conn = sqlite3.connect(db_name)
+ c = conn.cursor()
+
+ # Drops the index
+ c.execute('DROP INDEX {}'.format(index_name))
+
+ # Save index and close the connection to the database
+ conn.commit()
+ conn.close()
+
+
+
+def write_from_query(db_name, table_name, condition, content_column, out_file, fetchmany=False):
+ '''
+ Writes contents from a SQLite database column to an output file
+
+ Keyword arguments:
+ db_name (str): Path of the .sqlite database file.
+ table_name (str): Name of the target table in the SQLite file.
+ condition (str): Condition for querying the SQLite database table.
+ content_column (str): Name of the column that contains the content for the output file.
+ out_file (str): Path of the output file that will be written.
+
+ '''
+ # Connecting to the database file
+ conn = sqlite3.connect(db_name)
+ c = conn.cursor()
+
+ # Querying the database and writing the output file
+
+
+ # A) using .fetchmany(); recommended for larger databases
+ if fetchmany:
+ c.execute('SELECT ({}) FROM {} WHERE {}'.format(content_column, table_name, condition))
+ with open(out_file, 'w') as outf:
+ results = c.fetchmany(fetchmany)
+ while results:
+ for row in results:
+ outf.write(row[0])
+ results = c.fetchmany(fetchmany)
+
+ # B) simple .execute() loop
+ else:
+ c.execute('SELECT ({}) FROM {} WHERE {}'.format(content_column, table_name, condition))
+ with open(out_file, 'w') as outf:
+ for row in c:
+ outf.write(row[0])
+
+ # Closing the connection to the database
+ conn.close()
+
+if __name__ == '__main__':
+ write_from_query(
+ db_name='my_db.sqlite',
+ table_name='my_table',
+ condition='variable1=1 AND variable2<=5 AND variable3="Zinc_Plus"',
+ content_column='variable4',
+ out_file='sqlite_out.txt'
+ )
+
+
+
diff --git a/tutorials/table_of_contents_ipython.ipynb b/tutorials/table_of_contents_ipython.ipynb
new file mode 100644
index 0000000..1245132
--- /dev/null
+++ b/tutorials/table_of_contents_ipython.ipynb
@@ -0,0 +1,281 @@
+{
+ "metadata": {
+ "name": "",
+ "signature": "sha256:34307c4f0973ebef511e97c036657231fc4e230e7627cfe073d89f4046f9ce9f"
+ },
+ "nbformat": 3,
+ "nbformat_minor": 0,
+ "worksheets": [
+ {
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[Sebastian Raschka](http://sebastianraschka.com) \n",
+ "last updated: 05/29/2014"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "I would be happy to hear your comments and suggestions. \n",
+ "Please feel free to drop me a note via\n",
+ "[twitter](https://twitter.com/rasbt), [email](mailto:bluewoodtree@gmail.com), or [google+](https://plus.google.com/118404394130788869227).\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ ""
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Creating a table of contents with internal links in IPython Notebooks and Markdown documents"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Many people have asked me how I create the table of contents with internal links for my IPython notebooks and Markdown documents on GitHub. \n",
+ "Well, no (IPython) magic is involved, it is just a little bit of HTML, but I thought it might be worthwhile to write this little how-to tutorial."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ ""
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
\n",
+ "For example, [click this link](#bottom) to jump to the bottom of the page.\n",
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "## The two components to create an internal link"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "So how does it work? Basically, all you need are those two components: \n",
+ "1. the destination\n",
+ "2. an internal hyperlink to the destination"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ ""
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "###1. The destination"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "To define the destination (i.e., the section on the page or the cell you want to jump to), you just need to insert an empty HTML anchor tag and give it an **`id`**, \n",
+ "e.g., **``** \n",
+ "\n",
+ "This anchor tag will be invisible if you render it as Markdown in the IPython notebook. \n",
+ "Note that it would also work if we use the **`name`** attribute instead of **`id`**, but since the **`name`** attribute is not supported by HTML5 anymore, I would suggest to just use the **`id`** attribute, which is also shorter to type."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "###2. The internal hyperlink"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Now we have to create the hyperlink to the **``** anchor tag that we just created. \n",
+ "We can either do this in ye goode olde HTML where we put a fragment identifier in form of a hash mark (`#`) in front of the name, \n",
+ "for example, **`Link to the destination'`**\n",
+ "\n",
+ "Or alternatively, we can just use the slightly more convenient Markdown syntax: \n",
+ "**`[Link to the destination](#the_destination)`**\n",
+ "\n",
+ "**That's all!**\n"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# One more piece of advice"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Of course it would make sense to place the empty anchor tags for you table of contents just on top of each cell that contains a heading. \n",
+ "E.g., "
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "`` \n",
+ "`###Section 2` \n",
+ "`some text ...` "
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "And I did this for a very long time ... until I figured out that it wouldn't render the Markdown properly if you convert the IPython Notebook into HTML (for example, for printing via the print preview option). \n",
+ "\n",
+ "But instead of "
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "###Section 2"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "it would be rendered as"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "`###Section 2`"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "which is certainly not what we want (note that it looks normal in the IPython notebook, but not in the converted HTML version). So my favorite remedy would be to put the `id`-anchor tag into a separate cell just above the section, ideally with some line breaks for nicer visuals.\n",
+ "\n",
+ "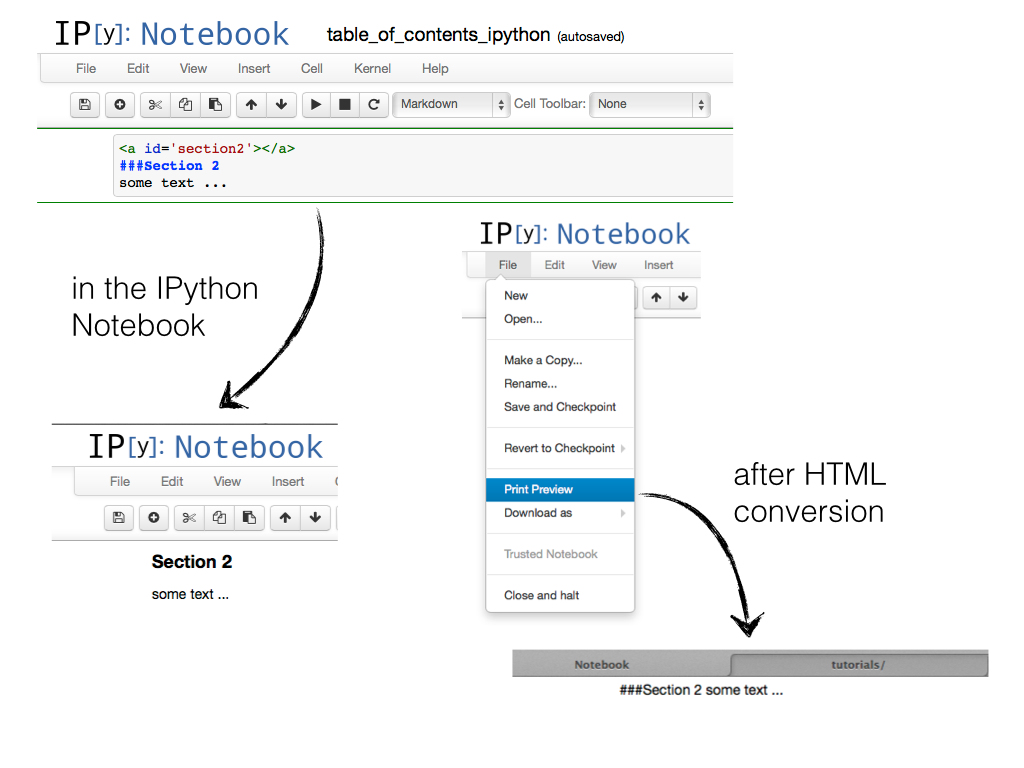\n",
+ "\n",
+ "### Solution 1: id-anchor tag in a separate cell\n",
+ "\n",
+ "\n",
+ "\n",
+ "
\n",
+ "
\n",
+ "\n",
+ "### Solution 2: line break between the id-anchor and text:\n",
+ "\n",
+ "\n",
+ "\n",
+ "(this alternative workaround was kindly submitted by [Ryan Morshead](https://github.com/rmorshea))\n",
+ "\n",
+ "
\n",
+ "
\n",
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Solution 3: using header cells"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Alternatively, and I think this is an even better solution, is to use header cells.\n",
+ "
\n",
+ "
\n",
+ ""
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "To define the hyperlink anchor tag to this \"header cell\" is just the text content of the \"header cell\" connected by dashes. E.g.,\n",
+ "\n",
+ "`[link to another section](#Another-section)`\n",
+ "
\n",
+ "
\n",
+ "
\n",
+ "
\n",
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[[Click this link and jump to the top of the page](#top)]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "You can't see it, but this cell contains a \n",
+ "`` \n",
+ "anchor tag just below this text.\n",
+ ""
+ ]
+ }
+ ],
+ "metadata": {}
+ }
+ ]
+}
\ No newline at end of file
diff --git a/tutorials/things_in_pandas.ipynb b/tutorials/things_in_pandas.ipynb
new file mode 100644
index 0000000..968d734
--- /dev/null
+++ b/tutorials/things_in_pandas.ipynb
@@ -0,0 +1,3201 @@
+{
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[Back to the GitHub repository](https://github.com/rasbt/python_reference)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 1,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [
+ {
+ "name": "stdout",
+ "output_type": "stream",
+ "text": [
+ "Sebastian Raschka 28/01/2015 \n",
+ "\n",
+ "CPython 3.4.2\n",
+ "IPython 2.3.1\n",
+ "\n",
+ "pandas 0.15.2\n"
+ ]
+ }
+ ],
+ "source": [
+ "%load_ext watermark\n",
+ "%watermark -a 'Sebastian Raschka' -v -d -p pandas"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[More information](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/ipython_magic/watermark.ipynb) about the `watermark` magic command extension."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Things in Pandas I Wish I'd Known Earlier"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "This is just a small but growing collection of pandas snippets that I find occasionally and particularly useful -- consider it as my personal notebook. Suggestions, tips, and contributions are very, very welcome!"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Sections"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "- [Loading Some Example Data](#Loading-Some-Example-Data)\n",
+ "- [Renaming Columns](#Renaming-Columns)\n",
+ " - [Converting Column Names to Lowercase](#Converting-Column-Names-to-Lowercase)\n",
+ " - [Renaming Particular Columns](#Renaming-Particular-Columns)\n",
+ "- [Applying Computations Rows-wise](#Applying-Computations-Rows-wise)\n",
+ " - [Changing Values in a Column](#Changing-Values-in-a-Column)\n",
+ " - [Adding a New Column](#Adding-a-New-Column)\n",
+ " - [Applying Functions to Multiple Columns](#Applying-Functions-to-Multiple-Columns)\n",
+ "- [Missing Values aka NaNs](#Missing-Values-aka-NaNs)\n",
+ " - [Counting Rows with NaNs](#Counting-Rows-with-NaNs)\n",
+ " - [Selecting NaN Rows](#Selecting-NaN-Rows)\n",
+ " - [Selecting non-NaN Rows](#Selecting-non-NaN-Rows)\n",
+ " - [Filling NaN Rows](#Filling-NaN-Rows)\n",
+ "- [Appending Rows to a DataFrame](#Appending-Rows-to-a-DataFrame)\n",
+ "- [Sorting and Reindexing DataFrames](#Sorting-and-Reindexing-DataFrames)\n",
+ "- [Updating Columns](#Updating-Columns)\n",
+ "- [Chaining Conditions - Using Bitwise Operators](#Chaining-Conditions---Using-Bitwise-Operators)\n",
+ "- [Column Types](#Column-Types)\n",
+ " - [Printing Column Types](#Printing-Column-Types)\n",
+ " - [Selecting by Column Type](#Selecting-by-Column-Type)\n",
+ " - [Converting Column Types](#Converting-Column-Types)\n",
+ "- [If-tests](#If-tests)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Loading Some Example Data"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[[back to section overview](#Sections)]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "I am heavily into sports prediction (via a machine learning approach) these days. So, let us use a (very) small subset of the soccer data that I am just working with."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 2,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [
+ {
+ "data": {
+ "text/html": [
+ "\n",
+ "\n",
+ " \n",
+ " \n",
+ " PLAYER \n",
+ " SALARY \n",
+ " GP \n",
+ " G \n",
+ " A \n",
+ " SOT \n",
+ " PPG \n",
+ " P \n",
+ " \n",
+ " \n",
+ " \n",
+ " \n",
+ " 0 \n",
+ " Sergio Agüero\\n Forward — Manchester City \n",
+ " $19.2m \n",
+ " 16 \n",
+ " 14 \n",
+ " 3 \n",
+ " 34 \n",
+ " 13.12 \n",
+ " 209.98 \n",
+ " \n",
+ " \n",
+ " 1 \n",
+ " Eden Hazard\\n Midfield — Chelsea \n",
+ " $18.9m \n",
+ " 21 \n",
+ " 8 \n",
+ " 4 \n",
+ " 17 \n",
+ " 13.05 \n",
+ " 274.04 \n",
+ " \n",
+ " \n",
+ " 2 \n",
+ " Alexis Sánchez\\n Forward — Arsenal \n",
+ " $17.6m \n",
+ " NaN \n",
+ " 12 \n",
+ " 7 \n",
+ " 29 \n",
+ " 11.19 \n",
+ " 223.86 \n",
+ " \n",
+ " \n",
+ " 3 \n",
+ " Yaya Touré\\n Midfield — Manchester City \n",
+ " $16.6m \n",
+ " 18 \n",
+ " 7 \n",
+ " 1 \n",
+ " 19 \n",
+ " 10.99 \n",
+ " 197.91 \n",
+ " \n",
+ " \n",
+ " 4 \n",
+ " Ángel Di María\\n Midfield — Manchester United \n",
+ " $15.0m \n",
+ " 13 \n",
+ " 3 \n",
+ " NaN \n",
+ " 13 \n",
+ " 10.17 \n",
+ " 132.23 \n",
+ " \n",
+ " \n",
+ " 5 \n",
+ " Santiago Cazorla\\n Midfield — Arsenal \n",
+ " $14.8m \n",
+ " 20 \n",
+ " 4 \n",
+ " NaN \n",
+ " 20 \n",
+ " 9.97 \n",
+ " NaN \n",
+ " \n",
+ " \n",
+ " 6 \n",
+ " David Silva\\n Midfield — Manchester City \n",
+ " $14.3m \n",
+ " 15 \n",
+ " 6 \n",
+ " 2 \n",
+ " 11 \n",
+ " 10.35 \n",
+ " 155.26 \n",
+ " \n",
+ " \n",
+ " 7 \n",
+ " Cesc Fàbregas\\n Midfield — Chelsea \n",
+ " $14.0m \n",
+ " 20 \n",
+ " 2 \n",
+ " 14 \n",
+ " 10 \n",
+ " 10.47 \n",
+ " 209.49 \n",
+ " \n",
+ " \n",
+ " 8 \n",
+ " Saido Berahino\\n Forward — West Brom \n",
+ " $13.8m \n",
+ " 21 \n",
+ " 9 \n",
+ " 0 \n",
+ " 20 \n",
+ " 7.02 \n",
+ " 147.43 \n",
+ " \n",
+ " \n",
+ " 9 \n",
+ " Steven Gerrard\\n Midfield — Liverpool \n",
+ " $13.8m \n",
+ " 20 \n",
+ " 5 \n",
+ " 1 \n",
+ " 11 \n",
+ " 7.50 \n",
+ " 150.01 \n",
+ " \n",
+ " \n",
+ "
\n",
+ ""
+ ],
+ "text/plain": [
+ " PLAYER SALARY GP G A SOT \\\n",
+ "0 Sergio Agüero\\n Forward — Manchester City $19.2m 16 14 3 34 \n",
+ "1 Eden Hazard\\n Midfield — Chelsea $18.9m 21 8 4 17 \n",
+ "2 Alexis Sánchez\\n Forward — Arsenal $17.6m NaN 12 7 29 \n",
+ "3 Yaya Touré\\n Midfield — Manchester City $16.6m 18 7 1 19 \n",
+ "4 Ángel Di María\\n Midfield — Manchester United $15.0m 13 3 NaN 13 \n",
+ "5 Santiago Cazorla\\n Midfield — Arsenal $14.8m 20 4 NaN 20 \n",
+ "6 David Silva\\n Midfield — Manchester City $14.3m 15 6 2 11 \n",
+ "7 Cesc Fàbregas\\n Midfield — Chelsea $14.0m 20 2 14 10 \n",
+ "8 Saido Berahino\\n Forward — West Brom $13.8m 21 9 0 20 \n",
+ "9 Steven Gerrard\\n Midfield — Liverpool $13.8m 20 5 1 11 \n",
+ "\n",
+ " PPG P \n",
+ "0 13.12 209.98 \n",
+ "1 13.05 274.04 \n",
+ "2 11.19 223.86 \n",
+ "3 10.99 197.91 \n",
+ "4 10.17 132.23 \n",
+ "5 9.97 NaN \n",
+ "6 10.35 155.26 \n",
+ "7 10.47 209.49 \n",
+ "8 7.02 147.43 \n",
+ "9 7.50 150.01 "
+ ]
+ },
+ "execution_count": 2,
+ "metadata": {},
+ "output_type": "execute_result"
+ }
+ ],
+ "source": [
+ "import pandas as pd\n",
+ "\n",
+ "df = pd.read_csv('https://raw.githubusercontent.com/rasbt/python_reference/master/Data/some_soccer_data.csv')\n",
+ "df"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Renaming Columns"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[[back to section overview](#Sections)]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Converting Column Names to Lowercase"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 3,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [
+ {
+ "data": {
+ "text/html": [
+ "\n",
+ "\n",
+ " \n",
+ " \n",
+ " player \n",
+ " salary \n",
+ " gp \n",
+ " g \n",
+ " a \n",
+ " sot \n",
+ " ppg \n",
+ " p \n",
+ " \n",
+ " \n",
+ " \n",
+ " \n",
+ " 7 \n",
+ " Cesc Fàbregas\\n Midfield — Chelsea \n",
+ " $14.0m \n",
+ " 20 \n",
+ " 2 \n",
+ " 14 \n",
+ " 10 \n",
+ " 10.47 \n",
+ " 209.49 \n",
+ " \n",
+ " \n",
+ " 8 \n",
+ " Saido Berahino\\n Forward — West Brom \n",
+ " $13.8m \n",
+ " 21 \n",
+ " 9 \n",
+ " 0 \n",
+ " 20 \n",
+ " 7.02 \n",
+ " 147.43 \n",
+ " \n",
+ " \n",
+ " 9 \n",
+ " Steven Gerrard\\n Midfield — Liverpool \n",
+ " $13.8m \n",
+ " 20 \n",
+ " 5 \n",
+ " 1 \n",
+ " 11 \n",
+ " 7.50 \n",
+ " 150.01 \n",
+ " \n",
+ " \n",
+ "
\n",
+ ""
+ ],
+ "text/plain": [
+ " player salary gp g a sot ppg \\\n",
+ "7 Cesc Fàbregas\\n Midfield — Chelsea $14.0m 20 2 14 10 10.47 \n",
+ "8 Saido Berahino\\n Forward — West Brom $13.8m 21 9 0 20 7.02 \n",
+ "9 Steven Gerrard\\n Midfield — Liverpool $13.8m 20 5 1 11 7.50 \n",
+ "\n",
+ " p \n",
+ "7 209.49 \n",
+ "8 147.43 \n",
+ "9 150.01 "
+ ]
+ },
+ "execution_count": 3,
+ "metadata": {},
+ "output_type": "execute_result"
+ }
+ ],
+ "source": [
+ "# Converting column names to lowercase\n",
+ "\n",
+ "df.columns = [c.lower() for c in df.columns]\n",
+ "\n",
+ "# or\n",
+ "# df.rename(columns=lambda x : x.lower())\n",
+ "\n",
+ "df.tail(3)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Renaming Particular Columns"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 4,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [
+ {
+ "data": {
+ "text/html": [
+ "\n",
+ "\n",
+ " \n",
+ " \n",
+ " player \n",
+ " salary \n",
+ " games \n",
+ " goals \n",
+ " assists \n",
+ " shots_on_target \n",
+ " points_per_game \n",
+ " points \n",
+ " \n",
+ " \n",
+ " \n",
+ " \n",
+ " 7 \n",
+ " Cesc Fàbregas\\n Midfield — Chelsea \n",
+ " $14.0m \n",
+ " 20 \n",
+ " 2 \n",
+ " 14 \n",
+ " 10 \n",
+ " 10.47 \n",
+ " 209.49 \n",
+ " \n",
+ " \n",
+ " 8 \n",
+ " Saido Berahino\\n Forward — West Brom \n",
+ " $13.8m \n",
+ " 21 \n",
+ " 9 \n",
+ " 0 \n",
+ " 20 \n",
+ " 7.02 \n",
+ " 147.43 \n",
+ " \n",
+ " \n",
+ " 9 \n",
+ " Steven Gerrard\\n Midfield — Liverpool \n",
+ " $13.8m \n",
+ " 20 \n",
+ " 5 \n",
+ " 1 \n",
+ " 11 \n",
+ " 7.50 \n",
+ " 150.01 \n",
+ " \n",
+ " \n",
+ "
\n",
+ ""
+ ],
+ "text/plain": [
+ " player salary games goals assists \\\n",
+ "7 Cesc Fàbregas\\n Midfield — Chelsea $14.0m 20 2 14 \n",
+ "8 Saido Berahino\\n Forward — West Brom $13.8m 21 9 0 \n",
+ "9 Steven Gerrard\\n Midfield — Liverpool $13.8m 20 5 1 \n",
+ "\n",
+ " shots_on_target points_per_game points \n",
+ "7 10 10.47 209.49 \n",
+ "8 20 7.02 147.43 \n",
+ "9 11 7.50 150.01 "
+ ]
+ },
+ "execution_count": 4,
+ "metadata": {},
+ "output_type": "execute_result"
+ }
+ ],
+ "source": [
+ "df = df.rename(columns={'p': 'points', \n",
+ " 'gp': 'games',\n",
+ " 'sot': 'shots_on_target',\n",
+ " 'g': 'goals',\n",
+ " 'ppg': 'points_per_game',\n",
+ " 'a': 'assists',})\n",
+ "\n",
+ "df.tail(3)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Applying Computations Rows-wise"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[[back to section overview](#Sections)]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Changing Values in a Column"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 5,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [
+ {
+ "data": {
+ "text/html": [
+ "\n",
+ "\n",
+ " \n",
+ " \n",
+ " player \n",
+ " salary \n",
+ " games \n",
+ " goals \n",
+ " assists \n",
+ " shots_on_target \n",
+ " points_per_game \n",
+ " points \n",
+ " \n",
+ " \n",
+ " \n",
+ " \n",
+ " 5 \n",
+ " Santiago Cazorla\\n Midfield — Arsenal \n",
+ " 14.8 \n",
+ " 20 \n",
+ " 4 \n",
+ " NaN \n",
+ " 20 \n",
+ " 9.97 \n",
+ " NaN \n",
+ " \n",
+ " \n",
+ " 6 \n",
+ " David Silva\\n Midfield — Manchester City \n",
+ " 14.3 \n",
+ " 15 \n",
+ " 6 \n",
+ " 2 \n",
+ " 11 \n",
+ " 10.35 \n",
+ " 155.26 \n",
+ " \n",
+ " \n",
+ " 7 \n",
+ " Cesc Fàbregas\\n Midfield — Chelsea \n",
+ " 14.0 \n",
+ " 20 \n",
+ " 2 \n",
+ " 14 \n",
+ " 10 \n",
+ " 10.47 \n",
+ " 209.49 \n",
+ " \n",
+ " \n",
+ " 8 \n",
+ " Saido Berahino\\n Forward — West Brom \n",
+ " 13.8 \n",
+ " 21 \n",
+ " 9 \n",
+ " 0 \n",
+ " 20 \n",
+ " 7.02 \n",
+ " 147.43 \n",
+ " \n",
+ " \n",
+ " 9 \n",
+ " Steven Gerrard\\n Midfield — Liverpool \n",
+ " 13.8 \n",
+ " 20 \n",
+ " 5 \n",
+ " 1 \n",
+ " 11 \n",
+ " 7.50 \n",
+ " 150.01 \n",
+ " \n",
+ " \n",
+ "
\n",
+ ""
+ ],
+ "text/plain": [
+ " player salary games goals assists \\\n",
+ "5 Santiago Cazorla\\n Midfield — Arsenal 14.8 20 4 NaN \n",
+ "6 David Silva\\n Midfield — Manchester City 14.3 15 6 2 \n",
+ "7 Cesc Fàbregas\\n Midfield — Chelsea 14.0 20 2 14 \n",
+ "8 Saido Berahino\\n Forward — West Brom 13.8 21 9 0 \n",
+ "9 Steven Gerrard\\n Midfield — Liverpool 13.8 20 5 1 \n",
+ "\n",
+ " shots_on_target points_per_game points \n",
+ "5 20 9.97 NaN \n",
+ "6 11 10.35 155.26 \n",
+ "7 10 10.47 209.49 \n",
+ "8 20 7.02 147.43 \n",
+ "9 11 7.50 150.01 "
+ ]
+ },
+ "execution_count": 5,
+ "metadata": {},
+ "output_type": "execute_result"
+ }
+ ],
+ "source": [
+ "# Processing `salary` column\n",
+ "\n",
+ "df['salary'] = df['salary'].apply(lambda x: x.strip('$m'))\n",
+ "df.tail()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Adding a New Column"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 6,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [
+ {
+ "data": {
+ "text/html": [
+ "\n",
+ "\n",
+ " \n",
+ " \n",
+ " player \n",
+ " salary \n",
+ " games \n",
+ " goals \n",
+ " assists \n",
+ " shots_on_target \n",
+ " points_per_game \n",
+ " points \n",
+ " position \n",
+ " team \n",
+ " \n",
+ " \n",
+ " \n",
+ " \n",
+ " 7 \n",
+ " Cesc Fàbregas\\n Midfield — Chelsea \n",
+ " 14.0 \n",
+ " 20 \n",
+ " 2 \n",
+ " 14 \n",
+ " 10 \n",
+ " 10.47 \n",
+ " 209.49 \n",
+ " \n",
+ " \n",
+ " \n",
+ " \n",
+ " 8 \n",
+ " Saido Berahino\\n Forward — West Brom \n",
+ " 13.8 \n",
+ " 21 \n",
+ " 9 \n",
+ " 0 \n",
+ " 20 \n",
+ " 7.02 \n",
+ " 147.43 \n",
+ " \n",
+ " \n",
+ " \n",
+ " \n",
+ " 9 \n",
+ " Steven Gerrard\\n Midfield — Liverpool \n",
+ " 13.8 \n",
+ " 20 \n",
+ " 5 \n",
+ " 1 \n",
+ " 11 \n",
+ " 7.50 \n",
+ " 150.01 \n",
+ " \n",
+ " \n",
+ " \n",
+ " \n",
+ "
\n",
+ ""
+ ],
+ "text/plain": [
+ " player salary games goals assists \\\n",
+ "7 Cesc Fàbregas\\n Midfield — Chelsea 14.0 20 2 14 \n",
+ "8 Saido Berahino\\n Forward — West Brom 13.8 21 9 0 \n",
+ "9 Steven Gerrard\\n Midfield — Liverpool 13.8 20 5 1 \n",
+ "\n",
+ " shots_on_target points_per_game points position team \n",
+ "7 10 10.47 209.49 \n",
+ "8 20 7.02 147.43 \n",
+ "9 11 7.50 150.01 "
+ ]
+ },
+ "execution_count": 6,
+ "metadata": {},
+ "output_type": "execute_result"
+ }
+ ],
+ "source": [
+ "df['team'] = pd.Series('', index=df.index)\n",
+ "\n",
+ "# or\n",
+ "df.insert(loc=8, column='position', value='') \n",
+ "\n",
+ "df.tail(3)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 7,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [
+ {
+ "data": {
+ "text/html": [
+ "\n",
+ "\n",
+ " \n",
+ " \n",
+ " player \n",
+ " salary \n",
+ " games \n",
+ " goals \n",
+ " assists \n",
+ " shots_on_target \n",
+ " points_per_game \n",
+ " points \n",
+ " position \n",
+ " team \n",
+ " \n",
+ " \n",
+ " \n",
+ " \n",
+ " 7 \n",
+ " Cesc Fàbregas \n",
+ " 14.0 \n",
+ " 20 \n",
+ " 2 \n",
+ " 14 \n",
+ " 10 \n",
+ " 10.47 \n",
+ " 209.49 \n",
+ " Midfield \n",
+ " Chelsea \n",
+ " \n",
+ " \n",
+ " 8 \n",
+ " Saido Berahino \n",
+ " 13.8 \n",
+ " 21 \n",
+ " 9 \n",
+ " 0 \n",
+ " 20 \n",
+ " 7.02 \n",
+ " 147.43 \n",
+ " Forward \n",
+ " West Brom \n",
+ " \n",
+ " \n",
+ " 9 \n",
+ " Steven Gerrard \n",
+ " 13.8 \n",
+ " 20 \n",
+ " 5 \n",
+ " 1 \n",
+ " 11 \n",
+ " 7.50 \n",
+ " 150.01 \n",
+ " Midfield \n",
+ " Liverpool \n",
+ " \n",
+ " \n",
+ "
\n",
+ ""
+ ],
+ "text/plain": [
+ " player salary games goals assists shots_on_target \\\n",
+ "7 Cesc Fàbregas 14.0 20 2 14 10 \n",
+ "8 Saido Berahino 13.8 21 9 0 20 \n",
+ "9 Steven Gerrard 13.8 20 5 1 11 \n",
+ "\n",
+ " points_per_game points position team \n",
+ "7 10.47 209.49 Midfield Chelsea \n",
+ "8 7.02 147.43 Forward West Brom \n",
+ "9 7.50 150.01 Midfield Liverpool "
+ ]
+ },
+ "execution_count": 7,
+ "metadata": {},
+ "output_type": "execute_result"
+ }
+ ],
+ "source": [
+ "# Processing `player` column\n",
+ "\n",
+ "def process_player_col(text):\n",
+ " name, rest = text.split('\\n')\n",
+ " position, team = [x.strip() for x in rest.split(' — ')]\n",
+ " return pd.Series([name, team, position])\n",
+ "\n",
+ "df[['player', 'team', 'position']] = df.player.apply(process_player_col)\n",
+ "\n",
+ "# modified after tip from reddit.com/user/hharison\n",
+ "#\n",
+ "# Alternative (inferior) approach:\n",
+ "#\n",
+ "#for idx,row in df.iterrows():\n",
+ "# name, position, team = process_player_col(row['player'])\n",
+ "# df.ix[idx, 'player'], df.ix[idx, 'position'], df.ix[idx, 'team'] = name, position, team\n",
+ " \n",
+ "df.tail(3)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Applying Functions to Multiple Columns"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 8,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [
+ {
+ "data": {
+ "text/html": [
+ "\n",
+ "\n",
+ " \n",
+ " \n",
+ " player \n",
+ " salary \n",
+ " games \n",
+ " goals \n",
+ " assists \n",
+ " shots_on_target \n",
+ " points_per_game \n",
+ " points \n",
+ " position \n",
+ " team \n",
+ " \n",
+ " \n",
+ " \n",
+ " \n",
+ " 0 \n",
+ " sergio agüero \n",
+ " 19.2 \n",
+ " 16 \n",
+ " 14 \n",
+ " 3 \n",
+ " 34 \n",
+ " 13.12 \n",
+ " 209.98 \n",
+ " forward \n",
+ " manchester city \n",
+ " \n",
+ " \n",
+ " 1 \n",
+ " eden hazard \n",
+ " 18.9 \n",
+ " 21 \n",
+ " 8 \n",
+ " 4 \n",
+ " 17 \n",
+ " 13.05 \n",
+ " 274.04 \n",
+ " midfield \n",
+ " chelsea \n",
+ " \n",
+ " \n",
+ " 2 \n",
+ " alexis sánchez \n",
+ " 17.6 \n",
+ " NaN \n",
+ " 12 \n",
+ " 7 \n",
+ " 29 \n",
+ " 11.19 \n",
+ " 223.86 \n",
+ " forward \n",
+ " arsenal \n",
+ " \n",
+ " \n",
+ " 3 \n",
+ " yaya touré \n",
+ " 16.6 \n",
+ " 18 \n",
+ " 7 \n",
+ " 1 \n",
+ " 19 \n",
+ " 10.99 \n",
+ " 197.91 \n",
+ " midfield \n",
+ " manchester city \n",
+ " \n",
+ " \n",
+ " 4 \n",
+ " ángel di maría \n",
+ " 15.0 \n",
+ " 13 \n",
+ " 3 \n",
+ " NaN \n",
+ " 13 \n",
+ " 10.17 \n",
+ " 132.23 \n",
+ " midfield \n",
+ " manchester united \n",
+ " \n",
+ " \n",
+ "
\n",
+ ""
+ ],
+ "text/plain": [
+ " player salary games goals assists shots_on_target \\\n",
+ "0 sergio agüero 19.2 16 14 3 34 \n",
+ "1 eden hazard 18.9 21 8 4 17 \n",
+ "2 alexis sánchez 17.6 NaN 12 7 29 \n",
+ "3 yaya touré 16.6 18 7 1 19 \n",
+ "4 ángel di maría 15.0 13 3 NaN 13 \n",
+ "\n",
+ " points_per_game points position team \n",
+ "0 13.12 209.98 forward manchester city \n",
+ "1 13.05 274.04 midfield chelsea \n",
+ "2 11.19 223.86 forward arsenal \n",
+ "3 10.99 197.91 midfield manchester city \n",
+ "4 10.17 132.23 midfield manchester united "
+ ]
+ },
+ "execution_count": 8,
+ "metadata": {},
+ "output_type": "execute_result"
+ }
+ ],
+ "source": [
+ "cols = ['player', 'position', 'team']\n",
+ "df[cols] = df[cols].applymap(lambda x: x.lower())\n",
+ "df.head()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Missing Values aka NaNs"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[[back to section overview](#Sections)]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Counting Rows with NaNs"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 9,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [
+ {
+ "name": "stdout",
+ "output_type": "stream",
+ "text": [
+ "3 rows have missing values\n"
+ ]
+ }
+ ],
+ "source": [
+ "nans = df.shape[0] - df.dropna().shape[0]\n",
+ "\n",
+ "print('%d rows have missing values' % nans)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Selecting NaN Rows"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 10,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [
+ {
+ "data": {
+ "text/html": [
+ "\n",
+ "\n",
+ " \n",
+ " \n",
+ " player \n",
+ " salary \n",
+ " games \n",
+ " goals \n",
+ " assists \n",
+ " shots_on_target \n",
+ " points_per_game \n",
+ " points \n",
+ " position \n",
+ " team \n",
+ " \n",
+ " \n",
+ " \n",
+ " \n",
+ " 4 \n",
+ " ángel di maría \n",
+ " 15.0 \n",
+ " 13 \n",
+ " 3 \n",
+ " NaN \n",
+ " 13 \n",
+ " 10.17 \n",
+ " 132.23 \n",
+ " midfield \n",
+ " manchester united \n",
+ " \n",
+ " \n",
+ " 5 \n",
+ " santiago cazorla \n",
+ " 14.8 \n",
+ " 20 \n",
+ " 4 \n",
+ " NaN \n",
+ " 20 \n",
+ " 9.97 \n",
+ " NaN \n",
+ " midfield \n",
+ " arsenal \n",
+ " \n",
+ " \n",
+ "
\n",
+ ""
+ ],
+ "text/plain": [
+ " player salary games goals assists shots_on_target \\\n",
+ "4 ángel di maría 15.0 13 3 NaN 13 \n",
+ "5 santiago cazorla 14.8 20 4 NaN 20 \n",
+ "\n",
+ " points_per_game points position team \n",
+ "4 10.17 132.23 midfield manchester united \n",
+ "5 9.97 NaN midfield arsenal "
+ ]
+ },
+ "execution_count": 10,
+ "metadata": {},
+ "output_type": "execute_result"
+ }
+ ],
+ "source": [
+ "# Selecting all rows that have NaNs in the `assists` column\n",
+ "\n",
+ "df[df['assists'].isnull()]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Selecting non-NaN Rows"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 11,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [
+ {
+ "data": {
+ "text/html": [
+ "\n",
+ "\n",
+ " \n",
+ " \n",
+ " player \n",
+ " salary \n",
+ " games \n",
+ " goals \n",
+ " assists \n",
+ " shots_on_target \n",
+ " points_per_game \n",
+ " points \n",
+ " position \n",
+ " team \n",
+ " \n",
+ " \n",
+ " \n",
+ " \n",
+ " 0 \n",
+ " sergio agüero \n",
+ " 19.2 \n",
+ " 16 \n",
+ " 14 \n",
+ " 3 \n",
+ " 34 \n",
+ " 13.12 \n",
+ " 209.98 \n",
+ " forward \n",
+ " manchester city \n",
+ " \n",
+ " \n",
+ " 1 \n",
+ " eden hazard \n",
+ " 18.9 \n",
+ " 21 \n",
+ " 8 \n",
+ " 4 \n",
+ " 17 \n",
+ " 13.05 \n",
+ " 274.04 \n",
+ " midfield \n",
+ " chelsea \n",
+ " \n",
+ " \n",
+ " 2 \n",
+ " alexis sánchez \n",
+ " 17.6 \n",
+ " NaN \n",
+ " 12 \n",
+ " 7 \n",
+ " 29 \n",
+ " 11.19 \n",
+ " 223.86 \n",
+ " forward \n",
+ " arsenal \n",
+ " \n",
+ " \n",
+ " 3 \n",
+ " yaya touré \n",
+ " 16.6 \n",
+ " 18 \n",
+ " 7 \n",
+ " 1 \n",
+ " 19 \n",
+ " 10.99 \n",
+ " 197.91 \n",
+ " midfield \n",
+ " manchester city \n",
+ " \n",
+ " \n",
+ " 6 \n",
+ " david silva \n",
+ " 14.3 \n",
+ " 15 \n",
+ " 6 \n",
+ " 2 \n",
+ " 11 \n",
+ " 10.35 \n",
+ " 155.26 \n",
+ " midfield \n",
+ " manchester city \n",
+ " \n",
+ " \n",
+ " 7 \n",
+ " cesc fàbregas \n",
+ " 14.0 \n",
+ " 20 \n",
+ " 2 \n",
+ " 14 \n",
+ " 10 \n",
+ " 10.47 \n",
+ " 209.49 \n",
+ " midfield \n",
+ " chelsea \n",
+ " \n",
+ " \n",
+ " 8 \n",
+ " saido berahino \n",
+ " 13.8 \n",
+ " 21 \n",
+ " 9 \n",
+ " 0 \n",
+ " 20 \n",
+ " 7.02 \n",
+ " 147.43 \n",
+ " forward \n",
+ " west brom \n",
+ " \n",
+ " \n",
+ " 9 \n",
+ " steven gerrard \n",
+ " 13.8 \n",
+ " 20 \n",
+ " 5 \n",
+ " 1 \n",
+ " 11 \n",
+ " 7.50 \n",
+ " 150.01 \n",
+ " midfield \n",
+ " liverpool \n",
+ " \n",
+ " \n",
+ "
\n",
+ ""
+ ],
+ "text/plain": [
+ " player salary games goals assists shots_on_target \\\n",
+ "0 sergio agüero 19.2 16 14 3 34 \n",
+ "1 eden hazard 18.9 21 8 4 17 \n",
+ "2 alexis sánchez 17.6 NaN 12 7 29 \n",
+ "3 yaya touré 16.6 18 7 1 19 \n",
+ "6 david silva 14.3 15 6 2 11 \n",
+ "7 cesc fàbregas 14.0 20 2 14 10 \n",
+ "8 saido berahino 13.8 21 9 0 20 \n",
+ "9 steven gerrard 13.8 20 5 1 11 \n",
+ "\n",
+ " points_per_game points position team \n",
+ "0 13.12 209.98 forward manchester city \n",
+ "1 13.05 274.04 midfield chelsea \n",
+ "2 11.19 223.86 forward arsenal \n",
+ "3 10.99 197.91 midfield manchester city \n",
+ "6 10.35 155.26 midfield manchester city \n",
+ "7 10.47 209.49 midfield chelsea \n",
+ "8 7.02 147.43 forward west brom \n",
+ "9 7.50 150.01 midfield liverpool "
+ ]
+ },
+ "execution_count": 11,
+ "metadata": {},
+ "output_type": "execute_result"
+ }
+ ],
+ "source": [
+ "df[df['assists'].notnull()]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Filling NaN Rows"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 12,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [
+ {
+ "data": {
+ "text/html": [
+ "\n",
+ "\n",
+ " \n",
+ " \n",
+ " player \n",
+ " salary \n",
+ " games \n",
+ " goals \n",
+ " assists \n",
+ " shots_on_target \n",
+ " points_per_game \n",
+ " points \n",
+ " position \n",
+ " team \n",
+ " \n",
+ " \n",
+ " \n",
+ " \n",
+ " 0 \n",
+ " sergio agüero \n",
+ " 19.2 \n",
+ " 16 \n",
+ " 14 \n",
+ " 3 \n",
+ " 34 \n",
+ " 13.12 \n",
+ " 209.98 \n",
+ " forward \n",
+ " manchester city \n",
+ " \n",
+ " \n",
+ " 1 \n",
+ " eden hazard \n",
+ " 18.9 \n",
+ " 21 \n",
+ " 8 \n",
+ " 4 \n",
+ " 17 \n",
+ " 13.05 \n",
+ " 274.04 \n",
+ " midfield \n",
+ " chelsea \n",
+ " \n",
+ " \n",
+ " 2 \n",
+ " alexis sánchez \n",
+ " 17.6 \n",
+ " 0 \n",
+ " 12 \n",
+ " 7 \n",
+ " 29 \n",
+ " 11.19 \n",
+ " 223.86 \n",
+ " forward \n",
+ " arsenal \n",
+ " \n",
+ " \n",
+ " 3 \n",
+ " yaya touré \n",
+ " 16.6 \n",
+ " 18 \n",
+ " 7 \n",
+ " 1 \n",
+ " 19 \n",
+ " 10.99 \n",
+ " 197.91 \n",
+ " midfield \n",
+ " manchester city \n",
+ " \n",
+ " \n",
+ " 4 \n",
+ " ángel di maría \n",
+ " 15.0 \n",
+ " 13 \n",
+ " 3 \n",
+ " 0 \n",
+ " 13 \n",
+ " 10.17 \n",
+ " 132.23 \n",
+ " midfield \n",
+ " manchester united \n",
+ " \n",
+ " \n",
+ " 5 \n",
+ " santiago cazorla \n",
+ " 14.8 \n",
+ " 20 \n",
+ " 4 \n",
+ " 0 \n",
+ " 20 \n",
+ " 9.97 \n",
+ " 0.00 \n",
+ " midfield \n",
+ " arsenal \n",
+ " \n",
+ " \n",
+ " 6 \n",
+ " david silva \n",
+ " 14.3 \n",
+ " 15 \n",
+ " 6 \n",
+ " 2 \n",
+ " 11 \n",
+ " 10.35 \n",
+ " 155.26 \n",
+ " midfield \n",
+ " manchester city \n",
+ " \n",
+ " \n",
+ " 7 \n",
+ " cesc fàbregas \n",
+ " 14.0 \n",
+ " 20 \n",
+ " 2 \n",
+ " 14 \n",
+ " 10 \n",
+ " 10.47 \n",
+ " 209.49 \n",
+ " midfield \n",
+ " chelsea \n",
+ " \n",
+ " \n",
+ " 8 \n",
+ " saido berahino \n",
+ " 13.8 \n",
+ " 21 \n",
+ " 9 \n",
+ " 0 \n",
+ " 20 \n",
+ " 7.02 \n",
+ " 147.43 \n",
+ " forward \n",
+ " west brom \n",
+ " \n",
+ " \n",
+ " 9 \n",
+ " steven gerrard \n",
+ " 13.8 \n",
+ " 20 \n",
+ " 5 \n",
+ " 1 \n",
+ " 11 \n",
+ " 7.50 \n",
+ " 150.01 \n",
+ " midfield \n",
+ " liverpool \n",
+ " \n",
+ " \n",
+ "
\n",
+ ""
+ ],
+ "text/plain": [
+ " player salary games goals assists shots_on_target \\\n",
+ "0 sergio agüero 19.2 16 14 3 34 \n",
+ "1 eden hazard 18.9 21 8 4 17 \n",
+ "2 alexis sánchez 17.6 0 12 7 29 \n",
+ "3 yaya touré 16.6 18 7 1 19 \n",
+ "4 ángel di maría 15.0 13 3 0 13 \n",
+ "5 santiago cazorla 14.8 20 4 0 20 \n",
+ "6 david silva 14.3 15 6 2 11 \n",
+ "7 cesc fàbregas 14.0 20 2 14 10 \n",
+ "8 saido berahino 13.8 21 9 0 20 \n",
+ "9 steven gerrard 13.8 20 5 1 11 \n",
+ "\n",
+ " points_per_game points position team \n",
+ "0 13.12 209.98 forward manchester city \n",
+ "1 13.05 274.04 midfield chelsea \n",
+ "2 11.19 223.86 forward arsenal \n",
+ "3 10.99 197.91 midfield manchester city \n",
+ "4 10.17 132.23 midfield manchester united \n",
+ "5 9.97 0.00 midfield arsenal \n",
+ "6 10.35 155.26 midfield manchester city \n",
+ "7 10.47 209.49 midfield chelsea \n",
+ "8 7.02 147.43 forward west brom \n",
+ "9 7.50 150.01 midfield liverpool "
+ ]
+ },
+ "execution_count": 12,
+ "metadata": {},
+ "output_type": "execute_result"
+ }
+ ],
+ "source": [
+ "# Filling NaN cells with default value 0\n",
+ "\n",
+ "df.fillna(value=0, inplace=True)\n",
+ "df"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Appending Rows to a DataFrame"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[[back to section overview](#Sections)]"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 13,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [
+ {
+ "data": {
+ "text/html": [
+ "\n",
+ "\n",
+ " \n",
+ " \n",
+ " player \n",
+ " salary \n",
+ " games \n",
+ " goals \n",
+ " assists \n",
+ " shots_on_target \n",
+ " points_per_game \n",
+ " points \n",
+ " position \n",
+ " team \n",
+ " \n",
+ " \n",
+ " \n",
+ " \n",
+ " 8 \n",
+ " saido berahino \n",
+ " 13.8 \n",
+ " 21 \n",
+ " 9 \n",
+ " 0 \n",
+ " 20 \n",
+ " 7.02 \n",
+ " 147.43 \n",
+ " forward \n",
+ " west brom \n",
+ " \n",
+ " \n",
+ " 9 \n",
+ " steven gerrard \n",
+ " 13.8 \n",
+ " 20 \n",
+ " 5 \n",
+ " 1 \n",
+ " 11 \n",
+ " 7.50 \n",
+ " 150.01 \n",
+ " midfield \n",
+ " liverpool \n",
+ " \n",
+ " \n",
+ " 10 \n",
+ " NaN \n",
+ " NaN \n",
+ " NaN \n",
+ " NaN \n",
+ " NaN \n",
+ " NaN \n",
+ " NaN \n",
+ " NaN \n",
+ " NaN \n",
+ " NaN \n",
+ " \n",
+ " \n",
+ "
\n",
+ ""
+ ],
+ "text/plain": [
+ " player salary games goals assists shots_on_target \\\n",
+ "8 saido berahino 13.8 21 9 0 20 \n",
+ "9 steven gerrard 13.8 20 5 1 11 \n",
+ "10 NaN NaN NaN NaN NaN NaN \n",
+ "\n",
+ " points_per_game points position team \n",
+ "8 7.02 147.43 forward west brom \n",
+ "9 7.50 150.01 midfield liverpool \n",
+ "10 NaN NaN NaN NaN "
+ ]
+ },
+ "execution_count": 13,
+ "metadata": {},
+ "output_type": "execute_result"
+ }
+ ],
+ "source": [
+ "# Adding an \"empty\" row to the DataFrame\n",
+ "\n",
+ "import numpy as np\n",
+ "\n",
+ "df = df.append(pd.Series(\n",
+ " [np.nan]*len(df.columns), # Fill cells with NaNs\n",
+ " index=df.columns), \n",
+ " ignore_index=True)\n",
+ "\n",
+ "df.tail(3)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 14,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [
+ {
+ "data": {
+ "text/html": [
+ "\n",
+ "\n",
+ " \n",
+ " \n",
+ " player \n",
+ " salary \n",
+ " games \n",
+ " goals \n",
+ " assists \n",
+ " shots_on_target \n",
+ " points_per_game \n",
+ " points \n",
+ " position \n",
+ " team \n",
+ " \n",
+ " \n",
+ " \n",
+ " \n",
+ " 8 \n",
+ " saido berahino \n",
+ " 13.8 \n",
+ " 21 \n",
+ " 9 \n",
+ " 0 \n",
+ " 20 \n",
+ " 7.02 \n",
+ " 147.43 \n",
+ " forward \n",
+ " west brom \n",
+ " \n",
+ " \n",
+ " 9 \n",
+ " steven gerrard \n",
+ " 13.8 \n",
+ " 20 \n",
+ " 5 \n",
+ " 1 \n",
+ " 11 \n",
+ " 7.50 \n",
+ " 150.01 \n",
+ " midfield \n",
+ " liverpool \n",
+ " \n",
+ " \n",
+ " 10 \n",
+ " new player \n",
+ " 12.3 \n",
+ " NaN \n",
+ " NaN \n",
+ " NaN \n",
+ " NaN \n",
+ " NaN \n",
+ " NaN \n",
+ " NaN \n",
+ " NaN \n",
+ " \n",
+ " \n",
+ "
\n",
+ ""
+ ],
+ "text/plain": [
+ " player salary games goals assists shots_on_target \\\n",
+ "8 saido berahino 13.8 21 9 0 20 \n",
+ "9 steven gerrard 13.8 20 5 1 11 \n",
+ "10 new player 12.3 NaN NaN NaN NaN \n",
+ "\n",
+ " points_per_game points position team \n",
+ "8 7.02 147.43 forward west brom \n",
+ "9 7.50 150.01 midfield liverpool \n",
+ "10 NaN NaN NaN NaN "
+ ]
+ },
+ "execution_count": 14,
+ "metadata": {},
+ "output_type": "execute_result"
+ }
+ ],
+ "source": [
+ "# Filling cells with data\n",
+ "\n",
+ "df.loc[df.index[-1], 'player'] = 'new player'\n",
+ "df.loc[df.index[-1], 'salary'] = 12.3\n",
+ "df.tail(3)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Sorting and Reindexing DataFrames"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[[back to section overview](#Sections)]"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 15,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [
+ {
+ "data": {
+ "text/html": [
+ "\n",
+ "\n",
+ " \n",
+ " \n",
+ " player \n",
+ " salary \n",
+ " games \n",
+ " goals \n",
+ " assists \n",
+ " shots_on_target \n",
+ " points_per_game \n",
+ " points \n",
+ " position \n",
+ " team \n",
+ " \n",
+ " \n",
+ " \n",
+ " \n",
+ " 0 \n",
+ " sergio agüero \n",
+ " 19.2 \n",
+ " 16 \n",
+ " 14 \n",
+ " 3 \n",
+ " 34 \n",
+ " 13.12 \n",
+ " 209.98 \n",
+ " forward \n",
+ " manchester city \n",
+ " \n",
+ " \n",
+ " 2 \n",
+ " alexis sánchez \n",
+ " 17.6 \n",
+ " 0 \n",
+ " 12 \n",
+ " 7 \n",
+ " 29 \n",
+ " 11.19 \n",
+ " 223.86 \n",
+ " forward \n",
+ " arsenal \n",
+ " \n",
+ " \n",
+ " 8 \n",
+ " saido berahino \n",
+ " 13.8 \n",
+ " 21 \n",
+ " 9 \n",
+ " 0 \n",
+ " 20 \n",
+ " 7.02 \n",
+ " 147.43 \n",
+ " forward \n",
+ " west brom \n",
+ " \n",
+ " \n",
+ " 1 \n",
+ " eden hazard \n",
+ " 18.9 \n",
+ " 21 \n",
+ " 8 \n",
+ " 4 \n",
+ " 17 \n",
+ " 13.05 \n",
+ " 274.04 \n",
+ " midfield \n",
+ " chelsea \n",
+ " \n",
+ " \n",
+ " 3 \n",
+ " yaya touré \n",
+ " 16.6 \n",
+ " 18 \n",
+ " 7 \n",
+ " 1 \n",
+ " 19 \n",
+ " 10.99 \n",
+ " 197.91 \n",
+ " midfield \n",
+ " manchester city \n",
+ " \n",
+ " \n",
+ "
\n",
+ ""
+ ],
+ "text/plain": [
+ " player salary games goals assists shots_on_target \\\n",
+ "0 sergio agüero 19.2 16 14 3 34 \n",
+ "2 alexis sánchez 17.6 0 12 7 29 \n",
+ "8 saido berahino 13.8 21 9 0 20 \n",
+ "1 eden hazard 18.9 21 8 4 17 \n",
+ "3 yaya touré 16.6 18 7 1 19 \n",
+ "\n",
+ " points_per_game points position team \n",
+ "0 13.12 209.98 forward manchester city \n",
+ "2 11.19 223.86 forward arsenal \n",
+ "8 7.02 147.43 forward west brom \n",
+ "1 13.05 274.04 midfield chelsea \n",
+ "3 10.99 197.91 midfield manchester city "
+ ]
+ },
+ "execution_count": 15,
+ "metadata": {},
+ "output_type": "execute_result"
+ }
+ ],
+ "source": [
+ "# Sorting the DataFrame by a certain column (from highest to lowest)\n",
+ "\n",
+ "df.sort('goals', ascending=False, inplace=True)\n",
+ "df.head()"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 16,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [
+ {
+ "data": {
+ "text/html": [
+ "\n",
+ "\n",
+ " \n",
+ " \n",
+ " player \n",
+ " salary \n",
+ " games \n",
+ " goals \n",
+ " assists \n",
+ " shots_on_target \n",
+ " points_per_game \n",
+ " points \n",
+ " position \n",
+ " team \n",
+ " \n",
+ " \n",
+ " \n",
+ " \n",
+ " 1 \n",
+ " sergio agüero \n",
+ " 19.2 \n",
+ " 16 \n",
+ " 14 \n",
+ " 3 \n",
+ " 34 \n",
+ " 13.12 \n",
+ " 209.98 \n",
+ " forward \n",
+ " manchester city \n",
+ " \n",
+ " \n",
+ " 2 \n",
+ " alexis sánchez \n",
+ " 17.6 \n",
+ " 0 \n",
+ " 12 \n",
+ " 7 \n",
+ " 29 \n",
+ " 11.19 \n",
+ " 223.86 \n",
+ " forward \n",
+ " arsenal \n",
+ " \n",
+ " \n",
+ " 3 \n",
+ " saido berahino \n",
+ " 13.8 \n",
+ " 21 \n",
+ " 9 \n",
+ " 0 \n",
+ " 20 \n",
+ " 7.02 \n",
+ " 147.43 \n",
+ " forward \n",
+ " west brom \n",
+ " \n",
+ " \n",
+ " 4 \n",
+ " eden hazard \n",
+ " 18.9 \n",
+ " 21 \n",
+ " 8 \n",
+ " 4 \n",
+ " 17 \n",
+ " 13.05 \n",
+ " 274.04 \n",
+ " midfield \n",
+ " chelsea \n",
+ " \n",
+ " \n",
+ " 5 \n",
+ " yaya touré \n",
+ " 16.6 \n",
+ " 18 \n",
+ " 7 \n",
+ " 1 \n",
+ " 19 \n",
+ " 10.99 \n",
+ " 197.91 \n",
+ " midfield \n",
+ " manchester city \n",
+ " \n",
+ " \n",
+ "
\n",
+ ""
+ ],
+ "text/plain": [
+ " player salary games goals assists shots_on_target \\\n",
+ "1 sergio agüero 19.2 16 14 3 34 \n",
+ "2 alexis sánchez 17.6 0 12 7 29 \n",
+ "3 saido berahino 13.8 21 9 0 20 \n",
+ "4 eden hazard 18.9 21 8 4 17 \n",
+ "5 yaya touré 16.6 18 7 1 19 \n",
+ "\n",
+ " points_per_game points position team \n",
+ "1 13.12 209.98 forward manchester city \n",
+ "2 11.19 223.86 forward arsenal \n",
+ "3 7.02 147.43 forward west brom \n",
+ "4 13.05 274.04 midfield chelsea \n",
+ "5 10.99 197.91 midfield manchester city "
+ ]
+ },
+ "execution_count": 16,
+ "metadata": {},
+ "output_type": "execute_result"
+ }
+ ],
+ "source": [
+ "# Optional reindexing of the DataFrame after sorting\n",
+ "\n",
+ "df.index = range(1,len(df.index)+1)\n",
+ "df.head()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Updating Columns"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[[back to section overview](#Sections)]"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 17,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [
+ {
+ "data": {
+ "text/html": [
+ "\n",
+ "\n",
+ " \n",
+ " \n",
+ " player \n",
+ " salary \n",
+ " games \n",
+ " goals \n",
+ " assists \n",
+ " shots_on_target \n",
+ " points_per_game \n",
+ " points \n",
+ " position \n",
+ " team \n",
+ " \n",
+ " \n",
+ " \n",
+ " \n",
+ " 1 \n",
+ " sergio agüero \n",
+ " 20 \n",
+ " 16 \n",
+ " 14 \n",
+ " 3 \n",
+ " 34 \n",
+ " 13.12 \n",
+ " 209.98 \n",
+ " forward \n",
+ " manchester city \n",
+ " \n",
+ " \n",
+ " 2 \n",
+ " alexis sánchez \n",
+ " 15 \n",
+ " 0 \n",
+ " 12 \n",
+ " 7 \n",
+ " 29 \n",
+ " 11.19 \n",
+ " 223.86 \n",
+ " forward \n",
+ " arsenal \n",
+ " \n",
+ " \n",
+ " 3 \n",
+ " saido berahino \n",
+ " 13.8 \n",
+ " 21 \n",
+ " 9 \n",
+ " 0 \n",
+ " 20 \n",
+ " 7.02 \n",
+ " 147.43 \n",
+ " forward \n",
+ " west brom \n",
+ " \n",
+ " \n",
+ "
\n",
+ ""
+ ],
+ "text/plain": [
+ " player salary games goals assists shots_on_target \\\n",
+ "1 sergio agüero 20 16 14 3 34 \n",
+ "2 alexis sánchez 15 0 12 7 29 \n",
+ "3 saido berahino 13.8 21 9 0 20 \n",
+ "\n",
+ " points_per_game points position team \n",
+ "1 13.12 209.98 forward manchester city \n",
+ "2 11.19 223.86 forward arsenal \n",
+ "3 7.02 147.43 forward west brom "
+ ]
+ },
+ "execution_count": 17,
+ "metadata": {},
+ "output_type": "execute_result"
+ }
+ ],
+ "source": [
+ "# Creating a dummy DataFrame with changes in the `salary` column\n",
+ "\n",
+ "df_2 = df.copy()\n",
+ "df_2.loc[0:2, 'salary'] = [20.0, 15.0]\n",
+ "df_2.head(3)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 18,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [
+ {
+ "data": {
+ "text/html": [
+ "\n",
+ "\n",
+ " \n",
+ " \n",
+ " salary \n",
+ " games \n",
+ " goals \n",
+ " assists \n",
+ " shots_on_target \n",
+ " points_per_game \n",
+ " points \n",
+ " position \n",
+ " team \n",
+ " \n",
+ " \n",
+ " player \n",
+ " \n",
+ " \n",
+ " \n",
+ " \n",
+ " sergio agüero \n",
+ " 19.2 \n",
+ " 16 \n",
+ " 14 \n",
+ " 3 \n",
+ " 34 \n",
+ " 13.12 \n",
+ " 209.98 \n",
+ " forward \n",
+ " manchester city \n",
+ " \n",
+ " \n",
+ " alexis sánchez \n",
+ " 17.6 \n",
+ " 0 \n",
+ " 12 \n",
+ " 7 \n",
+ " 29 \n",
+ " 11.19 \n",
+ " 223.86 \n",
+ " forward \n",
+ " arsenal \n",
+ " \n",
+ " \n",
+ " saido berahino \n",
+ " 13.8 \n",
+ " 21 \n",
+ " 9 \n",
+ " 0 \n",
+ " 20 \n",
+ " 7.02 \n",
+ " 147.43 \n",
+ " forward \n",
+ " west brom \n",
+ " \n",
+ " \n",
+ "
\n",
+ ""
+ ],
+ "text/plain": [
+ " salary games goals assists shots_on_target \\\n",
+ "player \n",
+ "sergio agüero 19.2 16 14 3 34 \n",
+ "alexis sánchez 17.6 0 12 7 29 \n",
+ "saido berahino 13.8 21 9 0 20 \n",
+ "\n",
+ " points_per_game points position team \n",
+ "player \n",
+ "sergio agüero 13.12 209.98 forward manchester city \n",
+ "alexis sánchez 11.19 223.86 forward arsenal \n",
+ "saido berahino 7.02 147.43 forward west brom "
+ ]
+ },
+ "execution_count": 18,
+ "metadata": {},
+ "output_type": "execute_result"
+ }
+ ],
+ "source": [
+ "# Temporarily use the `player` columns as indices to \n",
+ "# apply the update functions\n",
+ "\n",
+ "df.set_index('player', inplace=True)\n",
+ "df_2.set_index('player', inplace=True)\n",
+ "df.head(3)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 19,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [
+ {
+ "data": {
+ "text/html": [
+ "\n",
+ "\n",
+ " \n",
+ " \n",
+ " salary \n",
+ " games \n",
+ " goals \n",
+ " assists \n",
+ " shots_on_target \n",
+ " points_per_game \n",
+ " points \n",
+ " position \n",
+ " team \n",
+ " \n",
+ " \n",
+ " player \n",
+ " \n",
+ " \n",
+ " \n",
+ " \n",
+ " sergio agüero \n",
+ " 20 \n",
+ " 16 \n",
+ " 14 \n",
+ " 3 \n",
+ " 34 \n",
+ " 13.12 \n",
+ " 209.98 \n",
+ " forward \n",
+ " manchester city \n",
+ " \n",
+ " \n",
+ " alexis sánchez \n",
+ " 15 \n",
+ " 0 \n",
+ " 12 \n",
+ " 7 \n",
+ " 29 \n",
+ " 11.19 \n",
+ " 223.86 \n",
+ " forward \n",
+ " arsenal \n",
+ " \n",
+ " \n",
+ " saido berahino \n",
+ " 13.8 \n",
+ " 21 \n",
+ " 9 \n",
+ " 0 \n",
+ " 20 \n",
+ " 7.02 \n",
+ " 147.43 \n",
+ " forward \n",
+ " west brom \n",
+ " \n",
+ " \n",
+ "
\n",
+ ""
+ ],
+ "text/plain": [
+ " salary games goals assists shots_on_target \\\n",
+ "player \n",
+ "sergio agüero 20 16 14 3 34 \n",
+ "alexis sánchez 15 0 12 7 29 \n",
+ "saido berahino 13.8 21 9 0 20 \n",
+ "\n",
+ " points_per_game points position team \n",
+ "player \n",
+ "sergio agüero 13.12 209.98 forward manchester city \n",
+ "alexis sánchez 11.19 223.86 forward arsenal \n",
+ "saido berahino 7.02 147.43 forward west brom "
+ ]
+ },
+ "execution_count": 19,
+ "metadata": {},
+ "output_type": "execute_result"
+ }
+ ],
+ "source": [
+ "# Update the `salary` column\n",
+ "df.update(other=df_2['salary'], overwrite=True)\n",
+ "df.head(3)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 20,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [
+ {
+ "data": {
+ "text/html": [
+ "\n",
+ "\n",
+ " \n",
+ " \n",
+ " player \n",
+ " salary \n",
+ " games \n",
+ " goals \n",
+ " assists \n",
+ " shots_on_target \n",
+ " points_per_game \n",
+ " points \n",
+ " position \n",
+ " team \n",
+ " \n",
+ " \n",
+ " \n",
+ " \n",
+ " 0 \n",
+ " sergio agüero \n",
+ " 20 \n",
+ " 16 \n",
+ " 14 \n",
+ " 3 \n",
+ " 34 \n",
+ " 13.12 \n",
+ " 209.98 \n",
+ " forward \n",
+ " manchester city \n",
+ " \n",
+ " \n",
+ " 1 \n",
+ " alexis sánchez \n",
+ " 15 \n",
+ " 0 \n",
+ " 12 \n",
+ " 7 \n",
+ " 29 \n",
+ " 11.19 \n",
+ " 223.86 \n",
+ " forward \n",
+ " arsenal \n",
+ " \n",
+ " \n",
+ " 2 \n",
+ " saido berahino \n",
+ " 13.8 \n",
+ " 21 \n",
+ " 9 \n",
+ " 0 \n",
+ " 20 \n",
+ " 7.02 \n",
+ " 147.43 \n",
+ " forward \n",
+ " west brom \n",
+ " \n",
+ " \n",
+ "
\n",
+ ""
+ ],
+ "text/plain": [
+ " player salary games goals assists shots_on_target \\\n",
+ "0 sergio agüero 20 16 14 3 34 \n",
+ "1 alexis sánchez 15 0 12 7 29 \n",
+ "2 saido berahino 13.8 21 9 0 20 \n",
+ "\n",
+ " points_per_game points position team \n",
+ "0 13.12 209.98 forward manchester city \n",
+ "1 11.19 223.86 forward arsenal \n",
+ "2 7.02 147.43 forward west brom "
+ ]
+ },
+ "execution_count": 20,
+ "metadata": {},
+ "output_type": "execute_result"
+ }
+ ],
+ "source": [
+ "# Reset the indices\n",
+ "df.reset_index(inplace=True)\n",
+ "df.head(3)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Chaining Conditions - Using Bitwise Operators"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[[back to section overview](#Sections)]"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 21,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [
+ {
+ "data": {
+ "text/html": [
+ "\n",
+ "\n",
+ " \n",
+ " \n",
+ " player \n",
+ " salary \n",
+ " games \n",
+ " goals \n",
+ " assists \n",
+ " shots_on_target \n",
+ " points_per_game \n",
+ " points \n",
+ " position \n",
+ " team \n",
+ " \n",
+ " \n",
+ " \n",
+ " \n",
+ " 1 \n",
+ " alexis sánchez \n",
+ " 15 \n",
+ " 0 \n",
+ " 12 \n",
+ " 7 \n",
+ " 29 \n",
+ " 11.19 \n",
+ " 223.86 \n",
+ " forward \n",
+ " arsenal \n",
+ " \n",
+ " \n",
+ " 3 \n",
+ " eden hazard \n",
+ " 18.9 \n",
+ " 21 \n",
+ " 8 \n",
+ " 4 \n",
+ " 17 \n",
+ " 13.05 \n",
+ " 274.04 \n",
+ " midfield \n",
+ " chelsea \n",
+ " \n",
+ " \n",
+ " 7 \n",
+ " santiago cazorla \n",
+ " 14.8 \n",
+ " 20 \n",
+ " 4 \n",
+ " 0 \n",
+ " 20 \n",
+ " 9.97 \n",
+ " 0.00 \n",
+ " midfield \n",
+ " arsenal \n",
+ " \n",
+ " \n",
+ " 9 \n",
+ " cesc fàbregas \n",
+ " 14.0 \n",
+ " 20 \n",
+ " 2 \n",
+ " 14 \n",
+ " 10 \n",
+ " 10.47 \n",
+ " 209.49 \n",
+ " midfield \n",
+ " chelsea \n",
+ " \n",
+ " \n",
+ "
\n",
+ ""
+ ],
+ "text/plain": [
+ " player salary games goals assists shots_on_target \\\n",
+ "1 alexis sánchez 15 0 12 7 29 \n",
+ "3 eden hazard 18.9 21 8 4 17 \n",
+ "7 santiago cazorla 14.8 20 4 0 20 \n",
+ "9 cesc fàbregas 14.0 20 2 14 10 \n",
+ "\n",
+ " points_per_game points position team \n",
+ "1 11.19 223.86 forward arsenal \n",
+ "3 13.05 274.04 midfield chelsea \n",
+ "7 9.97 0.00 midfield arsenal \n",
+ "9 10.47 209.49 midfield chelsea "
+ ]
+ },
+ "execution_count": 21,
+ "metadata": {},
+ "output_type": "execute_result"
+ }
+ ],
+ "source": [
+ "# Selecting only those players that either playing for Arsenal or Chelsea\n",
+ "\n",
+ "df[ (df['team'] == 'arsenal') | (df['team'] == 'chelsea') ]"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 22,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [
+ {
+ "data": {
+ "text/html": [
+ "\n",
+ "\n",
+ " \n",
+ " \n",
+ " player \n",
+ " salary \n",
+ " games \n",
+ " goals \n",
+ " assists \n",
+ " shots_on_target \n",
+ " points_per_game \n",
+ " points \n",
+ " position \n",
+ " team \n",
+ " \n",
+ " \n",
+ " \n",
+ " \n",
+ " 1 \n",
+ " alexis sánchez \n",
+ " 15 \n",
+ " 0 \n",
+ " 12 \n",
+ " 7 \n",
+ " 29 \n",
+ " 11.19 \n",
+ " 223.86 \n",
+ " forward \n",
+ " arsenal \n",
+ " \n",
+ " \n",
+ "
\n",
+ ""
+ ],
+ "text/plain": [
+ " player salary games goals assists shots_on_target \\\n",
+ "1 alexis sánchez 15 0 12 7 29 \n",
+ "\n",
+ " points_per_game points position team \n",
+ "1 11.19 223.86 forward arsenal "
+ ]
+ },
+ "execution_count": 22,
+ "metadata": {},
+ "output_type": "execute_result"
+ }
+ ],
+ "source": [
+ "# Selecting forwards from Arsenal only\n",
+ "\n",
+ "df[ (df['team'] == 'arsenal') & (df['position'] == 'forward') ]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Column Types"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[[back to section overview](#Sections)]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Printing Column Types"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 23,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [
+ {
+ "data": {
+ "text/plain": [
+ "{dtype('float64'): ['games',\n",
+ " 'goals',\n",
+ " 'assists',\n",
+ " 'shots_on_target',\n",
+ " 'points_per_game',\n",
+ " 'points'],\n",
+ " dtype('O'): ['player', 'salary', 'position', 'team']}"
+ ]
+ },
+ "execution_count": 23,
+ "metadata": {},
+ "output_type": "execute_result"
+ }
+ ],
+ "source": [
+ "types = df.columns.to_series().groupby(df.dtypes).groups\n",
+ "types"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Selecting by Column Type"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 24,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [
+ {
+ "data": {
+ "text/html": [
+ "\n",
+ "\n",
+ " \n",
+ " \n",
+ " player \n",
+ " salary \n",
+ " position \n",
+ " team \n",
+ " \n",
+ " \n",
+ " \n",
+ " \n",
+ " 0 \n",
+ " sergio agüero \n",
+ " 20 \n",
+ " forward \n",
+ " manchester city \n",
+ " \n",
+ " \n",
+ " 1 \n",
+ " alexis sánchez \n",
+ " 15 \n",
+ " forward \n",
+ " arsenal \n",
+ " \n",
+ " \n",
+ " 2 \n",
+ " saido berahino \n",
+ " 13.8 \n",
+ " forward \n",
+ " west brom \n",
+ " \n",
+ " \n",
+ " 3 \n",
+ " eden hazard \n",
+ " 18.9 \n",
+ " midfield \n",
+ " chelsea \n",
+ " \n",
+ " \n",
+ " 4 \n",
+ " yaya touré \n",
+ " 16.6 \n",
+ " midfield \n",
+ " manchester city \n",
+ " \n",
+ " \n",
+ "
\n",
+ ""
+ ],
+ "text/plain": [
+ " player salary position team\n",
+ "0 sergio agüero 20 forward manchester city\n",
+ "1 alexis sánchez 15 forward arsenal\n",
+ "2 saido berahino 13.8 forward west brom\n",
+ "3 eden hazard 18.9 midfield chelsea\n",
+ "4 yaya touré 16.6 midfield manchester city"
+ ]
+ },
+ "execution_count": 24,
+ "metadata": {},
+ "output_type": "execute_result"
+ }
+ ],
+ "source": [
+ "# select string columns\n",
+ "df.loc[:, (df.dtypes == np.dtype('O')).values].head()"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "### Converting Column Types"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 25,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [],
+ "source": [
+ "df['salary'] = df['salary'].astype(float)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 26,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [
+ {
+ "data": {
+ "text/plain": [
+ "{dtype('float64'): ['salary',\n",
+ " 'games',\n",
+ " 'goals',\n",
+ " 'assists',\n",
+ " 'shots_on_target',\n",
+ " 'points_per_game',\n",
+ " 'points'],\n",
+ " dtype('O'): ['player', 'position', 'team']}"
+ ]
+ },
+ "execution_count": 26,
+ "metadata": {},
+ "output_type": "execute_result"
+ }
+ ],
+ "source": [
+ "types = df.columns.to_series().groupby(df.dtypes).groups\n",
+ "types"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# If-tests"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[[back to section overview](#Sections)]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "I was recently asked how to do an if-test in pandas, that is, how to create an array of 1s and 0s depending on a condition, e.g., if `val` less than 0.5 -> 0, else -> 1. Using the boolean mask, that's pretty simple since `True` and `False` are integers after all."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 1,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [
+ {
+ "data": {
+ "text/plain": [
+ "1"
+ ]
+ },
+ "execution_count": 1,
+ "metadata": {},
+ "output_type": "execute_result"
+ }
+ ],
+ "source": [
+ "int(True)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 2,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [
+ {
+ "data": {
+ "text/html": [
+ "\n",
+ "\n",
+ " \n",
+ " \n",
+ " 0 \n",
+ " 1 \n",
+ " 2 \n",
+ " 3 \n",
+ " \n",
+ " \n",
+ " \n",
+ " \n",
+ " 0 \n",
+ " 2.0 \n",
+ " 0.30 \n",
+ " 4.00 \n",
+ " 5 \n",
+ " \n",
+ " \n",
+ " 1 \n",
+ " 0.8 \n",
+ " 0.03 \n",
+ " 0.02 \n",
+ " 5 \n",
+ " \n",
+ " \n",
+ "
\n",
+ ""
+ ],
+ "text/plain": [
+ " 0 1 2 3\n",
+ "0 2.0 0.30 4.00 5\n",
+ "1 0.8 0.03 0.02 5"
+ ]
+ },
+ "execution_count": 2,
+ "metadata": {},
+ "output_type": "execute_result"
+ }
+ ],
+ "source": [
+ "import pandas as pd\n",
+ "\n",
+ "a = [[2., .3, 4., 5.], [.8, .03, 0.02, 5.]]\n",
+ "df = pd.DataFrame(a)\n",
+ "df"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 3,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [
+ {
+ "data": {
+ "text/html": [
+ "\n",
+ "\n",
+ " \n",
+ " \n",
+ " 0 \n",
+ " 1 \n",
+ " 2 \n",
+ " 3 \n",
+ " \n",
+ " \n",
+ " \n",
+ " \n",
+ " 0 \n",
+ " False \n",
+ " False \n",
+ " False \n",
+ " False \n",
+ " \n",
+ " \n",
+ " 1 \n",
+ " False \n",
+ " True \n",
+ " True \n",
+ " False \n",
+ " \n",
+ " \n",
+ "
\n",
+ ""
+ ],
+ "text/plain": [
+ " 0 1 2 3\n",
+ "0 False False False False\n",
+ "1 False True True False"
+ ]
+ },
+ "execution_count": 3,
+ "metadata": {},
+ "output_type": "execute_result"
+ }
+ ],
+ "source": [
+ "df = df <= 0.05\n",
+ "df"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": 4,
+ "metadata": {
+ "collapsed": false
+ },
+ "outputs": [
+ {
+ "data": {
+ "text/html": [
+ "\n",
+ "\n",
+ " \n",
+ " \n",
+ " 0 \n",
+ " 1 \n",
+ " 2 \n",
+ " 3 \n",
+ " \n",
+ " \n",
+ " \n",
+ " \n",
+ " 0 \n",
+ " 0 \n",
+ " 0 \n",
+ " 0 \n",
+ " 0 \n",
+ " \n",
+ " \n",
+ " 1 \n",
+ " 0 \n",
+ " 1 \n",
+ " 1 \n",
+ " 0 \n",
+ " \n",
+ " \n",
+ "
\n",
+ ""
+ ],
+ "text/plain": [
+ " 0 1 2 3\n",
+ "0 0 0 0 0\n",
+ "1 0 1 1 0"
+ ]
+ },
+ "execution_count": 4,
+ "metadata": {},
+ "output_type": "execute_result"
+ }
+ ],
+ "source": [
+ "df.astype(int)"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "execution_count": null,
+ "metadata": {
+ "collapsed": true
+ },
+ "outputs": [],
+ "source": []
+ }
+ ],
+ "metadata": {
+ "kernelspec": {
+ "display_name": "Python 3",
+ "language": "python",
+ "name": "python3"
+ },
+ "language_info": {
+ "codemirror_mode": {
+ "name": "ipython",
+ "version": 3
+ },
+ "file_extension": ".py",
+ "mimetype": "text/x-python",
+ "name": "python",
+ "nbconvert_exporter": "python",
+ "pygments_lexer": "ipython3",
+ "version": "3.4.3"
+ }
+ },
+ "nbformat": 4,
+ "nbformat_minor": 0
+}
diff --git a/tutorials/unit_testing.md b/tutorials/unit_testing.md
new file mode 100644
index 0000000..9fa3bbf
--- /dev/null
+++ b/tutorials/unit_testing.md
@@ -0,0 +1,290 @@
+
+## Unit testing in Python - Why we want to make it a habit
+
+
+
+#### Sections
+
+Advantages of unit testing
+Main components a typical unit test
+The different unit test frameworks in Python
+Installing py.test)
+A py.test example walkthrough
+ Writing some code we want to test
+ Creating a "test" file
+ Testing edge cases and refining our code
+
+* * *
+
+
+
+## Advantages of unit testing
+
+Traditionally, for every piece of code we write (let it be a single function
+or class method), we would feed it some arbitrary inputs to make sure that it
+works the way we have expected. And this might sound like a reasonable
+approach given that everything works as it should and if we do not plan to
+make any changes to the code until the end of days. Of course, this is rarely
+the case.
+Suppose we want to modify our code by refactoring it, or by tweaking it for
+improved efficiency: Do we really want to manually type the previous test
+cases all over again to make sure we didn't break anything? Or suppose we are
+planning to pass our code along to our co-workers: What reason do they have to
+trust it? How can we make their life easier by providing evidence that
+everything was tested and is supposed to work properly?
+Surely, no one wants to spend hours or even days of mundane work to test code
+that was inherited before it can be put to use in good conscience.
+There must be a cleverer way, an automated and more systematic approach…
+This is where unit tests come into play. Once we designed the interface
+(_here:_ the in- and outputs of our functions and methods), we can write down
+several test cases and let them be checked every time we make changes to our
+code - without the tedious work of typing everything all over again, and
+without the risk of forgetting anything or by omitting crucial tests simply
+due to laziness.
+**This is especially important in scientific research, where your whole project depends on the correct analysis and assessment of any data - and there is probably no more convenient way to convince both you and the rightly skeptical reviewer that you just made a(nother) groundbreaking discovery.**
+
+
+
+
+## Main components a typical unit test
+
+In principle, unit testing is really no more than a more systematic way to
+automate code testing process. Where the term "unit" is typically defined as
+an isolated test case that consists of a the following components:
+
+\- a so-called "fixture" (e.g., a function, a class or class method, or even a
+data file)
+\- an action on the fixture (e.g., calling a function with a particular input)
+\- an expected outcome (e.g., the expected return value of a function)
+\- the actual outcome (e.g., the actual return value of a function call)
+\- a verification message (e.g., a report whether the actual return value
+matches the expected return value or not)
+
+
+
+
+## The different unit test frameworks in Python
+
+In Python, we have the luxury to be able to choose from a variety of good and
+capable unit testing frameworks. Probably, the most popular and most widely
+used ones are:
+
+\- the [unittest](http://docs.python.org/3.3/library/unittest.html) module -
+part of the Python Standard Library
+\- [nose](https://nose.readthedocs.org/en/latest/index.html)
+\- [py.test](http://pytest.org/latest/index.html)
+
+All of them work very well, and they are all sufficient for basic unit
+testing. Some people might prefer to use _nose_ over the more "basic"
+_unittest_ module. And many people are moving to the more recent _py.test_
+framework, since it offers some nice extensions and even more advanced and
+useful features. However, it shall not be the focus of this tutorial to
+discuss all the details of the different unit testing frameworks and weight
+them against each other. The screenshot below shows how the simple execution
+of _py.test_ and _nose_ may look like. To provide you with a little bit more
+background information: Both _nose_ and _py.test_ are crawling a subdirectory
+tree while looking for Python script files that start with the naming prefix
+"test". If those script files contain functions, classes, and class methods
+that also start with the prefix "test", the contained code will be executed by
+the unit testing frameworks.
+
+
+
+
+
+* * *
+
+Command line syntax:
+`py.test ` \- default unit testing with detailed report
+`py.test -q ` \- default unit testing with summarized report
+(quiet mode)
+`nosetests` \- default unit testing with summarized report
+`nosetests -v` \- default unit testing with detailed report (verbose mode)
+
+* * *
+
+
+
+For the further sections of this tutorial, we will be using _py.test_, but
+everything is also compatible to the _nose_ framework, and for the simple
+examples below it would not matter which framework we picked.
+However, there is one little difference in the default behavior, though, and
+it might also answer the question: "How does the framework know where to find
+the test code to execute?"
+By default, _py.test_ descends into all subdirectories (from the current
+working directory or a particular folder that you provided as additional
+argument) looking for Python scripts that start with the prefix "test". If
+there are functions, classes, or class methods contained in these scripts that
+also start with the prefix "test", those will be executed by the unit testing
+framework. The basic behavior of _nose_ is quite similar, but in contrast to
+browsing through all subdirectories, it will only consider those that start
+with the prefix "test" to look for the respective Python unit test code. Thus,
+it is a good habit to put all your test code under a directory starting with
+the prefix "test" even if you use _py.test_ \- your _nose_ colleagues will
+thank you!
+The figure below shows how the _nose_ and _py.test_ unit test frameworks would
+descend the subdirectory tree looking for Python script files that start with
+the prefix "test".
+
+
+_Note: Interestingly,_ nose _seems to be twice as fast as_ py.test _in the
+example above, and I was curious if it is due to the fact that_ py.test
+_searches all subdirectories (_ nose _only searches those that start with
+"test"). Although there is a tiny speed difference when I specify the test
+code containing folder directly,_ nose _still seems to be faster. However, I
+don't know how it scales, and it might be an interesting experiment to test
+for much larger projects._
+
+
+
+
+
+
+## Installing py.test
+
+Installing py.test is pretty straightforward. We can install it directly from
+the command line via
+
+
+
+ pip install -U pytest
+
+
+
+or
+
+
+
+ easy_install -U pytest
+
+
+
+If this doesn't work for you, you can visit the _py.test_ website
+(
+
+
+
+or
+
+
+
+ python -m pytest
+
+
+
+
+
+
+
+## A py.test example walkthrough
+
+For the following example we will be using _py.test_, however, _nose_ works
+pretty similarly, and as I mentioned in the previous section, I only want to
+focus on the essentials of unit testing here. Note that _py.test_ has a lot of
+advanced and useful features to offer that we won't touch in this tutorial,
+e.g., setting break points for debugging, etc. (if you want to learn more,
+please take a look at the complete _py.test_ documentation:
+).
+
+
+
+### Writing some code we want to test
+
+Assume we wrote two very simple functions that we want to test, either as
+small scripts or part of a larger package. The first function,
+"multiple_of_three", is supposed to check whether a number is a multiple of
+the number 3 or not. We want the function to return the boolean value True if
+this is the case, and else it should return False. The second function,
+"filter_multiples_of_three", takes a list as input argument and is supposed to
+return a subset of the input list containing only those numbers that are
+multiples of 3.
+
+
+
+
+
+### Creating a "test" file
+
+Next, we write a small unit test to check if our function works for some
+simple input cases:
+
+
+
+
+Great, when we run our py.test unit testing framework, we see that everything
+works as expected!
+
+
+
+
+But what about edge cases?
+
+
+
+
+### Testing edge cases and refining our code
+
+In order to check if our function is yet robust enough to handle special
+cases, e.g., 0 as input, we extend our unit test code. Here, assume that we
+don't want 0 to evaluate to True, since we don't consider 3 to be a factor of
+0.
+
+
+
+
+As we can see from the _py.test report_, our test just failed. So let us go
+back and fix our code to handle this special case.
+
+
+
+So far so good, when we execute _py.test_ again (image not shown) we see that
+our codes handles 0 correctly now. Let us add some more edge cases: Negative
+integers, decimal floating-point numbers, and large integers.
+
+
+
+
+According to the unit test report, we face another problem here: Our code
+considers 3 as a factor of -9 (negative 9). For the sake of this example,
+let's assume that we don't want this to happen: We'd like to consider only
+positive numbers to be multiples of 3. In order to account for those cases, we
+need to make another small modification to our code by changing `!=0` to `>0`
+in the if-statement.
+
+
+
+After running the _py.test_ utility again, we are certain that our code can
+also handle negative numbers correctly now. And once we are satisfied with the
+general behavior of our current code, we can move on to testing the next
+function "filter_multiples_of_three", which depends on the correctness of
+"multiple_of_three".
+
+
+
+
+This time, our test seems to be "bug"-free, and we are confident that it can
+handle all the scenarios we could currently think of. If we plan to make any
+further modifications to the code in future, nothing can be more convenient to
+just re-run our previous tests in order to make sure that we didn't break
+anything.
+
+If you have any questions or need more explanations, you are welcome to
+provide feedback in the comment section below.
+
+
+
+
diff --git a/tutorials/useful_regex.ipynb b/tutorials/useful_regex.ipynb
new file mode 100644
index 0000000..24bcf14
--- /dev/null
+++ b/tutorials/useful_regex.ipynb
@@ -0,0 +1,1070 @@
+{
+ "metadata": {
+ "name": "",
+ "signature": "sha256:237609a5ef934bf65a93a410c9e5107b808049dd04b0faf2b30f9b423699ba6c"
+ },
+ "nbformat": 3,
+ "nbformat_minor": 0,
+ "worksheets": [
+ {
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[Sebastian Raschka](http://sebastianraschka.com) \n",
+ "\n",
+ "- [Link to this IPython notebook on Github](https://github.com/rasbt/python_reference/blob/master/tutorials/useful_regex.ipynb) "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "%load_ext watermark"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [],
+ "prompt_number": 1
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "%watermark -d -v -u -t -z"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [
+ {
+ "output_type": "stream",
+ "stream": "stdout",
+ "text": [
+ "Last updated: 06/07/2014 22:50:23 EDT\n",
+ "\n",
+ "CPython 3.4.1\n",
+ "IPython 2.1.0\n"
+ ]
+ }
+ ],
+ "prompt_number": 2
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[More information](http://nbviewer.ipython.org/github/rasbt/python_reference/blob/master/ipython_magic/watermark.ipynb) about the `watermark` magic command extension."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "I would be happy to hear your comments and suggestions. \n",
+ "Please feel free to drop me a note via\n",
+ "[twitter](https://twitter.com/rasbt), [email](mailto:bluewoodtree@gmail.com), or [google+](https://plus.google.com/+SebastianRaschka).\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "heading",
+ "level": 1,
+ "metadata": {},
+ "source": [
+ "A collection of useful regular expressions"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "heading",
+ "level": 2,
+ "metadata": {},
+ "source": [
+ "Sections"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "- [About the `re` module](#About-the-re-module)\n",
+ "- [Identify files via file extensions](#Identify-files-via-file-extensions)\n",
+ "- [Username validation](#Username-validation)\n",
+ "- [Checking for valid email addresses](#Checking-for-valid-email-addresses)\n",
+ "- [Check for a valid URL](#Check-for-a-valid-URL)\n",
+ "- [Checking for numbers](#Checking-for-numbers)\n",
+ "- [Validating dates](#Validating-dates)\n",
+ "- [Time](#Time)\n",
+ "- [Checking for HTML tags](#Checking-for-HTML-tags)\n",
+ "- [Checking for IP addresses](#Checking-for-IP-addresses)\n",
+ "- [Checking for MAC addresses](#Checking-for-MAC-addresses)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "heading",
+ "level": 2,
+ "metadata": {},
+ "source": [
+ "About the `re` module"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[[back to top](#Sections)]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "The purpose of this IPython notebook is not to rewrite a detailed tutorial about regular expressions or the in-built Python `re` module, but to collect some useful regular expressions for copy&paste purposes."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "The complete documentation of the Python `re` module can be found here [https://docs.python.org/3.4/howto/regex.html](https://docs.python.org/3.4/howto/regex.html). Below, I just want to list the most important methods for convenience:"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "- `re.match()` : Determine if the RE matches at the beginning of the string.\n",
+ "- `re.search()` : Scan through a string, looking for any location where this RE matches.\n",
+ "- `re.findall()` : Find all substrings where the RE matches, and returns them as a list.\n",
+ "- `re.finditer()` : Find all substrings where the RE matches, and returns them as an iterator."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "If you are using the same regular expression multiple times, it is recommended to compile it for improved performance.\n",
+ "\n",
+ " compiled_re = re.compile(r'some_regexpr') \n",
+ " for word in text:\n",
+ " match = comp.search(compiled_re))\n",
+ " # do something with the match\n",
+ " \n",
+ "**E.g., if we want to check if a string ends with a substring:**"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "import re\n",
+ "\n",
+ "needle = 'needlers'\n",
+ "\n",
+ "# Python approach\n",
+ "print(bool(any([needle.endswith(e) for e in ('ly', 'ed', 'ing', 'ers')])))\n",
+ "\n",
+ "# On-the-fly Regular expression in Python\n",
+ "print(bool(re.search(r'(?:ly|ed|ing|ers)$', needle)))\n",
+ "\n",
+ "# Compiled Regular expression in Python\n",
+ "comp = re.compile(r'(?:ly|ed|ing|ers)$') \n",
+ "print(bool(comp.search(needle)))"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [
+ {
+ "output_type": "stream",
+ "stream": "stdout",
+ "text": [
+ "True\n",
+ "True\n",
+ "True\n"
+ ]
+ }
+ ],
+ "prompt_number": 3
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "%timeit -n 10000 -r 50 bool(any([needle.endswith(e) for e in ('ly', 'ed', 'ing', 'ers')]))\n",
+ "%timeit -n 10000 -r 50 bool(re.search(r'(?:ly|ed|ing|ers)$', needle))\n",
+ "%timeit -n 10000 -r 50 bool(comp.search(needle))"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [
+ {
+ "output_type": "stream",
+ "stream": "stdout",
+ "text": [
+ "10000 loops, best of 50: 2.74 \u00b5s per loop\n",
+ "10000 loops, best of 50: 2.93 \u00b5s per loop"
+ ]
+ },
+ {
+ "output_type": "stream",
+ "stream": "stdout",
+ "text": [
+ "\n",
+ "10000 loops, best of 50: 1.28 \u00b5s per loop"
+ ]
+ },
+ {
+ "output_type": "stream",
+ "stream": "stdout",
+ "text": [
+ "\n"
+ ]
+ }
+ ],
+ "prompt_number": 4
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "heading",
+ "level": 2,
+ "metadata": {},
+ "source": [
+ "Identify files via file extensions"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[[back to top](#Sections)]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "A regular expression to check for file extensions. \n",
+ "\n",
+ "Note: This approach is not recommended for thorough limitation of file types (parse the file header instead). However, this regex is still a useful alternative to e.g., a Python's `endswith` approach for quick pre-filtering for certain files of interest."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "pattern = r'(?i)(\\w+)\\.(jpeg|jpg|png|gif|tif|svg)$'\n",
+ "\n",
+ "# remove `(?i)` to make regexpr case-sensitive\n",
+ "\n",
+ "str_true = ('test.gif', \n",
+ " 'image.jpeg', \n",
+ " 'image.jpg',\n",
+ " 'image.TIF'\n",
+ " )\n",
+ "\n",
+ "str_false = ('test.pdf',\n",
+ " 'test.gif.pdf',\n",
+ " )\n",
+ "\n",
+ "for t in str_true:\n",
+ " assert(bool(re.match(pattern, t)) == True), '%s is not True' %t\n",
+ "for f in str_false:\n",
+ " assert(bool(re.match(pattern, f)) == False), '%s is not False' %f"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [],
+ "prompt_number": 5
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "heading",
+ "level": 2,
+ "metadata": {},
+ "source": [
+ "Username validation"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[[back to top](#Sections)]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Checking for a valid user name that has a certain minimum and maximum length.\n",
+ "\n",
+ "Allowed characters:\n",
+ "- letters (upper- and lower-case)\n",
+ "- numbers\n",
+ "- dashes\n",
+ "- underscores"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "min_len = 5 # minimum length for a valid username\n",
+ "max_len = 15 # maximum length for a valid username\n",
+ "\n",
+ "pattern = r\"^(?i)[a-z0-9_-]{%s,%s}$\" %(min_len, max_len)\n",
+ "\n",
+ "# remove `(?i)` to only allow lower-case letters\n",
+ "\n",
+ "\n",
+ "\n",
+ "str_true = ('user123', '123_user', 'Username')\n",
+ " \n",
+ "str_false = ('user', 'username1234_is-way-too-long', 'user$34354')\n",
+ "\n",
+ "for t in str_true:\n",
+ " assert(bool(re.match(pattern, t)) == True), '%s is not True' %t\n",
+ "for f in str_false:\n",
+ " assert(bool(re.match(pattern, f)) == False), '%s is not False' %f"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [],
+ "prompt_number": 6
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "heading",
+ "level": 2,
+ "metadata": {},
+ "source": [
+ "Checking for valid email addresses"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[[back to top](#Sections)]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "A regular expression that captures most email addresses."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "pattern = r\"(^[a-zA-Z0-9_.+-]+@[a-zA-Z0-9-]+\\.[a-zA-Z0-9-.]+$)\"\n",
+ "\n",
+ "str_true = ('test@mail.com',)\n",
+ " \n",
+ "str_false = ('testmail.com', '@testmail.com', 'test@mailcom')\n",
+ "\n",
+ "for t in str_true:\n",
+ " assert(bool(re.match(pattern, t)) == True), '%s is not True' %t\n",
+ "for f in str_false:\n",
+ " assert(bool(re.match(pattern, f)) == False), '%s is not False' %f"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [],
+ "prompt_number": 7
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "source: [http://stackoverflow.com/questions/201323/using-a-regular-expression-to-validate-an-email-address](http://stackoverflow.com/questions/201323/using-a-regular-expression-to-validate-an-email-address)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "heading",
+ "level": 2,
+ "metadata": {},
+ "source": [
+ "Check for a valid URL"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[[back to top](#Sections)]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Checks for an URL if a string ...\n",
+ "\n",
+ "- starts with `https://`, or `http://`, or `www.`\n",
+ "- or ends with a dot extension"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "pattern = '^(https?:\\/\\/)?([\\da-z\\.-]+)\\.([a-z\\.]{2,6})([\\/\\w \\.-]*)*\\/?$'\n",
+ "\n",
+ "str_true = ('https://github.com', \n",
+ " 'http://github.com',\n",
+ " 'www.github.com',\n",
+ " 'github.com',\n",
+ " 'test.de',\n",
+ " 'https://github.com/rasbt',\n",
+ " 'test.jpeg' # !!! \n",
+ " )\n",
+ " \n",
+ "str_false = ('testmailcom', 'http:testmailcom', )\n",
+ "\n",
+ "for t in str_true:\n",
+ " assert(bool(re.match(pattern, t)) == True), '%s is not True' %t\n",
+ "\n",
+ "for f in str_false:\n",
+ " assert(bool(re.match(pattern, f)) == False), '%s is not False' %f"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [],
+ "prompt_number": 8
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "source: [http://code.tutsplus.com/tutorials/8-regular-expressions-you-should-know--net-6149](http://code.tutsplus.com/tutorials/8-regular-expressions-you-should-know--net-6149)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "heading",
+ "level": 2,
+ "metadata": {},
+ "source": [
+ "Checking for numbers"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[[back to top](#Sections)]"
+ ]
+ },
+ {
+ "cell_type": "heading",
+ "level": 3,
+ "metadata": {},
+ "source": [
+ "Positive integers"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "pattern = '^\\d+$'\n",
+ "\n",
+ "str_true = ('123', '1', )\n",
+ " \n",
+ "str_false = ('abc', '1.1', )\n",
+ "\n",
+ "for t in str_true:\n",
+ " assert(bool(re.match(pattern, t)) == True), '%s is not True' %t\n",
+ "\n",
+ "for f in str_false:\n",
+ " assert(bool(re.match(pattern, f)) == False), '%s is not False' %f"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [],
+ "prompt_number": 9
+ },
+ {
+ "cell_type": "heading",
+ "level": 3,
+ "metadata": {},
+ "source": [
+ "Negative integers"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "pattern = '^-\\d+$'\n",
+ "\n",
+ "str_true = ('-123', '-1', )\n",
+ " \n",
+ "str_false = ('123', '-abc', '-1.1', )\n",
+ "\n",
+ "for t in str_true:\n",
+ " assert(bool(re.match(pattern, t)) == True), '%s is not True' %t\n",
+ "\n",
+ "for f in str_false:\n",
+ " assert(bool(re.match(pattern, f)) == False), '%s is not False' %f"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [],
+ "prompt_number": 10
+ },
+ {
+ "cell_type": "heading",
+ "level": 3,
+ "metadata": {},
+ "source": [
+ "All integers"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "pattern = '^-{0,1}\\d+$'\n",
+ "\n",
+ "str_true = ('-123', '-1', '1', '123',)\n",
+ " \n",
+ "str_false = ('123.0', '-abc', '-1.1', )\n",
+ "\n",
+ "for t in str_true:\n",
+ " assert(bool(re.match(pattern, t)) == True), '%s is not True' %t\n",
+ "\n",
+ "for f in str_false:\n",
+ " assert(bool(re.match(pattern, f)) == False), '%s is not False' %f"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [],
+ "prompt_number": 11
+ },
+ {
+ "cell_type": "heading",
+ "level": 3,
+ "metadata": {},
+ "source": [
+ "Positive numbers"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "pattern = '^\\d*\\.{0,1}\\d+$'\n",
+ "\n",
+ "str_true = ('1', '123', '1.234', )\n",
+ " \n",
+ "str_false = ('-abc', '-123', '-123.0')\n",
+ "\n",
+ "for t in str_true:\n",
+ " assert(bool(re.match(pattern, t)) == True), '%s is not True' %t\n",
+ "\n",
+ "for f in str_false:\n",
+ " assert(bool(re.match(pattern, f)) == False), '%s is not False' %f"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [],
+ "prompt_number": 12
+ },
+ {
+ "cell_type": "heading",
+ "level": 3,
+ "metadata": {},
+ "source": [
+ "Negative numbers"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "pattern = '^-\\d*\\.{0,1}\\d+$'\n",
+ "\n",
+ "str_true = ('-1', '-123', '-123.0', )\n",
+ " \n",
+ "str_false = ('-abc', '1', '123', '1.234', )\n",
+ "\n",
+ "for t in str_true:\n",
+ " assert(bool(re.match(pattern, t)) == True), '%s is not True' %t\n",
+ "\n",
+ "for f in str_false:\n",
+ " assert(bool(re.match(pattern, f)) == False), '%s is not False' %f"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [],
+ "prompt_number": 13
+ },
+ {
+ "cell_type": "heading",
+ "level": 3,
+ "metadata": {},
+ "source": [
+ "All numbers"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "pattern = '^-{0,1}\\d*\\.{0,1}\\d+$'\n",
+ "\n",
+ "str_true = ('1', '123', '1.234', '-123', '-123.0')\n",
+ " \n",
+ "str_false = ('-abc')\n",
+ "\n",
+ "for t in str_true:\n",
+ " assert(bool(re.match(pattern, t)) == True), '%s is not True' %t\n",
+ "\n",
+ "for f in str_false:\n",
+ " assert(bool(re.match(pattern, f)) == False), '%s is not False' %f"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [],
+ "prompt_number": 14
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "source: [http://stackoverflow.com/questions/1449817/what-are-some-of-the-most-useful-regular-expressions-for-programmers](http://stackoverflow.com/questions/1449817/what-are-some-of-the-most-useful-regular-expressions-for-programmers)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "heading",
+ "level": 2,
+ "metadata": {},
+ "source": [
+ "Validating dates"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[[back to top](#Sections)]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Validates dates in `mm/dd/yyyy` format."
+ ]
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "pattern = '^(0[1-9]|1[0-2])\\/(0[1-9]|1\\d|2\\d|3[01])\\/(19|20)\\d{2}$'\n",
+ "\n",
+ "str_true = ('01/08/2014', '12/30/2014', )\n",
+ " \n",
+ "str_false = ('22/08/2014', '-123', '1/8/2014', '1/08/2014', '01/8/2014')\n",
+ "\n",
+ "for t in str_true:\n",
+ " assert(bool(re.match(pattern, t)) == True), '%s is not True' %t\n",
+ "\n",
+ "for f in str_false:\n",
+ " assert(bool(re.match(pattern, f)) == False), '%s is not False' %f"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [],
+ "prompt_number": 15
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[[back to top](#Sections)]"
+ ]
+ },
+ {
+ "cell_type": "heading",
+ "level": 3,
+ "metadata": {},
+ "source": [
+ "12-Hour format"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "pattern = r'^(1[012]|[1-9]):[0-5][0-9](\\s)?(?i)(am|pm)$'\n",
+ "\n",
+ "str_true = ('2:00pm', '7:30 AM', '12:05 am', )\n",
+ " \n",
+ "str_false = ('22:00pm', '14:00', '3:12', '03:12pm', )\n",
+ "\n",
+ "for t in str_true:\n",
+ " assert(bool(re.match(pattern, t)) == True), '%s is not True' %t\n",
+ "\n",
+ "for f in str_false:\n",
+ " assert(bool(re.match(pattern, f)) == False), '%s is not False' %f"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [],
+ "prompt_number": 29
+ },
+ {
+ "cell_type": "heading",
+ "level": 3,
+ "metadata": {},
+ "source": [
+ "24-Hour format"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "pattern = r'^([0-1]{1}[0-9]{1}|20|21|22|23):[0-5]{1}[0-9]{1}$'\n",
+ "\n",
+ "str_true = ('14:00', '00:30', )\n",
+ " \n",
+ "str_false = ('22:00pm', '4:00', )\n",
+ "\n",
+ "for t in str_true:\n",
+ " assert(bool(re.match(pattern, t)) == True), '%s is not True' %t\n",
+ "for f in str_false:\n",
+ " assert(bool(re.match(pattern, f)) == False), '%s is not False' %f"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [],
+ "prompt_number": 18
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "heading",
+ "level": 2,
+ "metadata": {},
+ "source": [
+ "Checking for HTML tags"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[[back to top](#Sections)]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Also this regex is only recommended for \"filtering\" purposes and not a ultimate way to parse HTML. For more information see this excellent discussion on StackOverflow: \n",
+ "[http://stackoverflow.com/questions/1732348/regex-match-open-tags-except-xhtml-self-contained-tags/](http://stackoverflow.com/questions/1732348/regex-match-open-tags-except-xhtml-self-contained-tags/) "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "pattern = r\"\"\"\\s]+))?)+\\s*|\\s*)/?>\"\"\"\n",
+ "\n",
+ "str_true = ('', '', '', '![]() ')\n",
+ " \n",
+ "str_false = ('a>', '')\n",
+ "\n",
+ "for t in str_true:\n",
+ " assert(bool(re.match(pattern, t)) == True), '%s is not True' %t\n",
+ "\n",
+ "for f in str_false:\n",
+ " assert(bool(re.match(pattern, f)) == False), '%s is not False' %f"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [],
+ "prompt_number": 16
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "source: [http://haacked.com/archive/2004/10/25/usingregularexpressionstomatchhtml.aspx/](http://haacked.com/archive/2004/10/25/usingregularexpressionstomatchhtml.aspx/)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
')\n",
+ " \n",
+ "str_false = ('a>', '')\n",
+ "\n",
+ "for t in str_true:\n",
+ " assert(bool(re.match(pattern, t)) == True), '%s is not True' %t\n",
+ "\n",
+ "for f in str_false:\n",
+ " assert(bool(re.match(pattern, f)) == False), '%s is not False' %f"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [],
+ "prompt_number": 16
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "source: [http://haacked.com/archive/2004/10/25/usingregularexpressionstomatchhtml.aspx/](http://haacked.com/archive/2004/10/25/usingregularexpressionstomatchhtml.aspx/)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "heading",
+ "level": 2,
+ "metadata": {},
+ "source": [
+ "Checking for IP addresses"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[[back to top](#Sections)]"
+ ]
+ },
+ {
+ "cell_type": "heading",
+ "level": 3,
+ "metadata": {},
+ "source": [
+ "IPv4"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ ""
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Image source: http://en.wikipedia.org/wiki/File:Ipv4_address.svg"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "pattern = r'^(?:(?:25[0-5]|2[0-4][0-9]|[01]?[0-9][0-9]?)\\.){3}(?:25[0-5]|2[0-4][0-9]|[01]?[0-9][0-9]?)$'\n",
+ "\n",
+ "str_true = ('172.16.254.1', '1.2.3.4', '01.102.103.104', )\n",
+ " \n",
+ "str_false = ('17216.254.1', '1.2.3.4.5', '01 .102.103.104', )\n",
+ "\n",
+ "for t in str_true:\n",
+ " assert(bool(re.match(pattern, t)) == True), '%s is not True' %t\n",
+ "\n",
+ "for f in str_false:\n",
+ " assert(bool(re.match(pattern, f)) == False), '%s is not False' %f"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [],
+ "prompt_number": 8
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "source: [http://answers.oreilly.com/topic/318-how-to-match-ipv4-addresses-with-regular-expressions/](http://answers.oreilly.com/topic/318-how-to-match-ipv4-addresses-with-regular-expressions/)"
+ ]
+ },
+ {
+ "cell_type": "heading",
+ "level": 3,
+ "metadata": {},
+ "source": [
+ "Ipv6"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ ""
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Image source: http://upload.wikimedia.org/wikipedia/commons/1/15/Ipv6_address.svg"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "pattern = r'^\\s*((([0-9A-Fa-f]{1,4}:){7}([0-9A-Fa-f]{1,4}|:))|(([0-9A-Fa-f]{1,4}:){6}(:[0-9A-Fa-f]{1,4}|((25[0-5]|2[0-4]\\d|1\\d\\d|[1-9]?\\d)(\\.(25[0-5]|2[0-4]\\d|1\\d\\d|[1-9]?\\d)){3})|:))|(([0-9A-Fa-f]{1,4}:){5}(((:[0-9A-Fa-f]{1,4}){1,2})|:((25[0-5]|2[0-4]\\d|1\\d\\d|[1-9]?\\d)(\\.(25[0-5]|2[0-4]\\d|1\\d\\d|[1-9]?\\d)){3})|:))|(([0-9A-Fa-f]{1,4}:){4}(((:[0-9A-Fa-f]{1,4}){1,3})|((:[0-9A-Fa-f]{1,4})?:((25[0-5]|2[0-4]\\d|1\\d\\d|[1-9]?\\d)(\\.(25[0-5]|2[0-4]\\d|1\\d\\d|[1-9]?\\d)){3}))|:))|(([0-9A-Fa-f]{1,4}:){3}(((:[0-9A-Fa-f]{1,4}){1,4})|((:[0-9A-Fa-f]{1,4}){0,2}:((25[0-5]|2[0-4]\\d|1\\d\\d|[1-9]?\\d)(\\.(25[0-5]|2[0-4]\\d|1\\d\\d|[1-9]?\\d)){3}))|:))|(([0-9A-Fa-f]{1,4}:){2}(((:[0-9A-Fa-f]{1,4}){1,5})|((:[0-9A-Fa-f]{1,4}){0,3}:((25[0-5]|2[0-4]\\d|1\\d\\d|[1-9]?\\d)(\\.(25[0-5]|2[0-4]\\d|1\\d\\d|[1-9]?\\d)){3}))|:))|(([0-9A-Fa-f]{1,4}:){1}(((:[0-9A-Fa-f]{1,4}){1,6})|((:[0-9A-Fa-f]{1,4}){0,4}:((25[0-5]|2[0-4]\\d|1\\d\\d|[1-9]?\\d)(\\.(25[0-5]|2[0-4]\\d|1\\d\\d|[1-9]?\\d)){3}))|:))|(:(((:[0-9A-Fa-f]{1,4}){1,7})|((:[0-9A-Fa-f]{1,4}){0,5}:((25[0-5]|2[0-4]\\d|1\\d\\d|[1-9]?\\d)(\\.(25[0-5]|2[0-4]\\d|1\\d\\d|[1-9]?\\d)){3}))|:)))(%.+)?\\s*$'\n",
+ "\n",
+ "str_true = ('2001:470:9b36:1::2',\n",
+ " '2001:cdba:0000:0000:0000:0000:3257:9652', \n",
+ " '2001:cdba:0:0:0:0:3257:9652', \n",
+ " '2001:cdba::3257:9652', )\n",
+ " \n",
+ "str_false = ('1200::AB00:1234::2552:7777:1313', # uses `::` twice\n",
+ " '1200:0000:AB00:1234:O000:2552:7777:1313', ) # contains an O instead of 0\n",
+ "\n",
+ "for t in str_true:\n",
+ " assert(bool(re.match(pattern, t)) == True), '%s is not True' %t\n",
+ "\n",
+ "for f in str_false:\n",
+ " assert(bool(re.match(pattern, f)) == False), '%s is not False' %f"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [],
+ "prompt_number": 21
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "source: [http://snipplr.com/view/43003/regex--match-ipv6-address/](http://snipplr.com/view/43003/regex--match-ipv6-address/)"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "
\n",
+ "
"
+ ]
+ },
+ {
+ "cell_type": "heading",
+ "level": 2,
+ "metadata": {},
+ "source": [
+ "Checking for MAC addresses"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "[[back to top](#Sections)]"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ ""
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Image source: http://upload.wikimedia.org/wikipedia/en/3/37/MACaddressV3.png "
+ ]
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "pattern = r'^(?i)([0-9A-F]{2}[:-]){5}([0-9A-F]{2})$'\n",
+ "\n",
+ "str_true = ('94-AE-70-A0-66-83', \n",
+ " '58-f8-1a-00-44-c8',\n",
+ " '00:A0:C9:14:C8:29'\n",
+ " , )\n",
+ " \n",
+ "str_false = ('0:00:00:00:00:00', \n",
+ " '94-AE-70-A0 -66-83', ) \n",
+ "\n",
+ "for t in str_true:\n",
+ " assert(bool(re.match(pattern, t)) == True), '%s is not True' %t\n",
+ "\n",
+ "for f in str_false:\n",
+ " assert(bool(re.match(pattern, f)) == False), '%s is not False' %f"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [],
+ "prompt_number": 29
+ }
+ ],
+ "metadata": {}
+ }
+ ]
+}
\ No newline at end of file
diff --git a/useful_scripts/combinations.py b/useful_scripts/combinations.py
new file mode 100755
index 0000000..5dbe91d
--- /dev/null
+++ b/useful_scripts/combinations.py
@@ -0,0 +1,71 @@
+#!/usr/bin/env python
+
+# Sebastian Raschka 2014
+# Functions to calculate factorial, combinations, and permutations
+# bundled in an simple command line interface.
+
+def factorial(n):
+ if n == 0:
+ return 1
+ else:
+ return n * factorial(n-1)
+
+def combinations(n, r):
+ numerator = factorial(n)
+ denominator = factorial(r) * factorial(n-r)
+ return int(numerator/denominator)
+
+def permutations(n, r):
+ numerator = factorial(n)
+ denominator = factorial(n-r)
+ return int(numerator/denominator)
+
+assert(factorial(3) == 6)
+assert(combinations(20, 8) == 125970)
+assert(permutations(30, 3) == 24360)
+
+
+
+
+if __name__ == '__main__':
+
+ import argparse
+ parser = argparse.ArgumentParser(
+ description='Script to calculate the number of combinations or permutations ("n choose r")',
+ formatter_class=argparse.RawTextHelpFormatter,
+
+ prog='Combinations',
+ epilog='Example: ./combinations.py -c 20 3'
+ )
+
+ parser.add_argument('-c', '--combinations', type=int, metavar='NUMBER', nargs=2,
+ help='Combinations: Number of ways to combine n items with sequence length r where the item order does not matter.')
+
+ parser.add_argument('-p', '--permutations', type=int, metavar='NUMBER', nargs=2,
+ help='Permutations: Number of ways to combine n items with sequence length r where the item order does not matter.')
+
+ parser.add_argument('-f', '--factorial', type=int, metavar='NUMBER', help='n! e.g., 5! = 5*4*3*2*1 = 120.')
+
+ parser.add_argument('--version', action='version', version='%(prog)s 1.0')
+
+ args = parser.parse_args()
+
+ if not any((args.combinations, args.permutations, args.factorial)):
+ parser.print_help()
+ quit()
+
+ if args.factorial:
+ print(factorial(args.factorial))
+
+ if args.combinations:
+ print(combinations(args.combinations[0], args.combinations[1]))
+
+ if args.permutations:
+ print(permutations(args.permutations[0], args.permutations[1]))
+
+ if args.factorial:
+ print(factorial(args.factorial))
+
+
+
+
\ No newline at end of file
diff --git a/useful_scripts/conc_gzip_files.py b/useful_scripts/conc_gzip_files.py
index da849c9..b8d9b33 100644
--- a/useful_scripts/conc_gzip_files.py
+++ b/useful_scripts/conc_gzip_files.py
@@ -13,7 +13,7 @@ def conc_gzip_files(in_dir, out_file, append=False, print_progress=True):
Keyword arguments:
in_dir (str): Path of the directory with the gzip-files
out_file (str): Path to the resulting file
- append (bool): If true, it appends contents to an exisiting file,
+ append (bool): If true, it appends contents to an existing file,
else creates a new output file.
print_progress (bool): prints progress bar if true.
diff --git a/useful_scripts/find_file.py b/useful_scripts/find_file.py
new file mode 100644
index 0000000..8cbcc4d
--- /dev/null
+++ b/useful_scripts/find_file.py
@@ -0,0 +1,18 @@
+# Sebastian Raschka 2014
+#
+# A Python function to find files in a directory based on a substring search.
+
+
+import os
+
+def find_files(substring, path):
+ results = []
+ for f in os.listdir(path):
+ if substring in f:
+ results.append(os.path.join(path, f))
+ return results
+
+# E.g.
+# find_files('Untitled', '/Users/sebastian/Desktop/')
+# returns
+# ['/Users/sebastian/Desktop/Untitled0.ipynb']
\ No newline at end of file
diff --git a/useful_scripts/fix_tab_csv.ipynb b/useful_scripts/fix_tab_csv.ipynb
new file mode 100644
index 0000000..496f89f
--- /dev/null
+++ b/useful_scripts/fix_tab_csv.ipynb
@@ -0,0 +1,94 @@
+{
+ "metadata": {
+ "name": "",
+ "signature": "sha256:996358a25da6fc77c66d183e79209307af06bd2f9abb0656d3bb70cfc2fe597a"
+ },
+ "nbformat": 3,
+ "nbformat_minor": 0,
+ "worksheets": [
+ {
+ "cells": [
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Sebastian Raschka 05/09/2014"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "# Fixing CSV files"
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "We have a directory `../CSV_files_raw/` with CSV files where some of them have 'tab-separated' and some of them 'comma-separated' columns. \n",
+ "Here, we will 'fix' them, i.e., have them all comma-separated, and save them to a new directory `../CSV_fixed`."
+ ]
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "First, we create a dictionary with the file basenames as keys. The values are lists of the file paths to the raw and new fixed CSV files. e.g., \n",
+ "\n",
+ " {\n",
+ " 'abc.csv': ['../CSV_files_raw/abc.csv', '../CSV_fixed/abc.csv'], \n",
+ " 'def.csv': ['../CSV_files_raw/def.csv', '../CSV_fixed/def.csv'], \n",
+ " ...\n",
+ " }"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "import sys\n",
+ "import os\n",
+ "\n",
+ "raw_dir = '../CSV_files_raw/'\n",
+ "fixed_dir = '../CSV_fixed'\n",
+ "\n",
+ "if not os.path.exists(fixed_dir):\n",
+ " os.mkdir(fixed_dir)\n",
+ "\n",
+ "f_dict = {os.path.basename(f):[os.path.join(raw_dir, f),\n",
+ " os.path.join(fixed_dir, f)]\n",
+ " for f in os.listdir(raw_dir) if f.endswith('.csv')} "
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [],
+ "prompt_number": 8
+ },
+ {
+ "cell_type": "markdown",
+ "metadata": {},
+ "source": [
+ "Now, we can replace the tabs with commas for the new files very easily:"
+ ]
+ },
+ {
+ "cell_type": "code",
+ "collapsed": false,
+ "input": [
+ "for f in f_dict.keys():\n",
+ " with open(f_dict[f][0], 'r') as raw, open(f_dict[f][1], 'w') as fixed:\n",
+ " for line in raw:\n",
+ " line = line.strip().split('\\t')\n",
+ " fixed.write(','.join(line) + '\\n')"
+ ],
+ "language": "python",
+ "metadata": {},
+ "outputs": [],
+ "prompt_number": 11
+ }
+ ],
+ "metadata": {}
+ }
+ ]
+}
\ No newline at end of file
diff --git a/useful_scripts/large_csv_to_sqlite.py b/useful_scripts/large_csv_to_sqlite.py
new file mode 100644
index 0000000..9932f9c
--- /dev/null
+++ b/useful_scripts/large_csv_to_sqlite.py
@@ -0,0 +1,48 @@
+# This is a workaround snippet for reading very large CSV that exceed the
+# machine's memory and dump it into an SQLite database using pandas.
+#
+# Sebastian Raschka, 2015
+#
+# Tested in Python 3.4.2 and pandas 0.15.2
+
+import pandas as pd
+import sqlite3
+from pandas.io import sql
+import subprocess
+
+# In and output file paths
+in_csv = '../data/my_large.csv'
+out_sqlite = '../data/my.sqlite'
+
+table_name = 'my_table' # name for the SQLite database table
+chunksize = 100000 # number of lines to process at each iteration
+
+# columns that should be read from the CSV file
+columns = ['molecule_id','charge','db','drugsnow','hba','hbd','loc','nrb','smiles']
+
+# Get number of lines in the CSV file
+nlines = subprocess.check_output(['wc', '-l', in_csv])
+nlines = int(nlines.split()[0])
+
+# connect to database
+cnx = sqlite3.connect(out_sqlite)
+
+# Iteratively read CSV and dump lines into the SQLite table
+for i in range(0, nlines, chunksize): # change 0 -> 1 if your csv file contains a column header
+
+ df = pd.read_csv(in_csv,
+ header=None, # no header, define column header manually later
+ nrows=chunksize, # number of rows to read at each iteration
+ skiprows=i) # skip rows that were already read
+
+ # columns to read
+ df.columns = columns
+
+ sql.to_sql(df,
+ name=table_name,
+ con=cnx,
+ index=False, # don't use CSV file index
+ index_label='molecule_id', # use a unique column from DataFrame as index
+ if_exists='append')
+cnx.close()
+
diff --git a/useful_scripts/prepend_python_shebang.sh b/useful_scripts/prepend_python_shebang.sh
new file mode 100644
index 0000000..686225f
--- /dev/null
+++ b/useful_scripts/prepend_python_shebang.sh
@@ -0,0 +1,21 @@
+#!/usr/bin/env bash
+# Sebastian Raschka 05/21/2014
+# Shell script that prepends a Python shebang
+# '#!/usr/bin/env python' to all
+# Python script files in the current directory
+# so that script files can be executed via
+# >> myscript.py
+# instead of
+# >> python myscript.py
+
+# prepends '#!/usr/bin/env python' to all .py files
+
+find ./ -maxdepth 1 -name "*.py" -exec sed -i.bak '1i\
+#!/usr/bin/env python
+' {} \;
+
+# removes temporary files
+find . -name "*.bak" -exec rm -rf {} \;
+
+# makes Python scripts executable
+chmod ug+x *.py
diff --git a/useful_scripts/preprocess_first_last_names.py b/useful_scripts/preprocess_first_last_names.py
new file mode 100644
index 0000000..b0957c2
--- /dev/null
+++ b/useful_scripts/preprocess_first_last_names.py
@@ -0,0 +1,79 @@
+# Sebastian Raschka 2014
+#
+# A Python function to generalize first and last names.
+# The typical use case of such a function to merge data that have been collected
+# from different sources (e.g., names of soccer players as shown in the doctest.)
+#
+
+import unicodedata
+import string
+import re
+
+def preprocess_names(name, output_sep=' ', firstname_output_letters=1):
+ """
+ Function that outputs a person's name in the format
+ (all lowercase)
+
+ >>> preprocess_names("Samuel Eto'o")
+ 'etoo s'
+
+ >>> preprocess_names("Eto'o, Samuel")
+ 'etoo s'
+
+ >>> preprocess_names("Eto'o,Samuel")
+ 'etoo s'
+
+ >>> preprocess_names('Xavi')
+ 'xavi'
+
+ >>> preprocess_names('Yaya Touré')
+ 'toure y'
+
+ >>> preprocess_names('José Ángel Pozo')
+ 'pozo j'
+
+ >>> preprocess_names('Pozo, José Ángel')
+ 'pozo j'
+
+ >>> preprocess_names('Pozo, José Ángel', firstname_output_letters=2)
+ 'pozo jo'
+
+ >>> preprocess_names("Eto'o, Samuel", firstname_output_letters=2)
+ 'etoo sa'
+
+ >>> preprocess_names("Eto'o, Samuel", firstname_output_letters=0)
+ 'etoo'
+
+ >>> preprocess_names("Eto'o, Samuel", output_sep=', ')
+ 'etoo, s'
+
+ """
+
+ # set first and last name positions
+ last, first = 'last', 'first'
+ last_pos = -1
+
+ if ',' in name:
+ last, first = first, last
+ name = name.replace(',', ' ')
+ last_pos = 1
+
+ spl = name.split()
+ if len(spl) > 2:
+ name = '%s %s' % (spl[0], spl[last_pos])
+
+ # remove accents
+ name = ''.join(x for x in unicodedata.normalize('NFKD', name) if x in string.ascii_letters+' ')
+
+ # get first and last name if applicable
+ m = re.match('(?P\w+)\W+(?P\w+)', name)
+ if m:
+ output = '%s%s%s' % (m.group(last), output_sep, m.group(first)[:firstname_output_letters])
+ else:
+ output = name
+ return output.lower().strip()
+
+
+if __name__ == "__main__":
+ import doctest
+ doctest.testmod()
diff --git a/useful_scripts/principal_eigenvector.py b/useful_scripts/principal_eigenvector.py
new file mode 100644
index 0000000..913cf62
--- /dev/null
+++ b/useful_scripts/principal_eigenvector.py
@@ -0,0 +1,20 @@
+# Select a principal eigenvector via NumPy
+# to be used as a template (copy & paste) script
+
+import numpy as np
+
+# set A to be your matrix
+A = np.array([[1, 2, 3],
+ [4, 5, 6],
+ [7, 8, 9]])
+
+
+eig_vals, eig_vecs = np.linalg.eig(A)
+idx = np.absolute(eig_vals).argsort()[::-1] # decreasing order
+sorted_eig_vals = eig_vals[idx]
+sorted_eig_vecs = eig_vecs[:, idx]
+
+principal_eig_vec = sorted_eig_vecs[:, 0] # eigvec with largest eigval
+
+normalized_pr_eig_vec = np.real(principal_eig_vec / np.sum(principal_eig_vec))
+print(normalized_pr_eig_vec) # eigvec that sums up to one
diff --git a/useful_scripts/random_string_generator.py b/useful_scripts/random_string_generator.py
new file mode 100644
index 0000000..15cfe51
--- /dev/null
+++ b/useful_scripts/random_string_generator.py
@@ -0,0 +1,19 @@
+import string
+import random
+
+def rand_string(length):
+ """ Generates a random string of numbers, lower- and uppercase chars. """
+ return ''.join(random.choice(
+ string.ascii_lowercase + string.ascii_uppercase + string.digits)
+ for i in range(length)
+ )
+
+if __name__ == '__main__':
+ print("Example1:", rand_string(length=4))
+ print("Example2:", rand_string(length=8))
+ print("Example2:", rand_string(length=16))
+
+
+ # Example1: 5bVL
+ # Example2: oIIg37xl
+ # Example2: 7IqDbrf506TatFO9
diff --git a/useful_scripts/sparsify_matrix.py b/useful_scripts/sparsify_matrix.py
new file mode 100644
index 0000000..ef5e141
--- /dev/null
+++ b/useful_scripts/sparsify_matrix.py
@@ -0,0 +1,38 @@
+# Sebastian Raschka 2014
+#
+# Sparsifying a matrix by Zeroing out all elements but the top k elements in a row.
+# The matrix could be a distance or similarity matrix (e.g., kernel matrix in kernel PCA),
+# where we are interested to keep the top k neighbors.
+
+import numpy as np
+
+print('Sparsify a matrix by zeroing all elements but the top 2 values in a row.\n')
+
+A = np.array([[1,2,3,4,5],[9,8,6,4,5],[3,1,7,8,9]])
+
+print('Before:\n%s\n' %A)
+
+
+k = 2 # keep top k neighbors
+for row in A:
+ sort_idx = np.argsort(row)[::-1] # get indexes of sort order (high to low)
+ for i in sort_idx[k:]:
+ row[i]=0
+
+print('After:\n%s\n' %A)
+
+
+"""
+Sparsify a matrix by zeroing all elements but the top 2 values in a row.
+
+Before:
+[[1 2 3 4 5]
+ [9 8 6 4 5]
+ [3 1 7 8 9]]
+
+After:
+[[0 0 0 4 5]
+ [9 8 0 0 0]
+ [0 0 0 8 9]]
+
+"""
\ No newline at end of file
diff --git a/useful_scripts/univariate_poisson_pdf.py b/useful_scripts/univariate_poisson_pdf.py
new file mode 100644
index 0000000..30d3ae4
--- /dev/null
+++ b/useful_scripts/univariate_poisson_pdf.py
@@ -0,0 +1,48 @@
+import numpy as np
+import math
+
+
+def poisson_lambda_mle(d):
+ """
+ Computes the Maximum Likelihood Estimate for a given 1D training
+ dataset from a Poisson distribution.
+
+ """
+ return sum(d) / len(d)
+
+def likelihood_poisson(x, lam):
+ """
+ Computes the class-conditional probability for an univariate
+ Poisson distribution
+
+ """
+ if x // 1 != x:
+ likelihood = 0
+ else:
+ likelihood = math.e**(-lam) * lam**(x) / math.factorial(x)
+ return likelihood
+
+
+if __name__ == "__main__":
+
+ # Plot Probability Density Function
+ from matplotlib import pyplot as plt
+
+ training_data = [0, 1, 1, 3, 1, 0, 1, 2, 1, 2, 2, 1, 2, 0, 1, 4]
+ mle_poiss = poisson_lambda_mle(training_data)
+ true_param = 1.0
+
+ x_range = np.arange(0, 5, 0.1)
+ y_true = [likelihood_poisson(x, true_param) for x in x_range]
+ y_mle = [likelihood_poisson(x, mle_poiss) for x in x_range]
+
+ plt.figure(figsize=(10,8))
+ plt.plot(x_range, y_true, lw=2, alpha=0.5, linestyle='--', label='true parameter ($\lambda={}$)'.format(true_param))
+ plt.plot(x_range, y_mle, lw=2, alpha=0.5, label='MLE ($\lambda={}$)'.format(mle_poiss))
+ plt.title('Poisson probability density function for the true and estimated parameters')
+ plt.ylabel('p(x|theta)')
+ plt.xlim([-1,5])
+ plt.xlabel('random variable x')
+ plt.legend()
+
+ plt.show()
diff --git a/zen_of_python.py b/zen_of_python.py
deleted file mode 100644
index d82cacd..0000000
--- a/zen_of_python.py
+++ /dev/null
@@ -1,24 +0,0 @@
->>> import this
-"""
-The Zen of Python, by Tim Peters
-
-Beautiful is better than ugly.
-Explicit is better than implicit.
-Simple is better than complex.
-Complex is better than complicated.
-Flat is better than nested.
-Sparse is better than dense.
-Readability counts.
-Special cases aren't special enough to break the rules.
-Although practicality beats purity.
-Errors should never pass silently.
-Unless explicitly silenced.
-In the face of ambiguity, refuse the temptation to guess.
-There should be one-- and preferably only one --obvious way to do it.
-Although that way may not be obvious at first unless you're Dutch.
-Now is better than never.
-Although never is often better than *right* now.
-If the implementation is hard to explain, it's a bad idea.
-If the implementation is easy to explain, it may be a good idea.
-Namespaces are one honking great idea -- let's do more of those!
-"""